code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 5 -- Univariate OLS Regression
#
# This week we will explore a different approach at evaluating the results of the Likert survey. We will assume an equal distance between each label as we did in assessing the **overall** score in week 2:
#
# Label | Description | Percent Score
# --|--|--
# 1 | strongly disagree | 0.20
# 2 | disagree | 0.40
# 3 | neutral | 0.60
# 4 | agree | 0.80
# 5 | strongly agree | 1.00
#
# To make matters more interesting, we have a new dataset this week:
#
# ```demographic_detail.csv```
#
# This dataset comprises the following metrics:
#
# Attribute Name | Field Name | Type | Categorical | Restrictions | Description
# --|--|--|--|--|--
# Employee ID | ```employee_id``` |```int``` | No | 1 and above | Employee ID assigned at start of employment
# Year of Birth | ```year_of_birth``` |```int``` | No | | Calendar year in which the employee was born
# Time on the Job | ```time_on_the_job``` |```int``` | No | | Number of months since the employee joined the company
#
#
# ## Q1: Merge Datasets
#
# Merge the new dataset with the ```roster_with_score.csv``` data. Save the resulting dataframe as a new csv: ```roster_with_score_2.csv```
# ## Q2: Correlation Coefficients
#
# Compute the correlation coefficients for the two new variables in the dataset against **overall** satisfaction. (**HINT**: you may want to transform the year of birth before computing the correlation coefficient).
# ## Q3: OLS Regression
#
# Using the ```statsmodels``` OLS regression package perform and OLS regression of ```overall``` satisfaction vs. the two variables available in the in the dataset.
#
# (a) which of the two regressions produces better results?
#
# (b) what metric(s) are you using to make that assessment?
# ## Q4: Residual Analysis
#
# Inspect the residuals of the two regressions.
#
# (a) Are they normally distributed?
#
# (b) Do you notice any significant outliers?
#
# (c) Do you observe anything else of concern?
# # Q5: Business Insights
#
# How do you interpret the results of the regressions?
#
# What do you recommend based on this analysis?
| 1_class_prep/week_5/loantronic_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MATH6005 Python Lab. Week 1.
# ## Introduction
#
# Welcome to the labs for MATH6005, Introduction to Python.
#
# This lab work assumes you are using `spyder 3` and `Python 3.x`
#
# You might find that the University labs have both spyder 2 and spyder 3 installed. **Please make sure you are using `spyder 3`**
#
# * Week 1 YouTube playlist: https://www.youtube.com/watch?v=u0yDOsUYxZg&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE
# ## Setup
# Python 3 is already installed and ready to use in University computer rooms.
#
# You may use your own laptop if you want. Python is free. Installation may take 10 - 15 minutes, so please *don't* do it right now (although you may want to download the installer on campus - it is not small).
#
# * If you are installing Python we recommend installing it via Anaconda:
# * https://www.youtube.com/watch?v=u0yDOsUYxZg
#
# For the labs, you will need to open `spyder`. On the bench PC, go to the Start menu -> All Programs -> Programming Languages -> Anaconda3 (64 bit), and then choose `spyder`. You may try searching for it but there may be multiple versions installed: you need to ensure that the version you use has **Python version 3**, not version 2.
#
# * We recommend that you view our introduction to Spyder video:
# * https://www.youtube.com/watch?v=bb-Y5ylTAZM&index=2&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE
# ## The print() function
# One of the most useful functions in Python is the `print()` function. It is used to display information to the user. It can be used to present the result of computations, intermediate calculations, general text and used for debugging.
#
# Once you have opened `spyder`, we will first look at the *console* in the bottom right part of the screen. That allows us to type in Python commands and get an *immediate* response. Let's use it to learn how to use `print()`
#
# Let's use `print` to display a message on the console. Type in the following to console:
print('hello world')
# If we want to include the result of a computation in the output from print we use the following format:
#
# ```python
# print('1 + 1 = {0}'.format(1+1))
# ```
#
# Try running the code above. The value in the curly brackets '{}' is replaced by the value 2 (i.e. 1+1).
#
# `print()` can include the output from multiple computations if needed e.g.
#
# ```python
# print('1 + 1 = {0} and 2 + 2 = {1}'.format(1+1, 2+2))
# ```
# The numeric value in the curly brakets corresponds to the index of value found in `.format(1+1, 2+2)` I.e. {0} is linked to 1+1 and {1} is linked to 2+2.
#
# You don't have to use the indexes in acsending order, but if you don't do make sure that your code makes sense! e.g.
print('1 + 1 = {1}!? and 2 + 2 = {0}!?'.format(1+1, 2+2))
# * You can also format your output to a specified number of decimal places using `{0:.2f}` instead of `{0}`
# * The .2f after the : tells python that the number is a floating point and that you would like it shortened to 2dp.
print('The number {0} given to 2 decimal places is {0:.2f}'.format(3.14159))
# * Similarly if you wanted to show the number to 3 decimal places you would use
print('The number {0} given to 3 decimal places is {0:.3f}'.format(3.14159))
# ## Basic mathematics in the console
# We have already seen that python can be used for basic mathematics when learning how to use `print()`
#
# Let's use the iPython console as a calculator. Try the following calculations:
1 + 1
1 / 2
(1 + 2.3 * 4.5) / 6.7
# * the `**` operator raises one number to the power of another.
#
3**2
# As before we can mix basic mathematics with `print()` e.g.
#
# (Note: if you are unfamiliar with the mod operator, it operates like a remainder function. if we type `15 % 4`, it will return the remainder after dividing 15 by 4.)
print('Addition: {0}'.format(2+2))
print('Substraction: {0}'.format(7-4))
print('Multiplication: {0}'.format(2*5))
print('Division: {0}'.format(10/2))
print('Exponentiation: {0}'.format(3**2))
print('Modulo: {0}'.format(15%4))
# ## The Spyder editor
# Using the console is fine, but has problems. We want to keep our work for future re-use. We want to systematically build up a lot of code. Neither of these is easy within the console. Instead, we can use the *editor* on the left half of the screen. This acts like a word-processor, allowing us to type in commands that we can then use or run later.
#
# In the editor, remove any existing text and type in some of the commands you've previously used:
#
# ```python
# 1 + 1
# 1 / 2
# print("Hello {0}".format(1 + 2))
# ```
#
# Save the file in a sensible location (your filestore, or the Desktop) under the name `lab1.py`.
#
# We then want to run the commands in the file. To do this, choose "Run" from the Run menu, *or* press the big green play button on the toolbar, *or* press `F5`.
#
# The output you see should look like:
1 + 1
1 / 2
print('1 + 1 = {0}'.format(1+1))
# Notice that the output only shows the result of the print function. This shows one key difference between files and the console: only output that is *explicitly* printed appears on the screen.
# ### Exercise 1: Calculate a factorial in the iPython console and editor
#
# Explicitly compute $6! = 6 \times 5 \times 4 \times 3 \times 2 \times 1$ in the console. Then do the same in the editor, printing it out with explanatory text.
# ## Variables
# We want to be able to store data and results of calculations in ways we can re-use. For this we define variables names.
#
# * Variables names can only contain letters, numbers, and underscores.
# Spaces are not allowed in variable names, so we use underscores instead of spaces. For example, use `student_name` instead of `student name`.
#
# * Variable names should be descriptive, without being too long. For example `mc_wheels` is better than just `wheels`, and `number_of_wheels_on_a_motorycle`.
#
# Here is an example variable called `salary`
salary = 30000
print(salary)
print('The value of the variable called salary is {0}'.format(salary))
# A variable name is a **label** that's points to a location in your computer's memory.
#
# In our example above, think of the variable as a post-it note with `salary` written on it. This points to the integer `30000` in the computer's memory. Then, when asked to `print` the value of `salary`, it `print`s the value with that label on it.
#
# In Python we can move that label to any other variable of any type e.g.
number = 1.2
print(number)
number = 'One point two'
print(number)
# There are certain rules and conventions for variable names:
#
# * always start with a letter;
# * only use lower case Latin letters, or numbers, or underscores;
# * in particular, never use spaces or hyphens (which can be interpreted as a new variable or a minus sign respectively).
#
#
# Mathematical functions also work on variables.
# +
salary = 30000
tax_rate = 0.2
salary_after_tax = salary * (1 - tax_rate)
print(salary_after_tax)
# -
# * Each variable has a **data type**.
# * We can check the data type of a variable using the built-in function `type`
# * For example, `salary` has the data type `int` (short for integer)
# * and `salary_after_tax` is of type `float` (short for floating point number)
# +
salary = 30000
tax_rate = 0.2
salary_after_tax = salary * (1 - tax_rate)
print(type(salary))
print(type(salary_after_tax))
# -
# * Notice that we didn't need to tell (or declare to) Python the data type of each variable
# * This is because Python is a **dynamically typed** programming language.
# * Python infers the type of a variable at runtime (when the code is run)
# * The most common primitive data types in python are:
# +
foo = True # bool (Boolean)
bar = False # bool (Boolean)
spam = 3.142 # float (floating point)
eggs = 10000000 # int (integer)
foobar = 'el<PASSWORD>ys' # str (string)
print(type(foo))
print(type(bar))
print(type(spam))
print(type(eggs))
print(type(foobar))
# -
# ## Strings
# We have already used strings extensively.
# Strings are sets of characters. Strings are easier to understand by looking at some examples.
# Strings are contained by either single, double quotes or triple quotes.
my_string = "This is a double-quoted string"
my_string = 'This is a single-quoted string'
# Double quotes lets us make strings that contain quotations
quote = "<NAME> said, 'Hope for the best, plan for the worst'"
print(quote)
# +
multi_line_string = '''triple quotes let us split strings
over mulitple lines'''
print(multi_line_string)
# -
# ### Exercise 2: Creating and using variables
#
# A rectangular box has width 2, height 3, and depth 2.
#
# Task:
# * Create a variable for width, height and depth.
# * Compute the volume of the box, assigning that to a fourth variable.
# * Print the result along with formattted explanatory text.
# ## Comments in code
# Comments allow you to write in you native language (e.g. English), within your program. In Python, any line that starts with a hash (#) symbol is ignored by the Python interpreter.
# +
# This an inline comment.
print("This line is not a comment, it is code.")
print("Python will ignore comments") #comments can appear after code
# -
# #### What makes a good comment?
#
# * It is short and to the point, but a complete thought. Most comments should be written in complete sentences.
# * It explains your thinking, so that when you return to the code later you will understand how you were approaching the problem.
# * It explains your thinking, so that others who work with your code will understand your overall approach to a problem.
# * It explains particularly difficult sections of code in detail.
# ## Functions and import
# We won't get very far with just basic algebraic operations. We'll want to perform more complex computations. For that we need python functions.
# Python has built-in mathematical functions. For example,
# * abs()
# * round()
# * max()
# * min()
# * sum()
#
# These functions all act as you would expect, given their names. Calling `abs()` on a number will return its absolute value. The `round()` function will round a number to specified number of decimal points (the default is 0).
#
# Additional functionality can be added in with using various packages such as `math` or `numpy`. We will explore `numpy` in more detail later in the course.
#
# To use these packages you need to first `import` them into your code.
import math
# The math library adds a long list of new mathematical functions to Python. It is documentated here: https://docs.python.org/3.7/library/math.html
print('pi: {0}'.format(math.pi))
print("Euler's Constant: {0}".format(math.e))
# Python's Math module includes some mathematical constants as seen above as well as commonly used mathematical functions.
print('Cosine of pi: {0}'.format(math.cos(math.pi)))
# We can import specific constants and functions from python modules
# +
from math import pi, cos
print('Cosine of pi: {0}'.format(cos(pi)))
# -
# ### Exercise 3: Use a function to calculate a factorial
#
# * Import the `math` library.
# * If required use `help(math)` or https://docs.python.org/3.7/library/math.html to explore the math module
# * Use `help(math.factorial)` to explore how you use the math factorial function.
# * use `math.factorial()` to check your calculation of $6!$
# ## Defining Python Functions
# So far we have used functions built-in to python such as `print()` and `math.cos()`
#
# You will also need to define your own functions in Python.
#
# ### Functions. Example 1: adding two numbers together
def my_add(a, b):
"""
Returns the sum of two numeric values
Keyword arguments:
a -- first number
b -- second number
"""
return a + b
# The Python keyword to *define* a function is `def`. Each function has a name: in this case `my_add`.
#
# The keyword arguments to the function are then a comma-separated list between round brackets `()`. This is in the same format as calling the function, but we are inventing the variable names to refer to the input within the function. So, however the user calls the function, the first argument that is passed in will be assigned the label `a` within the function itself.
#
# Finally there is a colon `:` to end the line. That says that whatever follows is the content, or body, of the function: the lines that will be executed when the function is called. All lines within the function to be executed **must** then be indented by four spaces. `spyder` should start doing this automatically.
#
# The three quotes are *documentation* for the function: they have no effect. However, **any undocumented function is broken**. We can see the documentation by using the help function:
help(my_add)
# We then include all the commands with the function that we want to run each time the function is called. Once we have a result that we want to send back to the place that called the function, we `return` it: this send back the appropriate value(s).
#
# We can now call our function:
print(my_add(1, 1))
print(my_add(1.2, 4.5))
# ### Functions: Example 2: Implementing a formula
#
# Suppose that you are promised a payment of £2000 in 5 years time.
#
# Assuming a compound interest *rate* of 3.5\% what is the **present value (PV)** of this future value (FV)?
#
# * We can calculate this with the forumla: PV = FV / ( 1 + rate)^n
# * We do not want type the code for this calculation each time we need it.
# * Instead we create a reusable function that we can call to do this for different FV, rate and n.
# * The code is below. The function follows the same basic pattern as the simple `my_add` function.
def pv(future_value, rate, n):
'''
Discount a value at defined rate n time periods into the future.
Forumula:
PV = FV / (1 + r)^n
Where
FV = future value
r = the comparator (interest) rate
n = number of years in the future
Keyword arguments:
future value -- the value to discount
rate -- the rate at which to do the discounting
n -- the number of time periods into the future
'''
return future_value / (1 + rate)**n
# +
#Test case 1
future_value = 2000
rate = 0.035
years = 5
result = pv(future_value, rate, years)
msg = 'Using an interest rate of {0}, a payment of £{1:.2f}' \
' in {2} years time is worth £{3:.2f} today'
print(msg.format(rate,future_value, years,result))
#Test case 2
future_value = 350
rate = 0.01
years = 10
result = pv(future_value, rate, years)
print(msg.format(rate,future_value, years,result))
# -
# ### Exercise 4: Write a function to convert fahrenheit to celsius
#
# Open Spyder and use the code editor to do the following:
#
# Define a function `convert_fahrenheit_to_celsius` that converts degrees fahrenheit to degrees celsius. The function should have a keyword argument for temperature in degrees fahrenheit and `return` a numeric value for temperature in degrees celsius.
#
# Store the answer in a variable and then print the answer to the user. Answers should be shown to **2 decimal places**.
#
# Conversion formula:
#
# ```python
# deg_celsius = (deg_fahrenheit - 32) / (9.0 / 5.0)
# ```
#
# Test data
#
# 1. Fahrenheit = 20; Celsius = -6.67
#
# 2. Fahrenheit = 100; Celsius = 37.78
# ### Exercise 5: Write a function to calculate velocity
#
# * Define a function that calculates and returns velocity (metres per second).
# * The function should accept two parameters: distance travelled (metres) and time (seconds)
#
# ```python
# velocity (m/s) = metres travelled (m) / time taken (s)
# ```
#
# Test data
#
# 1. distance travelled = 10m; time taken = 5s. (Velocity = 2.00 m/s)
#
# 2. distance travelled = 100m; time taken = 0.12s. (Velocity = 833.33m/s 2dp)
# ### Creating your own Python modules and importing functions
#
# In the same way we imported functions from `math` we can import functions from our own python modules
#
# * Open `py_finance.py` and `test_finance.py`
# * `test_finance.py` imports functions from the `py_finance` module
# * Watch the Youtube video that explains how they work:
# * https://www.youtube.com/watch?v=l0qE5_01bzw&t=170s
# ## Lists
# Variables are useful, but in nearly all real programming problems we need to store and manipulate **lots** of data in memory.
#
# For example, if we were developing a music streaming service we might need to hold the list of song's on a album or an artists back catalog.
#
# Or, if we were developing a software to manage the geographic routing of a fleet of delivery vehicles we might need to hold a matrix of travel distances between postcodes.
#
# A Python `List` is a simple and flexible way to store lots of variables (of any type of data).
foo = [0, 1, 2, 3]
print(foo)
# The square brackets `[]` say that what follows will be a list: a collection of objects. The commas separate the different objects contained within the list.
#
# In Python, a list can hold *anything*. For example:
bar = [0, 1.2, "hello", [3, 4]]
print(bar)
# This list holds an integer, a real number (or at least a floating point number), a string, and another list.
# We can find the length of a list using `len`:
print(len(foo))
# To access individual elements of a list, use square brackets again. The elements are ordered left-to-right, and the first element has number `0`:
print(foo[0])
print(foo[3])
print(bar[1])
print(bar[3])
# If we try to access an element that isn't in the list we get an error:
print(foo[4])
# We can assign the value of elements of a list in the same way as any variable:
foo[1] = 10
print(foo)
# The number in brackets is called the index of the item. Because lists start at zero, the index of an item is always one less than its position in the list. So to get the second item in the list, we need to use an index of 1.
# We can work with multiple elements of a list at once using *slicing* notation:
print(foo)
print(foo[0:2])
print(foo[:2])
print(foo[1:])
print(foo[0:4:2])
print(foo[::2])
# The notation `[start:end:step]` means to return the entries from the `start`, up to **but not including** the `end`, in steps of length `step`. If the `start` is not included (e.g. `[:2]`) it defaults to the start, i.e. `0`. If the `end` is not included (e.g. `[1:]`) it defaults to the end (i.e., `len(...)`). If the `step` is not included it defaults to `1`.
# To get the last item in a list, no matter how long the list is, you can use an index of -1. This syntax also works for the second to last item, the third to last, and so forth. You can't use a negative number larger than the length of the list, however.
print(foo[-1])
print(foo[-2])
print(foo[-1:0:-1])
# If you want to find out the position of an element in a list, you can use the index() function. This method returns a ValueError if the requested item is not in the list.
print(foo.index(10))
# You can test whether an item is in a list using the "in" keyword. This will become more useful after learning how to use if-else statements.
print(10 in foo)
print(11 in foo)
# We can add an item to a list using the append() method. This method adds the new item to the end of the list.
foo.append(12)
print(foo)
# We can also insert items anywhere we want in a list, using the insert() function. We specify the position we want the item to have, and everything from that point on is shifted one position to the right. In other words, the index of every item after the new item is increased by one.
foo.insert(1,13)
print(foo)
# We can remove an item from a list using the `del` statement. You need to specify the index you wish to remove.
del foo[2]
print(foo)
# ### Exercise 6: Marvel Comics
#
# You are given a list of comics:
#
# ```python
# comics = ['Iron-man', 'Captain America', 'Spider-man', 'Thor', 'Deadpool']
# ```
# Tasks:
#
# * slice and then print the first and second list items
# * slices and then print the second to fourth list items
# * slice and then print the fourth and fifth list items
# * append "Doctor Strange" to the list. Print the updated list
# * insert "Headpool" before "Deadpool" in the list. Print the updated list
# * delete "Iron-man". Print the updated list
#
# ### Week 1: Debug Challenge
#
# Each laboratory will have a debug challenge. You will be given a pre-existing script containing Python code. The catch is that the code doesn't run!
#
# Your challenge is to find and correct the errors so that the script correctly executes.
#
# The challenges are based around common problems students have when writing code. If you do the exercises it will help you debug your own code and maybe even avoid the mistakes in the first place!
# #### Challenge 1:
# Instructions:
#
# * open `week1_debug_challenge1.py` in `spyder 3`.
# * Attempt to run the code.
# * Fix the bugs!
# Hints:
# * Read the Python interpreter output.
# * The errors reported can look confusing at first, but read them carefully and they will point you to the lines of code with problems.
# * The `Spyder` IDE may give you some hints about formatting errors
# * It can be useful to use `print()` to display intermediate calculations and variable values.
# * Remember that `Spyder` has a variable viewer where you can look at the value of all variables created.
# * There might be multiple bugs! When you fix one and try to run the code you might find another!
#
# Have a go **yourself** and then watch our approach:
#
# * https://www.youtube.com/watch?v=XCuD59bYKx0
# ### Week 1: Do something we haven't taught you challenge
#
# Each laboratory will challenge you to do something that we haven't taught you before.
#
# No course can teach you everything you need for all programming problems. Being a competent Python programmer means that you need to learn how to find solutions to problems yourself. These challenges are designed to help you begin to use internet resources in order to solve your problem.
#
# **Before** you try the challenges it it worth watching our video on using StackOverflow:
# * https://www.youtube.com/watch?v=9WziNfkTRZ0&index=3&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE
# #### Challenge 1:
#
# You are given a unsorted `List` of integers.
#
# ```python
# [5, 7, 6, 4, 3, 2, 1]
# ```
#
# Find a command to sort the list into ascending order i.e.
#
# ```python
# [1, 2, 3, 4, 5, 6, 7]
# ```
#
# Once you have tried **yourself**. Watch our example strategy:
# * https://www.youtube.com/watch?v=369Ydv0wrGU&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE&index=5
unsorted = [5, 7, 6, 4, 3, 2, 1]
print(unsorted)
# #### Challenge 2:
#
# Sometimes a Python function needs to accept a variable number of arguments. For example, the built-in function `max()`
#
# ```python
# max(1, 2, 3)
# max(1, 2, 3, 4, 5, 6, 7, 8)
# ```
#
# Write a function that accepts a variable number of integer arguments and returns the number of arguments e.g.
#
# ```python
# result = number_of_arguments(1, 2, 3) # result = 3
# result = number_of_arguments(1, 2, 3, 4, 5) # result = 5
# ```
#
# **Try to solve this yourself.** Then watch our example strategy:
# * https://www.youtube.com/watch?v=ClyHbrkpqJU&index=6&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE
# #### Challenge 3:
#
# We have seen multiple Python functions that have required parameters.
#
# For example, the function `add_two(a, b)` **required** the user to provide two parameters `a` and `b`
#
# It is also possible in Python to have **default values** for the parameters.
#
# Write a function called `super_hero_name` that accepts two parameters of type string: `firstname` and `super_surname`.
#
# The parameter `super_surname` should have a default value of 'the spider'. The function should concatonate the names and return the resulting superhero name.
#
# E.g.
#
# ```python
# super_hero_name("tom") #returns "tom the spider"
# super_hero_name("tom", "ant-man") #returns "tom ant man"
# ```
#
# **Try this yourself first.** Then watch our approach:
# * https://www.youtube.com/watch?v=mny4iKtT21s&index=7&list=PLU2JUjGUsUm7R-3VdQA5S6IIaTK6YN1xE
# +
def super_hero_name(firstname, super_surname):
return firstname + ' ' + super_surname
super_hero_name("tom", "the spider")
# -
super_hero_name("tom", "ant-man")
# ## Optional Learning Material
#
# Download and open:
# * `string_manipulation.py` for detailed examples of how to manipulate and format Python strings.
# * `if_statement_preview.py` for a preview of if-statements that are covered in detail next week
| Labs/wk1/lab_wk1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="knOigRU1UJ9Y"
# # Estimating Auto Ownership
#
# This notebook illustrates how to re-estimate ActivitySim's auto ownership model. The steps in the process are:
# - Run ActivitySim in estimation mode to read household travel survey files, run the ActivitySim submodels to write estimation data bundles (EDB) that contains the model utility specifications, coefficients, chooser data, and alternatives data for each submodel.
# - Read and transform the relevant EDB into the format required by the model estimation package [larch](https://larch.newman.me) and then re-estimate the model coefficients. No changes to the model specification will be made.
# - Update the ActivitySim model coefficients and re-run the model in simulation mode.
#
# The basic estimation workflow is shown below and explained in the next steps.
#
# 
# -
# # Load libraries
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="s53VwlPwtNnr" outputId="d1208b7a-c1f2-4b0b-c439-bf312fe12be0"
import os
import larch # !conda install larch #for estimation
import larch.util.activitysim
import pandas as pd
# -
# # Required Inputs
#
# In addition to a working ActivitySim model setup, estimation mode requires an ActivitySim format household travel survey. An ActivitySim format household travel survey is very similar to ActivitySim's simulation model tables:
#
# - households
# - persons
# - tours
# - joint_tour_participants
# - trips (not yet implemented)
#
# Examples of the ActivitySim format household travel survey are included in the [example_estimation data folders](https://github.com/RSGInc/activitysim/tree/develop/activitysim/examples/example_estimation). The user is responsible for formatting their household travel survey into the appropriate format.
#
# After creating an ActivitySim format household travel survey, the `scripts/infer.py` script is run to append additional calculated fields. An example of an additional calculated field is the `household:joint_tour_frequency`, which is calculated based on the `tours` and `joint_tour_participants` tables.
#
# The input survey files are below.
# ### Survey households
pd.read_csv("../data_sf/survey_data/override_households.csv")
# ### Survey persons
pd.read_csv("../data_sf/survey_data/override_persons.csv")
# ### Survey tours
pd.read_csv("../data_sf/survey_data/override_tours.csv")
# ### Survey joint tour participants
pd.read_csv("../data_sf/survey_data/survey_joint_tour_participants.csv")
# # Example Setup if Needed
#
# To avoid duplication of inputs, especially model settings and expressions, the `example_estimation` depends on the `example`. The following commands create an example setup for use. The location of these example setups (i.e. the folders) are important because the paths are referenced in this notebook. The commands below download the skims.omx for the SF county example from the [activitysim resources repository](https://github.com/RSGInc/activitysim_resources).
# !activitysim create -e example_estimation_sf -d test
# + [markdown] colab_type="text" id="5lxwxkOuZvIy"
# # Run the Estimation Example
#
# The next step is to run the model with an `estimation.yaml` settings file with the following settings in order to output the EDB for all submodels:
#
# ```
# enable=True
#
# bundles:
# - school_location
# - workplace_location
# - auto_ownership
# - free_parking
# - cdap
# - mandatory_tour_frequency
# - mandatory_tour_scheduling
# - joint_tour_frequency
# - joint_tour_composition
# - joint_tour_participation
# - joint_tour_destination
# - joint_tour_scheduling
# - non_mandatory_tour_frequency
# - non_mandatory_tour_destination
# - non_mandatory_tour_scheduling
# - tour_mode_choice
# - atwork_subtour_frequency
# - atwork_subtour_destination
# - atwork_subtour_scheduling
# - atwork_subtour_mode_choice
#
# survey_tables:
# households:
# file_name: survey_data/override_households.csv
# index_col: household_id
# persons:
# file_name: survey_data/override_persons.csv
# index_col: person_id
# tours:
# file_name: survey_data/override_tours.csv
# joint_tour_participants:
# file_name: survey_data/override_joint_tour_participants.csv
# ```
#
# This enables the estimation mode functionality, identifies which models to run and their output estimation data bundles (EDBs), and the input survey tables, which include the override settings for each model choice.
#
# With this setup, the model will output an EBD with the following tables for this submodel:
# - model settings - auto_ownership_model_settings.yaml
# - coefficients - auto_ownership_coefficients.csv
# - utilities specification - auto_ownership_SPEC.csv
# - chooser and alternatives data - auto_ownership_values_combined.csv
#
# The following code runs the software in estimation mode, inheriting the settings from the simulation setup and using the San Francisco county data setup. It produces the EDB for all submodels but runs all the model steps identified in the inherited settings file.
# -
# %cd test
# !activitysim run -c configs_estimation/configs -c configs -o output -d data_sf
# # Read the EDB
#
# The next step is to read the EDB, including the coefficients, model settings, utilities specification, and chooser and alternative data.
# +
edb_directory = "output/estimation_data_bundle/auto_ownership/"
def read_csv(filename, **kwargs):
return pd.read_csv(os.path.join(edb_directory, filename), **kwargs)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="cBqPPkBpnaUZ" outputId="bd780019-c200-4cf6-844a-991c4d026480"
coefficients = read_csv("auto_ownership_coefficients.csv", index_col='coefficient_name')
spec = read_csv("auto_ownership_SPEC.csv")
chooser_data = read_csv("auto_ownership_values_combined.csv")
# -
# ### Coefficients
coefficients
# #### Utility specification
spec
# ### Chooser and alternatives data
chooser_data
# ### Remove choosers with invalid observed choice
chooser_data = chooser_data[chooser_data['override_choice'] >= 0]
# # Data Processing and Estimation Setup
#
# The next step is to transform the EDB for larch for model re-estimation.
# +
from larch import P, X
altnames = list(spec.columns[3:])
altcodes = range(len(altnames))
# -
m = larch.Model()
# One of the alternatives is coded as 0, so
# we need to explicitly initialize the MNL nesting graph
# and set to root_id to a value other than zero.
m.initialize_graph(alternative_codes=altcodes, root_id=99)
# ### Utility specifications
m.utility_co = larch.util.activitysim.dict_of_linear_utility_from_spec(
spec, 'Label', dict(zip(altnames,altcodes)),
)
m.utility_co
larch.util.activitysim.apply_coefficients(coefficients, m)
# ### Coefficients
m.pf
# ### Availability
av = True # all alternatives are available
d = larch.DataFrames(
co=chooser_data,
alt_codes=altcodes,
alt_names=altnames,
av=av,
)
m.dataservice = d
# ### Survey choice
m.choice_co_code = 'override_choice'
# # Estimate
#
# With the model setup for estimation, the next step is to estimate the model coefficients. Make sure to use a sufficiently large enough household sample and set of zones to avoid an over-specified model, which does not have a numerically stable likelihood maximizing solution. Larch has two built-in estimation methods: BHHH and SLSQP. BHHH is the default and typically runs faster, but does not follow constraints on parameters. SLSQP is safer, but slower, and may need additional iterations.
# m.estimate(method='SLSQP', options={'maxiter':1000})
m.estimate(method='BHHH', options={'maxiter':1000})
# ### Estimated coefficients
m.parameter_summary()
# + [markdown] colab_type="text" id="TojXWivZsx7M"
# # Output Estimation Results
# -
est_names = [j for j in coefficients.index if j in m.pf.index]
coefficients.loc[est_names,'value'] = m.pf.loc[est_names, 'value']
os.makedirs(os.path.join(edb_directory,'estimated'), exist_ok=True)
# ### Write the re-estimated coefficients file
coefficients.reset_index().to_csv(
os.path.join(edb_directory,'estimated',"auto_ownership_coefficients_revised.csv"),
index=False,
)
# ### Write the model estimation report, including coefficient t-statistic and log likelihood
m.to_xlsx(
os.path.join(edb_directory,'estimated',"auto_ownership_model_estimation.xlsx"), data_statistics=False
)
# # Next Steps
#
# The final step is to either manually or automatically copy the `auto_ownership_coefficients_revised.csv` file to the configs folder, rename it to `auto_ownership_coeffs.csv`, and run ActivitySim in simulation mode.
pd.read_csv(os.path.join(edb_directory,'estimated',"auto_ownership_coefficients_revised.csv"))
| activitysim/examples/example_estimation/notebooks/estimating_auto_ownership.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sprintenv
# language: python
# name: sprintenv
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as seabornInstance
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
from sklearn import metrics
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.datasets import make_regression
# %matplotlib inline
# # simple linear regression using SciKit Learn
# ## Create a simple random dataset
X, Y, w = make_regression(n_samples=30, n_features=1, coef=True,
random_state=1)
# +
plt.scatter(X, Y, s=10)
plt.title("random samples with one feature")
plt.xlabel('X')
plt.ylabel('Y')
plt.show()
print("coefficient :", w)
# +
# Split the data into training/testing sets
X_train = X[:-5]
X_test = X[-5:]
# Split the targets into training/testing sets
Y_train = Y[:-5]
Y_test = Y[-5:]
# -
# train model
model = LinearRegression()
model.fit(X_train, Y_train)
# +
# evaluating model
Y_pred = model.predict(X_test)
# make predictions
pred_df = pd.DataFrame({'Actual': Y_test.flatten(), 'Predicted': Y_pred.flatten()})
pred_df.head()
# -
# compute RMSE and determination coeficient (R2)
rmse = np.sqrt(mean_squared_error(Y_test,y_pred))
r2 = r2_score(Y_test,y_pred)
print("RMSE:", rmse)
print("R2:", r2)
plt.scatter(X, Y, s=10)
plt.plot(X, y_pred, color='r')
plt.title("random samples with one feature")
plt.xlabel('X')
plt.ylabel('Y')
plt.show()
print('predicted coef:', model.coef_[0])
# # Multiple linear regression using SciKit Learn
# ## create random dataset
# +
# random dataset with 5 linear features
X, Y, w = make_regression(n_samples=30, n_features=5, coef=True,
random_state=1, bias=3.5)
X_df = pd.DataFrame(X)
X_df.head()
# +
# Split the data into training/testing sets
X_train = X[:-5]
X_test = X[-5:]
# Split the targets into training/testing sets
Y_train = Y[:-5]
Y_test = Y[-5:]
# -
model = LinearRegression()
model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
# +
# evaluating model
Y_pred = model.predict(X_test)
# make predictions
pred_df = pd.DataFrame({'Actual': Y_test.flatten(), 'Predicted': Y_pred.flatten()})
pred_df.head()
# -
rmse = np.sqrt(mean_squared_error(Y_test,Y_pred))
r2 = r2_score(Y_test,Y_pred)
print("RSME:", rmse)
print("R2:", r2)
# look at predicted coefficients
predC_df = pd.DataFrame({'Actual': w.flatten(), 'Predicted': model.coef_.flatten()})
predC_df.head()
# # Linear regression on red wine dataset
# ## Import dataset
# import text documents from wikipedia abstracts
wine_data=pd.read_csv('winequality/winequality-red.csv',delimiter=';')
len(wine_data)
wine_data.head()
wine_data.describe()
# Look at Null values in datasets
wine_data.isnull().any()
# all the columns should give False, In case for any column you find True result,
# then remove all the null values from that column using :
# wine_data = wine_data.fillna(method='ffill')
plt.figure(figsize=(15,10))
plt.tight_layout()
seabornInstance.distplot(wine_data['quality'])
# +
# shuffle the rows of the dataframe
wine_data = wine_data.sample(frac=1).reset_index(drop=True)
# define the data/predictors as the pre-set feature names
df = wine_data.drop("quality", axis=1)
# Put the target (wine quality -- quality) in another DataFrame
target = pd.DataFrame(wine_data, columns=["quality"])
# -
# ## Compute simple regression
plt.scatter(x=wine_data['chlorides'], y=wine_data['quality'], s=10)
plt.title('Chlorides vs quality')
plt.xlabel('Chlorides')
plt.ylabel('quality')
plt.show()
# +
# compute a simple regression on a single input
X = df["chlorides"].values.reshape(-1,1)
y = target["quality"].values.reshape(-1,1)
# Split the data into training/testing sets
X_train = X[:-50]
X_test = X[-50:]
# Split the targets into training/testing sets
Y_train = y[:-50]
Y_test = y[-50:]
# +
# Create linear regression object
regr = LinearRegression()
# Train the model using the training sets
regr.fit(X_train, Y_train)
# Make predictions using the testing set
Y_pred = regr.predict(X_test)
# Print the coefficient
print('Coefficients:', regr.coef_[0])
# Print the intercept
print('Intercept:', regr.intercept_[0])
rmse = np.sqrt(mean_squared_error(Y_test,Y_pred))
r2 = r2_score(Y_test,Y_pred)
print("RSME:", rmse)
print("R2:", r2)
# +
# Plot outputs
plt.scatter(x=wine_data['chlorides'], y=wine_data['quality'], s=10)
plt.plot(X_test, Y_pred, color='blue', linewidth=3)
plt.title('chlorides vs quality')
plt.xlabel('chlorides')
plt.ylabel('quality')
plt.show()
# -
# make predictions
pred_df = pd.DataFrame({'Actual': Y_test.flatten(), 'Predicted': Y_pred.flatten()})
pred_df.head()
# # Multiple regression
# +
# compute a multiple regression
X = df.values
y = target["quality"].values
columns_name = ['fixed acidity', 'volatile acidity', 'citric acid', 'residual sugar', 'chlorides', 'free sulfur dioxide',
'total sulfur dioxide', 'density', 'pH', 'sulphates','alcohol']
# Split the data into training/testing sets
X_train = X[:-50]
X_test = X[-50:]
# Split the targets into training/testing sets
Y_train = y[:-50]
Y_test = y[-50:]
# Create linear regression object
regr = LinearRegression()
# Train the model using the training sets
regr.fit(X_train, Y_train)
# Make predictions using the testing set
Y_pred = regr.predict(X_test)
df_pred = pd.DataFrame({'Actual': Y_test, 'Predicted': Y_pred})
df_pred.head()
# -
# The coefficients
coeff_df = pd.DataFrame(regr.coef_, columns_name)
print(coeff_df)
# +
df1=df_pred.head(25)
df1.plot(kind='bar',figsize=(10,8))
plt.show()
# -
rmse = np.sqrt(mean_squared_error(Y_test,Y_pred))
r2 = r2_score(Y_test,Y_pred)
print("RSME:", rmse)
print("R2:", r2)
| sprint2_classification/.ipynb_checkpoints/sprint1_demo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import wandb
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from dotenv import load_dotenv
load_dotenv("../.env")
# + pycharm={"name": "#%%\n"}
########################################################################################
# query data from wandb
# add data to frames
api = wandb.Api()
data_description_map = {
"tiny-few": "tiny-few (50k files shared between 100 speakers)",
"tiny-high": "tiny-high (8 files from 8 sessions for 5994 speakers)",
}
def load_runs(name: str):
runs = api.runs(name)
df = pd.DataFrame(columns=["eer", "data", "ablation"])
for r in runs:
tags = r.tags
if r.state != "finished":
continue
eer = r.summary["test_eer_hard"]
if "tiny_few" in tags:
tags.remove("tiny_few")
data = "tiny_few"
elif "tiny_many_high" in tags:
tags.remove("tiny_many_high")
data = "tiny_many_high"
else:
raise ValueError(f"undetermined dataset from {tags=}")
ablation = tags[0]
df = pd.concat(
[
df,
pd.DataFrame(
{
"ablation": [ablation],
"eer": [eer],
"data": [data],
}
),
],
ignore_index=True,
)
return df
df = load_runs("wav2vec2-ablation")
# + pycharm={"name": "#%%\n"}
df
# + pycharm={"name": "#%%\n"}
df_grouped = df.groupby(by=["data", "ablation",])
# + pycharm={"name": "#%%\n"}
df_agg = df_grouped.agg(
eer_min=("eer", "min"),
eer_max=("eer", "max"),
eer_mean=("eer", "mean"),
eer_std=("eer", "std"),
count=("eer", "count")
)
# + pycharm={"name": "#%%\n"}
df_agg
# + pycharm={"name": "#%%\n"}
df_agg['count'].sum()
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
| result_analysis/plot_ablation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# name: python3
# ---
# # Interval List Intersections
# You are given two lists of closed intervals, first_list and second_list, where first_list[i] = [starti, endi] and second_list[j] = [startj, endj]. Each list of intervals is pairwise disjoint and in sorted order.
# Return the intersection of these two interval lists.
# A closed interval [a, b] (with a <= b) denotes the set of real numbers x with a <= x <= b.
# The intersection of two closed intervals is a set of real numbers that are either empty or represented as a closed interval. For example, the intersection of [1, 3] and [2, 4] is [2, 3].
# +
'''
Example 1
Input: first_list = [[0,2],[5,10],[13,23],[24,25]], second_list = [[1,5],[8,12],[15,24],[25,26]]
Output: [[1,2],[5,5],[8,10],[15,23],[24,24],[25,25]]
Example 2:
Input: first_list = [[1,3],[5,9]], second_list = []
Output: []
'''
def interval_intersection(first_list: list, second_list: list) -> list:
def find_intersection(first_list, second_list):
first, second = first_list[0], second_list[0]
left, right = (first, second) if first[0] < second[0] else (second, first)
if left[1] >= right[0]:
return [right[0], min(left[1], right[1])]
else:
return None
res = []
while first_list and second_list:
intersect = find_intersection(first_list, second_list)
if intersect:
res.append(intersect)
if first_list[0][1] < second_list[0][1]:
first_list = first_list[1:]
else:
second_list = second_list[1:]
return res
| python-data-structures/interview-fb/interval-list-intersections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example - Interpolate Missing Data
# +
import rioxarray # for the extension to load
import xarray
# %matplotlib inline
# -
# ## Load in xarray dataset
xds = xarray.open_dataarray("MODIS_ARRAY.nc")
xds
xds.isel(x=slice(0, 20), y=slice(0, 20)).plot()
# ## Fill misssing with interpolate_na
#
# API Reference:
#
# - DataArray: [rio.interpolate_na()](../rioxarray.rst#rioxarray.raster_array.RasterArray.interpolate_na)
# - Dataset: [rio.interpolate_na()](../rioxarray.rst#rioxarray.raster_dataset.RasterDataset.interpolate_na)
filled = xds.rio.interpolate_na()
filled
filled.isel(x=slice(0, 20), y=slice(0, 20)).plot()
| docs/examples/interpolate_na.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "88399a27-9868-485f-9cb1-7b225a16e33a"}
# # Short Introduction
# They showed people two of 84 candies and ask them which one they would choose. For each candy, they calculated how often it was chosen (in percent). So the dataset includes the name of each candy, some attributes like sugar content, whether it contains nougat or caramel and so on, and the probability that it was chosen in the survey.
#
# Therefore we focus on this probability (name of the feature in the dataset: winpercent).
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "86fd4150-0af0-41d0-b80d-b9b0c5fe711a"}
# # Overview
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "03610d76-4eec-43b7-b495-5324d7fd84fa"}
# First of all we load all needed packages.
# + _kg_hide-input=false application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "108046d0-324d-48ad-9dd7-c360b2ccdbb3"} _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
# %run ./Libraries
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "a0685391-afd2-4e49-b14f-c49a4f8441ad"}
# # Multivariate Analysis
#
# Now we are using machine learning to check if we find some non-linear dependencies.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "f86906cd-543b-4365-9af9-02bca385c580"}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/candy-power-ranking/candy-data.csv')
df['competitorname'] = df.competitorname.apply(lambda l: l.replace('Õ', '\''))
display(df)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "12fb8460-3112-4530-8d2f-dfb94cdd3baf"}
# ## Baseline
#
# A baseline model is a simple estimator, which give us an idea, how good our predictores should be at least and to see how much better they are.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "14fefdc8-5b6d-4298-8bcc-68de2cd39a06"}
features = df.columns[1:-1]
target = 'winpercent'
class MeanEstimator(BaseEstimator):
mean = None
def fit(self, X, y):
if isinstance(y, pd.Series) or isinstance(y, pd.DataFrame):
self.mean = np.mean(y).iloc[0]
else:
self.mean = np.mean(y)
def predict(self, X):
if self.mean is None:
raise ValueError('Estimator is not fitted yet')
return np.ones(X.shape[0])*self.mean
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4a2826bf-63ef-4019-ad17-bad86c7256b8"}
model_mean = MeanEstimator()
pred_mean = cross_val_predict(model_mean, df[features], df[[target]], cv=4, n_jobs=4)
dict(r2=r2_score(df[[target]], pred_mean),
rmse=sqrt(mean_squared_error(df[[target]], pred_mean))
)
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "be9c27de-6c5a-4255-bd73-34ba862fdd54"}
# The RSME of a better estimator should be less than the above value. And r² should be better than 0 of course,
#
# Perhapse you wonder why r² is negative. In general r² is between minus infinity and 1 (the square in r² is misleading). If you compute the r² of the target and the mean of the target you get exactly 0:
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "854e8f2a-190a-4e96-825c-e3062bb397fc"}
r2_score(df[[target]], np.ones(df.shape[0])*np.mean(df[[target]]).iloc[0])
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "203a6fe5-052f-4b53-9b7d-777153ca0fe2"}
# But we are using cross validation. So we compute the mean over the target of a train set and using this value as an estimator for the test set. This value can be a worse estimator than the mean of the target in the test set (which we do not know beforehand). In this case the r² score is less than 0.
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "6037af53-5885-4c73-a6f2-0e747bd31534"}
# ## XGBRegressor
#
# The XGBRegressor is the regressor of the xgboost package, a famous implementation of a gradient boosting algorithm. Unlike a classifier a regressor takes an interval variable as target.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "1d0acd54-bd54-474b-927d-858b191049c7"}
param_grid = dict(
max_depth=[3, 4, 5],
learning_rate=[0.05, 0.1, 0.2],
n_estimators=[32, 33, 34],
min_child_weight=[5, 6, 7],
subsample=[0.4, 0.5, 0.6],
)
clf = xgboost.XGBRegressor(objective='reg:squarederror')
model_xgb = GridSearchCV(estimator=clf, param_grid=param_grid, n_jobs=4, iid=True, refit=True, cv=4, scoring='r2')
model_xgb.fit(df[features], df[[target]])
pred_xgb = cross_val_predict(model_xgb.best_estimator_, df[features], df[[target]], cv=4, n_jobs=4)
dict(r2=r2_score(df[[target]], pred_xgb), rmse=sqrt(mean_squared_error(df[[target]], pred_xgb)))
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d6e7ef2f-a82e-4731-8a85-418619693e5d"}
# The best parameters are the following parameters
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7106ee46-5cac-4de3-a1fc-4b4f38653ace"}
model_xgb.best_params_
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "d0a1adfa-2618-4e18-a7bb-2d16afe951a4"}
# The feature importance:
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "dad5694b-4d4e-4bb1-b5c5-5ae44adaefac"}
imp = pd.DataFrame({'features': features, 'importance': model_xgb.best_estimator_.feature_importances_})
imp.sort_values('importance', ascending=False, inplace=True)
sns.barplot(x='importance', y='features', data=imp, palette="YlGn_r")
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "23a91947-3d60-40eb-a3fc-577be09190a9"}
# We see that chocolate is the most important feature
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "98be3eef-2320-41a5-88b8-03103c2cb8f2"}
# ## RandomForestRegressor
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "0756d1a8-bb87-4df1-9a9d-d59251efe8c1"}
features = df.columns[1:-1]
target = 'winpercent'
param_grid = {
'max_depth': [None, 10, 15, 20],
'max_features': ['auto', 'sqrt', 'log2'],
'min_samples_leaf': [6, 7, 8, 9],
'min_samples_split': [10, 11, 12, 13],
'n_estimators': [130, 140, 150, 160]
}
clf_rf = RandomForestRegressor()
model_rf = GridSearchCV(estimator=clf_rf, param_grid=param_grid, n_jobs=4, iid=True, refit=True, cv=4, scoring='r2')
model_rf.fit(df[features], np.ravel(df[[target]]))
pred_rf = cross_val_predict(model_rf.best_estimator_, df[features], df[[target]], cv=4, n_jobs=4)
dict(r2=r2_score(df[[target]], pred_rf), rmse=sqrt(mean_squared_error(df[[target]], pred_rf)))
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "7b26c7ec-002c-4d59-ab15-b591d29ac8fd"}
# The best parameters are the following parameters:
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "e368d927-9f3e-4a63-a30d-497e45941466"}
model_rf.best_params_
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "76004811-d4b1-4c4b-a759-fb2a009432dd"}
# The feature importance:
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "93e27f57-ff43-4af5-8ae9-1bb69f7e036d"}
imp = pd.DataFrame({'features': features, 'importance': model_rf.best_estimator_.feature_importances_})
imp.sort_values('importance', ascending=False, inplace=True)
sns.barplot(x='importance', y='features', data=imp, palette="YlGn_r")
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "c2926c12-25c0-477c-ae48-c858f87dfd72"}
# In the following we take the look on the performance on the random forest prediction.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "13bf0056-0380-415f-8177-0dcf0d3db0df"}
df_rf = df.copy()
df_rf['pred_rf'] = pred_rf
sns_dt = df_rf.sort_values('pred_rf', ascending=False).reset_index(drop=True).winpercent.expanding().mean()
ax = sns.lineplot(x=sns_dt.index, y=sns_dt, color='green', palette="YlGn")
ax.set(xlabel='Candies (ordered by predicted winpercent)', ylabel='Winpercent (cumulative mean)')
plt.show()
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "77c28f57-eb59-4277-87a0-45cc54ef3f77"}
# The performance is not really good. In particular if we take the findings from the univariate analysis we get better results.
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "72b0ad6d-e387-42ee-a7a3-d2073ced43c0"}
# ## XGBRegressor without chocolate
#
# If we remove chocolate from the features the model performs really bad and it performs similar to the baseline model above.
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "3b7eac7c-83a2-4307-9c73-ca191c1e50bd"}
features_no_choc = df.columns[2:-1]
param_grid = dict(
max_depth=[3, 4, 5],
learning_rate=[0.05, 0.1, 0.2],
n_estimators=[32, 33, 34],
min_child_weight=[5, 6, 7],
subsample=[0.4, 0.5, 0.6],
)
clf = xgboost.XGBRegressor(objective='reg:squarederror')
model_xgb_no_choc = GridSearchCV(estimator=clf, param_grid=param_grid, n_jobs=4, iid=True, refit=True, cv=4, scoring='r2')
model_xgb_no_choc.fit(df[features_no_choc], df[[target]])
pred_xgb_no_choc = cross_val_predict(model_xgb_no_choc.best_estimator_, df[features_no_choc], df[[target]], cv=4, n_jobs=4)
dict(r2=r2_score(df[[target]], pred_xgb_no_choc),
rmse=sqrt(mean_squared_error(df[[target]], pred_xgb_no_choc)))
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "4e478906-da8c-4c17-b16c-91c08a8e12b9"}
imp = pd.DataFrame({'features': features_no_choc, 'importance': model_xgb_no_choc.best_estimator_.feature_importances_})
imp.sort_values('importance', ascending=False, inplace=True)
sns.barplot(x='importance', y='features', data=imp, palette="YlGn_r")
# + [markdown] application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "aefd8179-7e49-4051-b4f4-9211f18add16"}
# # Summary
#
# Multivariate analysis also confirms the findings of the univariate analysis and does not lead to more relevant insights. Chocolate is the most important feature and if taken out it is peanuty almond
# + application/vnd.databricks.v1+cell={"title": "", "showTitle": false, "inputWidgets": {}, "nuid": "cd4f0904-9ddf-4695-b5f0-c6d8bc706296"}
| notebooks/3. Single Dataset - Multivariate Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Семинар по numpy
#
# В этом семинаре мы познакомимся с первой библиотекой, которая понадобится нам в курсе - библиотекой numpy. Это библиотека для работы с матрицами, и она может быть также полезна для матричных вычислений в других дисциплинах. Но сначала познакомимся с интерфейсом jupyter notebook. Это инструмент, позволяющий писать код в интерактивном режиме, рисовать графики и оформлять материалы.
#
# ## Очень краткий обзор интерфейса jupyter notebook
# Jupyter notebook состоит из ячеек.
# В ячейку можно записывать код, выполнять, а потом использовать созданные переменные / определенные функции и т. д. в других ячейках:
x = [1, 2, 3]
print(x)
x = 9
y = 3
# Выполнить ячейку: shift + enter, показать аргументы функции: shift + tab. Есть много других горячих клавиш, см. Help -> Keyboard Shortcuts (вверху интерфейса). Там же есть Help -> User Interface Tool!
#
# Обратите внимание на кнопки + (добавить ячейку), ножницы (удалить ячейку), стрелки, меняющие ячейки местами, кнопку остановить выполнение.
#
# Ноутбук сохраняется автоматически. Чтобы скачать: File -> Download as -> ipynb.
#
# Этот текст написан в ячейке типа Markdown, она позволяет красиво оформлять код. Переключение типа ячейки справа от кнопки стоп (черный квадрат). Ячейки с кодом имеют тип Code.
# ## Создание массивов в numpy
#
# Numpy - библиотека для работы с матрицами. Импортируем библиотеку:
import numpy as np
# Предположим, что мы провели опрос, в котором было четыре вопроса, на каждый можно ответить Да, Нет, Воздержался. В итоге получим таблицу размера 4 на 3. Создадим такую таблицу в numpy:
l = [[30, 2, 0], [3, 27, 1], [28, 1, 1], [6, 17, 5]]
ar = np.array(l)
ar # выводится созданное в последней строке, если в ней нет присваивания (знак =)
# Можно создавать массивы из нулей (np.zeros) или из единиц (np.ones):
A = np.ones((7, 9))
A
# Также часто используется создание векторов из натуральных чисел:
vec = np.arange(15)
vec
# И случайно сгенерированные массивы:
r = np.random.rand(3, 5)
r
# ### Размерности массивов
# Размерности:
ar.shape[0]
A.shape
vec.shape
# В numpy нулевая размерность отвечает за строки, первая - за столбцы. В нашей матрице ar 4 строки и 3 столбца. Вообще в numpy можно создавать массивы любых размерностей точно таким же образом, как мы сделали выше - numpy не различает векторы, матрицы и тензоры, все они называются массивами. Например, vec имеет длину 15 только по одной размерности, поэтому его shape равен (15,). Shape - это обычный кортеж языка python.
# Можно вытянуть все ответы в одну строку:
ar.ravel()
# Можно наоборот: преобразовать вектор в матрицу:
vec.reshape(3, 5)
# Обратите внимание, что числа записываются по строкам.
vec.reshape(3, 5).shape
# По одной из осей можно указывать -1, тогда библиотека сама посчитает число элементов по этой размерности:
vec.reshape(3, -1)
# Аналогичным образом можно дублироать функционал функции ravel:
vec.reshape(-1)
# ### Операции с массивами
# Можно выделить три группы операций с массивами в numpy:
# * поэлементные
# * матричные
# * агрегирующие
# Поэлементные выполняются между массивами одной формы (shape), хотя ниже мы обсудим некое обощение правила одной формы.
vec + vec + vec
A * 10
x = np.array([1, 2, 3])
y = np.array([-1, 1, -1])
x * y
# Обратите внимание, что * - это поэлементное умножение, а не матричное!
np.exp(x)
# Матричные операции - операции из линейной алгебры. Например, матричное произведение:
A = np.random.rand(7, 8)
B = np.random.rand(8, 3)
A.dot(B)
# Можно писать и так:
np.dot(A, B)
# И так:
A @ B
# Проверим форму:
A.dot(B).shape
# Обращение матрицы:
np.linalg.inv(np.random.rand(3, 3))
# Модуль np.linalg содержит много полезных матричных функций, их можно посмотреть в документации модуля.
# Агрегирующие операции агрерируют информацию в троках, столбцах, во всем массиве и т. д. Самые популярные такие операции - суммирование np.sum, усреднение np.mean, медиана np.median, максимум np.max и минимум np.min.
# Число полученных ответов на вопросы (всего):
np.sum(ar)
# Пробуем выяснить число респондентов. Для этого просуммируем матрицу по строкам (это делается с помощью указания axis=1):
np.sum(ar, axis = 1)
np.sum(ar, axis = 1).shape
# По столбцам: axis=0, по строкам: axis=1.
#
# В результате суммирования получился вектор (размерность на 1 меньше, чем у исходной матрицы). Можно указать keepdims=True, чтобы сохранть размерности:
np.sum(ar, axis = 1, keepdims=True).shape
# Задание для студентов: посчитать сумму по строкам, используя матричное произведение.
np.sum(A.dot(B), axis=1)
# Считаем число ответов "да", "нет", "воздержался" двумя способами:
np.sum(ar, axis=0)
ones = np.ones(4)
np.dot(ones, ar)
# ### Индексация
# Для индексации ставим [ ] и через запятую перечисляем действия с осями. В матрице 0 - по вертикали, 1 - по горизонтали
ar[1, 1] # выбрать 1 элемент
ar # вывели для проверки
ar[:, 2] # выделить столбец
ar[:, -1] # выделить последний столбец
ar[0] # выделить строку
ar[:, ::2] # выделить все столбы с четными номерами
# Можно делать логическую индексацию, чтобы выбирались только те элементы, для которых стоит True. Выберем ответы на вопросы с номерами, кратными 2:
ar[np.arange(ar.shape[0])%2==0]
# ### Добавление оси
#
# Для удобного перехода между размерностями используют добавление оси. Вектор можно сделать матрицей с размером 1 по одной из размерностей.
ones[:, np.newaxis]
ones[:, np.newaxis].shape
# вместо вектора с формой (4,) стала матрциа с формой (4, 1)
ones[np.newaxis, :]
# вместо вектора с формой (4,) стала матрциа с формой (1, 4)
ones[np.newaxis, :].shape
# ### Добавление оси в поэлементных операциях
# В поэлементных операциях можно использовать не только массивы в одинаковым в точности размером. В общем виде условие такое: по каждой размерности либо размер совпадает, либо в одном из массивов размер 1. Например, матрица размера (4, 3) и вектор размера (4, 1). В этом случае при выполнении операции столбец будет как бы "дублироваться" для каждого столбца в первой матрице. Воспользуемся этим, чтобы найти долю каждого ответа на все вопросы:
sums = ar.sum(axis=1) # всего ответов на каждый вопрос
sums.shape
sums[:, np.newaxis].shape # добавили ось
ar / sums[:, np.newaxis] # поделили число каждого варианта на общее число ответов на вопрос
# ### Объединение массивов
# Добавляем новый вопрос в таблицу:
row = np.array([5, 12, 15])
row = row[np.newaxis, :]
# конкретно тут можно без увеличения размерности
# но в других случаях может быть ошибка, лучше добавлять
ar = np.vstack((ar, row))
# Добавляем новый столбец в таблицу - максимальное число ответов:
mx = np.max(ar, 1)
mx
mx.shape
mx = mx[:, np.newaxis]
ar = np.hstack ((ar, mx))
ar
# Удаление строки (аналогично можно удалять столбец):
np.delete(ar, np.arange(3, 4), axis=1)
# ### Задания для студентов
# Выделите строки, у которых ответов "нет" больше, чем ответов "да":
ar[ar[:, 0]<ar[:, 1]]
# Вывести квадраты первых десяти натуральных чисел:
np.arange(10)**2
# Перемешать числа натуральные числа от 1 до 10 (воспользуйтесь np.random.permutation):
np.random.permutation(10)
# student's code here
# Составить таблицу умножения от 1 до 10:
p = np.arange(1, 10)
p * p[:, None]
| numpy_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Training and Deploying the Fraud Detection Model
#
# In this notebook, we will take the outputs from the Processing Job in the previous step and use it and train and deploy an XGBoost model. Our historic transaction dataset is initially comprised of data like timestamp, card number, and transaction amount and we enriched each transaction with features about that card number's recent history, including:
#
# - `num_trans_last_10m`
# - `num_trans_last_1w`
# - `avg_amt_last_10m`
# - `avg_amt_last_1w`
#
# Individual card numbers may have radically different spending patterns, so we will want to use normalized ratio features to train our XGBoost model to detect fraud.
# ### Imports
from sklearn.model_selection import train_test_split
from sagemaker.inputs import TrainingInput
from sagemaker.session import Session
from sagemaker import image_uris
import pandas as pd
import numpy as np
import sagemaker
import boto3
import io
# ### Essentials
# +
LOCAL_DIR = './data'
BUCKET = sagemaker.Session().default_bucket()
PREFIX = 'training'
sagemaker_role = sagemaker.get_execution_role()
s3_client = boto3.Session().client('s3')
# -
# First, let's load the results of the SageMaker Processing Job ran in the previous step into a Pandas dataframe.
df = pd.read_csv(f'{LOCAL_DIR}/aggregated/processing_output.csv')
#df.dropna(inplace=True)
df['cc_num'] = df['cc_num'].astype(np.int64)
df['fraud_label'] = df['fraud_label'].astype(np.int64)
df.head()
len(df)
# ### Split DataFrame into Train & Test Sets
# The artifically generated dataset contains transactions from `2020-01-01` to `2020-06-01`. We will create a training and validation set out of transactions from `2020-01-15` and `2020-05-15`, discarding the first two weeks in order for our aggregated features to have built up sufficient history for cards and leaving the last two weeks as a holdout test set.
# +
training_start = '2020-01-15'
training_end = '2020-05-15'
training_df = df[(df.datetime > training_start) & (df.datetime < training_end)]
test_df = df[df.datetime >= training_end]
test_df.to_csv(f'{LOCAL_DIR}/test.csv', index=False)
# -
# Although we now have lots of information about each transaction in our training dataset, we don't want to pass everything as features to the XGBoost algorithm for training because some elements are not useful for detecting fraud or creating a performant model:
# - A transaction ID and timestamp is unique to the transaction and never seen again.
# - A card number, if included in the feature set at all, should be a categorical variable. But we don't want our model to learn that specific card numbers are associated with fraud as this might lead to our system blocking genuine behaviour. Instead we should only have the model learn to detect shifting patterns in a card's spending history.
# - Individual card numbers may have radically different spending patterns, so we will want to use normalized ratio features to train our XGBoost model to detect fraud.
#
# Given all of the above, we drop all columns except for the normalised ratio features and transaction amount from our training dataset.
training_df.drop(['tid','datetime','cc_num','num_trans_last_10m', 'avg_amt_last_10m',
'num_trans_last_1w', 'avg_amt_last_1w'], axis=1, inplace=True)
# The [built-in XGBoost algorithm](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html) requires the label to be the first column in the training data:
training_df = training_df[['fraud_label', 'amount', 'amt_ratio1','amt_ratio2','count_ratio']]
training_df.head()
train, val = train_test_split(training_df, test_size=0.3)
train.to_csv(f'{LOCAL_DIR}/train.csv', header=False, index=False)
val.to_csv(f'{LOCAL_DIR}/val.csv', header=False, index=False)
# !aws s3 cp {LOCAL_DIR}/train.csv s3://{BUCKET}/{PREFIX}/
# !aws s3 cp {LOCAL_DIR}/val.csv s3://{BUCKET}/{PREFIX}/
# +
# initialize hyperparameters
hyperparameters = {
"max_depth":"5",
"eta":"0.2",
"gamma":"4",
"min_child_weight":"6",
"subsample":"0.7",
"objective":"binary:logistic",
"num_round":"100"}
output_path = 's3://{}/{}/output'.format(BUCKET, PREFIX)
# this line automatically looks for the XGBoost image URI and builds an XGBoost container.
# specify the repo_version depending on your preference.
xgboost_container = sagemaker.image_uris.retrieve("xgboost", sagemaker.Session().boto_region_name, "1.2-1")
# construct a SageMaker estimator that calls the xgboost-container
estimator = sagemaker.estimator.Estimator(image_uri=xgboost_container,
hyperparameters=hyperparameters,
role=sagemaker.get_execution_role(),
instance_count=1,
instance_type='ml.m5.2xlarge',
volume_size=5, # 5 GB
output_path=output_path)
# define the data type and paths to the training and validation datasets
content_type = "csv"
train_input = TrainingInput("s3://{}/{}/{}".format(BUCKET, PREFIX, 'train.csv'), content_type=content_type)
validation_input = TrainingInput("s3://{}/{}/{}".format(BUCKET, PREFIX, 'val.csv'), content_type=content_type)
# execute the XGBoost training job
estimator.fit({'train': train_input, 'validation': validation_input})
# -
# Ideally we would perform hyperparameter tuning before deployment, but for the purposes of this example will deploy the model that resulted from the Training Job directly to a SageMaker hosted endpoint.
predictor = estimator.deploy(
initial_instance_count=1,
instance_type='ml.t2.medium',
serializer=sagemaker.serializers.CSVSerializer(), wait=True)
endpoint_name=predictor.endpoint_name
#Store the endpoint name for later cleanup
# %store endpoint_name
endpoint_name
# Now to check that our endpoint is working, let's call it directly with a record from our test hold-out set.
payload_df = test_df.drop(['tid','datetime','cc_num','fraud_label','num_trans_last_10m', 'avg_amt_last_10m',
'num_trans_last_1w', 'avg_amt_last_1w'], axis=1)
payload = payload_df.head(1).to_csv(index=False, header=False).strip()
payload
float(predictor.predict(payload).decode('utf-8'))
# ## Show that the model predicts FRAUD / NOT FRAUD
count_ratio = 0.30
payload = f'1.00,1.0,1.0,{count_ratio:.2f}'
is_fraud = float(predictor.predict(payload).decode('utf-8'))
print(f'With transaction count ratio of: {count_ratio:.2f}, fraud score: {is_fraud:.3f}')
count_ratio = 0.06
payload = f'1.00,1.0,1.0,{count_ratio:.2f}'
is_fraud = float(predictor.predict(payload).decode('utf-8'))
print(f'With transaction count ratio of: {count_ratio:.2f}, fraud score: {is_fraud:.3f}')
| 11_stream/archive/99_streaming_feature_store/notebooks/3_train_and_deploy_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Francisco-Dan/daa_2021_/blob/master/7_Diciembre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Wrt6ySh1uQQb" outputId="308670fb-8c42-4559-bebc-66f93ffe0320"
def fibonacci(n):
print("Llamada", n)
if n == 1 or n == 0:
return n
else:
return (fibonacci(n-1)+fibonacci(n-2))
print(fibonacci(6))
# + id="qzWbd1aKzYhg"
"""Merge Sort"""
| 7_Diciembre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://alacip.org/wp-content/uploads/2014/03/logoEscalacip1.png" width="500"></center>
#
#
# <center> <h1>Curso: Introducción al Python</h1> </center>
#
# <br></br>
#
# * Profesor: <a href="http://www.pucp.edu.pe/profesor/jose-manuel-magallanes/" target="_blank">Dr. <NAME>, PhD</a> ([<EMAIL>](mailto:<EMAIL>))<br>
# - Profesor del **Departamento de Ciencias Sociales, Pontificia Universidad Católica del Peru**.<br>
# - Senior Data Scientist del **eScience Institute** and Visiting Professor at **Evans School of Public Policy and Governance, University of Washington**.<br>
# - Fellow Catalyst, **Berkeley Initiative for Transparency in Social Sciences, UC Berkeley**.
#
#
# ## Parte 4: Data Cleaning en Python
# El pre procesamiento de datos es la parte más tediosa del proceso de investigación.
#
# Esta primera parte delata diversos problemas que se tienen con los datos reales que están en la web, como la que vemos a continuación:
import IPython
wikiLink="https://en.wikipedia.org/wiki/List_of_freedom_indices"
iframe = '<iframe src=' + wikiLink + ' width=700 height=350></iframe>'
IPython.display.HTML(iframe)
# Recuerda inspeccionar la tabla para encontrar algun atributo que sirva para su descarga. De ahí, continua.
# +
# antes instala'beautifulsoup4' y 'html5lib'
# es posible que necesites salir y volver a cargar notebook
import pandas as pd
wikiTables=pd.read_html(wikiLink,header=0,flavor='bs4',attrs={'class': 'wikitable sortable'})
# -
# cuantas tenemos?
len(wikiTables)
# Hasta aquí todo parece bien. Como solo hay uno, lo traigo y comienzo a verificar 'suciedades'.
# +
DF=wikiTables[0]
#primera mirada
DF.head()
# -
# La limpieza requiere estrategia. Lo primero que salta a la vista, son los _footnotes_ que están en los títulos:
DF.columns
# aqui ves que pasa cuando divido cada celda usando el caracter '['
[element.split('[') for element in DF.columns]
# Te das cuenta que te puedes quedar con el primer elemento cada vez que partes:
[element.split('[')[0] for element in DF.columns]
# También hay que evitar espacios en blanco:
outSymbol=' '
inSymbol=''
[element.split('[')[0].replace(outSymbol,inSymbol) for element in DF.columns]
# Los números también molestan, pero están en diferentes sitios. Mejor intentemos expresiones regulares:
# +
import re # debe estar instalado.
