code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
hash('hola')
# $$
# zkh(a,b) = \left\{
# \begin{matrix}
# b & , & a\%b == 0\\
# zkh(b,a\%b) & , & a\%b \ne 0
# \end{matrix}\right.
# $$
#
# $$
# zkh(a,b) = \left\{
# \begin{matrix}
# b & , & a\%b == 0\\
# 1 & , & i=1\\
# fib(i-2)+fib(i-1) & , & i>1
# \end{matrix}\right.
# $$
#
# $$
# \begin{matrix}
# 0 & 0 & 0 & 3 & 0 & 0\\
# 0 & 1 & 0 & 0 & 0 & 0\\
# 0 & 0 & 0 & 0 & 0 & 0\\
# 0 & 0 & 4 & 0 & 0 & 0
# \end{matrix}
# $$
#
| Gardenkiak/Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Directory and Files Management
# 1.1 [Get Current Directory](#section1)
# 1.2 [Changing Directory](#section2)
# 1.3 [List Directories and Files](#section3)
# 1.4 [Making a New Directory](#section4)
# 1.5 [Renaming a Directory or a File](#section5)
# 1.6 [Removing Directory or File](#section6)
# <a id="section1"></a>
# **1. Get Current Directory**
# We can get the present working directory using the **getcwd()** method.
import os
os.getcwd()
# <a id="section2"></a>
# **2. Changing Directory**
# We can change the current working directory using the **chdir()** method.
#
# The new path that we want to change to must be supplied as a string to this method. We can use both forward slash (/) or the backward slash (\) to separate path elements.
import os
os.chdir('D:\code\Python Data Science')
os.getcwd()
# <a id="section3"></a>
# **3. List Directories and Files**
# All files and sub directories inside a directory can be known using the **listdir()** method.
import os
os.listdir() # returns in form of list
# <a id="section4"></a>
# **4. Making a New Directory**
# We can make a new directory using the **mkdir()** method.
#
# This method takes in the path of the new directory. If the full path is not specified, the new directory is created in the current working directory.
import os
os.mkdir("test")
os.listdir()
# <a id="section5"></a>
# **5. Renaming a Directory or a File**
# The **rename()** method can rename a directory or a file.
#
# The first argument is the old name and the new name must be supplies as the second argument.
import os
print(os.listdir())
os.rename('test', 'list')
print(os.listdir())
# <a id="section5"></a>
# **6. Removing Directory or File**
# A file can be removed (deleted) using the remove() method.
#
# Similarly, the rmdir() method removes an empty directory.
import os
os.remove("list")
os.dirlist()
# In order to remove a non-empty directory we can use the rmtree() method inside the shutil module.
>>> os.listdir()
['test']
>>> os.rmdir('test')
Traceback (most recent call last):
...
OSError: [WinError 145] The directory is not empty: 'test'
>>> import shutil
>>> shutil.rmtree('test')
>>> os.listdir()
[]
| 16. Adv Python File Handling/08. Directories in Python.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Procédure de test DF sur un processus simulé
#
# On commence par la simulation du processus:
# +
T <- 1000
u <- rnorm(T)
e <- rep(NA,T)
y <- rep(NA,T)
dy <- rep(NA,500)
y[1] <- 0
e[1] <- 0
for (t in 2:T){
e[t] <- 0.2*e[t-1] +u[t]
y[t] <- y[t-1] + e[t]
}
# -
# * Etape 1: On fait un test de racine unitaire sur le modèle le plus général
#
# $$\Delta y_t = c + \rho y_{t-1} + \beta t + \varepsilon_t$$
#
dy=diff(y)
T=length(dy)
lags = 0
y.lag.1=y[(lags+1):T]
t = (lags+1):T
model_nc3 <- lm(dy~1+t+y.lag.1 )
summary(model_nc3)
# La statistique de test associée à $\rho$ est -0.970. On compare cette valeur à la valeur critique d'une table <NAME>. Par exemple: http://homes.chass.utoronto.ca/~floyd/statabs.pdf
# Cette valeur critique est égale à -3.45. La t-stat est au dessus, on accepte l'hypothèse nulle de racine unitaire.
#
# * Etape 2: Il faut maintenant vérifier que le modèle 3 est le bon modèle.
# Validité du modèle 3
ssrnc3 = sum(residuals(model_nc3)^2)
model_c3 <- lm(dy~1)
ssrc3 = sum(residuals(model_c3)^2)
stat_f3 = ((ssrc3 - ssrnc3)/2)/(ssrnc3/(T-3))
print(stat_f3)
# La statistique de Fisher est de 1.77. On compare à la statistique de la table de DF. On trouve F critique = 6.49
# On accepte l'hypothèse nulle, le modèle 3 n' est pas le bon.
#
# * Étape 3: On recommence avec le modèle 2
#
# $$ \Delta y_t = c + \rho y_{t-1} + \varepsilon_t $$
# +
# On estime le modèle 2 car modèle 3 mal spécifié
model_nc2 <- lm(dy~1+y.lag.1 )
summary(model_nc2)
# -
# La statistique de test associée à $\rho$ est -0.970. On compare cette valeur à la valeur critique d'une table Dickey Fuller. La statistique associée est t = -1.778. La valeur critique lue dans la table DF est égale à -2.8. On accepte l'hypothèse nulle de racine unitaire.
#
# * Étape 4: On vérifie que le modèle 2 est le bon modèle
# Validité du modèle 2
ssrnc2 = sum(residuals(model_nc2)^2)
ssrc2 = sum(dy^2)
stat_f2 = ((ssrc2 - ssrnc2)/2)/(ssrnc2/(T-2))
print(stat_f2)
# La statistique de Fisher est F=1.62. La valeur critique est de 4.71. On accepte l'hypothèse nulle, le modèle 2 n'est pas le bon.
#
# * Étape 5: On finit avec le modèle le plus simple
model_nc1 <- lm(dy~0 + y.lag.1 )
summary(model_nc1)
# La t-stat est de -1.40. La valeur critique lue dans la table de DF est de -1.95. Le modèle est I(1)
# Refaisons le même exercice mais avec une tendance déterministe.
#
#
# +
T <- 1000
u <- rnorm(T)
e <- rep(NA,T)
y <- rep(NA,T)
dy <- rep(NA,500)
y[1] <- 0
e[1] <- 0
for (t in 2:T){
e[t] <- 0.2*e[t-1] +u[t]
y[t] <- y[t-1] + e[t] + 0.01*t
}
dy=diff(y)
T=length(dy)
lags = 0
y.lag.1=y[(lags+1):T]
t = (lags+1):T
model_nc3 <- lm(dy~1+t+y.lag.1 )
summary(model_nc3)
# Validité du modèle 3
ssrnc3 = sum(residuals(model_nc3)^2)
model_c3 <- lm(dy~1)
ssrc3 = sum(residuals(model_c3)^2)
stat_f3 = ((ssrc3 - ssrnc3)/2)/(ssrnc3/(T-3))
print(stat_f3)
# -
# La t-value est de 0.98, la valeur critique de -3.45. On accepte la racine unitaire. La stat de Fisher est de 4043, on rejette l'hypothèse nulle. On a bien le modèle simulé.
# Entrainez-vous avec différents modèles!!!!
| code/DF_test_using_R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# General imports
import numpy as np
import torch
import matplotlib.pyplot as plt
# DeepMoD stuff
from deepymod import DeepMoD
from deepymod.model.func_approx import NN
from deepymod.model.library import Library1D
from deepymod.model.constraint import LeastSquares
from deepymod.model.sparse_estimators import Threshold, PDEFIND
from deepymod.training import train
from deepymod.training.sparsity_scheduler import TrainTestPeriodic
from deepymod.data import Dataset
from deepymod.data.burgers import BurgersDelta
#if torch.cuda.is_available():
# device = 'cuda'
#else:
device = 'cpu'
# Settings for reproducibility
np.random.seed(44)
torch.manual_seed(0)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# Making dataset
noise = 0.01
A = 1
v = 0.25
# +
runs = 1
dataset = Dataset(BurgersDelta, A=A, v=v)
x = np.linspace(-3.9, 4.2, 120)
t = np.linspace(0.1, 1.1, 100)
t_grid, x_grid = np.meshgrid(t, x, indexing='ij')
X, y = dataset.create_dataset(x_grid.reshape(-1, 1), t_grid.reshape(-1, 1), n_samples=0, noise=noise, random=False, normalize=False)
X.shape
xt = X.reshape(x_grid.shape[0],x_grid.shape[1],2)
yt = y.reshape(x_grid.shape[0],x_grid.shape[1])
# -
xt.shape
delta_x = 6
shift = np.mod(np.arange(0,xt.shape[0]),delta_x)
shifted_X, shifted_Y = [], []
cut_off = np.int((xt.shape[1]/delta_x)-1)
ytrain = np.empty([yt.shape[0],cut_off])
xtrain = np.empty([xt.shape[0],cut_off,xt.shape[2]])
for i in np.arange(yt.shape[0]):
ytrain[i,:] = yt[i,shift[i]:-1:delta_x][:cut_off]
xtrain[i,:,0] = xt[i,shift[i]:-1:delta_x,0][:cut_off]
xtrain[i,:,1] = xt[i,shift[i]:-1:delta_x,1][:cut_off]
# +
delta_x = 10
shift = np.mod(np.arange(0,xt.shape[0]),delta_x)
shifted_X, shifted_Y = [], []
cut_off = np.int((xt.shape[1]/delta_x)-1)
ytrain = np.empty([yt.shape[0],cut_off])
xtrain = np.empty([xt.shape[0],cut_off,xt.shape[2]])
for i in np.arange(yt.shape[0]):
ytrain[i,:] = yt[i,shift[i]:-1:delta_x][:cut_off]
xtrain[i,:,0] = xt[i,shift[i]:-1:delta_x,0][:cut_off]
xtrain[i,:,1] = xt[i,shift[i]:-1:delta_x,1][:cut_off]
# -
fig = plt.figure(figsize=(4,4))
ax = fig.add_subplot(111)
ax.plot(xtrain[:,:,1],'bo', markersize=2.5)
ax.set_aspect(aspect=3)
ax.set_ylim([-3.0,3.0])
ax.set_xlim([0,40.0])
plt.savefig('shifted.pdf')
plt.show()
(np.random.rand(1,n_x)-0.5)*10
number_of_samples= 10
# +
ytrain = np.empty([yt.shape[0],number_of_samples])
xtrain = np.empty([xt.shape[0],number_of_samples,xt.shape[2]])
for i in np.arange(yt.shape[0]):
idx = np.random.permutation(yt.shape[1])
ytrain[i,:] = yt[i,idx][:number_of_samples]
xtrain[i,:,0] = xt[i,idx,0][:number_of_samples]
xtrain[i,:,1] = xt[i,idx,1][:number_of_samples]
# -
fig = plt.figure(figsize=(4,4))
ax = fig.add_subplot(111)
ax.plot(xtrain[:,:,1],'bo', markersize=2.5)
ax.set_aspect(aspect=3)
ax.set_ylim([-3.0,3.0])
ax.set_xlim([0,40.0])
plt.savefig('random.pdf')
plt.show()
x = np.linspace(-4, 4, 12)
t = np.linspace(0.1, 1.1, 100)
t_grid, x_grid = np.meshgrid(t, x, indexing='ij')
fig = plt.figure(figsize=(4,4))
ax = fig.add_subplot(111)
ax.plot(x_grid,'bo', markersize=2.5)
ax.set_aspect(aspect=3)
ax.set_ylim([-3.0,3.0])
ax.set_xlim([0,40.0])
plt.savefig('grid.pdf')
plt.show()
# +
plt.plot(xtrain[:,:,1],'bo', markersize=2)
plt.ylim([-3.0,3.0])
plt.xlim([0, 100])
# -
plt.plot(xtrain[19,:,1],ytrain[19],'o-')
plt.plot(xtrain[40,:,1],ytrain[40],'o-')
plt.plot(xtrain[81,:,1],ytrain[21],'o-')
for nx in [16,14]:
delta_x = nx
shift = np.mod(np.arange(0,xt.shape[0]),delta_x)
shifted_X, shifted_Y = [], []
cut_off = np.int((xt.shape[1]/delta_x)-1)
ytrain = np.empty([yt.shape[0],cut_off])
xtrain = np.empty([xt.shape[0],cut_off,xt.shape[2]])
for i in np.arange(yt.shape[0]):
ytrain[i,:] = yt[i,shift[i]:-1:delta_x][:cut_off]
xtrain[i,:,0] = xt[i,shift[i]:-1:delta_x,0][:cut_off]
xtrain[i,:,1] = xt[i,shift[i]:-1:delta_x,1][:cut_off]
xtrain = np.transpose(xtrain,axes=(1,0,2))
ytrain = ytrain.T
X = torch.tensor(xtrain.reshape(-1,2), dtype=torch.float32, requires_grad=True)
y = torch.tensor(ytrain.reshape(-1,1), dtype=torch.float32)
idx = np.random.permutation(y.shape[0])
y = y[idx]
X[:,0] = X[idx,0]
X[:,1] = X[idx,1]
X, y = X.to(device), y.to(device)
network = NN(2, [30, 30, 30, 30], 1)
library = Library1D(poly_order=2, diff_order=3) # Library function
estimator = Threshold(0.2) # Sparse estimator
constraint = LeastSquares() # How to constrain
model = DeepMoD(network, library, estimator, constraint).to(device) # Putting it all in the model
sparsity_scheduler = TrainTestPeriodic(periodicity=50, patience=500, delta=1e-7) # in terms of write iterations
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True, lr=1e-3) # Defining optimizer
train(model, X, y, optimizer, sparsity_scheduler, log_dir='runs/shifted_grid_'+str(cut_off), split=0.8, write_iterations=25, max_iterations=100000, delta=1e-4, patience=500)
print(model.constraint_coeffs(sparse=True, scaled=True))
# +
for n_x in np.array([2]):
number_of_samples = n_x
ytrain = np.empty([yt.shape[0],number_of_samples])
xtrain = np.empty([xt.shape[0],number_of_samples,xt.shape[2]])
for i in np.arange(yt.shape[0]):
idx = np.random.permutation(yt.shape[1])
ytrain[i,:] = yt[i,idx][:number_of_samples]
xtrain[i,:,0] = xt[i,idx,0][:number_of_samples]
xtrain[i,:,1] = xt[i,idx,1][:number_of_samples]
xtrain_b = np.transpose(xtrain,axes=(1,0,2))
ytrain_b = ytrain.T
X = torch.tensor(xtrain_b.reshape(-1,2), dtype=torch.float32, requires_grad=True)
y = torch.tensor(ytrain_b.reshape(-1,1), dtype=torch.float32)
print(y.shape)
X, y = X.to(device), y.to(device)
network = NN(2, [30, 30, 30, 30], 1)
library = Library1D(poly_order=2, diff_order=3) # Library function
estimator = Threshold(0.2) # Sparse estimator
constraint = LeastSquares() # How to constrain
model = DeepMoD(network, library, estimator, constraint).to(device) # Putting it all in the model
sparsity_scheduler = TrainTestPeriodic(periodicity=50, patience=500, delta=1e-7) # in terms of write iterations
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True, lr=1e-3) # Defining optimizer
# train(model, X, y, optimizer, sparsity_scheduler, log_dir='runs/random_grid'+str(n_x), split=0.8, write_iterations=25, max_iterations=100000, delta=1e-4, patience=500)
print(model.constraint_coeffs(sparse=True, scaled=True))
# -
for n_x in np.array([4,6,8,10,12,14,16]):
for run in np.arange(runs):
number_of_samples = n_x * 100
idx = np.random.permutation(y.shape[0])
X = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
y = torch.tensor(y[idx, :][:number_of_samples], dtype=torch.float32)
X, y = X.to(device), y.to(device)
network = NN(2, [30, 30, 30, 30], 1)
library = Library1D(poly_order=2, diff_order=3) # Library function
estimator = Threshold(0.2) # Sparse estimator
constraint = LeastSquares() # How to constrain
model = DeepMoD(network, library, estimator, constraint).to(device) # Putting it all in the model
sparsity_scheduler = TrainTestPeriodic(periodicity=50, patience=500, delta=1e-7) # in terms of write iterations
optimizer = torch.optim.Adam(model.parameters(), betas=(0.99, 0.99), amsgrad=True, lr=1e-3) # Defining optimizer
train(model, X, y, optimizer, sparsity_scheduler, log_dir='runs/rand'+str(n_x), split=0.8, write_iterations=25, max_iterations=100000, delta=1e-4, patience=500)
print(model.constraint_coeffs(sparse=True, scaled=True))
| paper/Burgers/.ipynb_checkpoints/shifted_grid_loop-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Business Understanding
# We would like to track Corona VIRUS spread across countries and with personal local information
#
# The general information is not so relevant for me I would like to have a deep dive local development of the spread
# # Goals
# - We would like to understand the data quality
# - Everything should be automated as much as possible: how many clicks do we need to execute the full pipeline
# # Constraints:
# - Each notebook should be left clean and ready for full execution
| notebooks/Business_Understanding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="OU4AAstR0HG6"
# # Descritores de Imagens via Histograma de Cor
#
# Exemplos baseados em https://github.com/xn2333/OpenCV/blob/master/Seminar_Image_Processing_in_Python.ipynb*
#
#
#
#
#
#
# + [markdown] id="HFHge950Mca1"
# # Instalando Bibliotecas
# + id="6dy-iP-VTibt"
import numpy as np
import pandas as pd
import cv2 as cv
from google.colab.patches import cv2_imshow # for image display
from skimage import io
from PIL import Image
import matplotlib.pylab as plt
# + [markdown] id="mf4fV9OiPRM7"
# # Vamos construir nosso dataset
# + colab={"base_uri": "https://localhost:8080/"} id="6vIbaraW9cIn" outputId="e31261a9-fa3c-4ce0-8bd6-5fe8a838efbf"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="tamKtShzqe3n" outputId="c6b07639-fee0-4432-a89f-703735612ccc"
import shutil, sys
# move used images
dataset = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv')
print(dataset)
for index, row in dataset.iterrows():
if row['post']:
shutil.move("/content/drive/MyDrive/hourlyloonacatcher/" + row['group'] + "/" + row['filename'], "/content/drive/MyDrive/hourlyloonacatcher/" + row['group'] + "/posted/")
dataset = dataset.drop([index])
dataset = dataset.reset_index().drop(columns='index')
print(dataset)
dataset.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="mRbpONE69Cf4" outputId="327c1e16-89a5-4363-f578-87a9234b7f32"
# percentagens taken by each girl
loona = [
'heejin',
'hyunjin',
'haseul',
'yeojin',
'vivi',
'kimlip',
'jinsoul',
'choerry',
'yves',
'chuu',
'gowon',
'oliviahye'
]
deukae = [
"jiu",
"sua",
"siyeon",
"handong",
"yoohyeon",
"dami",
"gahyeon"
]
images = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/images_to_post.csv')
for l in loona:
print('{0}: {1}'.format(l,(100 * len(images[images['loona'].str.contains(l)].index) / len(images.index))))
print()
for d in deukae:
print('{0}: {1}'.format(d,(100 * len(images[images['deukae'].str.contains(d)].index) / len(images.index))))
# + id="cLfL1Ixt2ZmT"
loona = [
'heejin',
'hyunjin',
'haseul',
'yeojin',
'vivi',
'kimlip',
'jinsoul',
'choerry',
'yves',
'chuu',
'gowon',
'oliviahye'
]
deukae = [
"jiu",
"sua",
"siyeon",
"handong",
"yoohyeon",
"dami",
"gahyeon"
]
pairs = {}
for l in loona:
for d in deukae:
pairs[l + '_' + d] = 0
post_log = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher/post_log.csv')
for index, row in post_log.iterrows():
pairs[row['loona'] + '_' + row['deukae']] += 1
pairs_df = {'girls': [], 'values': []}
for key, value in pairs.items():
pairs_df['girls'].append(key)
pairs_df['values'].append(100 * (value / len(post_log.index)))
pairs_df = pd.DataFrame(pairs_df)
pairs_df.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/log_stats.csv')
# + id="BdCc7sN0Q8nc"
def load_zero():
import os
path = '/content/drive/MyDrive/hourlyloonacatcher/'
dataset = {"group": [], "filename": [], "image": [], "post": []}
for filename in os.listdir(path + 'loona/'):
img = cv.imread(path + 'loona/' + filename)
if img is not None:
dataset["group"].append('loona')
dataset["filename"].append(filename)
dataset["image"].append(img)
dataset["post"].append(False)
for filename in os.listdir(path + 'deukae/'):
img = cv.imread(path + 'deukae/' + filename)
if img is not None:
dataset["group"].append('deukae')
dataset["filename"].append(filename)
dataset["image"].append(img)
dataset["post"].append(False)
return dataset
def load_flying():
import os
path = '/content/drive/MyDrive/hourlyloonacatcher/'
dataset = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv').drop(columns='Unnamed: 0').reset_index().drop(columns='index').to_dict(orient='list')
dataset["image"] = []
for filename in os.listdir(path + 'loona/'):
img = cv.imread(path + 'loona/' + filename)
if img is not None and filename not in dataset["filename"]:
dataset["group"].append('loona')
dataset["filename"].append(filename)
dataset["image"].append(img)
dataset["post"].append(False)
elif img is not None:
dataset["image"].append(img)
for filename in os.listdir(path + 'deukae/'):
img = cv.imread(path + 'deukae/' + filename)
if img is not None and filename not in dataset["filename"]:
dataset["group"].append('deukae')
dataset["filename"].append(filename)
dataset["image"].append(img)
dataset["post"].append(False)
elif img is not None:
dataset["image"].append(img)
return dataset
# + [markdown] id="OinBn2XWVEUw"
# # Extraindo características do dataset usando Histograma de Cor
# + id="C9R7ZuPrVISf"
def prepareX(dataset):
color = ('b','g','r')
dataset_hist_r = []
dataset_hist_g = []
dataset_hist_b = []
counter = 0
for image in dataset["image"]:
hists = {}
for i,col in enumerate(color):
histr = cv.calcHist([image],[i],None,[256],[0,256])
if col == 'r': dataset_hist_r.append(histr)
if col == 'g': dataset_hist_g.append(histr)
if col == 'b': dataset_hist_b.append(histr)
X_r = np.array(dataset_hist_r)
length = np.sqrt((X_r**2).sum(axis=1))[:,None]
X_r = X_r / length
X_g = np.array(dataset_hist_g)
length = np.sqrt((X_g**2).sum(axis=1))[:,None]
X_g = X_g / length
X_b = np.array(dataset_hist_b)
length = np.sqrt((X_b**2).sum(axis=1))[:,None]
X_b = X_b / length
X = np.concatenate((X_r,X_g,X_g),axis=1)
X.shape
X = X.reshape(X.shape[0],X.shape[1])
X.shape
return X
# + [markdown] id="n0KJ7fVqnQEx"
# # Agrupamento de Imagens
# + id="v2BJaaKinRXV"
from sklearn.cluster import KMeans
import numpy as np
def do_kmeans(X, n_clusters=100):
kmeans = KMeans(n_clusters=n_clusters).fit(X)
return kmeans
# + [markdown] id="RUlfDz2RSrED"
# ## Escolher a imagem
# + id="9Ndoeo90S2qT"
def get_probs(loona_prob = [100/12] * 12, deukae_prob = [100/7] * 7):
loona = [
'heejin',
'hyunjin',
'haseul',
'yeojin',
'vivi',
'kimlip',
'jinsoul',
'choerry',
'yves',
'chuu',
'gowon',
'oliviahye'
]
deukae = [
"jiu",
"sua",
"siyeon",
"handong",
"yoohyeon",
"dami",
"gahyeon"
]
return loona, loona_prob, deukae, deukae_prob
# + id="pyxgY2_2Yfpc"
def update_probs(loona_prob, deukae_prob, loona, deukae, loona_choice, deukae_choice):
for i in range(len(loona)):
if loona[i] not in loona_choice:
loona_prob[i] += 100/12
else:
loona_prob[i] = 100/12
for i in range(len(deukae)):
if deukae[i] not in deukae_choice:
deukae_prob[i] += 100/7
else:
deukae_prob[i] = 100/7
return loona_prob, deukae_prob
# + id="E7WLTcT7xa6H"
import random
from sklearn.metrics import silhouette_samples
n_clusters = 50
def first_run():
dataset = load_zero()
X = prepareX(dataset)
kmeans = do_kmeans(X, n_clusters)
cluster_labels = kmeans.labels_
sample_silhouette_values = silhouette_samples(X, cluster_labels)
means_lst = []
for label in range(n_clusters):
means_lst.append(sample_silhouette_values[cluster_labels == label].mean())
clusterList = [x for _,x in sorted(zip(means_lst,range(n_clusters)))]
imageLoona = None
imageDeukae = None
images_to_post = {'loona': [], 'deukae': []}
# first time
loona, loona_prob, deukae, deukae_prob = get_probs()
loona_choice = random.choices(loona, weights=loona_prob, k=1)
deukae_choice = random.choices(deukae, weights=deukae_prob, k=1)
loona_prob, deukae_prob = update_probs(loona_prob, deukae_prob, loona, deukae, loona_choice, deukae_choice)
for i in range(192):
print(loona_choice[0], deukae_choice[0])
imageLoona, imageDeukae = -1, -1
#get image pair for post
for c in clusterList:
for image_id, cluster in enumerate(cluster_labels):
if not dataset["post"][image_id]:
if cluster == c:
if loona_choice[0] in dataset["filename"][image_id]:
print(image_id)
imageLoona = image_id
break
for image_id, cluster in enumerate(cluster_labels):
if not dataset["post"][image_id]:
if cluster == c:
if deukae_choice[0] in dataset["filename"][image_id]:
print(image_id)
imageDeukae = image_id
break
if (imageLoona != -1) and (imageDeukae != -1):
dataset["post"][imageLoona] = True
dataset["post"][imageDeukae] = True
# new pair
loona_choice = random.choices(loona, weights=loona_prob, k=1)
deukae_choice = random.choices(deukae, weights=deukae_prob, k=1)
loona_prob, deukae_prob = update_probs(loona_prob, deukae_prob, loona, deukae, loona_choice, deukae_choice)
# get out and choose right image
break
if (imageLoona != -1) and (imageDeukae != -1):
images_to_post['loona'].append(dataset["filename"][imageLoona])
images_to_post['deukae'].append(dataset["filename"][imageDeukae])
else:
print("No more image combinations for the pair")
#cv2_imshow(dataset["image"][imageLoona])
#cv2_imshow(dataset["image"][imageDeukae])
# generate dfs for saving
del dataset['image']
df_dataset = pd.DataFrame(dataset)
df_images_to_post = pd.DataFrame(images_to_post)
df_loona = {'name': [], 'prob': []}
df_loona['name'] = loona
df_loona['prob'] = loona_prob
df_loona = pd.DataFrame(df_loona)
df_deukae = {'name': [], 'prob': []}
df_deukae['name'] = deukae
df_deukae['prob'] = deukae_prob
df_deukae = pd.DataFrame(df_deukae)
df_dataset.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv')
df_images_to_post.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/images_to_post.csv')
df_loona.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_loona.csv')
df_deukae.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_deukae.csv')
def flying_run():
dataset = load_flying()
X = prepareX(dataset)
kmeans = do_kmeans(X, n_clusters)
cluster_labels = kmeans.labels_
sample_silhouette_values = silhouette_samples(X, cluster_labels)
means_lst = []
for label in range(n_clusters):
means_lst.append(sample_silhouette_values[cluster_labels == label].mean())
clusterList = [x for _,x in sorted(zip(means_lst,range(n_clusters)))]
imageLoona = None
imageDeukae = None
images_to_post = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher/images_to_post.csv').drop(columns='Unnamed: 0').reset_index().drop(columns='index').to_dict(orient='list')
#df_loona = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_loona.csv').drop(columns='Unnamed: 0').reset_index().drop(columns='index')
#loona, loona_prob = list(df_loona['name']), list(df_loona['prob'])
#df_deukae = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_deukae.csv').drop(columns='Unnamed: 0').reset_index().drop(columns='index')
#deukae, deukae_prob = list(df_deukae['name']), list(df_deukae['prob'])
loona, loona_prob, deukae, deukae_prob = get_probs()
loona_choice = random.choices(loona, weights=loona_prob, k=1)
deukae_choice = random.choices(deukae, weights=deukae_prob, k=1)
loona_prob, deukae_prob = update_probs(loona_prob, deukae_prob, loona, deukae, loona_choice, deukae_choice)
for i in range(192):
print(loona_choice[0], deukae_choice[0])
imageLoona, imageDeukae = -1, -1
#get image pair for post
for c in clusterList:
for image_id, cluster in enumerate(cluster_labels):
if not dataset["post"][image_id]:
if cluster == c:
if loona_choice[0] in dataset["filename"][image_id]:
print(image_id)
imageLoona = image_id
break
for image_id, cluster in enumerate(cluster_labels):
if not dataset["post"][image_id]:
if cluster == c:
if deukae_choice[0] in dataset["filename"][image_id]:
print(image_id)
imageDeukae = image_id
break
if (imageLoona != -1) and (imageDeukae != -1):
dataset["post"][imageLoona] = True
dataset["post"][imageDeukae] = True
# new pair
loona_choice = random.choices(loona, weights=loona_prob, k=1)
deukae_choice = random.choices(deukae, weights=deukae_prob, k=1)
loona_prob, deukae_prob = update_probs(loona_prob, deukae_prob, loona, deukae, loona_choice, deukae_choice)
# get out and choose right image
break
if (imageLoona != -1) and (imageDeukae != -1):
images_to_post['loona'].append(dataset["filename"][imageLoona])
images_to_post['deukae'].append(dataset["filename"][imageDeukae])
else:
print("No more image combinations for the pair")
#cv2_imshow(dataset["image"][imageLoona])
#cv2_imshow(dataset["image"][imageDeukae])
# generate dfs for saving
del dataset['image']
df_dataset = pd.DataFrame(dataset)
df_images_to_post = pd.DataFrame(images_to_post)
df_loona = {'name': [], 'prob': []}
df_loona['name'] = loona
df_loona['prob'] = loona_prob
df_loona = pd.DataFrame(df_loona)
df_deukae = {'name': [], 'prob': []}
df_deukae['name'] = deukae
df_deukae['prob'] = deukae_prob
df_deukae = pd.DataFrame(df_deukae)
df_dataset.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv')
df_images_to_post.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/images_to_post.csv')
df_loona.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_loona.csv')
df_deukae.to_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_deukae.csv')
# + id="nFcOO_Er-XJk"
# when no images pairs were generated previously
#first_run()
# when image pairs are already being posted
flying_run()
# + id="8TqnKTryNrv3"
df = pd.read_csv('/content/drive/MyDrive/hourlyloonacatcher_dfs/df_dataset.csv').drop(columns='Unnamed: 0').reset_index().drop(columns='index')
# + id="If2zm-vzN806" colab={"base_uri": "https://localhost:8080/"} outputId="24975240-01ed-40ef-db51-96408b1a18ba"
len(df[df['post'] == True])
| Cluster_hloonacatcher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 10 Ordinary differential equations – Integrators
# More about integrators to solve ODEs.
#
# * Euler's Rule (and it's shortcomings)
# * Runge-Kutta methods
# + [markdown] slideshow={"slide_type": "slide"}
# ## Euler's Rule
#
# From the last lecture: Given the $n$-dimensional vectors from the ODE standard form
#
# $$
# \frac{d\mathbf{y}}{dt} = \mathbf{f}(t, \mathbf{y})
# $$
# the **Euler rule** amounts to
#
# \begin{align}
# \mathbf{f}(t, \mathbf{y}) = \frac{d\mathbf{y}(t)}{dt} &\approx \frac{\mathbf{y}(t_{n+1}) - \mathbf{y}(t_n)}{h}\\
# \mathbf{y}_{n+1} &\approx \mathbf{y}_n + h \mathbf{f}(t_n, \mathbf{y}_n) \quad \text{with} \quad \mathbf{y}_n := \mathbf{y}(t_n)
# \end{align}
# -
# Euler's rule cannot obtain high precision without very small $h$ but this leads to higher round-off error accumulation.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Euler's Rule applied to the 1D harmonic oscillator (theory)
#
# \begin{alignat}{3}
# f^{(0)}(t, \mathbf{y}) &= y^{(1)},
# &\quad y^{(0)}_0 &= x_0,\\
# f^{(1)}(t, \mathbf{y}) &= - \frac{k}{m} y^{(0)},
# &\quad y^{(1)}_0 &= v_0.
# \end{alignat}
#
# with $k=1$; $x_0 = 0$ and $v_0 = +1$.
#
# First step from $t=0$ to $t=h$:
# + [markdown] slideshow={"slide_type": "subslide"}
# \begin{align}
# y^{(0)}_1 &= y^{(0)}_0 + h y^{(1)}_0 = x_0 + v_0 h\\
# y^{(1)}_1 &= y^{(1)}_0 + h \left(-\frac{k}{m}y^{(0)}_0\right) = v_0 + \frac{-k x_0}{m} h
# \end{align}
# + [markdown] slideshow={"slide_type": "fragment"}
# Compare to the equations of movement under a constant force (*ballistic motion*):
#
# $$
# x = x_0 + v_0 h + \frac{1}{2} a h^2, \quad v = v_0 + a h
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# In Euler, the position does not even contain the $h^2$ term, i.e., the acceleration only contributes *indirectly* via the velocity of the *next* step. (Remember: Euler's error is $\mathcal{O}(h^2)$!)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Euler's Rule: Application to 3 oscillators (numerical)
# `git pull` new resources.
#
# Put the notebook `10-ODE_integrators-students.ipynb` *and* the module `integrators.py` into the same directory, for instance a work directory `~/PHY494/10_ODEs`.
#
# Open [integrators.py](https://github.com/ASU-CompMethodsPhysics-PHY494/PHY494-resources/blob/master/10_ODEs/integrators.py) to have a look at it: it contains code to integrate Newton's equations of motions for different forces and with different integrators (based on the work done in the last lecture).
# + slideshow={"slide_type": "subslide"}
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
matplotlib.style.use('ggplot')
import integrators_solution as integrators
# -
t_max = 20
t_harm, y_harm = integrators.integrate_newton(t_max=t_max)
t_anharm, y_anharm = integrators.integrate_newton(t_max=t_max,
force=integrators.F_anharmonic)
t_power, y_power = integrators.integrate_newton(t_max=t_max,
force=integrators.F_power)
# + slideshow={"slide_type": "subslide"}
plt.plot(t_harm, y_harm[:, 0], label=r"harmonic $\frac{k}{2} x^2$")
plt.plot(t_anharm, y_anharm[:, 0], label=r"anharmonic $\frac{k}{2} x^2(1 - \frac{2}{3} \alpha x)$")
plt.plot(t_power, y_power[:, 0], label=r"power $\frac{k}{6} x^6$")
plt.legend(loc="best")
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Phase-space portrait
# Plot position against momentum ($x, p$):
# -
m = 1
x = y_harm[:, 0]
p = m * y_harm[:, 1]
ax = plt.subplot(1,1,1)
ax.plot(x, p)
ax.set_aspect(1)
# + slideshow={"slide_type": "subslide"}
t, y = integrators.integrate_newton(t_max=40,
force=integrators.F_harmonic,
h=0.01)
t2, y2 = integrators.integrate_newton(t_max=40, h=0.001,
force=integrators.F_harmonic)
# + slideshow={"slide_type": "-"}
ax = plt.subplot(1,1,1)
ax.plot(y[:, 0], m * y[:, 1], 'b-', label=r"$h=0.01$")
ax.plot(y2[:, 0], m * y2[:, 1], 'r--', label="$h=0.001$")
ax.set_xlabel("position $x$")
ax.set_ylabel("momentum $p$")
ax.set_aspect(1)
ax.legend(loc="best")
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Energy
# Energy is conserved: stringent test for the integrator!
#
# $$
# E = T_\text{kin} + U\\
# \frac{dE}{dt} = 0
# $$
# with $T_\text{kin} = \frac{1}{2} m v^2$.
#
# Calculate and plot the three energy terms:
# + slideshow={"slide_type": "subslide"}
KE = 0.5 * m * y_harm[:, 1]**2
PE = integrators.U_harmonic(y_harm[:, 0])
energy = KE + PE
plt.plot(t_harm, KE, 'r-', t_harm, PE, 'b-', t_harm, energy, 'k--')
# + [markdown] slideshow={"slide_type": "subslide"}
# Harmonic oscillator with larger timestep $h=0.01$
# + slideshow={"slide_type": "-"}
m = 1
t, y = integrators.integrate_newton(t_max=500, force=integrators.F_harmonic, h=0.01, mass=m)
x, v = y.T
KE = 1/2 * m * v**2
PE = integrators.U_harmonic(x)
energy = KE + PE
# + slideshow={"slide_type": "subslide"}
ax = plt.subplot(1,1,1)
ax.plot(t, KE, 'r-', t, PE, 'b-', t, energy, 'k--')
ax.set_xlim(0, 20)
ax.set_ylim(0, 0.7)
ax.set_xlabel("time")
ax.set_ylabel("energy")
ax.set_title("Euler: harmonic oscillator")
# -
# Note how energy conservation becomes worse when $h$ is increased.
# + [markdown] slideshow={"slide_type": "subslide"}
# For longer time scales, the energy just keeps on increasing — clearly, we are not solving the equations of motions in a physically meaningful way.
# + slideshow={"slide_type": "-"}
ax.set_ylim(0, 80)
ax.set_xlim(0,500)
ax.figure
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Quantify energy drift
# Measure the relative departure from energy conservation
#
# $$
# \epsilon_E(t) = \log_{10}\left|\frac{E(t) - E(t=0)}{E(t=0)}\right|
# $$
#
# (By using the decadic logarithm, $-\epsilon_E$ is a direct measure of the precision.)
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### Harmonic oscillator
# Analyze energy drift for the last harmonic oscillator example ($h=0.01$):
# + slideshow={"slide_type": "-"}
energy_precision = np.log10(energy/energy[0] - 1)
plt.plot(t, energy_precision)
plt.ylabel("log(relative error in $E$)")
plt.xlabel("time")
# -
# As we already know, with such a large time step, energy conservation is very bad with the *Euler* integrator: The relative (!) error quickly becomes 100% (after $t=200$).
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Code re-use: energy analysis as a function
# The module `integrators` contains the function `integrators.analyze_energies(t, y, U)` to automate the task of analysing the energy. You *must* provide the appropriate potential energy function `U`.
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Problem: Make `analyze_energies()` work and analyze the power law oscillator
# The code is *incomplete*.
# 1. Open `integrators.py` in your editor.
# 2. Complete the `kinetic_energy()` function (remember: $T_\text{kin} = \frac{1}{2} m v^2$)
#
# Then `reload()` the module:
# -
# when working on the integrator module you need to reload() whenever you make changes
from importlib import reload # for Python 3
reload(integrators);
# and try it out for power law oscillator with a large time step $h = 0.01$. Can you find a value of $h$ that will give you energy precision to 4th decimal over 40 time units?
# ###### Energy stability analysis of the 6-th power oscillator
# $x^6$ potential with $h=0.01$:
# + slideshow={"slide_type": "subslide"}
t, y = integrators.integrate_newton(t_max=40, force=integrators.F_power,
h=0.01)
# -
integrators.analyze_energies(t, y, integrators.U_power)
# + [markdown] slideshow={"slide_type": "subslide"}
# Trying 100 times smaller $h$ (takes 100 times longer!!)
# -
t, y = integrators.integrate_newton(t_max=40, force=integrators.F_power,
h=0.0001)
integrators.analyze_energies(t, y, integrators.U_power)
# + [markdown] slideshow={"slide_type": "fragment"}
# *Reducing $h$ further does not seem to help and takes order of magnitude longer to run*. **We need a better integrator!**
# + [markdown] slideshow={"slide_type": "slide"}
# ## Runge-Kutta methods
# Explicit RK methods propagate with the general form
#
# $$
# y_{n+1} = y_n + h \sum_{i=1}^s b_i k_i, \quad t_{n+1} = t_n + h
# $$
#
# where $s$ is the number of *stages* and the coefficients $a_{ij}$, $b_i$ and $c_i$ define the method.
#
# \begin{align}
# k_1 &= f(t_n, y_n)\\
# k_2 &= f(t_n + c_2 h, y_n + h(a_{21} k_1))\\
# k_3 &= f(t_n + c_3 h, y_n + h(a_{31} k_1 + a_{32} k_2))\\
# \vdots& \\
# k_s &= f(t_n + c_s h, y_n + h(a_{s1} k_1 + a_{s2} k_2 + \cdots + a_{s,s-1} k_{s-1})).
# \end{align}
# -
# Basic idea: improve estimate for the slope by using weighted averages of slopes over the interval.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### RK4
# General purpose ODE solver.
#
# $$
# y_{n+1} = y_n + \frac{h}{6}\Big(k_1 + 2 k_2 + 2 k_3 + k_4\Big), \quad t_{n+1} = t_n + h
# $$
#
# and
#
# \begin{align}
# k_1 &= f(t_n, y_n)\\
# k_2 &= f(t_n + \frac{1}{2} h, y_n + \frac{1}{2} h k_1)\\
# k_3 &= f(t_n + \frac{1}{2} h, y_n + \frac{1}{2} h k_2)\\
# k_4 &= f(t_n + h, y_n + h k_3)
# \end{align}
# -
# `rk4` can obtain $\mathcal{O}(h^4)$ precision.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### RK2
# `rk2` is simpler to derive and understand than `rk4` (which we will use more often).
# -
# Start with the exact solution
#
# $$
# \frac{dy}{dt} = f(t, y) \quad\text{and}\quad y(t) = \int f(t, y)dt\\
# y_{n+1} = y_n + \int_{t_n}^{t_{n+1}} f(t, y)dt
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# Expand $f(t, y)$ around *midpoint* $t_{n+1/2} = t + \frac{h}{2}$ of integration interval
#
# \begin{gather}
# f(t, y) \approx f(t_{n+1/2}, y_{n+1/2}) + (t - t_{n+1/2})\, f^{(1)}(t_{n+1/2}) +
# \mathcal{O}(h^2)
# \end{gather}
# + [markdown] slideshow={"slide_type": "fragment"}
# Integrate the *expansion* (Taylor series around midpoint):
# \begin{align}
# \int_{t_n}^{t_{n+1}} f(t, y) \, dt &\approx h\, f(t_{n+1/2}, y_{n+1/2}) + \mathcal{O}(h^3)\\
# y_{n+1} &= y_n + h\, f(t_{n+1/2}, y_{n+1/2}) + \mathcal{O}(h^3)
# \end{align}
#
# I.e., use **slope at midpoint**! (The $\mathcal{O}(h^2)$ term in the integral is zero because $[(h/2)^2 - (h/2)^2] = 0$.)
# + [markdown] slideshow={"slide_type": "subslide"}
# But we don't have $y_{n+1/2}$ so we use Euler's rule to estimate it to $\mathcal{O}(h^2)$:
#
# $$
# y_{n+1/2} \approx y_n + \frac{1}{2} h\, f(t_n, y_n)
# $$
#
# and insert into $y_{n+1} = y_n + h\, f(t_{n+1/2}, y_{n+1/2})$ (so it is still $\mathcal{O}(h^3)$ accurate).
#
# + [markdown] slideshow={"slide_type": "subslide"}
# Altogether: **RK2** algorithm
#
# $$
# \mathbf{y}_{n+1} = \mathbf{y}_n + h \mathbf{k}_2, \quad t_{n+1} = t_n + h
# $$
#
# and
#
# \begin{align}
# \mathbf{k}_1 &= \mathbf{f}(t_n, \mathbf{y}_n)\\
# \mathbf{k}_2 &= \mathbf{f}(t_n + \frac{1}{2} h, \mathbf{y}_n + \frac{1}{2} h \mathbf{k}_1)
# \end{align}
#
# with $\mathcal{O}(h^3)$ accuracy.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### RK2 applied to the 1D harmonic oscillator (theory)
#
# \begin{alignat}{3}
# f^{(0)}(t, \mathbf{y}) &= y^{(1)},
# &\quad y^{(0)}_0 &= x_0,\\
# f^{(1)}(t, \mathbf{y}) &= - \frac{k}{m} y^{(0)},
# &\quad y^{(1)}_0 &= v_0.
# \end{alignat}
#
# with $k=1$; $x_0 = 0$ and $v_0 = +1$.
# + [markdown] slideshow={"slide_type": "subslide"}
# First step from $t=0$ to $t=h$:
# + [markdown] slideshow={"slide_type": "-"}
# \begin{align}
# y^{(0)}_{1} &=
# y^{(0)} _{0} + hf^{(0)} \Big(\frac{h}{2}, y^{(0)}_{0} + hk_{1} \Big)\\
# &= x_{0} + h\left[v_{0} + \frac{h}{2} \left(-\frac{k}{m} x_{0}\right) \right]\\
# &= x_0 + v_0 h + \frac{1}{2} \left(-\frac{k}{m} x_{0}\right) h^2\\
# y^{(1)}_{1} &= y^{(1)}_{0} +
# h f^{(1)}\left(\frac{h}{2},
# \mathbf{y}_{0} + \frac{h}{2} \mathbf{f}(0, \mathbf{y}_{0})\right) \\
# &= v_{0} + \frac{h}{m} \left[-k\left(x_{0} - \frac{h}{2} k x_{0}\right)\right]
# \end{align}
# + [markdown] slideshow={"slide_type": "-"}
# Note that `rk2` contains a $h^2$ term in the position update.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### RK2 applied in practice
# Let's implement `rk2`
#
# \begin{align}
# \mathbf{y}_{n+1} &= \mathbf{y}_n + h \mathbf{k}_2\\
# \mathbf{k}_1 &= \mathbf{f}(t_n, \mathbf{y}_n)\\
# \mathbf{k}_2 &= \mathbf{f}(t_n + \frac{1}{2} h, \mathbf{y}_n + \frac{1}{2} h \mathbf{k}_1)
# \end{align}
#
# in our `integrators.py`.
# -
from importlib import reload
reload(integrators);
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### RK2 for the power potential
# + slideshow={"slide_type": "-"}
t, y = integrators.integrate_newton(t_max=40, force=integrators.F_power,
integrator=integrators.rk2, h=0.01)
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### phase-space portrait
# -
plt.plot(y[:, 0], m*y[:, 1])
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### energy drift
# -
integrators.analyze_energies(t, y, integrators.U_power)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### RK4 applied in practice
# Implement `rk4` in `integrator.py`:
#
# \begin{align}
# y_{n+1} &= y_n + \frac{h}{6}\Big(k_1 + 2 k_2 + 2 k_3 + k_4\Big)\\
# k_1 &= f(t_n, y_n)\\
# k_2 &= f(t_n + \frac{1}{2} h, y_n + \frac{1}{2} h k_1)\\
# k_3 &= f(t_n + \frac{1}{2} h, y_n + \frac{1}{2} h k_2)\\
# k_4 &= f(t_n + h, y_n + h k_3)
# \end{align}
# -
reload(integrators)
# and test the 6th power potential again. What level of energy precision can you obtain over 40 time units?
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### RK4 for the 6th power potential
# Note that we can easily run `rk4` with a $h=0.01$!
# + slideshow={"slide_type": "-"}
t, y = integrators.integrate_newton(t_max=40, force=integrators.F_power,
integrator=integrators.rk4, h=0.01)
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### phase-space protrait
# -
plt.plot(y[:, 0], m*y[:, 1])
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### energy drift
# -
integrators.analyze_energies(t, y, integrators.U_power)
# You should be able to get 8 - 11 decimals precision from `rk4` with a smaller $h$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### High precision RK4 for the 6th power potential
# Try a 10 times smaller step $h=0.001$
# + slideshow={"slide_type": "-"}
t, y = integrators.integrate_newton(t_max=40, force=integrators.F_power,
integrator=integrators.rk4, h=0.001)
integrators.analyze_energies(t, y, integrators.U_power)
# -
# This looks like stable integration with an error of not worse than $10^{-12}$: *Looking very good!* (Especially when you compare it to *Euler* with $h=0.001$!)
#
# It looks as if the main source of error comes from the force evaluation because the error spikes when the potential energy is dominant.
| 10_ODEs/10-ODE_integrators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Data Pre-processing
import numpy as np
import re
# +
import pandas as pd
df = pd.read_csv('Conversational_Pos_Politeness_LIWC_data.csv')
df.rename(columns={'A':'Text Content',
'B':'Document',
'C':'Codes',
'D':'author_association',
'E':'is_poster',
'F':'quote_len',
'G':'percent_len_thread',
'H':'percent_len_comment',
'I':'percent_pos_thread',
'J':'percent_pos_comment',
'K':'is_first_comment',
'L':'is_last_comment',
'M':'percent_start_time_thread',
'N':'percent_end_time_thread',
'O':'percent_previous_time_thread',
'P':'percent_next_time_thread',
'Q':'contain_code',
'R':'pos_str',
'S':'polite_score'}, inplace=True)
# -
# check if any row contains NaN
df1 = df[df.isnull().any(axis=1)]
print(df1.shape[0])
display(df)
def flag_category(text, flag):
if flag == 'argu_part':
if text.find('claim') > -1:
return 1
elif text.find('warrant') > -1:
return 2
elif text.find('ground') > -1:
return 3
else:
return 0
elif flag =='argu_count':
argu_count = 0
if text.find('claim') > -1:
argu_count += 1
if text.find('warrant') > -1:
argu_count += 1
if text.find('ground') > -1:
argu_count += 1
return argu_count
else:
return text.find(flag) > -1
df['is_non_argumentative'] = df.apply(lambda x: flag_category(x['Codes'], 'non-argumentative'), axis=1)
df['is_against'] = df.apply(lambda x: flag_category(x['Codes'], 'against'), axis=1)
df['argu_part'] = df.apply(lambda x: flag_category(x['Codes'], 'argu_part'), axis=1)
df['argu_count'] = df.apply(lambda x: flag_category(x['Codes'], 'argu_count'), axis=1)
# ## 1.2. Check if argu_part is assigned correctly
# Quotation take one and only one of 'claim', 'warrant' or 'data'
#
# this dataframe needs to be empty
df[(df.is_non_argumentative == False) & (df.argu_part == 0)]
# this dataframe needs to be empty
df[(df.is_non_argumentative == False) & (df.argu_count > 1)]
# ## 1.3. Write to file
# remove argu_count column, so it does not write to the file
df = df.drop(['argu_count'], axis=1)
df.to_csv('Preprocessed.csv', index=False)
| Argument Extraction Program and Results/Preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preface
# Welcome! Allow me to be the first to offer my congratulations on your decision to take an interest in Applied Predictive Modeling with Python! This is a collection of IPython Notebooks that provides an interactive way to reproduce this awesome [book](http://www.amazon.com/Applied-Predictive-Modeling-Max-Kuhn/dp/1461468485) by Kuhn and Johnson.
#
# If you experience any problems along the way or have any feedback at all, please reach out to me.
#
# Best Regards, <br />
# <NAME> <br />
# Email: <EMAIL> <br />
# Twitter: @_LeiG
# ## Setups
import numpy
import scipy
import pandas
import sklearn
import matplotlib
import rpy2
import pyearth
import statsmodels
# ## Prepare Datasets
# Thanks to the authors, all datasets that are necessary in order to reproduce the examples in the book are available in the *.RData* format from their R package $\texttt{caret}$ and $\texttt{AppliedPredictiveModeling}$. To prepare them for our purpose, I did a little hack so that you can download all the datasets and convert them from *.RData* to *.csv* by running this script "[fetch_data.py](https://github.com/LeiG/Applied-Predictive-Modeling-with-Python/blob/master/fetch_data.py)".
# %run ../fetch_data.py
# # 1. Introduction
# **Predictive modeling**: the process of developing a mathematical tool or model that generates an accurate prediction.
# There are a number of common reasons why predictive models fail, e.g,
#
# - inadequante pre-processing of the data
# - inadequate model validation
# - unjustified extrapolation
# - over-fitting the model to the existing data
# - explore relatively few models when searching for relationships
# ## 1.1 Prediction Versus Interpretation
# The trade-off between prediction and interpretation depends on the primary goal of the task. The unfortunate reality is that as we push towards higher accuracy, models become more complex and their interpretability becomes more difficult.
# ## 1.2 Key Ingredients of Predictive Models
# The foundation of an effective predictive model is laid with *intuition* and *deep knowledge of the problem context*, which are entirely vital for driving decisions about model development. The process begins with *relevant* data.
# ## 1.3 Terminology
# - The *sample*, *data point*, *observation*, or *instance* refer to a single independent unit of data
# - The *training* set consists of the data used to develop models while the *test* or *validation* set is used solely for evaluating the performance of a final set of candidate models. **NOTE**: usually people refer to the *validation* set for evaluating candidates and divide *training* set using cross-validation into several sub-*training* and *test* sets to tune parameters in model development.
# - The *predictors*, *independent variables*, *attributes*, or *descriptors* are the data used as input for the prediction equation.
# - The *outcome*, *dependent variable*, *target*, *class*, or *response* refer to the outcome event or quantity that is being predicted.
# ## 1.4 Example Data Sets and Typical Data Scenarios
# ## 1.5 Overview
# - **Part I General Strategies**
# - Ch.2 A short tour of the predictive modeling process
# - Ch.3 Data pre-processing
# - Ch.4 Over-fitting and model tuning
# - **Part II Regression Models**
# - Ch.5 Measuring performance in regression models
# - Ch.6 Linear regression and its cousins
# - Ch.7 Nonlinear regression models
# - Ch.8 Regression trees and rule-based models
# - Ch.9 A summary of solubility models
# - Ch.10 Case study: compressive strength of concrete
# - **Part III Classification Models**
# - Ch.11 Measuring performance in classification models
# - Ch.12 Discriminant analysis and other linear classification models
# - Ch.13 Nonlinear classification models
# - Ch.14 Classification trees and rule-based models
# - Ch.15 A summary of grant application models
# - Ch.16 Remedies for severe class imbalance
# - Ch.17 Case study: job scheduling
# - **Part IV Other Considerations**
# - Ch.18 Measuring predictor importance
# - Ch.19 An introduction to feature selection
# - Ch.20 Factors that can affect model performance
| notebooks/.ipynb_checkpoints/Chapter 1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Source of the materials**: Biopython cookbook (adapted)
# <font color='red'>Status: Draft</font>
# BLAST and other sequence search tools (*experimental code*) {#chapter:searchio}
# ===========================================================
#
# *WARNING*: This chapter of the Tutorial describes an *experimental*
# module in Biopython. It is being included in Biopython and documented
# here in the tutorial in a pre-final state to allow a period of feedback
# and refinement before we declare it stable. Until then the details will
# probably change, and any scripts using the current `Bio.SearchIO` would
# need to be updated. Please keep this in mind! For stable code working
# with NCBI BLAST, please continue to use Bio.Blast described in the
# preceding Chapter \[chapter:blast\].
#
# Biological sequence identification is an integral part of
# bioinformatics. Several tools are available for this, each with their
# own algorithms and approaches, such as BLAST (arguably the most
# popular), FASTA, HMMER, and many more. In general, these tools usually
# use your sequence to search a database of potential matches. With the
# growing number of known sequences (hence the growing number of potential
# matches), interpreting the results becomes increasingly hard as there
# could be hundreds or even thousands of potential matches. Naturally,
# manual interpretation of these searches’ results is out of the question.
# Moreover, you often need to work with several sequence search tools,
# each with its own statistics, conventions, and output format. Imagine
# how daunting it would be when you need to work with multiple sequences
# using multiple search tools.
#
# We know this too well ourselves, which is why we created the
# `Bio.SearchIO` submodule in Biopython. `Bio.SearchIO` allows you to
# extract information from your search results in a convenient way, while
# also dealing with the different standards and conventions used by
# different search tools. The name `SearchIO` is a homage to BioPerl’s
# module of the same name.
#
# In this chapter, we’ll go through the main features of `Bio.SearchIO` to
# show what it can do for you. We’ll use two popular search tools along
# the way: BLAST and BLAT. They are used merely for illustrative purposes,
# and you should be able to adapt the workflow to any other search tools
# supported by `Bio.SearchIO` in a breeze. You’re very welcome to follow
# along with the search output files we’ll be using. The BLAST output file
# can be downloaded
# [here](http://biopython.org/SRC/Tests/Tutorial/my_blast.xml), and the
# BLAT output file
# [here](http://biopython.org/SRC/Tests/Tutorial/my_blat.psl). Both output
# files were generated using this sequence:
#
#
# ```
# >mystery_seq
# CCCTCTACAGGGAAGCGCTTTCTGTTGTCTGAAAGAAAAGAAAGTGCTTCCTTTTAGAGGG
#
# ```
#
# The BLAST result is an XML file generated using `blastn` against the
# NCBI `refseq_rna` database. For BLAT, the sequence database was the
# February 2009 `hg19` human genome draft and the output format is PSL.
#
# We’ll start from an introduction to the `Bio.SearchIO` object model. The
# model is the representation of your search results, thus it is core to
# `Bio.SearchIO` itself. After that, we’ll check out the main functions in
# `Bio.SearchIO` that you may often use.
#
# Now that we’re all set, let’s go to the first step: introducing the core
# object model.
#
# The SearchIO object model {#sec:searchio-model}
# -------------------------
#
# Despite the wildly differing output styles among many sequence search
# tools, it turns out that their underlying concept is similar:
#
# - The output file may contain results from one or more search queries.
#
# - In each search query, you will see one or more hits from the given
# search database.
#
# - In each database hit, you will see one or more regions containing
# the actual sequence alignment between your query sequence and the
# database sequence.
#
# - Some programs like BLAT or Exonerate may further split these regions
# into several alignment fragments (or blocks in BLAT and possibly
# exons in exonerate). This is not something you always see, as
# programs like BLAST and HMMER do not do this.
#
# Realizing this generality, we decided use it as base for creating the
# `Bio.SearchIO` object model. The object model consists of a nested
# hierarchy of Python objects, each one representing one concept outlined
# above. These objects are:
#
# - `QueryResult`, to represent a single search query.
#
# - `Hit`, to represent a single database hit. `Hit` objects are
# contained within `QueryResult` and in each `QueryResult` there is
# zero or more `Hit` objects.
#
# - `HSP` (short for high-scoring pair), to represent region(s) of
# significant alignments between query and hit sequences. `HSP`
# objects are contained within `Hit` objects and each `Hit` has one or
# more `HSP` objects.
#
# - `HSPFragment`, to represent a single contiguous alignment between
# query and hit sequences. `HSPFragment` objects are contained within
# `HSP` objects. Most sequence search tools like BLAST and HMMER unify
# `HSP` and `HSPFragment` objects as each `HSP` will only have a
# single `HSPFragment`. However there are tools like BLAT and
# Exonerate that produce `HSP` containing multiple `HSPFragment`.
# Don’t worry if this seems a tad confusing now, we’ll elaborate more
# on these two objects later on.
#
# These four objects are the ones you will interact with when you use
# `Bio.SearchIO`. They are created using one of the main `Bio.SearchIO`
# methods: `read`, `parse`, `index`, or `index_db`. The details of these
# methods are provided in later sections. For this section, we’ll only be
# using read and parse. These functions behave similarly to their
# `Bio.SeqIO` and `Bio.AlignIO` counterparts:
#
# - `read` is used for search output files with a single query and
# returns a `QueryResult` object
#
# - `parse` is used for search output files with multiple queries and
# returns a generator that yields `QueryResult` objects
#
# With that settled, let’s start probing each `Bio.SearchIO` object,
# beginning with `QueryResult`.
#
# ### QueryResult {#sec:searchio-qresult}
#
# The QueryResult object represents a single search query and contains
# zero or more Hit objects. Let’s see what it looks like using the BLAST
# file we have:
#
#
from Bio import SearchIO
blast_qresult = SearchIO.read('data/my_blast.xml', 'blast-xml')
print(blast_qresult)
#
# We’ve just begun to scratch the surface of the object model, but you can
# see that there’s already some useful information. By invoking `print` on
# the `QueryResult` object, you can see:
#
# - The program name and version (blastn version 2.2.27+)
#
# - The query ID, description, and its sequence length (ID is 42291,
# description is ‘mystery\_seq’, and it is 61 nucleotides long)
#
# - The target database to search against (refseq\_rna)
#
# - A quick overview of the resulting hits. For our query sequence,
# there are 100 potential hits (numbered 0–99 in the table). For each
# hit, we can also see how many HSPs it contains, its ID, and a
# snippet of its description. Notice here that `Bio.SearchIO`
# truncates the hit table overview, by showing only hits numbered
# 0–29, and then 97–99.
#
# Now let’s check our BLAT results using the same procedure as above:
#
#
blat_qresult = SearchIO.read('data/my_blat.psl', 'blat-psl')
print(blat_qresult)
#
# You’ll immediately notice that there are some differences. Some of these
# are caused by the way PSL format stores its details, as you’ll see. The
# rest are caused by the genuine program and target database differences
# between our BLAST and BLAT searches:
#
# - The program name and version. `Bio.SearchIO` knows that the program
# is BLAT, but in the output file there is no information regarding
# the program version so it defaults to ‘<unknown version>’.
#
# - The query ID, description, and its sequence length. Notice here that
# these details are slightly different from the ones we saw in BLAST.
# The ID is ‘mystery\_seq’ instead of 42991, there is no known
# description, but the query length is still 61. This is actually a
# difference introduced by the file formats themselves. BLAST
# sometimes creates its own query IDs and uses your original ID as the
# sequence description.
#
# - The target database is not known, as it is not stated in the BLAT
# output file.
#
# - And finally, the list of hits we have is completely different. Here,
# we see that our query sequence only hits the ‘chr19’ database entry,
# but in it we see 17 HSP regions. This should not be surprising
# however, given that we are using a different program, each with its
# own target database.
#
# All the details you saw when invoking the `print` method can be accessed
# individually using Python’s object attribute access notation (a.k.a. the
# dot notation). There are also other format-specific attributes that you
# can access using the same method.
#
#
print("%s %s" % (blast_qresult.program, blast_qresult.version))
print("%s %s" % (blat_qresult.program, blat_qresult.version))
blast_qresult.param_evalue_threshold # blast-xml specific
#
# For a complete list of accessible attributes, you can check each
# format-specific documentation. Here are the ones [for
# BLAST](http://biopython.org/DIST/docs/api/Bio.SearchIO.BlastIO-module.html)
# and for
# [BLAT](http://biopython.org/DIST/docs/api/Bio.SearchIO.BlatIO-module.html).
#
# Having looked at using `print` on `QueryResult` objects, let’s drill
# down deeper. What exactly is a `QueryResult`? In terms of Python
# objects, `QueryResult` is a hybrid between a list and a dictionary. In
# other words, it is a container object with all the convenient features
# of lists and dictionaries.
#
# Like Python lists and dictionaries, `QueryResult` objects are iterable.
# Each iteration returns a `Hit` object:
#
#
for hit in blast_qresult:
hit
#
# To check how many items (hits) a `QueryResult` has, you can simply
# invoke Python’s `len` method:
#
#
len(blast_qresult)
len(blat_qresult)
#
# Like Python lists, you can retrieve items (hits) from a `QueryResult`
# using the slice notation:
#
#
blast_qresult[0] # retrieves the top hit
blast_qresult[-1] # retrieves the last hit
#
# To retrieve multiple hits, you can slice `QueryResult` objects using the
# slice notation as well. In this case, the slice will return a new
# `QueryResult` object containing only the sliced hits:
#
#
blast_slice = blast_qresult[:3] # slices the first three hits
print(blast_slice)
#
# Like Python dictionaries, you can also retrieve hits using the hit’s ID.
# This is particularly useful if you know a given hit ID exists within a
# search query results:
#
#
blast_qresult['gi|731339628|ref|XM_010682658.1|']
#
# You can also get a full list of `Hit` objects using `hits` and a full
# list of `Hit` IDs using `hit_keys`:
#
#
blast_qresult.hits
blast_qresult.hit_keys
#
# What if you just want to check whether a particular hit is present in
# the query results? You can do a simple Python membership test using the
# `in` keyword:
#
#
'gi|262205317|ref|NR_030195.1|' in blast_qresult
'gi|731339628|ref|XM_010682658.1|' in blast_qresult
#
# Sometimes, knowing whether a hit is present is not enough; you also want
# to know the rank of the hit. Here, the `index` method comes to the
# rescue:
#
#
blast_qresult.index('gi|595807351|ref|XM_007202530.1|')
#
# Remember that we’re using Python’s indexing style here, which is
# zero-based. This means our hit above is ranked at no. 23, not 22.
#
# Also, note that the hit rank you see here is based on the native hit
# ordering present in the original search output file. Different search
# tools may order these hits based on different criteria.
#
# If the native hit ordering doesn’t suit your taste, you can use the
# `sort` method of the `QueryResult` object. It is very similar to
# Python’s `list.sort` method, with the addition of an option to create a
# new sorted `QueryResult` object or not.
#
# Here is an example of using `QueryResult.sort` to sort the hits based on
# each hit’s full sequence length. For this particular sort, we’ll set the
# `in_place` flag to `False` so that sorting will return a new
# `QueryResult` object and leave our initial object unsorted. We’ll also
# set the `reverse` flag to `True` so that we sort in descending order.
#
#
for hit in blast_qresult[:5]: # id and sequence length of the first five hits
print("%s %i" % (hit.id, hit.seq_len))
sort_key = lambda hit: hit.seq_len
sorted_qresult = blast_qresult.sort(key=sort_key, reverse=True, in_place=False)
for hit in sorted_qresult[:5]:
print("%s %i" % (hit.id, hit.seq_len))
#
# The advantage of having the `in_place` flag here is that we’re
# preserving the native ordering, so we may use it again later. You should
# note that this is not the default behavior of `QueryResult.sort`,
# however, which is why we needed to set the `in_place` flag to `True`
# explicitly.
#
# At this point, you’ve known enough about `QueryResult` objects to make
# it work for you. But before we go on to the next object in the
# `Bio.SearchIO` model, let’s take a look at two more sets of methods that
# could make it even easier to work with `QueryResult` objects: the
# `filter` and `map` methods.
#
# If you’re familiar with Python’s list comprehensions, generator
# expressions or the built in `filter` and `map` functions, you’ll know
# how useful they are for working with list-like objects (if you’re not,
# check them out!). You can use these built in methods to manipulate
# `QueryResult` objects, but you’ll end up with regular Python lists and
# lose the ability to do more interesting manipulations.
#
# That’s why, `QueryResult` objects provide its own flavor of `filter` and
# `map` methods. Analogous to `filter`, there are `hit_filter` and
# `hsp_filter` methods. As their name implies, these methods filter its
# `QueryResult` object either on its `Hit` objects or `HSP` objects.
# Similarly, analogous to `map`, `QueryResult` objects also provide the
# `hit_map` and `hsp_map` methods. These methods apply a given function to
# all hits or HSPs in a `QueryResult` object, respectively.
#
# Let’s see these methods in action, beginning with `hit_filter`. This
# method accepts a callback function that checks whether a given `Hit`
# object passes the condition you set or not. In other words, the function
# must accept as its argument a single `Hit` object and returns `True` or
# `False`.
#
# Here is an example of using `hit_filter` to filter out `Hit` objects
# that only have one HSP:
#
#
filter_func = lambda hit: len(hit.hsps) > 1 # the callback function
len(blast_qresult) # no. of hits before filtering
filtered_qresult = blast_qresult.hit_filter(filter_func)
len(filtered_qresult) # no. of hits after filtering
for hit in filtered_qresult[:5]: # quick check for the hit lengths
print("%s %i" % (hit.id, len(hit.hsps)))
#
# `hsp_filter` works the same as `hit_filter`, only instead of looking at
# the `Hit` objects, it performs filtering on the `HSP` objects in each
# hits.
#
# As for the `map` methods, they too accept a callback function as their
# arguments. However, instead of returning `True` or `False`, the callback
# function must return the modified `Hit` or `HSP` object (depending on
# whether you’re using `hit_map` or `hsp_map`).
#
# Let’s see an example where we’re using `hit_map` to rename the hit IDs:
#
#
# +
def map_func(hit):
hit.id = hit.id.split('|')[3] # renames 'gi|301171322|ref|NR_035857.1|' to 'NR_035857.1'
return hit
mapped_qresult = blast_qresult.hit_map(map_func)
for hit in mapped_qresult[:5]:
print(hit.id)
# -
#
# Again, `hsp_map` works the same as `hit_map`, but on `HSP` objects
# instead of `Hit` objects.
#
# ### Hit {#sec:searchio-hit}
#
# `Hit` objects represent all query results from a single database entry.
# They are the second-level container in the `Bio.SearchIO` object
# hierarchy. You’ve seen that they are contained by `QueryResult` objects,
# but they themselves contain `HSP` objects.
#
# Let’s see what they look like, beginning with our BLAST search:
#
#
from Bio import SearchIO
blast_qresult = SearchIO.read('data/my_blast.xml', 'blast-xml')
blast_hit = blast_qresult[3] # fourth hit from the query result
print(blast_hit)
#
# You see that we’ve got the essentials covered here:
#
# - The query ID and description is present. A hit is always tied to a
# query, so we want to keep track of the originating query as well.
# These values can be accessed from a hit using the `query_id` and
# `query_description` attributes.
#
# - We also have the unique hit ID, description, and full
# sequence lengths. They can be accessed using `id`, `description`,
# and `seq_len`, respectively.
#
# - Finally, there’s a table containing quick information about the HSPs
# this hit contains. In each row, we’ve got the important HSP details
# listed: the HSP index, its e-value, its bit score, its span (the
# alignment length including gaps), its query coordinates, and its
# hit coordinates.
#
# Now let’s contrast this with the BLAT search. Remember that in the BLAT
# search we had one hit with 17 HSPs.
#
#
blat_qresult = SearchIO.read('data/my_blat.psl', 'blat-psl')
blat_hit = blat_qresult[0] # the only hit
print(blat_hit)
#
# Here, we’ve got a similar level of detail as with the BLAST hit we saw
# earlier. There are some differences worth explaining, though:
#
# - The e-value and bit score column values. As BLAT HSPs do not have
# e-values and bit scores, the display defaults to ‘?’.
#
# - What about the span column? The span values is meant to display the
# complete alignment length, which consists of all residues and any
# gaps that may be present. The PSL format do not have this
# information readily available and `Bio.SearchIO` does not attempt to
# try guess what it is, so we get a ‘?’ similar to the e-value and bit
# score columns.
#
# In terms of Python objects, `Hit` behaves almost the same as Python
# lists, but contain `HSP` objects exclusively. If you’re familiar with
# lists, you should encounter no difficulties working with the `Hit`
# object.
#
# Just like Python lists, `Hit` objects are iterable, and each iteration
# returns one `HSP` object it contains:
#
#
for hsp in blast_hit:
hsp
#
# You can invoke `len` on a `Hit` to see how many `HSP` objects it has:
#
#
len(blast_hit)
len(blat_hit)
#
# You can use the slice notation on `Hit` objects, whether to retrieve
# single `HSP` or multiple `HSP` objects. Like `QueryResult`, if you slice
# for multiple `HSP`, a new `Hit` object will be returned containing only
# the sliced `HSP` objects:
#
#
blat_hit[0] # retrieve single items
sliced_hit = blat_hit[4:9] # retrieve multiple items
len(sliced_hit)
print(sliced_hit)
#
# You can also sort the `HSP` inside a `Hit`, using the exact same
# arguments like the sort method you saw in the `QueryResult` object.
#
# Finally, there are also the `filter` and `map` methods you can use on
# `Hit` objects. Unlike in the `QueryResult` object, `Hit` objects only
# have one variant of `filter` (`Hit.filter`) and one variant of `map`
# (`Hit.map`). Both of `Hit.filter` and `Hit.map` work on the `HSP`
# objects a `Hit` has.
#
# ### HSP {#sec:searchio-hsp}
#
# `HSP` (high-scoring pair) represents region(s) in the hit sequence that
# contains significant alignment(s) to the query sequence. It contains the
# actual match between your query sequence and a database entry. As this
# match is determined by the sequence search tool’s algorithms, the `HSP`
# object contains the bulk of the statistics computed by the search tool.
# This also makes the distinction between `HSP` objects from different
# search tools more apparent compared to the differences you’ve seen in
# `QueryResult` or `Hit` objects.
#
# Let’s see some examples from our BLAST and BLAT searches. We’ll look at
# the BLAST HSP first:
#
#
from Bio import SearchIO
blast_qresult = SearchIO.read('data/my_blast.xml', 'blast-xml')
blast_hsp = blast_qresult[0][0] # first hit, first hsp
print(blast_hsp)
#
# Just like `QueryResult` and `Hit`, invoking `print` on an `HSP` shows
# its general details:
#
# - There are the query and hit IDs and descriptions. We need these to
# identify our `HSP`.
#
# - We’ve also got the matching range of the query and hit sequences.
# The slice notation we’re using here is an indication that the range
# is displayed using Python’s indexing style (zero-based, half open).
# The number inside the parenthesis denotes the strand. In this case,
# both sequences have the plus strand.
#
# - Some quick statistics are available: the e-value and bitscore.
#
# - There is information about the HSP fragments. Ignore this for now;
# it will be explained later on.
#
# - And finally, we have the query and hit sequence alignment itself.
#
# These details can be accessed on their own using the dot notation, just
# like in `QueryResult` and `Hit`:
#
#
blast_hsp.query_range
#
#
blast_hsp.evalue
#
# They’re not the only attributes available, though. `HSP` objects come
# with a default set of properties that makes it easy to probe their
# various details. Here are some examples:
#
#
blast_hsp.hit_start # start coordinate of the hit sequence
def map_func(hit):
hit.id = hit.id.split('|')[3] # renames 'gi|301171322|ref|NR_035857.1|' to 'NR_035857.1'
return hitblast_hsp.query_span # how many residues in the query sequence
blast_hsp.aln_span # how long the alignment is
#
# Check out the `HSP`
# [documentation](http://biopython.org/DIST/docs/api/Bio.SearchIO._model.hsp-module.html)
# for a full list of these predefined properties.
#
# Furthermore, each sequence search tool usually computes its own
# statistics / details for its `HSP` objects. For example, an XML BLAST
# search also outputs the number of gaps and identical residues. These
# attributes can be accessed like so:
#
#
blast_hsp.gap_num # number of gaps
blast_hsp.ident_num # number of identical residues
#
# These details are format-specific; they may not be present in other
# formats. To see which details are available for a given sequence search
# tool, you should check the format’s documentation in `Bio.SearchIO`.
# Alternatively, you may also use `.__dict__.keys()` for a quick list of
# what’s available:
#
#
blast_hsp.__dict__.keys()
#
# Finally, you may have noticed that the `query` and `hit` attributes of
# our HSP are not just regular strings:
#
#
blast_hsp.query
blast_hsp.hit
#
# They are `SeqRecord` objects you saw earlier in
# Section \[chapter:SeqRecord\]! This means that you can do all sorts of
# interesting things you can do with `SeqRecord` objects on `HSP.query`
# and/or `HSP.hit`.
#
# It should not surprise you now that the `HSP` object has an `alignment`
# property which is a `MultipleSeqAlignment` object:
#
#
print(blast_hsp.aln)
#
# Having probed the BLAST HSP, let’s now take a look at HSPs from our BLAT
# results for a different kind of HSP. As usual, we’ll begin by invoking
# `print` on it:
#
#
blat_qresult = SearchIO.read('data/my_blat.psl', 'blat-psl')
blat_hsp = blat_qresult[0][0] # first hit, first hsp
print(blat_hsp)
#
# Some of the outputs you may have already guessed. We have the query and
# hit IDs and descriptions and the sequence coordinates. Values for evalue
# and bitscore is ‘?’ as BLAT HSPs do not have these attributes. But The
# biggest difference here is that you don’t see any sequence alignments
# displayed. If you look closer, PSL formats themselves do not have any
# hit or query sequences, so `Bio.SearchIO` won’t create any sequence or
# alignment objects. What happens if you try to access `HSP.query`,
# `HSP.hit`, or `HSP.aln`? You’ll get the default values for these
# attributes, which is `None`:
#
#
blat_hsp.hit is None
blat_hsp.query is None
blat_hsp.aln is None
#
# This does not affect other attributes, though. For example, you can
# still access the length of the query or hit alignment. Despite not
# displaying any attributes, the PSL format still have this information so
# `Bio.SearchIO` can extract them:
#
#
blat_hsp.query_span # length of query match
blat_hsp.hit_span # length of hit match
#
# Other format-specific attributes are still present as well:
#
#
blat_hsp.score # PSL score
blat_hsp.mismatch_num # the mismatch column
#
# So far so good? Things get more interesting when you look at another
# ‘variant’ of HSP present in our BLAT results. You might recall that in
# BLAT searches, sometimes we get our results separated into ‘blocks’.
# These blocks are essentially alignment fragments that may have some
# intervening sequence between them.
#
# Let’s take a look at a BLAT HSP that contains multiple blocks to see how
# `Bio.SearchIO` deals with this:
#
#
blat_hsp2 = blat_qresult[0][1] # first hit, second hsp
print(blat_hsp2)
#
# What’s happening here? We still some essential details covered: the IDs
# and descriptions, the coordinates, and the quick statistics are similar
# to what you’ve seen before. But the fragments detail is all different.
# Instead of showing ‘Fragments: 1’, we now have a table with two data
# rows.
#
# This is how `Bio.SearchIO` deals with HSPs having multiple fragments. As
# mentioned before, an HSP alignment may be separated by intervening
# sequences into fragments. The intervening sequences are not part of the
# query-hit match, so they should not be considered part of query nor hit
# sequence. However, they do affect how we deal with sequence coordinates,
# so we can’t ignore them.
#
# Take a look at the hit coordinate of the HSP above. In the `Hit range:`
# field, we see that the coordinate is `[54233104:54264463]`. But looking
# at the table rows, we see that not the entire region spanned by this
# coordinate matches our query. Specifically, the intervening region spans
# from `54233122` to `54264420`.
#
# Why then, is the query coordinates seem to be contiguous, you ask? This
# is perfectly fine. In this case it means that the query match is
# contiguous (no intervening regions), while the hit match is not.
#
# All these attributes are accessible from the HSP directly, by the way:
#
#
blat_hsp2.hit_range # hit start and end coordinates of the entire HSP
blat_hsp2.hit_range_all # hit start and end coordinates of each fragment
blat_hsp2.hit_span # hit span of the entire HSP
blat_hsp2.hit_span_all # hit span of each fragment
blat_hsp2.hit_inter_ranges # start and end coordinates of intervening regions in the hit sequence
blat_hsp2.hit_inter_spans # span of intervening regions in the hit sequence
#
# Most of these attributes are not readily available from the PSL file we
# have, but `Bio.SearchIO` calculates them for you on the fly when you
# parse the PSL file. All it needs are the start and end coordinates of
# each fragment.
#
# What about the `query`, `hit`, and `aln` attributes? If the HSP has
# multiple fragments, you won’t be able to use these attributes as they
# only fetch single `SeqRecord` or `MultipleSeqAlignment` objects.
# However, you can use their `*_all` counterparts: `query_all`, `hit_all`,
# and `aln_all`. These properties will return a list containing
# `SeqRecord` or `MultipleSeqAlignment` objects from each of the HSP
# fragment. There are other attributes that behave similarly, i.e. they
# only work for HSPs with one fragment. Check out the `HSP`
# [documentation](http://biopython.org/DIST/docs/api/Bio.SearchIO._model.hsp-module.html)
# for a full list.
#
# Finally, to check whether you have multiple fragments or not, you can
# use the `is_fragmented` property like so:
#
#
blat_hsp2.is_fragmented # BLAT HSP with 2 fragments
blat_hsp.is_fragmented # BLAT HSP from earlier, with one fragment
#
# Before we move on, you should also know that we can use the slice
# notation on `HSP` objects, just like `QueryResult` or `Hit` objects.
# When you use this notation, you’ll get an `HSPFragment` object in
# return, the last component of the object model.
#
# ### HSPFragment {#sec:searchio-hspfragment}
#
# `HSPFragment` represents a single, contiguous match between the query
# and hit sequences. You could consider it the core of the object model
# and search result, since it is the presence of these fragments that
# determine whether your search have results or not.
#
# In most cases, you don’t have to deal with `HSPFragment` objects
# directly since not that many sequence search tools fragment their HSPs.
# When you do have to deal with them, what you should remember is that
# `HSPFragment` objects were written with to be as compact as possible. In
# most cases, they only contain attributes directly related to sequences:
# strands, reading frames, alphabets, coordinates, the sequences
# themselves, and their IDs and descriptions.
#
# These attributes are readily shown when you invoke `print` on an
# `HSPFragment`. Here’s an example, taken from our BLAST search:
#
#
from Bio import SearchIO
blast_qresult = SearchIO.read('data/my_blast.xml', 'blast-xml')
blast_frag = blast_qresult[0][0][0] # first hit, first hsp, first fragment
print(blast_frag)
#
# At this level, the BLAT fragment looks quite similar to the BLAST
# fragment, save for the query and hit sequences which are not present:
#
#
blat_qresult = SearchIO.read('data/my_blat.psl', 'blat-psl')
blat_frag = blat_qresult[0][0][0] # first hit, first hsp, first fragment
print(blat_frag)
#
# In all cases, these attributes are accessible using our favorite dot
# notation. Some examples:
#
#
blast_frag.query_start # query start coordinate
blast_frag.hit_strand # hit sequence strand
blast_frag.hit # hit sequence, as a SeqRecord object
#
# A note about standards and conventions {#sec:searchio-standards}
# --------------------------------------
#
# Before we move on to the main functions, there is something you ought to
# know about the standards `Bio.SearchIO` uses. If you’ve worked with
# multiple sequence search tools, you might have had to deal with the many
# different ways each program deals with things like sequence coordinates.
# It might not have been a pleasant experience as these search tools
# usually have their own standards. For example, one tools might use
# one-based coordinates, while the other uses zero-based coordinates. Or,
# one program might reverse the start and end coordinates if the strand is
# minus, while others don’t. In short, these often creates unnecessary
# mess must be dealt with.
#
# We realize this problem ourselves and we intend to address it in
# `Bio.SearchIO`. After all, one of the goals of `Bio.SearchIO` is to
# create a common, easy to use interface to deal with various search
# output files. This means creating standards that extend beyond the
# object model you just saw.
#
# Now, you might complain, “Not another standard!”. Well, eventually we
# have to choose one convention or the other, so this is necessary. Plus,
# we’re not creating something entirely new here; just adopting a standard
# we think is best for a Python programmer (it is Biopython, after all).
#
# There are three implicit standards that you can expect when working with
# `Bio.SearchIO`:
#
# - The first one pertains to sequence coordinates. In `Bio.SearchIO`,
# all sequence coordinates follows Python’s coordinate style:
# zero-based and half open. For example, if in a BLAST XML output file
# the start and end coordinates of an HSP are 10 and 28, they would
# become 9 and 28 in `Bio.SearchIO`. The start coordinate becomes 9
# because Python indices start from zero, while the end coordinate
# remains 28 as Python slices omit the last item in an interval.
#
# - The second is on sequence coordinate orders. In `Bio.SearchIO`,
# start coordinates are always less than or equal to end coordinates.
# This isn’t always the case with all sequence search tools, as some
# of them have larger start coordinates when the sequence strand
# is minus.
#
# - The last one is on strand and reading frame values. For strands,
# there are only four valid choices: `1` (plus strand), `-1` (minus
# strand), `0` (protein sequences), and `None` (no strand). For
# reading frames, the valid choices are integers from `-3` to `3` and
# `None`.
#
# Note that these standards only exist in `Bio.SearchIO` objects. If you
# write `Bio.SearchIO` objects into an output format, `Bio.SearchIO` will
# use the format’s standard for the output. It does not force its standard
# over to your output file.
#
# Reading search output files {#sec:searchio-input}
# ---------------------------
#
# There are two functions you can use for reading search output files into
# `Bio.SearchIO` objects: `read` and `parse`. They’re essentially similar
# to `read` and `parse` functions in other submodules like `Bio.SeqIO` or
# `Bio.AlignIO`. In both cases, you need to supply the search output file
# name and the file format name, both as Python strings. You can check the
# documentation for a list of format names `Bio.SearchIO` recognizes.
#
# `Bio.SearchIO.read` is used for reading search output files with only
# one query and returns a `QueryResult` object. You’ve seen `read` used in
# our previous examples. What you haven’t seen is that `read` may also
# accept additional keyword arguments, depending on the file format.
#
# Here are some examples. In the first one, we use `read` just like
# previously to read a BLAST tabular output file. In the second one, we
# use a keyword argument to modify so it parses the BLAST tabular variant
# with comments in it:
#
#
from Bio import SearchIO
qresult = SearchIO.read('data/tab_2226_tblastn_003.txt', 'blast-tab')
qresult
qresult2 = SearchIO.read('data/tab_2226_tblastn_007.txt', 'blast-tab', comments=True)
qresult2
#
# These keyword arguments differs among file formats. Check the format
# documentation to see if it has keyword arguments that modifies its
# parser’s behavior.
#
# As for the `Bio.SearchIO.parse`, it is used for reading search output
# files with any number of queries. The function returns a generator
# object that yields a `QueryResult` object in each iteration. Like
# `Bio.SearchIO.read`, it also accepts format-specific keyword arguments:
#
#
from Bio import SearchIO
qresults = SearchIO.parse('data/tab_2226_tblastn_001.txt', 'blast-tab')
for qresult in qresults:
print(qresult.id)
qresults2 = SearchIO.parse('data/tab_2226_tblastn_005.txt', 'blast-tab', comments=True)
for qresult in qresults2:
print(qresult.id)
#
# Dealing with large search output files with indexing {#sec:searchio-index}
# ----------------------------------------------------
#
# Sometimes, you’re handed a search output file containing hundreds or
# thousands of queries that you need to parse. You can of course use
# `Bio.SearchIO.parse` for this file, but that would be grossly
# inefficient if you need to access only a few of the queries. This is
# because `parse` will parse all queries it sees before it fetches your
# query of interest.
#
# In this case, the ideal choice would be to index the file using
# `Bio.SearchIO.index` or `Bio.SearchIO.index_db`. If the names sound
# familiar, it’s because you’ve seen them before in
# Section \[sec:SeqIO-index\]. These functions also behave similarly to
# their `Bio.SeqIO` counterparts, with the addition of format-specific
# keyword arguments.
#
# Here are some examples. You can use `index` with just the filename and
# format name:
#
#
from Bio import SearchIO
idx = SearchIO.index('data/tab_2226_tblastn_001.txt', 'blast-tab')
sorted(idx.keys())
idx['gi|16080617|ref|NP_391444.1|']
idx.close()
#
# Or also with the format-specific keyword argument:
#
#
idx = SearchIO.index('data/tab_2226_tblastn_005.txt', 'blast-tab', comments=True)
sorted(idx.keys())
idx['gi|16080617|ref|NP_391444.1|']
idx.close()
#
# Or with the `key_function` argument, as in `Bio.SeqIO`:
#
#
key_function = lambda id: id.upper() # capitalizes the keys
idx = SearchIO.index('data/tab_2226_tblastn_001.txt', 'blast-tab', key_function=key_function)
sorted(idx.keys())
idx['GI|16080617|REF|NP_391444.1|']
idx.close()
#
# `Bio.SearchIO.index_db` works like as `index`, only it writes the query
# offsets into an SQLite database file.
#
# Writing and converting search output files {#sec:searchio-write}
# ------------------------------------------
#
# It is occasionally useful to be able to manipulate search results from
# an output file and write it again to a new file. `Bio.SearchIO` provides
# a `write` function that lets you do exactly this. It takes as its
# arguments an iterable returning `QueryResult` objects, the output
# filename to write to, the format name to write to, and optionally some
# format-specific keyword arguments. It returns a four-item tuple, which
# denotes the number or `QueryResult`, `Hit`, `HSP`, and `HSPFragment`
# objects that were written.
#
#
from Bio import SearchIO
qresults = SearchIO.parse('data/mirna.xml', 'blast-xml') # read XML file
SearchIO.write(qresults, 'results.tab', 'blast-tab') # write to tabular file
#
# You should note different file formats require different attributes of
# the `QueryResult`, `Hit`, `HSP` and `HSPFragment` objects. If these
# attributes are not present, writing won’t work. In other words, you
# can’t always write to the output format that you want. For example, if
# you read a BLAST XML file, you wouldn’t be able to write the results to
# a PSL file as PSL files require attributes not calculated by BLAST (e.g.
# the number of repeat matches). You can always set these attributes
# manually, if you really want to write to PSL, though.
#
# Like `read`, `parse`, `index`, and `index_db`, `write` also accepts
# format-specific keyword arguments. Check out the documentation for a
# complete list of formats `Bio.SearchIO` can write to and their
# arguments.
#
# Finally, `Bio.SearchIO` also provides a `convert` function, which is
# simply a shortcut for `Bio.SearchIO.parse` and `Bio.SearchIO.write`.
# Using the convert function, our example above would be:
#
#
from Bio import SearchIO
SearchIO.convert('data/mirna.xml', 'blast-xml', 'results.tab', 'blast-tab')
| notebooks/08 - BLAST and other sequence search tools (experimental code).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""The Lotka-Volterra model where:
x is the number of preys
y is the number of predators
"""
#Credits:
#http://visual.icse.us.edu.pl/NPB/notebooks/Lotka_Volterra_with_SAGE.html
#as implemented in K3D_Animations/Lotka-Volterra.ipynb
#https://en.wikipedia.org/wiki/Lotka%E2%80%93Volterra_equations
import numpy as np
from scipy.integrate import odeint
def rhs(y0, t, a):
x, y = y0[0], y0[1]
return [x-x*y, a*(x*y-y)]
a_1 = 1.2
x0_1, x0_2, x0_3 = 2.0, 1.2, 1.0
y0_1, y0_2, y0_3 = 4.2, 3.7, 2.4
T = np.arange(0, 8, 0.02)
sol1 = odeint(rhs, [x0_1, y0_1], T, args=(a_1,))
sol2 = odeint(rhs, [x0_2, y0_2], T, args=(a_1,))
sol3 = odeint(rhs, [x0_3, y0_3], T, args=(a_1,))
limx = np.linspace(np.min(sol1[:,0]), np.max(sol1[:,0]), 20)
limy = np.linspace(np.min(sol1[:,1]), np.max(sol1[:,1]), 20)
vx, vy = np.meshgrid(limx, limy)
vx, vy = np.ravel(vx), np.ravel(vy)
vec = rhs([vx, vy], t=0.01, a=a_1)
origins = np.stack([np.zeros(np.shape(vx)), vx, vy]).T
vectors = np.stack([np.zeros(np.shape(vec[0])), vec[0], vec[1]]).T
vectors /= np.stack([np.linalg.norm(vectors, axis=1)]).T * 5
curve_points1 = np.vstack([np.zeros(sol1[:,0].shape), sol1[:,0], sol1[:,1]]).T
curve_points2 = np.vstack([np.zeros(sol2[:,0].shape), sol2[:,0], sol2[:,1]]).T
curve_points3 = np.vstack([np.zeros(sol3[:,0].shape), sol3[:,0], sol3[:,1]]).T
########################
from vedo import *
Arrows(origins, origins+vectors, c='r')
Line(curve_points1, c='o', lw=8)
Line(np.vstack([T, sol1[:,0], sol1[:,1]]).T, c='o')
Line(curve_points2, c='g', lw=8)
Line(np.vstack([T, sol2[:,0], sol2[:,1]]).T, c='g')
Line(curve_points3, c='b', lw=8)
Line(np.vstack([T, sol3[:,0], sol3[:,1]]).T, c='b')
show(..., viewup='x') # ... means all sofar created objects
# -
| examples/notebooks/advanced/volterra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preprocessing and Vocabulary Building
# ## Importing libraries
# <hr>
# +
import nltk
import random
import warnings
import numpy as np
import matplotlib.pyplot as plt # library for visualization
from nltk.corpus import twitter_samples # sample Twitter dataset from NLTK
# import our convenience functions
import utils
warnings.filterwarnings("ignore")
# -
# ## Load the NLTK sample dataset
# <hr>
nltk.download('twitter_samples')
# select the set of positive and negative tweets
all_positive_tweets = twitter_samples.strings('positive_tweets.json')
all_negative_tweets = twitter_samples.strings('negative_tweets.json')
# +
print('Number of positive tweets: ', len(all_positive_tweets))
print('Number of negative tweets: ', len(all_negative_tweets))
print('\nThe type of all_positive_tweets is: ', type(all_positive_tweets))
print('The type of a tweet entry is: ', type(all_negative_tweets[0]))
# +
# Declare a figure with a custom size
fig = plt.figure(figsize=(5, 5))
# labels for the two classes
labels = 'Positives', 'Negative'
# Sizes for each slide
sizes = [len(all_positive_tweets), len(all_negative_tweets)]
# Declare pie chart, where the slices will be ordered and plotted counter-clockwise:
plt.pie(sizes, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
# Equal aspect ratio ensures that pie is drawn as a circle.
plt.axis('equal')
# Display the chart
plt.show()
# -
# ## Looking at raw texts
# <hr>
# +
# print positive in greeen
print('\033[92m' + all_positive_tweets[random.randint(0,5000)])
# print negative in red
print('\033[91m' + all_negative_tweets[random.randint(0,5000)])
# -
# Our selected sample. Complex enough to exemplify each step
tweet = all_positive_tweets[2277]
print(tweet)
# ## Preprocess raw text for Sentiment analysis
# <hr>
# +
print('\033[92m')
print(tweet)
print('\033[94m')
# call the imported function
tweets_stem = utils.process_text(tweet); # Preprocess a given tweet
print('preprocessed tweet:')
print(tweets_stem) # Print the result
# -
| material/Preprocessing and Vocabulary Building.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Asymptotic Analysis
# ___
# It is a method to describe the limiting behaviour of algorithms when subjected to very large input datasets.
# * *Complexity class*: set of problems with related complexity
# * *Reduction*: transformation of one problem into another problem which is at least as difficult as the original problem (e.g. polynomial time reduction)
# * *Hard problem*: class of problems if the problem can be reduced to original problem
#
# <img align="center" src="./images/Complexity_chart_1.jpg" alt="complexity chart" width="800" height="800"/>
# Big O notation is used in Computer Science to describe the performance or complexity of an algorithm. Big O specifically describes the worst-case scenario, and can be used to describe the execution time required or the space used (e.g. in memory or on disk) by an algorithm.