# espacios: \\s+
# uno o mas numeros \\d+
# bracket que abre \\[
# bracket que cierra \\]
pattern='\\s+|\\d+|\\[|\\]'
nothing=''
#substituyendo 'pattern' por 'nothing':
[re.sub(pattern,nothing,element) for element in DF.columns]
# -
# Ya tengo nuevos titulos de columna (headers)!!
newHeaders=[re.sub(pattern,nothing,element) for element in DF.columns]
# Preparemos los cambios:
list(zip(DF.columns,newHeaders))
# veamos los cambios:
{old:new for old,new in zip(DF.columns,newHeaders)}
# Uso un dict por si hubieses querido cambiar solo algunas columnas:
changes={old:new for old,new in zip(DF.columns,newHeaders)}
DF.rename(columns=changes,inplace=True)
# ahora tenemos:
DF.head()
# Las columnas son categorías, veamos si todas se han escrito de la manera correcta:
DF.FreedomintheWorld.value_counts()
DF.IndexofEconomicFreedom.value_counts()
DF.PressFreedomIndex.value_counts()
DF.DemocracyIndex.value_counts()
# ### Ejercicio
#
# Traer y limpiar la tabla que se ubica es este [link](https://www.cia.gov/library/publications/resources/the-world-factbook/fields/349.html)
import IPython
ciaLink="https://www.cia.gov/library/publications/resources/the-world-factbook/fields/349.html"
ciaTables=pd.read_html(ciaLink,header=0,flavor='bs4',attrs={'id': 'fieldListing'})
ciaTable=ciaTables[0]
ciaTable.head(3)
# +
ciaTable['urbPop']=[v.split('rate of ')[0].split('%')[0].split(': ')[1] for v in ciaTable.Urbanization]
# -
ciaTable['rateUrb']=[v.split('rate of ')[1].split('%')[0].split(': ')[1] for v in ciaTable.Urbanization]
ciaTable
# ____
#
# [Ir a inicio](#beginning)
# _____
#
# **AUSPICIO**:
#
# * El desarrollo de estos contenidos ha sido posible gracias al grant del Berkeley Initiative for Transparency in the Social Sciences (BITSS) at the Center for Effective Global Action (CEGA) at the University of California, Berkeley
#
#
# <center>
# <img src="https://www.bitss.org/wp-content/uploads/2015/07/bitss-55a55026v1_site_icon.png" style="width: 200px;"/>
# </center>
#
# * Este curso cuenta con el auspicio de:
#
#
# <center>
# <img src="https://www.python.org/static/img/psf-logo@2x.png" style="width: 500px;"/>
# </center>
#
#
#
# **RECONOCIMIENTO**
#
#
# EL Dr. Magallanes agradece a la Pontificia Universidad Católica del Perú, por su apoyo en la participación en la Escuela ALACIP.
#
# <center>
# <img src="https://dci.pucp.edu.pe/wp-content/uploads/2014/02/Logotipo_colores-290x145.jpg" style="width: 400px;"/>
# </center>
#
#
# El autor reconoce el apoyo que el eScience Institute de la Universidad de Washington le ha brindado desde el 2015 para desarrollar su investigación en Ciencia de Datos.
#
# <center>
# <img src="https://escience.washington.edu/wp-content/uploads/2015/10/eScience_Logo_HR.png" style="width: 500px;"/>
# </center>
#
# <br>
# <br>
| Parte4_P_DataCleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Naive Bayes
#
# Basic Naive Bayes assumes categorical data. A simple extension for real- valued data is called Gaussian Naive Bayes.
#
# In machine learning we are often interested in selecting the best hypothesis (h) given data (d).
# In a classification problem, our hypothesis (h) may be the class to assign for a new data instance
# (d). One of the easiest ways of selecting the most probable hypothesis given the data that we
# have that we can use as our prior knowledge about the problem. Bayes Theorem provides a
# way that we can calculate the probability of a hypothesis given our prior knowledge. Bayes
# Theorem is stated as:
#
#
# P(h|d) = P(d|h) x P(h) / P(d)
#
#
# Where:
#
# - P(h|d) is the probability of hypothesis h
# given the data d. This is called the posterior
# probability.
#
# - P(d|h) is the probability of data d given that the hypothesis h
# was true
#
# - P(h) is the probability of hypothesis h being true (regardless of the data). This is called
# the prior probability of h.
#
# - P(d) is the probability of the data regardless of the hypothesis
# 20.1 Tutorial Dataset
#
# The dataset describes two categorical input variables and a class variable that has two outputs.
#
from io import StringIO
import pandas as pd
import numpy as np
dataset = StringIO("""Weather Car Class
sunny working go-out
rainy broken go-out
sunny working go-out
sunny working go-out
sunny working go-out
rainy broken stay-home
rainy broken stay-home
sunny working stay-home
sunny broken stay-home
rainy broken stay-home
""")
def clean_cols(cols): return cols.lower().strip()
nb = pd.read_csv(dataset, sep=" ").rename(columns = clean_cols)
nb.sample(3)
# encode categorical variable
nb['class'] = nb['class'].eq('go-out').astype(int)
nb['weather'] = nb['weather'].eq('sunny').astype(int)
nb['car'] = nb['car'].eq('working').astype(int)
nb
# There are two types of quantities that need to be calculated from the dataset for the naive
# Bayes model:
#
# - Class Probabilities.
#
# - Conditional Probabilities.
#
# Let’s start with the class probabilities.
# +
# class probabilities count of each class / total examples
p_go = nb['class'].eq(1).sum() / nb.shape[0]
p_sh = nb['class'].eq(0).sum() / nb.shape[0]
p_go, p_sh
# -
# p(d|h) The conditional probabilities are the probability of each input value given each class value. The
# conditional probabilities for the dataset can be calculated as follows:
#
# +
# example p(weather=sunny| class=go-out) = count(weather=sunny & class=go-out) / count(class=go-out)
p_s_go = np.sum(nb['class'].eq(1) & nb['weather'].eq(1)) / nb['class'].eq(1).sum()
p_r_go = np.sum(nb['class'].eq(1) & nb['weather'].eq(0)) / nb['class'].eq(1).sum()
p_s_sh = np.sum(nb['class'].eq(0) & nb['weather'].eq(1)) / nb['class'].eq(0).sum()
p_r_sh = np.sum(nb['class'].eq(0) & nb['weather'].eq(0)) / nb['class'].eq(0).sum()
# and for car variable
p_wk_go = np.sum(nb['class'].eq(1) & nb['car'].eq(1)) / nb['class'].eq(1).sum()
p_bk_go = np.sum(nb['class'].eq(1) & nb['car'].eq(0)) / nb['class'].eq(1).sum()
p_wk_sh = np.sum(nb['class'].eq(0) & nb['car'].eq(1)) / nb['class'].eq(0).sum()
p_bk_sh = np.sum(nb['class'].eq(0) & nb['car'].eq(0)) / nb['class'].eq(0).sum()
# -
p_s_go
# ### Making predictions
#
# We don't need a probability to predict the most likely class for a new data instance.
# We only need the numerator and the class that gives the largest response, which will be the
# predicted output
#
# MAP
# (
# h
# ) =
# max
# (
# P
# (
# d
# |
# h
# )
# ×
# P
# (
# h
# ))
#
# lets make a prediction for the first row of data
nb.loc[0]
# p(d|h) we have the data but we need to access which hypothesis yeilds a bigger probability
go_out = p_s_go * p_wk_go * p_go
stay_home = p_s_sh * p_wk_sh * p_sh
# we correctly predicted go-out
go_out, stay_home
nb.loc[1]
# incorrect predictions for row 2
go_out = p_bk_go * p_r_go * p_go
stay_home = p_bk_sh * p_r_sh * p_sh
go_out, stay_home
nb.loc[2]
# correctly predicted go out
go_out = p_s_go * p_wk_go * p_go
stay_home = p_s_sh * p_wk_sh * p_sh
go_out, stay_home
# ### Gaussian Naive Bayes
# A simple dataset was contrived for our purposes. It is comprised of two input variables X1 and
# X2 and one output variable Y . The input variables are drawn from a Gaussian distribution,
# which is one assumption made by Gaussian Naive Bayes. The class variable has two values, 0
# and 1, therefore the problem is a binary classi
# cation problem.
# +
dataset = StringIO("""X1 X2 Y
3.393533211 2.331273381 0
3.110073483 1.781539638 0
1.343808831 3.368360954 0
3.582294042 4.67917911 0
2.280362439 2.866990263 0
7.423436942 4.696522875 1
5.745051997 3.533989803 1
9.172168622 2.511101045 1
7.792783481 3.424088941 1
7.939820817 0.791637231 1
""")
# -
gnb = pd.read_csv(dataset, sep=' ').rename(columns = clean_cols)
gnb
from probability import normal_pdf
xs = [x/ 10 for x in range(-50,51)]
pdf_example = pd.Series(index=xs, data = [normal_pdf(x) for x in xs])
pdf_example
pdf_example.plot();
# There are two types of probabilities that we need to summarize from our training data for the
# naive Bayes model:
#
# - Class Probabilities.
# - Conditional Probabilities.
# class probs
p_y_1 = gnb['y'].eq(1).sum()/ gnb['y'].count()
p_y_0 = gnb['y'].eq(0).sum()/ gnb['y'].count()
# The X1 and X2 input variables are real values. As such we will model them as having being
# drawn from a Gaussian distribution. This will allow us to estimate the probability of a given
# value using the Gaussian PDF described above.
#
# The Gaussian PDF requires two parameters in
# addition to the value for which the probability is being estimated: the mean and the standard
# deviation. Therefore we must estimate the mean and the standard deviation for each group of
# conditional probabilities that we require.
# conditional probabilities
# P(X1|Y=0); P(X1|Y=1)...
p_x1_y0_mean = gnb.loc[gnb['y'].eq(0), 'x1'].mean()
p_x1_y1_mean = gnb.loc[gnb['y'].eq(1), 'x1'].mean()
p_x2_y0_mean = gnb.loc[gnb['y'].eq(0), 'x2'].mean()
p_x2_y1_mean = gnb.loc[gnb['y'].eq(1), 'x2'].mean()
p_x1_y0_std = gnb.loc[gnb['y'].eq(0), 'x1'].std()
p_x1_y1_std = gnb.loc[gnb['y'].eq(1), 'x1'].std()
p_x2_y0_std = gnb.loc[gnb['y'].eq(0), 'x2'].std()
p_x2_y1_std = gnb.loc[gnb['y'].eq(1), 'x2'].std()
# #### Make Prediction with Gaussian Naive Bayes
# We can make predictions using Bayes Theorem, introduced and explained in a previous chapter.
# We don't need a probability to predict the most likely class for a new data instance. We only
# need the numerator and the class that gives the largest response is the predicted response.
#
# MAP(h) = max(P(d|h) x P(h))
#
# Let's take the first record from our dataset and use our learned model to predict which class
# we think it belongs. Instance: X1 = 3:393533211, X2 = 2:331273381, Y = 0. We can plug the
# probabilities for our model in for both classes and calculate the response. Starting with the
# response for the output class 0. We multiply the conditional probabilities together and multiply
# it by the probability of any instance belonging to the class.
# looking at first data example
gnb.loc[0]
# +
# class 0 = P(pdf(X1)|class = 0) x P(pdf(X2)|class = 0) x P(class = 0)
# for response y equal to 0
p1 = normal_pdf(gnb.loc[0, 'x1'], mu=p_x1_y0_mean, sigma=p_x1_y0_std)
p2 = normal_pdf(gnb.loc[0, 'x2'], mu=p_x2_y0_mean, sigma=p_x2_y0_std)
y_0 = p1 * p2 * p_y_0
# for response y equal to 1
p1 = normal_pdf(gnb.loc[0, 'x1'], mu=p_x1_y1_mean, sigma=p_x1_y1_std)
p2 = normal_pdf(gnb.loc[0, 'x2'], mu=p_x2_y1_mean, sigma=p_x2_y1_std)
y_1 = p1 * p2 * p_y_1
# prediction is 0
y_0, y_1
# +
from typing import Tuple
def gaussian_nb_prediction(index: int)-> Tuple[float, float]:
"""
Based on x1 and x2 index returns output for
output class y = 0 and y= 1
"""
# hardcoded probabilities for naive bayes
p_x1_y0_mean = gnb.loc[gnb['y'].eq(0), 'x1'].mean()
p_x1_y1_mean = gnb.loc[gnb['y'].eq(1), 'x1'].mean()
p_x2_y0_mean = gnb.loc[gnb['y'].eq(0), 'x2'].mean()
p_x2_y1_mean = gnb.loc[gnb['y'].eq(1), 'x2'].mean()
p_x1_y0_std = gnb.loc[gnb['y'].eq(0), 'x1'].std()
p_x1_y1_std = gnb.loc[gnb['y'].eq(1), 'x1'].std()
p_x2_y0_std = gnb.loc[gnb['y'].eq(0), 'x2'].std()
p_x2_y1_std = gnb.loc[gnb['y'].eq(1), 'x2'].std()
# example class 0 = P(pdf(X1)|class = 0) x P(pdf(X2)|class = 0) x P(class = 0)
# for response y equal to 0
p1 = normal_pdf(gnb.loc[index, 'x1'], mu=p_x1_y0_mean, sigma=p_x1_y0_std)
p2 = normal_pdf(gnb.loc[index, 'x2'], mu=p_x2_y0_mean, sigma=p_x2_y0_std)
y_0 = p1 * p2 * p_y_0
# for response y equal to 1
p1 = normal_pdf(gnb.loc[index, 'x1'], mu=p_x1_y1_mean, sigma=p_x1_y1_std)
p2 = normal_pdf(gnb.loc[index, 'x2'], mu=p_x2_y1_mean, sigma=p_x2_y1_std)
y_1 = p1 * p2 * p_y_1
return y_0, y_1
# -
outputs = pd.DataFrame(data = [gaussian_nb_prediction(idx) for idx in gnb.index.values],
columns=['output_y0','output_y1'])
gnb_predictions = pd.concat([gnb, outputs], axis=1)
gnb_predictions
# based on max value by row calculate prediction value
gnb_predictions['prediction'] = outputs.idxmax(axis='columns').eq('output_y1').astype(int)
# 100% accuracy
gnb_predictions
# This section provides some tips for preparing your data for Naive Bayes.
#
# - Categorical Inputs: Naive Bayes assumes label attributes such as binary, categorical or
# nominal.
# - Gaussian Inputs: If the input variables are real-valued, a Gaussian distribution is
# assumed. In which case the algorithm will perform better if the univariate distributions of
# your data are Gaussian or near-Gaussian. This may require removing outliers (e.g. values
# that are more than 3 or 4 standard deviations from the mean).
# - Classification Problems: Naive Bayes is a classification algorithm suitable for binary
# and multi-class classification.
# - Log Probabilities: The calculation of the likelihood of different class values involves
# multiplying a lot of small numbers together. This can lead to an under
# ow of numerical precision. As such it is good practice to use a log transform of the probabilities to avoid
# this underflow.
# - Kernel Functions: Rather than assuming a Gaussian distribution for numerical input
# values, more complex distributions can be used such as a variety of kernel density functions.
# - Update Probabilities: When new data becomes available, you can simply update the
# probabilities of your model. This can be helpful if the data changes frequently
# In this chapter you discovered how to implement the Gaussian Naive Bayes classifier from
# scratch. You learned about:
#
# - The Gaussian Probability Density Function for estimating the probability of any given
# real value.
# - How to estimate the probabilities required by the Naive Bayes model from a training
# dataset.
# - How to use the learned Naive Bayes model to make predictions.
| old_dsfs/naive_bayes_mlm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="OIom1X-v0TGc"
# # Assignment - Basic Pandas
# <sup>Created by <NAME>, Department of Computer Engineering, Chulalongkorn University</sup>
#
# Using pandas to explore youtube trending data from GB (GBvideos.csv and GB_category_id.json) and answer the questions.
# + id="_ooeQeBn0TGf"
import pandas as pd
import numpy as np
# + [markdown] id="RNyAGpWT0TGh"
# To simplify data retrieval process on Colab, we heck if we are in the Colab environment and download data files from a shared drive and save them in folder "data".
#
# For those using jupyter notebook on the local computer, you can read data directly assuming you save data in the folder "data".
# + id="qro_9JWV0TGi" colab={"base_uri": "https://localhost:8080/"} outputId="3e7f5c45-98ec-4160-bcc1-1697a048d29b"
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !wget https://github.com/kaopanboonyuen/2110446_DataScience_2021s2/raw/main/datasets/data.tgz -O data.tgz
# !tar -xzvf data.tgz
# + [markdown] id="rkNi2LHT0TGj"
# ## How many rows are there in the GBvideos.csv after removing duplications?
# + id="PFzYyW7V0TGj"
# + [markdown] id="mSc2U7HJ0TGk"
# ## How many VDO that have "dislikes" more than "likes"? Make sure that you count only unique title!
# + id="Q0NF1dI40TGk"
# + [markdown] id="oNxh-5CL0TGk"
# ## How many VDO that are trending on 22 Jan 2018 with comments more than 10,000 comments?
# + id="07-y5sNw0TGl"
# + [markdown] id="jnBQwfD70TGl"
# ## Which date that has the minimum average number of comments per VDO?
# + id="KIDZyavc0TGl"
# + [markdown] id="wL7iTiic0TGl"
# ## Compare "Sports" and "Comady", how many days that there are more total daily views of VDO in "Sports" category than in "Comady" category?
# + id="UIzvYqdA0TGm"
| code/week1_numpy_pandas/PandasAssignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="width:1000 px">
#
# <div style="float:right; width:98 px; height:98px;">
# <img src="https://raw.githubusercontent.com/Unidata/MetPy/master/metpy/plots/_static/unidata_150x150.png" alt="Unidata Logo" style="height: 98px;">
# </div>
#
# <h1>Using numpy and KD-trees with netCDF data</h1>
# <h3>Unidata Python Workshop</h3>
#
# <div style="clear:both"></div>
# </div>
#
# <hr style="height:2px;">
#
# <div style="float:right; width:250 px"><img src="https://upload.wikimedia.org/wikipedia/commons/b/b6/3dtree.png" alt="Example Image" style="height: 300px;"></div>
#
# There is now a Unidata Developer's [Blog entry](http://www.unidata.ucar.edu/blogs/developer/en/entry/accessing_netcdf_data_by_coordinates) accompanying this iPython notebook.
#
# The goal is to demonstrate how to quickly access netCDF data based on geospatial coordinates instead of array indices.
#
# - First we show a naive and slow way to do this, in which we also have to worry about longitude anomalies
# - Then we speed up access with numpy arrays
# - Next, we demonstrate how to eliminate longitude anomalies
# - Finally, we use a kd-tree data structure to significantly speed up access by coordinates for large problems
# ## Getting data by coordinates from a netCDF File
#
# Let's look at a netCDF file from the *Atlantic Real-Time Ocean Forecast System*. If you have cloned the [Unidata 2015 Python Workshop](https://github.com/Unidata/unidata-python-workshop), this data is already available in '../data/rtofs_glo_3dz_f006_6hrly_reg3.nc'. Otherwise you can get it from [rtofs_glo_3dz_f006_6hrly_reg3.nc](https://github.com/Unidata/tds-python-workshop/blob/master/data/rtofs_glo_3dz_f006_6hrly_reg3.nc).
# ### Looking at netCDF metadata from Python
#
# In iPython, we could invoke the **ncdump** utility like this:
#
# filename = '../data/rtofs_glo_3dz_f006_6hrly_reg3.nc'
# # !ncdump -h $filename
#
# *if* we know that a recent version of **ncdump** is installed that
# can read compressed data from netCDF-4 classic model files.
#
# Alternatively, we'll use the netCDF4python package to show information about
# the file in a form that's somewhat less familiar, but contains the information
# we need for the subsequent examples. This works for any netCDF file format:
import netCDF4
filename = '../../data/rtofs_glo_3dz_f006_6hrly_reg3.nc'
ncfile = netCDF4.Dataset(filename, 'r')
print(ncfile) # shows global attributes, dimensions, and variables
ncvars = ncfile.variables # a dictionary of variables
# print information about specific variables, including type, shape, and attributes
for varname in ['temperature', 'salinity', 'Latitude', 'Longitude']:
print(ncvars[varname])
# Here's a sparse picture (every 25th point on each axis) of what the grid looks like on which Latitude, Longitude, Temperature, Salinity, and other variables are defined:
#
# 
# ## Example query: sea surface temperature and salinity at 50N, 140W?
#
# - So **Longitude** and **Latitude** are 2D netCDF variables of shape 850 x 712, indexed by **Y** and **X** dimensions
# - That's 605200 values for each
# - There's no _direct_ way in this file (and many netCDF files) to compute grid indexes from coordinates via a coordinate system and projection parameters. Instead, we have to rely on the latitude and longitude auxiliary coordinate variables, as required by the CF conventions for data not on a simple lat,lon grid.
# - To get the temperature at 50N, 140W, we need to find **Y** and **X** indexes **iy** and **ix** such that (**Longitude[iy, ix]**, **Latitude[iy, ix]**) is "close" to (50.0, -140.0).
# ### Naive, slow way using nested loops
#
# - Initially, for simplicity, we just use Euclidean distance squared, as if the Earth is flat, latitude and longitude are $x$- and $y$-coordinates, and the distance squared between points $(lat_1,lon_1)$ and $(lat_0,lon_0)$ is $( lat_1 - lat_0 )^2 + ( lon_1 - lon_0 )^2$.
# - Note: these assumptions are wrong near the poles and on opposite sides of longitude boundary discontinuity.
# - So, keeping things simple, we want to find **iy** and **ix** to minimize
#
# ``(Latitude[iy, ix] - lat0)**2 + (Longitude[iy, ix] - lon0)**2``
#
# 
# ## Reading netCDF data into numpy arrays
#
# To access netCDF data, rather than just metadata, we will also need NumPy:
#
# - A Python library for scientific programming.
# - Supports n-dimensional array-based calculations similar to Fortran and IDL.
# - Includes fast mathematical functions to act on scalars and arrays.
#
# With the Python netCDF4 package, using "[ ... ]" to index a netCDF variable object reads or writes a numpy array from the associated netCDF file.
#
# The code below reads latitude and longitude values into 2D numpy arrays named **latvals** and **lonvals**:
# ### First version: slow and spatially challenged
# Here's a function that uses simple nested loops to find indices that minimize the distance to the desired coordinates, written as if using Fortran or C rather than Python. We'll call this function in the cell following this definition ...
# +
import numpy as np
import netCDF4
def naive_slow(latvar,lonvar,lat0,lon0):
'''
Find "closest" point in a set of (lat,lon) points to specified point
latvar - 2D latitude variable from an open netCDF dataset
lonvar - 2D longitude variable from an open netCDF dataset
lat0,lon0 - query point
Returns iy,ix such that
(lonval[iy,ix] - lon0)**2 + (latval[iy,ix] - lat0)**2
is minimum. This "closeness" measure works badly near poles and
longitude boundaries.
'''
# Read from file into numpy arrays
latvals = latvar[:]
lonvals = lonvar[:]
ny,nx = latvals.shape
dist_sq_min = 1.0e30
for iy in range(ny):
for ix in range(nx):
latval = latvals[iy, ix]
lonval = lonvals[iy, ix]
dist_sq = (latval - lat0)**2 + (lonval - lon0)**2
if dist_sq < dist_sq_min:
iy_min, ix_min, dist_sq_min = iy, ix, dist_sq
return iy_min,ix_min
# -
# When we call the function above it takes several seconds to run, because it calculates distances one point at a time, for each of the 605200 $(lat, lon)$ points. Note that once indices for the point nearest to (50, -140) are found, they can be used to access temperature, salinity, and other netCDF variables that use the same dimensions.
ncfile = netCDF4.Dataset(filename, 'r')
latvar = ncfile.variables['Latitude']
lonvar = ncfile.variables['Longitude']
iy,ix = naive_slow(latvar, lonvar, 50.0, -140.0)
print('Closest lat lon:', latvar[iy,ix], lonvar[iy,ix])
tempvar = ncfile.variables['temperature']
salvar = ncfile.variables['salinity']
print('temperature:', tempvar[0, 0, iy, ix], tempvar.units)
print('salinity:', salvar[0, 0, iy, ix], salvar.units)
ncfile.close()
# ### NumPy arrays instead of loops: fast, but still assumes flat earth
#
# The above function is slow, because it doesn't make good use of NumPy arrays. It's much faster to use whole array operations to eliminate loops and element-at-a-time computation. NumPy functions that help eliminate loops include:
#
# - The `argmin()` method that returns a 1D index of the minimum value of a NumPy array
# - The `unravel_index()` function that converts a 1D index back into a multidimensional index
# +
import numpy as np
import netCDF4
def naive_fast(latvar,lonvar,lat0,lon0):
# Read latitude and longitude from file into numpy arrays
latvals = latvar[:]
lonvals = lonvar[:]
ny,nx = latvals.shape
dist_sq = (latvals-lat0)**2 + (lonvals-lon0)**2
minindex_flattened = dist_sq.argmin() # 1D index of min element
iy_min,ix_min = np.unravel_index(minindex_flattened, latvals.shape)
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
latvar = ncfile.variables['Latitude']
lonvar = ncfile.variables['Longitude']
iy,ix = naive_fast(latvar, lonvar, 50.0, -140.0)
print('Closest lat lon:', latvar[iy,ix], lonvar[iy,ix])
ncfile.close()
# -
# ### Spherical Earth with tunnel distance: fast _and_ correct
#
# Though assuming a flat Earth may work OK for this example, we'd like to not worry about whether longitudes are from 0 to 360 or -180 to 180, or whether points are close to the poles.
# The code below fixes this by using the square of "tunnel distance" between (lat,lon) points. This version is both fast and correct (for a _spherical_ Earth).
# +
import numpy as np
import netCDF4
from math import pi
from numpy import cos, sin
def tunnel_fast(latvar,lonvar,lat0,lon0):
'''
Find closest point in a set of (lat,lon) points to specified point
latvar - 2D latitude variable from an open netCDF dataset
lonvar - 2D longitude variable from an open netCDF dataset
lat0,lon0 - query point
Returns iy,ix such that the square of the tunnel distance
between (latval[it,ix],lonval[iy,ix]) and (lat0,lon0)
is minimum.
'''
rad_factor = pi/180.0 # for trignometry, need angles in radians
# Read latitude and longitude from file into numpy arrays
latvals = latvar[:] * rad_factor
lonvals = lonvar[:] * rad_factor
ny,nx = latvals.shape
lat0_rad = lat0 * rad_factor
lon0_rad = lon0 * rad_factor
# Compute numpy arrays for all values, no loops
clat,clon = cos(latvals),cos(lonvals)
slat,slon = sin(latvals),sin(lonvals)
delX = cos(lat0_rad)*cos(lon0_rad) - clat*clon
delY = cos(lat0_rad)*sin(lon0_rad) - clat*slon
delZ = sin(lat0_rad) - slat;
dist_sq = delX**2 + delY**2 + delZ**2
minindex_1d = dist_sq.argmin() # 1D index of minimum element
iy_min,ix_min = np.unravel_index(minindex_1d, latvals.shape)
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
latvar = ncfile.variables['Latitude']
lonvar = ncfile.variables['Longitude']
iy,ix = tunnel_fast(latvar, lonvar, 50.0, -140.0)
print('Closest lat lon:', latvar[iy,ix], lonvar[iy,ix])
ncfile.close()
# -
# ### KD-Trees: faster data structure for lots of queries
#
# We can still do better, by using a data structure designed to support efficient nearest-neighbor queries: the [KD-tree](http://en.wikipedia.org/wiki/K-d_tree). It works like a multidimensional binary tree, so finding the point nearest to a query point is _much_ faster than computing all the distances to find the minimum. It takes some setup time to load all the points into the data structure, but that only has to be done once for a given set of points.
#
# For a single point query, it's still more than twice as fast as the naive slow version above, but building the KD-tree for 605,200 points takes more time than the fast numpy search through all the points, so in this case using the KD-tree for a _single_ point query is sort of pointless ...
# +
import numpy as np
import netCDF4
from math import pi
from numpy import cos, sin
from scipy.spatial import cKDTree
def kdtree_fast(latvar,lonvar,lat0,lon0):
rad_factor = pi/180.0 # for trignometry, need angles in radians
# Read latitude and longitude from file into numpy arrays
latvals = latvar[:] * rad_factor
lonvals = lonvar[:] * rad_factor
ny,nx = latvals.shape
clat,clon = cos(latvals),cos(lonvals)
slat,slon = sin(latvals),sin(lonvals)
# Build kd-tree from big arrays of 3D coordinates
triples = list(zip(np.ravel(clat*clon), np.ravel(clat*slon), np.ravel(slat)))
kdt = cKDTree(triples)
lat0_rad = lat0 * rad_factor
lon0_rad = lon0 * rad_factor
clat0,clon0 = cos(lat0_rad),cos(lon0_rad)
slat0,slon0 = sin(lat0_rad),sin(lon0_rad)
dist_sq_min, minindex_1d = kdt.query([clat0*clon0, clat0*slon0, slat0])
iy_min, ix_min = np.unravel_index(minindex_1d, latvals.shape)
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
latvar = ncfile.variables['Latitude']
lonvar = ncfile.variables['Longitude']
iy,ix = kdtree_fast(latvar, lonvar, 50.0, -140.0)
print('Closest lat lon:', latvar[iy,ix], lonvar[iy,ix])
ncfile.close()
# -
# ### Timing the functions
#
# If you're curious about actual times for the versions above, the iPython notebook "%%timeit" statement gets accurate timings of all of them. Below, we time just a single query point, in this case (50.0, -140.0). To get accurate timings, the "%%timeit" statement lets us do untimed setup first on the same line, before running the function call in a loop.
ncfile = netCDF4.Dataset(filename,'r')
latvar = ncfile.variables['Latitude']
lonvar = ncfile.variables['Longitude']
# %%timeit
naive_slow(latvar, lonvar, 50.0, -140.0)
# %%timeit
naive_fast(latvar, lonvar, 50.0, -140.0)
# %%timeit
tunnel_fast(latvar, lonvar, 50.0, -140.0)
# %%timeit
kdtree_fast(latvar, lonvar, 50.0, -140.0)
ncfile.close()
# ## Separating setup from query
#
# The above use of the KD-tree data structure is not the way it's meant to be used. Instead, it should be initialized _once_ with all the k-dimensional data for which nearest-neighbors are desired, then used repeatedly on each query, amortizing the work done to build the data structure over all the following queries. By separately timing the setup and the time required per query, the threshold for number of queries beyond which the KD-tree is faster can be determined.
#
# That's exactly what we'll do now. We split each algorithm into two functions, a setup function and a query function. The times per query go from seconds (the naive version) to milliseconds (the array-oriented numpy version) to microseconds (the turbo-charged KD-tree, once it's built).
#
# Rather than just using functions, we define a Class for each algorithm, do the setup in the class constructor, and provide a query method.
# +
# Split naive_slow into initialization and query, so we can time them separately
import numpy as np
import netCDF4
class Naive_slow(object):
def __init__(self, ncfile, latvarname, lonvarname):
self.ncfile = ncfile
self.latvar = self.ncfile.variables[latvarname]
self.lonvar = self.ncfile.variables[lonvarname]
# Read latitude and longitude from file into numpy arrays
self.latvals = self.latvar[:]
self.lonvals = self.lonvar[:]
self.shape = self.latvals.shape
def query(self,lat0,lon0):
ny,nx = self.shape
dist_sq_min = 1.0e30
for iy in range(ny):
for ix in range(nx):
latval = self.latvals[iy, ix]
lonval = self.lonvals[iy, ix]
dist_sq = (latval - lat0)**2 + (lonval - lon0)**2
if dist_sq < dist_sq_min:
iy_min, ix_min, dist_sq_min = iy, ix, dist_sq
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
ns = Naive_slow(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
print('Closest lat lon:', ns.latvar[iy,ix], ns.lonvar[iy,ix])
ncfile.close()
# +
# Split naive_fast into initialization and query, so we can time them separately
import numpy as np
import netCDF4
class Naive_fast(object):
def __init__(self, ncfile, latvarname, lonvarname):
self.ncfile = ncfile
self.latvar = self.ncfile.variables[latvarname]
self.lonvar = self.ncfile.variables[lonvarname]
# Read latitude and longitude from file into numpy arrays
self.latvals = self.latvar[:]
self.lonvals = self.lonvar[:]
self.shape = self.latvals.shape
def query(self,lat0,lon0):
dist_sq = (self.latvals-lat0)**2 + (self.lonvals-lon0)**2
minindex_flattened = dist_sq.argmin() # 1D index
iy_min, ix_min = np.unravel_index(minindex_flattened, self.shape) # 2D indexes
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
ns = Naive_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
print('Closest lat lon:', ns.latvar[iy,ix], ns.lonvar[iy,ix])
ncfile.close()
# +
# Split tunnel_fast into initialization and query, so we can time them separately
import numpy as np
import netCDF4
from math import pi
from numpy import cos, sin
class Tunnel_fast(object):
def __init__(self, ncfile, latvarname, lonvarname):
self.ncfile = ncfile
self.latvar = self.ncfile.variables[latvarname]
self.lonvar = self.ncfile.variables[lonvarname]
# Read latitude and longitude from file into numpy arrays
rad_factor = pi/180.0 # for trignometry, need angles in radians
self.latvals = self.latvar[:] * rad_factor
self.lonvals = self.lonvar[:] * rad_factor
self.shape = self.latvals.shape
clat,clon,slon = cos(self.latvals),cos(self.lonvals),sin(self.lonvals)
self.clat_clon = clat*clon
self.clat_slon = clat*slon
self.slat = sin(self.latvals)
def query(self,lat0,lon0):
# for trignometry, need angles in radians
rad_factor = pi/180.0
lat0_rad = lat0 * rad_factor
lon0_rad = lon0 * rad_factor
delX = cos(lat0_rad)*cos(lon0_rad) - self.clat_clon
delY = cos(lat0_rad)*sin(lon0_rad) - self.clat_slon
delZ = sin(lat0_rad) - self.slat;
dist_sq = delX**2 + delY**2 + delZ**2
minindex_1d = dist_sq.argmin() # 1D index
iy_min, ix_min = np.unravel_index(minindex_1d, self.shape) # 2D indexes
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
ns = Tunnel_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
print('Closest lat lon:', ns.latvar[iy,ix], ns.lonvar[iy,ix])
ncfile.close()
# +
# Split kdtree_fast into initialization and query, so we can time them separately
import numpy as np
import netCDF4
from math import pi
from numpy import cos, sin
from scipy.spatial import cKDTree
class Kdtree_fast(object):
def __init__(self, ncfile, latvarname, lonvarname):
self.ncfile = ncfile
self.latvar = self.ncfile.variables[latvarname]
self.lonvar = self.ncfile.variables[lonvarname]
# Read latitude and longitude from file into numpy arrays
rad_factor = pi/180.0 # for trignometry, need angles in radians
self.latvals = self.latvar[:] * rad_factor
self.lonvals = self.lonvar[:] * rad_factor
self.shape = self.latvals.shape
clat,clon = cos(self.latvals),cos(self.lonvals)
slat,slon = sin(self.latvals),sin(self.lonvals)
clat_clon = clat*clon
clat_slon = clat*slon
triples = list(zip(np.ravel(clat*clon), np.ravel(clat*slon), np.ravel(slat)))
self.kdt = cKDTree(triples)
def query(self,lat0,lon0):
rad_factor = pi/180.0
lat0_rad = lat0 * rad_factor
lon0_rad = lon0 * rad_factor
clat0,clon0 = cos(lat0_rad),cos(lon0_rad)
slat0,slon0 = sin(lat0_rad),sin(lon0_rad)
dist_sq_min, minindex_1d = self.kdt.query([clat0*clon0,clat0*slon0,slat0])
iy_min, ix_min = np.unravel_index(minindex_1d, self.shape)
return iy_min,ix_min
ncfile = netCDF4.Dataset(filename, 'r')
ns = Kdtree_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
print('Closest lat lon:', ns.latvar[iy,ix], ns.lonvar[iy,ix])
ncfile.close()
# -
# ### Setup times for the four algorithms
ncfile = netCDF4.Dataset(filename, 'r')
# %%timeit
ns = Naive_slow(ncfile,'Latitude','Longitude')
# %%timeit
ns = Naive_fast(ncfile,'Latitude','Longitude')
# %%timeit
ns = Tunnel_fast(ncfile,'Latitude','Longitude')
# %%timeit
ns = Kdtree_fast(ncfile,'Latitude','Longitude')
# ### Query times for the four algorithms
# %%timeit ns = Naive_slow(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
# %%timeit ns = Naive_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
# %%timeit ns = Tunnel_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
# %%timeit ns = Kdtree_fast(ncfile,'Latitude','Longitude')
iy,ix = ns.query(50.0, -140.0)
ncfile.close()
# In the next cell, we copy the results of the %%timeit runs into Python variables. _(Is there a way to capture %%timeit output, so we don't have to do this manually?)_
ns0,nf0,tf0,kd0 = 3.76, 3.8, 27.4, 2520 # setup times in msec
nsq,nfq,tfq,kdq = 7790, 2.46, 5.14, .0738 # query times in msec
# ### Summary of timings
#
# The naive_slow method is always slower than all other methods. The naive_fast method would only be worth considering if non-flatness of the Earth is irrelevant, for example in a relatively small region not close to the poles and not crossing a longitude discontinuity.
#
# Total time for running initialization followed by N queries is:
#
# - naive_slow: $ns0 + nsq * N$
# - naive_fast: $nf0 + nfq * N$
# - tunnel_fast: $nt0 + ntq * N$
# - kdtree_fast: $kd0 + kdq * N$
N = 10000
print(N, "queries using naive_slow:", round((ns0 + nsq*N)/1000,1), "seconds")
print(N, "queries using naive_fast:", round((nf0 + nfq*N)/1000,1), "seconds")
print(N, "queries using tunnel_fast:", round((tf0 + tfq*N)/1000,1), "seconds")
print(N, "queries using kdtree_fast:", round((kd0 + kdq*N)/1000,1), "seconds")
print('')
print("kd_tree_fast outperforms naive_fast above:", int((kd0-nf0)/(nfq-kdq)), "queries")
print("kd_tree_fast outperforms tunnel_fast above:", int((kd0-tf0)/(tfq-kdq)), "queries")
# The advantage of using KD-trees is much greater for more search set points, as KD-tree query complexity is O(log(N)), but the other algorithms are O(N), the same as the difference between using binary search versus linear search.
| notebooks/netCDF/netcdf-by-coordinates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="j9ZMig7m1Cn4" colab_type="text"
# # PySpark Analytics for Crowd Dynamics
# # Harvard University
# # CS 205, Spring 2019
# # Group 1
# + id="VtVHM3oEbixW" colab_type="code" outputId="b31f29fb-1486-4e28-de2e-6d73716e1b73" colab={"base_uri": "https://localhost:8080/", "height": 214}
# !pip install pyspark
# + id="dU6MHYtKbnLX" colab_type="code" outputId="64ce014f-dfbb-4a64-a5cf-b9af15d4f4b9" colab={"base_uri": "https://localhost:8080/", "height": 119}
from google.colab import drive
import pandas as pd
import numpy as np
from functools import reduce
from pyspark import SparkConf, SparkContext
from pyspark.sql import SQLContext, SparkSession
from pyspark.mllib.linalg import Vectors
from pyspark.sql.functions import udf, array, avg, col, size, struct, lag, window, countDistinct, monotonically_increasing_id, collect_list
import pyspark.sql.functions as F
from pyspark.sql.types import DoubleType, IntegerType, StringType, ArrayType, LongType, FloatType
from pyspark.sql.window import Window
import re
drive.mount('/content/gdrive', force_remount=True)
# + id="3iHxYt95bqDW" colab_type="code" colab={}
spark = SparkSession.builder.getOrCreate()
directory = '/content/gdrive/My Drive/CS 205 Project/records/'
# + [markdown] id="fmASLabX8UjX" colab_type="text"
# # Load data
# + id="v8BzcE-P-RSQ" colab_type="code" outputId="087cae63-a979-4493-df78-f082386d4aa1" colab={"base_uri": "https://localhost:8080/", "height": 445}
df = spark.read.json(directory + '1-min')
df.show()
# + [markdown] id="xL46J7P0u1tx" colab_type="text"
# # Add Frame Index (Timestep) and Display Schema
# + id="icllunSMu1Tp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="7bb27104-d03c-4aea-c930-8a2b88bf7ec3"
df = df.withColumn("frame", monotonically_increasing_id().cast("timestamp")).select("frame", "bboxes", "scores")
df.show()
# + id="thMWSpX2xCe2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 132} outputId="e43736af-ef81-41d1-c759-b0de840b3ecb"
df.printSchema()
# + [markdown] id="R9IEO0298Z3C" colab_type="text"
# # Functions used for UDFs (including velocity and group size)
# + id="2exW6frCbtP7" colab_type="code" colab={}
def count(column):
return len(column)
def sum_vals(column):
return float(sum(column))
def avg_vals(columns):
return float(columns[0] / columns[1])
def get_center(values):
res = []
for i in range(int(len(values)/4)):
y1, x1, y2, x2 = values[4*i:4*i+4]
x_mean, y_mean = round(float(x1+x2)/2, 3), round(float(y1+y2)/2, 3)
res.extend([x_mean, y_mean])
return res
def get_x(values):
return [values[i] for i in range(0, len(values), 2)]
def get_y(values):
return [values[i] for i in range(1, len(values), 2)]
def fudf(val):
return reduce(lambda x, y:x+y, val)
# + id="JwRM-h2B6jzE" colab_type="code" colab={}
def compute_velocities(cols, fps=0.7, threshold=0.3):
# Assume frame_1 = [x_i, y_i, x_{i+1}, y_{i+1}, ...]
f_1 = iter(cols[0])
f_2 = iter(cols[1])
frame_1 = list(zip(f_1, f_1))
frame_2 = list(zip(f_2, f_2))
val_1 = {k: v for k, v in enumerate(frame_1)}
val_2 = {k: v for k, v in enumerate(frame_2)}
# Compute pairwise distances
distances = {}
for i in range(len(frame_1)):
for j in range(len(frame_2)):
# Euclidean distance between two people
distances[i, j] = np.sqrt(
(val_1[i][0] - val_2[j][0]) ** 2 + (val_1[i][1] - val_2[j][1]) ** 2)
# Assigned ids from frame 1 (reference frame), {id_i: vel_i}
velocities = dict()
# Assigned ids from frame 2 (with values as match in frame 1)
targets = dict()
num_assigned = 0
num_ids = min(len(frame_1), len(frame_2))
# Sort distances by key: (id in frame 1, id in frame 2)
pairs = sorted(distances.items(), key=lambda v:v[1])
for p, dist in pairs:
# Stop assigning ids when the distance exceeds a user-defined threshold
# i.e. this covers the case when a person leaves one end of the image
# and another person enters at the opposite side. We should not match
# these ids to each other.
if dist > threshold:
break
# Found closest ids between frames
if p[0] not in velocities and p[1] not in targets and num_assigned < num_ids:
num_assigned += 1
# Velocity (distance units per second)
velocities[p[0]] = dist * fps
targets[p[1]] = p[0]
return [float(v) for v in velocities.values()]
def dfs_all(graph):
def dfs(node, graph):
stack = [node]
cc = [node]
while stack:
u = stack.pop()
for v in graph[u]:
if not visited[v]:
visited[v] = True
cc.append(v)
stack.append(v)
return cc
ccs = []
visited = [False for _ in range(len(graph))]
for i in range(len(graph)):
if not visited[i]:
visited[i] = True
cc = dfs(i, graph)
ccs.append(cc)
return list(map(len, ccs))
def compute_groups(positions, threshold=0.1):
p_1 = iter(positions)
positions = list(zip(p_1, p_1))
# Compute pairwise distances
graph = {i: set() for i in range(len(positions))}
for i in range(len(positions)):
for j in range(i, len(positions)):
# Euclidean distance between two people
dist = np.sqrt(
(positions[i][0] - positions[j][0]) ** 2 + (positions[i][1] - positions[j][1]) ** 2)
# Add edge to graph
if dist < threshold:
graph[i].add(j)
graph[j].add(i)
lengths = dfs_all(graph)
return lengths
# + [markdown] id="8YzPdjAQ8YIN" colab_type="text"
# # UDFs
# + id="DS2CsxSUggwh" colab_type="code" colab={}
count_udf = udf(count, IntegerType())
sum_udf = udf(sum_vals, DoubleType())
avg_udf = udf(lambda arr: avg_vals(arr), DoubleType())
center_udf = udf(get_center, ArrayType(FloatType()))
velocity_udf = udf(lambda arr: compute_velocities(arr), ArrayType(DoubleType()))
group_udf = udf(compute_groups, ArrayType(IntegerType()))
x_udf = udf(get_x, ArrayType(DoubleType()))
y_udf = udf(get_y, ArrayType(DoubleType()))
flattenUdf = udf(fudf, ArrayType(DoubleType()))
# + [markdown] id="YRJEGj8dWHac" colab_type="text"
# # Window size for pairwise shifting
# + id="czV9xoexWKka" colab_type="code" colab={}
w_pair = Window().partitionBy().orderBy(col("frame"))
# + [markdown] id="A88Tq50vWEJY" colab_type="text"
# # Create and Modify Columns
# + id="mW5QnXx-clQW" colab_type="code" outputId="2cf11ee5-23ec-490e-e7a5-576ea9f732b2" colab={"base_uri": "https://localhost:8080/", "height": 465}
df = (df.withColumn('num_people', count_udf('scores'))
.withColumn('centers', center_udf('bboxes'))
.withColumn('x_centers', x_udf('centers'))
.withColumn('y_centers', y_udf('centers'))
.withColumn('group_sizes', group_udf('centers'))
.withColumn('num_groups', count_udf('group_sizes'))
.withColumn('next_frame_centers', lag("centers", -1).over(w_pair)).na.drop()
.withColumn('velocities', velocity_udf(struct('centers', 'next_frame_centers')))
.withColumn('num_velocities', count_udf('velocities'))
.withColumn('sum_velocities', sum_udf('velocities')))
df.show()
# + [markdown] id="bKvkxJXdnTml" colab_type="text"
# # Aggregate each 5 minute window to compute:
# - average number of people detected
# - average group size
# - average velocity
# + id="41j9407Kf92N" colab_type="code" colab={}
# 0.7 frames/second => 210 frames for 300 seconds (5 minutes)
# 2.4 frames/second => 720 frames for 300 seconds (5 minutes)
agg_df = (df.groupBy(window("frame", windowDuration="3 seconds", slideDuration="3 seconds"))
.agg(F.sum('num_people'),
F.sum('num_groups'),
F.sum('sum_velocities'),
F.sum('num_velocities'),
avg('num_people'),
collect_list('x_centers'),
collect_list('y_centers'))
.withColumn('avg_group_size', avg_udf(struct('sum(num_people)', 'sum(num_groups)')))
.withColumn('avg_velocity', avg_udf(struct('sum(sum_velocities)', 'sum(num_velocities)')))
.withColumnRenamed('avg(num_people)', 'avg_num_people')
.withColumn('x_centers', flattenUdf('collect_list(x_centers)'))
.withColumn('y_centers', flattenUdf('collect_list(y_centers)'))
.drop('collect_list(x_centers)')
.drop('collect_list(y_centers)')
.drop('sum(num_people)')
.drop('sum(num_groups)')
.drop('sum(sum_velocities)')
.drop('sum(num_velocities)')
.orderBy('window'))
# + id="AgoVmE0Uz8jg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 330} outputId="232ecfce-3587-4fe9-dcc0-084fee00d80e"
agg_df.show()
# + id="Ooj4MwGl4fgT" colab_type="code" colab={}
pdf = agg_df.toPandas()
# + id="fTTlmoT342Q-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="bc1ef2ca-7eeb-4bdd-8b79-05524489d886"
pdf
# + [markdown] id="6luoywH15o0o" colab_type="text"
# # Visualization
# + id="3Q83pCXv4AXJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 597} outputId="ac044d95-1a40-4fae-f55d-b0ee8ff767c7"
from pathlib import Path
from bokeh.plotting import figure, show, output_notebook
from bokeh.models import ColumnDataSource, HoverTool
from bokeh.models.widgets import Tabs, Panel
output_notebook()
pdf.index.name = 'index'
colors = ['red', 'blue', 'green']
colnames = ['avg_velocity', 'avg_group_size', 'avg_num_people']
plot_titles = ['Average Velocities During the Day',
'Average Group Size During the Day',
'Average Number of People During the Day']
tab_titles = ['Velocity', 'Group Size', 'Number of People']
ylabels = ['Average Velocity (m/s)',
'Average Group size (number of people)',
'Average Number of People']
# Create panels for each tab of the visualization
panels = []
for i in range(len(colors)):
hover = HoverTool()
hover.tooltips = [('Timestamp', '@{}'.format(colnames[i]))]
p = figure(title=plot_titles[i],
plot_height=500,
plot_width=500,
tools=[hover, "pan,reset,wheel_zoom"])
p.vbar(x='index',
top=colnames[i],
width=0.9,
color=colors[i],
source=pdf)
p.xaxis.axis_label = "Time of Day (5 minute windows)"
p.yaxis.axis_label = ylabels[i]
panels.append(Panel(child=p, title=tab_titles[i]))
tabs = Tabs(tabs=panels)
show(tabs)
| notebooks/FullAnalytics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
# !{sys.executable} -m pip install --upgrade --no-cache-dir pyfiglet
# +
import pyfiglet
name = input("What is your name? ")
print("Your name is: ", name)
print(pyfiglet.figlet_format('Hello %s' % name, font='standard'))
# -
# !{sys.executable} -m pip install --upgrade --no-cache-dir colorama
# +
from colorama import Fore
from pyfiglet import figlet_format as fformat
font = 'standard'
subfont = 'cybermedium'
red, white, blue, green = Fore.RED, Fore.WHITE, Fore.BLUE, Fore.GREEN
line1 = fformat('Hello %s' % name, font=font)
line2 = fformat('Pick the blue pill', font=subfont, width=200)
line3 = fformat('or the red pill.', font=subfont, width=200)
print('%s%s%s' % (green, line1, white))
print('%s%s%s' % (blue, line2, white))
print('%s%s%s' % (red, line3, white))
# -
# !{sys.executable} -m pip install --upgrade --no-cache-dir logzero
# +
import logzero
from logzero import logger
# Setup rotating logfile with 3 rotations, each with a maximum filesize of 1MB:
logzero.logfile("rotating-logfile.log", maxBytes=1e6, backupCount=3)
logger.info("This log message goes to the console and the logfile")
# These log messages are sent to the console
logger.debug("This is blue from console.")
logger.info("This is green from console.")
logger.warning("This is yellow from console.")
logger.error("This is red from console.")
# This is how you'd log an exception
try:
raise Exception("this is a demo exception")
except Exception as e:
logger.exception(e)
# -
# !{sys.executable} -m pip install --upgrade --no-cache-dir pipulate
try:
gurl
except:
gurl = input("Enter a Google Sheet URL ")
# !{sys.executable} -m pip install --upgrade --no-cache-dir pyppeteer
| examples/LESSON12-Installing_Packages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # HVAC with Amazon SageMaker RL
#
# ---
# ## Introduction
#
#
# HVAC stands for Heating, Ventilation and Air Conditioning and is responsible for keeping us warm and comfortable indoors. HVAC takes up a whopping 50% of the energy in a building and accounts for 40% of energy use in the US [1, 2]. Several control system optimizations have been proposed to reduce energy usage while ensuring thermal comfort.
#
# Modern buildings collect data about the weather, occupancy and equipment use. All of this can be used to optimize HVAC energy usage. Reinforcement Learning (RL) is a good fit because it can learn how to interact with the environment and identify strategies to limit wasted energy. Several recent research efforts have shown that RL can reduce HVAC energy consumption by 15-20% [3, 4].
#
# As training an RL algorithm in a real HVAC system can take time to converge as well as potentially lead to hazardous settings as the agent explores its state space, we turn to a simulator to train the agent. [EnergyPlus](https://energyplus.net/) is an open source, state of the art HVAC simulator from the US Department of Energy. We use a simple example with this simulator to showcase how we can train an RL model easily with Amazon SageMaker RL.
#
# 1. Objective: Control the data center HVAC system to reduce energy consumption while ensuring the room temperature stays within specified limits.
# 2. Environment: We have a small single room datacenter that the HVAC system is cooling to ensure the compute equipment works properly. We will train our RL agent to control this HVAC system for one day subject to weather conditions in San Francisco. The agent takes actions every 5 minutes for a 24 hour period. Hence, the episode is a fixed 120 steps.
# 3. State: The outdoor temperature, outdoor humidity and indoor room temperature.
# 4. Action: The agent can set the heating and cooling setpoints. The cooling setpoint tells the HVAC system that it should start cooling the room if the room temperature goes above this setpoint. Likewise, the HVAC systems starts heating if the room temperature goes below the heating setpoint.
# 5. Reward: The rewards has two components which are added together with coefficients:
# 1. It is proportional to the energy consumed by the HVAC system.
# 2. It gets a large penalty when the room temperature exceeds pre-specified lower or upper limits (as defined in `data_center_env.py`).
#
# References
#
# 1. [sciencedirect.com](https://www.sciencedirect.com/science/article/pii/S0378778807001016)
# 2. [environment.gov.au](https://www.environment.gov.au/system/files/energy/files/hvac-factsheet-energy-breakdown.pdf)
# 3. Wei, Tianshu, <NAME>, and <NAME>. "Deep reinforcement learning for building hvac control." In Proceedings of the 54th Annual Design Automation Conference 2017, p. 22. ACM, 2017.
# 4. Zhang, Zhiang, and <NAME>. "Practical implementation and evaluation of deep reinforcement learning control for a radiant heating system." In Proceedings of the 5th Conference on Systems for Built Environments, pp. 148-157. ACM, 2018.
# ## Pre-requisites
#
# ### Imports
#
# To get started, we'll import the Python libraries we need, set up the environment with a few prerequisites for permissions and configurations.
import sagemaker
import boto3
import sys
import os
import glob
import re
import subprocess
import numpy as np
from IPython.display import HTML
import time
from time import gmtime, strftime
sys.path.append("common")
from misc import get_execution_role, wait_for_s3_object
from docker_utils import build_and_push_docker_image
from sagemaker.rl import RLEstimator, RLToolkit, RLFramework
# ### Setup S3 bucket
#
# Create a reference to the default S3 bucket that will be used for model outputs.
sage_session = sagemaker.session.Session()
s3_bucket = sage_session.default_bucket()
s3_output_path = 's3://{}/'.format(s3_bucket)
print("S3 bucket path: {}".format(s3_output_path))
# ### Define Variables
#
# We define a job below that's used to identify our jobs.
# create unique job name
job_name_prefix = 'rl-hvac'
# ### Configure settings
#
# You can run your RL training jobs locally on the SageMaker notebook instance or on SageMaker training. In both of these scenarios, you can run in either 'local' (where you run the commands) or 'SageMaker' mode (on SageMaker training instances). 'local' mode uses the SageMaker Python SDK to run your code in Docker containers locally. It can speed up iterative testing and debugging while using the same familiar Python SDK interface. Just set `local_mode = True`. And when you're ready move to 'SageMaker' mode to scale things up.
# +
# run local (on this machine)?
# or on sagemaker training instances?
local_mode = False
if local_mode:
instance_type = 'local'
else:
# choose a larger instance to avoid running out of memory
instance_type = "ml.m4.4xlarge"
# -
# ### Create an IAM role
#
# Either get the execution role when running from a SageMaker notebook instance `role = sagemaker.get_execution_role()` or, when running from local notebook instance, use utils method `role = get_execution_role()` to create an execution role.
# +
try:
role = sagemaker.get_execution_role()
except:
role = get_execution_role()
print("Using IAM role arn: {}".format(role))
# -
# ### Install docker for `local` mode
#
# In order to work in `local` mode, you need to have docker installed. When running from your local machine, please make sure that you have docker or docker-compose (for local CPU machines) and nvidia-docker (for local GPU machines) installed. Alternatively, when running from a SageMaker notebook instance, you can simply run the following script to install dependencies.
#
# Note, you can only run a single local notebook at one time.
# Only run from SageMaker notebook instance
if local_mode:
# !/bin/bash ./common/setup.sh
# ## Build docker container
#
# Since we're working with a custom environment with custom dependencies, we create our own container for training. We:
#
# 1. Fetch the base MXNet and Coach container image,
# 2. Install EnergyPlus and its dependencies on top,
# 3. Upload the new container image to AWS ECR.
cpu_or_gpu = 'gpu' if instance_type.startswith('ml.p') else 'cpu'
repository_short_name = "sagemaker-hvac-coach-%s" % cpu_or_gpu
docker_build_args = {
'CPU_OR_GPU': cpu_or_gpu,
'AWS_REGION': boto3.Session().region_name,
}
custom_image_name = build_and_push_docker_image(repository_short_name, build_args=docker_build_args)
print("Using ECR image %s" % custom_image_name)
# ## Setup the environment
#
# The environment is defined in a Python file called `data_center_env.py` and for SageMaker training jobs, the file will be uploaded inside the `/src` directory.
#
# The environment implements the init(), step() and reset() functions that describe how the environment behaves. This is consistent with Open AI Gym interfaces for defining an environment.
#
# 1. `init()` - initialize the environment in a pre-defined state
# 2. `step()` - take an action on the environment
# 3. `reset()` - restart the environment on a new episode
# ## Configure the presets for RL algorithm
#
# The presets that configure the RL training jobs are defined in the “preset-energy-plus-clipped-ppo.py” file which is also uploaded as part of the `/src` directory. Using the preset file, you can define agent parameters to select the specific agent algorithm. You can also set the environment parameters, define the schedule and visualization parameters, and define the graph manager. The schedule presets will define the number of heat up steps, periodic evaluation steps, training steps between evaluations, etc.
#
# All of these can be overridden at run-time by specifying the `RLCOACH_PRESET` hyperparameter. Additionally, it can be used to define custom hyperparameters.
# !pygmentize src/preset-energy-plus-clipped-ppo.py
# ## Write the Training Code
#
# The training code is written in the file “train-coach.py” which is uploaded in the /src directory.
# First import the environment files and the preset files, and then define the main() function.
# !pygmentize src/train-coach.py
# ## Train the RL model using the Python SDK Script mode
#
# If you are using local mode, the training will run on the notebook instance. When using SageMaker for training, you can select a GPU or CPU instance. The RLEstimator is used for training RL jobs.
#
# 1. Specify the source directory where the environment, presets and training code is uploaded.
# 2. Specify the entry point as the training code
# 3. Specify the choice of RL toolkit and framework. This automatically resolves to the ECR path for the RL Container.
# 4. Define the training parameters such as the instance count, job name, S3 path for output and job name.
# 5. Specify the hyperparameters for the RL agent algorithm. The RLCOACH_PRESET can be used to specify the RL agent algorithm you want to use.
# 6. [optional] Define the metrics definitions that you are interested in capturing in your logs. These can also be visualized in CloudWatch and SageMaker Notebooks.
# +
estimator = RLEstimator(entry_point="train-coach.py",
source_dir='src',
dependencies=["common/sagemaker_rl"],
image_name=custom_image_name,
role=role,
train_instance_type=instance_type,
train_instance_count=1,
output_path=s3_output_path,
base_job_name=job_name_prefix,
hyperparameters = {
'save_model': 1
}
)
estimator.fit(wait=local_mode)
# -
# ## Store intermediate training output and model checkpoints
#
# The output from the training job above is stored on S3. The intermediate folder contains gifs and metadata of the training.
# +
job_name=estimator._current_job_name
print("Job name: {}".format(job_name))
s3_url = "s3://{}/{}".format(s3_bucket,job_name)
if local_mode:
output_tar_key = "{}/output.tar.gz".format(job_name)
else:
output_tar_key = "{}/output/output.tar.gz".format(job_name)
intermediate_folder_key = "{}/output/intermediate/".format(job_name)
output_url = "s3://{}/{}".format(s3_bucket, output_tar_key)
intermediate_url = "s3://{}/{}".format(s3_bucket, intermediate_folder_key)
print("S3 job path: {}".format(s3_url))
print("Output.tar.gz location: {}".format(output_url))
print("Intermediate folder path: {}".format(intermediate_url))
tmp_dir = "/tmp/{}".format(job_name)
os.system("mkdir {}".format(tmp_dir))
print("Create local folder {}".format(tmp_dir))
# -
# ## Visualization
# ### Plot metrics for training job
# We can pull the reward metric of the training and plot it to see the performance of the model over time.
# +
# %matplotlib inline
import pandas as pd
csv_file_name = "worker_0.simple_rl_graph.main_level.main_level.agent_0.csv"
key = os.path.join(intermediate_folder_key, csv_file_name)
wait_for_s3_object(s3_bucket, key, tmp_dir)
csv_file = "{}/{}".format(tmp_dir, csv_file_name)
df = pd.read_csv(csv_file)
df = df.dropna(subset=['Training Reward'])
x_axis = 'Episode #'
y_axis = 'Training Reward'
plt = df.plot(x=x_axis,y=y_axis, figsize=(12,5), legend=True, style='b-')
plt.set_ylabel(y_axis);
plt.set_xlabel(x_axis);
# -
# ## Evaluation of RL models
#
# We use the last checkpointed model to run evaluation for the RL Agent.
#
# ### Load checkpointed model
#
# Checkpointed data from the previously trained models will be passed on for evaluation / inference in the checkpoint channel. In local mode, we can simply use the local directory, whereas in the SageMaker mode, it needs to be moved to S3 first.
# +
wait_for_s3_object(s3_bucket, output_tar_key, tmp_dir)
if not os.path.isfile("{}/output.tar.gz".format(tmp_dir)):
raise FileNotFoundError("File output.tar.gz not found")
os.system("tar -xvzf {}/output.tar.gz -C {}".format(tmp_dir, tmp_dir))
if local_mode:
checkpoint_dir = "{}/data/checkpoint".format(tmp_dir)
else:
checkpoint_dir = "{}/checkpoint".format(tmp_dir)
print("Checkpoint directory {}".format(checkpoint_dir))
# -
if local_mode:
checkpoint_path = 'file://{}'.format(checkpoint_dir)
print("Local checkpoint file path: {}".format(checkpoint_path))
else:
checkpoint_path = "s3://{}/{}/checkpoint/".format(s3_bucket, job_name)
if not os.listdir(checkpoint_dir):
raise FileNotFoundError("Checkpoint files not found under the path")
os.system("aws s3 cp --recursive {} {}".format(checkpoint_dir, checkpoint_path))
print("S3 checkpoint file path: {}".format(checkpoint_path))
# ### Run the evaluation step
#
# Use the checkpointed model to run the evaluation step.
# +
estimator_eval = RLEstimator(entry_point="evaluate-coach.py",
source_dir='src',
dependencies=["common/sagemaker_rl"],
image_name=custom_image_name,
role=role,
train_instance_type=instance_type,
train_instance_count=1,
output_path=s3_output_path,
base_job_name=job_name_prefix,
hyperparameters = {
"RLCOACH_PRESET": "preset-energy-plus-clipped-ppo",
"evaluate_steps": 288*2, #2 episodes, i.e. 2 days
}
)
estimator_eval.fit({'checkpoint': checkpoint_path})
# -
# # Model deployment
# Since we specified a custom image when configuring the RLEstimator, we have to manually choose our deployment method. We'll choose to deploy with the MXNet Model since we used MXNet as the base container for our training.
from sagemaker.mxnet import MXNetModel
model = MXNetModel(model_data=estimator.model_data,
role=role,
entry_point='deploy-mxnet-coach.py',
source_dir='src',
dependencies=["common/sagemaker_rl"],
framework_version='1.3.0')
predictor = model.deploy(initial_instance_count=1,
instance_type=instance_type,
endpoint_name=job_name_prefix)
# We can test the endpoint with a samples observation, where the current room temperature is high. Since the environment vector was of the form `[outdoor_temperature, outdoor_humidity, indoor_humidity]` and we used observation normalization in our preset, we choose an observation of `[0, 0, 2]`. Since we're deploying a PPO model, our model returns both state value and actions.
action, action_mean, action_std = predictor.predict(np.array([0., 0., 2.,]))
action_mean
# We can see heating and cooling setpoints are returned from the model, and these can be used to control the HVAC system for efficient energy usage. More training iterations will help improve the model further.
# ### Clean up endpoint
predictor.delete_endpoint()
| reinforcement_learning/rl_hvac_coach_energyplus/rl_hvac_coach_energyplus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="top"></a>
# # Composites
# <hr>
#
# ## Background
#
# Composites are 2-dimensional representations of 3-dimensional data.
# There are many cases in which this is desired. Sometimes composites are used in visualization - such as showing an RGB image of an area. Other times they are used for convenience, such as reducing the run time of an analysis by reducing the amount of data to be processed in a task by working with composites instead of full datasets. Other times they are required by an algorithm.
#
# There are several kinds of composites that can be made. This notebook provides an overview of several of them and shows how to create them in the context of Open Data Cube.
# <hr>
#
# ## Index
#
# * [Import Dependencies and Connect to the Data Cube](#Composites_import)
# * [Load Data from the Data Cube](#Composites_retrieve_data)
# * [Most Common Composites](#Composites_most_common)
# * Mean composites
# * Median composites
# * Geometric median (geomedian) composites
# * Geometric medoid (geomedoid) composites
# * [Other Composites](#Composites_other_composites)
# * Most-recent composites
# * Least-recent composites
# ## <span id="Composites_import">Import Dependencies and Connect to the Data Cube [▴](#top)</span>
# +
import sys
import os
sys.path.append(os.environ.get('NOTEBOOK_ROOT'))
from utils.data_cube_utilities.clean_mask import landsat_clean_mask_full
# landsat_qa_clean_mask, landsat_clean_mask_invalid
from utils.data_cube_utilities.dc_mosaic import create_hdmedians_multiple_band_mosaic
from utils.data_cube_utilities.dc_mosaic import create_mosaic
from datacube.utils.aws import configure_s3_access
configure_s3_access(requester_pays=True)
import datacube
dc = datacube.Datacube()
# -
# ## <span id="Composites_retrieve_data">Load Data from the Data Cube [▴](#top)</span>
product = 'ls8_usgs_sr_scene'
platform = 'LANDSAT_8'
collection = 'c1'
level = 'l2'
landsat_ds = dc.load(platform=platform, product=product,
time=("2017-01-01", "2017-12-31"),
lat=(-1.395447, -1.172343),
lon=(36.621306, 37.033980),
group_by='solar_day',
dask_chunks={'latitude':500, 'longitude':500,
'time':5})
# clean_mask = (landsat_qa_clean_mask(landsat_ds, platform) &
# (landsat_ds != -9999).to_array().all('variable') &
# landsat_clean_mask_invalid(landsat_ds))
clean_mask = landsat_clean_mask_full(dc, landsat_ds, product=product, platform=platform,
collection=collection, level=level)
landsat_ds = landsat_ds.where(clean_mask)
# ## <span id="Composites_most_common">Most Common Composites [▴](#top)</span>
# ### Mean composites
# > A mean composite is obtained by finding the mean (average) value of each band for each pixel. To create mean composites, we use the built-in `mean()` method of xarray objects.
mean_composite = landsat_ds.mean('time', skipna=True)
# ### Median composites
# > A median composite is obtained by finding the median value of each band for each pixel. Median composites are quick to obtain and are usually fairly representative of their data, so they are acceptable for visualization as images. To create median composites, we use the built-in `median()` method of xarray objects.