# <br>
# **O(1)** has the lowest algorithmic complexity and **O(n!)** has the highest algorithmic complexity.<br>
# Note: For small input datasets, the algorithmic complexity or behaviour is different.
# <img align="center" src="./images/Complexity_chart_2.png" alt="complexity chart" width="400" height="400"/>
# ### O(1)
# O(1) describes an algorithm that will always execute in the same time (or space) regardless of the size of the input data set.
def IsFirstElementNull(elements):
return elements[0] == null
# In this case, we are just looking at elements[0] - only one element.
# ___
# ### O(N)
# O(N) describes an algorithm which grows linearly in proportion to the size of the input.
def ContainsValue(elements, value):
for element in elements:
if element == value:
return true
return false
# In this case, we are looking for matching a string value in elements. String match may occur at the first comparison itself but in the worst case, it will happen at the last part of the loop thereby iterating over whole loop.
# ___
# ### O(N<sup>2</sup>)
# O(N<sup>2</sup>) describes an algorithm which grows proportionally to the square of the size of the input dataset. Algorithms with nested iterations may result in O(N<sup>3</sup>), O(N<sup>4</sup>) etc.
def ContainsDuplicates(elements):
outer = 0
for outer in range(len(elements)):
inner = 0
for inner in range(len(elements)):
# Don't compare with self
if outer == inner:
continue
if elements[outer] == elements[inner]:
return true
return false
# ___
# ### O(2<sup>N</sup>)
# O(2<sup>N</sup>) describes an algorithm which grows exponentially in proportion to the size of the input dataset. With every addtion to the input, complexity doubles. The curve is like a plateau in the starting but rises at a rapid rate.
def Fibonacci(number):
if number <= 1:
return number
return Fibonacci(number - 2) + Fibonacci(number - 1)
# In this Fibonacci example, calculation of Fibonacci numbers via recursion makes the algorithm complexity grow exponentially.
# ___
# ### Logarithms (Log N)
# Binary search is searching algorithms which operates on sorted datasets. It halves the dataset in each iteration thereby improving at each step. It produces a growth curve that peaks at the beginning and slowly flattens out as the size of the data sets increases. E.g. an input data set containing 10 items takes one second to complete, a data set containing 100 items takes two seconds, and a data set containing 1000 items will take three seconds.
# <br><br>Doubling the size of the input data set has little effect on its growth as after a single iteration of the algorithm the data set will be halved and therefore on a par with an input data set half the size. This makes algorithms like binary search extremely efficient when dealing with large data sets.
# <img align="center" src="./images/data_structure_complexity.jpg" alt="complexity chart" width="800" height="800"/>
# <img align="center" src="./images/Complexity_chart_3.jpg" alt="complexity chart" width="600" height="600"/>
# ___
# ## Class of problems
# ### P
# It is the complexity class of decision problems that can be solved on a deterministic Turing machine in polynomial time(in the worst case). If we can turn a problem into decision problem, the result would belong to P.
# ### NP
# The complexity class of decision problems that can be solved on a non-deterministic Turing machine in polynomial time(in the worst case). <br>
# All *yes* instances of NP problems can be solved in polynomial time with non-deterministic Turing machine. *Complete* problems are those who can be reduced to NP.
# <br>
# ### NP Hard
# The set of problems which are at least as hard as NP but need not be itself in NP. e.g. Travelling salesman problem - finding shortest route in a graph.
# ___
# ## Recursion
# There are three laws of recursion:
# 1. Every recursion problem must have a base case.
# 2. Every recursion problem must proceed towards base case.
# 3. Every recursion problem must call itself, recursively.
#
# For every recursive call, the recursive function has to allocate memory on the stack for arguments, return address, and local variables, costing time to push and pop these data onto the stack. Recursive algorithms take at least O(n) space where n is the depth of the recursive call.
# Note: Recursive approach to problems are costly when sub problems are overlapping with each other. An iterative approach will run in O(n) time while recursive approach will run in exponential time.
# #### Recursive relations
# For describing the running time of recursive functions, we use recursive relations:<br>
# $$ T (n) = a · T (g(n)) + f(n) $$
# * $a$:- number of recursive calls
# * $g(n)$:- size of each sub-problem to be solved recursively
# * $f(n)$:- extra work done by the function
# | Equation | Complexity | Function |
# | :-----------------------: | :----------: | :---------------------: |
# | <img width=150/> | <img width=150/> | <img width=150/> |
# | $T(n) = T(n − 1) + 1$ | O(n) | Processing a sequence |
# | $T(n) = T(n − 1) + n$ | O(n<sup>2</sup>)| Handshake problem |
# | $T(n) = 2T(n − 1) + 1$| O(2<sup>n</sup>)| Towers of Hanoi |
# | $T(n) = T(n=2) + 1$| O(ln n)| Binary search |
# | $T(n) = T(n=2) + n$| O(n)| Randomized select |
# | $T(n) = 2T(n=2) + 1$| O(n)| Tree transversal |
# | $T(n) = 2T(n=2) + n$| O(n\*ln n)| Sort by divide and conquer |
# ___
# #### Divide and Conquer algorithms
# Recurrences for divide and conquer have the following form:
# $$T(n)=a.T(n/b)+f(n)$$
# * $a$:- number of recursive calls
# * $1/b$:- percentage of the dataset
# * $f(n)$:- work done by function
# Height of the binary search tree (balanced) is h = ln<sub>b</sub> n
# <img align="center" src="./images/binary_search_tree.png" alt="binary search tree" width="600" height="600"/>
# ___
# ### Runtime in functions
# If the algorithm doesn't involve recursive calling, the run time of algorithm can be calculated using the complexity of data structure and flow blocks.<br>
# Note: A function containing recursive calling for finding nth term of fibonacci series will take exponential time.
def find_fibonacci_seq_rec(n):
if n < 2:
return n
return find_fibonacci_seq_rec(n - 1) + find_fibonacci_seq_rec(n - 2)
# <p style="text-align:center;"><b>Algorithmic complexity calculation for different algorithms</b></p>
# | Big O notation | Complexity | Algorithm |
# |:---:|:---:|:---:|
# | O(n<sup>2</sup>) | quadratic | insertion, selection sort |
# | O(n ln n) | log linear | algorithms breaking problem into smaller chunks per invocation and then sticking together results, merge sort, quick sort |
# | O(n) | linear | iteration over a list |
# | O(ln n) | log | algorithm breaking problem into smaller chunks per invocation, searching a binary search tree |
# | O(1) | constant | hash table lookup or modification |
# | O(n<sup>k</sup>) | polynomial | k-nested loops over n |
# | O(k<sup>n</sup>) | exponential | producing every subset of n items |
# | O(n!) | factorial | producing every ordering of n values |
# $$ --The End-- $$
| python_learnings/9. Asymptotic Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1. Load Libraries
import os
os.environ["CUDA_VISIBLE_DEVICES"]="1"
# +
PATH = '/mnt/project/grp_202/khan74/i3d_keras/tmp'
import sys
sys.path.append(PATH+'/../')
sys.path.append(PATH+'/../src/')
# -
PATH = '/mnt/project/grp_202/khan74/i3d_keras/'
# +
from __future__ import print_function
from __future__ import absolute_import
import warnings
import numpy as np
from keras.models import Model
from keras import layers
from keras.layers import Activation
from keras.layers import Dense
from keras.layers import Input
from keras.layers import BatchNormalization
from keras.layers import Conv3D
from keras.layers import MaxPooling3D
from keras.layers import AveragePooling3D
from keras.layers import Dropout
from keras.layers import Reshape
from keras.layers import Lambda
from keras.layers import GlobalAveragePooling3D
from keras.engine.topology import get_source_inputs
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras import backend as K
from keras import optimizers
import keras
# -
import matplotlib.pyplot as plt
import seaborn as sn
from ipywidgets import *
import cv2
from i3d_inception import Inception_Inflated3d
# +
# Animate
#@title Import the necessary modules
# TensorFlow and TF-Hub modules.
from absl import logging
#import tensorflow as tf
#import tensorflow_hub as hub
logging.set_verbosity(logging.ERROR)
# Some modules to help with reading the UCF101 dataset.
import random
import re
import os
import tempfile
import cv2
import numpy as np
# Some modules to display an animation using imageio.
import imageio
from IPython import display
from urllib import request # requires python3
# +
# Utilities to open video files using CV2
def crop_center_square(frame):
y, x = frame.shape[0:2]
min_dim = min(y, x)
start_x = (x // 2) - (min_dim // 2)
start_y = (y // 2) - (min_dim // 2)
return frame[start_y:start_y+min_dim,start_x:start_x+min_dim]
def load_video(path, max_frames=0, resize=(224, 224)):
cap = cv2.VideoCapture(path)
frames = []
try:
while True:
ret, frame = cap.read()
if not ret:
break
frame = crop_center_square(frame)
frame = cv2.resize(frame, resize)
frame = frame[:, :, [2, 1, 0]]
frames.append(frame)
if len(frames) == max_frames:
break
finally:
cap.release()
return np.array(frames) / 255.0
def animate(images):
converted_images = np.clip(images * 255, 0, 255).astype(np.uint8)
imageio.mimsave('./animation.gif', converted_images, fps=25)
with open('./animation.gif','rb') as f:
display.display(display.Image(data=f.read(), height=300))
# -
def show_rgb(video_path_npy):
rgb = np.load(video_path_npy)
rgb = (rgb + 1) / 2
#rgb = np.squeeze(rgb, axis=0) # remove the batch dimension
nb_frames = rgb.shape[0]
for i in range(0, nb_frames):
rgb_1st_img = rgb[i, :, :, :]
cv2.imshow('img_flow', rgb_1st_img)
cv2.waitKey(0)
def conv3d_bn(x,
filters,
num_frames,
num_row,
num_col,
padding='same',
strides=(1, 1, 1),
use_bias = False,
use_activation_fn = True,
use_bn = True,
name=None):
"""Utility function to apply conv3d + BN.
# Arguments
x: input tensor.
filters: filters in `Conv3D`.
num_frames: frames (time depth) of the convolution kernel.
num_row: height of the convolution kernel.
num_col: width of the convolution kernel.
padding: padding mode in `Conv3D`.
strides: strides in `Conv3D`.
use_bias: use bias or not
use_activation_fn: use an activation function or not.
use_bn: use batch normalization or not.
name: name of the ops; will become `name + '_conv'`
for the convolution and `name + '_bn'` for the
batch norm layer.
# Returns
Output tensor after applying `Conv3D` and `BatchNormalization`.
"""
if name is not None:
bn_name = name + '_bn'
conv_name = name + '_conv'
else:
bn_name = None
conv_name = None
x = Conv3D(
filters, (num_frames, num_row, num_col),
strides=strides,
padding=padding,
use_bias=use_bias,
name=conv_name)(x)
if use_bn:
if K.image_data_format() == 'channels_first':
bn_axis = 1
else:
bn_axis = 4
x = BatchNormalization(axis=bn_axis, scale=False, name=bn_name)(x)
if use_activation_fn:
x = Activation('relu', name=name)(x)
return x
# ### 2. Load Model
# +
FRAME_HEIGHT = 224
FRAME_WIDTH = 224
NUM_RGB_CHANNELS = 3
NUM_FLOW_CHANNELS = 2
NUM_CLASSES = 2
#LABEL_MAP_PATH = PATH+'data/label_map.txt'
# -
rgb_sample = np.load(PATH + '/data/fall_no_fall/HS_data/processed_dataset_win20_thre18/fall/10229.npy')
rgb_sample = np.expand_dims(rgb_sample, axis=0)
INPUT_SHAPE = rgb_sample.shape[1]; INPUT_SHAPE
rgb_model = Inception_Inflated3d(
include_top=False,
weights='rgb_imagenet_and_kinetics',
input_shape=(INPUT_SHAPE, FRAME_HEIGHT, FRAME_WIDTH, NUM_RGB_CHANNELS),
classes=NUM_CLASSES)
# +
x = rgb_model.output
#x = AveragePooling3D((2, 7, 7), strides=(1, 1, 1), padding='valid', name='global_avg_pool')(x)
x = Dropout(0.5)(x)
x = conv3d_bn(x, NUM_CLASSES, 1, 1, 1, padding='same',
use_bias=True, use_activation_fn=False, use_bn=False, name='Conv3d_6a_1x1')
num_frames_remaining = int(x.shape[1])
x = Reshape((num_frames_remaining, NUM_CLASSES))(x)
# logits (raw scores for each class)
x = Lambda(lambda x: K.mean(x, axis=1, keepdims=False),
output_shape=lambda s: (s[0], s[2]))(x)
x = Activation('softmax', name='prediction')(x)
model = Model(rgb_model.input, x, name='i3d_inception')
# -
model.compile(loss = "categorical_crossentropy", optimizer = optimizers.Adam(lr=0.0001), metrics=["accuracy"])
model.summary()
# ### Load Data + Train
# +
# data_path = '/home/khan74/project/i3d_keras/data/fall_no_fall/HS_data/processed_dataset_win20_thre15/'
# data_path = '/home/khan74/project/i3d_keras/data/fall_no_fall/HS_data/processed_dataset_win20_thre18_2/'
data_path = '/home/khan74/project/i3d_keras/data/fall_no_fall/HS_data/processed_dataset_win20_thre18_3/'
# +
import glob
train_falls = []
for file in glob.glob(data_path+"train/fall/*.npy"):
train_falls.append(file)
train_no_falls = []
for file in glob.glob(data_path+"train/nofall/*.npy"):
train_no_falls.append(file)
test_falls = []
for file in glob.glob(data_path+"test/fall/*.npy"):
test_falls.append(file)
test_no_falls = []
for file in glob.glob(data_path+"test/nofall/*.npy"):
test_no_falls.append(file)
# -
print(len(train_falls)); print(len(train_no_falls)); print(len(test_falls)); print(len(test_no_falls))
# +
x_train, y_train = [], []
for f in train_falls:
rgb_sample = np.load(f)
x_train.append(rgb_sample)
y_train.append(1)
for f in train_no_falls:
rgb_sample = np.load(f)
x_train.append(rgb_sample)
y_train.append(0)
# +
x_test, y_test = [], []
for f in test_falls:
rgb_sample = np.load(f)
x_test.append(rgb_sample)
y_test.append(1)
for f in test_no_falls:
rgb_sample = np.load(f)
x_test.append(rgb_sample)
y_test.append(0)
# +
x_train = np.array(x_train); x_train = x_train/255
y_train = np.array(y_train); y_train = keras.utils.to_categorical(y_train, 2)
x_test = np.array(x_test); x_test = x_test/255
y_test = np.array(y_test); y_test = keras.utils.to_categorical(y_test, 2)
# -
hist = model.fit(x=x_train, y=y_train, batch_size=16, epochs=10, shuffle=True, validation_data=(x_test, y_test))#validation_split=0.2)
model.save_weights('train_2.h5')
# +
#Training Accuracy/Loss
plt.figure( figsize=(11,4) )
# Plot training & validation accuracy values
plt.subplot(1,2,1)
plt.plot(hist.history['acc'])
plt.plot(hist.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
#plt.show()
# Plot training & validation loss values
plt.subplot(1,2,2)
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# -
# ### Predict
# + [markdown] heading_collapsed=true
# ###### cf matrix
# + hidden=true
import itertools
from sklearn import metrics
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_recall_curve
from inspect import signature
# + hidden=true
def plot_prec_recall_curve(Predictions, Actuals):
precision, recall, _ = precision_recall_curve(Actuals, Predictions)
average_precision = average_precision_score(Actuals, Predictions)
# In matplotlib < 1.5, plt.fill_between does not have a 'step' argument
step_kwargs = ({'step': 'post'}
if 'step' in signature(plt.fill_between).parameters
else {})
plt.step(recall, precision, color='b', alpha=0.2,
where='post')
plt.fill_between(recall, precision, alpha=0.2, color='b', **step_kwargs)
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title('2-class Precision-Recall curve: AP={0:0.2f}'.format(
average_precision))
# + hidden=true
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues,
figure_size = (11, 6),
save=0,
save_path='/home/khan74'):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
#if normalize:
#cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
#print("Normalized confusion matrix")
#else:
#print('Confusion matrix, without normalization')
#print(cm)
plt.figure(figsize=figure_size)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
#plt.yticks(tick_marks, classes, rotation=45)
plt.yticks([-.5, 0.00, 1.00, 1.5], ['', 'No Fall', 'Fall', ''])
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
if save:
plt.savefig(save_path)
#plt.savefig('confusion_matrix.png')
# + hidden=true
Actuals, Predictions = [], []
# + hidden=true
for f in test_no_falls:
test_sample = np.load(f)
test_sample = np.expand_dims(test_sample, axis=0)
test_sample= test_sample/255
p = model.predict(test_sample)
v = np.argmax(p)
#print(p)
if v == 1:
print('Fall')
Actuals.append(0)
Predictions.append(v)
elif v == 0:
print('No Fall')
Actuals.append(0)
Predictions.append(v)
# + hidden=true
for f in test_falls:
test_sample = np.load(f)
test_sample = np.expand_dims(test_sample, axis=0)
test_sample= test_sample/255
p = model.predict(test_sample)
v = np.argmax(p)
#print(p)
if v == 1:
print('Fall')
Actuals.append(1)
Predictions.append(v)
elif v == 0:
print('No Fall')
Actuals.append(1)
Predictions.append(v)
# + hidden=true
Predictions = np.array(Predictions); Actuals = np.array(Actuals)
# -
# ###### Results
cm = confusion_matrix(Actuals, Predictions)
plot_confusion_matrix(cm, classes=['No Fall', 'Fall'], title="Confusion Matrix", figure_size=(8,5), save=0)
print('Test Accuracy: {:.2f}'.format(np.sum(Predictions==Actuals)/len(Actuals)))
print('Test Precision: {:.2f}'.format(precision_score(Actuals, Predictions)))
print('Test Recall: {:.2f}'.format(recall_score(Actuals, Predictions)))
plot_prec_recall_curve(Predictions, Actuals)
| i3d/nbs/Train_fall-Non_Overlap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import the dependencies#
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import csv
#import our CSV data files and set the paths#
ride_csv= "raw_data/ride_data.csv"
city_csv= "raw_data/city_data.csv"
#read csvs into df for reference#
ride_df = pd.read_csv(ride_csv)
city_df = pd.read_csv(city_csv)
#merge dataframes, using an outer merge#
data = pd.merge(ride_df,city_df, on='city', how='outer')
# -
#Reset our index to the "City" column#
data = data.set_index('city')
data
# +
# Create the GroupBy object based on the "city column
fare_by_city = data.groupby(["city"])
# Calculate averages for fares from each city using the .mean() method
fare_by_city.mean()
# +
# ********** SECOND SECTION: BUBBLE CHARTS ************
# could not complete the set up of scatter/pie charts#
data_df = data
fare_by_city_df = fare_by_city.mean()
# -
plt.style.use('ggplot')
>>> df = pd.DataFrame(data_df, columns=['a', 'b', 'c'])
>>> df.plot(kind='scatter', x='a', y='b', s=df['c']*400);
# +
data.plot(kind="bar", facecolor="red")
# -
plt.show()
| Instructions/Pyber/Pyber.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Q0_wToNFHMN3"
# # **Decision Trees**
#
# The Wisconsin Breast Cancer Dataset(WBCD) can be found here(https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/breast-cancer-wisconsin.data)
#
# This dataset describes the characteristics of the cell nuclei of various patients with and without breast cancer. The task is to classify a decision tree to predict if a patient has a benign or a malignant tumour based on these features.
#
# Attribute Information:
# ```
# # Attribute Domain
# -- -----------------------------------------
# 1. Sample code number id number
# 2. Clump Thickness 1 - 10
# 3. Uniformity of Cell Size 1 - 10
# 4. Uniformity of Cell Shape 1 - 10
# 5. Marginal Adhesion 1 - 10
# 6. Single Epithelial Cell Size 1 - 10
# 7. Bare Nuclei 1 - 10
# 8. Bland Chromatin 1 - 10
# 9. Normal Nucleoli 1 - 10
# 10. Mitoses 1 - 10
# 11. Class: (2 for benign, 4 for malignant)
# ```
# + id="qYdlWpUVHMOB"
import pandas as pd
headers = ["ID","CT","UCSize","UCShape","MA","SECSize","BN","BC","NN","Mitoses","Diagnosis"]
data = pd.read_csv('./data/breast-cancer-wisconsin.data', na_values='?',
header=None, index_col=['ID'], names = headers)
data = data.reset_index(drop=True)
data = data.fillna(0)
data.describe()
# + [markdown] id="x_gQq5qrHMOG"
# 1. a) Implement a decision tree (you can use decision tree implementation from existing libraries).
# + id="g6R3GmzBHMOH"
# + [markdown] id="VZ7N9m_mHMOJ"
# 1. b) Train a decision tree object of the above class on the WBC dataset using misclassification rate, entropy and Gini as the splitting metrics.
# + id="eHFij6PaHMOJ"
# + [markdown] id="eXEjInvmHMOK"
# 1. c) Report the accuracies in each of the above splitting metrics and give the best result.
# + id="49QZvmgNHMOL"
# + [markdown] id="Bz_7nYxPHMON"
# 1. d) Experiment with different approaches to decide when to terminate the tree (number of layers, purity measure, etc). Report and give explanations for all approaches.
# + id="yLRI0niJHMOP"
# + [markdown] id="uWAN_wWXHMOQ"
# 2. What is boosting, bagging and stacking?
# Which class does random forests belong to and why?
# + [markdown] id="LnO5uqHlHMOR"
# Answer:
# + [markdown] id="pihvGbqLHMOS"
# 3. Implement random forest algorithm using different decision trees .
# + id="dXdPP2aIHMOT"
# + [markdown] id="zJOn5nNZHMOU"
# 4. Report the accuracies obtained after using the Random forest algorithm and compare it with the best accuracies obtained with the decision trees.
# + id="Ce4KiiIGHMOV"
# + [markdown] id="yj-vNvsYHMOX"
# 5. Submit your solution as a separate pdf in the final zip file of your submission
#
#
# Compute a decision tree with the goal to predict the food review based on its smell, taste and portion size.
#
# (a) Compute the entropy of each rule in the first stage.
#
# (b) Show the final decision tree. Clearly draw it.
#
# Submit a handwritten response. Clearly show all the steps.
#
#
| Assignment3/.ipynb_checkpoints/DecisionTreesAndRandomForests-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
from transformers import MT5ForConditionalGeneration, MT5Config, MT5EncoderModel, MT5Tokenizer, Trainer, TrainingArguments
from progeny_tokenizer import TAPETokenizer
import numpy as np
import math
import random
import scipy
import time
import pandas as pd
from torch.utils.data import DataLoader, RandomSampler, Dataset, BatchSampler
import typing
from pathlib import Path
import argparse
from collections import OrderedDict
import pickle
import matplotlib.pyplot as plt
# -
# # Analyze generated sequences
tsv_name = 'generated_seqs/baseline_gen/basegen_seqs260000.tsv'
df = pd.read_table(tsv_name)
df
# ## filter out sequences without constant region
wt_seq = 'STIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNTNITEENVQNMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQ'
constant_region = 'NTNITEEN'
wt_cs_ind = wt_seq.index(constant_region)
indices_to_drop = []
dropped_seqs = []
for index, row in df.iterrows():
seq = row['MT_seq']
if constant_region not in seq:
indices_to_drop.append(index)
dropped_seqs.append(seq)
else:
cs_ind = seq.index(constant_region)
if cs_ind != wt_cs_ind:
indices_to_drop.append(index)
dropped_seqs.append(seq)
df = df.drop(indices_to_drop)
print(len(df))
# ## filter out sequences with invalid token
rejected_tokens = ["<pad>", "<sep>", "<cls>", "<mask>", "<unk>"]
indices_to_drop = []
dropped_seqs = []
for index, row in df.iterrows():
seq = row['MT_seq']
for rejected_token in rejected_tokens:
if rejected_token in seq:
indices_to_drop.append(index)
dropped_seqs.append(seq)
break
df = df.drop(indices_to_drop)
print(len(df))
df
# ## Analyze hamming distance
# Compute hamming distance between MT and WT
def hamming_dist(str1, str2):
i = 0
count = 0
while(i < len(str1)):
if(str1[i] != str2[i]):
count += 1
i += 1
return count
hamming_dist_list = []
wt_seq = df.iloc[0]['WT_seq']
wt_seq
for index, row in df.iterrows():
gen_seq = row['MT_seq']
h_dist = hamming_dist(gen_seq, wt_seq)
hamming_dist_list.append(h_dist)
hamming_dist_list
print("Hamming distance stats")
print("max: ", np.max(hamming_dist_list))
print("min: ", np.min(hamming_dist_list))
print("median: ", np.median(hamming_dist_list))
print("mean: ", np.mean(hamming_dist_list))
plt.figure(figsize=(8,6))
plt.hist(hamming_dist_list, label='generated', bins=[i for i in range(46)])
plt.xlabel("Hamming Distance", size=14)
plt.ylabel("Count", size=14)
plt.title("Hamming Distance from WT seq")
# plt.legend(loc='upper left')
# hamming distance for generator training data
gen_train_data = 'data/gen_train_data/top_half_ddG/train_ddG.pkl'
gen_train_df = pd.read_pickle(gen_train_data)
wt_seq = gen_train_df.iloc[0]['WT_seq']
wt_seq
gen_train_hamming_dist_list = []
for index, row in gen_train_df.iterrows():
train_seq = row['MT_seq']
h_dist = hamming_dist(train_seq, wt_seq)
gen_train_hamming_dist_list.append(h_dist)
plt.figure(figsize=(8,6))
plt.hist(gen_train_hamming_dist_list, label='train_data', bins=[i for i in range(46)])
plt.xlabel("Hamming Distance", size=14)
plt.ylabel("Count", size=14)
plt.title("Hamming Distance from WT seq")
# plt.legend(loc='upper left')
len(hamming_dist_list)
# +
plt.figure(figsize=(8,6))
plt.hist(hamming_dist_list, density=True, label='generated', bins=[i for i in range(46)], alpha=0.4)
# plt.xlabel("Hamming Distance", size=14)
# plt.ylabel("Count", size=14)
# plt.title("Hamming Distance from WT seq")
plt.hist(gen_train_hamming_dist_list, density=True, label='train_data', bins=[i for i in range(46)], alpha=0.4)
plt.xlabel("Hamming Distance", size=14)
plt.ylabel("Density", size=14)
plt.title("Top Half Generator")
plt.legend(loc='upper left')
# +
plt.figure(figsize=(8,6))
plt.hist(hamming_dist_list, density=True, label='generated', bins=[i for i in range(16)], alpha=0.4)
# plt.xlabel("Hamming Distance", size=14)
# plt.ylabel("Count", size=14)
# plt.title("Hamming Distance from WT seq")
plt.hist(gen_train_hamming_dist_list, density=True, label='train_data', bins=[i for i in range(16)], alpha=0.4)
plt.xlabel("Hamming Distance", size=14)
plt.ylabel("Density", size=14)
plt.title("Hamming Distance from WT seq")
plt.legend(loc='upper left')
# -
# # Analyze ddG values of gen 10K
from tape.metrics import spearmanr
results_tsv_name = "../../prot5_alvin/utils/foldx_sim_results/tophalf-basegen_top10K-Dscore_250Kgen/results_full.tsv"
ddG_df = pd.read_table(results_tsv_name)
ddG_df
disc_pred_list = ddG_df['disc_pred']
ddG_list = ddG_df['ddG']
disc_ddG_cor = spearmanr(disc_pred_list, ddG_list)
disc_ddG_cor
# Disc-predicted most stable ones first
ddG_df = ddG_df.sort_values(by='disc_pred', ascending=True)
ddG_df
top_half_stable_df = ddG_df[:len(ddG_df)//2]
top_half_stable_df
gen_train_ddG_list = gen_train_df['ddG']
len(ddG_list)
# +
plt.figure(figsize=(8,6))
plt.hist(ddG_list, density=True, label='generated', bins=[i for i in range(-10, 10)], alpha=0.4)
# plt.xlabel("Hamming Distance", size=14)
# plt.ylabel("Count", size=14)
# plt.title("Hamming Distance from WT seq")
plt.hist(gen_train_ddG_list, density=True, label='train_data', bins=[i for i in range(-10, 10)], alpha=0.4)
plt.xlabel("ddG", size=14)
plt.ylabel("Density", size=14)
plt.title("full 10k Gen")
plt.legend(loc='upper left')
# -
len(top_half_stable_df['ddG'])
# +
plt.figure(figsize=(8,6))
plt.hist(top_half_stable_df['ddG'], density=True, label='generated_top_half_pred', bins=[i for i in range(-10, 10)], alpha=0.4)
# plt.xlabel("Hamming Distance", size=14)
# plt.ylabel("Count", size=14)
# plt.title("Hamming Distance from WT seq")
plt.hist(gen_train_ddG_list, density=True, label='train_data', bins=[i for i in range(-10, 10)], alpha=0.4)
plt.xlabel("ddG", size=14)
plt.ylabel("Density", size=14)
plt.title("top half disc-scored Gen")
plt.legend(loc='upper left')
# -
np.max(top_half_stable_df['ddG'])
np.min(top_half_stable_df['ddG'])
np.mean(top_half_stable_df['ddG'])
np.median(top_half_stable_df['ddG'])
np.max(gen_train_ddG_list)
np.min(gen_train_ddG_list)
np.mean(gen_train_ddG_list)
np.median(gen_train_ddG_list)
# ## plot top 100, 1k and 10k distribution
# +
topk_hist_list = [10000, 1000, 100]
plt.figure(figsize=(8,6))
for topk in topk_hist_list:
topk_ddG_list = ddG_list[:topk]
topk_disc_pred_list = disc_pred_list[:topk]
topk_disc_ddG_cor = spearmanr(topk_disc_pred_list, topk_ddG_list)
print("topk: ", topk)
print("topk_disc_ddG_cor: ", topk_disc_ddG_cor)
plt.hist(topk_ddG_list, density=True, label='gen top {}'.format(topk), bins=[i for i in range(-10, 10)], alpha=0.2)
plt.hist(gen_train_ddG_list, density=True, label='train_data', bins=[i for i in range(-10, 10)], alpha=0.2)
plt.xlabel("ddG", size=14)
plt.ylabel("Density", size=14)
plt.title("full 10k Gen")
plt.legend(loc='upper left')
# +
topk_hist_list = [10000, 100]
plt.figure(figsize=(8,6))
for topk in topk_hist_list:
topk_ddG_list = ddG_list[:topk]
topk_disc_pred_list = disc_pred_list[:topk]
topk_disc_ddG_cor = spearmanr(topk_disc_pred_list, topk_ddG_list)
print("topk: ", topk)
print("topk_disc_ddG_cor: ", topk_disc_ddG_cor)
plt.hist(topk_ddG_list, density=True, label='gen top {}'.format(topk), bins=[i for i in range(-10, 10)], alpha=0.2)
plt.hist(gen_train_ddG_list, density=True, label='train_data', bins=[i for i in range(-10, 10)], alpha=0.2)
plt.xlabel("ddG", size=14)
plt.ylabel("Density", size=14)
plt.title("Top-Half Gen")
plt.legend(loc='upper left')
# -
# top 10, 100, 1000 by disc ranking
ddG_df
topK_list = [10, 100, 1000, 10000]
topK_df = ddG_df[:10]
topK_df
train_75pct = np.percentile(gen_train_ddG_list, 25)
# +
for topK in topK_list:
topK_df = ddG_df[:topK]
print("top K: ", len(topK_df))
print("max: ", np.max(topK_df['ddG']))
print("min: ", np.min(topK_df['ddG']))
print("mean: ", np.mean(topK_df['ddG']))
print("median: ", np.median(topK_df['ddG']))
PCI_75pct = np.sum(topK_df['ddG'] < train_75pct) / len(topK_df['ddG'])
print("PCI_75pct: ", PCI_75pct)
PCI_WT = np.sum(topK_df['ddG'] < 0) / len(topK_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
tophalf_df = ddG_df[:len(ddG_df)//2]
print("top half: ", len(tophalf_df))
print("max: ", np.max(tophalf_df['ddG']))
print("min: ", np.min(tophalf_df['ddG']))
print("mean: ", np.mean(tophalf_df['ddG']))
print("median: ", np.median(tophalf_df['ddG']))
PCI_75pct = np.sum(tophalf_df['ddG'] < train_75pct) / len(tophalf_df['ddG'])
print("PCI_75pct: ", PCI_75pct)
PCI_WT = np.sum(tophalf_df['ddG'] < 0) / len(tophalf_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
# training data distribution
print("train dataset: ", len(gen_train_ddG_list))
print("max: ", np.max(gen_train_ddG_list))
print("min: ", np.min(gen_train_ddG_list))
print("mean: ", np.mean(gen_train_ddG_list))
print("median: ", np.median(gen_train_ddG_list))
PCI_75pct = np.sum(gen_train_ddG_list < train_75pct) / len(gen_train_ddG_list)
print("PCI_75pct: ", PCI_75pct)
PCI_WT = np.sum(gen_train_ddG_list < 0) / len(gen_train_ddG_list)
print("PCI_WT: ", PCI_WT)
print("_"*20)
# -
topK_list = [10, 100, 1000, 10000]
percentile_list = [100, 95, 90, 85, 80, 75]
# +
for topK in topK_list:
topK_df = ddG_df[:topK]
print("top K: ", len(topK_df))
print("max: ", np.max(topK_df['ddG']))
print("min: ", np.min(topK_df['ddG']))
print("mean: ", np.mean(topK_df['ddG']))
print("median: ", np.median(topK_df['ddG']))
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(topK_df['ddG'] < pct) / len(topK_df['ddG'])
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(topK_df['ddG'] < 0) / len(topK_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
tophalf_df = ddG_df[:len(ddG_df)//2]
print("top half: ", len(tophalf_df))
print("max: ", np.max(tophalf_df['ddG']))
print("min: ", np.min(tophalf_df['ddG']))
print("mean: ", np.mean(tophalf_df['ddG']))
print("median: ", np.median(tophalf_df['ddG']))
# PCI_75pct = np.sum(tophalf_df['ddG'] < train_75pct) / len(tophalf_df['ddG'])
# print("PCI_75pct: ", PCI_75pct)
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(tophalf_df['ddG'] < pct) / len(tophalf_df['ddG'])
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(tophalf_df['ddG'] < 0) / len(tophalf_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
# training data distribution
print("train dataset: ", len(gen_train_ddG_list))
print("max: ", np.max(gen_train_ddG_list))
print("min: ", np.min(gen_train_ddG_list))
print("mean: ", np.mean(gen_train_ddG_list))
print("median: ", np.median(gen_train_ddG_list))
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(gen_train_ddG_list < pct) / len(gen_train_ddG_list)
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(gen_train_ddG_list < 0) / len(gen_train_ddG_list)
print("PCI_WT: ", PCI_WT)
print("_"*20)
# +
topK_list = [10, 100, 1000, 10000]
percentile_list = [100]
for topK in topK_list:
topK_df = ddG_df[:topK]
print("top K: ", len(topK_df))
print("min: ", np.min(topK_df['ddG']))
print("mean: ", np.mean(topK_df['ddG']))
print("median: ", np.median(topK_df['ddG']))
print("max: ", np.max(topK_df['ddG']))
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(topK_df['ddG'] < pct) / len(topK_df['ddG'])
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(topK_df['ddG'] < 0) / len(topK_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
tophalf_df = ddG_df[:len(ddG_df)//2]
print("top half: ", len(tophalf_df))
print("min: ", np.min(tophalf_df['ddG']))
print("mean: ", np.mean(tophalf_df['ddG']))
print("median: ", np.median(tophalf_df['ddG']))
print("max: ", np.max(tophalf_df['ddG']))
# PCI_75pct = np.sum(tophalf_df['ddG'] < train_75pct) / len(tophalf_df['ddG'])
# print("PCI_75pct: ", PCI_75pct)
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(tophalf_df['ddG'] < pct) / len(tophalf_df['ddG'])
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(tophalf_df['ddG'] < 0) / len(tophalf_df['ddG'])
print("PCI_WT: ", PCI_WT)
print("_"*20)
# training data distribution
print("train dataset: ", len(gen_train_ddG_list))
print("min: ", np.min(gen_train_ddG_list))
print("mean: ", np.mean(gen_train_ddG_list))
print("median: ", np.median(gen_train_ddG_list))
print("max: ", np.max(gen_train_ddG_list))
for percentile in percentile_list:
pct = np.percentile(gen_train_ddG_list, 100-percentile)
PCI_pct = np.sum(gen_train_ddG_list < pct) / len(gen_train_ddG_list)
print("PCI_{}pct: ".format(percentile), PCI_pct)
PCI_WT = np.sum(gen_train_ddG_list < 0) / len(gen_train_ddG_list)
print("PCI_WT: ", PCI_WT)
print("_"*20)
# -
| ACE2/Analyze foldx results tophalf-baselineGenDisc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Working with Order Book Data: NASDAQ ITCH
# The primary source of market data is the order book, which is continuously updated in real-time throughout the day to reflect all trading activity. Exchanges typically offer this data as a real-time service and may provide some historical data for free.
#
# The trading activity is reflected in numerous messages about trade orders sent by market participants. These messages typically conform to the electronic Financial Information eXchange (FIX) communications protocol for real-time exchange of securities transactions and market data or a native exchange protocol.
# ## Background
# ### The FIX Protocol
# Just like SWIFT is the message protocol for back-office (example, for trade-settlement) messaging, the [FIX protocol](https://www.fixtrading.org/standards/) is the de facto messaging standard for communication before and during, trade execution between exchanges, banks, brokers, clearing firms, and other market participants. Fidelity Investments and Salomon Brothers introduced FIX in 1992 to facilitate electronic communication between broker-dealers and institutional clients who by then exchanged information over the phone.
#
# It became popular in global equity markets before expanding into foreign exchange, fixed income and derivatives markets, and further into post-trade to support straight-through processing. Exchanges provide access to FIX messages as a real-time data feed that is parsed by algorithmic traders to track market activity and, for example, identify the footprint of market participants and anticipate their next move.
# ### Nasdaq TotalView-ITCH Order Book data
# While FIX has a dominant large market share, exchanges also offer native protocols. The Nasdaq offers a [TotalView ITCH direct data-feed protocol](http://www.nasdaqtrader.com/content/technicalsupport/specifications/dataproducts/NQTVITCHspecification.pdf) that allows subscribers to track
# individual orders for equity instruments from placement to execution or cancellation.
#
# As a result, it allows for the reconstruction of the order book that keeps track of the list of active-limit buy and sell orders for a specific security or financial instrument. The order book reveals the market depth throughout the day by listing the number of shares being bid or offered at each price point. It may also identify the market participant responsible for specific buy and sell orders unless it is placed anonymously. Market depth is a key indicator of liquidity and the potential price impact of sizable market orders.
# The ITCH v5.0 specification declares over 20 message types related to system events, stock characteristics, the placement and modification of limit orders, and trade execution. It also contains information about the net order imbalance before the open and closing cross.
# ## Imports
import warnings
warnings.filterwarnings('ignore')
# +
# %matplotlib inline
import gzip
import shutil
from struct import unpack
from collections import namedtuple, Counter, defaultdict
from pathlib import Path
from urllib.request import urlretrieve
from urllib.parse import urljoin
from datetime import timedelta
from time import time
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
import seaborn as sns
# + pycharm={"name": "#%%\n"}
sns.set_style('whitegrid')
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
def format_time(t):
"""Return a formatted time string 'HH:MM:SS
based on a numeric time() value"""
m, s = divmod(t, 60)
h, m = divmod(m, 60)
return f'{h:0>2.0f}:{m:0>2.0f}:{s:0>5.2f}'
# -
# ## Get NASDAQ ITCH Data from FTP Server
# The Nasdaq offers [samples](ftp://emi.nasdaq.com/ITCH/) of daily binary files for several months.
#
# We are now going to illustrates how to parse a sample file of ITCH messages and reconstruct both the executed trades and the order book for any given tick.
# The data is fairly large and running the entire example can take a lot of time and require substantial memory (16GB+). Also, the sample file used in this example may no longer be available because NASDAQ occasionaly updates the sample files.
# The following table shows the frequency of the most common message types for the sample file date March 29, 2018:
# | Name | Offset | Length | Value | Notes |
# |-------------------------|---------|---------|------------|--------------------------------------------------------------------------------------|
# | Message Type | 0 | 1 | S | System Event Message |
# | Stock Locate | 1 | 2 | Integer | Always 0 |
# | Tracking Number | 3 | 2 | Integer | Nasdaq internal tracking number |
# | Timestamp | 5 | 6 | Integer | Nanoseconds since midnight |
# | Order Reference Number | 11 | 8 | Integer | The unique reference number assigned to the new order at the time of receipt. |
# | Buy/Sell Indicator | 19 | 1 | Alpha | The type of order being added. B = Buy Order. S = Sell Order. |
# | Shares | 20 | 4 | Integer | The total number of shares associated with the order being added to the book. |
# | Stock | 24 | 8 | Alpha | Stock symbol, right padded with spaces |
# | Price | 32 | 4 | Price (4) | The display price of the new order. Refer to Data Types for field processing notes. |
# | Attribution | 36 | 4 | Alpha | Nasdaq Market participant identifier associated with the entered order |
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Set Data paths
# -
# We will store the download in a `data` subdirectory and convert the result to `hdf` format (discussed in the last section of chapter 2).
data_path = Path('data') # set to e.g. external harddrive
itch_store = str(data_path / 'itch.h5')
order_book_store = data_path / 'order_book.h5'
# You can find several sample files on the [NASDAQ ftp server](ftp://emi.nasdaq.com/ITCH/).
#
# The FTP address, filename and corresponding date used in this example:
FTP_URL = 'ftp://emi.nasdaq.com/ITCH/'
SOURCE_FILE = '10302019.NASDAQ_ITCH50.gz'
# ### Download & unzip
def may_be_download(url):
"""Download & unzip ITCH data if not yet available"""
if not data_path.exists():
print('Creating directory')
data_path.mkdir()
else:
print('Directory exists')
filename = data_path / url.split('/')[-1]
if not filename.exists():
print('Downloading...', url)
urlretrieve(url, filename)
else:
print('File exists')
unzipped = data_path / (filename.stem + '.bin')
if not unzipped.exists():
print('Unzipping to', unzipped)
with gzip.open(str(filename), 'rb') as f_in:
with open(unzipped, 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
else:
print('File already unpacked')
return unzipped
# This will download 5.1GB data that unzips to 12.9GB.
# + pycharm={"name": "#%%\n"}
file_name = may_be_download(urljoin(FTP_URL, SOURCE_FILE))
date = file_name.name.split('.')[0]
# -
# ## ITCH Format Settings
# ### The `struct` module for binary data
# The ITCH tick data comes in binary format. Python provides the `struct` module (see [docs])(https://docs.python.org/3/library/struct.html) to parse binary data using format strings that identify the message elements by indicating length and type of the various components of the byte string as laid out in the specification.
# From the docs:
#
# > This module performs conversions between Python values and C structs represented as Python bytes objects. This can be used in handling binary data stored in files or from network connections, among other sources. It uses Format Strings as compact descriptions of the layout of the C structs and the intended conversion to/from Python values.
# Let's walk through the critical steps to parse the trading messages and reconstruct the order book:
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Defining format strings
# + [markdown] pycharm={"name": "#%% md\n"}
# The parser uses format strings according to the following formats dictionaries:
# -
event_codes = {'O': 'Start of Messages',
'S': 'Start of System Hours',
'Q': 'Start of Market Hours',
'M': 'End of Market Hours',
'E': 'End of System Hours',
'C': 'End of Messages'}
# + pycharm={"name": "#%%\n"}
encoding = {'primary_market_maker': {'Y': 1, 'N': 0},
'printable' : {'Y': 1, 'N': 0},
'buy_sell_indicator' : {'B': 1, 'S': -1},
'cross_type' : {'O': 0, 'C': 1, 'H': 2},
'imbalance_direction' : {'B': 0, 'S': 1, 'N': 0, 'O': -1}}
# + pycharm={"name": "#%%\n"}
formats = {
('integer', 2): 'H', # int of length 2 => format string 'H'
('integer', 4): 'I',
('integer', 6): '6s', # int of length 6 => parse as string, convert later
('integer', 8): 'Q',
('alpha', 1) : 's',
('alpha', 2) : '2s',
('alpha', 4) : '4s',
('alpha', 8) : '8s',
('price_4', 4): 'I',
('price_8', 8): 'Q',
}
# -
# ### Create message specs for binary data parser
# The ITCH parser relies on message specifications that we create in the following steps.
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Load Message Types
# + [markdown] pycharm={"name": "#%% md\n"}
# The file `message_types.xlxs` contains the message type specs as laid out in the [documentation](https://www.nasdaqtrader.com/content/technicalsupport/specifications/dataproducts/NQTVITCHSpecification.pdf)
# + pycharm={"name": "#%%\n"}
message_data = (pd.read_excel('message_types.xlsx',
sheet_name='messages')
.sort_values('id')
.drop('id', axis=1))
# + pycharm={"name": "#%%\n"}
message_data.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Basic Cleaning
# + [markdown] pycharm={"name": "#%% md\n"}
# The function `clean_message_types()` just runs a few basic string cleaning steps.
# + pycharm={"name": "#%%\n"}
def clean_message_types(df):
df.columns = [c.lower().strip() for c in df.columns]
df.value = df.value.str.strip()
df.name = (df.name
.str.strip() # remove whitespace
.str.lower()
.str.replace(' ', '_')
.str.replace('-', '_')
.str.replace('/', '_'))
df.notes = df.notes.str.strip()
df['message_type'] = df.loc[df.name == 'message_type', 'value']
return df
# + pycharm={"name": "#%%\n"}
message_types = clean_message_types(message_data)
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Get Message Labels
# -
# We extract message type codes and names so we can later make the results more readable.
# + pycharm={"name": "#%%\n"}
message_labels = (message_types.loc[:, ['message_type', 'notes']]
.dropna()
.rename(columns={'notes': 'name'}))
message_labels.name = (message_labels.name
.str.lower()
.str.replace('message', '')
.str.replace('.', '')
.str.strip().str.replace(' ', '_'))
# message_labels.to_csv('message_labels.csv', index=False)
message_labels.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Finalize specification details
# + [markdown] pycharm={"name": "#%% md\n"}
# Each message consists of several fields that are defined by offset, length and type of value. The `struct` module will use this format information to parse the binary source data.
# + pycharm={"name": "#%%\n"}
message_types.message_type = message_types.message_type.ffill()
message_types = message_types[message_types.name != 'message_type']
message_types.value = (message_types.value
.str.lower()
.str.replace(' ', '_')
.str.replace('(', '')
.str.replace(')', ''))
message_types.info()
# -
message_types.head()
# + [markdown] pycharm={"name": "#%% md\n"}
# Optionally, persist/reload from file:
# + pycharm={"name": "#%%\n"}
message_types.to_csv('message_types.csv', index=False)
# -
message_types = pd.read_csv('message_types.csv')
# The parser translates the message specs into format strings and `namedtuples` that capture the message content. First, we create `(type, length)` formatting tuples from ITCH specs:
message_types.loc[:, 'formats'] = (message_types[['value', 'length']]
.apply(tuple, axis=1).map(formats))
# Then, we extract formatting details for alphanumerical fields
alpha_fields = message_types[message_types.value == 'alpha'].set_index('name')
alpha_msgs = alpha_fields.groupby('message_type')
alpha_formats = {k: v.to_dict() for k, v in alpha_msgs.formats}
alpha_length = {k: v.add(5).to_dict() for k, v in alpha_msgs.length}
# + [markdown] pycharm={"name": "#%% md\n"}
# We generate message classes as named tuples and format strings
# -
message_fields, fstring = {}, {}
for t, message in message_types.groupby('message_type'):
message_fields[t] = namedtuple(typename=t, field_names=message.name.tolist())
fstring[t] = '>' + ''.join(message.formats.tolist())
# + pycharm={"name": "#%%\n"}
alpha_fields.info()
# -
alpha_fields.head()
# Fields of `alpha` type (alphanumeric) require post-processing as defined in the `format_alpha` function:
# + pycharm={"name": "#%%\n"}
def format_alpha(mtype, data):
"""Process byte strings of type alpha"""
for col in alpha_formats.get(mtype).keys():
if mtype != 'R' and col == 'stock':
data = data.drop(col, axis=1)
continue
data.loc[:, col] = data.loc[:, col].str.decode("utf-8").str.strip()
if encoding.get(col):
data.loc[:, col] = data.loc[:, col].map(encoding.get(col))
return data
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Process Binary Message Data
# + [markdown] pycharm={"name": "#%% md\n"}
# The binary file for a single day contains over 350,000,000 messages worth over 12 GB.
# + pycharm={"name": "#%%\n"}
def store_messages(m):
"""Handle occasional storing of all messages"""
with pd.HDFStore(itch_store) as store:
for mtype, data in m.items():
# convert to DataFrame
data = pd.DataFrame(data)
# parse timestamp info
data.timestamp = data.timestamp.apply(int.from_bytes, byteorder='big')
data.timestamp = pd.to_timedelta(data.timestamp)
# apply alpha formatting
if mtype in alpha_formats.keys():
data = format_alpha(mtype, data)
s = alpha_length.get(mtype)
if s:
s = {c: s.get(c) for c in data.columns}
dc = ['stock_locate']
if m == 'R':
dc.append('stock')
try:
store.append(mtype,
data,
format='t',
min_itemsize=s,
data_columns=dc)
except Exception as e:
print(e)
print(mtype)
print(data.info())
print(pd.Series(list(m.keys())).value_counts())
data.to_csv('data.csv', index=False)
return 1
return 0
# + pycharm={"name": "#%%\n"}
messages = defaultdict(list)
message_count = 0
message_type_counter = Counter()
# + [markdown] pycharm={"name": "#%% md\n"}
# The script appends the parsed result iteratively to a file in the fast HDF5 format using the `store_messages()` function we just defined to avoid memory constraints (see last section in chapter 2 for more on this format).
# -
# The following code processes the binary file and produces the parsed orders stored by message type:
# + pycharm={"name": "#%%\n"}
start = time()
with file_name.open('rb') as data:
while True:
# determine message size in bytes
message_size = int.from_bytes(data.read(2), byteorder='big', signed=False)
# get message type by reading first byte
message_type = data.read(1).decode('ascii')
message_type_counter.update([message_type])
# read & store message
record = data.read(message_size - 1)
message = message_fields[message_type]._make(unpack(fstring[message_type], record))
messages[message_type].append(message)
# deal with system events
if message_type == 'S':
seconds = int.from_bytes(message.timestamp, byteorder='big') * 1e-9
print('\n', event_codes.get(message.event_code.decode('ascii'), 'Error'))
print(f'\t{format_time(seconds)}\t{message_count:12,.0f}')
if message.event_code.decode('ascii') == 'C':
store_messages(messages)
break
message_count += 1
if message_count % 2.5e7 == 0:
seconds = int.from_bytes(message.timestamp, byteorder='big') * 1e-9
d = format_time(time() - start)
print(f'\t{format_time(seconds)}\t{message_count:12,.0f}\t{d}')
res = store_messages(messages)
if res == 1:
print(pd.Series(dict(message_type_counter)).sort_values())
break
messages.clear()
print('Duration:', format_time(time() - start))
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Summarize Trading Day
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Trading Message Frequency
# + pycharm={"name": "#%%\n"}
counter = pd.Series(message_type_counter).to_frame('# Trades')
counter['Message Type'] = counter.index.map(message_labels.set_index('message_type').name.to_dict())
counter = counter[['Message Type', '# Trades']].sort_values('# Trades', ascending=False)
counter
# -
with pd.HDFStore(itch_store) as store:
store.put('summary', counter)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Top Equities by Traded Value
# + pycharm={"name": "#%%\n"}
with pd.HDFStore(itch_store) as store:
stocks = store['R'].loc[:, ['stock_locate', 'stock']]
trades = store['P'].append(store['Q'].rename(columns={'cross_price': 'price'}), sort=False).merge(stocks)
trades['value'] = trades.shares.mul(trades.price)
trades['value_share'] = trades.value.div(trades.value.sum())
trade_summary = trades.groupby('stock').value_share.sum().sort_values(ascending=False)
trade_summary.iloc[:50].plot.bar(figsize=(14, 6), color='darkblue', title='Share of Traded Value')
plt.gca().yaxis.set_major_formatter(FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
sns.despine()
plt.tight_layout()
# +
with pd.HDFStore(itch_store) as store:
stocks = store['R'].loc[:, ['stock_locate', 'stock']]
trades = store['P'].append(store['Q'].rename(columns={'cross_price': 'price'}), sort=False).merge(stocks)
trades['value'] = trades.shares.mul(trades.price)
trades['value_share'] = trades.value.div(trades.value.sum())
trade_summary = trades.groupby('stock').value_share.sum().sort_values(ascending=False)
trade_summary.iloc[:50].plot.bar(figsize=(14, 6), color='darkblue', title='Share of Traded Value')
plt.gca().yaxis.set_major_formatter(FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
sns.despine()
plt.tight_layout()
| 02_market_and_fundamental_data/01_NASDAQ_TotalView-ITCH_Order_Book/01_parse_itch_order_flow_messages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import nltk
import multiprocessing
import difflib
import time
import gc
import xgboost as xgb
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from collections import Counter
from sklearn.metrics import log_loss
from scipy.optimize import minimize
from sklearn.cross_validation import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from scipy.spatial.distance import cosine, correlation, canberra, chebyshev, minkowski, jaccard, euclidean
from xgb_utils import *
# +
def get_train():
feats_src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/'
keras_q1 = np.load(feats_src + 'train_q1_transformed.npy')
keras_q2 = np.load(feats_src + 'train_q2_transformed.npy')
feats_src2 = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/scripts/features/NER_features/'
keras_q1 = np.load(feats_src2 + 'q1train_NER_128len.npy')
keras_q2 = np.load(feats_src2 + 'q2train_NER_128len.npy')
xgb_feats = pd.read_csv(feats_src + '/the_1owl/owl_train.csv')
abhishek_feats = pd.read_csv(feats_src + 'abhishek/train_features.csv',
encoding = 'ISO-8859-1').iloc[:, 2:]
text_feats = pd.read_csv(feats_src + 'other_features/text_features_train.csv',
encoding = 'ISO-8859-1')
img_feats = pd.read_csv(feats_src + 'other_features/img_features_train.csv')
srk_feats = pd.read_csv(feats_src + 'srk/SRK_grams_features_train.csv')
mephisto_feats = pd.read_csv('../../data/features/spacylemmat_fullclean/train_mephistopeheles_features.csv').iloc[:, 6:]
#turkewitz_feats = pd.read_csv('../../data/features/lemmat_spacy_features/train_turkewitz_features.csv')
turkewitz_feats = pd.read_csv(feats_src + 'other_features/train_turkewitz_feats_orig.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
turkewitz_feats['freq_sum'] = turkewitz_feats.q1_freq + turkewitz_feats.q2_freq
turkewitz_feats['freq_diff'] = turkewitz_feats.q1_freq - turkewitz_feats.q2_freq
turkewitz_feats['freq_mult'] = turkewitz_feats.q1_freq * turkewitz_feats.q2_freq
turkewitz_feats['freq_div'] = turkewitz_feats.q1_freq / turkewitz_feats.q2_freq
xgb_feats.drop(['z_len1', 'z_len2', 'z_word_len1', 'z_word_len2'], axis = 1, inplace = True)
y_train = xgb_feats['is_duplicate']
xgb_feats = xgb_feats.iloc[:, 8:]
df = pd.concat([xgb_feats, abhishek_feats, text_feats, img_feats,
turkewitz_feats, mephisto_feats], axis = 1)
df = pd.DataFrame(df)
dfc = df.iloc[0:1000,:]
dfc = dfc.T.drop_duplicates().T
duplicate_cols = sorted(list(set(df.columns).difference(set(dfc.columns))))
print('Dropping duplicate columns:', duplicate_cols)
df.drop(duplicate_cols, axis = 1, inplace = True)
print('Final shape:', df.shape)
X = np.concatenate([keras_q1, keras_q2, df.values], axis = 1)
X = X.astype('float32')
print('Training data shape:', X.shape)
return X, y_train
def labelcount_encode(df2, cols):
df = df2.copy()
categorical_features = cols
new_df = pd.DataFrame()
for cat_feature in categorical_features:
cat_feature_value_counts = df[cat_feature].value_counts()
value_counts_list = cat_feature_value_counts.index.tolist()
value_counts_range_rev = list(reversed(range(len(cat_feature_value_counts)))) # for ascending ordering
value_counts_range = list(range(len(cat_feature_value_counts))) # for descending ordering
labelcount_dict = dict(zip(value_counts_list, value_counts_range))
new_df[cat_feature] = df[cat_feature].map(labelcount_dict)
return new_df
def count_encode(df2, cols):
df = df2.copy()
categorical_features = cols
new_df = pd.DataFrame()
for i in categorical_features:
new_df[i] = df[i].astype('object').replace(df[i].value_counts())
return new_df
def bin_numerical(df2, cols, step):
df = df2.copy()
numerical_features = cols
new_df = pd.DataFrame()
for i in numerical_features:
feature_range = np.arange(0, np.max(df[i]), step)
new_df[i] = np.digitize(df[i], feature_range, right=True)
return new_df
# +
def train_xgb(cv = False):
t = time.time()
params = {
'seed': 1337,
'colsample_bytree': 0.48,
'silent': 1,
'subsample': 0.74,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 12,
'min_child_weight': 20,
'nthread': 8,
'tree_method': 'hist',
#'updater': 'grow_gpu',
}
X_train, y_train = get_train()
if cv:
dtrain = xgb.DMatrix(X_train, y_train)
hist = xgb.cv(params, dtrain, num_boost_round = 100000, nfold = 5,
stratified = True, early_stopping_rounds = 350, verbose_eval = 250,
seed = 1337)
del X_train, y_train
gc.collect()
print('Time it took to train in CV manner:', time.time() - t)
return hist
else:
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, stratify = y_train,
test_size = 0.2, random_state = 111)
del X_train, y_train
gc.collect()
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
print('Start training...')
gbm = xgb.train(params, dtrain, 100000, watchlist,
early_stopping_rounds = 350, verbose_eval = 100)
print('Start predicting...')
val_pred = gbm.predict(xgb.DMatrix(X_val), ntree_limit=gbm.best_ntree_limit)
score = log_loss(y_val, val_pred)
print('Final score:', score, '\n', 'Time it took to train and predict:', time.time() - t)
del X_tr, X_val, y_tr, y_val
gc.collect()
return gbm
def run_xgb(model_name, train = True, test = False, cv = False):
if cv:
gbm_hist = train_xgb(True)
return gbm_hist
if train:
gbm = train_xgb()
gbm.save_model('saved_models/XGB/{}.txt'.format(model_name))
if test:
predict_test('{}'.format(model_name))
return gbm
# +
def get_transformations_features(transformations_src, mode = 'train'):
print('Adding features based on data transformations.')
lsa10tr_3grams_q1 = np.load(transformations_src + '{}_lsa10_3grams.npy'.format(mode))[0]
lsa10tr_3grams_q2 = np.load(transformations_src + '{}_lsa10_3grams.npy'.format(mode))[1]
transforms_feats = pd.DataFrame()
transforms_feats['cosine'] = [cosine(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['correlation'] = [correlation(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['jaccard'] = [jaccard(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['euclidean'] = [euclidean(x, y) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
transforms_feats['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(lsa10tr_3grams_q1, lsa10tr_3grams_q2)]
return transforms_feats
def get_doc2vec_features(doc2vec_src, mode = 'train'):
print('Adding features based on Doc2Vec distances.')
doc2vec_pre_q1 = np.load(doc2vec_src + '{}_q1_doc2vec_vectors_pretrained.npy'.format(mode))
doc2vec_pre_q2 = np.load(doc2vec_src + '{}_q2_doc2vec_vectors_pretrained.npy'.format(mode))
doc2vec_quora_q1 = np.load(doc2vec_src + '{}_q1_doc2vec_vectors_trainquora.npy'.format(mode))
doc2vec_quora_q2 = np.load(doc2vec_src + '{}_q2_doc2vec_vectors_trainquora.npy'.format(mode))
d2v_feats_pretrained = pd.DataFrame()
d2v_feats_pretrained['cosine'] = [cosine(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['correlation'] = [correlation(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['jaccard'] = [jaccard(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['euclidean'] = [euclidean(x, y) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_pretrained['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(doc2vec_pre_q1, doc2vec_pre_q2)]
d2v_feats_quora = pd.DataFrame()
d2v_feats_quora['cosine'] = [cosine(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['correlation'] = [correlation(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['jaccard'] = [jaccard(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['euclidean'] = [euclidean(x, y) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
d2v_feats_quora['minkowski'] = [minkowski(x, y, 3) for (x,y) in zip(doc2vec_quora_q1, doc2vec_quora_q2)]
return d2v_feats_pretrained, d2v_feats_quora
def labelcount_encode(df2, cols):
df = df2.copy()
categorical_features = cols
new_df = pd.DataFrame()
for cat_feature in categorical_features:
cat_feature_value_counts = df[cat_feature].value_counts()
value_counts_list = cat_feature_value_counts.index.tolist()
value_counts_range_rev = list(reversed(range(len(cat_feature_value_counts)))) # for ascending ordering
value_counts_range = list(range(len(cat_feature_value_counts))) # for descending ordering
labelcount_dict = dict(zip(value_counts_list, value_counts_range))
new_df[cat_feature] = df[cat_feature].map(labelcount_dict)
return new_df
def count_encode(df2, cols):
df = df2.copy()
categorical_features = cols
new_df = pd.DataFrame()
for i in categorical_features:
new_df[i] = df[i].astype('object').replace(df[i].value_counts())
return new_df
def bin_numerical(df2, cols, step):
df = df2.copy()
numerical_features = cols
new_df = pd.DataFrame()
for i in numerical_features:
feature_range = np.arange(0, np.max(df[i]), step)
new_df[i] = np.digitize(df[i], feature_range, right=True)
return new_df
def bin_numerical2(df2, cols, step):
df = df2.copy()
numerical_features = cols
for i in numerical_features:
feature_range = np.arange(0, np.max(df[i]), step)
df[i] = pd.cut(df[i], feature_range, right=True)
df[i] = pd.factorize(df[i], sort = True)[0]
return df
# +
src = '/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/scripts/features/'
train_orig = pd.read_csv(src + 'df_train_lemmatfullcleanSTEMMED.csv').iloc[:, :-1]
test_orig = pd.read_csv(src + 'df_test_lemmatfullcleanSTEMMED.csv').iloc[:, 4:]
full = pd.concat([train_orig, test_orig], ignore_index = True)
dflc = labelcount_encode(full, ['question1', 'question2'])
lc_cols = ['q1_lc', 'q2_lc']
dflc.columns = lc_cols
dflc_bin = bin_numerical(dflc, lc_cols, 5000)
dflc_bin.columns = ['q1_lc_bin', 'q2_lc_bin']
dflc['q1_lc'] = dflc['q1_lc'] / np.max(dflc['q1_lc'])
dflc['q2_lc'] = dflc['q2_lc'] / np.max(dflc['q2_lc'])
dflc_full = pd.concat([dflc, dflc_bin], axis = 1)
dflc_train = dflc_full.iloc[:train_orig.shape[0], :]
dflc_test = dflc_full.iloc[train_orig.shape[0]:, :]
dflc_test.to_csv('dflc_test.csv', index = False)
# -
dflc_train
# +
#turkewitz_feats = pd.read_csv('/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/uncleaned_data/other_features/train_turkewitz_feats_orig.csv')
turkewitz_feats = pd.read_csv('/media/w/1c392724-ecf3-4615-8f3c-79368ec36380/DS Projects/Kaggle/Quora/data/features/spacylemmat_fullclean/train_turkewitz_features.csv')
turkewitz_feats = turkewitz_feats[['q1_freq', 'q2_freq']]
tf_dflc = pd.concat([dflc_train, turkewitz_feats], axis = 1)
tf_dflc['q1gr1'] = tf_dflc.groupby(['q1_freq'])['q1_lc_bin'].transform('mean')
tf_dflc['q2gr2'] = tf_dflc.groupby(['q2_freq'])['q2_lc_bin'].transform('mean')
tf_dflc['q1gr1'] = tf_dflc['q1gr1'] / np.max(tf_dflc['q1gr1'])
tf_dflc['q2gr2'] = tf_dflc['q2gr2'] / np.max(tf_dflc['q2gr2'])
ff1 = turkewitz_feats.groupby(['q2_freq'])['q1_freq'].transform('sum')
ff2 = turkewitz_feats.groupby(['q1_freq'])['q2_freq'].transform('sum')
ff1 = ff1 / np.max(ff1)
ff2 = ff2 / np.max(ff2)
ff1m = turkewitz_feats.groupby(['q2_freq'])['q1_freq'].transform('mean')
ff2m = turkewitz_feats.groupby(['q1_freq'])['q2_freq'].transform('mean')
ff1m = ff1m / np.max(ff1m)
ff2m = ff2m / np.max(ff2m)
gr_feats = pd.DataFrame()
gr_feats['ff1'] = ff1
gr_feats['ff2'] = ff2
gr_feats['ff1m'] = ff1m
gr_feats['ff2m'] = ff2m
train_lc3 = labelcount_encode(turkewitz_feats, ['q1_freq', 'q2_freq'])
train_c = count_encode(turkewitz_feats, ['q1_freq', 'q2_freq'])
train_c.q1_freq = train_c.q1_freq / np.max(train_c.q1_freq)
train_c.q2_freq = train_c.q2_freq / np.max(train_c.q2_freq)
new_feats = np.concatenate([train_c, train_lc3, gr_feats, tf_dflc, dflc_train], axis = 1)
# +
X_train, y_train = get_train()
d2v_pre = np.load('train_doc2vec_pretrained_distances.npy')
d2v_quora = np.load('train_doc2vec_quoratrain_distances.npy')
transforms = np.load('train_transformations_distances.npy')
X_train = np.concatenate([X_train, d2v_pre, d2v_quora, transforms, new_feats], axis = 1)
X_train = X_train.astype('float32')
# +
params = {
'seed': 1337,
'colsample_bytree': 0.48,
'silent': 1,
'subsample': 0.74,
'eta': 0.05,
'objective': 'binary:logistic',
'eval_metric': 'logloss',
'max_depth': 12,
'min_child_weight': 20,
'nthread': 8,
'tree_method': 'hist',
}
t = time.time()
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, stratify = y_train,
test_size = 0.2, random_state = 111)
dtrain = xgb.DMatrix(X_tr, label = y_tr)
dval = xgb.DMatrix(X_val, label = y_val)
watchlist = [(dtrain, 'train'), (dval, 'valid')]
print('Start training...')
gbm = xgb.train(params, dtrain, 100000, watchlist,
early_stopping_rounds = 150, verbose_eval = 100)
print('Start predicting...')
val_pred = gbm.predict(xgb.DMatrix(X_val), ntree_limit=gbm.best_ntree_limit)
score = log_loss(y_val, val_pred)
print('Final score:', score, '\n', 'Time it took to train and predict:', time.time() - t)
gbm.save_model('saved_models/XGB/XGB_turkewitz_Doc2Vec2_LSA_GroupedFeats_experiments_sortedBIN.txt')
# -
| models/.ipynb_checkpoints/14.05 - XGB Magic Feats Experiments 0.27-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Input and Output
# %cd ~/Desktop/11072018/bioinformatics-tool-paper
import pandas as pd
infile = 's5-tool_similarity.dir/related_tools.txt'
# ### Code
# +
# Read dataframe
related_tool_dataframe = pd.read_table(infile).sort_values(['source_tool_name', 'similarity'], ascending=False)
# Initialize I
occurrence = 0
# Set source tool name
previous_source_tool_name = None
# Get top 2 similar tools
for index, rowData in related_tool_dataframe.head(50).iterrows():
# Set occurrence
if rowData['source_tool_name'] == previous_source_tool_name:
occurrence += 1
else:
occurrence = 0
# Set previous tool name
previous_source_tool_name = rowData['source_tool_name']
# Remove extra edges
if occurrence > 1:
related_tool_dataframe.drop(index, inplace=True)
# -
related_tool_dataframe
| old_code/pipeline/notebooks/Code Development.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %precision 2
# # Nonparametric Latent Dirichlet Allocation
#
# _Latent Dirichlet Allocation_ is a [generative](https://en.wikipedia.org/wiki/Generative_model) model for topic modeling. Given a collection of documents, an LDA inference algorithm attempts to determined (in an unsupervised manner) the topics discussed in the documents. It makes the assumption that each document is generated by a probability model, and, when doing inference, we try to find the parameters that best fit the model (as well as unseen/latent variables generated by the model). If you are unfamiliar with LDA, <NAME> has a [friendly introduction](http://blog.echen.me/2011/08/22/introduction-to-latent-dirichlet-allocation/) you should read.
#
#
# Because LDA is a _generative_ model, we can simulate the construction of documents by forward-sampling from the model. The generative algorithm is as follows (following [Heinrich](http://www.arbylon.net/publications/text-est.pdf)):
#
# * for each topic $k\in [1,K]$ do
# * sample term distribution for topic $\overrightarrow \phi_k \sim \text{Dir}(\overrightarrow \beta)$
# * for each document $m\in [1, M]$ do
# * sample topic distribution for document $\overrightarrow\theta_m\sim \text{Dir}(\overrightarrow\alpha)$
# * sample document length $N_m\sim\text{Pois}(\xi)$
# * for all words $n\in [1, N_m]$ in document $m$ do
# * sample topic index $z_{m,n}\sim\text{Mult}(\overrightarrow\theta_m)$
# * sample term for word $w_{m,n}\sim\text{Mult}(\overrightarrow\phi_{z_{m,n}})$
#
# You can implement this with [a little bit of code](https://gist.github.com/tdhopper/521006b60e1311d45509) and start to simulate documents.
#
# In LDA, we assume each word in the document is generated by a two-step process:
#
# 1. Sample a topic from the topic distribution for the document.
# 2. Sample a word from the term distribution from the topic.
#
# When we fit the LDA model to a given text corpus with an inference algorithm, our primary objective is to find the set of topic distributions $\underline \Theta$, term distributions $\underline \Phi$ that generated the documents, and latent topic indices $z_{m,n}$ for each word.
#
# To run the generative model, we need to specify each of these parameters:
vocabulary = ['see', 'spot', 'run']
num_terms = len(vocabulary)
num_topics = 2 # K
num_documents = 5 # M
mean_document_length = 5 # xi
term_dirichlet_parameter = 1 # beta
topic_dirichlet_parameter = 1 # alpha
# The term distribution vector $\underline\Phi$ is a collection of samples from a Dirichlet distribution. This describes how our 3 terms are distributed across each of the two topics.
from scipy.stats import dirichlet, poisson
from numpy import round
from collections import defaultdict
from random import choice as stl_choice
term_dirichlet_vector = num_terms * [term_dirichlet_parameter]
term_distributions = dirichlet(term_dirichlet_vector, 2).rvs(size=num_topics)
print(term_distributions)
# Each document corresponds to a categorical distribution across this distribution of topics (in this case, a 2-dimensional categorical distribution). This categorical distribution is a _distribution of distributions_; we could look at it as a Dirichlet process!
#
# The base base distribution of our Dirichlet process is a uniform distribution of topics (remember, topics are term distributions).
base_distribution = lambda: stl_choice(term_distributions)
# A sample from base_distribution is a distribution over terms
# Each of our two topics has equal probability
from collections import Counter
for topic, count in Counter([tuple(base_distribution()) for _ in range(10000)]).most_common():
print("count:", count, "topic:", [round(prob, 2) for prob in topic])
# Recall that a sample from a Dirichlet process is a distribution that approximates (but varies from) the base distribution. In this case, a sample from the Dirichlet process will be a distribution over topics that varies from the uniform distribution we provided as a base. If we use the stick-breaking metaphor, we are effectively breaking a stick one time and the size of each portion corresponds to the proportion of a topic in the document.
#
# To construct a sample from the DP, we need to [again define our DP class](/dirichlet-distribution/):
# +
from scipy.stats import beta
from numpy.random import choice
class DirichletProcessSample():
def __init__(self, base_measure, alpha):
self.base_measure = base_measure
self.alpha = alpha
self.cache = []
self.weights = []
self.total_stick_used = 0.
def __call__(self):
remaining = 1.0 - self.total_stick_used
i = DirichletProcessSample.roll_die(self.weights + [remaining])
if i is not None and i < len(self.weights) :
return self.cache[i]
else:
stick_piece = beta(1, self.alpha).rvs() * remaining
self.total_stick_used += stick_piece
self.weights.append(stick_piece)
new_value = self.base_measure()
self.cache.append(new_value)
return new_value
@staticmethod
def roll_die(weights):
if weights:
return choice(range(len(weights)), p=weights)
else:
return None
# -
# For each document, we will draw a topic distribution from the Dirichlet process:
topic_distribution = DirichletProcessSample(base_measure=base_distribution,
alpha=topic_dirichlet_parameter)
# A sample from this _topic_ distribution is a _distribution over terms_. However, unlike our base distribution which returns each term distribution with equal probability, the topics will be unevenly weighted.
for topic, count in Counter([tuple(topic_distribution()) for _ in range(10000)]).most_common():
print("count:", count, "topic:", [round(prob, 2) for prob in topic])
# To generate each word in the document, we draw a sample topic from the topic distribution, and then a term from the term distribution (topic).
# +
topic_index = defaultdict(list)
documents = defaultdict(list)
for doc in range(num_documents):
topic_distribution_rvs = DirichletProcessSample(base_measure=base_distribution,
alpha=topic_dirichlet_parameter)
document_length = poisson(mean_document_length).rvs()
for word in range(document_length):
topic_distribution = topic_distribution_rvs()
topic_index[doc].append(tuple(topic_distribution))
documents[doc].append(choice(vocabulary, p=topic_distribution))
# -
# Here are the documents we generated:
for doc in documents.values():
print(doc)
# We can see how each topic (term-distribution) is distributed across the documents:
for i, doc in enumerate(Counter(term_dist).most_common() for term_dist in topic_index.values()):
print("Doc:", i)
for topic, count in doc:
print(5*" ", "count:", count, "topic:", [round(prob, 2) for prob in topic])
# To recap: for each document we draw a _sample_ from a Dirichlet _Process_. The base distribution for the Dirichlet process is a categorical distribution over term distributions; we can think of the base distribution as an $n$-sided die where $n$ is the number of topics and each side of the die is a distribution over terms for that topic. By sampling from the Dirichlet process, we are effectively reweighting the sides of the die (changing the distribution of the topics).
#
# For each word in the document, we draw a _sample_ (a term distribution) from the distribution (over term distributions) _sampled_ from the Dirichlet process (with a distribution over term distributions as its base measure). Each term distribution uniquely identifies the topic for the word. We can sample from this term distribution to get the word.
#
# Given this formulation, we might ask if we can roll an _infinite_ sided die to draw from an unbounded number of topics (term distributions). We can do exactly this with a _Hierarchical_ Dirichlet process. Instead of the base distribution of our Dirichlet process being a _finite_ distribution over topics (term distributions) we will instead make it an infinite Distribution over topics (term distributions) by using yet another Dirichlet process! This base Dirichlet process will have as its base distribution a Dirichlet _distribution_ over terms.
#
# We will again draw a _sample_ from a Dirichlet _Process_ for each document. The base distribution for the Dirichlet process is itself a Dirichlet process whose base distribution is a Dirichlet distribution over terms. (Try saying that 5-times fast.) We can think of this as a countably infinite die each side of the die is a distribution over terms for that topic. The sample we draw is a topic (distribution over terms).
#
# For each word in the document, we will draw a _sample_ (a term distribution) from the distribution (over term distributions) _sampled_ from the Dirichlet process (with a distribution over term distributions as its base measure). Each term distribution uniquely identifies the topic for the word. We can sample from this term distribution to get the word.
#
# These last few paragraphs are confusing! Let's illustrate with code.
# +
term_dirichlet_vector = num_terms * [term_dirichlet_parameter]
base_distribution = lambda: dirichlet(term_dirichlet_vector).rvs(size=1)[0]
base_dp_parameter = 10
base_dp = DirichletProcessSample(base_distribution, alpha=base_dp_parameter)
# -
# This sample from the base Dirichlet process is our infinite sided die. It is a probability distribution over a countable infinite number of topics.
#
# The fact that our die is countably infinite is important. The sampler `base_distribution` draws topics (term-distributions) from an uncountable set. If we used this as the base distribution of the Dirichlet process below each document would be constructed from a _completely unique set of topics_. By feeding `base_distribution` into a Dirichlet Process (stochastic memoizer), we allow the topics to be shared across documents.
#
# In other words, `base_distribution` will never return the same topic twice; however, every topic sampled from `base_dp` would be sampled an infinite number of times (if we sampled from `base_dp` forever). At the same time, `base_dp` will also return an _infinite number_ of topics. In our formulation of the the LDA sampler above, our base distribution only ever returned a finite number of topics (`num_topics`); there is no `num_topics` parameter here.
#
# Given this setup, we can generate documents from the _hierarchical Dirichlet process_ with an algorithm that is essentially identical to that of the original _latent Dirichlet allocation_ generative sampler:
# +
nested_dp_parameter = 10
topic_index = defaultdict(list)
documents = defaultdict(list)
for doc in range(num_documents):
topic_distribution_rvs = DirichletProcessSample(base_measure=base_dp,
alpha=nested_dp_parameter)
document_length = poisson(mean_document_length).rvs()
for word in range(document_length):
topic_distribution = topic_distribution_rvs()
topic_index[doc].append(tuple(topic_distribution))
documents[doc].append(choice(vocabulary, p=topic_distribution))
# -
# Here are the documents we generated:
for doc in documents.values():
print(doc)
# And here are the latent topics used:
for i, doc in enumerate(Counter(term_dist).most_common() for term_dist in topic_index.values()):
print("Doc:", i)
for topic, count in doc:
print(5*" ", "count:", count, "topic:", [round(prob, 2) for prob in topic])
# Our documents were generated by an unspecified number of topics, and yet the topics were shared across the 5 documents. This is the power of the hierarchical Dirichlet process!
#
# This non-parametric formulation of Latent Dirichlet Allocation was first published by [<NAME> et al](http://www.cs.berkeley.edu/~jordan/papers/hdp.pdf).
#
# Unfortunately, forward sampling is the easy part. Fitting the model on data requires [complex MCMC](http://psiexp.ss.uci.edu/research/papers/sciencetopics.pdf) or [variational inference](http://www.cs.princeton.edu/~chongw/papers/WangPaisleyBlei2011.pdf). There are a [limited](http://www.stats.ox.ac.uk/~teh/software.html) [number](https://github.com/shuyo/iir/blob/master/lda/hdplda2.py) of [implementations](https://github.com/renaud/hdp-faster) [of HDP-LDA](http://www.arbylon.net/resources.html) available, and none of them are great.
| pages/2015-08-03-nonparametric-latent-dirichlet-allocation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
# C Crash Course
# ====
#
# C functions are typically split into header files (`.h`) where things are declared but not defined, and implementation files (`.c`) where they are defined. When we run the C compiler, a complex sequence of events is triggered with the usual successful outcome begin an executable file as illustrated at http://www.codingunit.com/
#
# 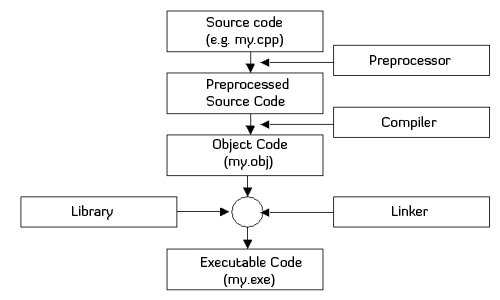
#
# The preprocessor merges the contents of the header and implementation files, and also expands any macros. The compiler then translates these into low level object code (`.o`) for each file, and the linker then joins together the newly generated object code with pre-compiled object code from libraries to form an executable. Sometimes we just want to generate object code and save it as a library (e.g. so that we can use it in Python).
# ## A tutorial example - coding a Fibonacci function in C
# #### Python version
def fib(n):
a, b = 0, 1
for i in range(n):
a, b = a+b, a
return a
fib(100)
# #### C version
# ##### Header file
# +
# %%file fib.h
double fib(int n);
# -
# ##### Implemetnation file
# +
# %%file fib.c
double fib(int n) {
double a = 0, b = 1;
for (int i=0; i<n; i++) {
double tmp = b;
b = a;
a += tmp;
}
return a;
}
# -
# ##### Driver program
# +
# %%file main.c
#include <stdio.h> // for printf()
#include <stdlib.h> // for atoi())
#include "fib.h" // for fib()
int main(int argc, char* argv[]) {
int n = atoi(argv[1]);
printf("%f", fib(n));
}
# -
# ##### Makefile
# +
# %%file Makefile
CC=gcc
CFLAGS=-std=c99 -Wall
fib: main.o fib.o
$(CC) $(CFLAGS) -o fib main.o fib.o
main.o: main.c fib.h
$(CC) $(CFAGS) -c main.c
fib.o: fib.c
$(CC) $(CFLAGS) -c fib.c
clean:
rm -f *.o
# -
# ##### Compile
# ! make
# #### Run executable file
# ! ./fib 100
# C Basics
# ----
# ### Types in C
#
# The basic types are very simple - use int, float and double for numbers. In general, avoid float for plain C code as its lack of precision may bite you unless you are writing CUDA code. Strings are quite nasty to use in C - I would suggest doing all your string processing in Python ...