#
# 
median_composite = landsat_ds.median('time', skipna=True)
# ### Geometric median (geomedian) composites
# > Geometric median (or "geomedian") composites are the best composites to use for most applications for which a representative, **synthetic** (calculated, not selected from the data) time slice is desired. They are essentiall median composites, but instead of finding the median on a per-band basis, they find the median for all bands together. If a composite will be used for analysis - not just visualization - it should be a geomedian composite. The only downside of this composite type is that it takes much longer to obtain than other composite types. For more information, see the [Geomedians_and_Geomedoids notebook](Geomedians_and_Geomedoids.ipynb).
geomedian_composite = create_hdmedians_multiple_band_mosaic(landsat_ds)
# ### Geometric medoid (geomedoid) composites
# > Geometric medoid (or "geomedoid") composites are the best composites to use for most applications for which a representative, **non-syntheic** (selected from the data, not calculated) time slice is desired. For more information, see the [Geomedians_and_Geomedoids notebook](Geomedians_and_Geomedoids.ipynb).
geomedoid_composite = create_hdmedians_multiple_band_mosaic(landsat_ds, operation='medoid')
# ## <span id="Composites_other_composites">Other Composites [▴](#top)</span>
# ### Most-recent composites
# > Most-recent composites use the most recent cloud-free pixels in an image. To create, a most-recent composite, we use the **create_mosaic** utility function.
#
# 
most_recent_composite = create_mosaic(landsat_ds)
# ### Least-recent composites
# > Least-recent composites are simply the opposite of most-recent composites. To create, a least-recent composite, we use the **create_mosaic** utility function, specifying `reverse_time=True`.
most_recent_composite = create_mosaic(landsat_ds, reverse_time=True)
| notebooks/compositing/Composites.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import matplotlib as plt
import matplotlib.pyplot as plt
# %matplotlib inline
sess = tf.InteractiveSession()
image = np.array([[[[1],[2],[3]],[[4],[5],[6]],[[7],[8],[9]]]], dtype = np.float32)
print(image.shape)
plt.imshow(image.reshape(3,3), cmap='Greys') #시각화
#1 => n개의 이미지#
# 3*3 size
#1 color
print("image.shape", image.shape)
weight = tf.constant([[[[1.]],[[1.]]],[[[1.]],[[1.]]]])
print("weight.shape", weight.shape)
conv2d = tf.nn.conv2d(image, weight, strides=[1,1,1,1],padding = 'SAME') #1*1 stride줌 #same을 했기때문에 input과 같은 사이즈나옴
conv2d_img = conv2d.eval()
print("conv2d_img.shape", conv2d_img.shape)
#시각화하기위한코드
conv2d_img = np.swapaxes(conv2d_img,0,3) #transposing 2D array
for i, one_img in enumerate(conv2d_img):
print(one_img.reshape(3,3))
plt.subplot(1,3,i+1), plt.imshow(one_img.reshape(3,3), cmap = 'gray')
print("image.shape", image.shape)
weight = tf.constant([[[[1.,10,-1.]],[[1.,10.,-1]]],[[[1.,10.,-1.]],[[1.,10.,-1.]]]])
print("weight.shape", weight.shape) #마지막 숫자 3은 필터의 갯수==나올 이미지의 갯수
#필터를 몇장을 쓰는가에 따라 하나의 이미지에 대하여 여러 이미지가 나옴.
conv2d = tf.nn.conv2d(image, weight, strides=[1,1,1,1],padding = 'SAME')
conv2d_img = conv2d.eval()
print("conv2d_img.shape", conv2d_img.shape)
#시각화 하기 위한 코드
conv2d_img = np.swapaxes(conv2d_img,0,3)
for i, one_img in enumerate(conv2d_img):
print(one_img.reshape(3,3))
plt.subplot(1,3,i+1), plt.imshow(one_img.reshape(3,3), cmap= 'gray')
#max pooling_어떤 데이터를 서브 샘플링한다.
# %matplotlib inline
image = np.array([[[[4],[3]],[[2],[1]]]],dtype = np.float32)
pool = tf.nn.max_pool(image, ksize= [1,2,2,1], strides = [1,1,1,1], padding = 'SAME')
#필터사이즈 #max_pool이 CNN과 잘 동작함
print(pool.shape)
print(pool.eval())
#최댓값 뽑아내는 것
#max pooling2
# %matplotlib inline
image = np.array([[[[4],[3]],[[2],[1]]]],dtype = np.float32)
pool = tf.nn.max_pool(image, ksize= [1,2,2,1], strides = [1,1,1,1], padding = 'VALID')
print(pool.shape)
print(pool.eval())
#실전 이미지에 넣기
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/",one_hot=True)
#데이터 읽어오기
img = mnist.train.images[5].reshape(28,28)
#가장 첫번째에 있는 거 불러와서 쉐잎을 28*28로 잡
plt.imshow(img, cmap = 'gray') #출력
# +
sess = tf.InteractiveSession()
img = img.reshape(-1,28,28,1) #28*28의 한 색깔, n개의 이미지일 때는 -1=(컴퓨터에게 알아서 계산해~라는 reshape의 방법)
W1 = tf.Variable(tf.random_normal([3, 3, 1, 5], stddev=0.01)) #칼라에 신경써야함.(1) (3*3은 필터의 사이즈) (5개의 필터사용)
conv2d = tf.nn.conv2d(img, W1, strides=[1, 2, 2, 1], padding='SAME')# (2*2는 필터를 2칸씩 옮기겠다.그래서 출력이 14*14)
print(conv2d)
sess.run(tf.global_variables_initializer())
#그림 출력하기는 code
conv2d_img = conv2d.eval()
conv2d_img = np.swapaxes(conv2d_img, 0, 3)
for i, one_img in enumerate(conv2d_img):
plt.subplot(1,5,i+1), plt.imshow(one_img.reshape(14,14), cmap='gray')
#5개의 서로다른 필터를 사용했기 때문에 서로다른 5개의 이미지가 나옴.
# +
#max pooling
pool = tf.nn.max_pool(conv2d, ksize=[1, 2, 2, 1], strides=[
1, 2, 2, 1], padding='SAME')
#입력 이미지 14*14인데 또 필터가 두탄씩 움직이니까 7*7로 또 줄어듦.
print(pool)
#실행코드
sess.run(tf.global_variables_initializer())
pool_img = pool.eval()
#출력 code
pool_img = np.swapaxes(pool_img, 0, 3)
for i, one_img in enumerate(pool_img):
plt.subplot(1,5,i+1), plt.imshow(one_img.reshape(7, 7), cmap='gray')
#7*7의 이미지가 나옴. 이미지가 서브 샘플링되어서 해상도가 떨어져있음. max pooling을 이용해서 간단한 서브 샘플링을 해봄.
# +
import tensorflow as tf
import random
# import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
tf.set_random_seed(777) # reproducibility
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True) #vector표현가능하게one_hot=True
# Check out https://www.tensorflow.org/get_started/mnist/beginners for
# more information about the mnist dataset
# hyper parameters
learning_rate = 0.001
training_epochs = 15
batch_size = 100
# dropout (keep_prob) rate 0.7~0.5 on training, but should be 1 for testing
keep_prob = tf.placeholder(tf.float32)
#Conv layer1
#input의 이미지를 우리가 원하는대로 만들어야
# input place holders
X = tf.placeholder(tf.float32, [None, 784]) #784인 이유는 MNIST가 28*28이기 때문
X_img = tf.reshape(X, [-1, 28, 28, 1]) # img 28x28x1(칼라) (black/white) #이미지로 넣기 위해서 #x_img가 입력이 될 것.
Y = tf.placeholder(tf.float32, [None, 10])
#첫번째 conv layer1
# L1 ImgIn shape=(?, 28, 28, 1)
W1 = tf.Variable(tf.random_normal([3, 3, 1, 32], stddev=0.01)) #3*3size의 필터 #1은 칼라 #32 필터
# Conv -> (?, 28, 28, 32)
# Pool -> (?, 14, 14, 32)
L1 = tf.nn.conv2d(X_img, W1, strides=[1, 1, 1, 1], padding='SAME')
L1 = tf.nn.relu(L1)
L1 = tf.nn.max_pool(L1, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
#필터 2*2 # stride 2*2 # 28*28->14*14
L1 = tf.nn.dropout(L1, keep_prob=keep_prob)
'''
Tensor("Conv2D:0", shape=(?, 28, 28, 32), dtype=float32)
Tensor("Relu:0", shape=(?, 28, 28, 32), dtype=float32)
Tensor("MaxPool:0", shape=(?, 14, 14, 32), dtype=float32)
Tensor("dropout/mul:0", shape=(?, 14, 14, 32), dtype=float32)
'''
#conv layer2
# L2 ImgIn shape=(?, 14, 14, 32)
W2 = tf.Variable(tf.random_normal([3, 3, 32, 64], stddev=0.01)) #32는 필터의 갯수와 같아야함. 64개의 필터를 쓸 것
# Conv ->(?, 14, 14, 64)
# Pool ->(?, 7, 7, 64)
L2 = tf.nn.conv2d(L1, W2, strides=[1, 1, 1, 1], padding='SAME')
L2 = tf.nn.relu(L2)
L2 = tf.nn.max_pool(L2, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
L2 = tf.nn.dropout(L2, keep_prob=keep_prob)
'''
Tensor("Conv2D_1:0", shape=(?, 14, 14, 64), dtype=float32)
Tensor("Relu_1:0", shape=(?, 14, 14, 64), dtype=float32)
Tensor("MaxPool_1:0", shape=(?, 7, 7, 64), dtype=float32)
Tensor("dropout_1/mul:0", shape=(?, 7, 7, 64), dtype=float32)
'''
#conv layer3
# L3 ImgIn shape=(?, 7, 7, 64)
W3 = tf.Variable(tf.random_normal([3, 3, 64, 128], stddev=0.01))
# Conv ->(?, 7, 7, 128)
# Pool ->(?, 4, 4, 128)
# Reshape ->(?, 4 * 4 * 128) # Flatten them for FC
L3 = tf.nn.conv2d(L2, W3, strides=[1, 1, 1, 1], padding='SAME')
L3 = tf.nn.relu(L3)
L3 = tf.nn.max_pool(L3, ksize=[1, 2, 2, 1], strides=[
1, 2, 2, 1], padding='SAME')
L3 = tf.nn.dropout(L3, keep_prob=keep_prob)
#입체적인 모양을 쭉 펼쳐야됨. 128*4*4만크므이 길이를 갖는게 n개 있게됨.
L3_flat = tf.reshape(L3, [-1, 128 * 4 * 4])
'''
Tensor("Conv2D_2:0", shape=(?, 7, 7, 128), dtype=float32)
Tensor("Relu_2:0", shape=(?, 7, 7, 128), dtype=float32)
Tensor("MaxPool_2:0", shape=(?, 4, 4, 128), dtype=float32)
Tensor("dropout_2/mul:0", shape=(?, 4, 4, 128), dtype=float32)
Tensor("Reshape_1:0", shape=(?, 2048), dtype=float32)
'''
#FC를 2번하겠다.그럼 정확도가 높아짐.
#Conv layer4
# L4 FC 4x4x128 inputs -> 625 outputs
#벡터 입력의 값 = 128*4*4 출력의 값 625개
W4 = tf.get_variable("W4", shape=[128 * 4 * 4, 625],
initializer=tf.contrib.layers.xavier_initializer())
#bias를 출력의 값과 똑같게 줌 (625)
b4 = tf.Variable(tf.random_normal([625]))
#곱하고 더한다.
L4 = tf.nn.relu(tf.matmul(L3_flat, W4) + b4)
#학습할때는 dropout을 0.5나 0.7로
L4 = tf.nn.dropout(L4, keep_prob=keep_prob)
'''
Tensor("Relu_3:0", shape=(?, 625), dtype=float32)
Tensor("dropout_3/mul:0", shape=(?, 625), dtype=float32)
'''
#Fully connecter(fc,dense) layer
# L5 Final FC 625 inputs 입력받아서 -> 10 outputs
W5 = tf.get_variable("W5", shape=[625, 10],
initializer=tf.contrib.layers.xavier_initializer())
b5 = tf.Variable(tf.random_normal([10]))
logits = tf.matmul(L4, W5) + b5
'''
Tensor("add_1:0", shape=(?, 10), dtype=float32)
'''
# define cost/loss & optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
#Training and Evaluation
# initialize
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# 학습시키기
print('Learning started. It takes sometime.')
for epoch in range(training_epochs):
avg_cost = 0
total_batch = int(mnist.train.num_examples / batch_size)
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
feed_dict = {X: batch_xs, Y: batch_ys, keep_prob: 0.7}
c, _ = sess.run([cost, optimizer], feed_dict=feed_dict)
avg_cost += c / total_batch
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.9f}'.format(avg_cost))
print('Learning Finished!')
# 학습이 잘 되었는지 평가하기
# if you have a OOM error, please refer to lab-11-X-mnist_deep_cnn_low_memory.py
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print('Accuracy:', sess.run(accuracy, feed_dict={
X: mnist.test.images, Y: mnist.test.labels, keep_prob: 1}))
#test할 때는 반드시 dropout을 1로
# Get one and predict
r = random.randint(0, mnist.test.num_examples - 1)
print("Label: ", sess.run(tf.argmax(mnist.test.labels[r:r + 1], 1)))
print("Prediction: ", sess.run(
tf.argmax(logits, 1), feed_dict={X: mnist.test.images[r:r + 1], keep_prob: 1}))
# plt.imshow(mnist.test.images[r:r + 1].
# reshape(28, 28), cmap='Greys', interpolation='nearest')
# plt.show(
#cost가 떨어지면서99프로까지 정확성을 얻을 수 있음.
# +
#위의 코드는 관리하기 불편함.그래서 python의 class으로 보다 효과적으로 관리.
# Lab 11 MNIST and Deep learning CNN
import tensorflow as tf
# import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
tf.set_random_seed(777) # reproducibility
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# Check out https://www.tensorflow.org/get_started/mnist/beginners for
# more information about the mnist dataset
# hyper parameters
learning_rate = 0.001
training_epochs = 15
batch_size = 100
class Model:
#초기화
def __init__(self, sess, name):
self.sess = sess
self.name = name
self._build_net()
#네트워크를 빌드하는 건 다 넣음.
def _build_net(self):
with tf.variable_scope(self.name):
# dropout (keep_prob) rate 0.7~0.5 on training, but should be 1
# for testing
self.keep_prob = tf.placeholder(tf.float32)
# input place holders
self.X = tf.placeholder(tf.float32, [None, 784])
# img 28x28x1 (black/white)
X_img = tf.reshape(self.X, [-1, 28, 28, 1])
self.Y = tf.placeholder(tf.float32, [None, 10])
# L1 ImgIn shape=(?, 28, 28, 1)
W1 = tf.Variable(tf.random_normal([3, 3, 1, 32], stddev=0.01))
# Conv -> (?, 28, 28, 32)
# Pool -> (?, 14, 14, 32)
L1 = tf.nn.conv2d(X_img, W1, strides=[1, 1, 1, 1], padding='SAME')
L1 = tf.nn.relu(L1)
L1 = tf.nn.max_pool(L1, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
L1 = tf.nn.dropout(L1, keep_prob=self.keep_prob)
'''
Tensor("Conv2D:0", shape=(?, 28, 28, 32), dtype=float32)
Tensor("Relu:0", shape=(?, 28, 28, 32), dtype=float32)
Tensor("MaxPool:0", shape=(?, 14, 14, 32), dtype=float32)
Tensor("dropout/mul:0", shape=(?, 14, 14, 32), dtype=float32)
'''
# L2 ImgIn shape=(?, 14, 14, 32)
W2 = tf.Variable(tf.random_normal([3, 3, 32, 64], stddev=0.01))
# Conv ->(?, 14, 14, 64)
# Pool ->(?, 7, 7, 64)
L2 = tf.nn.conv2d(L1, W2, strides=[1, 1, 1, 1], padding='SAME')
L2 = tf.nn.relu(L2)
L2 = tf.nn.max_pool(L2, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
L2 = tf.nn.dropout(L2, keep_prob=self.keep_prob)
'''
Tensor("Conv2D_1:0", shape=(?, 14, 14, 64), dtype=float32)
Tensor("Relu_1:0", shape=(?, 14, 14, 64), dtype=float32)
Tensor("MaxPool_1:0", shape=(?, 7, 7, 64), dtype=float32)
Tensor("dropout_1/mul:0", shape=(?, 7, 7, 64), dtype=float32)
'''
# L3 ImgIn shape=(?, 7, 7, 64)
W3 = tf.Variable(tf.random_normal([3, 3, 64, 128], stddev=0.01))
# Conv ->(?, 7, 7, 128)
# Pool ->(?, 4, 4, 128)
# Reshape ->(?, 4 * 4 * 128) # Flatten them for FC
L3 = tf.nn.conv2d(L2, W3, strides=[1, 1, 1, 1], padding='SAME')
L3 = tf.nn.relu(L3)
L3 = tf.nn.max_pool(L3, ksize=[1, 2, 2, 1], strides=[
1, 2, 2, 1], padding='SAME')
L3 = tf.nn.dropout(L3, keep_prob=self.keep_prob)
L3_flat = tf.reshape(L3, [-1, 128 * 4 * 4])
'''
Tensor("Conv2D_2:0", shape=(?, 7, 7, 128), dtype=float32)
Tensor("Relu_2:0", shape=(?, 7, 7, 128), dtype=float32)
Tensor("MaxPool_2:0", shape=(?, 4, 4, 128), dtype=float32)
Tensor("dropout_2/mul:0", shape=(?, 4, 4, 128), dtype=float32)
Tensor("Reshape_1:0", shape=(?, 2048), dtype=float32)
'''
# L4 FC 4x4x128 inputs -> 625 outputs
W4 = tf.get_variable("W4", shape=[128 * 4 * 4, 625],
initializer=tf.contrib.layers.xavier_initializer())
b4 = tf.Variable(tf.random_normal([625]))
L4 = tf.nn.relu(tf.matmul(L3_flat, W4) + b4)
L4 = tf.nn.dropout(L4, keep_prob=self.keep_prob)
'''
Tensor("Relu_3:0", shape=(?, 625), dtype=float32)
Tensor("dropout_3/mul:0", shape=(?, 625), dtype=float32)
'''
# L5 Final FC 625 inputs -> 10 outputs
W5 = tf.get_variable("W5", shape=[625, 10],
initializer=tf.contrib.layers.xavier_initializer())
b5 = tf.Variable(tf.random_normal([10]))
self.logits = tf.matmul(L4, W5) + b5
'''
Tensor("add_1:0", shape=(?, 10), dtype=float32)
'''
# define cost/loss & optimizer
self.cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(
logits=self.logits, labels=self.Y))
self.optimizer = tf.train.AdamOptimizer(
learning_rate=learning_rate).minimize(self.cost)
correct_prediction = tf.equal(
tf.argmax(self.logits, 1), tf.argmax(self.Y, 1))
self.accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
#예측하는거
def predict(self, x_test, keep_prop=1.0):
return self.sess.run(self.logits, feed_dict={self.X: x_test, self.keep_prob: keep_prop})
#정확도를 얻는 것
def get_accuracy(self, x_test, y_test, keep_prop=1.0):
return self.sess.run(self.accuracy, feed_dict={self.X: x_test, self.Y: y_test, self.keep_prob: keep_prop})
#학습하는 서
def train(self, x_data, y_data, keep_prop=0.7):
return self.sess.run([self.cost, self.optimizer], feed_dict={
self.X: x_data, self.Y: y_data, self.keep_prob: keep_prop})
# initialize
sess = tf.Session()
#model1을 만듦.
m1 = Model(sess, "m1")
sess.run(tf.global_variables_initializer())
print('Learning Started!')
# train my model
for epoch in range(training_epochs):
avg_cost = 0
total_batch = int(mnist.train.num_examples / batch_size)
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
#session run할 필요없이 ml에 train 함수를 호출해버리면 됨.
#깔끔하게 관리가 됨.
c, _ = m1.train(batch_xs, batch_ys)
avg_cost += c / total_batch
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.9f}'.format(avg_cost))
print('Learning Finished!')
# Test model and check accuracy
print('Accuracy:', m1.get_accuracy(mnist.test.images, mnist.test.labels))
| 2017/CS_20SI/code/CNN_Using_MNIST_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
import boto3
from multiprocessing import Pool
metadata = pd.read_csv("/Users/phoenix.logan/code/skeeters/data/CMS001_CMS002_MergedAnnotations_190325.csv")
metadata.head(n=2)
metadata = pd.read_csv("/Users/phoenix.logan/code/skeeters/data/CMS001_CMS002_MergedAnnotations_190325.csv")
metadata_new = metadata[["czbiohub-mosquito_sequences_id", "raw_sequence_run_directory"]]
metadata_new = metadata_new.rename(
index=str,
columns={
"czbiohub-mosquito_sequences_id": "id",
"raw_sequence_run_directory": "read1"
},
)
# +
def fix_filepaths(seq_id, path):
# change czb-seqs bucket to czbiohub-seqs
path_new = path.replace("czbiohub-seqbot", "czb-seqbot")
if path_new.endswith(".gz"):
# change R2 to R1 for read1_path (fix data inconsistencies)
return path_new.replace("_R2_", "_R1_")
# elif path_new.endswith("/"):
# return f"{path_new}{seq_id}_R1_001.fastq.gz"
else:
return os.path.join(path_new, f"{seq_id}_R1_001.fastq.gz")
def read_pair(read1):
return read1.replace("_R1_", "_R2_")
metadata_new["read1"] = metadata_new.apply(lambda row: fix_filepaths(row['id'], row['read1']), axis=1)
metadata_new["read1"] = metadata_new["read1"].apply(lambda x: f"s3://{x}")
metadata_new["read2"] = metadata_new["read1"].apply(lambda x: read_pair(x))
metadata_new[["id", "read1", "read2"]].to_csv("metadata_formatted.csv", index=False)
# +
import boto3
from botocore.exceptions import ClientError
s3 = boto3.resource('s3')
def check(bucket, key):
try:
# try to load file
s3.Object(bucket, key).load()
except ClientError as e:
# if file DNE (404),
print(f"{bucket}, {key} could not load, error message: {e.response['Error']['Code']}")
#return int(e.response['Error']['Code']) != 404
return False
print(f"{bucket} with key {key} already loaded, skipping...")
return True
# +
def run_s3_command(cmd):
cmd_strip = cmd.strip("s3://")
cmd_split = cmd_strip.split("/")
bucket_name = cmd_split.pop(0)
r1_key= ("/").join(cmd_split)
r2_key = r1_key.replace("_R1_", "_R2_")
for key in [r1_key, r2_key]:
copy_source = {
'Bucket': 'czbiohub-seqbot',
'Key': key
}
if not check('czb-seqbot', copy_source["Key"]):
print(f"copy source: ", copy_source)
try:
print(f"uploading {copy_source['Key']} to new bucket")
s3.meta.client.copy(copy_source, 'czb-seqbot', key)
except:
print(f"ERROR UPLOADING: {copy_source}")
return False
return True
fps = [i for i in metadata_new["read1"]][5:]
with Pool(5) as p:
res = p.map(run_s3_command, fps)
# -
metadata_new[["id", "read1", "read2"]].to_csv("metadata_formatted.csv", index=False)
fps = [i for i in metadata_new["read1_path"]][5:]
data = {'sequence_id': ["CMS_001_RNA_A_S1", "CMS_002_RNA_A_S1", "CMS_003_RNA_A_S2"],
'read1_path': [
's3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_001_RNA_A_S1_R1_001.fasta.gz',
's3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_002_RNA_A_S1_R1_001.fasta.gz',
's3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_003_RNA_A_S2_R1_001.fasta.gz'
],
'read2_path': ['s3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_001_RNA_A_S1_R2_001.fasta.gz',
's3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_002_RNA_A_S1_R2_001.fasta.gz',
's3://czbiohub-mosquito/sequences/CMS001_fasta.gz/CMS_003_RNA_A_S2_R2_001.fasta.gz'
]
}
skeeter_metadata = pd.DataFrame.from_dict(data)
skeeter_metadata.to_csv("metadata_formatted_skeeters.csv", index=False)
metadata_new[["id", "read1", "read2"]]
| file_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.5 Levenshtein bis
# ## Problema: calcular el cost d'edició entre un patró i un text.
# Suposem que estem fent un problema de Levenshtein clàssic on els costos de substitució, inserció i eliminació són tots iguals a 1. Volem comparar les paraules el patró 'VERVE' amb el text 'BARBER'. Tenim la següent taula:
# | | * | B | A | R | B | E | R |
# | :-: | :-: | :-: | :-: | :-: | :-: | :-: | :-:|
# | * | 0 | 1 | 2 | 3 | 4 | 5 | 6 |
# | **V** | 1 | | | | | | |
# | **E** | 2 | | | | | | |
# | **R** | 3 | | | | | | |
# | **V** | 4 | | | | | | |
# | **E** | 5 | | | | | | |
# **Pregunta 1**. El patró i el text són intercanviables? Prova de fer la taula amb els seus rols intercanviats i observa si es fa el mateix tipus de canvis (per ex. insercions i eliminacions [deletions])
#
# Sí.
# **Pregunta 2**. Explica raonadament d'on surten els valors de la primera fila i de la primera columna. Explica el cas concret del valor 4 de la primera columna i quarta fila (al costat de la V).
#
# Els valors de la primera fila i la primera columna són els costos d'edició des d'una cadena buida. Per exemple el valor 4 de la primera columna i quarta fila és el cost d'inserir les 4 lletres, ja que partim d'una cadena buida.
# Suposem que hem omplert la taula tal i com s'indica
#
#
# | | * | B | A | R | B | E | R |
# | :-: | :-: | :-: | :-: | :-: | :-: | :-: | :-:|
# | * | 0 | 1 | 2 | 3 | 4 | 5 | 6 |
# | **V** | 1 | 1 | 2 | 3 | 4 | 5 | 6 |
# | **E** | 2 | 2 | 2 | 3 | 4 | **4** | |
# | **R** | 3 | | | | | | |
# | **V** | 4 | | | | | | |
# | **E** | 5 | | | | | | |
# **Pregunta 3**. Si ens parem a la casella assenyalada en negreta, quin és el text i quin el patró en aquest moment? Quina és la distància d'edició entre ells? Esmenta quines operacions (substitució, inserció, eliminació) has seguit per arribar a aquesta casella (si hi ha més d'una possibilitat, només cal que expliquis una).
#
# El text és BARBE i el patró és VE. La distància entre ells és de 4.
#
# Substituïm V per B (BE) -> +1
#
# Inserim A (BAE) -> +1
#
# Inserim R (BARE) -> +1
#
# Inserim B (BARBE) -> +1
# ## Problema 2: calcular la distància entre un patró i el substring més semblant d'un text.
# En aquest problema, no es penalitza moure el patró al llarg del text, ja que el substring pot estar situat en qualsevol posició. A més, treballarem amb els costos que us indica la pràctica, és a dir, les operacions de substitució i inserció costen 1, però l'eliminació costa 2. Suposa que volem comparar el patró 'VERVE' al text 'BARBER'. Per això, tenim la següent taula:
# | | * | B | A | R | B | E | R |
# | :-: | :-: | :-: | :-: | :-: | :-: | :-: | :-:|
# | * | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
# | **V** | 2 | | | | | | |
# | **E** | 4 | | | | | | |
# | **R** | 6 | | | | | | |
# | **V** | 8 | | | | | | |
# | **E** | 10 | | | | | | |
# **Pregunta 4**. Explica raonadament d'on surten els valors de la primera fila i la primera columna.
#
# Els valors de la primera fila son tots 0 perquè considerem que el cost de desplaçar la cadena és nul.
#
# Els valors de la primera columna van de 2 en 2 perquè ara considerem que el cost d'eliminació és 2.
# Suposem que hem omplert la taula tal i com s'indica
#
# | | * | B | A | R | B | E | R |
# | :-: | :-: | :-: | :-: | :-: | :-: | :-: | :-:|
# | * | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
# | **V** | 2 | 2 | 1 | 1 | 1 | 1 | 1 |
# | **E** | 4 | 3 | 2 | 2 | 2 | 1 | 2 |
# | **R** | | | | | | | |
# | **V** | | | | | | | |
# | **E** | | | | | | | |
# **Pregunta 5.** Hem calculat ja diverses possibilitats i substrings del text amb part del patró. Quina és la distància mínima trobada entre el patró analitzat fins ara i el substring més semblant del text? Quin és aquest patró i el substring corresponent?
#
# La distància mínima entre el patró i el substring més semblant del text és de només 1.
#
# El patró és VER i el substring és BER. El cost és 1 perquè només substituïm la V per la B.
| Algorismica/4.5_teoric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 11: Transformers
#
# In today's lab, we will learn how to create a transformer from scratch, then
# we'll take a look at ViT (the visual transformer). Some of the material in this
# lab comes from the following online sources:
#
# - https://medium.com/the-dl/transformers-from-scratch-in-pytorch-8777e346ca51
# - https://towardsdatascience.com/how-to-code-the-transformer-in-pytorch-24db27c8f9ec
# - https://towardsdatascience.com/implementing-visualttransformer-in-pytorch-184f9f16f632
# - https://github.com/lucidrains/vit-pytorch#vision-transformer---pytorch
# - https://medium.com/mlearning-ai/vision-transformers-from-scratch-pytorch-a-step-by-step-guide-96c3313c2e0c
#
# <img src="img/optimus_prime.jpg" title="Transformer" style="width: 600px;" />
#
# The above photo needs a credit!
# ## Transformers and machine learning trends
#
# Before the arrival of transformers, CNNs were most often used in the visual domain, while RNNs like LSTMs were most often used in NLP.
# There were many attempts at crossover, without much real success. Neither approach seemed capable of dealing with very large complex
# natural language datasets effectively.
#
# In 2017, the Transformer was introduced. "Attention is all you need" has been cited more than 38,000 times.
#
# The main concept in a Transformer is self-attention, which replaces the sequential processing of RNNs and the local
# processing of CNNs with the ability to adaptively extract arbitrary relationships between different elements of its input,
# output, and memory state.
#
# ## Transformer architecture
#
# We will use [<NAME>'s implementation of the Transformer in PyTorch](https://github.com/fkodom/transformer-from-scratch/tree/main/src).
#
# The architecture of the transformer looks like this:
#
# <img src="img/Transformer.png" title="Transformer" style="width: 600px;" />
#
# Here is a summary of the Transformer's details and mathematics:
#
# <img src="img/SummaryTransformer.PNG" title="Transformer Details" style="width: 1000px;" />
#
# There are several processes that we need to implement in the model. We go one by one.
# ## Attention
#
# Before Transformers, the standard model for sequence-to-sequence learning was seq2seq, which combines an RNN for encoding with
# an RNN for decoding. The encoder processes the input and retains important information in a sequence or block of memory,
# while the decoder extracts the important information from the memory in order to produce an output.
#
# One problem with seq2seq is that some information may be lost while processing a long sequence.
# Attention allows us to focus on specific inputs directly.
#
# An attention-based decoder, when we want to produce the output token at a target position, will calculate an attention score
# with the encoder's memory at each input position. A high score for a particular encoder position indicates that it is more important
# than another position. We essentially use the decoder's input to select which encoder output(s) should be used to calculate the
# current decoder output. Given decoder input $q$ (the *query*) and encoder outputs $p_i$, the attention operation calculates dot
# products between $q$ and each $p_i$. The dot products give the similarity of each pair. The dot products are softmaxed to get
# positive weights summing to 1, and the weighted average $r$ is calculated as
#
# $$r = \sum_i \frac{e^{p_i\cdot q}}{\sum_j e^{p_j\cdot q}}p_i .$$
#
# We can think of $r$ as an adaptively selected combination of the inputs most relevant to producing an output.
#
# ### Multi-head self attention
#
# Transformers use a specific type of attention mechanism, referred to as multi-head self attention.
# This is the most important part of the model. An illustration from the paper is shown below.
#
# <img src="img/MultiHeadAttention.png" title="Transformer" style="width: 600px;" />
#
# The multi-head attention layer is described as:
#
# $$\text{Attention}(Q,K,V)=\text{softmax}(\frac{QK^T}{\sqrt{d_k}})V$$
#
# $Q$, $K$, and $V$ are batches of matrices, each with shape <code>(batch_size, seq_length, num_features)</code>.
# When we are talking about *self* attention, each of the three matrices in
# each batch is just a separate linear projection of the same input $\bar{h}_t^{l-1}$.
#
# Multiplying the query $Q$ with the key $K$ arrays results in a <code>(batch_size, seq_length, seq_length)</code> array,
# which tells us roughly how important each element in the sequence is to each other element in the sequence. These dot
# products are converted to normalized weights using a softmax across rows, so that each row of weights sums to one.
# Finally, the weight matrix attention is applied to the value ($V$) array using matrix multiplication. We thus get,
# for each token in the input sequence, a weighted average of the rows of $V$, each of which corresponds to one of the
# elements in the input sequence.
#
# Here is code for the scaled dot-product operation that is part of a multi-head attention layer:
# +
import torch
import torch.nn.functional as f
from torch import Tensor, nn
def scaled_dot_product_attention(query: Tensor, key: Tensor, value: Tensor) -> Tensor:
# MatMul operations are translated to torch.bmm in PyTorch
temp = query.bmm(key.transpose(1, 2))
scale = query.size(-1) ** 0.5
softmax = f.softmax(temp / scale, dim=-1)
return softmax.bmm(value)
# -
# A multi-head attention module is composed of several identical
# *attention head* modules.
# Each attention head contains three linear transformations for $Q$, $K$, and $V$ and combines them using scaled dot-product attention.
# Note that this attention head could be used for self attention or another type of attention such as decoder-to-encoder attention, since
# we keep $Q$, $K$, and $V$ separate.
class AttentionHead(nn.Module):
def __init__(self, dim_in: int, dim_q: int, dim_k: int):
super().__init__()
self.q = nn.Linear(dim_in, dim_q)
self.k = nn.Linear(dim_in, dim_k)
self.v = nn.Linear(dim_in, dim_k)
def forward(self, query: Tensor, key: Tensor, value: Tensor) -> Tensor:
return scaled_dot_product_attention(self.q(query), self.k(key), self.v(value))
# Multiple attention heads can be combined with the output concatenation and linear transformation to construct a multi-head attention layer:
class MultiHeadAttention(nn.Module):
def __init__(self, num_heads: int, dim_in: int, dim_q: int, dim_k: int):
super().__init__()
self.heads = nn.ModuleList(
[AttentionHead(dim_in, dim_q, dim_k) for _ in range(num_heads)]
)
self.linear = nn.Linear(num_heads * dim_k, dim_in)
def forward(self, query: Tensor, key: Tensor, value: Tensor) -> Tensor:
return self.linear(
torch.cat([h(query, key, value) for h in self.heads], dim=-1)
)
# Each attention head computes its own transformation of the query, key, and value arrays,
# and then applies scaled dot-product attention. Conceptually, this means each head can attend to a different part of the input sequence, independent of the others. Increasing the number of attention heads allows the model to pay attention to more parts of the sequence at
# once, which makes the model more powerful.
# ### Positional Encoding
#
# To complete the transformer encoder, we need another component, the *position encoder*.
# The <code>MultiHeadAttention</code> class we just write has no trainable components that depend on a token's position
# in the sequence (axis 1 of the input tensor). Meaning all of the weight matrices we have seen so far
# *perform the same calculation for every input position*; that is, we don't have any position-dependent weights.
# All of the operations so far operate over the *feature dimension* (axis 2). This is good in that the model is compatible with any sequence
# length. But without *any* information about position, our model is going to be unable to differentiate between different orderings of
# the input -- we'll get the same result regardless of the order of the tokens in the input.
#
# Since order matters ("Ridgemont was in the store" has a different
# meaning from "The store was in Ridgemont"), we need some way to provide the model with information about tokens' positions in the input sequence.
# Whatever strategy we use should provide information about the relative position of data points in the input sequences.
# In the Transformer, positional information is encoded using trigonometric functions in a constant 2D matrix $PE$:
#
# $$PE_{(pos,2i)}=\sin (\frac{pos}{10000^{2i/d_{model}}})$$
# $$PE_{(pos,2i+1)}=\cos (\frac{pos}{10000^{2i/d_{model}}}),$$
#
# where $pos$ refers to a position in the input sentence sequence and $i$ refers to the position along the embedding vector dimension.
# This matrix is *added* to the matrix consisting of the embeddings of each of the input tokens:
#
# <img src="img/positionalencoder.png" title="Positional Encoder" style="width: 400px;" />
#
# Position encoding can implemented as follows (put this in `utils.py`):
def position_encoding(seq_len: int, dim_model: int, device: torch.device = torch.device("cpu")) -> Tensor:
pos = torch.arange(seq_len, dtype=torch.float, device=device).reshape(1, -1, 1)
dim = torch.arange(dim_model, dtype=torch.float, device=device).reshape(1, 1, -1)
phase = pos / (1e4 ** (dim // dim_model))
return torch.where(dim.long() % 2 == 0, torch.sin(phase), torch.cos(phase))
# These sinusoidal encodings allow us to work with arbirary length sequences because the sine and cosine functions are periodic in the range
# $[-1, 1]$. One hope is that if during inference we are provided with an input sequence longer than any found during training.
# The position encodings of the last elements in the sequence would be different from anything the model has seen before, but with the
# periodic sine/cosine encoding, there will still be some similar structure, with the new encodings being very similar to neighboring encodings the model has seen before. For this reason, despite the fact that learned embeddings appeared to perform equally as well, the authors chose
# this fixed sinusoidal encoding.
# ### The complete encoder
#
# The transformer uses an encoder-decoder architecture. The encoder processes the input sequence and returns a sequence of
# feature vectors or memory vectors, while the decoder outputs a prediction of the target sequence,
# incorporating information from the encoder memory.
#
# First, let's complete the transformer layer with the two-layer feed forward network. Put this in `utils.py`:
def feed_forward(dim_input: int = 512, dim_feedforward: int = 2048) -> nn.Module:
return nn.Sequential(
nn.Linear(dim_input, dim_feedforward),
nn.ReLU(),
nn.Linear(dim_feedforward, dim_input),
)
# Let's create a residual module to encapsulate the feed forward network or attention
# model along with the common dropout and LayerNorm operations (also in `utils.py`):
class Residual(nn.Module):
def __init__(self, sublayer: nn.Module, dimension: int, dropout: float = 0.1):
super().__init__()
self.sublayer = sublayer
self.norm = nn.LayerNorm(dimension)
self.dropout = nn.Dropout(dropout)
def forward(self, *tensors: Tensor) -> Tensor:
# Assume that the "query" tensor is given first, so we can compute the
# residual. This matches the signature of 'MultiHeadAttention'.
return self.norm(tensors[0] + self.dropout(self.sublayer(*tensors)))
# Now we can create the complete encoder! Put this in `encoder.py`. First, the encoder layer
# module, which comprised a self attention residual block followed by a fully connected residual block:
class TransformerEncoderLayer(nn.Module):
def __init__(
self,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
dim_q = dim_k = max(dim_model // num_heads, 1)
self.attention = Residual(
MultiHeadAttention(num_heads, dim_model, dim_q, dim_k),
dimension=dim_model,
dropout=dropout,
)
self.feed_forward = Residual(
feed_forward(dim_model, dim_feedforward),
dimension=dim_model,
dropout=dropout,
)
def forward(self, src: Tensor) -> Tensor:
src = self.attention(src, src, src)
return self.feed_forward(src)
# Then the Transformer encoder just encapsulates several transformer encoder layers:
class TransformerEncoder(nn.Module):
def __init__(
self,
num_layers: int = 6,
dim_model: int = 512,
num_heads: int = 8,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
self.layers = nn.ModuleList(
[
TransformerEncoderLayer(dim_model, num_heads, dim_feedforward, dropout)
for _ in range(num_layers)
]
)
def forward(self, src: Tensor) -> Tensor:
seq_len, dimension = src.size(1), src.size(2)
src += position_encoding(seq_len, dimension)
for layer in self.layers:
src = layer(src)
return src
# ### The decoder
#
# The decoder module is quite similar to the encoder, with just a few small differences:
# - The decoder accepts two inputs (the target sequence and the encoder memory), rather than one input.
# - There are two multi-head attention modules per layer (the target sequence self-attention module and the decoder-encoder attention module) rather than just one.
# - The second multi-head attention module, rather than strict self attention, expects the encoder memory as $K$ and $V$.
# - Since accessing future elements of the target sequence would be "cheating," we need to mask out future elements of the input target sequence.
#
# First, we have the decoder version of the transformer layer and the decoder module itself:
# +
class TransformerDecoderLayer(nn.Module):
def __init__(
self,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
dim_q = dim_k = max(dim_model // num_heads, 1)
self.attention_1 = Residual(
MultiHeadAttention(num_heads, dim_model, dim_q, dim_k),
dimension=dim_model,
dropout=dropout,
)
self.attention_2 = Residual(
MultiHeadAttention(num_heads, dim_model, dim_q, dim_k),
dimension=dim_model,
dropout=dropout,
)
self.feed_forward = Residual(
feed_forward(dim_model, dim_feedforward),
dimension=dim_model,
dropout=dropout,
)
def forward(self, tgt: Tensor, memory: Tensor) -> Tensor:
tgt = self.attention_1(tgt, tgt, tgt)
tgt = self.attention_2(tgt, memory, memory)
return self.feed_forward(tgt)
class TransformerDecoder(nn.Module):
def __init__(
self,
num_layers: int = 6,
dim_model: int = 512,
num_heads: int = 8,
dim_feedforward: int = 2048,
dropout: float = 0.1,
):
super().__init__()
self.layers = nn.ModuleList(
[
TransformerDecoderLayer(dim_model, num_heads, dim_feedforward, dropout)
for _ in range(num_layers)
]
)
self.linear = nn.Linear(dim_model, dim_model)
def forward(self, tgt: Tensor, memory: Tensor) -> Tensor:
seq_len, dimension = tgt.size(1), tgt.size(2)
tgt += position_encoding(seq_len, dimension)
for layer in self.layers:
tgt = layer(tgt, memory)
return torch.softmax(self.linear(tgt), dim=-1)
# -
# Note that there is not, as of yet, any masked attention implementation here!
# Making this version of the Transformer work in practice would require at least that.
#
# ### Putting it together
#
# Now we can put the encoder and decoder together:
class Transformer(nn.Module):
def __init__(
self,
num_encoder_layers: int = 6,
num_decoder_layers: int = 6,
dim_model: int = 512,
num_heads: int = 6,
dim_feedforward: int = 2048,
dropout: float = 0.1,
activation: nn.Module = nn.ReLU(),
):
super().__init__()
self.encoder = TransformerEncoder(
num_layers=num_encoder_layers,
dim_model=dim_model,
num_heads=num_heads,
dim_feedforward=dim_feedforward,
dropout=dropout,
)
self.decoder = TransformerDecoder(
num_layers=num_decoder_layers,
dim_model=dim_model,
num_heads=num_heads,
dim_feedforward=dim_feedforward,
dropout=dropout,
)
def forward(self, src: Tensor, tgt: Tensor) -> Tensor:
return self.decoder(tgt, self.encoder(src))
# Let’s create a simple test, as a sanity check for our implementation. We can construct random tensors for the input and target sequences, check that our model executes without errors, and confirm that the output tensor has the correct shape:
src = torch.rand(64, 32, 512)
tgt = torch.rand(64, 16, 512)
out = Transformer()(src, tgt)
print(out.shape)
# torch.Size([64, 16, 512])
# You could try implementing masked attention and training this Transformer model on a
# sequence-to-sequence problem. However, to understand masking, you might first find
# the [PyTorch Transformer tutorial](https://pytorch.org/tutorials/beginner/transformer_tutorial.html)
# useful. Note that this model is only a Transformer encoder for language modeling, but it uses
# masking in the encoder's self attention module.
# ## Vision Transformer (ViT)
#
# The Vision Transformer (ViT) is a transformer targeted at vision processing tasks. It has achieved state-of-the-art performance in image classification and (with some modification) other tasks. The ViT concept for image classification is as follows:
#
# <img src="img/vit.gif" title="ViT" />
#
# ### How does ViT work?
#
# The steps of ViT are as follows:
#
# 1. Split input image into patches
# 2. Flatten the patches
# 3. Produce linear embeddings from the flattened patches
# 4. Add position embeddings
# 5. Feed the sequence preceeded by a `[class]` token as input to a standard transformer encoder
# 6. Pretrain the model to ouptut image labels for the `[class]` token (fully supervised on a huge dataset such as ImageNet-22K)
# 7. Fine-tune on the downstream dataset for the specific image classification task
# ### ViT architecture
#
# ViT is a Transformer encoder. In detail, it looks like this:
#
# <img src="img/ViTArchitecture.png" title="ViT architecture" />
#
# In the figure we see four main parts:
# <ol style="list-style-type:lower-alpha">
# <li> The high-level architecture of the model.</li>
# <li> The Transformer module.</li>
# <li> The multiscale self-attention (MSA) head.</li>
# <li> An individual self-attention (SA) head.</li>
# </ol>
# ### Let's start
#
# Let's do a small scale implementation with the MNIST dataset. The
# code here is based on [<NAME>'s paper reimplementation repository](https://github.com/BrianPulfer/PapersReimplementations).
# +
import numpy as np
import torch
import torch.nn as nn
from torch.nn import CrossEntropyLoss
from torch.optim import Adam
from torch.utils.data import DataLoader
from torchvision.datasets.mnist import MNIST
from torchvision.transforms import ToTensor
# -
# Import the MNIST dataset:
# +
# Loading data
transform = ToTensor()
train_set = MNIST(root='./../datasets', train=True, download=True, transform=transform)
test_set = MNIST(root='./../datasets', train=False, download=True, transform=transform)
train_loader = DataLoader(train_set, shuffle=True, batch_size=16)
test_loader = DataLoader(test_set, shuffle=False, batch_size=16)
# -
# ### Train and test functions
#
# Next, let's create the train and test functions:
# +
def train_ViT_classify(model, optimizer, N_EPOCHS, train_loader, device="cpu"):
criterion = CrossEntropyLoss()
for epoch in range(N_EPOCHS):
train_loss = 0.0
for batch in train_loader:
x, y = batch
x = x.to(device)
y = y.to(device)
y_hat = model(x)
loss = criterion(y_hat, y) / len(x)
train_loss += loss.item()
optimizer.zero_grad()
loss.backward()
optimizer.step()
print(f"Epoch {epoch + 1}/{N_EPOCHS} loss: {train_loss:.2f}")
def test_ViT_classify(model, optimizer, test_loader):
criterion = CrossEntropyLoss()
correct, total = 0, 0
test_loss = 0.0
for batch in test_loader:
x, y = batch
x = x.to(device)
y = y.to(device)
y_hat = model(x)
loss = criterion(y_hat, y) / len(x)
test_loss += loss
correct += torch.sum(torch.argmax(y_hat, dim=1) == y).item()
total += len(x)
print(f"Test loss: {test_loss:.2f}")
print(f"Test accuracy: {correct / total * 100:.2f}%")
# -
# ### Multi-head Self Attention (MSA) Model
#
# As with the basic transformer above, to build the ViT model, we need to create a MSA module and put it
# together with the other elements.
#
# For a single image, self attention means that each patch's representation
# is updated based on its input token's similarity with those of the other patches.
# As before, we perform a linear mapping of each patch to three distinct vectors $q$, $k$, and $v$ (query, key, value).
#
# For each patch, we need to compute the dot product of its $q$ vector with all of the $k$ vectors, divide by the square root of the dimension
# of the vectors, then apply softmax to the result. The resulting matrix is called the matrix of attention cues.
# We multiply the attention cues with the $v$ vectors associated with the different input tokens and sum them all up.
#
# The input for each patch is transformed to a new value based on its similarity (after the linear mapping to $q$, $k$, and $v$) with other patches.
#
# However, the whole procedure is carried out $H$ times on $H$ sub-vectors of our current 8-dimensional patches, where $H$ is the number of heads.
#
# Once all results are obtained, they are concatenated together then passed through a linear layer.
#
# The MSA model looks like this:
class MSA(nn.Module):
def __init__(self, d, n_heads=2):
super(MSA, self).__init__()
self.d = d
self.n_heads = n_heads
assert d % n_heads == 0, f"Can't divide dimension {d} into {n_heads} heads"
d_head = int(d / n_heads)
self.q_mappings = [nn.Linear(d_head, d_head) for _ in range(self.n_heads)]
self.k_mappings = [nn.Linear(d_head, d_head) for _ in range(self.n_heads)]
self.v_mappings = [nn.Linear(d_head, d_head) for _ in range(self.n_heads)]
self.d_head = d_head
self.softmax = nn.Softmax(dim=-1)
def forward(self, sequences):
# Sequences has shape (N, seq_length, token_dim)
# We go into shape (N, seq_length, n_heads, token_dim / n_heads)
# And come back to (N, seq_length, item_dim) (through concatenation)
result = []
for sequence in sequences:
seq_result = []
for head in range(self.n_heads):
q_mapping = self.q_mappings[head]
k_mapping = self.k_mappings[head]
v_mapping = self.v_mappings[head]
seq = sequence[:, head * self.d_head: (head + 1) * self.d_head]
q, k, v = q_mapping(seq), k_mapping(seq), v_mapping(seq)
attention = self.softmax(q @ k.T / (self.d_head ** 0.5))
seq_result.append(attention @ v)
result.append(torch.hstack(seq_result))
return torch.cat([torch.unsqueeze(r, dim=0) for r in result])
# **Note**: for each head, we create distinct Q, K, and V mapping functions (square matrices of size 4x4 in our example).
#
# Since our inputs will be sequences of size (N, 50, 8), and we only use 2 heads, we will at some point have an (N, 50, 2, 4) tensor, use a nn.Linear(4, 4) module on it, and then come back, after concatenation, to an (N, 50, 8) tensor.
# ### Position encoding
#
# The position encoding allows the model to understand where each patch is in the original image. While it is theoretically possible to learn
# such positional embeddings, the original Vaswani et al. Transformer uses a fixed position embedding representation that adds
# low-frequency values to the first dimension and higher-frequency values to the later dimensions, resulting in a code that is
# more similar for nearby tokens than far away tokens. For each token, we add to its j-th coordinate the value
#
# $$ p_{i,j} =
# \left\{\begin{matrix}
# \sin (\frac{i}{10000^{j/d_{embdim}}})\\
# \cos (\frac{i}{10000^{j/d_{embdim}}})
# \end{matrix}\right.
# $$
#
# We can visualize the position encoding matrix thusly:
#
# <img src="img/peimages.png" title="" style="width: 800px;" />
#
# Here is an implementation:
def get_positional_embeddings(sequence_length, d, device="cpu"):
result = torch.ones(sequence_length, d)
for i in range(sequence_length):
for j in range(d):
result[i][j] = np.sin(i / (10000 ** (j / d))) if j % 2 == 0 else np.cos(i / (10000 ** ((j - 1) / d)))
return result.to(device)
# ### ViT Model
#
# Create the ViT model as below. The explaination is later.
class ViT(nn.Module):
def __init__(self, input_shape, n_patches=7, hidden_d=8, n_heads=2, out_d=10):
# Super constructor
super(ViT, self).__init__()
# Input and patches sizes
self.input_shape = input_shape
self.n_patches = n_patches
self.n_heads = n_heads
assert input_shape[1] % n_patches == 0, "Input shape not entirely divisible by number of patches"
assert input_shape[2] % n_patches == 0, "Input shape not entirely divisible by number of patches"
self.patch_size = (input_shape[1] / n_patches, input_shape[2] / n_patches)
self.hidden_d = hidden_d
# 1) Linear mapper
self.input_d = int(input_shape[0] * self.patch_size[0] * self.patch_size[1])
self.linear_mapper = nn.Linear(self.input_d, self.hidden_d)
# 2) Classification token
self.class_token = nn.Parameter(torch.rand(1, self.hidden_d))
# 3) Positional embedding
# (In forward method)
# 4a) Layer normalization 1
self.ln1 = nn.LayerNorm((self.n_patches ** 2 + 1, self.hidden_d))
# 4b) Multi-head Self Attention (MSA) and classification token
self.msa = MSA(self.hidden_d, n_heads)
# 5a) Layer normalization 2
self.ln2 = nn.LayerNorm((self.n_patches ** 2 + 1, self.hidden_d))
# 5b) Encoder MLP
self.enc_mlp = nn.Sequential(
nn.Linear(self.hidden_d, self.hidden_d),
nn.ReLU()
)
# 6) Classification MLP
self.mlp = nn.Sequential(
nn.Linear(self.hidden_d, out_d),
nn.Softmax(dim=-1)
)
def forward(self, images):
# Dividing images into patches
n, c, w, h = images.shape
patches = images.reshape(n, self.n_patches ** 2, self.input_d)
# Running linear layer for tokenization
tokens = self.linear_mapper(patches)
# Adding classification token to the tokens
tokens = torch.stack([torch.vstack((self.class_token, tokens[i])) for i in range(len(tokens))])
# Adding positional embedding
tokens += get_positional_embeddings(self.n_patches ** 2 + 1, self.hidden_d, device).repeat(n, 1, 1)
# TRANSFORMER ENCODER BEGINS ###################################
# NOTICE: MULTIPLE ENCODER BLOCKS CAN BE STACKED TOGETHER ######
# Running Layer Normalization, MSA and residual connection
self.msa(self.ln1(tokens.to("cpu")).to(device))
out = tokens + self.msa(self.ln1(tokens))
# Running Layer Normalization, MLP and residual connection
out = out + self.enc_mlp(self.ln2(out))
# TRANSFORMER ENCODER ENDS ###################################
# Getting the classification token only
out = out[:, 0]
return self.mlp(out)
# #### Step 1: Patchifying and the linear mapping
#
# The transformer encoder was developed with sequence data in mind, such as English sentences. However, an image is not a sequence. Thus, we break it into multiple sub-images and map each sub-image to a vector.
#
# We do so by simply reshaping our input, which has size $(N, C, H, W)$ (in our example $(N, 1, 28, 28)$), to size (N, #Patches, Patch dimensionality), where the dimensionality of a patch is adjusted accordingly.
#
# In MNIST, we break each $(1, 28, 28)$ into 7x7 patches (hence, each of size 4x4). That is, we are going to obtain 7x7=49 sub-images out of a single image.
#
# $$(N,1,28,28) \rightarrow (N,P\times P, H \times C/P \times W \times C/P) \rightarrow (N, 7\times 7, 4\times 4) \rightarrow (N, 49, 16)$$
#
# <img src="img/patch.png" title="an image is split into patches" />
# #### Step 2: Adding the classification token
#
# When information about all other tokens will be present here, we will be able to classify the image using only this special token. The initial value of the special token (the one fed to the transformer encoder) is a parameter of the model that needs to be learned.
#
# We can now add a parameter to our model and convert our (N, 49, 8) tokens tensor to an (N, 50, 8) tensor (we add the special token to each sequence).
#
# Passing from (N,49,8) → (N,50,8) is probably sub-optimal. Also, notice that the classification token is put as the first token of each sequence. This will be important to keep in mind when we will then retrieve the classification token to feed to the final MLP.
# #### Step 3: Positional encoding
#
# See above, as we mentioned.
#
# #### Step 4: LN, MSA, and Residual Connection
#
# The step is to apply layer normalization to the tokens, then apply MSA, and add a residual connection (add the input we had before applying LN).
# - **Layer normalization** is a popular block that, given an input, subtracts its mean and divides by the standard deviation.
# - **MSA**: same as the vanilla transformer.
# - **A residual connection** consists in just adding the original input to the result of some computation. This, intuitively, allows a network to become more powerful while also preserving the set of possible functions that the model can approximate.
#
# The residual connection is added at the original (N, 50, 8) tensor to the (N, 50, 8) obtained after LN and MSA.
# #### Step 5: LN, MLP, and Residual Connection
# All that is left to the transformer encoder is just a simple residual connection between what we already have and what we get after passing the current tensor through another LN and an MLP.
#
# #### Step 6: Classification MLP
# Finally, we can extract just the classification token (first token) out of our N sequences, and use each token to get N classifications.
# Since we decided that each token is an 8-dimensional vector, and since we have 10 possible digits, we can implement the classification MLP as a simple 8x10 matrix, activated with the SoftMax function.
#
# The output of our model is now an (N, 10) tensor.
# device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
# print('Using device', device)
# We haven't gotten CUDA too work yet -- the kernels always die!
device = "cpu"
# +
model = ViT((1, 28, 28), n_patches=7, hidden_d=20, n_heads=2, out_d=10)
model = model.to(device)
N_EPOCHS = 5
LR = 0.01
optimizer = Adam(model.parameters(), lr=LR)
# -
train_ViT_classify(model, optimizer, N_EPOCHS, train_loader, device)
test_ViT_classify(model, optimizer, test_loader)
# The testing accuracy is over 90%. Our implementation is done!
# ### Pytorch ViT
#
# [Here](https://github.com/lucidrains/vit-pytorch#vision-transformer---pytorch) is the link of the full version of ViT using pytorch.
# !pip install vit-pytorch
# ## ViT Pytorch implementation
# +
import torch
from vit_pytorch import ViT
v = ViT(
image_size = 256,
patch_size = 32,
num_classes = 1000,
dim = 1024,
depth = 6,
heads = 16,
mlp_dim = 2048,
dropout = 0.1,
emb_dropout = 0.1
)
img = torch.randn(1, 3, 256, 256)
preds = v(img) # (1, 1000)
# -
# The implementation also contains a distillable ViT:
# +
import torch
from torchvision.models import resnet50
from vit_pytorch.distill import DistillableViT, DistillWrapper
teacher = resnet50(pretrained = True)
v = DistillableViT(
image_size = 256,
patch_size = 32,
num_classes = 1000,
dim = 1024,
depth = 6,
heads = 8,
mlp_dim = 2048,
dropout = 0.1,
emb_dropout = 0.1
)
distiller = DistillWrapper(
student = v,
teacher = teacher,
temperature = 3, # temperature of distillation
alpha = 0.5, # trade between main loss and distillation loss
hard = False # whether to use soft or hard distillation
)
img = torch.randn(2, 3, 256, 256)
labels = torch.randint(0, 1000, (2,))
loss = distiller(img, labels)
loss.backward()
# after lots of training above ...
pred = v(img) # (2, 1000)
# -
# and so on...
# ## To do on your own
#
# If we can manage the resources, let's try pre-training a ViT on ImageNet 1K then fine tuning it on another dataset such as CIFAR-10.
#
| Labs/11-Transformers/11-Transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import syft
import syft.nn as nn
from syft.controller import tensors, models
import imp
imp.reload(syft.controller)
imp.reload(syft.nn)
imp.reload(syft)
import numpy as np
from syft import FloatTensor
# -
# create tensors
data = np.array([[-1,-2,3,4,5,-6]]).astype('float')
a = FloatTensor(data, autograd=True)
b = FloatTensor(data+5, autograd=True)
np.random.seed(1)
a_ = np.array([[1,2,3,4,5]]).astype('float')
b_ = np.array([[5,1,3,8,2],[2,5,3,2,5],[3,5,2,3,6]]).astype('float').transpose()
at = Variable(torch.FloatTensor(a_),requires_grad=True)
# bt = Variable(torch.FloatTensor(b_),requires_grad=True)
ct = at ** 2
ct.backward(torch.ones(1,5))
ct.data.numpy()
at.grad
a = FloatTensor(a_,autograd=True)
a.grad()
| notebooks/tests/PySyft Autograd Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:Georg_animal_feces-phyloseq]
# language: R
# name: conda-env-Georg_animal_feces-phyloseq-r
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Goal" data-toc-modified-id="Goal-1"><span class="toc-item-num">1 </span>Goal</a></span></li><li><span><a href="#Var" data-toc-modified-id="Var-2"><span class="toc-item-num">2 </span>Var</a></span></li><li><span><a href="#Init" data-toc-modified-id="Init-3"><span class="toc-item-num">3 </span>Init</a></span></li><li><span><a href="#Load" data-toc-modified-id="Load-4"><span class="toc-item-num">4 </span>Load</a></span><ul class="toc-item"><li><span><a href="#Checks" data-toc-modified-id="Checks-4.1"><span class="toc-item-num">4.1 </span>Checks</a></span></li></ul></li><li><span><a href="#Merging-&-pruning-tree" data-toc-modified-id="Merging-&-pruning-tree-5"><span class="toc-item-num">5 </span>Merging & pruning tree</a></span><ul class="toc-item"><li><span><a href="#Checks" data-toc-modified-id="Checks-5.1"><span class="toc-item-num">5.1 </span>Checks</a></span></li><li><span><a href="#Writing-tree" data-toc-modified-id="Writing-tree-5.2"><span class="toc-item-num">5.2 </span>Writing tree</a></span></li></ul></li><li><span><a href="#sessionInfo" data-toc-modified-id="sessionInfo-6"><span class="toc-item-num">6 </span>sessionInfo</a></span></li></ul></div>
# -
# # Goal
#
# * Create phylogeny for all genome reps used for the Struo2 database
# * merging & filtering GTDB MLSA phylogenies
# # Var
# +
work_dir = '/ebio/abt3_projects/databases_no-backup/GTDB/release95/Struo/phylogeny/'
# species-rep genomes selected
genomes_file = file.path(dirname(work_dir),'metadata_1per-GTDB-Spec_gte50comp-lt5cont_wtaxID_wPath.tsv')
# trees from gtdb
arc_tree_file = '/ebio/abt3_projects/databases_no-backup/GTDB/release95/phylogeny/ar122_r95.tree'
bac_tree_file = '/ebio/abt3_projects/databases_no-backup/GTDB/release95/phylogeny/bac120_r95.tree'
# full gtdb metadata
gtdb_meta_dir = '/ebio/abt3_projects/databases_no-backup/GTDB/release95/metadata/'
gtdb_meta_arc_file = file.path(gtdb_meta_dir, 'ar122_metadata_r95.tsv')
gtdb_meta_bac_file = file.path(gtdb_meta_dir, 'bac120_metadata_r95.tsv')
# -
# # Init
library(dplyr)
library(tidyr)
library(ggplot2)
library(data.table)
library(tidytable)
library(ape)
library(LeyLabRMisc)
df.dims()
# # Load
tax_levs = c('Domain', 'Phylum', 'Class', 'Order', 'Family', 'Genus', 'Species')
# genomes used for struo
genomes = Fread(genomes_file) %>%
select.(ncbi_organism_name, accession, gtdb_taxonomy) %>%
separate.(gtdb_taxonomy, tax_levs, sep = ';') %>%
mutate.(Species = gsub(' ', '_', Species))
genomes %>% unique_n('genomes', ncbi_organism_name)
genomes
# arc tree
arc_tree = read.tree(arc_tree_file)
arc_tree
# bac tree
bac_tree = read.tree(bac_tree_file)
bac_tree
# metadata: archaea
gtdb_meta_arc = Fread(gtdb_meta_arc_file) %>%
select.(accession, gtdb_taxonomy) %>%
filter.(accession %in% arc_tree$tip.label) %>%
separate.(gtdb_taxonomy, tax_levs, sep = ';')
gtdb_meta_arc
# metadata: bacteria
gtdb_meta_bac = Fread(gtdb_meta_bac_file) %>%
select.(accession, gtdb_taxonomy) %>%
filter.(accession %in% bac_tree$tip.label) %>%
separate.(gtdb_taxonomy, tax_levs, sep = ';')
gtdb_meta_bac
# combined
gtdb_meta = rbind(gtdb_meta_arc, gtdb_meta_bac)
gtdb_meta_arc = gtdb_meta_bac = NULL
gtdb_meta
# ## Checks
arc_tree$edge.length %>% summary
bac_tree$edge.length %>% summary
# # Merging & pruning tree
# binding trees at root
tree = ape::bind.tree(arc_tree, bac_tree)
tree
# renaming as species
idx = gtdb_meta %>%
filter.(accession %in% tree$tip.label) %>%
select.(accession, Species) %>%
mutate.(Species = gsub(' ', '_', Species)) %>%
as.data.frame
rownames(idx) = idx$accession
tree$tip.label = idx[tree$tip.label,'Species']
tree
# checking overlap
overlap(genomes$Species, tree$tip.label)
# purning
to_rm = setdiff(tree$tip.label, genomes$Species)
to_rm %>% length
tree_f = ape::drop.tip(tree, to_rm)
tree_f
# ## Checks
# branch lengths
tree_f$edge.length %>% summary
# checking overlap
overlap(genomes$Species, tree_f$tip.label)
# ## Writing tree
F = file.path(work_dir, 'ar122-bac120_r89_1per-GTDB-Spec_gte50comp-lt5cont.nwk')
write.tree(tree_f, F)
cat('File writen:', F, '\n')
# # sessionInfo
sessionInfo()
| notebooks/GTDB_release95/03_GTDBr95_db/01_phylogeny.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Defining function
# We define add function with two input a,b and later when we call add function it should print addition
# a : Parameter
# b : Parameter
# C : output
def add(a,b):
c = a + b
print(c)
# +
# Calling the previously defined function
add(4,5)
# -
def hollywood(actor,movie):
print (str(actor)+" acted in the movie "+ str(movie))
hollywood("<NAME>", "To Kill a Mocking Bird")
# +
# Here we define a function containing return statement which returns the results
# Return function breaks the code and returns the value back
def cube(integer):
result = integer * integer * integer
return result
# -
cube(9)
# +
# If statement
Marvel_movie = True
if Marvel_movie:
print("Watch Avengers")
# +
# If-else statement
Marvel_movie = False
if Marvel_movie:
print("Watch Avengers")
else:
print("Watch Aquaman")
# +
# If-else statement
Marvel_movie = True
DC_movie = True
if Marvel_movie and DC_movie:
print(" Watch Avengers and Aquaman")
else:
print("Watch Jurassic Park")
# +
# If-else-elif statement
# Change the boolean of Marvel_movie and DC_movie
Marvel_movie = True
DC_movie = False
if Marvel_movie and DC_movie:
print("Watch Avengers and Aquaman")
elif Marvel_movie and not(DC_movie):
print("Watch Doctor Strange")
elif not(Marvel_movie) and DC_movie:
print("Watch Justice League")
else:
print("Watch Jurassic Park")
# +
# Comparision operators
# Here "=","<",">","!=" are comparision operators
# Compare three integers
def max_num(int1,int2,int3):
if int1 >= int2 and int1 >= int3:
return int1
elif int2 >= int1 and int2 >= int3:
return int2
else:
return int3
# -
max_num(8,6,5)
| Python-Tutorial_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install dipy
# +
import numpy as np
import matplotlib.pyplot as plt
from dipy.tracking.local import LocalTracking, ThresholdTissueClassifier
from dipy.tracking.utils import random_seeds_from_mask
from dipy.reconst.dti import TensorModel
from dipy.reconst.csdeconv import (ConstrainedSphericalDeconvModel,
auto_response)
from dipy.direction import peaks_from_model
from dipy.data import fetch_stanford_hardi, read_stanford_hardi, get_sphere
from dipy.segment.mask import median_otsu
from dipy.viz import actor, window
from dipy.io.image import save_nifti
from nibabel.streamlines import save as save_trk
from nibabel.streamlines import Tractogram
from dipy.tracking.streamline import Streamlines
# -
from dipy.viz import *
from dipy.io.image import save_nifti
from nibabel.streamlines import save as save_trk
from nibabel.streamlines import Tractogram
from dipy.tracking.streamline import Streamlines
def show_slice(mr_img, mask = None):
slice_id = mr_img.shape[1] // 2
slice_img = mr_img[:, slice_id, :]
plt.figure(figsize=(16, 8))
plt.subplot(1, 2, 1)
plt.imshow(
slice_img.T, vmax=0.5 * np.max(slice_img),
origin='lower', cmap='gray', interpolation='nearest'
)
if mask is not None:
slice_mask = mask[:, slice_id]
plt.subplot(1, 2, 2)
plt.imshow(slice_mask.T, origin='lower', cmap='gray', interpolation='nearest')
plt.show()
# # Load sample dMRI data
# +
fetch_stanford_hardi()
img, gtab = read_stanford_hardi()
data = img.get_data()
show_slice(data[...,0])
# -
# # Model of the diffusion signal
# +
_, mask = median_otsu(data, 3, 1, False, vol_idx=range(10, 50), dilate=2)
response, _ = auto_response(gtab, data, roi_radius=10, fa_thr=0.7)
csd_model = ConstrainedSphericalDeconvModel(gtab, response)
csd_peaks = peaks_from_model(model=csd_model,
data=data,
sphere=get_sphere('symmetric724'),
mask=mask,
relative_peak_threshold=.5,
min_separation_angle=25,
parallel=True)
# -
# # Fractional anisotropy
# +
tensor_model = TensorModel(gtab, fit_method='WLS')
tensor_fit = tensor_model.fit(data, mask)
fa = tensor_fit.fa
show_slice(fa)
# -
# # White matter mask
# TASK: Find optimal threshold value for FA in order to obtain the most accurate white matter mask.
wm_mask = fa > 0.01
wm_mask1 = fa > 0.19
show_slice(data[...,0], wm_mask)
show_slice(wm_mask1, data[...,0]*wm_mask1)
# # Run deterministic tractography
# +
tissue_classifier = ThresholdTissueClassifier(fa, 0.1)
seeds = random_seeds_from_mask(wm_mask, seeds_count=1)
streamline_generator = LocalTracking(csd_peaks, tissue_classifier,
seeds, affine=np.eye(4),
step_size=0.5)
streamlines = Streamlines(streamline_generator)
# +
ren = window.Renderer()
ren.clear()
ren.add(actor.line(streamlines))
window.show(ren, size=(900, 900))
# -
show_slice(data[...,0])
| 1/deterministic_tractography.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# # Preparation steps before running this notebook
# 1. Run cwr_etl.ipynb first
# 2. Paste the IBM COS (Cloud Object Store) configuration below, either bei following the tutorial mentioned below or by taking the already existing config from cwr_etl.ipynb
# !pip install tensorflow==2.5.0
# +
# In order to obtain the correct values for "credentias", "bucket_name" and "endpoint"
# please follow the tutorial at https://github.com/IBM/skillsnetwork/wiki/Cloud-Object-Storage-Setup
credentials = {
# your credentials go here
}
bucket_name = # your bucket name goes here
endpoint = # your endpoint goes here
# +
import base64
from ibm_botocore.client import Config
import ibm_boto3
import time
# Create client
client = ibm_boto3.client(
's3',
aws_access_key_id = credentials["cos_hmac_keys"]['access_key_id'],
aws_secret_access_key = credentials["cos_hmac_keys"]["secret_access_key"],
endpoint_url=endpoint
)
client.download_file(bucket_name,'result_healthy_pandas.csv', 'result_healthy_pandas.csv')
client.download_file(bucket_name,'result_faulty_pandas.csv', 'result_faulty_pandas.csv')
# -
import pandas as pd
df_healthy = pd.read_csv('result_healthy_pandas.csv', engine='python', header=None)
df_healthy.head()
df_healthy.loc[df_healthy[1] == 100]
df_faulty = pd.read_csv('result_faulty_pandas.csv', engine='python', header=None)
df_faulty.head()
import numpy as np
from numpy import concatenate
from matplotlib import pyplot
from pandas import read_csv
from pandas import DataFrame
from pandas import concat
import sklearn
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_squared_error
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.callbacks import Callback
from keras.models import Sequential
from keras.layers import LSTM, Dense, Activation
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import time
# %matplotlib inline
def get_recording(df,file_id):
return np.array(df.sort_values(by=0, ascending=True).loc[df[1] == file_id].drop(0,1).drop(1,1))
import numpy as np
healthy_sample = get_recording(df_healthy,100)
faulty_sample = get_recording(df_faulty,105)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_sample)
ax.plot(range(0,size), healthy_sample[:,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,size), healthy_sample[:,1], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(faulty_sample)
ax.plot(range(0,size), faulty_sample[:,1], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,size), faulty_sample[:,0], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
ax.plot(range(0,500), healthy_sample[:500,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,500), healthy_sample[:500,1], '-', color='blue', animated = True, linewidth=1)
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
ax.plot(range(0,500), faulty_sample[:500,0], '-', color='red', animated = True, linewidth=1)
ax.plot(range(0,500), faulty_sample[:500,1], '-', color='blue', animated = True, linewidth=1)
class LossHistory(Callback):
def on_train_begin(self, logs={}):
self.losses = []
def on_batch_end(self, batch, logs={}):
self.losses.append(logs.get('loss'))
# +
timesteps = 100
dim = 2
lossHistory = LossHistory()
# design network
model = Sequential()
model.add(LSTM(50,input_shape=(timesteps,dim),return_sequences=True))
model.add(Dense(2))
model.compile(loss='mae', optimizer='adam')
def train(data):
model.fit(data, data, epochs=20, batch_size=72, validation_data=(data, data), verbose=1, shuffle=False,callbacks=[lossHistory])
def score(data):
yhat = model.predict(data)
return yhat
# +
#some learners constantly reported 502 errors in Watson Studio.
#This is due to the limited resources in the free tier and the heavy resource consumption of Keras.
#This is a workaround to limit resource consumption
import os
# reduce number of threads
os.environ['TF_NUM_INTEROP_THREADS'] = '1'
os.environ['TF_NUM_INTRAOP_THREADS'] = '1'
import tensorflow
# -
def create_trimmed_recording(df,file_id):
recording = get_recording(df,file_id)
samples = len(recording)
trim = samples % 100
recording_trimmed = recording[:samples-trim]
recording_trimmed.shape = (int((samples-trim)/timesteps),timesteps,dim)
return recording_trimmed
#pd.unique()
#df_healthy.drop(0,1).drop(2,1).drop(3,1)
pd.unique(df_healthy.iloc[:,1])
# +
file_ids = pd.unique(df_healthy.iloc[:,1])
start = time.time()
for file_id in file_ids:
recording_trimmed = create_trimmed_recording(df_healthy,file_id)
print("Staring training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id,time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
healthy_losses = lossHistory.losses
# -
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_losses)
plt.ylim(0,0.008)
ax.plot(range(0,size), healthy_losses, '-', color='blue', animated = True, linewidth=1)
# +
#file_ids = spark.sql('select distinct _c1 from df_healhty').rdd.map(lambda row : row._c1).collect()
start = time.time()
for file_id in [105]:
recording_trimmed = create_trimmed_recording(df_faulty,file_id)
print("Staring training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id,time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
faulty_losses = lossHistory.losses
# +
file_ids = pd.unique(df_faulty.iloc[:,1])
start = time.time()
for file_id in file_ids:
recording_trimmed = create_trimmed_recording(df_faulty,file_id)
print("Staring training on %s" % (file_id))
train(recording_trimmed)
print("Finished training on %s after %s seconds" % (file_id,time.time()-start))
print("Finished job on after %s seconds" % (time.time()-start))
faulty_losses = lossHistory.losses
# -
fig, ax = plt.subplots(num=None, figsize=(14, 6), dpi=80, facecolor='w', edgecolor='k')
size = len(healthy_losses+faulty_losses)
plt.ylim(0,0.008)
ax.plot(range(0,size), healthy_losses+faulty_losses, '-', color='blue', animated = True, linewidth=1)
| coursera_ai/week3/cwr_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="PKdCkYCAGsvw"
import numpy as np
import pandas as pd
import io
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 221} id="6kVmJ_9THjaZ" outputId="011388f8-9d76-4e26-a056-95c325b8ba41"
from google.colab import drive
drive.mount('/content/drive')
data = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/ML/Lab6/BuyComputer.csv')
data.drop(columns=['User ID',], axis = 1, inplace = True)
data.head()
# + id="9yhzb68c7rNN"
#Declare label as last column in the source file
label = data.iloc[:,-1].values
# print(label)
#Declare X as all columns excluding last
x = data.iloc[:,:-1].values
# print('\n', x)
# + id="cIDtGMDhIP0W"
#splitting data
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, label, test_size = 0.40, random_state = 90)
# print(x_train, '\n', x_test, '\n\n', y_train, '\n', y_test)
# + id="ChWM7Bd7IbPA"
#scaling data
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
x_train = sc.fit_transform(x_train)
x_test = sc.transform(x_test)
# + colab={"base_uri": "https://localhost:8080/"} id="DuhNMM3g8kgU" outputId="7cee3cd8-4321-4a5d-80e8-7ded32e3e2a0"
#variables to calculate sigmoid function
y_pred = []
x_length = len(x_train[0])
w = []
b = 0.2
print(x_length)
# + colab={"base_uri": "https://localhost:8080/"} id="jM4Yn0cD80oa" outputId="0dadcbfc-6059-4fb3-8af9-360e1ebef458"
entries = len(x_train[:, 0])
print(entries)
# + colab={"base_uri": "https://localhost:8080/"} id="43mVmUdG9LqA" outputId="81bf093a-e9df-4838-84b6-8000fa1cacf0"
for weight in range(x_length):
w.append(0)
w
# + id="-TfrMka89VN2"
#sigmoid function
def sigmoid(z):
return 1 / (1 + np.exp(-z))
# + id="034AnZWc9dNO"
def predict(inputs):
z = np.dot(w, inputs) + b
temp = sigmoid(z)
return temp
# + id="jOwYZs_k9kmK"
#Loss function
def loss_func(y, a):
J = -(y * np.log(a) + (1-y) * np.log(1-a))
return J
# + id="G79fmcF69wyM"
dw = []
db = 0
J = 0
alpha = 0.1
for x in range(x_length):
dw.append(0)
# + id="F7q9BF1_90qL"
#Repeating the process 3000 times
for iter in range(3000):
for i in range(entries):
local_x = x_train[i]
a = predict(local_x)
dz = a - y_train[i]
J += loss_func(y_train[i],a)
for j in range(x_length):
dw[j] = dw[j] + (local_x[j] * dz)
db += dz
J = J / entries
db = db / entries
for x in range(x_length):
dw[x] = dw[x] / entries
for x in range(x_length):
w[x] = w[x] - (alpha * dw[x])
b = b - (alpha * db)
J=0
# + colab={"base_uri": "https://localhost:8080/"} id="VSgiu1jR98Dj" outputId="8ab4af1a-fc77-4aab-c218-a426d94e5d04"
#Print weight
print(w)
#print bias
print('\n', b)
# + id="dN6DHqpp_W_9"
#predicting the label
for x in range(len(y_test)):
y_pred.append(predict(x_test[x]))
# + colab={"base_uri": "https://localhost:8080/"} id="xJ4uOkGBbsQ0" outputId="b9ce0836-d342-4169-a38a-11fab21e8b37"
#print actual and predicted values in a table
print("Actual\t\tPredicted")
for x in range(len(y_pred)):
print(y_test[x] ,y_pred[x], sep="\t\t")
if(y_pred[x] >= 0.5):
y_pred[x] = 1
else:
y_pred[x] = 0
# + colab={"base_uri": "https://localhost:8080/"} id="PPSUOwaQ_wHJ" outputId="ce29b13f-64eb-43b3-ade8-81a4eef6acfb"
# Calculating accuracy of prediction
cnt = 0
for x in range(len(y_pred)):
if(y_pred[x] == y_test[x]):
cnt += 1
print('Accuracy : ', (cnt / (len(y_pred))) * 100)
# + [markdown] id="x6nmajpzhAEn"
# #Using sklearn LogisticRegression model
# + colab={"base_uri": "https://localhost:8080/"} id="SklvAV_wcrL3" outputId="eb4217fe-5ac0-4717-ec9b-2858b060a32f"
# Fitting Logistic Regression to the Training set
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression(random_state = 90)
#Fit
lr.fit(x_train, y_train)
#predicting the test label with lr. Predict always takes X as input
y_predict = lr.predict(x_test)
from sklearn.metrics import accuracy_score
print('Accuracy : ', accuracy_score(y_predict, y_test))
| Lab6/090_Lab6_LogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis - Video Game Sales - <NAME> - 12/16/2019
import pandas as pd
import numpy as np
df = pd.read_csv('vgsales.csv')
df
df[['Publisher']].mode()
# the most common video game publisher is electronic arts
df[['Platform']].mode()
# the most common platform is the DS
df[['Genre']].mode()
# The most common Genre is Action
df.sort_values('Global_Sales', ascending= False)['Name'].head(21)
# The top twenty games are ^ (dont want to type out 20 games)
df[['NA_Sales']].median()
# the median North American sales is .08
around_median = df['NA_Sales'].between(0.8, 0.8,)
df[around_median]
# Values around the median North American Sales ^
gsmean = df[['NA_Sales']].mean()
other = df.loc[:, 'NA_Sales'].std()
other-gsmean
# the top selling NA game is .55 above the mean
nintendo_sales = df[df['Publisher'].between('Nintendo', 'Nintendo')].mean()
other_sales = df[df['Publisher'].str.contains('Nintendo') == False].mean()
nintendo_sales.Global_Sales-other_sales.Global_Sales
# Nintendo's mean sales is 2.09 above all other platforms
# 1st question: What are the top 5 worst selling Games?
df.sort_values('Global_Sales', ascending= False)['Name'].tail(5)
# question 2: what genre is the least popular?
df.sort_values('Genre', ascending= False)['Genre'].head(1)
# question 3: what are the top 5 grossing games in Japan
df.sort_values('JP_Sales', ascending= False)['Name'].head(5)
| vg-stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/JBNU-2021/blob/main/Preprocessing/classification_feature.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="basic-nature"
# ### Classification Feature Selection:
# (Numerical Input, Categorical Output)
# + [markdown] id="recorded-gregory"
# Feature selection is performed using ANOVA F measure via the f_classif() function.
# + id="chemical-minute"
#@title 기본 제목 텍스트
# ANOVA feature selection for numeric input and categorical output
from sklearn.datasets import make_classification
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_classif
# + colab={"base_uri": "https://localhost:8080/"} id="t_z26i8-YQ4N" outputId="ddb07e6b-6b4c-47da-dc1e-ed566aaaf4c9"
# generate dataset
X, y = make_classification(n_samples=100, n_features=20, n_informative=2)
# define feature selection
fs = SelectKBest(score_func=f_classif, k=2)
# apply feature selection
X_selected = fs.fit_transform(X, y)
print(X_selected.shape)
# + id="angry-angola"
| Preprocessing/classification_feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Creating a Segmentation App with MONAI Deploy App SDK
#
# This tutorial shows how to create an organ segmentation application for a PyTorch model that has been trained with MONAI.
#
# Deploying AI models requires the integration with clinical imaging network, even if in a for-research-use setting. This means that the AI deploy application will need to support standards-based imaging protocols, and specifically for Radiological imaging, DICOM protocol.
#
# Typically, DICOM network communication, either in DICOM TCP/IP network protocol or DICOMWeb, would be handled by DICOM devices or services, e.g. MONAI Deploy Informatics Gateway, so the deploy application itself would only need to use DICOM Part 10 files as input and save the AI result in DICOM Part10 file(s). For segmentation use cases, the DICOM instance file could be a DICOM Segmentation object or a DICOM RT Structure Set, and for classification, DICOM Structure Report and/or DICOM Encapsulated PDF.
#
# During model training, input and label images are typically in non-DICOM volumetric image format, e.g., NIfTI and PNG, converted from a specific DICOM study series. Furthermore, the voxel spacings most likely have been re-sampled to be uniform for all images. When integrated with imaging networks and receiving DICOM instances from modalities and Picture Archiving and Communications System, PACS, an AI deploy application may have to deal with a whole DICOM study with multiple series, whose images' spacing may not be the same as expected by the trained model. To address these cases consistently and efficiently, MONAI Deploy Application SDK provides classes, called operators, to parse DICOM studies, select specific series with application-defined rules, and convert the selected DICOM series into domain-specific image format along with meta-data representing the pertinent DICOM attributes.
#
# In the following sections, we will demonstrate how to create a MONAI Deploy application package using the MONAI Deploy App SDK.
#
# :::{note}
# For local testing, if there is a lack of DICOM Part 10 files, one can use open source programs, e.g. 3D Slicer, to convert NIfTI to DICOM files.
#
# :::
#
# ## Creating Operators and connecting them in Application class
#
# We will implement an application that consists of five Operators:
#
# - **DICOMDataLoaderOperator**:
# - **Input(dicom_files)**: a folder path ([`DataPath`](/modules/_autosummary/monai.deploy.core.domain.DataPath))
# - **Output(dicom_study_list)**: a list of DICOM studies in memory (List[[`DICOMStudy`](/modules/_autosummary/monai.deploy.core.domain.DICOMStudy)])
# - **DICOMSeriesSelectorOperator**:
# - **Input(dicom_study_list)**: a list of DICOM studies in memory (List[[`DICOMStudy`](/modules/_autosummary/monai.deploy.core.domain.DICOMStudy)])
# - **Input(selection_rules)**: a selection rule (Dict)
# - **Output(study_selected_series_list)**: a DICOM series object in memory ([`StudySelectedSeries`](/modules/_autosummary/monai.deploy.core.domain.StudySelectedSeries))
# - **DICOMSeriesToVolumeOperator**:
# - **Input(study_selected_series_list)**: a DICOM series object in memory ([`StudySelectedSeries`](/modules/_autosummary/monai.deploy.core.domain.StudySelectedSeries))
# - **Output(image)**: an image object in memory ([`Image`](/modules/_autosummary/monai.deploy.core.domain.Image))
# - **SpleenSegOperator**:
# - **Input(image)**: an image object in memory ([`Image`](/modules/_autosummary/monai.deploy.core.domain.Image))
# - **Output(seg_image)**: an image object in memory ([`Image`](/modules/_autosummary/monai.deploy.core.domain.Image))
# - **DICOMSegmentationWriterOperator**:
# - **Input(seg_image)**: a segmentation image object in memory ([`Image`](/modules/_autosummary/monai.deploy.core.domain.Image))
# - **Input(study_selected_series_list)**: a DICOM series object in memory ([`StudySelectedSeries`](/modules/_autosummary/monai.deploy.core.domain.StudySelectedSeries))
# - **Output(dicom_seg_instance)**: a file path ([`DataPath`](/modules/_autosummary/monai.deploy.core.domain.DataPath))
#
#
# :::{note}
# The `DICOMSegmentationWriterOperator` needs both the segmentation image as well as the original DICOM series meta-data in order to use the patient demographics and the DICOM Study level attributes.
# :::
#
# The workflow of the application would look like this.
#
# ```{mermaid}
# %%{init: {"theme": "base", "themeVariables": { "fontSize": "16px"}} }%%
#
# classDiagram
# direction TB
#
# DICOMDataLoaderOperator --|> DICOMSeriesSelectorOperator : dicom_study_list...dicom_study_list
# DICOMSeriesSelectorOperator --|> DICOMSeriesToVolumeOperator : study_selected_series_list...study_selected_series_list
# DICOMSeriesToVolumeOperator --|> SpleenSegOperator : image...image
# DICOMSeriesSelectorOperator --|> DICOMSegmentationWriterOperator : study_selected_series_list...study_selected_series_list
# SpleenSegOperator --|> DICOMSegmentationWriterOperator : seg_image...seg_image
#
#
# class DICOMDataLoaderOperator {
# <in>dicom_files : DISK
# dicom_study_list(out) IN_MEMORY
# }
# class DICOMSeriesSelectorOperator {
# <in>dicom_study_list : IN_MEMORY
# <in>selection_rules : IN_MEMORY
# study_selected_series_list(out) IN_MEMORY
# }
# class DICOMSeriesToVolumeOperator {
# <in>study_selected_series_list : IN_MEMORY
# image(out) IN_MEMORY
# }
# class SpleenSegOperator {
# <in>image : IN_MEMORY
# seg_image(out) IN_MEMORY
# }
# class DICOMSegmentationWriterOperator {
# <in>seg_image : IN_MEMORY
# <in>study_selected_series_list : IN_MEMORY
# dicom_seg_instance(out) DISK
# }
# ```
#
# ### Setup environment
#
# +
# Install MONAI and other necessary image processing packages for the application
# !python -c "import monai" || pip install -q "monai"
# !python -c "import torch" || pip install -q "torch>=1.5"
# !python -c "import numpy" || pip install -q "numpy>=1.20"
# !python -c "import nibabel" || pip install -q "nibabel>=3.2.1"
# !python -c "import pydicom" || pip install -q "pydicom>=1.4.2"
# !python -c "import SimpleITK" || pip install -q "SimpleITK>=2.0.0"
# !python -c "import typeguard" || pip install -q "typeguard>=2.12.1"
# Install MONAI Deploy App SDK package
# !python -c "import monai.deploy" || pip install --upgrade -q "monai-deploy-app-sdk"
# Install Clara Viz package
# !python -c "import clara.viz" || pip install --upgrade -q "clara-viz"
# -
# ### Download/Extract ai_spleen_seg_data from Google Drive
# +
# Download ai_spleen_seg_data test data zip file
# !pip install gdown
# !gdown https://drive.google.com/uc?id=1GC_N8YQk_mOWN02oOzAU_2YDmNRWk--n
# After downloading ai_spleen_seg_data zip file from the web browser or using gdown,
# !unzip -o "ai_spleen_seg_data_updated_1203.zip"
# -
# ### Setup imports
#
# Let's import necessary classes/decorators to define Application and Operator.
# +
import logging
from os import path
from numpy import uint8
import monai.deploy.core as md
from monai.deploy.core import ExecutionContext, Image, InputContext, IOType, Operator, OutputContext
from monai.deploy.operators.monai_seg_inference_operator import InMemImageReader, MonaiSegInferenceOperator
from monai.transforms import (
Activationsd,
AsDiscreted,
Compose,
CropForegroundd,
EnsureChannelFirstd,
Invertd,
LoadImaged,
SaveImaged,
ScaleIntensityRanged,
Spacingd,
ToTensord,
)
from monai.deploy.core import Application, resource
from monai.deploy.operators.dicom_data_loader_operator import DICOMDataLoaderOperator
from monai.deploy.operators.dicom_seg_writer_operator import DICOMSegmentationWriterOperator
from monai.deploy.operators.dicom_series_selector_operator import DICOMSeriesSelectorOperator
from monai.deploy.operators.dicom_series_to_volume_operator import DICOMSeriesToVolumeOperator
from monai.deploy.operators.clara_viz_operator import ClaraVizOperator
# -
# ### Creating Model Specific Inference Operator classes
#
# Each Operator class inherits [Operator](/modules/_autosummary/monai.deploy.core.Operator) class and input/output properties are specified by using [@input](/modules/_autosummary/monai.deploy.core.input)/[@output](/modules/_autosummary/monai.deploy.core.output) decorators.
#
# Business logic would be implemented in the <a href="../../modules/_autosummary/monai.deploy.core.Operator.html#monai.deploy.core.Operator.compute">compute()</a> method.
#
# The App SDK provides a `MonaiSegInferenceOperator` class to perform segmentation prediction with a Torch Script model. For consistency, this class uses MONAI dictionary-based transforms, as `Compose` object, for pre and post transforms. The model-specific inference operator will then only need to create the pre and post transform `Compose` based on what has been used in the model training and validation. Note that for deploy application, `ignite` is not needed nor supported.
#
# #### SpleenSegOperator
#
# The `SpleenSegOperator` gets as input an in-memory [Image](/modules/_autosummary/monai.deploy.core.domain.Image) object that has been converted from a DICOM CT series by the preceding `DICOMSeriesToVolumeOperator`, and as output in-memory segmentation [Image](/modules/_autosummary/monai.deploy.core.domain.Image) object.
#
# The `pre_process` function creates the pre-transforms `Compose` object. For `LoadImage`, a specialized `InMemImageReader`, derived from MONAI `ImageReader`, is used to convert the in-memory pixel data and return the `numpy` array as well as the meta-data. Also, the DICOM input pixel spacings are often not the same as expected by the model, so the `Spacingd` transform must be used to re-sample the image with the expected spacing.
#
# The `post_process` function creates the post-transform `Compose` object. The `SaveImageD` transform class is used to save the segmentation mask as NIfTI image file, which is optional as the in-memory mask image will be passed down to the DICOM Segmentation writer for creating a DICOM Segmentation instance. The `Invertd` must also be used to revert the segmentation image's orientation and spacing to be the same as the input.
#
# When the `MonaiSegInferenceOperator` object is created, the `ROI` size is specified, as well as the transform `Compose` objects. Furthermore, the dataset image key names are set accordingly.
#
# Loading of the model and performing the prediction are encapsulated in the `MonaiSegInferenceOperator` and other SDK classes. Once the inference is completed, the segmentation [Image](/modules/_autosummary/monai.deploy.core.domain.Image) object is created and set to the output (<a href="../../modules/_autosummary/monai.deploy.core.OutputContext.html#monai.deploy.core.OutputContext.set">op_output.set(value, label)</a>), by the `MonaiSegInferenceOperator`.
@md.input("image", Image, IOType.IN_MEMORY)
@md.output("seg_image", Image, IOType.IN_MEMORY)
@md.env(pip_packages=["monai==0.6.0", "torch>=1.5", "numpy>=1.20", "nibabel"])
class SpleenSegOperator(Operator):
"""Performs Spleen segmentation with a 3D image converted from a DICOM CT series.
"""
def __init__(self):
self.logger = logging.getLogger("{}.{}".format(__name__, type(self).__name__))
super().__init__()
self._input_dataset_key = "image"
self._pred_dataset_key = "pred"
def compute(self, op_input: InputContext, op_output: OutputContext, context: ExecutionContext):
input_image = op_input.get("image")
if not input_image:
raise ValueError("Input image is not found.")
output_path = context.output.get().path
# This operator gets an in-memory Image object, so a specialized ImageReader is needed.
_reader = InMemImageReader(input_image)
pre_transforms = self.pre_process(_reader)
post_transforms = self.post_process(pre_transforms, path.join(output_path, "prediction_output"))
# Delegates inference and saving output to the built-in operator.
infer_operator = MonaiSegInferenceOperator(
(
160,
160,
160,
),
pre_transforms,
post_transforms,
)
# Setting the keys used in the dictironary based transforms may change.
infer_operator.input_dataset_key = self._input_dataset_key
infer_operator.pred_dataset_key = self._pred_dataset_key
# Now let the built-in operator handles the work with the I/O spec and execution context.
infer_operator.compute(op_input, op_output, context)
def pre_process(self, img_reader) -> Compose:
"""Composes transforms for preprocessing input before predicting on a model."""
my_key = self._input_dataset_key
return Compose(
[
LoadImaged(keys=my_key, reader=img_reader),
EnsureChannelFirstd(keys=my_key),
Spacingd(keys=my_key, pixdim=[1.0, 1.0, 1.0], mode=["blinear"], align_corners=True),
ScaleIntensityRanged(keys=my_key, a_min=-57, a_max=164, b_min=0.0, b_max=1.0, clip=True),
CropForegroundd(keys=my_key, source_key=my_key),
ToTensord(keys=my_key),
]
)
def post_process(self, pre_transforms: Compose, out_dir: str = "./prediction_output") -> Compose:
"""Composes transforms for postprocessing the prediction results."""
pred_key = self._pred_dataset_key
return Compose(
[
Activationsd(keys=pred_key, softmax=True),
AsDiscreted(keys=pred_key, argmax=True),
Invertd(
keys=pred_key, transform=pre_transforms, orig_keys=self._input_dataset_key, nearest_interp=True
),
SaveImaged(keys=pred_key, output_dir=out_dir, output_postfix="seg", output_dtype=uint8, resample=False),
]
)
# ### Creating Application class
#
# Our application class would look like below.
#
# It defines `App` class, inheriting [Application](/modules/_autosummary/monai.deploy.core.Application) class.
#
# The requirements (resource and package dependency) for the App can be specified by using [@resource](/modules/_autosummary/monai.deploy.core.resource) and [@env](/modules/_autosummary/monai.deploy.core.env) decorators.
#
# The base class method, `compose`, is overridden. Objects required for DICOM parsing, series selection (selecting the first series for the current release), pixel data conversion to volume image, and segmentation instance creation are created, so is the model-specific `SpleenSegOperator`. The execution pipeline, as a Directed Acyclic Graph, is created by connecting these objects through <a href="../../modules/_autosummary/monai.deploy.core.Application.html#monai.deploy.core.Application.add_flow">self.add_flow()</a>.
@resource(cpu=1, gpu=1, memory="7Gi")
class AISpleenSegApp(Application):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
def compose(self):
study_loader_op = DICOMDataLoaderOperator()
series_selector_op = DICOMSeriesSelectorOperator()
series_to_vol_op = DICOMSeriesToVolumeOperator()
# Creates DICOM Seg writer with segment label name in a string list
dicom_seg_writer = DICOMSegmentationWriterOperator(seg_labels=["Spleen"])
# Creates the model specific segmentation operator
spleen_seg_op = SpleenSegOperator()
# Creates the DAG by linking the operators
self.add_flow(study_loader_op, series_selector_op, {"dicom_study_list": "dicom_study_list"})
self.add_flow(series_selector_op, series_to_vol_op, {"study_selected_series_list": "study_selected_series_list"})
self.add_flow(series_to_vol_op, spleen_seg_op, {"image": "image"})
self.add_flow(series_selector_op, dicom_seg_writer, {"study_selected_series_list": "study_selected_series_list"})
self.add_flow(spleen_seg_op, dicom_seg_writer, {"seg_image": "seg_image"})
viz_op = ClaraVizOperator()
self.add_flow(series_to_vol_op, viz_op, {"image": "image"})
self.add_flow(spleen_seg_op, viz_op, {"seg_image": "seg_image"})
# ## Executing app locally
#
# We can execute the app in the Jupyter notebook. Note that the DICOM files of the CT Abdomen series must be present in the `dcm` and the Torch Script model at `model.ts`. Please use the actual path in your environment.
#
# +
app = AISpleenSegApp()
app.run(input="dcm", output="output", model="model.ts")
# -
# Once the application is verified inside Jupyter notebook, we can write the above Python code into Python files in an application folder.
#
# The application folder structure would look like below:
#
# ```bash
# my_app
# ├── __main__.py
# ├── app.py
# └── spleen_seg_operator.py
# ```
#
# :::{note}
# We can create a single application Python file (such as `spleen_app.py`) that includes the content of the files, instead of creating multiple files.
# You will see such an example in <a href="./02_mednist_app.html#executing-app-locally">MedNist Classifier Tutorial</a>.
# :::
# Create an application folder
# !mkdir -p my_app
# ### spleen_seg_operator.py
# +
# %%writefile my_app/spleen_seg_operator.py
import logging
from os import path
from numpy import uint8
import monai.deploy.core as md
from monai.deploy.core import ExecutionContext, Image, InputContext, IOType, Operator, OutputContext
from monai.deploy.operators.monai_seg_inference_operator import InMemImageReader, MonaiSegInferenceOperator
from monai.transforms import (
Activationsd,
AsDiscreted,
Compose,
CropForegroundd,
EnsureChannelFirstd,
Invertd,
LoadImaged,
SaveImaged,
ScaleIntensityRanged,
Spacingd,
ToTensord,
)
@md.input("image", Image, IOType.IN_MEMORY)
@md.output("seg_image", Image, IOType.IN_MEMORY)
@md.env(pip_packages=["monai==0.6.0", "torch>=1.5", "numpy>=1.20", "nibabel", "typeguard"])
class SpleenSegOperator(Operator):
"""Performs Spleen segmentation with a 3D image converted from a DICOM CT series.
"""
def __init__(self):
self.logger = logging.getLogger("{}.{}".format(__name__, type(self).__name__))
super().__init__()
self._input_dataset_key = "image"
self._pred_dataset_key = "pred"
def compute(self, op_input: InputContext, op_output: OutputContext, context: ExecutionContext):
input_image = op_input.get("image")
if not input_image:
raise ValueError("Input image is not found.")
output_path = context.output.get().path
# This operator gets an in-memory Image object, so a specialized ImageReader is needed.
_reader = InMemImageReader(input_image)
pre_transforms = self.pre_process(_reader)
post_transforms = self.post_process(pre_transforms, path.join(output_path, "prediction_output"))
# Delegates inference and saving output to the built-in operator.
infer_operator = MonaiSegInferenceOperator(
(
160,
160,
160,
),
pre_transforms,
post_transforms,
)
# Setting the keys used in the dictironary based transforms may change.
infer_operator.input_dataset_key = self._input_dataset_key
infer_operator.pred_dataset_key = self._pred_dataset_key
# Now let the built-in operator handles the work with the I/O spec and execution context.
infer_operator.compute(op_input, op_output, context)
def pre_process(self, img_reader) -> Compose:
"""Composes transforms for preprocessing input before predicting on a model."""
my_key = self._input_dataset_key
return Compose(
[
LoadImaged(keys=my_key, reader=img_reader),
EnsureChannelFirstd(keys=my_key),
Spacingd(keys=my_key, pixdim=[1.0, 1.0, 1.0], mode=["blinear"], align_corners=True),
ScaleIntensityRanged(keys=my_key, a_min=-57, a_max=164, b_min=0.0, b_max=1.0, clip=True),
CropForegroundd(keys=my_key, source_key=my_key),
ToTensord(keys=my_key),
]
)
def post_process(self, pre_transforms: Compose, out_dir: str = "./prediction_output") -> Compose:
"""Composes transforms for postprocessing the prediction results."""
pred_key = self._pred_dataset_key
return Compose(
[
Activationsd(keys=pred_key, softmax=True),
AsDiscreted(keys=pred_key, argmax=True),
Invertd(
keys=pred_key, transform=pre_transforms, orig_keys=self._input_dataset_key, nearest_interp=True
),
SaveImaged(keys=pred_key, output_dir=out_dir, output_postfix="seg", output_dtype=uint8, resample=False),
]
)
# -
# ### app.py
# +
# %%writefile my_app/app.py
import logging
from spleen_seg_operator import SpleenSegOperator
from monai.deploy.core import Application, resource
from monai.deploy.operators.dicom_data_loader_operator import DICOMDataLoaderOperator
from monai.deploy.operators.dicom_seg_writer_operator import DICOMSegmentationWriterOperator
from monai.deploy.operators.dicom_series_selector_operator import DICOMSeriesSelectorOperator
from monai.deploy.operators.dicom_series_to_volume_operator import DICOMSeriesToVolumeOperator
@resource(cpu=1, gpu=1, memory="7Gi")
class AISpleenSegApp(Application):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
def compose(self):
study_loader_op = DICOMDataLoaderOperator()
series_selector_op = DICOMSeriesSelectorOperator(Sample_Rules_Text)
series_to_vol_op = DICOMSeriesToVolumeOperator()
# Creates DICOM Seg writer with segment label name in a string list
dicom_seg_writer = DICOMSegmentationWriterOperator(seg_labels=["Spleen"])
# Creates the model specific segmentation operator
spleen_seg_op = SpleenSegOperator()
# Creates the DAG by link the operators
self.add_flow(study_loader_op, series_selector_op, {"dicom_study_list": "dicom_study_list"})
self.add_flow(series_selector_op, series_to_vol_op, {"study_selected_series_list": "study_selected_series_list"})
self.add_flow(series_to_vol_op, spleen_seg_op, {"image": "image"})
self.add_flow(series_selector_op, dicom_seg_writer, {"study_selected_series_list": "study_selected_series_list"})
self.add_flow(spleen_seg_op, dicom_seg_writer, {"seg_image": "seg_image"})
# This is a sample series selection rule in JSON, simply selecting CT series.
# If the study has more than 1 CT series, then all of them will be selected.
# Please see more detail in DICOMSeriesSelectorOperator.
Sample_Rules_Text = """
{
"selections": [
{
"name": "CT Series",
"conditions": {
"StudyDescription": "(.*?)",
"Modality": "(?i)CT",
"SeriesDescription": "(.*?)"
}
}
]
}
"""
if __name__ == "__main__":
# Creates the app and test it standalone. When running is this mode, please note the following:
# -i <DICOM folder>, for input DICOM CT series folder
# -o <output folder>, for the output folder, default $PWD/output
# -m <model file>, for model file path
# e.g.
# python3 app.py -i input -m model.ts
#
AISpleenSegApp(do_run=True)
# -
# ```python
# if __name__ == "__main__":
# AISpleenSegApp(do_run=True)
# ```
#
# The above lines are needed to execute the application code by using `python` interpreter.
#
# ### \_\_main\_\_.py
#
# \_\_main\_\_.py is needed for <a href="../../developing_with_sdk/packaging_app.html#required-arguments">MONAI Application Packager</a> to detect the main application code (`app.py`) when the application is executed with the application folder path (e.g., `python simple_imaging_app`).
# +
# %%writefile my_app/__main__.py
from app import AISpleenSegApp
if __name__ == "__main__":
AISpleenSegApp(do_run=True)
# -
# !ls my_app
# In this time, let's execute the app in the command line.
# !python my_app -i dcm -o output -m model.ts
# Above command is same with the following command line:
import os
os.environ['MKL_THREADING_LAYER'] = 'GNU'
# !monai-deploy exec my_app -i dcm -o output -m model.ts
# !ls output
# ## Packaging app
# Let's package the app with [MONAI Application Packager](/developing_with_sdk/packaging_app).
# !monai-deploy package -b nvcr.io/nvidia/pytorch:21.11-py3 my_app --tag my_app:latest -m model.ts
# :::{note}
# Building a MONAI Application Package (Docker image) can take time. Use `-l DEBUG` option if you want to see the progress.
# :::
#
# We can see that the Docker image is created.
# !docker image ls | grep my_app
# ## Executing packaged app locally
#
# The packaged app can be run locally through [MONAI Application Runner](/developing_with_sdk/executing_packaged_app_locally).
# +
# Copy DICOM files are in 'dcm' folder
# Launch the app
# !monai-deploy run my_app:latest dcm output
# -
# !ls output
| notebooks/tutorials/03_segmentation_viz_app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# @Date : 2018-04-16 18:24:00
# @Author : guanglinzhou (<EMAIL>)
# @Link : https://github.com/GuanglinZhou
# @Version : $Id$
# 获取一个表格,列名如下:
# 路名,WayID,类型(secondary),user,version,timestamp
import os
import csv
import xml.etree.cElementTree as ET
from collections import defaultdict
tree = ET.ElementTree(file='hefei_highways.osm')
root = tree.getroot()
# -
root.attrib
for elem in tree.iter(tag='node'):
if (elem.attrib['id'] == '251187830'):
print(elem.attrib['lat'])
| processDataFromOSM/jupyter/getRoadTable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#load os
import os
#load Flask
#pip install flask==0.12.4
import flask
app = flask.Flask(__name__)
#comment out line before production, only needed during testing:
#app.config['TESTING'] = True
from flask import Flask, render_template,request
#load model preprocessing
import numpy as np
import pandas as pd
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import keras.models
from keras.models import model_from_json
from keras.layers import Input
# -
# watch:
#
# https://www.youtube.com/watch?v=MwZwr5Tvyxo
#
# https://www.youtube.com/watch?v=f6Bf3gl4hWY&t=1743s
#
# https://www.youtube.com/watch?v=IIi6e5oDZ68
#
# https://www.youtube.com/watch?v=RbejfDTHhhg
#
# see code: https://github.com/llSourcell/how_to_deploy_a_keras_model_to_production
# Load pre-trained model into memory
json_file = open('model.json','r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
#load weights into new model
loaded_model.load_weights("model.h5")
print("Loaded Model from disk")
# Helper function for tokenizing text to feed through pre-trained deep learning network
def prepDataForDeepLearning(text):
trainWordFeatures = tokenizer.texts_to_sequences(text)
textTokenized = pad_sequences(trainWordFeatures, 201, padding='post')
return textTokenized
# +
# Load files needed to create proper matrix using tokens from training data
inputDataTrain = pd.DataFrame(pd.read_csv("train_DrugExp_Text.tsv", "\t", header=None))
trainText = [item[1] for item in inputDataTrain.values.tolist()]
trainingLabels = [0 if item[0] == -1 else 1 for item in inputDataTrain.values.tolist()]
VOCABULARY_SIZE=10000
tokenizer = Tokenizer(num_words=VOCABULARY_SIZE)
tokenizer.fit_on_texts(trainText)
## convert words into word ids
meanLength = np.mean([len(item.split(" ")) for item in trainText])
textTokenized = prepDataForDeepLearning(trainText)
# -
loaded_model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
#Test that model works based on accuracy in-sample (comment out to run more quickly)
#Note: this model performs at about 82% out-of-sample
loss, accuracy = loaded_model.evaluate(textTokenized,trainingLabels)
print('loss:', loss)
print('accuracy:', accuracy)
#Test with some text (note: lower to zero = more severe):
textDataTest = ['I had a severe reaction to my medication and it was not fun. I developed a severe rash and was not able to sleep. Terrible! I hate the doctor that gave this to me and I am never taking this drug again.']
textTokenizedTest = prepDataForDeepLearning(textDataTest)
#Note: subtract to get things into severity probability:
out = 1-np.asscalar(loaded_model.predict(textTokenizedTest))
out
#Test with some text (note: lower to zero = more severe):
textDataTest = ['I love my medication!']
textTokenizedTest = prepDataForDeepLearning(textDataTest)
#Note: subtract to get things into severity probability:
out = 1-np.asscalar(loaded_model.predict(textTokenizedTest))
out
# See above. The first message had a 92% probability being severe and the second had a 20% chance.
# Appears to be working!
# define a predict function as an endpoint
@app.route('/', methods=['GET','POST'])
def predict():
#whenever the predict method is called, we're going
#to input the user entered text into the model
#and return a prediction
if request.method=='POST':
textData = request.form.get('text_entered')
print(textData)
textDataArray = [textData]
print(textDataArray)
textTokenized = prepDataForDeepLearning(textDataArray)
print(textTokenized)
prediction = int((1-np.asscalar(loaded_model.predict(textTokenized)))*100)
print(prediction)
#return prediction in new page
return render_template('prediction.html', prediction=prediction)
else:
return render_template("search_page.html")
# Note: This code likely will return error message. Follow instructions below to correct error.
#
# You need to edit the echo function definition at ../Lib/site-packages/click/utils.py the default value for the file parameter must be sys.stdout instead of None.
#
# Do the same for the secho function definition at ../Lib/site-packages/click/termui.py
if __name__ == "__main__":
# start the flask app, allow remote connections
#decide what port to run the app in
port = int(os.environ.get('PORT', 5000))
#this ensures that updates to html/css/js will come through
app.jinja_env.auto_reload = True
#run the app locally on the givn port
app.run(host='0.0.0.0', port=port)
| main (for local run).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparkify Project Workspace
# This workspace contains a tiny subset (128MB) of the full dataset available (12GB). Feel free to use this workspace to build your project, or to explore a smaller subset with Spark before deploying your cluster on the cloud. Instructions for setting up your Spark cluster is included in the last lesson of the Extracurricular Spark Course content.
#
# You can follow the steps below to guide your data analysis and model building portion of this project.
# +
# import libraries
import pyspark
from pyspark.sql import functions as SF
import pyspark.sql.types as pst
from pyspark.sql import Window
from pyspark.sql.types import ArrayType, BooleanType, LongType, FloatType, IntegerType, DoubleType, StringType, DateType
from pyspark.ml.feature import RegexTokenizer, VectorAssembler
from pyspark.ml.classification import RandomForestClassifier, GBTClassifier
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.feature import MinMaxScaler
from pyspark.ml.feature import StringIndexer
from pyspark.mllib.evaluation import MulticlassMetrics
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml import Pipeline
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
import os
import pandas
import matplotlib.pyplot as plt
import numpy as np
import operator
from datetime import datetime as dt
import datetime
from sklearn.feature_extraction import DictVectorizer
from collections import Counter, OrderedDict
pandas.options.display.max_columns = None
pandas.options.display.max_rows = None
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:70% !important; }</style>"))
# -
# I guess in the local mode there is no need to set some of the configs, however I am using the same config as if we were running spark in yarn mode.
# +
# create a Spark session
spark = pyspark.sql.SparkSession.builder.appName("sparkify_ms").enableHiveSupport().\
config("spark.executor.instances","5").config("spark.executor.memory","10g").\
config("spark.executor.cores","7").config("spark.driver.cores","7").\
config("spark.driver.memory","20g").config("spark.driver.maxResultSize","18g").getOrCreate()
sc = spark.sparkContext
print(spark.version, sc.master)
# +
# sc.stop()
# -
# # Load and Clean Dataset
# In this workspace, the mini-dataset file is `mini_sparkify_event_data.json`. Load and clean the dataset, checking for invalid or missing data - for example, records without userids or sessionids.
# Load json to spark dataframe
df = spark.read.json('mini_sparkify_event_data.json')
type(df)
# Check data-types
df.printSchema()
# Data Preview
pdf = df.limit(100).toPandas()
pdf.head()
# Prepare a filturing function
def filter_values(data, spark_functions, col_apply_on, values, isin=True):
"""
:param data :type Spark datafram
:param spark_functions: type spark_functions:
:param col_apply_on :type str - column name for filtering implementation
:param values :type list - of values within col_apply_on
:param isin :type bool - True: implement isin(select values), False: implement ~isin(filter values)
:return: spark dataframe
"""
if isin:
return data.where(spark_functions.col(col_apply_on).isin(values))
else:
return data.where(~spark_functions.col(col_apply_on).isin(values))
# Check volume & missing id values
print(df.count())
df = df.fillna('', subset=['sessionId', 'userId'])
print(filter_values(df, SF, 'sessionId', '', isin=False).count())
print(filter_values(df, SF, 'userId', '', isin=False).count())
# CLEANING STEPS: drop unwanted columns, drop rows where userId value is missing ...
df = df.drop(*['artist','song','id_copy','firstName', 'lastName'])
df = df.dropna(how = 'any', subset = ['userId'])
df = filter_values(df, SF, 'userId', '', isin=False)
df = df.withColumn('userId', df['userId'].cast(IntegerType()))
# # Exploratory Data Analysis
# When you're working with the full dataset, perform EDA by loading a small subset of the data and doing basic manipulations within Spark. In this workspace, you are already provided a small subset of data you can explore.
# ### Define Churn
#
# Once you've done some preliminary analysis, create a column `Churn` to use as the label for your model. I suggest using the `Cancellation Confirmation` events to define your churn, which happen for both paid and free users. As a bonus task, you can also look into the `Downgrade` events.
# Create churn column
df = df.withColumn('churn', SF.when(SF.col('page') == 'Cancellation Confirmation', 1).otherwise(0))
df.groupBy("churn").count().orderBy('count').sort(SF.desc("count")).show()
# +
# Spread the churn value to all rows of "churn" users
window = Window.partitionBy("userId").rangeBetween(Window.unboundedPreceding,
Window.unboundedFollowing)
df = df.withColumn("churn", SF.sum("churn").over(window))
df.groupBy("churn").count().orderBy('count').sort(SF.desc("count")).show()
# -
# ### Explore Data
# Once you've defined churn, perform some exploratory data analysis to observe the behavior for users who stayed vs users who churned. You can start by exploring aggregates on these two groups of users, observing how much of a specific action they experienced per a certain time unit or number of songs played.
# #### Preview values in some of the columns:
# Obviously one of the most interesting columns is the 'page' - will be a source of many usefull features.
print(df.groupBy("userId").count().count(), '\n')
df.groupBy("auth").count().orderBy('count').sort(SF.desc("count")).show(10)
df.groupBy("method").count().orderBy('count').sort(SF.desc("count")).show(10)
df.groupBy("level").count().orderBy('count').sort(SF.desc("count")).show(10)
df.groupBy("page").count().orderBy('count').sort(SF.desc("count")).show(23)
# #### With a help of regexTokenizer extract userAgent words - these details can be transformed into a valuable features
#
df = df.withColumn('userAgent_reg', SF.lower(SF.regexp_replace('userAgent', "[^0-9a-zA-Z\\s]", "")))
regexTokenizer = RegexTokenizer(inputCol="userAgent_reg", outputCol="userAgent_words")
df = regexTokenizer.transform(df)
print(df.groupBy("userAgent_words").count().count())
df.groupBy("userAgent_words").count().orderBy('count').sort(SF.desc("count")).show(10, False)
# ##### Words Correlation heatmap
# In the heatmap below - created out of the word-matrix (from userAgent word) - we can see that some of the words are perfectly correlated with other ones (yellow dots). We'll need to remove some words accordingly - as we do not want to have perfectly correlated features in the training df.
# +
words_list = df.select('userAgent_words').distinct().rdd.map(lambda r: r[0]).collect()
v = DictVectorizer()
X = v.fit_transform(Counter(f) for f in (_l for _l in words_list))
print(X.shape)
words_df = pandas.DataFrame(X.toarray())
words_df.columns = words_df.columns.map(dict([(val, key) for key, val in v.vocabulary_.items()]))
f = plt.figure(figsize=(8, 6))
plt.matshow(words_df.corr(), fignum=f.number)
plt.show()
# -
# #### In the next steps let's create some date columns which will help us with research and/or with new features:
# - "temp_" & "hlp_" prefixes are used for temporary or helping columns
# - "fe_" prefix will be used strictly for features to train the model on.
# Convert timestamp to date
d_fromtmstp = SF.udf(lambda x: dt.fromtimestamp(x/1000).date(), DateType())
df = df.withColumn("hlp_date", d_fromtmstp(SF.col('ts')))
# Get MIN & MAX date for each user
df = df.withColumn("hlp_date_max", SF.max("hlp_date").over(Window.partitionBy("userId")))
df = df.withColumn("hlp_date_min", SF.min("hlp_date").over(Window.partitionBy("userId")))
# Total days per user
timeFmt = "yyyy-MM-dd"
df = df.withColumn('fe_total_days', SF.datediff(SF.to_date('hlp_date_max', timeFmt), SF.to_date('hlp_date_min', timeFmt)))
# #### Check the mean of days customers are using the page - people who cancelled vs others
# This is a tricky feature, as the numbers in first table may change by time.
# However the second table tells us which day could be critical and so this feature could help our model too ...
df.filter(df.churn!=1).select(['fe_total_days']).describe().show()
df.filter(df.churn==1).select(['fe_total_days']).describe().show()
# #### Increase / Decrease of songs played daily by user per quarters - can we use it as a feature?
#
# This is an experiment with a categorical feature which represents how number of played songs by user increases or decreases from quarter to quarter.
# Let's say we have 20 days of sessions for a user. These 20 days are divided into quarters and we calculate a mean of played songs for each quarter. Than Q1:5, Q2:10, Q3:7, Q4:3 is converted to a value IDD (increase, decrease, decrease).
# +
join_col = 'temp_join_key'
df = df.withColumn(join_col, SF.concat(SF.col('userId'),SF.lit('_'), SF.col('hlp_date')))
temp_df = df.groupby(join_col).agg(SF.count(SF.when(SF.col('page')=='NextSong', True)).alias('hlp_songs_daily'))
df_churn = df.filter(df.churn==1)
df_churn = df_churn.join(temp_df, 'temp_join_key', how='left')
temp_df = df_churn.dropDuplicates(subset=['temp_join_key'])\
.withColumn('temp_inc_dec', SF.collect_list('hlp_songs_daily').over(Window.partitionBy('userId').orderBy('hlp_date')))
temp_df = temp_df.orderBy(SF.desc('ts')).dropDuplicates(subset=['userId']).select(['userId', 'temp_inc_dec'])
df_churn = df_churn.join(temp_df, 'userId', how='left')
def increase_decrease(_list):
arrs = np.array_split(_list, 4)
quarters = [arrs[1].mean() - arrs[0].mean(),
arrs[2].mean() - arrs[1].mean(),
arrs[3].mean() - arrs[2].mean()]
def _repl(x):
if x < 0: return 'D'
else: return 'I'
return ''.join([_repl(x) for x in quarters])
increase_decrease_udf = SF.udf(increase_decrease, StringType())
df_churn = df_churn.withColumn("fe_songs_daily_inc_dec", increase_decrease_udf(df_churn['temp_inc_dec']))
# -
df_churn.select('fe_songs_daily_inc_dec').distinct().rdd.map(lambda r: r[0]).collect()
df_churn.dropDuplicates(subset=['userId']).groupBy("fe_songs_daily_inc_dec").count().orderBy('count')
.sort(SF.desc("count")).show(10)
# Even though I expected to have more D's at the end of the triplets for churn customers, I will use this feature for modeling.
# # Feature Engineering
# In following step I will create couple of basic features (total counts or averages) and couple of experimental features.
# #### total sessions per user
temp_df = df.groupby('userId').agg(SF.count('userId').alias('fe_sessions_total'))
df = df.join(temp_df, 'userId', how='left')
# #### FE level switch count
# 'fe_level_switch_count' feature tells us how many times user switched from PAID to FREE or vice versa
# +
df = df.withColumn("hlp_level", SF.collect_set("level").over(Window.partitionBy("userId", "hlp_date").orderBy(["ts"])))
udf_paid_count = SF.udf(lambda x: int(len(set(x))>1), IntegerType())
df = df.withColumn('fe_hlp_level', udf_paid_count('hlp_level'))
temp_df = df.select('userId', 'hlp_date', 'fe_hlp_level').filter(df.fe_hlp_level==1).groupby('userId', 'hlp_date').agg(SF.max('hlp_date'))
temp_df = temp_df.drop(*['hlp_date', 'max(hlp_date)']).groupby('userId').agg(SF.count('userId').alias('fe_level_switch_count')).drop('fe_hlp_level')
df = df.join(temp_df, 'userId', how='left')
# -
# #### FE error & help count per user
# +
# Error
temp_df = df.groupby('userId').agg(SF.count(SF.when(SF.col('page')=='Error', True)).alias('fe_error'))
df = df.join(temp_df, 'userId', how='left')
# Help
temp_df = df.groupby('userId').agg(SF.count(SF.when(SF.col('page')=='Help', True)).alias('fe_help'))
df = df.join(temp_df, 'userId', how='left')
# -
# #### Avg Session length
df = df.withColumn("fe_session_len", SF.avg("length").over(Window.partitionBy("userId").orderBy(["ts"])))
# #### Songs played daily
# +
# df = df.withColumn("fe_songs_daily", SF.count("length").over(Window.partitionBy("userId", "hlp_date").orderBy(["ts"])))
# join_col = 'temp_join_key'
# df = df.withColumn(join_col, SF.concat(SF.col('userId'),SF.lit('_'), SF.col('hlp_date')))
# temp_df = df.groupby(join_col).agg(SF.count(SF.when(SF.col('page')=='NextSong', True)).alias('fe_songs_daily'))
# df = df.join(temp_df, 'temp_join_key', how='left').drop(join_col)
# -
# #### Total Songs played by user
temp_df = df.groupby('userId').agg(SF.count(SF.when(SF.col('page')=='NextSong', True)).alias('fe_songs_total'))
df = df.join(temp_df, 'userId', how='left')
# #### Total Songs / Total Days ratio
df = df.withColumn('fe_days_songs_ratio', SF.col('fe_total_days')/SF.col('fe_songs_total'))
# #### Total sessions / Total days ratio
df = df.withColumn('fe_sessions_total', SF.col('fe_total_days')/SF.col('fe_songs_total'))
# #### Location State
# Extracting state abbreviations from location details
def extract_state(loc_col):
try:
return str(loc_col).split(',')[-1].strip()
except:
return ''
udf_extract_state = SF.udf(extract_state, StringType())
df = df.withColumn('fe_state', udf_extract_state('location'))
# #### Tunes played daily - increase/decrease by quarters
#
# Based on the experiment from the the research part I created a function which can be easily used not only for songs played increase/decrease but also for Error or Thumbs Down page:
# +
def page_inc_dec_quarterly_FE(df, page, new_feature_name):
""" Returns the quarter increase/decrease categories
for a selected page per user.
param: df, type: spark dataframe
param: page, type: string
param: new_feature_name, type: string
return: spark dataframe
"""
# Count the daily usage of selected page per user (temp_page_daily)
join_col = 'temp_join_key'
df = df.withColumn(join_col, SF.concat(SF.col('userId'),SF.lit('_'), SF.col('hlp_date')))
temp_df = df.groupby(join_col).agg(SF.count(SF.when(SF.col('page')==page, True)).alias('temp_page_daily'))
df = df.join(temp_df, 'temp_join_key', how='left')
# Collect the temp_page_daily numbers into lists
temp_df = df.dropDuplicates(subset=['temp_join_key'])\
.withColumn('temp_inc_dec', SF.collect_list('temp_page_daily').over(Window.partitionBy('userId').orderBy('hlp_date')))
# Get the major temp_page_daily list - the one from last day & join with df
temp_df = temp_df.orderBy(SF.desc('ts')).dropDuplicates(subset=['userId']).select(['userId', 'temp_inc_dec'])
df = df.join(temp_df, 'userId', how='left').drop('temp_page_daily')
def increase_decrease(_list):
"""Splits the list into 4 ongoing parts,
calculates the mean and identifies
increase or decrease between these quarters.
Returns values like DDI, DID, III, etc ..."""
arrs = np.array_split(_list, 4)
quarters = [arrs[1].mean() - arrs[0].mean(),
arrs[2].mean() - arrs[1].mean(),
arrs[3].mean() - arrs[2].mean()]
def _repl(x):
if x < 0: return 'D'
else: return 'I'
return ''.join([_repl(x) for x in quarters])
# Turn the lists of numbers into the Increase/Decrease categories
increase_decrease_udf = SF.udf(increase_decrease, StringType())
return df.withColumn(new_feature_name, increase_decrease_udf(df['temp_inc_dec'])).drop('temp_inc_dec')
df = page_inc_dec_quarterly_FE(df, 'NextSong', 'fe_songs_daily_inc_dec')
# -
# #### Errors daily - increase/decrease by quarters
df = page_inc_dec_quarterly_FE(df, 'Error', 'fe_error_daily_inc_dec')
# #### Thumbs Down daily - increase/decrease by quarters
df = page_inc_dec_quarterly_FE(df, 'Thumbs Down', 'fe_Thumbs_Down_daily_inc_dec')
# #### Number of downgrades per user
temp_df = df.groupby('userId').agg(SF.count(SF.when(SF.col('page')=='Submit Downgrade', True))
.alias('fe_downgrades_total'))
df = df.join(temp_df, 'userId', how='left')
# #### Add fe_ prefix to existing features (level, gender, status)
as_is_features = ['level', 'gender', 'status']
for col in as_is_features:
df = df.withColumnRenamed(col, 'fe_{}'.format(col))
# Lets finalize the lazy function runs - before we start the complicated part of feature engineering
df.cache()
# #### FE userAgent
# In this part we tokenize the userAgent words, reduce them to not get perfectly correlated features and create a new feature/column which holds the information about the total usage of each word per user.
# Each word represents a system, version or even a hardware used by customer and it can become a good feature when it comes to predict churn.
# +
def regex_tokenize(df):
""" Creates new field containing the list of words
returned by RegexTokenizer from 'userAgent' column.
param: df, type: spark dataframe
return: spark dataframe
"""
df = df.withColumn('userAgent_reg', SF.lower(SF.regexp_replace('userAgent', "[^0-9a-zA-Z\\s]", "")))
regexTokenizer = RegexTokenizer(inputCol="userAgent_reg", outputCol="userAgent_words")
return regexTokenizer.transform(df)
def intersect(_lists):
"""Returns intersection of multiple lists.
param: list of lists
"""
intrs = list(set(words_list[0]).intersection(set(words_list[1])))
for x in range(2,len(_lists)):
intrs = list(set(intrs).intersection(set(words_list[x])))
if not intrs: break
return intrs
def remove_perf_corr_columns(words_list):
"""Removes words from the Tokenizer output lists
to get rid of perfectly correlated couples.
param: list of lists
return: list of lists
"""
v = DictVectorizer()
while True:
X = v.fit_transform(Counter(f) for f in (_l for _l in words_list))
words_df = pandas.DataFrame(X.toarray())
words_df.columns = words_df.columns.map(dict([(value, key) for key, value in v.vocabulary_.items()]))
df_corr = words_df.corr().stack().reset_index()
df_corr.columns = ['FEATURE_1', 'FEATURE_2', 'CORRELATION']
df_corr = df_corr[((df_corr.CORRELATION==1)) & (df_corr.FEATURE_1!=df_corr.FEATURE_2)]
if df_corr.shape[0] > 0:
def select_feature(df):
return str(sorted([df['FEATURE_1'], df['FEATURE_2']]))
df_corr['Feature_X'] = df_corr.apply(select_feature, axis=1)
df_corr.drop_duplicates(subset='Feature_X', inplace=True)
remove_cols = df_corr.FEATURE_1.tolist()
new_words_list = []
for l in words_list:
l = [item for item in l if item not in remove_cols]
new_words_list.append(l)
words_list = new_words_list
else:
break
return words_list
def merge_lists(_lists):
"""Merges multiple lists together.
param: list of lists
return: list
"""
result = _lists[0]
for x in range(1, len(_lists)):
result.extend(_lists[x])
return list(set(result))
# df = regex_tokenize(df) - already done in exploration part
words_list = df.select('userAgent_words').distinct().rdd.map(lambda r: r[0]).collect()
words_to_remove = intersect(words_list)
all_words = merge_lists(words_list)
all_words = [item for item in all_words if item not in words_to_remove]
# -
# Finally: Create the columns with summarized usage of certain words
i = 1
for col_name in all_words:
i += 1
udf_contains = SF.udf(lambda x: int(col_name in x), IntegerType())
temp_col = 'temp_ua_{}'.format(col_name)
df = df.withColumn(temp_col, udf_contains(df.userAgent_words))
df = df.withColumn('fe_ua_{}'.format(col_name), SF.sum(temp_col)
.over(Window.partitionBy('userId')))
if (i % 3) == 0: df.cache()
# #### Save / Load preprocessed dataframe for more efficiency
# +
# df.write.parquet(os.path.join('{}/'.format(os.path.abspath(os.getcwd())), 'etl_df_output'))
df = spark.read.parquet(os.path.join('{}/etl_df_output'.format(os.path.abspath(os.getcwd()))))
# -
# # Modeling
# Let's start with the final preprocessing of our dataframe. For this purpose I have combined all the necessary steps into one function.
# For modelling we need a dataframe containing 1 row per user. Extracting these rows in the first step of the function will save us a lot of time as every other operation will be applied to reduced dataframe. In the next steps, except for fillna, we extract the bunch of features we plan to use, create the "label" column out of "churn", we convert categorical features to numerical (this is what pyspark.ml classifiers like RandomForest or GradientBoostedTrees are expecting) and finally we split the data into 3 parts: Train, Test & Validation.
# +
def get_cols_containing_null(df):
"""Returns a list of columns containing null values & Na's
param: df, type: spark dataframe
return: python list
"""
check = df.select(*[col for col in df.columns if col.startswith('fe_')])
check = check.select([SF.count(SF.when(SF.isnan(c) | SF.isnull(c), c)).alias(c)
for (c,c_type) in check.dtypes if c_type not in ('timestamp', 'string', 'date', ' array')])\
.toPandas().rename(index={0: '_count'}).T.reset_index()
return check[check._count>0]['index'].values.tolist()
def preprocess_and_split(df, split_ratio=[0.6, 0.4]):
""" - Keeps 1 row per user
- fills na's & nulls
- selects appropriate features
- Converts categorical to numerical columns
- splits the dataframe by provided ratio
param: df, type: spark dataframe
param: split_ration, type: list (optional)
return: train, test and validation df (spark dataframes)
"""
# select 1 row per user
window = Window.partitionBy('userId').orderBy(SF.col('ts').desc())
dff = df.withColumn('row',SF.row_number().over(window)).where(SF.col('row')==2).drop('row')
# Fillna
na_subset = get_cols_containing_null(dff)
dff = dff.fillna(0, subset=na_subset)
# select only features & label fields
features = [col for col in df.columns if col.startswith('fe_')]
dff = dff.select(*features + ['churn'])
dff = dff.withColumn('label', df['churn'].cast(DoubleType()))
dff = dff.drop('churn')
# Convert categorical columns to numerical
str_cols = [f.name for f in dff.schema.fields if isinstance(f.dataType, StringType)]
for col in str_cols:
dff = dff.withColumnRenamed(col, '{}_temp'.format(col))
stringIndexer = StringIndexer(inputCol='{}_temp'.format(col), outputCol=col)
si_model = stringIndexer.fit(dff)
dff = si_model.transform(dff).drop('{}_temp'.format(col))
train_df, test_val = dff.randomSplit(split_ratio, seed=42)
test_val.cache()
test_df, val_df = test_val.randomSplit(split_ratio, seed=5)
return train_df, test_df, val_df
# +
# Check if we have an appropriate sample of "churn" occasions in each of the datasets.
train_df, test_df, val_df = preprocess_and_split(df)
print('Train: {}/{}'.format(train_df.count(), train_df.filter(train_df.label==1).count()))
print('Test: {}/{}'.format(test_df.count(), test_df.filter(test_df.label==1).count()))
print('Validation: {}/{}'.format(val_df.count(), val_df.filter(val_df.label==1).count()))
# -
# Finally we can start with the training!
# Here is the sequence of our steps:
# - Create a dictionary with classifiers as the keys and appropriate paramGrids as values
# - Prepare Vectorizer and Scaler for the pipeline
# (I decided to use RandomForest and GradientBoostedTrees and therefore Scaling is not necessary,
# however we can keep it in the pipeline in case we decide to add LinearRegression too.
# Scaling will not do any damage to the results anyway.)
# - Next we loop over the classifiers and get the best one of each grid by using F1 score metric
# of the MulticlassClassificationEvaluator.
# - We save the results and models to a dictionary.
# - In the last step we select the best of the best models by comparing the results of each iteration.
#
# Why did we use the F1 score as the key metric?
# As we are using a small dataset, we need to use a bit more demanding metric than precision or recall. F-score is calculated from the precision and recall of testing.