#
# Structs are sort of like classes in Python
#
# ```c
# struct point {
# double x;
# double y;
# double z;
# };
#
# struct point p1 = {.x = 1, .y = 2, .z = 3};
# struct point p2 = {1, 2, 3};
# struct point p3;
# p3.x = 1;
# p3.y = 2;
# p3.z = 3;
# ```
#
# You can define your own types using `typedef` -.e.g.
# ```c
# #include <stdio.h>
# struct point {
# double x;
# double y;
# double z;
# };
#
# typedef struct point point;
#
# int main() {
# point p = {1, 2, 3};
# printf("%.2f, %.2f, %.2f", p.x, p.y, p.z);
# };
# ```
#
# ### Operators
# Most of the operators in C are the same in Python, but an important difference is the increment/decrement operator. That is
# ```c
# int c = 10;
# c++; // same as c = c + 1, i.e., c is now 11
# c--; // same as c = c - 1, i.e.. c is now 10 again
# ```
#
# There are two forms of the increment operator - postfix ```c++``` and prefix ```++c```. Both increment the variable, but in an expression, the postfix version returns the value before the increment and the prefix returns the value after the increment.
#
# +
# %%file increment.c
#include <stdio.h>
#include <stdlib.h>
int main()
{
int x = 3, y;
y = x++; // x is incremented and y takes the value of x before incrementation
printf("x = %d, y = %d\n", x, y);
y = ++x; // x is incremented and y takes the value of x after incrementation
printf("x = %d, y = %d\n", x, y);
}
# + language="bash"
#
# gcc -std=c99 -Wall increment.c -o increment
# ./increment
# -
# #### Ternary operator
#
# The ternary operator `expr = condition ? expr1 : expr2` allows an if-else statement to be put in a single line. In English, this says that if condition is True, expr1 is assigned to expr, otherwise expr2 is assigned to expr. We used it in the tutorial code to print a comma between elements in a list unless the element was the last one, in which case we printed a new line '\n'.
#
# Note: There is a similar ternary construct in Python `expr = expr1 if condition else epxr2`.
# ### Control of program flow
#
# Very similar to Python or R. The examples below should be self-explanatory.
# #### if-else
# ```C
# // Interpretation of grades by Asian parent
# if (grade == 'A') {
# printf("Acceptable\n");
# } else if (grade == 'B') {
# printf("Bad\n");
# } else if (grade == 'C') {
# printf("Catastrophe\n");
# } else if (grade == 'D') {
# printf("Disowned\n");
# } else {
# printf("Missing child report filed with local police\n")
# }
# ```
# #### for, while, do
# ```C
# // Looping variants
#
# // the for loop in C consists of the keyword for followed by
# // (initializing statement; loop condition statement; loop update statement)
# // followed by the body of the loop in curly braces
# int arr[3] = {1, 2, 3};
# for (int i=0; i<sizeof(arr)/sizeof(arr[0]); i++) {
# printf("%d\n", i);
# }
#
# int i = 3;
# while (i > 0) {
# i--;
# }
#
# int i = 3;
# do {
# i==;
# } while (i > 0);
# ```
# ### Arrays and pointers
# #### Automatic arrays
# If you know the size of the arrays at initialization (i.e. when the program is first run), you can usually get away with the use of fixed size arrays for which C will automatically manage memory for you.
#
# ```c
# int len = 3;
#
# // Giving an explicit size
# double xs[len];
# for (int i=0; i<len; i++) {
# xs[i] = 0.0;
# }
#
# // C can infer size if initializer is given
# double ys[] = {1, 2, 3};
# ```
#
# #### Pointers and dynamic memory management
#
# Otherwise, we have to manage memory ourselves using pointers. Basically, memory in C can be automatic, static or dynamic. Variables in automatic memory are managed by the computer, when it goes out of *scope*, the variable disappears. Static variables essentially live forever. Dynamic memory is allocated in the *stack*, and you manage its lifetime.
#
# Mini-glossary:
# * **scope**: Where a variable is visible - basically C variables have *block* scope - variables either live within a pair of curly braces (including variables in parentheses just before block such as function arguments and the counter in a for loop), or they are visible throughout the file.
# * **stack**: Computer memory is divided into a stack (small) and a heap (big). Automatic variables are put on the stack; dynamic variables are put in the heap. Hence if you have a very large array, you would use dynamic memory allocation even if you know its size at initialization.
#
# Any variable in memory has an address represented as a 64-bit integer in most operating systems. A pointer is basically an integer containing the address of a block of memory. This is what is returned by functions such as `malloc`. In C, a pointer is denoted by '*'. However, the '*' notation is confusing because its interpretation depends on whenever you are using it in a declaration or not. In a declaration
# ```c
# int *p = malloc(sizeof(int)); // p is a pointer to an integer
# *p = 5; // *p is an integer
# ```
#
# To get the actual address value, we can use the `&` address operator. This is often used so that a function can alter the value of an argument passed in (e.g. see address.c below).
# +
# %%file pointers.c
#include <stdio.h>
int main()
{
int i = 2;
int j = 3;
int *p;
int *q;
*p = i;
q = &j;
printf("p = %p\n", p);
printf("*p = %d\n", *p);
printf("&p = %p\n", &p);
printf("q = %p\n", q);
printf("*q = %d\n", *q);
printf("&q = %p\n", &q);
}
# + language="bash"
#
# gcc -std=c99 -Wall -Wno-uninitialized pointers.c -o pointers
# ./pointers
# +
# %%file address.c
#include <stdio.h>
void change_arg(int *p) {
*p *= 2;
}
int main()
{
int x = 5;
change_arg(&x);
printf("%d\n", x);
}
# + language="bash"
#
# gcc -std=c99 -Wall address.c -o address
# ./address
# -
# ### Pointer arithmetic
# If we want to store a whole sequence of ints, we can do so by simply allocating more memory:
#
# ```c
# int *ps = malloc(5 * sizeof(int)); // ps is a pointer to an integer
# for (int i=0; i<5; i++) {
# ps[i] = i;
# }
# ```
#
# The computer will find enough space in the heap to store 5 consecutive integers in a **contiguous** way. Since C arrays are all fo the same type, this allows us to do **pointer arithmetic** - i.e. the pointer `ps` is the same as `&ps[0]` and `ps + 2` is the same as `&ps[2]`. An example at this point is helpful.
# +
# %%file pointers2.c
#include <stdio.h>
#include <stdlib.h>
int main()
{
int *ps = malloc(5 * sizeof(int));
for (int i =0; i < 5; i++) {
ps[i] = i + 10;
}
printf("%d, %d\n", *ps, ps[0]); // remmeber that *ptr is just a regular variable outside of a declaration, in this case, an int
printf("%d, %d\n", *(ps+2), ps[2]);
printf("%d, %d\n", *(ps+4), *(&ps[4])); // * and & are inverses
}
# + language="bash"
#
# gcc -std=c99 -Wall pointers2.c -o pointers2
# ./pointers2
# -
# #### Pointers and arrays
#
# An array name is actually just a constant pointer to the address of the beginning of the array. Hence, we can dereference an array name just like a pointer. We can also do pointer arithmetic with array names - this leads to the following legal but weird syntax:
#
# ```c
# arr[i] = *(arr + i) = i[arr]
# ```
# +
# %%file array_pointer.c
#include <stdio.h>
int main()
{
int arr[] = {1, 2, 3};
printf("%d\t%d\t%d\t%d\t%d\t%d\n", *arr, arr[0], 0[arr], *(arr + 2), arr[2], 2[arr]);
}
# + language="bash"
#
# gcc -std=c99 -Wall array_pointer.c -o array_pointer
# ./array_pointer
# -
# #### More on pointers
#
# **Different kinds of nothing**: There is a special null pointer indicated by the keyword NULL that points to nothing. It is typically used for pointer comparisons, since NULL pointers are guaranteed to compare as not equal to any other pointer (including another NULL). In particular, it is often used as a sentinel value to mark the end of a list. In contrast a void pointer (void \*) points to a memory location whose type is not declared. It is used in C for generic operations - for example, `malloc` returns a void pointer. To totally confuse the beginning C student, there is also the NUL keyword, which refers to the `'\0'` character used to terminate C strings. NUL and NULL are totally different beasts.
#
# **Deciphering pointer idioms**: A common C idiom that you should get used to is `*q++ = *p++` where p and q are both pointers. In English, this says
#
# * \*q = \*p (copy the variable pointed to by p into the variable pointed to by q)
# * increment q
# * increment p
# +
# %%file pointers3.c
#include <stdio.h>
#include <stdlib.h>
int main()
{
// example 1
typedef char* string;
string s[] = {"mary ", "had ", "a ", "little ", "lamb", NULL};
for (char **sp = s; *sp != NULL; sp++) {
printf("%s", *sp);
}
printf("\n");
// example 2
char *src = "abcde";
char *dest = malloc(5); // char is always 1 byte by C99 definition
char *p = src + 4;
char *q = dest;
while ((*q++ = *p--)); // put the string in src into dest in reverse order
for (int i = 0; i < 5; i++) {
printf("i = %d, src[i] = %c, dest[i] = %c\n", i, src[i], dest[i]);
}
}
# + language="bash"
#
# gcc -std=c99 -Wall pointers3.c -o pointers3
# ./pointers3
# -
# ### Functions
# +
# %%file square.c
#include <stdio.h>
double square(double x)
{
return x * x;
}
int main()
{
double a = 3;
printf("%f\n", square(a));
}
# + language="bash"
#
# gcc -std=c99 -Wall square.c -o square
# ./square
# -
# ### Function pointers
#
# How to make a nice function pointer: Start with a regular function declaration func, for example, here func is a function that takes a pair of ints and returns an int
#
# ```
# int func(int, int);
# ```
#
# To turn it to a function pointer, just add a `*` and wrap in parenthesis like so
#
# ```
# int (*func)(int, int);
# ```
#
# Now `func` is a pointer to a function that takes a pair of ints and returns an int. Finally, add a typedef so that we can use `func` as a new type
# ```
# typedef int (*func)(int, int);
# ```
# which allows us to create arrays of function pointers, higher order functions etc as shown in the following example.
# +
# %%file square2.c
#include <stdio.h>
#include <math.h>
// Create a function pointer type that takes a double and returns a double
typedef double (*func)(double x);
// A higher order function that takes just such a function pointer
double apply(func f, double x)
{
return f(x);
}
double square(double x)
{
return x * x;
}
double cube(double x)
{
return pow(x, 3);
}
int main()
{
double a = 3;
func fs[] = {square, cube, NULL};
for (func *f=fs; *f; f++) {
printf("%.1f\n", apply(*f, a));
}
}
# + language="bash"
#
# gcc -std=c99 -Wall -lm square2.c -o square2
# ./square2
# -
# ### Using make to compile C programs
#
# As you have seen, the process of C program compilation can be quite messy, with all sorts of different compiler and linker flags to specify, libraries to add and so on. For this reason, most C programs are compiled using the `make` built tool that you are already familiar with. Here is a simple generic makefile that you can customize to compile your own programs adapted from the book 21st Century C by <NAME> (O'Reilly Media).
#
# * **TARGET**: Typically the name of the executable
# * **OBJECTS**: The intermediate object files - typically there is one file.o for every file.c
# * **CFLAGS**: Compiler flags, e.g. -Wall (show all warnings), -g (add debug information), -O3 (use level 3 optimization). Also used to indicate paths to headers in non-standard locations, e.g. -I/opt/include
# * **LDFLAGS**: Linker flags, e.g. -lm (link against the libmath library). Also used to indicate path to libraries in non-standard locations, e.g. -L/opt/lib
# * **CC**: Compiler, e.g. gcc or clang or icc
#
# In addition, there are traditional dummy flags
# * **all**: Builds all targets (for example, you may also have html and pdf targets that are optional)
# * **clean**: Remove intermediate and final products generated by the makefile
# +
# %%file makefile
TARGET =
OBJECTS =
CFLAGS = -g -Wall -O3
LDLIBS =
CC = c99
all: TARGET
clean:
rm $(TARGET) $(OBJECTS)
$(TARGET): $(OBJECTS)
# -
# Just fill in the blanks with whatever is appropriate for your program. Here is a simple example where the main file `test_main.c` uses a function from `stuff.c` with declarations in `stuff.h` and also depends on the libm C math library.
# +
# %%file stuff.h
#include <stdio.h>
#include <math.h>
void do_stuff();
# +
# %%file stuff.c
#include "stuff.h"
void do_stuff() {
printf("The square root of 2 is %.2f\n", sqrt(2));
}
# +
# %%file test_make.c
#include "stuff.h"
int main()
{
do_stuff();
}
# +
# %%file makefile
TARGET = test_make
OBJECTS = stuff.o
CFLAGS = -std=c99 -g -Wall -O3
LDLIBS = -lm
CC = gcc
all: $(TARGET)
clean:
rm $(TARGET) $(OBJECTS)
$(TARGET): $(OBJECTS)
# -
# ! make
# ! ./test_make
# Make is clever enough to recompile only what has been changed since the last time it was called
# ! make
# ! make clean
# ! make
# ### Exercise
# ### Debugging programs (understanding compiler warnings and errors)
#
# Try to fix the following buggy program.
# +
# %%file buggy.c
# Create a function pointer type that takes a double and returns a double
double *func(double x);
# A higher order function that takes just such a function pointer
double apply(func f, double x)
{
return f(x);
}
double square(double x)
{
return x * x;
}
double cube(double x)
{
return pow(3, x);
}
double mystery(double x)
{
double y = 10;
if (x < 10)
x = square(x);
else
x += y;
x = cube(x);
return x;
}
int main()
{
double a = 3;
func fs[] = {square, cube, mystery, NULL}
for (func *f=fs, f != NULL, f++) {
printf("%d\n", apply(f, a));
}
}
# -
# ! gcc -std=c99 -g -Wall buggy.c -o buggy
# ### Why not C?
#
# What other language has an annual Obfuscated Code Contest <http://www.ioccc.org/>? In particular, the following features of C are very conducive to writing unreadable code:
#
# * lax rules for identifiers (e.g. _o, _0, _O, O are all valid identifiers)
# * chars are bytes and pointers are integers
# * pointer arithmetic means that `array[index]` is the same as `*(array+index)` which is the same as `index[array]`!
# * lax formatting rules especially with respect to whitespace (or lack of it)
# * Use of the comma operator to combine multiple expressions together with the ?: operator
# * Recursive function calls - e.g. main calling main repeatedly is legal C
#
# Here is one winning entry from the 2013 IOCCC [entry](http://www.ioccc.org/2013/dlowe/hint.html) that should warm the heart of statisticians - it displays sparklines (invented by Tufte).
#
# ```c
# main(a,b)char**b;{int c=1,d=c,e=a-d;for(;e;e--)_(e)<_(c)?c=e:_(e)>_(d)?d=e:7;
# while(++e<a)printf("\xe2\x96%c",129+(**b=8*(_(e)-_(c))/(_(d)-_(c))));}
# ```
# %%file sparkl.c
main(a,b)char**b;{int c=1,d=c,e=a-d;for(;e;e--)_(e)<_(c)?c=e:_(e)>_(d)?d=e:7;
while(++e<a)printf("\xe2\x96%c",129+(**b=8*(_(e)-_(c))/(_(d)-_(c))));}
# ! gcc -Wno-implicit-int -include stdio.h -include stdlib.h -D'_(x)=strtof(b[x],0)' sparkl.c -o sparkl
import numpy as np
np.set_printoptions(linewidth=np.infty)
print(' '.join(map(str, (100*np.sin(np.linspace(0, 8*np.pi, 30))).astype('int'))))
# + language="bash"
#
# ./sparkl 0 76 98 51 -31 -92 -88 -21 60 99 68 -10 -82 -96 -41 41 96 82 10 -68 -99 -60 21 88 92 31 -51 -98 -76 0
# -
# ### Learning Obfuscated C
# If you have too much time on your hands and really want to know how **not** to write C code (unless you are crafting an entry for the IOCCC), I recommend this tutorial
# <http://www.dreamincode.net/forums/topic/38102-obfuscated-code-a-simple-introduction/>
| notebooks/S09B_C.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
from scipy import signal
from numpy.fft import fft, rfft, ifft
from audlib.sig.fbanks import Gammatone
from audlib.plot import magresp
plt.rc('text', usetex=True)
# +
sr = 16000
nfft = 2**10
nyquist = nfft//2+1 # Boundary for Nyquist frequency
num_chans = 40
gtbank = Gammatone(sr, num_chans)
### Plot filter responses
fig = plt.figure(figsize=(16, 12), dpi= 100)
ax1 = fig.add_subplot(211)
total = np.zeros((nyquist), dtype=np.complex_)
for kk in range(len(gtbank)):
ww, hh = gtbank.freqz(kk, nfft=nfft, powernorm=True)
total += hh[:nyquist]
magresp(ww[:nyquist]*(sr/2), hh[:nyquist], ax1, units=('hz', 'mag'))
# Plot total filter response
ax2 = fig.add_subplot(212)
magresp(ww[:nyquist], total, ax2, units=('rad', 'mag'))
# -
# The implementation is a direct translation of <NAME>'s `ERBFilteBank` and `MakeERBFilters` in the *Auditory Toolbox*. Below is a 40-channel Gammatone filterbank extracted using his MATLAB code:
#
# 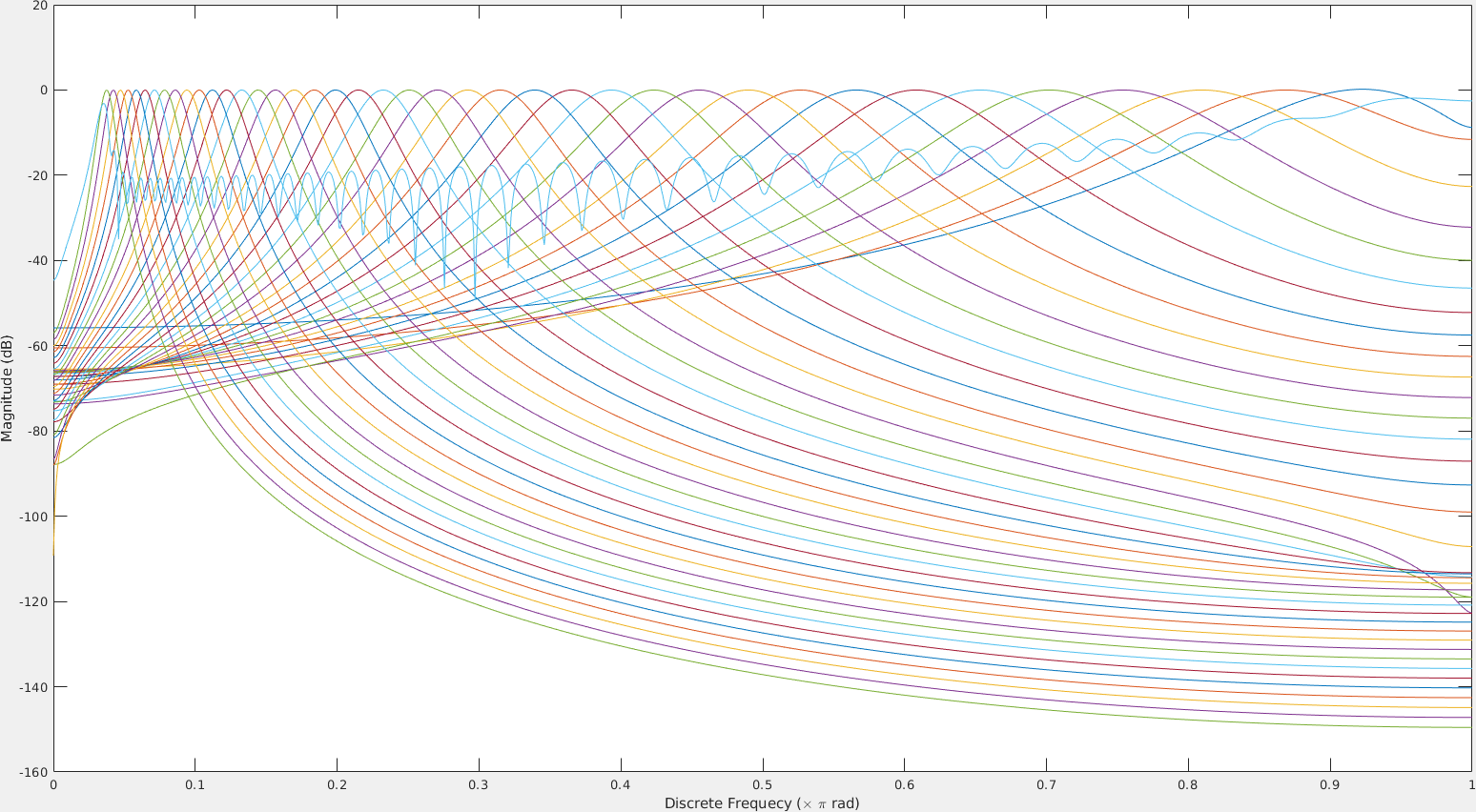
# +
# Obtain Gammatone-weighted short-time power spectr
from audlib.quickstart import welcome
from audlib.sig.transform import stpowspec
from audlib.sig.window import hamming
from audlib.plot import specgram
sig, sr = welcome()
wlen = .025
hop = .01
nfft = 1024
wind = hamming(int(wlen*sr))
powerspec = stpowspec(sig, wind, int(hop*sr), nfft, synth=False)
wts = gtbank.gammawgt(nfft, powernorm=True, squared=True)
gammaspec = powerspec @ wts
print(gammaspec.shape)
fig = plt.figure(figsize=(16, 12), dpi= 100)
ax1 = fig.add_subplot(211)
specgram(10*np.log10(gammaspec), ax1, time_axis=np.arange(gammaspec.shape[0])*hop, freq_axis=gtbank.cf)
| examples/dsp/Gammatone Filterbank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Let us first import all the necessary packages.
from __future__ import division, print_function
import torch
from torch.autograd import Variable
from torch_submod.graph_cuts import TotalVariation2dWeighted
from matplotlib import pyplot as plt
# %matplotlib inline
from tqdm import tqdm_notebook as tqdm
from pandas import DataFrame
# We will plot the images using the following function.
def plot(x):
plt.figure(figsize=(5, 10))
plt.imshow(x.numpy())
# -
# ### Data creation
#
# We will create first a simple image ``x``, and corrupt it with normal noise to get ``x_noisy``.
# +
device = 'cpu'
torch.manual_seed(0) # To ensure reproducibility.
m, n = 50, 100 # The image dimensions.
std = 1e-1 # The standard deviation of noise.
x = torch.zeros((m, n), device=device)
x[:m//2, :n//2] += 1
x_noisy = x + torch.normal(torch.zeros(x.size(), device=device))
plot(x.cpu())
plot(x_noisy.cpu())
# +
# The learnable parameters.
# First, we will fix the column and row-weights to be equal.
# We parametrize the log of these weights.
log_w_row = (- 3 * torch.ones(1, device=device)).requires_grad_(True)
log_w_col = (- 3 * torch.ones(1, device=device)).requires_grad_(True)
# We also have a scale weight.
scale = (- torch.ones(1, device=device)).requires_grad_(True)
tv2d = TotalVariation2dWeighted()
optimizer = torch.optim.SGD([log_w_row, log_w_col, scale], lr=.5)
# We keep data of the form (w_row, w_col, scale) at each iteration.
data = []
losses = []
for iter_no in tqdm(range(1000)):
w_row = torch.exp(log_w_row)
w_col = torch.exp(log_w_col)
y = tv2d(scale * x_noisy, w_row.expand((m, n-1)), w_col.expand((m - 1, n)))
optimizer.zero_grad()
loss = torch.mean((y - x)**2)
loss.backward()
if iter_no % 100 == 0:
losses.append(loss.item())
plot(y.detach().cpu())
data.append((w_row.item(), w_col.item(), scale.item()))
optimizer.step()
# -
df = DataFrame(data, columns=('w_col', 'w_row', 'scale'))
axes = df.plot(subplots=True)
axes[-1].set_xlabel('iteration');
| notebooks/denoising.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ipycytoscape: Grafos interativos
#
# ## Uma Jupyter - Cytoscape ponte
#
# ## https://github.com/cytoscape/ipycytoscape
#
# Um widget que permite a visualização de grafos interativos com cytoscape.js em JupyterLab e Jupyter notebook.
#
# - BSD-3-Clause License
#
# **Instalação:**
#
# ```bash
# conda install -c conda-forge ipycytoscape
# ```
# ### ipycytoscape walkthrough
#
# - cytoscape json
# - layouts
# - pandas
# - styling
# - networkx
# - grafos direcionados e não direcionados
# - criando elementos personalizados
# - neo4j
# - interatividade com ipywidgets
import ipycytoscape as cy
cyGraph = cy.CytoscapeWidget()
# ### **cytoscape json**
cytoscape_json = {
'nodes': [
{ 'data': { 'id': '0', 'name': 'Cytoscape', 'classes': 'node' }},
{ 'data': { 'id': '1', 'name': 'Grid', 'classes': 'node' }},
{ 'data': { 'id': '2', 'name': 'Cola', 'classes': 'node' }},
{ 'data': { 'id': '3', 'name': 'Popper', 'classes': 'node' }},
{ 'data': { 'id': '4', 'name': 'Cytoscape.js', 'classes': 'node'}}
],
'edges': [
{'data': { 'source': '4', 'target': '0' }},
{'data': { 'source': '1', 'target': '2' }},
{'data': { 'source': '1', 'target': '3' }},
{'data': { 'source': '2', 'target': '3' }},
{'data': { 'source': '4', 'target': '4' }},
{'data': { 'source': '4', 'target': '3' }},
]
}
# Todos os exemplos neste notebook são criados in loco para tornar o processo mais simples. Mas também é possível carregá-los de muitos outras formas diferentes.
#
# Para uma descrição mais extensa de todos os atributos disponíveis em um grafo cytoscape: [documentação cytoscape](https://js.cytoscape.org/#cy.data).
cyGraph.graph.add_graph_from_json(cytoscape_json)
cyGraph
# #### Layouts:
#
# - cola
# - dagre
# - euler
# - cose
# - breadthfirst
# - circle
# - grid
# - random
# - null
#
#
# <img src="attachment:16109f49-1053-42ab-b02d-9685e0743e6e.png" width="500"/>
cyGraph.set_layout(name='random')
cyGraph
# ### **pandas**
import pandas as pd
pdCy = cy.CytoscapeWidget()
robots = ['marvin','c3po','r2d2','data']
universe = ['douglas adams','star wars','star wars','star trek']
cooleness_lvl = ['10', '3', '10', '10']
robotsRates = list(zip(robots, universe, cooleness_lvl))
df = pd.DataFrame(data = robotsRates, columns=['robot', 'universe', 'cooleness_lvl'])
pdCy.graph.add_graph_from_df(df, ['universe'], ['robot', 'cooleness_lvl'])
pdCy
pdCy.graph.nodes
edge = cy.Edge(data={"source": 1, "target": 2})
pdCy.graph.add_edge(edge)
edge = cy.Edge(data={"source": 'parent-2', "target": 'parent-1'})
pdCy.graph.add_edge(edge)
pdCy.graph.edges
# #### **styling**
#
# - layout
# - style
# - tooltips
pdCy.set_style([
{
'selector': 'node[name] *= ""',
'style': {
'background-color': 'blue',
}},
{
'selector': '[id *= "parent-1"]',
'style': {
'background-color': 'yellow',
}},
{
'selector': '[id *= "parent-2"]',
'style': {
'background-color': 'red',
'color': 'blue'
}
}])
pdCy
# Mais informação sobre seletores cytoscape estão disponíveis [aqui](https://js.cytoscape.org/#selectors).
pdCy.set_tooltip_source('name')
# Mais exemplos em como personalizar labels e tooltips [aqui](https://github.com/cytoscape/ipycytoscape/blob/master/examples/Text%20on%20node.ipynb) e [aqui](https://github.com/cytoscape/ipycytoscape/blob/master/examples/Tooltips%20example.ipynb). Ou online via [binder](https://mybinder.org/v2/gh/QuantStack/ipycytoscape/stable?filepath=examples).
# ### **networkx**
import networkx as nx
nxCyGraph = cy.CytoscapeWidget()
nxGraph = nx.complete_graph(5)
nxCyGraph.graph.add_graph_from_networkx(nxGraph)
nxCyGraph
# #### **grafos direcionados e não direcionados**
directCyGraph = cy.CytoscapeWidget()
directNxGraph = nx.complete_graph(4)
directCyGraph.graph.add_graph_from_networkx(directNxGraph, directed=True)
directCyGraph
from random import random
# +
mixedNxGraph = nx.complete_graph(5)
for s, t, data in mixedNxGraph.edges(data=True):
if random() > .5:
mixedNxGraph[s][t]['classes'] = 'directed'
mixedGraph = cy.CytoscapeWidget()
mixedGraph.graph.add_graph_from_networkx(mixedNxGraph)
mixedGraph
# -
# #### **Criando elementos personalizados**
# Nós e arestas personalizados podem ser criados para qualquer tipo de grafo, isso signfica que apesar dos exemplos vistos aqui, não é apenas restrito à objetos networkx. Para isso, basta usar a API do ipycytoscape. Nós vemos um exemplo de como usar essa API no exemplo a seguir, sobre Pandas.
class CustomNode(cy.Node):
def __init__(self, name, classes = ''):
super().__init__()
self.data['id'] = name
self.classes = classes
# +
n1 = CustomNode("node 1", classes = 'first')
n2 = CustomNode("node 2", classes = 'second')
G = nx.Graph()
G.add_node(n1)
G.add_node(n2)
G.add_edge(n1, n2)
# -
customInheritedGraph = cy.CytoscapeWidget()
customInheritedGraph.graph.add_graph_from_networkx(G)
customInheritedGraph.graph.nodes
# ### **Neo4j**
#
# Exemplos para como executar código ipycytoscape com Neo4j podem ser encontrados [aqui](https://github.com/sbl-sdsc/neo4j-ipycytoscape) binder e um notebook [aqui](https://binder.pangeo.io/v2/gh/sbl-sdsc/neo4j-ipycytoscape/master).
# +
#from py2neo import Graph
#cy.add_graph_from_neo4j(neo4j_graph)
# -
# ### **interatividade e interoperabilidade com ipywidgets**
#
# - eventos javascript
# - DOM (elements, events)
# - interaçâo com widgets
import ipywidgets as widgets
cyGraph
cyGraph.set_style([{
"selector": "edge.highlighted",
"css": {
"line-color": "red"
}
}])
# +
btn = widgets.Button(description = "red edges", disabled = False)
def btn_callback(b):
for edge in cyGraph.graph.edges:
edge.classes = " highlighted"
btn.on_click(callback = btn_callback)
display(btn)
# -
def paint_blue(event):
auxNode = cyGraph.graph.nodes[int(event['data']['id'])]
auxNode.classes += ' blue'
cyGraph.on('node', 'click', paint_blue)
cyGraph.set_style([{
"selector": "edge.highlighted",
"css": {
"line-color": "red"
}
},
{
"selector": "node.blue",
"css": {
"background-color": "blue"
},
}])
cyGraph
# **Lista de eventos**
#
# Esses eventos podem ser aplicado a objetos `nodes` ou `edges`:
#
# * **mousedown :** quando o botão do mouse é pressionado
# * **mouseup :** quando o botão do mouse é solto
# * **click :** mousedown então mouseup
# * **mouseover :** quando o cursor é colocado em cima do
# * **mouseout :** quando o cursor é movido para longe do targe
# * **mousemove :** quando o cursor é movido para o target
# * **touchstart :** quando um ou mais dedos começam a tocar a tela
# * **touchmove :** quando um ou mais dedos movem sobre a tela
# * **touchend :** quando um ou mais dedos são removidos da tela
# * **tapstart :** tap para começar um evento
# * **vmousedown :** alias para 'tapstart'
# * **tapdrag :** evento de movimento
# * **vmousemove :** alias para 'tapdrag'
#
# E muitos outros! Cheque a documentação do cytoscape para a lista completa.
# # Exercísio
# ### 1) Create another instance of the cyGraph object
# ### 2) Create a button widget, that when clicked, paints the graph's nodes yellow
| notebooks/07.Mais-bibliotecas/07.02-ipycytoscape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import the function BeautifulSoup from the package bs4
# Import packages
import requests
from bs4 import BeautifulSoup
# ### Assign the URL of interest to the variable url.
# Specify url: url
url = 'https://www.python.org/~guido/'
# ### Package the request to the URL, send the request and catch the response with a single function requests.get(), assigning the response to the variable r.
# Package the request, send the request and catch the response: r
r= requests.get(url)
# ### Use the text attribute of the object r to return the HTML of the webpage as a string; store the result in a variable html_doc.
# Extracts the response as html: html_doc
html_doc= r.text
# ### Create a BeautifulSoup object soup from the resulting HTML using the function BeautifulSoup().
#
# Create a BeautifulSoup object from the HTML: soup
soup=BeautifulSoup(html_doc,"lxml")
# ### Use the method prettify() on soup and assign the result to pretty_soup.
# Prettify the BeautifulSoup object: pretty_soup
pretty_soup = soup.prettify()
# Print the response
print(pretty_soup)
| import data in python part 2/1.7 Parsing HTML with BeautifulSoup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Preliminaries
# Data preparation
import os
import pandas as pd
import numpy as np
HOUSING_PATH = "datasets/housing"
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
housing = load_housing_data()
housing["income_cat"] = np.ceil(housing["median_income"] / 1.5)
housing["income_cat"].where(housing["income_cat"] < 5, 5.0, inplace=True)
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing["income_cat"]):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
for set_ in (strat_train_set, strat_test_set):
set_.drop(["income_cat"], axis=1, inplace=True)
housing = strat_train_set.drop("median_house_value", axis=1)
housing_labels = strat_train_set["median_house_value"].copy()
housing_num = housing.drop("ocean_proximity", axis=1)
from sklearn.preprocessing import Imputer
from future_encoders import OneHotEncoder
from sklearn.base import BaseEstimator, TransformerMixin
# column index
rooms_ix, bedrooms_ix, population_ix, household_ix = 3, 4, 5, 6
class CombinedAttributesAdder(BaseEstimator, TransformerMixin):
def __init__(self, add_bedrooms_per_room = True): # no *args or **kargs
self.add_bedrooms_per_room = add_bedrooms_per_room
def fit(self, X, y=None):
return self # nothing else to do
def transform(self, X, y=None):
rooms_per_household = X[:, rooms_ix] / X[:, household_ix]
population_per_household = X[:, population_ix] / X[:, household_ix]
if self.add_bedrooms_per_room:
bedrooms_per_room = X[:, bedrooms_ix] / X[:, rooms_ix]
return np.c_[X, rooms_per_household, population_per_household,
bedrooms_per_room]
else:
return np.c_[X, rooms_per_household, population_per_household]
attr_adder = CombinedAttributesAdder(add_bedrooms_per_room=False)
housing_extra_attribs = attr_adder.transform(housing.values)
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
num_pipeline = Pipeline([
('imputer', Imputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler())
])
housing_num_tr = num_pipeline.fit_transform(housing_num)
from future_encoders import ColumnTransformer
num_attribs = list(housing_num)
cat_attribs = ["ocean_proximity"]
full_pipeline = ColumnTransformer([
("num", num_pipeline, num_attribs),
("cat", OneHotEncoder(), cat_attribs)
])
housing_prepared = full_pipeline.fit_transform(housing)
# -
# ### 1.Try a Support Vector Machine regressor (`sklearn.svm.SVR`), with various hyperparameters such as `kernel="linear"` (with various values for the `C` hyperparameter) or `kernel="rbf"` (with various values for the `C` and `gamma` hyperparameters). Don’t worry about what these hyperparameters mean for now. How does the best `SVR` predictor perform?
# +
from sklearn.svm import SVR
# default SVM with linear kernel
svm_reg = SVR(kernel="linear")
svm_reg.fit(housing_prepared, housing_labels)
# -
housing_predictions = svm_reg.predict(housing_prepared)
svm_mse = mean_squared_error(housing_labels, housing_predictions)
svm_rmse = np.sqrt(svm_mse)
svm_rmse
# +
param_grid = [
{'kernel': ['linear'], 'C': [1, 3, 10, 30, 100, 300, 1000, 3000, 10000, 30000]}
]
svm_reg = SVR()
grid_search = GridSearchCV(svm_reg, param_grid, cv=5, scoring='neg_mean_squared_error')
grid_search.fit(housing_prepared, housing_labels)
# -
grid_search.best_params_
np.sqrt(-grid_search.best_score_)
# +
param_grid = [
{'kernel': ['rbf'], 'C': [10, 30, 100, 300, 1000, 3000, 10000, 30000],
'gamma': [0.01, 0.03, 0.1, 0.3, 1, 3]}
]
svm_reg = SVR()
grid_search = GridSearchCV(svm_reg, param_grid, cv=5, scoring='neg_mean_squared_error')
grid_search.fit(housing_prepared, housing_labels)
# -
feature_importances = grid_search.best_estimator_.feature_importances_
# ### 2. Try replacing `GridSearchCV` with `RandomizedSearchCV`.
# +
# replacing in the context code
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import randint
param_distribs = {
'n_estimators': randint(low=1, high=100),
'max_features': randint(low=1, high=8)
}
forest_reg = RandomForestRegressor()
rnd_search = RandomizedSearchCV(forest_reg, param_distributions=param_distribs,
n_iter=10, cv=5, scoring='neg_mean_squared_error', random_state=42)
rnd_search.fit(housing_prepared, housing_labels)
# -
rnd_search.best_params_
np.sqrt(-rnd_search.best_score_)
# +
# the following code is referring to the author's solution
# https://github.com/ageron/handson-ml/blob/master/02_end_to_end_machine_learning_project.ipynb
from scipy.stats import expon, reciprocal
param_distribs = {
'kernel':['linear', 'rbf'],
'C': reciprocal(20, 200000),
'gamma': expon(scale=1.0)
}
svm_reg = SVR()
rnd_search = RandomizedSearchCV(svm_reg, param_distributions=param_distribs,
n_iter=50, cv=5, scoring='neg_mean_squared_error',
verbose=2, n_jobs=4, random_state=42)
rnd_search.fit(housing_prepared, housing_labels)
# -
np.sqrt(-rnd_search.best_score_)
rnd_search.best_params_
# ### 3. Try adding a transformer in the preparation pipeline to select only the most important attributes.
# ### 4. Try creating a single pipeline that does the full data preparation plus the final prediction.
# ### 5. Automatically explore some preparation options using `GridSearchCV`.
| Chapter 2/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from PIL import Image
import PIL
import os
import cv2 as cv
BACKGROUND_DIR = os.getcwd() + "/Backgrounds/"
DIR = os.getcwd() + "/Icons/"
# -
# ### Icon
#
# Run this document when you want to add new icons and process them. A few things to keep in mind:
# 1. Make sure the picture you have taken, has the icon somewhat centered. It makes things easier.
# 2. Adjust the processed image, by ysing the 5 parameters left, right, up, down and margin.
# 3. Be careful not to overwrite old images, by forgetting to change the filename.
# +
def white_to_transparency(img):
image_array = np.asarray(img.convert('RGBA')).copy()
HEIGHT = np.shape(image_array)[0]
INFLECTION_POINT = int(np.sum(image_array[0,:,0]) / HEIGHT) - 10
INFLECTION_POINT = 100
image_array[image_array > INFLECTION_POINT] = 255
image_array[image_array < INFLECTION_POINT] = 0
image_array[:, :, 3] = (255 * (image_array[:, :, :3] != 255).any(axis=2)).astype(np.uint8)
return Image.fromarray(image_array)
def extract_icon(image_array, LOCATION, MARGIN):
HEIGHT, WIDTH = np.shape(image_array)
up, down = LOCATION[0]
left, right = LOCATION[1]
offset_x, offset_y = LOCATION
newimgarr = image_array[int(HEIGHT/2) - MARGIN - up:int(HEIGHT/2) + MARGIN + down,
int(WIDTH/2) - MARGIN + left:int(WIDTH/2) + MARGIN + right]
return Image.fromarray(newimgarr)#.resize((40, 40))
def get_mask(FILENAME):
img = cv.imread(FILENAME, 0)
_, th1 = cv.threshold(img, 100, 1, cv.THRESH_BINARY_INV)
return th1
def save_mask(FILENAME):
mask = get_mask("mask_" + FILENAME)
x_vals = mask[:, 0]
y_vals = mask[:, 1]
FILENAME = FILENAME.split(".")[0]
np.savez(FILENAME + '_mask.npz', x=x_vals, y=y_vals)
# +
FILENAME = "Home/Home2.png"
img = Image.open(DIR + FILENAME).convert("L")
imgarr = np.array(img)
UP = 150
DOWN = 140
LEFT = -190
RIGHT = 120
MARGIN = 50
_, th1 = cv.threshold(imgarr, 100, 1, cv.THRESH_BINARY_INV)
print(Image.fromarray(th1))
#icon_image = extract_icon(imgarr, [(UP, DOWN), (LEFT, RIGHT)], MARGIN)
icon_image.show()
#th1 = get_mask(FILENAME)
#img = Image.fromarray(th1)
#img.show()
#print(np.sum(get_mask(FILENAME)))
# -
icon_image = icon_image.resize((40, 40))
icon_image.save(DIR + "/" + FILENAME)
# + [markdown] heading_collapsed=true
# ### Save Icon
# Only run this when you need to save the icon image
# + hidden=true
# -
# ### Resize Backgrounds
#
# Resizes the backgrounds to (900, 1200) and makes sure the image is in landscape orientation. Only input is the filename.
FILENAME = 'background11.png'
TARGET_WIDTH, TARGET_HEIGHT = (900, 1200)
image = Image.open(BACKGROUND_DIR + FILENAME).convert("L")
imgarr = np.array(image)
HEIGHT, WIDTH = np.shape(imgarr)
if WIDTH > HEIGHT:
imgarr = imgarr.T
Image.fromarray(imgarr)
image = image.resize((TARGET_WIDTH, TARGET_HEIGHT), Image.ANTIALIAS)
image.show()
# ### Save Background
image.save(BACKGROUND_DIR + "/" + FILENAME)
# ### Project: Better results
import matplotlib.pyplot as plt
import cv2 as cv
FILENAME = "D4.png"
img = cv.imread(DIR + FILENAME,0)
img
plt.imshow(img,'gray')
ret,th1 = cv.threshold(img,100,255,cv.THRESH_BINARY_INV)
plt.imshow(-1 * th1,'gray')
th1
a, b = cv.findContours(th1, cv.RETR_TREE, cv.CHAIN_APPROX_SIMPLE)
a[4]
cv.drawContours(th1, a[3], -1, (0,255,0), 3)
plt.imshow(th2,'gray')
# Steps should be:
# * Find background image
# * Select icon
# * Select icon mask file
# * Insert icon on background
# * Use that position to alter the mask location
# * Insert the mask location into the via_regions file.
| samples/wireframe/Data_Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from utils import save
from sklearn.model_selection import train_test_split
# -
# ! ls -l ../data/
TRAIN_PATH = '../data/train.csv'
train_df = pd.read_csv(TRAIN_PATH)
train_df.head()
# Let's see how long these questions are.
# +
train_df['q1_len'] = train_df['question1'].str.len()
train_df['q2_len'] = train_df['question2'].str.len()
train_df.loc[:, ['q1_len', 'q2_len']].describe()
# -
# There seems to be some really small questions. Let's take a look at these questions.
train_df[train_df['q1_len'] == 1]
train_df[train_df['q2_len'] == 1]
# Let's confirm did not make a mistake reading the csv file.
# ! cat {TRAIN_PATH} | head -n 47060 | tail -n 1
# The number of questions do not seem to be matching the lines in `train.csv`. Let's confirm this.
print('Rows:',train_df.shape[0])
# ! wc -l {TRAIN_PATH}
# Let's confirm the last row id is present in both.
print(train_df['id'].tail(1))
# ! cat {TRAIN_PATH} | tail -n 1
train_df['id'].min()
# This seems accurate. The `id` in the train.csv is auto-incrementing and starts at 0, and the maximum value of the `id` column matches the last row in the flat file.
#
# I am going to remove all single character questions as this will be added noise in my classifier.
# +
train_df = train_df[train_df['q1_len'] != 1]
train_df = train_df[train_df['q2_len'] != 1]
train_df.describe()
# -
train_df.info()
# There appears to be some null values for questions. Let's take a look at this data and confirm if we can drop the data.
train_df[train_df['question1'].isna()]
train_df[train_df['question2'].isna()]
# !cat {TRAIN_PATH} | head -n 105785 | tail -n 1
# Appears there are indeed some blank questions. Any row with a blank question for either pair will be dropped.
train_df = train_df.dropna()
train_df.info()
# How many pairs of questions were flagged as duplicates in the dataset?
train_df[train_df['is_duplicate'] == 1].shape[0] / train_df.shape[0]
# About 37% of the question pairs are identified as duplicates. The training data is not that imbalanced.
#
# Let's now sort by id, perform train / test split and save the altered data frame.
# +
train_df = train_df.sort_values('id')
X, y = (train_df.loc[:,['id', 'question1', 'question2']], train_df.is_duplicate.values)
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state = 42)
# need to sort by id to keep consistent with the stacking and unstacking of questions
temp_y = pd.DataFrame(np.concatenate([X_train.id.values.reshape(-1, 1), y_train.reshape(-1, 1)], axis=1))
temp_y = temp_y.sort_values(0)
y_train = temp_y.loc[:, 1].values
X_train = X_train.sort_values('id')
temp_y = pd.DataFrame(np.concatenate([X_test.id.values.reshape(-1, 1), y_test.reshape(-1, 1)], axis=1))
temp_y = temp_y.sort_values(0)
y_test = temp_y.loc[:, 1].values
X_test = X_test.sort_values('id')
# -
save(train_df, 'train')
save(X_train, 'X_train')
save(X_test, 'X_test')
save(y_train, 'y_train')
save(y_test, 'y_test')
| py_files/00. Data Loading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + outputHidden=false inputHidden=false
import subprocess, re, json
from IPython.display import JSON, display
proc = subprocess.Popen(
'../node_modules/.bin/jest --json',
stdout=subprocess.PIPE, stderr=subprocess.PIPE, shell=True
)
raw = proc.stdout.read()
data = json.loads(raw)
display(JSON(data))
# + outputHidden=false inputHidden=false
from vdom import table, tr, td
runtimes = [
(re.split("packages/", x['name'])[1], x['endTime'] - x['startTime'])
for x in data['testResults']
]
display(
table(
[
tr(
td(name),
td(f"{time / 1000.}s")
) for (name, time) in
sorted(runtimes, key=lambda x: x[1], reverse=True)
if time > 1000
]
)
)
| scripts/evaluate-tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GSD: Assessing ambiguous nts in 1011 collection genomes (SMALL SET)
#
# **DEVELOPMENT WITH JUST TWO OF THE 1011 genomes collection (SMALL SET)**
#
# Checks a collection of sequenced cerevisiae genomes from [Peter et al 2018](https://www.ncbi.nlm.nih.gov/pubmed/29643504) for ambiguous/gap-representing residues. Compares them to a roughly dozen genomes sequenced by the PacBio method from [Yue et al., 2017](https://www.ncbi.nlm.nih.gov/pubmed/28416820) as well as the SGD reference sequence from [here](https://downloads.yeastgenome.org/sequence/S288C_reference/chromosomes/fasta/).
#
# References for sequence data:
# - [Genome evolution across 1,011 Saccharomyces cerevisiae isolates. <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Nature. 2018 Apr;556(7701):339-344. doi: 10.1038/s41586-018-0030-5. Epub 2018 Apr 11. PMID: 29643504](https://www.ncbi.nlm.nih.gov/pubmed/29643504)
#
# - [Contrasting evolutionary genome dynamics between domesticated and wild yeasts.
# Yue JX, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Nat Genet. 2017 Jun;49(6):913-924. doi: 10.1038/ng.3847. Epub 2017 Apr 17. PMID: 28416820](https://www.ncbi.nlm.nih.gov/pubmed/28416820)
#
#
# - [Life with 6000 genes. <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Science. 1996 Oct 25;274(5287):546, 563-7. PMID: 8849441](https://www.ncbi.nlm.nih.gov/pubmed/8849441)
#
#
# -----
# ## Preparation
#
# Get scripts and sequence data necessary.
#
# **Before doing `Run All`, make sure the fifth cell is set to get the desired set from the 1011 genomes collection.**
#
# !pip install pyfaidx
# Get the genomes data by running these commands.
import pandas as pd
# Prepare for getting PacBio (Yue et al 2017 sequences)
#make a list of the strain designations
yue_et_al_strains = ["S288C","DBVPG6044","DBVPG6765","SK1","Y12",
"YPS128","UWOPS034614","CBS432","N44","YPS138",
"UFRJ50816","UWOPS919171"]
# Get & unpack the genome sequences from strains
for s in yue_et_al_strains:
# !curl -LO http://yjx1217.github.io/Yeast_PacBio_2016/data/Nuclear_Genome/{s}.genome.fa.gz
# !curl -OL http://yjx1217.github.io/Yeast_PacBio_2016/data/Mitochondrial_Genome/{s}.mt.genome.fa.gz
# !gunzip -f {s}.genome.fa.gz
# !gunzip -f {s}.mt.genome.fa.gz
# add the mitochondrial genome content onto the nuclear
# !cat {s}.genome.fa {s}.mt.genome.fa > temp.genome.fa
# !mv temp.genome.fa {s}.genome.fa
# !rm *.mt.genome.fa
# +
# add identifiers to each `chr` so results for each strain clear later
chromosome_id_prefix = "chr"
def add_strain_id_to_description_line(file,strain_id):
'''
Takes a file and edits every description line to add
strain_id after the caret.
Saves the fixed file
'''
import sys
output_file_name = "temp.txt"
# prepare output file for saving so it will be open and ready
with open(output_file_name, 'w') as output_file:
# read in the input file
with open(file, 'r') as input_handler:
# prepare to give feeback later or allow skipping to certain start
lines_processed = 0
for line in input_handler:
lines_processed += 1
if line.startswith(">"):
rest_o_line = line.split(">")
new_line = ">"+strain_id + rest_o_line[1]
else:
new_line = line
# Send text to output
output_file.write(new_line)
# replace the original file with edited
# !mv temp.txt {file}
# Feedback
sys.stderr.write("\n{} chromosome identifiers tagged.".format(file))
for s in yue_et_al_strains:
add_strain_id_to_description_line(s+".genome.fa",s)
# -
# Get SGD reference sequence that includes nuclear and mitochondrial sequence as one file,
# among others. I'll use file name for the reference genome worked out
# in `GSD Assessing_ambiguous_nts_in_nuclear_PB_genomes.ipynb`, so more of the
# previously worked out code will work.
# !curl -OL https://downloads.yeastgenome.org/sequence/S288C_reference/genome_releases/S288C_reference_genome_Current_Release.tgz
# !tar -xzf S288C_reference_genome_Current_Release.tgz
# !rm S288C_reference_genome_Current_Release.tgz
# !mv S288C_reference_genome_R64-2-1_20150113/S288C_reference_sequence_R64-2-1_20150113.fsa ./SGD_REF.genome.fa
# !rm -rf S288C_reference_genome_R64-2-1_20150113
# **Now to get the entire collection or a subset of the 1011 genomes, the next cell will need to be edited.** I'll probably leave it with a small set for typical running purposes. However, to make it run fast, try the 'super-tiny' set with just two.
# +
# Method to get ALL the genomes. TAKES A WHILE!
# #!curl -O http://1002genomes.u-strasbg.fr/files/1011Assemblies.tar.gz
# #!tar xzf 1011Assemblies.tar.gz
# Small development set
# !curl -OL https://www.dropbox.com/s/f42tiygq9tr1545/medium_setGENOMES_ASSEMBLED.tar.gz
# !tar xzf medium_setGENOMES_ASSEMBLED.tar.gz
# Tiny development set
# #!curl -OL https://www.dropbox.com/s/txufq2jflkgip82/tiny_setGENOMES_ASSEMBLED.tar.gz
# #!tar xzf tiny_setGENOMES_ASSEMBLED.tar.gz
# #!mv tiny_setGENOMES_ASSEMBLED GENOMES_ASSEMBLED
# -
# Before process the list of all of them, fix one that has an file name mismatch with what the description lines have.
# Specifically, the assembly file name is `CDH.re.fa`, but the FASTA-entries inside begin `CDH-3`.
# Simple file name mismatch. So next cell will change that file name to match.
#
# (I have been adding this particular file into the unpacked 'GENOMES_ASSEMBLED' directory from the 'SMALL' set via drag and drop from my local computer at this point in order to test the next cell. GIVE IT A MINUTE TO UPLOAD BECAUSE IF YOU RUN THE NEXT CELL TOO SOON, IT WILL ONLY MOVE WHAT IS UPLOADED AND THEN THERE WILL STILL BE A FILE WITH ORIGINAL NAME AS REST UPLOADS.)
import os
import sys
file_with_issues = "CDH.re.fa"
if os.path.isfile("GENOMES_ASSEMBLED/"+file_with_issues):
sys.stderr.write("\nFile with name non-matching entries ('{}') observed and"
" fixed.".format(file_with_issues))
# !mv GENOMES_ASSEMBLED/CDH.re.fa GENOMES_ASSEMBLED/CDH_3.re.fa
#pause and then check if file with original name is there still because
# it means this was attempted too soon and need to start over.
import time
time.sleep(12) #12 seconds
if os.path.isfile("GENOMES_ASSEMBLED/"+file_with_issues):
sys.stderr.write("\n***PROBLEM. TRIED THIS CELL BEFORE FINISHED UPLOADING.\n"
"DELETE FILES ASSOCIATED AND START ALL OVER AGAIN WITH UPLOAD STEP***.")
else:
sys.stderr.write("\nFile '{}' not seen and so nothing done"
". OKAY??".format(file_with_issues))
# Make a list of all `genome.fa` files, excluding `genome.fa.nhr` and `genome.fa.nin` and `genome.fansq`
# The excluding was only necessary because I had run some BLAST queries preiminarily in development. Normally,
# it would just be the `.re.fa` at the outset. (But keeping because removal ability could be useful.)
fn_to_check = "genome.fa"
genomes = []
import os
import fnmatch
for file in os.listdir('.'):
if fnmatch.fnmatch(file, '*'+fn_to_check):
if not file.endswith(".nhr") and not file.endswith(".nin") and not file.endswith(".nsq"):
genomes.append(file)
for file in os.listdir('GENOMES_ASSEMBLED'):
if fnmatch.fnmatch(file, '*'+".re.fa"):
if not file.endswith(".nhr") and not file.endswith(".nin") and not file.endswith(".nsq"):
# plus skip hidden files
if not file.startswith("._"):
genomes.append("GENOMES_ASSEMBLED/"+file)
len(genomes)
#
# Now you are prepared to analyze each genome.
# ## Assessing the genomes in regards to stretches of N base calls
#
# ### Total number of Ns present
# +
from pyfaidx import Fasta
query = "N"
#query = "X"
#query = "A" # CONTROL. should occur!
genomes_with_ambiguous = [] # This to be used for reporting when amount to analyze starts
# to get large
cut_off_for_printing_status_each = 100
for g in genomes:
print(g)
chrs = Fasta(g)
ambiguous = []
for x in chrs:
#print(x.name)
if query in str(x):
ambiguous.append(x.name)
if ambiguous:
genomes_with_ambiguous.append(g)
#print(ambiguous)
if len(genomes) < cut_off_for_printing_status_each:
if not ambiguous:
print ("No ambiguous nucleotides or gaps in {}.".format(g))
else:
print("There are occurences of '{}' in {}.".format(query,g))
if genomes_with_ambiguous and (len(genomes) >= cut_off_for_printing_status_each):
print("There are occurences of '{}' in {} genomes.".format(query,len(genomes_with_ambiguous)))
if len(genomes_with_ambiguous) < 250:
print("The following are the ones with '{}':\n{}.".format(query,genomes_with_ambiguous))
else:
print("That is a lot, and so they won't be listed here.\nAnalyses below will reveal more specifics.")
else:
print ("No ambiguous nucleotides or gaps were found in any the genomes.")
# -
# Another way to assess, just count all the letters present:
# %%time
from pyfaidx import Fasta
import pandas as pd
import collections
nt_counts = {}
for g in genomes:
if ".genome.fa" in g:
strain_id = g.split(".genome.fa")[0]
else:
strain_id = g.split(".re.fa")[0][18:]
concatenated_seqs = ""
chrs = Fasta(g)
for x in chrs:
#print(x.name)
concatenated_seqs += str(x)
nt_counts[strain_id] = collections.Counter(concatenated_seqs)
nt_count_df = pd.DataFrame.from_dict(nt_counts, orient='index').fillna(0)
nt_count_df["Total_nts"] = nt_count_df.sum(1)
def percent_calc(items):
'''
takes a list of two items and calculates percentage of first item
within total (second item)
'''
return items[0]/items[1]
nt_count_df['% N'] = nt_count_df[['N','Total_nts']].apply(percent_calc, axis=1)
nt_count_df = nt_count_df.sort_values(['% N', 'Total_nts'],ascending=[0,0])
nt_count_df_styled = nt_count_df.style.format({'Total_nts':'{:.2E}','% N':'{:.2%}'})
nt_count_df.to_csv('PB_n_1011_collection_nt_count_small.tsv', sep='\t',index = True)
nt_count_df.to_pickle("PB_n_1011_collection_nt_count_small.pkl")
nt_count_df_styled
# In development, I was suprised at first to see so letters that weren't 'A','G','T','C','N', or lowercase version of those. However, the extras match up with ones identified in [PatMatch documentation](https://www.yeastgenome.org/nph-patmatch) for pattern syntax for nucleotides. And so I assume they refer to that here. But I was still wondering why are a handfull are lowercase?
#
# As an example, this code below shows displaying an example where the `t` lowercase is seen.
#
# ```
# jovyan@jupyter-fomightez-2dcl-5fsq-5fdemo-2dbinder-2d24azsfxa:~/notebooks/GSD$ cat GENOMES_ASSEMBLED/CEF_4.re.fa|grep "t"
# TTCGTARGAYAGACTCWTTCCCGTGtAAATRTTTGTGACAGYTACGTCTATTTTCTACTM
# AYAGACTCWTTCCCGTGtAAATRTTTGTGACAGYTACGTCTATTTTCTACTMKATRTTTA
# CCCCGAAAGGAGAAATATAatATATATATATATAATATGCATCCTTATTATAATATTATT
# AATTAAATTAAATTAAATTAAATTAAATTAAATTAaattaGATGTTCATTAAATAAAAAT
# jovyan@jupyter-fomightez-2dcl-5fsq-5fdemo-2dbinder-2d24azsfxa:~/notebooks/GSD$
# ```
#
# Why lowercase?
# Seeing them in place doesn't spark many ideas as to why there'd be a few cases, except those that seem along with other lowercases letters in a block, such as 'aatta'. Sometimes lowercase is used for those where identity might be questionable because support limiting or stretch in which it occurs offers low information content.
# The small nature of the block of lowercase though suggests maybe just comes from quality scores from the Illumina reads?
# ### Examining the number and size of stretches of Ns present
# Make a dataframe of the data and also make a summary one with the say, top five(?) number instances, for each. (Idea not yet implemented, yet.)
# +
# %%time
# count frequency of blocks of Ns in the genomes
import re
from collections import defaultdict
from pyfaidx import Fasta
import pandas as pd
import collections
min_number_Ns_in_row_to_collect = 3
pattern_obj = re.compile("N{{{},}}".format(min_number_Ns_in_row_to_collect), re.I) # adpated from
# code worked out in `collapse_large_unknown_blocks_in_DNA_sequence.py`, which relied heavily on
# https://stackoverflow.com/a/250306/8508004
len_match_dict_by_strain = {}
#genomes = ["N44.genome.fa"]
for g in genomes:
len_match_dict = defaultdict(int)
if ".genome.fa" in g:
strain_id = g.split(".genome.fa")[0]
else:
strain_id = g.split(".re.fa")[0][18:]
records = Fasta(g)
for record in records:
for m in pattern_obj.finditer(str(record)):
len_match_dict[len(m.group())] += 1
#before passing final dictionary for strain to
#collection, add an entry of zero for size of 1 so the
#strain with no stretches will be in final dataframe
if not len_match_dict:
len_match_dict[1]=0
len_match_dict_by_strain[strain_id] = len_match_dict
#stretches_size_freq_df = pd.DataFrame.from_dict(len_match_dict_by_strain, orient='index').fillna(0)
#stretches_size_freq_df = pd.DataFrame.from_dict(len_match_dict_by_strain).fillna(0).stack().reset_index() # if wanted all stretches of size to show count even if none present
stretches_size_freq_df = pd.DataFrame.from_dict(len_match_dict_by_strain).stack().reset_index()
stretches_size_freq_df.columns = ['stretch_size','strain','#_instances']
stretches_size_freq_df = stretches_size_freq_df[['strain','stretch_size','#_instances']]
stretches_size_freq_df = stretches_size_freq_df.sort_values(['stretch_size','#_instances'],ascending=[0,0]).reset_index(drop=True)
# -
# Because of use of `%%time` in above cell, easiest to display resulting dataframe using next cell.
stretches_size_freq_df
stretches_size_freq_df = stretches_size_freq_df.drop(stretches_size_freq_df[(stretches_size_freq_df["stretch_size"] > 17000) | (stretches_size_freq_df["#_instances"] == 0.0)].index)
#df = df.drop(df[(df.score < 50) & (df.score > 20)].index)
stretches_size_freq_df
# Than can be plotted for visualization.
#
#
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
# Visualization
import seaborn as sns
sns.set() # from https://seaborn.pydata.org/examples/scatterplot_sizes.html; seems to restore style if it gets altered
p = sns.scatterplot(x="stretch_size", y="#_instances", hue="strain",data=stretches_size_freq_df)
# Set the scale of the x-and y-axes
p.set(xscale="log", yscale="log");
p.get_legend().set_visible(False);
# -
# Starting to go torwards the representation seen [here in Figure 1](https://www.researchgate.net/publication/12094921_Computational_comparison_of_two_draft_sequences_of_the_human_genome). For example, you can see the banding caused by using the log scale for 'number of instances' axis. However, as there aren't that many and that wide of a range of stretches of N among the 1011 collection (at least with out what I have seen in development), I don't think this path is going to yield a nice visual for the 1011 *cerevisiae genomes* collection.
# ### Examining the number and size of stretches of Ns present for the 1011 collection (Different tack)
#
# Because of the extremes of the PacBio and the reference sequence, even if just concern yourself, with 'stetch_size' the data don't visualize well with those from the 1011 collection. That can be summarized best with a plot by running the next cell. (Code reduced to minimum but fleshe out in this section.)
#
#
# %matplotlib inline
import seaborn as sns
sns.set()
# collect blocks of Ns in the genomes
import re
from collections import defaultdict
from pyfaidx import Fasta
import pandas as pd
import collections
min_number_Ns_in_row_to_collect = 3
pattern_obj = re.compile("N{{{},}}".format(min_number_Ns_in_row_to_collect), re.I)
matches = []
for g in genomes:
len_match_dict = defaultdict(int)
if ".genome.fa" in g:
strain_id = g.split(".genome.fa")[0]
else:
strain_id = g.split(".re.fa")[0][18:]
records = Fasta(g)
for record in records:
for m in pattern_obj.finditer(str(record)):
matches.append((strain_id,len(m.group())))
matches.append(("SGD_ref",0))
labels = ['strain', 'stretch_size']
stretch_df = pd.DataFrame.from_records(matches, columns=labels)
sns.distplot(stretch_df["stretch_size"], kde=False, rug=True);
# See how most stuff is squashed all on low end relative the PacBio sequences at around 17500.
#
# For visualization of the 1011 collection data adequately, probably best to separate out PacBio and SGD references and start treating strains as category and let Seaborn handle aggregating the data, as it does this so well. So we need a simpler dataframe than one in above section, one for each instance of a stretch of Ns found.
# +
# %%time
# collect blocks of Ns in the genomes
import re
from collections import defaultdict
from pyfaidx import Fasta
import pandas as pd
import collections
min_number_Ns_in_row_to_collect = 3
pattern_obj = re.compile("N{{{},}}".format(min_number_Ns_in_row_to_collect), re.I) # adpated from
# code worked out in `collapse_large_unknown_blocks_in_DNA_sequence.py`, which relied heavily on
# https://stackoverflow.com/a/250306/8508004
matches = []
#genomes = ["N44.genome.fa"]
for g in genomes:
len_match_dict = defaultdict(int)
if ".genome.fa" in g:
strain_id = g.split(".genome.fa")[0]
else:
strain_id = g.split(".re.fa")[0][18:]
records = Fasta(g)
for record in records:
for m in pattern_obj.finditer(str(record)):
matches.append((strain_id,len(m.group())))
#before using list of matches to make a dataframe, add
# entry for reference sequence that has no actual instancesto
#collection, add an entry of zero for size of 1 so the
#strain with no stretches will be in final dataframe
matches.append(("SGD_ref",0))
#make a dataframe from the list
labels = ['strain', 'stretch_size']
stretch_df = pd.DataFrame.from_records(matches, columns=labels)
# Note that later, when trying to compare the distribution to the entire sequence set to what I see
# from just the collected mitochondrial genomes sequences, I realized I had not included a sort for the `stretch_df`
# and when there is a lot of data they come out ranging all over the place. The next command adds that.
stretch_df = stretch_df.sort_values(['stretch_size'],ascending=[0]).reset_index(drop=True)
# -
len(stretch_df)
stretch_df.to_csv('PB_n_1011_collection_all_stretchesN_df.tsv', sep='\t',index = False)
stretch_df.to_pickle("PB_n_1011_collection_all_stretchesN_df.pkl")
# (If running on that whole set, store those files so you can read them in later, or elsewhere, as needed.)
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
# Visualization
import seaborn as sns
sns.set()
p = sns.stripplot(x="stretch_size", y="strain", data=stretch_df);
# Set the scale of the x-and y-axes
p.set(xscale="log");
#p.get_legend().set_visible(False);
#p = sns.swarmplot(x="stretch_size", y="strain", data=df);
# -
# With about 60 it is too crowded to read, but I haven't tried adjusting height yet because hoping other methods below better suited. Although this illustrates the distribution of N stretches in the 1011 collection data more convincingly than the same code when there was only two from the 1011 collection.
#
# But if you try swarmplot, you'll see it isn't showing the distibution well for those in the 1011 collection as just with these few it is too many, it seems.
#
# DOES THIS NEXT CELL WORK WHEN THERE ARE 60 strains?!?!? Seemed to get hung up when I tried with the 'small set!?!? Fluke?
# Commented pit block-styple all the swarmplots just below because seemed to take forever to run and in fact gave up before any completed. It seems something is happening because some image is made when stop and so would be good to explore more if curious.
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
'''
# Visualization
import seaborn as sns
sns.set()
p = sns.swarmplot(x="stretch_size", y="strain", data=stretch_df);
# Set the scale of the x-and y-axes
p.set(xscale="log");
#p.get_legend().set_visible(False);
#p = sns.swarmplot(x="stretch_size", y="strain", data=df);
'''
# -
# When there was only two in the tiny set, could see that better by limiting to those from the 1011 collection. Possible here?
stretch_df = stretch_df.drop(stretch_df[(stretch_df["stretch_size"] > 17000) | (stretch_df["strain"] == "SGD_ref")].index)
# Later by alternatively running the code above instead of below & counting length of df, I confirmed that the above line doesn't remove any from the
# 1011 collection and that `(stretch_df["strain"] == "SGD_ref")` and `(stretch_df["stretch_size"] == 0)` result in the same set. <--Turned
# out not the case when dealing with just derived mito data. Went with more explicit command now and moved original to next line.
#stretch_df = stretch_df.drop(stretch_df[(stretch_df["stretch_size"] > 17000) | (stretch_df["stretch_size"] == 0)].index)
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
'''
# Visualization
import seaborn as sns
sns.set()
p = sns.swarmplot(x="stretch_size", y="strain", data=stretch_df);
# Set the scale of the x-and y-axes
p.set(xscale="log");
#p.get_legend().set_visible(False);
#p = sns.swarmplot(x="stretch_size", y="strain", data=df);
'''
# -
# Many more at 50 than can be handled so get artificats like discussed [here](https://stackoverflow.com/questions/50817607/variable-spacing-of-discrete-categories-in-seaborn-swarmplot). And it can be made better by increasing the height, although still seems to happen no matter how tall it is finally made as even the use of `plt.figure(figsize=(8,13))` causes curling.
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
import matplotlib.pyplot as plt
plt.figure(figsize=(8,13))
'''
# Visualization
import seaborn as sns
sns.set()
p = sns.swarmplot(x="stretch_size", y="strain", data=stretch_df);
# Set the scale of the x-and y-axes
p.set(xscale="log");
#p.get_legend().set_visible(False);
#p = sns.swarmplot(x="stretch_size", y="strain", data=df);
'''
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
# Visualization
import seaborn as sns
sns.set()
stretch_df = stretch_df.drop(stretch_df[(stretch_df["stretch_size"] > 17000) | (stretch_df["stretch_size"] == 0)].index)
sns.distplot(stretch_df["stretch_size"], kde=False, rug=True);
# -
# Nicer, but what pertains to which strain?
# +
# %matplotlib inline
# above line works for JupyterLab which I was developing in. Try `%matplotlib notebook` for when in classic.
# Visualization - based on https://towardsdatascience.com/histograms-and-density-plots-in-python-f6bda88f5ac0
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
#Limit to subset, if not set already
stretch_df = stretch_df.drop(stretch_df[(stretch_df["stretch_size"] > 17000) | (stretch_df["stretch_size"] == 0)].index)
# List of strains to plot
strains = set(stretch_df["strain"].tolist())
# Iterate through the strains
for strain in strains:
# Subset to the strain
subset = stretch_df[stretch_df['strain'] == strain]
# Draw the density plot
sns.distplot(subset['stretch_size'], hist = False, kde = True,# rug = True,
kde_kws = {'shade': True, 'linewidth': 3},
#rug_kws={'color': 'black'},
label = strain)
# Plot formatting
plt.legend(prop={'size': 16}, title = 'Strain')
plt.title('Density Plot with strains')
plt.xlabel('Stretch size (base)')
plt.ylabel('Density');
# -
# As discussed at the end of [here](https://towardsdatascience.com/histograms-and-density-plots-in-python-f6bda88f5ac0), uncomment the two portions dealing with `rug` to see the actual data points. I don't leave them on here bey default becuase many and cannot see which belong to which strain.
#
# While this was nice with very foew, it became difficult once I add a lot more in order to do anything with the 1011 collection or even a subset. Alternatives? [Raincloud plots](https://nbviewer.jupyter.org/github/pog87/PtitPrince/blob/master/RainCloud_Plot.ipynb) look promising for this situation.
# !pip install ptitprince
# +
import seaborn as sns
import matplotlib.pyplot as plt
#sns.set(style="darkgrid")
#sns.set(style="whitegrid")
#sns.set_style("white")
sns.set(style="whitegrid",font_scale=2)
#Limit to subset, if not set already
stretch_df = stretch_df.drop(stretch_df[(stretch_df["stretch_size"] > 17000) | (stretch_df["stretch_size"] == 0)].index)
#Want an image file of the figure saved?
savefigs = True
savefig_fn = 'n_stetch_distrbution_raincloud_small_subset.png'
f, ax = plt.subplots(figsize=(18, 46))
import ptitprince as pt
ort = "h"
ax=pt.half_violinplot(data=stretch_df, palette="Set2", bw=0.2, linewidth=1,cut=0.,scale="area", width=2.0, inner=None,orient=ort,x='stretch_size', y="strain")
ax=sns.stripplot(data=stretch_df, palette="Set2", edgecolor="white",size=2,orient=ort,x='stretch_size',y="strain",jitter=1,zorder=0)
ax=sns.boxplot(data=stretch_df, color="black",orient=ort,width=.15,x='stretch_size',y="strain",zorder=10,showcaps=True,boxprops={'facecolor':'none', "zorder":10},showfliers=True,whiskerprops={'linewidth':2, "zorder":10},saturation=1)
# Despite above code working, trying the 'shorter' version illustrated at https://nbviewer.jupyter.org/github/RainCloudPlots/RainCloudPlots/blob/master/tutorial_python/raincloud_tutorial_python.ipynb
# fails. Feel free to comment out the three `ax` lines just above and uncomment the next line to try yourself. Maybe cannot handle numbers? Because
# cause wasn't different version because the demo notebook using the single line method worked when run in same session.
#ax=pt.RainCloud(x='stretch_size', y="strain", data = stretch_df, palette = "Set2", bw = 0.2, width_viol = .6, figsize = (18,46), orient = ort)
# Finalize the figure
#ax.set(xlim=(0,1000)) #This can be used to 'zoom in', if you accept, i.e., note in the legend, that anything with a
# line continuing off means there are more points off the scale. Could indicating symbols be added like I did in
# plot expression plots?
#ax.set(ylim=(3.5, -.7)) # this can be used to actually plot fractions of them 'kind of'. `ax.set(ylim=(3.5, -.7))` results
# in plots of four when there were 48 from the 1011 collection, although they are spaced out.
sns.despine(left=True)
if savefigs:
f.savefig(savefig_fn, bbox_inches='tight')
# -
# A little small but it is the most insightful plot so far for the group of 48 from the 1011 collection. While the height set via `figsize` has to be high enough for sure and adjusted when there are more, once it reaches a certain point increasing height in `figsize` setting doesn't help. Turns out can get the violoinplot part to look taller by changing the `width` setting for the `half_violinplot` plot part. Demo [here](https://github.com/pog87/PtitPrince/blob/master/RainCloud_Plot.ipynb) had `width=.8`. With 47 genomes, I noted improvement setting that higher, currently trying `width=2.0`, with 48 genomes. The density curve for 'YAM' starts overlapping one above it if increases much beyond that. (Related note: `bw` setting seems to factor into the smoothed histogram curve because it makes it less acute, and more spread out, when increased from `.2` to `1.0`; referenced in relation to `sigma` [here](https://nbviewer.jupyter.org/github/RainCloudPlots/RainCloudPlots/blob/master/tutorial_python/raincloud_tutorial_python.ipynb). 'smoothed histogram' language based on [here](https://nbviewer.jupyter.org/github/RainCloudPlots/RainCloudPlots/blob/master/tutorial_python/raincloud_tutorial_python.ipynb).)
#
# I wish the individual strain plots would take up more of the lower space too, but this is probably a good compromise and at least it works with almost 48 samples.
# Actually, later, I found when limiting to only 1000 for the stretch size with this group, things looked better in spacing regard between them too. I used `ax.set(xlim=(0,1000))` with the knowledge that anything with a line continuing off means there are more points off the scale. That 'zoom in' view really looks nice, but I left the code defaulted to all points.
# Synopsis: Be sure to run with `ax.set(xlim=(0,1000))`!
# ----
# +
import time
def executeSomething():
#code here
print ('.')
time.sleep(480) #60 seconds times 8 minutes
while True:
executeSomething()
# -
| notebooks/GSD/GSD Assessing_ambiguous_nts_in_1011_collection_genomesSMALL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
f22_lidar= np.genfromtxt('/Users/arcturus/Documents/PhD/f22/python_zi/xaa_12mil_zi.xyz',
skip_header=1)
f22_kp= np.genfromtxt('/Users/arcturus/Documents/PhD/f22/python_zi/xaa_12milkeypoints.xyz')
min_x = min(f22_lidar[:,1])
min_y = min(f22_lidar[:,2])
# +
f22_s = f22_lidar[(f22_lidar[:,2] >= min_y + 35000) &
(f22_lidar[:,2] <= min_y + 50000), :]
f22_kp_s = f22_kp[(f22_kp[:,2] >= min_y + 35000) &
(f22_kp[:,2] <= min_y + 50000), :]
# +
#f22_s[1:10,:]
# +
cmap = plt.get_cmap('gist_earth_r')
cmap2 = plt.get_cmap('Oranges_r', 16)
cmap3 = plt.get_cmap('Greys', 16)
#cmap4 = plt.get_cmap('cubehelix', 16)
cmap4 = plt.get_cmap('gist_earth_r')
#plot_yticks = np.arange(min(f22_s[:,1]), max(f22_s[:,1])+200, 200)
plot_yticks = np.arange(400, 1400, 200)
fig = plt.figure(figsize=(22/2.51, 16/2.51))
ax0 = fig.add_subplot(4,1,1)
a0 = ax0.scatter(f22_s[:,2], f22_s[:,1], c=f22_s[:,3], cmap=cmap4, lw=0, s = 3,
vmin=0, vmax=3)
#plt.colorbar()
ax0.scatter(f22_kp_s[:,1], f22_kp_s[:,0], c=f22_kp_s[:,2], cmap=cmap2, lw=0, s = 4)
#ax0.set_xlim([-6100, 2700])
#ax0.set_ylim([-250, 250])
ax0.set_ylabel('Across swath (m)')
ax0.grid()
ax0.text(-0.06, 0.99,'A', ha='center', va='center', transform=ax0.transAxes, fontsize=12, weight='bold')
ax0.set_yticks(plot_yticks)
ax0.set_xticklabels([])
cbaxes0= fig.add_axes([0.99, 0.78, 0.02, 0.18])
cb0 = plt.colorbar(a0, cax=cbaxes0, cmap=cmap4)
cb0.set_ticks(np.arange(0,3.5,0.5))
cb0.set_ticklabels(np.arange(0,3.5,0.5))
ax1 = fig.add_subplot(4,1,2)
a1 = ax1.scatter(f22_s[:,2], f22_s[:,1], c=f22_s[:,4], cmap=cmap, lw=0, s = 3,
vmin=0, vmax=0.25)
#ax1.set_xlim([-6100, 2700])
#ax1.set_ylim([-250, 250])
ax1.set_ylabel('Across swath (m)')
ax1.grid()
ax1.text(-0.06, 0.99,'B', ha='center', va='center', transform=ax1.transAxes, fontsize=12, weight='bold')
ax1.set_yticks(plot_yticks)
ax1.set_xticklabels([])
cbaxes1= fig.add_axes([0.99, 0.55, 0.02, 0.18])
cb1 = plt.colorbar(a1, cax=cbaxes1, cmap=cmap)
cb1.set_ticks(np.arange(0,0.26,0.05))
cb1.set_ticklabels(np.arange(0,0.26,0.05))
ax2 = fig.add_subplot(4,1,3)
a2 = ax2.scatter(f22_s[:,2], f22_s[:,1], c=f22_s[:,5], cmap=cmap4, lw=0, s = 3,
vmin=0, vmax=10)
#ax2.set_xlim([-6100, 2700])
#ax2.set_ylim([-250, 250])
ax2.set_ylabel('Across swath (m)')
ax2.grid()
ax2.text(-0.06, 0.99,'C', ha='center', va='center', transform=ax2.transAxes, fontsize=12, weight='bold')
ax2.set_yticks(plot_yticks)
ax2.set_xticklabels([])
cbaxes2= fig.add_axes([0.99, 0.32, 0.02, 0.18])
cb2 = plt.colorbar(a2, cax=cbaxes2, cmap=cmap4)
cb2.set_ticks(np.arange(0,12,2))
cb2.set_ticklabels(np.arange(0,12,2))
ax3 = fig.add_subplot(4,1,4)
a3 = ax3.scatter(f22_s[:,2], f22_s[:,1], c=f22_s[:,6], cmap=cmap, lw=0, s=3, vmin=0, vmax=5)
#ax3.set_xlim([-6100, 2700])
#ax3.set_ylim([-250, 250])
ax3.set_xlabel('Distance along swath (m)')
ax3.set_ylabel('Across swath (m)')
ax3.set_yticks(plot_yticks)
ax3.grid()
ax3.text(-0.06, 0.99,'D', ha='center', va='center', transform=ax3.transAxes, fontsize=12, weight='bold')
cbaxes3= fig.add_axes([0.99, 0.09, 0.02, 0.18])
cb3 = plt.colorbar(a3, cax=cbaxes3, cmap=cmap)
cb3.set_ticks(np.arange(0,6,1))
cb3.set_ticklabels(np.arange(0,6,1))
plt.tight_layout()
plt.savefig('f22_lidar_map.png', bbox_inches='tight', dpi=300)
# -
print('total freeboard from this LiDAR segment: {0}, std: {1}'.format(np.mean(f22_s[:,3]), np.std(f22_s[:,3])))
print('total freeboard range this LiDAR segment: {0}, max: {1}'.format(np.min(f22_s[:,3]), np.max(f22_s[:,3])))
print('uncertainty surrounding total freeboard from this LiDAR segment: {0} to {1}'.format(np.min(f22_s[:,4]), np.max(f22_s[:,4])))
print('ice thickness from this LiDAR segment: {0}, std: {1}'.format(np.mean(f22_s[:,5]), np.std(f22_s[:,5])))
print('ice thickness range this LiDAR segment: {0}, max: {1}'.format(np.min(f22_s[:,5]), np.max(f22_s[:,5])))
print('uncertainty surrounding ice thickness from this LiDAR segment: {0} to {1}'.format(np.min(f22_s[:,6]), np.max(f22_s[:,6])))
print('snow depth from this LiDAR segment: {0}, std: {1}'.format(np.mean(f22_s[:,11]), np.std(f22_s[:,11])))
print('snow depth range this LiDAR segment: {0}, max: {1}'.format(np.min(f22_s[:,11]), np.max(f22_s[:,11])))
print('uncertainty surrounding snow depth from this LiDAR segment: {0} to {1}'.format(np.min(f22_s[:,12]), np.max(f22_s[:,12])))
# +
plt.scatter(f22_s[:,2], f22_s[:,1], f22_s[:,6], lw=0, vmin=0, vmax=5)
#ax3.set_xlim([-6100, 2700])
# -
| f22 chunks map and basic stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import Libraries
import csv
import json
import os
import glob
# ## Read Config File
import configparser
config = configparser.ConfigParser()
config.read('config.ini')
csvFolderPath = config['DEFAULT']['CSV-Folder-Path']
contain_string = config['DEFAULT']['Contain-String']
jsonFolderPath = config['DEFAULT']['JSON-Folder-Path']
# ## Read CSV File into Array & Write into JSON Array
for file in glob.glob("{}/*{}*.csv".format(csvFolderPath, contain_string)):
file_basename = os.path.basename(file)
filename = os.path.splitext(file_basename)[0]
# Read CSV File into Array
arr = []
with open (file) as csvFile:
csvReader = csv.DictReader(csvFile)
for csvRow in csvReader:
arr.append(csvRow)
# Write into JSON Array
jsonFilePath = "{}/{}.json".format(jsonFolderPath,filename)
with open(jsonFilePath, "w") as jsonFile:
jsonFile.write(json.dumps(arr,indent=4))
| f0017/f0017.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
from pymongo import MongoClient
# +
myclient= MongoClient("mongodb+srv://Derek:<EMAIL>/API?retryWrites=true&w=majority")
db = myclient["API"]
with open('semD.json', 'r') as s1:
semD = json.load(s1)
with open('comList.json', 'r') as s2:
comList = json.load(s2)
with open('lessonList.json','r') as s3:
lessonList = json.load(s3)
Collection = db["semD"]
Collection1 = db["comList"]
#Collection2 = db['lessonList']
#Collection3 = db['recList']
# -
Collection.insert_many(semD)
Collection1.insert_many(comList)
#Collection2.insert_many(lessonList)
semD
| Orbital data/Data manipulation/2020-2021 sem 1/import.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import os.path as p
import pandas as pd
from PIL import Image
from tqdm import tqdm_notebook as tqdm
import os
from joblib import Parallel, delayed
import cv2
# ### COCO Format
# 
# COCO Object Detection Format
# 
# ## Open Images
OUT_DIR = '/APL/Datasets/open-images-challenge-2019-COCO-format'
os.makedirs(OUT_DIR, exist_ok=True)
OPEN_IMG = p.join('/APL/Datasets/open-images-challenge-2019')
# #### Read original CSV data
SET = 'train'
#SET = 'validation'
#SET = 'test'
# Boxes
bbox_annotation = pd.read_csv(p.join(OPEN_IMG, 'challenge-2019-' + SET + '-detection-bbox-expanded.csv'))
bbox_annotation
# Define basic JSON data structure
new_data_format = {'info': {'description': 'Open Images Challenge Dataset',
'url': 'https://storage.googleapis.com/openimages/web/index.html',
'version': '1.0',
'year': 2019,
'contributor': 'Open Image authors, this format: <NAME> @McGill University',
'date_created': '2019/07/03'},
'images': [],
'annotations': [],
'categories': []}
# Get all Image IDs
unique_image_ids = bbox_annotation['ImageID'].unique()
unique_image_ids
# +
# Create image str ID to int ID mapping
im_id_mapping = []
i = 0
for im in tqdm(unique_image_ids):
im_id_mapping.append((im,i))
i += 1
# -
im_id_mapping
# Write mapping for image ID (string) to int to disk
df = pd.DataFrame(data=im_id_mapping, columns=['ImageID','IntIndex'])
df.to_csv(os.path.join(OPEN_IMG, 'challenge-2019-' + SET + '-image-ID-mapping.csv'), index=False)
# Get all image sizes
results = []
for originalID,newID in tqdm(im_id_mapping):
im_filename = originalID + '.jpg'
with Image.open(p.join(OPEN_IMG, SET, im_filename)) as image:
im_size = image.size
results.append({'file_name': im_filename,
'height': im_size[1],
'width': im_size[0],
'id': newID})
# Make sure above is done
print(len(results))
print(len(im_id_mapping))
# List of images is done -> save
new_data_format['images'] = results
# ## Class mapping stuff
# + active=""
# # Create list of images with width + height and filename
#
# def create_im_entry(i, v):
# im_filename = i + '.jpg'
#
# with Image.open(p.join(OPEN_IMG, SET, im_filename)) as image:
# im_size = image.size
#
# return {'file_name': im_filename,
# 'height': im_size[1],
# 'width': im_size[0],
# 'id': v}
#
# results = Parallel()(delayed(create_im_entry)(i,v) for i,v in tqdm(im_id_mapping.items()))
# -
#Get category names
category_csv = pd.read_csv(os.path.join(OPEN_IMG,'challenge-2019-classes-description-500.csv'),header=None,names=['MID','Description'])
category_csv
# Get label hierarchy (not used)
with open(os.path.join(OPEN_IMG, 'challenge-2019-label500-hierarchy.json')) as f:
label_hierarchy = json.load(f)
label_hierarchy
# Create mapping from name to MID
category_list = []
category_MID_mapping = []
i = 0
for row in category_csv.itertuples():
# Supercategory = None because we already processed everything using the OID script
category_list.append({'id': i, 'name': row.Description, 'supercategory': None})
# Save mapping from original name to new ID
category_MID_mapping.append((row.MID, i))
i += 1
# Save mapping to disk
MID_mapping_df = pd.DataFrame(data=category_MID_mapping, columns=['MID','IntIndex'])
MID_mapping_df.to_csv(os.path.join(OPEN_IMG, 'challenge-2019-category-ID-mapping.csv'), index=False)
# Save category data
new_data_format['categories'] = category_list
category_MID_mapping_dict = {}
for i in category_MID_mapping:
category_MID_mapping_dict[i[0]] = i[1]
# ## End category stuff
# Create dictinary mapping from Open Images string ID to image info
im_info_map = {p.splitext(i['file_name'])[0]:i for i in new_data_format['images']}
im_info_map
for r in bbox_annotation.itertuples():
print(r)
break
# load line profiler
# %load_ext line_profiler
# %lprun -f create_annotations create_annotations(bbox_annotation,im_info_map,MID_mapping_df)
# Create annotations
# def create_annotations(bbox_annotation,im_info_map,MID_mapping_df):
def create_annotations():
# iter_count = 0
list_annot = []
for r in tqdm(bbox_annotation.itertuples(), total=len(bbox_annotation)):
im_info = im_info_map[r.ImageID]
im_w = im_info['width']
im_h = im_info['height']
x = r.XMin * im_w
w = (r.XMax - r.XMin) * im_w
y = r.YMin * im_h
h = (r.YMax - r.YMin) * im_h
#Get real category name (iloc = 0 is first match)
category_real_name = category_MID_mapping_dict[r.LabelName]
list_annot.append({'id': r.Index,
'image_id': im_info['id'],
'category_id': category_real_name,
'bbox': [x, y, w, h],
'segmentation': [[x, y, x, y + h, x + w, y + h, x + w, y]],
'area': w * h,
'iscrowd': 0}) # Set to 0 because otherwise it gets ignored
# iter_count += 1
# if iter_count == 5000:
# return list_annot
return list_annot
list_annot = create_annotations()
new_data_format['annotations'] = list_annot
new_data_format
OUTPUT_JSON = p.join(OUT_DIR, SET + '.json')
#Output to file
with open(OUTPUT_JSON, 'w') as f:
json.dump(new_data_format,f)
# +
## Fix for is group of
JSON_FILE_FIX='/APL/Datasets/open-images-challenge-2019-COCO-format/validation.json'
with open(JSON_FILE_FIX) as f:
dt = json.load(f)
annotations = dt['annotations']
# -
new_annot = []
for i in tqdm(annotations):
if 'iscrowd' in i:
i['iscrowd'] = 0
new_annot.append(i)
dt['annotations'] = new_annot
with open(JSON_FILE_FIX, 'w') as f:
json.dump(dt,f)
| Notebooks/convert_coco_format_mike_open_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
# ## Does nn.Conv2d init work well?
# [Jump_to lesson 9 video](https://course.fast.ai/videos/?lesson=9&t=21)
# +
#export
from exp.nb_02 import *
def get_data():
path = Config().data_path()/'mnist'
with gzip.open(path/'mnist.pkl.gz', 'rb') as f:
((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding='latin-1')
return map(tensor, (x_train,y_train,x_valid,y_valid))
def normalize(x, m, s): return (x-m)/s
# -
??torch.nn.modules.conv._ConvNd.reset_parameters
x_train,y_train,x_valid,y_valid = get_data()
train_mean,train_std = x_train.mean(),x_train.std()
x_train = normalize(x_train, train_mean, train_std)
x_valid = normalize(x_valid, train_mean, train_std)
x_train = x_train.view(-1,1,28,28)
x_valid = x_valid.view(-1,1,28,28)
x_train.shape,x_valid.shape
n,*_ = x_train.shape
c = y_train.max()+1
nh = 32
n,c
l1 = nn.Conv2d(1, nh, 5)
x = x_valid[:100]
x.shape
def stats(x): return x.mean(),x.std()
l1.weight.shape
stats(l1.weight)
stats(l1.bias)
t = l1(x)
stats(t)
init.kaiming_normal_(l1.weight, a=1.)
stats(l1(x))
import torch.nn.functional as F
def f1(x,a=0): return F.leaky_relu(l1(x),a)
init.kaiming_normal_(l1.weight, a=0)
stats(f1(x))
init.kaiming_normal_(l1.weight, a=0.01)
stats(f1(x))
l1 = nn.Conv2d(1, nh, 5)
stats(f1(x))
l1.weight.shape
# receptive field size
rec_fs = l1.weight[0,0].numel()
rec_fs
nf,ni,*_ = l1.weight.shape
nf,ni
fan_in = ni*rec_fs
fan_out = nf*rec_fs
fan_in,fan_out
def gain(a): return math.sqrt(2.0 / (1 + a**2))
gain(1),gain(0),gain(0.01),gain(0.1),gain(math.sqrt(5.))
torch.zeros(10000).uniform_(-1,1).std()
1/math.sqrt(3.)
def kaiming2(x,a, use_fan_out=False):
nf,ni,*_ = x.shape
rec_fs = x[0,0].shape.numel()
fan = nf*rec_fs if use_fan_out else ni*rec_fs
std = gain(a) / math.sqrt(fan)
bound = math.sqrt(3.) * std
x.data.uniform_(-bound,bound)
kaiming2(l1.weight, a=0);
stats(f1(x))
kaiming2(l1.weight, a=math.sqrt(5.))
stats(f1(x))
class Flatten(nn.Module):
def forward(self,x): return x.view(-1)
m = nn.Sequential(
nn.Conv2d(1,8, 5,stride=2,padding=2), nn.ReLU(),
nn.Conv2d(8,16,3,stride=2,padding=1), nn.ReLU(),
nn.Conv2d(16,32,3,stride=2,padding=1), nn.ReLU(),
nn.Conv2d(32,1,3,stride=2,padding=1),
nn.AdaptiveAvgPool2d(1),
Flatten(),
)
y = y_valid[:100].float()
t = m(x)
stats(t)
l = mse(t,y)
l.backward()
stats(m[0].weight.grad)
# +
# init.kaiming_uniform_??
# -
for l in m:
if isinstance(l,nn.Conv2d):
init.kaiming_uniform_(l.weight)
l.bias.data.zero_()
t = m(x)
stats(t)
l = mse(t,y)
l.backward()
stats(m[0].weight.grad)
# ## Export
# !./notebook2script.py 02a_why_sqrt5.ipynb
| nbs/dl2/02a_why_sqrt5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
# %matplotlib inline
import os
import numpy as np
# -
# First open all series (SC, LC and WN) and plot their autocorrelation
ISIseq_path = "/Users/nicolasfarrugia/Dropbox/Manip/Individual_ISI_series/series_manip_512stim/series txt/"
allfiles = os.listdir(ISIseq_path)
# To make sure the files are taken for the right train
allfiles = sorted(allfiles)
# +
SC_files=allfiles[0:75:3]
print("All SC files : ")
print(SC_files)
print("All LC files : ")
LC_files=allfiles[1:75:3]
print(LC_files)
print("All WN files : ")
WN_files=allfiles[2:75:3]
print(WN_files)
# -
SC_series = [np.recfromtxt(os.path.join(ISIseq_path,X)) for X in SC_files]
LC_series = [np.recfromtxt(os.path.join(ISIseq_path,X)) for X in LC_files]
WN_series = [np.recfromtxt(os.path.join(ISIseq_path,X)) for X in WN_files]
# Plot just a few subjects
# +
from pandas.tools.plotting import autocorrelation_plot
nbsubjtoplot = 25
for cur_sc,cur_lc,cur_wn,subjnum in zip(SC_series[0:nbsubjtoplot],
LC_series[0:nbsubjtoplot],
WN_series[0:nbsubjtoplot],
SC_files[0:nbsubjtoplot]):
subjid = subjnum[0:2]
plt.figure(figsize=(10,5))
ax1=plt.subplot(1,3,1)
autocorrelation_plot(cur_sc,ax1)
plt.xlim([0,15])
plt.ylim([-1,1])
plt.title("SC subject %s" % subjid)
ax2=plt.subplot(1,3,2)
autocorrelation_plot(cur_lc,ax2)
plt.xlim([0,15])
plt.ylim([-1,1])
plt.title("LC subject %s" % subjid)
ax3=plt.subplot(1,3,3)
autocorrelation_plot(cur_wn,ax3)
plt.xlim([0,15])
plt.ylim([-1,1])
plt.title("WN subject %s" % subjid)
plt.tight_layout()
plt.show()
# -
# Neural Networks using [KERAS](www.keras.io)
#
# +
from keras.models import Sequential
from keras.layers import LSTM, Dense
import keras.backend as K
from keras import metrics
import numpy as np
data_dim = 1
timesteps = None
hidden_dim = 128
# expected input data shape: (batch_size, timesteps, data_dim)
model = Sequential()
model.add(LSTM(hidden_dim, return_sequences=True,
input_shape=(timesteps, data_dim))) # returns a sequence of vectors of dimension 32
model.add(LSTM(hidden_dim, return_sequences=True)) # returns a sequence of vectors of dimension 32
model.add(LSTM(hidden_dim, return_sequences=True))
model.add(LSTM(1,return_sequences=True))
# model.add(Dense(1, activation='relu'))
def mean_pred(y_true, y_pred):
return K.mean(y_pred)
model.compile(loss='mean_squared_error',
optimizer='rmsprop',
metrics=[metrics.mae])
model.summary()
# +
def norm(v):
return((v-min(v))/(max(v)-min(v)))
def norm_batch(b):
return np.array([norm(v) for v in b])
# +
X_train = np.vstack(SC_series)
Y_train = norm_batch(X_train[:22,1:])
Y_val = norm_batch(X_train[22:,1:])
X_val = norm_batch(X_train[22:,:511])
X_train = norm_batch(X_train[:22,:511])
print(X_train[0][:10])
print(X_train.shape,Y_train.shape,X_val.shape,Y_val.shape)
# -
model.fit(X_train.reshape((22,511,1)), Y_train.reshape((22, 511,1)),
batch_size=8, epochs=25)
# +
test = model.evaluate(X_val.reshape(3,511,1), Y_val.reshape(3, 511, 1))
print("Accuracy: ", test[1])
pred = model.predict(X_val.reshape(3,511,1))
print("Prediction: ", pred[0, -6:, 0])
print("Solution: ", Y_val[0, -6:])
print("Distance: ", sum([abs(x-y) for x, y in zip(pred[0, 1:, 0], Y_val[0, 1:])])/len(Y_val[0, 1:]))
# -
# New model using sequence-to-sequence autoencoders
# +
from keras.layers import Input, LSTM, RepeatVector
from keras.models import Model
from keras import regularizers
timesteps = 512
input_dim = 1
latent_dim = 50
inputs = Input(shape=(timesteps, input_dim))
encoded = LSTM(latent_dim,kernel_regularizer=regularizers.l2(0.01),
activity_regularizer=regularizers.l1(0.01))(inputs)
decoded = RepeatVector(timesteps)(encoded)
decoded = LSTM(input_dim, return_sequences=True)(decoded)
sequence_autoencoder = Model(inputs, decoded)
#Create the encoder model
encoder = Model(inputs, encoded)
# create the decoder model
#decoder = Model(encoded, decoded)
# -
sequence_autoencoder.compile(loss='cosine_proximity',
optimizer='RMSprop',lr = 0.01)
# +
X_train = (np.vstack(SC_series[0:20]).reshape((20,512,1)))
X_test = (np.vstack(SC_series[20:]).reshape((5,512,1)))
print(X_train.shape,X_test.shape)
sequence_autoencoder.fit(X_train,X_train,epochs=10,validation_data=(X_test,X_test))
# -
# encode and decode some digits
# note that we take them from the *test* set
#encoded_seqs = encoder.predict(X_test)
decoded_seqs = sequence_autoencoder.predict(X_test)
plt.figure()
ax1=plt.subplot(1,2,1)
autocorrelation_plot(decoded_seqs[0],ax1)
plt.xlim([0,15])
ax2 = plt.subplot(1,2,2)
autocorrelation_plot(X_test[0],ax2)
plt.xlim([0,15])
X_test[0]
| Tests_SC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
from matplotlib import pyplot as plt
# +
#Created by <NAME> On Jupyter Notebook
# -
def pixelVal(pix, r1, s1, r2, s2):
if (0 <= pix and pix <= r1):
return (s1 / r1)*pix
elif (r1 < pix and pix <= r2):
return ((s2 - s1)/(r2 - r1)) * (pix - r1) + s1
else:
return ((255 - s2)/(255 - r2)) * (pix - r2) + s2
img = cv2.imread('test2.jpg',0)
r1 = 70
s1 = 0
r2=140
s2 = 255
pixelVal_vec = np.vectorize(pixelVal)
contrast_stretched = pixelVal_vec(img, r1, s1, r2, s2)
equ=np.hstack((img,contrast_stretched))
plt.title("Original/Contrast Stretched")
plt.imshow(equ,'gray')
plt.show()
# +
histr = cv2.calcHist([img],[0],None,[256],[0,256])
# show the plotting graph of an image
plt.plot(histr)
plt.show()
# -
stretch_near = cv2.resize(img, (3000,3000),
interpolation = cv2.INTER_NEAREST)
plt.title('original')
plt.imshow(img,'gray')
plt.show()
plt.title('Zoomed Image')
plt.imshow(stretch_near,'gray')
plt.show()
img = cv2.imread('test1.jpg',0)
equ = cv2.equalizeHist(img)
res = np.hstack((img, equ))
plt.title("original/enhaced stacked side by side")
plt.imshow(res,'gray')
plt.show()
image=cv2.imread("test.jpg",0)
x,y=image.shape
th=np.sum(image)/(x*y)
binary=np.zeros((x,y),np.double)
binary=(image>=th)*255
binary=binary.astype(np.uint8)
plt.title("original/binary")
equ=np.hstack((image,binary))
plt.imshow(equ,'gray')
plt.show()
image=cv2.imread('test2.jpg',0)
x,y=image.shape
z=np.zeros((x,y))
for i in range(0,x):
for j in range(0,y):
if(image[i][j]>50 and image[i][j]<150):
z[i][j]=255
else:
z[i][j]=image[i][j]
equ=np.hstack((image,z))
plt.title('Original\Graylevel slicing with background')
plt.imshow(equ,'gray')
plt.show()
image=cv2.imread('test2.jpg',0)
x,y=image.shape
z=np.zeros((x,y))
for i in range(0,x):
for j in range(0,y):
if(image[i][j]>50 and image[i][j]<150):
z[i][j]=255
else:
z[i][j]=0
equ=np.hstack((image,z))
plt.title('Original\Graylevel slicing w/o background')
plt.imshow(equ,'gray')
plt.show()
image=cv2.imread('test.jpg',0)
x,y=image.shape
z=255-image
equ=np.hstack((image,z))
plt.title('Original\Image Negative')
plt.imshow(equ,'gray')
plt.show()
image=cv2.imread('test.png',0)
x,y=image.shape
c=255/(np.log(1+np.max(image)))
z=c*np.log(1+image)
z=np.array(z,dtype=np.uint8)
equ=np.hstack((image,z))
plt.title('Log Transformation')
plt.imshow(equ,'gray')
plt.show()
img=cv2.imread('saltandpep.jpg',0)
x,y=img.shape
z=cv2.blur(img,(3,3))
z1=cv2.blur(img,(5,5))
equ=np.hstack((img,z))
plt.title('original/Averaging filter3X3')
plt.imshow(equ,'gray')
plt.show()
equ=np.hstack((img,z1))
plt.title('original/averging filter 5X5')
plt.imshow(equ,'gray')
plt.show()
z=cv2.medianBlur(img,5)
equ=np.hstack((img,z))
plt.title('original/Median Blur')
plt.imshow(equ,'gray')
plt.show()
| OpenCV DIP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
if not os.path.isdir('bio_segmentation_keras'):
# !git clone https://github.com/izikeros/bio_segmentation_keras
from bio_segmentation_keras.data import train_generator, test_generator, save_result
from bio_segmentation_keras.model import unet
else:
from data import train_generator, test_generator, save_result
from model import unet
# -
# ## Train your Unet with membrane data
# membrane data is in folder membrane/, it is a binary classification task.
#
# The input shape of image and mask are the same :(batch_size,rows,cols,channel = 1)
# ### Train with data generator
# +
from keras.callbacks import ModelCheckpoint
data_gen_args = dict(rotation_range=0.2,
width_shift_range=0.05,
height_shift_range=0.05,
shear_range=0.05,
zoom_range=0.05,
horizontal_flip=True,
fill_mode='nearest')
myGene = train_generator(2,'data/membrane/train','image','label',data_gen_args,save_to_dir = None)
model = unet()
model_checkpoint = ModelCheckpoint('unet_membrane.hdf5', monitor='loss',verbose=1, save_best_only=True)
model.fit_generator(myGene,steps_per_epoch=2000,epochs=5,callbacks=[model_checkpoint])
# -
# ### Train with npy file
# +
#imgs_train,imgs_mask_train = gen_train("data/membrane/train/aug/","data/membrane/train/aug/")
#model.fit(imgs_train, imgs_mask_train, batch_size=2, nb_epoch=10, verbose=1,validation_split=0.2, shuffle=True, callbacks=[model_checkpoint])
# -
# ### test your model and save predicted results
# + pycharm={"is_executing": false}
testGene = test_generator("data/membrane/test")
model = unet()
model.load_weights("unet_membrane.hdf5")
results = model.predict_generator(testGene,30,verbose=1)
save_result("data/membrane/test",results)
# + pycharm={"name": "#%%\n"}
| trainUnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + code_folding=[19, 29, 38, 101, 128, 138, 164] deletable=true editable=true
import os
os.chdir('/home/yash/Desktop/Decision-Flip-Experiments')
from scipy.misc import imread
import matplotlib.patches as mpatches
from models import *
from plotter import *
from saveloader import *
from fgsm_cifar import fgsm
from fgsm_cifar_wrt_class_flipstop import fgsm_wrt_class
from helper import *
from helper import _to_categorical
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import numpy as np
sd = 'shape_dict'
def make_data(X_test, X_train, sess, n, env, n_classes=10):
X_test_sub = X_test[:n]
X_train_sub = X_train[:n]
y_train_sub = sess.run(env.ybar, feed_dict={env.x: X_train_sub, env.training: False})
y_train_sub = _to_categorical(np.argmax(y_train_sub, axis=1), n_classes)
y_test_sub = sess.run(env.ybar, feed_dict={env.x: X_test_sub, env.training: False})
y_test_sub = _to_categorical(np.argmax(y_test_sub, axis=1), n_classes)
return X_test_sub, y_test_sub, X_train_sub, y_train_sub
def plot_all_data_graph(method, epochs, n):
for from_cls in range(2):
for to_cls in range(n_classes):
if (from_cls != to_cls):
l2_test, l2_train, l2_random, l2_random_normal = restore_flip(method, epochs, n, from_cls, to_cls)
#There might be a prob. here! since lens are diff, solved it inside the func
plot_data_graph_without_random(l2_test, l2_train, l2_random, l2_random_normal, n, from_cls, to_cls)
plot_hists_without_random(l2_test, l2_train, n, from_cls, to_cls)
def random_normal_func(X, n, save, lr, lrn):
X = X.reshape(-1, img_rows * img_cols * img_chas)
mean, std = np.mean(X, axis=0), np.std(X, axis=0)
randomX = np.zeros([n, X[0].size])
for i in range(X[0].size):
randomX[:, i] = np.random.normal(mean[i], std[i], n)
randomX = randomX.reshape(-1, img_rows, img_cols, img_chas)
X_random_normal = randomX
ans = sess.run(env.ybar, feed_dict={env.x: randomX, env.training: False})
y_random_normal = _to_categorical(np.argmax(ans, axis=1), n_classes)
X_random = np.random.rand(n, img_rows, img_cols, img_chas)
y_random = sess.run(env.ybar, feed_dict={env.x: X_random, env.training: False})
y_random = _to_categorical(np.argmax(y_random, axis=1), n_classes)
if (save):
save_as_txt(get_flip_path(lr), X_random)
save_as_txt(get_flip_path(lrn), X_random_normal)
return X_random, y_random, X_random_normal, y_random_normal
def run_flip(method, epochs, n, cls=-1):
save_obj({},sd)
test_label = make_label("test", method, epochs, n, False)
train_label = make_label("train", method, epochs, n, False)
random_label = make_label("random", method, epochs, n, False)
random_normal_label = make_label("random_normal", method, epochs, n, False)
data_label_random, data_label_random_normal = make_label("_", method, epochs, n, True)
X_test_sub, y_test_sub, X_train_sub, y_train_sub = make_data(X_test, X_train, sess, n, env)
X_random, y_random, X_random_normal, y_random_normal = random_normal_func(X_train, n, True,
data_label_random,
data_label_random_normal)
if (cls < -2 or cls > 9):
print("Invalid Params")
return
if (method == 1):
X_flip_test = create_adv(X_test_sub, y_test_sub, test_label)
X_flip_train = create_adv(X_train_sub, y_train_sub, train_label)
X_flip_random = create_adv(X_random, y_random, random_label)
X_flip_random_normal = create_adv(X_random_normal, y_random_normal, random_normal_label)
if (method == 2):
X_flip_per_class_test = create_adv_wrt_class(X_test_sub, y_test_sub, test_label)
X_flip_per_class_train = create_adv_wrt_class(X_train_sub, y_train_sub, train_label)
X_flip_per_class_random = create_adv_wrt_class(X_random, y_random, random_label)
X_flip_per_class_random_normal = create_adv_wrt_class(X_random_normal, y_random_normal, random_normal_label)
_, X_flip_test = give_m2_ans(X_test_sub, X_flip_per_class_test, cls)
_, X_flip_train = give_m2_ans(X_train_sub, X_flip_per_class_train, cls)
_, X_flip_random = give_m2_ans(X_random, X_flip_per_class_random, cls)
_, X_flip_random_normal = give_m2_ans(X_random_normal, X_flip_per_class_random_normal, cls)
# a = _predict(X_flip_test, env)
# print(np.argmax(a,axis=1))
l2_test = find_l2(X_flip_test, X_test_sub)
l2_train = find_l2(X_flip_train, X_train_sub)
l2_random = find_l2(X_flip_random, X_random)
l2_random_normal = find_l2(X_flip_random_normal, X_random_normal)
return l2_test, l2_train, l2_random, l2_random_normal
def restore_random_data(lr, lrn):
Xr = load_from_txt(get_flip_path(lr))
Xrn = load_from_txt(get_flip_path(lrn))
y_random_normal = sess.run(env.ybar, feed_dict={env.x: Xrn, env.training: False})
y_random_normal = _to_categorical(np.argmax(y_random_normal, axis=1), n_classes)
y_random = sess.run(env.ybar, feed_dict={env.x: Xr, env.training: False})
y_random = _to_categorical(np.argmax(y_random, axis=1), n_classes)
return Xr, y_random, Xrn, y_random_normal
def restore_flip(method, epochs, n, from_cls=-1, to_cls=-1):
test_label = make_label("test", method, epochs, n, False)
train_label = make_label("train", method, epochs, n, False)
random_label = make_label("random", method, epochs, n, False)
random_normal_label = make_label("random_normal", method, epochs, n, False)
data_label_random, data_label_random_normal = make_label("_", method, epochs, n, True)
X_test_sub, y_test_sub, X_train_sub, y_train_sub = make_data(X_test, X_train, sess, n, env)
X_random, y_random, X_random_normal, y_random_normal = restore_random_data(data_label_random,
data_label_random_normal)
if (method == 1):
X_flip_test = load_from_txt(get_flip_path(test_label))
X_flip_train = load_from_txt(get_flip_path(train_label))
X_flip_random = load_from_txt(get_flip_path(random_label))
X_flip_random_normal = load_from_txt(get_flip_path(random_normal_label))
if (method == 2):
X_flip_per_class_test = load_from_txt(get_flip_path(test_label))
X_flip_per_class_train = load_from_txt(get_flip_path(train_label))
X_flip_per_class_random = load_from_txt(get_flip_path(random_label))
X_flip_per_class_random_normal = load_from_txt(get_flip_path(random_normal_label))
if (from_cls != -1):
print('From Class ' + str(from_cls) + '\n')
X_test_sub, y_test_sub, X_flip_per_class_test = get_class(X_test_sub, y_test_sub,
X_flip_per_class_test, from_cls)
X_train_sub, y_train_sub, X_flip_per_class_train = get_class(X_train_sub, y_train_sub,
X_flip_per_class_train, from_cls)
X_random, y_random, X_flip_per_class_random = get_class(X_random, y_random,
X_flip_per_class_random, from_cls)
X_random_normal, y_random_normal, X_flip_per_class_random_normal = get_class(
X_random_normal, y_random_normal, X_flip_per_class_random_normal, from_cls)
print('Test Data:' + str(y_test_sub.shape[0]))
print('Train Data: ' + str(y_train_sub.shape[0]))
print('Random Data: ' + str(y_random.shape[0]))
print('Random Normal Data: ' + str(y_random_normal.shape[0]))
_, X_flip_test = give_m2_ans(X_test_sub, X_flip_per_class_test, to_cls)
_, X_flip_train = give_m2_ans(X_train_sub, X_flip_per_class_train, to_cls)
_, X_flip_random = give_m2_ans(X_random, X_flip_per_class_random, to_cls)
_, X_flip_random_normal = give_m2_ans(X_random_normal, X_flip_per_class_random_normal, to_cls)
l2_test = find_l2(X_flip_test, X_test_sub)
l2_train = find_l2(X_flip_train, X_train_sub)
l2_random = find_l2(X_flip_random, X_random)
l2_random_normal = find_l2(X_flip_random_normal, X_random_normal)
return l2_test, l2_train, l2_random, l2_random_normal
X_train, y_train, X_test, y_test, X_valid, y_valid = load_data(os)
class Dummy:
pass
env = Dummy()
# + code_folding=[0] deletable=true editable=true
# We need a scope since the inference graph will be reused later
with tf.variable_scope('model'):
env.x = tf.placeholder(tf.float32, (None, img_rows, img_cols,
img_chas), name='x')
env.y = tf.placeholder(tf.float32, (None, n_classes), name='y')
env.training = tf.placeholder(bool, (), name='mode')
env.ybar, logits = model(env.x, logits=True,
training=env.training)
z = tf.argmax(env.y, axis=1)
zbar = tf.argmax(env.ybar, axis=1)
env.count = tf.cast(tf.equal(z, zbar), tf.float32)
env.acc = tf.reduce_mean(env.count, name='acc')
xent = tf.nn.softmax_cross_entropy_with_logits(labels=env.y,
logits=logits)
env.loss = tf.reduce_mean(xent, name='loss')
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(extra_update_ops):
env.optim = tf.train.AdamOptimizer(beta1=0.9, beta2=0.999, epsilon=1e-08,).minimize(env.loss)
with tf.variable_scope('model', reuse=True):
for i in range(n_classes):
if (i == 0):
env.x_adv_wrt_class = (fgsm_wrt_class(model, env.x, i, step_size=.05, bbox_semi_side=10))
else:
x = (fgsm_wrt_class(model, env.x, i, step_size=.05, bbox_semi_side=10))
env.x_adv_wrt_class = tf.concat([env.x_adv_wrt_class, x], axis=0)
env.x_adv, env.all_flipped = fgsm(model, env.x, step_size=.05, bbox_semi_side=10) # epochs is redundant now!
# + deletable=true editable=true
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
sess.run(tf.local_variables_initializer())
# + code_folding=[0, 4] deletable=true editable=true
def save_model(label):
saver = tf.train.Saver()
saver.save(sess, './models/cifar/' + label)
def restore_model(label):
saver = tf.train.Saver()
saver.restore(sess, './models/cifar/' + label)
# + code_folding=[0] deletable=true editable=true
def _evaluate(X_data, y_data, env):
print('\nEvaluating')
n_sample = X_data.shape[0]
batch_size = 128
n_batch = int(np.ceil(n_sample/batch_size))
loss, acc = 0, 0
ns = 0
for ind in range(n_batch):
print(' batch {0}/{1}'.format(ind+1, n_batch), end='\r')
start = ind*batch_size
end = min(n_sample, start+batch_size)
batch_loss, batch_count, batch_acc = sess.run(
[env.loss, env.count, env.acc],
feed_dict={env.x: X_data[start:end],
env.y: y_data[start:end],
env.training: False})
loss += batch_loss*batch_size
ns+=batch_size
acc += batch_acc*batch_size
loss /= ns
acc /= ns
print(' loss: {0:.4f} acc: {1:.4f}'.format(loss, acc))
return loss, acc
# + code_folding=[0, 15, 37] deletable=true editable=true
def _predict(X_data, env):
print('\nPredicting')
n_sample = X_data.shape[0]
batch_size = 128
n_batch = int(np.ceil(n_sample/batch_size))
yval = np.empty((X_data.shape[0], n_classes))
for ind in range(n_batch):
print(' batch {0}/{1}'.format(ind+1, n_batch), end='\r')
start = ind*batch_size
end = min(n_sample, start+batch_size)
batch_y = sess.run(env.ybar, feed_dict={
env.x: X_data[start:end], env.training: False})
yval[start:end] = batch_y
return yval
def train(label):
print('\nTraining')
n_sample = X_train.shape[0]
batch_size = 128
n_batch = int(np.ceil(n_sample/batch_size))
n_epoch = 50
for epoch in range(n_epoch):
print('Epoch {0}/{1}'.format(epoch+1, n_epoch))
for ind in range(n_batch):
print(' batch {0}/{1}'.format(ind+1, n_batch), end='\r')
start = ind*batch_size
end = min(n_sample, start+batch_size)
sess.run(env.optim, feed_dict={env.x: X_train[start:end],
env.y: y_train[start:end],
env.training: True})
if(epoch%5 == 0):
model_label = label+ '{0}'.format(epoch)
print("saving model " + model_label)
save_model(model_label)
save_model(label)
def train_again(X, y, epochs):
#Not making batches, do that if size > 128
for i in range(X.shape[0]):
for e in range(epochs):
sess.run(env.optim, feed_dict={env.x: [X[i]],
env.y: [y[i]],
env.training: True})
# + code_folding=[] deletable=true editable=true
def create_adv_wrt_class(X, Y, label = None):
print('\nCrafting adversarial')
n_sample = X.shape[0]
pred = np.argmax(Y,axis=1)
batch_size = 1
n_batch = int(np.ceil(n_sample/batch_size))
n_epoch = 20
x_adv_shape = list(X.shape)[1:]
x_adv_shape = np.append(np.append(n_sample,n_classes),x_adv_shape)
X_adv = np.empty(x_adv_shape)
for ind in range(n_batch):
print(' batch {0}/{1}'.format(ind+1, n_batch), end='\r')
start = ind*batch_size
end = min(n_sample, start+batch_size)
tmp = sess.run(env.x_adv_wrt_class, feed_dict={env.x: X[start:end],
env.y: Y[start:end],
env.training: False})
tmp[pred[start]] = X[start]
X_adv[start:end] = tmp
if(label != None):
print('\nSaving adversarial')
os.makedirs('data', exist_ok=True)
save_as_txt(get_flip_path(label), X_adv)
return X_adv
# + code_folding=[0] deletable=true editable=true
def create_adv(X, Y, label):
print('\nCrafting adversarial')
n_sample = X.shape[0]
batch_size = 1
n_batch = int(np.ceil(n_sample/batch_size))
n_epoch = 20
X_adv = np.empty_like(X)
for ind in range(n_batch):
print(' batch {0}/{1}'.format(ind+1, n_batch), end='\r')
start = ind*batch_size
end = min(n_sample, start+batch_size)
tmp, all_flipped = sess.run([env.x_adv, env.all_flipped], feed_dict={env.x: X[start:end],
env.y: Y[start:end],
env.training: False})
X_adv[start:end] = tmp
print('\nSaving adversarial')
os.makedirs('data', exist_ok=True)
save_as_txt(get_flip_path(label), X_adv)
return X_adv
# + deletable=true editable=true
method = 2
n = 1000
epochs = 100
label="cifar_with_cnn"
cls = -1
# train(label)
restore_model(label + str(epochs))
X = X_train[:50]
y = y_train[:50]
lbl = "temp_exp_flipstop100e"
Xf1 = load_from_txt(get_flip_path(lbl))
lbl = "temp_exp_flipstop200e"
Xf2 = load_from_txt(get_flip_path(lbl))
# + deletable=true editable=true
for i in range (len(Xf1)):
restore_model(label + str(100))
pred1 = _predict(Xf1[i],env)
restore_model(label + str(200))
pred2 = _predict(Xf2[i],env)
a1 = (np.argmax(pred1, axis = 1))
a2 = np.argmax(pred2, axis=1)
print ()
x=(np.count_nonzero(a1-[0,1,2,3,4,5,6,7,8,9]))
y=(np.count_nonzero(a2-[0,1,2,3,4,5,6,7,8,9]))
print(x)
print(y)
print('e100-e200: ' + str(x-y))
# -
| CIFAR/.ipynb_checkpoints/multiple-hits-experiment-diff-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
model_name = "microsoft/CodeGPT-small-py-adaptedGPT2"
model_name = "microsoft/CodeGPT-small-py"
model_name="jaron-maene/gpt2-medium-nl2bash"
model_name="facebook/bart-large-mnli"
model_name="bert-base-uncased"
model_name = "distilgpt2"
model_name = "gpt2"
model_name = "gpt2-large"
# -
# +
from transformers import pipeline, set_seed
>>> generator = pipeline('text-generation', model=model_name)
>>> set_seed(142)
#>>> generator("Hello, I'm a language model,", max_length=30, num_return_sequences=5)
# -
generator("Anaconda is a Python distribution,", max_length=30, num_return_sequences=5)
>>> set_seed(14243)
generator(" Anaconda is a Python distribution,", max_length=30, num_return_sequences=5)
generator(" Anaconda ... a Python distribution\n\n\n", max_length=30, num_return_sequences=5)
generator(" Anaconda is a Python ", max_length=30, num_return_sequences=5)
generator(" Anaconda is a snake ", max_length=30, num_return_sequences=5)
generator(" Anaconda is a ", max_length=30, num_return_sequences=5)
| hf-gpt2-2 - Copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Basic control structures and variables
#
# ## Nails for the hammer
# + [markdown] slideshow={"slide_type": "notes"}
# As one of the prereqs for this course was some knowledge of MATLAB or a decent understanding of programming, we won't spend a huge amount of time on concepts, assuming you have a fair idea, and focus on the Python context. If you find yourself stuck on a fundamental question, do ask others around you, and do help others around you, especially if I'm on the far side of the room. I'll not feel left out.
#
# These first few tasks are all pretty unapplied and guided - this should get us underway, and we will then start learning Python by using actual science applications. Since we only have two part-days, and you already have used some coding, I will move through this introductory session fairly quickly, so you have the tools to explore a wide variety of Python aspects by the end of Day 2 - please discuss with each other and if you are still wondering about things during the break, do ask.
#
# My recommendation is that the first time I run a cell, you do so too - I will show you how in a mo. This means everything I define and later use, you will have defined and can later use too.
# + [markdown] slideshow={"slide_type": "slide"}
# # First experiment with Jupyter
# + [markdown] slideshow={"slide_type": "fragment"}
# * Find which line number you are in Etherpad
# * Click the blank box below saying `In [ ]`
# * Change the box below to read `x = LINENUM` (e.g. `x = 15`)
# * Press *Ctrl+Return*
# +
#x = ???
# + [markdown] slideshow={"slide_type": "notes"}
# Note that `#` is the comment symbol in Python - all text after it is ignored.
#
# This lets you to make notes-to-self in your code. However, above, we want to Python to run that line, so don't forget to delete the `#` symbol before pressing _Ctrl+Return_
# + [markdown] slideshow={"slide_type": "fragment"}
# This just set the variable `x` (globally for this notebook) to the number of your Etherpad line
# + [markdown] slideshow={"slide_type": "slide"}
# Click to edit the next `In [ ]` and press *Ctrl+Return* (without changing anything this time)
# -
2 * x
# + [markdown] slideshow={"slide_type": "notes"}
# Hopefully, you should notice that *x* is indeed what you input. This should all be pretty familiar, unless you come from statically-typed languages (C/C++/Java) where you would have to declare a variable. Python lets you effectively declare it, and its type, by assigning to it.
# + [markdown] slideshow={"slide_type": "fragment"}
# We can refer to this output as `Out[2]`, using it like any variable, without having to re-run everything so far.
# + [markdown] slideshow={"slide_type": "slide"}
# Do the same trick in the next `In [ ]` (edit and *Ctrl+Return*)
# -
if Out[2] / 2 == x:
print("Something else")
print(2/3)
# + [markdown] slideshow={"slide_type": "notes"}
# There are a couple of things here to dissect. Many languages need braces `{}`, brackets `[]` or parentheses `()` to draw boundaries around the argument of the `if` statement, or the body. In Python, symbols are reduced in favour of words and spaces, for readability. To achieve the same effect, Python separates the condition from the body with the colon `:` and the indentation (four spaces, by convention)
#
# Indentation is Python's Marmite for other programmers. As they are keen to point out, most whitespace does *not* matter: e.g. I could write
#
# `Out[2] / 2 == x :`
#
# and it's the same as
#
# `Out[2] / 2 = x:`
#
# However, indentation (whitespace at the start of a line) does. It is an extremely visually clear way to show where blocks of code, like the body of this `if` statement, start and end. For scientists, it forces a very good programming practice - basic code style - your code doesn't run without if it isn't visually clear (well... neatly indented). One of the reasons I got into Python was that my PhD C++ code was so unreadable, dodgily indented and unwieldy, it was easier to rewrite in Python than try and extend it - once I had modification was easy, I could see how everything was laid out, where functions started and ended, before even reading a character.
#
# Another thing to note is that you can print with `print`. Here we are running Python3, the latest version. If you've played with Python2, you will notice this syntax is a bit different - it has parentheses (aka. parens) around the argument. A whole raft of changes came in with Python3 and, like XP, Python2 will be around for some time to come through sheer momentum. However by 2018, almost all important libraries are now Python3-ready and I, for one, develop new software in Python3 only, where possible. Good porting tools exist for moving old code.
#
# To summarize those key points:
# + [markdown] slideshow={"slide_type": "skip"}
# * Indentation separates bodies of functions, conditionals, loops from the outside code
# * Basic boolean operators are `==`, `or`, `and`, `not`, `>`, `<`, etc.
# * This is Python3 syntax (very slightly different to Python2)
# + [markdown] slideshow={"slide_type": "slide"}
# # Debugging
# ## Ladder for the hole
# + [markdown] slideshow={"slide_type": "notes"}
# Now we can edit and run code, the inevitable consequence is that we get a bug!
# + [markdown] slideshow={"slide_type": "slide"}
# Try running the line below (without correcting it):
# -
if (x < 2) or (x > 100):
print("Do I know you?')
else:
print('Yes I do')
# + [markdown] slideshow={"slide_type": "fragment"}
# Great. Pretty colours! What do they mean?
# + [markdown] slideshow={"slide_type": "notes"}
# In general, the most immediately useful information is on the very last line: `SyntaxError`. Above it, is the last "frame" called - that is, level of function. Say you call a function, and it calls another, and so forth until the innermost function hits an error. Here, it will list all the functions in that sequence, so you can track exactly how you got to the dodgy line.
#
# In Jupyter, the filename is not very informative - but you can see the input number (e.g. `ipython-input-4` ⇆ `In [4]`).
#
# Bonus mark if you spotted the error (not that we actually have marks, but hey) - have a look and see if you can fix it and re-run the command, the same way you ran it the first time. Feel free to discuss, use Etherpad chat, whichever. Note that the `In [4]` changes to `In [5]`, and so forth, each time you run that "cell".
#
# In general, "a string" and 'a string' are equivalent - Python does not mind whether single or double quotes are used, as long as they match at either end.
# + [markdown] slideshow={"slide_type": "slide"}
# * Check the error at the bottom
# * Check the caret (^) showing where Python sees a problem
# * Look at the syntax highlighting in the cell
# + [markdown] slideshow={"slide_type": "slide"}
# # Functions
#
# ## Painting the Black Box
# + [markdown] slideshow={"slide_type": "notes"}
# Like every language, Python has functions... they are defined like so:
# -
def tell_me_if_im_old(name, age):
print("...checking if", name, "is old...")
if age < 100:
return False
else:
return True
# + [markdown] slideshow={"slide_type": "notes"}
# Couple things to note: (i) this has nested blocks, we just indented twice for the inner ones (`if` and `else` bodies). (ii) this *didn't* specify a type for `age` (or `name`) - all it cares is that we can compare `age` to an *int*. Similarly, we don't specify a return type - it is up to us to return something intuitive and sensible. Let's use it:
# + slideshow={"slide_type": "fragment"}
phils_age = 32
name = "Phil"
print("I am", phils_age, "- am I old?")
if tell_me_if_im_old(name, phils_age):
print("Yup, it's a fact.")
else:
print("Not at all, spring chicken.")
# + [markdown] slideshow={"slide_type": "notes"}
# This one I'm saving for future use. You can see that this is used pretty much like every function in many languages - name, parentheses (aka "parens") containing arguments.
# + [markdown] slideshow={"slide_type": "notes"}
# There are other types of function - those on objects. Rather than cramming an introduction to object-oriented programming (OOP) now, especially as a number of you have done Computer Science, I have added an Appendix to these notes about my non-existent dogs, "Freddie" and "Nitwit". If OOP confuses you, we can use a bit of the break if you're brain isn't fried to go through those.
#
# For the moment though, what matters is that you can add a dot to most things in Python and access a number of properties of that variable or things it can do. A couple of examples to make it clearer:
# + slideshow={"slide_type": "slide"}
"just a normal string".upper()
# + slideshow={"slide_type": "fragment"}
if "another string".islower():
print("The string was lowercase")
# + [markdown] slideshow={"slide_type": "slide"}
# # Lists and dicts
# ## Conveyor belts and storage units
# + [markdown] slideshow={"slide_type": "notes"}
# Like any language, often need to be able to group things in Python.
# + [markdown] slideshow={"slide_type": "slide"}
# We can create a `list`, which is like a one-dimensional C or FORTRAN array, as follows (don't forget to execute it!):
# -
things_to_do = ['Learn Python', 'Finish PhD', 'Publish research',
'Accept Nobel prize', 'Inspire a new generation']
# + [markdown] slideshow={"slide_type": "notes"}
# The syntax here is the same as for the more basic types we saw so far, `variable = value`, and our list is a comma-separated series of *things*, bounded by brackets `[]`.
# + [markdown] slideshow={"slide_type": "fragment"}
# Once you execute the cell above, you should be able to try this:
# -
sorted(things_to_do)
# ...and get an alphabetical list of tasks. Note that `sorted` doesn't update the original object, so if you re-run without `sorted`, it's back to the previous order.
# + [markdown] slideshow={"slide_type": "notes"}
# To do this, you use a slightly different technique *or* you just assign: `things_to_do = sorted(things_to_do)`.
#
# `sorted` guesses the ordering you want, based on the type of data. It doesn't all have to be strings, you could have mixture of strings and integers, say. `sorted` can take optional arguments, letting you force it to do things the way you want, or even provide your own ordering function.
# + [markdown] slideshow={"slide_type": "slide"}
# # So how do we get things out of a list?
# + [markdown] slideshow={"slide_type": "fragment"}
# Lists are ordered, so you can say, "I'd like the fifth item, please". You put it in brackets after the list name:
# + slideshow={"slide_type": "fragment"}
things_to_do[4]
# + [markdown] slideshow={"slide_type": "slide"}
# However, note that...
# + [markdown] slideshow={"slide_type": "fragment"}
# ## PYTHON IS ZERO-INDEXED
# + [markdown] slideshow={"slide_type": "fragment"}
# Like C, but unlike MATLAB or FORTRAN. The first item in a list is "`things_to_do[0]`"
# + [markdown] slideshow={"slide_type": "fragment"}
# By the way, did you notice that `In[]` is a list?
# -
type(In)
# + [markdown] slideshow={"slide_type": "notes"}
# You can try `len` as well!
# + [markdown] slideshow={"slide_type": "slide"}
# # Slicing
#
# ## Sounds cool, is cool
# + [markdown] slideshow={"slide_type": "fragment"}
# Suppose I want a bit of a list, like the middle 3 items, or the first 2...
# + slideshow={"slide_type": "fragment"}
print(things_to_do)
things_to_do[1:4]
# + [markdown] slideshow={"slide_type": "notes"}
# ...if you wonder why `things_to_do[4]` (the fifth item) didn't appear, note that that number after the colon means 'up-to-but-not-including'...
# + slideshow={"slide_type": "fragment"}
things_to_do[:2]
# + slideshow={"slide_type": "fragment"}
things_to_do[3:]
# + slideshow={"slide_type": "fragment"}
things_to_do[-2:]
# + [markdown] slideshow={"slide_type": "notes"}
# We can use negative numbers to count back from the end instead...
# + [markdown] slideshow={"slide_type": "notes"}
# Which is "start two from the end, and give me everything from there onwards"
# + [markdown] slideshow={"slide_type": "slide"}
# # Dictionaries
# + [markdown] slideshow={"slide_type": "notes"}
# This is fine - a list is like a conveyer belt, with a load of arbitrarily filled boxes passing by, one after the other. However, sometimes we care which item is which, and want to have a name for it, so we can drop in and grab it wherever it may be. That's where dictionaries (= dicts) come in. Try running this:
# + slideshow={"slide_type": "fragment"}
my_meetup_dot_com_profile = {
"first name": "Ignatius",
"favourite number": 9,
"favourite programming language": "FORTRAN66",
3: "is the magic number"
}
# + [markdown] slideshow={"slide_type": "notes"}
# The syntax is slightly different. Instead of brackets, we use braces `{}`.
#
# Now each element has a name, followed by a colon and the element itself, which can still be basically anything. Like storage units, we now have a group of things we can address. That address could be a number, a string, or any other basic type. And keys can be different types - check the last entry.
# + [markdown] slideshow={"slide_type": "fragment"}
# This is *completely unsorted*. However, you can get any element back by its name (key):
# -
print("My favourite number is")
my_meetup_dot_com_profile['favourite Numbr']
# + [markdown] slideshow={"slide_type": "notes"}
# Hmm... I'll let you fix that...
# + [markdown] slideshow={"slide_type": "notes"}
# And even though it is unsorted, it can have integers or so forth as keys - note the very last entry in the definition...
# + slideshow={"slide_type": "fragment"}
my_meetup_dot_com_profile[3]
# + [markdown] slideshow={"slide_type": "notes"}
# ...but not...
# -
my_meetup_dot_com_profile[2]
# + [markdown] slideshow={"slide_type": "notes"}
# Because it doesn't exist.
# + [markdown] slideshow={"slide_type": "slide"}
# # Extending lists and dicts
# + [markdown] slideshow={"slide_type": "fragment"}
# Extending lists is a little unwieldy, but dicts are more intuitive:
# +
things_to_do.append("Find a nice retirement village in the Galapagos Islands")
my_meetup_dot_com_profile["Interests"] = ["Python2", "Python3", "Scientific Python", "Pottery"]
print("TODO:", things_to_do, "\n\nMEETUP:", my_meetup_dot_com_profile)
# -
# Here, we added an `Interests` key to the `my_meetup_dot_com_profile` dictionary.
#
# Note that anything can be an *item* in a dictionary or list - in this case, a list is an item in our dict, whose key is `Interests`. Also note that, as with other languages, `\n` signifies a newline.
# + [markdown] slideshow={"slide_type": "slide"}
# # Join
#
# ## The most useful method in the str object
# + [markdown] slideshow={"slide_type": "notes"}
# Different languages have different ways of collapsing a string with a separator - it is something you will need to do again and again. In Python, the approach is a little surprising - we have already seen that strings are objects (of type `str`), but it turns out Python uses this to provide a way of joining iterables (like `lists` or `dicts`):
# + slideshow={"slide_type": "fragment"}
', '.join(things_to_do)
# + [markdown] slideshow={"slide_type": "fragment"}
# This way around, all the string needs to do is add itself before each item in the iterable (except the first) - as long as it can keep getting items, it doesn't care what the iterable is, a dict or anything (iterable == thing you iterate/step through).
# + [markdown] slideshow={"slide_type": "notes"}
# Can you see the benefit of defining this method as part of `str`, being able to pass it any possible iterable to join up, instead of having to define a method on every type of iterable and passing it a string?
# + [markdown] slideshow={"slide_type": "slide"}
# # Quick aside
#
# ## Save early, save often
# + [markdown] slideshow={"slide_type": "fragment"}
# Jupyter does do some automatic saving, but click the disk icon at the top left now to force an update...
# + [markdown] slideshow={"slide_type": "slide"}
# # ReDebugging
#
# ## Bringing it together
# + [markdown] slideshow={"slide_type": "slide"}
# Here's another one to see if you can fix - try running and then spot the issue:
# -
y = 120
for z in range(5):
y = y / z
print("When z is", z, "then y is", y)
# Output should be:
# ```
# When z is 1 then y is 120.0
# When z is 2 then y is 60.0
# When z is 3 then y is 20.0
# When z is 4 then y is 5.0
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# Couple of bits of useful information:
#
# * `for` is, unsurprisingly, a `for` loop, as in other languages
# * `for` doesn't have limit arguments, like in C, say - it takes a set/list/(anything iterable) and goes through each element
# * `range` just returns a list of integers
# * `range` can take one argument or two arguments (or more, but not so relevant now)
# * `range` documentation is here : [Python3 range syntax](https://docs.python.org/3/library/functions.html#func-range) (link opens in new window, so click it!)
#
# The easiest way, IMO, to info on any Python function or library is to Google "Python3 funcname" and click the first python.org link. If even something seems wrong, make sure you are looking at the Python3, not Python2 docs (or v.v.)
#
# If it isn't there already, when you solve this (i.e. get the output above), please make a note in Etherpad. If you are still solving the puzzle - don't look at Etherpad unless you want the answer.
# + [markdown] slideshow={"slide_type": "notes"}
# Note that the `in` operator can separately be used to return a boolean: e.g. "`list_item in alist`", "`key in adict`" can be used for `if` clauses
# + [markdown] slideshow={"slide_type": "slide"}
# To see what range actually returns, you can run:
# -
type(range(5))
# + [markdown] slideshow={"slide_type": "notes"}
# Well, that wasn't very descriptive. It turns out the `range` function returns something with (a confusingly named) type `range` - this is what `for` gets handed. Don't worry about the ins-and-outs of that type just yet - what matters is: things of type `range` look like, act like and sound like a `list`.
#
# As far as `for` is concerned, that's good enough - this is the [**duck test**](https://simple.wikipedia.org/wiki/Duck_typing), and is a key paradigm in Python - when writing code, don't require your input to be of a specific type `float`, or type `int` (for example), just complain if it doesn't *do* what you want (for example, it refuses to be added to something, or printed, by giving an error).
#
# Here, `range` can iterate, like a list, which is all `for` wants - it doesn't care that it isn't _technically_ a list, as it can do what `for` needs. Similarly with, say, the `==` comparison operator - you can compare any objects and as long as they define their comparison behaviour, the operator itself doesn't care what they are.
#
# To prove `range(5)` is what we expect it is, lets turn it into a list and see what happens...
# + slideshow={"slide_type": "fragment"}
list(range(5))
# + [markdown] slideshow={"slide_type": "fragment"}
# Essentially, `for` sees what we see - a sequence of five numbers.
# + [markdown] slideshow={"slide_type": "slide"}
# Bear in mind that you can cast like this to various types. `str` will make something a string...
# -
str(range(5))
3.14 + " is almost pi"
# + [markdown] slideshow={"slide_type": "fragment"}
# However, unlike some languages, Python does care about type, so you must concatenate strings with strings or add numerical types to numerical types. To check the type of any number or variable, you can use the built-in, `type`:
# -
type(str(3.14))
# + [markdown] slideshow={"slide_type": "slide"}
# A string is, well, kind of like a list of characters, right? Certainly true in C and FORTRAN. So can "`for`" iterate over that too?
# -
d = ""
for i in range(2):
d += "Let me hear you say "
j = 4
for c in "YMCA":
d += c + '.'
d += " "
print(d)
# + [markdown] slideshow={"slide_type": "notes"}
# Yes, yes it can: check out the '`for c in "YMCA"`'. We also snuck in a nested loop - you just keep indenting each time you nest - no more complicated than that.
#
# And, admittedly, a new operator has appeared: "`+=`". Familiar to most languages it is equivalent to `d = d + blah`. Note that we therefore have to have `d` defined before the first `+=`, as "`d = d + anything`" doesn't make sense if `d` does not already exist. This is the reason for our first line: "`d = ""`".
#
# Finally, note that we have looped twice using `range`, just as above, but our loop variable `i` is never used - it is effectively a placeholder. All we want from that line is to have the body below run twice.
#
# *Sidenote*: some people use an underscore "`_`" as a loop variable in this case, to indicate to someone reading that the variable is nothing more than that and never used - as far as Python is concerned, "`_`" is as good a name for a variable as any other, so this is entirely about readability.
# + [markdown] slideshow={"slide_type": "slide"}
# Just to prove that "`for`" will iterate over any list, numeric, string or otherwise, we can try it with our todo list:
# -
total_months = 0
for task in things_to_do:
total_months += len(task)
print(task, ":")
print(" using Python, this task will take", len(task), "months to complete")
print("You cannot retire for at least", total_months // 12, "years")
# + [markdown] slideshow={"slide_type": "notes"}
# Here we have added one or two minor surprises. One is the *horrendously* misleading use of *len* (hint: you are unlikely to get a Nobel Prize by 2020), which in reality calculates the number of items in any "`iterable`". Reminder: this is the general term for something sort-of-list-like. According to my script, tasks take as long as the number of items in their "list", i.e. characters in the string.
#
# Another aspect is the double slash. This is an important difference between Python3 and Python2 (and many other languages). [Dividing non-divisible integers](https://en.wikipedia.org/wiki/Division_%28mathematics%29#Of_integers) with one slash, as normal, gives a float in Python3. If you want to get an integer (that is, whatever the float answer is with the decimal chopped off), you can use double-slash. Try removing the second slash and re-running to see the exact non-integer answer. In Python2, using one slash *always* gives an integer, unless you cast the bottom or top to a float (as in C, say).
# + [markdown] slideshow={"slide_type": "subslide"}
# # Exercise
# + [markdown] slideshow={"slide_type": "notes"}
# ...if we have time...
# -
# Use the cell below to create a function that takes a dictionary as an argument (like `name` and `age` were in an earlier example), and returns a new dictionary, with the same keys, but where each original item has been converted to a string.
#
# Any items of a ``list`` type should become a single string with entries separated by commas and no brackets at either side (e.g. "one, two, kumquat, four").
#
# Call your function with "**`my_meetup_dot_com_profile`**" as an argument and print each entry on a separate line like so:
#
# FAVOURITE PROGRAMMING LANGUAGE : FORTRAN66
# FIRST NAME : Ignatius
# ...
#
# where each key is capitalized and item value printed beside it. The order of lines is not important.
# + [markdown] slideshow={"slide_type": "notes"}
# Note that you can create a blank dictionary with "`= {}`".
# Extension: if you get that done quickly, try making your function call itself on any (nested) items it finds of `dict` type, and turn the output into the string-value you need for your new dictionary. This is "recursion".
#
# Set
# ```
# my_meetup_dot_com_profile['dogs'] = {"Freddie": 5, "Nitwit": "also a dog"}
# ```
# and run your function on "**`my_meetup_dot_com_profile`**" again to test your improvement.
# + slideshow={"slide_type": "notes"}
# Try the exercise in here
# + [markdown] slideshow={"slide_type": "slide"}
# # Modules
#
# ## Gotta catch em all
# + [markdown] slideshow={"slide_type": "slide"}
# ## Introduction to modules
# + [markdown] slideshow={"slide_type": "fragment"}
# * Where Python gets its power
# + [markdown] slideshow={"slide_type": "notes"}
# Wealth of modules providing everything from full GUI toolkits to astrophysics simulations. Mature and reliable numerics libraries, and an evolving ecosystem that grows day by day
# + [markdown] slideshow={"slide_type": "fragment"}
# * Some modules come bundled, but you can install many more
# + [markdown] slideshow={"slide_type": "notes"}
# Python has tools so you can just specify the name and it will get it from the appropriate online repository. On Windows, this can be tricky, but a recent Python distribution, Anaconda, makes it easy, and includes thousands of packages out of the box. This seems your best bet if on Windows or Mac and I have USB sticks here for you to get going at the end of the day.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Some are part of Python, lots are third-party
# + [markdown] slideshow={"slide_type": "notes"}
# It is good to have an idea of which is which, as you can get (normally free) help, often at pretty short notice, by heading to the project's forums - remember, always be polite and, bear in mind, it isn't a commercial service, so an answer isn't guaranteed. For reference, when someone is really desperate they can offer a bounty for a solution or a bit of code on certain websites, not to mention people offering consultancy support, so that doesn't entirely mean there are no other options, but most of the time what's freely available is more than adequate.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Using modules
# + [markdown] slideshow={"slide_type": "fragment"}
# First off, you *import* a module. This tells Python that it should hoke it out of its cave of treasures for upcoming use...
# + slideshow={"slide_type": "-"}
import math
# + [markdown] slideshow={"slide_type": "notes"}
# This brings in a whole set of tools for dealing with basic mathematics. (so execute it!)
# + [markdown] slideshow={"slide_type": "fragment"}
# We reach into this with a dot...
# + slideshow={"slide_type": "-"}
math.pi
# + [markdown] slideshow={"slide_type": "fragment"}
# It has functions and constants...
# -
math.sin(math.pi / 2)
# + [markdown] slideshow={"slide_type": "notes"}
# So how do you find out what is available in `math`? Like before, Google `python3 math`... try it now:
# https://www.google.ie/search?q=python3+math (this is just the Google search link)
# + [markdown] slideshow={"slide_type": "notes"}
# For me, the closest version of Python was actually the second link (for Python 3.5) - normally, minor version differences are rarely a problem, but if what you see seems different to the manual, just add Google for "`python 3.4 math`", or whatever. To check the current version that Jupyter is using, go to `Help->About`.
# + [markdown] slideshow={"slide_type": "slide"}
# Another useful one is `os`...
# -
import os
print(os.path.exists('/usr/bin/python'))
# + [markdown] slideshow={"slide_type": "notes"}
# This is in fact using a submodule, `os.path`, we can reach in twice to get functions inside that.
# + [markdown] slideshow={"slide_type": "fragment"}
# Also `sys`...
# -
import sys
print(sys.path)
# sys.exit(1) # Exit with error code 1
# + [markdown] slideshow={"slide_type": "notes"}
# The last command tells your script to exit, with code 1. I don't really want to try that right now, as whatever happens won't be good.
# + [markdown] slideshow={"slide_type": "slide"}
# ## There are four key variants of importing
#
# ### You will see all four, so check back here
# + slideshow={"slide_type": "fragment"}
import math
print(math.e)
# + [markdown] slideshow={"slide_type": "notes"}
# This is the most succint as it just dumps everything in the `math` module into the global namespace, which saves much typing, but it can make debugging a headache, especially if there are some strange functions or variables in there that happen to have the same name as something else you later decide to use (this does happen from time to time)
# + slideshow={"slide_type": "fragment"}
import math as m
print(m.e)
# + [markdown] slideshow={"slide_type": "notes"}
# ...we've seen that...
# + slideshow={"slide_type": "fragment"}
from math import e
print(e)
# + [markdown] slideshow={"slide_type": "notes"}
# ...in other words, give `math` an alias...
# + slideshow={"slide_type": "fragment"}
from math import *
print(e)
# + [markdown] slideshow={"slide_type": "notes"}
# ...which saves some typing if you're *sure* you don't want to use any other variable called `e`...
# + [markdown] slideshow={"slide_type": "slide"}
# # Pythonic
# ## Walks like a Python...
# + [markdown] slideshow={"slide_type": "notes"}
# This is a fundamental principle of Python and, unusually for something for fundamental, it isn't part of the code. Python is a bit like Ultimate Frisbee - in case you haven't had the pleasure, referees are not required, even in international competitions. This is on the basis that, if you're playing the sport, then you've decided you're there with the ethos, and the players can sort out those decisions amongst themselves and get on with the game. If you're trying to take advantage of that, then the fundamental question is, why play Ultimate Frisbee?
# + [markdown] slideshow={"slide_type": "notes"}
# Good Python practice frequently diverges from what is commended in other languages. Succinctness or efficiency is not necessarily good - clarity comes before cleverness. However, using a pattern from previous experience when Python has a neater, more beautiful, more *Pythonic* way - that's something to try to avoid. The upshot of all this is that, when you come to what the community considers "good code", it is simple, elegant and clear.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Not-so-Pythonic way:
# -
for i in range(len(things_to_do)):
print(things_to_do[i])
# + [markdown] slideshow={"slide_type": "fragment"}
# ## More Pythonic way:
# -
for task in things_to_do:
print(task)
# Same output, but one feels more like pseudocode, more like how you would verbally convey the idea if somebody asked.
# + [markdown] slideshow={"slide_type": "slide"}
# If you want to *get* what Python is about, try this (literally):
# -
import this
# + [markdown] slideshow={"slide_type": "notes"}
# This is one of the hardest things to get when switching to Python, particularly from compiled languages, but it is one of Python's greatest rewards. Nothing much beats opening up a script you hacked together two years ago to see zen-like transparency - no confusion, beautifully professional, and thanks to Python's design, no extra time spent writing it, just a feel for what Pythonic *is*.
# + [markdown] slideshow={"slide_type": "slide"}
# # A couple of resources...
# + [markdown] slideshow={"slide_type": "fragment"}
# * If there is one link from this course to click: [What is Pythonic](http://blog.startifact.com/posts/older/what-is-pythonic.html)
# + [markdown] slideshow={"slide_type": "fragment"}
# * [PEP](https://www.python.org/dev/peps/pep-0008/) (Python Enhancement Proposal) 8 - code style
# + [markdown] slideshow={"slide_type": "notes"}
# These are publicly accessible design documents and nearly everything new of importance passes through here. PEP8 is, in fact, a style guide; unlike many languages, there is more or less a definitive recommended style, and this spells it out. Since I discovered *flake8* and *pylint*, I have my IDE set to highlight style errors and it has reaped dividends to the moon and back - I strongly recommend you do so too, especially if we ever have to code together.
# + [markdown] slideshow={"slide_type": "slide"}
# # Exceptions
# ## Are more than ordinary
# + [markdown] slideshow={"slide_type": "notes"}
# In the words of one of early computing's most idiosyncratic legends, Rear Admiral Dr <NAME>, "It is often easier to ask for forgiveness than to ask for permission". A distinguishing feature of Python is that it is built from the ground up with this maxim in mind, known as EAFP. In practice, this means preferring exceptions over tests, so using a `try-except` block (or, elsewhere, `try-catch`, etc.) instead of an `if` statement when checking whether you can perform an action.
# + [markdown] slideshow={"slide_type": "subslide"}
# Simple examples where you are more likely to use an exception in Python than another language:
# + [markdown] slideshow={"slide_type": "fragment"}
# * When making a directory
# + [markdown] slideshow={"slide_type": "notes"}
# Can you see why this might be useful? If you use an `if` to check for non-existence, there is a potential race condition: by the time you reach the body, it could be made. If you catch the exception, you know it existed exactly when you tried to make it. (Although, the recommended routine, `os.makedirs`, has an optional don't-complain-if-dir-exists argument, which is probably even better)
# + [markdown] slideshow={"slide_type": "fragment"}
# * When testing file existence
# + [markdown] slideshow={"slide_type": "notes"}
# Again with the race condition - this is in fact a security issue, as an attacker can create the file between you checking for existence and opening it for writing. If they create it, of course, then they set the permissions and can see the content regardless of your attempts to block reading.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Checking type
# + [markdown] slideshow={"slide_type": "notes"}
# We talked a lot about duck-typing... the underlying principle is that you never reject input types *as long as they work*. Now imagine you have a routine with an argument `x`, where you want to do one thing if `x` is numeric type and another if it's, say, a string... if you use `if` and check their type, well what if this is some weird subclass of float that your routine has been sent, or the author of this type has carefully implemented all the necessary magic functions to make it quack like a float, but it's a completely unrelated class?
#
# Push the boat out and see if it floats. Try casting to a float and if it doesn't work catch the exception. Then try casting to a string. Everything should cast to a string somehow. Now you have made checking numeric-ness a problem of `float`, which is infinitely more qualified to do this than you are.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Checking for a dictionary key
# + [markdown] slideshow={"slide_type": "notes"}
# Maybe what you think is a dictionary isn't - it's something that, when you request `thingy["something"]` will check whether `"something"` is something it might dynamically add, and, if so, will gladly return the result. If you check first (`if "something" in thingy:`), either the answer is misleading ("no"), or what looks like an innocent if-clause is modifying your dictionary. Moreover, even for an ordinary dictionary, having an if-clause followed by a retrieval hits your dictionary scan twice - try-catch only searches once.
# + [markdown] slideshow={"slide_type": "notes"}
# Hopefully this motivates the idea of exceptions before tests - EAFP. Why then have you been repeatedly told not to do this in, as Python calls them, LBYL (Look Before You Leap) languages? The answer is usually that exceptions are horrendously slow and inefficient. In Python this isn't true, by design. They are so fundamental to the language that every loop in fact ends, not with a failing test, but when the iterable throws a specific exception (the `StopIteration` exception).
# + [markdown] slideshow={"slide_type": "subslide"}
# # Try a try
# + [markdown] slideshow={"slide_type": "notes"}
# The actual syntax is similar to what you will have seen elsewhere:
# -
try:
something_stupid()
except:
print('Doh!')
# + [markdown] slideshow={"slide_type": "notes"}
# Well, the first thing is that we didn't define *something_stupid*. So we get an exception and our extremely generic `except` provides no useful information.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An improvement
# + slideshow={"slide_type": "-"}
try:
something_stupid()
except Exception as e:
print('Doh! You forgot that', e)
# + [markdown] slideshow={"slide_type": "notes"}
# Definitely better, and note that `print` will cast `e`, the exception, to a string - it is actually a more complex object:
# + slideshow={"slide_type": "fragment"}
try:
something_stupid()
except Exception as e:
print(', '.join(dir(e)))
# + [markdown] slideshow={"slide_type": "notes"}
# Note in particular that `e` can have arguments when it is thrown - retrieved via `e.args`. Suppose we don't actually want to stop the exception bubbling up, just to some logging or tidy-up on the way through.
# + slideshow={"slide_type": "subslide"}
try:
something_stupid()
except Exception as e:
print('I am *not* cleaning your mess for you, deal with it yourself!')
raise e
# + [markdown] slideshow={"slide_type": "notes"}
# Now we get the same exception we would have got the last time, but the `raise` keyword has kept it moving on past. This is quite useful, as Python exceptions tend to have lots of juicy info we wouldn't want to lose by ending our except block with a boring `print('oh noes')` and program exit.
#
# In fact this is how we can throw a brand new exception - we take one of the standard types, e.g. "`RuntimeError`" and `raise` it with an explanatory argument:
# + slideshow={"slide_type": "subslide"}
raise RuntimeError("Something quite generic went wrong")
# + [markdown] slideshow={"slide_type": "subslide"}
# # Better yet
# + [markdown] slideshow={"slide_type": "notes"}
# This is a very broad error, and a parent class of most of the exceptions we have seen. It doesn't make it easy for calling functions to pin-point what went wrong...
#
# Looking at it from the other end, just as we should be more specific about what we catch, we should be more specific about what we catch. This is a good idea, as, usually when we catch an exception, it is because we expect a particular thing to go wrong. As a case in point:
# -
try:
os.makedirs(dirname)
except:
# Great, someone has already created that directory
# We can carry on!
pass
# lalalalala...
print("just mucking about with my friend dirname")
# + [markdown] slideshow={"slide_type": "notes"}
# First note `pass`. This is required because every block must have at least one non-comment line - if nothing else is there, we can use `pass` - it is Python's `no-op` if that helps.
#
# Now ask yourself, "did we ever actually define dirname?" No? Then as soon as we use it after our supposed check, we will get an unhandled NameError exception.
# + [markdown] slideshow={"slide_type": "subslide"}
# We can specify what type of exceptions we catch... dir-already-exists have type `OSError`.
# -
try:
os.makedirs(dirname)
except OSError:
print("We had an error creating the directory")
# lalalalala...
print("just mucking about with my friend dirname")
# + [markdown] slideshow={"slide_type": "notes"}
# Now we have stepped out of the way of genuine errors coming through. We can actually handle different exception types in different ways - there are plenty of cases this might be useful, if you want to do something awkward which could fail in five different directions.
# + slideshow={"slide_type": "subslide"}
try:
os.makedirs(dirname)
except OSError:
# Great, someone has already created that directory
# We can carry on!
pass
except NameError:
print("""
This is the third code example
where you haven't defined `dirname`.
Seriously, catch yourself on.
""")
# lalalalala...
print("just mucking about with my friend dirname")
# + [markdown] slideshow={"slide_type": "notes"}
# Yeah. Great.
#
# Useful tool just slotted in there: multi-line strings. If you start a string with three quotes, you can keep going on and on until you hit another three.
# + [markdown] slideshow={"slide_type": "notes"}
# One final example to show how, for this particular case, you really can be a little more specific.
# + slideshow={"slide_type": "subslide"}
dirname = "/etc/passwd"
try:
os.makedirs(dirname)
except OSError:
# Great, someone has already created that directory
# We can carry on!
pass
# lalalalala...
print("just mucking about with my friend", dirname)
# + [markdown] slideshow={"slide_type": "notes"}
# Emm...we tried to make a directory with the same name as a key system file and are blithely assuming that every failure in doing so is simply because it is an existing directory. Not so good.
# + slideshow={"slide_type": "subslide"}
import errno
dirname = "/etc/pass"
try:
os.makedirs(dirname)
except OSError as e:
if e.errno != errno.EEXIST:
raise
# Great, someone has already created that directory
# We can carry on!
# lalalalala...
print("just mucking about with my friend", dirname)
# + [markdown] slideshow={"slide_type": "notes"}
# This emphasises the fact that subclasses of `Exception` often have additional contextually-relevant properties, which you should use. It also points out that `if` statements still have their place in error-handling!
#
# When `raise` has no argument, it re-raises whichever exception it was that got us into this mess.
# + [markdown] slideshow={"slide_type": "subslide"}
# # A few important Exceptions
# ## That you might want to catch
#
# * KeyError - `box_of_tricks["not here"]`
# * TypeError - `1 + "banana"`
# * IOError - `open('/etc/passwd', 'w')`
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Exceptions are often more *pythonic* than pre-checking. [This Python2 doc](https://docs.python.org/2/howto/doanddont.html#exceptions) is a good start on that road. CRUCIAL READING!
#
# + [markdown] slideshow={"slide_type": "notes"}
# ## We are near the end of the walk-through
#
# Feel free to go back and forward through this. I am going to give you one final task to get you started on this topic...
# + [markdown] slideshow={"slide_type": "slide"}
# Remember your line number from Etherpad? If not, it should be in variable `x`...
# -
print("I am on line number", x)
# + [markdown] slideshow={"slide_type": "notes"}
# But double-check Etherpad...
# + [markdown] slideshow={"slide_type": "notes"}
# Remember the post-its? So, everybody stick your star post-it somewhere I can see it...
# + [markdown] slideshow={"slide_type": "slide"}
# Your task is to write a Jupyter cell to give you the $x^{th}$ digit of $pi$ and **write the digit at the start of that Etherpad line**, the one with your name on it. In particular, try and do this, just using `x`, without typing the number in directly. Then you can experiment to see if other numbers work too.
#
# There are several ways to do this... I've given you enough tools above to find at least one. If, when you get it working, no-one has described your approach on Etherpad, write a short description at the bottom of the notepad.
#
# When done, swap your star for an arrow.
# +
# Overwrite this cell with your code and run it
# + [markdown] slideshow={"slide_type": "notes"}
# So, hopefully, you now have a calculator for the $x^{th}$ digit of $\pi$. How can you use this without going back and editing the cell?
# + [markdown] slideshow={"slide_type": "slide"}
# Well... copy and paste your text below the line in the cell below:
# -
def get_xth_digit_of_pi(x):
# + [markdown] slideshow={"slide_type": "fragment"}
# Now indent everything beneath the `def` line by four spaces and run it. You have just defined a function! If so, this should give you a `1`:
# -
get_xth_digit_of_pi(5)
# + [markdown] slideshow={"slide_type": "notes"}
# If you had errors that don't make sense, try the chat window, try someone beside you (my normal approach in life) or ask me.
# + [markdown] slideshow={"slide_type": "slide"}
# By the by, here are a couple of approaches - which is more Pythonic?
# ```
# 1.
# scaled_up = math.pi * (10 ** (x - 1))
# higher_digits = 10 * (scaled_up // 10)
# return int(scaled_up - higher_digits)
#
# 2.
# pi_as_str = str(math.pi)
# if x == 1:
# x = 0
# return pi_as_str[x]
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# # Classes
# + [markdown] slideshow={"slide_type": "fragment"}
# The basic syntax for classes is pretty succinct:
# -
class MyClass(object):
some_attribute = "default_value"
def __init__(self, some_attribute, arg2, arg3):
# Some initialization (over-riding default)
self.some_attribute = some_attribute
def some_method(self):
return self.some_attribute
def another_method(self, arg1):
if arg1 == "example":
output = self.some_method()
return output
# + [markdown] slideshow={"slide_type": "notes"}
# A few points to note: `class` is just a block like any other. Default values can be supplied at its top level, but be careful - often it is better to set defaults in the initializer. Methods are defined as functions, but inside the class block. All of them have a first argument `self`, which is the object itself. Like, say, `this` in C++ (but without dereferencing required).
#
# Python classes don't have a particularly strong approach to private/public access, but starting with an underscore is a conventional approach to indicate a member shouldn't be touched by anything outside the class. We subclass from `object` - additional classes may be added by comma-separation. Subclassing from `object` is actually the default in Python3 if no parentheses (or parent class) are provided - so-called *new-style classes*.
#
# You also see your first magic method - the `__init__` function. This is the constructor, called when the class is created. Here we use it to override a default from the arguments it is passed.
# + slideshow={"slide_type": "subslide"}
aninstance = MyClass("non-default value", 1, 2)
# + [markdown] slideshow={"slide_type": "notes"}
# Here, you see the arguments to the `__init__` function.
# + slideshow={"slide_type": "fragment"}
aninstance.another_method("example")
# + [markdown] slideshow={"slide_type": "notes"}
# And, one method has called the next and returned the value we asked. We can check the type and dir of this like any other object.
# + slideshow={"slide_type": "fragment"}
print(type(aninstance))
print(", ".join(dir(aninstance)))
# + [markdown] slideshow={"slide_type": "notes"}
# Note that you can call a override a method in a parent class simply by defining a method with the same name. If you want to effectively wrap it, you can use the format
#
# ```python
# class MyClass(myparentclass):
# ...
# def mymethod(self, arg1, arg2):
# val = myparentclass.mymethod(self, arg1, arg2)
# # Do something to val
# return val
# ```
#
# Note that you must explicitly pass "`self`" in this case.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Exercise
# + [markdown] slideshow={"slide_type": "notes"}
# ...if we have time...
# -
# Below, we subclass `str` and call it `MyString`. It has one method, called `join` to override the usual `join` functionality that strings have - otherwise, `MyString` strings behaves exactly as any other Python strings.
#
# Exercise: make this overridden `join` method check if the iterable you are joining has "Pottery" in it and, if so, make it raise a RuntimeError.
#
# Extension: combine this with your previous dictionary-to-strings function. Then update your dictionary-to-strings function to have a try-except block, which quietly swaps "Pottery" for "More Python" when the RuntimeError above error occurs (and then continues as normal).
# +
class MyString(str):
def join(self, iterable):
# Replace this with your Pottery-checker statement
# ...
# then continue as normal:
return str.join(self, iterable)
delimiter = MyString(",")
delimiter.join(my_meetup_dot_com_profile["Interests"])
# + [markdown] slideshow={"slide_type": "slide"}
# # What about all the other structures??
# + [markdown] slideshow={"slide_type": "fragment"}
# Well, they are not so hard once you get this far:
# + [markdown] slideshow={"slide_type": "notes"}
# but you'll come across them throughout today
# + [markdown] slideshow={"slide_type": "fragment"}
# * Loops can be `while [SOME BOOLEAN]:`, as well as for
# + [markdown] slideshow={"slide_type": "fragment"}
# * We will explore some actual modules after lunch
# + [markdown] slideshow={"slide_type": "fragment"}
# * Generators put loops inside lists
# + [markdown] slideshow={"slide_type": "notes"}
# I'll let that boggle your mind for a moment. Our future sessions are not going to be so closely led, partially because it *cannot* be as interesting experiment with my code as it is getting your hands dirty with your own. Now you've got plenty of tools at your disposal.
# + [markdown] slideshow={"slide_type": "slide"}
# # Finally...
# ## After a whistle-stop tour, we pull into a Python siding...
#
# * Please write one great thing about this morning on your arrow and leave it beside the door
# * Please write one dire (or just not so good) thing about this morning on your star and leave it beside the door
# + [markdown] slideshow={"slide_type": "notes"}
# Now, I'm going to do a quick tour of scripting applications of Python, a few ideas you might want to ask me about over break/lunch, to give you an idea of what Python can do, so sit back relax and let me fiddle with projector technology at the front for a couple of minutes...
# + [markdown] slideshow={"slide_type": "slide"}
# # PS: Tuples
# ## The Ice-List
# + [markdown] slideshow={"slide_type": "fragment"}
# Tuples are basically frozen lists...
# -
target_coordinates = (56, -5)
print(target_coordinates[0], "N", target_coordinates[1], "E")
# ...but it's fixed, so you can't do this...
# + slideshow={"slide_type": "fragment"}
target_coordinates[1] = 38
# + [markdown] slideshow={"slide_type": "notes"}
# If you alter it to set the whole variable (try it), that's fine though - you can alter what `target_coordinates` points to, you just can't alter the tuple itself.
# + [markdown] slideshow={"slide_type": "notes"}
# Lists are constantly changing - length, content, type of content. Really, they aren't much like arrays in compiled languages like C or FORTRAN. They are far too flightly to be used as something like a dictionary index for example. However, supose I have a pair - exactly two elements, one a known `int` and one a known `string`, and it can't change. Well, if you stick two basic types together then why can't you do the same thing with it you can do with basic types? There's no weird changing behaviour going on, so if you freeze a list into something like that, can you use it as a dictionary index?
# + [markdown] slideshow={"slide_type": "subslide"}
# In short - yes you can!
# -
battleships = {} # new dict
coordinates = [3, 2]
battleships[coordinates] = "HIT"
# + [markdown] slideshow={"slide_type": "fragment"}
# So it doesn't work with a list - try it now with a tuple - swap the square brackets on the *second* line for parens () and re-run
# + slideshow={"slide_type": "fragment"}
print("At", coordinates, "we have a", battleships[coordinates])
# + [markdown] slideshow={"slide_type": "fragment"}
# **Aside**: A *hashable* type can be used as an index in a dict - tuples are, lists aren't (because they can change, aka are *mutable*)
# + [markdown] slideshow={"slide_type": "slide"}
# # PS: an introduction to objects...
# + [markdown] slideshow={"slide_type": "subslide"}
# # Objects
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Totally class
# + [markdown] slideshow={"slide_type": "notes"}
# If you haven't done object oriented coding before, I am afraid I am not going to give you the magic bullet now. It is an important concept to get to grips with, but for the moment, we will keep it fairly functional. Do take time to look through it later - Python is an excellent language for playing around with it and getting an intuition for what an object is. A number of you will be very familiar with this so please bear with me and the very rough introductory description I will give - for those who are not so familiar, this is worth getting the basics down early.
#
# In real life, everything has properties, and many things have things they can do. In Python these properties are called attributes and the things an object can do are called methods. Together these are called members and, in Python, are accessed by putting a dot after the object and the attribute or method name. Everything else is just like any other variable or function, respectively.
#
# For instance, my dog Freddie has properties - he has a colour, which is black (technically, he is imaginary, but in my head he is definitely black). In Python terms, this would be:
# + [markdown] slideshow={"slide_type": "fragment"}
# **Attribute**: `freddie.colour = black`
# + [markdown] slideshow={"slide_type": "notes"}
# I can tell Freddie to roll-over. When I do so, I am, in some sense, calling Freddie's method:
# + [markdown] slideshow={"slide_type": "fragment"}
# **Method**: `freddie.roll_over()`
# + [markdown] slideshow={"slide_type": "notes"}
# I have another dog, Nitwit. Nitwit can also roll over...
# + [markdown] slideshow={"slide_type": "fragment"}
# **Method**: `nitwit.roll_over()`
# + [markdown] slideshow={"slide_type": "notes"}
# Both dogs can shake hands, but I need to tell them which paw...
# + [markdown] slideshow={"slide_type": "fragment"}
# **Method**: `freddie.shake_paw('left')`
# + [markdown] slideshow={"slide_type": "notes"}
# At this point, you're probably wondering what exactly *is* the set of attributes and methods that my dogs have? This template, showing what attributes and methods a dog of mine can be expected to have, is called a class.
# + [markdown] slideshow={"slide_type": "subslide"}
# # A simple class
# -
class PhilsDog:
name = ""
colour = ""
def shake_paw(self, side):
print("My name is", self.name, "and I am shaking my", side, "paw like a good dog")
# + [markdown] slideshow={"slide_type": "notes"}
# Here we create a class called `PhilsDog` that all of my dogs *implement* (that is, they are of that type). It indicates that they will have a colour and that they have a method called `shake_paw`. Think of this from the perspective of the dog - the first argument, `self`, is a little Python magic that refers to the dog itself. This lets the dog use it's name and colour (and any other methods) in the `shake_paw` method. The second argument is which paw I told my dog to shake. In response, any dog of mine says "Shaking left/right paw like a good dog". That's quite impressive, but I'd rather they just learned to zip it and actually shake their paws instead.
# + [markdown] slideshow={"slide_type": "fragment"}
# Now, that's just the template for one of my dogs, so how do I use it?
# + slideshow={"slide_type": "fragment"}
freddie = PhilsDog()
freddie.name = "Freddie"
freddie.colour = "black"
nitwit = PhilsDog()
nitwit.name = "Nitwit"
nitwit.colour = "brown"
print("Freddie is", freddie.colour, "while Nitwit is", nitwit.colour)
# + [markdown] slideshow={"slide_type": "notes"}
# I call the class like a function. This creates a new PhilsDog object - in computing terminology, I **instantiate** the class, creating a new **instance** of PhilsDog. Freddie is an instance of PhilsDog and so is Nitwit. But PhilsDog is just a template - as you can see, both Freddie and Nitwit have their own colour and name. I can update these just as any variable, and Freddie's doesn't affect Nitwit's, and I can read both back out again.
# + slideshow={"slide_type": "subslide"}
freddie.shake_paw('left')
nitwit.shake_paw('right')
# + [markdown] slideshow={"slide_type": "fragment"}
# Good dogs. Here we see the `shake_paw` method being called for each dog and with a different `side` parameter. To remind you, the body of the method was:
#
# ```python
# def shake_paw(self, side):
# print("My name is", self.name, "and I am shaking my", side, "paw like a good dog")
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# You can see how the `self.name` matches the name I gave each dog on the previous slide.
#
# If that was familiar to you, then that was probably a very unexciting few minutes - if not, it's probably come and gone very quickly. Thankfully, we only need to know that, if we have an object, we can get its attributes and methods by adding a dot and the attribute/method name.
#
# Now why is that important...
# + [markdown] slideshow={"slide_type": "subslide"}
# # In Python
# ## Everything is an object (nearly)
# + slideshow={"slide_type": "subslide"}
"just a normal string".upper()
# + slideshow={"slide_type": "fragment"}
"Another string".islower()
# + [markdown] slideshow={"slide_type": "subslide"}
# # Two useful tools
# + slideshow={"slide_type": "fragment"}
type(freddie)
# + [markdown] slideshow={"slide_type": "notes"}
# "`type`" lets you examine what class an object is. Don't worry about that `__main__` just for the moment.
# + slideshow={"slide_type": "fragment"}
dir(freddie)[-3:]
# + [markdown] slideshow={"slide_type": "notes"}
# This is a list of all the members of `freddie`. If he hasn't had any added on the fly (which is possible in Python), this is the same as the members of the `PhilsDog` class. That syntax in the brackets is coming up in a couple of slides, but basically it means, the last three items. Python denotes somewhat magic functions with double-underscores on either side (they are all the previous items I'm hiding away) - there is good reason for these being here, but they aren't essential just now. Try removing the bit in brackets, including the brackets, to see what you get.
| .ipynb_checkpoints/Basic control structures 2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Programming Assignment: Data Science Basics
# -
# In this programming assignment you need to apply your new `numpy`, `pandas` and `matplotlib` knowledge. You will need to do several [`groupby`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html)s and [`join`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.join.html)`s to solve the task.
# + slideshow={"slide_type": "fragment"}
# Import Packages
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Pandas allows you to load the required data sets directly from github:
# + slideshow={"slide_type": "fragment"}
# Load Data
transactions = pd.read_csv('https://github.com/NikoStein/pds_data/raw/main/data/sales_train.csv.gz')
items = pd.read_csv('https://github.com/NikoStein/pds_data/raw/main/data/items.csv')
item_categories = pd.read_csv('https://github.com/NikoStein/pds_data/raw/main/data/item_categories.csv')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Get to know the data
# Print the **shape** of the loaded dataframes and use [`df.head`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.head.html) function to print several rows. Examine the features you are given.
# +
# Write your code here
# + [markdown] slideshow={"slide_type": "slide"}
# ## Maximum total revenue among all the shops
#
# Now use your `pandas` skills to get answers for the following questions.
# The first question is:
#
# What was the maximum total revenue among all the shops in July, 2013?
#
# * Hereinafter *revenue* refers to total sales minus value of goods returned.
#
# *Hints:*
#
# * Sometimes items are returned, find such examples in the dataset.
# * It is handy to split `date` field into [`day`, `month`, `year`] components and use `df.year == 13` and `df.month == 6` in order to select target subset of dates.
# * You may work with `date` feature as with strings, or you may first convert it to `pd.datetime` type with `pd.to_datetime` function, but do not forget to set correct `format` argument.
# +
# Write your code here
# -
max_revenue = # Save the final value in this variable
# In addition we can find the index (shop id) of the row with the max value in the column using ```idxmax()```
# +
# Write your code here
# + [markdown] slideshow={"slide_type": "slide"}
# ## Constant price
#
# How many items are there, such that their price stays constant (to the best of our knowledge) during the whole period of time?
#
# * Let's assume, that the items are returned for the same price as they had been sold.
# +
# Write your code here
# -
num_items_constant_price = # Save the final value in this variable
# + [markdown] slideshow={"slide_type": "slide"}
# ## Variance of sold items per day
#
# What was the variance of the number of sold items per day sequence for the shop with `shop_id = 25` in December, 2014?
#
# * Do not count the items that were sold but returned back later.
# * Fill `total_num_items_sold`: An (ordered) array that contains the total number of items sold on each day
# * Then compute variance of the of `total_num_items_sold`
# * If there were no sales at a given day, ***do not*** impute missing value with zero, just ignore that day
# +
# Write your code here
# -
total_num_items_sold = # Save the final value in this variable
total_num_items_sold_var = # Save the final value in this variable
# + [markdown] slideshow={"slide_type": "slide"}
# ## Vizualization of the daily revenue
#
# Use `total_num_items_sold` to plot the daily revenue of `shop_id = 25` in December, 2014.
#
# * y-label: 'Num items'
# * x-label: 'Day'
# * plot-title: 'Daily revenue for shop_id = 25'
# * use plt.show() to display the plot in the end
# +
# Write your code here
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Final submission
# Only for the submission (do not change the code).
# -
print(max_revenue)
print(num_items_constant_price)
print(total_num_items_sold)
print(total_num_items_sold_var)
# ## Bonus
#
# What item category that generated the highest revenue in spring 2014?</b></li>
#
# * Submit the `id` of the category found.
#
# * Here, spring is the period from March to Mai.
# +
# Write your code here
# -
category_id_with_max_revenue = # Save the final value in this variable
print(category_id_with_max_revenue)
| assignments/01_PA_Data Science Basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# ## Load the model
from keras.models import Model, load_model
model = load_model('model.h5')
# ## Load and predict a random image
# +
from random import choice
from IPython.display import display, Image, clear_output
from keras.preprocessing.image import img_to_array, load_img
from keras.applications.imagenet_utils import preprocess_input, decode_predictions
import numpy as np
labels = ['vegan','dairy','meat']
def get_random_file(label, subset='test'):
path = 'data/' + subset + '/' + label
# files = !ls $path
return path + '/' + choice(files)
def get_image(img_path):
img = load_img(img_path)
img = img.resize((224,224))
return img
def predict_diet(img):
#the model expects a batch, so send it a batch with a size of 1
y_prob = model.predict(np.expand_dims(img, axis=0))
labels = ['dairy', 'meat', 'vegan']
return labels[y_prob.argmax(axis=-1)[0]]
true_label = choice(labels)
f = get_random_file(true_label)
img = get_image(f)
display(img)
print(f)
print('actual:',true_label)
print('prediction:',predict_diet(img))
# +
#save model structure to an image file
from keras.utils import plot_model
plot_model(model, to_file='model.png', show_shapes=True)
| predict_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# + [markdown] id="fOtTTDK8k7kG"
# Exercise 11 - Recurrent Neural Networks
# ========
#
# A recurrent neural network (RNN) is a class of neural network that excels when your data can be treated as a sequence - such as text, music, speech recognition, connected handwriting, or data over a time period.