# Precision is the number of correctly identified positive results divided by the number of all positive results, including those not identified correctly, and the recall is the number of correctly identified positive results divided by the number of all samples that should have been identified as positive.
# Our goal is to predict true positives as accurate as possible while false positive predictions could lead to wasted resources. Therefore the F1 score became our metric.
# +
# Create a dictionary of classifiers & their paramGrids
eval_dict = {}
rf_clf = RandomForestClassifier()
gb_clf = GBTClassifier()
paramGrid_rf = ParamGridBuilder() \
.addGrid(rf_clf.numTrees, [15]) \
.addGrid(rf_clf.maxDepth, [7, 15]) \
.build()
paramGrid_gb = ParamGridBuilder()\
.addGrid(gb_clf.maxDepth, [7, 12])\
.addGrid(gb_clf.maxBins, [20])\
.addGrid(gb_clf.maxIter, [6]).build()
classifiers = {rf_clf: paramGrid_rf, gb_clf: paramGrid_gb}
# Prepare vectorizer & scaler for the pipeline
features = [col for col in train_df.columns if col.startswith('fe_')]
assembler = VectorAssembler(inputCols=features, outputCol='features_vect')
minmaxscaler = MinMaxScaler(inputCol="features_vect", outputCol="features")
# Run the CrossValidator for each classifier and its paramGrid
for clf, paramGrid in classifiers.items():
pipeline =Pipeline(stages=[assembler, minmaxscaler, clf])
evaluator=MulticlassClassificationEvaluator(metricName="f1")
cross_val = CrossValidator(estimator=pipeline,
estimatorParamMaps=paramGrid,
evaluator=evaluator,
numFolds=3)
# Evaluate the model and get the F1 score
model = cross_val.fit(train_df)
eval_result = evaluator.evaluate(model.transform(test_df))
# Create a dictionary of models and F1 scores
eval_dict[model] = eval_result
print('{}: {}'.format(str(model), eval_result))
# Select the model with highest evaluation score
best_model = max(eval_dict, key=eval_dict.get)
# -
# We achieved F1 training score of 0.94 with GradientBoostedTrees.
# Let's have a look at the parameters and testing score of the best model:
param_dict = {}
for m, p in zip(best_model.avgMetrics, best_model.getEstimatorParamMaps()):
param_dict[str(p)] = m
print('Test F1 Score: {}'.format(round(max(best_model.avgMetrics), 2)))
max(param_dict, key=param_dict.get)
# +
# model_path = '{}/best_model/'.format(os.path.abspath(os.getcwd()))
# best_model.save(model_path)
# -
# We can now validate our best model on the validation dataset.
# I've added couple of more details/metrics to get a better picture of how good our model really is.
# +
pred_df = best_model.transform(val_df)
print('Total correct predictions: {} / 39'.format(pred_df.filter(pred_df.label == pred_df.prediction).count()))
print('Churn correct predictions: {} / 13'.format(pred_df.filter((pred_df.label == pred_df.prediction) &
(pred_df.label == 1)).count()))
print('Churn incorrect predictions: {}'.format(pred_df.filter((pred_df.label == 0) &
(pred_df.prediction == 1)).count()))
# convert dataframe to rdd to be able to use MulticlassMetrics
prd_rdd = pred_df.select(['label', 'prediction']).rdd
mc_metrics = MulticlassMetrics(prd_rdd)
print('Precision: {}'.format(round(mc_metrics.precision(1.0), 2)))
print('Recall: {}'.format(round(mc_metrics.recall(1.0), 2)))
print('F1-Score: {}'.format(round(mc_metrics.fMeasure(1.0), 2)))
print('Accuracy: {}'.format(round(mc_metrics.accuracy, 2)))
# -
# ### We achieved quite a good score!
| Sparkify_capstone_project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">Registration: Memory-Time Trade-off</h1>
#
# When developing a registration algorithm or when selecting parameter value settings for an existing algorithm our choices are dictated by two, often opposing, constraints:
# <ul>
# <li>Required accuracy.</li>
# <li>Alloted time.</li>
# </ul>
#
# As the goal of registration is to align multiple data elements into the same coordinate system, it is only natural that the primary focus is on accuracy. In most cases the reported accuracy is obtained without constraining the algorithm's execution time. Don't forget to provide the running times even if they are not critical for your particular application as they may be critical for others.
#
# With regard to the emphasis on execution time, on one end of the spectrum we have longitudinal studies where time constraints are relatively loose. In this setting a registration taking an hour may be perfectly acceptable. At the other end of the spectrum we have intra-operative registration. In this setting, registration is expected to complete within seconds or minutes. The underlying reasons for the tight timing constraints in this setting have to do with the detrimental effects of prolonged anesthesia and with the increased costs of operating room time. While short execution times are important, simply completing the registration on time without sufficient accuracy is also unacceptable.
#
# This notebook illustrates a straightforward approach for reducing the computational complexity of registration for intra-operative use via preprocessing and increased memory usage, a case of the [memory-time trade-off](https://en.wikipedia.org/wiki/Space%E2%80%93time_tradeoff).
#
# The computational cost of registration is primarily associated with interpolation, required for evaluating the similarity metric. Ideally we would like to use the fastest possible interpolation method, nearest neighbor. Unfortunately, nearest neighbor interpolation most often yields sub-optimal results. A straightforward solution is to pre-operatively create a super-sampled version of the moving-image using higher order interpolation*. We then perform registration using the super-sampled image, with nearest neighbor interpolation.
#
# Tallying up time and memory usage we see that:
#
# <table>
# <tr><td></td> <td><b>time</b></td><td><b>memory</b></td></tr>
# <tr><td><b>pre-operative</b></td> <td>increase</td><td>increase</td></tr>
# <tr><td><b>intra-operative</b></td> <td>decrease</td><td>increase</td></tr>
# </table><br><br>
#
#
# <font size="-1">*A better approach is to use single image super resolution techniques such as the one described in <NAME>, <NAME>, <NAME>,"Single-image super-resolution of brain MR images using overcomplete dictionaries", <i>Med Image Anal.</i>, 17(1):113-132, 2013.</font>
#
# +
import SimpleITK as sitk
import numpy as np
import matplotlib.pyplot as plt
from __future__ import print_function
#utility method that either downloads data from the MIDAS repository or
#if already downloaded returns the file name for reading from disk (cached data)
# %run update_path_to_download_script
from downloaddata import fetch_data as fdata
import registration_utilities as ru
from ipywidgets import interact, fixed
# %matplotlib inline
def register_images(fixed_image, moving_image, initial_transform, interpolator):
registration_method = sitk.ImageRegistrationMethod()
registration_method.SetMetricAsMattesMutualInformation(numberOfHistogramBins=50)
registration_method.SetMetricSamplingStrategy(registration_method.REGULAR)
registration_method.SetMetricSamplingPercentage(0.01)
registration_method.SetInterpolator(interpolator)
registration_method.SetOptimizerAsGradientDescent(learningRate=1.0, numberOfIterations=1000)
registration_method.SetOptimizerScalesFromPhysicalShift()
registration_method.SetInitialTransform(initial_transform, inPlace=False)
final_transform = registration_method.Execute(fixed_image, moving_image)
return( final_transform, registration_method.GetOptimizerStopConditionDescription())
# -
# ## Load data
#
# We use the the training data from the Retrospective Image Registration Evaluation (<a href="http://www.insight-journal.org/rire/">RIRE</a>) project.
#
# The RIRE reference, ground truth, data consists of a set of corresponding points in the fixed and moving coordinate systems. These points were obtained from fiducials embedded in the patient's skull and are thus sparse (eight points). We use these to compute the rigid transformation between the two coordinate systems, and then generate a dense reference. This generated reference data is more similar to the data you would use for registration evaluation.
# +
fixed_image = sitk.ReadImage(fdata("training_001_ct.mha"), sitk.sitkFloat32)
moving_image = sitk.ReadImage(fdata("training_001_mr_T1.mha"), sitk.sitkFloat32)
fixed_fiducial_points, moving_fiducial_points = ru.load_RIRE_ground_truth(fdata("ct_T1.standard"))
R, t = ru.absolute_orientation_m(fixed_fiducial_points, moving_fiducial_points)
reference_transform = sitk.Euler3DTransform()
reference_transform.SetMatrix(R.flatten())
reference_transform.SetTranslation(t)
# Generate a reference dataset from the reference transformation (corresponding points in the fixed and moving images).
fixed_points = ru.generate_random_pointset(image=fixed_image, num_points=1000)
moving_points = [reference_transform.TransformPoint(p) for p in fixed_points]
interact(lambda image1_z, image2_z, image1, image2:ru.display_scalar_images(image1_z, image2_z, image1, image2),
image1_z=(0,fixed_image.GetSize()[2]-1),
image2_z=(0,moving_image.GetSize()[2]-1),
image1 = fixed(fixed_image),
image2=fixed(moving_image));
# -
# ## Invest time and memory in exchange for future time savings
#
# We now resample our moving image to a finer spatial resolution.
# +
# Isotropic voxels with 1mm spacing.
new_spacing = [1.0]*moving_image.GetDimension()
# Create resampled image using new spacing and size.
original_size = moving_image.GetSize()
original_spacing = moving_image.GetSpacing()
resampled_image_size = [int(spacing/new_s*size)
for spacing, size, new_s in zip(original_spacing, original_size, new_spacing)]
resampled_moving_image = sitk.Image(resampled_image_size, moving_image.GetPixelID())
resampled_moving_image.SetSpacing(new_spacing)
resampled_moving_image.SetOrigin(moving_image.GetOrigin())
resampled_moving_image.SetDirection(moving_image.GetDirection())
# Resample original image using identity transform and the BSpline interpolator.
resample = sitk.ResampleImageFilter()
resample.SetReferenceImage(resampled_moving_image)
resample.SetInterpolator(sitk.sitkBSpline)
resample.SetTransform(sitk.Transform())
resampled_moving_image = resample.Execute(moving_image)
print('Original image size and spacing: {0} {1}'.format(original_size, original_spacing))
print('Resampled image size and spacing: {0} {1}'.format(resampled_moving_image.GetSize(),
resampled_moving_image.GetSpacing()))
print('Memory ratio: 1 : {0}'.format((np.array(resampled_image_size)/np.array(original_size).astype(float)).prod()))
# -
# Another option for resampling an image, without any transformation, is to use the ExpandImageFilter or
# in its functional form SimpleITK::Expand. This filter accepts the interpolation method and an integral expansion factor. This is less flexible than the resample filter as we have less control over the resulting image's spacing.
# On the other hand this requires less effort from the developer, a single line of code as compared to the cell above:
#
# resampled_moving_image = sitk.Expand(moving_image,
# [int(original_s/new_s + 0.5) for original_s, new_s in zip(original_spacing, new_spacing)],
# sitk.sitkBSpline)
#
# ## Registration
#
# ### Initial Alignment
#
# We will use the same initial alignment for both registrations.
initial_transform = sitk.CenteredTransformInitializer(fixed_image,
moving_image,
sitk.Euler3DTransform(),
sitk.CenteredTransformInitializerFilter.GEOMETRY)
# ### Original Resolution
#
# For this registration we use the original resolution and linear interpolation.
# +
# %%timeit -r1 -n1
# The arguments to the timeit magic specify that this cell should only be run once.
# We define this variable as global so that it is accessible outside of the cell (timeit wraps the code in the cell
# making all variables local, unless explicitly declared global).
global original_resolution_errors
final_transform, optimizer_termination = register_images(fixed_image, moving_image, initial_transform, sitk.sitkLinear)
final_errors_mean, final_errors_std, _, final_errors_max, original_resolution_errors = ru.registration_errors(final_transform, fixed_points, moving_points)
print(optimizer_termination)
print('After registration, errors in millimeters, mean(std): {:.2f}({:.2f}), max: {:.2f}'.format(final_errors_mean, final_errors_std, final_errors_max))
# -
# ### Higher Resolution
#
# For this registration we use the higher resolution image and nearest neighbor interpolation.
# +
# %%timeit -r1 -n1
# The arguments to the timeit magic specify that this cell should only be run once.
# We define this variable as global so that it is accessible outside of the cell (timeit wraps the code in the cell
# making all variables local, unless explicitly declared global).
global resampled_resolution_errors
final_transform, optimizer_termination = register_images(fixed_image, resampled_moving_image, initial_transform, sitk.sitkNearestNeighbor)
final_errors_mean, final_errors_std, _, final_errors_max, resampled_resolution_errors = ru.registration_errors(final_transform, fixed_points, moving_points)
print(optimizer_termination)
print('After registration, errors in millimeters, mean(std): {:.2f}({:.2f}), max: {:.2f}'.format(final_errors_mean, final_errors_std, final_errors_max))
# -
# ### Compare the error distributions
#
# To fairly compare the two registration above we look at their running times (see results above) and their
# error distributions (plotted below).
plt.hist(original_resolution_errors, bins=20, alpha=0.5, label='original resolution', color='blue')
plt.hist(resampled_resolution_errors, bins=20, alpha=0.5, label='higher resolution', color='green')
plt.legend()
plt.title('TRE histogram');
# ## Conclusions
#
# It appears that the memory-time trade-off works in our favor, but is this always the case? Well, you will have to answer that for yourself.
#
# Some immediate things you can try:
# * Change the interpolation method for the "original resolution" registration to nearest neighbor.
# * Change the resolution of the resampled image - will a higher resolution always result in faster running times?
| Python/64_Registration_Memory_Time_Tradeoff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualize vanilla LIME and robust LIME in the PCA and t-SNE spaces
#
# 1. Load COMPAS dataset
# 2. Generate synthetic neighborhood via vanilla LIME
# * Can be done via [this LIME tabular method](https://github.com/marcotcr/lime/blob/2ba75c188dcffe3e926c093efc5d03a0d51692b6/lime/lime_tabular.py#L468)
# 3. Generate synthetic neighborhood via robust LIME (CTGAN)
# 4. Reduce the original dataset, vanilla LIME neighborhood, and robust LIME neighborhood to 2-D using [PCA](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) and [t-SNE](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html)
# * Tips on t-SNE: https://distill.pub/2016/misread-tsne/
# 5. Plot
# +
# %matplotlib inline
import os
import sys
sys.path.append('../')
from experiments.utils.datasets import get_dataset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Explainers
from lime.lime_tabular import LimeTabularExplainer
from faster_lime.explainers.numpy_robust_tabular_explainer import NumpyRobustTabularExplainer
# d reduction utilities
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
# 1. Load dataset
data = get_dataset('compas', {})
# +
# 2. Generate vanilla LIME synthetic neighborhood
data_synthetic_vanilla = ...
# +
# 3. Generate robust LIME synthetic neighborhood
data_synthetic_robust = ...
# +
# 4. Reduce to 2-D
# +
# 5. Plot
| notebooks/visualize_neighborhood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Dfz1Z0z8tndQ" colab_type="text"
# # Input Data Preparation
# + id="ZA0oN9b7tndR" colab_type="code" colab={}
from glob import glob
from imageio import imread
from tqdm import tqdm
import tensorflow as tf
import json
import os.path as osp
import numpy as np
import numpy.random as npr
import cv2
import os
from tensorflow.contrib.learn.python.learn.datasets import base
# + id="kkeXsn3ztndW" colab_type="code" colab={}
class DetectionDataset(object):
def __init__(self, data_type, class_names, images, gt_box_sets):
self._num_samples = len(images)
self._data_type = data_type
self._class_names = class_names
self._num_classes = len(self._class_names)
self._images =images
self._gt_box_sets = gt_box_sets
self._image_shapes = np.asarray([img.shape for img in images], dtype=np.float32)
self._augment = True
self._indices = None
self._cursor = 0
self._epoch_count = 0
@property
def class_names(self):
return self._class_names
@property
def num_classes(self):
return self._num_classes
@property
def images(self):
return self._images
@property
def gt_box_sets(self):
return self._gt_box_sets
def set_augment(self, augment):
self._augment = augment
def get_image(self, index):
return self._images[index]
def get_gt_box_set(self, index):
return self._gt_box_sets[index]
def get_image_shape(self, index):
return self._image_shapes[index]
@property
def num_samples(self):
return self._num_samples
@property
def num_epochs(self):
return self._epoch_count
def _start_next_epoch(self, shuffle):
is_initial = self._indices is None
self._indices = npr.permutation(self._num_samples) if shuffle else np.arange(self._num_samples)
self._cursor = 0
if not is_initial:
self._epoch_count += 1
def next_batch(self, batch_size, shuffle=True, get_idx=False):
if self._indices is None:
self._start_next_epoch(shuffle)
stride = min(batch_size, self._num_samples - self._cursor)
indices = self._indices[self._cursor:self._cursor + stride]
self._cursor += stride
while len(indices) < batch_size:
self._start_next_epoch(shuffle)
stride = min(batch_size, self.num_samples - self._cursor)
indices = np.concatenate([indices, self._indices[self._cursor:self._cursor + stride]])
self._cursor += stride
batch_images, batch_gt_box_sets = self._images[indices], self._gt_box_sets[indices]
if batch_size == 1:
batch_images = np.asarray([batch_images[0]], dtype=np.float32)
batch_gt_box_sets = np.asarray([batch_gt_box_sets[0]], dtype=np.float32)
if get_idx:
return batch_images, batch_gt_box_sets, indices
else:
return batch_images, batch_gt_box_sets
def get_batch(self, idx):
indices = [idx]
batch_images, batch_gt_box_sets = self._images[indices], self._gt_box_sets[indices]
batch_images = np.asarray([batch_images[0]], dtype=np.float32)
batch_gt_box_sets = np.asarray([batch_gt_box_sets[0]], dtype=np.float32)
return batch_images, batch_gt_box_sets
def read_data(data_dir, load_rate=1.0):
# Get a list of the class names.
class_names = []
with open(osp.join(data_dir, 'classes.json'), 'r') as f:
class_dict = json.load(f)
for i in sorted(class_dict.keys(), key=int):
class_names.append(class_dict[str(i)])
# Load image and annotation files.
images, ground_truth_box_sets = [], []
image_files = sorted(glob(osp.join(data_dir, 'images/*.jpg')))
annotation_files = sorted(glob(osp.join(data_dir, 'annotations/*.anno')))
num_samples = len(image_files)
if load_rate < 1:
num_samples = int(num_samples * load_rate)
image_files = image_files[:num_samples]
annotation_files = annotation_files[:num_samples]
assert len(image_files) == len(annotation_files)
for img_fp, anno_fp in zip(tqdm(image_files), annotation_files):
img = imread(img_fp)
if len(img.shape) < 3:
img = cv2.cvtColor(img, cv2.COLOR_GRAY2RGB)
images.append(np.asarray(img / 255, dtype=np.float32))
with open(anno_fp, 'r') as f:
anno = json.load(f)
box_set = []
for class_names, boxes in anno.items():
box_set.extend([box + [class_names.index(class_names)] for box in boxes])
ground_truth_box_sets.append(np.asarray(box_set, dtype=np.float32))
return [class_names], images, ground_truth_box_sets, num_samples
def read_train_data_sets(data_dir, load_rate=1.0):
class_names, train_images, train_gt_box_sets, data_size = read_data(data_dir + 'train', load_rate)
train_size = int(data_size * (1 - 0.2))
if train_size <= 0:
print('No data to read')
return base.Datasets(train=None, validation=None, test=None)
validation_images = np.array(train_images[train_size:])
validation_gt_box_sets = np.array(train_gt_box_sets[train_size:])
train_images = np.array(train_images[:train_size])
train_gt_box_sets = np.array(train_gt_box_sets[:train_size])
train = DetectionDataset('train', class_names, train_images, train_gt_box_sets)
validation = DetectionDataset('validation', class_names, validation_images, validation_gt_box_sets)
return base.Datasets(train=train, validation=validation, test=None)
def read_test_data_sets(data_dir, load_rate=1.0):
class_names, test_images, test_gt_box_sets, data_size = read_data(data_dir + 'test', load_rate)
try:
test_images = np.array(test_images)
except:
print('No data to read')
return base.Datasets(train=None, validation=None, test=None)
test_gt_box_sets = np.array(test_gt_box_sets)
test = DetectionDataset('test', class_names, test_images, test_gt_box_sets)
return base.Datasets(train=None, validation=None, test=test)
def read_data_sets(data_dir, load_rate=1.0):
class_names, train_images, train_gt_box_sets, data_size = read_data(data_dir + 'train', load_rate)
train_size = int(data_size * (1 - 0.2))
if train_size <= 0:
print('No data to read')
return base.Datasets(train=None, validation=None, test=None)
validation_images = np.array(train_images[train_size:])
validation_gt_box_sets = np.array(train_gt_box_sets[train_size:])
train_images = np.array(train_images[:train_size])
train_gt_box_sets = np.array(train_gt_box_sets[:train_size])
class_names, test_images, test_gt_box_sets, data_size = read_data(data_dir + 'test', load_rate)
try:
test_images = np.array(test_images)
except:
print('No data to read')
return base.Datasets(train=None, validation=None, test=None)
test_gt_box_sets = np.array(test_gt_box_sets)
test = DetectionDataset('test', class_names, test_images, test_gt_box_sets)
train = DetectionDataset('train', class_names, train_images, train_gt_box_sets)
validation = DetectionDataset('validation', class_names, validation_images, validation_gt_box_sets)
test = DetectionDataset('test', class_names, test_images, test_gt_box_sets)
return base.Datasets(train=train, validation=validation, test=test)
# + id="oCnFBCv_tndZ" colab_type="code" colab={} outputId="79cc8e02-0318-415c-f7d1-0b0a50fb1875"
loadtest1 = read_train_data_sets('./FDDB/', load_rate=0.2)
loadtest2 = read_test_data_sets('./FDDB/', load_rate=0.2)
# + [markdown] id="YSsljZvXtndc" colab_type="text"
# # 1. Train
# + [markdown] id="xGrUUE7rtndd" colab_type="text"
# <br />
# ## Global Variables
# + id="FI2p0YQVtnde" colab_type="code" colab={}
is_train = True
# + [markdown] id="q90Y5LqHtndg" colab_type="text"
# <br />
# # 1.1 Load Data
# + id="kfiZ-BTttndh" colab_type="code" colab={} outputId="29ccb881-74f0-4f9f-d3ba-a13cd006d496"
data1 = read_train_data_sets('../test_data/FDDB/', load_rate=0.2)
data2 = read_test_data_sets('../test_data/FDDB/', load_rate=1.0)
data_train = data1.train
data_test = data2.test
num_classes = data_train.num_classes
# + id="oFHG3Uxotndl" colab_type="code" colab={}
tf_input_image = tf.placeholder(tf.float32, shape=[1, None, None, 3])
tf_gt_boxes = tf.placeholder(tf.float32, shape=[1, None, 5])
# + [markdown] id="ZrVA_L5otndn" colab_type="text"
# <br />
# # 1.2 Hyperparameters
# + id="OKJeaqG6tndo" colab_type="code" colab={}
class ModelHyperParameter():
#----------------------------------------------------------------------
# Train
#----------------------------------------------------------------------
# Training loop
batch_size = 1
num_epochs = 200
if is_train:
num_batches_per_epoch = data_train.num_samples // batch_size
#----------------------------------------------------------------------
# Train Backpropagation: Gradient descent optimization
#----------------------------------------------------------------------
# momentum
momentum = 0.9
# decaying learning rate
init_learning_rate = 0.01 # default learning rate (epsilon)
patience_of_no_improvement_epochs = 30
learning_rate_decay = 0.1
lower_bound_learning_rate = 1e-8
current_learning_rate_value = 0.001
#----------------------------------------------------------------------
# Validation & Testing
#----------------------------------------------------------------------
# Validation & Testing
num_evaluations = 500 # 100
# Evaluator: check is better score
is_better_score_threshold = 1e-4
#----------------------------------------------------------------------
# RPN Model
#----------------------------------------------------------------------
rpn_channels = 512
# stride for images
# 이미지 내 gride 계산 시, 한 grid에 total stride = 16 pixel 씩 되도록 등분함
total_stride = 16
# Anchor box generation
anchor_base_size = 16 # initial size = 16
anchor_scales = [8, 16, 24] # 2 scales x 3 ratios = 6 anchor boxes for each grid
anchor_ratios = [0.5, 1, 2]
num_anchors_per_grid = 9 # |anchor_scales| x |anchor_ratios|
# Anchor box's y true targeting & batch sampling
anchor_positive_rate = 0.5
anchor_batch_size = 128
anchor_positive_threshold = 0.7
anchor_negative_threshold = 0.3
# NMS (non maximum suppression)
nms_top_k = 2000
nms_iou_threshold = 0.7
#----------------------------------------------------------------------
# FRCNN Model
#----------------------------------------------------------------------
# Proposal box's y true targeting & batch sampling
proposal_positive_rate = 0.25
proposal_batch_size = 128
proposal_positive_threshold = 0.7
proposal_negative_threshold = 0.5
# ROI pooling
# from Fast R-CNN, speical version of spatial pyramid pooling from SPP-Net)
# R-CNN -> SPP-Net -> Fast R-CNN -> Faster R-CNN
roi_pool_size = 7
_hp = ModelHyperParameter()
# + [markdown] id="CW32VfqKtndp" colab_type="text"
# <br />
# # 1.3 Build ConvNet Model (VGG16)
# + id="7IOZMknHtndq" colab_type="code" colab={}
info = '' # log printing variable
# + [markdown] id="Z2vjmICQtndr" colab_type="text"
# #### layer building functions
# + id="vhQYCb5Rtnds" colab_type="code" colab={}
def input_feature_extraction_only(input):
x = input
global info
info = ' Inputs'
info += '\n {:12s}: {:17s} {}'.format('x', str(x.shape), 'input images')
return x
def output_labels(out_channel):
y = tf.placeholder(tf.float32, [None, out_channel])
global info
info += '\n {:12s}: {:17s} {}\n\n Feature Extraction'.format('y', str(y.shape), 'target value (answer label)')
return y
# + id="yZlVCT2Etndu" colab_type="code" colab={}
def conv(name, inputs, filter_size, stride, num_filters, padding='SAME', is_print=True):
bias=0.0
in_channel = int(inputs.get_shape()[-1])
out_channel = num_filters
weights = tf.get_variable(name=name + '_weights',
shape=[filter_size, filter_size, in_channel, out_channel],
dtype=tf.float32,
initializer=tf.contrib.layers.xavier_initializer())
biases = tf.get_variable(name + '_biases',
[out_channel], tf.float32,
tf.constant_initializer(value=bias))
conv = tf.nn.conv2d(inputs, weights,
strides=[1, stride, stride, 1],
padding=padding) + bias
conv = tf.nn.relu(conv) # activation (non-linearizing)
if is_print:
global info
# skip if this is an inception layer's sub layer
info += '\n {:12s}: {:17s} -> {:17s}'.format(
name, str(inputs.shape), str(conv.shape) )
return conv
# + id="u7HIyEB4tndv" colab_type="code" colab={}
def pool(name, inputs, filter_size, stride, padding='SAME', is_print=True):
pool = tf.nn.max_pool(inputs,
ksize = [1, filter_size, filter_size, 1],
strides = [1,stride,stride,1], padding=padding)
if is_print:
global info
# skip if this is an inception layer's sub layer
info += '\n {:12s}: {:17s} -> {:17s}'.format(
name, str(inputs.shape), str(pool.shape) )
return pool
# + id="4MC20XBZtndx" colab_type="code" colab={}
def fc(name, inputs, output_size, is_print=True):
bias=0.1
in_dim = int(inputs.get_shape()[-1])
out_dim = output_size
weights = tf.get_variable(name=name + '_weights',
shape=[in_dim, out_dim],
initializer=tf.contrib.layers.xavier_initializer())
biases = tf.get_variable(name + '_biases',
[out_dim], tf.float32,
tf.constant_initializer(value=bias))
fc = tf.matmul(inputs, weights) + biases
fc = tf.nn.relu(fc) # activation
# The probability of keeping each unit for dropout layers
keep_prob_value = tf.cond(tf.cast(is_train, tf.bool),
lambda: train_dropout_rate,
lambda: 1.0)
fc = tf.nn.dropout(fc, keep_prob=keep_prob_value)
if is_print:
global info
info += '\n {:12s}: {:17s} -> {:17s}'.format(
name, str(inputs.shape), str(fc.shape) )
return fc
# + id="rhPNLhPqtndy" colab_type="code" colab={}
def fc_last(name, inputs, output_size, is_print=True):
bias=0.0
in_dim = int(inputs.get_shape()[-1])
out_dim = output_size
weights = tf.get_variable(name=name + '_weights',
shape=[in_dim, out_dim],
initializer=tf.contrib.layers.xavier_initializer())
biases = tf.get_variable(name + '_biases',
[out_dim], tf.float32,
tf.constant_initializer(value=bias))
logits = tf.matmul(inputs, weights) + biases
if is_print:
global info
info += '\n {:12s}: {:17s} -> {:17s}'.format(
name, str(inputs.shape), str(logits.shape) )
return logits
# + id="dfsM-u6Dtndy" colab_type="code" colab={}
def logits_to_softmax(name, inputs):
# hypothesis (prediction) of target value y
y_hat = tf.nn.softmax(inputs)
global info
info += '\n\n Output\n {:12s}: {:8s}hypothesis (prediction) of target value y'.format(name, str(y_hat.shape))
return y_hat
# + [markdown] id="92hmdqEVtnd0" colab_type="text"
# #### network layers
# + id="xV4KXIQOtnd0" colab_type="code" colab={}
class ConvModel:
# input
x = None
y = None
# feature extraction
conv1 = None
conv2 = None
pool2 = None
conv3 = None
conv4 = None
pool4 = None
conv5 = None
conv6 = None
conv7 = None
pool7 = None
conv8 = None
conv9 = None
conv10 = None
pool10 = None
conv11 = None
conv12 = None
conv13 = None
pool13 = None
output_feature_map = None
#flat = None
# classification
#fc14 = None
#fc15 = None
#logits = None
# hypothesis (prediction) of target value y
#y_prediction = None
m_conv = ConvModel()
# + id="jCwwDTSmtnd1" colab_type="code" colab={}
tf_input_image = tf.placeholder(tf.float32, shape=[1, None, None, 3])
m_conv.input_image = input_feature_extraction_only(tf_input_image)
m_conv.conv1 = conv('conv1' , m_conv.input_image, 3, 1, 64 )
m_conv.conv2 = conv('conv2' , m_conv.conv1 , 3, 1, 64 )
m_conv.pool2 = pool('pool2' , m_conv.conv2 , 2, 2)
m_conv.conv3 = conv('conv3' , m_conv.pool2 , 3, 1, 128)
m_conv.conv4 = conv('conv4' , m_conv.conv3 , 3, 1, 128)
m_conv.pool4 = pool('pool4' , m_conv.conv4 , 2, 2)
m_conv.conv5 = conv('conv5' , m_conv.pool4 , 3, 1, 256)
m_conv.conv6 = conv('conv6' , m_conv.conv5 , 3, 1, 256)
m_conv.conv7 = conv('conv7' , m_conv.conv6 , 3, 1, 256)
m_conv.pool7 = pool('pool7' , m_conv.conv7 , 2, 2)
m_conv.conv8 = conv('conv8' , m_conv.pool7 , 3, 1, 512)
m_conv.conv9 = conv('conv9' , m_conv.conv8 , 3, 1, 512)
m_conv.conv10 = conv('conv10', m_conv.conv9 , 3, 1, 512)
m_conv.pool10 = pool('pool10', m_conv.conv10, 2, 2)
m_conv.conv11 = conv('conv11', m_conv.pool10, 3, 1, 512)
m_conv.conv12 = conv('conv12', m_conv.conv11, 3, 1, 512)
m_conv.conv13 = conv('conv13', m_conv.conv12, 3, 1, 512)
m_conv.output_feature_map = m_conv.conv13
# + id="DX8IsfWBtnd2" colab_type="code" colab={} outputId="e2d70124-0fa7-4c65-cb92-542bfa5a7f41"
print(info)
# + [markdown] id="4DsEmPUHtnd3" colab_type="text"
# <br />
# # 1.4 Build RPN Model
# + [markdown] id="auzfBGE4tnd4" colab_type="text"
# ### 1.4.1 RPN Preprocessing 1 - Anchorbox Generation
# + id="XuUZMDnmtnd4" colab_type="code" colab={}
current_image_h = 0
current_image_w = 0
num_grids_h = 0
num_grids_w = 0
num_grids = 0
# + id="V0jOCZ8Dtnd6" colab_type="code" colab={}
def anchorbox_generation():
input_image = tf_input_image
image_shape = tf.shape(input_image)
global current_image_h
global current_image_w
global num_grids_h
global num_grids_w
global num_grids
# Calculate the number of grid cells.
current_image_h = image_shape[1]
current_image_w = image_shape[2]
# Calculate the number of grid cells.
num_grids_h = tf.to_int32(tf.ceil(current_image_h / np.float32(_hp.total_stride)))
num_grids_w = tf.to_int32(tf.ceil(current_image_w / np.float32(_hp.total_stride)))
num_grids = num_grids_h * num_grids_w
# Create a base anchor.
base_anchor = np.array([0, 0, _hp.anchor_base_size - 1, _hp.anchor_base_size - 1]) # [4]
# Expand anchors by ratios.
anchors = expand_anchors_by_ratios(base_anchor, _hp.anchor_ratios) # [num_ratios, 4]
# Expand anchors by scales.
anchors = np.vstack( # [num_ratios * num_scales, 4]
[expand_anchors_by_scales(anchors[i, :], _hp.anchor_scales) for i in range(anchors.shape[0])]
)
# Expand anchors by shifts.
_hp.num_anchors_per_grid = anchors.shape[0]
anchors = tf.constant(anchors.reshape((1, _hp.num_anchors_per_grid, 4)), dtype=tf.int32) # [1, num_anchors_per_grid, 4]
# [1, num_anchors_per_grid, 4]
shift_x, shift_y = tf.meshgrid(
tf.range(num_grids_w) * _hp.total_stride,
tf.range(num_grids_h) * _hp.total_stride
)
shift_x, shift_y = tf.reshape(shift_x, (-1,)), tf.reshape(shift_y, (-1,)) # [num_grids], [num_grids]
shifts = tf.transpose(tf.stack([shift_x, shift_y, shift_x, shift_y])) # [num_grids, 4]
shifts = tf.transpose(tf.reshape(shifts, shape=[1, num_grids, 4]), perm=(1, 0, 2)) # [num_grids, 1, 4]
anchors = tf.add(anchors, shifts) # [num_grids, num_anchors_per_grid, 4]
anchors = tf.reshape(anchors, (-1, 4)) # [num_anchors, 4], where num_grids * num_anchors_per_grid = num_anchors
return tf.cast(anchors, dtype=tf.float32)
def expand_anchors_by_scales(anchor, scales):
"""
Enumerate a set of anchors for each scale wrt an anchor.
"""
scales = np.array(scales)
w, h, x_ctr, y_ctr = get_whxy_format(anchor)
ws = w * scales
hs = h * scales
anchors = get_anchor_format(ws, hs, x_ctr, y_ctr)
return anchors
def expand_anchors_by_ratios(anchor, ratios):
"""
Enumerate a set of anchors for each aspect ratio wrt an anchor.
"""
ratios = np.array(ratios)
w, h, x_ctr, y_ctr = get_whxy_format(anchor)
size = w * h
size_ratios = size / ratios
ws = np.round(np.sqrt(size_ratios))
hs = np.round(ws * ratios)
anchors = get_anchor_format(ws, hs, x_ctr, y_ctr)
return anchors
def get_whxy_format(anchor):
"""
Return width, height, x center, and y center for an anchor (window).
"""
w = anchor[2] - anchor[0] + 1
h = anchor[3] - anchor[1] + 1
x_ctr = anchor[0] + 0.5 * (w - 1)
y_ctr = anchor[1] + 0.5 * (h - 1)
return w, h, x_ctr, y_ctr
def get_anchor_format(ws, hs, x_ctr, y_ctr):
"""
Given a vector of widths (ws) and heights (hs) around a center (x_ctr, y_ctr), output a set of anchors (windows).
"""
ws = ws[:, np.newaxis]
hs = hs[:, np.newaxis]
anchors = np.hstack(
(x_ctr - 0.5 * (ws - 1), y_ctr - 0.5 * (hs - 1), x_ctr + 0.5 * (ws - 1), y_ctr + 0.5 * (hs - 1))
)
return anchors
# + id="BHXTt5wUtnd6" colab_type="code" colab={}
init_anchor_boxes = anchorbox_generation()
# + [markdown] id="r8Ne-Arbtnd7" colab_type="text"
# #### (test) visualization anchor box
# + id="jTGXrTfttnd8" colab_type="code" colab={}
graph = tf.get_default_graph()
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(graph=graph, config=config)
sess.run(tf.global_variables_initializer()) # initialize all weights
# + id="EIS7CU1Ttnd8" colab_type="code" colab={}
batch_image, batch_gt_boxes = data_train.get_batch(idx=3)
# + id="z_TcjMQ-tnd9" colab_type="code" colab={}
anchor_boxes, image_h, image_w, n_grids_h, n_grids_w, n_grids = sess.run([
init_anchor_boxes,
current_image_h, current_image_w,
num_grids_h, num_grids_w, num_grids],
feed_dict={tf_input_image: batch_image})
# + id="N_m9xElptnd-" colab_type="code" colab={} outputId="fb22ce18-1e8e-4e41-a93a-9fd1a9624e81"
print('Image shape : {} x {}'.format(image_h, image_w))
print('Total stride : {}'.format(_hp.total_stride))
print('H grids : {} / {} = {}'.format(image_h, _hp.total_stride, n_grids_h))
print('W grids : {} / {} = {}'.format(image_w, _hp.total_stride, n_grids_w))
print('Number of grids : {} x {} = {}'.format(n_grids_h, n_grids_w, n_grids))
print('Number of anchors: {} x {} = {}'.format(n_grids, _hp.num_anchors_per_grid, anchor_boxes.shape[0]))
# + [markdown] id="RoteHr92tnd_" colab_type="text"
# # anchor boxes for one grid
# + id="RofiTCyVtneA" colab_type="code" colab={} outputId="3c7b841b-98cc-4193-fba0-2d03798c2c9c"
# %matplotlib inline
import matplotlib.pyplot as plt
import cv2
# FIXME
MARGIN = 200
GRID_IDX = (5, 5)
SHOW_ALL = False
PRINT_BOX_INFO = False
K = len(_hp.anchor_scales) * len(_hp.anchor_ratios)
img_h, img_w = batch_image.shape[1:3]
img_with_margin = np.zeros((img_h + MARGIN * 2, img_w + MARGIN * 2, 3)).astype(np.uint8)
img_with_margin[
MARGIN:MARGIN + img_h, MARGIN:MARGIN + img_w, :
] = np.asarray(batch_image[0] * 255, dtype=np.uint8)
for idx, box in enumerate(anchor_boxes):
if not SHOW_ALL and idx // K != GRID_IDX[0] * n_grids_w + GRID_IDX[1]:
continue
box = box + MARGIN
cv2.rectangle(img_with_margin, (box[0], box[1]), (box[2], box[3]), color=(255, 255, 0), thickness=2)
if PRINT_BOX_INFO:
print('[Box # {}] Shape: ({:3}, {:3}) -> Size: {:6d}, Scale: {:.3f}'.format(
idx % K, int(box[2] - box[0]), int(box[3] - box[1]),
int((box[2] - box[0]) * (box[3] - box[1])), (box[2] - box[0]) / (box[3] - box[1])
))
plt.figure(figsize=(16, 16))
plt.title('Anchor boxes at {} grid cell'.format(GRID_IDX), fontdict={'fontsize': 16})
plt.imshow(img_with_margin)
plt.show()
# + [markdown] id="H5hh19kRtneB" colab_type="text"
# all anchor boxes
# + id="AiIEbGyVtneB" colab_type="code" colab={} outputId="951c54a3-d0ff-439e-b1d9-1b1797f141bd"
# %matplotlib inline
import matplotlib.pyplot as plt
import cv2
# FIXME
MARGIN = 200
GRID_IDX = (5, 5)
SHOW_ALL = True
PRINT_BOX_INFO = False
K = len(_hp.anchor_scales) * len(_hp.anchor_ratios)
img_h, img_w = batch_image.shape[1:3]
img_with_margin = np.zeros((img_h + MARGIN * 2, img_w + MARGIN * 2, 3)).astype(np.uint8)
img_with_margin[
MARGIN:MARGIN + img_h, MARGIN:MARGIN + img_w, :
] = np.asarray(batch_image[0] * 255, dtype=np.uint8)
for idx, box in enumerate(anchor_boxes):
if not SHOW_ALL and idx // K != GRID_IDX[0] * n_grids_w + GRID_IDX[1]:
continue
box = box + MARGIN
cv2.rectangle(img_with_margin, (box[0], box[1]), (box[2], box[3]), color=(255, 255, 0), thickness=2)
if PRINT_BOX_INFO:
print('[Box # {}] Shape: ({:3}, {:3}) -> Size: {:6d}, Scale: {:.3f}'.format(
idx % K, int(box[2] - box[0]), int(box[3] - box[1]),
int((box[2] - box[0]) * (box[3] - box[1])), (box[2] - box[0]) / (box[3] - box[1])
))
plt.figure(figsize=(16, 16))
plt.title('Anchor boxes at {} grid cell'.format(GRID_IDX), fontdict={'fontsize': 16})
plt.imshow(img_with_margin)
plt.show()
# + [markdown] id="ax1bwrF8tneD" colab_type="text"
# ### 1.4.2 RPN Preprocessing 2 - Target Value Generation
# + id="cjcFjFKNtneD" colab_type="code" colab={}
from cython_bbox import bbox_overlaps # 두 박스 간 IOU 계산
# + id="ESs4uFd7tneE" colab_type="code" colab={}
def target_value_generation(init_anchor_boxes, gt_boxes, current_image_h, current_image_w, num_grids_h, num_grids_w, num_grids):
gt_boxes = gt_boxes[0]
anchor_boxes = init_anchor_boxes
num_init_anchor_boxes = anchor_boxes.shape[0]
# 이미지 밖으로 넘어나간 anchor box는 없애
indices_within_image_boundary = np.where(
(anchor_boxes[:, 0] >= 0) &
(anchor_boxes[:, 1] >= 0) &
(anchor_boxes[:, 2] < current_image_w) &
(anchor_boxes[:, 3] < current_image_h))[0]
anchor_boxes = anchor_boxes[indices_within_image_boundary, :]
num_anchor_boxes = len(indices_within_image_boundary)
# anchor box, gt box 간 모든 combination 에 대해 IOU 계산
# 결과 matrix 는 M x N 행렬 (M = ancchor box size, N = gt box size)
iou_matrix = bbox_overlaps( # [num_anchor_boxes, num_gt_boxes]-D matrix
np.ascontiguousarray(anchor_boxes, dtype=np.float64),
np.ascontiguousarray(gt_boxes, dtype=np.float64) # (np.ascontiguousarray 는 성능 향상용, 배열 최적화)
)
# 각각의 anchor box 마다 가장 값이 큰 IOU 를 갖는 gt box를 찾는다.
# for i = 1 ~ |anchor boxes|
# abox_max_overlaps[i] = 0
# for j = 1 ~ |gt boxes|
# if abox_max_overlaps < iou_matrix[i][j]:
# abox_max_overlaps[i] = iou_matrix[i][j] 가장 큰 값 갖는 gt box와의 IOU 값
# abox_max_overlaps_indices[i] = j 그 gt box 의 위치
row_wise = 1 # 행단위
abox_max_overlaps_indices = iou_matrix.argmax(axis=row_wise) # [num_anchor_boxes]
abox_max_overlaps = iou_matrix.max(axis=row_wise) # [num_anchor_boxes]
# 이제 gt box 기준, gt box 마다 가장 가까운 anchor box를 찾는다.
column_wise = 0
gtbox_max_overlaps_indices = iou_matrix.argmax(axis=column_wise) # [num_gt_boxes]
gtbox_max_overlaps = iou_matrix.max(axis=column_wise) # [num_gt_boxes]
# create anchor boxes' label matrix N x N (N = num_anchor_boxes)
label_matrix = np.empty((num_anchor_boxes,), dtype=np.float32)
ignore = -1
positive = 1
negative = 0
label_matrix.fill(ignore) # Default to be ignored
label_matrix[abox_max_overlaps < _hp.anchor_negative_threshold] = negative # IOU < 0.3
label_matrix[abox_max_overlaps >= _hp.anchor_positive_threshold] = positive # IOU >= 0.7
# Assign positive labels to anchor boxes which best matches to any GT boxes.
# gt box 에 대해서 나랑 가장 잘 맞는 애로 선택된 anchor box가 있으면
# 그 anchor box는 무조건 positive
label_matrix[gtbox_max_overlaps_indices] = positive
# Subsample the positive samples if we have too many.
# |negative| + |positive| > batch size 이면 안돼
# positive ratio = 0.5
# batch size = 128 이면
# |posi| = 64, |nega| = 128 -64 되도록 dropout 처리해줘야
num_positive_bound = int(_hp.anchor_positive_rate * _hp.anchor_batch_size)
positive_indices = np.where(label_matrix == positive)[0]
if len(positive_indices) > num_positive_bound:
# sampling 된 anchor box 중 positive (예 90개) 개수가
# num_positive_bound (예 64개) 넘었으면
# 90 -64 = 26개만큼 random sampling 해서 ignore (-1)로 변경
drop_out_indices = npr.choice(positive_indices,
size=(len(positive_indices) - num_positive_bound),
replace=False)
label_matrix[drop_out_indices] = ignore
# Subsampe the negative samples if we have too many.
# negative 경우도 마찬가지로 처리
num_negative_bound = _hp.anchor_batch_size - np.sum(label_matrix == positive)
negative_indices = np.where(label_matrix == negative)[0]
if len(negative_indices) > num_negative_bound:
drop_out_indices = npr.choice(negative_indices,
size=(len(negative_indices) - num_negative_bound),
replace=False)
label_matrix[drop_out_indices] = ignore
# classification 을 위한 positive, negative anchor box 선정은 이제 끝
# regression 은 positive 에 대해 얼만큼 옮겨야 gt box로 갈지도 정해줘야
# t parameter 계산
# RPN sub func[2] compute_bbox_deltas 함수 사용
# Compute bbox deltas of all the inside anchor boxes.
# anchor box 가 gtbox 로 가려면 얼마나 움직여야하는지 계산
# positive 뿐 아니라 걍 다 집어넣음 -> 나중에 positive 인 anchor box index 로 selection 해서 사용
bbox_targets = compute_bbox_deltas(anchor_boxes, gt_boxes[abox_max_overlaps_indices, :])
# Map up to original set of anchor boxes (only inside anchor boxes -> all anchor boxes)
labels = unmap_data(label_matrix,
num_init_anchor_boxes,
indices_within_image_boundary, fill=-1)
bbox_targets = unmap_data(bbox_targets,
num_init_anchor_boxes,
indices_within_image_boundary, fill=0)
# Reshape tensors into a grid cell form.
labels = labels.reshape((1, num_grids_h, num_grids_w, _hp.num_anchors_per_grid))
bbox_targets = bbox_targets.reshape((1, num_grids_h, num_grids_w, _hp.num_anchors_per_grid * 4))
return labels, bbox_targets
def compute_bbox_deltas(src_bboxes, dst_bboxes):
src_widths = src_bboxes[:, 2] - src_bboxes[:, 0] + 1.0
src_heights = src_bboxes[:, 3] - src_bboxes[:, 1] + 1.0
src_ctr_x = src_bboxes[:, 0] + 0.5 * src_widths
src_ctr_y = src_bboxes[:, 1] + 0.5 * src_heights
dst_widths = dst_bboxes[:, 2] - dst_bboxes[:, 0] + 1.0
dst_heights = dst_bboxes[:, 3] - dst_bboxes[:, 1] + 1.0
dst_ctr_x = dst_bboxes[:, 0] + 0.5 * dst_widths
dst_ctr_y = dst_bboxes[:, 1] + 0.5 * dst_heights
targets_dx = (dst_ctr_x - src_ctr_x) / src_widths
targets_dy = (dst_ctr_y - src_ctr_y) / src_heights
targets_dw = np.log(dst_widths / src_widths)
targets_dh = np.log(dst_heights / src_heights)
targets = np.vstack((targets_dx, targets_dy, targets_dw, targets_dh)).transpose()
return targets
def unmap_data(data, count, indices, fill=0):
"""Unmap a subset of item (data) back to the original set of items (of size count)
"""
if len(data.shape) == 1:
ret = np.empty((count,), dtype=np.float32)
ret.fill(fill)
ret[indices] = data
else:
ret = np.empty((count,) + data.shape[1:], dtype=np.float32)
ret.fill(fill)
ret[indices, :] = data
return ret
# + id="hqomKTLNtneF" colab_type="code" colab={}
tf_gt_boxes = tf.placeholder(tf.float32, shape=[1, None, 5])
target_labels, target_t_parameters = tf.py_func(
target_value_generation,
[init_anchor_boxes,
tf_gt_boxes,
current_image_h,
current_image_w,
num_grids_h,
num_grids_w,
num_grids],
[tf.float32, tf.float32])
# + id="CxmKNYCRtneF" colab_type="code" colab={} outputId="45f8d5f9-8905-48e3-ee79-189e68764af4"
print(target_labels)
print(target_t_parameters)
# + [markdown] id="x4P7Ei3UtneH" colab_type="text"
# #### (test) check target value generation
# + id="mCmyglyYtneH" colab_type="code" colab={}
graph = tf.get_default_graph()
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(graph=graph, config=config)
sess.run(tf.global_variables_initializer()) # initialize all weights
# + id="_F1wjJvitneI" colab_type="code" colab={}
batch_image, batch_gt_boxes = data_train.get_batch(idx=3)
# + id="WYuHElmltneI" colab_type="code" colab={}
rpn_target_labels, rpn_target_t_parameters = sess.run([
target_labels,
target_t_parameters],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="6cLp6Cq6tneJ" colab_type="code" colab={} outputId="aa3fab79-bed4-418a-b995-0c0fc52a3c15"
print('RPN target labels : {}'.format(rpn_target_labels.shape))
print('RPN target t params: {}'.format(rpn_target_t_parameters.shape))
print('Batch size: {:4}'.format(_hp.anchor_batch_size))
print(' Positive: {:4}'.format(np.count_nonzero(rpn_target_labels == 1)))
print(' Negative: {:4}'.format(np.count_nonzero(rpn_target_labels == 0)))
print(' Ignored : {:4}'.format(np.count_nonzero(rpn_target_labels == -1)))
# + [markdown] id="lvHflmAjtneK" colab_type="text"
# ### 1.4.3 Build RPN Model
# + id="OCKuo7EktneK" colab_type="code" colab={}
class RPN_Model:
# y_true (target value)
classification_y_true = None
regression_y_true = None
# rpn network layers
input_layer = None # layer 1 (conv feature)
intermediate_layer = None # layer 2
classification_logits = None # layer 3-1 (object or bg)
classification_probabilities = None
classification_y_predictions = None
regression_y_predictions = None # layer 3-2 (t parameters)
# rpn train output
proposal_boxes = None
proposal_scores = None
m_rpn = RPN_Model()
# + [markdown] id="sYNwFnMMtneL" colab_type="text"
# #### target values
# + id="KONDCK_JtneL" colab_type="code" colab={}
m_rpn.classification_y_true = target_labels
m_rpn.regression_y_true = target_t_parameters
# + [markdown] id="sl5WOTW6tneM" colab_type="text"
# #### input layer
# + id="4zWUZHt1tneM" colab_type="code" colab={}
m_rpn.input_layer = m_conv.output_feature_map
# + [markdown] id="91xlyBnOtneN" colab_type="text"
# #### intermediate layer
# + id="fRfEofZWtneN" colab_type="code" colab={}
import tensorflow.contrib.slim as slim
m_rpn.intermediate_layer = slim.conv2d(
inputs=m_rpn.input_layer,
num_outputs=_hp.rpn_channels,
kernel_size=[3, 3], # 1 stride, 1 padding
trainable=is_train)
# + [markdown] id="2hKo5asntneO" colab_type="text"
# #### output layers
# + id="XmiAVDpCtneO" colab_type="code" colab={}
def classification_layer(inputs):
scores = slim.conv2d(
inputs=inputs, # [H/16, W/16, 2 * num_anchors_per_grid]
num_outputs=_hp.num_anchors_per_grid * 2, # output 2 K = is object [T/F] per |anchors| of a grid
kernel_size=[1, 1], # 1 x 1 filter, 1 stride, 0 padding
trainable=is_train,
weights_initializer=tf.random_normal_initializer(mean=0.0, stddev=0.01),
weights_regularizer=tf.contrib.layers.l2_regularizer(0.0001),
padding='VALID', activation_fn=None)
probs = tf.reshape(tf.nn.softmax(tf.reshape(scores, (-1, 2))), tf.shape(scores))
y_predictions = tf.argmax(tf.reshape(probs, (-1, 2)), axis=1)
m_rpn.classification_logits = scores
m_rpn.classification_probabilities = probs
m_rpn.classification_y_predictions = y_predictions
# + id="uaCowOeHtneP" colab_type="code" colab={}
def regression_layer(inputs):
t_parameters_from_anchorbox_to_gtbox = slim.conv2d(
inputs=inputs,
num_outputs=_hp.num_anchors_per_grid * 4, # output 4 K = [tx, ty, tw, th] per |anchors| of a grid
kernel_size=[1, 1], # 1 x 1 filter, 1 stride, 0 padding
trainable=is_train,
weights_initializer=tf.random_normal_initializer(mean=0.0, stddev=0.01),
weights_regularizer=tf.contrib.layers.l2_regularizer(0.0001),
padding='VALID', activation_fn=None)
m_rpn.regression_y_predictions = t_parameters_from_anchorbox_to_gtbox
# + id="UXfLR2zLtneQ" colab_type="code" colab={}
classification_layer(m_rpn.intermediate_layer)
regression_layer(m_rpn.intermediate_layer)
# + id="L9JQ_2CktneQ" colab_type="code" colab={} outputId="47c2f1d9-5a25-463b-db59-21595b6ba2b9"
print(m_rpn.input_layer.shape)
print(m_rpn.intermediate_layer.shape)
print(m_rpn.classification_logits.shape) # |anchor box| 9 x |object or not| 2 = 18
print(m_rpn.regression_y_predictions.shape) # |ancho box| 9 x |t param| 4 = 36
# + id="fGvP7xjytneR" colab_type="code" colab={}
sess.run(tf.global_variables_initializer())
conv_features, rpn_intermediate_layer = sess.run(
[m_conv.output_feature_map,
m_rpn.intermediate_layer],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="M62omSyVtneS" colab_type="code" colab={} outputId="f360c2bc-8780-4c83-f672-8e3a24239151"
print('Image shape: {}'.format(batch_image.shape))
print('Total stride: {}'.format(_hp.total_stride))
print('conv_feats shape: {}'.format(conv_features.shape))
print('rpn_intermediate shape: {}'.format(rpn_intermediate_layer.shape))
# + id="zBvTi5c8tneT" colab_type="code" colab={} outputId="909fa838-4ffa-44a2-ee8b-e9831a64463b"
print(m_rpn.classification_logits.shape)
print(m_rpn.regression_y_predictions.shape)
# + id="2P1KfO_PtneV" colab_type="code" colab={}
sess.run(tf.global_variables_initializer())
rpn_class_scores, rpn_t_parameters = sess.run(
[m_rpn.classification_logits,
m_rpn.regression_y_predictions],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="Tz4I6MtvtneV" colab_type="code" colab={} outputId="1174ebfc-6f4b-4de0-b1d6-573ecc246edb"
print(rpn_class_scores.shape) # 2 x 9 = 18
print(rpn_t_parameters.shape) # 4 x 9 = 36
# + [markdown] id="QZQz5MXytneX" colab_type="text"
# ### 1.4.4 RPN Postprocessing - Anchorbox Transformation
#
# <br />
#
# RPN regression 학습 결과로 얻은 t parameter 로 anchor box 의 위치 이동
# + id="lBhpRyZstneX" colab_type="code" colab={}
def transform_bboxes(anchor_boxes, regression_y_predictions):
deltas = tf.reshape(regression_y_predictions, (-1, 4))
boxes = anchor_boxes
boxes = tf.cast(boxes, deltas.dtype)
# Compute size and center coordinates of the boxex.
widths = tf.subtract(boxes[:, 2], boxes[:, 0]) + 1.0
heights = tf.subtract(boxes[:, 3], boxes[:, 1]) + 1.0
ctr_x = tf.add(boxes[:, 0], widths * 0.5)
ctr_y = tf.add(boxes[:, 1], heights * 0.5)
tx, ty, tw, th = deltas[:, 0], deltas[:, 1], deltas[:, 2], deltas[:, 3]
transformed_ctr_x = tf.add(tf.multiply(tx, widths), ctr_x)
transformed_ctr_y = tf.add(tf.multiply(ty, heights), ctr_y)
transformed_w = tf.multiply(tf.exp(tw), widths)
transformed_h = tf.multiply(tf.exp(th), heights)
transformed_xmin = tf.subtract(transformed_ctr_x, transformed_w * 0.5)
transformed_ymin = tf.subtract(transformed_ctr_y, transformed_h * 0.5)
transformed_xmax = tf.add(transformed_ctr_x, transformed_w * 0.5)
transformed_ymax = tf.add(transformed_ctr_y, transformed_h * 0.5)
return tf.stack([transformed_xmin, transformed_ymin, transformed_xmax, transformed_ymax], axis=1)
# + id="26gpFUgItneY" colab_type="code" colab={}
transformed_anchor_boxes = transform_bboxes(init_anchor_boxes, m_rpn.regression_y_predictions)
# + id="BfNyzRvwtneY" colab_type="code" colab={}
def clip_bboxes(input_image, boxes):
img_shape = tf.shape(input_image)
img_shape = tf.to_float(img_shape)
b0 = tf.maximum(tf.minimum(boxes[:, 0], img_shape[2] - 1), 0)
b1 = tf.maximum(tf.minimum(boxes[:, 1], img_shape[1] - 1), 0)
b2 = tf.maximum(tf.minimum(boxes[:, 2], img_shape[2] - 1), 0)
b3 = tf.maximum(tf.minimum(boxes[:, 3], img_shape[1] - 1), 0)
return tf.stack([b0, b1, b2, b3], axis=1)
# + id="ImtwtRSPtneZ" colab_type="code" colab={}
clipped_anchor_boxes = clip_bboxes(tf_input_image, transformed_anchor_boxes)
# + id="gA16tjmttneZ" colab_type="code" colab={}
def nms(cliped_boxes, class_probs):
K = _hp.num_anchors_per_grid
object_scores = tf.reshape(tf.transpose(tf.reshape(class_probs, (-1, K, 2)), (0, 2, 1)),
tf.shape(class_probs))[..., K:]
nms_indices = tf.image.non_max_suppression(
cliped_boxes, tf.reshape(object_scores, (-1,)), max_output_size=_hp.nms_top_k,
iou_threshold=_hp.nms_iou_threshold
)
boxes = tf.gather(cliped_boxes, nms_indices)
scores = tf.gather(tf.reshape(object_scores, (-1,)), nms_indices)
return boxes, scores
# + id="SUrfzljytnea" colab_type="code" colab={}
m_rpn.proposal_boxes, m_rpn.proposal_scores = nms(clipped_anchor_boxes, m_rpn.classification_probabilities)
# + [markdown] id="YxUPsU7etneb" colab_type="text"
# #### test postporcessing
# + id="2srSug9Wtneb" colab_type="code" colab={} outputId="a479b4f1-afe3-461c-803f-0df4aeaa8156"
print(transformed_anchor_boxes.shape)
print(clipped_anchor_boxes.shape)
# + id="P74S0N0Atnec" colab_type="code" colab={}
_transformed_anchor_boxes, _clipped_anchor_boxes = sess.run(
[transformed_anchor_boxes, clipped_anchor_boxes],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="VJCYhsEytned" colab_type="code" colab={} outputId="ba355aa2-7c02-4ff7-b439-be9a3da834f5"
print(_transformed_anchor_boxes.shape)
print(_clipped_anchor_boxes.shape)
# + id="vNYfTSgZtnee" colab_type="code" colab={} outputId="2b09fd31-ea87-488c-c9d0-ae5eb89f0bf2"
print(m_rpn.proposal_boxes.shape)
print(m_rpn.proposal_scores.shape)
# + id="2zY7qZbEtnef" colab_type="code" colab={}
proposal_boxes, proposal_scores = sess.run(
[m_rpn.proposal_boxes, m_rpn.proposal_scores],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + [markdown] id="ZVeDacF4tneg" colab_type="text"
# #### RPN 네트워크의 최종 결과 (proposal boxes)
#
# 이 proposal box가 frcnn (roi pooling)의 입력값이 된다!
# + id="N9SpzmaZtneg" colab_type="code" colab={} outputId="28906c70-4f74-4aae-b6f3-19b2e013cd82"
print(proposal_boxes.shape) # dl==this value is now input of frcnn!