#
# RNN's can analyse or predict a word based on the previous words in a sentence - they allow a connection between previous information and current information.
#
# This exercise looks at implementing a LSTM RNN to generate new characters after learning from a large sample of text. LSTMs are a special type of RNN which dramatically improves the model’s ability to connect previous data to current data where there is a long gap.
#
# We will train an RNN model using a novel written by <NAME> - The Time Machine.
# + [markdown] id="fqrEn9TKrB9M"
# Conexion a Google Drive
# + id="VB_YPfl7qswE" outputId="43359286-ac4f-45b7-e2a0-e8f2382fe86f" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="8V0ql8Phk7kH"
# Step 1
# ------
#
# Let's start by loading our libraries and text file. This might take a few minutes.
#
# #### Run the cell below to import the necessary libraries.
# + id="SqGMyFtZk7kI"
# %%capture
# Run this!
from keras.models import load_model
from keras.models import Sequential
from keras.layers import Dense, Activation, LSTM
from keras.callbacks import LambdaCallback, ModelCheckpoint
import numpy as np
import random, sys, io, string
# + [markdown] id="g3Vbpc78k7kN"
# #### Replace the `<addFileName>` with `The Time Machine`
# + id="R0JzyQybk7kO" outputId="276088ae-fa3b-48a2-bf5e-d6ed45a1125c" colab={"base_uri": "https://localhost:8080/"}
###
# REPLACE THE <addFileName> BELOW WITH The Time Machine
###
text = io.open('/content/drive/My Drive/azure/Data/The Time Machine.txt', encoding = 'UTF-8').read()
###
# Let's have a look at some of the text
print(text[0:198])
# This cuts out punctuation and make all the characters lower case
text = text.lower().translate(str.maketrans("", "", string.punctuation))
# Character index dictionary
charset = sorted(list(set(text)))
index_from_char = dict((c, i) for i, c in enumerate(charset))
char_from_index = dict((i, c) for i, c in enumerate(charset))
print('text length: %s characters' %len(text))
print('unique characters: %s' %len(charset))
# + [markdown] id="Q5316i4ak7kV"
# Expected output:
# ```The Time Traveller (for so it will be convenient to speak of him) was expounding a recondite matter to us. His pale grey eyes shone and twinkled, and his usually pale face was flushed and animated.
# text length: 174201 characters
# unique characters: 39```
#
# Step 2
# -----
#
# Next we'll divide the text into sequences of 40 characters.
#
# Then for each sequence we'll make a training set - the following character will be the correct output for the test set.
#
# ### In the cell below replace:
# #### 1. `<sequenceLength>` with `40`
# #### 2. `<step>` with `4`
# #### and then __run the code__.
# + id="ITWfM-Xnk7kW" outputId="29de0c1c-5f10-461c-c503-39f045f81188" colab={"base_uri": "https://localhost:8080/"}
###
# REPLACE <sequenceLength> WITH 40 AND <step> WITH 4
###
sequence_length = 40
step = 4
###
sequences = []
target_chars = []
for i in range(0, len(text) - sequence_length, step):
sequences.append([text[i: i + sequence_length]])
target_chars.append(text[i + sequence_length])
print('number of training sequences:', len(sequences))
# + [markdown] id="-sDAlc36k7kZ"
# Expected output:
# `number of training sequences: 43541`
#
# #### Replace `<addSequences>` with `sequences` and run the code.
# + id="5S8aeAaRk7ka"
# One-hot vectorise
X = np.zeros((len(sequences), sequence_length, len(charset)), dtype=np.bool)
y = np.zeros((len(sequences), len(charset)), dtype=np.bool)
###
# REPLACE THE <addSequences> BELOW WITH sequences
###
for n, sequence in enumerate(sequences):
###
for m, character in enumerate(list(sequence[0])):
X[n, m, index_from_char[character]] = 1
y[n, index_from_char[target_chars[n]]] = 1
# + [markdown] id="Tyv_tlpWk7kd"
# Step 3
# ------
#
# Let's build our model, using a single LSTM layer of 128 units. We'll keep the model simple for now, so that training does not take too long.
#
# ### In the cell below replace:
# #### 1. `<addLSTM>` with `LSTM`
# #### 2. `<addLayerSize>` with `128`
# #### 3. `<addSoftmaxFunction>` with `'softmax`
# #### and then __run the code__.
# + id="g_wHrXv4k7ke"
model = Sequential()
###
# REPLACE THE <addLSTM> BELOW WITH LSTM (use uppercase) AND <addLayerSize> WITH 128
###
model.add(LSTM(128, input_shape = (X.shape[1], X.shape[2])))
###
###
# REPLACE THE <addSoftmaxFunction> with 'softmax' (INCLUDING THE QUOTES)
###
model.add(Dense(y.shape[1], activation = 'softmax'))
###
model.compile(loss = 'categorical_crossentropy', optimizer = 'Adam')
# + [markdown] id="zo5UJnszk7ki"
# The code below generates text at the end of an epoch (one training cycle). This allows us to see how the model is performing as it trains. If you're making a large neural network with a long training time it's useful to check in on the model as see if the text generating is legible as it trains, as overtraining may occur and the output of the model turn to nonsense.
#
# The code below will also save a model if it is the best performing model, so we can use it later.
#
# #### Run the code below, but don't change it
# + id="M9DXb8Lpk7ki"
# Run this, but do not edit.
# It helps generate the text and save the model epochs.
# Generate new text
def on_epoch_end(epoch, _):
diversity = 0.5
print('\n### Generating text with diversity %0.2f' %(diversity))
start = random.randint(0, len(text) - sequence_length - 1)
seed = text[start: start + sequence_length]
print('### Generating with seed: "%s"' %seed[:40])
output = seed[:40].lower().translate(str.maketrans("", "", string.punctuation))
print(output, end = '')
for i in range(500):
x_pred = np.zeros((1, sequence_length, len(charset)))
for t, char in enumerate(output):
x_pred[0, t, index_from_char[char]] = 1.
predictions = model.predict(x_pred, verbose=0)[0]
exp_preds = np.exp(np.log(np.asarray(predictions).astype('float64')) / diversity)
next_index = np.argmax(np.random.multinomial(1, exp_preds / np.sum(exp_preds), 1))
next_char = char_from_index[next_index]
output = output[1:] + next_char
print(next_char, end = '')
print()
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
# Save the model
checkpoint = ModelCheckpoint('Models/model-epoch-{epoch:02d}.hdf5',
monitor = 'loss', verbose = 1, save_best_only = True, mode = 'min')
# + [markdown] id="6yHJfxNYk7ko"
# The code below will start the model to train. This may take a long time. Feel free to stop the training with the `square stop button` to the right of the `Run button` in the toolbar.
#
# Later in the exercise, we will load a pretrained model.
#
# ### In the cell below replace:
# #### 1. `<addPrintCallback>` with `print_callback`
# #### 2. `<addCheckpoint>` with `checkpoint`
# #### and then __run the code__.
# + id="ZsSjVzNrk7kp" outputId="da9611e7-e6ed-4068-9e2a-7238bea44fa4" colab={"base_uri": "https://localhost:8080/"}
###
# REPLACE <addPrintCallback> WITH print_callback AND <addCheckpoint> WITH checkpoint
###
model.fit(X, y, batch_size = 128, epochs = 3, callbacks = [print_callback, checkpoint])
###
# + [markdown] id="vP7THHvxk7ku"
# The output won't appear to be very good. But then, this dataset is small, and we have trained it only for a short time using a rather small RNN. How might it look if we upscaled things?
#
# Step 5
# ------
#
# We could improve our model by:
# * Having a larger training set.
# * Increasing the number of LSTM units.
# * Training it for longer
# * Experimenting with difference activation functions, optimization functions etc
#
# Training this would still take far too long on most computers to see good results - so we've trained a model already for you.
#
# This model uses a different dataset - a few of the King Arthur tales pasted together. The model used:
# * sequences of 50 characters
# * Two LSTM layers (512 units each)
# * A dropout of 0.5 after each LSTM layer
# * Only 30 epochs (we'd recomend 100-200)
#
# Let's try importing this model that has already been trained.
#
# #### Replace `<addLoadModel>` with `load_model` and run the code.
# + id="zT8ueupKk7ku" outputId="f3e2ea92-b9d2-431a-c53a-e6a361deb301" colab={"base_uri": "https://localhost:8080/"}
from keras.models import load_model
print("loading model... ", end = '')
###
# REPLACE <addLoadModel> BELOW WITH load_model
###
model = load_model('/content/drive/My Drive/azure/Models/arthur-model-epoch-30.hdf5')
###
model.compile(loss = 'categorical_crossentropy', optimizer = 'Adam')
###
print("model loaded")
# + [markdown] id="5jjCc8iAk7ky"
# Step 6
# -------
#
# Now let's use this model to generate some new text!
#
# #### Replace `<addFilePath>` with `'Data/Arthur tales.txt'`
# + id="bL2mIxdYk7ky" outputId="b0fc0fee-f9cb-47af-914a-f5af5f33458e" colab={"base_uri": "https://localhost:8080/"}
###
# REPLACE <addFilePath> BELOW WITH 'Data/Arthur tales.txt' (INCLUDING THE QUOTATION MARKS)
###
text = io.open('/content/drive/My Drive/azure/Data/Arthur tales.txt', encoding='UTF-8').read()
###
# Cut out punctuation and make lower case
text = text.lower().translate(str.maketrans("", "", string.punctuation))
# Character index dictionary
charset = sorted(list(set(text)))
index_from_char = dict((c, i) for i, c in enumerate(charset))
char_from_index = dict((i, c) for i, c in enumerate(charset))
print('text length: %s characters' %len(text))
print('unique characters: %s' %len(charset))
# + [markdown] id="VOUv8xI8k7k1"
# ### In the cell below replace:
# #### 1. `<sequenceLength>` with `50`
# #### 2. `<writeSentence>` with a sentence of your own, at least 50 characters long.
# #### 3. `<numCharsToGenerate>` with the number of characters you want to generate (choose a large number, like 1500)
# #### and then __run the code__.
# + id="RPfUDlC-k7k2" outputId="481b5d1b-42f7-40bf-b004-7d4cffce6305" colab={"base_uri": "https://localhost:8080/"}
# Generate text
diversity = 0.5
print('\n### Generating text with diversity %0.2f' %(diversity))
###
# REPLACE <sequenceLength> BELOW WITH 50
###
sequence_length = 50
###
# Next we'll make a starting point for our text generator
###
# REPLACE <writeSentence> WITH A SENTENCE OF AT LEAST 50 CHARACTERS
###
seed = " I finished work at 5 PM. I went home and took a shower"
###
seed = seed.lower().translate(str.maketrans("", "", string.punctuation))
###
# OR, ALTERNATIVELY, UNCOMMENT THE FOLLOWING TWO LINES AND GRAB A RANDOM STRING FROM THE TEXT FILE
###
#start = random.randint(0, len(text) - sequence_length - 1)
#seed = text[start: start + sequence_length]
###
print('### Generating with seed: "%s"' %seed[:40])
output = seed[:sequence_length].lower().translate(str.maketrans("", "", string.punctuation))
print(output, end = '')
###
# REPLACE THE <numCharsToGenerate> BELOW WITH THE NUMBER OF CHARACTERS WE WISH TO GENERATE, e.g. 1500
###
for i in range(50):
###
x_pred = np.zeros((1, sequence_length, len(charset)))
for t, char in enumerate(output):
x_pred[0, t, index_from_char[char]] = 1.
predictions = model.predict(x_pred, verbose=0)[0]
exp_preds = np.exp(np.log(np.asarray(predictions).astype('float64')) / diversity)
next_index = np.argmax(np.random.multinomial(1, exp_preds / np.sum(exp_preds), 1))
next_char = char_from_index[next_index]
output = output[1:] + next_char
print(next_char, end = '')
print()
# + [markdown] id="OW73pTfak7k5"
# How does it look? Does it seem intelligible?
#
# Conclusion
# --------
#
# We have trained an RNN that learns to predict characters based on a text sequence. We have trained a lightweight model from scratch, as well as imported a pre-trained model and generated new text from that.
| 11. Recurrent Neural Networks - Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get started
# Fill in your host name and get started exploring your catalog with the `datapath` API.
from deriva.core import ErmrestCatalog, get_credential
# Fill in your desired scheme, hostname and catalog number.
scheme = 'https'
hostname = None
catalog_number = 1
# Use DERIVA-Auth to get a `credential` or use `None` if your catalog allows anonymous access.
credential = None
#credential = get_credential(hostname)
# Now, connect to your catalog and the `pathbuilder` interface for the catalog.
assert scheme == 'http' or scheme == 'https', "Invalid http scheme used."
assert isinstance(hostname, str), "Hostname not set."
assert isinstance(catalog_number, int), "Invalid catalog number"
catalog = ErmrestCatalog(scheme, hostname, catalog_number, credential)
pb = catalog.getPathBuilder()
# Start using the pathbuilder, like this to get a list of your catalog's schema names.
list(pb.schemas)
| docs/get-started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jraval/DS-Unit-2-Kaggle-Challenge/blob/master/JAY_RAVAL_LS_DSPT9_224_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2XUX_-Nw6Ehb"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 4*
#
# ---
# + [markdown] id="nCc3XZEyG3XV"
# # Classification Metrics
#
# ## Assignment
# - [ ] If you haven't yet, [review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
# - [ ] Plot a confusion matrix for your Tanzania Waterpumps model.
# - [ ] Continue to participate in our Kaggle challenge. Every student should have made at least one submission that scores at least 70% accuracy (well above the majority class baseline).
# - [ ] Submit your final predictions to our Kaggle competition. Optionally, go to **My Submissions**, and _"you may select up to 1 submission to be used to count towards your final leaderboard score."_
# - [ ] Commit your notebook to your fork of the GitHub repo.
# - [ ] Read [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](http://archive.is/DelgE), by Lambda DS3 student <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
#
#
# ## Stretch Goals
#
# ### Reading
#
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
# - [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
#
#
# ### Doing
# - [ ] Share visualizations in our Slack channel!
# - [ ] RandomizedSearchCV / GridSearchCV, for model selection. (See module 3 assignment notebook)
# - [ ] Stacking Ensemble. (See module 3 assignment notebook)
# - [ ] More Categorical Encoding. (See module 2 assignment notebook)
# + id="lsbRiKBoB5RE"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="BVA1lph8CcNX"
import pandas as pd
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# + id="6LjzQO_E6Ehf"
# Import necessary modules
import category_encoders as ce
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import FunctionTransformer
from sklearn.ensemble import RandomForestClassifier
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import plot_confusion_matrix
# + id="t4eGLYrKMHa5"
def wrangle(X):
"""Wrangles train, validate, and test sets in the same way"""
X = X.copy()
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Drop duplicate columns
duplicate_columns = ['quantity_group']
X = X.drop(columns=duplicate_columns)
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these like null values
X['latitude'] = X['latitude'].replace(-2e-08, np.nan)
# When columns have zeros and shouldn't, they are like null values
cols_with_zeros = ['construction_year', 'longitude', 'latitude', 'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
return X
# + id="I63OvHLJNKoD"
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# Split train into train & val. Make val the same size as test.
target = 'status_group'
train, val = train_test_split(train, test_size=len(test),
stratify=train[target], random_state=42)
# Arrange data into X features matrix and y target vector
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
X_test = test
# + colab={"base_uri": "https://localhost:8080/"} id="AL_uF1TcNSBw" outputId="0b822d35-c1ed-4bb7-e442-fd5222d6f16d"
#Make pipeline!
pipeline = make_pipeline(
FunctionTransformer(wrangle, validate=False),
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
RandomForestClassifier(n_estimators=100, random_state=42, n_jobs=-1)
)
#Fit on train, score on val
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_val)
print('Validation Accuracy', accuracy_score(y_val, y_pred ))
# + colab={"base_uri": "https://localhost:8080/", "height": 389} id="ipujMdjBhDFH" outputId="0918e20e-b0b9-4abf-97ff-e0cb6658cd61"
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(pipeline, X_val, y_val, values_format='.0f', xticks_rotation='vertical');
# + [markdown] id="JWQxWnqZh8Ya"
# ## **How many correct predictions were made? Total Predictions? Classification Accuracy?**
# + colab={"base_uri": "https://localhost:8080/"} id="FoYxZHnxhZZd" outputId="18aeb9d6-193b-4040-b1e4-cb6fb63c5970"
correct_predictions = 7005 + 4351 + 332
correct_predictions
# + colab={"base_uri": "https://localhost:8080/"} id="1xSG3nWIifwv" outputId="a624f7dd-a3d2-4a7e-d80c-ab57b65e5926"
total_predictions = 7005 + 171 + 622 + 555 + 332 + 156 + 1098 + 68 + 4351
total_predictions
# + colab={"base_uri": "https://localhost:8080/"} id="qkfCOf0WjItj" outputId="08142110-35e7-42dd-d86f-1e7487407e68"
correct_predictions/total_predictions#Manual accuracy score
# + colab={"base_uri": "https://localhost:8080/"} id="3BRGB4h2jaXt" outputId="bc6ff04f-82d3-4bcb-9518-e36bf307c4d7"
accuracy_score(y_val, y_pred)
# + colab={"base_uri": "https://localhost:8080/"} id="c74x7m1JjuZg" outputId="14d9d919-63d5-4a0e-945e-3448278dd54a"
sum(y_pred == y_val)/ len(y_pred)
# + [markdown] id="GVBLWGHjkDOH"
# # Use classification metrics: precision, recall
# + [markdown] id="m3Sh1IlckPfg"
# ## Overview
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="3Y2lEExfWOm8" outputId="f63211fe-ce08-4836-ab84-8fe0299e830d"
pd.__version__
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="7ABIbcoCWb8m" outputId="346f27a3-ddc3-4d4d-fc22-32d3017fe245"
import sklearn
sklearn.__version__
# + id="X1L4pDCQ7iTR"
# Load the digits data
# The deafult with 10 classes (digits 0-9)
digits = datasets.load_digits(n_class=10)
# Create the feature matrix
X = digits.data
# Create the target array
y = digits.target
# Split the data into a training set and a test set
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Instantiate and train a decision tree classifier
dt_classifier = DecisionTreeClassifier().fit(X_train, y_train)
# + id="UCITooiTBacX"
# Load modules
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# Create the data (feature, target)
X, y = make_classification(n_samples=10000, n_features=5,
n_classes=2, n_informative=3,
random_state=42)
# Split the data into a training set and a test set
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create and fit the model
logreg_classifier = LogisticRegression().fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 493} id="kKlH37cB8Fux" outputId="afb9afec-ec37-4ac7-fb73-244b7e5231de"
# Plot the decision matrix
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1,1,figsize=(8,8))
plot_confusion_matrix(dt_classifier, X_test, y_test,
display_labels=digits.target_names,
cmap=plt.cm.Blues, ax=ax)
ax.set_title('Confusion Matrix: Digits decision tree model')
#fig.clf();
# + colab={"base_uri": "https://localhost:8080/"} id="yZ_U-Mb6GToa" outputId="261f3df0-1b35-43a6-bfb9-fee7d7f4570e"
# The value of the 10th observation
print('The 10th observation: ', X_test[10:11])
# Print out the probability for the 10th observation
print('Predicted probability for the 10th observation: ',
logreg_classifier.predict_proba(X_test)[10:11])
# Print the two classes
print('The two classes: ')
logreg_classifier.classes_
# + colab={"base_uri": "https://localhost:8080/"} id="tDOflQbcGs_v" outputId="77d0788a-f364-46cd-c753-206c483dec9a"
# Check the class of the 10th observation
logreg_classifier.predict(X_test[10:11])
# + id="io4qiwoZIpeJ"
# Load modules
import pandas as pd
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve
# Create the data (feature, target)
X, y = make_classification(n_samples=10000, n_features=5,
n_classes=2, n_informative=3,
random_state=42)
# Split the data into a training set and a test set
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# Create and fit the model
logreg_classifier = LogisticRegression().fit(X_train, y_train)
# Create predicted probabilities
y_pred_prob = logreg_classifier.predict_proba(X_test)[:,1]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="7sq94LMkI0-t" outputId="734f2b21-9b19-4548-9231-088aba9378a7"
# Create the data for the ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_pred_prob)
# See the results in a table
roccurve_df = pd.DataFrame({
'False Positive Rate': fpr,
'True Positive Rate': tpr,
'Threshold': thresholds
})
roccurve_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 312} id="QZCFkM6gJZyu" outputId="c48f24ba-c2e6-42b1-977d-3337a3d730f5"
# Plot the ROC curve
import matplotlib.pyplot as plt
plt.plot(fpr, tpr)
plt.plot([0,1], ls='--')
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
# + colab={"base_uri": "https://localhost:8080/"} id="9jQuo0dlJrvd" outputId="319b14ac-e51a-48c9-c2a6-fa2ca30498e9"
# Calculate the area under the curve
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_pred_prob)
# + id="Gm3WOGrtJuuQ"
| JAY_RAVAL_LS_DSPT9_224_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .download-project
# language: python
# name: .download-project
# ---
# +
import os
import sys
import time
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from src.data import load_annotation, load_data
from src.utils.train import train_test_split
import warnings
warnings.filterwarnings("ignore", category=RuntimeWarning)
# +
# config
DATAFRAME_PATH = "../data/raw/data_frames"
ANNOTATION_PATH = "../data/processed/Annotation.csv"
CATEGORY = "Defecation"
THRESHOLD = 0.3
FEATURE_NAMES = ['Min', 'Max', 'Median', 'Mean', 'Variance', 'LinearTrend']
SOURCES = ['TotalWeight', 'WaterDistance', 'RadarSum']
ANNOTATIONS = load_annotation.get_annotation(ANNOTATION_PATH)
USER_IDS = load_annotation.get_complete_ids(ANNOTATION_PATH, CATEGORY)
TRAIN_IDS, TEST_IDS = train_test_split(USER_IDS[USER_IDS < 2000])
print ("Training on {} cases, {} ...".format(len(TRAIN_IDS), TRAIN_IDS[:5]))
print ("Testing on {} cases, {} ...".format(len(TEST_IDS), TEST_IDS[:5]))
# +
rf_train_config = {
'USE_IDS': TRAIN_IDS,
'ANNOTATION_PATH': ANNOTATION_PATH,
'FEATURE_NAMES': FEATURE_NAMES,
'SOURCES': SOURCES,
'CATEGORY': CATEGORY
}
rf_test_config = {
'USE_IDS': TEST_IDS,
'ANNOTATION_PATH': ANNOTATION_PATH,
'FEATURE_NAMES': FEATURE_NAMES,
'SOURCES': SOURCES,
'CATEGORY': CATEGORY
}
dataset = {}
dataset['train'] = load_data.RandomForestDataset(rf_train_config)
dataset['test'] = load_data.RandomForestDataset(rf_test_config)
# -
train_x, train_y = dataset['train'].get_all_features_and_labels()
test_x, test_y = dataset['test'].get_all_features_and_labels()
train_x[train_x['TotalWeight_LogVariance'] == - np.inf]
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report, plot_roc_curve
rf = RandomForestClassifier(n_estimators = 5, max_features = 3)
rf.fit(train_x, train_y)
# +
import pickle
import time
timestr = time.strftime("%Y%m%d-%H%M%S")
with open(f"../randomforest-defecate-{timestr}.pkl", "wb") as f:
pickle.dump(rf, f)
# -
def classification_result(model, testX, testY, threshold = 0.5):
testYPredProb = model.predict_proba(testX)
testYPred = (testYPredProb[:, 1] > threshold).astype(int)
print (f"threshold = {threshold}", "\n")
print (classification_report(testY, testYPred))
classification_result(
model = rf,
testX = train_x,
testY = train_y,
threshold = 0.3
)
train_x
print (TRAIN_IDS)
print (TEST_IDS)
classification_result(
model = rf,
testX = test_x,
testY = test_y,
threshold = 0.5
)
def variable_importance(trainX, model):
plt.bar(x = range(trainX.shape[1]), height = model.feature_importances_)
xticks_pos = np.arange(trainX.shape[1])
plt.xticks(xticks_pos, trainX.columns, rotation=45, ha = 'right')
pass
# variable importance
plt.figure(figsize = (12, 6))
variable_importance(train_x, rf)
| notebooks/archive/002-run_randomforest_defecate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: '''Python Interactive'''
# language: python
# name: b8ef8e07-421c-4aa3-9034-7419e8c5e615
# ---
# # Network3 #
# ## Preliminaries ##
# %conda install theano
# %pip install mnist
# +
import numpy as np
import gzip
import _pickle
from src.nndl.network3 import Network, FullyConnectedLayer, SoftmaxLayer
import matplotlib.pyplot as plt
import theano
import theano.tensor as T
with gzip.open('./data/mnist.pkl.gz', 'rb') as fp:
training_inputs, training_results, testing_inputs, testing_results = \
_pickle.load(fp)
def shared(data):
shared_x = theano.shared(
np.asarray(data[0], dtype=theano.config.floatX),
borrow=True)
shared_y = theano.shared(
np.asarray(data[1], dtype=theano.config.floatX),
borrow=True)
return shared_x, T.cast(shared_y, "int32")
training_data = shared((training_inputs, training_results))
testing_data = shared((testing_inputs, testing_results))
networks = []
monitor = []
# -
training_x, training_y = training_data
training_y[0]
# ## Network 1 `[784, 100, 10]` ##
networks.append(
Network([FullyConnectedLayer(n_in=784, n_out=100),
SoftmaxLayer(n_in=100, n_out=10)],
10)
)
net = networks[0]
net.stochastic_gradient_descent(training_data, 60, 10, 0.1, testing_data)
# ## Network 2 `[20x1x5x5, 2880, 100, 10]` ##
networks.append(
Network([ConvolutionalPoolLayer(image_shape=(10, 1, 28, 28),
filter_shape=(20, 1, 5, 5),
poolsize=(2, 2)),
FullyConnectedLayer(n_in=2880, n_out=100),
SoftmaxLayer(n_in=100, n_out=10)], 10)
)
net = networks[1]
net.stochastic_gradient_descent(training_data, 60, 10, 0.1, test_data)
# ## Network 3 `[20x1x5x5, 40x20x5x5, 640, 100, 10]` ##
networks.append(
Network([
ConvolutionalPoolLayer(image_shape=(10, 1, 28, 28),
filter_shape=(20, 1, 5, 5),
poolsize=(2, 2)),
ConvolutionalPoolLayer(image_shape=(10, 20, 12, 12),
filter_shape=(40, 20, 5, 5),
poolsize=(2, 2)),
FullyConnectedLayer(n_in=640, n_out=100),
SolftmaxLayer(n_in=100, n_out=10)], 10)
)
net = networks[2]
net.stochastic_gradient_descent(training_data, 60, 10, 0.1, test_data)
# ## Network 4 `[20x1x5x5, 40x20x5x5, 640, 100, 10]` with ReLU Activations ##
networks.append(
Network([
ConvolutionalPoolLayer(image_shape=(10, 1, 28, 28),
filter_shape=(20, 1, 5, 5),
poolsize=(2, 2),
activation_fn=ReLU),
ConvolutionalPoolLayer(image_shape=(10, 20, 12, 12),
filter_shape=(40, 20, 5, 5),
poolsize=(2, 2),
activation_fn=ReLU),
FullyConnectedLayer(n_in=640, n_out=100, activation_fn=ReLU),
SoftmaxLayer(n_in=100, n_out=10)
], 10)
)
net = networks[3]
net.stochastic_gradient_descent(training_data, 60, 10, 0.03, test_data,
lambda_=0.1)
# ## Network 5 `[20x1x5x5, 40x20x5x5, 640, 1000, 1000, 10]` with ReLU Activations and Dropout ##
networks.append(
Network([
ConvolutionalPoolLayer(image_shape=(10, 1, 28, 28),
filter_shape=(20, 1, 5, 5),
poolsize=(2, 2),
activation_fn=ReLU),
ConvolutionalPoolLayer(image_shape=(10, 20, 12, 12),
filter_shape=(40, 20, 5, 5),
poolsize=(2, 2),
activation_fn=ReLU),
FullyConnectedLayer(n_in=640, n_out=1000, activation_fn=ReLU,
p_dropout = 0.5),
FullyConnectedLayer(n_in=1000, n_out=1000, activation_fn=ReLU,
p_dropout = 0.5),
SoftmaxLayer(n_in=1000, n_out=10, p_dropout = 0.5)
], 10)
)
net = networks[4]
net.stochastic_gradient_descent(training_data, 60, 10, 0.03, test_data)
| Network3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
# +
# Declaration des constantes
DATA_DIR = '../data/external/'
PROCESS_DIR = '../data/processed/'
CITIES_FILENAME = 'base-cc-filosofi-2015.xls'
# -
filo_df = pd.read_excel(os.path.join(DATA_DIR, CITIES_FILENAME), skiprows=5)
filo_df.head()
filo_df.NBPERSMENFISC15.isnull().sum()
| notebooks/preparation_filosofi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 3: Joining dataframes with `cptac`
#
# In this tutorial, we provide several examples of how to use the built-in `cptac` functions for joining different dataframes.
import cptac
cptac.download(dataset="endometrial", version="latest")
en = cptac.Endometrial()
# ## General format
#
# In all of the join functions, you specify the dataframes you want to join by passing their names to the appropriate parameters in the function call. The function will automatically check that the dataframes whose names you provided are valid for the join function, and print an error message if they aren't.
#
# Whenever a column from an -omics dataframe is included in a joined table, the name of the -omics dataframe it came from is joined to the column header, to avoid confusion.
#
# If you wish to only include particular columns in the join, pass them to the appropriate parameters in the join function. All such parameters will accept either a single column name as a string, or a list of column name strings. In this use case, we will usually only select specific columns for readability, but you could select the whole dataframe in all these cases, except for the mutations dataframe.
#
# The join functions use logic analogous to an SQL INNER JOIN.
# ## `join_omics_to_omics`
#
# The `join_omics_to_omics` function joins two -omics dataframes to each other. Types of -omics data valid for use with this function are acetylproteomics, CNV, phosphoproteomics, phosphoproteomics_gene, proteomics, and transcriptomics.
prot_and_phos = en.join_omics_to_omics(df1_name="proteomics", df2_name="phosphoproteomics")
prot_and_phos.head()
# Joining only specific columns.
# (Note that when a gene is selected from the phosphoproteomics dataframe, data for all sites of the gene are selected. The same is done for acetylproteomics data.)
# +
prot_and_phos_selected = en.join_omics_to_omics(
df1_name="proteomics",
df2_name="phosphoproteomics",
genes1="A1BG",
genes2="PIK3CA")
prot_and_phos_selected.head()
# -
# ## `join_metadata_to_omics`
#
# The `join_metadata_to_omics` function joins a metadata dataframe (e.g. clinical or derived_molecular) with an -omics dataframe:
clin_and_tran = en.join_metadata_to_omics(metadata_df_name="clinical", omics_df_name="transcriptomics")
clin_and_tran.head()
# Joining only specific columns:
# +
clin_and_tran = en.join_metadata_to_omics(
metadata_df_name="clinical",
omics_df_name="transcriptomics",
metadata_cols = ["Age", "Histologic_type"],
omics_genes="ZZZ3")
clin_and_tran.head()
# -
# ## `join_metadata_to_metadata`
#
# The `join_metadata_to_metadata` function joins two metadata dataframes (e.g. clinical or derived_molecular) to each other. Note how we passed a column name to select from the clinical dataframe, but passing `None` for the column parameter for the derived_molecular dataframe caused the entire dataframe to be selected. We could have omitted the `cols2` parameter altogether, as it is assigned to None by default.
# +
hist_and_derived_molecular = en.join_metadata_to_metadata(
df1_name="clinical",
df2_name="derived_molecular",
cols1="Histologic_type") # Note that we can omit the cols2 parameter, and it will by default select all of df2.
# We could have also omitted cols1, if we wanted to select all of df1.
hist_and_derived_molecular.head()
# -
# ## `join_omics_to_mutations`
#
# The `join_omics_to_mutations` function joins an -omics dataframe with the mutation data for a specified gene or genes. Because there may be multiple mutations for one gene in a single sample, the mutation type and location data are returned in lists by default, even if there is only one mutation. If there is no mutation for the gene in a particular sample, the list contains either "Wildtype_Tumor" or "Wildtype_Normal", depending on whether it's a tumor or normal sample. The mutation status column contains either "Single_mutation", "Multiple_mutation", "Wildtype_Tumor", or "Wildtype_Normal", for help with parsing.
#
# (Note: You can hide the Location columns by passing `False` to the optional `show_location` parameter.)
# +
selected_acet_and_PTEN_mut = en.join_omics_to_mutations(
omics_df_name="proteomics",
mutations_genes="PTEN",
omics_genes=["AURKA", "TP53"])
selected_acet_and_PTEN_mut.head(10)
# -
# ### Filtering multiple mutations
#
# The function has the ability to filter multiple mutations down to just one mutation. It allows you to specify particular mutation types or locations to prioritize, and also provides a default sorting hierarchy for all other mutations. The default hierarchy chooses truncation mutations over missense mutations, and silent mutations last of all. If there are multiple mutations of the same type, it chooses the mutation occurring earlier in the sequence.
#
# To filter all mutations based on this default hierarchy, simply pass an empty list to the optional `mutations_filter` parameter. Notice how in sample S001, the nonsense mutation was chosen over the missense mutation, because it's a type of trucation mutation, even though the missense mutation occurs earlier in the peptide sequence. In sample S008, both mutations were types of truncation mutations, so the function just chose the earlier one.
PTEN_default_filter = en.join_omics_to_mutations(omics_df_name="proteomics", mutations_genes="PTEN",
omics_genes=["AURKA", "TP53"],
mutations_filter=[])
PTEN_default_filter.loc[["C3L-00006", "C3L-00137"]]
# To prioritize a particular type of mutation, or a particular location, include it in the `mutations_filter` list. Below, we tell the function to prioritize nonsense mutations over all other mutations. Notice how in sample S008, the nonsense mutation is now selected instead of the frameshift insertion, even though the nonsense mutation occurs later in the peptide sequence.
PTEN_simple_filter = en.join_omics_to_mutations(omics_df_name="proteomics", mutations_genes="PTEN",
omics_genes=["AURKA", "TP53"],
mutations_filter=["Nonsense_Mutation"])
PTEN_simple_filter.loc[["C3L-00006", "C3L-00137"]]
# You can include multiple mutation types and/or locations in the `mutations_filter` list. Values earlier in the list will be prioritized over values later in the list. For example, with the filter we specify below, the function first selects sample S001's missense mutation over its nonsense mutation, because we put the location of S001's missense mutation as the first value in our filter list. We still included Nonsense_Mutation in the filter list, but it comes after the location of S001's missense mutation, which is why S001's missense mutation is still prioritized. However, on all other samples, unless they also have a mutation at that same location, the function will continue prioritizing nonsense mutations, as we see in sample S008.
PTEN_complex_filter = en.join_omics_to_mutations(omics_df_name="proteomics", mutations_genes="PTEN",
omics_genes=["AURKA", "TP53"],
mutations_filter=["p.R130Q", "Nonsense_Mutation"])
PTEN_complex_filter.loc[["C3L-00006", "C3L-00137"]]
# ## `join_metadata_to_mutations`
#
# The `join_metadata_to_mutations` function works exactly like `join_omics_to_mutations`, except that it works with metadata dataframes (e.g. clinical and derived molecular) instead of omics dataframes. It also can filter multiple mutations, which you control through the `mutations_filter` parameter, and has the ability to hide the location colunms.
# +
hist_and_PTEN = en.join_metadata_to_mutations(
metadata_df_name="clinical",
mutations_genes="PTEN",
metadata_cols="Histologic_type")
hist_and_PTEN.head()
# -
# With multiple mutations filtered:
# +
hist_and_PTEN = en.join_metadata_to_mutations(
metadata_df_name="clinical",
mutations_genes="PTEN",
metadata_cols="Histologic_type",
mutations_filter=["Nonsense_Mutation"])
hist_and_PTEN.head()
# -
# # Exporting dataframes
#
# If you wish to export a dataframe to a file, simply call the dataframe's to_csv method, passing the path you wish to save the file to, and the value separator you want:
hist_and_PTEN.to_csv(path_or_buf="histologic_type_and_PTEN_mutation.tsv", sep='\t')
| notebooks/tutorial03_joining_dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
fig = plt.figure()
ax = Axes3D(fig)
#列出实验数据
point=[[2,3,48],[4,5,50],[5,7,51],[8,9,55],[9,12,56]]
plt.xlabel("X1")
plt.ylabel("X2")
#表示矩阵中的值
ISum = 0.0
X1Sum = 0.0
X2Sum = 0.0
X1_2Sum = 0.0
X1X2Sum = 0.0
X2_2Sum = 0.0
YSum = 0.0
X1YSum = 0.0
X2YSum = 0.0
#在图中显示各点的位置
for i in range(0,len(point)):
x1i=point[i][0]
x2i=point[i][1]
yi=point[i][2]
ax.scatter(x1i, x2i, yi, color="red")
show_point = "["+ str(x1i) +","+ str(x2i)+","+str(yi) + "]"
ax.text(x1i,x2i,yi,show_point)
ISum = ISum+1
X1Sum = X1Sum+x1i
X2Sum = X2Sum+x2i
X1_2Sum = X1_2Sum+x1i**2
X1X2Sum = X1X2Sum+x1i*x2i
X2_2Sum = X2_2Sum+x2i**2
YSum = YSum+yi
X1YSum = X1YSum+x1i*yi
X2YSum = X2YSum+x2i*yi
# 进行矩阵运算
# _mat1 设为 mat1 的逆矩阵
m1=[[ISum,X1Sum,X2Sum],[X1Sum,X1_2Sum,X1X2Sum],[X2Sum,X1X2Sum,X2_2Sum]]
mat1 = np.matrix(m1)
m2=[[YSum],[X1YSum],[X2YSum]]
mat2 = np.matrix(m2)
_mat1 =mat1.getI()
mat3 = _mat1*mat2
# 用list来提取矩阵数据
m3=mat3.tolist()
a0 = m3[0][0]
a1 = m3[1][0]
a2 = m3[2][0]
# 绘制回归线
x1 = np.linspace(0,9)
x2 = np.linspace(0,12)
y = a0+a1*x1+a2*x2
ax.plot(x1,x2,y)
show_line = "y="+str(a0)+"+"+str(a1)+"x1"+"+"+str(a2)+"x2"
plt.title(show_line)
plt.show()
# -
plt.show()
import xlrd
import pandas as pd
df = pd.read_excel("./3d.xlsx")
print(df)
# +
# coding= utf-8
import numpy as np
import tensorflow as tf
import time
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# 参数、迭代阈值
w1 = 2.4
w2 = 8.8
b = 0.5
threshold = 0.00001
# 1.生成数据
# 这里需要强制转换成float32,否则后面在matmul时会提示float32和float64类型不匹配
# 这里的x_data包含x和y坐标
x_coor = df['R1']
y_coor = df['R2']
x_data = np.float32(np.zeros([2, 22]))
x_data[0, :] = df['R1']
x_data[1, :] = df['R2']
y_data = df['G2']
ax = plt.figure().add_subplot(111, projection='3d')
ax.scatter(x_coor, y_coor, y_data, c='r', marker='o')
# 显示图像
plt.show()
# 2.定义模型变量
# random_uniform用于返回一个指定大小,数值介于指定范围的矩阵
W = tf.Variable(tf.random_uniform([1, 2], -10, 10))
# 这里b的shape是(1,),并不是1×1,第二维是任意大小,所以才能在下面和1×100的矩阵相加
b = tf.Variable(tf.zeros([1]))
y = tf.matmul(W, x_data) + b
# 3.定义评价指标
# 采用方差的定义,求取平均方差
loss = tf.reduce_mean(tf.square(y - y_data))
# 4.构造运行图
optimizer = tf.train.GradientDescentOptimizer(0.5)
train = optimizer.minimize(loss)
# 5.启动图
init = tf.global_variables_initializer()
sess = tf.Session()
# 运行之前必须要先运行这行代码,进行初始化
sess.run(init)
t1 = time.time()
count = 0
while sess.run(loss) > threshold:
sess.run(train)
count += 1
# 注意,直接打印loss并不是它的内容,必须sess.run(loss)打印出来的才是它的数值
# 因为我们在建立graph的时候,只建立tensor的结构形状信息,并没有执行数据的操作
print(count, sess.run(loss), sess.run(W), sess.run(b))
t2 = time.time()
print(t2 - t1, "seconds.")
# +
import pandas as pd
def excel_one_line_to_list(i):
df = pd.read_excel("./3d.xlsx", usecols=[i],names=None) # 读取项目名称列,不要列名
df_li = df.values.tolist()
result = []
for s_li in df_li:
result.append(s_li[0])
return result
r1=[]
g1=[]
r2=[]
g2=[]
if __name__ == '__main__':
r1=excel_one_line_to_list(0)
g1=excel_one_line_to_list(1)
r2=excel_one_line_to_list(2)
g2=excel_one_line_to_list(3)
# -
r1
# +
X=[]
for i in range(len(r1)):
X.append([r1[i],g1[i]])
print(X)
Y=[]
for i in range(len(r1)):
Y.append([r2[i],g2[i]])
print(Y)
# +
#实验一
# 方法一:use keras to get weight in learning process
from keras.models import Sequential
from keras.layers import Dense
model = Sequential()
model.add(Dense(input_dim=2, units=2))
model.compile(loss='mse', optimizer='adam')
for step in range(20000):
cost = model.train_on_batch(X, Y)
if step % 1000 == 0:
print("After %d trainings, the cost: %f" % (step, cost))
print('\nTesting ------------')
cost = model.evaluate(X, Y, batch_size=40)
print('test cost:', cost)
W, b = model.layers[0].get_weights()
print('Weights=', W, '\nbiases=', b)
# -
#最小二乘法线性回归
from sklearn import linear_model
model = linear_model.LinearRegression()
model.fit(X,Y)
print(model.score(X,Y))
print('W='+str(model.coef_))
print('b='+str(model.intercept_))
# +
import numpy as np
X=[]
for i in range(len(r1)):
X.append([r1[i],g1[i]])
X = np.array(X)
Y=[]
for i in range(len(r1)):
Y.append([r2[i],g2[i]])
Y = np.array(Y)
# +
from sklearn import linear_model
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from sklearn.preprocessing import PolynomialFeatures
polynomial = PolynomialFeatures(degree = 3)
X_transformed = polynomial.fit_transform(X)#x每个数据对应的多项式系数
regr = linear_model.LinearRegression()
regr.fit(X_transformed, Y)#训练数据
# regr.fit(X, Y)
Y_hat = regr.predict(X_transformed)
# +
import numpy as np
X=[]
for i in range(len(r1)):
X.append([r1[i],g1[i]])
X = np.array(X)
Y=[]
for i in range(len(r1)):
Y.append([r2[i],g2[i]])
Y = np.array(Y)
# -
import numpy as np
x = np.hsplit(X,2)[0]
y = np.hsplit(X,2)[1]
z = np.hsplit(Y_hat,2)[0]
xx,yy = np.meshgrid(x,y)
print(x.shape)
print(y.shape)
print(z.shape)
print(Y_hat.shape)
# +
from sklearn import linear_model
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from sklearn.preprocessing import PolynomialFeatures
polynomial = PolynomialFeatures(degree = 3)
X_transformed = polynomial.fit_transform(X)#x每个数据对应的多项式系数
regr = linear_model.LinearRegression()
regr.fit(X_transformed, Y)#训练数据
# regr.fit(X, Y)
Y_hat = regr.predict(X_transformed)
# -
z_ = regr.predict(polynomial.fit_transform(np.concatenate([xx.reshape(22*22,1),yy.reshape(22*22,1)],axis = 1)))
fig = plt.figure()
ax = plt.gca(projection='3d') # 返回的对象就是导入的axes3d类型对象
plt.title('r1,g1<->r2', fontsize=20)
ax.set_xlabel('r1', fontsize=14)
ax.set_ylabel('g1', fontsize=14)
ax.set_zlabel('r2', fontsize=14)
plt.tick_params(labelsize=10)
ax.scatter(xx,yy,np.hsplit(z_,2)[0],cmap=plt.cm.hot)
surf = ax.plot_surface(xx, yy, np.hsplit(z_,2)[0].reshape(22,22), rstride=10, cstride=10, cmap=plt.cm.hot,alpha=0.8)
fig.colorbar(surf, shrink=0.8, aspect=6)
plt.show()
fig = plt.figure()
ax = plt.gca(projection='3d') # 返回的对象就是导入的axes3d类型对象
plt.title('r1,g1<->g2', fontsize=20)
ax.set_xlabel('r1', fontsize=14)
ax.set_ylabel('g1', fontsize=14)
ax.set_zlabel('g2', fontsize=14)
plt.tick_params(labelsize=10)
ax.scatter(xx,yy,np.hsplit(z_,2)[1],cmap=plt.cm.hot)
surf = ax.plot_surface(xx, yy, np.hsplit(z_,2)[1].reshape(22,22), rstride=10, cstride=10, cmap=plt.cm.hot)
fig.colorbar(surf, shrink=0.8, aspect=6)
plt.show()
def depress(data):
for i in data:
for j in range(len(i)):
if i[j] > 255:
i[j] = 255
return data
#实验五
#给定RGB值,预测测得RGB值
import math
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size=0.3,random_state=10)
model = linear_model.LinearRegression()
model.fit(X_train,Y_train)
result = depress(model.predict(X_test))#depress value > 255
print("RMSE:"+str(math.sqrt(np.sum((Y_test-result)**2)/result.size)))
| Lab2/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
# %matplotlib inline
import matplotlib.pyplot as plt
import librosa
data = pd.read_csv('features_30_sec.csv')
data.head()
# +
# music_genres = 'blues classical country disco hiphop jazz metal pop reggae rock'.split()
# -
data[data['label']=='jazz'][0:10]
dataset = data[data['label'].isin(['blues', 'classical', 'country', 'disco', 'hiphop', 'jazz', 'metal', 'pop', 'reggae', 'rock'])].drop(['filename','length'],axis=1)
dataset.shape
# +
# K Neighbors Classification
# -
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, StandardScaler,MinMaxScaler
from sklearn import preprocessing
from sklearn.neighbors import KNeighborsClassifier
y = LabelEncoder().fit_transform(dataset.iloc[:,-1])
y.shape
X = MinMaxScaler().fit_transform(np.array(dataset.iloc[:, :-1], dtype = float))
print(X)
X.shape
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.20,random_state=42)
print ('Train set:', X_train.shape, y_train.shape)
print ('Test set:', X_test.shape, y_test.shape)
knn = KNeighborsClassifier(n_neighbors=6,weights='distance',metric='minkowski',p=2)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
from sklearn.metrics import classification_report,confusion_matrix
print(classification_report(y_test,pred))
print(y_test)
print(pred)
cf_matrix = confusion_matrix(y_test,pred)
print(cf_matrix)
import seaborn as sns
# %matplotlib inline
classes=['blues', 'classical', 'country', 'disco', 'hiphop', 'jazz', 'metal', 'pop', 'reggae', 'rock']
sns.heatmap(cf_matrix, annot=True , cmap='Blues',xticklabels=classes,yticklabels=classes)
# +
#LogisticRegression
# -
X = StandardScaler().fit_transform(np.array(dataset.iloc[:, :-1], dtype = float))
y = LabelEncoder().fit_transform(dataset.iloc[:,-1])
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.30,random_state=42)
from sklearn.linear_model import LogisticRegression
LR = LogisticRegression(C=0.12, solver='lbfgs', verbose=0 ,multi_class='auto').fit(X_train,y_train)
LR
y_pred = LR.predict(X_test)
print (classification_report(y_test, y_pred))
from sklearn.metrics import jaccard_similarity_score
jaccard_similarity_score(y_test, y_pred)
# +
#Support Vector Machines
# -
from sklearn import svm
X = StandardScaler().fit_transform(np.array(dataset.iloc[:, :-1], dtype = float))
y = LabelEncoder().fit_transform(dataset.iloc[:,-1])
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.20,random_state=42)
rbf = svm.SVC(kernel='rbf',degree=1,decision_function_shape='ovo' ).fit(X_train, y_train)
poly = svm.SVC(kernel='poly', degree=1,decision_function_shape='ovo' ).fit(X_train, y_train)
poly_pred = poly.predict(X_test)
rbf_pred = rbf.predict(X_test)
print("Poly_pred:")
print (classification_report(y_test, poly_pred))
print("Rbf_pred:")
print (classification_report(y_test, rbf_pred))
# +
# ax = dataset[dataset['label'] == 'jazz'][0:50].plot(kind='scatter', x='chroma_stft', y='rmse', color='DarkBlue', label='jazz');
# dataset[dataset['label'] == 'metal'][0:50].plot(kind='scatter', x='chroma_stft', y='rmse', color='Yellow', label='metal', ax=ax);
# dataset[dataset['label'] == 'classical'][0:50].plot(kind='scatter', x='chroma_stft', y='rmse', color='Green', label='classical', ax=ax);
# dataset[dataset['label'] == 'blues'][0:50].plot(kind='scatter', x='chroma_stft', y='rmse', color='Red', label='blues', ax=ax);
# dataset[dataset['label'] == 'pop'][0:50].plot(kind='scatter', x='chroma_stft', y='rmse', color='Pink', label='pop', ax=ax);
# plt.show()
# +
#Decsission Trees
# -
from sklearn.tree import DecisionTreeClassifier
from sklearn.externals.six import StringIO
import pydotplus
import matplotlib.image as mpimg
from sklearn import tree
musicTree = DecisionTreeClassifier(criterion="entropy", max_depth = 10)
musicTree
musicTree.fit(X_train,y_train)
predTree = musicTree.predict(X_test)
from sklearn import metrics
print("DecisionTrees's Accuracy: ", metrics.accuracy_score(y_test, predTree))
dot_data = StringIO()
filename = "musictree_30sec_10.png"
featureNames = dataset.columns[0:57]
targetNames = dataset["label"].unique().tolist()
out=tree.export_graphviz(musicTree,feature_names=featureNames, out_file=dot_data, filled=True, special_characters=True,rotate=False)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
graph.write_png(filename)
img = mpimg.imread(filename)
plt.figure(figsize=(200, 400))
plt.imshow(img,interpolation='nearest')
# +
#RandomForest
# -
from sklearn.ensemble import RandomForestClassifier
rforest = RandomForestClassifier(n_estimators=1000, max_depth=10, random_state=0)
rforest.fit(X_train,y_train)
predForest = rforest.predict(X_test)
print("Random Forrest Accuracy: ", metrics.accuracy_score(y_test, predForest))
| Project_ML_STCET/GTZAN_MGD/MGD_sklearn_30sec_10 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Rizwan-Ahmed-Surhio/Accountants/blob/main/5_OperatorArithmetic_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="mdDu-WmWHMl5"
# ## Arithmetic Operations
# + id="rxX8MDDJHZXO"
2 + 2
# + id="t7i83QVBiSUw"
4 - 2
# + id="lrMEIxQ4iVa0"
4 * 3
# + id="yQPLupyR67WR"
16 / 4
# + id="qXiTsKkW6BEN"
2 ** 3
# + [markdown] id="9Zua-AIKl7TY"
# ### Modulo Operator
# + id="yPSDLyYL7MON"
7 % 2
# + [markdown] id="dWXDxpbNlxZ_"
# ### Integer Division
#
# + id="7jD191gbGbS3"
-16 / 3
# + id="DUe_gfuqGJG7"
16 // 3
# + id="YolrkdvlGW2P"
-16 // 3
| 5_OperatorArithmetic_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
import json
import numpy as np
from PIL import Image
import torch
import torchvision.models as models
import torchvision.transforms as transforms
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# -
model = models.resnet18(pretrained=True)
model.eval()
class_idx = json.load(open("network_vis/imagenet_class_index.json"))
# +
transform = transforms.Compose([
transforms.Scale(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])])
# +
image = transform(Image.open('network_vis/dog.jpg'))
plt.imshow(plt.imread('network_vis/dog.jpg'))
predictions = model(image.unsqueeze(0)).detach().numpy()[0]
print('Ranking: class (score)')
for i, idx in enumerate(np.argsort(predictions)[-10:][::-1]):
print('%d: %s (%.04f)' % (
i,
class_idx[str(idx)][1],
predictions[idx]))
# +
def visualize_layer(weight):
fig=plt.figure(figsize=(8, 8))
for i in range(64):
x = weight[i, ...].transpose([1, 2, 0])
x = (x - np.min(x))/np.ptp(x)
fig.add_subplot(8, 8, i + 1)
if x.shape[2] == 3:
imshow(x)
else:
imshow(x[:,:,0])
axis('off')
show()
visualize_layer(model.conv1.weight.data.cpu().numpy())
visualize_layer(model.layer1[0].conv1.weight.data.cpu().numpy())
# -
import sys
sys.path.append('..')
sys.path.append('../..')
from data import load
valid_data = load.get_dogs_and_cats('valid', resize=None, batch_size=128, is_resnet=True)
def undo_transform(x):
mean = np.array([0.485, 0.456, 0.406])[np.newaxis, :, np.newaxis, np.newaxis]
std = np.array([0.229, 0.224, 0.225])[np.newaxis, :, np.newaxis, np.newaxis]
return (x * std) + mean
# +
all_data = []
C1 = []
C2 = []
C3 = []
def store_activation(L, m, grad_in, grad_out):
L.append(grad_out.detach().cpu().numpy())
model.to(device)
try:
h1 = model.conv1.register_forward_hook(lambda *args: store_activation(C1, *args))
h2 = model.layer1[0].conv1.register_forward_hook(lambda *args: store_activation(C2, *args))
h3 = model.layer2[0].conv1.register_forward_hook(lambda *args: store_activation(C3, *args))
for it, (data, label) in enumerate(valid_data):
all_data.append(undo_transform(data.numpy()))
if device is not None:
data, label = data.to(device), label.to(device)
result = model(data)
finally:
h1.remove()
h2.remove()
h3.remove()
all_data = np.vstack(all_data)
C1 = np.vstack(C1)
C2 = np.vstack(C2)
C3 = np.vstack(C3)
# +
def to_pil(x):
x = x.transpose((1, 2, 0))
x -= x.min(0).min(0)[None,None]
x /= x.max(0).max(0)[None,None]
return Image.fromarray((x * 255).astype('uint8'))
def viz_act(img, act):
num_acts = len(act)
img = to_pil(img)
fig=plt.figure(figsize=(20, 20))
fig.add_subplot(1, 1+num_acts, 1)
imshow(img)
axis('off')
for i, c in enumerate(act):
fig.add_subplot(1, 1+num_acts, i + 2)
imshow(c)
axis('off')
show()
for i in range(3):
viz_act(all_data[i], C1[i, :3])
for i in range(3):
viz_act(all_data[i], C2[i, :3])
for i in range(3):
viz_act(all_data[i], C3[i, :3])
# +
def find_max_act(C, filter, W=10, img_size=[224, 224]):
fig=plt.figure(figsize=(8, 8))
filter_act = C[:, filter]
for i, idx in enumerate(np.argsort(-filter_act, axis=None)[:64]):
max_pos = list(np.unravel_index(idx, filter_act.shape))
max_pos[1] = int(max_pos[1] * float(img_size[0]) / filter_act.shape[1])
max_pos[2] = int(max_pos[2] * float(img_size[1]) / filter_act.shape[2])
img = to_pil(all_data[max_pos[0],
:,
max(max_pos[1] - W, 0) : max_pos[1] + W,
max(max_pos[2] - W, 0) : max_pos[2] + W])
fig.add_subplot(8, 8, i + 1)
imshow(img)
axis('off')
show()
find_max_act(C1, filter=0, W=4)#, img_size=img.size)
find_max_act(C1, filter=1, W=4)#, img_size=img.size)
find_max_act(C1, filter=2, W=4)#, img_size=img.size)
# -
find_max_act(C2, filter=0, W=12)#, img_size=img.size)
find_max_act(C2, filter=1, W=12)#, img_size=img.size)
find_max_act(C2, filter=2, W=12)#, img_size=img.size)
find_max_act(C3, filter=0, W=12)#, img_size=img.size)
find_max_act(C3, filter=1, W=12)#, img_size=img.size)
find_max_act(C3, filter=2, W=12)#, img_size=img.size)
| deep_learning_notes/4.13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## More Jupyter Widgets
# One of the nice things about using the Jupyter notebook systems is that there is a
# rich set of contributed plugins that seek to extend this system. In this lecture I
# want to introduce you to one such plugin, call ipy web rtc. Webrtc is a fairly new
# protocol for real time communication on the web. Yup, I'm talking about chatting.
# The widget brings this to the Jupyter notebook system. Lets take a look.
#
# First, lets import from this library two different classes which we'll use in a
# demo, one for the camera and one for images.
from ipywebrtc import CameraStream, ImageRecorder
# Then lets take a look at the camera stream object
help(CameraStream)
# We see from the docs that it's east to get a camera facing the user, and we can have
# the audio on or off. We don't need audio for this demo, so lets create a new camera
# instance
camera = CameraStream.facing_user(audio=False)
# The next object we want to look at is the ImageRecorder
help(ImageRecorder)
# The image recorder lets us actually grab images from the camera stream. There are features
# for downloading and using the image as well. We see that the default format is a png file.
# Lets hook up the ImageRecorder to our stream
image_recorder = ImageRecorder(stream=camera)
# Now, the docs are a little unclear how to use this within Jupyter, but if we call the
# download() function it will actually store the results of the camera which is hooked up
# in image_recorder.image. Lets try it out
# First, lets tell the recorder to start capturing data
image_recorder.recording=True
# Now lets download the image
image_recorder.download()
# Then lets inspect the type of the image
type(image_recorder.image)
# Ok, the object that it stores is an ipywidgets.widgets.widget_media.Image. How do we do
# something useful with this? Well, an inspection of the object shows that there is a handy
# value field which actually holds the bytes behind the image. And we know how to display
# those.
# Lets import PIL Image
import PIL.Image
# And lets import io
import io
# And now lets create a PIL image from the bytes
img = PIL.Image.open(io.BytesIO(image_recorder.image.value))
# And render it to the screen
display(img)
# +
# Great, you see a picture! Hopefully you are following along in one of the notebooks
# and have been able to try this out for yourself!
#
# What can you do with this? This is a great way to get started with a bit of computer vision.
# You already know how to identify a face in the webcam picture, or try and capture text
# from within the picture. With OpenCV there are any number of other things you can do, simply
# with a webcam, the Jupyter notebooks, and python!
| course_5/module_3-lecoptional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="jY-fw0HlOOEC" colab_type="text"
# # Palmer Penguins Dataset
# This is a very simple project to actually explore the relatively new dataset open-sources sometimes ago.
# It's a light dataset and slightly noisy and thus provide enough working room and complexity for a beginner in ML. It requires some cleaning, and can be used to learn the classification models. This dataset is also great for nice visualization and 3D plotting.
# + [markdown] id="YB6n2ckkOOEE" colab_type="text"
# ## Importing the dataset
# + id="U1wHG4eYOOEF" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123625430, "user_tz": -330, "elapsed": 2791, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib notebook
# + id="eBdc8LSROgQt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} executionInfo={"status": "ok", "timestamp": 1594123845764, "user_tz": -330, "elapsed": 66779, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="b4827d8f-36b6-4657-8f74-d50f4acfd38f"
# To get the dataset from google drive
# you need to save the dataset in your google drive
from google.colab import drive
drive.mount('/content/gdrive')
# + id="Voq1rtfIO5kM" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123854523, "user_tz": -330, "elapsed": 3015, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
#Read the data from drive
df=pd.read_csv('/content/gdrive/My Drive/Datasets/palmerpenguins.csv')
# + id="3rmzjKEmOOEL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 270} executionInfo={"status": "ok", "timestamp": 1594123854959, "user_tz": -330, "elapsed": 1776, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="d1ad1ec2-ccdd-422b-8cd8-058330212c3d"
#df = pd.read_csv("Location_of_dataset_locally")
df.head(2)
# + [markdown] id="j11kHEIBOOES" colab_type="text"
# ## Understanding the dataset (not really), cleaning and feature engineering.
# + id="jHB2FEUJOOET" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 270} executionInfo={"status": "ok", "timestamp": 1594123860105, "user_tz": -330, "elapsed": 1969, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="9a6b44bf-6257-416e-bb38-1f054fd45c81"
columns_to_keep = ['Sample Number',
'Species',
'Region',
'Island',
'Stage',
'Culmen Length (mm)',
'Culmen Depth (mm)',
'Flipper Length (mm)',
'Body Mass (g)'
]
df = df[columns_to_keep]
df.head(2)
# + id="aw1vVeKrOOEX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 270} executionInfo={"status": "ok", "timestamp": 1594123879576, "user_tz": -330, "elapsed": 2787, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="484ed9bb-9324-4e6b-86a8-64b8593969d6"
df.set_index("Sample Number")
#df = df.dropna(inplace = True) # Not a good idea for this dataset
df.head(2)
# + [markdown] id="Eeu-gDp2OOEc" colab_type="text"
# #### Names of the species
# The names are too long (also contains scientific names) , so just seperating the names and keeping the general specie name (pronounce-able names).
# + id="0Op4M6o-OOEd" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123883932, "user_tz": -330, "elapsed": 1078, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
new = df["Species"].str.split("(", n = 1, expand = True)
df["Species"] = new[0]
df["Name"] = new[1]
#df.head()
# + [markdown] id="r7K132oCOOEh" colab_type="text"
# Also inputing the names to the model is not a great idea so I'm assigning an integer to the repective specie which will be the label.
#
# There can be better ways to do this which I don't know about, corrections are welcome.
# + id="gsh0-BITOOEi" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123888242, "user_tz": -330, "elapsed": 1086, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
df["Label"] = new[0]
species = {'Adelie Penguin ':1, 'Chinstrap penguin ':2, 'Gentoo penguin ':3}
df.Label = [species[item] for item in df.Label]
#df.head()
# + [markdown] id="alqXdiG1OOEn" colab_type="text"
# It's time to split the dataset
# + id="CzYzoh0hOOEo" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123890584, "user_tz": -330, "elapsed": 1318, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
from sklearn.model_selection import train_test_split
# + [markdown] id="oEfNJ1umOOEs" colab_type="text"
# Remember we left the NaN fields untouched, so I'll just fill those fields with the average of that column. (I hope it's a nice idea).
# + [markdown] id="-KlZ-kSEOOEt" colab_type="text"
# #### I decided to choose these four features , although there are many other useful features in the dataset.
# + id="LNzKQuMxOOEu" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123894560, "user_tz": -330, "elapsed": 1042, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
X = df[['Culmen Length (mm)','Culmen Depth (mm)','Flipper Length (mm)','Body Mass (g)']]
X = X.fillna(X.mean())
y = df['Label']
y = y.fillna(y.mean())
# + id="yOlB3cu4OOEx" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123895853, "user_tz": -330, "elapsed": 705, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# + [markdown] id="pqRKVR_QOOE1" colab_type="text"
# ## Visualizing the cleaned datset
# Now we have to visulize the data to see if it's actually differentiable and in what ways.
# + id="QmQ9lL2bOOE2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} executionInfo={"status": "ok", "timestamp": 1594123900746, "user_tz": -330, "elapsed": 3102, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="92994196-6240-4e2c-db74-76efde343ec4"
# plotting a scatter matrix
from matplotlib import cm
from pandas.plotting import scatter_matrix
cmap = cm.get_cmap('gnuplot')
scatter = scatter_matrix(X_train, c= y_train, marker = 'o', s=40, hist_kwds={'bins':15}, figsize=(9,9), cmap=cmap)
# + [markdown] id="ZQChHcHtOOE6" colab_type="text"
# ### Attractive bits of the project (^_^)
# plotting the 3D plot gives a clear idea of how the data is clearly differentiable.
# + id="5gXiEq1bOOE7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} executionInfo={"status": "ok", "timestamp": 1594123926563, "user_tz": -330, "elapsed": 1466, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="26b26c0f-613e-4925-ef67-7e7ab00d00b9"
# plotting a 3D scatter plot
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection = '3d')
ax.scatter(X_train['Culmen Length (mm)'], X_train['Culmen Depth (mm)'], X_train['Flipper Length (mm)'], c = y_train, marker = 'o', s=20)
ax.set_xlabel('Culem length (mm)')
ax.set_ylabel('Culem Depth (mm)')
ax.set_zlabel('Flipper Lenth (mm)')
plt.show();
# + [markdown] id="BpsFi14EOOFC" colab_type="text"
# ### Training the model(s)
# #### KNN-Classifier
# At this point I remember the syntax for KNN-Classifier word to word, so I'm going to implement it :)
# + id="dKrPNIlBOOFD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} executionInfo={"status": "ok", "timestamp": 1594123934194, "user_tz": -330, "elapsed": 1167, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="89c798fe-5ef4-402d-a0db-77ae52b2816f"
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
knn = KNeighborsClassifier(n_neighbors = 7)
knn.fit(X_train_scaled, y_train)
print('Accuracy of K-NN classifier on training set: {:.2f}'
.format(knn.score(X_train_scaled, y_train)))
print('Accuracy of K-NN classifier on test set: {:.2f}'
.format(knn.score(X_test_scaled, y_test)))
# + [markdown] id="Z9aCvnpwOOFH" colab_type="text"
# #### DecisionTree-Classifier
# ##### Using decision tree without max decision tree depth, the model is likely to overfit
#
# And now I'll train a Decision Tree Classifier , it's handy and also very nice when plotted.
# + id="J0qdjdJcOOFI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} executionInfo={"status": "ok", "timestamp": 1594123942266, "user_tz": -330, "elapsed": 689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="0f201542-e1b5-491e-a0f7-435cbb308755"
from sklearn.tree import DecisionTreeClassifier
#creating a model instance
clf = DecisionTreeClassifier().fit(X_train, y_train)
print('Accuracy of Decision Tree classifier on training set: {:.2f}'
.format(clf.score(X_train, y_train)))
print('Accuracy of Decision Tree classifier on test set: {:.2f}'
.format(clf.score(X_test, y_test)))
# + [markdown] id="jjahrqv8OOFM" colab_type="text"
# #### I'm going to plot the tree(s)
# I could use " tree.plot_tree(tree_instance) " but it would be very hard to comprehend.
# So we are providing the features and the labels to the graph and also keeping the "fill" on to colore the boxes, it makes the plots far better and readable.
# + id="Ggk6QJa7OOFN" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1594123946028, "user_tz": -330, "elapsed": 1197, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}}
from sklearn import tree
#tree.plot_tree(clf);
# + id="-bC30eFXOOFR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} executionInfo={"status": "ok", "timestamp": 1594123951520, "user_tz": -330, "elapsed": 3298, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="9e5822d8-a14d-4b8d-cfec-9612086c3bdb"
fn=['Culmen Length (mm)','Culmen Depth (mm)','Flipper Length (mm)','Body Mass (g)']
cn=['Adelie Penguin ', 'Chinstrap penguin ', 'Gentoo penguin ']
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (6, 6), dpi=300)
tree.plot_tree(clf,
feature_names = fn,
class_names=cn,
filled = True);
#fig.savefig('Bad_tree.png')
# + [markdown] id="OCe3gnHgOOFW" colab_type="text"
# I'm plotting the decision tree just to understand hoe the "max_depth" is helping to avoid overfit.
# ##### Also it looks cool :)
# + [markdown] id="OPKvuVJ9OOFX" colab_type="text"
# #### Avoiding Overfitting
# To avoide overfitting we will use max_depth, in this case I tried 3 to 7 and 4 seems to work best (0-4).
# Also I don't know if there is a way to "calculate" the the optimum value for max_depth, point out if there is.
# + id="8EmlT254OOFY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} executionInfo={"status": "ok", "timestamp": 1594123960229, "user_tz": -330, "elapsed": 1038, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="e0aaf807-7590-4cc7-8005-355ce1871c78"
clf2 = DecisionTreeClassifier(max_depth = 4).fit(X_train, y_train)
print('Accuracy of Decision Tree classifier on training set: {:.2f}'
.format(clf2.score(X_train, y_train)))
print('Accuracy of Decision Tree classifier on test set: {:.2f}'
.format(clf2.score(X_test, y_test)))
# + [markdown] id="pz4s3sOHOOFc" colab_type="text"
# See how the train accuracy decreased and the test accuracy increased by avoiding overfitting.
# + id="4yUDcoOzOOFd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} executionInfo={"status": "ok", "timestamp": 1594123978355, "user_tz": -330, "elapsed": 2967, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gir4BHm7l_si7-mbqrlPR3lMF-BmJiJC7P87ZHc3tY=s64", "userId": "13755018619219626781"}} outputId="ab9accb0-1f4f-48ea-94a0-24432500433f"
fn=['Culmen Length (mm)','Culmen Depth (mm)','Flipper Length (mm)','Body Mass (g)']
cn=['Adelie Penguin ', 'Chinstrap penguin ', 'Gentoo penguin ']
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (6, 6), dpi=300)
tree.plot_tree(clf2,
feature_names = fn,
class_names=cn,
filled = True);
# This line will download the image
#fig.savefig('Improve_tree.png')
# + [markdown] id="vuvsp81FOOFh" colab_type="text"
# ### It was a nice experience working with the dataset.
# + id="57Y5fIDmUAaD" colab_type="code" colab={}
| Projects/Palmer_Penguins/Penguins data 1.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Libraries import
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from statsmodels.stats.outliers_influence import variance_inflation_factor
import warnings
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression,Lasso,LassoCV,Ridge,RidgeCV,ElasticNet,ElasticNetCV
from sklearn.svm import SVR
from sklearn.neighbors import KNeighborsRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import pickle
# -
#Dataset load
datafile = pd.read_csv("marriage.csv")
df = datafile.copy()
# Datafile exploration
df.info()
df.isnull().sum()
df.describe()
df.head(5)
# ## Exploratory Data Analysis
# Gender: Lets drop the na value rows since imputating them is not a good idea.
df.dropna(subset = ['gender'], inplace = True)
# Height: First change the value type and convert it into integer and then do imputation.
df['height'] = df['height'].astype(str)
df['height'] = df['height'].apply(lambda x: x.replace("'",'.'))
df['height'] = df['height'].apply(lambda x: x.replace('"' ,''))
df['height'] = df['height'].astype(float)
df['height'].fillna(df.height.median(),inplace=True)
# Religion
df.religion.value_counts()
df['religion'].fillna("Hindu", inplace = True)
# Caste
df.caste.value_counts()
df['caste'].fillna("others", inplace = True)
#mother_tongue
df.mother_tongue.value_counts()
df.dropna(subset = ['mother_tongue'], inplace = True)
# profession
df.profession.value_counts()
df.dropna(subset = ['profession'], inplace = True)
#location
df.location.value_counts()
df['location'].fillna("Mumbai", inplace = True)
# age of marriage
df.age_of_marriage.describe()
df['age_of_marriage'].fillna(df.age_of_marriage.median(),inplace=True)
#Dropping of the id column
df.drop('id',inplace=True,axis=1)
# Renaming of the age of marriage column
df.rename(columns={"age_of_marriage": "age"}, inplace = True)
df.info()
df.head()
# ### Visualisation
# gender
plt.figure(figsize=(10,5))
print(df.gender.value_counts())
sns.countplot(df.gender)
plt.show()
warnings.filterwarnings('ignore')
#height
sns.distplot(df.height)
plt.show()
#religion
plt.figure(figsize=(15,5))
print(df.religion.value_counts())
sns.countplot(df.religion)
plt.show()
#caste
plt.figure(figsize=(15,5))
print(df.caste.value_counts())
sns.countplot(df.caste)
plt.show()
#mother_tongue
plt.figure(figsize=(15,5))
print(df.mother_tongue.value_counts())
sns.countplot(df.mother_tongue)
plt.show()
#country
plt.figure(figsize=(15,5))
print(df.country.value_counts())
sns.countplot(df.country)
plt.show()
# ### Pre-model data preparations
# Depedent and independent variable split
y = df['age']
x =df.drop(columns = ['age'])
x.drop(["profession","location"],axis = 1, inplace = True)
x
# Label encoding
le = LabelEncoder()
x.loc[:,['gender','religion','caste','mother_tongue','country']]= \
x.loc[:,['gender','religion','caste','mother_tongue','country']].apply(le.fit_transform)
# +
# To check for multicollinearity
# VIF dataframe
vif_data = pd.DataFrame()
vif_data["feature"] = x.columns
# calculating VIF for each feature
vif_data["VIF"] = [variance_inflation_factor(x.values, i)
for i in range(len(x.columns))]
print(vif_data)
# -
# Scaling
scalar = StandardScaler()
x_scaled = scalar.fit_transform(x)
# +
# VIF check after scaling
vif = pd.DataFrame()
vif["vif"] = [variance_inflation_factor(x_scaled,i) for i in range(x_scaled.shape[1])]
vif["Features"] = x.columns
#let's check the values
vif
# -
# Train test dataset split
x_train,x_test,y_train,y_test = train_test_split(x_scaled,y, test_size= 0.25, random_state = 101)
# ## Model Building
# ### Linear Regression
# +
model_lr = LinearRegression()
model_lr.fit(x_train, y_train)
print(f"Model score on train data: {round(model_lr.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(model_lr.score(x_test, y_test),2)}")
# -
# ### Regularisation
# +
#Lets try regularising our linear model to reduce overfitting
## LASSO regularisation
lasscv = LassoCV(alphas = None,cv =10, max_iter = 100000, normalize = True)
lasscv.fit(x_train, y_train)
# -
# best alpha parameter
alpha = lasscv.alpha_
alpha
lasso_model = Lasso(alpha)
lasso_model.fit(x_train, y_train)
print(f"Model score on train data: {round(lasso_model.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(lasso_model.score(x_test, y_test),2)}")
## RIDGE Regularisation
alphas = np.random.uniform(low=0, high=10, size=(50,))
ridgecv = RidgeCV(alphas = alphas,cv=10,normalize = True)
ridgecv.fit(x_train, y_train)
# best alpha parameter
alpha = ridgecv.alpha_
alpha
ridge_model = Ridge(alpha=ridgecv.alpha_)
ridge_model.fit(x_train, y_train)
print(f"Model score on train data: {round(ridge_model.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(ridge_model.score(x_test, y_test),2)}")
# ELASTIC NET Regularisation
elasticcv = ElasticNetCV(alphas = None, cv =10)
elasticcv.fit(x_train, y_train)
alpha = elasticcv.alpha_
alpha
elasticcv.l1_ratio
elasticnet_model = ElasticNet(alpha = elasticcv.alpha_,l1_ratio=0.5)
elasticnet_model.fit(x_train, y_train)
print(f"Model score on train data: {round(elasticnet_model.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(elasticnet_model.score(x_test, y_test),2)}")
# +
# Thus regularisation gives no improvement in model score.Lets try other models.
# -
# ### Support Vector Regressor
# +
model_svr = SVR()
model_svr.fit(x_train, y_train)
print(f"Model score on train data: {round(model_svr.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(model_svr.score(x_test, y_test),2)}")
# -
# ### K Nearest Neighbour
# +
model_knn = KNeighborsRegressor()
model_knn.fit(x_train, y_train)
print(f"Model score on train data: {round(model_knn.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(model_knn.score(x_test, y_test),2)}")
# -
# ### Decision Tree
#
# +
model_dt = DecisionTreeRegressor()
model_dt.fit(x_train, y_train)
print(f"Model score on train data: {round(model_dt.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(model_dt.score(x_test, y_test),2)}")
# -
# ### Random Forest
# +
model_rf = RandomForestRegressor()
model_rf.fit(x_train, y_train)
print(f"Model score on train data: {round(model_rf.score(x_train, y_train),2)}")
print(f"Model score on test data: {round(model_rf.score(x_test, y_test),2)}")
# +
# Thus for this usecase, linear regression seems to give us the highets score of 90% on out test datatset.
# -
# Best model saving
pickle.dump(model_lr, open('model.pkl','wb'))
model=pickle.load(open('model.pkl','rb'))
| Year of marriage prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Our three global feature descriptors are
#1. Color Histogram that quantifies color of the flower.
#2. Hu Moments that quantifies shape of the flower.
#3. Haralick Texture that quantifies texture of the flower.
# imports the necessary libraries we need to work with.
from sklearn.metrics import confusion_matrix, precision_score, recall_score, accuracy_score, classification_report
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
from sklearn import model_selection
from sklearn import linear_model
import matplotlib.pyplot as plt
from sklearn import metrics
# from skimage import feature
from numpy import asarray
import seaborn as sn
import pandas as pd
import numpy as np
import mahotas
import h5py
import cv2
import os
print("[Status]: Done")
# +
#image per class
images_per_class = 80
#to fixed our image size
fixed_size = tuple((500, 500))
dataset_path = "Dataset\FlowerSpecies"
csv_data = "output\data.csv"
bins = 8
print("[Status]: Done")
# +
# feature-descriptor-1: Hu Moments(7)
# it can be used to describe,characterize, and quantify the shape of an object in an image
def feature_hu_moments(image):
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
feature = cv2.HuMoments(cv2.moments(image)).flatten()
return feature
# feature-descriptor-2: Haralick Texture (13)
# Texture analysis. Haralick texture features are calculated
# from a Gray Level Co-occurrence Matrix, (GLCM),
def feature_haralick(image):
# convert the image to grayscale
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# compute the haralick texture feature vector
haralick = mahotas.features.haralick(gray).mean(axis=0)
# return the result
return haralick
# feature-descriptor-3: Color Histogram (512)
def feature_histogram(image, mask=None):
# convert the image to HSV color-space
image = cv2.cvtColor(image, cv2.COLOR_BGR2HSV)
# compute the color histogram
hist = cv2.calcHist([image], [0, 1, 2], None, [bins, bins, bins], [0, 256, 0, 256, 0, 256])
# normalize the histogram
cv2.normalize(hist, hist)
# return the histogram
return hist.flatten()
print("[Status]: Done")
# +
data_labels = os.listdir(dataset_path)
# sort the training labels
data_labels.sort()
print(data_labels)
# # empty lists to hold feature vectors and labels
features = []
labels = []
print("Now it is time to extract important features from image")
# +
# loop over the dataset sub-folders
for image_type in data_labels:
# join the data path and each species folder
dir = os.path.join(dataset_path, image_type)
print(dir)
# get the current label
current_label = image_type
# print(os.listdir(dir))
for image_name in os.listdir(dir):
# read the image and resize it to a fixed-size
image = cv2.imread(os.path.join(dir,image_name))
image = cv2.resize(image, fixed_size)
# print(image.shape, image_name)
####################################
# Global Feature extraction
####################################
fv_hu_moments = fd_hu_moments(image)
fv_haralick = fd_haralick(image)
fv_histogram = fd_histogram(image)
###################################
# Concatenate global features
###################################
feature = np.hstack([fv_histogram, fv_haralick, fv_hu_moments])
# update the list of labels and feature vectors
labels.append(current_label)
features.append(feature)
print("[STATUS] processed folder: {}".format(current_label))
print("[STATUS] completed Global Feature Extraction...")
# -
features = np.array(features)
labels = np.array(labels)
# print(labels)
print("Number of Features Extraction : ", features.shape[1])
print("Number of Sample : ", features.shape[0])
print(features)
df = pd.DataFrame(features)
df['target'] = labels
df.to_csv(csv_data, index = False)
print("[Status] write the dataset in csv file")
# Total we have 532 features
print("Dataset")
dataset = pd.read_csv(csv_data, index_col=False)
dataset
# +
X = dataset.iloc[ : , :-1]
print("store features values into x ", X.shape)
Y = dataset.target
print("store features values into y ", Y.shape)
print("No of the features: ", np.array(X).shape[1])
print("Fearture vector size: ", np.array(X).shape)
print("Lables size[Target]: ", np.array(Y).shape)
# print("Lables [Target]: ", np.array(Y))
# -
x1 = X['0']
y1 = X['1']
x1 = x1.sort_values()
y1 = y1.sort_values()
plt.scatter(x1, y1)
plt.plot(x1, y1)
plt.show()
scaler = MinMaxScaler(feature_range=(0, 1))
sd_scaler = StandardScaler()
x_sd_scaler = sd_scaler.fit_transform(X)
x_rescaled = scaler.fit_transform(X)
print("[STATUS] feature vector normalized...")
print(pd.DataFrame(x_rescaled).shape)
# x_plot = pd.DataFrame(x_rescaled)
# x_plot.plot.kde()
label_encoder = LabelEncoder()
target = label_encoder.fit_transform(Y)
# print(target)
# define one hot encoding
encoder = OneHotEncoder(sparse=False)
# transform data
onehot = encoder.fit_transform(np.reshape(target, (-1, 1)))
# print(onehot)
# temp = encoder.inverse_transform(onehot)
# print(temp)
targetNames = np.unique(Y)
print("Target values are: ", target)
print("Target Shape: ", target.shape)
print("Target Names are: ", targetNames)
print("[STATUS] training labels encoded...")
# +
from sklearn.model_selection import train_test_split
train_frac = 0.6 #60% left ->: 100-60 = 40%
valid_frac = 0.5 #(100/40)*20
test_frac = 0.5 #(100/40)*20
x_train, x_rem, y_train, y_rem = train_test_split(x_rescaled, target, train_size=train_frac)
# x_train, x_rem, y_train, y_rem = train_test_split(x_rescaled, target, train_size=train_frac)
x_valid, x_test, y_valid, y_test = train_test_split(x_rem, y_rem, test_size=test_frac)
print("Train vector size: ", x_train.shape, y_train.shape)
print("Test vector size: ", x_test.shape, y_test.shape)
print("Validation vector size: ", x_valid.shape, y_valid.shape)
# -
# +
lm = linear_model.LogisticRegression(multi_class='ovr', solver='liblinear')
lm.fit(x_train, y_train)
print("Training done!")
print('Predicted value is =', lm.predict([x_test[12]]))
print('Actual value from test data is %s and corresponding image is as below' % (y_test[12]) )
# print(lm.score(x_test, y_test))
y_pred_test = lm.predict(x_test)
y_pred_train = lm.predict(x_train)
df = pd.DataFrame(x_test)
df['Actual'] = y_test
df['Predicted'] = y_pred_test
df.to_csv('Result_exp/Logistic_Regression_Result/test_result.csv')
print("Over all train accuracy % : ", accuracy_score(y_train, y_pred_train)*100)
print("Over all test accuracy % : ", accuracy_score(y_test, y_pred_test)*100)
# print("Over all test accuracy : ", accuracy_score(y_test, y_pred))
# print('Confusion Matrix For Test Data set : \n' + str(confusion_matrix(y_test, y_pred)))
print('Confusion Matrix For Test Data set : \n')
cm = confusion_matrix(y_test, y_pred_test)
plt.figure(figsize = (10,7))
# cmap= "Greens_r"
# cmap= "OrRd_r"
# cmap= "OrRd"
sn.heatmap(cm, annot=True,cmap="Greens_r")
print("Classification Report for 3-classes: ")
print(classification_report(y_test, y_pred_test))
# -
# +
sgd = linear_model.SGDClassifier()
sgd.fit(x_train, y_train)
print("Training done!")
print('Predicted value is =', sgd.predict([x_test[12]]))
print('Actual value from test data is %s and corresponding image is as below' % (y_test[12]) )
# print(lm.score(x_test, y_test))
y_pred_test = sgd.predict(x_test)
y_pred_train = sgd.predict(x_train)
df = pd.DataFrame(x_test)
df['Actual'] = y_test
df['Predicted'] = y_pred_test
df.to_csv('Result_exp/Logistic_Regression_Result/test_result.csv')
print("Over all train accuracy % : ", accuracy_score(y_train, y_pred_train)*100)
print("Over all test accuracy % : ", accuracy_score(y_test, y_pred_test)*100)
# print("Over all test accuracy : ", accuracy_score(y_test, y_pred))
# print('Confusion Matrix For Test Data set : \n' + str(confusion_matrix(y_test, y_pred)))
print('Confusion Matrix For Test Data set : \n')
cm = confusion_matrix(y_test, y_pred_test)
plt.figure(figsize = (10,7))
# cmap= "Greens_r"
# cmap= "OrRd_r"
# cmap= "OrRd"
sn.heatmap(cm, annot=True,cmap="Greens_r")
print("Classification Report for 3-classes: ")
print(classification_report(y_test, y_pred_test))
# +
from sklearn.model_selection import train_test_split
train_frac = 0.6 #60% left ->: 100-60 = 40%
valid_frac = 0.5 #(100/40)*20
test_frac = 0.5 #(100/40)*20
x_train, x_rem, y_train, y_rem = train_test_split(x_sd_scaler, target, train_size=train_frac)
# x_train, x_rem, y_train, y_rem = train_test_split(x_rescaled, target, train_size=train_frac)
x_valid, x_test, y_valid, y_test = train_test_split(x_rem, y_rem, test_size=test_frac)
print("Train vector size: ", x_train.shape, y_train.shape)
print("Test vector size: ", x_test.shape, y_test.shape)
print("Validation vector size: ", x_valid.shape, y_valid.shape)
# +
lm = linear_model.LogisticRegression(multi_class='ovr', solver='liblinear')
lm.fit(x_train, y_train)
print("Training done!")
print('Predicted value is =', lm.predict([x_test[12]]))
print('Actual value from test data is %s and corresponding image is as below' % (y_test[12]) )
# print(lm.score(x_test, y_test))
y_pred_test = lm.predict(x_test)
y_pred_train = lm.predict(x_train)
df = pd.DataFrame(x_test)
df['Actual'] = y_test
df['Predicted'] = y_pred_test
df.to_csv('Result_exp/Logistic_Regression_Result/test_result.csv')
print("Over all train accuracy % : ", accuracy_score(y_train, y_pred_train)*100)
print("Over all test accuracy % : ", accuracy_score(y_test, y_pred_test)*100)
# print("Over all test accuracy : ", accuracy_score(y_test, y_pred))
# print('Confusion Matrix For Test Data set : \n' + str(confusion_matrix(y_test, y_pred)))
print('Confusion Matrix For Test Data set : \n')
cm = confusion_matrix(y_test, y_pred_test)
plt.figure(figsize = (10,7))
# cmap= "Greens_r"
# cmap= "OrRd_r"
# cmap= "OrRd"
sn.heatmap(cm, annot=True,cmap="Greens_r")
print("Classification Report for 3-classes: ")
print(classification_report(y_test, y_pred_test))
# +
sgd = linear_model.SGDClassifier()
sgd.fit(x_train, y_train)
print("Training done!")
print('Predicted value is =', sgd.predict([x_test[12]]))
print('Actual value from test data is %s and corresponding image is as below' % (y_test[12]) )
# print(lm.score(x_test, y_test))
y_pred_test = sgd.predict(x_test)
y_pred_train = sgd.predict(x_train)
df = pd.DataFrame(x_test)
df['Actual'] = y_test
df['Predicted'] = y_pred_test
df.to_csv('Result_exp/Logistic_Regression_Result/test_result.csv')
print("Over all train accuracy % : ", accuracy_score(y_train, y_pred_train)*100)
print("Over all test accuracy % : ", accuracy_score(y_test, y_pred_test)*100)
# print("Over all test accuracy : ", accuracy_score(y_test, y_pred))
# print('Confusion Matrix For Test Data set : \n' + str(confusion_matrix(y_test, y_pred)))
print('Confusion Matrix For Test Data set : \n')
cm = confusion_matrix(y_test, y_pred_test)
plt.figure(figsize = (10,7))
# cmap= "Greens_r"
# cmap= "OrRd_r"
# cmap= "OrRd"
sn.heatmap(cm, annot=True,cmap="Greens_r")
print("Classification Report for 3-classes: ")
print(classification_report(y_test, y_pred_test))
# -
# Total we have 532 features
print("Dataset")
dataset = pd.read_csv(csv_data, index_col=False)
dataset
| features_extraction_and_Logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from keras.preprocessing.image import load_img, img_to_array
from keras.models import load_model
import os
import pandas as pd
from tqdm.notebook import tqdm
# +
longitud, altura = 100, 100
modelo='./modelo_vgg16/modelo_vgg16.h5'
pesos='./modelo_vgg16/pesos_vgg16.h5'
cnn=load_model(modelo)
cnn.load_weights(pesos)
lista_categorias = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K', 'L', 'M',
'N', 'O', 'P', 'Q', 'R', 'S', 'T', 'U', 'V', 'W', 'X', 'Y', 'Z',
'del', 'nothing', 'space']
categoria=[]
filenames=[]
for file in tqdm(os.listdir('./test')):
filenames.append(file)
file=load_img('./test/' + file, target_size=(longitud, altura))
file=img_to_array(file)
file=np.expand_dims(file, axis=0)/255
arreglo=cnn.predict(file)
resultado=arreglo[0]
respuesta=np.argmax(resultado)
categoria.append(lista_categorias[respuesta])
# -
submission = pd.DataFrame(
{'archivo': filenames,
'target': categoria,
})
submission.to_csv(r'./submissions_vgg16/submission_vgg16_2.csv', index=False)
| Modelo_3_validacion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building Pulse Schedules
#
# Building `Schedule`s is very straightforward, yet there are a few ways to compose them. We're going to explore these different methods here.
#
# In this tutorial we use the `Play` instruction, with a `ConstantPulse` pulse argument. To learn about these and other `Instruction`s, first check out [this page](2_building_pulse_instructions.ipynb).
#
# Tip: `Instruction`s can be treated just like `Schedule`s -- that means all the methods we use here work equally well for composing `Schedule`s with `Schedule`s, `Instruction`s with `Instruction`s, or between the two.
#
# As the basis for trying out the composition methods, let's initialize a couple dummy `Schedule`s.
# +
from qiskit.pulse import Schedule, Play, Constant, Delay, DriveChannel
sched_a = Schedule(name="A")
sched_b = Schedule(name="B")
# -
# ### `insert` or `|`
#
# The `insert` method schedules an `Instruction` or `Schedule` at a particular time. Let's use it to give each of our schedules a different instruction.
#
# The insert schedule method overloads the `|` (or) operator for `Schedule`s, so we will use this special syntax on schedule `B`. When using this syntactic sugar, the `time` argument is implicitly zero.
# +
sched_a = sched_a.insert(0, Play(Constant(duration=5, amp=1), DriveChannel(0)))
sched_b |= Play(Constant(duration=5, amp=-1), DriveChannel(0))
sched_a.draw()
# -
# ### `shift` or `<<`
#
# Sometimes we need to offset one schedule from `time=0`. For instance, when using `|`, we use `shift` to add an instruction at a time besides zero.
# +
sched_a |= Play(Constant(duration=5, amp=0.5), DriveChannel(0)).shift(10)
sched_b |= Play(Constant(duration=5, amp=-0.5), DriveChannel(0)) << 10
sched_b.draw()
# -
# Let's see how we can use `insert` to compose schedules `A` and `B` together.
# +
sched_a_and_b = sched_a.insert(20, sched_b) # A followed by B at time 20
sched_b_and_a = sched_a | sched_b << 20 # B followed by A at time 20
sched_a_and_b.draw()
# -
# ### `append` or `+`
#
# The `append` method is like `insert`, but the insertion time is determined for us. The `Instruction` or `Schedule` being added will begin when all the channels common to the two become free. If they contain no common channels, then the `Schedule` will be appended at `time=0`. In psuedocode:
#
# ```
# time = 0
# for channel in intersection(original_sched_channels, appended_sched_channels):
# time = max(time, original_sched.duration(channel))
# ```
#
# The append schedule method overloads the `+` (add) operator for Schedules.
#
# Let's continue with schedule `A` and schedule `B`. Remember that they both have instructions scheduled on the same channel, `DriveChannel(0)`. Schedule `A` contains positive amplitude pulses, and schedule `B` contains negative amplitude pulses.
# +
sched_a_plus_b = sched_a.append(sched_b)
sched_a_plus_b.draw()
# -
# Let's see what happens when there are no common channels.
#
# Warning: Common channels is not the same as common qubits. Measurements require different channels than gate operations, even on one qubit. If you simply append a measurement to a schedule, you'll likely schedule your measurement at `time=0`, not what you were expecting! Instead, use the shift operation, with `schedule.duration` as the time argument.
# +
sched_a_plus_c = sched_a + Play(Constant(duration=20, amp=0.2), DriveChannel(1))
sched_a_plus_c.draw()
# -
# This time, the new instruction being appended was added at `time=0`, because it did not have any instructions on `DriveChannel(0)`.
#
# Be wary: `append` only "slides" the appended schedule into the original around channels that are not common between them. The following example demonstrates this point.
# +
sched_d = Play(Constant(duration=20, amp=-0.2), DriveChannel(0)) \
+ (Play(Constant(duration=10, amp=-0.2), DriveChannel(1)) << 10)
(sched_a_plus_c + sched_d).draw()
# -
# Congrats, you've mastered schedule composition!
#
# On the next page, we will see how to [add measurements to your schedule](4_adding_measurements.ipynb).
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| tutorials/pulse/3_building_pulse_schedules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# Imports
########################################################################
# Python Standard Libraries
import os
import multiprocessing
from timeit import default_timer as timer
########################################################################
# Numpy Library
import numpy as np # linear algebra
########################################################################
# Pandas Library
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
########################################################################
# MATPLOT Library
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.ticker import MaxNLocator
# %matplotlib inline
########################################################################
# SKLearn Library
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
from sklearn import metrics
from sklearn.metrics import accuracy_score, precision_recall_curve, classification_report, confusion_matrix, average_precision_score, roc_curve, auc, multilabel_confusion_matrix
########################################################################
# SCIPY Library
from scipy.stats import gaussian_kde
import scipy.stats as st
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Utility functions
########################################################################
# Print system information
def print_system_info():
mem_bytes = os.sysconf('SC_PAGE_SIZE') * os.sysconf('SC_PHYS_PAGES') # e.g. 4015976448
mem_gib = mem_bytes/(1024.**3) # e.g. 3.74
print("{:<23}{:f} GB".format('RAM:', mem_gib))
print("{:<23}{:d}".format('CORES:', multiprocessing.cpu_count()))
# !lscpu
########################################################################
# Walk through input files
def print_input_files():
# Input data files are available in the "../input/" directory.
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
########################################################################
# Dump text files
def dump_text_file(fname):
with open(fname, 'r') as f:
print(f.read())
########################################################################
# Dump CSV files
def dump_csv_file(fname, count=5):
# count: 0 - column names only, -1 - all rows, default = 5 rows max
df = pd.read_csv(fname)
if count < 0:
count = df.shape[0]
return df.head(count)
########################################################################
# Dataset related functions
ds_nbaiot = '/kaggle/input/nbaiot-dataset'
dn_nbaiot = ['Danmini_Doorbell', 'Ecobee_Thermostat', 'Ennio_Doorbell', 'Philips_B120N10_Baby_Monitor', 'Provision_PT_737E_Security_Camera', 'Provision_PT_838_Security_Camera', 'Samsung_SNH_1011_N_Webcam', 'SimpleHome_XCS7_1002_WHT_Security_Camera', 'SimpleHome_XCS7_1003_WHT_Security_Camera']
def fname(ds, f):
if '.csv' not in f:
f = f'{f}.csv'
return os.path.join(ds, f)
def fname_nbaiot(f):
return fname(ds_nbaiot, f)
def get_nbaiot_device_files():
nbaiot_all_files = dump_csv_file(fname_nbaiot('data_summary'), -1)
nbaiot_all_files = nbaiot_all_files.iloc[:,0:1].values
device_id = 1
indices = []
for j in range(len(nbaiot_all_files)):
if str(device_id) not in str(nbaiot_all_files[j]):
indices.append(j)
device_id += 1
nbaiot_device_files = np.split(nbaiot_all_files, indices)
return nbaiot_device_files
def get_nbaiot_device_data(device_id, count_norm=-1, count_anom=-1):
if device_id < 1 or device_id > 9:
assert False, "Please provide a valid device ID 1-9, both inclusive"
if count_anom == -1:
count_anom = count_norm
device_index = device_id -1
device_files = get_nbaiot_device_files()
device_file = device_files[device_index]
df = pd.DataFrame()
y = []
for i in range(len(device_file)):
fname = str(device_file[i][0])
df_c = pd.read_csv(fname_nbaiot(fname))
count = count_anom
if 'benign' in fname:
count = count_norm
rows = count if count >=0 else df_c.shape[0]
print("processing", fname, "rows =", rows)
y_np = np.ones(rows) if 'benign' in fname else np.zeros(rows)
y.extend(y_np.tolist())
df = pd.concat([df.iloc[:,:].reset_index(drop=True),
df_c.iloc[:rows,:].reset_index(drop=True)], axis=0)
X = df.iloc[:,:].values
y = np.array(y)
Xdf = df
return (X, y, Xdf)
def get_nbaiot_devices_data():
devices_data = []
for i in range(9):
device_id = i + 1
(X, y) = get_nbaiot_device_data(device_id)
devices_data.append((X, y))
return devices_data
#print_input_files()
print_system_info()
# -
def remove_correlated_features(df, threshold):
df = df.copy()
# Create correlation matrix
corr_matrix = df.corr().abs()
# Select upper triangle of correlation matrix
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
# Find features with correlation greater than a threshold
to_drop = [column for column in upper.columns if any(upper[column] > threshold)]
# Drop features
df.drop(to_drop, axis=1, inplace=True)
return df.iloc[:,:].values
def classify_lof(X):
state = np.random.RandomState(42)
detector = LocalOutlierFactor(n_neighbors=20, algorithm='auto', leaf_size=30, metric='minkowski', p=2, metric_params=None, contamination=0.5)
start = timer()
y_pred = detector.fit_predict(X)
end = timer()
execution_time = end - start
y_pred_1_m1 = y_pred # the y_pred is in 1 = normal, -1 = anomaly
return (execution_time, y_pred_1_m1)
for i in range(9):
device_index = i
device_id = device_index + 1
device_name = dn_nbaiot[device_index]
(X, y, Xdf) = get_nbaiot_device_data(device_id)
features_before = X.shape[1]
X = remove_correlated_features(Xdf, 0.95)
features_after = X.shape[1]
X_std = StandardScaler().fit_transform(X)
print(device_name)
print('total features', features_before)
print('final features', features_after)
print("method,execution_time,acc,tn,fp,fn,tp")
(execution_time, y_pred_1_m1) = classify_lof(X_std)
name = "LOF"
y_pred = y_pred_1_m1.copy()
y_pred[y_pred == -1] = 0 # convert -1 as 0 for anomaly
tn, fp, fn, tp = confusion_matrix(y, y_pred, labels=[0,1]).ravel()
acc = accuracy_score(y, y_pred)
method = name + '-1'
print(f'{method},{execution_time:.2f},{acc:.2f},{tn},{fp},{fn},{tp}')
# The actual prediction may be wrong because of data imbalance,
# need to swap the results
y_pred = y_pred_1_m1.copy()
y_pred[y_pred == 1] = 0 # convert 1 as 0 for anomaly
y_pred[y_pred == -1] = 1 # convert -1 as 1 for normal
tn, fp, fn, tp = confusion_matrix(y, y_pred, labels=[0,1]).ravel()
acc = accuracy_score(y, y_pred)
method = name + '-2'
print(f'{method},{execution_time:.2f},{acc:.2f},{tn},{fp},{fn},{tp}')
| comparison-lof nbv5.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # Workfow Management and Meta Job Scheduler
//
// This tutorial illustrates how computations can be executed on large
// (or small) compute servers, aka.
// high-performance-computers (HPC), aka. compute clusters, aka. supercomputers, etc.
//
// This means, the computation is not executed on the local workstation
// (or laptop) but on some other computer.
// This approach is particulary handy for large computations, which run
// for multiple hours or days, since a user can
// e.g. shutdown or restart his personal computer without killing the compute job.
//
// *BoSSS* features a set of classes and routines
// (an API, application programing interface) for communication with
// compute clusters. This is especially handy for **scripting**,
// e.g. for parameter studies, where dozens of computations
// have to be started and monitored.
// First, we initialize the new worksheet;
// Note:
// 1. This tutorial can be found in the source code repository as as `MetaJobManager.ipynb`.
// One can directly load this into Jupyter to interactively work with the following code examples.
// 2. **In the following line, the reference to `BoSSSpad.dll` is required**.
// You must either set `#r "BoSSSpad.dll"` to something which is appropirate for your computer
// (e.g. `C:\Program Files (x86)\FDY\BoSSS\bin\Release\net5.0\BoSSSpad.dll` if you installed the binary distribution),
// or, if you are working with the source code, you must compile `BoSSSpad` and put it side-by-side to this worksheet file
// (from the original location in the repository, you can use the scripts `getbossspad.sh`, resp. `getbossspad.bat`).
// + dotnet_interactive={"language": "csharp"}
#r "BoSSSpad.dll"
//#r "../../../src/L4-application/BoSSSpad/bin/Debug/net5.0/BoSSSpad.dll"
using System;
using System.Collections.Generic;
using System.Linq;
using ilPSP;
using ilPSP.Utils;
using BoSSS.Platform;
using BoSSS.Foundation;
using BoSSS.Foundation.Grid;
using BoSSS.Foundation.Grid.Classic;
using BoSSS.Foundation.IO;
using BoSSS.Solution;
using BoSSS.Solution.Control;
using BoSSS.Solution.GridImport;
using BoSSS.Solution.Statistic;
using BoSSS.Solution.Utils;
using BoSSS.Solution.Gnuplot;
using BoSSS.Application.BoSSSpad;
using BoSSS.Application.XNSE_Solver;
using static BoSSS.Application.BoSSSpad.BoSSSshell;
Init();
// -
//
// ## Batch Processing
//
// First, we have to select a *batch system* (aka.*execution queue*, aka. *queue*) that we want to use.
// Batch systems are a common approach to organize workloads (aka. compute jobs)
// on compute clusters.
// On such systems, a user typically does **not** starts a simulation manually/interactively.
// Instead, he specifies a so-called *compute job*. The *scheduler*
// (i.e. the batch system) collects
// compute jobs from all users on the compute cluster, sorts them according to
// some priority and puts the jobs into some queue, also called *batch*.
// The jobs in the batch are then executed in order, depending on the
// available hardware and the scheduling policies of the system.
//
// The *BoSSS* API provides front-ends (clients) for the following
// batch system software:
//
// - `BoSSS.Application.BoSSSpad.SlurmClient` for the
// Slurm Workload Manager (very prominent on Linux HPC systems)
// - `BoSSS.Application.BoSSSpad.MsHPC2012Client`
// for the Microsoft HPC Pack 2012 and higher
// - `BoSSS.Application.BoSSSpad.MiniBatchProcessorClient` for the
// mini batch processor, a minimalistic, *BoSSS*-internal batch system which mimiks
// a supercomputer batch system on the local machine.
//
// A list of clients for various batch systems, which are loaded at the
// `Init()` command can be configured through the
// `~/.BoSSS/etc/BatchProcessorConfig.json`-file.
// If this file is missing, a default setting, containing a
// mini batch processor, is initialized.
//
// The list of all execution queues can be accessed through:
// + dotnet_interactive={"language": "csharp"}
ExecutionQueues
// -
// For the sake of simplicuity, we pick the first one to execute the examples in this tutorial:
// + dotnet_interactive={"language": "csharp"}
var myBatch = ExecutionQueues[0];
// -
// ### Note on the Mini Batch Processor:
// The batch processor for local jobs can be started separately (by launching
// `MiniBatchProcessor.exe` or `dotnet MiniBatchProcessor.dll`), or from the worksheet;
// In the latter case, it depends on the operating system, whether the
// `MiniBatchProcessor.exe` is terminated with the notebook kernel, or not.
// If no mini-batch-processor is running, it is started
// at the initialization of the workflow management.
//
// ```csharp
// MiniBatchProcessor.Server.StartIfNotRunning(false);
// ```
//
// ## Initializing the workflow management
//
// In order to use the workflow management,
// the very first thing we have to do is to initialize it by defineing
// a **project name**, here it is `MetaJobManager_Tutorial`.
// This is used to generate names for the compute jobs and to
// identify sessions in the database:
// + dotnet_interactive={"language": "csharp"}
BoSSSshell.WorkflowMgm.Init("MetaJobManager_Tutorial");
// -
// We verify that we have no jobs defined so far ...
// + dotnet_interactive={"language": "csharp"}
BoSSSshell.WorkflowMgm.AllJobs
// + dotnet_interactive={"language": "csharp"}
// the folloowing line is part of the trest system and not neccesary in User worksheets:
NUnit.Framework.Assert.IsTrue(BoSSSshell.WorkflowMgm.AllJobs.Count == 0, "MetaJobManager tutorial: expecting 0 jobs on entry.");
// -
// The initialization of the Workflow Management environment already creates, resp. opens
// a *BoSSS database* whith the same name as the project name as the project.
// + dotnet_interactive={"language": "csharp"}
// From previous versions of the code, not required anymore:
//var myLocalDb = myBatch.CreateTempDatabase();
// -
// ### Notes on databases:
// * For expensive simulations, which run for days or longer, temporaray databases,
// which are created with a different name every time the notebook is executed,
// are typically **not** desired.
// Hence, one wants the compute jobs to persist, i.e. if the worksheet is re-executed (maybe on another day),
// but the computation
// has been successful somewhen in the past, this result is recovered from the database.
// In such a scenario, one cannot use a temporary database, but the default project database
// should be used instead.
// * Under certain circumstances, one could also create additional databases:
// Using the methods `BatchProcessorClient.CreateTempDatabase()`,
// resp. `BatchProcessorClinet.CreateOrOpenCompatibleDatabase(...)`.
// (as demonstrated below) ensures that the database is in a directory which can be accessed by the batch system.
// (Alternative functions, i.e. `BoSSSshell.CreateTempDatabase()` or `BoSSSshell.OpenOrCreateDatabase(...)`
// do not guarantee this and the user has to ensure an appropriate location.
// + dotnet_interactive={"language": "csharp"}
databases
// -
var myLocalDb = databases[0];
// ## Loading a BoSSS-Solver and Setting up a Simulation
//
// As an example, we use the workflow management tools to simulate
// incompressible channel flow, therefore we have to import the namespace,
// and repeat the steps from the IBM example (Tutorial 2) in order to setup the
// control object:
// + dotnet_interactive={"language": "csharp"}
using BoSSS.Application.XNSE_Solver;
// -
// We create a grid with boundary conditions:
// + dotnet_interactive={"language": "csharp"}
var xNodes = GenericBlas.Linspace(0, 10 , 41);
var yNodes = GenericBlas.Linspace(-1, 1, 9);
GridCommons grid = Grid2D.Cartesian2DGrid(xNodes, yNodes);
grid.DefineEdgeTags(delegate (double[] X) {
double x = X[0];
double y = X[1];
if (Math.Abs(y - (-1)) <= 1.0e-8)
return "wall"; // lower wall
if (Math.Abs(y - (+1)) <= 1.0e-8)
return "wall"; // upper wall
if (Math.Abs(x - (0.0)) <= 1.0e-8)
return "Velocity_Inlet"; // inlet
if (Math.Abs(x - (+10.0)) <= 1.0e-8)
return "Pressure_Outlet"; // outlet
throw new ArgumentOutOfRangeException("unknown domain");
});
// -
// One can save this grid explicitly to a database, but it is not a must;
// The grid should be saved automatically, when the job is activated.
// + dotnet_interactive={"language": "csharp"}
//myLocalDb.SaveGrid(ref grid);
// -
// Next, we create the control object for the incompressible simulation:
// + dotnet_interactive={"language": "csharp"}
var c = new XNSE_Control();
// general description:
int k = 1;
string desc = "Steady state, channel, k" + k;
c.SessionName = "SteadyStateChannel";
c.ProjectDescription = desc;
c.savetodb = true;
c.Tags.Add("k" + k);
// setting the grid:
c.SetGrid(grid);
// DG polynomial degree
c.SetDGdegree(k);
// Physical parameters:
double reynolds = 20;
c.PhysicalParameters.rho_A = 1;
c.PhysicalParameters.mu_A = 1.0/reynolds;
// Timestepping properties:
c.TimesteppingMode = AppControl._TimesteppingMode.Steady;
// -
// The specification of boundary conditions and initial values
// is a bit more complicated if the job manager is used:
//
// Since the solver is executed in an external program, the control object
// has to be saved in a file. For lots of complicated objects,
// especially for delegates, C\# does not support serialization
// (converting the object into a form that can be saved on disk, or
// transmitted over a network), so a workaround is needed.
// This is achieved e.g. by the **Formula** object, where a C\#-formula
// is saved as a string.
// + dotnet_interactive={"language": "csharp"}
var WallVelocity = new Formula("X => 0.0", false); // 2nd Argument=false says that its a time-indep. formula.
// -
// Testing the formula:
// + dotnet_interactive={"language": "csharp"}
WallVelocity.Evaluate(new[]{0.0, 0.0}, 0.0) // evaluationg at (0,0), at time 0
// + dotnet_interactive={"language": "csharp"}
// [Deprecated]
/// A disadvantage of string-formulas is that they look a bit ``alien''
/// within the worksheet; therefore, there is also a little hack which allows
/// the conversion of a static memeber function of a static class into a
/// \code{Formula} object:
// + dotnet_interactive={"language": "csharp"}
// Deprecated, this option is no longer supported in .NET5
static class StaticFormulas {
public static double VelX_Inlet(double[] X) {
//double x = X[0];
double y = X[0];
double UX = 1.0 - y*y;
return UX;
}
public static double VelY_Inlet(double[] X) {
return 0.0;
}
}
// + dotnet_interactive={"language": "csharp"}
// InletVelocityX = GetFormulaObject(StaticFormulas.VelX_Inlet);
//var InletVelocityY = GetFormulaObject(StaticFormulas.VelY_Inlet);
// + dotnet_interactive={"language": "csharp"}
var InletVelocityX = new Formula("X => 1 - X[0]*X[0]", false);
var InletVelocityY = new Formula("X => 0.0", false);
// -
// Finally, we set boundary values for our simulation. The initial values
// are set to zero per default; for the steady-state simulation initial
// values are irrelevant anyway:
//
// Initial Values are set to 0
// + dotnet_interactive={"language": "csharp"}
c.BoundaryValues.Clear();
c.AddBoundaryValue("wall", "VelocityX", WallVelocity);
c.AddBoundaryValue("Velocity_Inlet", "VelocityX", InletVelocityX);
c.AddBoundaryValue("Velocity_Inlet", "VelocityY", InletVelocityY);
c.AddBoundaryValue("Pressure_Outlet");
// -
// ## Activation and Monitoring of the the Job
//
// Finally, we are ready to deploy the job at the batch processor;
// In a usual work flow scenario, we **do not** want to (re-) submit the
// job every time we run the worksheet -- usually, one wants to run a job once.
//
// The concept to overcome this problem is job activation. If a job is
// activated, the meta job manager first checks the databases and the batch
// system, if a job with the respective name and project name is already
// submitted. Only if there is no information that the job was ever submitted
// or started anywhere, the job is submitted to the respective batch system.
//
// First, a `Job* -object is created from the control object:
// + dotnet_interactive={"language": "csharp"}
var JobLocal = c.CreateJob();
// -
// This job is not activated yet, it can still be configured:
// + dotnet_interactive={"language": "csharp"}
JobLocal.Status
// + dotnet_interactive={"language": "csharp"}
// Test:
NUnit.Framework.Assert.IsTrue(JobLocal.Status == JobStatus.PreActivation);
// -
// ### Starting the compute Job
// One can change e.g. the number of MPI processes:
// + dotnet_interactive={"language": "csharp"}
JobLocal.NumberOfMPIProcs = 1;
// -
// Note that these jobs are desigend to be **persistent**:
// This means the computation is only started
// **once for a given control object**, no matter how often the worksheet
// is executed.
//
// Such a behaviour is useful for expensive simulations, which run on HPC
// servers over days or even weeks. The user (you) can close the worksheet
// and maybe open and execute it a few days later, and he can access
// the original job which he submitted a few days ago (maybe it is finished
// now).
//
// Then, the job is activated, resp. submitted, resp. deployed
// to one batch system.
// If job persistency is not wanted, traces of the job can be removed
// on request during activation, causing a fresh job deployment at the
// batch system:
// + dotnet_interactive={"language": "csharp"}
JobLocal.Activate(myBatch); // execute the job in 'myBatch'
// -
// All jobs can be listed using the workflow management:
// + dotnet_interactive={"language": "csharp"}
BoSSSshell.WorkflowMgm.AllJobs
// -
// Check the present job status:
// + dotnet_interactive={"language": "csharp"}
JobLocal.Status
// + dotnet_interactive={"language": "csharp"}
/// BoSSScmdSilent BoSSSexeSilent
NUnit.Framework.Assert.IsTrue(
JobLocal.Status == JobStatus.PendingInExecutionQueue
|| JobLocal.Status == JobStatus.InProgress
|| JobLocal.Status == JobStatus.FinishedSuccessful);
// -
// ### Evaluation of Job
// Here, we block until both of our jobs have finished:
// + dotnet_interactive={"language": "csharp"}
BoSSSshell.WorkflowMgm.BlockUntilAllJobsTerminate(1000);
// -
// We examine the output and error stream of the job:
// This directly accesses the *\tt stdout*-redirection of the respective job
// manager, which may contain a bit more information than the
// `Stdout`-copy in the session directory.
// + dotnet_interactive={"language": "csharp"}
JobLocal.Stdout
// -
// Additionally we display the error stream and hope that it is empty:
// + dotnet_interactive={"language": "csharp"}
JobLocal.Stderr
// -
// We can also obtain the session
// which was stored during the execution of the job:
// + dotnet_interactive={"language": "csharp"}
var Sloc = JobLocal.LatestSession;
Sloc
// -
// We can also list all attempts to run the job at the assigend processor:
// + dotnet_interactive={"language": "csharp"}
JobLocal.AllDeployments
// + dotnet_interactive={"language": "csharp"}
NUnit.Framework.Assert.IsTrue(JobLocal.AllDeployments.Count == 1, "MetaJobManager tutorial: Found more than one deployment.");
// -
// Finally, we check the status of our jobs:
// + dotnet_interactive={"language": "csharp"}
JobLocal.Status
// -
// If anything failed, hints on the reason why are provides by the `GetStatus` method:
// + dotnet_interactive={"language": "csharp"}
JobLocal.GetStatus(WriteHints:true)
// + dotnet_interactive={"language": "csharp"}
NUnit.Framework.Assert.IsTrue(JobLocal.Status == JobStatus.FinishedSuccessful, "MetaJobManager tutorial: Job was not successful.");
// -
// ## Exporting Plots
// Each run of the solver corresponds to one **session** in the database. A session is basically a collection of information on the entire solver run, i.e. the simulation result, input and solver settings as well as meta-data such as computer and daten and time.
//
// Since in this tutorial only one solver run was executed, there is only one session in the Workflow Management (`wmg` is just an alias for `BoSSSshell.WorkflowMgm.`):
wmg.Sessions
// We select the first (and only) session and create an **export instruction** object.
// The **supersampling** setting increases the output resolution.
// This is required to vizualize high-order DG ploynomials with the low-order Tecplot-format.
// Tecplot can only vizualize a linear interpolation within a cell. With a second-degree supersampling,
// each cell is subdivided twice (in 2D, one subdivision is 4 cells, i.e. 2 subdivisions are $4^2 = 16$ cells).
// In this way, the curve of e.g. a secondd order polynomial can be represented with the linear interpolation over 16 cells.
var outPath = wmg.Sessions[0].Export().WithSupersampling(2).Do();
// On the respective directory (see output above) one should finaaly find plot-files which than can be used
// for further post processing in third-party software such as Paraview, LLNL Visit or Tecplot.
//
// The `Do()` command returns the location of the output files:
outPath
// To finalize this tutorial, we list all files in the plot output directory:
System.Threading.Thread.Sleep(10000); // just wait for the external plot application to finish
System.IO.Directory.GetFiles(outPath, "*").Select(fullPath => System.IO.Path.GetFileName(fullPath))
| doc/handbook/MetaJobManager/MetaJobManager.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Install dependencies for this example
# Note: This does not include itkwidgets, itself
import sys
# !{sys.executable} -m pip install itk-io plotly>=3.0.0 bqplot
# +
try:
from urllib.request import urlretrieve
except ImportError:
from urllib import urlretrieve
import os
import itk
import numpy as np
from itkwidgets import line_profile
# -
# Download data
file_name = 'statueLeg.nrrd'
if not os.path.exists(file_name):
url = 'https://data.kitware.com/api/v1/file/5b8446868d777f43cc8d5ec1/download'
urlretrieve(url, file_name)
# Create a line profile from pre-specified end points.
image = itk.imread(file_name)
line_profile(image, point1=[35.3, 169.7, 113.6], point2=[325.1, 197.3, 204.6], ui_collapsed=True)
# If the line profile is not initialized with end points, a plane view is presented. Click twice in the plane to select the end points.
#
# In any view, click and drag the end points to change the line profile location.
line_profile(image)
# Multiple plotting libraries are supported, including *ipympl*, a.k.a. jupyter-matplotlib, *bqplot*, and *plotly*.
line_profile(image, plotter='ipympl', point1=[35.3, 169.7, 113.6], point2=[325.1, 197.3, 204.6], ui_collapsed=True)
line_profile(image, plotter='bqplot', point1=[35.3, 169.7, 113.6], point2=[325.1, 197.3, 204.6], ui_collapsed=True)
# Compare multiple images along the same line with a *comparisons* dictionary of *label, image* pairs.
image = itk.imread(file_name)
processed = {'Sigma ' + str(sigma): itk.SmoothingRecursiveGaussianImageFilter(image, Sigma=sigma) for sigma in np.linspace(1.0, 3.0, 3) }
line_profile(image, comparisons=processed, point1=[35.3, 169.7, 113.6], point2=[325.1, 197.3, 204.6], ui_collapsed=True)
| examples/LineProfile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Growler
#
# > COhPy - Jan 25, 2016
#
# > <NAME>
# ## Python 3.4
#
# * Includes a new module : `asyncio`
# * "Single-threaded concurrent code using coroutines"
# * Already available for python via external libraries like Tornado, Diesel, Greenlets… etc.
# * These do not currently use the built-in library (backwards compatible)
# * There are a few new frameworks emerging which do utilize asyncio: Vase, Rainfall
# ## Growler
#
# * Yet another python web framework
# * A web framework which utilizes the asyncio module at its core
# * Is that enough?
# * Use a middleware-based style similar to nodejs' Connect/Express frameworks
#
# > People like Python; people like Express… why not?
# ## Extensive use of Middleware
#
# * Web application designed as a pipe
# * Add middleware functions which process the request and response and pass to next function
#
# * ```python
# app.use(_middleware_)
# app.get('/path', endpoint)
# app.post('/user/logout', logout)
# ```
#
# * middleware can be run to redirect requests, render templates, parse specific requests (websockets/authentication, etc…)
# ## Running
# ```python
# import growler
#
# app = growler.App()
# …
# app.get('/', get_index)
# app.post('/', post_index)
#
# app.run()
# ```
# ## Running with Decorators
# ```python
# @app.get('/')
# def index(req, res, next):
# if req.username:
# user = req.username
# res.send("hello" + user)
# else:
# res.send("who are you?")
#
# app.run()
# ```
# ## Things I’ve learned
# * Asyncio coroutines are not “normal” python coroutines
# * Cannot 'send' a value to a yield
# * Wrap functions in tasks! (yield from them)
# ## Wishlist
#
# * Decorated classes
# * Class-Based middleware
# * Route "overlap"
# * /user/id
# * /user/name
| 2014-09/Growler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# #### This notebook demonstrates how to run QIIME2 on Biowulf for development of pipeline for Nephele
#
# ##### (Optional) Update to the latest qiime 2
# * wget https://data.qiime2.org/distro/core/qiime2-2020.11-py36-linux-conda.yml
# * conda env create -n qiime2-2020.11 --file qiime2-2020.11-py36-linux-conda.yml
#
# ##### Step 1: in terminal window 1
# ```
# ssh -X <EMAIL>
# source /data/$USER/conda/etc/profile.d/conda.sh
# conda activate base
# conda activate qiime2-2020.11
# jupyter serverextension enable --py qiime2 --sys-prefix
# cd /data/quinonesm/
# ```
#
# ##### follow https://hpc.nih.gov/apps/jupyter.html
# ```
# module load tmux
# tmux
# sinteractive --gres=lscratch:5 --mem=20g --tunnel
# ```
#
# ##### Step 2: in terminal window 2: open the tunnel per instructions (for example)
# ```
# ssh -L 38772:localhost:38772 <EMAIL>
# ```
#
# ##### Step 3: back in the terminal window 1
# ```
# module load jupyter && jupyter lab --ip localhost --port $PORT1 --no-browser
# ```
#
#
# #### Info for pipeline users:
# ##### diagrams: https://docs.qiime2.org/2020.11/tutorials/overview/#let-s-get-oriented-flowcharts
# in particular https://docs.qiime2.org/2020.11/tutorials/overview/#derep-denoise
# ##### basic tutorial: https://docs.qiime2.org/2020.11/tutorials/moving-pictures/
#
# ##### To launch this notebook locally (in Mac), first activate the environment, then pass it to jupyter
# ```
# source /opt/miniconda3/bin/activate*
# conda activate qiime2-2020.11
# jupyter serverextension enable --py qiime2 --sys-prefix
# jupyter notebook
# ```
# _______
# #### The pipeline below will use the same dataset available in the Nephele User Guide <https://nephele.niaid.nih.gov/user_guide/>
cd /data/quinonesm/Nephele/
qiime info
# ### Part 1a: Import of demultiplexed PairedEnd reads using indicated filepaths
# ***The validation of the mapping file will be different for this pipeline because it needs the sample-id and column headers as forward-absolute-filepath, reverse-absolute-filepath. For single end, it only needs sample-id and absolute-filepath***
#
# ***There will be 3 options for import (paired, single and previously joined)***
# +
# The manifest file (mapping file) must have the absolute path as below. For the single end, the column header should say "absolute-filepath" instead of "forward-absolute-filepath".
#sample-id forward-absolute-filepath reverse-absolute-filepath Antibiotic Animal Day Tissue
#A22831 $PWD/N2_16S_example_data/22831_S41_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22831_S41_R2_subsample.fastq.gz Control RhDCBC 347 SwabJejunum
#A22833 $PWD/N2_16S_example_data/22833_S45_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22833_S45_R2_subsample.fastq.gz Control RhDCBC 347 SwabRectum
#A22349 $PWD/N2_16S_example_data/22349_S26_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22349_S26_R2_subsample.fastq.gz Control RhDCVf 274 SwabCecum
#A22192 $PWD/N2_16S_example_data/22192_S22_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22192_S22_R2_subsample.fastq.gz Vancomycin RhCL4c 239 SwabRectum
#A22187 $PWD/N2_16S_example_data/22187_S19_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22187_S19_R2_subsample.fastq.gz Vancomycin RhCL4c 239 SwabIleum
#A22061 $PWD/N2_16S_example_data/22061_S5_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22061_S5_R2_subsample.fastq.gz Vancomycin RhDCAV 178 SwabTransverseColon
#A22057 $PWD/N2_16S_example_data/22057_S2_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22057_S2_R2_subsample.fastq.gz Vancomycin RhDCAV 178 SwabJejunum
#A22145 $PWD/N2_16S_example_data/22145_S14_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22145_S14_R2_subsample.fastq.gz Control RhDCKj 239 SwabIleum
#A22350 $PWD/N2_16S_example_data/22350_S27_R1_subsample.fastq.gz $PWD/N2_16S_example_data/22350_S27_R2_subsample.fastq.gz Control RhDCVf 274 SwabTransverseColon
#7pRecSw478.1 $PWD/N2_16S_example_data/23572_S307_R1_subsample.fastq.gz $PWD/N2_16S_example_data/23572_S307_R2_subsample.fastq.gz Vancomycin RhCL7p 478 SwabRectum
# -
# ##### PairedEnd import
# import paired end data - sample data from Nephele User Guide. The mapping file how has a different format of column headers.
qiime tools import \
--type 'SampleData[PairedEndSequencesWithQuality]' \
--input-path N2_16S_mapping_file.txt \
--output-path paired-end-demux.qza \
--input-format PairedEndFastqManifestPhred33V2
# view summary of demultiplexed imported paired end data
qiime demux summarize \
--i-data paired-end-demux.qza \
--o-visualization paired-end-demux.qzv
# join pairs
qiime vsearch join-pairs \
--i-demultiplexed-seqs paired-end-demux.qza \
--o-joined-sequences paired-end-joined.qza
# view summary of joined pairs
qiime demux summarize \
--i-data paired-end-joined.qza \
--o-visualization paired-end-joined.qzv
qiime quality-filter q-score \
--i-demux paired-end-joined.qza \
--p-min-quality 20 \
--o-filtered-sequences paired-end-joined-filtered.qza \
--o-filter-stats paired-end-joined-filter-stats.qza
# view summary of joined pairs and filtered pairs
qiime demux summarize \
--i-data paired-end-joined-filtered.qza \
--o-visualization paired-end-joined-filtered.qzv
# create summary table for paired end
qiime metadata tabulate \
--m-input-file paired-end-joined-filter-stats.qza \
--o-visualization paired-end-joined-filter-stats.qzv
# dereplicate sequences
qiime vsearch dereplicate-sequences \
--i-sequences paired-end-joined-filtered.qza \
--o-dereplicated-table table.qza \
--o-dereplicated-sequences rep-seqs.qza
# ### Part 1b: Import of demultiplexed SingleEnd reads using indicated filepaths
# ##### SingleEnd import
# import - here is the import of Single-end but this should be done for the Paired-End as well
qiime tools import \
--type 'SampleData[SequencesWithQuality]' \
--input-path N2_16S_mapping_file_singleend.txt \
--output-path single-end-demux.qza \
--input-format SingleEndFastqManifestPhred33V2
# view summary of demultiplexed
qiime demux summarize \
--i-data single-end-demux.qza \
--o-visualization single-end-demux.qzv
# filter
qiime quality-filter q-score \
--i-demux single-end-demux.qza \
--p-min-quality 20 \
--o-filtered-sequences single-end-demux-filtered.qza \
--o-filter-stats single-end-demux-filter-stats.qza
# dereplicate sequences
qiime vsearch dereplicate-sequences \
--i-sequences single-end-demux-filtered.qza \
--o-dereplicated-table table.qza \
--o-dereplicated-sequences rep-seqs.qza
# ### Part 1c: Import of demultiplexed prejoined reads using indicated filepaths
# Note: For pre-joined reads (as in the output of the QC pipe), we will need to add another option
qiime tools import \
--input-path N2_16S_mapping_file_joined.txt \
--output-path fj-joined-demux.qza \
--type SampleData[JoinedSequencesWithQuality] \
--input-format SingleEndFastqManifestPhred33V2
# view summary of demultiplexed
qiime demux summarize \
--i-data fj-joined-demux.qza \
--o-visualization fj-joined-demux.qzv
# filter
qiime quality-filter q-score \
--i-demux fj-joined-demux.qza \
--p-min-quality 20 \
--o-filtered-sequences joined-demux-filtered.qza \
--o-filter-stats joined-demux-filter-stats.qza
# dereplicate sequences
qiime vsearch dereplicate-sequences \
--i-sequences joined-demux-filtered.qza \
--o-dereplicated-table table.qza \
--o-dereplicated-sequences rep-seqs.qza
# ### Part 2: Clustering
# ***This can be done for both SingleEnd or PairedEnd. We will give the denovo, closed and open reference options but give the closed as default because it is the lowest compute***
# ##### Part 2a: Denovo - Let's do 0.97 default for --p-perc-identity
qiime vsearch cluster-features-de-novo \
--i-table table.qza \
--i-sequences rep-seqs.qza \
--p-perc-identity 0.97 \
--o-clustered-table table-dn-97.qza \
--o-clustered-sequences rep-seqs-dn-97.qza
# create stats and summary of rep-seqs data from denovo clustering
qiime feature-table tabulate-seqs \
--i-data rep-seqs-dn-97.qza \
--o-visualization rep-seqs-dn-97.qzv
# ##### Import OTU reference sets for clustering
# +
# import otu tables from greengenes
qiime tools import \
--input-path /Users/quinonesm/OneDrive_National_Institutes_of_Health/Nephele/qiime2_MiSeq_paired/gg_13_8_otus/rep_set/99_otus.fasta \
--output-path 99_otus.qza \
--type 'FeatureData[Sequence]'
# import otu tables
qiime tools import \
--input-path /Users/quinonesm/OneDrive_National_Institutes_of_Health/Nephele/qiime2_MiSeq_paired/gg_13_8_otus/rep_set/97_otus.fasta \
--output-path 97_otus.qza \
--type 'FeatureData[Sequence]'
# import otu tables
qiime tools import \
--input-path /Users/quinonesm/OneDrive_National_Institutes_of_Health/Nephele/qiime2_MiSeq_paired/gg_13_8_otus/rep_set/85_otus.fasta \
--output-path 85_otus.qza \
--type 'FeatureData[Sequence]'
# -
# ##### Part 2b: Open Reference
# use 85, 97, or 99 OTU
qiime vsearch cluster-features-open-reference \
--i-table table.qza \
--i-sequences rep-seqs.qza \
--i-reference-sequences 97_otus.qza \
--p-perc-identity 0.97 \
--o-clustered-table table-or-97.qza \
--o-clustered-sequences rep-seqs-or-97.qza \
--o-new-reference-sequences new-ref-seqs-or-97.qza
# create stats and summary of rep-seqs data from open-reference clustering
qiime feature-table tabulate-seqs \
--i-data rep-seqs-or-97.qza \
--o-visualization rep-seqs-or-97.qzv
# rename files
mv rep-seqs-or-97.qza rep-seqs.qza
mv table-or-97.qza table.qza
# ##### Part 2c: Closed reference
qiime vsearch cluster-features-closed-reference \
--i-table table.qza \
--i-sequences rep-seqs.qza \
--i-reference-sequences 97_otus.qza \
--p-perc-identity 0.97 \
--o-clustered-table table-cr-97.qza \
--o-clustered-sequences rep-seqs-cr-97.qza \
--o-unmatched-sequences unmatched-cr-97.qza
# create stats and summary of rep-seqs data from closed-reference clustering
qiime feature-table tabulate-seqs \
--i-data rep-seqs-cr-97.qza \
--o-visualization rep-seqs-cr-97.qzv
# rename files
mv rep-seqs-cr-97.qza rep-seqs.qza
mv table-cr-97.qza table.qza
# summarized denoised sequence variants
# ### Part 3: Denoising with Deblur
# ***Perform sequence quality control for Illumina data using the Deblur
# workflow with a 16S reference as a positive filter. Only forward reads are
# supported at this time. The specific reference used is the 88% OTUs from
# Greengenes 13_8. This mode of operation should only be used when data were
# generated from a 16S amplicon protocol on an Illumina platform. The
# reference is only used to assess whether each sequence is likely to be 16S
# by a local alignment using SortMeRNA with a permissive e-value; the
# reference is not used to characterize the sequences.Reference: docs.qiiime2.org)***
qiime deblur denoise-16S \
--i-demultiplexed-seqs single-end-demux-filtered.qza \
--p-trim-length 200 \
--o-representative-sequences single-end-rep-seqs-deblur.qza \
--o-table single-end-table-deblur.qza \
--p-sample-stats \
--o-stats deblur-single-end-denoised-stats.qza
# summarize stats from deblur processing
qiime metadata tabulate \
--m-input-file deblur-single-end-qfilter-stats.qza \
--o-visualization deblur-single-end-qfilter-stats.qzv
qiime deblur visualize-stats \
--i-deblur-stats deblur-single-end-denoised-stats.qza \
--o-visualization deblur-single-end-denoised-stats.qzv
# summarized denoised sequence variants
qiime feature-table summarize \
--i-table single-end-table-deblur.qza \
--o-visualization single-end-table-deblur.qzv \
--m-sample-metadata-file N2_16S_mapping_file_singleend.txt
qiime feature-table tabulate-seqs \
--i-data single-end-rep-seqs-deblur.qza \
--o-visualization single-end-rep-seqs-deblur.qzv
# +
# rename - This one I didn't run since I wanted to use the open clustering for now
# mv single-end-rep-seqs-deblur.qza rep-seqs.qza
# mv single-end-table-deblur.qza table.qza
# -
# ### Part 4: Create a phylogenetic tree to be used in downstream diversity steps
# ***This step will take input from the Denoising steps above (either Deblur or vsearch clustering)***
# +
# Option A: (default) inferring phylogeny using an reference based fragment insertion approach
# download both files (greengenes and silva) from https://docs.qiime2.org/2020.11/data-resources/#sepp-reference-databases
# sepp-refs-silva-128.qza or sepp-refs-gg-13-8.qza
qiime fragment-insertion sepp \
--i-representative-sequences rep-seqs.qza \
--i-reference-database sepp-refs-silva-128.qza \
--o-tree insertion-tree.qza \
--p-threads 2 \
--o-placements insertion-placements.qza
# -
# Option B: denovo phylogenetic tree - tested with closed reference
qiime phylogeny align-to-tree-mafft-fasttree \
--i-sequences rep-seqs.qza \
--o-alignment aligned-rep-seqs.qza \
--o-masked-alignment masked-aligned-rep-seqs.qza \
--o-tree unrooted-tree.qza \
--o-rooted-tree rooted-tree.qza
# ### Part 5: Alpha and Beta Diversity - This is similar as the Downstream Analysis pipeline except that it uses the phylogenetic tree
# ##### Use the sampling depth function as in here https://github.niaid.nih.gov/bcbb/nephele2/blob/next_release/pipelines/DS_analysis_16S/Qiime_2.0_Core_Diversity.md
#
# ##### For the metadata, only a sample ID column is required. Columns are inferred as numerical or categorical but optionally we could add a line to indicate column type.
# https://docs.google.com/spreadsheets/d/1hBo_NWijLILEFYJrYs7R_Bwc7i_n3ZmPKUQetpVk-pk/edit#gid=0
# core metrics using phylogeny - similar to what is available in the downstream analysis pipeline.
rm -rf core-metrics-results_5k_insertion
qiime diversity core-metrics-phylogenetic \
--i-phylogeny insertion-tree.qza \
--i-table table.qza \
--p-sampling-depth 5000 \
--m-metadata-file N2_16S_mapping_file.txt \
--output-dir core-metrics-results_5k_insertion
# core metrics using phylogeny - similar to what is available in the downstream analysis pipeline.
rm -rf core-metrics-results_5k
qiime diversity core-metrics-phylogenetic \
--i-phylogeny rooted-tree.qza \
--i-table table.qza \
--p-sampling-depth 5000 \
--m-metadata-file N2_16S_mapping_file.txt \
--output-dir core-metrics-results_5k
# +
# alpha diversity group significance
qiime diversity alpha-group-significance \
--i-alpha-diversity core-metrics-results_5k/faith_pd_vector.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--o-visualization core-metrics-results_5k/faith-pd-group-significance.qzv
qiime diversity alpha-group-significance \
--i-alpha-diversity core-metrics-results_5k/evenness_vector.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--o-visualization core-metrics-results_5k/evenness-group-significance.qzv
# -
# ### Part 6: Alpha rarefaction
qiime diversity alpha-rarefaction \
--i-table table.qza \
--i-phylogeny rooted-tree.qza \
--p-max-depth 10000 \
--m-metadata-file N2_16S_mapping_file.txt \
--o-visualization alpha-rarefaction.qzv
# ### Part 7: Taxonomy classification
# +
# get the classifiers (Silva and Greengenes) - These are the ones trained only to the V4 region.
#Naive Bayes classifiers trained on:
#Silva 138 99% OTUs full-length sequences (MD5: fddefff8bfa2bbfa08b9cad36bcdf709)
#Silva 138 99% OTUs from 515F/806R region of sequences (MD5: 28105eb0f1256bf38b9bb310c701dc4e)
#Greengenes 13_8 99% OTUs full-length sequences (MD5: 03078d15b265f3d2d73ce97661e370b1)
#Greengenes 13_8 99% OTUs from 515F/806R region of sequences (MD5: 682be39339ef36a622b363b8ee2ff88b)
# we could make additional classifiers for other 16S regions or the entire 16S
# classifier for full 16S using OTUs 99%
# wget https://data.qiime2.org/2020.11/common/silva-138-99-nb-classifier.qza
# wget https://data.qiime2.org/2020.11/common/gg-13-8-99-nb-classifier.qza
# classifier for V4 region using OTUs 99%
#wget https://data.qiime2.org/2020.11/common/gg-13-8-99-515-806-nb-classifier.qza
#wget https://data.qiime2.org/2020.11/common/silva-138-99-515-806-nb-classifier.qza
# (optional) - make a classifier by following https://docs.qiime2.org/2020.11/tutorials/feature-classifier/
# +
# First allow user to select option to classify (sklearn or vsearch), then allow selection of classifier (silva or greengenes).
# The reads-per-batch allow for faster processing
# +
# classify below (tested on biowulf interactive session as: sinteractive --mem=20g --cpus-per-task=4)
# -
# ##### Part 7a: Classify method sklearn
# +
# Select to classify with (slow: full length or faster: only for V4 region)
# option 1: SILVA V4 region
qiime feature-classifier classify-sklearn \
--i-reads rep-seqs.qza \
--i-classifier silva-138-99-515-806-nb-classifier.qza \
--o-classification taxonomy.qza \
--p-reads-per-batch 100
# option 2: Greengenes V4 region
qiime feature-classifier classify-sklearn \
--i-reads rep-seqs.qza \
--i-classifier gg-13-8-99-515-806-nb-classifier.qza \
--o-classification taxonomy.qza \
--p-reads-per-batch 100
# option 3: SILVA full length
qiime feature-classifier classify-sklearn \
--i-reads rep-seqs.qza \
--i-classifier silva-138-99-nb-classifier.qza \
--o-classification taxonomy.qza \
--p-reads-per-batch 100
# option 4: Greengenes full length
qiime feature-classifier classify-sklearn \
--i-reads rep-seqs.qza \
--i-classifier gg-13-8-99-nb-classifier.qza \
--o-classification taxonomy.qza \
--p-reads-per-batch 100
# -
# ##### Part 7b:Classify method vsearch
# +
# two options (silva and greengenes)
qiime feature-classifier classify-consensus-vsearch \
--i-query rep-seqs.qza \
--i-reference-reads silva-138-99-seqs.qza \
--i-reference-taxonomy silva-138-99-tax.qza \
--o-classification taxonomy.qza
# -
# view taxonomy assignments
qiime metadata tabulate \
--m-input-file taxonom.qza \
--o-visualization taxonomy.qzv
# ### Part 8:Make Barplots
# make table of rep sequences
qiime feature-table tabulate-seqs \
--i-data rep-seqs.qza \
--o-visualization rep_set.qzv
# start by filtering or subsample the table to 2000 for example
qiime feature-table filter-samples \
--i-table table.qza \
--p-min-frequency 2000 \
--o-filtered-table table_2k.qza
# make barplot
qiime taxa barplot \
--i-table table_2k.qza \
--i-taxonomy taxonomy.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--o-visualization taxa_barplot.qzv
# ___________
# #### Part 9: Optional analyses - Let's not worry about this yet
# ***(I wonder if these can be done here or in the downstream analysis pipeline). It will need metadata file columns headers to be presented to the user for the user to select from. For each of the metadata colums, the beta-group-significance function will be run.***
# beta group significance - This will be an optional step
qiime diversity beta-group-significance \
--i-distance-matrix core-metrics-results_5k/unweighted_unifrac_distance_matrix.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--m-metadata-column Antibiotic \
--o-visualization core-metrics-results_5k/unweighted-unifrac-Antibiotic-significance.qzv \
--p-pairwise
# permanova for exploring large differences between groups even with large within group variances
qiime diversity beta-group-significance \
--i-distance-matrix core-metrics-results_5k/unweighted_unifrac_distance_matrix.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--m-metadata-column Antibiotic \
--o-visualization core-metrics-results_5k/unweighted-unifrac-Antibiotic-significance_disp.qzv \
--p-method permdisp
# adonis
qiime diversity adonis \
--i-distance-matrix core-metrics-results_5k/unweighted_unifrac_distance_matrix.qza \
--m-metadata-file N2_16S_mapping_file.txt \
--o-visualization core-metrics-results_5k/unweighted_Antibiotic+Animal_adonis.qzv \
--p-formula Antibiotic+Animal
# +
#Other analyses that can be added:
#1) To determine if the continuous sample metadata is correlated with sample composition, an association test can be run using:
#qiime metadata distance-matrix in combination with qiime diversity mantel and qiime diversity bioenv commands.
#2) differential abundance
# -
# create stats and summary of rep-seqs data from denovo clustering
qiime feature-table tabulate-seqs \
--i-data rep-seqs-dn-99.qza \
--o-visualization rep-seqs-dn-99.qzv
| pipelines/QIIME2_16S/qiime2_nephele_paired_pipe_nodada.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Installs
pip install pymongo==3.11.0
# # Imports
from os import environ
from pprint import pprint as pp
from pymongo import MongoClient
# # Database Link
client = MongoClient(environ["MONGO_PORT_27017_TCP_ADDR"], 27017)
database = client.polyplot
collection = database["graticules"]
# # Basic Functions
# +
def drop_collection():
collection.drop()
def find_graticule(query={}, filter={"_id": 0}):
return dict(collection.find_one(query, filter))
def find_graticules(query={}, filter={"_id": 0}, sort=[("step", -1)], limit=0):
collection.create_index(sort)
return list(collection.find(query, filter).sort(sort).limit(limit))
| notes/collections/graticules.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="2T1wWDPa5bO_"
# # Formati dati 1 - Introduzione
#
# ## [Scarica zip esercizi](../_static/generated/formats.zip)
#
# [Naviga file online](https://github.com/DavidLeoni/softpython-it/tree/master/formats)
#
# -
# In questi tutorial parleremo di formati dei dati:
#
# File testuali:
#
# * File a linee
# * CSV
# * breve panoramica sui cataloghi open data
# * menzione licenze (Creative Commons CC-Zero)
# * menzione JSON
#
# Menzione file binari:
#
# * fogli Excel
#
# ### Che fare
#
#
# - scompatta lo zip in una cartella, dovresti ottenere qualcosa del genere:
#
# ```
#
# formats
# formats1-lines.ipynb
# formats1-lines-sol.ipynb
# formats2.ipynb
# formats2-csv-sol.ipynb
# formats3-json.ipynb
# formats3-json-sol.ipynb
# formats4-chal.ipynb
# jupman.py
# ```
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Per essere visualizzato correttamente, il file del notebook DEVE essere nella cartella szippata.
# </div>
#
# - apri il Jupyter Notebook da quella cartella. Due cose dovrebbero aprirsi, prima una console e poi un browser. Il browser dovrebbe mostrare una lista di file: naviga la lista e apri il notebook `formats.ipynb`
# - Prosegui leggendo il file degli esercizi, ogni tanto al suo interno troverai delle scritte **ESERCIZIO**, che ti chiederanno di scrivere dei comandi Python nelle celle successive.
#
# Scorciatoie da tastiera:
#
# * Per eseguire il codice Python dentro una cella di Jupyter, premi `Control+Invio`
# * Per eseguire il codice Python dentro una cella di Jupyter E selezionare la cella seguente, premi `Shift+Invio`
# * Per eseguire il codice Python dentro una cella di Jupyter E creare una nuova cella subito dopo, premi `Alt+Invio`
# * Se per caso il Notebook sembra inchiodato, prova a selezionare `Kernel -> Restart`
#
#
#
# + [markdown] colab_type="text" id="fC_te77v5bPE"
# ## File a linee
#
# I file a linee tipicamente sono file di testo che contengono informazioni raggruppate per linee. Un esempio usando personaggi storici potrebbe essere un file così:
#
# ```
# Leonardo
# <NAME>
# Sandro
# Botticelli
# Niccolò
# Macchiavelli
# ```
#
# Si nota subito una regolarità: le prime due linee contengono i dati di <NAME>, prima il nome e poi il cognome. Le successive due linee hanno invece i dati di <NAME>, di nuovo prima il nome e poi il cognome, e così via
#
# Un programma che potremo voler fare potrebbe essere leggere le linee e stampare a video nomi e cognomi così:
#
# ```
# <NAME>
# <NAME>
# <NAME>
# ```
#
# Per iniziare ad avere un'approssimazione del risultato finale, possiamo aprire il file, leggere solo la prima linea e stamparla:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="-XsMLCyU5bPH" outputId="824ecd67-35e8-4b79-bde5-ddf7f70334f4"
with open('people-simple.txt', encoding='utf-8') as f:
linea=f.readline()
print(linea)
# + [markdown] colab_type="text" id="C0eSCHzw5bPZ"
# Che è successo? Esaminiamo le varie linee:
#
# **il comando open**
#
# Il comando
#
# ```python
# open('people-simple.txt', encoding='utf-8')
# ```
#
# ci permette di aprire il file di testo dicendo a Python il percorso del file `'people-simple.txt'` e la codifica con cui è stato scritto (`encoding='utf-8'`).
#
# **La codifica**
#
# La codifica dipende dal sistema operativo e dell'editor con cui è stato scritto il file. Quando apriamo un file, Python non può divinare la codifica, e se non gliela specifichiamo potrebbe aprirlo assumendo una codifica diversa dall'originale - tradotto, affidandoci al caso o sbagliando codifica in seguito potremmo vedere dei caratteri strani (tipo quadratini invece di lettere accentate).
#
# In genere, quando apri un file, prova prima a specificare la codifica `utf-8` che è la più comune scrivendo `encoding='utf-8'`, e se per caso non va bene prova invece `encoding='latin-1'` (solitamente utile se il file è stato scritto su sistemi Windows). Se apri file scritti in posti più esotici, tipo in Cina, potresti dover usare un'altro encoding. Per approfondire queste questioni, quando hai tempo leggi [Immersione in Python - Cap 4 - Stringhe](http://gpiancastelli.altervista.org/dip3-it/stringhe.html) e [Immersione in Python - Cap 11 - File](http://gpiancastelli.altervista.org/dip3-it/stringhe.html), **entrambe letture caldamente consigliate**.
#
# **il with**
#
# Il `with` definisce un blocco con all'interno le istruzioni:
#
# ```python
# with open('people-simple.txt', encoding='utf-8') as f:
# linea=f.readline()
# print(linea)
# ```
#
# Abbiamo usato il `with` per dire a Python che in ogni caso, anche se accadono errori, vogliamo che dopo aver usato il file, e cioè eseguito le istruzioni nel blocco interno (il `linea=f.readline()` e `print(linea)`) Python deve chiudere automaticamente il file. Chiudere propriamente un file evita di sprecare risorse di memoria e creare errori paranormali. Se vuoi evitare di andare a caccia di file zombie mai chiusi, ricordati sempre di aprire i file nei blocchi `with`! Inoltre, alla fine della riga nella parte `as f:` abbiamo assegnato il file ad una variabile chiamata qui `f`, ma potevamo usare un qualunque altro nome.
#
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Per indentare il codice, usa SEMPRE sequenze di 4 spazi bianchi. Sequenze di 2 soli spazi per quanto consentite non sono raccomandate.
# </div>
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: A seconda dell'editor che usi, premendo TAB potresti ottenere una sequenza di spazi bianchi come accade in Jupyter (4 spazi che sono raccomandati), oppure un carattere speciale di tabulazione (da evitare)! Per quanto noiosa questa distinzione ti possa apparire, ricordatela perchè potrebbe generare errori molto difficili da scoprire.
# </div>
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Nei comandi che creano blocchi come il `with`, ricordati di mettere sempre il carattere dei doppi punti `:` alla fine della linea !
# </div>
#
# + [markdown] colab_type="text" id="en84Abg_5bPb"
# Il comando
#
# ```
# linea=f.readline()
# ```
# mette nella variabile `linea` l'intera linea, come una stringa. Attenzione: la stringa conterrà alla fine anche il carattere speciale di ritorno a capo !
#
# Ti chiederai da dove venga fuori quel `readline`. Come quasi tutto in Python, la nostra variabile `f` che rappresenta il file appena aperto è un oggetto, e ogni oggetto, a seconda del suo tipo, ha dei _metodi_ particolari che possiamo usare su di esso. In questo caso il metodo è `readline`.
#
# Il comando seguente stampa il contenuto della stringa:
#
# ```python
# print(linea)
# ```
#
# + [markdown] colab_type="text" id="wzyIGFfP5bPd"
# **✪ 1.1 ESERCIZIO**: Prova a riscrivere nella cella qua il blocco with appena visto, ed esegui la cella premendo Control-Invio. Riscrivi il codice con le dita, non con il copia e incolla ! Fai attenzione ad indentare correttamente con gli spazi il blocco.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Z26Xnnfm5bPg"
# scrivi qui
with open('people-simple.txt', encoding='utf-8') as f:
linea=f.readline()
print(linea)
# + [markdown] colab_type="text" id="0zsZD7-N5bPo"
# **✪ 1.2 ESERCIZIO**: immagino ti starai chiedendo che cosa è esattamente quella `f`, e cosa faccia esattamente il metodo `readline`. Quando ti trovi in queste situazioni, puoi aiutarti con le funzioni `type` e `help`. Questa volta, copia e incolla direttamente sempre lo stesso codice qua sotto, ma aggiungi a mano dentro il blocco `with` i comandi:
#
# * `print(type(f))`
# * `help(f.readline) # Attenzione: ricordati il 'f.' prima del readline !!`
# * `help(f)`
#
# Ogni volta che aggiungi qualcosa, prova ad eseguire con Control+Invio e vedere cosa succede.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="B5vALRz55bPq"
# scrivi qui il codice (copia e incolla)
with open('people-simple.txt', encoding='utf-8') as f:
linea=f.readline()
print(linea)
print('type(f):', type(f))
print()
help(f.readline)
#help(f) # lunga ...
# + [markdown] colab_type="text" id="I53T_-LV5bPx"
# Prima abbiamo messo il contenuto della prima linea nella variabile `line`, ora potremmo metterlo in una variabile dal nome più significativo, come `nome`. Non solo, possiamo anche direttamente leggere la linea successiva nella variabile `cognome` e poi stampare la concatenazione delle due:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="nK3hZrCq5bP0" outputId="49e2a028-ba09-4d96-be97-43a08ef3686b"
with open('people-simple.txt', encoding='utf-8') as f:
nome=f.readline()
cognome=f.readline()
print(nome + ' ' + cognome)
# + [markdown] colab_type="text" id="7UwUBQnc5bP-"
# **PROBLEMA !** La stampa mette comunque uno strano ritorno a capo. Come mai? Se ti ricordi, prima ho detto che `readline` legge il contenuto della linea in una stringa aggiungendo alla fine anche il carattere speciale di ritorno a capo. Per eliminarlo, puoi usare il comando `rstrip()`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="cNvkN5BD5bQB" outputId="d9dd006e-6bd9-4671-c98c-3af68cce2dc7"
with open('people-simple.txt', encoding='utf-8') as f:
nome=f.readline().rstrip()
cognome=f.readline().rstrip()
print(nome + ' ' + cognome)
# + [markdown] colab_type="text" id="1XWpXow25bQM"
# **✪ 1.3 ESERCIZIO**: Di nuovo, riscrivi il blocco qua sopra nella cella sotto, ed esegui la cella con Control+Invio. Domanda: che succede se usi `strip()` invece di `rstrip()`? Ed `lstrip()`? Riesci a dedurre il significato di `r` e `l` ? Se non ci riesci, prova ad usare il comando python `help` chiamando `help(string.rstrip)`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="cMnM_Ywm5bQP"
# scrivi qui
with open('people-simple.txt', encoding='utf-8') as f:
nome=f.readline().rstrip()
cognome=f.readline().rstrip()
print(nome + ' ' + cognome)
# + [markdown] colab_type="text" id="O7vruQTu5bQb"
# Benissimo, abbiamo la prima linea ! Adesso possiamo leggere tutte le linee in sequenza. A tal fine possiamo usare un ciclo `while`:
#
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="dXZBiWN-5bQh" outputId="607b53ee-8c4d-48cc-dc06-c8d6c0996223"
with open('people-simple.txt', encoding='utf-8') as f:
linea=f.readline()
while linea != "":
nome = linea.rstrip()
cognome=f.readline().rstrip()
print(nome + ' ' + cognome)
linea=f.readline()
# -
# <div class="alert alert-info">
#
# **NOTA:** In Python ci sono [metodi più concisi](https://thispointer.com/5-different-ways-to-read-a-file-line-by-line-in-python/) per leggere un file tdi testo linea per linea, ma abbiamo usato questo approccio per esplicitare tutti i passaggi
# </div>
# + [markdown] colab_type="text" id="Gs1Fthi-5bQr"
# Cosa abbiamo fatto? Per prima cosa, abbiamo aggiunto un ciclo `while` in un nuovo blocco
#
# <div class="alert alert-warning">
#
# **Attenzione**: nel nuovo blocco while, dato che è già all'interno del blocco esterno `with`, le istruzioni sono indentate di 8 spazi e non più 4! Se per caso sbagli gli spazi, possono succedere brutti guai!
# </div>
#
# Prima leggiamo una linea, e due casi sono possibili:
#
# a. siamo alla fine del file (o il file è vuoto): in questo caso la chiamata a `readline()` ritorna una stringa vuota
#
# b. non siamo alla fine del file: la prima linea è messa come una stringa dentro la variabile `linea`. Dato che Python internamente usa un puntatore per tenere traccia della posizione a cui si è durante la lettura del file, dopo una lettura questo puntatore è mosso all'inizio della riga successiva. In questo modo una chiamata successiva a `readline()` leggerà la linea dalla nuova posizione.
#
#
# Nel blocco `while` diciamo a Python di continuare il ciclo fintanto che `linea` _non_ è vuota. In questo caso, dentro il blocco `while` estraiamo il nome dalla linea e lo mettiamo nella variabile `nome` (rimuovendo il carattere extra di ritorno a capo con la `rstrip()` come fatto in precedenza), e poi procediamo leggendo la nuova riga ed estraendo il risultato dentro la variabile `cognome`. Infine, leggiamo di nuovo la linea dentro la variabile `linea` così sarà pronta per la prossima iterazione di estrazione nome. Se la `linea` è vuota il ciclo terminerà:
#
#
# ```python
# while linea != "": # entra il ciclo se la linea contiene caratteri
# nome = linea.rstrip() # estrae il nome
# cognome=f.readline().rstrip() # legge la nuova linea ed estrae il cognome
# print(nome + ' ' + cognome)
# linea=f.readline() # legge la prossima linea
# ```
#
# + [markdown] colab_type="text" id="z2RxmnlK5bQu"
# **✪ 1.4 ESERCIZIO**: Di nuovo come prima, riscrivi nella cella qua sotto il codice col while appena spiegato, facendo MOLTA attenzione all'indentazione (per la linea del with esterno fai pure copia e incolla):
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="rElpI-gY5bQw"
# scrivi qui il codice col while interno
with open('people-simple.txt', encoding='utf-8') as f:
linea=f.readline()
while linea != "":
nome = linea.rstrip()
cognome=f.readline().rstrip()
print(nome + ' ' + cognome)
linea=f.readline()
# + [markdown] colab_type="text" id="6pP7nWFc5bQ4"
# ## File a linee people-complex:
#
# Guarda il file `people-complex.txt`, più complesso:
# ```
# nome: Leonardo
# cognome: <NAME>
# data di nascita: 1452-04-15
# nome: Sandro
# cognome: Botticelli
# data di nascita: 1445-03-01
# nome: Niccolò
# cognome: Macchiavelli
# data di nascita: 1469-05-03
# ```
#
# Supponendo di leggere il file per voler stampare questo output, come faresti ?
#
# ```
# <NAME>, 1452-04-15
# <NAME>, 1445-03-01
# <NAME>, 1469-05-03
# ```
# + [markdown] colab_type="text" id="c76qOu-25bQ7"
#
# **Suggerimento 1**: Per ottenere dalla stringa `'abcde'` la sottostringa `'cde'`, che inizia all'indice 2, puoi usare l'operatore di parentesi quadre, indicando l'indice di inizio seguito dai doppi punti `:`
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="wbGZhl2w5bQ-" outputId="d42be766-a32c-44c6-c44f-b29b34817cd8"
x = 'abcde'
x[2:]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="Nr3E0n-d5bRI" outputId="33bdbff3-051a-4c6f-86b0-209fa7706cd3"
x[3:]
# + [markdown] colab_type="text" id="HlDQzVWG5bRS"
# **Suggerimento 2**: Per sapere la lunghezza di una stringa, usa la funzione `len`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="G5-AXund5bRT" outputId="2bc3c444-04d0-4053-d3c0-60fe33ed6099"
len('abcde')
# + [markdown] colab_type="text" id="KVA0FPut5bRe"
# **✪ 1.5 ESERCIZIO**: Scrivi qua sotto la soluzione dell'esercizio 'People complex':
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="XlS5VSI05bRg"
# scrivi qui
with open('people-complex.txt', encoding='utf-8') as f:
linea=f.readline()
while linea != "":
nome = linea.rstrip()[len("nome: "):]
cognome= f.readline().rstrip()[len("cognome: "):]
nato = f.readline().rstrip()[len("data di nascita: "):]
print(nome + ' ' + cognome + ', ' + nato)
linea=f.readline()
# + [markdown] colab_type="text" id="Q_F0X5jm5bRo"
# ## Esercizio file a linee immersione-in-python-toc
#
# Questo esercizio è più difficile, se sei alle prime armi puoi saltarlo e passare ai CSV.
#
# Il libro Immersione in Python in Italiano è bello e ti fornisce un PDF, che però ha un problema: se provi a stamparlo scoprirai che manca l'indice. Senza perderci d'animo, abbiamo trovato un programmino che ci ha estratto i titoli in un file come segue, che però come scoprirai non è esattamente bello da vedere. Dato che siamo Python ninja, abbiamo deciso di trasformare i titoli grezzi in una [tabella dei contenuti vera e propria](http://it.softpython.org/_static/toc-immersione-in-python-3.txt). Sicuramente ci sono metodi più furbi per compiere questa operazione, come caricare il pdf in Python con una apposita libreria per i pdf, ma pareva un esercizio interessante.
#
# Ti viene consegnato il file immersione-in-python-toc.txt:
#
# ```
# BookmarkBegin
# BookmarkTitle: Il vostro primo programma Python
# BookmarkLevel: 1
# BookmarkPageNumber: 38
# BookmarkBegin
# BookmarkTitle: Immersione!
# BookmarkLevel: 2
# BookmarkPageNumber: 38
# BookmarkBegin
# BookmarkTitle: Dichiarare funzioni
# BookmarkLevel: 2
# BookmarkPageNumber: 41
# BookmarkBeginint
# BookmarkTitle: Argomenti opzionali e con nome
# BookmarkLevel: 3
# BookmarkPageNumber: 42
# BookmarkBegin
# BookmarkTitle: Scrivere codice leggibile
# BookmarkLevel: 2
# BookmarkPageNumber: 44
# BookmarkBegin
# BookmarkTitle: Stringhe di documentazione
# BookmarkLevel: 3
# BookmarkPageNumber: 44
# BookmarkBegin
# BookmarkTitle: Il percorso di ricerca di import
# BookmarkLevel: 2
# BookmarkPageNumber: 46
# BookmarkBegin
# BookmarkTitle: Ogni cosa è un oggetto
# BookmarkLevel: 2
# BookmarkPageNumber: 47
# ```
#
# Scrivi un programma python per stampare e video il seguente output:
#
# ```
# Il vostro primo programma Python 38
# Immersione! 38
# Dichiarare funzioni 41
# Argomenti opzionali e con nome 42
# Scrivere codice leggibile 44
# Stringhe di documentazione 44
# Il percorso di ricerca di import 46
# Ogni cosa è un oggetto 47
# ```
#
# Per questo esercizio, dovrai inserire nell'output degli spazi artificiali, in una quantità determinata dalle righe `BookmarkLevel`.
#
# **DOMANDA**: che cos'è lo strano valore `è` alla fine del file originale ? Dobbiamo riportarlo nell'output ?
#
# **SUGGERIMENTO 1**: Per convertire una stringa in numero intero, usa la funzione `int`:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="XWlxnhpX5bRy"
x = '5'
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="GdJd09mZ5bR_" outputId="dc19c940-e1af-4c84-aa60-012cfb985133"
x
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="nETKnztF5bSF" outputId="118522fe-186c-452e-9218-223508605154"
int(x)
# + [markdown] colab_type="text" id="wwfLN6lv5bSQ"
#
# <div class="alert alert-warning">
#
# **Attenzione**: `int(x)` ritorna un valore, e non modifica mai l'argomento `x` !
# </div>
#
# + [markdown] colab_type="text" id="w06zsF1a5bSS"
# **SUGGERIMENTO 2**: Per sostituire un sottostringa in una stringa, si può usare la funzione replace:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="sGPUddlf5bST" outputId="4af45183-e44d-4fad-a1ba-16c6722c62ce"
x = 'abcde'
x.replace('cd', 'HELLO' )
# + [markdown] colab_type="text" id="7OBEbHya5bSd"
# **SUGGERIMENTO 4**: Finchè c'è una sola sequenza da sostituire, va bene il replace, ma se avessimo un milione di sequenze orribili come `>`, `>`, `&x3e;`, che faremmo? Da bravi data cleaner, possiamo riconoscere che sono [sequenze di escape HTML](https://corsidia.com/materia/web-design/caratterispecialihtml), perciò potremmo usare dei metodi specifici per queste sequenze come [html.unescape](https://docs.python.org/3/library/html.html#html.unescape). Provalo invece del replace e guarda se funziona!
#
# NOTA: Prima di usare il metodo `html.unescape`, importa il modulo `html` con il comando:
#
# ```python
# import html
# ```
# + [markdown] colab_type="text" id="m2scPeP15bSe"
# **SUGGERIMENTO 3**: Per scrivere _n_ copie di un carattere, usa `*` come qua:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="0mOU41SC5bSf" outputId="74391ce6-fef1-4d9e-a630-804af033f3fd"
"b" * 3
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="NK5lFuks5bSm" outputId="56bed93f-878f-4d92-88fc-2afef788218c"
"b" * 7
# + [markdown] colab_type="text" id="58yyJFaf5bSu"
# **✪✪✪ 1.6 ESERCIZIO**: Scrivi qua sotto la soluzione per file a linee immersione-in-python-toc, e prova ad eseguirla premendo Control + Invio:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ndZarkM45bSv"
# scrivi qui
import html
with open("immersione-in-python-toc.txt", encoding='utf-8') as f:
linea=f.readline()
while linea != "":
linea = f.readline().strip()
titolo = html.unescape(linea[len("BookmarkTitle: "):])
linea=f.readline().strip()
livello = int(linea[len("BookmarkLevel: "):])
linea=f.readline().strip()
pagina = linea[len("BookmarkPageNumber: "):]
print((" " * livello) + titolo + " " + pagina)
linea=f.readline()
# -
# ## Prosegui
#
# Continua con i [file tabellari CSV](https://it.softpython.org/formats/formats2-csv-sol.html)
| formats/formats1-lines-sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: right;" src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAOIAAAAjCAYAAACJpNbGAAAABHNCSVQ<KEY>zAAALEgAACxIB0t1+/AAAABR0RVh0Q3JlYXRpb24gVGltZQAzLzcvMTNND4u/AAAAHHRFWHRTb2Z0d2FyZQBBZG9iZSBGaXJld29ya3MgQ1M26LyyjAAACMFJREFUeJztnD1y20gWgD+6nJtzAsPhRqKL3AwqwQdYDpXDZfoEppNNTaWbmD7BUEXmI3EPMFCR2YI1UDQpdAPqBNzgvRZA/BGUZEnk9FeFIgj0z2ugX7/XP+jGer2mLv/8b6d+4Efgf/8KG0+Zn8XyXLx+bgEslqegcfzxSY3Irrx6bgEsFssBWsRGowGufwHAYtq7u+H6fUCOxTTWax4wBAbr+SRqNDKesOv3gN/133sW0yh927j1mucIaFWINl7PJ+OcvMcfW8Bol3iN44+mLIOsTCp3UJFfAETr+WRQcG8EOJpunEnTyDlYzycbeWr5xxq3jOF6PglK8ix9buv5xCsrAzBkMV1l5OwD/aJ4BXzV3+8F9z4gz/hTSbz8cxc84FuNvDc4VIsYA7+qohmGwAnycA194G22YqUYlZxv4vpN4AuwBv4oON5m8k3TVLnK4sYFcRyN86dWvCwnlCvFCeUVvwX8CkSZZ5eWs5mLJWE/VZThBMgpfirPk5J4f1SU4QsQ6LNP4+j9OkSUKdRiGlD87CWe3PcyR5PFdAhc1cz/joOziMoIeVF95GX1EGVY6bWhvsAeZQrm+kON80PDneD6PRbTi4LQpmJfsZieFaR1qXlXURh3y2BaBPyG63sspv0t6e+CKJTrf2YxHe8Qr6z8AXBdGbMoHgCTshgr4AiItfxljenPJGv5roCi+rGVw1TExTTWl99ThRsglfYHUnF7SMv+Bhjn4idxbhFLGiAu6gjXD3LuUBF5VzWi3CoAfMP1kxe7mNYZMT5DLFgf13eAXi3ZtvMOsUb3V3J5/mmqy+/66RbnTC1LFdfIu/kd8Qx2bTQeg2GBTPfiUF1TgHNE0QaIq/JDX9RKr/WBy/V8EhfEHWncWMO2EKV8S7UypYnYdE2r+o8gyj5MHXVYsZh+JnG7A+3LPQxR5g9II/UJ148ockmrybqm2+Qapo6gppwB8J7EM6jqaz8u0lhfkXgB58BKPam6rvEdh2kRARbTMa7/HXEfVqnW8hxxWwE+5+JJRTYd9CM90gxw/XFuMKMo/yTNDzUkLnbr6rCYnuH6N8igQ3CvNPJproDPuH6MKMd4Z5kMUjnrh98tn1if72/Ie729Vzq708L0YV3/HGmgB4iHsjOProhhd1lrEr4zaz/FvM4lolTnqWum/6jKmeuDmFb1jHylNg96hPQbhcU0wPVBXESvQI4W5aNshsK4jeOPhSOcOaThMVb48dhU8m2UlR+29ZHzrqyhLL0EaTROteGt67EYIsT6F1HXC/ikcvS00dl51PRwLaIwQtzCxGWRFnRMkT8v/SyAy8I+iliHJtDUsHHq7imipE42GtJanxdcB6mgQcm9MmKNs1m5F9MI13+n+cXZSEpAeV8mQgZqNkmU/HsuT7kf4PrGhXcK0h1SXv7iPKsJKCrDYvoV17+meMqhiDFlll7GEb4U3iseAf+k7mqksmU9qUoaj73E7TEtol3iZnks7Moai8WylUN3TS0WANbzyYv2rqxFtFheANYi7iGNRoPOrO2QGTQIu8vhU8vSmbWNDAHQD7vLYWfWbgFx2F3ee3FBZ9ZuIgMpTWAQdpeRXm9pPoPOrD3UMCtkQM4BRmF3ubG6ZZdxkOfCWsT9pU96CuX56KfOjeIFVC8Ar8NI0xuyOQJsVkWl8xzptQGPNY/6xFiLuL+0gIu0FVTrNESmbK7C7tLrzNpmPW0EeGF32UyFN19UnCAT4ZHGWWnYqDNrB4jViZBK/kbD9sLuMiBZSD8AVp1Z+0LD/NmZta+BIzOS3pm1xwBhd9kvkeEGUbQeqSmIdHhkXnGs5fIQRUxPV1x0Zm2zMuoq7C69rU/yBWAt4v7iAd86s/ZaDweZP+wBvwBOZ9b2SCrrmPzk+AWizA09j1QxMK4gZumcWKUWMvkdA56mfxN2l7GmHWk6V2F32Qi7yxaIsmnYHvkJ9zEQqAwBotQXwK2m0c+EN/Kk8zPTZiOkIWrp/xNTnpeOtYh7iFauN+k5W+0vXab6UsbyecAw229SxWiG3aVZ7NBCKrGHuneazy2iyBeIuxkjk9UDE1bzOtJ4IzbdwysNN0D6dnf9Rk3/iKSBWOnhUbASSWW+DbvLWM+HKreZ3O/r77gza5u842w6LxFrEfcTj+Jv3mK4q7Co63hE+fI6E94hUaT0cry+XushSuvoNZO2CdsCrlXJHDYVMUIUJso2BmhfL+wuV6rMvVR6AXnS1428XupaE7Hwnrqkg4cMGD0lr3NfpVegrUw1m2sN0+crNirEX1uTqiPbPoyI/QSKKmqA9I9aer+fcR2zxIj7GiMV+EYVIkZc3r5eH2rYI+0vnpBYIE/vGwUCdYM7s3agbqXJu58VIOwug86sfd2ZtSPNKwi7S9PHy4UnscCmXKuUZQRdsqbPwCHp2754pKYnW0akcZBO/x2df29XnvA//6iV8T3TSluBmOQlR+v5JNvaHixlDZRalRZifbZaAg3vIIrkmP6YVu6owI1M9x2r0vVIFCBGXNLS96Ph45IGY2ey6e1DY20UMaLGItUXoIhVvCv5tvDg2MWLqYNaoKBKWe6Z7gBR8OwAzZOyD4poBmtidlwt/gIxw/QHz0+oWKIoj19fRz8p3YOjoV8195F5l31ltZ5PfnluISyW+/IK6SPstRIiH/FaLHvLa2R+6F6f978AVsD7v0vf0HK4vNK9VfbVojSBceP4o/PcglgsD8GMmjaRbRCc1PEQIrbv45nlIfleIrs778XkrcWSZXMcXPZyqbvfxy7ckuyqHJPslJzH9c3We2ZRbx1O/07ziJbDI1FE2Qwp4n4DNzHJhkZF16+3bnwrCmi40U2eWoj7KZvobn7+YtKO1vPJVyyWPSZrER1kNU0TqfienpvlaWZR7oX+3tba6lxcX7MK3tNfo2RlpNc8tthsIFbAKYtpsA+TtRbLNp5/H4/EFXX0MOfbOGUxvbCKaDkEnl8Rq0jc1ayFjhFFjKwiWg6B/wNk+JCXXNBIXQAAAABJRU5ErkJggg==">
#
#
# # An Jupyter notebook for running PCSE/WOFOST on a CGMS8 database
#
# This Jupyter notebook will demonstrate how to connect and read data from a CGMS8 database for a single grid. Next the data will be used to run a PCSE/WOFOST simulation for potential and water-limited conditions, the latter is done for all soil types present in the selected grid. Results are visualized and exported to an Excel file.
#
# Note that no attempt is made to *write* data to a CGMS8 database as writing data to a CGMS database can be tricky and slow. In our experience it is better to first dump simulation results to a CSV file and use specialized loading tools for loading data into the database such as [SQLLoader](http://www.oracle.com/technetwork/database/enterprise-edition/sql-loader-overview-095816.html) for ORACLE or [pgloader](http://pgloader.io/) for PostgreSQL databases.
#
# A dedicated package is now available for running WOFOST simulations using a CGMS database: [pyCGMS](https://github.com/ajwdewit/pycgms). The steps demonstrated in this notebook are implemented in as well in the pyCGMS package which provides a nicer interface to run simulations using a CGMS database.
#
# **Prerequisites for running this notebook**
#
# Several packages need to be installed for running PCSE/WOFOST on a CGMS8 database:
#
# 1. PCSE and its dependencies. See the [PCSE user guide](http://pcse.readthedocs.io/en/stable/installing.html) for more information;
# 2. The database client software for the database that will be used, this depends on your database of choice. For SQLite no client software is needed as it is included with python. For Oracle you will need the [Oracle client software](http://www.oracle.com/technetwork/database/features/instant-client/index-097480.html) as well as the [python bindings for the Oracle client (cx_Oracle)](http://sourceforge.net/projects/cx-oracle/files/)). See [here](https://wiki.python.org/moin/DatabaseInterfaces) for an overview of database connectors for python;
# 3. The `pandas` module for processing and visualizing WOFOST output;
# 4. The `matplotlib` module, although we will mainly use it through pandas;
# ## Importing the relevant modules
#
# First the required modules need to be imported. These include the CGMS8 data providers for PCSE as well as other relevant modules.
# +
# %matplotlib inline
import os, sys
data_dir = os.path.join(os.getcwd(), "data")
import matplotlib as mpl
import matplotlib.pyplot as plt
plt.style.use('ggplot')
import sqlalchemy as sa
import pandas as pd
import pcse
from pcse.db.cgms8 import GridWeatherDataProvider, AgroManagementDataProvider, SoilDataIterator, \
CropDataProvider, STU_Suitability, SiteDataProvider
from pcse.models import Wofost71_WLP_FD, Wofost71_PP
from pcse.util import DummySoilDataProvider, WOFOST71SiteDataProvider
from pcse.base import ParameterProvider
print("This notebook was built with:")
print("python version: %s " % sys.version)
print("PCSE version: %s" % pcse.__version__)
# -
# ## Building the connection to a CGMS8 database
#
# The connection to the database will be made using SQLAlchemy. This requires a database URL to be provided, the format of this URL depends on the database of choice. See the SQLAlchemy documentation on [database URLs](http://docs.sqlalchemy.org/en/latest/core/engines.html#database-urls) for the different database URL formats.
#
# For this example we will use a database that was created for Anhui province in China. This database can be downloaded [here](https://drive.google.com/open?id=1nk-juuxzBOLoHDJl_s-YnGiHKIdSFc5f).
cgms8_db = "c:/data/nobackup/CGMS8_Anhui/CGMS_Anhui_complete.db"
if not os.path.exists(cgms8_db):
print("CGMS DB not found!")
else:
dbURL = "sqlite:///%s" % cgms8_db
engine = sa.create_engine(dbURL)
# ## Defining what should be simulated
#
# For the simulation to run, some IDs must be provided that refer to the location (`grid_no`), crop type (`crop_no`) and year (`campaign_year`) for which the simulation should be carried out. These IDs refer to columns in the CGMS database that are used to define the relationships.
grid_no = 81159
crop_no = 1 # Winter-wheat
campaign_year = 2008
# ## Retrieving data for the simulation from the database
#
# ### Weather data
#
# Weather data will be derived from the GRID_WEATHER table in the database. By default, the entire time-series of weather data available for this grid cell will be fetched from the database.
weatherdata = GridWeatherDataProvider(engine, grid_no)
print(weatherdata)
# ### Agromanagement information
#
# Agromanagement in CGMS mainly refers to the cropping calendar for the given crop and location.
agromanagement = AgroManagementDataProvider(engine, grid_no, crop_no, campaign_year)
agromanagement
# ### Soil information
#
# A CGMS grid cell can contain multiple soils which may or may not be suitable for a particular crop. Moreover, a complicating factor is the arrangement of soils in many soil maps which consist of *Soil Mapping Units* `(SMUs)` which are soil associations whose location on the map is known. Within an SMU, the actual soil types are known as *Soil Typological Units* `(STUs)` whose spatial delination is not known, only the percentage area within the SMU is known.