print(proposal_scores.shape) #
# + [markdown] id="L17ZqK3ytneh" colab_type="text"
# <br />
# # 1.5 Build RCNN Model
#
# faster rcnn 의 roi pooling 네트워크 모델은 fast rcnn의 것과 동일하다
# (RPN 부분만 다름)
#
# + [markdown] id="nxI5CVRbtneh" colab_type="text"
# ### 1.5.1 (F)RCNN Preprocessing - Target Value Generation, and Batch Sampling
# + id="zbji8biitnei" colab_type="code" colab={}
import numpy as np
import numpy.random as npr
from cython_bbox import bbox_overlaps
def rcnn_target_and_sample(proposal_boxes, proposal_scores, gt_boxes, num_object_classes):
gt_boxes = gt_boxes[0]
num_classes = num_object_classes + 1 # add 1 count for the background
num_proposal_boxes = len(proposal_boxes)
iou_matrix = bbox_overlaps(
np.ascontiguousarray(proposal_boxes, dtype=np.float64),
np.ascontiguousarray(gt_boxes, dtype=np.float64)
)
row_wise = 1 # 행단위
pbox_max_overlaps_indices = iou_matrix.argmax(axis=row_wise) # [num_proposal_boxes]
pbox_max_overlaps = iou_matrix.max(axis=row_wise) # [num_proposal_boxes]
label_matrix = np.empty((num_proposal_boxes,), dtype=np.float32)
ignore = -1
positive = 1
negative = 0
label_matrix.fill(ignore) # Default to be ignored
label_matrix[pbox_max_overlaps < _hp.proposal_negative_threshold] = negative # IOU < 0.5
label_matrix[pbox_max_overlaps >= _hp.proposal_positive_threshold] = positive # IOU >= 0.7
# Subsample the positive samples if we have too many.
# |negative| + |positive| > batch size 이면 안돼
# positive ratio = 0.5
# batch size = 128 이면
# |posi| = 64, |nega| = 128 -64 되도록 dropout 처리해줘야
num_positive_bound = int(_hp.proposal_positive_rate * _hp.proposal_batch_size)
positive_indices = np.where(label_matrix == positive)[0]
if len(positive_indices) > num_positive_bound:
# sampling 된 proposal box 중 positive (예 90개) 개수가
# num_positive_bound (예 64개) 넘었으면
# 90 -64 = 26개만큼 random sampling 해서 ignore (-1)로 변경
drop_out_indices = npr.choice(positive_indices,
size=(len(positive_indices) - num_positive_bound),
replace=False)
label_matrix[drop_out_indices] = ignore
positive_indices = np.where(label_matrix == positive)[0]
# Subsampe the negative samples if we have too many.
# negative 경우도 마찬가지로 처리
num_negative_bound = _hp.proposal_batch_size - np.sum(label_matrix == positive)
negative_indices = np.where(label_matrix == negative)[0]
if len(negative_indices) > num_negative_bound:
drop_out_indices = npr.choice(negative_indices,
size=(len(negative_indices) - num_negative_bound),
replace=False)
label_matrix[drop_out_indices] = ignore
negative_indices = np.where(label_matrix == negative)[0]
# Collect the sampled proposal boxes.
sampled_indices = np.append(positive_indices, negative_indices)
sampled_proposal_boxes = proposal_boxes[sampled_indices]
sampled_proposal_scores = proposal_scores[sampled_indices]
labels = label_matrix[sampled_indices]
bbox_targets = compute_bbox_deltas(proposal_boxes[sampled_indices],
gt_boxes[pbox_max_overlaps_indices[sampled_indices]])
return sampled_proposal_boxes, sampled_proposal_scores, labels, bbox_targets
def compute_bbox_deltas(src_bboxes, dst_bboxes):
src_widths = src_bboxes[:, 2] - src_bboxes[:, 0] + 1.0
src_heights = src_bboxes[:, 3] - src_bboxes[:, 1] + 1.0
src_ctr_x = src_bboxes[:, 0] + 0.5 * src_widths
src_ctr_y = src_bboxes[:, 1] + 0.5 * src_heights
dst_widths = dst_bboxes[:, 2] - dst_bboxes[:, 0] + 1.0
dst_heights = dst_bboxes[:, 3] - dst_bboxes[:, 1] + 1.0
dst_ctr_x = dst_bboxes[:, 0] + 0.5 * dst_widths
dst_ctr_y = dst_bboxes[:, 1] + 0.5 * dst_heights
targets_dx = (dst_ctr_x - src_ctr_x) / src_widths
targets_dy = (dst_ctr_y - src_ctr_y) / src_heights
targets_dw = np.log(dst_widths / src_widths)
targets_dh = np.log(dst_heights / src_heights)
targets = np.vstack((targets_dx, targets_dy, targets_dw, targets_dh)).transpose()
return targets
# + id="N2URg1Lwtnei" colab_type="code" colab={}
sampled_proposal_boxes, sampled_proposal_scores, classification_y_true, regression_y_true = tf.py_func(
rcnn_target_and_sample,
[m_rpn.proposal_boxes,
m_rpn.proposal_scores, tf_gt_boxes, num_classes],
[tf.float32, tf.float32, tf.float32, tf.float32])
# + id="zxWoRoe4tnej" colab_type="code" colab={} outputId="81f01b01-ccd5-4645-a8ef-49397c2aca86"
print(sampled_proposal_boxes.shape)
print(sampled_proposal_scores.shape)
print(classification_y_true.shape)
print(regression_y_true.shape)
# + id="cpishv0vtnek" colab_type="code" colab={}
pboxes, pbox_scores, frcnn_target_labels, frcnn_target_t_parameters = sess.run(
[sampled_proposal_boxes,
sampled_proposal_scores,
classification_y_true,
regression_y_true
],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="m0TrfvEvtnek" colab_type="code" colab={} outputId="9df143ce-a9be-4b90-eec2-9f<PASSWORD>"
print('sampled_proposal_boxes: {}'.format(pboxes.shape))
print('sampled_proposal_scores: {}'.format(pbox_scores.shape))
print('frcnn_labels: {}'.format(frcnn_target_labels.shape))
print('frcnn_bbox_targets: {}'.format(frcnn_target_t_parameters.shape))
# + id="2VMQGdCvtnel" colab_type="code" colab={} outputId="0190c2b5-8cba-4be9-8102-f4e88ff45918"
print('Batch size: {:4}'.format(_hp.anchor_batch_size))
print(' Positive: {:4}'.format(np.count_nonzero(frcnn_target_labels == 1)))
print(' Negative: {:4}'.format(np.count_nonzero(frcnn_target_labels == 0)))
print(' Ignored : {:4}'.format(np.count_nonzero(frcnn_target_labels == -1)))
# + [markdown] id="P4BRB_qstnem" colab_type="text"
# ### 1.5.2 Build FRCNN Model
# + id="bAwRPjHytnen" colab_type="code" colab={}
class FRCNN_Model:
# ROI Pooling
# sampled proposal boxes for input
sampled_proposal_boxes = None
sampled_proposal_scores = None
# y_true (target value)
classification_y_true = None
regression_y_true = None
# roi pooling train output
roi_pooled_features = None
# fully connected feature extraction
fc_features = None
# output layer: classification
classification_logits = None
classification_probabilities = None
classification_y_predictions = None
# output layer: regression layer
regression_y_predictions = None
m_frcnn = FRCNN_Model()
# + [markdown] id="yeSXiJXbtnen" colab_type="text"
# #### save preprocessing results
# + id="_M_z3TOhtnen" colab_type="code" colab={}
m_frcnn.sampled_proposal_boxes = sampled_proposal_boxes
m_frcnn.sampled_proposal_scores = sampled_proposal_scores
m_frcnn.classification_y_true = classification_y_true
m_frcnn.regression_y_true = regression_y_true
# + id="CHfQO8zttneo" colab_type="code" colab={} outputId="b68e7b17-01c2-4b52-bf33-634702c7fa9c"
print(m_frcnn.sampled_proposal_boxes)
print(m_frcnn.sampled_proposal_scores)
print(m_frcnn.classification_y_true)
print(m_frcnn.regression_y_true)
# + [markdown] id="j_b3jC3rtnep" colab_type="text"
# #### ROI pooling layer
# + id="8-myMBWDtnep" colab_type="code" colab={}
def roi_pooling_layer(conv_feats, proposal_boxes):
pool_size = _hp.roi_pool_size
# Get normalized ROI coordinates.
ceiled_img_h = tf.to_float(tf.shape(conv_feats)[1]) * np.float32(_hp.total_stride)
ceiled_img_w = tf.to_float(tf.shape(conv_feats)[2]) * np.float32(_hp.total_stride)
xmin = proposal_boxes[:, 0:1] / ceiled_img_w
ymin = proposal_boxes[:, 1:2] / ceiled_img_h
xmax = proposal_boxes[:, 2:3] / ceiled_img_w
ymax = proposal_boxes[:, 3:4] / ceiled_img_h
normalized_boxes = tf.concat([ymin, xmin, ymax, xmax], axis=1) # [num_proposal_boxes, 4]
batch_indices = tf.zeros((tf.shape(normalized_boxes)[0],), dtype=tf.int32)
pre_pool_size = pool_size * 2
cropped_feats = tf.image.crop_and_resize(
conv_feats, normalized_boxes, batch_indices, [pre_pool_size, pre_pool_size], name='roi_pooled_feats'
)
pooled_feats = slim.max_pool2d(cropped_feats, [2, 2], padding='SAME')
return pooled_feats
# + id="8M4TwdA-tneq" colab_type="code" colab={}
m_frcnn.roi_pooled_features = roi_pooling_layer(
m_conv.output_feature_map,
m_frcnn.sampled_proposal_boxes)
# + id="IxVm84Uqtneq" colab_type="code" colab={} outputId="c496ecc4-279d-4e50-dfdf-ce65a081a74a"
print(m_frcnn.roi_pooled_features.shape) # 7 = roi pool size
# + [markdown] id="zQHFcaortner" colab_type="text"
# #### FC Feature Extraction Layers
# + id="CUjBe2lntner" colab_type="code" colab={}
def fc_layers(inputs):
pooled_rois_flat = slim.flatten(inputs, scope='pooled_rois_flat')
fc6 = slim.fully_connected(pooled_rois_flat, 4096, scope='fc6')
dropout6 = slim.dropout(fc6, keep_prob=0.5, is_training=is_train, scope='dropout6')
fc7 = slim.fully_connected(fc6, 4096, scope='fc7')
dropout7 = slim.dropout(fc7, keep_prob=0.5, is_training=is_train, scope='dropout7')
fc_feats = dropout7
return fc_feats
# + id="tfp9bLPqtnes" colab_type="code" colab={}
m_frcnn.fc_features = fc_layers(m_frcnn.roi_pooled_features)
# + id="C4ZW3I-Ptnet" colab_type="code" colab={} outputId="50ae8d1e-a505-4fc9-8665-18b9e435169f"
print(m_frcnn.fc_features.shape) # number of ? should be 128 (frcnn batch size)
# + id="IjIyO4A0tneu" colab_type="code" colab={} outputId="68300574-eaf3-483e-bbf0-d4e407906775"
sess.run(tf.global_variables_initializer())
fc_features = sess.run(m_frcnn.fc_features,
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
print(fc_features.shape)
# + [markdown] id="zvB5gEaetnev" colab_type="text"
# #### frcnn output layers: classification and regression
# + id="Bi5ipXn2tnev" colab_type="code" colab={}
def classification_layer(inputs, num_object_classes):
num_classes = num_object_classes + 1 # 1 is for background
scores = slim.fully_connected(
inputs, num_classes,
weights_initializer=tf.random_normal_initializer(mean=0.0, stddev=0.01),
trainable=is_train, activation_fn=None, scope='class_scores'
)
probs = tf.nn.softmax(scores)
y_predictions = tf.argmax(probs, axis=1)
m_frcnn.classification_logits = scores
m_frcnn.classification_probabilities = probs
m_frcnn.classification_y_predictions = y_predictions
# + id="2t1uwOhltnev" colab_type="code" colab={}
def regression_layer(inputs, num_object_classes):
num_classes = num_object_classes
t_parameters_from_proposalbox_to_gtbox = slim.fully_connected(
inputs, num_classes * 4,
weights_initializer=tf.random_normal_initializer(mean=0.0, stddev=0.001),
trainable=is_train, activation_fn=None, scope='bbox_params'
)
m_frcnn.regression_y_predictions = t_parameters_from_proposalbox_to_gtbox
# + id="69poNY-Mtnew" colab_type="code" colab={}
classification_layer(m_frcnn.fc_features, num_classes)
regression_layer(m_frcnn.fc_features, num_classes)
# + id="9VdjbrrKtnew" colab_type="code" colab={} outputId="28e079c6-937a-43ec-8892-7a4ee05f9630"
print(m_frcnn.classification_logits.shape)
print(m_frcnn.classification_probabilities.shape)
print(m_frcnn.classification_y_predictions.shape)
print(m_frcnn.regression_y_predictions.shape)
# + id="j-ETQ3AEtnex" colab_type="code" colab={}
sess.run(tf.global_variables_initializer())
sampled_proposal_boxes, proposal_boxes, y_true_cls, y_true_reg, roi_pooled_features, fc_features, frcnn_class_scores, frcnn_class_probs, frcnn_class_preds, frcnn_t_parameters = sess.run(
[m_frcnn.sampled_proposal_boxes, m_rpn.proposal_boxes,
m_frcnn.classification_y_true,
m_frcnn.regression_y_true,
m_frcnn.roi_pooled_features,
m_frcnn.fc_features,
m_frcnn.classification_logits,
m_frcnn.classification_probabilities,
m_frcnn.classification_y_predictions,
m_frcnn.regression_y_predictions],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="OtVbCU4qtney" colab_type="code" colab={} outputId="39bb8e89-fe8e-4dff-c954-49ec764aea3f"
print(proposal_boxes.shape)
print(sampled_proposal_boxes.shape)
print(y_true_cls.shape)
print(y_true_reg.shape)
print(roi_pooled_features.shape)
print(fc_features.shape)
print(frcnn_class_scores.shape)
print(frcnn_class_probs.shape)
print(frcnn_class_preds.shape)
print(frcnn_t_parameters.shape)
# + [markdown] id="dbZ_EEqutnez" colab_type="text"
# <br />
# # Set Model Propagation
#
# #### 1. Forward propagation
# model에 input data 넣어서 model로 구한 y_prediction 값을 구하는 과정
# - rpn 모델의 1) classification, 2) regression
# - frcnn 모델의 3) classification, 4) regression
# - 4 가지에 대한 prediction 계산 진행
#
# #### 2. Loss computation
# y_prediction 값과 y_true 값을 비교해서 loss (error)를 구하는 과정
# - 위 4 가지 prediction 에 대한 loss를 계산하고 하나의 loss로 합침
#
# #### 3. Backpropagation
# loss 를가지고 model 의 train weight 를 update, optimize 시킴
#
#
# 1 > 2 > 3 과정을 반복하며 y_true값과 유사한 결과를 내도록 모델을 학습시킨다.
# + id="DBu1a68rtnez" colab_type="code" colab={}
forward_propagation = None
compute_loss = None
back_propagation = None
# + id="d7N_PPlStnez" colab_type="code" colab={}
def set_forward_propagation():
forward_output = {}
forward_output['y_true'] = m_frcnn.classification_y_true
forward_output['y_pred'] = m_frcnn.classification_y_predictions
forward_output['proposal_boxes'] = m_frcnn.sampled_proposal_boxes
forward_output['t_parameters'] = m_frcnn.regression_y_predictions
forward_output['gt_boxes'] = tf_gt_boxes[0][:,:4]
return forward_output
# + id="ufE9ZXP2tne0" colab_type="code" colab={}
forward_propagation = set_forward_propagation()
# + id="kNGizGXstne0" colab_type="code" colab={} outputId="c1c51257-5673-4315-c49c-99f9ef0aafd4"
print(forward_propagation['y_true'])
# + id="jsAJT49itne1" colab_type="code" colab={}
def define_loss_function():
# rpn losses
rpn_classification_logits = m_rpn.classification_logits
rpn_classification_y_true = m_rpn.classification_y_true
rpn_regression_prediction = m_rpn.regression_y_predictions
rpn_regression_y_true = m_rpn.regression_y_true
rpn_classification_y_true = tf.reshape(rpn_classification_y_true, (-1,))
idx_selected = tf.reshape(tf.where(tf.not_equal(rpn_classification_y_true, -1)), (-1,))
idx_positive = tf.reshape(tf.where(tf.equal(rpn_classification_y_true, 1)), (-1,))
rpn_classification_logits = tf.gather(tf.reshape(rpn_classification_logits, (-1, 2)), idx_selected)
rpn_classification_y_true = tf.gather(rpn_classification_y_true, idx_selected)
rpn_classification_loss = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=rpn_classification_logits,
labels=tf.cast(rpn_classification_y_true, tf.int64)))
rpn_regression_prediction = tf.gather(tf.reshape(rpn_regression_prediction, (-1, 4)), idx_positive)
rpn_regression_y_true = tf.gather(tf.reshape(rpn_regression_y_true, (-1, 4)), idx_positive)
rpn_huber_loss = tf.losses.huber_loss(
rpn_regression_y_true,
rpn_regression_prediction,
reduction=tf.losses.Reduction.NONE)
rpn_regression_loss = tf.reduce_mean(
tf.reduce_sum(rpn_huber_loss, axis=1))
# frcnn losses
frcnn_classification_logits = m_frcnn.classification_logits
frcnn_classification_y_true = m_frcnn.classification_y_true
frcnn_regression_prediction = m_frcnn.regression_y_predictions
frcnn_regression_y_true = m_frcnn.regression_y_true
frcnn_classification_loss = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=frcnn_classification_logits,
labels=tf.cast(frcnn_classification_y_true, tf.int64)))
idx_not_ignore = tf.reshape(tf.where(tf.not_equal(frcnn_classification_y_true, 0)), (-1,))
frcnn_regression_prediction = tf.reshape(frcnn_regression_prediction, (_hp.proposal_batch_size, -1, 4)) # [batch_size, num_classes, 4]
frcnn_regression_y_true = tf.reshape(frcnn_regression_y_true, (_hp.proposal_batch_size, 1, 4)) # [batch_size, 1, 4]
frcnn_regression_mask = tf.one_hot(tf.cast(frcnn_classification_y_true, tf.int64), tf.shape(frcnn_regression_prediction)[1])
frcnn_regression_mask = tf.expand_dims(frcnn_regression_mask, axis=2)
frcnn_huber_loss = tf.losses.huber_loss(
frcnn_regression_y_true,
frcnn_regression_prediction,
reduction=tf.losses.Reduction.NONE)
frcnn_huber_loss_masked = frcnn_regression_mask * frcnn_huber_loss
# 계산은 다 하지만 mask로 필요한 애들만 적용
frcnn_regression_loss = tf.reduce_mean(tf.reduce_sum(
frcnn_huber_loss_masked[:, 1:, :], axis=[1, 2]))
loss_result = {}
loss_result['rpn_cls'] = rpn_classification_loss
loss_result['rpn_reg'] = rpn_regression_loss
loss_result['frcnn_cls'] = frcnn_classification_loss
loss_result['frcnn_reg'] = frcnn_regression_loss
loss_result['total'] = rpn_classification_loss + rpn_regression_loss + frcnn_classification_loss + frcnn_regression_loss
return loss_result
# + id="eUww8q_ktne2" colab_type="code" colab={}
compute_loss = define_loss_function()
# + id="ww2DIDtItne2" colab_type="code" colab={}
current_learning_rate = 0.0
def set_backward_propagation():
variables_to_update = tf.trainable_variables()
global current_learning_rate
current_learning_rate = tf.placeholder(tf.float32)
# momentum , learning rate 설명
optimizer = tf.train.MomentumOptimizer(
current_learning_rate,
_hp.momentum,
use_nesterov=False).minimize(
compute_loss['total'], var_list=variables_to_update)
return optimizer
# + id="VoW_erIbtne3" colab_type="code" colab={} outputId="c24cc613-37fb-4c3d-ce62-de74eb0bd5e0"
back_propagation = set_backward_propagation()
# + id="yuPCegmctne4" colab_type="code" colab={}
sess.run(tf.global_variables_initializer())
forward_output = sess.run(forward_propagation,
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
# + id="KGZz4BW1tne4" colab_type="code" colab={} outputId="fbaa89c1-064b-4f64-ec6f-e090af5279f0"
print('FRCNN y_true batch set: ', forward_output['y_true'].shape)
print('FRCNN y_pred batch set: ', forward_output['y_pred'].shape)
print('FRCNN proposal_boxes : ', forward_output['proposal_boxes'].shape)
print('FRCNN t_parameters : ', forward_output['t_parameters'].shape)
print('FRCNN gt_boxes : ', forward_output['gt_boxes'].shape)
# + [markdown] id="e8qqbB27tne5" colab_type="text"
# <br />
# # Run Train
#
# #### functions to help batch training/testing
# + id="5eWiP-O0tne5" colab_type="code" colab={}
def execute_train(sess):
global current_learning_rate
for batch_step in range(_hp.num_batches_per_epoch):
batch_image, batch_gt_boxes = data_train.next_batch(_hp.batch_size, shuffle=True)
train_forward, train_loss, _ = sess.run(
[forward_propagation,
compute_loss,
back_propagation],
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes,
current_learning_rate: _hp.current_learning_rate_value})
return train_loss
# + id="ybc2k1Aitne6" colab_type="code" colab={}
import os
import time
base_path = 'trained_model_result/' # result saving location
if not os.path.exists(base_path):
os.makedirs(base_path)
os.chown(base_path, uid=1000, gid=1000)
timestamp = time.strftime("%Y%m%d_%H%M%S")
output_path = os.path.join(base_path, timestamp + '/')
os.makedirs(output_path)
os.chown(output_path, uid=1000, gid=1000)
# + id="rhnv7skQtne6" colab_type="code" colab={} outputId="3c24bb1a-ab43-49b8-ed44-b65a3ba71eb1"
bad_epochs = 0
min_loss = 0.0
current_learning_rate_value = _hp.init_learning_rate
graph = tf.get_default_graph()
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(graph=graph, config=config)
sess.run(tf.global_variables_initializer()) # initialize all weights
saver = tf.train.Saver() # to save trained model
output_model_path = os.path.join(output_path, 'model.ckpt')
train_results = dict() # dictionary to contain training(, evaluation) results and details
total_steps = _hp.num_epochs * _hp.num_batches_per_epoch
str = '\n------------------------------------------------------------------------' + \
'\n execute train' + \
'\n------------------------------------------------------------------------' + \
'\n train data size : {:10}'.format(data_train.num_samples) + \
'\n batch size : {:10}'.format(_hp.batch_size) + \
'\n batche loop per epoch : {:10} = |train data| {} / |batch| {}'.format(_hp.num_batches_per_epoch, data_train.num_samples, _hp.batch_size) + \
'\n epoches : {:10}'.format(_hp.num_epochs) + \
'\n total iterations : {:10} = |batch loop| {} * |epoch| {}\n\n'.format(total_steps, _hp.num_batches_per_epoch, _hp.num_epochs)
print(str)
start_time = time.time()
# + id="bDHyzu7ptne7" colab_type="code" colab={}
def is_better(current_loss, min_loss):
return current_loss['total'] < min_loss
# + id="o9dxOfTrtne8" colab_type="code" colab={}
def update_learning_rate():
# decaying learning rate (epsilon)
global bad_epochs
if bad_epochs > _hp.patience_of_no_improvement_epochs:
new_learning_rate = _hp.current_learning_rate_value * _hp.learning_rate_decay
# Decay learning rate only when the difference is higher than lower bound epsilon.
if _hp.current_learning_rate_value - new_learning_rate > _hp.lower_bound_learning_rate:
_hp.current_learning_rate_value = new_learning_rate
bad_epochs = 0
# + id="PmXtv6zGtne8" colab_type="code" colab={}
_hp.num_epochs = 10 # to test
_hp.num_batches_per_epoch = 1 # to test
# + id="e9bTI4Vctne9" colab_type="code" colab={} outputId="5d3f5073-a0b9-4dc6-a1c2-a6ff73a95735"
# start training loop
for epoch_step in range(1, _hp.num_epochs + 1):
# perform a gradient update of the current epoch
current_loss = execute_train(sess)
str = '[epoch{:4}] loss: {:.6f} | learning rate: {:.6f}'\
.format(epoch_step, current_loss['total'], current_learning_rate_value)
print(str)
# Keep track of the current best model,
if is_better(current_loss, min_loss):
min_loss = current_loss
bad_epochs = 0
saver.save(sess, output_model_path) # save current weights
else:
bad_epochs += 1
update_learning_rate()
if current_learning_rate_value < 0.000001:
print(' exit train: learning rate is too small (< 0.000001)')
break
# + id="EqiL_IARtne-" colab_type="code" colab={} outputId="589a1414-2d1c-4aaa-88bf-b980d74859d2"
saver.save(sess, output_model_path) # to test
# + [markdown] id="o2Mn_YBmtne_" colab_type="text"
# # Test
# + id="YGp8__l5tne_" colab_type="code" colab={}
is_train = False
# + id="mzW3IY_XtnfA" colab_type="code" colab={}
def execute_test(sess):
batch_image, batch_gt_boxes = data_test.next_batch(_hp.batch_size, shuffle=True)
test_forward = sess.run(forward_propagation,
feed_dict={tf_input_image: batch_image,
tf_gt_boxes: batch_gt_boxes})
return test_forward
# + id="2oaC_GcDtnfB" colab_type="code" colab={}
data_test = data2.test
num_classes = data_test.num_classes
# + id="nv7aZmRptnfC" colab_type="code" colab={} outputId="710bb8c7-b241-4bb9-aa67-28d1e6f57f33"
graph = tf.get_default_graph()
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(graph=graph, config=config)
saver = tf.train.Saver()
saver.restore(sess, output_model_path)
str = '\n------------------------------------------------------------------------' + \
'\nexecute test' + \
'\n------------------------------------------------------------------------' + \
'\n test data size : {:10}'.format(data_test.num_samples) + \
'\n batch size : {:10}'.format(_hp.batch_size)
print(str)
# + id="2wetEbKWtnfD" colab_type="code" colab={}
# + id="tMjzXaBEtnfD" colab_type="code" colab={}
| 5.CNN_Advanced/RCNN/01_faster_rcnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Single Variable Calculus
# :label:`sec_single_variable_calculus`
#
# In :numref:`sec_calculus`, we saw the basic elements of differential calculus. This section takes a deeper dive into the fundamentals of calculus and how we can understand and apply it in the context of machine learning.
#
# ## Differential Calculus
# Differential calculus is fundamentally the study of how functions behave under small changes. To see why this is so core to deep learning, let us consider an example.
#
# Suppose that we have a deep neural network where the weights are, for convenience, concatenated into a single vector $\mathbf{w} = (w_1, \ldots, w_n)$. Given a training dataset, we consider the loss of our neural network on this dataset, which we will write as $\mathcal{L}(\mathbf{w})$.
#
# This function is extraordinarily complex, encoding the performance of all possible models of the given architecture on this dataset, so it is nearly impossible to tell what set of weights $\mathbf{w}$ will minimize the loss. Thus, in practice, we often start by initializing our weights *randomly*, and then iteratively take small steps in the direction which makes the loss decrease as rapidly as possible.
#
# The question then becomes something that on the surface is no easier: how do we find the direction which makes the weights decrease as quickly as possible? To dig into this, let us first examine the case with only a single weight: $L(\mathbf{w}) = L(x)$ for a single real value $x$.
#
# Let us take $x$ and try to understand what happens when we change it by a small amount to $x + \epsilon$. If you wish to be concrete, think a number like $\epsilon = 0.0000001$. To help us visualize what happens, let us graph an example function, $f(x) = \sin(x^x)$, over the $[0, 3]$.
#
# + origin_pos=2 tab=["pytorch"]
# %matplotlib inline
import torch
from IPython import display
from d2l import torch as d2l
torch.pi = torch.acos(torch.zeros(1)).item() * 2 # Define pi in torch
# Plot a function in a normal range
x_big = torch.arange(0.01, 3.01, 0.01)
ys = torch.sin(x_big**x_big)
d2l.plot(x_big, ys, 'x', 'f(x)')
# + [markdown] origin_pos=4
# At this large scale, the function's behavior is not simple. However, if we reduce our range to something smaller like $[1.75,2.25]$, we see that the graph becomes much simpler.
#
# + origin_pos=6 tab=["pytorch"]
# Plot a the same function in a tiny range
x_med = torch.arange(1.75, 2.25, 0.001)
ys = torch.sin(x_med**x_med)
d2l.plot(x_med, ys, 'x', 'f(x)')
# + [markdown] origin_pos=8
# Taking this to an extreme, if we zoom into a tiny segment, the behavior becomes far simpler: it is just a straight line.
#
# + origin_pos=10 tab=["pytorch"]
# Plot a the same function in a tiny range
x_small = torch.arange(2.0, 2.01, 0.0001)
ys = torch.sin(x_small**x_small)
d2l.plot(x_small, ys, 'x', 'f(x)')
# + [markdown] origin_pos=12
# This is the key observation of single variable calculus: the behavior of familiar functions can be modeled by a line in a small enough range. This means that for most functions, it is reasonable to expect that as we shift the $x$ value of the function by a little bit, the output $f(x)$ will also be shifted by a little bit. The only question we need to answer is, "How large is the change in the output compared to the change in the input? Is it half as large? Twice as large?"
#
# Thus, we can consider the ratio of the change in the output of a function for a small change in the input of the function. We can write this formally as
#
# $$
# \frac{L(x+\epsilon) - L(x)}{(x+\epsilon) - x} = \frac{L(x+\epsilon) - L(x)}{\epsilon}.
# $$
#
# This is already enough to start to play around with in code. For instance, suppose that we know that $L(x) = x^{2} + 1701(x-4)^3$, then we can see how large this value is at the point $x = 4$ as follows.
#
# + origin_pos=13 tab=["pytorch"]
# Define our function
def L(x):
return x**2 + 1701*(x-4)**3
# Print the difference divided by epsilon for several epsilon
for epsilon in [0.1, 0.001, 0.0001, 0.00001]:
print(f'epsilon = {epsilon:.5f} -> {(L(4+epsilon) - L(4)) / epsilon:.5f}')
# + [markdown] origin_pos=14
# Now, if we are observant, we will notice that the output of this number is suspiciously close to $8$. Indeed, if we decrease $\epsilon$, we will see value becomes progressively closer to $8$. Thus we may conclude, correctly, that the value we seek (the degree a change in the input changes the output) should be $8$ at the point $x=4$. The way that a mathematician encodes this fact is
#
# $$
# \lim_{\epsilon \rightarrow 0}\frac{L(4+\epsilon) - L(4)}{\epsilon} = 8.
# $$
#
# As a bit of a historical digression: in the first few decades of neural network research, scientists used this algorithm (the *method of finite differences*) to evaluate how a loss function changed under small perturbation: just change the weights and see how the loss changed. This is computationally inefficient, requiring two evaluations of the loss function to see how a single change of one variable influenced the loss. If we tried to do this with even a paltry few thousand parameters, it would require several thousand evaluations of the network over the entire dataset! It was not solved until 1986 that the *backpropagation algorithm* introduced in :cite:`Rumelhart.Hinton.Williams.ea.1988` provided a way to calculate how *any* change of the weights together would change the loss in the same computation time as a single prediction of the network over the dataset.
#
# Back in our example, this value $8$ is different for different values of $x$, so it makes sense to define it as a function of $x$. More formally, this value dependent rate of change is referred to as the *derivative* which is written as
#
# $$\frac{df}{dx}(x) = \lim_{\epsilon \rightarrow 0}\frac{f(x+\epsilon) - f(x)}{\epsilon}.$$
# :eqlabel:`eq_der_def`
#
# Different texts will use different notations for the derivative. For instance, all of the below notations indicate the same thing:
#
# $$
# \frac{df}{dx} = \frac{d}{dx}f = f' = \nabla_xf = D_xf = f_x.
# $$
#
# Most authors will pick a single notation and stick with it, however even that is not guaranteed. It is best to be familiar with all of these. We will use the notation $\frac{df}{dx}$ throughout this text, unless we want to take the derivative of a complex expression, in which case we will use $\frac{d}{dx}f$ to write expressions like
# $$
# \frac{d}{dx}\left[x^4+\cos\left(\frac{x^2+1}{2x-1}\right)\right].
# $$
#
# Oftentimes, it is intuitively useful to unravel the definition of derivative :eqref:`eq_der_def` again to see how a function changes when we make a small change of $x$:
#
# $$\begin{aligned} \frac{df}{dx}(x) = \lim_{\epsilon \rightarrow 0}\frac{f(x+\epsilon) - f(x)}{\epsilon} & \implies \frac{df}{dx}(x) \approx \frac{f(x+\epsilon) - f(x)}{\epsilon} \\ & \implies \epsilon \frac{df}{dx}(x) \approx f(x+\epsilon) - f(x) \\ & \implies f(x+\epsilon) \approx f(x) + \epsilon \frac{df}{dx}(x). \end{aligned}$$
# :eqlabel:`eq_small_change`
#
# The last equation is worth explicitly calling out. It tells us that if you take any function and change the input by a small amount, the output would change by that small amount scaled by the derivative.
#
# In this way, we can understand the derivative as the scaling factor that tells us how large of change we get in the output from a change in the input.
#
# ## Rules of Calculus
# :label:`sec_derivative_table`
#
# We now turn to the task of understanding how to compute the derivative of an explicit function. A full formal treatment of calculus would derive everything from first principles. We will not indulge in this temptation here, but rather provide an understanding of the common rules encountered.
#
# ### Common Derivatives
# As was seen in :numref:`sec_calculus`, when computing derivatives one can oftentimes use a series of rules to reduce the computation to a few core functions. We repeat them here for ease of reference.
#
# * **Derivative of constants.** $\frac{d}{dx}c = 0$.
# * **Derivative of linear functions.** $\frac{d}{dx}(ax) = a$.
# * **Power rule.** $\frac{d}{dx}x^n = nx^{n-1}$.
# * **Derivative of exponentials.** $\frac{d}{dx}e^x = e^x$.
# * **Derivative of the logarithm.** $\frac{d}{dx}\log(x) = \frac{1}{x}$.
#
# ### Derivative Rules
# If every derivative needed to be separately computed and stored in a table, differential calculus would be near impossible. It is a gift of mathematics that we can generalize the above derivatives and compute more complex derivatives like finding the derivative of $f(x) = \log\left(1+(x-1)^{10}\right)$. As was mentioned in :numref:`sec_calculus`, the key to doing so is to codify what happens when we take functions and combine them in various ways, most importantly: sums, products, and compositions.
#
# * **Sum rule.** $\frac{d}{dx}\left(g(x) + h(x)\right) = \frac{dg}{dx}(x) + \frac{dh}{dx}(x)$.
# * **Product rule.** $\frac{d}{dx}\left(g(x)\cdot h(x)\right) = g(x)\frac{dh}{dx}(x) + \frac{dg}{dx}(x)h(x)$.
# * **Chain rule.** $\frac{d}{dx}g(h(x)) = \frac{dg}{dh}(h(x))\cdot \frac{dh}{dx}(x)$.
#
# Let us see how we may use :eqref:`eq_small_change` to understand these rules. For the sum rule, consider following chain of reasoning:
#
# $$
# \begin{aligned}
# f(x+\epsilon) & = g(x+\epsilon) + h(x+\epsilon) \\
# & \approx g(x) + \epsilon \frac{dg}{dx}(x) + h(x) + \epsilon \frac{dh}{dx}(x) \\
# & = g(x) + h(x) + \epsilon\left(\frac{dg}{dx}(x) + \frac{dh}{dx}(x)\right) \\
# & = f(x) + \epsilon\left(\frac{dg}{dx}(x) + \frac{dh}{dx}(x)\right).
# \end{aligned}
# $$
#
# By comparing this result with the fact that $f(x+\epsilon) \approx f(x) + \epsilon \frac{df}{dx}(x)$, we see that $\frac{df}{dx}(x) = \frac{dg}{dx}(x) + \frac{dh}{dx}(x)$ as desired. The intuition here is: when we change the input $x$, $g$ and $h$ jointly contribute to the change of the output by $\frac{dg}{dx}(x)$ and $\frac{dh}{dx}(x)$.
#
#
# The product is more subtle, and will require a new observation about how to work with these expressions. We will begin as before using :eqref:`eq_small_change`:
#
# $$
# \begin{aligned}
# f(x+\epsilon) & = g(x+\epsilon)\cdot h(x+\epsilon) \\
# & \approx \left(g(x) + \epsilon \frac{dg}{dx}(x)\right)\cdot\left(h(x) + \epsilon \frac{dh}{dx}(x)\right) \\
# & = g(x)\cdot h(x) + \epsilon\left(g(x)\frac{dh}{dx}(x) + \frac{dg}{dx}(x)h(x)\right) + \epsilon^2\frac{dg}{dx}(x)\frac{dh}{dx}(x) \\
# & = f(x) + \epsilon\left(g(x)\frac{dh}{dx}(x) + \frac{dg}{dx}(x)h(x)\right) + \epsilon^2\frac{dg}{dx}(x)\frac{dh}{dx}(x). \\
# \end{aligned}
# $$
#
#
# This resembles the computation done above, and indeed we see our answer ($\frac{df}{dx}(x) = g(x)\frac{dh}{dx}(x) + \frac{dg}{dx}(x)h(x)$) sitting next to $\epsilon$, but there is the issue of that term of size $\epsilon^{2}$. We will refer to this as a *higher-order term*, since the power of $\epsilon^2$ is higher than the power of $\epsilon^1$. We will see in a later section that we will sometimes want to keep track of these, however for now observe that if $\epsilon = 0.0000001$, then $\epsilon^{2}= 0.0000000000001$, which is vastly smaller. As we send $\epsilon \rightarrow 0$, we may safely ignore the higher order terms. As a general convention in this appendix, we will use "$\approx$" to denote that the two terms are equal up to higher order terms. However, if we wish to be more formal we may examine the difference quotient
#
# $$
# \frac{f(x+\epsilon) - f(x)}{\epsilon} = g(x)\frac{dh}{dx}(x) + \frac{dg}{dx}(x)h(x) + \epsilon \frac{dg}{dx}(x)\frac{dh}{dx}(x),
# $$
#
# and see that as we send $\epsilon \rightarrow 0$, the right hand term goes to zero as well.
#
# Finally, with the chain rule, we can again progress as before using :eqref:`eq_small_change` and see that
#
# $$
# \begin{aligned}
# f(x+\epsilon) & = g(h(x+\epsilon)) \\
# & \approx g\left(h(x) + \epsilon \frac{dh}{dx}(x)\right) \\
# & \approx g(h(x)) + \epsilon \frac{dh}{dx}(x) \frac{dg}{dh}(h(x))\\
# & = f(x) + \epsilon \frac{dg}{dh}(h(x))\frac{dh}{dx}(x),
# \end{aligned}
# $$
#
# where in the second line we view the function $g$ as having its input ($h(x)$) shifted by the tiny quantity $\epsilon \frac{dh}{dx}(x)$.
#
# These rule provide us with a flexible set of tools to compute essentially any expression desired. For instance,
#
# $$
# \begin{aligned}
# \frac{d}{dx}\left[\log\left(1+(x-1)^{10}\right)\right] & = \left(1+(x-1)^{10}\right)^{-1}\frac{d}{dx}\left[1+(x-1)^{10}\right]\\
# & = \left(1+(x-1)^{10}\right)^{-1}\left(\frac{d}{dx}[1] + \frac{d}{dx}[(x-1)^{10}]\right) \\
# & = \left(1+(x-1)^{10}\right)^{-1}\left(0 + 10(x-1)^9\frac{d}{dx}[x-1]\right) \\
# & = 10\left(1+(x-1)^{10}\right)^{-1}(x-1)^9 \\
# & = \frac{10(x-1)^9}{1+(x-1)^{10}}.
# \end{aligned}
# $$
#
# Where each line has used the following rules:
#
# 1. The chain rule and derivative of logarithm.
# 2. The sum rule.
# 3. The derivative of constants, chain rule, and power rule.
# 4. The sum rule, derivative of linear functions, derivative of constants.
#
# Two things should be clear after doing this example:
#
# 1. Any function we can write down using sums, products, constants, powers, exponentials, and logarithms can have its derivate computed mechanically by following these rules.
# 2. Having a human follow these rules can be tedious and error prone!
#
# Thankfully, these two facts together hint towards a way forward: this is a perfect candidate for mechanization! Indeed backpropagation, which we will revisit later in this section, is exactly that.
#
# ### Linear Approximation
# When working with derivatives, it is often useful to geometrically interpret the approximation used above. In particular, note that the equation
#
# $$
# f(x+\epsilon) \approx f(x) + \epsilon \frac{df}{dx}(x),
# $$
#
# approximates the value of $f$ by a line which passes through the point $(x, f(x))$ and has slope $\frac{df}{dx}(x)$. In this way we say that the derivative gives a linear approximation to the function $f$, as illustrated below:
#
# + origin_pos=16 tab=["pytorch"]
# Compute sin
xs = torch.arange(-torch.pi, torch.pi, 0.01)
plots = [torch.sin(xs)]
# Compute some linear approximations. Use d(sin(x))/dx = cos(x)
for x0 in [-1.5, 0.0, 2.0]:
plots.append(torch.sin(torch.tensor(x0)) + (xs - x0) *
torch.cos(torch.tensor(x0)))
d2l.plot(xs, plots, 'x', 'f(x)', ylim=[-1.5, 1.5])
# + [markdown] origin_pos=18
# ### Higher Order Derivatives
#
# Let us now do something that may on the surface seem strange. Take a function $f$ and compute the derivative $\frac{df}{dx}$. This gives us the rate of change of $f$ at any point.
#
# However, the derivative, $\frac{df}{dx}$, can be viewed as a function itself, so nothing stops us from computing the derivative of $\frac{df}{dx}$ to get $\frac{d^2f}{dx^2} = \frac{df}{dx}\left(\frac{df}{dx}\right)$. We will call this the second derivative of $f$. This function is the rate of change of the rate of change of $f$, or in other words, how the rate of change is changing. We may apply the derivative any number of times to obtain what is called the $n$-th derivative. To keep the notation clean, we will denote the $n$-th derivative as
#
# $$
# f^{(n)}(x) = \frac{d^{n}f}{dx^{n}} = \left(\frac{d}{dx}\right)^{n} f.
# $$
#
# Let us try to understand *why* this is a useful notion. Below, we visualize $f^{(2)}(x)$, $f^{(1)}(x)$, and $f(x)$.
#
# First, consider the case that the second derivative $f^{(2)}(x)$ is a positive constant. This means that the slope of the first derivative is positive. As a result, the first derivative $f^{(1)}(x)$ may start out negative, becomes zero at a point, and then becomes positive in the end. This tells us the slope of our original function $f$ and therefore, the function $f$ itself decreases, flattens out, then increases. In other words, the function $f$ curves up, and has a single minimum as is shown in :numref:`fig_positive-second`.
#
# 
# :label:`fig_positive-second`
#
#
# Second, if the second derivative is a negative constant, that means that the first derivative is decreasing. This implies the first derivative may start out positive, becomes zero at a point, and then becomes negative. Hence, the function $f$ itself increases, flattens out, then decreases. In other words, the function $f$ curves down, and has a single maximum as is shown in :numref:`fig_negative-second`.
#
# 
# :label:`fig_negative-second`
#
#
# Third, if the second derivative is a always zero, then the first derivative will never change---it is constant! This means that $f$ increases (or decreases) at a fixed rate, and $f$ is itself a straight line as is shown in :numref:`fig_zero-second`.
#
# 
# :label:`fig_zero-second`
#
# To summarize, the second derivative can be interpreted as describing the way that the function $f$ curves. A positive second derivative leads to a upwards curve, while a negative second derivative means that $f$ curves downwards, and a zero second derivative means that $f$ does not curve at all.
#
# Let us take this one step further. Consider the function $g(x) = ax^{2}+ bx + c$. We can then compute that
#
# $$
# \begin{aligned}
# \frac{dg}{dx}(x) & = 2ax + b \\
# \frac{d^2g}{dx^2}(x) & = 2a.
# \end{aligned}
# $$
#
# If we have some original function $f(x)$ in mind, we may compute the first two derivatives and find the values for $a, b$, and $c$ that make them match this computation. Similarly to the previous section where we saw that the first derivative gave the best approximation with a straight line, this construction provides the best approximation by a quadratic. Let us visualize this for $f(x) = \sin(x)$.
#
# + origin_pos=20 tab=["pytorch"]
# Compute sin
xs = torch.arange(-torch.pi, torch.pi, 0.01)
plots = [torch.sin(xs)]
# Compute some quadratic approximations. Use d(sin(x)) / dx = cos(x)
for x0 in [-1.5, 0.0, 2.0]:
plots.append(torch.sin(torch.tensor(x0)) + (xs - x0) *
torch.cos(torch.tensor(x0)) - (xs - x0)**2 *
torch.sin(torch.tensor(x0)) / 2)
d2l.plot(xs, plots, 'x', 'f(x)', ylim=[-1.5, 1.5])
# + [markdown] origin_pos=22
# We will extend this idea to the idea of a *Taylor series* in the next section.
#
# ### Taylor Series
#
#
# The *Taylor series* provides a method to approximate the function $f(x)$ if we are given values for the first $n$ derivatives at a point $x_0$, i.e., $\left\{ f(x_0), f^{(1)}(x_0), f^{(2)}(x_0), \ldots, f^{(n)}(x_0) \right\}$. The idea will be to find a degree $n$ polynomial that matches all the given derivatives at $x_0$.
#
# We saw the case of $n=2$ in the previous section and a little algebra shows this is
#
# $$
# f(x) \approx \frac{1}{2}\frac{d^2f}{dx^2}(x_0)(x-x_0)^{2}+ \frac{df}{dx}(x_0)(x-x_0) + f(x_0).
# $$
#
# As we can see above, the denominator of $2$ is there to cancel out the $2$ we get when we take two derivatives of $x^2$, while the other terms are all zero. Same logic applies for the first derivative and the value itself.
#
# If we push the logic further to $n=3$, we will conclude that
#
# $$
# f(x) \approx \frac{\frac{d^3f}{dx^3}(x_0)}{6}(x-x_0)^3 + \frac{\frac{d^2f}{dx^2}(x_0)}{2}(x-x_0)^{2}+ \frac{df}{dx}(x_0)(x-x_0) + f(x_0).
# $$
#
# where the $6 = 3 \times 2 = 3!$ comes from the constant we get in front if we take three derivatives of $x^3$.
#
#
# Furthermore, we can get a degree $n$ polynomial by
#
# $$
# P_n(x) = \sum_{i = 0}^{n} \frac{f^{(i)}(x_0)}{i!}(x-x_0)^{i}.
# $$
#
# where the notation
#
# $$
# f^{(n)}(x) = \frac{d^{n}f}{dx^{n}} = \left(\frac{d}{dx}\right)^{n} f.
# $$
#
#
# Indeed, $P_n(x)$ can be viewed as the best $n$-th degree polynomial approximation to our function $f(x)$.
#
# While we are not going to dive all the way into the error of the above approximations, it is worth mentioning the infinite limit. In this case, for well behaved functions (known as real analytic functions) like $\cos(x)$ or $e^{x}$, we can write out the infinite number of terms and approximate the exactly same function
#
# $$
# f(x) = \sum_{n = 0}^\infty \frac{f^{(n)}(x_0)}{n!}(x-x_0)^{n}.
# $$
#
# Take $f(x) = e^{x}$ as am example. Since $e^{x}$ is its own derivative, we know that $f^{(n)}(x) = e^{x}$. Therefore, $e^{x}$ can be reconstructed by taking the Taylor series at $x_0 = 0$, i.e.,
#
# $$
# e^{x} = \sum_{n = 0}^\infty \frac{x^{n}}{n!} = 1 + x + \frac{x^2}{2} + \frac{x^3}{6} + \cdots.
# $$
#
# Let us see how this works in code and observe how increasing the degree of the Taylor approximation brings us closer to the desired function $e^x$.
#
# + origin_pos=24 tab=["pytorch"]
# Compute the exponential function
xs = torch.arange(0, 3, 0.01)
ys = torch.exp(xs)
# Compute a few Taylor series approximations
P1 = 1 + xs
P2 = 1 + xs + xs**2 / 2
P5 = 1 + xs + xs**2 / 2 + xs**3 / 6 + xs**4 / 24 + xs**5 / 120
d2l.plot(xs, [ys, P1, P2, P5], 'x', 'f(x)', legend=[
"Exponential", "Degree 1 Taylor Series", "Degree 2 Taylor Series",
"Degree 5 Taylor Series"])
# + [markdown] origin_pos=26
# Taylor series have two primary applications:
#
# 1. *Theoretical applications*: Often when we try to understand a too complex function, using Taylor series enables us to turn it into a polynomial that we can work with directly.
#
# 2. *Numerical applications*: Some functions like $e^{x}$ or $\cos(x)$ are difficult for machines to compute. They can store tables of values at a fixed precision (and this is often done), but it still leaves open questions like "What is the 1000-th digit of $\cos(1)$?" Taylor series are often helpful to answer such questions.
#
#
# ## Summary
#
# * Derivatives can be used to express how functions change when we change the input by a small amount.
# * Elementary derivatives can be combined using derivative rules to create arbitrarily complex derivatives.
# * Derivatives can be iterated to get second or higher order derivatives. Each increase in order provides more fine grained information on the behavior of the function.
# * Using information in the derivatives of a single data example, we can approximate well behaved functions by polynomials obtained from the Taylor series.
#
#
# ## Exercises
#
# 1. What is the derivative of $x^3-4x+1$?
# 2. What is the derivative of $\log(\frac{1}{x})$?
# 3. True or False: If $f'(x) = 0$ then $f$ has a maximum or minimum at $x$?
# 4. Where is the minimum of $f(x) = x\log(x)$ for $x\ge0$ (where we assume that $f$ takes the limiting value of $0$ at $f(0)$)?
#
# + [markdown] origin_pos=28 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1088)
#
| python/d2l-en/pytorch/chapter_appendix-mathematics-for-deep-learning/single-variable-calculus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aayushkumar20/ML-based-projects./blob/main/Counts%20number%20of%20persons%20in%20a%20given%20image%2C%20video%20and%20realtime/counts_number_of_person_in_the_video_and_images.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fNUj9lo3eusi"
# # Important modules (please install before executing the code)..
# + id="f8Ir1SVWe8iH"
# !pip install opencv-python
# !pip install imutils
# !pip install numpy
# + [markdown] id="FDN9X43xfMC-"
# ### Importing all required modules and assigning a simple name to the numpy.
# + id="H-V0XXg9fKiX"
from csv import writer
from tabnanny import check
import cv2
import numpy as np
import imutils
import argparse
# + [markdown] id="7lsSnE_AfdtM"
# ### Creating a model for detecting Humans.
# + id="zLalWzWkfj0m"
HOGCV=cv2.HOGDescriptor()
HOGCV.setSVMDetector(cv2.HOGDescriptor_getDefaultPeopleDetector())
# + [markdown] id="lKX-CtpNfoxw"
# ##### 👆 Here `cv2.HOGDescriptor_getDefaultPeopleDetector()` is a pre-trained model with approx 98% efficiency for detecting Human being in a real time situations.
# + [markdown] id="BQSfyBUlgxYR"
# # 1) Detect method
# ## It'll create a box around the 🧑.
# + [markdown] id="7w_i0mbLhuvf"
# **It'll returns two values**
# <br>
# 1. List containing coordinates of bounding Box of person in the form of X,Y,W,H.
# 2. Accuracy.
# + id="i5Ix5I-GgsM7"
def detect(frame):
bounding_boxes_coordinates, weights = HOGCV.detectMultiScale(frame, winStride=(4,4), padding=(8,8), scale=1.05)
person = 1
for (x,y,w,h) in bounding_boxes_coordinates:
cv2.rectangle(frame, (x,y), (x+w,y+h), (0,0,255), 2)
cv2.putText(frame, "Person {}".format(person), (x,y-5), cv2.FONT_HERSHEY_SIMPLEX, 0.5, (0,0,255), 2)
person += 1
cv2.putText(frame, 'Status: Detecting', (10,20), cv2.FONT_HERSHEY_SIMPLEX, 0.5, (0,0,255), 2)
cv2.putText(frame, "Total People: {}".format(person-1), (10,30), cv2.FONT_HERSHEY_SIMPLEX, 0.5, (0,0,255), 2)
cv2.imshow("Frame", frame)
return frame
# + [markdown] id="wrkcRVqwiaAB"
# # 2) HumanDetector()
# + [markdown] id="Ef8tfQtfiiXJ"
# **It'll access video via**
# <br>
# 1. WebCamera
# 2. Stored in the local PC.
# + id="QTJvlSqYhV8R"
def human_detection(args):
image_path = args["image"]
video_path = args["video"]
if str(args["camera"])=='true':camera=True
else:camera=False
writer = None
if args["output"] is not None and image_path is None:
writer=cv2.VideoWriter(args['output'],cv2.VideoWriter_fourcc(*'MJPG'),30,(640,480))
if camera:
print("[INFO] opening camera...")
detectByCamera(writer)
elif video_path is not None:
print("[INFO] opening video file (path)...")
detectByPathVideo(video_path,writer)
elif image_path is not None:
print("[INFO] opening image file (path)...")
detectByPathImage(image_path,args["output"])
# + [markdown] id="4eJwbTd1jgYE"
# # 3. Detect By Camera()
# + [markdown] id="19AVnqiAjyKN"
# **Here `cv2.VideoCapture(0)` records the video from webCamera and `video.read()` reads frame by frame.**
# + id="nXX26WMVjoe-"
def detectByCamera(writer):
video=cv2.VideoCapture(0)
print('Detecting by camera...')
while True:
check,frame=video.read()
if writer is not None:
writer.write(frame)
key=cv2.waitKey(1)
if key==ord('q'):
break
video_release()
cv2.destroyAllWindows()
# + [markdown] id="9Re3PlIek1-F"
# # 4. DetectByPathVideo()
# + id="FD04M6Vdkrf3"
def detectByPathVideo(video_path,writer):
video=cv2.VideoCapture(video_path)
check,frame=video.read()
if check==False:
print('[ERROR] Video not found')
return
print('Detecting by video...')
while video.isOpened():
check,frame=video.read()
if check:
frame=imutils.resize(frame,width=min(800,frame.shape[1]))
frame=detect(frame)
if writer is not None:
writer.write(frame)
key=cv2.waitKey(1)
if key==ord('q'):
break
else:
break
video.realease()
cv2.destroyAllWindows()
def detectByCamera(writer):
video=cv2.VideoCapture(0)
print('Detecting by camera...')
while True:
check,frame=video.read()
if writer is not None:
writer.write(frame)
key=cv2.waitKey(1)
if key==ord('q'):
break
video.realease()
cv2.destroyAllWindows()
# + [markdown] id="OqIsySFwlJKV"
# # 5. DetectByPathImage()
# + [markdown] id="O4_Mx4wLladh"
# **This method only works when we have to count the people form the image selected by the user or given by user.**
# + id="0H85P2RIlZix"
def detectByPathImage(image_path,writer):
image=cv2.imread(image_path)
if image is None:
print('[ERROR] Image not found')
return
print('Detecting by image...')
frame=imutils.resize(image,width=min(800,image.shape[1]))
frame=detect(frame)
if writer is not None:
writer.write(frame)
cv2.imshow("Frame",frame)
cv2.waitKey(0)
cv2.destroyAllWindows()
# + [markdown] id="CWahX7h1mdux"
# # 6. Argparse()
# + [markdown] id="3E3e_1oenlrm"
# ***This function simply parses and returns as a dictionary the argumnets passed throug the terminal.***
# 1. Image - Path of image file in the local host.
# 2. Video - Path of video file in the local host.
# 3. Camera - It'll call cameraDetect().
#
# + id="2wg1lVzRnk6K"
def argsParser():
ap=argparse.ArgumentParser()
ap.add_argument('-i','--image',default=None,help='Path to image')
ap.add_argument('-v','--video',default=None,help='Path to video')
ap.add_argument('-c','--camera',default=None,help='Detect by camera')
ap.add_argument('-o','--output',default=None,help='Path to output video')
args=vars(ap.parse_args())
return args
# + [markdown] id="-5tWaLKSoVdC"
# # Main function.
# + id="mDLdUPX2oatJ"
if __name__=="__main__":
HOGCV=cv2.HOGDescriptor()
HOGCV.setSVMDetector(cv2.HOGDescriptor_getDefaultPeopleDetector())
args=argsParser()
human_detection(args)
| Counts number of persons in a given image, video and realtime/counts_number_of_person_in_the_video_and_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
import spacy
nlp = spacy.load('en_core_web_md')
working = pd.read_csv("../data/interim/working.csv")
working = working.drop(['Unnamed: 0'], axis=1)
working.head()
## Note: this notebook takes 30 minutes to run, primarily because of cell 5. Please feel free to skip it if you
## have downloaded the repository, as all of the work done by this notebook has been saved in the export file.
# -
from gensim.models import Phrases
# +
# Generate a list of strings from the database
corpus = working['NARRATIVE'].tolist()
(corpus[0][0:100], corpus[1][0:100], corpus[2][300:400],),
(type(corpus), type(corpus[0]))
# +
# This function will convert a string into a list of unigrams and remove whitespace, punctuation, and stopwords.
def unigrammize(input_text):
for word in nlp.Defaults.stop_words:
for w in (word, word[0].upper() + word[1:], word.upper()):
lex = nlp.vocab[w]
lex.is_stop = True
lower = nlp(input_text.lower())
unigrams = [token.lemma_ for token in lower
if (not token.is_stop) and (not token.is_punct) and (not token.is_space)]
return unigrams
# +
# %%time
# Convert the corpus into a list of lists of unigrams
unigrams = [unigrammize(document) for document in corpus]
# -
# View a portion of the unigrams
[unigrams[0][0:10], unigrams[1][0:10], unigrams[2][0:10]]
# +
# This function adds bigrams and trigrams to a list of list of unigrams without deleting the original unigrams.
def add_trigrams(unigrams):
result = []
bigram_model = Phrases(unigrams, min_count=1, delimiter=b'_')
trigram_model = Phrases(bigram_model[unigrams], min_count=1, delimiter=b'_')
for document in unigrams:
bigrams_ = [b for b in bigram_model[document] if b.count('_') == 1]
trigrams_ = [t for t in trigram_model[bigram_model[document]] if t.count('_') == 2]
merged_ = trigrams_ + bigrams_ + document
result.append(merged_)
return result
# +
# View a sample of the list of lists that includes trigrams
trigrams = add_trigrams(unigrams)
[trigrams[0][0:30], trigrams[1][0:30], trigrams[2][0:30], trigrams[3][0:30]]
# -
# View another sample of the list of lists to make sure it still includes unigrams and not just trigrams
trigrams[0][40:]
# Add the trigrams to the database and save it to disk.
working.loc[:, 'TRIGRAMS'] = trigrams
working.to_csv("../data/interim/trigrams3.csv")
| notebooks/02 Natural Language Processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# [TOC]
# ## 0. 汉诺塔问题
# 
#
# <img src='./pictures/02_hanoi.png' style='zoom:40%'/>
# 当$n=2$时:
# 1. 把小圆盘从A移动到B;
# 2. 把大圆盘从A移动到C;
# 3. 把小圆盘从B移动到C;
#
# 当$n>2$时:
# 1. 把上面$n-1$个盘子从A经C移动到B;
# 2. 把第$n$个圆盘从A移动到C;
# 3. 把$n-1$个小圆盘从B经A移动到C;
def hanoi(n, a, b, c):
'''
n: 剩下盘子数
a: 第一个柱子名
b: 第二个柱子名
c: 第三个柱子名
'''
if n > 0:
hanoi(n-1, a, c, b)
print('moving from %s to %s' % (a, c))
hanoi(n-1, b, a, c)
hanoi(3, 'A', 'B', 'C')
hanoi(2, 'A', 'B', 'C')
# ## 1. 查找算法
# 1.1 顺序查找
# 1.2 二分查找
# ### 1.1 顺序查找(Linear Search)
# 顺序查找: 也叫`线性查找`,从列表第一个元素开始,顺序进行搜索,直到找到元素或搜索到列表最后一个元素为止
# 时间复杂度:$O(n)$
def linear_search(li, target):
for idx, val in enumerate(li):
if val == target:
return idx
return None
idx = linear_search([1,4,2,5], 5)
print(idx)
# ### 1.2 二分查找(Binary Search)
# 二分查找:又叫`折半查找`,从**有序**列表的初始候选区li\[0:n\]开始,通过对待查找的值与候选区中间值的比较,可以使候选区减少一半。
# 时间复杂度:$O(\log{n})$
def bianry_search(li, target):
begin, end = 0, len(li)-1
while end >= begin:
if target == li[(begin+end)//2]:
return (begin+end)//2
elif target < li[(begin+end)//2]:
end = (begin+end)//2 - 1
else:
begin = (begin+end)//2 + 1
return None
# +
idx = bianry_search([1,3,5,6,8,10], 4)
print(idx)
idx = bianry_search([1,3,5,6,8,10], 6)
print(idx)
# -
def bin_search(li, target):
left, right = 0, len(li)-1
while left <= right:
mid = (left + right) >> 1
if target == li[mid]:
return mid
elif target < li[mid]:
right = mid - 1
else:
left = mid + 1
return None
# +
import random
li = [random.randint(0, 9) for _ in range(10)]
li.sort()
print(li)
idx = bin_search(li, 3)
print(idx)
# -
# ### 1.3 内置indx()函数
# 内部用的是线性查找
li = ['a', 'v', 'd', 'g', 'b']
idx = list.index(li, 'g')
print(idx)
li = list(range(15))
idx = list.index(li, 6)
print(idx)
# ## 2. 排序算法
# 2.1 冒泡排序
# 2.2 选择排序
# 2.3 插入排序
# 2.4 快速排序
# 2.5 堆排序
# 2.6 归并排序
# 2.7 希尔排序
# 2.8 计数排序
# 2.9 基数排序
# 2.10 内置sort()
# ### 2.1 冒泡排序(Bubble Sort)
# - 列表每两个相邻的数,如果前面比后面大(默认升序),则交换这两个数;
# - 一趟排序完成后,则无序区减少一个数,有序去增加一个数。
# - 时间复杂度:$O(n^2)$ ,有两层循环
def bubble_sort(li):
for i in range(len(li)-1): # 第i趟比较
for j in range(len(li)-i-1):
if li[j] > li[j+1]:
li[j], li[j+1] = li[j+1], li[j] # 交换两个元素
return li
li = bubble_sort([4,2,7,0,6,8,1,10,0])
li
# 改进1:
# 若一趟比较没有发生元素交换,则认为无序区元素已经有序,可以停止排序
def bubble_sort(li):
for i in range(len(li)-1): # 第i趟比较
exchange = False # 标志位
for j in range(len(li)-i-1):
if li[j] > li[j+1]:
li[j], li[j+1] = li[j+1], li[j] # 交换两个元素
exchange = True
if not exchange:
return li
return li
# +
import random
li = list(range(50))
random.shuffle(li)
print(li)
li = bubble_sort(li)
print(li)
# -
# ### 2.2 选择排序(Select Sort)
def select_sort_simple(li):
sorted_li = []
for i in range(len(li)): # 一共n趟查找
min_val = min(li) # O(n)
sorted_li.append(min_val)
li.remove(min_val) # O(n)
return sorted_li
# +
import random
li = [random.randint(0,100) for i in range(10)]
print(li)
print(select_sort_simple(li))
# -
# 上面的选择排序不推荐:
# 1. 会新生成一个列表,占用内存空间;
# 2. 虽然只有一层循环,但是min()和remove()操作都是$O(n)$,因此总的时间复杂度是$O(n^2)$
def select_sort(li):
for i in range(len(li)-1): # 第i趟选择,共需n-1趟
min_idx = i
for j in range(i+1, len(li)):
if li[j] < li[min_idx]:
min_idx = j
li[i], li[min_idx] = li[min_idx], li[i]
return li
# +
import random
li = [random.randint(0,100) for i in range(10)]
print(li)
print(select_sort(li))
# -
# ### 2.3 插入排序(Insert Sort)
# 1. 初始时手里(有序区)只有一张牌;
# 2. 每次(从无序区)摸一张牌,插入到手里已有牌的正确位置
# 3. 时间复杂度:$O(n^2)$
#
# <img src='./pictures/03_插入排序.gif' style='zoom:40%'/>
def insert_sort(li):
for i in range(1, len(li)): #第i次插入,共需n-1次插入,从第二个元素开始,表示摸到的牌
tmp = li[i] # 摸到的牌
j = i - 1 # 从摸到的牌的前一张牌开始,进行比较和挪动
while j >= 0 and li[j] > tmp:
li[j+1] = li[j]
j -= 1
li[j+1] = tmp
return li
# +
import random
li = [random.randint(0,20) for i in range(10)]
print(li)
print(insert_sort(li))
# +
import time
li = list(range(10000))
random.shuffle(li)
begin_time = time.time()
print(bubble_sort(li))
end_time = time.time()
print(end_time - begin_time)
# -
# ### 2.4 快速排序
# 思想:
# 1. 取一个元素p(第一个元素),使元素p归位;
# 2. 列表被p分成两部分,左边都比p小,右边都比p大;
# 3. 递归完成排序。
#
# 时间复杂度:$\color{red}{n\log n}$
# <img src='./pictures/04_quick_sort.gif' style='zoom:40%' align='left'></img>
def partition1(li, left, right):
'''
归位函数
'''
tmp = li[left] # 缓存第一个元素,也就是将第一个元素归位
flag = True
while left < right:
if li[right] < tmp and flag:
li[left] = li[right]
left += 1
flag = False
elif li[right] >= tmp and flag:
right -= 1
elif li[left] < tmp and not flag:
left += 1
elif li[left] >= tmp and not flag:
li[right] = li[left]
right -= 1
flag = True
li[left] = tmp
return left
def partition(li, left, right):
tmp = li[left]
while left < right:
while right > left and li[right] >= tmp:
right -= 1
li[left] = li[right]
while left < right and li[left] <= tmp:
left += 1
li[right] = li[left]
li[left] = tmp
return left
li = [random.randint(0,20) for i in range(15)]
li2 = li
print(li)
partition1(li,0,len(li)-1)
partition(li2,0,len(li2)-1)
print(li)
print(li2)
# +
def _quick_sort(li, left, right):
'''
利用递归实现快速排序
'''
if left < right:
mid = partition(li, left, right)
_quick_sort(li, left, mid-1)
_quick_sort(li, mid+1, right)
def quick_sort(li):
_quick_sort(li, 0, len(li)-1)
# +
import random
li = list(range(10))
random.shuffle(li)
print(li)
quick_sort(li)
print(li)
# +
import time
from copy import deepcopy
li = list(range(10000))
random.shuffle(li)
l1 = deepcopy(li)
l2 = deepcopy(li)
begin_time = time.time()
bubble_sort(l1)
end_time = time.time()
print('冒泡排序时间:', end_time - begin_time)
begin_time = time.time()
quick_sort(l2)
end_time = time.time()
print('快速排序时间:', end_time - begin_time)
# -
# **快速排序的两个问题:**
# 1. 最坏情况:
# > 当待排序列表是逆序列表时,不能很好的partition,此时时间复杂度为$O(n^2)$;
# > **解决办法:**随机选择一个位置的元素和起始位置元素交换,开始快排
#
# 2. 递归
# > python解释器为了避免内存溢出和性能影响,设置了`最大递归深度为998`,当调用栈超过998层就会报错;
# > **解决办法:** 手动修改最大递归深度,sys.setrecursionlimit(3000)
# ### 2.5 堆排序(Heap Sort)
# **树**
#
# <img src='./pictures/05_树.jpg' style='zoom:70%'/>
# 1. 根节点:A
# 2. 树的深度(高度):4
# 3. <font color=red>树的度:</font>6 `树中最多子节点的个数`
# 4. 孩子节点/父节点
# 5. 子树
# **二叉树**
# 1. 度不超过2的树;
# 2. 每个节点最多有2个孩子节点;
# 3. 两个孩子节点被区分为`左孩子节点`和`右孩子节点`
#
# <img src='./pictures/06_二叉树.jpg' style='zoom:45%'/>
# **满二叉树**
#
# 一个二叉树,如果每一层的结点数都达到最大值,则这个二叉树就是满二叉树
#
# <img src='./pictures/07_满二叉树.jpg' style='zoom:45%'/>
# **完全二叉树**
#
# 叶节点只能出现在最下层和次下层,并且最下面一层的节点都集中在该层最左边的若干位置的二叉树
#
# <img src='./pictures/08_完全二叉树.jpg' style='zoom:70%'/>
# **二叉树的存储方式**
#
# 1. <font color=red>链式存储方式</font>
#
# 2. <font color=red>顺序存储方式</font>
# **二叉树的顺序存储方式:**
#
# <img src='./pictures/09_顺序存储二叉树.png' style='zoom:40%'/>
# 1. 父节点与左孩子节点的编号下标关系:
# i --> 2i+1
#
# 2. 父节点与右孩子节点的编号下标关系:
# i --> 2i+2
# **大根堆、小根堆**
# 1. 一棵`完全二叉树`,满足任一节点都比其孩子节点大;
# 1. 一棵`完全二叉树`,满足任一节点都比其孩子节点小;
#
# <img src='./pictures/10_大根堆_小根堆.jpg' style='zoom:50%'/>
# **堆的向下调整:**
#
# 假设根节点的左右子树都是堆,但根节点不满足堆的性质,可以通过一次向下的调整来将其变成一个堆。
#
# <img src='./pictures/11_堆的向下调整.jpg' style='zoom:50%'/>
# #### 2.5.1 堆排序过程
# 1. 构造堆
# <font color=red>从下往上构建大根堆</font>
#
#
# 2. 挨个出数
# - 将根节点元素拿出;
# - <font color='DarkOrchid'>将最后一个叶节点挪到根节点位置</font>;
# - 运用堆的向下调整,实现大根堆;
# - 重复上面的操作,直至所有元素被排好序
#
# **堆排序过程:**
#
# <img src='./pictures/12_堆排序.gif' style='zoom:70%' align='center'/>
#
# **构建大根堆**
#
# <img src='./pictures/13_堆构造.png' style='zoom:50%' align='center'/>
def sift(li, low, high):
'''
堆的向下调整
low: 堆的根节点位置
high: 堆最后一个元素位置
'''
i = low # 根节点
j = 2 * i + 1 # 左子节点
tmp = li[i]
while j <= high:
# 判断右子节点
if j+1 <= high and li[j+1] > li[j]: # 右子节点存在,且大于左子节点
j += 1
if li[j] > tmp: # 子节点 比 根节点大
li[i] = li[j]
i = j
j = 2 * i + 1
else:
break
else:
li[i] = tmp
# <img src='./pictures/11_堆的向下调整.jpg' style='zoom:50%'/>
li = [2,9,7,8,5,0,1,6,4,3]
sift(li,0,len(li)-1)
print(li)
def heap_sort(li):
'''
1. 构造大根堆
2. 将根节点元素取出
3. 将最后一个叶子节点挪到根节点位置
4. 堆的向下调整,实现大根堆
5. 重复上面2~4步,直到所有元素已排好序
'''
# 1. 构造大根堆
n = len(li)
idx = (n-1-1)//2 # 获取最后一个非叶子节点编号
for i in range(idx, -1, -1):
sift(li, i, n-1) # 始终将n-1作为high
for i in range(n-1, 0, -1):
# 2. 将根节点元素取出,和第i个节点交换元素
li[0], li[i] = li[i], li[0]
# 3. 对剩下的堆进行向下调整
sift(li, 0, i-1)
# +
import random
li = [random.randint(0, 15) for _ in range(10)]
print(li)
heap_sort(li)
print(li)
print()
li = [random.randint(0, 15) for _ in range(9)]
print(li)
heap_sort(li)
print(li)
# -
# Heap Sort时间复杂度:
# sift()的时间复杂度为$O(\log{n})$
# 堆排序时间复杂度:$O(n\log{n})$
# 堆的内置模块heapq
# +
import heapq
import random
li = list(range(24))
random.shuffle(li)
print(li)
# 建堆,默认是小根堆
heapq.heapify(li)
print(li)
# 依次弹出元素
for i in range(len(li)):
print(heapq.heappop(li), end=', ')
# -
# #### 2.5.2 topk问题
# 现有n个数,设计算法得到前k大的数。(k<n)
# 解决思路:
# 1. 排序后切片:$O(n\log{n})$
# 2. 排序LowB三人组(冒泡、插入、选择):$O(kn)$
# 3. <font color=red>堆排序思路:</font>$\color{red}{O(n\log{k})}$
#
# **堆排序实现topk解决思路:**
# 1. 取列表前k个元素建立一个小根堆,则堆顶便是目前第k大的数;
# 2. 依次向后遍历原列表,对于列表中的每个元素,如果小于堆顶,则忽略该元素;如果大于堆顶,则将该元素更换为堆顶元素,并对堆进行一次调整;
# 3. 遍历列表所有元素,倒序弹出堆顶。
# +
'''
堆排序实现topk
'''
def sift(li, low, high):
'''
向下调整为小根堆
'''
i = low # i代表子树的根
j = 2 * i + 1 # 左子节点
tmp = li[low]
while j <= high:
if j+1 <= high and li[j+1] < li[j]:
j += 1 # 切到右子节点
if li[j] < li[i]:
li[i], li[j] = li[j], li[i]
i = j
j = 2 * i + 1
else:
break
def topk(li, k):
'''
获取li列表中前k大的数
'''
heap = li[:k]
# 1. 取前k个元素,构建小根堆
i = (k - 2) // 2 # 最后一个非叶子节点
for j in range(i, -1, -1):
sift(heap, j, k-1)
# 2. 遍历剩余len(li)-k个元素
for i in range(k, len(li)):
if li[i] > heap[0]: # 若第k个元素比堆顶元素大,则替换调
heap[0] = li[i]
sift(heap, 0 ,k-1)
# 3. 倒序取出前k个数
for i in range(k-1, -1, -1):
heap[0], heap[i] = heap[i], heap[0]
sift(heap, 0, i - 1 )
return heap
# +
import random
li = [random.randint(0,50) for i in range(10)]
# li = [25, 25, 2, 14, 1, 15, 1, 18]
print('原列表:', li, sep='\n')
top = topk(li, 4)
print(top)