#
# Therefore, fetching soil information works in two steps:
# 1. First of all the `SoilDataIterator` will fetch all soil information for the given grid cell. It presents it as a list which contains all the SMUs that are present in the grid cell with their internal STU representation. The soil information is organized in such a way that the system can iterate over the different soils including information on soil physical properties as well as SMU area and STU percentage with the SMU.
# 2. Second, the `STU_Suitability` will contain all soils that are suitable for a given crop. The 'STU_NO' of each crop can be used to check if a particular STU is suitable for that crop.
#
# The example grid cell used here only contains a single SMU/STU combination.
soil_iterator = SoilDataIterator(engine, grid_no)
soil_iterator
suitable_stu = STU_Suitability(engine, crop_no)
# ### Crop parameters
#
# Crop parameters are needed for parameterizing the crop simulation model. The `CropDataProvider` will retrieve them from the database for the given crop_no, grid_no and campaign_year.
cropd = CropDataProvider(engine, grid_no, crop_no, campaign_year)
# ### Site parameters
#
# Site parameters are an ancillary class of parameters that are related to a given site. For example, an important parameter is the initial amount of moisture in the soil profile (WAV) and the Atmospheric CO$_2$ concentration (CO2). Site parameters will be fetched for each soil type within the soil iteration loop.
# ## Simulating with WOFOST
#
# ### Place holders for storing simulation results
daily_results = {}
summary_results = {}
# ### Potential production
# For potential production we can provide site data directly
sited = WOFOST71SiteDataProvider(CO2=360, WAV=25)
# We do not need soildata for potential production so we provide some dummy values here
soild = DummySoilDataProvider()
# Start WOFOST, run the simulation
parameters = ParameterProvider(sitedata=sited, soildata=soild, cropdata=cropd)
wofost = Wofost71_PP(parameters, weatherdata, agromanagement)
wofost.run_till_terminate()
# convert output to Pandas DataFrame and store it
daily_results['Potential'] = pd.DataFrame(wofost.get_output()).set_index("day")
summary_results['Potential'] = wofost.get_summary_output()
# ### Water-limited production
#
# Water-limited simulations will be carried out for each soil type. First we will check that the soil type is suitable. Next we will retrieve the site data and run the simulation. Finally, we will collect the output and store the results.
for smu_no, area, stu_no, percentage, soild in soil_iterator:
# Check if this is a suitable STU
if stu_no not in suitable_stu:
continue
# retrieve the site data for this soil type
sited = SiteDataProvider(engine, grid_no, crop_no, campaign_year, stu_no)
# Start WOFOST, run the simulation
parameters = ParameterProvider(sitedata=sited, soildata=soild, cropdata=cropd)
wofost = Wofost71_WLP_FD(parameters, weatherdata, agromanagement)
wofost.run_till_terminate()
# Store simulation results
runid = "smu_%s-stu_%s" % (smu_no, stu_no)
daily_results[runid] = pd.DataFrame(wofost.get_output()).set_index("day")
summary_results[runid] = wofost.get_summary_output()
# ## Visualizing and exporting simulation results
#
# ### We can visualize the simulation results using pandas and matplotlib
# Generate a figure with 10 subplots
fig, axes = plt.subplots(nrows=5, ncols=2, figsize=(12, 30))
# Plot results
for runid, results in daily_results.items():
for var, ax in zip(results, axes.flatten()):
results[var].plot(ax=ax, title=var, label=runid)
ax.set_title(var)
fig.autofmt_xdate()
r = axes[0][0].legend(loc='upper left')
# ### Exporting the simulation results
#
# A pandas DataFrame or panel can be easily export to a [variety of formats](http://pandas.pydata.org/pandas-docs/stable/io.html) including CSV, Excel or HDF5. First we convert the results to a Panel, next we will export to an Excel file.
excel_fname = os.path.join(data_dir, "output", "cgms8_wofost_results.xls")
with pd.ExcelWriter(excel_fname) as writer:
for name, df in daily_results.items():
df.to_excel(writer, sheet_name=name)
# ## Simulating with a different start date waterbalance
# By default CGMS starts the simulation when the crop is planted. Particularly in dry climates this can be problematic because the results become very sensitive to the initial value of the soil water balance. In such scenarios, it is more realistic to start the water balance with a dry soil profile well before the crop is planted and let the soil 'fill up' as a result of rainfall.
#
# To enable this option, the column `GIVEN_STARTDATE_WATBAL` in the table `INITIAL_SOIL_WATER` should be set to the right starting date for each grid_no, crop_no, year and stu_no. Moreover, the other parameters in the table should be set to the appropriate values (particularly the initial soil moisture `WAV`).
#
# The start date of the water balance should then be used to update the agromanagement data during the simulation loop, see the example below.
for smu_no, area, stu_no, percentage, soild in soil_iterator:
# Check if this is a suitable STU
if stu_no not in suitable_stu:
continue
# retrieve the site data for this soil type
sited = SiteDataProvider(engine, grid_no, crop_no, campaign_year, stu_no)
# update the campaign start date in the agromanagement data
agromanagement.set_campaign_start_date(sited.start_date_waterbalance)
# Start WOFOST, run the simulation
parameters = ParameterProvider(sitedata=sited, soildata=soild, cropdata=cropd)
wofost = Wofost71_WLP_FD(parameters, weatherdata, agromanagement)
wofost.run_till_terminate()
# Store simulation results
runid = "smu_%s-stu_%s" % (smu_no, stu_no)
daily_results[runid] = pd.DataFrame(wofost.get_output()).set_index("day")
summary_results[runid] = wofost.get_summary_output()
# ## Let's show the results
#
# As you can see, the results from the simulation are slightly different because of a different start date of the water balance.
#
# Generate a figure with 10 subplots
fig, axes = plt.subplots(nrows=5, ncols=2, figsize=(12, 30))
# Plot results
for runid, results in daily_results.items():
for var, ax in zip(results, axes.flatten()):
results[var].plot(ax=ax, title=var, label=runid)
fig.autofmt_xdate()
r = axes[0][0].legend(loc='upper left')
| 05 Using PCSE WOFOST with a CGMS8 database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: f45869f7d29be55bc11001e8f9741479e5cb8688820e4b86bb9403b134f57a38
# name: 'Python 3.7.6 64-bit (''base'': conda)'
# ---
# # Sesión 2 - Scientific computing in Python
# ### Instalar paquetes/librerias
#
# Los paquetes que vamos a usar en este curso ya los tenenemos instalados con Anaconda en el entorno `base`. Para ver que paquetes tenemos instalados en el entorno `base` podemos mirarlo por el entorno gráfico que nos proporciona Anaconda, o a traves de la linea de comandos con el siguiente comando:
#
# ```
# conda list
# ```
#
# Para instalar paquetes que no vengan ya de serie, de nuevo dos opciones, usar la interfaz gráfica o la linea de comandos con alguno de los siguientes comandos:
# ```
# conda install scikit-learn
# pip install scikit-learn
# ```
# ## Numpy
#
# * Numpy (numerical python) es uno de los paquetes más importantes de Python.
#
# * Muchos otros paquetes usan funcionalidades de este paquete de base. Por ese motivo, es importante conocer los conceptos básicos de Numpy.
#
# * Numpy tiene un tipo de estructura especifico denominado `ndarray` que hace referencia a un vector/matríz N-dimensional.
# Para importar el módulo Numpy a la sesión de Python en la que estamos trabajando:
import numpy as np
# Importante: Los import de los módulos se suelen hacer al principio en los script de python.
# Podemos ver todos los métodods asociados que tiene Numpy con la función `dir`:
dir(np)
# Defininimos un `ndarray` con la función `np.array`:
a = np.array([1, 2, 3])
a
type(a)
dir(a)
# Cada `ndarray` está asociado a un tipo de dato (float, float32, float64, int, ...) y todos los objetos que lo forman tienen que ser de ese mismo tipo. Podemos ver que tipo de dato esta asociado a un `ndarray` con `dtype`:
a.dtype
# Vamos a definir ahora una bi dimensional:
b = np.array([[1.3, 2.4],[0.3, 4.1]])
b
b.dtype
b.shape
b.ndim
b.size
# Podemos definir `ndarray`s con más tipos de elementos:
c = np.array([['a', 'b'],['c', 'd']])
c
d = np.array([[1, 2, 3],[4, 5, 6]], dtype=complex)
d
# ### Diferentes tipos de funciones para crear `ndarrays`:
np.zeros((3, 3))
np.ones((3, 3))
np.arange(0, 10)
np.arange(4, 10)
np.arange(0, 12, 3)
np.arange(0, 6, 0.6)
np.linspace(0, 10, 5)
np.random.random(3)
np.random.random((3, 3))
# La función `reshape`:
np.arange(0,12).reshape(3, 4)
# ### Operaciones aritméticas
a = np.arange(4)
a
a+4
a*2
b = np.arange(4, 8)
b
a + b
a - b
a * b
A = np.arange(0, 9).reshape(3, 3)
A
B = np.ones((3, 3))
B
A * B
# ### Multiplicación de matrices
#
# Hasta ahora, solo hemos hecho operaciones por elementos de los `ndarray`s. Ahora vamos a ver otro tipo de operaciones, como la multiplicación de matrices.
np.dot(A, B)
A.dot(B)
np.dot(B, A)
# ### Más funciones para las `ndarrays`
a = np.arange(1, 10)
a
np.sqrt(a)
np.log(a)
np.sin(a)
a.sum()
a.min()
a.max()
a.mean()
a.std()
# ### Manipular vectores y matrices
a = np.arange(1, 10)
a
a[4]
a[-4]
a[:5]
a[[1,2,8]]
A = np.arange(1, 10).reshape((3, 3))
A
A[1, 2]
A[:,0]
A[0:2, 0:2]
A[[0,2], 0:2]
# ### Iterando un array
a
for i in a:
print(i)
A
for row in A:
print(row)
for i in A.flat:
print(i)
# Si necesitamos aplicar una función en las columnas o filas de una matriz, hay una forma más elegante y eficaz de hacerlo que usando un `for`.
np.apply_along_axis(np.mean, axis=0, arr=A)
np.apply_along_axis(np.mean, axis=1, arr=A)
# ### Condiciones y arrays de Booleanos
A = np.random.random((4, 4))
A
A < 0.5
A[A < 0.5]
# ### Unir arrays
A = np.ones((3, 3))
B = np.zeros((3, 3))
np.vstack((A, B))
np.hstack((A, B))
# Funciones más especificas para unir arrays de una sola dimensión y crear así arrays bidimensionales:
a = np.array([0, 1, 2])
b = np.array([3, 4, 5])
c = np.array([6, 7, 8])
np.column_stack((a, b, c))
np.row_stack((a, b, c))
# ### Importante: Copía o vista de un elemento `ndarray`
a = np.array([1, 2, 3, 4])
a
b = a
b
a[2] = 0
a
b
c = a[0:2]
c
a[0] = 0
c
# Evitamos esto usando la funcion `copy`.
a = np.array([1, 2, 3, 4])
b = a.copy()
b
a[2] = 0
b
c = a[0:2].copy()
c
a[0] = 0
c
# ## Pandas
#
# * Pandas es el paquete de referencia para el análisis de datos en Python.
#
# * Pandas proporciona estructuras de datos complejas y funciones especificas para trabajar con ellas.
#
# * El concenpto fundamental de Pandas son los `DataFrame`, una estructura de datos con dos dimensiones. También están las `Series`, que son de una dimensión.
#
# * Pandas usa Numpy
import numpy as np
import pandas as pd
# ### DataFrame
#
# Un DataFrame es basicamente una tabla. Esta formado por filas y columnas, que son arrays con valores individuales (pueden ser números o no).
pd.DataFrame({'Yes': [50, 21], 'No': [131, 2]})
pd.DataFrame({'Juan': ['Sopa', 'Pescado'], 'Ana': ['Pasta', 'Solomillo']})
# Estamos usando `pd.DataFrame()` para construir objetos `DataFrame`. Como argumento le pasamos un diccionario con los `keys` `['Juan', 'Ana']` y sus respectivos valores. Aunque este es el método más común para construir un objeto `DataFrame`, no es el único.
# El método para construir `DataFrames` que hemos usado le asigna una etiqueta a cada columna que va desde el 0 hasta el número de columnas ascendentemente. Algunas veces esto está bien, pero otras veces puede que queramos asignar una etiqueta específica a cada columna.
pd.DataFrame({'Juan': ['Sopa', 'Pescado', 'Yogurt'], 'Ana': ['Pasta', 'Solomillo', 'Fruta']}, index=['1 Plato', '2 Plato', 'Postre'])
# ### Series
#
# Las `Series` son una sequencia de datos. Si los `DataFrames` son tablas de datos, las `Series` son listas de datos.
pd.Series([1, 2, 3, 4, 5])
pd.Series([30, 35, 40], index=['2015 matriculas', '2016 matriculas', '2017 matriculas'], name='Matriculas en máster de modelización')
# Las `Series` y los `DataFrames` están estrechamente relacionados. De hecho, podemos pensas que los `DataFrames` son simplemente un puñado de `Series` juntados.
# ### Leer ficheros de datos
#
# Aunque exista la opción de crear los `DataFrames` y las `Series` a mano, lo más habitual va a ser que trabajemos con datos que ya existen y están recogidos en algún tipo de fichero (.xls, .csv, .json, ...)
#
# El formato más habitual para guardar datos el el CSV. Los ficheros CSV contienen valores separados por comas.
reviews = pd.read_csv("data/winemag-data-130k-v2.csv")
reviews.shape
reviews.head()
reviews = pd.read_csv("data/winemag-data-130k-v2.csv", index_col=0)
reviews.head()
# ### Seleccionar subconjuntos del `DataFrame` o `Series`
# Podemos seleccionar los valores de una o varias columnas de varias maneras.
reviews.country
reviews['country']
reviews[['country', 'province']]
reviews[['country', 'province']][:5]
# También podemos usar los indices para seleccionar los subconjuntos usando el método `iloc`.
reviews.iloc[0]
reviews.iloc[0,0]
reviews.iloc[:,-1]
reviews.iloc[-3:, :3]
reviews.iloc[[0, 10, 100], 0]
# Por último, tambien podemos usar el método `loc` para usar las etiquetas de las filas y columnas.
reviews.loc[:, ['taster_name', 'taster_twitter_handle', 'points']]
# CUIDADO! En este caso, las etiquetas de las filas son números, pero `iloc` y `loc` no funciónan igual.
reviews.iloc[:5, 0]
reviews.loc[:5, 'country']
# ### Manipular el índice
reviews.set_index("title")
# ### Selección condicional
# Podemos buscar los vinos de Italia.
reviews['country'] == 'Italy'
# La anterior expresión nos ha devuelto una `Series` con los booleanos que nos dicen cuando el vino es Italiano. Para encontrar esas instancias devueltas por los booleanos hacemos:
reviews.loc[reviews['country'] == 'Italy']
# Si además de que sea Italiano, también queremos que nuestro vino tenga una puntuación mayor o igual a 90:
reviews.loc[(reviews['country'] == 'Italy') & (reviews['points'] >= 90)]
# Si queremos un vino Italiano o con puntuación mayor o igual a 90:
reviews.loc[(reviews['country'] == 'Italy') | (reviews['points'] >= 90)]
# Si queremos un vino Italiano o Español:
reviews.loc[reviews['country'].isin(['Italy', 'Spain'])]
# Si queremos deshacernos de las instancias en las que no tenemos el valor del precio:
reviews.loc[reviews['price'].notnull()]
# ### Añadir datos
#
# Añadir datos a nuestros `DataFrames` es fácil. Por ejemplo, podemos asignar el mismo valor a todas las instancias con el siguiente comando:
reviews['critic'] = 'everyone'
reviews['critic']
reviews['index_backwards'] = range(len(reviews), 0, -1)
reviews['index_backwards']
# ### Describir nuestro dataset
#
# Pandas nos proporciona herramientas para facilmente conocer un poco por encima como es el dataset con el que estamos trabajando a traves de valores estadísticos.
reviews.describe()
reviews.describe(include='all')
reviews.dtypes
reviews['points'].mean()
reviews['points'].quantile(0.25)
reviews['country'].unique()
reviews['country'].value_counts()
# ### Modificar los valores de una columna
#
# Por ejemplo, vamos a normalizar los datos de la columna points.
(reviews['points'] - reviews['points'].mean()) / reviews['points'].std()
reviews['province - region'] = reviews['province'] + ' - ' + reviews['region_1']
reviews
# ### Eliminar columnas
reviews.columns
reviews.pop('province - region')
reviews = reviews.drop(columns=['critic', 'index_backwards'])
reviews.columns
# ### Agrupar datos
reviews.groupby('points')['points'].count()
# Lo que ha ocurrido es que la función `groupby()` ha creado diferentes grupos dependiendo de la puntuación y luego a contado cuantos vinos hay en cada grupo.
#
# Ahora, vamos a calcular el precio medio de los vinos dependiendo la puntuación:
reviews.groupby('points')['price'].mean()
# Podemos agrupar usando más de un criterio y devolver más de un valor con `agg()`.
reviews.groupby(['price', 'country']).agg(['count', 'min', 'mean', 'max'])
# ### Ordenar instancias
reviews.sort_values(by='points')
reviews.sort_values(by='points', ascending=False).reset_index()
# ### Missing data
#
# Tratar con los datos que faltan es muy importante. Pandas nos ofrece funciones como `isnull, notnull y fillna` para localizar y rellenar los valores perdidos.
reviews['country'].isnull()
reviews[reviews['country'].isnull()]
reviews['region_2'].fillna("Unknown")
reviews['price'][reviews['price'].isnull()]
reviews['price'].fillna(method='bfill')
reviews['region_2'].fillna("Unknown")
# ### Sustituir valores
reviews['taster_twitter_handle'].replace("@kerinokeefe", "@kerino")
# ### Renombrar
#
# Podemos renombrar los nombres de los indices o columnas con la función `rename`.
reviews.rename(columns={'points': 'score'})
reviews.rename(index={0: 'firstEntry', 1: 'secondEntry'})
# ### Combinar datasets
df1 = reviews.iloc[:50000, :]
df2 = reviews.iloc[50000:, :]
df2.shape
pd.concat([df1, df2])
dfl = reviews.iloc[:, :8]
dfr = reviews.iloc[:, 8:]
dfr.shape
dfl.join(dfr)
| content/course/Sesion02/clase02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fake News Classication
# # Introduction
# - Fake news is false or misleading information presented as news. It often has the aim of damaging the reputation of a person or entity, or making money through advertising revenue.
# - The prevalence of fake news has increased with the rise of social media, especially the Facebook News Feed. Political polarization, post-truth politics, confirmation bias and social media algorithms have been implicated in the spread of fake news.
# - Fake news can reduce the impact of real news by competing with it. As a data scientist, our goal is to harness the good of the information age to catalyze change in society and hopefully progress within this field and likeminded researchers over time will reduce the negatives.
# - So, in this project we are going to design a classifier to classify whether a news is fake or not fake.
# # Dataset Description
# - The dataset has been downloaded from https://www.kaggle.com/c/fake-news/data
# - It consist of following :
# - train.csv: A full training dataset with the following attributes:
#
# 1. id: unique id for a news article
# 2. title: the title of a news article
# 3. author: author of the news article
# 4. text: the text of the article; could be incomplete
# 5. label: a label that marks the article as potentially unreliable
# - 1: unreliable
# - 0: reliable
#
# - test.csv: A testing training dataset with all the same attributes at train.csv without the label.
# # 1. Importing Dependencies
# +
# Basic Libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Sk-learn Libraries
from sklearn.naive_bayes import MultinomialNB
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
# Modelling Algorithms
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from xgboost import XGBClassifier
# Metrics
from sklearn.metrics import classification_report , accuracy_score , confusion_matrix, roc_auc_score
from sklearn.metrics import accuracy_score,f1_score
from sklearn.model_selection import cross_val_score
from sklearn import metrics
# NLP Libraries
import re
import itertools
import nltk
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
# Other Libraries
import string
from collections import Counter
# Keras Libraries
from tensorflow.keras.models import Model
from tensorflow.keras.layers import LSTM, Activation, Dense, Dropout, Input, Embedding
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.preprocessing.text import one_hot
from tensorflow.keras.preprocessing.sequence import pad_sequences
# %matplotlib inline
# -
# # 2. Data Reading & Analysis
# Reading the data
train_data = pd.read_csv('train.csv')
test_data = pd.read_csv('test.csv')
# ## Working with Train Data
# Looking at train data
train_data.head()
# Shape of train_data dataframe
train_data.shape
# Collecting info about the data
train_data.info()
# **Observations:**
# - It seems there are some null values. Lets pre-process the data
# Computing sum of Null values
train_data.isnull().sum()
# **Observations:**
# - Here we have two choices:
# 1. Remove the NaN entries
# 2. Fill the NaN with empty string ' '
# - It will be good to choose 2nd choice as if model is trained on a concatenation of the title, the author and the main text, the model would be more generalized because adding more words to the input might increase the reliablity of the model.
# Handling Missing Values
train_data = train_data.fillna(' ')
# Validating
train_data.isnull().sum()
# Dropping duplicates
train_data.drop_duplicates(inplace = True)
# Create a column with all the data available
train_data['total'] = train_data['title']+' '+train_data['author']+' '+train_data['text']
train_data.head()
# +
## Let's use groupby to use describe by label, this way we can begin to think about the features that separate
# Fake or Not Fake!
train_data.groupby('label').describe()
# -
# **Observations:**
# - Data looks balanced
# Let's make a new column to detect how news are:
train_data['news_length'] = train_data['text'].apply(len)
train_data.head()
# Analysing the 'news_length' feature
train_data['news_length'].describe()
# Max is 142961 characters, let's use masking to find this news
train_data[train_data['news_length'] == 142961]['title'].iloc[0]
# +
# Lets see if news length is a distinguishing feature between Fake and Not Fake:
fig,ax = plt.subplots(nrows = 1, ncols = 2, figsize = (12,6))
train_data[train_data['label'] == 1].hist(column = 'news_length', bins = 150, ax = ax[0], color = 'green')
ax[0].set(xlabel = 'Length of news', ylabel = 'Frequency', title = 'Non - Reliable / Fake')
train_data[train_data['label'] == 0].hist(column = 'news_length', bins = 50, ax = ax[1], color = 'red')
ax[1].set(xlabel = 'Length of news', ylabel = 'Frequency', title = 'Reliable / Not Fake')
# -
# **Observations:**
# - As expected Through the basic EDA we've seen that no seperation or trend has observed using 'news_length' feature.
# # 3. Pre-processing
# - Our main issue with our data is that it is all in text format (strings). The classification algorithms that we've learned about so far will need some sort of numerical feature vector in order to perform the classification task.
# - There are actually many methods to convert a corpus to a vector format. The simplest is the the bag-of-words approach, where each unique word in a text will be represented by one number.
# ## Train-Test Splitting
# Dividing the training set by using train_test_split
x_train, x_test, y_train, y_test = train_test_split(train_data['total'], train_data.label, test_size = 0.20, random_state = 0)
# ## Vectorizing our Data
# ### 1. Bag-of-Words (Count Vectorizer)
# +
# Initialize the 'count_vectorizer'
cv = CountVectorizer(ngram_range = (1, 2), stop_words = 'english')
# fit and transform the training data
count_train = cv.fit_transform(x_train)
# Transform the test data
count_test = cv.transform(x_test)
# -
print('Shape of Sparse Matrix: ', count_train.shape)
print('Amount of Non-Zero occurences: ', count_train.nnz)
print('Shape of Sparse Matrix: ', count_test.shape)
print('Amount of Non-Zero occurences: ', count_test.nnz)
# ### 2. TF-IDF (TFIDF Vectorizer)
# +
# Initialize the 'tfidf_vectorizer'
tfidf = TfidfVectorizer(ngram_range = (1, 2), stop_words = 'english')
# fit and transform the training data
tfidf_train = tfidf.fit_transform(x_train)
# transform the test set
tfidf_test = tfidf.transform(x_test)
# -
print('Shape of Sparse Matrix: ', tfidf_train.shape)
print('Amount of Non-Zero occurences: ', tfidf_train.nnz)
print('Shape of Sparse Matrix: ', tfidf_test.shape)
print('Amount of Non-Zero occurences: ', tfidf_test.nnz)
# ### 3. Pipelined (BOW-TFIDF Transformer)
# +
# Initialize the 'tfidf_transformer'
tv = TfidfTransformer()
# Learn the training data vocabulary, then use it to create a document-term matrix
pipelined_train = tv.fit_transform(count_train)
# Transform test data (using fitted vocabulary) into a document-term matrix
pipelined_test = tv.transform(count_test)
# -
print('Shape of Sparse Matrix: ', pipelined_train.shape)
print('Amount of Non-Zero occurences: ', pipelined_train.nnz)
print('Shape of Sparse Matrix: ', pipelined_test.shape)
print('Amount of Non-Zero occurences: ', pipelined_test.nnz)
# # 4. Building and Evaluating a Model
# +
# Defining a function that outputs a confusion matrix
def plot_confusion_matrix(cm, classes, normalize = False, title = 'Confusion matrix', cmap = plt.cm.Blues):
plt.imshow(cm, interpolation = 'nearest', cmap = cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation = 45)
plt.yticks(tick_marks, classes)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j], horizontalalignment = "center", color = "white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# -
# ## 1. Multinomial Naive Bayes
# ### 1.1 Multinomial Naive Bayes with Count Vectorizer (Bag-of-Words)
# +
# Tuning Hyper-Parameters
grid_parameters = {'alpha' : [0.01, 0.1, 0.15, 0.2]}
# Instantiate the classifier
nb_classifier_bow = GridSearchCV(MultinomialNB(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using count_train
nb_classifier_bow.fit(count_train, y_train)
# Finding best_estimator
nb_classifier_bow_best = nb_classifier_bow.best_estimator_
# Predicting for Test data using count_test
y_pred_nb = nb_classifier_bow_best.predict(count_test)
print ("Best Parameters For Multinomial Naive Bayes for Bag-of-Words : ", nb_classifier_bow.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy Multinomial for Naive Bayes for Bag-of-Words :")
print(nb_classifier_bow_best.score(count_train, y_train))
print('-'*110)
print("Test Accuracy Multinomial for Naive Bayes for Bag-of-Words :")
print(accuracy_score(y_pred_nb , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_nb, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a MultinomialNB with Count Vectorizer')
# -
# ### 1.2 Multinomial Naive Bayes with TF-IDF Vectorizer
# +
# Tuning Hyper-Parameters
grid_parameters = {'alpha' : [0.01, 0.1, 0.15, 0.2]}
# Instantiate the classifier
nb_classifier_tfidf = GridSearchCV(MultinomialNB(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using tfidf_train
nb_classifier_tfidf.fit(tfidf_train, y_train)
# Finding best_estimator
nb_classifier_tfidf_best = nb_classifier_tfidf.best_estimator_
# Predicting for Test data using tfidf_test
y_pred_nb_tfidf = nb_classifier_tfidf_best.predict(tfidf_test)
print ("Best Parameters For Multinomial Naive Bayes for TFIDF vectorizer : ", nb_classifier_tfidf.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy Multinomial Naive Bayes for TFIDF vectorizer :")
print(nb_classifier_tfidf_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy Multinomial Naive Bayes for TFIDF vectorizer :")
print(accuracy_score(y_pred_nb_tfidf , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_nb_tfidf, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Multinomial Naive Bayes for TFIDF vectorizer')
# -
# ### 1.3 Multinomial Naive Bayes with Pipelined(BOW-TF-IDF Transformer)
# +
# Tuning Hyper-Parameters
grid_parameters = {'alpha' : [0.01, 0.1, 0.15, 0.2]}
# Instantiate the classifier
nb_classifier_pipelined = GridSearchCV(MultinomialNB(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using pipelined_train
nb_classifier_pipelined.fit(pipelined_train, y_train)
# Finding best_estimator
nb_classifier_pipelined_best = nb_classifier_pipelined.best_estimator_
# Predicting for Test data using pipelined_test
y_pred_nb_pipelined = nb_classifier_pipelined_best.predict(pipelined_test)
print ("Best Parameters For Multinomial Naive Bayes for Pipelined Transformer : ", nb_classifier_pipelined.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy Multinomial Naive Bayes for pipelined Transformer :")
print(nb_classifier_pipelined_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy Multinomial Naive Bayes for Pipelined Transformer :")
print(accuracy_score(y_pred_nb_pipelined , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_nb_pipelined, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Multinomial Naive Bayes for Pipelined Transformer')
# -
# ## 2. Passive Agressive Classifier
# ### 2.1 Passive Agressive Classifier With Count Vectorizer (Bag-of-Words)
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1]}
# Instantiate the classifier
pac_classifier_bow = GridSearchCV(PassiveAggressiveClassifier(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using count_train
pac_classifier_bow.fit(count_train, y_train)
# Finding best_estimator
pac_classifier_bow_best = pac_classifier_bow.best_estimator_
# Predicting for Test data using count_test
y_pred_pac = pac_classifier_bow_best.predict(count_test)
print ("Best Parameters For Passive Agressive Classifier for Bag-of-Words : ", pac_classifier_bow.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy Passive Agressive Classifier for Bag-of-Words :")
print(pac_classifier_bow_best.score(count_train, y_train))
print('-'*110)
print("Test Accuracy Passive Agressive Classifier for Bag-of-Words :")
print(accuracy_score(y_pred_pac , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_pac, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Passive Agressive Classifier with Count Vectorizer')
# -
# ### 2.2 Passive Agressive Classifier With TFIDF Vectorizer
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1, 1.5]}
# Instantiate the classifier
pact_classifier_tfidf = GridSearchCV(PassiveAggressiveClassifier(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using tfidf_train
pact_classifier_tfidf.fit(tfidf_train, y_train)
# Finding best_estimator
pact_classifier_tfidf_best = pact_classifier_tfidf.best_estimator_
# Predicting for Test data using tfidf_test
y_pred_pact_tfidf = pact_classifier_tfidf_best.predict(tfidf_test)
print ("Best Parameters For Passive Agressive Classifier With TFIDF Vectorizer : ", pact_classifier_tfidf.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy for Passive Agressive Classifier With TFIDF Vectorizer :")
print(pact_classifier_tfidf_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy for Passive Agressive Classifier With TFIDF Vectorizer :")
print(accuracy_score(y_pred_pact_tfidf , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_pact_tfidf, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Passive Agressive Classifier With TFIDF Vectorizer')
# -
# ### 2.3 Passive Agressive Classifier With Pipelined (BOW-TFIDF Transformer)
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1, 1.5]}
# Instantiate the classifier
pacp_classifier_pipelined = GridSearchCV(PassiveAggressiveClassifier(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using pipelined_train
pacp_classifier_pipelined.fit(pipelined_train, y_train)
# Finding best_estimator
pacp_classifier_pipelined_best = pacp_classifier_pipelined.best_estimator_
# Predicting for Test data using pipelined_test
y_pred_pacp_pipelined = pacp_classifier_pipelined_best.predict(pipelined_test)
print ("Best Parameters For Passive Agressive Classifier With Pipelined Transformer : ", pacp_classifier_pipelined.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy Passive Agressive Classifier With Pipelined Transformer :")
print(pacp_classifier_pipelined_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy Passive Agressive Classifier With Pipelined Transformer :")
print(accuracy_score(y_pred_pacp_pipelined , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_pacp_pipelined, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Passive Agressive Classifier With Pipelined Transformer')
# -
# ## 3. Logistic Regression
# ### 3.1 Logistic Regression with CountVectorizer (Bag-of-Words)
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1, 1.5], 'penalty': ['l1','l2']}
# Instantiate the classifier
lr_classifier_bow = GridSearchCV(LogisticRegression(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using count_train
lr_classifier_bow.fit(count_train, y_train)
# Finding best_estimator
lr_classifier_bow_best = lr_classifier_bow.best_estimator_
# Predicting for Test data using count_test
y_pred_lr = lr_classifier_bow_best.predict(count_test)
print ("Best Parameters For Logistic Regression for Bag-of-Words : ", lr_classifier_bow.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy for Logistic Regression for Bag-of-Words :")
print(lr_classifier_bow_best.score(count_train, y_train))
print('-'*110)
print("Test Accuracy for Logistic Regression for Bag-of-Words :")
print(accuracy_score(y_pred_lr , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_lr, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Logistic Regression with Count Vectorizer')
# -
# ### 3.2 Logistic Regression with TFIDF Vectorizer
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1, 1.5], 'penalty': ['l1','l2']}
# Instantiate the classifier
lrt_classifier_tfidf = GridSearchCV(LogisticRegression(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using tfidf_train
lrt_classifier_tfidf.fit(tfidf_train, y_train)
# Finding best_estimator
lrt_classifier_tfidf_best = lrt_classifier_tfidf.best_estimator_
# Predicting for Test data using tfidf_test
y_pred_lrt_tfidf = lrt_classifier_tfidf_best.predict(tfidf_test)
print ("Best Parameters For Logistic Regression With TFIDF Vectorizer : ", lrt_classifier_tfidf.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy for Logistic Regression With TFIDF Vectorizer :")
print(lrt_classifier_tfidf_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy for Logistic Regression With TFIDF Vectorizer :")
print(accuracy_score(y_pred_lrt_tfidf , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_lrt_tfidf, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Logistic Regression With TFIDF Vectorizer')
# -
# ### 3.3 Logistic Regression with Pipelined (BOW-TFIDF Transformer)
# +
# Tuning Hyper-Parameters
grid_parameters = {'C' : [0.01, 0.1, 0.5, 1, 1.5], 'penalty': ['l1','l2']}
# Instantiate the classifier
lrp_classifier_pipelined = GridSearchCV(LogisticRegression(), grid_parameters, verbose = 1, cv = 3, n_jobs = -1)
# Train the model using pipelined_train
lrp_classifier_pipelined.fit(pipelined_train, y_train)
# Finding best_estimator
lrp_classifier_pipelined_best = lrp_classifier_pipelined.best_estimator_
# Predicting for Test data using pipelined_test
y_pred_lrp_pipelined = lrp_classifier_pipelined_best.predict(pipelined_test)
print ("Best Parameters For Logistic Regression With Pipelined Transformer : ", lrp_classifier_pipelined.best_params_)
print('-'*110)
# Evaluation
print("Training Accuracy for Logistic Regression With Pipelined Transformer :")
print(lrp_classifier_pipelined_best.score(tfidf_train, y_train))
print('-'*110)
print("Test Accuracy for Logistic Regression With Pipelined Transformer :")
print(accuracy_score(y_pred_lrp_pipelined , y_test))
print('-'*110)
# Plotting Confusion matrix
cm = metrics.confusion_matrix(y_test, y_pred_lrp_pipelined, labels = [0,1])
plot_confusion_matrix(cm, classes = ['TRUE','FAKE'], title ='Confusion Matrix for a Logistic Regression With Pipelined Transformer')
# -
# ## Conclusions For Machine Learning Algorithms
# - We get highest Accuracy with **Passive Agressive Classifier With Pipelined (BOW-TFIDF Transformer)** about 98.02% on test data. But when we analyse confusion matrix we have more number of fake news labeled as true news i.e. 58.
# - We get good Accuracy with **Logistic Regression with CountVectorizer (Bag-of-Words)** about 97.88% on test data. Even though the accuracy score is a bit lower, we have less fake news labeled as true news ie. only 41.
# - Therefore, this model can be chosen for Submitting the actual test file because it seems to maximize the accuracy while minimizing the false negative rate!
# - But yet before reaching to final conclusion lets analyze by using Deep Learning Techniques
# ## 4. LSTMs
# Lets copy the original data into new variable
new_train_data = train_data.copy()
new_train_data.head()
# Lets sort the news based on 'news_length'
new_train_data.sort_values(by = 'news_length', inplace = True)
new_train_data
# **Note:**
# - As the Lengths of 'text' i.e description of news is in lakhs of words so due to lack of computational resources we will restrict our analysis to only 'title' column.
# Lets copy the original data into new variable
new_train_data = train_data.copy()
new_train_data.head()
# Considering only One Feature i.e 'title' as author will not make sense initially.
x = new_train_data['title']
y = new_train_data['label']
x.shape
y.shape
# ### Pre-processing
ps = PorterStemmer()
corpus = []
for i in range(0, len(new_train_data)):
title = re.sub('[^a-zA-Z]', ' ', new_train_data['title'][i])
title = title.lower()
title = title.split()
STOPWORDS = set(stopwords.words('english')) - set(['not', "aren't", "couldn't", "didn't", "doesn't", "don't"
"hadn't", "haven't", "isn't", "shan't", "needn't", "shouldn't"
"wasn't", "wouldn't"])
title = [word for word in title if not word in STOPWORDS]
title = ' '.join(title)
corpus.append(title)
# Lets look for some news title
corpus[0]
corpus[30]
# +
# Consider vocabulory size as 7000
vocab_size = 7000
# Lets perform one hot representation
one_hot_encode = [one_hot(words, vocab_size)for words in corpus]
# -
# Consider length of first title
len(one_hot_encode[0])
# +
# Here we are specifying a sentence length so that each sentence in the corpus will be of same length. Consider 30 arbitarily
sentence_length = 30
# Lets perform padding for creating equal length sentences
embedded_titles = pad_sequences(one_hot_encode, padding = 'pre', maxlen = sentence_length)
print(embedded_titles)
# -
# ## Train - Test Splitting
z = np.array(embedded_titles)
y = np.array(y)
X_train, X_test, y_train, y_test = train_test_split(z, y, test_size = 0.20, random_state = 1)
# ## Modeling
# +
# Initializing the Model
model = Sequential()
# Adding Embedded Layer
model.add(Embedding(input_dim = vocab_size, output_dim = 100, input_length = sentence_length))
# Adding Dropout Layer
model.add(Dropout(0.3))
# Adding LSTM Layer
model.add(LSTM(200))
# Adding Dropout Layer
model.add(Dropout(0.3))
# Fully Connected Layer
model.add(Dense(1, activation = 'sigmoid'))
# Compiling the Model
model.compile(loss = 'binary_crossentropy', optimizer = 'adam', metrics = ['accuracy'])
# -
model.summary()
model_result = model.fit(X_train, y_train, validation_data = (X_test, y_test), epochs = 20, batch_size = 64, workers = -1)
model_evaluation = model.evaluate(X_test, y_test)
print('Test Loss is : {} and the Test Accuracy is: {}'.format(model_evaluation[0], model_evaluation[1]))
# this function is used to update the plots for each epoch and error
def plt_dynamic(x, vy, ty, ax, colors = ['b']):
ax.plot(x, vy, 'b', label = "Validation Loss")
ax.plot(x, ty, 'r', label = "Train Loss")
plt.legend()
plt.grid()
fig.canvas.draw()
# +
# Plotting Train & Validation Loss
fig,ax = plt.subplots(1,1)
ax.set_xlabel('epochs')
ax.set_ylabel('Binary Crossentropy Loss')
# list of epoch numbers
p = list(range(1, 21))
vy = model_result.history['val_loss']
ty = model_result.history['loss']
# Calling Function
plt_dynamic(p, vy, ty, ax)
# -
# **Observations:**
# - As seen from the above graph, our model is overfitting due to usage of only one Feature & limited vocab. Yet we have achieved the accuracy about 92.67% which is not bad.
# - If we utilized all the features then we could hope for better results. For now lets stop here.
# # End Results
# - We get highest Accuracy with **Passive Agressive Classifier With Pipelined (BOW-TFIDF Transformer)** about 98.02% on test data. But when we analyse confusion matrix we have more number of fake news labeled as true news i.e. 58.
# - We get good Accuracy with **Logistic Regression with CountVectorizer (Bag-of-Words)** about 97.88% on test data. Even though the accuracy score is a bit lower, we have less fake news labeled as true news ie. only 41.
# - Therefore, this model can be chosen for Submitting the actual test file because it seems to maximize the accuracy while minimizing the false negative rate!
# # Working with Test Data
# Looking at test data
test_data.head()
# Shape of train_data dataframe
test_data.shape
# Computing sum of Null values
test_data.isnull().sum()
# Handling Missing Values
test_data = test_data.fillna(' ')
test_data.head()
# Dropping duplicates
test_data.drop_duplicates(inplace = True)
# Create a column with all the data available
test_data['total'] = test_data['title']+' '+test_data['author']+' '+test_data['text']
test_data.head()
test = test_data['total']
# +
# Initialize the 'count_vectorizer'
cv = CountVectorizer(ngram_range = (1, 2), stop_words = 'english')
# fit and transform the training data
count_train = cv.fit_transform(x_train)
# Transform the test data
test_bow = cv.transform(test)
# -
print('Shape of Sparse Matrix: ', count_train.shape)
print('Amount of Non-Zero occurences: ', count_train.nnz)
print('Shape of Sparse Matrix: ', test_bow.shape)
print('Amount of Non-Zero occurences: ', test_bow.nnz)
pred_test_data = lr_classifier_bow_best.predict(test_bow)
final_submission = test_data
final_submission['Predicted_Label'] = pred_test_data
final_submission.to_csv('Fake_News_Prediction.csv', index = False)
# # End of File
| Fake News Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/siqitoday/Playground/blob/main/HODL%20Homework1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="h5kWWfR8z9uZ"
import tensorflow as tf
from tensorflow import keras
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# initialize the seeds of different random number generators so that the
# results will be the same every time the notebook is run
keras.utils.set_random_seed(42)
# + [markdown] id="oWOYqBF03aBE"
# # **Introduction**
#
# Your goal in this exercise is to recognize sentiment from a facial image. To that end, we will use the 2013 FER ('Facial Expression Recognition'). The dataset consists of ~ 36,000 images, each annotated with one of seven labels: 'angry', 'disgust', 'fear', 'happy', 'sad', 'surprise', 'neutral'. Our goals will be to:
#
# 1) Build a conv-net from scratch to recognize images.
# 2) Use transfer learning as an alternative.
#
# But first, lets start with grabbing the data itself from Dropbox
# + id="kklgssFnY8Ob"
# !wget -q -P ./ https://www.dropbox.com/s/ia62dg6kpp3q8wb/fer2013.csv
# + colab={"base_uri": "https://localhost:8080/"} id="U5S29hLu7djK" outputId="c620eb73-4184-4f7b-a569-24f6189c2c63"
data = pd.read_csv('/content/fer2013.csv')
print(data.head())
# + [markdown] id="BtEmyeIL7o9V"
# As you can see, each (gray-scale) image is encoded as a list of (2304) pixels that we will reshape into an (48x48) image. This keeps the size of the dataset manageable. Moreover, each image is associated with an emotion label between 0 and 6 that map respectively to the emotions 'angry', 'disgust', 'fear', 'happy', 'sad', 'surprise', and 'neutral'. Let's convert these pixel lists into actual images that we can work with next.
# + id="sG4mAu4k-k86"
pixels = data['pixels'].tolist()
width, height = 48, 48
faces = []
for pixel_sequence in pixels:
face = [int(pixel) for pixel in pixel_sequence.split(' ')] # read each face as a 1-d array
face = np.asarray(face).reshape(width, height) # reshape the length 2304 1-d array into an 48x48 array
face = np.stack((face,)*3, axis=-1) # convert single channel to three channels simply by replicating the single channel we have.
faces.append(face.astype('float32'))
faces = np.asarray(faces)
emotions = pd.get_dummies(data['emotion']).to_numpy() # each emotion is 'one-hot' encoded as a 7-dim vector
emotions_names = ('angry', 'disgust', 'fear', 'happy', 'sad', 'surprise', 'neutral')
# + [markdown] id="4WZRh-dDBvWw"
# Lets take a look at some of these fun images!
# + colab={"base_uri": "https://localhost:8080/", "height": 591} id="iqmEpir1B3Mu" outputId="7627b790-da45-4139-de04-aa730c6c751e"
fig = plt.figure(figsize=(10, 10))
for i in range(9):
ax = fig.add_subplot(3, 3, i+1)
ax.set_title(f"{emotions_names[np.argmax(emotions[i])]}")
ax.imshow(faces[i].astype('uint8'))
ax.axis('off')
# + [markdown] id="Sts35VbQGJMf"
# As in the original dataset, we will reserve the first 28,709 images for training and the rest for testing.
# + id="ux8sgM1kptNb"
train_faces, train_emotions = faces[:28709], emotions[:28709]
test_faces, test_emotions = faces[28709:], emotions[28709:]
# + colab={"base_uri": "https://localhost:8080/"} id="Rs8BTG252Oh9" outputId="727befc2-fde6-4570-e8e3-c55253f1924f"
print(train_faces.shape, train_emotions.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="kr2jXspqqs7n" outputId="31703ae1-ef2d-4bcc-f381-5bb1b48fd15e"
print(test_faces.shape, test_emotions.shape)
# + [markdown] id="-aQhn5et0LoL"
# # **Construct a CNN Model**
#
# We will try a simple CNN on this dataset with three convolutional + pooling layers + one dense layer
# + id="dJD-ejmW0naX"
input = keras.Input(shape=train_faces.shape[1:])
x = keras.layers.Rescaling(1./255)(input) #normalizing
x = keras.layers.Conv2D(16, kernel_size=(2, 2), activation="relu", name="Conv_1")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Conv2D(16, kernel_size=(2, 2), activation="relu", name="Conv_2")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Conv2D(16, kernel_size=(2, 2), activation="relu", name="Conv_3")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Flatten()(x)
x = keras.layers.Dense(256, activation="relu")(x)
output = keras.layers.Dense(7, activation="softmax")(x)
model = keras.Model(input, output, name='CNN_model')
# + id="oQ9uJx7YtU4H" outputId="46d3f497-0933-416e-c5c5-b755ffe51539" colab={"base_uri": "https://localhost:8080/"}
model.summary()
# + id="yHAHKo0W1fym"
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="tGd1oX0N1kfJ" outputId="475097b9-a7e5-4164-c229-2ecbe98cbae3"
epochs = 30
history = model.fit(train_faces, train_emotions,
batch_size=64,
epochs=epochs,
validation_split=0.2)
# + colab={"base_uri": "https://localhost:8080/"} id="Q2S3SGwF1bXW" outputId="c1e50710-64a4-4ab3-efec-bf9c891d19dc"
score = model.evaluate(test_faces, test_emotions)
print("Test accuracy:", score[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="8YHzLOsNuwbE" outputId="255bcc68-ed63-49ae-d199-aa92df6eccdd"
history_dict = history.history
acc = history_dict["accuracy"]
val_acc = history_dict["val_accuracy"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
# + [markdown] id="UIGCf1_zA5G1"
# # **Data Augmented CNN Model**
#
# + [markdown] id="3C4PGf4X1Ib5"
# ***Data Augmentation:***
#
# The basic idea of augmentation is to slightly alter the image so that the value of the dependent variable (i.e. the category that it belongs to) doesn't change.
# + id="A18vDhSv6RWK"
data_augmentation = keras.Sequential(
[
keras.layers.RandomFlip("horizontal"),
keras.layers.RandomZoom(0.2),
]
)
# + [markdown] id="0ehLSXqP1b5P"
# Lets quickly visualize what an augmentation does ...
# + colab={"base_uri": "https://localhost:8080/", "height": 578} id="gTDMmESZ6TZ_" outputId="8c6ee894-41e8-4622-9432-7c5994c53f73"
augmented_images = [data_augmentation(np.expand_dims(train_faces[0],axis=0)) for i in range(9)]
fig = plt.figure(figsize=(10, 10))
for i in range(9):
ax = fig.add_subplot(3, 3, i+1, xticks=[], yticks=[])
ax.imshow(tf.keras.preprocessing.image.array_to_img(augmented_images[i][0]))
# + id="9Yk6UhMrBSVt"
input = keras.Input(shape=train_faces.shape[1:])
x = data_augmentation(input)
x = keras.layers.Rescaling(1./255)(x)
x = keras.layers.Conv2D(32, kernel_size=(2, 2), activation="relu", name="Conv_1")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Conv2D(32, kernel_size=(2, 2), activation="relu", name="Conv_2")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Conv2D(32, kernel_size=(2, 2), activation="relu", name="Conv_3")(x) # convolutional layer!
x = keras.layers.MaxPool2D()(x) # pooling layer
x = keras.layers.Flatten()(x)
x = keras.layers.Dense(512, activation="relu")(x)
output = keras.layers.Dense(7, activation="softmax")(x)
model = keras.Model(input, output, name='augmented_CNN_model')
# + colab={"base_uri": "https://localhost:8080/"} id="oFDrN4GfBiRx" outputId="9b5a1ce2-d635-4641-841b-30a5ce2aac81"
model.summary()
# + id="mJCy1tCyBhxb"
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="PMW5t9ZOBlyC" outputId="520367c2-3a82-4ed6-8e4a-0d0872f92867"
epochs = 20
history = model.fit(train_faces, train_emotions,
batch_size=64,
epochs=epochs,
validation_split=0.2)
# + colab={"base_uri": "https://localhost:8080/"} id="terijZIKBod8" outputId="8bdaee17-63f6-45d9-abc8-63c2d196280d"
score = model.evaluate(test_faces, test_emotions)
print("Test accuracy:", score[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="xhXMussbBqTQ" outputId="5452146f-529c-4f72-9193-0423a1ce8793"
history_dict = history.history
acc = history_dict["accuracy"]
val_acc = history_dict["val_accuracy"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
# + [markdown] id="4Ddqer_8KbwB"
# # **Constructing a Transfer Learning Model**
#
# Next, we architect a basic transfer learning model. We will take a slightly different approach from what we saw in class: as opposed to generating feature vectors by running each image through a pre-trained model, we will simply (a) remove the top from an existing pre-trained model, and (b) add a custom designed top. Why are we doing this? Its so that we can 'tweak' the weights of the original pre-trained model to suit our problem.
# + [markdown] id="0GdCS59whVDu"
# ***Fine Tune Model:***
#
# In the stub below, we define a 'fine-tuned' model to stack on top of a pre-trained base model. In contrast with what we did in class, we do not explicitly use the pre-trained model to generate stored features but incorporate it as part of the overall network.
# + id="FSUlxaZvKVBd"
top_model = keras.Sequential(
[
keras.layers.Flatten(),
keras.layers.Dense(1024, activation='relu'),
keras.layers.Dense(1024, activation='relu'),
]
)
# + [markdown] id="dHKqB9r1iH9W"
# ***Overall Model:***
#
# Below, we take a base-model (the VGG19 model: https://keras.io/api/applications/vgg/) and perform the following steps:
#
#
# 1. Remove the top of the model (include_top=False)
# 2. 'Freeze' the model's parameters so they do not get impacted by training
# 3. Feed the output of the base model to our fine-tune model
# 4. Feed the output of our fine-tune model to a soft-max output layer.
# 5. **Unfreeze the last 10 layers of the VGG19 model so we can update those weights as well.**
#
#
# + id="CYOu-5lMPZRX"
def construct_model(no_classes, input_shape, metrics=['accuracy']):
base_model = keras.applications.VGG19(
include_top=False,
weights="imagenet",
input_tensor=None,
input_shape=None,
pooling=None,
classes=1000,
classifier_activation="softmax",
)
# Freeze the base_model
base_model.trainable = False
#
# Create new model on top
#
inputs = keras.Input(shape=input_shape)
x = keras.layers.Rescaling(1./255)(inputs) #normalizing
# Apply random data augmentation
x = data_augmentation(x)
# The base model contains batchnorm layers. We want to keep them in inference mode
# when we unfreeze the base model for fine-tuning, so we make sure that the
# base_model is running in inference mode here. We didn't cover batchnorm
# layers in class so just take our word for it :-)
x = base_model(x, training=False)
# Next we feed the output from our base model to the top model we designed.
x = top_model(x)
outputs = keras.layers.Dense(no_classes, activation='softmax')(x)
model = keras.Model(inputs, outputs)
model.summary()
#unfreeze the last 10 layers of the model so that some tweaks can be done to the weights of the VGG19 model.
for layer in model.layers[-10:]:
if not isinstance(layer, keras.layers.BatchNormalization): #the batch normalization layer is untouched
layer.trainable = True
model.compile(loss='categorical_crossentropy', optimizer=keras.optimizers.Adam(0.2*1e-4), metrics=metrics) #here we choose a different rate for Adam than default for a better convergence
return model
# + [markdown] id="-OlMcWW6ji1Z"
# ***Training the Overall Model***
# + colab={"base_uri": "https://localhost:8080/"} id="g0Ahpg7vQ519" outputId="6ce605e6-a469-4c9f-aa4e-092effacff93"
no_classes = 7
NO_EPOCHS = 30
model = construct_model(no_classes,(48,48,3))
history = model.fit(train_faces, train_emotions, epochs=NO_EPOCHS, validation_split=0.2)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="_XWZv3tCeKWA" outputId="e2cbc730-c6c1-4795-f588-272b83561047"
history_dict = history.history
acc = history_dict["accuracy"]
val_acc = history_dict["val_accuracy"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
# + id="kshP4LwZ9TV2"
MODEL_FILE_NAME = 'fer2013VGGTransferFinal.h5'
model.save(MODEL_FILE_NAME)
# + [markdown] id="Trh8XLTcXnlr"
# The accuracy for the testing set is improved significantly (the state-of-the-art with tremendous efforts is around 73.3%)
# + colab={"base_uri": "https://localhost:8080/"} id="mzg5PNAf1BiX" outputId="ec87ca32-c254-4ff0-f069-1a8dfa0009b4"
score = model.evaluate(test_faces, test_emotions)
print("Test accuracy:", score[1])
# + id="z1W3a8u68GVo"
predictions_index = model.predict(test_faces).argmax(axis=1)
actuals_index = test_emotions.argmax(axis=1)
actuals = [emotions_names[i] for i in actuals_index]
predictions = [emotions_names[i] for i in predictions_index]
# + [markdown] id="yYkyOqTgl2oT"
# Check out the confusion matrix
# + id="1aHxeB2w85LM"
#df = pd.DataFrame({'Predictions': predictions, 'Actuals': actuals})
#pd.crosstab(df.Predictions, df.Actuals)
| HODL Homework1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Plotting with matplotlib
# + slideshow={"slide_type": "-"}
# most common way to import matplotlib
from matplotlib import pyplot as plt
# -
# import numpy to work with data to plot
import numpy as np
# Magic Command that sets how matplotlib integrates into the ipython notebook.
# %matplotlib inline
# Setup for darkmode browser
plt.style.use('dark_background')
# + slideshow={"slide_type": "slide"}
x = np.linspace(-2,2, 50)
y = np.cos(x)
plt.figure(figsize=(7,5))
plt.plot(x,y, ".-")
plt.xlabel("time (s)")
plt.ylabel("Voltage (V)")
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is matplotlib?
#
# - Matplotlib is basically Matlabs plotting translated into Python. (Similar to numpy which is also heavily based on Matlab.)|
# - It's cumbersome but versatile
# - It can make paper quality plots but is maybe not the right choice for live plotting.
# + [markdown] slideshow={"slide_type": "slide"}
# ## How to best use Matplotlib
#
# - create a style file
# - import that file in different scripts and notebooks for consistency
#
# - if you want to do things quickly
# - use subplot and tight_layout
# - if you want to create polished paper figures
# - use gridspec for multi-axis plots
# - use subplots_adjust to manage whitespace around figure
# + slideshow={"slide_type": "slide"}
# Do not run %matplotlib inline after these definitions, else they will get overwritten
import matplotlib
fnt_size = 13.
full_fig_width = 12.2
full_fig_width_in = full_fig_width/2.54
my_font = {'sans-serif':'Arial'}
my_font.update({'size':fnt_size})
matplotlib.rc('font', **my_font)
my_axes = {'formatter.useoffset':False,
'labelsize':fnt_size,
'titlesize':fnt_size}
matplotlib.rc('axes',**my_axes)
# my_xtick = {'major.pad':8,
# 'minor.pad':8}
my_xtick = {'direction':'in',
'top':True}
matplotlib.rc('xtick',**my_xtick)
my_ytick = {'direction':'in',
'right':True}
matplotlib.rc('ytick',**my_ytick)
# ax.ticklabel_format(useOffset=False)
# figure = {'subplot.bottom': 0.1,
# 'subplot.left': 0.15}
# figure.subplot.hspace: 0.2
# figure.subplot.left: 0.125
# figure.subplot.right: 0.9
# figure.subplot.top: 0.9
# figure.subplot.wspace: 0.2}
# matplotlib.rc('figure',**figure)
# legend = {'frameon':False,
# 'fontsize':'small'}
legend = {'numpoints':1,
'handlelength':1,
'frameon':'False',
'fontsize':fnt_size,
'loc':'best'}
matplotlib.rc('legend', **legend)
my_line = {'linewidth':0.75,
'markersize':3}
matplotlib.rc('lines', **my_line)
# -
x = np.linspace(-2,2, 50)
y = np.cos(x)
plt.figure(figsize=(7,5))
plt.plot(x,y, ".-")
plt.xlabel("time (s)")
plt.ylabel("Voltage (V)")
# ## Paper figures
| Plotting with matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# %matplotlib inline
# Import dependencies
import matplotlib.pyplot as plt
import statistics
# +
# Set the x-axis to a list of strings for each month.
x_axis = ["Jan", "Feb", "Mar", "April", "May", "June", "July", "Aug", "Sept", "Oct", "Nov", "Dec"]
# Set the y-axis to a list of floats as the total fare in US dollars accumulated for each month.
y_axis = [10.02, 23.24, 39.20, 35.42, 32.34, 27.04, 43.82, 10.56, 11.85, 27.90, 20.71, 20.09]
# -
# Get the standard deviation of the values in the y-axis
stdev = statistics.stdev(y_axis)
stdev
plt.errorbar(x_axis, y_axis, yerr=stdev)
plt.errorbar(x_axis, y_axis, yerr=stdev, capsize=3)
fig, ax = plt.subplots()
ax.errorbar(x_axis, y_axis, yerr=stdev, capsize=3)
plt.show()
# adding errors to a bar chart
plt.bar(x_axis, y_axis, yerr=stdev, capsize=3)
# adding errors to bar chart using OOP method
fig, ax = plt.subplots()
ax.bar(x_axis, y_axis, yerr=stdev, capsize=3)
plt.show()
# Adjusting x-ticks via NumPy
import numpy as np
plt.barh(x_axis, y_axis)
plt.xticks(np.arange(0, 51, step=5.0))
plt.gca().invert_yaxis()
# +
# adding minor tickers w/ NumPy
from matplotlib.ticker import MultipleLocator
# Increase the size of the plot figure
fig, ax = plt.subplots(figsize=(8,8))
ax.barh(x_axis, y_axis)
ax.set_xticks(np.arange(0, 51, step=5.0))
# Create minor ticks at an increment of 1
ax.xaxis.set_minor_locator(MultipleLocator(1))
plt.show()
# -
| Chart_Extras.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] id="55C63pB8Va-v"
# # Классические методы и критерии статистики
# + [markdown] id="udKXx2oGVa0n"
# ## Гипотеза о равенстве средних двух генеральных совокупностей
# + [markdown] id="YbwkC7WaWc5A"
# ### Тест Шапиро-Уилка
# + colab={"base_uri": "https://localhost:8080/", "height": 522} id="Ii2khqlvWcha" outputId="d013eac1-d796-4893-efd5-4254b1f193e0"
d.intake <- c(5260,5470,5640,6180,6390,6515,6805,7515,7515,8230,8770)
shapiro.test(d.intake)
hist(d.intake)
# + [markdown] id="r113CVjVVag3"
# ### Одновыборочный t-критерий
# + id="KNzQMRm3VAY1"
d.intake <- c(5260,5470,5640,6180,6390,6515,6805,7515,7515,8230,8770)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="G6aBXinrWIpP" outputId="eef2e764-6036-4082-edf0-1006661837ec"
# среднее
mean(d.intake)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="1GVeDR-dWZgu" outputId="aab56346-6bdb-4adb-fc87-75372cd7b817"
# Отличается ли это выборочное среднее значение от установленной нормы 7725?
t.test(d.intake, mu=7725)
# + id="p-OpLLnOXJWF"
# Вероятность получить такое (либо большее) значение t при условии, что проверяемая нулевая гипотеза верна, оказалась весьма мала.
# Следовательно, мы можем отклонить проверяемую нулевую гипотезу о равенстве выборочного среднего значения нормативу и принять альтернативную гипотезу.
# Принимая это предположение, мы рискуем ошибиться с вероятностью менее 5%.
# + [markdown] id="PbHqy8rmYQ7P"
# ### Сравнение двух независимых выборок
# + colab={"base_uri": "https://localhost:8080/", "height": 255} id="eDdGmKA9X-Ks" outputId="f35753cc-23eb-4953-8c08-0d9a6f462a18"
# При сравнении двух выборок проверяемая нулевая гипотеза состоит в том, что обе эти выборки происходят из нормально распределенных
# генеральных совокупностей с одинаковыми средними значениями.
install.packages("ISwR")
library(ISwR)
data(energy)
attach(energy)
head(energy)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="nTAXNFzuZzjn" outputId="8af80136-b40a-4ac2-f4c8-8b111457be2c"
# Соответствующие средние значения
tapply(expend, stature, mean)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="syJcVwM9aBfF" outputId="ea53d33e-3b37-4e09-c628-455c54469179"
# t-test (знак ~ это обозначение зависимости между переменными)
t.test(expend ~ stature)
# + [markdown] id="RuD7gYQ0bmUk"
# ### Сравнение двух зависимых выборок
# + id="wren4NlNbfXu"
# Парный критерий Стьюдента
# t.test (x, y, paired = TRUE)
# + [markdown] id="5Waa0ow6cD_P"
# ## Использование рангового критерия Уилкоксона-Манна-Уитни
# + [markdown] id="lku8ZqXlcdmt"
# ### Одновыборочный критерий Уилкоксона
# + id="5kWd1vDtb7Jc"
d.intake <- c(5260, 5470, 5640, 6180, 6390, 6515, 6805, 7515, 7515, 8230, 8770)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} id="mYATAxtOeLNj" outputId="f14aeace-8e78-4f87-f9f1-ec496f774893"
wilcox.test(d.intake, mu = 7725)
# + [markdown] id="zdb6Tp_fezt-"
# ### Сравнение двух независимых выборок
# + colab={"base_uri": "https://localhost:8080/"} id="Zv_4tDhsetUb" outputId="e182e7c8-3e4b-415a-b1c2-728997286a8b"
data (energy)
attach (energy)
str(energy)
# + colab={"base_uri": "https://localhost:8080/", "height": 153} id="laIFCDfbfIzj" outputId="11ae8eb5-ff3b-4f7b-b877-1053cf577e24"
# Отвергаем нулевую гипотезу о равенстве средних
wilcox.test (expend ~ stature, paired = FALSE)
# + [markdown] id="U4j4QUNXfmDr"
# ### Сравнение двух независимых выборок
# + id="altivSGyftl7"
# wilcox.test(x, y, paired = TRUE )
# + [markdown] id="5bq7eKlOhrAh"
# ## Гипотеза об однородности дисперсии
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="ZHm5MYmQhx_b" outputId="48458747-76b6-4b9e-b771-4a0f6438aa12"
# F-критер<NAME>
data(energy, package="ISwR")
attach (energy)
colnames(energy)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="h2_jBAWLGrgS" outputId="6d19347d-5a9d-4f6f-a366-757a18cd88b4"
var.test(expend ~ stature)
| google_colab_statistics2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: literature
# language: python
# name: literature
# ---
# **Packages**
# +
import re
import numpy as np
import spacy
# -
import gensim
import gensim.corpora as corpora
from gensim.utils import simple_preprocess
from gensim.models import CoherenceModel
# +
import pyLDAvis
import pyLDAvis.gensim
import matplotlib.pyplot as plt
# %matplotlib inline
from nltk.corpus import stopwords
# +
import imp
tools = imp.load_source('tools', '../lda_tools.py')
# -
import tqdm
# **The data**
import pandas as pd
df = pd.read_csv('../data/prisoners_dilemma_articles_meta_data_clean.csv')
data = df[['abstract', 'unique_key', 'title', 'date']]
data = data.drop_duplicates()
data = data.reset_index(drop=True)
# We want to perform the analysis yearly. We will write a function to perform the analysis but also clean the data
# and create the corpus each time.
stop_words = stopwords.words('english')
mallet_path = '/Users/storm/rsc/mallet-2.0.8/bin/mallet'
# +
def get_model_and_opt_num_of_topics(yearly_dataset, limit, step=1, start=2):
# clean
words = list(tools.sentences_to_words(yearly_data['abstract'].values))
# bigram = gensim.models.Phrases(words, min_count=5, threshold=100)
# bigram_mod = gensim.models.phrases.Phraser(bigram)
lemmatized_words = tools.clean_words(words, stop_words)
# corpus
id2word = corpora.Dictionary(lemmatized_words)
texts = lemmatized_words
corpus = [id2word.doc2bow(text) for text in texts]
model_list, coherence_values = tools.compute_coherence_values(limit=limit,
mallet_path=mallet_path,
dictionary=id2word,
corpus=corpus,
texts=lemmatized_words,
step=step, start=start)
max_coherence_value = max(coherence_values)
max_index = coherence_values.index(max_coherence_value)
# num_of_topics = list(range(start, limit, step))
return model_list, max_index, coherence_values
# -
years = sorted(data.date.unique())
len(years)
results = []
for year in tqdm.tqdm_notebook(years[:5]):
yearly_data = data[data['date'] <= year]
result = get_model_and_opt_num_of_topics(yearly_data, limit=15, start=3)
results.append(result)
len(yearly_data['title'].unique())
len(df['title'].unique())
num_of_topics = list(range(2, 7, 1))
models, max_indices, coh_val = zip(*results)
coh_val[1]
plt.plot([num_of_topics[index] for index in max_indices], '-o');
| src/nbs/LDA Yearly analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import data analysis packages
import numpy as np
import pandas as pd
# import sklearn packages
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import LinearRegression
import pickle
# -
batting = pd.read_csv('data/baseballdatabank-master/core/batting.csv')
batting = batting[batting['yearID'] >= 2010]
batting['OBP'] = (batting['H'] + batting['BB'] + batting['HBP'])/(batting['AB'] + batting['BB'] + batting['HBP'] + batting['SF'])
batting = batting.drop(columns=['playerID','stint','teamID','lgID','CS','G','yearID'])
batting = batting.dropna()
batting
| Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import Data
import Models
from Models import nGrams as n
# ## Import Data
poems = Data.PoetryFoundationPoems()
poems = Data.Poem_words_dict(poems)
poem_syllables = Data.PoemOfSyllables(poems)
# ## Ngram model for both Syllables and Words
a = Models.nGrams()
# +
WordsTri = {}
for i in poems:
WordsTri = a.NGram(WordsTri, i,3)
Words4 = {}
for i in poems:
Words4 = a.NGram(Words4, i, 4)
# +
Syllables3 = {}
for i in poem_syllables:
Syllables3 = a.NGram(Syllables3, i, 3)
Syllables4 = {}
for i in poem_syllables:
Syllables4 = a.NGram(Syllables4, i, 4)
# -
#
words_by_syllables = Data.words_by_syllables()
n.next_word_prob(["ALONG","THAT"], WordsTri, 3, 0)
WordsTri["ALONG"]["THAT"]
n.next_word_prob([2,1], Syllables3, 3, 0)
Syllables3[2][1]
from Models import nGrams as nn
def nextSyllabeWord(words, words_nGram, syllable_nGram, syllable_dic, n):
syllables = [syllable_dic[word] for word in words]
wordsDic = words_nGram
syllaDic = syllable_nGram
nn.next_word_prob(words, wordsDic, n, 0)
nn.next_word_prob(syllables, syllaDic, 0.5)
for word in words:
wordsDic = wordsDic[word]
for syllable in syllables:
syllaDic = syllaDic[syllable]
Max = 0
next_word = "Error"
#print(words)
for word,probablity_word in list(wordsDic.items()):
final_prob = 0
if(word in syllable_dic):
word_syllable_count = syllable_dic[word]
final_prob = probablity_word * syllaDic[word_syllable_count]
#print(word,probablity_word,syllaDic[word_syllable_count], final_prob)
if(final_prob>Max):
Max = final_prob
next_word = word
return(next_word)
nextSyllabeWord(["ALONG","THAT"], WordsTri, Syllables3,Data.SyllablesDict(),3 )
prev_word1 = "WHY"#words_by_syllables[2][random.randint(0,len(words_by_syllables[3]))]
temp = list(WordsTri[prev_word1].keys())
prev_word2 = "DREAM"#temp[random.randint(0,len(temp)-1)]
generate_str = prev_word1+" "+prev_word2
generate_str
for i in range(50):
next_word = nextSyllabeWord([prev_word1, prev_word2], WordsTri, Syllables3,Data.SyllablesDict(),3)
prev_word1 = prev_word2
prev_word2 = next_word
generate_str = generate_str+" "+ next_word
print(generate_str.replace("NEWLINE ","\n"))
import random
# +
prev_word1 = "I"#words_by_syllables[2][random.randint(0,len(words_by_syllables[3]))]
temp = list(Words4[prev_word1].keys())
prev_word2 = "DO"#temp[random.randint(0,len(temp)-1)]
temp2 = list(Words4[prev_word1][prev_word2].keys())
prev_word3 = "NOT"#temp2[random.randint(0,len(temp2)-1)]
generate_str = prev_word1+" "+prev_word2 + " " +prev_word3
#generate_str
for i in range(50):
#print(generate_str)
next_words = nextSyllabeWord([prev_word1, prev_word2, prev_word3], Words4, Syllables4,Data.SyllablesDict(),4)
prev_word1 = prev_word2
prev_word2 = prev_word3
prev_word3 = next_words
generate_str = generate_str+" "+ next_words
print(generate_str.replace("NEWLINE ","\n"))
# +
prev_word1 = "I"#words_by_syllables[2][random.randint(0,len(words_by_syllables[3]))]
temp = list(Words4[prev_word1].keys())
prev_word2 = "DO"#temp[random.randint(0,len(temp)-1)]
temp2 = list(Words4[prev_word1][prev_word2].keys())
prev_word3 = "NOT"#temp2[random.randint(0,len(temp2)-1)]
generate_str = prev_word1+" "+prev_word2 + " " +prev_word3
for i in range(50):
#print(generate_str)
next_words = n.next_word([prev_word1, prev_word2, prev_word3], Words4,4)
prev_word1 = prev_word2
prev_word2 = prev_word3
prev_word3 = next_words
generate_str = generate_str+" "+ next_words
print(generate_str.replace("NEWLINE ","\n"))
# -
prev_word1 = "S"#words_by_syllables[2][random.randint(0,len(words_by_syllables[3]))]
temp = list(Words4[prev_word1].keys())
prev_word2 = temp[random.randint(0,len(temp)-1)]
temp2 = list(Words4[prev_word1][prev_word2].keys())
prev_word3 = temp2[random.randint(0,len(temp2)-1)]
generate_str = prev_word1+" "+prev_word2 + " " +prev_word3
generate_str
import random
Words4["I"]["DO"]["NOT"]
| .ipynb_checkpoints/Adding Syllables to Ngram Words-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sklearn
import matplotlib.pyplot as plt
import sklearn.datasets
train_X, train_Y = sklearn.datasets.make_moons(n_samples=300, noise=.2)
plt.scatter(train_X[:, 0], train_X[:, 1], c=train_Y, s=40, cmap=plt.cm.Spectral);
train_X = train_X.T
train_Y = train_Y.reshape((1, train_Y.shape[0]))
# +
from typing import Iterator
class Layer:
def __init__(self, ID: int):
self.ID = ID
def __repr__(self):
return f'Layer(ID={self.ID})'
class SomeModel:
def __init__(self, *args):
self.layers = [layer for layer in args]
self.index = 0
def __iter__(self) -> Iterator:
return self
def __next__(self) -> Layer:
if self.index < len(self.layers):
layer = self.layers[self.index]
self.index += 1
return layer
else:
raise StopIteration
# -
from typing import Protocol
class NameSayer(Protocol):
first = 'John'
last = 'Doe'
def say(self) -> str:
raise NotImplementedError
cls = SomeBaseClass()
class Person(NameSayer):
def say(self) -> str:
return f'{self.first} {self.last}'
def change(self, first: str, last: str) -> None:
self.first = first
self.last = last
cls = Person()
def make_say_name(sayer: NameSayer) -> None:
print(sayer.say())
make_say_name(cls)
cls.change(first='Johny', last='Appleseed')
make_say_name(cls)
| TestBed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ptlesson] *
# language: python
# name: conda-env-ptlesson-py
# ---
# +
# 분류용 가상 데이터 생성
# https://datascienceschool.net/view-notebook/ec26c797cec646e295d737c522733b15/
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import make_classification
from matplotlib import rc
import warnings
warnings.filterwarnings(action="ignore")
rc('font', family="AppleGothic")
# %matplotlib inline
plt.title("1개의 독립변수를 가진 가상 데이터")
X, y = make_classification(
n_features=1, # 독립변수의 수
n_informative=1, # 종속변수와 상관관계가 있는 성분의 수
n_redundant=0, # 다른 독립 변수의 선형 조합으로 나타나는 성분의 수
n_clusters_per_class=1, # 클래스 당 클러스터의 수
random_state=4,
#n_samples=100, # 표본 데이터의 수
#n_repeated=0, # 단순 중복된 성분의 독립변수의 수
#n_classes=2 # 종속변수의 클래수 수
#weights #각 클래스에 할당된 표본 수
)
plt.scatter(X, y, marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlabel("$X$")
plt.ylabel("$y$")
plt.show()
# -
sns.distplot(X[y==0], label="y=0")
sns.distplot(X[y==1], label="y=1")
plt.legend()
plt.show()
# +
plt.title("하나의 독립변수만 클래스와 상관관계가 있는 가상 데이터")
X, y = make_classification(n_features=2, n_informative=1, n_redundant=0,
n_clusters_per_class=1, random_state=4)
plt.scatter(X[:, 0], X[:, 1], marker='o', c=y,
s=100, edgecolor="k", linewidth=2)
plt.xlabel("$X_1$")
plt.ylabel("$X_2$")
plt.show()
# -
plt.subplot(121)
sns.distplot(X[y==0,0], label="y=0")
sns.distplot(X[y==1,0], label="y=1")
plt.legend()
plt.xlabel("$x_1$")
plt.subplot(122)
sns.distplot(X[y==0,1], label="y=0")
sns.distplot(X[y==1,1], label="y=1")
plt.legend()
plt.xlabel("$x_2$")
plt.show()
plt.figure(figsize=(8,8))
plt.title("두개의 독립변수 모두 클래스와 상관관계가 있는 가상데이터")
X, y = make_classification(n_samples=500, n_features=2, n_informative=2, n_redundant=0,
n_clusters_per_class=1, random_state=6)
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlim(-4,4)
plt.ylim(-4,4)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
plt.subplot(121)
sns.distplot(X[y==0, 0], label="y=0")
sns.distplot(X[y==1, 1], label="y=1")
plt.legend()
plt.xlabel("$x_1$")
plt.subplot(122)
sns.distplot(X[y==0, 1], label="y=0")
sns.distplot(X[y==1, 1], label="y=1")
plt.legend()
plt.xlabel("$x_2$")
plt.show()
l, V = np.linalg.eig(X.T @ X)
X2 = -X @ V
plt.figure(figsize=(8,8))
plt.scatter(X2[:,0], X2[:,1], marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlim(-4,4)
plt.ylim(-4,4)
plt.xlabel("$x'_1$")
plt.ylabel("$x'_2$")
plt.show()
# PCA
from sklearn.decomposition import PCA
pca = PCA(n_components=1)
X_pca = pca.fit_transform(X)
sns.distplot(X_pca[y==0], label="y=0")
sns.distplot(X_pca[y==1], label="y=1")
plt.legend()
plt.xlabel("x'")
plt.xlim(-3,5)
plt.show()
# 클래스별 데이터 갯수를 달리 줄때: weights
plt.title("비대칭 데이터")
X, y = make_classification(n_features=2, n_informative=2, n_redundant=0,
n_clusters_per_class=1, weights=[0.9,0.1], random_state=6)
val, cnt = np.unique(y, return_counts=True)
print("각 클래스별 데이터의 갯수 - {}클래스: {} / {}클래스: {}".format(val[0],cnt[0],val[1],cnt[1]))
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
# 클래스 당 여러개의 클래스: n_clusters_per_class
# (n_classes) x (n_clusters_per_class) <= 2^(n_informative)
plt.title("클래스 내에 2개의 클러스터가 있는 가상 데이터")
X2, Y2 = make_classification(n_samples=400, n_features=2, n_informative=2, n_redundant=0,
n_clusters_per_class=2, random_state=0)
plt.scatter(X2[:,0], X2[:,1], marker='o', c=Y2, s=100, edgecolor="k", linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
# 다중 클래스: n_classes
plt.title("다중 클래스를 가진 가상 데이터")
X, y = make_classification(n_samples=300, n_features=2, n_informative=2, n_redundant=0,
n_clusters_per_class=1, n_classes=3, random_state=0)
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
# 등방성 가우시안 분포: make_blobs
from sklearn.datasets import make_blobs
plt.title("세개의 클러스터를 가진 가상 데이터")
X, y = make_blobs(n_samples=300, n_features=2, centers=3, random_state=1,)
#cluster_std=1.0, center_box=(-10.,10.))
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor="k", linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
# 초승달 모양 클러스터: make_moons
from sklearn.datasets import make_moons
plt.title("초승달 모양 클러스터를 가진 가상 데이터")
X, y = make_moons(n_samples=400, noise=0.1, random_state=0)
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor='k', linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
# 다차원 가우시안 분포: make_gaussian_quantiles
from sklearn.datasets import make_gaussian_quantiles
plt.title("등고선으로 구분되는 두 개의 클러스터를 가진 가상 데이터")
X, y = make_gaussian_quantiles(n_samples=400, n_features=2, n_classes=2, random_state=0)
plt.scatter(X[:,0], X[:,1], marker='o', c=y, s=100, edgecolor='k', linewidth=2)
plt.xlabel("$x_1$")
plt.ylabel("$x_2$")
plt.show()
| P4_Supervised_Learning/01_dataset_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function
import tensorflow as tf
from tensorflow import keras
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# +
NUM_WORDS = 10000
(train_data, train_labels), (test_data, test_labels) = keras.datasets.imdb.load_data(num_words=NUM_WORDS)
# -
def multi_hot_sequences(sequences, dimension):
# Create an all-zero matrix of shape (len(sequences), dimension)
results = np.zeros((len(sequences), dimension))
for i, word_indices in enumerate(sequences):
results[i, word_indices] = 1.0 # set specific indices of results[i] to 1s
return results
train_data = multi_hot_sequences(train_data, dimension=NUM_WORDS)
test_data = multi_hot_sequences(test_data, dimension=NUM_WORDS)
plt.plot(train_data[0])
train_data[0]
# +
baseline_model = keras.Sequential([
# `input_shape` is only required here so that `.summary` works.
keras.layers.Dense(16, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
baseline_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
baseline_model.summary()
# -
baseline_history = baseline_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
# +
smaller_model = keras.Sequential([
keras.layers.Dense(4, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(4, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
smaller_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
smaller_model.summary()
# -
smaller_history = smaller_model.fit(train_data,
train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
# +
bigger_model = keras.models.Sequential([
keras.layers.Dense(512, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(512, activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
bigger_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
bigger_model.summary()
# -
bigger_history = bigger_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
# +
def plot_history(histories, key='binary_crossentropy'):
plt.figure(figsize=(16,10))
for name, history in histories:
val = plt.plot(history.epoch, history.history['val_'+key],
'--', label=name.title()+' Val')
plt.plot(history.epoch, history.history[key], color=val[0].get_color(),
label=name.title()+' Train')
plt.xlabel('Epochs')
plt.ylabel(key.replace('_',' ').title())
plt.legend()
plt.xlim([0,max(history.epoch)])
plot_history([('baseline', baseline_history),
('smaller', smaller_history),
('bigger', bigger_history)])
# +
l2_model = keras.models.Sequential([
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dense(16, kernel_regularizer=keras.regularizers.l2(0.001),
activation=tf.nn.relu),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
l2_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_crossentropy'])
l2_model_history = l2_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
# -
plot_history([('baseline', baseline_history),
('l2', l2_model_history)])
# +
dpt_model = keras.models.Sequential([
keras.layers.Dense(16, activation=tf.nn.relu, input_shape=(NUM_WORDS,)),
keras.layers.Dropout(0.5),
keras.layers.Dense(16, activation=tf.nn.relu),
keras.layers.Dropout(0.5),
keras.layers.Dense(1, activation=tf.nn.sigmoid)
])
dpt_model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy','binary_crossentropy'])
dpt_model_history = dpt_model.fit(train_data, train_labels,
epochs=20,
batch_size=512,
validation_data=(test_data, test_labels),
verbose=2)
# -
plot_history([('baseline', baseline_history),
('dropout', dpt_model_history)])
| Tensorflow/Explore overfitting and underfitting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from operator import itemgetter
import networkx as nx
import matplotlib.pyplot as plt
# %matplotlib inline
G = nx.generators.barabasi_albert_graph(1000, 2)
node_and_degree = dict(G.degree())
largest_hub, degree = sorted(node_and_degree.items(), key=itemgetter(1))[-1]
hub_ego = nx.ego_graph(G, largest_hub)
pos = nx.spring_layout(hub_ego)
nx.draw(hub_ego,pos,node_color='b', node_size=50, with_labels=False)
nx.draw_networkx_nodes(hub_ego, pos, nodelist=[largest_hub], node_size=300, node_color='r')
| jupyter_notebooks/D4/supplementaries/notebooks/drawing/ego_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={}
import os, glob, re, pickle
from functools import partial
from collections import OrderedDict
from cytoolz import compose
import operator as op
import pandas as pd
import seaborn as sns
import numpy as np
import scanpy as sc
import anndata as ad
import matplotlib.pyplot as plt
from pyscenic.export import export2loom, add_scenic_metadata
from pyscenic.utils import load_motifs
from pyscenic.transform import df2regulons
from pyscenic.aucell import aucell
from pyscenic.rss import regulon_specificity_scores
from pyscenic.plotting import plot_rss
import matplotlib.pyplot as plt
import matplotlib as mpl
from adjustText import adjust_text
import seaborn as sns
from pyscenic.binarization import binarize
# + pycharm={}
# Helper function
def derive_regulons(folder):
# Load enriched motifs.
motifs = load_motifs(folder+'motifs.csv')
motifs.columns = motifs.columns.droplevel(0)
def contains(*elems):
def f(context):
return any(elem in context for elem in elems)
return f
# For the creation of regulons we only keep the 10-species databases and the activating modules. We also remove the
# enriched motifs for the modules that were created using the method 'weight>50.0%' (because these modules are not part
# of the default settings of modules_from_adjacencies anymore.
motifs = motifs[
np.fromiter(map(compose(op.not_, contains('weight>50.0%')), motifs.Context), dtype=np.bool) & \
np.fromiter(map(contains('mm9-tss-centered-10kb-10species.mc9nr',
'mm9-500bp-upstream-10species.mc9nr',
'mm9-tss-centered-5kb-10species.mc9nr'), motifs.Context), dtype=np.bool) & \
np.fromiter(map(contains('activating'), motifs.Context), dtype=np.bool)]
# We build regulons only using enriched motifs with a NES of 3.0 or higher; we take only directly annotated TFs or TF annotated
# for an orthologous gene into account; and we only keep regulons with at least 10 genes.
regulons = list(filter(lambda r: len(r) >= 10, df2regulons(motifs[(motifs['NES'] >= 3.0)
& ((motifs['Annotation'] == 'gene is directly annotated')
| (motifs['Annotation'].str.startswith('gene is orthologous to')
& motifs['Annotation'].str.endswith('which is directly annotated for motif')))
])))
# Rename regulons, i.e. remove suffix.
regulons = list(map(lambda r: r.rename(r.transcription_factor), regulons))
# Pickle these regulons.
with open(folder+'regulons.dat', 'wb') as f:
pickle.dump(regulons, f)
# -
# # SCENIC
# ### Load the expression matrix
SC_EXP_FNAME = "Data/Cells/CD4/cd4.matrix.csv"
ex_matrix = pd.read_csv(SC_EXP_FNAME, sep=' ', header=0, index_col=0).T
ex_matrix.index = ex_matrix.index.str.replace('.', '-')
ex_matrix
# + [markdown] pycharm={}
# __METADATA CLEANING__
# -
BASE_FOLDER = "Data/Cells/CD4/"
df_cluster = pd.read_csv(BASE_FOLDER+"cd4.clusters.csv")
df_tumor = pd.read_csv(BASE_FOLDER+"cd4.tumor.csv")
df_cluster['x'] = pd.Series(df_cluster['x'], dtype="category")
df_metadata = df_cluster.merge(df_tumor, on="Unnamed: 0")
df_metadata.columns = ["Cell ID", "Cluster", "Tumor"]
df_metadata = df_metadata.set_index("Cell ID")
df_metadata.to_csv(BASE_FOLDER+"SCENIC/cd4.metadata.csv", index=False)
df_metadata
# +
# Regulons
BASE_FOLDER = "Data/Cells/CD4/SCENIC/"
derive_regulons(BASE_FOLDER)
with open(BASE_FOLDER+'regulons.dat', 'rb') as f:
regulons = pickle.load(f)
# -
BASE_FOLDER = "Data/Cells/CD4/SCENIC/"
REGULONS_DAT_FNAME = os.path.join(BASE_FOLDER, 'regulons.dat')
with open(REGULONS_DAT_FNAME, 'rb') as f:
regulons = pickle.load(f)
regulon_df = []
for i,regulon in enumerate(regulons):
gene2weight = pd.Series(dict(regulon.gene2weight)).sort_values(ascending=False).round(2)
regulon_df.append([regulon.name, len(regulon.genes), regulon.score, list(zip(gene2weight.index, gene2weight))])
regulon_df = pd.DataFrame(regulon_df, columns=["Regulon", "Size", "Score", "Genes"]).sort_values(by="Score", ascending=False).set_index('Regulon')
regulon_df.to_excel(BASE_FOLDER+"regulon_df.xlsx")
regulon_df
# Calculate AUC Matrix
auc_mtx = aucell(ex_matrix, regulons, seed=42)
auc_mtx.to_csv(BASE_FOLDER+"auc.csv")
auc_mtx
# +
# # %%capture
# from tqdm import tqdm_notebook as tqdm
# tqdm().pandas()
# for start in tqdm(range(0, len(auc_mtx.columns))):
# print(auc_mtx.iloc[:,start:start+1].columns)
# binary_mtx, auc_thresholds = binarize(auc_mtx.iloc[:,start:start+1], seed=42, num_workers=1)
# +
# auc_mtx['Klf2'].to_csv(BASE_FOLDER+"error_column.txt")
# auc_mtx = auc_mtx.drop('Klf2', axis=1)
# -
# Calculate Binary Matrix
binary_mtx, auc_thresholds = binarize(auc_mtx, seed=42, num_workers=1)
binary_mtx.to_csv(BASE_FOLDER+"binary.csv")
binary_mtx
| Code/CD4SCENICAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jupyter Markdown
# ### hslksadslkhdashj
# In this lesson, we start the module discussing Markdown to create written reports with embedded working code. Markdown is an authoring format that enables easy creation of dynamic documents, presentations, and reports. It combines the core syntax of markdown (an easy-to-write plain text format) with embedded python code chunks that are run so their output can be included in the final document. Markdown documents can be regenerated whenever underlying code or data changes.