# -
# ### 2.6 归并排序
# <img src='./pictures/14_归并排序.webp' style='zoom:100%' align='left'/>
# **归并**
# 假设现在的列表分两段有序,如何将其合成为一个有序列表?
#
# <img src='./pictures/15_一次归并.png' style='zoom:50%'></img>
#
#
# 这种操作称为一次归并
def merge(li, low, mid, high):
'''
一次归并操作
'''
i = low
j = mid + 1
new_li = []
while i <= mid:
if j <= high:
if li[i] <= li[j]:
new_li.append(li[i])
i += 1
else:
new_li.append(li[j])
j += 1
else:
new_li.extend(li[i:mid+1])
break
if j <= high:
new_li.extend(li[j:high+1])
# 把new_li重新写会li,后面递归会用到
li[low:high+1] = new_li
li = [1,3,4,6,2,5,7,8,9]
merge(li,0,3,len(li)-1)
print(li)
# <img src='./pictures/16_归并排序.jpg' style='zoom:40%'/>
#
# 时间复杂度:$O(n\log{n})$
# > 有$\log{n}$层,每一层都需遍历,所以是$n\log{n}$
#
# 空间复杂度:$O(n)$
# > 在merge过程中有新开辟一个列表,所以归并排序不是原地排序,空间复杂度为$O(n)$
def merge_sort(li, low, high):
'''
归并排序: 递归实现
'''
if low < high:
mid = (low + high) // 2
merge_sort(li, low, mid) # 左边部分
merge_sort(li, mid+1, high) # 右边部分
merge(li, low, mid, high) # 归并左右两边
li = [8,4,5,7,1,3,6,2]
merge_sort(li, 0, len(li)-1)
print(li)
# ### NB三人组总结
# 1. 三种排序算法(快速排序、堆排序、归并排序)的时间复杂度都是$n\log{n}$;
# 2. 一般情况下,就运行时间而言:
# 快速排序 < 归并排序 < 堆排序
# 3. 三种排序算法的缺点:
# - 快速排序:极端情况下排序效率低(比如原始列表为倒序)
# - 归并排序:需要额外的内存开销;
# - 堆排序:在快的排序算法中相对较慢
#
# <img src='./pictures/17_排序算法总结.jpg' style='zoom:60%'/>
#
# 快速排序空间复杂度说明:
# > 快排中涉及到递归调用,递归调用会给函数开辟空间。快排中平均递归深度为$\log{n}$,最坏情况递归深度为$n$
#
# 稳定性说明:
# > 如果是相邻位置元素比较交换,则是稳定的,比如冒泡排序;
# > 如果是隔着位置对元素进行比较交换,则是不稳定的,比如选择排序。0,2,4,2,1,4
#
# python内置排序是改进的归并排序,因为它是稳定的
# ### 2.7 希尔排序
#
# 1. 希尔排序(Shell Sort)是一种`分组插入排序`算法;
#
# 2. 首先取一个整数$d_{1} = n / 2$,将元素分为$d_{1}$个组,每组相邻元素之间距离为$d_{1}$,在各组内进行直接插入排序;
#
# 3. 取第二个整数$d_{2} = d_{1} / 2$,重复上述分组排序过程,直到$d_{i}=1$,即所有元素在同一组内进行直接插入排序;
#
# 4. 希尔排序每趟并不使某些元素有序,而是使整体数据越来越接近有序;最后一趟排序使得所有数据有序。
#
# <img src='./pictures/18_Shell_Sort.gif' style='zoom:80%'/>
# +
def insert_sort_gap(li, gap):
'''
分组插入排序
gap: 组间距
'''
for i in range(gap, len(li)):
tmp = li[i]
j = i - gap
while j >= 0 and li[j] >= tmp:
li[j+gap] = li[j] # 因为第j位置元素比tmp大,所以将该位置元素往后挪gap位
j -= gap
li[j+gap] = tmp # while终止条件1)j<0; 2)li[j]<tmp 所以需将tmp写回到li[j+gap]位置上
def shell_sort(li):
'''
希尔排序
'''
gap = len(li) // 2
while gap >= 1:
insert_sort_gap(li, gap)
gap //= 2
# -
li = [3,5,2,0,7,6,4,1]
shell_sort(li)
print(li)
# **希尔排序时间复杂度:**
# 希尔排序的时间复杂度与gap序列的选取有关。
# ### 2.8 计数排序
# **排序前提:**
#
# `已知列表中数据的范围`,比如都在0到100之间,设计时间复杂度为$O(n)$的算法。
# <img src='./pictures/19_计数排序.gif' style='zoom:60%'/>
def count_sort(li, max_count=100):
'''
计数排序
max_count: 元素的最大值
数值范围:[0,max_count]
'''
count = [0 for _ in range(max_count+1)]
for val in li:
# 计数
count[val] += 1
li.clear()
for idx, val in enumerate(count):
for i in range(val):
li.append(idx)
# +
import random
li = [random.randint(0,20) for i in range(15)]
print(li)
count_sort(li)
print(li)
# -
# ### 2.9 桶排序(Bucket Sort)
#
# 1. 在计数排序中,如果元素的范围比较大(比如1到1亿之间),我们需要开辟一个很大的列表,如何改造算法?
#
# **桶排序(Bucket Sort):**首先将元素分在不同的桶中,再对桶中的元素进行排序,最后按桶的顺序,依次输出所有元素,即得到有序数列。
#
# <img src='./pictures/20_桶排序.png' style='zoom:60%'/>
def bucket_sort(li, n=100, max_num=10000):
'''
桶排序
n: 桶的个数
max_num: 最大的元素
'''
# 1. 创建n个桶
buckets = [[] for _ in range(n)]
for val in li:
# 2. 将元素依次放入对应的桶中
idx = min(val // (max_num // 100), n-1) # min()的作用,将max_num元素放入最后一个桶,因为索引越界了
buckets[idx].append(val)
# 3. 放入的同时,对桶内元素进行插入排序
for i in range(len(buckets[idx])-1, 0, -1):
# 1,3,4,5,2
if buckets[idx][i-1] > buckets[idx][i]:
buckets[idx][i-1], buckets[idx][i] = buckets[idx][i], buckets[idx][i-1]
else:
break
# 4. 将所有桶中元素依次取出
sorted_li = []
for buc in buckets:
sorted_li.extend(buc)
return sorted_li
# +
import random
li = [random.randint(0,100000) for _ in range(10000)]
sorted_li = bucket_sort(li, max_num=100000)
print(sorted_li)
# -
# **桶排序性能总结:**
# 1. 桶排序的表现取决于数据的分布。也就是需要对不同的数据排序时采用不同的分桶策略。(比方说,数据是均匀分布的,桶的大小可以一样;如果数据是正态分布的,那均值附近应分得更精细);
# 2. 平均时间复杂度:$O(n+k)$;
# 3. 最坏情况时间复杂度:$O(n^{2}k)$
# 4. 空间复杂度:$O(nk)$
# ### 2.10 基数排序
#
# **多关键字排序:**
#
# 比如现有一员工表,要求按照年龄排序,年龄相同的按照薪资进行排序。
#
# **基数排序:**
#
# 对数据的排序也可以看做是多关键字排序,先按个位排,然后按十位排,依次进行
#
# <img src='./pictures/21_基数排序.gif' style='zoom:60%'/>
def radix_sort(li):
max_num = max(li) # 获取最大元素,决定进行几次桶排序
length = len(str(max_num))
buckets = [[] for _ in range(10)] # 创建10个桶
for i in range(length):
for val in li:
# 将元素放入对应的桶中
i_val = (val % (10**(i+1)))//(10**i)
buckets[i_val].append(val)
# 将桶中元素拿出
li.clear()
for buc in buckets:
li.extend(buc)
buc.clear() # 拿出后记得清空桶
# +
import random
li = [random.randint(0,200) for _ in range(15)]
print(li)
radix_sort(li)
print(li)
# -
# **基数排序性能**
# 1. 时间复杂度:$O(kn)$,其中k表示最大元素的位数;
# 2. 空间复杂度:$O(k+n)$
#
# 与NB三人组比较:
# $k = math.floor(math.log(10, length))$ 以10为底,length为数字的位数;
# $n\log{n}$,其中$\log{n} = math.log(2,n)$ 以2为底,n为元素个数
#
# 所以,如果元素位数小,个数多,基数排序要比NB三人组快。
# ## 3. 数据结构
# 3.1 列表
# 3.2 栈
# 3.3 队列
# 3.4 链表
# 3.5 哈希表
# 3.6 树
# ### 3.1 数据结构介绍
#
# 数据结构按照其逻辑结构可分为**线性结构**、**树结构**、**图结构**
# - 线性结构:数据结构中的元素存在一对一的相互关系;
# - 树结构:数据结构中的元素存在一对多的关系;
# - 图结构:数据结构中的元素存在多对多的关系。
# ### 3.2 列表
#
# 列表(其它语言称数组)是一种基本数据类型。
# 关于列表的问题:
# - 列表中的元素是如何存储的?
# - 列表的基本操作:按下标查找、插入元素、删除元素...
# - 这些操作的时间复杂度是多少?
# - Python的列表是如何实现的?
# 其它语言中的数组与列表的两点不同:
#
# 1. 数组中的元素类型必须相同,而Python中列表元素类型无限制;
# 2. 创建数组时需指定长度,而Python列表无需指定长度。
#
# <img src='./pictures/22_数组与列表.png' style='zoom:50%'/>
# 1. 数组连续内存空间中存的是元素,查找元素时通过计算地址值直接得到元素,因此元素类型需相同(即所占内存空间长度相同);
# 2. 列表连续内存空间中存的是元素的地址值,因而元素类型可以不相同;
# 3. 按下标查找元素操作是$O(1)$;
# 4. 插入和删除元素操作是$O(n)$,因为涉及挪动元素。
# ### 3.3 栈
# 栈(Stack)是一个数据集合,可以理解为只能在一端进行插入或删除操作的列表。
#
# 栈的特点:后进先出LIFO(Last In,First Out)
#
# 栈的概念: 栈顶、栈底
#
# 栈的基本操作:
# - 进栈(压栈):push
# - 出栈:pop
# - 取栈顶:gettop(只看不取)
class Stack():
def __init__(self):
self.stack = []
def push(self, element):
self.stack.append(element)
def pop(self):
return self.stack.pop()
def get_top(self):
if len(self.stack) > 0:
return self.stack[-1]
else:
return None
def is_empty(self):
return len(self.stack) == 0
# +
stack = Stack()
stack.push(1)
stack.push(1)
stack.push('good')
stack.push(9)
print(stack.get_top())
print(stack.pop())
print(stack.get_top())
# -
# **括号匹配问题:**
# 1. (){}\[()\] 匹配
# 2. [(]) 不匹配
# 3. {}) 不匹配
def brace_match(string):
'''
用栈来解决括号匹配问题
'''
braces = {'}':'{',']':'[',')':'(','>':'<'}
stack = Stack()
for s in string:
top = stack.get_top()
if s in braces.values(): # 如果是左括号,直接放入栈顶
stack.push(s)
continue
elif stack.is_empty(): # 如果不是左括号,且栈为空,返回False
return False
elif top == braces[s]: # 如果匹配,则弹出
stack.pop()
else:
return False
if stack.is_empty():
return True
else:
return False
s1 = '{}()'
s2 = '[)]'
s3 = '{(})'
s4 = '{([{()}])}'
print(brace_match(s1))
print(brace_match(s2))
print(brace_match(s3))
print(brace_match(s4))
# ### 3.4 队列
# 1. 队列(Queue)是一个数据集合,仅允许在列表的一端进行插入,另一端进行删除;
# 2. 进行插入的一端称为队尾(rear),插入动作称为进队或入队;
# 3. 进行删除的一端称为对头(front),删除操作称为出队;
# 4. 队列的性质:先进先出FIFO(First-in,First-out)
#
# <img src='./pictures/23_环形队列.jpeg' style='zoom:70%'/>
# 环形队列:当队尾指针front == Maxsize - 1时,再前进一个位置就自动到0.
# - 队首指针前进1:front = (front + 1) % Maxsize
# - 队尾指针前进1:rear = (rear + 1) % Maxsize
# - 队空条件:rear == front
# - 队满条件:(rear + 1) % Maxsize == front
class Queue():
def __init__(self, size=100):
self.queue = [0 for _ in range(size)]
self.size = size
self.rear = 0 # 队尾
self.front = 0 # 队首
def push(self, element):
if (self.rear + 1) % self.size == self.front:
raise Exception('队已满')
self.rear = (self.rear + 1) % self.size
self.queue[self.rear] = element
def pop(self):
if self.rear == self.front:
return None
self.front = (self.front + 1) % self.size
return self.queue[self.front]
def is_empty(self):
return self.rear == self.front
def is_full(self):
return (self.rear + 1) % self.size == self.front
# +
queue = Queue(12)
queue.push(0)
queue.push(1)
queue.push(2)
queue.push(3)
queue.push(4)
queue.push(5)
queue.push(6)
queue.push(7)
queue.push(8)
queue.push(9)
queue.push(10)
# queue.push(11)
# queue.push(12)
print(queue.pop())
print(queue.pop())
print(queue.is_empty())
# -
# **双向队列:**两端都支持进队和出队操作
#
# Python内置队列模块
# from collections import deque
# - 创建队列:queue = deque()
# - 进队:append()
# - 出队:popleft()
# - 双向队列首进队:appendleft()
# - 双向队列尾出队:pop()
# +
from collections import deque
# 同时支持单向和双向队列
queue = deque()
queue.append(12) # 队尾进队
queue.append(1024) # 队尾进队
print(queue.popleft()) # 队首出队
# 双向队列操作
queue.appendleft(99) # 队首进队
print(queue.pop()) # 队尾出队
# 当队列满了后,deque会自动将队首元素出队
queue2 = deque([1,2,3,4,5,6,7,8,9], 5)
print(queue2.popleft())
# -
# **迷宫问题:**
#
# <img src='./pictures/24_迷宫问题.jpg' style='zoom:50%'/>
# **栈——深度优先搜索算法**
#
# 1. 又叫`回朔法`
# 2. 思路:从一个节点开始,任意找下一个能走的点,当找不到能走的点时,退回上一个点寻找是否有其它方向的点。
# 3. 使用栈存储当前路径。
# +
maze = [
[1,1,1,1,1,1,1,1,1,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,0,0,1,1,0,0,1],
[1,0,1,1,1,0,0,0,0,1],
[1,0,0,0,1,0,0,0,0,1],
[1,0,1,0,0,0,1,0,0,1],
[1,0,1,1,1,0,1,1,0,1],
[1,1,0,0,0,0,0,0,0,1],
[1,1,1,1,1,1,1,1,1,1],
]
def path_search(x1,y1,x2,y2):
'''
x1,y1: 起始点坐标
x2,y2: 终点坐标
'''
stack = []
stack.append((x1,y1))
px = x1
py = y1
while px != x2 or py != y2:
# 按 上——>右——>下——>左 的步骤搜索
if maze[px-1][py] != 1 and (px-1, py) not in stack:
px -= 1
stack.append((px, py))
elif maze[px][py+1] != 1 and (px, py+1) not in stack:
py += 1
stack.append((px, py))
elif maze[px+1][py] != 1 and (px+1, py) not in stack:
px += 1
stack.append((px, py))
elif maze[px][py-1] != 1 and (px, py-1) not in stack:
py -= 1
stack.append((px, py))
else:
maze[px][py] = 1
stack.pop()
px, py = stack[-1]
if px == x1 and py == y1:
raise Exception('No path available!')
return stack
# +
maze = [
[1,1,1,1,1,1,1,1,1,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,0,0,1,1,0,0,1],
[1,0,1,1,1,0,0,0,0,1],
[1,0,0,0,1,0,0,0,0,1],
[1,0,1,0,0,0,1,0,0,1],
[1,0,1,1,1,0,1,1,0,1],
[1,1,0,0,0,0,0,0,0,1],
[1,1,1,1,1,1,1,1,1,1],
]
print(path_search(1,1,8,8))
# -
# **队列——广度优先搜索**
#
# 广度优先搜索的路径是最短的。
#
# <img src='./pictures/25_迷宫问题_队列解决.jpg' style='zoom:45%'/>
# +
from collections import deque
maze = [
[1,1,1,1,1,1,1,1,1,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,1,0,0,0,1,0,1],
[1,0,0,0,0,1,1,0,0,1],
[1,0,1,1,1,0,0,0,0,1],
[1,0,0,0,1,0,0,0,0,1],
[1,0,1,0,0,0,1,0,0,1],
[1,0,1,1,1,0,1,1,0,1],
[1,1,0,0,0,0,0,0,0,1],
[1,1,1,1,1,1,1,1,1,1],
]
dirs = [
lambda x,y:(x-1, y), # 上
lambda x,y:(x, y+1), # 右
lambda x,y:(x+1, y), # 下
lambda x,y:(x, y-1) # 左
]
def output_path(path):
'''
输出路径
'''
node = path[-1] # 取出终点
out_path = []
while node[2] != -1:
out_path.append(node[0:2])
node = path[node[2]]
out_path.append(path[0][0:2])
out_path.reverse()
return out_path
def maze_path(x1,y1,x2,y2):
'''
x1,y1: 起始点坐标
x2,y2: 终点坐标
'''
queue = deque() # 记录搜索到的最新结点
queue.append((x1, y1, -1)) # 把起点加入队列, 第3个元素表示该节点的上一节点在path中的索引
path = [] # 记录历史路径的列表
while len(queue) != 0: # 只要队列里还有值,就代表还有路可走
cur_node = queue.popleft() # 取出当前节点
path.append(cur_node) # 将当前节点放入历史路径列表中
if cur_node[0] == x2 and cur_node[1] == y2: # 表示已走到终点
output = output_path(path)
return output
for next_node in dirs:
next_x, next_y = next_node(cur_node[0], cur_node[1])
if maze[next_x][next_y] == 0: # 路是通的
# 将next_node节点入队
queue.append((next_x, next_y, len(path)-1))
# 将next_node标记为已走过
maze[next_x][next_y] = 2
else:
raise Exception('无路可走了!')
# -
path = maze_path(1,1,8,8)
for p in path:
print(p)
# ### 3.5 链表
#
# 链表是由一系列节点组成的元素的集合。每个节点包含两部分,数据域item和指向下一节点的指针next。通过节点之间的相互连接,最终串成一个链表。
#
# <img src='./pictures/26_链表.jpg' style='zoom:100%'/>
class Node():
def __init__(self, item):
self.item = item
self.next = None
# +
node1 = Node(11)
node2 = Node(22)
node3 = Node(33)
node1.next = node2
node2.next = node3
print(node1.item)
print(node1.next.item)
print(node1.next.next.item)
# -
# #### 3.5.1 链表的创建和遍历
# **头插法和尾插法:**
#
# <img src='./pictures/27_头插法和尾插法.png' style='zoom:50%'/>
# +
def creat_linklist_head(li):
'''
头插法创建链表,
遍历链表得到的是倒序的
'''
head = Node(li[0])
for ele in li[1:]:
node = Node(ele)
node.next = head
head = node
return head
def creat_linklist_tail(li):
'''
尾插法创建链表
遍历链表得到的是倒序的
'''
head = Node(li[0])
tail = head
for ele in li[1:]:
node = Node(ele)
tail.next = node
tail = node
return head
def print_linklist(head):
'''
遍历打印链表
'''
while head and head.next:
print(head.item, end = ', ')
head = head.next
else:
print(head.item)
# +
lk = creat_linklist_head([1,2,3,4,5,6])
print_linklist(lk)
lk = creat_linklist_tail([1,2,3,4,5,6])
print_linklist(lk)
# -
# #### 3.5.2 链表的插入和删除
#
# <img src='./pictures/28_链表的插入和删除.png' style='zoom:50%'/>
# +
def insert_node(cur_node, new_node):
'''
在cur_node之后插入新节点new_node
'''
new_node.next = cur_node.next
cur_node.next = new_node
def del_node(cur_node):
'''
将cur_node的后一节点从链表中删除
'''
p = cur_node.next
cur_node.next = p.next
del p
# +
lk = creat_linklist_tail([1,2,3,4,5,6,7])
print_linklist(lk)
cur_node = lk.next
new_node = Node(88)
insert_node(cur_node, new_node)
print_linklist(lk)
del_node(cur_node)
print_linklist(lk)
# -
# #### 3.5.3 链表总结
#
# | 操作 | 顺序表 | 链表 |
# | ---- | ---- | ---- |
# | 按元素查找 | <img width=100/> $O(n)$ | <img width=100/> $O(n)$ |
# | 按下标查找 | $O(1)$ | $O(n)$ |
# | 在某元素后插入 | $O(n)$ | $O(1)$ |
# | 删除某元素 | $O(n)$ | $O(1)$ |
#
# 1. 链表的插入和删除操作明显优于顺序表;
# 2. 链表的内存空间可以更灵活的分配。
# ### 3.6 哈希表(散列表)
# 哈希表是一个通过`哈希函数`来计算数据存储位置的数据结构,通常支持如下操作:
# - insert(key, value)
# - get(key)
# - delete(key)
#
# **直接寻址表:**
#
# U是所有可能key的集合
#
# <img src='./pictures/29_直接寻址表.jpeg' style='zoom:100%'/>
# 直接寻址技术缺点:
# 1. 当域$U$很大时,创建$T$列表需要大量内存,很不实际;
# 2. 如果$U$很大,而实际$K$很少,则大量空间被浪费;
# 3. 无法处理关键字不是数字的情况。
#
# 改进直接寻址表:<font color=red>哈希(Hashing)</font>
# - 构建大小为$m$的寻址表$T$;
# - key为k的元素放到$h(k)$位置上;
# - $h(k)$是一个函数,其将域$U$映射到表T\[0, 1, ..., m-1\]
#
# **哈希表**
#
# 1. 哈希表(Hash Table, 又称散列表),是一种线性表的存储结构。哈希表由一个<font color=red>直接寻址表</font>和一个<font color=red>哈希函数</font>组成。哈希函数$h(k)$将关键字k作为自变量,返回元素的存储下标。
#
# 2. 假如有一个长度为7的哈希表,哈希函数 h(k) = k%7. 元素集合{14, 22, 3, 5}的存储方式如下图:
#
# <img src='./pictures/30_哈希表示例.png' style='zoom:70%'/>
# **<font color=red>哈希冲突</font>**
# 1. 哈希表的大小是有限的,而要存的值的总数量是无限的,因此对于任何哈希函数,都会出现两个不同元素映射到同一位置的情况,称为哈希冲突;
# 2. 比如 h(k)=k%7 ,h(0)=h(7)=h(14)=...
#
# **解决哈希冲突——开放寻址法**
#
# 如果哈希函数返回的位置已经有值,则可以向后探查新的位置来存储这个值。
# - 线性探查:如果位置i被占用,则依次向后探查i+1, i+2, ...直到找到空位,进行存储;
# - 二次探查:如果位置i被占用,则探查i+1², i-1², i+2², i-2², ...
# - 二度哈希:有n个哈希函数,当使用第1个哈希函数n1发生冲突时,则尝试使用h2, h3, ...
#
#
# **解决哈希冲突——拉链法**
#
# 哈希表每个位置都连接一个链表,当冲突发生时,冲突元素将被加到该位置链表的最后。
#
# <img src='./pictures/31_哈希冲突_拉链法.jpeg' style='zoom:70%'/>
class Linklist():
class Node():
def __init__(self, item):
self.item = item
self.next = None
class LinklistIterator():
def __init__(self, node):
self.node = node
def __next__(self):
if self.node:
cur_node = self.node
self.node = cur_node.next
return cur_node.item
else:
raise StopIteration
def __iter__(self):
return self
def __init__(self, iterable=None):
self.head = None
self.tail = None
if iterable:
self.extend(iterable)
def extend(self, iterable):
for obj in iterable:
self.append(obj)
def append(self, obj):
node = Linklist.Node(obj)
if not self.head: # 链表为空
self.head = node
self.tail = node
else: # 追加节点
self.tail.next = node
self.tail = node
def find(self, obj):
for n in self:
if n == obj:
return True
else:
return False
def __iter__(self):
return self.LinklistIterator(self.head)
def __repr__(self):
return "<" + ", ".join(map(str, self)) + ">"
lk = Linklist([1, 2, 3, 4, 5])
print(lk)
class HashTable():
def __init__(self, size=101):
self.size = size
self.T = [Linklist() for _ in range(size)]
def h(self, k):
return k % self.size
def insert(self, k):
h = self.h(k)
# 不能重复插入
if self.find(k):
raise Exception('不能重复插入')
else:
self.T[h].append(k)
def find(self, k):
h = self.h(k)
return self.T[h].find(k)
# +
table = HashTable()
table.insert(0)
table.insert(101)
table.insert(4)
# table.insert(4)
print(table.T)
print(table.find(0))
print(table.find(1))
# -
# **哈希表的应用——md5算法**
#
# MD5(Message-Digest Algorithm 5)**曾经**是密码学中常用的哈希函数,可以把任意长度的数据映射为128位的哈希值,其曾经包含如下特征:
# - 同样的消息,其MD5值必定相同;
# - 可以快速计算出任意给定消息的MD5值;
# - 除非暴力枚举,否则不能根据哈希值反推出消息本身;
# - 两条消息之间即使只有微小的差别,其对应的MD5值也应是完全不同,完全不相关的;
# - 不能在有意义的时间内人工构造两个不同的消息,使得其具有相同的MD5值。
#
# 应用举例: 文件的哈希值
# 算出两个文件的哈希值,若两个文件的哈希值相同,则可认为这两个文件是相同的(哈希值相同,文件不同的概率太太太低),因此:
# - 用户可以利用它来验证下载的文件是否完整;
# - 云存储服务商可以利用它来判断用户要上传的文件是否已存在服务器上,从而实现秒传的功能,同时避免存储过多相同文件的副本。
#
#
# **哈希表的应用——SHA2算法**
# 1. 历史上MD5和SHA-1曾经是使用最为广泛的cryptographic hash function,但是随着密码学的发展,这两个哈希函数的安全性相继受到了各种挑战。
# 2. 因此现在安全性较重要的场合<font color=red>推荐使用SHA-2</font>等新的更安全的哈希函数;
# 3. SHA-2包含了一系列的哈希函数:SHA-224, SHA-256, SHA-384, SHA-512, SHA-512/224, SHA-512/256, 其对应的哈希值长度分别为224,256,384或512位。
# 4. SHA-2具有和MD5类似的性质。
# ### 3.7 树
# #### 3.7.1 二叉树
#
# 二叉树的链式存储:将二叉树的节点定义为一个对象,节点之间通过类似链表的连接方式来连接。
#
# <font color=red>因为二叉树可能不是完全二叉树,因此用列表存储不方便</font>
#
# <img src='./pictures/32_二叉树链式存储.jpeg' style='zoom:50%'/>
# +
class BiTreeNode():
def __init__(self, data):
self.data = data
self.lchild = None
self.rchild = None
a = BiTreeNode('A')
b = BiTreeNode('B')
c = BiTreeNode('C')
d = BiTreeNode('D')
e = BiTreeNode('E')
f = BiTreeNode('F')
g = BiTreeNode('G')
a.lchild = b
a.rchild = f
b.lchild = c
b.rchild = d
f.rchild = g
d.lchild = e
print(a.lchild.rchild.data)
# -
# #### 3.7.2 二叉树的遍历
#
# <img src='./pictures/32_二叉树链式存储.jpeg' style='zoom:40%'/>
#
# 二叉树的遍历方式:
# - 前序遍历:ABCDEFG 根-左-右
# - 中序遍历:CBEDAFG 左-根-右
# - 后续遍历:CEDBGFA 左-右-根
# - 层次遍历:ABFCDGE
# +
def pre_order(root):
'''
前序遍历
'''
if root:
print(root.data, end=', ')
pre_order(root.lchild)
pre_order(root.rchild)
def in_order(root):
'''
中序遍历
'''
if root:
in_order(root.lchild)
print(root.data, end=', ')
in_order(root.rchild)
def post_order(root):
'''
后序遍历
'''
if root:
post_order(root.lchild)
post_order(root.rchild)
print(root.data, end=', ')
from collections import deque
def level_order(root):
'''
层次遍历
'''
queue = deque()
queue.append(root)
while len(queue) > 0:
cur_node = queue.popleft()
print(cur_node.data, end=', ')
if cur_node.lchild:
queue.append(cur_node.lchild)
if cur_node.rchild:
queue.append(cur_node.rchild)
# -
pre_order(a)
print()
in_order(a)
print()
post_order(a)
print()
level_order(a)
# #### 3.7.3 二叉搜索树
#
# 二叉搜索树时一个二叉树且满足性质:
# > 设X是二叉树的一个节点。
# > 如果Y是X的左子树上的一个节点,那么Y.key <= X.key;
# > 如果Y是X的右子树上的一个节点,那么Y.key >= X.key;
#
# <img src='./pictures/33_搜索二叉树.jpeg' style='zoom:50%'/>
# +
class BiTreeNode():
def __init__(self, data):
self.data = data
self.lchild = None
self.rchild = None
self.parent = None
class BST():
'''
二叉搜索树
'''
def __init__(self, iterable):
self.root = None
if iterable:
for val in iterable:
self.insert_no_cur(val)
# 递归方式插入(不太好理解)
def insert(self, node, val):
if not node: # 比如对于上图,要插入0,1的左节点为空,需创建一个node
node = BiTreeNode(val)
elif val < node.data:
node.lchild = insert(node.lchild, val)
node.lchild.parent = node
elif val > node.data:
node.rchild = insert(node.rchild, val)
node.rchild.parent = node
return node
# 不用递归实现插入
def insert_no_cur(self, val):
if not self.root:
# 空二叉搜索树,则初始化根节点
self.root = BiTreeNode(val)
return
node = self.root
while True:
if val < node.data:
if not node.lchild: # 左子节点为空
child_node = BiTreeNode(val)
node.lchild = child_node
child_node.parent = node
return
else:
node = node.lchild
continue
elif val > node.data:
if not node.rchild:
child_node = BiTreeNode(val)
node.rchild = child_node
child_node.parent = node
return
else:
node = node.rchild
continue
else:
return
def __query(self, node, val):
'''
递归查询
'''
if not node:
return None
elif val == node.data:
return node
elif val < node.data:
return self.__query(node.lchild, val)
elif val > node.data:
return self.__query(node.rchild, val)
def query(self, val):
'''
递归查询
'''
return self.__query(self.root, val)
def __remove_leaf(self, node):
'''
情况一:删除叶子节点
'''
if not node.parent: # 根节点
self.root = None
elif node == node.parent.lchild: #
node.parent.lchild = None
elif node == node.parent.rchild:
node.parent.rchild = None
def __remove_node_with_single_child(self, node):
'''
情况二:要删除的节点只有一个孩子节点
'''
if not node.parent: # 要删除的节点是根节点
if node.lchild:
self.root = node.lchild
elif node.rchild:
self.root = node.rchild
self.root.parent = None
elif node == node.parent.lchild:
if node.lchild:
node.parent.lchild = node.lchild
node.lchild.parent = node.parent
else:
node.parent.lchild = node.rchild
node.rchild.parent = node.parent
elif node == node.parent.rchild:
if node.lchild:
node.parent.rchild = node.lchild
node.lchild.parent = node.parent
else:
node.parent.rchild = node.rchild
node.rchild.parent = node.parent
def remove(self, val):
'''
删除节点
'''
if not self.root: # 代表空树
return
node = self.query(val) # 查询要删除的节点
if not node: # 要删除的节点不存在
raise Exception('Error key!')
# 情况1:node是叶子节点
if not node.lchild and not node.rchild:
self.__remove_leaf(node)
# 情况2:node节点只有一个孩子
elif (node.lchild and not node.rchild) or (not node.lchild and node.rchild):
self.__remove_node_with_single_child(node)
# 情况3:node有两个子树
else:
# 找该节点右子树最小节点(一直往左找)
node_tmp = node.rchild
while node_tmp.lchild:
node_tmp = node_tmp.lchild
# 数据替换到要删除的节点
node.data = node_tmp.data
# 将右子树最小节点删除
if node_tmp.rchild:
self.__remove_node_with_single_child(node_tmp)
else:
self.__remove_leaf(node_tmp)
def in_order(self, root):
'''
中序遍历
'''
if root:
self.in_order(root.lchild)
print(root.data, end=' ')
self.in_order(root.rchild)
# +
import random
bst = BST([8,3,10,1,6,4,7,13,14,])
print('root =', bst.root.data)
# 中序遍历
bst.in_order(bst.root)
print()
# 查询
print(bst.query(11), bst.query(5),bst.query(13).data)
print()
# 删除节点
# bst.remove(1)
bst.remove(6)
bst.in_order(bst.root)
# -
# **二叉搜索树节点删除:**
# 1. 若是叶子节点,直接删除;
# 2. 若要删除的节点只有一个孩子,则将它的parent和child直接相连;
# 3. 若要删除的节点是根节点,则将右子树最小节点替换到该位置上。
#
# **二叉搜索树的效率:**
# 1. 平均情况下,二叉搜索树进行搜索的时间复杂度为$O(\log{n})$;
# 2. <font color=blue>最坏情况下,二叉搜索树可能非常偏斜,这样时间复杂度接近线性;</font>
# 3. 接近方案:
# - 随机化插入
# - AVL树
# #### 3.7.4 AVL树
#
# AVL树:是一棵自平衡的二叉搜索树。
# 1. 根的左右子树的高度差的绝对值不能超过1;
# 2. 根的左右子树都是平衡二叉树。
#
# <img src='./pictures/34_AVL树.png' style='zoom:50%'/>
# **AVL树插入:**
# 1. 插入一个节点可能会破坏AVL树的平衡,可以通过<font color=blue>旋转</font>操作来进行修正。
#
# 2. 插入一个节点后,只有从插入节点到根节点的路径上的节点的平衡可能被改变。我们需要找出第一个平衡条件被破坏的节点,称之为K。K的两棵子树高度差为2.
#
# <img src='./pictures/35_AVL旋转.gif' style='zoom:50%'/>
#
# 3. 不平衡的出现可能有4中情况:
#
# <img src='./pictures/36_LL旋转.png' style='zoom:50%'/>
# <img src='./pictures/37_RR旋转.png' style='zoom:50%'/>
# <img src='./pictures/38_LR旋转.png' style='zoom:50%'/>
# <img src='./pictures/39_RL旋转.png' style='zoom:50%'/>
# +
class AVLNode(BiTreeNode):
def __init__(self, data):
BiTreeNode.__init__(self, data)
self.bf = 0 # 平衡因子
class AVLTree(BST):
def __init__(self, li=None):
BST.__init__(self, li)
def rotate_left(self, k1, k2):
'''
k1 k2
n1 k2 k1 n3
n2 n3 n1 n2 new
new
'''
k1.lchild = k2.lchild
if k2.lchild:
k2.lchild.parent = k1
k2.lchild = k1
k1.parent = k2
# 更新bf(balance factor)
k1.bf = 0
k2.bf = 0
return k2
def rotate_right(self, k2, k1):
'''
k2 k1
k1 n1 n2 k2
n2 n3 new n3 n1
new
'''
k2.lchild = k1.rchild
if k1.rchild:
k1.rchild.parent = k2
k2.parent = k1
k1.rchild = k2
# 更新bf
k1.bf = 0
k2.bf = 0
return k1
def rotate_left_right(self, k3, k1):
'''
k3 | k3 | k2
k1 D | k2 D | k1 k3
A k2 | k1 C | A B C D
B C | A B |
'''
k2 = k1.rchild
# 先对k1子树进行左旋转
k2 = self.rotate_left(k1, k2)
k3.lchild = k2
k2.parent = k3
# 再对k3树进行右旋转
k3 = self.rotate_right(k3, k2)
# 更新bf
# case1: k2节点后插入B
if k2.lchild:
k1.bf = 0
k3.bf = 1
elif k2.rchild:
k1.bf = -1
k3.bf = 0
else: # 此时没有A、B、C、D节点,只是插入k2导致k3的平衡被破坏
k1.bf = 0
k3.bf = 0
return k3
def rotate_right_left(self, k1, k3):
'''
k1 | k1 | k2
A k3 | A k2 | k1 k3
k2 D | B k3 | A B C D
B C | C D |
'''
k2 = k3.lchild
# 先对k3子树进行右旋转
k2 = self.rotate_right(k3, k2)
k1.rchild = k2
k2.parent = k1
# 再对k1树进行左旋转
k3 = self.rotate_left(k1, k2)
# 更新bf
if k2.lchild: # case1: k2节点后插入的是B
k1.bf = 0
k3.bf = 1
elif k2.rchild: # case2: k2节点后插入的是C
k1.bf = -1
k3.bf = 0
else: # case3: 此时没有A、B、C、D节点,只是插入k2导致k3的平衡被破坏
k1.bf = 0
k3.bf = 0
return k2
def insert_no_cur(self, val):
#================1.插入val==================
if not self.root:
# 空二叉搜索树,则初始化根节点
self.root = AVLNode(val)
return
node = self.root
while True:
if val < node.data:
if not node.lchild: # 左子节点为空,则创建一个左子节点
child_node = AVLNode(val)
node.lchild = child_node
child_node.parent = node
else: # 若左子节点不为None, 则将左子节点作为根,与val进行比较
node = node.lchild
continue
elif val > node.data:
if not node.rchild:
child_node = AVLNode(val)
node.rchild = child_node
child_node.parent = node
else:
node = node.rchild
continue
else: # 若插入的值与已有节点相同,不做任何处理
return
#================2.更新bf==================
# node 为插入节点的父节点
while child_node.parent: # 直到更新到根节点位置,也就是节点的parent为None为止
if child_node == node.lchild: # 从node的左边插入child_node
node.bf -= 1 # 更新后的node.bf要么等于0,要么等于-1
if node.bf == 0: # 也就是新插入节点不会导致树的失衡
break
else: # 继续向上更新bf
parent_node = node.parent
if node == parent_node.lchild: # 说明新插入是left-left形式,要进行右旋
parent_node.bf -= 1 # 更新后的bf只可能是-2, -1, 0三种情况
if parent_node.bf == 0: # 平衡了,不用再往上更新
return
elif parent_node.bf == -1: # 继续往上更新
child_node = node
node = node.parent
continue
elif parent_node.bf == -2: # 进行右旋
root_node = self.rotate_left(node, child_node)
else: # 说明新插入是right-left,要进行左-右旋转
parent_node.bf += 1
if parent_node.bf == 0:
return
elif parent_node.bf == 1:
child_node = node
node = node.parent
continue
elif parent_node.bf == 2:
root_node = self.rotate_left_right(parent_node, node)
else: # 从node的右边插入child_node
node.bf += 1
if node.bf == 0:
break
else: # 继续向上更新
parent_node = node.parent
if node == parent_node.node.lchild: # 说明新插入是right-left形式
parent_noder.bf -= 1
if parent_node.bf == 0:
return
elif parent_node.bf == -1:
child_node = node
node = parent_node
continue
else: # 进行右-左旋转
root_node = self.rotate_right_left(parent, node)
#================3. 连接旋转后的子树==================
up_node = parent_node.parent
if up_node:
if parent_node == up_node.lchild:
up_node.lchild = root_node
root_node.parent = up_node
else:
up_node.rchild = root_node
root_node.parent = up_node
else:
self.root = root_node
return
def in_order(self, root):
'''
中序遍历
'''
if root:
self.in_order(root.lchild)
print(root.data, end=' ')
self.in_order(root.rchild)
# +
import random
tree = AVLTree(random.shuffle([i for i in range(1,11)]))
tree.in_order(tree.root)
print()
tree.insert_no_cur(11)
tree.in_order(tree.root)
# -
# #### 3.7.5 二叉搜索树扩展应用——B树
# B树(B-Tree):B树是一棵自平衡的多路搜索树。通常用于数据库的索引。
#
# <img src='./pictures/40_B树.png' style='zoom:50%'/>
#
# 因为数据库数据时分块存在硬盘上的,AVL树虽然查询效率也很高,但它是二路平衡树,树的高度为$\log{_{2}(n)}$,而B树是多路平衡树,可以大大降低树的高度,加快搜索效率。
# ## 4. 贪心算法
# 1. 贪心算法(又称贪婪算法)是指,在对问题求解时,总是做出在当前看来是最好的选择。也就是说,不从整体最优上加以考虑,他所做出的是在某种意义上的局部最优解。
# 2. 贪心算法并不保证会得到最优解,但是在某些问题上贪心算法的解就是最优解。要会判断一个问题能否用贪心算法来计算。
# ### 4.1 找零问题
# 假设商店老板需要找零n元钱,钱币的面额有:100元、50元、20元、5元、1元,如何找零使得所需钱币的数量最少?
# +
money = [100, 50, 20, 5, 1]
def change(li, n):
num = [0 for _ in range(len(li))]
for idx, m in enumerate(money):
num[idx] = n // m
n %= m
return num
# -
change(money, 321)
# ### 4.2 背包问题
# 一个小偷在某个商店发现有n个商品,第i个商品价值$\text{v}_{i}$元,重$\text{w}_{i}$千克。他希望拿走的价值尽量高,但他的背包最多只能容纳W千克的东西。他应该拿走哪些商品?
# - **0-1背包:** 对于一个商品,小偷要么把它完整拿走,那么留下。不能只拿走一部分,或把一个商品拿走多次。(商品为金条)
# - **分数背包:** 对于一个商品,小偷可以拿走其中任意一部分。(商品为金砂)
#
# 明显,贪心算法求解分数背包得到的是最优解。而0-1背包问题,则不一定。
# +
'''
商品1:v1=60, w1=10
商品2:v2=100, w2=20
商品3:v3=120, w3=30
背包容量:W=50
'''
def factional_backpack(goods, W):
num = [0 for _ in range(len(goods))]
total_value = 0
for idx, (price, weight) in enumerate(goods):
if W >= weight:
num[idx] = 1
total_value += price
W -= weight
else:
num[idx] = W / weight
total_value += price * W / weight
break
return num, total_value
# +
goods = [(60, 10), (120, 30),(100, 20)]
goods.sort(key=lambda x:x[0]/x[1], reverse=True)
num, total_value = factional_backpack(goods, 50)
print(num)
print(total_value)
num, total_value = factional_backpack(goods, 100)
print(num)
print(total_value)
# -
# ### 4.3 数字拼接问题
# 有n个非负整数,将其按照字符串拼接的方式拼接为一个整数。如何拼接可以使得到的整数最大?例:
# > 32, 94, 128, 1286, 6, 71可以拼接的最大整数为:94716321286128
def num_join(li):
li = list(map(str, li))
# 同冒泡排序
for i in range(len(li)-1):
for j in range(i+1, len(li)):
if li[i] + li[j] < li[j] + li[i]:
li[i], li[j] = li[j], li[i]
return ''.join(li)
li = [32, 94, 128, 1286, 6, 71]
num = num_join(li)
print(num)
# ### 4.4 活动选择问题
# 假设有n个活动,这些活动要占用同一块场地,而场地在某时刻只能供一个活动使用。
#
# 每个活动都有一个开始时间si和结束时间fi(题目中时间以整数表示),表示活动在\[si, fi\}区间占用场地。
#
# 问:安排哪些活动能够使该场地举办的活动个数最多?
#
# <img src='./pictures/41_活动选择问题.jpg' style='zoom:50%'/>
#
# 贪心结论:<font color=blue>最先结束的活动一定是最优解的一部分</font>
#
# 证明:假设a是所有活动中最先结束的活动,b是最优解中最先结束的多动。
# - 如果a = b, 结论成立;
# - 如果a ≠ b, 则b的结束时间一定晚于a,则此时用a替换掉b,a一定不与最优解中的其它活动时间重叠,因此替换掉b后也是最优解。
# +
activations = [(1,4),(3,5),(0,6),(5,7),(3,9),(5,9),(6,10),(8,11),(8,12),(2,14),(12,16)]
# 首先按活动结束时间进行排序
activations.sort(key=lambda x:x[1])
def act_selection(activations):
act = [activations[0]]
for i in range(1, len(activations)):
# 若时间不重叠,则加入
if activations[i][0] >= act[-1][1]:
act.append(activations[i])
return act
# -
print(act_selection(activations))
# ## 5. 动态规划
# +
'''
递归实现斐波那契数列,会出现子问题重复计算问题
'''
def fibonacci(n):
if n == 1 or n == 2:
return 1
else:
return fibonacci(n-2) + fibonacci(n-1)
def fibonacci_no_cur(n):
val = [0,1,1]
if n > 2:
for i in range(n-2):
num = val[-1] + val[-2]
val.append(num)
return val[-1]
else:
return val[n]
# +
import time
start = time.time()
print(fibonacci(35))
print('递归实现:', time.time()-start)
start = time.time()
print(fibonacci_no_cur(35))
print('非递归实现:', time.time()-start)
# -
# **动态规划(DP)** = <font color=red>递推式</font> + 重复子问题
# ### 5.1 钢条切割问题
#
# <img src='./pictures/42_钢条切割问题.jpg' style='zoom:50%'/>
#
# <img src='./pictures/43_钢条切割问题2.jpg' style='zoom:50%'/>
# 钢条切割问题的递推式1:
#
# <img src='./pictures/44_钢条切割问题_递推式1.jpg' style='zoom:50%'/>
# **<font color=red>最优子结构</font>**
#
# 1. 可以将求解规模为n的原问题,划分为规模更小的子问题:完成一次切割后,可以将产生的两段钢条看成两个独立的钢条切割问题。
#
# 2. 组合两个子问题的最优解,并在所有可能的两段切割方案中选取组合收益最大的,构成原问题的最优解;
#
# 3. 钢条切割满足**最优子结构:** 问题的最优解由相关子问题的最优解组合而成,这些子问题可以独立求解。
#
# 钢条切割问题的递推式2:
#
# <img src='./pictures/45_钢条切割问题_递推式2.jpg' style='zoom:50%'/>
# +
'''
递归实现递推式1
'''
def cut_rod_cur(p, n):
if n == 0: # 长度为0
return 0
else:
pn = p[n-1]
for i in range(1, n):
pn = max(pn, cut_rod_cur(p, i) + cut_rod_cur(p, n-i))
return pn
'''
递归实现递推式2
'''
def cut_rod_cur2(p, n):
if n == 0: # 长度为0
return 0
else:
max_val = 0
for i in range(0, n):
max_val = max(max_val, p[i] + cut_rod_cur2(p, n-i-1))
return max_val
# +
p = [1,5,8,9,10,17,17,20,24,30,32,36,40,42,44,48,50]
import time
start = time.time()
print(cut_rod_cur(p,15))
print('递推公式1:', time.time() - start)
start = time.time()
print(cut_rod_cur2(p,15))
print('递推公式2:', time.time() - start)
# -
# **自顶向下递归实现:**
# 上面两个实现都是自顶向下的实现方式,效率很差,时间复杂度是$O(2^{n})$
#
# **动态规划的思想:**
# - 每个子问题只求解一遍,保存求解结果;
# - 之后需要此问题,只需查找保存的结果。
'''
动态规划思想实现递推式2,也就是自底向上
'''
def cut_rod_dp(p, n):
ri = [0] # ri的临时存放表
for i in range(1, n+1): # 求ri,i从1到n
tmp = 0
for j in range(0, i): # r从1到n-i
tmp = max(tmp, p[j] + ri[i-j-1])
ri.append(tmp)
return ri[n]
# 时间复杂度:$O(n^{2})$
# +
p = [1,5,8,9,10,17,17,20,24,30,32,36,40,42,44,48,50]
import time
start = time.time()
print(cut_rod_cur(p,15))
print('递推公式1:', time.time() - start)
start = time.time()
print(cut_rod_cur2(p,15))
print('递推公式2:', time.time() - start)
start = time.time()
print(cut_rod_dp(p, 15))
print('DP实现:', time.time()-start)
# -
# **钢条切割问题——重构解:**
#
# <img src='./pictures/46_钢条切割问题_重构解.jpg' style='zoom:40%'/>
# +
'''
重构解
'''
def cut_rod_extend(p, n):
ri = [0] # ri的临时存放表
si = [0] # ri对应最优值对应切割方案中左边一段的长度
for i in range(1, n+1): # 求ri,i从1到n
tmp_r = 0
tmp_s = 0
for j in range(0, i): # r从1到n-i
if p[j] + ri[i-j-1] > tmp_r:
tmp_r = p[j] + ri[i-j-1]
tmp_s = j + 1
ri.append(tmp_r)
si.append(tmp_s)
return ri[n], si
def cut_rod_solution(p, n):
val, s = cut_rod_extend(p,n)
print(s)
ones = []
while n > 0:
ones.append(s[n])
n -= ones[-1]
return ones
# +
p = [1,5,8,9,10,17,17,20,24,30]
cut_rod_solution(p, 9)
# -
# ### 5.2 最长公共子序列LCS(Longest Common Subsequence)
#
# <img src='./pictures/47_最长公共子序列(LCS).jpg' style='zoom:50%'/>
# **定理:**令$X=<x_{1}, x_{2}, \cdots , x_{m}>$ 和 $Y=<y_{1}, y_{2}, \cdots , y_{n}$为两个序列,$Z=<z_{1}, z_{2}, \cdots , z_{k}>$ 为$X$ 和 $Y$ 的任意LCS,则:
# 1. 如果$x_m = y_{n}$,则$z_{k} = x_{m} = y_{n}$ 且$Z_{k-1}$是$X_{m-1}$和$Y_{n-1}$的一个LCS;
# 1. 如果$x_m \neq y_{n}$,那么$z_{k} \neq x_{m}$ 意味着$Z$ 是$X_{m-1}$和$Y$的一个LCS;
# 2. 如果$x_m \neq y_{n}$,那么$z_{k} \neq y_{n}$ 意味着$Z$ 是$X$和$Y_{n-1}$的一个LCS;
#
# 最优解的递推式:
# $$
# C[i, j]=\left\{\begin{array}{ll}
# 0, & \text { 当 } i=0 \text { 或 } j=0 \\
# C[i-1, j-1]+1, & \text { 当 } i, j>0 \text { 且 } x_{i}=y_{j} \\
# M A X(C[i, j-1], C[i-1, j]) & \text { 当 } i, j>0 \text { 且 } x_{i} \neq y_{j}
# \end{array}\right.
# $$
# 上面的定理和递推公式可用如下表格表示:
# 第一行和第一列表示空串
# <img src='./pictures/48_LCS_Table.jpg' style='zoom:50%'/>
# +
def LCS_lenght(x,y):
'''
求两个字符串的最长公共子序列的长度
'''
x_len = len(x)
y_len = len(y)
lcs_table = [[0 for _ in range(y_len+1)] for _ in range(x_len+1)]
for row in range(1, x_len+1):
for col in range(1, y_len+1):
if x[row-1] == y[col-1]: #两个字母相同
lcs_table[row][col] = lcs_table[row-1][col-1] + 1 # 等于左上方值 + 1
else: # 两个字母不想等,则取max(左边,右边)
lcs_table[row][col] = max(lcs_table[row-1][col], lcs_table[row][col-1])
return lcs_table[x_len][y_len]
'''
在计算lcs_table的同时,记录子串长度值的路径
规定:1-来自左上方,2-来自上方,3-来自左方
'''
def LCS(x, y):
x_len = len(x)
y_len = len(y)
lcs_table = [[0 for _ in range(y_len+1)] for _ in range(x_len+1)]
trace = [[0 for _ in range(y_len+1)] for _ in range(x_len+1)]
for row in range(1, x_len+1):
for col in range(1, y_len+1):
if x[row-1] == y[col-1]: #两个字母相同
lcs_table[row][col] = lcs_table[row-1][col-1] + 1 # 等于左上方值 + 1
trace[row][col] = 1
elif lcs_table[row-1][col] > lcs_table[row][col-1]: # 说明来自上方
lcs_table[row][col] = lcs_table[row-1][col]
trace[row][col] = 2
else: # 说明来自左方
lcs_table[row][col] = lcs_table[row][col-1]
trace[row][col] = 3
return lcs_table[x_len][y_len], trace
'''
根据路径求出最长子串
'''
def LCS_substr(x,y):
m = len(x)
n = len(y)
length, trace = LCS(x, y)
substr = []
while m > 0 and n > 0:
if trace[m][n] == 1: # 来自左上方,说明该位置字符是相同的
substr.append(x[m-1])
m -= 1
n -= 1
continue
elif trace[m][n] == 2: # 来自上方,该位置字符是不同的
m -= 1
continue
else: # 来自左方,该位置字符是不同的
n -= 1
continue
return ''.join(reversed(substr))
# -
LCS_length('ABCBDAB', 'BDCABA')
length, trace = LCS('ABCBDAB', 'BDCABA')
for _ in trace:
print(_)
LCS_substr('ABCBDAB', 'BDCABA')
# ## 6. 欧几里得算法
# 最大公约数(Greatest Common Divisor,GCD)
#
# 欧几里得算法:gcd(a, b) = gcd(b, a%b), 例:
#
# > gcd(60, 21) = gcd(21, 18) = gcd(18, 3) = gcd(3, 0) = 3
# +
'''
递归实现gcd
'''
def gcd_rec(a, b):
if a % b == 0:
return b
else:
return gcd_rec(b, a % b)
'''
非递归实现gcd
'''
def gcd_no_rec(a, b):
while b != 0:
tmp = b
b = a % b
a = tmp
else:
return a
'''
分数的运算
'''
class Fraction():
def __init__(self, molecular, denominator):
self.molecular = molecular # 分子
self.denominator = denominator
gcd = self.gcd(molecular, denominator)
self.molecular = molecular / gcd
self.denominator = denominator / gcd
def gcd(self, molecular, denominator):
while denominator != 0:
tmp = denominator
denominator = molecular % denominator
molecular = tmp
else:
return molecular
'''
分数加法
'''
def __add__(self, other):
m1 = self.molecular
d1 = self.denominator
m2 = other.molecular
d2 = other.denominator
# 先统一分母,且最小公倍数
# 6,8 --> gcd:2 --> 6/2=3,8/2=4 -->最小公倍数:2*3*4
gcd = self.gcd(d1, d2)
denominator = gcd * (d1 / gcd) * (d2 / gcd) # 分母的最小公倍数
molecular = m1 * (d1 / gcd) + m2 * (d2 / gcd)
return Fraction(molecular, denominator)
def __str__(self):
return '%d/%d'%(self.molecular, self.denominator)
# -
f1 = Fraction(1,2)
f2 = Fraction(1,3)
total = f1 + f2
print(total)
# ## 7. RSA算法
# 传统密码:加密算法是秘密的,比如凯撒码;
#
# 现代密码系统:加密算法是公开的,但秘钥是秘密的:
#
# - 对称加密(DES、3DES、Blowfish、IDEA、RC4、RC5、RC6 和 AES )
# - 非对称加密(RSA、ECC(移动设备用)、Diffie-Hellman、El Gamal、DSA(数字签名用))
#
# 公钥:用于加密
#
# 私钥:用于解密
# RSA加解密过程:
# 1. 随机选取两个质数 p 和 q ;
# 2. 计算 n = pq ;
# 3. 选取一个与 φ(n) 互质的小奇数e, 其中φ(n)=(p-1)(q-1)
# 4. 对φ(n),计算e的乘法逆元d,即满足 (e\*d) mod φ(n) = 1
# 5. 公钥(e,n) 私钥(d,n)
# 6. 加密过程:c=(m^e) mod n
# 7. 解密过程:m=(c^d) mod n
# ## 8. 字符串算法
# 1. 哈希法(最直观的方法)
# 2. KMP算法(最基础的方法)
# 3. 扩展KMP算法
# 4. Manacher算法(解决回文串问题(aba))
# 5. AC自动机(Trie+KMP)
# - 哈希:最简单直观,易实现,便于拓展;
# - KMP:本质是利用模板串自身的信息,去减少匹配时的冗余比较,达到优秀的时间复杂度;$O(n)$
# - 扩展KMP、Manacher算法:利用对称性,降低时间复杂度,$O(n)$
# - AC自动机:Trie树与KMP的结合,可以解决一个串和多个串的匹配问题。
# ### 8.1 哈希法
# **问题:**
# 有一个字符串集合,假设为 \['abc', 'bcd', 'adf', 'bce', 'edaf', 'adfc','ad'\], 问字符串'adfc'在该集合中吗?
# 该问题可以用字符串哈希实现,效率比暴力计算高。
#
# 字符串哈希就是将一个字符串映射为一个整数。
# 哈希公式:
# $$
# \text{hash}[i] = \text{hash}[i-1]*p + \text{id}(s[i])
# $$
# 其中,p为质数,id(x)为x-'a'+1或者x的ASCII码。
# +
class Node():
def __init__(self, value=None):
self.next = None
self.pre = None
self.value = value
'''计算hash值'''
def get_hash(string:str, p:int=31):
h = 0
for idx, char in enumerate(string):
if idx == 0:
h += ord(string[idx])
else:
h = h * p + ord(string[idx])
return h
class StrHash():
def __init__(self, mod=31, maxn=1e6+7):
self.mod = mod
self.li = [None for _ in range(int(maxn))]
def insert(self, string):
shash = get_hash(string, self.mod)
if self.li[shash] is None:
node = Node(string)
self.li[shash] = node
else:
# 若有hash冲突,采用头插法插入
root_node = self.li[shash]
if root_node.next is None:
node = Node(string)
root_node.next = node
node.pre = root_node
else:
node = Node(string)
root_node.next.pre = node
node.next = root_node.next
root_node.next = node
node.pre = root_node
def __contains__(self, item):
shash = get_hash(item, self.mod)
root_node = self.li[shash]
if root_node is None:
return False
else:
while root_node is not None:
if item == root_node.value:
return True
else:
root_node = root_node.next
else:
return False
# -
get_hash('ab')
# +
shash = StrHash()
shash.insert('ab')
shash.insert('a')
shash.insert('abc')
shash.insert('def')
print('ab' in shash)
print('abd' in shash)
# -
# ### 8.2 KMP
# 待匹配字符串T: 'ABACDEFBEDFACCD'
# 模式字符串P: 'ACCD'
#
# 字符串匹配问题就是看T中是否有P。
#
# **概念:**
# 1. 子串:
# > 'ABCDABC'的前缀子串有:\['A', 'AB', 'ABC', 'ABCD', 'ABCDA', 'ABCDAB', 'ABCDABC'\], 其中,除它本身外,其它称为真前缀子串;
# > 'ABCDABC'的后缀子串有:\['C', 'BC', 'ABC', 'DABC', 'CDABC', 'BCDABC', 'ABCDABC'\], 其中,除它本身外,其余称为真后缀子串。
#
# KMP算法是一种改进的字符串匹配算法,由D.E.Knuth,J.H.Morris和V.R.Pratt提出的,因此人们称它为克努特—莫里斯—普拉特操作(简称KMP算法)。KMP算法的核心是利用匹配失败后的信息,尽量减少模式串与主串的匹配次数以达到快速匹配的目的。
# 暴力解法
# +
'''
T: ABCACEBDACEABCB
P: ACEAB
'''
def str_match(text, pattern):
t_idx = 0 # 待匹配字符串指针
while t_idx <= len(text)-len(pattern):
for i in range(t_idx, t_idx+len(pattern)):
if text[i] == pattern[i-t_idx]:
if i-t_idx == len(pattern)-1:
return True
else:
continue
else:
t_idx += 1
break
return False
# +
t = 'ABCACEBDACEABCBEABCBDFSRKJNMHBVFGRTEWXCVBNMSDFGH'
p1 = 'TEWXCVBNMSD'
p2 = 'TEWXCVBNMSG'
print(str_match(t, p1))
print(str_match(t, p2))
# -
# KMP算法
# <img src='./pictures/51_KMP.png' style='zoom:20%'/>
#
# <img src='./pictures/52_KMP_next.png' style='zoom:50%'/>
'''
0 1 2 3 4 5 6 7 8 9 10 11
A B A B A A A B A B A A
0 1 1 2 3 4 2 2 3 4 5 6
'''
def next_idx(pattern):
# 前两个元素始终为0,1
idx = [1 for _ in range(len(pattern))]
idx[0] = 0
# 从第3个元素开始计算
for i in range(2, len(pattern)):
for j in range(i - 1, 0, -1):
if pattern[:j] == pattern[i-j:i]:
idx[i] = j + 1
break
return idx
print(next_idx('ABABAAABABAA'))
print(next_idx('ABCDEFG'))
def str_match_kmp(text, pattern):
p_next = next_idx(pattern)
t_idx = 0 # text指针当前位置
p_idx = 0 # pattern指针当前位置
while t_idx < len(text) and p_idx < len(pattern):
if text[t_idx] == pattern[p_idx]:
if p_idx == len(pattern) - 1:
return True
else:
t_idx += 1
p_idx += 1
continue
else:
next_id = p_next[p_idx]
if next_id == 0:
t_idx += 1
continue
else:
p_idx = next_id - 1
continue
else:
return False
t = 'ABCDESDSFGCDGABCDEF'
p = 'ABCDEF'
print(str_match(t, p))
# +
t = 'ABCACEBDACEABCBEABCBDFSRKJNMHBVFGRTEWXCVBNMSDFGH'
p1 = 'TEWXCVBNMSD'
p2 = 'TEWXCVBNMSG'
print(str_match(t, p1))
print(str_match(t, p2))
# +
text = ''
with open('kmp_text.txt') as f:
lines = f.readlines()
for line in lines:
text = text + line
print(text)
pattern = '所以复杂度'
pattern2 = '所以复杂度啊'
# +
import time
begin = time.time()
print(str_match(text, pattern))
print(str_match(text, pattern2))
print('暴力法总共耗时:',time.time()-begin)
begin = time.time()
print(str_match_kmp(text, pattern))
print(str_match_kmp(text, pattern2))
print('KMP总共耗时:',time.time()-begin)
# -
# **KMP的核心:** <font color=red>主要是通过next(),避免了待匹配字符串指针的回溯,从而节省匹配时间。</font>
# ### 8.3:字典树(trie树)
# 在字典分词算法中,我们需要判断当前字符串是否在字典中。如果用有序集合(TreeMap)的话,复杂度是$O(\log{n})$;如果用散列表(HashMap)的话,时间复杂度虽然下降了,但内存复杂度却上去了。
#
#
# 字典树,又叫trie树、前缀树,有如下性质:
# 1. 字典树每条边都对应一个字;
# 2. 从根节点往下的路径构成一个个字符串;
# 3. 字典树并不直接在节点上存储字符串,而是将词语视作根节点到某节点之间的一条路径,并在终点节点(蓝色)上做标记“该节点对应词语的结尾”;
# 4. 字符串就是一条路径,要查询一个单词,只需顺着这条路径从根节点往下走。
#
# <img src='./pictures/49_字典树示意图.jpg' style='zoom:45%'/>
#
# 当词典大小为$n$时,虽然最坏情况下字典树的复杂度依然是$O(\log{n})$(假设子节点用对数复杂度的数据结构存储,所有词语都是单字),但他的实际速度要比二分查找快。这是因为随着路径的深入,前缀匹配是递进的过程,算法不必比较字符串的前缀。
#
# <img src='./pictures/50_字典树实现.jpg' style='zoom:50%'/>
# +
class Node:
def __init__(self, value):
self._children = {}
self._value = value
def _add_child(self, char, value, overwrite=False):
child = self._children.get(char)
if child is None: # 对应子节点为None
child = Node(value)
self._children[char] = child
elif overwrite:
child._value = value
return child
class Trie(Node):
def __init__(self):
super().__init__(None) # 初始化一个根节点
# 覆写__contains__魔法方法
def __contains__(self, key):
return self[key] is not None # 等价于 self.__getitem__(key)
# 可以像对待dict一样操作字典树
def __getitem__(self, key):
root = self
for char in key:
root = root._children.get(char)
if root is None:
return None
return root._value
def __setitem__(self, key, value):
root = self
for idx, char in enumerate(key):
if idx < len(key) - 1:
root = root._add_child(char, None, False)
else: # 蓝色节点
root = root._add_child(char, value, True)
# +
trie = Trie()
trie['入门'] = 'introduction'
trie['自然人'] = 'human'
trie['自语'] = 'speak to oneself'
trie['自然'] = 'nature'
print('自然' in trie)
print('自然猪' in trie)
# 删
trie['自然'] = None
print('自然' in trie)
# 改
trie['自然人'] = 'nature human'
print(trie['自然人'])
# 查
print(trie['入门'])