#
#
# ## Loading python libraries
# +
# %matplotlib inline
# # %matplotlib inline is a magic function in IPython that displays images in the notebook
# Line magics are prefixed with the % character and work much like OS command-line calls
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import stats
import seaborn as sns
# Make plots larger
plt.rcParams['figure.figsize'] = (10, 6)
# -
# Markdown allows one to run code and graphs in the notebook.
# Plot two normal distributions
domain = np.arange(-22, 22, 0.1)
values = stats.norm(3.3, 5.5).pdf(domain)
plt.plot(domain, values, color='r', linewidth=2)
plt.fill_between(domain, 0, values, color='#ffb6c1', alpha=0.3)
values = stats.norm(4.4, 2.3).pdf(domain)
plt.plot(domain, values, color='b', linewidth=2)
plt.ylabel("Probability")
plt.title("Two Normal Distributions")
plt.show()
# ## Loading data
iris = pd.read_csv("http://nikbearbrown.com/YouTube/MachineLearning/DATA/iris.csv", sep=',')
iris.head()
# Let's do something with the iris data!
iris.describe()
# ## Syntax
#
# Plain text
#
# End a line with two spaces
# to start a new paragraph.
#
# ```
# Emphasis, aka italics, with *asterisks* or _underscores_.
#
# > Blockquote
#
# *italics* and _italics_
#
# **bold** and __bold__
#
# Strong emphasis, aka bold, with **asterisks** or __underscores__.
#
# Combined emphasis with **asterisks and _underscores_**.
#
# superscript^2^
#
# Strikethrough uses two tildes. ~~Scratch this.~~
#
# ~~strikethrough~~
#
# ```
#
# Emphasis, aka italics, with *asterisks* or _underscores_.
#
# > Blockquote
#
# *italics* and _italics_
#
# **bold** and __bold__
#
# Strong emphasis, aka bold, with **asterisks** or __underscores__.
#
# Combined emphasis with **asterisks and _underscores_**.
#
# superscript^2^
#
# Strikethrough uses two tildes. ~~Scratch this.~~
#
# ~~strikethrough~~
#
#
#
# ```
# Inline `code` has `back-ticks around` it.
#
# [link](www.rstudio.com)
#
# # Header 1
#
# ## Header 2
#
# ### Header 3
#
# #### Header 4
#
# ##### Header 5
#
# ###### Header 6
#
# endash: --
# emdash: ---
# ellipsis: ...
#
# inline equation: $A = \pi*r^{2}$
#
# horizontal rule (or slide break):
#
# ***
#
# > block quote
#
# * unordered list
#
# * item 2
# + sub-item 1
# + sub-item 2
#
# 1. ordered list
# 2. item 2
# + sub-item 1
# + sub-item 2
#
# Simple dot points:
#
# * Point 1
# * Point 2
# * Point 3
#
# and numeric dot points:
#
# 1. Number 1
# 2. Number 2
# 3. Number 3
#
# and nested dot points:
#
# * A
# * A.1
# * A.2
# * B
# * B.1
# * B.2
#
#
# Table Header | Second Header
# ------------- | -------------
# Table Cell | Cell 2
# Cell 3 | Cell 4
#
#
# ```
#
# Inline `code` has `back-ticks around` it.
#
# [link](www.rstudio.com)
#
# # Header 1
#
# ## Header 2
#
# ### Header 3
#
# #### Header 4
#
# ##### Header 5
#
# ###### Header 6
#
# endash: --
# emdash: ---
# ellipsis: ...
#
# inline equation: $A = \pi*r^{2}$
#
# horizontal rule (or slide break):
#
# ***
#
# > block quote
#
# * unordered list
#
# * item 2
# + sub-item 1
# + sub-item 2
#
# 1. ordered list
# 2. item 2
# + sub-item 1
# + sub-item 2
#
# Simple dot points:
#
# * Point 1
# * Point 2
# * Point 3
#
# and numeric dot points:
#
# 1. Number 1
# 2. Number 2
# 3. Number 3
#
# and nested dot points:
#
# * A
# * A.1
# * A.2
# * B
# * B.1
# * B.2
#
# Table Header | Second Header
# ------------- | -------------
# Table Cell | Cell 2
# Cell 3 | Cell 4
#
#
# ## Plots
# Images can be saved to a folder and/or plotted inline.
#
# ### Simple plot
# Here is a basic plot using seaborn graphics:
#
sns.pairplot(iris, hue="Species")
# ## Equations
#
# Multiple syntaxes exist:
#
# dollars: $inline$ and $$display$$. (same as LaTeX)
# single backslash: \(inline\) and \[display\] (same as LaTeX)
# double backslash: \\(inline\\) and \\[display\\]
# naked latex environment \begin{foo}...\end{foo}.
#
#
# Equations are included by using LaTeX notation and including them either between single dollar signs (inline equations) or double dollar signs (displayed equations).
# If you hang around the Q&A site [CrossValidated](http://stats.stackexchange.com) you'll be familiar with this idea.
#
# There are inline equations such as $y_i = \alpha + \beta x_i + e_i$.
#
# And displayed formulas:
#
# $$\frac{1}{1+\exp(-x)}$$
#
# # Typesetting Equations
#
# ## Inline vs. Display Material
#
# Equations can be formatted *inline* or as *displayed formulas*. In the latter case, they are centered and set off from the main text. In the former case, the mathematical material occurs smoothly in the line of text.
#
# In order to fit neatly in a line, summation expressions (and similar constructs) are formatted slightly differently in their inline and display versions.
#
# Inline mathematical material is set off by the use of single dollar-sign characters. Consequently, if you wish to use a dollar sign (for example, to indicate currency), you need to preface it with a back-slash. The following examples, followed by their typeset versions, should make this clear
#
# ```
# This summation expression $\sum_{i=1}^n X_i$ appears inline.
# ```
#
# This summation expression $\sum_{i=1}^n X_i$ appears inline.
#
# ```
# This summation expression is in display form.
#
# $$\sum_{i=1}^n X_i$$
# ```
# This summation expression is in display form.
#
# $$\sum_{i=1}^n X_i$$
#
# ## Some LaTeX Basics
#
# In this section, we show you some rudiments of the LaTeX typesetting language.
#
# ### Subscripts and Superscripts
# To indicate a subscript, use the underscore `_` character. To indicate a superscript, use a single caret character `^`. Note: this can be confusing, because the R Markdown language delimits superscripts with two carets. In LaTeX equations, a single caret indicates the superscript.
#
# If the subscript or superscript has just one character, there is no need to delimit with braces. However, if there is more than one character, braces must be used.
#
# The following examples illustrate:
#
# ```
# $$X_i$$
# $$X_{i}$$
# ```
# $$X_i$$
# $$X_{i}$$
#
# Notice that in the above case, braces were not actually needed.
#
# In this next example, however, failure to use braces creates an error, as LaTeX sets only the first character as a subscript
#
# ```
# $$X_{i,j}$$
# $$X_i,j$$
# ```
#
# $$X_{i,j}$$
# $$X_i,j$$
#
# Here is an expression that uses both subscripts and superscripts
#
# ```
# $$X^2_{i,j}$$
# ```
#
# $$X^2_{i,j}$$
#
# ### Square Roots
#
# We indicate a square root using the `\sqrt` operator.
#
# ```
# $$\sqrt{b^2 - 4ac}$$
# ```
#
# $$\sqrt{b^2 - 4ac}$$
#
# ### Fractions
#
# Displayed fractions are typeset using the `\frac` operator.
#
# ```
# $$\frac{4z^3}{16}$$
# ```
# $$\frac{4z^3}{16}$$
#
# ### Summation Expressions
#
# These are indicated with the `\sum' operator, followed by a subscript for the material appearing below the summation sign, and a superscript for any material appearing above the summation sign.
#
# Here is an example.
#
# ```
# $$\sum_{i=1}^{n} X^3_i$$
# ```
# $$\sum_{i=1}^{n} X^3_i$$
#
#
# ### Self-Sizing Parentheses
#
# In LaTeX, you can create parentheses, brackets, and braces which size themselves automatically to contain large expressions. You do this using the `\left` and `\right` operators. Here is an example
#
# ```
# $$\sum_{i=1}^{n}\left( \frac{X_i}{Y_i} \right)$$
# ```
#
# $$\sum_{i=1}^{n}\left( \frac{X_i}{Y_i} \right)$$
#
# ### Greek Letters
#
# Many statistical expressions use Greek letters. Much of the Greek alphabet is implemented in LaTeX, as indicated in the LaTeX cheat sheet available at the course website. There are both upper and lower case versions available for some letters.
#
# ```
# $$\alpha, \beta, \gamma, \Gamma$$
# ```
#
# $$\alpha, \beta, \gamma, \Gamma$$
#
#
#
# ### Special Symbols
#
# All common mathematical symbols are implemented, and you can find a listing on a LaTeX cheat sheet.
#
#
# ```
# * Subscripts to get \( a_{b} \) write: $a_{b}$
# * Superscripts write \( a^{b} \) write: $a^{b}$
# * Greek letters like \( \alpha, \beta, \ldots \) write: $\alpha, \beta, \ldots$
# * Sums like \( \sum_{n=1}^N \) write: $\sum_{n=1}^N$
# * Multiplication like \( \times \) write: $\times$
# * Products like \( \prod_{n=1}^N \) write: $\prod_{n=1}^N$
# * Inequalities like \( <, \leq, \geq \) write: $<, \leq, \geq$
# * Distributed like \( \sim \) write: $\sim$
# * Hats like \( \widehat{\alpha} \) write: $\widehat{\alpha}$
# * Averages like \( \bar{x} \) write: $\bar{x}$
# * Fractions like \( \frac{a}{b} \) write: $\frac{a}{b}$
#
# ```
#
# * Subscripts to get \( a_{b} \) write: $a_{b}$
# * Superscripts write \( a^{b} \) write: $a^{b}$
# * Greek letters like \( \alpha, \beta, \ldots \) write: $\alpha, \beta, \ldots$
# * Sums like \( \sum_{n=1}^N \) write: $\sum_{n=1}^N$
# * Multiplication like \( \times \) write: $\times$
# * Products like \( \prod_{n=1}^N \) write: $\prod_{n=1}^N$
# * Inequalities like \( <, \leq, \geq \) write: $<, \leq, \geq$
# * Distributed like \( \sim \) write: $\sim$
# * Hats like \( \widehat{\alpha} \) write: $\widehat{\alpha}$
# * Averages like \( \bar{x} \) write: $\bar{x}$
# * Fractions like \( \frac{a}{b} \) write: $\frac{a}{b}$
#
# Some examples. (Notice that, in the third example, I use the tilde character for a forced space. Generally LaTeX does spacing for you automatically, and unless you use the tilde character, R will ignore your attempts to add spaces.)
#
# ```
# $$a \pm b$$
# $$x \ge 15$$
# $$a_i \ge 0~~~\forall i$$
# ```
#
# $$a \pm b$$
# $$x \ge 15$$
# $$a_i \ge 0~~~\forall i$$
#
# ### Special Functions
#
# LaTeX typesets special functions in a different font from mathematical variables. These functions, such as $\sin$, $\cos$, etc. are indicated in LaTeX with a backslash. Here is an example that also illustrates how to typeset an integral.
#
# ```
# $$\int_0^{2\pi} \sin x~dx$$
# ```
# $$\int_0^{2\pi} \sin x~dx$$
#
# ### Matrices
#
# Matrics are presented in the `array` environment. One begins with the statement
# `\begin{array}` and ends with the statement `\end{array}`. Following the opening statement, a format code is used to indicate the formatting of each column. In the example below, we use the code `{rrr}` to indicate that each column is right justified. Each row is then entered, with cells separated by the `&` symbol, and each line (except the last) terminated by `\\`.
#
# ```
# $$\begin{array}
# {rrr}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9
# \end{array}
# $$
# ```
# $$\begin{array}
# {rrr}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9
# \end{array}
# $$
#
# In math textbooks, matrices are often surrounded by brackets, and are assigned to a boldface letter. Here is an example
#
# ```
# $$\mathbf{X} = \left[\begin{array}
# {rrr}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9
# \end{array}\right]
# $$
# ```
#
# $$\mathbf{X} = \left[\begin{array}
# {rrr}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9
# \end{array}\right]
# $$
#
#
# ### Tables
# Tables can be included using the following notation
#
# A | B | C
# --- | --- | ---
# 1 | Male | Blue
# 2 | Female | Pink
#
# ### Hyperlinks
#
# * Here is my blog nikbearbrown.com [my blog nikbearbrown.com](http://nikbearbrown.com).
#
# ### Images
#
# Here's some example images:
#
# 
#
#
# 
#
#
# <NAME>, known as "Student", British statistician. Picture taken in 1908
#
# <NAME> (1876 - 1937) was a chemist and statistician, better known by his pen name Student. He worked in a beer brewery and his testing of very small patches led him to discover certain small-sample distributions.This led to the development of Student's t-Test. His communications with Fisher on the subject are legendary.
#
# See
#
# * <NAME>osset [William Sealy Gosset] (https://en.wikipedia.org/wiki/William_Sealy_Gosset)
# * Famous Statisticians - Department of Statistics, GMU [Famous Statisticians - Department of Statistics, GMU] (http://statistics.gmu.edu/pages/famous.html)
#
#
# ### Quote
# Let's quote some stuff:
#
# > To be, or not to be, that is the question:
# > Whether 'tis nobler in the mind to suffer
# > The slings and arrows of outrageous fortune,
#
#
# ## Hotkeys
#
# There are a ton of useful hotkeys in Jupyter. Pressing "shift-m" puts you into **command mode**. Pushing another key then usually does something useful. For example,
# * `shift-m` then `a` inserts a new cell above this one
# * `shift-m` then `b` inserts a new cell below
# * Esc will take you into command mode where you can navigate around your notebook with arrow keys.
#
# While in command mode:
# * A to insert a new cell above the current cell, B to insert a new cell below.
# * M to change the current cell to Markdown, Y to change it back to code
# * D + D (press the key twice) to delete the current cell
# * Enter will take you from command mode back into edit mode for the given cell.
# * Shift = Tab will show you the Docstring (documentation) for the the object you have just typed in a code cell - you can keep pressing this short cut to cycle through a few modes of documentation.
# * Ctrl + Shift + - will split the current cell into two from where your cursor is.
# * Esc + F Find and replace on your code but not the outputs.
# * Esc + O Toggle cell output.
# * Select Multiple Cells:
# * Shift + J or Shift + Down selects the next sell in a downwards direction. You can also select sells in an upwards direction by using Shift + K or Shift + Up.
# * Once cells are selected, you can then delete / copy / cut / paste / run them as a batch. This is helpful when you need to move parts of a notebook.
# * You can also use Shift + M to merge multiple cells.
#
# You can find a list of hotkeys [here](https://www.dataquest.io/blog/jupyter-notebook-tips-tricks-shortcuts/).
# ## Conclusion
#
# * Markdown is awesome.
# * Analysis is much more readable and understandable when text, code, figures and equations can be shown together.
# * For journal articles, LaTeX will presumably still be required.
#
# Last update September 1, 2017
| Week_1/NBB_Jupyter_Markdown.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
df_minjust = pd.read_csv('data/Minjust2018.csv',
header=None,
usecols=[4, 27, 28, 33])
df_minjust.fillna('', inplace=True)
df_minjust.rename(columns={4: 'company_name',
27: 'director',
28: 'activity',
33: 'founders'},
inplace = True)
df_minjust
df_decl = pd.read_csv('data/declaration.csv',
header=None,
usecols=[1, 2, 3, 4, 5])
df_decl.fillna('', inplace=True)
df_decl.rename(columns={1: 'surname',
2: 'name',
3: 'patronymic',
4: 'workplace',
5: 'position'},
inplace = True)
df_decl
columns = ['company_name', 'director', 'activity', 'founders', 'surname', 'name', 'patronymic', 'workplace', 'position']
result = pd.DataFrame(columns=columns)
result1 = pd.DataFrame(columns=columns)
result2 = pd.DataFrame(columns=columns)
for index, row in df_decl.loc[: 10].iterrows():
search = search_df(df_minjust, 'founders', child_regex(row['surname'], row['name'])).copy()
for column in df_decl.columns:
search[column] = row[column]
result = result.append(search, ignore_index=True)
result
# +
def search_df(df, column, pattern):
return df.loc[df[column].str.contains(pattern)]
def child_regex(surname, name):
pattern = re.compile(surname +
r'а?\s+\w+\s+' + # First name
remove_vowels(name) + # Patronymic
r'|' +
r'\w+\s+' + # First name
remove_vowels(name) + r'\w+\s+' + # Patronymic
surname,
re.IGNORECASE)
return pattern
def remove_vowels(name):
while True:
if name[-1] not in ['а', 'о', 'и', 'е', 'ё', 'э', 'ы', 'у', 'ю', 'я', 'й']:
break
else:
name = name[: -1]
return name
# -
for index, row in df_decl.loc[355: 425].iterrows():
search = search_df(df_minjust, 'founders', child_regex(row['surname'], row['name'])).copy()
for column in df_decl.columns:
search[column] = row[column]
result1 = result1.append(search, ignore_index=True)
result1
# +
def brother_regex(surname, patronymic):
pattern = re.compile(surname +
r'\s+\w+\s+' + # First name
patronymic + # Patronymic
r'|' +
r'\w+\s+' +
patronymic + r'\s+' + # Patronymic
surname,
re.IGNORECASE)
return pattern
def count_letters(words):
return pd.Series([len(word) for word in words])
# -
for index, row in df_decl.loc[355: 425].iterrows():
search = search_df(df_minjust, 'director', brother_regex(row['surname'], row['patronymic'])).copy()
for column in df_decl.columns:
search[column] = row[column]
result2 = result2.append(search, ignore_index=True)
result2
result.to_pickle('result1.pkl')
result.to_pickle('result2.pkl')
| df_children_search_russian_lang_implemented-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.10 64-bit (''venv'': venv)'
# name: pythonjvsc74a57bd0a06f596174f42b43cb4a04750d8acb053248ec3389513c8276a84c804cff32f0
# ---
# # Cleaning the Titanic dataset
# As before we start by importing some libraries.
#
# We will use pandas again to handle our data.
#
# We will also import `matplotlib` and `seaborn`, these libraries are used to create some visualisations of our data.
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# -
titanic_df = pd.read_csv('datasets/titanic.csv')
titanic_df.shape
# ### Dataset description
#
# Here we have a description of the column headers of the CSV data.
#
# 1. PassengerId - Passenger unique Id
# 2. Survival - Survival (0 = No; 1 = Yes).
# 3. Pclass - Passenger Class (1 = 1st; 2 = 2nd; 3 = 3rd)
# 4. Name - Name
# 5. Sex - Sex
# 6. Age - Age
# 7. Sibsp - Number of Siblings/Spouses Aboard
# 8. Parch - Number of Parents/Children Aboard
# 9. Ticket - Ticket Number
# 10. Fare - Passenger Fare
# 11. Cabin - Cabin
# 12. Embarked - Port of Embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
#
#
# We will drop some of the columns that have no relevance to our model.
titanic_df.drop(['PassengerId', 'Name', 'Ticket', 'Cabin'], 'columns', inplace=True)
titanic_df.head(10)
titanic_df.shape
# When working wth large datasets in is quite common to have records with misssing data, we will check for any records that have missing fields.
#
# We can use the `.isnull()` function from the pandas library to check if a value is null.
#
# The following line will count how many null values there are in each column of our dataframe.
titanic_df[titanic_df.isnull().any(axis=1)].count()
# As we can see there are quite a few records with missing data. It is possible to use different techniques to predict what the missing data could be.
# For this example we will just omit any records with missing data.
titanic_df = titanic_df.dropna()
titanic_df.shape
# Lets just do a sanity check to see if there are any null values left in our dataframe.
titanic_df[titanic_df.isnull().any(axis=1)].count()
# We can use the `describe` method to get some statistical information about our dataset.
titanic_df.describe()
# Notice the mean for the 'Survived' column is 0.404494, this means only about 40% of the passengers in our data survived. (This is higher than the 31% of the total passenger survival rate)
#
#
# Let's see if we plot some of our data on a scatter plt can we see anything interesting.
# +
fig, ax = plt.subplots(figsize=(12, 8))
plt.scatter(titanic_df['Age'], titanic_df['Survived'])
plt.xlabel('Age')
plt.ylabel('Survived')
# -
# This shows us that there is very little to no correlation between the passenger's age and if the passenger survived
# +
fig, ax = plt.subplots(figsize=(12, 8))
plt.scatter(titanic_df['Fare'], titanic_df['Survived'])
plt.xlabel('Fare')
plt.ylabel('Survived')
# -
# We can see that there area few outliers of passengers that survived that paid much higher fares.
#
# Try to create a scatterplot for a different feature.
# +
# Write your code here
# -
# As we are looking at this in terms of binary classification a scatterplot is of very little use.
#
# Let's try looking at the data in a crosstab table(confusion matrix).
pd.crosstab(titanic_df['Sex'], titanic_df['Survived'])
# We can see from this that there was a much higher survival rate for women than men
pd.crosstab(titanic_df['Pclass'], titanic_df['Survived'])
# We can use one of pandas built in functions to view correlations between columns
# +
titanic_data_corr = titanic_df.corr()
titanic_data_corr
# -
# We can see from this that class is negatively correlated with survival, the lower the class the lower the chance of survival.
#
# Fare is positively correlated with survival, the higher the fare the higher the chance of survival.
#
# We can use the Seaborn library to show this correlation matrix as a heatmap.
# +
fig, ax = plt.subplots(figsize=(12,10))
sns.heatmap(titanic_data_corr, annot=True)
# -
# Now that we have cleaned our data up we can move onto to preproccessing, this is where we make sure the data is in a form that the model will understand so that it can be used for training.
#
# We'll save our cleaned data to disk for the next step.
titanic_df.to_csv('datasets/titanic_cleaned.csv', index=False)
| 1_classification/2_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# +
# %%capture
import os
import site
os.sys.path.insert(0, '/home/schirrmr/code/reversible/')
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
os.sys.path.insert(0, '/home/schirrmr/code/explaining/reversible//')
# %load_ext autoreload
# %autoreload 2
import numpy as np
import logging
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import cm
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
matplotlib.rcParams['figure.figsize'] = (12.0, 1.0)
matplotlib.rcParams['font.size'] = 14
import seaborn
seaborn.set_style('darkgrid')
from reversible2.sliced import sliced_from_samples
from numpy.random import RandomState
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
import copy
import math
import itertools
import torch as th
from braindecode.torch_ext.util import np_to_var, var_to_np
from reversible2.splitter import SubsampleSplitter
from reversible2.view_as import ViewAs
from reversible2.invert import invert
from reversible2.affine import AdditiveBlock
# -
from reversible2.bhno import create_inputs
# +
orig_train_cnt = load_file('/data/schirrmr/schirrmr/HGD-public/reduced/train/4.mat')
train_cnt = orig_train_cnt.reorder_channels(['C3', 'C4'])
train_inputs = create_inputs(train_cnt, final_hz=64, half_before=True)
# -
orig_test_cnt = load_file('/data/schirrmr/schirrmr/HGD-public/reduced/test/4.mat')
test_cnt = orig_test_cnt.reorder_channels(['C3', 'C4'])
test_inputs = create_inputs(test_cnt, final_hz=64, half_before=True)
cuda = True
if cuda:
train_inputs = [i.cuda() for i in train_inputs]
test_inputs = [i.cuda() for i in test_inputs]
# +
from reversible2.distribution import TwoClassDist
from reversible2.blocks import dense_add_block, conv_add_block_3x3
from reversible2.rfft import RFFT, Interleave
from reversible2.util import set_random_seeds
from torch.nn import ConstantPad2d
import torch as th
from reversible2.splitter import SubsampleSplitter
set_random_seeds(2019011641, cuda)
feature_model = nn.Sequential(
SubsampleSplitter(stride=[2,1],chunk_chans_first=False),# 2 x 32
conv_add_block_3x3(2,32),
conv_add_block_3x3(2,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 4 x 16
conv_add_block_3x3(4,32),
conv_add_block_3x3(4,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 8 x 8
conv_add_block_3x3(8,32),
conv_add_block_3x3(8,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 16 x 4
conv_add_block_3x3(16,32),
conv_add_block_3x3(16,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 32 x 2
conv_add_block_3x3(32,32),
conv_add_block_3x3(32,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 64 x 1
ViewAs((-1,64,1, 1), (-1,64)),
dense_add_block(64,64),
dense_add_block(64,64),
dense_add_block(64,64),
dense_add_block(64,64),
dense_add_block(64,64),
dense_add_block(64,64),
RFFT(),
)
if cuda:
feature_model.cuda()
device = list(feature_model.parameters())[0].device
from reversible2.ot_exact import ot_euclidean_loss_for_samples
class_dist = TwoClassDist()
if cuda:
class_dist.cuda()
# -
optim_model = th.optim.Adam(feature_model.parameters())
optim_dist = th.optim.Adam(class_dist.parameters(), lr=1e-2)
# +
# %%writefile plot.py
import torch as th
import matplotlib.pyplot as plt
import numpy as np
from reversible2.util import var_to_np
from reversible2.plot import display_close
from matplotlib.patches import Ellipse
import seaborn
def plot_outs(feature_model_a, train_inputs, test_inputs, class_dist):
# Compute dist for mean/std of encodings
data_cls_dists = []
for i_class in range(len(train_inputs)):
this_class_outs = feature_model_a(train_inputs[i_class])[:,:2]
data_cls_dists.append(
th.distributions.MultivariateNormal(th.mean(this_class_outs, dim=0),
covariance_matrix=th.diag(th.std(this_class_outs, dim=0))))
for setname, set_inputs in (("Train", train_inputs), ("Test", test_inputs)):
outs = [feature_model_a(ins) for ins in set_inputs]
c_outs = [o[:,:2] for o in outs]
c_outs_all = th.cat(c_outs)
cls_dists = []
for i_class in range(len(c_outs)):
mean, std = class_dist.get_mean_std(i_class)
cls_dists.append(
th.distributions.MultivariateNormal(mean[:2],covariance_matrix=th.diag(std[:2])))
preds = th.stack([cls_dists[i_cls].log_prob(c_outs_all)
for i_cls in range(len(cls_dists))],
dim=-1)
pred_labels = np.argmax(var_to_np(preds), axis=1)
labels = np.concatenate([np.ones(len(set_inputs[i_cls])) * i_cls
for i_cls in range(len(train_inputs))])
acc = np.mean(labels == pred_labels)
data_preds = th.stack([data_cls_dists[i_cls].log_prob(c_outs_all)
for i_cls in range(len(cls_dists))],
dim=-1)
data_pred_labels = np.argmax(var_to_np(data_preds), axis=1)
data_acc = np.mean(labels == data_pred_labels)
print("{:s} Accuracy: {:.2f}%".format(setname, acc * 100))
fig = plt.figure(figsize=(5,5))
ax = plt.gca()
for i_class in range(len(c_outs)):
o = var_to_np(c_outs[i_class]).squeeze()
plt.scatter(o[:,0], o[:,1], s=20, alpha=0.75)
means = var_to_np(class_dist.class_means).copy()
means[1-i_class] = 0
stds = var_to_np(th.exp(class_dist.class_log_stds))
stds[1-i_class] = 0.05
for sigma in [0.5,1,2,3]:
ellipse = Ellipse(means, stds[0]*sigma, stds[1]*sigma)
ax.add_artist(ellipse)
ellipse.set_edgecolor(seaborn.color_palette()[i_class])
ellipse.set_facecolor("None")
for i_class in range(len(c_outs)):
o = var_to_np(c_outs[i_class]).squeeze()
plt.scatter(np.mean(o[:,0]), np.mean(o[:,1]),
color=seaborn.color_palette()[i_class+2], s=80, marker="^")
plt.title("{:6s} Accuracy: {:.2f}%\n"
"From data mean/std: {:.2f}%".format(setname, acc * 100, data_acc * 100))
plt.legend(("Right", "Rest", "Right Mean", "Rest Mean"))
display_close(fig)
# -
from plot import plot_outs
n_epochs = 2001
for i_epoch in range(n_epochs):
optim_model.zero_grad()
optim_dist.zero_grad()
for i_class in range(len(train_inputs)):
class_ins = train_inputs[i_class]
samples = class_dist.get_samples(i_class, len(train_inputs[i_class]) * 5)
inverted = invert(feature_model, samples)
loss = ot_euclidean_loss_for_samples(class_ins.squeeze(), inverted.squeeze())
loss.backward()
optim_model.step()
optim_dist.step()
if i_epoch % (n_epochs // 20) == 0:
print("Epoch {:d} of {:d}".format(i_epoch, n_epochs))
print("Loss: {:E}".format(loss.item()))
plot_outs(feature_model, train_inputs, test_inputs,
class_dist)
fig = plt.figure(figsize=(8,2))
plt.plot(var_to_np(th.cat((th.exp(class_dist.class_log_stds),
th.exp(class_dist.non_class_log_stds)))),
marker='o')
display_close(fig)
for setname, set_inputs in (("Train", train_inputs), ("Test", test_inputs)):
total_loss = 0
total_spec_ot = 0
for i_class in range(2):
class_ins = set_inputs[i_class]
samples = class_dist.get_samples(i_class, len(train_inputs[i_class]) * 5)
inverted = invert(feature_model, samples)
loss = ot_euclidean_loss_for_samples(class_ins.squeeze(), inverted.squeeze())
total_loss += loss
spec_ot = ot_euclidean_loss_for_samples(
th.rfft(class_ins.squeeze(), signal_ndim=1, normalized=True).view(class_ins.shape[0],-1),
th.rfft(inverted.squeeze(), signal_ndim=1, normalized=True).view(inverted.shape[0],-1))
total_spec_ot += spec_ot
print(setname + " Loss: {:.1f}".format(total_loss.item() / 2))
print(setname + " SpecLoss: {:.1f}".format(spec_ot.item() / 2))
# Show losses between sets
for setname, set_inputs in (("Train", train_inputs),):
total_loss = 0
total_spec_ot = 0
for i_class in range(2):
class_ins = set_inputs[i_class]
samples = class_dist.get_samples(i_class, len(train_inputs[i_class]) * 5) # take same number of samples
inverted = test_inputs[i_class]
loss = ot_euclidean_loss_for_samples(class_ins.squeeze(), inverted.squeeze())
total_loss += loss
spec_ot = ot_euclidean_loss_for_samples(
th.rfft(class_ins.squeeze(), signal_ndim=1, normalized=True).view(class_ins.shape[0],-1),
th.rfft(inverted.squeeze(), signal_ndim=1, normalized=True).view(inverted.shape[0],-1))
total_spec_ot += spec_ot
print(setname + " Loss: {:.1f}".format(total_loss.item() / 2))
print(setname + " SpecLoss: {:.1f}".format(spec_ot.item() / 2))
def create_th_grid(dim_0_vals, dim_1_vals):
curves = []
for dim_0_val in dim_0_vals:
this_curves = []
for dim_1_val in dim_1_vals:
vals = th.stack((dim_0_val, dim_1_val))
this_curves.append(vals)
curves.append(th.stack(this_curves))
curves = th.stack(curves)
return curves
means = th.stack([class_dist.get_mean_std(i_cls,)[0][:2]
for i_cls in range(len(train_inputs))])
stds = th.stack([class_dist.get_mean_std(i_cls,)[1][:2]
for i_cls in range(len(train_inputs))])
mins = th.min(means - stds * 2, dim=1)[0]
maxs = th.max(means + stds * 2, dim=1)[0]
mean_for_plot = th.mean(th.stack((mins, maxs)), dim=0)
# +
#mins.data[0] = -0.2
# +
n_vals = 20
dim_0_vals = th.linspace(mins[0].item(), maxs[0].item(),n_vals, device=mins.device)
dim_1_vals = th.linspace(mins[1].item(), maxs[1].item(),n_vals, device=mins.device)
grid = create_th_grid(dim_0_vals, dim_1_vals)
full_grid = th.cat((grid, class_dist.non_class_means.repeat(grid.shape[0],grid.shape[1],1)), dim=-1)
inverted = invert(feature_model, full_grid.view(-1, full_grid.shape[-1])).squeeze()
inverted_grid = inverted.view(len(dim_0_vals), len(dim_1_vals),-1)
x_len, y_len = var_to_np(maxs - mins) / full_grid.shape[:-1]
max_abs = th.max(th.abs(inverted_grid))
y_factor = (y_len / (2*max_abs)).item() * 0.9
# +
plt.figure(figsize=(32,32))
for i_x in range(full_grid.shape[0]):
for i_y in range(full_grid.shape[1]):
x_start = mins[0].item() + x_len * i_x + 0.1 * x_len
x_end = mins[0].item() + x_len * i_x + 0.9 * x_len
y_center = mins[1].item() + y_len * i_y + 0.5 * y_len
curve = var_to_np(inverted_grid[i_x][i_y])
label = ''
if i_x == 0 and i_y == 0:
label = 'Generated data'
plt.plot(np.linspace(x_start, x_end, len(curve)),
curve * y_factor + y_center, color='black',
label=label)
outs = [feature_model(ins) for ins in train_inputs]
c_outs = [o[:,:2] for o in outs]
plt.gca().set_autoscale_on(False)
for i_cls, c_out in enumerate(c_outs):
plt.scatter(var_to_np(c_out[:,0]),
var_to_np(c_out[:,1]), s=200, alpha=0.75, label='Encodings {:s}'.format(
["Right", "Rest"][i_cls]))
plt.legend(fontsize=14)
# -
fig = plt.figure(figsize=(8,2))
for i_class in range(2):
cur_mean, cur_std = class_dist.get_mean_std(i_class, )
inverted = invert(feature_model, cur_mean.unsqueeze(0))
plt.plot(var_to_np(inverted.squeeze()))
plt.legend(("Right Hand", "Rest"))
display(fig)
plt.close(fig)
from plot import display_close
# +
mean_0, _ = class_dist.get_mean_std(0, )
mean_1, _ = class_dist.get_mean_std(1, )
n_interpolates = 100
alphas = th.linspace(1,0,n_interpolates, device=mean_0.device)
interpolates = mean_0.unsqueeze(0) * alphas.unsqueeze(1) + mean_1.unsqueeze(0) * (1-alphas.unsqueeze(1))
inverted = invert(feature_model,interpolates, )
from matplotlib import rcParams, cycler
cmap = plt.cm.coolwarm
with plt.rc_context(rc={'axes.prop_cycle':cycler(color=cmap(np.linspace(0, 1, len(inverted))))}):
fig = plt.figure(figsize=(8,2))
plt.plot(var_to_np(inverted).squeeze().T,);
display_close(fig)
| notebooks/bhno-classification/BhNo64.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Different orthotropic fitted closure approximations in simple shear flow.
#
# Model should reproduce Figure 6(a) in
# <NAME> and <NAME> (2001),
# 'Improved model of orthotropic closure approximation for flow induced fiber
# orientation', Polymer Composites, 22(5), 636-649, DOI: 10.1002/pc.10566
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
from fiberoripy.orientation import folgar_tucker_ode
# +
# geometric factor
xi = 1.0
# phenomenological fiber-fiber interaction coefficient
C_I = 0.01
# time steps
t = np.linspace(0, 30, 60)
# initial fiber orientation state
A0 = 1.0 / 3.0 * np.eye(3)
# -
# define a function that describes the (time-dependend) velocity gradient
def L(t):
"""Velocity gradient."""
return np.array([[0.0, 1.0, 0.0], [0.0, 0.0, 0.0], [0.0, 0.0, 0.0]])
# compute solutions by integrating the ODEs
a_ibof = odeint(
folgar_tucker_ode,
A0.ravel(),
t,
args=(xi, L, C_I, "HYBRID"),
)
a_orf = odeint(
folgar_tucker_ode,
A0.ravel(),
t,
args=(xi, L, C_I, "ORF"),
)
a_orw = odeint(
folgar_tucker_ode,
A0.ravel(),
t,
args=(xi, L, C_I, "ORW"),
)
a_orw3 = odeint(
folgar_tucker_ode,
A0.ravel(),
t,
args=(xi, L, C_I, "ORW3"),
)
# +
# plot components
plt.plot(t, a_ibof[:, 0], linestyle="-", label="$a_{11}$ Hybrid", color="b")
plt.plot(t, a_orf[:, 0], linestyle="-", label="$a_{11}$ ORF", color="r")
plt.plot(t, a_orw[:, 0], linestyle="-", label="$a_{11}$ ORW", color="g")
plt.plot(t, a_orw3[:, 0], linestyle="-.", label="$a_{11}$ ORW3", color="k")
plt.plot(t, a_ibof[:, 1], linestyle="--", label="$a_{12}$ Hybrid", color="b")
plt.plot(t, a_orf[:, 1], linestyle="--", label="$a_{12}$ ORF", color="r")
plt.plot(t, a_orw[:, 1], linestyle="--", label="$a_{12}$ ORW", color="g")
plt.plot(t, a_orw3[:, 1], linestyle="-.", label="$a_{12}$ ORW3", color="k")
# adjust some plot settings.
plt.ylim=([-0.2, 1])
plt.grid(b=True, which="major", linestyle="-")
plt.minorticks_on()
plt.grid(b=True, which="minor", linestyle="--", alpha=0.2)
plt.xlabel("Time $t$ in $s$")
plt.ylabel("$a_{11}, a_{12}$")
plt.legend(loc="center right")
plt.title(r"Simple shear flow, $\xi = 1$, $C_1 = 0,01$")
plt.show()
# -
| examples/orientation/orthofitclosures_perfectshear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
messages = pd.read_csv('C:\\Users\\<NAME>\\Downloads\\New folder (5)\\SMSSpamCollection', sep='\t',
names=["label", "message"])
nltk.download('stopwords')
# +
#Data cleaning and preprocessing
import re
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
ps = PorterStemmer()
corpus = []
for i in range(0, len(messages)):
review = re.sub('[^a-zA-Z]', ' ', messages['message'][i])
review = review.lower()
review = review.split()
review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
review = ' '.join(review)
corpus.append(review)
# -
corpus
# Creating the Bag of Words model
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features=5000)
X = cv.fit_transform(corpus).toarray()
X.shape
y=pd.get_dummies(messages['label'])
y=y.iloc[:,1].values
y.shape
# +
# Train Test Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 0)
# +
from sklearn.naive_bayes import MultinomialNB
spam_detect_model = MultinomialNB().fit(X_train, y_train)
y_pred=spam_detect_model.predict(X_test)
# -
from sklearn.metrics import confusion_matrix
conf=confusion_matrix(y_test,y_pred)
conf
from sklearn.metrics import accuracy_score
score=accuracy_score(y_test,y_pred)
score
| Spam Classifier With NLP/Implementing a Spam classifier in python NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pb111/Python-tutorials-and-projects/blob/master/_args_and_kwargs_in_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LUirywQGAPk5"
# # **`*args and **kwargs in Python`**
# + [markdown] id="QjG4stXZB1av"
# In Python, we can pass a variable number of arguments to a function using special symbols. There are two special symbols:
# + [markdown] id="o7pNLKKyCCmE"
# 
# + [markdown] id="f_o7eYUdCqJ1"
# Special Symbols Used for passing arguments:-
#
# 1.)*args (Non-Keyword Arguments)
#
# 2.)**kwargs (Keyword Arguments)
# + [markdown] id="XHMmRoaXDPz1"
# “We use *args and **kwargs as an argument when we have no doubt about the number of arguments we should pass in a function.”
# + [markdown] id="F2GoenWcvhqE"
# ### **1.) *args**
#
# The special syntax *args in function definitions in python is used to pass a variable number of arguments to a function. It is used to pass a non-key worded, variable-length argument list.
#
# - The syntax is to use the symbol * to take in a variable number of arguments; by convention, it is often used with the word args.
#
# - What *args allows you to do is take in more arguments than the number of formal arguments that you previously defined. With *args, any number of extra arguments can be tacked on to your current formal parameters (including zero extra arguments).
#
# - For example : we want to make a multiply function that takes any number of arguments and able to multiply them all together. It can be done using *args.
#
# - Using the *, the variable that we associate with the * becomes an iterable meaning you can do things like iterate over it, run some higher-order functions such as map and filter, etc.
# + [markdown] id="2PksEFIOF7c9"
# #### **Example for usage of *args:**
# + colab={"base_uri": "https://localhost:8080/"} id="9p2fBTm9wI3U" outputId="82513591-2734-43cc-c332-2e9a572fadec"
# Python program to illustrate
# *args for variable number of arguments
def myFun(*argv):
for arg in argv:
print(arg)
myFun('Hello','Welcome','to','GeeksforGeeks')
# + id="mnCkOanuxJ8T" colab={"base_uri": "https://localhost:8080/"} outputId="ff63c9eb-ab94-4257-8b33-7bf3b454f7f3"
# Python program to illustrate
# *args with first extra argument
def myFun(arg1,*argv):
print("First argument :", arg1)
for arg in argv:
print("Next argument through *argv :", arg)
myFun('Hello', 'Welcome', 'to', 'GeeksforGeeks')
# + [markdown] id="3gWBuJjSEHyD"
# ### **2.) **kwargs**
#
# The special syntax **kwargs in function definitions in python is used to pass a keyworded, variable-length argument list. We use the name kwargs with the double star. The reason is because the double star allows us to pass through keyword arguments (and any number of them).
#
# - A keyword argument is where you provide a name to the variable as you pass it into the function.
# - One can think of the *kwargs* as being a dictionary that maps each keyword to the value that we pass alongside it. That is why when we iterate over the *kwargs* there doesn’t seem to be any order in which they were printed out.
# + [markdown] id="qu5XxyqMFuqO"
# #### **Example for usage of **kwargs:**
# + colab={"base_uri": "https://localhost:8080/"} id="iFnUuhaUDgXd" outputId="c0021824-7ff9-4669-ff5e-b59516e4cc98"
# Python program to illustrate
# **kwargs for variable number of keyword arguments
def myFun1(**kwargs):
for key,value in kwargs.items():
print("%s == %s" %(key, value))
# Driver code
myFun1(first="Geeks", mid="for", last="Geeks")
# + colab={"base_uri": "https://localhost:8080/"} id="zJUoTVqdH_YE" outputId="da65d4ba-77db-48ec-fd0c-de80d37051f9"
# Python program to illustrate **kargs for
# variable number of keyword arguments with
# one extra argument.
def myFun2(arg1, **kwargs):
for key, value in kwargs.items():
print ("%s == %s" %(key, value))
# Driver code
myFun2("Hi", first ='Geeks', mid ='for', last='Geeks')
# + [markdown] id="0d9Gozw0LVTg"
# ### **3. Using `*args` and **kwargs to call a function**
# + colab={"base_uri": "https://localhost:8080/"} id="EbrHhFcvJ_I_" outputId="4f6ba8af-1024-4484-bbdf-8e2116bd097e"
def myFun(arg1, arg2, arg3):
print("arg1:", arg1)
print("arg2:", arg2)
print("arg3:", arg3)
# Now we can use *args or **kwargs to
# pass arguments to this function :
args = ("Geeks", "for", "Geeks")
myFun(*args)
kwargs = {"arg1" : "Geeks", "arg2" : "for", "arg3" : "Geeks"}
myFun(**kwargs)
# + [markdown] id="X1ra4ijENASj"
# ### **4. Using `*args` and **kwargs in same line to call a function**
# + colab={"base_uri": "https://localhost:8080/"} id="4MoD4lnXME2C" outputId="4356b895-188d-4a04-efb0-6ace52321728"
def myFun(*args,**kwargs):
print("args: ", args)
print("kwargs: ", kwargs)
# Now we can use both *args ,**kwargs
# to pass arguments to this function :
myFun('geeks','for','geeks', first="Geeks", mid="for", last="Geeks")
# + [markdown] id="BVAa2JCcBkg_"
# https://www.geeksforgeeks.org/args-kwargs-python/
| _args_and_kwargs_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Examples</h1>
# <strong>Example #1: Basic Math Operations</strong>
#
# I want to point out two things. The first is the print statements.
#
# Do you see how each variable is preceded by str? That is because those variables are either integers or floats. If we want to smoosh a number together with a string, we have to cast the number as a string and that is what the str method does.
#
# The second thing is the value of the division operation. If you are sharp you will see that we divided two integer values. Despite the fact that the result was an integer, the data type of division resulted in a float.
# +
x = 10
y = 5
addition = x + y
subtraction = x - y
multiplication = x * y
division = x / y
integer_division = x // y
modulo = x % y
exponent = x ** y
print('The result of addition was ' + str(addition))
print('The result of subtraction was ' + str(subtraction))
print('The result of multiplication was ' + str(multiplication))
print('The result of division was ' + str(division))
print('The result of integer_division was ' + str(integer_division))
print('The result of modulo was ' + str(modulo))
print('The result of exponent was ' + str(exponent))
# -
# Copyright © 2020, Mass Street Analytics, LLC. All Rights Reserved.
| 02 Basics/06-arithmetic-operators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RPReactor 3.8
# language: python
# name: rpreactor_3.8
# ---
# # Notebook 7 - Property modeling with mol2vec
# By <NAME>, <NAME> November 2021 - May 2022
# This notebook was developed using **RPReactor 3.8** kernel on [jprime.lbl.gov](https://gpu2.ese.lbl.gov/).
# In this notebook, we explore the application of [mol2vec embeddings](https://pubs.acs.org/doi/abs/10.1021/acs.jcim.7b00616) to model different molecular properties. The embeddings are used as a drop-in replacement of conventional molecular descriptors. Given the relatively high dimensionality of mol2vec embeddings (300-D), we apply the heuristic feature selection approach that was also used in notebooks 5 and 6.
# +
import numpy as np
import pandas as pd
from sklearn.model_selection import KFold, GridSearchCV
from sklearn.svm import SVR
from sklearn.linear_model import LinearRegression
from sklearn.feature_selection import SequentialFeatureSelector
# %run "./plotting.py"
# -
# %config InlineBackend.figure_format = 'retina'
# We already have the molecules embedded using mol2vec:
df_mol2vec = pd.read_csv('./data/mol2vec_all.csv', index_col=0)
df_mol2vec.head()
# ## Research Octane Number (RON)
# Let us load the data and look at the distribution:
df = pd.read_csv("./data/data_RON.csv")
smiles = df.Smiles
Y = df.RON
print(len(smiles))
plot_histogram(Y, xlabel="Research Octane Number (RON)")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='RON'].iloc[:,2:])
D.shape
# Define the partitions for cross-validation.
num_of_partitions = 10
kf = KFold(n_splits=num_of_partitions, shuffle=True, random_state=42)
# We want to select a subset of descriptors that we will use to train the model.
sfs = SequentialFeatureSelector(LinearRegression(), n_features_to_select=15)
# Define hyperparameters for the regression model:
param_grid = {
'C': [1, 5, 7, 10, 30, 50, 100, 300, 500],
'epsilon': [0.1, 0.3, 1, 3, 5, 10, 20],
'kernel': ['rbf']
}
# Perform cross-validation:
# + jupyter={"outputs_hidden": true} tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="Research Octane Number observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/RON_mol2vec.svg')
# -
# ## Cetane Number (CN)
df = pd.read_csv("./data/data_CN.csv")
smiles = df.Smiles
Y = df.CN
print(len(smiles))
plot_histogram(Y, xlabel="Cetane Number (CN)")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='CN'].iloc[:,2:])
D.shape
# + jupyter={"outputs_hidden": true} tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="Cetane Number observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/CN_mol2vec.svg')
# -
# ## Melting Point (MP)
df = pd.read_csv("./data/data_MP.csv")
smiles = df.Smiles
Y = df.mpC
print(len(smiles))
plot_histogram(Y, xlabel="Melting Point (Celsius)")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='MP'].iloc[:,2:])
print(D.shape)
# Define a smaller hyperparameter space as this is a large data set
param_grid = {
'C': [100, 300, 500],
'epsilon': [3, 5, 10, 20],
'kernel': ['rbf']
}
# + tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="Melting Point (Celsius) observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/MP_mol2vec.svg')
# -
# ## Flash Point (FP)
df = pd.read_csv("./data/data_FP.csv")
smiles = df.Smiles
Y = df.FP
print(len(smiles))
plot_histogram(Y, xlabel="Flash Point (Celsius)")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='FP'].iloc[:,2:])
print(D.shape)
param_grid = {
'C': [1, 5, 7, 10, 30, 50, 100, 300, 500],
'epsilon': [0.1, 0.3, 1, 3, 5, 10, 20],
'kernel': ['rbf']
}
# + tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="Flash Point (Celsius) observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/FP_mol2vec.svg')
# -
# ## Histamine receptor pKi
df = pd.read_csv("./data/data_H1.csv")
Y = df.pKi
smiles = df.Smiles
print(len(smiles))
plot_histogram(Y, xlabel="pKi binding to histamine H1 receptor")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='H1'].iloc[:,2:])
print(D.shape)
# + tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="pKi H1 observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/H1_mol2vec.svg')
# -
# ## Muscarinic receptor pKi
df = pd.read_csv("./data/data_M2.csv")
Y = df.pKi
smiles = df.Smiles
print(len(smiles))
plot_histogram(Y, xlabel="pKi binding to muscarinic M2 receptor")
D = np.array(df_mol2vec[df_mol2vec['dataset']=='M2'].iloc[:,2:])
print(D.shape)
# + tags=[]
# %%time
Y_cv_pred = []
Y_obs = []
i = 1
for train_index, val_index in kf.split(D):
print(f"Partition {i}/{num_of_partitions}")
i+=1
D_train , D_val = D[train_index, :], D[val_index, :]
y_train , y_val = Y[train_index], Y[val_index]
sfs.fit(D_train, y_train)
X_train = sfs.transform(D_train)
X_val = sfs.transform(D_val)
# Train the SVR model
# Optimize hyperparameters
grid = GridSearchCV(SVR(), param_grid, cv=5, refit=True)
grid.fit(X_train, y_train)
print(grid.best_params_)
# Test set predictions
y_cv_pred = grid.predict(X_val)
# Save corresponding validation instances
Y_cv_pred.extend(y_cv_pred)
Y_obs.extend(y_val)
# + tags=[]
# Parity plot
parity_plot(x=Y_obs,
y=Y_cv_pred,
xlabel="pKi M2 observations",
ylabel="Cross-validated predictions",
savetitle='./results/mol2vec/M2_mol2vec.svg')
# -
| notebooks/N7_mol2vec.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0
# language: julia
# name: julia-1.3
# ---
# # Performance
#
# This notebook measures performance of `Simulate.jl` functionality in order to compile the [performance section](https://pbayer.github.io/Simulate.jl/dev/performance/) of the documentation.
using Simulate, BenchmarkTools, Random
res = Dict(); # results dictionary
# ## Event-based simulations
# The following is a modification of the [channel example](https://pbayer.github.io/Simulate.jl/dev/approach/#Event-based-modeling-1). We simulate events
#
# 1. taking something from a common channel or waiting if there is nothing,
# 2. then taking a delay, doing a calculation and
# 3. returning three times to the first step.
#
# As calculation we take the following Machin-like sum:
#
# $$4 \sum_{k=1}^{n} \frac{(-1)^{k+1}}{2 k - 1}$$
#
# This gives a slow approximation to $\pi$. The benchmark creates long queues of timed and conditional events and measures how fast they are handled.
# ### Function calls as events
#
# The first implementation is based on events with `SimFunction`s.
# +
function take(id::Int64, qpi::Vector{Float64}, step::Int64)
if isready(ch)
take!(ch) # take something from common channel
event!(SF(put, id, qpi, step), after, rand()) # timed event after some time
else
event!(SF(take, id, qpi, step), SF(isready, ch)) # conditional event until channel is ready
end
end
function put(id::Int64, qpi::Vector{Float64}, step::Int64)
put!(ch, 1)
qpi[1] += (-1)^(id+1)/(2id -1) # Machin-like series (slow approximation to pi)
step > 3 || take(id, qpi, step+1)
end
function setup(n::Int) # a setup he simulation
reset!(𝐶)
Random.seed!(123)
global ch = Channel{Int64}(32) # create a channel
global qpi = [0.0]
si = shuffle(1:n)
for i in 1:n
take(si[i], qpi, 1)
end
for i in 1:min(n, 32)
put!(ch, 1) # put first tokens into channel 1
end
end
# -
# If we setup 250 summation elements, we get 1000 timed events and over 1438 sample steps with conditional events.
@time setup(250)
println(@time run!(𝐶, 500))
println("result=", qpi[1])
t = run(@benchmarkable run!(𝐶, 500) setup=setup(250) evals=1 seconds=15.0 samples=50)
res["Event based with SimFunctions"] = minimum(t).time * 1e-6 # ms
# ### Expressions as events
#
# The 2nd implementation does the same but with expressions, which are `eval`uated in global scope during runtime. This gives a one-time warning for beeing slow:
# +
function take(id::Int64, qpi::Vector{Float64}, step::Int64)
if isready(ch)
take!(ch) # take something from common channel
event!(:(put($id, qpi, $step)), after, rand()) # timed event after some time
else
event!(:(take($id, qpi, $step)), :(isready(ch))) # conditional event until channel is ready
end
end
function put(id::Int64, qpi::Vector{Float64}, step::Int64)
put!(ch, 1)
qpi[1] += (-1)^(id+1)/(2id -1) # Machin-like series (slow approximation to pi)
step > 3 || take(id, qpi, step+1)
end
# -
@time setup(250)
println(@time run!(𝐶, 500))
println("result=", sum(qpi))
t = run(@benchmarkable run!(𝐶, 500) setup=setup(250) evals=1 seconds=15.0 samples=50)
res["Event based with Expressions"] = minimum(t).time * 1e-6 #
res
res["Event based with Expressions"]/res["Event based with SimFunctions"]
# This takes much longer and shows that `eval` for Julia expressions, done in global scope is very expensive and should be avoided if performance is any issue.
# ### Involving a global variable
#
# The third implementation works with `Simfunction`s like the first but involves a global variable `A`:
# +
function take(id::Int64, qpi::Vector{Float64}, step::Int64)
if isready(ch)
take!(ch) # take something from common channel
event!(SF(put, id, qpi, step), after, rand()) # timed event after some time
else
event!(SF(take, id, qpi, step), SF(isready, ch)) # conditional event until channel is ready
end
end
function put(id::Int64, qpi::Vector{Float64}, step::Int64)
put!(ch, 1)
global A += (-1)^(id+1)/(2id -1) # Machin-like series (slow approximation to pi)
step > 3 || take(id, qpi, step+1)
end
function setup(n::Int) # a setup he simulation
reset!(𝐶)
Random.seed!(123)
global ch = Channel{Int64}(32) # create a channel
global A = 0
si = shuffle(1:n)
for i in 1:n
take(si[i], qpi, 1)
end
for i in 1:min(n, 32)
put!(ch, 1) # put first tokens into channel 1
end
end
# -
ch = Channel{Int64}(32)
@code_warntype put(1, qpi, 1)
@time setup(250)
println(@time run!(𝐶, 500))
println("result=", A)
t = run(@benchmarkable run!(𝐶, 500) setup=setup(250) evals=1 seconds=10.0 samples=30)
res["Event based with functions and a global variable"] = minimum(t).time * 1e-6 #
res
# In this case the compiler does well to infer the type of `A` and it runs only marginally slower than the first version.
| docs/notebooks/performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy.stats import gaussian_kde
from scipy.interpolate import interp1d
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import rc
rc('font', **{'family': 'serif', 'serif': ['Computer Modern']})
rc('text', usetex=True)
# # Building the joint prior
#
# In this repository there exists code to compute the conditional priors $p(\chi_\mathrm{eff}|q)$ and $p(\chi_q|q)$ (functions `chi_effective_prior_from_isotropic_spins` and `chi_p_prior_from_isotropic_spins`, respectively) on $\chi_\mathrm{eff}$ and $\chi_p$ corresponding to uniform and isotropic component spin priors. Each of these priors have been marginalized over all other spin degrees of freedom.
#
# In some circumstances, though, we might want the *joint* prior $p(\chi_\mathrm{eff},\chi_p|q)$ acting on the two effective spin parameters. Although we were able to derive closed-form expressions for $p(\chi_\mathrm{eff}|q)$ and $p(\chi_q|q)$, I personally lack the will-power and/or attention span to derive an analytic expression for $p(\chi_\mathrm{eff},\chi_p|q)$. Instead, let's build a function to do this numerically.
#
# First, note that the joint prior on $\chi_\mathrm{eff}$ and $\chi_p$ is *weird*. Demonstrate this by drawing random component spins, computing the corresponding effective spins, and plotting the resulting density.
# +
def chi_p(a1,a2,cost1,cost2,q):
sint1 = np.sqrt(1.-cost1**2)
sint2 = np.sqrt(1.-cost2**2)
return np.maximum(a1*sint1,((3.+4.*q)/(4.+3.*q))*q*a2*sint2)
def chi_eff(a1,a2,cost1,cost2,q):
return (a1*cost1 + q*a2*cost2)/(1.+q)
# Choose some fixed mass ratio
q = 0.5
# Draw random component spins and compute effective parameters
ndraws = 30000
random_a1s = np.random.random(ndraws)
random_a2s = np.random.random(ndraws)
random_cost1s = 2.*np.random.random(ndraws)-1.
random_cost2s = 2.*np.random.random(ndraws)-1.
# Plot!
random_chi_effs = chi_eff(random_a1s,random_a2s,random_cost1s,random_cost2s,q)
random_chi_ps = chi_p(random_a1s,random_a2s,random_cost1s,random_cost2s,q)
fig,ax = plt.subplots()
ax.hexbin(random_chi_effs,random_chi_ps,cmap='Blues',gridsize=30)
ax.set_xlabel('$\chi_\mathrm{eff}$',fontsize=14)
ax.set_ylabel('$\chi_p$',fontsize=14)
plt.show()
# -
# There are a few visible features we need to worry about.
# 1. First, the prior distribution comes to a sharp point at $\chi_\mathrm{eff} = \chi_p = 0$; this is related to the fact that the marginal $p(\chi_\mathrm{eff}|q)$ is quite sharply peaked about the origin (see `Demo.ipynb`)
# 2. The concentration about $\chi_\mathrm{eff} = 0$ also implies that vanishingly few of our prior draws occur in the distant wings of the joint prior, at very negative or very positive $\chi_\mathrm{eff}$.
# 3. In the vertical direction, we can see the same sharp drop and extended plateau as seen in the marginal $\chi_p$ prior in `Demo.ipynb`
#
# Naively, we could just draw a bunch of prior samples and form a KDE over this space. The first two features listed above, though, make this extremely difficult. The extreme narrowness of $p(\chi_\mathrm{eff},\chi_p|q)$ near the origin means we must use an extremely small KDE bandwidth to accurately capture this behavior, but such a small bandwidth will accentuate sampling fluctuations elsewhere. Meanwhile, the fact that very few samples occur at very positive or very negative $\chi_\mathrm{eff}$ means that we will need to perform a vast number of draws (like, many millions) if we wish to accurately estimate the prior on posterior samples falling in these areas.
#
# Recall that this prior remains *conditional* on $q$, and so we can't just build a single KDE (in which case we might tolerate having to perform a vast number of draws and slow KDE evaluation), but will need to build a new estimator every time we consider a different mass ratio.
#
# Instead, let's leverage our knowledge of the marginal prior $p(\chi_\mathrm{eff}|q)$ and factor the joint prior as
#
# \begin{equation}
# p(\chi_\mathrm{eff},\chi_p|q) = p(\chi_p|\chi_\mathrm{eff},q) p(\chi_\mathrm{eff},q),
# \end{equation}
#
# so that we only have to worry about numerically constructing the one-dimensional distribution $p(\chi_p|\chi_\mathrm{eff},q)$.
#
# Given $\chi_\mathrm{eff}$ and $q$, we will repeatedly draw $\{a_1,a_2,\cos t_1,\cos t_2\}$ consistent with $\chi_\mathrm{eff}$, and then construct the resulting distribution over $\chi_p$. In particular, we will regard
#
# \begin{equation}
# \cos t_1 = \frac{(1+q)\chi_\mathrm{eff} - q a_2 \cos t_2}{a_1}
# \end{equation}
#
# as a function of the $\chi_\mathrm{eff}$ and the three other component spin parameters. In making this choice, though, we are *really* drawing from a slice through
#
# \begin{equation}
# \frac{dP}{d a_1 da_2 d\chi_\mathrm{eff} d\cos t_2 } = \frac{dP}{d a_1 da_2 d \cos t_1 d\cos t_2} \frac{\partial \cos t_1}{\partial \chi_\mathrm{eff}}.
# \end{equation}
#
# Thus, in order to have properly sampled from the underlying uniform and isotropic distribution $dP/d a_1 da_2 d \cos t_1 d\cos t_2$, we will need to remember to divide out the Jacobian weights $\partial \cos t_1/\partial \chi_\mathrm{eff} = a_1/(1+q)$.
#
# Let's try this in the following cell:
# +
# Fix some value for chi_eff and q
# Feel free to change these!
aMax = 1.
Xeff = 0.2
q = 0.5
# Draw random spin magnitudes.
# Note that, given a fixed chi_eff, a1 can be no larger than (1+q)*chi_eff,
# and a2 can be no larger than (1+q)*chi_eff/q
ndraws = 100000
a1 = np.random.random(ndraws)*aMax
a2 = np.random.random(ndraws)*aMax
# Draw random tilts for spin 2
cost2 = 2.*np.random.random(ndraws)-1.
# Finally, given our conditional value for chi_eff, we can solve for cost1
# Note, though, that we still must require that the implied value of cost1 be *physical*
cost1 = (Xeff*(1.+q) - q*a2*cost2)/a1
# While any cost1 values remain unphysical, redraw a1, a2, and cost2, and recompute
# Repeat as necessary
while np.any(cost1<-1) or np.any(cost1>1):
to_replace = np.where((cost1<-1) | (cost1>1))[0]
a1[to_replace] = np.random.random(to_replace.size)*aMax
a2[to_replace] = np.random.random(to_replace.size)*aMax
cost2[to_replace] = 2.*np.random.random(to_replace.size)-1.
cost1 = (Xeff*(1.+q) - q*a2*cost2)/a1
Xp_draws = chi_p(a1,a2,cost1,cost2,q)
jacobian_weights = (1.+q)/a1
# -
# For comparison, let's also take a brute-force approach, drawing truly random component spins and saving those whose $\chi_\mathrm{eff}$ are "close to" the conditioned $\chi_\mathrm{eff}$ value specified above. This can take a while, depending on the values of $q$ and $\chi_\mathrm{eff}$ we've chosen...
# +
test_a1s = np.array([])
test_a2s = np.array([])
test_cost1s = np.array([])
test_cost2s = np.array([])
while test_a1s.size<30000:
test_a1 = np.random.random()*aMax
test_a2 = np.random.random()*aMax
test_cost1 = 2.*np.random.random()-1.
test_cost2 = 2.*np.random.random()-1.
test_xeff = chi_eff(test_a1,test_a2,test_cost1,test_cost2,q)
if np.abs(test_xeff-Xeff)<0.02:
test_a1s = np.append(test_a1s,test_a1)
test_a2s = np.append(test_a2s,test_a2)
test_cost1s = np.append(test_cost1s,test_cost1)
test_cost2s = np.append(test_cost2s,test_cost2)
# -
# Let's plot both approaches below. For completeness, also plot what happens if we *forget* the Jacobian factors, which gives a clear mismatch relative to the brute force draws.
fig,ax = plt.subplots()
ax.hist(Xp_draws,density=True,bins=30,weights=jacobian_weights,label='Our approach')
ax.hist(Xp_draws,density=True,bins=30,histtype='step',ls='--',color='black',label='Our approach (w/out Jacobians)')
ax.hist(chi_p(test_a1s,test_a2s,test_cost1s,test_cost2s,q),density=True,histtype='step',bins=30,color='black',
label='Brute force')
plt.legend()
ax.set_xlabel(r'$\chi_p$',fontsize=14)
ax.set_ylabel(r'$p(\chi_p|\chi_\mathrm{eff},q)$',fontsize=14)
plt.show()
# We could stop here, KDE our (appropriately weighted) draws, and evaluate the KDE at a $\chi_p$ of interest. We want to be a bit more careful with the end points, though. If we KDE directly, some of our probability will leak out past our boundaries at $\chi_p = 0$ and $\chi_p = 1$.
# +
demo_kde = gaussian_kde(Xp_draws,weights=jacobian_weights)
fig,ax = plt.subplots()
ax.hist(Xp_draws,density=True,bins=30,weights=jacobian_weights)
ax.plot(np.linspace(-0.1,1.1,50),demo_kde(np.linspace(-0.1,1.1,50)),color='black',label='KDE')
plt.legend()
ax.set_xlabel(r'$\chi_p$',fontsize=14)
ax.set_ylabel(r'$p(\chi_p|\chi_\mathrm{eff},q)$',fontsize=14)
plt.show()
# -
# Even if we truncate to the interval $0 \leq \chi_p \leq 1$, we will still generically end up in a situation where our prior does not go to zero at $\chi_p = 0$ and $\chi_p = 1$:
# +
# Integrate across (0,1) to obtain appropriate normalization
truncated_grid = np.linspace(0,1,100)
norm_constant = np.trapz(demo_kde(truncated_grid),truncated_grid)
fig,ax = plt.subplots()
ax.hist(Xp_draws,density=True,bins=30,weights=jacobian_weights)
ax.plot(truncated_grid,demo_kde(truncated_grid)/norm_constant,color='black',label='KDE')
plt.legend()
ax.set_xlabel(r'$\chi_p$',fontsize=14)
ax.set_ylabel(r'$p(\chi_p|\chi_\mathrm{eff},q)$',fontsize=14)
plt.show()
# -
# Instead, we will take a two step approach. First, use a KDE to evaluate $p(\chi_p|\chi_\mathrm{eff},q)$ across a grid of points well inside the boundaries at $0$ and $\mathrm{Max}(\chi_p)$. Then manually specificy the endpoints, with $p(\chi_p|\chi_\mathrm{eff},q) = 0$.
#
# Note that the maximum value of $\chi_p$ given some $\chi_\mathrm{eff}$ is
#
# \begin{equation}
# \begin{aligned}
# \mathrm{Max}(\chi_p) &= \mathrm{Max}\left[\mathrm{max}\left( s_{1p}, \frac{3+4q}{4+3q} q s_{2p}\right)\right] \\
# &= \mathrm{Max}(s_{1p}),
# \end{aligned}
# \end{equation}
#
# defining $s_p = a \sin t$ as the in-plane spin component. If we define $s_z = a \cos t$, then
#
# \begin{equation}
# \begin{aligned}
# \mathrm{Max}(\chi_p)
# &= \mathrm{Max}\sqrt{a^2_\mathrm{max}-s_{1z}^2} \\
# &= \sqrt{a^2_\mathrm{max}-\mathrm{Min}(s_{1z}^2)} \\
# &= \sqrt{a^2_\mathrm{max}-\mathrm{Min}\left[\left((1+q)\chi_\mathrm{eff} - q s_{2z}\right)^2\right]}
# \end{aligned}
# \end{equation}
#
# where the minimum is taken over possible $s_{2z}$. If $(1+q)\chi_\mathrm{eff} \leq a_\mathrm{max} q$, then there is always some $s_{2z}$ available such that the bracketed term is zero, giving $\mathrm{Max}(\chi_p) = a_\mathrm{max}$. If, on the other hand, $(1+q)\chi_\mathrm{eff} > a_\mathrm{max} q$ then the bracketed term will necessarily always be non-zero, with its smallest value occurring at $s_{2z} = a_\mathrm{max}$. In this case, $\mathrm{Max}(\chi_p) = \sqrt{a^2_\mathrm{max}-\left((1+q)\chi_\mathrm{eff} - a_\mathrm{max} q\right)^2}$.
# +
# Compute maximum chi_p
if (1.+q)*np.abs(Xeff)/q<aMax:
max_Xp = aMax
else:
max_Xp = np.sqrt(aMax**2 - ((1.+q)*np.abs(Xeff)-q)**2.)
# Set up a grid slightly inside (0,max chi_p) and evaluate KDE
reference_grid = np.linspace(0.05*max_Xp,0.95*max_Xp,30)
reference_vals = demo_kde(reference_grid)
# Manually prepend/append zeros at the boundaries
reference_grid = np.concatenate([[0],reference_grid,[max_Xp]])
reference_vals = np.concatenate([[0],reference_vals,[0]])
norm_constant = np.trapz(reference_vals,reference_grid)
# Interpolate!
prior_vals = [np.interp(Xp,reference_grid,reference_vals) for Xp in truncated_grid]
fig,ax = plt.subplots()
ax.hist(Xp_draws,density=True,bins=30,weights=jacobian_weights)
ax.plot(truncated_grid,prior_vals/norm_constant,color='black',label='Our interpolant')
plt.legend()
ax.set_xlabel(r'$\chi_p$',fontsize=14)
ax.set_ylabel(r'$p(\chi_p|\chi_\mathrm{eff},q)$',fontsize=14)
plt.show()
# -
# This procedure is implemented in the function `chi_p_prior_given_chi_eff_q` appearing in `priors.py`. For completeness, let's compare the output of this function against the result we got in this notebook.
from priors import *
ndraws=100000
priors_from_function = [chi_p_prior_given_chi_eff_q(q,aMax,Xeff,xp,ndraws=ndraws,bw_method=1.*ndraws**(-1./5.)) for xp in reference_grid]
fig,ax = plt.subplots()
ax.plot(reference_grid,priors_from_function,label='From priors.py')
ax.plot(reference_grid,reference_vals)
ax.set_xlabel(r'$\chi_p$',fontsize=14)
ax.set_ylabel(r'$p(\chi_p|\chi_\mathrm{eff},q)$',fontsize=14)
plt.legend()
plt.show()
| Joint-ChiEff-ChiP-Prior.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ai6ph/Loan-default-prediction/blob/main/loan_default_prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] papermill={"duration": 0.014043, "end_time": "2021-06-16T17:09:38.366043", "exception": false, "start_time": "2021-06-16T17:09:38.352000", "status": "completed"} tags=[] id="maritime-calculator"
# Objective of this project is to predict if the loanee will be a defaulter or not.
# We will use the custom dataset available on the Kaggle.
# We will use the simple and basic Data Science and machine learning methods to execute the solution.
# + papermill={"duration": 0.02206, "end_time": "2021-06-16T17:09:38.400413", "exception": false, "start_time": "2021-06-16T17:09:38.378353", "status": "completed"} tags=[] id="confident-warner"
#Including the basic libraries at the start
import pandas as pd
import numpy as np
# + [markdown] papermill={"duration": 0.011799, "end_time": "2021-06-16T17:09:38.424375", "exception": false, "start_time": "2021-06-16T17:09:38.412576", "status": "completed"} tags=[] id="olympic-motivation"
# Now let us impot the dataset to our notebook
# + papermill={"duration": 0.041448, "end_time": "2021-06-16T17:09:38.477814", "exception": false, "start_time": "2021-06-16T17:09:38.436366", "status": "completed"} tags=[] id="latest-agency"
#importing the dataset
df=pd.read_csv("https://raw.githubusercontent.com/ai6ph/Loan-default-prediction/main/Default_Fin.csv")
# + [markdown] papermill={"duration": 0.012079, "end_time": "2021-06-16T17:09:38.502245", "exception": false, "start_time": "2021-06-16T17:09:38.490166", "status": "completed"} tags=[] id="japanese-acquisition"
# Now let us have a simple look at our data.
# + papermill={"duration": 0.037453, "end_time": "2021-06-16T17:09:38.551694", "exception": false, "start_time": "2021-06-16T17:09:38.514241", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 204} id="least-andrew" outputId="60fac468-1992-46f7-b179-43af59008fac"
df.head()
# + [markdown] papermill={"duration": 0.012252, "end_time": "2021-06-16T17:09:38.576695", "exception": false, "start_time": "2021-06-16T17:09:38.564443", "status": "completed"} tags=[] id="faced-compression"
# Now before we move ahead and start using the dataset, let us see if there is any need of data pre-processing by counting null or NaN values if any.
# + papermill={"duration": 0.021825, "end_time": "2021-06-16T17:09:38.610887", "exception": false, "start_time": "2021-06-16T17:09:38.589062", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="peripheral-mixture" outputId="a02a3d57-126a-40f6-941f-d052f0927752"
df.isnull().sum()
# + [markdown] papermill={"duration": 0.012265, "end_time": "2021-06-16T17:09:38.635907", "exception": false, "start_time": "2021-06-16T17:09:38.623642", "status": "completed"} tags=[] id="rising-polish"
# So, we can clearly see that there is no NaN values present in our data, we can directly move ahead and see the interelation of our features present in dataset.
# + [markdown] papermill={"duration": 0.012261, "end_time": "2021-06-16T17:09:38.660737", "exception": false, "start_time": "2021-06-16T17:09:38.648476", "status": "completed"} tags=[] id="scientific-premises"
# As we know that our target variable is "Defaulted?", we will look at the correlation present amoung all of them so we can see it there are any features we don't need in model development.
# + papermill={"duration": 0.027796, "end_time": "2021-06-16T17:09:38.701578", "exception": false, "start_time": "2021-06-16T17:09:38.673782", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 204} id="enabling-vietnamese" outputId="c751959b-1612-41a5-b953-aac798bd8d89"
#looking at the inter-relation or correlation amoung all of the features
df.corr()
# + papermill={"duration": 0.021566, "end_time": "2021-06-16T17:09:38.736169", "exception": false, "start_time": "2021-06-16T17:09:38.714603", "status": "completed"} tags=[] id="native-philip"
#Dropping "Defaulted?" and taking remaining as independent variable X
X = df.drop(['Defaulted?'], axis="columns")
#Taking "Defaulted?" as dependent variable Y
Y = df["Defaulted?"]
# + id="AVbqebaf74D5"
# + [markdown] papermill={"duration": 0.012699, "end_time": "2021-06-16T17:09:38.761869", "exception": false, "start_time": "2021-06-16T17:09:38.749170", "status": "completed"} tags=[] id="reverse-sauce"
# Now, as we have only one dataset, we will split it into 2 parts, named test and train dataset.
# This will help us in giving more clear approach at model training and testing.
# + papermill={"duration": 0.87786, "end_time": "2021-06-16T17:09:39.652703", "exception": false, "start_time": "2021-06-16T17:09:38.774843", "status": "completed"} tags=[] id="ideal-motorcycle"
#Importing required libraries for splitting dataset
from sklearn.model_selection import train_test_split
#Diving dataset to X_train and Y_train as training variables and y_train and y_test as testing variables
#Making the ratio of 70% training and 30% testing as it is most efficient ratio.
x_train, x_test, y_train, y_test = train_test_split(X,Y, test_size = 0.30, random_state = 0)
# + [markdown] papermill={"duration": 0.013202, "end_time": "2021-06-16T17:09:39.679055", "exception": false, "start_time": "2021-06-16T17:09:39.665853", "status": "completed"} tags=[] id="lucky-enlargement"
# Once we have splitted the dataset, we will import the Logistic Regression into our notebook, as it only deals with 2 values, 0 or 1 and True or False which is the best fit for this case.
# + papermill={"duration": 0.101402, "end_time": "2021-06-16T17:09:39.794772", "exception": false, "start_time": "2021-06-16T17:09:39.693370", "status": "completed"} tags=[] id="eleven-designation"
#Importing the Logistic Regression
from sklearn.linear_model import LogisticRegression
#Creating Logistic Regression object
lr = LogisticRegression()
# + [markdown] papermill={"duration": 0.013257, "end_time": "2021-06-16T17:09:39.821776", "exception": false, "start_time": "2021-06-16T17:09:39.808519", "status": "completed"} tags=[] id="helpful-rough"
#
# Now we will just train our model using x_train and y_train variables which we got by splitting dataset.
# + papermill={"duration": 0.085836, "end_time": "2021-06-16T17:09:39.921214", "exception": false, "start_time": "2021-06-16T17:09:39.835378", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="placed-ethnic" outputId="7082a784-eb87-45e3-c288-42379391c392"
#Fitting model with x_train and y_train variables
lr.fit(x_train,y_train)
# + [markdown] papermill={"duration": 0.024948, "end_time": "2021-06-16T17:09:39.971708", "exception": false, "start_time": "2021-06-16T17:09:39.946760", "status": "completed"} tags=[] id="specialized-producer"
# Now we will predict the values over testing dataset and see what happens.
# + papermill={"duration": 0.058132, "end_time": "2021-06-16T17:09:40.055570", "exception": false, "start_time": "2021-06-16T17:09:39.997438", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="cellular-finger" outputId="4<PASSWORD>4<PASSWORD>-e<PASSWORD>-8<PASSWORD>"
Yhat=lr.predict(x_test)
Yhat
# + [markdown] papermill={"duration": 0.029041, "end_time": "2021-06-16T17:09:40.132427", "exception": false, "start_time": "2021-06-16T17:09:40.103386", "status": "completed"} tags=[] id="dangerous-bedroom"
# Now finally, let us see our model accuracy firstly on training datset
# + papermill={"duration": 0.029754, "end_time": "2021-06-16T17:09:40.188754", "exception": false, "start_time": "2021-06-16T17:09:40.159000", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="unique-ivory" outputId="a82d044e-9eb3-446a-ba73-981c406e1a93"
lr.score(x_train,y_train)
# + [markdown] papermill={"duration": 0.025823, "end_time": "2021-06-16T17:09:40.241494", "exception": false, "start_time": "2021-06-16T17:09:40.215671", "status": "completed"} tags=[] id="requested-aruba"
# and finally accuracy of our model for testing dataset
# + papermill={"duration": 0.042641, "end_time": "2021-06-16T17:09:40.310507", "exception": false, "start_time": "2021-06-16T17:09:40.267866", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="framed-frontier" outputId="134fc459-0aa8-40a3-ca08-5aab77407085"
lr.score(x_test,y_test)
# + [markdown] papermill={"duration": 0.026542, "end_time": "2021-06-16T17:09:40.370724", "exception": false, "start_time": "2021-06-16T17:09:40.344182", "status": "completed"} tags=[] id="printable-nightmare"
# We can conclude that, we have successfully built a Machine Learning model to predict the Loan Defaulters using a custom dataset with a training accuracy of 96.6% approximately and testing accuracy of 96.1%.
# + id="7N1aVAc6r-Sw"
| loan_default_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JohnAquino15/Linear-Algebra-58019/blob/main/Midterm_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="cuVQ_C1M3zr1"
# # **Midterm Exam**
# + [markdown] id="VNJeH01N37Fb"
# <NAME>ino Linear Algebra 58019
# + colab={"base_uri": "https://localhost:8080/"} id="-bxJtse14Cci" outputId="92b80ba6-5527-434b-cddf-a43c942871aa"
#Question 1. Create a Python code that displays a square matrix whose length is 5
import numpy as np
A = np.array([[2,4,5,6,7],[6,7,3,5,8],[3,6,7,8,9],[1,4,5,7,8],[3,5,7,8,1]])
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="QjLRjV8R4t6T" outputId="1a094409-f0fd-4b45-c19c-faa56a547218"
#Question 2. Create a Python code that displays a square matrix whose elements below the principal diagonal are zero
import numpy as np
A = np.array([[2,4,5,6,7],[0,7,3,5,8],[0,0,7,8,9],[0,0,0,7,8],[0,0,0,0,1]])
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="HcmWCHh55fxf" outputId="93876b42-81f2-407e-dac1-80de69b4f4d6"
#Question 3. Create a Python code that displays a square matrix which is symmetrical (10 points)
import numpy as np
A = np.array([[1,3,6],[3,7,5],[6,5,8]])
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="BvV4lu4G64Ij" outputId="c2524562-75fd-43d5-bb81-2080c7f29cbb"
#Question 4. What is the inverse of matrix C? Show your solution by python coding.
import numpy as np
C = np.array([[1,2,3],[2,3,3],[3,4,-2]]) #Matrix C
print(C)
# + colab={"base_uri": "https://localhost:8080/"} id="4_9WYviG7rfO" outputId="66cb71c6-10c9-49d2-ed64-1d970542771c"
C_inv=np.linalg.inv(C) #Inverse of Matrix C
print(C_inv)
# + colab={"base_uri": "https://localhost:8080/"} id="byBSjs0J9AKx" outputId="9950a6f8-87fc-45e4-dab6-243b6c71f375"
#Question 5. What is the determinant of the given matrix in Question 4? Show your solution by python coding.
import numpy as np
C = np.array([[1,2,3],[2,3,3],[3,4,-2]]) #Matrix C
print('Determinant =',round(np.linalg.det(C))) #Determinant of Matrix C
# + colab={"base_uri": "https://localhost:8080/"} id="JCgPGfBd9FVo" outputId="ccea303b-8438-4347-f727-53df4146ddf0"
#Question 6. Find the roots of the linear equations by showing its python codes
#5X1 + 4X2 + X3 = 3.4
#10X1 + 9X2 + 4X3 = 8.8
#10X1 + 13X2 + 15X3 = 19.2
import numpy as np
A=np.array([[5,4,1],[10,9,4],[10,13,15]]) #Matrix A of Linear Equations
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="OXzeSGfr-u8U" outputId="63f75e40-bd65-4586-fc3d-490ed50c155e"
B = np.array([[3.4],[8.8],[19.2]]) #Constants
print(B)
# + colab={"base_uri": "https://localhost:8080/"} id="Huahlg1U_QRL" outputId="6b545723-e70d-4d63-a63c-3bce3c431cdf"
A_inv=np.linalg.inv(A) #Inverse of matrix A
print(A_inv)
# + colab={"base_uri": "https://localhost:8080/"} id="P9TNks1B_eFV" outputId="244204ef-e512-44bf-eb0a-2fd5a2370673"
X=np.dot(A_inv,B) #Formula to get the missing variable X1, X2, AND X3
print(X)
# + [markdown] id="U0NFhgZLAaVh"
# X1=0.2, X2=0.4, X3=0.8
# + id="GLnWjneABxKt"
| Midterm_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from igraph import *
inputfile = open('graph.txt', 'r')
edges = []
for line in inputfile:
edges.append([int(x) for x in line.split()])
inputfile.close()
edges
g = Graph(edges)
g.delete_vertices(0)
summary(g)
adj = g.get_adjacency()
N = 300
g.es[g.incident(2)]
cur_edges = []
def dfs(x, rem):
if rem == 0:
return
for i in range(N):
if adj[x - 1][i] > 0:
cur_edges.append((x, i + 1))
adj[x - 1][i] = 0
adj[i][x - 1] = 0
dfs(i + 1, rem - 1)
adj[x - 1][i] = 1
adj[i][x - 1] = 1
degree = []
for v in range(N):
print(v)
cur_edges = []
dfs(v + 1, 9)
cur_edges = sorted(set(cur_edges))
degree.append(cur_edges)
d = [len(degree[i]) for i in range(N)]
f = open('degree.txt', 'w')
for i in range(N):
f.write("%s\n" % d[i])
f.close()
f = open('links.txt', 'w')
for i in range(N):
f.write("%s:\n" % (i + 1))
for item in degree[i]:
f.write("(%s, %s) " % item)
f.write("\n")
f.close()
for i in range(N):
for j in range(len(degree[i])):
degree[i][j] = sorted(set(degree[i][j]))
for i in range(N):
for j in range(len(degree[i])):
degree[i][j] = tuple(degree[i][j])
degree[i] = list(set(degree[i]))
for i in range(N):
degree[i] = sorted(degree[i])
degree[0]
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import urllib.parse
IMG_PATH = "../../www/assets/img"
filenames = os.listdir(IMG_PATH)
filename = filenames[3]
print(filename)
urllib.parse.unquote(filename)
for filename in filenames:
decoded = urllib.parse.unquote(filename)
if str(decoded) != str(filename):
mv_cmd = f"mv \"{IMG_PATH}/{filename}\" \"{IMG_PATH}/{decoded}\""
print(mv_cmd)
os.system(mv_cmd)
| notebooks/urldecode_images_filenames.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # National Plan and Provider Enumeration System (NPPES)
#
# a.k.a. NPI Downloadable File, or full replacement Monthly NPI File,
# National Provider Identifier Standard (NPI).
#
# [Source](https://www.cms.gov/Regulations-and-Guidance/HIPAA-Administrative-Simplification/NationalProvIdentStand/DataDissemination.html)
#
# Grab the current Full Replacement Monthly NPI File from
# [http://download.cms.gov/nppes/NPI_Files.html](http://download.cms.gov/nppes/NPI_Files.html)
url <- "http://download.cms.gov/nppes/NPPES_Data_Dissemination_Apr_2016.zip" # Update this URL as needed
# Download and unzip the file
f <- tempfile()
download.file(url, f, mode="wb")
path <- tempdir()
unzip(f, exdir = path)
unzip(f, list=TRUE)
# Read the data file
library(data.table)
filenames <- unzip(f, list = TRUE)$Name
hdrFile <- grep("FileHeader\\.csv$", filenames, value = TRUE)
hdrFile <- file.path(path, hdrFile)
datFile <- grep("[0-9]\\.csv$", filenames, value = TRUE)
datFile <- file.path(path, datFile)
D <- fread(datFile)
# Substitute spaces in column names
oldvar <- names(D)
newvar <- gsub("\\(|\\)", "", gsub("\\s", "_", oldvar))
setnames(D, oldvar, newvar)
# Show some examples
D[grep("OHSU", Provider_Other_Organization_Name), .(NPI, Provider_Other_Organization_Name)]
# Show all the variables
names(D)
| NPPES/NPPES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
import seaborn as sns4
import warnings
warnings.filterwarnings('ignore')
market_caps = pd.read_csv("../data/stock_mktcaps.csv")
missing = market_caps.isna().sum()
for i in missing.index:
if missing[i] >= 1:
market_caps = market_caps.drop(columns=[i])
# +
market_caps["date"] = pd.to_datetime(market_caps['date'])
market_caps = market_caps.set_index("date")
# -
market_caps
#use Oyuka's filtering here
filtered_market_cap = market_caps["2003-01-02":"2021-06-30"]
filtered_market_cap = filtered_market_cap.agg(np.mean).to_frame().reset_index()
filtered_market_cap = filtered_market_cap.rename(columns={"index":"stock", 0:"avg market cap"})
from sklearn.cluster import KMeans
# +
inertia = []
for k in range(1 , 21):
estimator = KMeans(n_clusters=k)
estimator.fit(filtered_market_cap[["avg market cap"]])
inertia.append(estimator.inertia_)
sns.pointplot(np.arange(1,21), inertia)
# -
km = KMeans(n_clusters=5)
km.fit(filtered_market_cap[["avg market cap"]])
clusters = pd.DataFrame(km.cluster_centers_, columns=["avg market cap"], index=['Cluster1', 'Cluster2', 'Cluster3', 'Cluster4', 'Cluster5'])
filtered_market_cap['Cluster'] = km.labels_ + 1
filtered_market_cap["Cluster"].unique()
filtered_market_cap
print("Min/Max for Cluster 1")
print(filtered_market_cap[filtered_market_cap["Cluster"] == 1]["avg market cap"].min())
print(filtered_market_cap[filtered_market_cap["Cluster"] == 1]["avg market cap"].max())
print()
print("Min/Max for Cluster 2")
print(filtered_market_cap[filtered_market_cap["Cluster"] == 2]["avg market cap"].min())
print(filtered_market_cap[filtered_market_cap["Cluster"] == 2]["avg market cap"].max())
print()
print("Min/Max for Cluster 3")
print(filtered_market_cap[filtered_market_cap["Cluster"] == 3]["avg market cap"].min())
print(filtered_market_cap[filtered_market_cap["Cluster"] == 3]["avg market cap"].max())
print()
print("Min/Max for Cluster 4")
print(filtered_market_cap[filtered_market_cap["Cluster"] == 4]["avg market cap"].min())
print(filtered_market_cap[filtered_market_cap["Cluster"] == 4]["avg market cap"].max())
print()
print("Min/Max for Cluster 5")
print(filtered_market_cap[filtered_market_cap["Cluster"] == 5]["avg market cap"].min())
print(filtered_market_cap[filtered_market_cap["Cluster"] == 5]["avg market cap"].max())
filtered_market_cap.boxplot(by="Cluster")
filtered_market_cap[filtered_market_cap["Cluster"] == 1]
filtered_market_cap[filtered_market_cap["Cluster"] == 2]
filtered_market_cap[filtered_market_cap["Cluster"] == 3]
filtered_market_cap[filtered_market_cap["Cluster"] == 4]
filtered_market_cap[filtered_market_cap["Cluster"] == 5]
| Clustering Avg Market Caps (K-means).ipynb |