# -
# ### 8.4 AC自动机
# Aho-Corasick automaton,该算法在1975年产生于贝尔实验室,是著名的多模匹配算法。
#
# **多模式匹配(multi-pattern matching):** 给定多个词语(也称模式串,pattern),从母文本中匹配它们的问题称多模式匹配,比如:
# 母文本: ushers
# 模式串集合: {he, she, his, hers}
#
# 前置知识:trie树、KMP、BFS
#
# KMP: 单对单文本匹配,比如母文本:ABACDEFADBCA, pattern:ADBCA
# trie树: 多对单文本匹配,比如母文本集合:{自然、自然人、自然语言、入门} pattern:自然语言
# AC自动机: 单对多文本匹配。
# AC自动机的实现原理:
# 1. 第模式串构建trie树;
# 2. 构建fail指针:(fail本质上当前单词的最长后缀,比如:this的最长后缀是his,所以this的s指向his的s.)
# - 它是BFS来构建;
# - 它的第一层全部指向root
# - fail指向:它的父亲的fail节点的同字符儿子,若没有找到,就继续跳fail,直到跳到root还没有,指向root
#
# 3. 匹配(本质上是在fail链上找单词)
# - 从根出发,从第一个文本首字母出发;
# - 进行trie匹配
# - 如果有,进入它
# - 如果没有跳fail,直到root
# - 如果跳到root,下一个文本串
# - 找到单词,不断跳fail,并进行回溯
# <img src='./pictures/53_AC自动机.png' style='zoom:45%'/>
# **匹配过程:**
# 1.
# +
class ACNode():
def __init__(self, character=None):
self.children = {}
self.pre = None
self.character = character
self.pattern = []
self.fail = None
'''根据character获取子节点'''
def child_node(self, character):
if character in self.children:
return self.children[character]
else:
return None
def extend_pattern(self, pattern):
self.pattern.extend(pattern)
def append_pattern(self, pattern):
self.pattern.append(pattern)
from collections import deque
class ACAutomaton():
def __init__(self, pattern_list=[]):
self.root = ACNode()
self.root.fail = self.root # root的fail指向它本身
self.fail_finished = False
for pattern in pattern_list:
self.add_pattern(pattern)
'''添加模式串'''
def add_pattern(self, pattern):
cur_node = self.root
for character in pattern: # 遍历模式串
child_node = cur_node.child_node(character)
if child_node:
cur_node = child_node
else:
child_node = ACNode(character)
cur_node.children[character] = child_node
child_node.pre = cur_node
cur_node = child_node
cur_node.append_pattern(pattern)
'''遍历所有pattern'''
def get_all(self, cur_node):
if len(cur_node.pattern) > 0:
print(cur_node.pattern)
for key in cur_node.children:
self.get_all(cur_node.children[key])
'''构建fail链'''
def __construct_fail(self):
if self.fail_finished:
return
# BFS遍历每个节点,构建fail链
queue = deque()
# 第一层的节点的fail指向root
for key in self.root.children:
self.root.children[key].fail = self.root
queue.append(self.root.children[key])
while len(queue) > 0:
cur_node = queue.popleft()
cur_char = cur_node.character
# 将当前节点的子节点添加到queue中
children = cur_node.children
for key, node in children.items():
queue.append(node)
# 设置当前节点的fail节点
if cur_node.fail is None:
parent_node = cur_node.pre
fail_node = parent_node.fail
while cur_char not in fail_node.children:
if fail_node == self.root:
cur_node.fail = fail_node
break
else:
fail_node = fail_node.fail
continue
else:
cur_node.fail = fail_node.children[cur_char]
cur_node.extend_pattern(fail_node.children[cur_char].pattern)
self.fail_finished = True
'''模式匹配'''
def match(self, text):
# 先检查并构建fail链
self.__construct_fail()
# 根据fail链进行匹配
root = self.root
out_pattern = []
for character in text:
if character in root.children: # 如果匹配成功,则沿着trie树继续向下匹配
root = root.children[character]
if len(root.pattern) > 0:
out_pattern.extend(root.pattern)
else: # 如果匹配不成功,则根据fail链进行匹配
fail_node = root.fail
while character not in fail_node.children:
fail_node = fail_node.fail
if fail_node == self.root:
break
else:
root = fail_node.children[character]
if len(root.pattern) > 0:
out_pattern.extend(root.pattern)
return out_pattern
# -
patterns = ['she', 'he', 'her', 'his', 'this', 'is']
ac = ACAutomaton(patterns)
ac.get_all(ac.root)
print(ac.match('sherthis'))
# ### 8.5 Manacher算法(回文串问题)
# a
# +
import time
def display_run_time(func):
def wrapper(*args):
t1 = time.time()
result = func(*args)
t2 = time.time()
print('Total Time:%.6fs'%(t2-t1))
return result
return wrapper
# -
# #### 暴力解法
# 遍历每一个子串,校验它们是否是回文串,时间复杂度为$O(n^{3})$
@display_run_time
def brute_fore(string):
size = len(string)
max_len = 0
start = 0
for i in range(size):
for j in range(i+1,size+1): # 两层循环,遍历所有的子串
sub_string = string[i:j]
sub_len = len(sub_string)
for k in range(sub_len):
if sub_string[k] != sub_string[sub_len-k-1]:
break
elif sub_len-k-1 < k and sub_len > max_len:
max_len = sub_len
start = i
break
return start, max_len
start, max_len = brute_fore('dababac')
print(start, max_len)
# #### 中心扩展法
# 以被考虑词为中心,向两边扩展,分两种情况讨论:
# 1. 长度为奇数,比如:aba
# 2. 长度为偶数,比如:abba
#
# 时间复杂度$O(n^{2})$
@display_run_time
def center_extend(string):
'''
abadc
abba
'''
match_string = ''
size = len(string)
# 按奇数考虑
for i in range(size):
j = i-1
k = i+1
while j >= 0 and k < size and string[j] == string[k]:
j -= 1
k += 1
else:
if k-j-1 > len(match_string):
match_string = string[j+1:k]
# 按偶数考虑
for i in range(size):
j = i
k = i+1
while j >= 0 and k < size and string[j] == string[k]:
j -= 1
k += 1
else:
if k-j-1 > len(match_string):
match_string = string[j+1:k]
return match_string
print(center_extend('a'))
print(center_extend('aba'))
print(center_extend('abba'))
print(center_extend('eabba'))
print(center_extend('acbdadbe'))
@display_run_time
def manacher(string):
max_len = 0
# 1. 字符串预处理
pre_str = '#'
for s in string:
pre_str = pre_str + s + '#'
length = len(pre_str)
# 2. 初始化半径列表和最右边解
rad = [0 for _ in range(len(pre_str))]
max_right = 0 # max_right对应的字符包含在回文串中
mid_point = 0 # max_right对应的回文中点
# 3. 遍历每个字符,找最长回文串
for i in range(length):
# 根据已有回文半径,得到当前i位置扩充前的回文半径
if i < max_right:
# 如果 i 在max_right的左边,则rad[i]要么等于i 关于 mid_pos 对称点 i' 的rad[i'] , 要么等于max_right - i
rad[i] = min(rad[2 * mid_point - i], max_right - i)
else:
rad[i] = 1 # 点i在max_right的右边
# 拓展rad[i]
while i - rad[i] >=0 and i + rad[i] < length and pre_str[i - rad[i]] == pre_str[i + rad[i]]:
rad[i] += 1
# 更新max_right和mid_pos
if rad[i] + i -1 > max_right:
max_right = rad[i] + i -1
mid_point = i
# 更新最长回文串的长度
max_len = max(rad[i], max_len)
return max_len - 1
print(manacher('abbcbb'))
print(manacher('ddcdabbcbbfafgag'))
print(manacher('aaaa'))
print(manacher('a'))
chr(98)
# +
import random
from copy import deepcopy
mid_str = [chr(random.randint(97, 123)) for i in range(100)]
pre_str = [chr(random.randint(97, 123)) for i in range(100)]
post_str = [chr(random.randint(97, 123)) for i in range(200)]
str1 = ''.join(mid_str)
str2 = str1[::-1]
mid_str = str1 + 'a' + str2
string = ''.join(pre_str) + mid_str + ''.join(post_str)
print(mid_str)
print('-'*10)
print(string)
# -
brute_fore(string)
center_extend(string)
manacher(string)
# ## 9. Dijkstra(迪杰斯特拉算法)
# 对于如下一个五向带权连通图,求一个顶点到剩余其它顶点的最短路径。
#
# <img src='./pictures/54_ Dijkstra.jpg' style='zoom:50%'/>
# Dijkstra算法的核心是维护三个数组:
# - **visited数组**:可以将顶点标记为已被访问的和未被访问的;
# - **path数组**:维护起点到任意顶点的路径;
# - **distance数组**:维护任意时刻各顶点到起点的距离。
#
# 实现步骤:
# 1. 将图转化为一个图邻接矩阵;
# 2. 初始化visited、path、distance三数组;
# 3. 对剩余n-1个顶点进行探寻:
# - 从distance中找出当前未被探寻(visit=false)的,距起点最近的顶点;
# - 更新visited、distance、path数组
# - 以当前顶点为中转点,更新剩余未被访问顶点到源点的距离,更新原则:
# - 未被访问的顶点;
# - 中转点 + 中转点到目标点的距离 < 源点到目标点的距离
#
# <img src='./pictures/55_Dijkstra_步骤分解.png' style='zoom:55%'/>
# **注意事项:**
# 1. 不能出现负权值边;
# 2. 时间复杂度为$O(n^{2})$
# +
from copy import deepcopy
inf = float('inf')
def dijkstra(weight, start):
count = len(weight)
# 定义visited、distance、path三个数组
visited = [False for _ in range(count)]
path = [[start] for _ in range(count)]
distance = [inf for _ in range(count)]
# 根据起点信息初始化visited、distance
visited[start] = True
for i in range(count):
distance[i] = weight[start][i]
# 循环更新三个数组
for _ in range(1, count): # 循环n-1次
# 寻找最近的未标记节点
mdis = inf
midx = 0
for idx, dis in enumerate(distance):
if dis != 0 and dis < mdis and visited[idx] == False:
mdis = dis
midx = idx
# 更新visited和path和当前顶点的distance
visited[midx] = True
distance[midx] = mdis
path[midx].append(midx)
# 循环更新起始顶点到剩余顶点的distance
for idx, val in enumerate(visited):
if val == True:
continue
elif mdis + weight[midx][idx] < distance[idx] and weight[midx][idx] != inf:
distance[idx] = mdis + weight[midx][idx]
path[idx] = deepcopy(path[midx])
return distance, path
# -
weight = [
[0, 12, inf, inf, inf, 16, 14],
[12, 0, 10, inf, inf, 7, inf],
[inf, 10, 0, 3, 5, 6, inf],
[inf, inf, 3, 0, 4, inf, inf],
[inf, inf, 5, 4, 0, 2, 8],
[16, 7, 6, inf, 2, 0, 9],
[14, inf, inf, inf, 8, 9, 0]
]
distance, path = dijkstra(weight, start=2)
print(distance)
print(path)
# ## 10. Floyd算法
# **解决的问题:**
#
# 求一个带权有向图(Weighted Directed Graph)的<font color=red>任意两点</font>的最短距离的计算,运用了动态规划的思想,<font color=red>算法的时间复杂度为$O(V^{3})$,空间复杂度为$O(V^{2})$。</font>
#
# <img src='./pictures/55_Floyd求解问题.png' style='zoom:50%'/>
# **算法思想:**
# 从任意节点$i$到任意节点$j$的最短路径不外乎有两种可能:
# 1. 直接从$i$到$j$;
# 2. 从$i$经过若干中间节点$k$到达$j$。
#
# 所以我们假设$Dis(i,j)$为节点$i$到节点$j$的最短路径的距离,对于每一个节点$k$, 我们检查$Dis(i,k) + Dis(k,j) < Dis(i,j)$是否成立,如果成立,证明从$i \rightarrow k \rightarrow j$比$i \rightarrow j$路径更短,我们便设置$Dis(i,j) = Dis(i,k) + Dis(k,j)$,这样一来,当我们遍历往所有节点$k$,$Dis(i,j)$中记录的便是$i \rightarrow j$最短路径的距离。
#
#
# **算法关键:**
# ```
# for(k=0;k<n;k++)//中转站0~k
# for(i=0;i<n;i++) //i为起点
# for(j=0;j<n;j++) //j为终点
# if(d[i][j]>d[i][k]+d[k][j])//松弛操作
# d[i][j]=d[i][k]+d[k][j];
# ```
# +
inf = 99999999 # 表示无穷远距离
def floyd(weight):
vex_num = len(weight)
# 定义路径矩阵、距离矩阵
path = [[-1 for _ in range(vex_num)] for _ in range(vex_num)]
# 将顶点i到它自身的路径标记为它本身
for i in range(vex_num):
path[i][i] = i
# 以任意顶点为中转点,更新所有两点之间的距离。
for k in range(vex_num): # 中间点
for i in range(vex_num): # 起点
for j in range(vex_num): # 终点
if weight[i][k] + weight[k][j] < weight[i][j]:
weight[i][j] = weight[i][k] + weight[k][j]
path[i][j] = [i,k,j]
return weight, path
# +
weight = [
[0, 6, 1, inf],
[inf, 0, inf, inf],
[inf, 4, 0, 1],
[inf, 1, inf, 0]
]
distance, path = floyd(weight)
for dis in distance:
print(dis)
print()
for p in path:
print(p)
# -
# ### 10.1 Dijkstra和Floyd算法比较
# | <img width=200/>Dijkstra | <img width=200/>Floyd |
# | :--- | :--- |
# | 不能处理负权图 | 能处理负权图 |
# | 处理单源最短路径 | 处理多源最短路径 |
# | 时间复杂度$O(n^{2})$ | 时间复杂度$O(n^{3})$ |
#
#
# 其实,也可以对每一个顶点执行一遍Dijkstra算法得到任意两点之间的距离,时间复杂度也是$O(n^{3})$;
# 对于稀疏的图,n次Dijkstra更出色;而对于稠密的图,可以使用Floyd算法。
| Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import tensorflow as tf
# +
output = None
hidden_layer_weights = [
[0.1, 0.2, 0.4],
[0.4, 0.6, 0.6],
[0.5, 0.9, 0.1],
[0.8, 0.2, 0.8]]
out_weights = [
[0.1, 0.6],
[0.2, 0.1],
[0.7, 0.9]]
# Weights and biases
weights = [
tf.Variable(hidden_layer_weights),
tf.Variable(out_weights)]
biases = [
tf.Variable(tf.zeros(3)),
tf.Variable(tf.zeros(2))]
# Input
features = tf.Variable([[1.0, 2.0, 3.0, 4.0],
[-1.0, -2.0, -3.0, -4.0],
[11.0, 12.0, 13.0, 14.0]])
# nn
layer1 = tf.add(tf.matmul(features, weights[0]), biases[0])
layer2 = tf.nn.relu(layer1)
output = tf.nn.softmax(tf.add(tf.matmul(layer2, weights[1]), biases[1]))
# session
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
print(sess.run(output))
# -
| labs/miniflow/toy_dnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Docker
#
# > The simplest way to deploy a Vespa app.
# ## Deploy application package created with pyvespa
# This section assumes you have an [ApplicationPackage](../../reference-api.rst#vespa.package.ApplicationPackage) instance assigned to `app_package` containing your app desired configuration. If that is not the case, you can learn how to do it by checking [some examples](../../howto/create_app_package/create_app_package.rst). For the purpose of this demonstration we are going to use a minimal (and useless) application package:
# +
from vespa.package import ApplicationPackage
app_package = ApplicationPackage(name="sample_app")
# -
# We can locally deploy our `app_package` using Docker without leaving the notebook, by creating an instance of [VespaDocker](../../reference-api.rst#vespa.deployment.VespaDocker), as shown below:
# +
import os
from vespa.deployment import VespaDocker
disk_folder = os.path.join(os.getenv("WORK_DIR"), "sample_application") # specify your desired absolute path here
vespa_docker = VespaDocker(
port=8080,
disk_folder=disk_folder
)
app = vespa_docker.deploy(
application_package = app_package,
)
# -
# `app` now holds a [Vespa](../../reference-api.rst#vespa.application.Vespa) instance, which we are going to use to interact with our application. Congratulations, you now have a Vespa application up and running.
# ## Learn Vespa by looking at underlying config files
# It is important to know that `pyvespa` simply provides a convenient API to define Vespa application packages from python. `vespa_docker.deploy` export Vespa configuration files to the `disk_folder` defined above. Going through those files is a nice way to start learning about Vespa syntax.
# It is also possible to export the Vespa configuration files representing an application package created with `pyvespa` without deploying the application by using the `export_application_package` method:
vespa_docker.export_application_package(
application_package=app_package,
)
# This will export the application files to an `application` folder within the `disk_folder`.
# ## Deploy application package from Vespa config files
# `pyvespa` provides a subset of the Vespa API, so there will be cases where we want to modify Vespa config files to implement Vespa features that are not yet available in `pyvespa`. We can then modify the files and continue to use pyvespa to deploy and interact with the Vespa application. To do that we can use the `deploy_from_disk` method:
app = vespa_docker.deploy_from_disk(
application_name="sample_app",
application_folder="application"
)
# + nbsphinx="hidden"
# this is a hidden cell. It will not show on the documentation HTML.
from shutil import rmtree
rmtree(disk_folder, ignore_errors=True)
vespa_docker.container.stop()
vespa_docker.container.remove()
| docs/sphinx/source/howto/deploy_app_package/deploy-docker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="37d0200e9bded9a4e9f7546a13f0c4ab35cb116e" _cell_guid="f14a6c39-a6a6-45a6-a6db-c2a95e4d9a40"
# # Bivariate plotting with pandas
#
# <table>
# <tr>
# <td><img src="https://i.imgur.com/bBj1G1v.png" width="350px"/></td>
# <td><img src="https://i.imgur.com/ChK9zR3.png" width="350px"/></td>
# <td><img src="https://i.imgur.com/KBloVHe.png" width="350px"/></td>
# <td><img src="https://i.imgur.com/C7kEWq7.png" width="350px"/></td>
# </tr>
# <tr>
# <td style="font-weight:bold; font-size:16px;">Scatter Plot</td>
# <td style="font-weight:bold; font-size:16px;">Hex Plot</td>
# <td style="font-weight:bold; font-size:16px;">Stacked Bar Chart</td>
# <td style="font-weight:bold; font-size:16px;">Bivariate Line Chart</td>
# </tr>
# <tr>
# <td>df.plot.scatter()</td>
# <td>df.plot.hex()</td>
# <td>df.plot.bar(stacked=True)</td>
# <td>df.plot.line()</td>
# </tr>
# <tr>
# <td>Good for interval and some nominal categorical data.</td>
# <td>Good for interval and some nominal categorical data.</td>
# <td>Good for nominal and ordinal categorical data.</td>
# <td>Good for ordinal categorical and interval data.</td>
# </tr>
# </table>
#
# In the previous notebook, we explored using `pandas` to plot and understand relationships within a single column. In this notebook, we'll expand this view by looking at plots that consider two variables at a time.
#
# Data without relationships between variables is the data science equivalent of a blank canvas. To paint the picture in, we need to understand how variables interact with one another. Does an increase in one variable correlate with an increase in another? Does it relate to a decrease somewhere else? The best way to paint the picture in is by using plots that enable these possibilities.
# + _uuid="3aed82c633067c88ccab2fd99f403211c019aec2" _cell_guid="09b3d35a-a0a3-400b-ba07-6e63d31c57a5"
import pandas as pd
reviews = pd.read_csv("../input/wine-reviews/winemag-data_first150k.csv", index_col=0)
reviews.head()
# + [markdown] _uuid="0e1d38092973a1cddd7deed2f0a8d62acf7eafb4" _cell_guid="73e25c94-3e15-483e-8104-78ca461b11eb"
# ## Scatter plot
#
# The simplest bivariate plot is the lowly **scatter plot**. A simple scatter plot simply maps each variable of interest to a point in two-dimensional space. This is the result:
# + _uuid="2af420e85bbbe6c53c990478a415e269a762ce74" _cell_guid="4265c95d-5d23-4e9d-96c9-99a23dbbf6b2"
reviews[reviews['price'] < 100].sample(100).plot.scatter(x='price', y='points')
# + [markdown] _uuid="424ed88967885b75428f6d5e6af7c5efd60447c0" _cell_guid="a2c42f84-3720-4cfd-ae13-0c8e9bb26b02"
# Note that in order to make effective use of this plot, we had to **downsample** our data, taking just 100 points from the full set. This is because naive scatter plots do not effectively treat points which map to the same place.
#
# For example, if two wines, both costing 100 dollars, get a rating of 90, then the second one is overplotted onto the first one, and we add just one point to the plot.
#
# This isn't a problem if it happens just a few times. But with enough points the distribution starts to look like a shapeless blob, and you lose the forest for the trees:
# + _uuid="2aaa5810e1066c5a7767003014e34da78e845945" _cell_guid="d0ba3d4c-3d5d-4e26-8635-02490690b5fa"
reviews[reviews['price'] < 100].plot.scatter(x='price', y='points')
# + [markdown] _uuid="9e853fc686a49318704cdb6ff9d8c2b977ece321" _cell_guid="2ce7a9bb-2c2c-43f9-a92f-9671a9e72b07"
# There are a few ways to treat this problem. We've already demonstrated one way: sampling the points. Another interesting way to do this that's built right into `pandas` is to use our next plot type, a hexplot.
# + [markdown] _uuid="fe3643a0454a3f21eb4275530f9fe2e38ae8ed4a" _cell_guid="2a04f27a-267c-4ed6-8dec-4a768c7e085e"
# ## Hexplot
#
# A hexplot aggregates points in space into hexagons, and then colorize those hexagons:
# + _uuid="d42c2e53d1c27bf3165067e0bab596dc3eb84614" _cell_guid="8b6c247a-517f-42cd-9c3e-50c85d84356f"
reviews[reviews['price'] < 100].plot.hexbin(x='price', y='points', gridsize=15)
# + [markdown] _uuid="f9f3f0acdf7fdfc426d25489afae9a5dcb1a5014" _cell_guid="9edaadda-3537-48f7-9e7c-cba259a11bc0"
# The data in this plot is directly comprable to the scatter plot from earlier, but the story it tells us is very different. The hexplot provides us with a much more useful view on the dataset, showing that the bottles of wine reviewed by Wine Magazine cluster around 87.5 points and around $20.
#
# Hexplots and scatter plots can by applied to combinations of interval variables or ordinal categorical variables. To help aleviate overplotting, scatter plots (and, to a lesser extent, hexplots) benefit from variables which can take on a wide range of unique values.
# + [markdown] _uuid="dd5ce3eca8388bfec03b200981c9c1a162c0d60e" _cell_guid="c6c7c234-fcdc-4164-99ee-fcc505699a58"
# ## Stacked plots
#
# Scatter plots and hex plots are new. But we can also use the simpler plots we saw in the last notebook.
#
# The easiest way to modify them to support another visual variable is by using stacking. A stacked chart is one which plots the variables one on top of the other.
#
# We'll use a supplemental selection of the five most common wines for this next section.
# + _uuid="25e5f61f206936edf8256325b840b0e1db603f7c" _cell_guid="7ac873e1-acf7-4ca6-9d5c-fc153c3a5f67"
wine_counts = pd.read_csv("../input/most-common-wine-scores/top-five-wine-score-counts.csv",
index_col=0)
# + [markdown] _uuid="01491b0974ae78679f81d46810c80946a9030ae6" _cell_guid="1708ba11-2d36-4bc5-b1a0-5d0df451e71a"
# `wine_counts` counts the number of times each of the possible review scores was received by the five most commonly reviewed types of wines:
# + _uuid="ac42b2097981f02252154e5a605b2f3949e26c31" _cell_guid="fe331c41-b9db-4f32-8d9b-85ac8c2a4874"
wine_counts.head()
# + [markdown] _uuid="e7387d16c3c8034693ef35a0bedbb409b82c9eef" _cell_guid="3660ff18-9a24-4bb2-bb52-45fbf1f5b44b"
# Many `pandas` multivariate plots expect input data to be in this format, with one categorical variable in the columns, one categorical variable in the rows, and counts of their intersections in the entries.
#
# Let's now look at some stacked plots. We'll start with the stacked bar chart.
# + _uuid="62e32c42570d07e74330147e8e15153c558a2b37" _cell_guid="ea937860-2007-462e-9256-ad04565cd92e"
wine_counts.plot.bar(stacked=True)
# + [markdown] _uuid="ced800e5c23e0bcbc681f6e262c77c9e9a995fa7" _cell_guid="5e809373-b709-4b95-b655-bb02237a1063"
# Stacked bar plots share the strengths and weaknesses of univariate bar charts. They work best for nominal categorical or small ordinal categorical variables.
#
# Another simple example is the area plot, which lends itself very naturally to this form of manipulation:
# + _uuid="58648a443b94b1330070dbe11447fa1c1c0f8740" _cell_guid="3cd35cef-6fd0-490b-848c-dd602d83f7c8"
wine_counts.plot.area()
# + [markdown] _uuid="4674d0194edbc4f2e70866f55da2645025101a23" _cell_guid="44b7ee06-5428-439e-9829-bb893e9e28e8"
# Like single-variable area charts, multivariate area charts are meant for nominal categorical or interval variables.
#
# Stacked plots are visually very pretty. However, they suffer from two major problems.
#
# The first limitation is that the second variable in a stacked plot must be a variable with a very limited number of possible values (probably an ordinal categorical, as here). Five different types of wine is a good number because it keeps the result interpretable; eight is sometimes mentioned as a suggested upper bound. Many dataset fields will not fit this critereon naturally, so you will have to "make do", as here, by selecting a group of interest.
#
# The second limitation is one of interpretability. As easy as they are to make, and as pretty as they look, stacked plots are really hard to distinguish values within. For example, looking at the plot above, can you tell which wine is the most common one to have gotten a score of approximately 87: the purple, the red, or the green? It's actually really hard to tell!
# + [markdown] _uuid="c17ecb009cd51ef2e671ae9580abd5b6da12166f" _cell_guid="bbc7dfdb-4090-47d7-85c0-4acccfc672f5"
# ## Bivariate line chart
#
# One plot type we've seen already that remains highly effective when made bivariate is the line chart. Because the line in this chart takes up so little visual space, it's really easy and effective to overplot multiple lines on the same chart.
# + _uuid="f3b3138890544d6654a5e27c5483bdb1a4470980" _cell_guid="742c43c9-885c-4969-9c23-a5a63d940af9"
wine_counts.plot.line()
# + [markdown] _uuid="f7ffee305a0cbdb41a16d4ab05e02f81fa89e7b1" _cell_guid="9b9c3c34-de96-4d2a-a1ce-ecbce18e8d69"
# Using a line chart this way makes inroads against the second limitation of stacked plotting. Bivariate line charts are much more interpretable: we can see in this chart fairly easily that the green wine (the Chardonnay) very slightly edges out the Pinot Noir around the 87-point scoreline.
# + [markdown] _uuid="9d066077eade3df7e43eeb980381a5e14f42a974" _cell_guid="b82632f4-5a21-43e7-9df7-2757e4dcfe0b"
# ## Exercises
#
# In this section we introduced and explored some common bivariate plot types:
#
# * Scatter plots
# * Hex plots
# * Stacked bar charts and area charts
# * Bivariate line charts
#
# Let's now put what we've learned to the test!
#
# Try answering the following questions:
#
# 1. A scatter plot or hex plot is good for what two types of data?
# 2. What type of data makes sense to show in a stacked bar chart, but not in a bivariate line chart?
# 3. What type of data makes sense to show in a bivariate line chart, but not in a stacked bar chart?
# 4. Suppose we create a scatter plot but find that due to the large number of points it's hard to interpret. What are two things we can do to fix this issue?
#
# To see the answers, click the "Output" button on the cell below.
# + _kg_hide-output=true _kg_hide-input=true _uuid="8edeb224c969e631b07d8aeeab7d8839cfa32ab5" _cell_guid="88c2e00f-1595-40f6-8099-e4c860b6ea03"
from IPython.display import HTML
HTML("""
<ol>
<li>Scatter plots and hex plots work best with a mixture of ordinal categorical and interval data.</li>
<br/>
<li>Nominal categorical data makes sense in a (stacked) bar chart, but not in a (bivariate) line chart.</li>
<br/>
<li>Interval data makes sense in a bivariate line chart, but not in a stacked bar chart.</li>
<br/>
<li>One way to fix this issue would be to sample the points. Another way to fix it would be to use a hex plot.</li>
</ol>
""")
# + [markdown] _uuid="b98a4ae7804eee0a2adc625d17b72fd6f0e7d1ae" _cell_guid="a614ee1c-41cb-4338-9fd7-a06a8d9dc0be"
# Next, let's replicate some plots. Recall the Pokemon dataset from earlier:
# + _uuid="423c0a457a6487d95b41785633fb2b00d7dc3cb9" _cell_guid="df5eca2b-aad2-4583-b3cd-48de4b6bb420"
pokemon = pd.read_csv("../input/pokemon/Pokemon.csv", index_col=0)
pokemon.head()
# + [markdown] _uuid="740558330ba5fba187515204a87261cd4d227f8a" _cell_guid="c65672c7-18f9-4465-90cb-d17264422611"
# For the exercises that follow, try forking this notebook and replicating the plots that follow. To see the answers, hit the "Input" button below to un-hide the code.
# + _kg_hide-input=true _uuid="028798b2b4cee37374fb53f3c6e5b4658f2033b3" _cell_guid="1fc74bae-d290-4f72-b481-78ead07a1c6f"
pokemon.plot.scatter(x='Attack',y='Defense')
# + _kg_hide-input=true _uuid="4027dac93bcf3d77eb4e07c49c831be30aa9c0cf" _cell_guid="9b9b3aa5-a8ce-4717-bc6f-ad8ef09282e9"
pokemon.plot.hexbin(x='Attack',y='Defense',gridsize=15)
# + [markdown] _uuid="1c385e7a8dbb16903f2ce25eac123dcc34ee35b5" _cell_guid="cc3defbf-88c7-4744-94e2-1df1254ac325"
# For thee next plot, use the following data:
# + _uuid="dc04a757fa939180bd36c340ba154a9d5bca9df1" _cell_guid="fb61f746-3c6b-4461-b89d-c31f7414d9e8"
pokemon_stats_legendary = pokemon.groupby(['Legendary', 'Generation']).mean()[['Attack', 'Defense']]
# + _kg_hide-input=true _uuid="c7f4927c36f617a15b56da12340c68d18a5daf35" _cell_guid="0f91be2b-c180-4097-8648-af9f105ae93c"
pokemon_stats_legendary.plot.bar(stacked='True')
# + [markdown] _uuid="7002d9f9c6b74c6c0d1ea352e126b22575e91483" _cell_guid="e58893f2-a0a8-4ae1-a919-dce6ba47c39a"
# For the next plot, use the following data:
# + _uuid="4c929c49bd752d1b047c9776f3296306bf896fc7" _cell_guid="ebb7cc3d-4b72-4564-a5df-64c15af2d49b"
pokemon_stats_by_generation = pokemon.groupby('Generation').mean()[['HP', 'Attack', 'Defense', 'Sp. Atk', 'Sp. Def', 'Speed']]
# + _kg_hide-input=true _uuid="406782d0963db63ca0d4b29cff8207256da62584" _cell_guid="77a0098d-7fbd-440a-a736-1232cf469256"
pokemon_stats_by_generation.plot.line()
# + [markdown] _uuid="73742ea40362bbe24bdf15d0303b3bc0a61501b5" _cell_guid="4974dc7f-17ee-45ef-8e77-cf14dc575da3"
# ## Conclusion
#
# In this section we introduced and explored some common bivariate plot types:
#
# * Scatter plots
# * Hex plots
# * Stacked bar charts and area charts
# * Bivariate line charts
#
# In the next section we will move on to exploring another plotting library, `seaborn`, which compliments `pandas` with many more advanced data visualization tools for you to use.
#
# [Click here to move on to the next section, "Plotting with seaborn"](https://www.kaggle.com/residentmario/plotting-with-seaborn/).
| bivariate_plotting_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get ChIP data
import pandas as pd
import glob
import os
import seaborn as sb
# %matplotlib inline
import matplotlib.pyplot as plt
import multiprocessing as mp
import subprocess
p = mp.Pool(64)
df = pd.read_csv('/home/shared/Data/encode/mouse/ChIP/metadata.tsv',sep='\t')
df
f_chmm = glob.glob('/home/shared/Data/encode/mouse/enhancer/replicated/*')
f_chmm_dic = {}
for f in f_chmm:
term = f.split('/')[-1].split('_')[0]
age = f.split('/')[-1].split('_')[1]
if term not in f_chmm_dic.keys():
f_chmm_dic[term] = []
f_chmm_dic[term].append(age)
else:
f_chmm_dic[term].append(age)
# ### download peaks of H3K27ac
def get_file(tup):
mark,url,acc,name = tup
os.system('wget -P /home/shared/Data/encode/mouse/ChIP/bigwig/{}/ {}'.format(mark,url))
os.system('mv /home/shared/Data/encode/mouse/ChIP/bigwig/{}/{}.bigWig /home/shared/Data/encode/mouse/ChIP/bigwig/{}/{}.bigWig'.format(mark,acc,mark,name))
# +
f_acc_dic = {}
meta_dic = {}
count = 0
meta_tups = []
for r in df.iterrows():
term = '-'.join(r[1]['Biosample term name'].split(' '))
age = 'e{}'.format(r[1]['Biosample Age'].split(' ')[0])
if age == "e0":
age = 'P0'
rep = r[1]['Biological replicate(s)']
f_acc = r[1]['File accession']
target = r[1]['Experiment target']
url = r[1]['File download URL']
if age not in ['e10.5','e8','eunknown']:
if term not in ['brain','brown-adipose-tissue']:
if target == 'H3K27ac-mouse':
mark = 'H3K27ac'
if rep == '1, 2':
if r[1]['Audit ERROR'] != "extremely low read depth":
if r[1]['Output type'] == 'fold change over control':
if term == 'embryonic-facial-prominence':
if term not in meta_dic.keys():
meta_dic[term] = []
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
else:
if age not in meta_dic[term]:
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
count += 1
meta_tups.append((mark,url,f_acc,'{}_{}_{}_{}_{}'.format(term,age,'1-2',target,f_acc)))
#os.system('wget -P /home/shared/Data/encode/mouse/ChIP/bigwig/H3K27ac/ {}'.format(url))
#os.system('mv /home/shared/Data/encode/mouse/ChIP/bigwig/H3K27ac/{}.bigWig /home/shared/Data/encode/mouse/ChIP/bigwig/H3K27ac/{}_{}_{}_{}.bigWig'.format(f_acc,term,age,target,f_acc))
# -
meta_tups
p.map(get_file,meta_tups)
# ### download peaks of K27me3, K9me3, K4me1, K4me2, K4me3, K36me3
targets = list(set(df['Experiment target']))
# +
f_acc_dic = {}
meta_dic = {}
count = 0
for indx,exp in enumerate(targets):
meta_dic = {}
plt.figure(indx)
data_df = pd.DataFrame(0,index=data_df.index, columns=data_df.columns)
mark = exp.split('-')[0]
if mark in ['H3K27me3','H3K9me3','H3K9ac','H3K4me1','H3K4me2','H3K4me3','H3K36me3','H3K27ac']:
#os.system('mkdir /home/shared/Data/encode/mouse/{}_rep_peaks'.format(exp.split('-')[0]))
count = 0
for r in df.iterrows():
term = '-'.join(r[1]['Biosample term name'].split(' '))
age = 'e{}'.format(r[1]['Biosample Age'].split(' ')[0])
if age == "e0":
age = 'P0'
rep = r[1]['Biological replicate(s)']
f_acc = r[1]['File accession']
target = r[1]['Experiment target']
url = r[1]['File download URL']
if age not in ['e10.5','e8','eunknown']:
if term not in ['brain','brown-adipose-tissue']:
if target == exp:
if rep == '1, 2':
if r[1]['Audit ERROR'] != "extremely low read depth":
if r[1]['Output type'] == 'replicated peaks':
if term not in meta_dic.keys():
meta_dic[term] = []
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
else:
if age not in meta_dic[term]:
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
count += 1
data_df.ix[term,age] += 1
print(mark, count)
#avail_data_df = get_data_filled(meta_dic,data_df)
sb.heatmap(data_df,annot=True,fmt="d",vmax=4,vmin=0)
#os.system('wget -P /home/shared/Data/encode/mouse/{}_rep_peaks/ {}'.format(mark,url))
#os.system('mv /home/shared/Data/encode/mouse/{}_rep_peaks/{}.bed.gz /home/shared/Data/encode/mouse/{}_rep_peaks/{}_{}_{}_{}.bed.gz'.format(mark,f_acc,mark,term,age,target,f_acc))
# -
meta_dic
# #### plot chIP data availability
# +
def get_data_filled(dic,zdf):
for i in dic.keys():
for t in dic[i]:
zdf.ix[i,t] = 1
return zdf
data_df = pd.DataFrame(0,index=data_df.index, columns=data_df.columns)
avail_data_df = get_data_filled(meta_dic,data_df)
sb.heatmap(avail_data_df)
# -
# #### plot chromHMM data availability
data_analysis = {'embryonic-facial-prominence': ['e11.5', 'e15.5', 'e12.5', 'e13.5', 'e14.5'],
'forebrain': ['e12.5', 'e11.5', 'e15.5', 'e13.5', 'e14.5', 'e16.5', 'P0'],
'heart': ['e11.5', 'e16.5', 'e13.5', 'e15.5', 'P0', 'e12.5', 'e14.5'],
'hindbrain': ['e14.5', 'e11.5', 'e15.5', 'e13.5', 'P0', 'e12.5', 'e16.5'],
'intestine': ['P0', 'e15.5', 'e14.5', 'e16.5'],
'kidney': ['e16.5', 'e14.5', 'e15.5', 'P0'],
'limb': ['e12.5', 'e13.5', 'e15.5', 'e14.5', 'e11.5'],
'liver': ['P0', 'e16.5', 'e11.5', 'e15.5', 'e12.5', 'e13.5', 'e14.5'],
'lung': ['e14.5', 'P0', 'e16.5', 'e15.5'],
'midbrain': ['e16.5', 'e13.5', 'e14.5', 'e11.5', 'e15.5', 'e12.5', 'P0'],
'neural-tube': ['e14.5', 'e13.5', 'e11.5', 'e15.5', 'e12.5'],
'stomach': ['e14.5', 'P0', 'e15.5', 'e16.5']}
time_pt = ['P0','e11.5','e12.5','e13.5','e14.5','e15.5','e16.5']
data_df = pd.DataFrame(0,index=data_analysis.keys(),columns=time_pt)
for i in data_analysis.keys():
for t in data_analysis[i]:
data_df.ix[i,t] = 1
data_df['sum'] = list(data_df.sum(axis=1))
data_df.sort_values(by='sum',inplace=True)
del(data_df['sum'])
sb.set_context('paper',font_scale=1.8)
sb.heatmap(data_df)
# ### download bam files
targets = list(set(df['Experiment target']))
def get_file(tup):
mark,url,acc,name = tup
os.system('wget -P /home/shared/Data/encode/mouse/{}_bam/ {}'.format(mark,url))
os.system('mv /home/shared/Data/encode/mouse/{}_bam/{}.bam /home/shared/Data/encode/mouse/{}_bam/{}.bam'.format(mark,acc,mark,name))
import multiprocessing as mp
p = mp.Pool(64)
# +
f_acc_dic = {}
meta_dic = {}
count = 0
for exp in targets:
mark = exp.split('-')[0]
if mark in ['H3K27ac']:
os.system('mkdir /home/shared/Data/encode/mouse/{}_bam'.format(exp.split('-')[0]))
count = 0
meta_tups = []
for r in df.iterrows():
term = '-'.join(r[1]['Biosample term name'].split(' '))
age = 'e{}'.format(r[1]['Biosample Age'].split(' ')[0])
if age == "e0":
age = 'P0'
rep = r[1]['Biological replicate(s)']
f_acc = r[1]['File accession']
target = r[1]['Experiment target']
url = r[1]['File download URL']
if age not in ['e10.5','e8','eunknown']:
if term not in ['brain','brown-adipose-tissue']:
if target == exp:
#if rep == '1, 2':
if r[1]['Audit ERROR'] != "extremely low read depth":
if r[1]['Output type'] == 'alignments':
if term not in meta_dic.keys():
meta_dic[term] = []
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
else:
if age not in meta_dic[term]:
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
meta_tups.append((mark,url,f_acc,'{}_{}_{}_{}_{}'.format(term,age,rep,target,f_acc)))
print(mark, len(meta_tups))
p.map(get_file,meta_tups)
# -
# ### download RNA-seq data
meta = pd.read_csv('/home/shared/Data/encode/mouse/RNA-seq/metadata.tsv',sep='\t')
meta.columns
meta[['File accession','Biosample term name','Biosample Age','Biological replicate(s)','Technical replicate','File format','Output type','Assay','Biosample term id','Experiment accession']]
# +
meta_dic = {}
meta_tups = []
data_df = pd.DataFrame(0,index=data_df.index, columns=data_df.columns)
for r in meta.iterrows():
term = '-'.join(r[1]['Biosample term name'].split(' '))
age = 'e{}'.format(r[1]['Biosample Age'].split(' ')[0])
sex = r[1]['Biosample sex']
if age == "e0":
age = 'P0'
rep = r[1]['Biological replicate(s)']
f_acc = r[1]['File accession']
url = r[1]['File download URL']
target = r[1]['Assay']
if term in data_df.index:
if target == 'RNA-seq':
if age not in ['e10.5','e10','e8','e14','e18','eunknown']:
if term not in ['brain','brown-adipose-tissue']:
if r[1]['Output type'] == 'gene quantifications':
if term not in meta_dic.keys():
meta_dic[term] = []
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
else:
if age not in meta_dic[term]:
meta_dic[term].append(age)
f_acc_dic[f_acc] = "{}_{}".format(term,age)
#print(term,age)
#break
data_df.ix[term,age] += 1
# #meta_tups.append(
# (
# target,
# url,
# f_acc,
# '{}_{}_{}_{}_{}_{}'.format(term,age,sex,rep,target,f_acc)
# )
# )
sb.heatmap(data_df,annot=True,fmt='d')
print(meta_dic)
# +
def get_file(tup):
mark,url,acc,name = tup
os.system('wget -P /home/shared/Data/encode/mouse/RNA-seq/gene_quantification/ {}'.format(url))
os.system('mv /home/shared/Data/encode/mouse/RNA-seq/gene_quantification/{}.tsv /home/shared/Data/encode/mouse/RNA-seq/gene_quantification/{}.tsv'.format(acc,name))
import multiprocessing as mp
p = mp.Pool(64)
out = p.map(get_file,meta_tups)
# -
gq_files = glob.glob('/home/shared/Data/encode/mouse/RNA-seq/gene_quantification/*')
def get_filtered_genes(df,name):
gene_list = list(df['gene_id'])
sel_g = ['ENS' in c for c in gene_list]
sel_df = pd.DataFrame(df[sel_g][['gene_id','TPM']])
sel_df.columns = ['gene_id',name]
return sel_df
gq_mat = pd.DataFrame()
for f in gq_files:
gq_df = pd.read_csv(f,sep='\t')
f_name = f.split('/')[-1].split('.tsv')[0]
gq_df_sel = get_filtered_genes(gq_df,f_name)
gq_df_sel.set_index('gene_id',inplace=True)
if gq_mat.shape == (0,0):
gq_mat = gq_df_sel
else:
gq_mat = gq_mat.join(gq_df_sel)
gq_mat.isnull().values.any()
gq_mat.fillna(value=0,inplace=True)
gq_mat.to_csv('/home/shared/Data/encode/mouse/RNA-seq/Expression_matrix.txt',sep='\t')
gq_mat = pd.read_csv('/home/shared/Data/encode/mouse/Expression_matrix.txt',sep='\t')
gq_mat.set_index('gene_id',inplace=True)
gq_mat_log2TPM = np.log2(gq_mat+1)
gq_mat_log2TPM.to_csv('/home/shared/Data/encode/mouse/gene_expression_log2TPM_signal_matrix.txt',sep='\t')
import seaborn as sb
# %matplotlib inline
import matplotlib.pyplot as plt
stages = ['P0','e11.5','e12.5','e13.5','e14.5','e15.5','e16.5']
heart_to_remove = ['ENCFF662WLV','ENCFF705YYN','unknown']
for i,s in enumerate(stages):
plt.figure(i)
heart = gq_mat.ix[:,['heart_{}'.format(s) in c for c in gq_mat.columns]]
heart = heart.ix[:,[all(sb not in c for sb in heart_to_remove) for c in heart.columns]]
sb.heatmap(heart.corr(),vmin=0,vmax=1)
# ### get TPM (transcript per millions) for each enhancers
# ##### To find total number of reads in bam
# +
import subprocess
output = subprocess.check_output("samtools view -c /home/shared/Data/encode/mouse/H3K36me3_bam/hindbrain_e13.5_1_H3K36me3-mouse_ENCFF669UYV.bam",shell=True)
print(int(output.strip()))
# -
# ##### Get TPMs
df = pd.read_csv('/home/shared/Data/encode/mouse/ChIP/metadata.tsv',sep='\t')
files = glob.glob('/home/shared/Data/encode/mouse/ChIP/bam/H3K27ac_bam/*')
def get_reads_regions(f):
name = f.split('/')[-1].split('.bam')[0]
n_arr = name.split('_')
tissue = n_arr[0]
age = n_arr[1]
sex = n_arr[2]
rep = n_arr[3]
assay = n_arr[4]
acc = n_arr[5]
subprocess.call('bedtools coverage -a /home/vamin/projects/epee/peak_overlapped_pc_tss_subtracted_enh_mouse_pc_tss_subtracted.bed -b {} > /home/shared/Data/encode/mouse/ChIP/{}_H3K27ac_reads.bed'.format(f,name),shell=True)
for f in files:
get_reads_regions(f)
coverage_files = glob.glob('/home/shared/Data/encode/mouse/ChIP/*_reads.bed')
coverage_files[0]
len(coverage_files)
import numpy as np
enh_signal_df = pd.DataFrame()
for f in coverage_files:
name = f.split('/')[-1].split('_H3K27ac_reads.bed')[0]
counts_df = pd.read_csv(f,sep='\t',header=None)
tpms = (counts_df[3]/counts_df[5])*(1/sum(counts_df[3]/counts_df[5]))*1e6
signal = np.log2(tpms+1)
if enh_signal_df.shape == (0,0):
enh_signal_df = counts_df.ix[:,[0,1,2]]
enh_signal_df.columns = ['chr','start','stop']
enh_signal_df[name] = signal
else:
enh_signal_df[name] = signal
enh_signal_df.to_csv('/home/shared/Data/encode/mouse/enh_H3K27ac_log2TPM_signal_matrix.txt',sep='\t')
import seaborn as sns
# %matplotlib inline
import numpy as np
sns.distplot(np.log2(tpms+1))
data_analysis = {'embryonic-facial-prominence': ['e11.5', 'e15.5', 'e12.5', 'e13.5', 'e14.5'],
'forebrain': ['e12.5', 'e11.5', 'e15.5', 'e13.5', 'e14.5', 'e16.5', 'P0'],
'heart': ['e11.5', 'e16.5', 'e13.5', 'e15.5', 'P0', 'e12.5', 'e14.5'],
'hindbrain': ['e14.5', 'e11.5', 'e15.5', 'e13.5', 'P0', 'e12.5', 'e16.5'],
'intestine': ['P0', 'e15.5', 'e14.5', 'e16.5'],
'kidney': ['e16.5', 'e14.5', 'e15.5', 'P0'],
'limb': ['e12.5', 'e13.5', 'e15.5', 'e14.5', 'e11.5'],
'liver': ['P0', 'e16.5', 'e11.5', 'e15.5', 'e12.5', 'e13.5', 'e14.5'],
'lung': ['e14.5', 'P0', 'e16.5', 'e15.5'],
'midbrain': ['e16.5', 'e13.5', 'e14.5', 'e11.5', 'e15.5', 'e12.5', 'P0'],
'neural-tube': ['e14.5', 'e13.5', 'e11.5', 'e15.5', 'e12.5'],
'stomach': ['e14.5', 'P0', 'e15.5', 'e16.5']}
time_pt = ['P0','e11.5','e12.5','e13.5','e14.5','e15.5','e16.5']
data_df = pd.DataFrame(0,index=data_analysis.keys(),columns=time_pt)
def get_data_filled(dic,zdf):
for i in dic.keys():
for t in dic[i]:
zdf.ix[i,t] = 1
return zdf
for i in data_analysis.keys():
for t in data_analysis[i]:
data_df.ix[i,t] = 1
data_df['sum'] = list(data_df.sum(axis=1))
data_df.sort_values(by='sum',inplace=True)
del(data_df['sum'])
sb.set_context('paper',font_scale=1.8)
sb.heatmap(data_df)
data_to_remove = ['ENCFF001KGS','ENCFF001KGT']
for indx,val in enumerate(data_analysis['heart']):
temp = enh_signal_df.ix[:,['heart_{}'.format(val) in c for c in enh_signal_df.columns]]
tempf = temp.ix[:,[all(sb not in c for sb in data_to_remove) for c in temp.columns]]
plt.figure(indx)
sb.heatmap(tempf.corr(),vmin=0,vmax=1)
data_to_remove = ['ENCFF001KGS','ENCFF001KGT']
| enh_gene/ENCODE_metadata_to_dataframes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from chatbot import CAChatBot
os.chdir(os.path.join(os.getcwd(), '..'))
chatbot = CAChatBot()
# -
# ### 在这个demo中,你将看到如下几个问题类型的回答和介绍:
# 1. index_value
# 2. index_overall
# 3. index_2_overall
# 4. indexes_m_compare
# 5. indexes_n_compare
# 6. indexes_g_compare
# 7. indexes_2m_compare
# 8. indexes_2n_compare
# ## 1. index_value
# 回答了某一个年度的某项(可以是一项也可以是多项)指标的值。
# + pycharm={"name": "#%%\n"}
chatbot.query('2013年的货邮周转量为?')
# + pycharm={"name": "#%%\n"}
chatbot.query('2011年货邮周转量和游客周转量是多少?')
# 名字近似但不对的,会对其进行模糊查询并匹配最相似的。例 “游客周转量” =》 “旅客周转量”。
# + pycharm={"name": "#%%\n"}
chatbot.query('2012年节能减排的值怎样?')
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 2. index_overall
# 回答某一个年度的某项(一或多项)指标占其总指标的百分比和倍数。
# + pycharm={"name": "#%%\n"}
chatbot.query('2013年旅客周转量占其总体百分之多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('2013年货邮周转量和全行业取得驾驶执照飞行员占总体多少?')
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 3. index_2_overall
# 回答两个年度的一项指标占比总指标的变化(主要指占比是增加还是减少)。
# + pycharm={"name": "#%%\n"}
chatbot.query('2012年游客周转量占总体的百分比比去年变化多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('2012年游客周转量和货邮周转量占总体的百分比比去年变化多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('13年的运输总周转量占父级的倍数比11年降低多少?')
# -
# ## 4. indexes_m_compare
# 回答某一个年度中某一项指标与另一项指标的倍数关系,只有单位相同的指标才可以比较。
# + pycharm={"name": "#%%\n"}
chatbot.query('2011年旅客周转量是货邮周转量的几倍?')
# + pycharm={"name": "#%%\n"}
chatbot.query('11年旅客周转量是新增机场数量的几倍?')
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 5. indexes_n_compare
# 回答某一个年度中某一项指标与另一项指标的比较关系(多少关系),同样也只有单位相同才可比较。
# + pycharm={"name": "#%%\n"}
chatbot.query('13年旅客周转量比货邮周转量多多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('12年旅客周转量比货邮运输量少多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('11年旅客周转量比新增机场数量多多少?')
# -
# ## 6. indexes_g_compare
# 回答某一个年度中某项(一或多项)指标的同比变化(同比只能是比去年的数据)。
# + pycharm={"name": "#%%\n"}
chatbot.query('2013年旅客运输量同比上升多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('12年游客运输量和货邮周转量同比变化多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('11年旅客周转量同比增长?')
# -
# ## 7. indexes_2m_compare
# 回答某两个年度的某一项指标之间的倍数关系,非数值类型无法比较。
# + pycharm={"name": "#%%\n"}
chatbot.query('13年游客周转量是11年的几倍?')
# + pycharm={"name": "#%%\n"}
chatbot.query('13年游客周转量和运输总周转量是11年的几倍?')
# + pycharm={"name": "#%%\n"}
chatbot.query('12年新增机场是11年的几倍?')
# -
# ## 8. indexes_2n_compare
# 回答某两个年度的某项指标之间的比较关系(多少关系),非数值类型无法比较
# + pycharm={"name": "#%%\n"}
chatbot.query('12年比13年货邮运输量增加了多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('13年同去年相比,货邮周转量变化了多少?')
# + pycharm={"name": "#%%\n"}
chatbot.query('2012年节能减排比去年变化了多少?')
| demo/demo2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using geoprocessing tools
#
# In ArcGIS API for Python, geoprocessing toolboxes and tools within them are represented as Python module and functions within that module. To learn more about this organization, refer to the page titled [Accessing geoprocessing tools](https://developers.arcgis.com/python/guide/accessing-geoprocessing-tools/).In this part of the guide, we will observe:
#
# - [Invoking geoprocessing tools](#invoking-geoprocessing-tools)
# - [Understanding tool input parameter and output return types](#understanding-tool-input-parameter-and-output-return-types)
# - [Using helper types](#using-helper-types)
# - [Using strings as input](#using-strings-as-input)
# - [Tools with multiple outputs](#tools-with-multiple-outputs)
# - [Invoking tools that create multiple outputs](#invoking-tools-that-create-multiple-outputs)
# - [Using named tuple to access multiple outputs](#using-named-tuple-to-access-multiple-outputs)
# - [Tools that export map image layer as output](#tools-that-export-map-image-layer-as-output)
#
# <a id="invoking-geoprocessing-tools"></a>
# ## Invoking Geoprocessing Tools
# You can execute a geoprocessing tool easily by importing its toolbox as a module and calling the function for the tool. Let us see how to execute the `extract_zion_data` tool from the Zion toolbox URL:
# +
# connect to ArcGIS Online
from arcgis.gis import GIS
from arcgis.geoprocessing import import_toolbox
gis = GIS()
# import the Zion toolbox
zion_toolbox_url = 'http://gis.ices.dk/gis/rest/services/Tools/ExtractZionData/GPServer'
zion = import_toolbox(zion_toolbox_url)
# -
result = zion.extract_zion_data()
# Thus, executing a geoprocessing tool is that simple. Let us learn a few more concepts that will help in using these tools efficiently.
#
# <a id="understanding-tool-input-parameter-and-output-return-types"></a>
# ## Understanding tool input parameter and output return types
#
# The functions for calling geoprocessing tools can accept and return built-in Python types such as str, int, bool, float, dicts, datetime.datetime as well as some helper types defined in the ArcGIS API for Python such as the following:
# * `arcgis.features.FeatureSet` - a set of features
# * `arcgis.geoprocessing.LinearUnit` - linear distance with specified units
# * `arcgis.geoprocessing.DataFile` - a url or item id referencing data
# * `arcgis.geoprocessing.RasterData` - url or item id and format of raster data
#
# The tools can also accept lists of the above types.
#
# **Note**: When the helper types are used an input, the function also accepts strings in their place. For example '5 Miles' can be passed as an input instead of LinearUnit(5, 'Miles') and a URL can be passed instead of a `DataFile` or `RasterData` input.
#
# Some geoprocessing tools are configured to return an `arcgis.mapping.MapImageLayer` for visualizing the results of the tool.
#
# In all cases, the documentation of the tool function indicates the type of input parameters and the output values.
# <a id="using-helper-types"></a>
# ### Using helper types
#
# The helper types (`LinearUnit`, `DataFile` and `RasterData`) defined in the `arcgis.geoprocessing` module are simple classes that hold strings or URLs and have a dictionary representation.
#
# The `extract_zion_data()` tool invoked above returns an output zip file as a `DataFile`:
type(result)
# The output `Datafile` can be queried as shown in the snippet below.
result
# The value types such as `DataFile` include helpful methods such as download:
result.download()
# <a id="using-strings-as-input"></a>
# ### Using strings as input
#
# Strings can also be used as inputs in place of the helper types such as `LinearUnit`, `RasterData` and `DataFile`.
#
# The example below calls the viewshed tool to compute and display the geographical area that is visible from a clicked location on the map. The function accepts an observation point as a `FeatureSet` and a viewshed distance as a `LinearUnit`, and returns a `FeatureSet`:
viewshed = import_toolbox('http://sampleserver1.arcgisonline.com/ArcGIS/rest/services/Elevation/ESRI_Elevation_World/GPServer')
help(viewshed.viewshed)
import arcgis
arcgis.env.out_spatial_reference = 4326
map = gis.map('South San Francisco', zoomlevel=12)
map
# 
# The code snippet below adds an event listener to the map, such that when clicked, `get_viewshed()` is called with the map widget and clicked point geometry as inputs. The event handler creates a `FeatureSet` from the clicked point geometry, and uses the string '5 Miles' as input for the viewshed_distance parameter instead of creating a `LinearUnit` object. These are passed into the viewshed function that returns the viewshed from the observation point. The map widget is able to draw the returned `FeatureSet` using its `draw()` method:
# +
from arcgis.features import Feature, FeatureSet
def get_viewshed(m, g):
res = viewshed.viewshed(FeatureSet([Feature(g)]),"5 Miles") # "5 Miles" or LinearUnit(5, 'Miles') can be passed as input
m.draw(res)
map.on_click(get_viewshed)
# -
# <a id="tools-with-multiple-outputs"></a>
# ## Tools with multiple outputs
#
# Some Geoprocessing tools can return multiple results. For these tools, the corresponding function returns the multiple output values as a [named tuple](https://docs.python.org/3/library/collections.html#namedtuple-factory-function-for-tuples-with-named-fields).
#
# The example below uses a tool that returns multiple outputs:
sandiego_toolbox_url = 'https://gis-public.co.san-diego.ca.us/arcgis/rest/services/InitialResearchPacketCSV_Phase2/GPServer'
multioutput_tbx = import_toolbox(sandiego_toolbox_url)
help(multioutput_tbx.initial_research_packet_csv)
# <a id="invoking-tools-that-create-multiple-outputs"></a>
# ### Invoking tools that create multple outputs
#
# The code snippet below shows how multiple outputs returned from a tool can be automatically unpacked by Python into multiple variables. Also, since we're not interested in the job status output, we can discard it using "_" as the variable name:
report_output_csv_file, output_map_flags_file, soil_output_file, _ = multioutput_tbx.initial_research_packet_csv()
report_output_csv_file
output_map_flags_file
soil_output_file
# <a id="using-named-tuple-to-access-multiple-outputs"></a>
# ### Using named tuple to access multiple tool outputs
# The code snippet below shows using a named tuple to access the multiple outputs returned from the tool:
results = multioutput_tbx.initial_research_packet_csv()
results.report_output_csv_file
results.job_status
# <a id="tools-that-export-map-image-layer-as-output"></a>
# ## Tools that export MapImageLayer as output
#
# Some Geoprocessing tools are configured to return their output as MapImageLayer for easier visualization of the results. The resultant layer can be added to a map or queried.
#
# An example of such a tool is below:
hotspots = import_toolbox('https://sampleserver6.arcgisonline.com/arcgis/rest/services/911CallsHotspot/GPServer')
help(hotspots.execute_911_calls_hotspot)
result_layer, output_features, hotspot_raster = hotspots.execute_911_calls_hotspot()
result_layer
hotspot_raster
# The resultant hotspot raster can be visualized in the Jupyter Notebook using the code snippet below:
from IPython.display import Image
Image(hotspot_raster['mapImage']['href'])
| guide/08-using-geoprocessing-tools/using-geoprocessing-tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="JbDHnhet8CWy"
# _Lambda School Data Science_
#
# # Sequence your narrative
#
# Today we will create a sequence of visualizations inspired by [<NAME>'s 200 Countries, 200 Years, 4 Minutes](https://www.youtube.com/watch?v=jbkSRLYSojo).
#
# Using this [data from Gapminder](https://github.com/open-numbers/ddf--gapminder--systema_globalis/):
# - [Income Per Person (GDP Per Capital, Inflation Adjusted) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--income_per_person_gdppercapita_ppp_inflation_adjusted--by--geo--time.csv)
# - [Life Expectancy (in Years) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--life_expectancy_years--by--geo--time.csv)
# - [Population Totals, by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv)
# - [Entities](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv)
# - [Concepts](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--concepts.csv)
# + [markdown] colab_type="text" id="zyPYtsY6HtIK"
# Objectives
# - sequence multiple visualizations
# - combine qualitative anecdotes with quantitative aggregates
#
# Links
# - [<NAME>’s TED talks](https://www.ted.com/speakers/hans_rosling)
# - [Spiralling global temperatures from 1850-2016](https://twitter.com/ed_hawkins/status/729753441459945474)
# - "[The Pudding](https://pudding.cool/) explains ideas debated in culture with visual essays."
# - [A Data Point Walks Into a Bar](https://lisacharlotterost.github.io/2016/12/27/datapoint-in-bar/): a thoughtful blog post about emotion and empathy in data storytelling
# + [markdown] colab_type="text" id="SxTJBgRAW3jD"
# ## Make a plan
#
# #### How to present the data?
#
# Variables --> Visual Encodings
# - Income --> x
# - Lifespan --> y
# - Region --> color
# - Population --> size
# - Year --> animation frame (alternative: small multiple)
# - Country --> annotation
#
# Qualitative --> Verbal
# - Editorial / contextual explanation --> audio narration (alternative: text)
#
#
# #### How to structure the data?
#
# | Year | Country | Region | Income | Lifespan | Population |
# |------|---------|----------|--------|----------|------------|
# | 1818 | USA | Americas | ### | ## | # |
# | 1918 | USA | Americas | #### | ### | ## |
# | 2018 | USA | Americas | ##### | ### | ### |
# | 1818 | China | Asia | # | # | # |
# | 1918 | China | Asia | ## | ## | ### |
# | 2018 | China | Asia | ### | ### | ##### |
#
# + [markdown] colab_type="text" id="S2dXWRTFTsgd"
# ## More imports
# + colab_type="code" id="y-TgL_mA8OkF" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# + [markdown] colab_type="text" id="CZGG5prcTxrQ"
# ## Load & look at data
# + colab_type="code" id="-uE25LHD8CW0" colab={}
income = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--income_per_person_gdppercapita_ppp_inflation_adjusted--by--geo--time.csv')
# + colab_type="code" id="gg_pJslMY2bq" colab={}
lifespan = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--life_expectancy_years--by--geo--time.csv')
# + colab_type="code" id="F6knDUevY-xR" colab={}
population = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv')
# + colab_type="code" id="hX6abI-iZGLl" colab={}
entities = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv')
# + colab_type="code" id="AI-zcaDkZHXm" colab={}
concepts = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--concepts.csv')
# + colab_type="code" id="EgFw-g0nZLJy" outputId="d1cb27e0-45f4-495e-b437-d8a242539af4" executionInfo={"status": "ok", "timestamp": 1568305871553, "user_tz": 360, "elapsed": 342, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
income.shape, lifespan.shape, population.shape, entities.shape, concepts.shape
# + colab_type="code" id="I-T62v7FZQu5" outputId="701c52e8-5752-4912-8af3-080d7fb7c44a" executionInfo={"status": "ok", "timestamp": 1568305882671, "user_tz": 360, "elapsed": 299, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
income.head()
# + colab_type="code" id="2zIdtDESZYG5" outputId="21244fb1-4f5b-4a51-8c7b-ca7d53d7132a" executionInfo={"status": "ok", "timestamp": 1568305883598, "user_tz": 360, "elapsed": 309, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
lifespan.head()
# + colab_type="code" id="58AXNVMKZj3T" outputId="a3044653-0b33-4228-a53d-45ed7ab4634a" executionInfo={"status": "ok", "timestamp": 1568305884339, "user_tz": 360, "elapsed": 316, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
population.head()
# + colab_type="code" id="0ywWDL2MZqlF" outputId="341d676f-1e9d-4494-bd18-be07ffbfb913" executionInfo={"status": "ok", "timestamp": 1568306369608, "user_tz": 360, "elapsed": 332, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 253}
pd.options.display.max_columns = 500
entities.head()
# + colab_type="code" id="mk_R0eFZZ0G5" outputId="c7ccb232-e6c2-4dfb-cd63-858a33286b1a" executionInfo={"status": "ok", "timestamp": 1568306373724, "user_tz": 360, "elapsed": 355, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 512}
concepts.head()
# + [markdown] colab_type="text" id="6HYUytvLT8Kf"
# ## Merge data
# + [markdown] colab_type="text" id="dhALZDsh9n9L"
# https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf
# + colab_type="code" id="A-tnI-hK6yDG" outputId="d189daba-d550-4bcb-d08a-50533493ded9" executionInfo={"status": "ok", "timestamp": 1568307200255, "user_tz": 360, "elapsed": 285, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
print(income.shape)
print(lifespan.shape)
# + id="aPqY3by9XR8L" colab_type="code" outputId="c3ef2677-b5df-4fee-a9df-aeae114d2248" executionInfo={"status": "ok", "timestamp": 1568307201026, "user_tz": 360, "elapsed": 627, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
merged = pd.merge(income, lifespan)
merged = pd.merge(income, lifespan, how='inner', on=['geo', 'time'])
merged.shape
# + id="f4um58T8X-Y6" colab_type="code" outputId="8a36aa3d-4e45-4057-d835-2d8995eeab0f" executionInfo={"status": "ok", "timestamp": 1568307753790, "user_tz": 360, "elapsed": 341, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 215}
pd.options.display.max_rows = 500
merged = pd.merge(income, lifespan, how='outer', on=['geo', 'time'])
print(merged.shape)
merged.head()
# + id="XmGA-QnbbVek" colab_type="code" outputId="54d3b9a6-2e65-4957-a854-c7d844251280" executionInfo={"status": "ok", "timestamp": 1568307765128, "user_tz": 360, "elapsed": 339, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
# how to check for duplicates using a specific subset of columns
merged.duplicated(subset=['geo', 'time']).value_counts()
# + id="Xr_tVTIwblId" colab_type="code" outputId="22e88384-6d19-4bba-bbbb-b81b1210f054" executionInfo={"status": "ok", "timestamp": 1568307846296, "user_tz": 360, "elapsed": 326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
# Count the number of unique values in a specific column
# if the final number does not match the num_rows of the column
# then you have duplicates
merged['geo'].nunique()
# + id="VgFH-7CKYsW2" colab_type="code" outputId="f120136d-4e19-47c9-ea77-fc8487643958" executionInfo={"status": "ok", "timestamp": 1568307202672, "user_tz": 360, "elapsed": 318, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 215}
merged = pd.merge(income, lifespan, how='left', on=['geo', 'time'])
print(merged.shape)
merged.head()
# + id="H30jJzO3ZGBo" colab_type="code" outputId="5303dda2-2db7-44f3-cc72-19539656db1c" executionInfo={"status": "ok", "timestamp": 1568307203984, "user_tz": 360, "elapsed": 317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 215}
merged = pd.merge(income, lifespan, how='right', on=['geo', 'time'])
print(merged.shape)
merged.head()
# + id="BdUP-TMkaNpj" colab_type="code" outputId="22af32be-4f2a-4bcf-8c3a-3d8a3c8e66ef" executionInfo={"status": "ok", "timestamp": 1568308919061, "user_tz": 360, "elapsed": 352, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 215}
merged = pd.merge(income, lifespan)
print(merged.shape)
merged.head()
# + id="-DZ0HXH_a5tE" colab_type="code" outputId="bc7444a2-b7e3-414e-a6c3-5d49791d0a68" executionInfo={"status": "ok", "timestamp": 1568307991485, "user_tz": 360, "elapsed": 294, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
merged.duplicated(subset=['geo', 'time']).value_counts()
# + id="2qX65__jcCFa" colab_type="code" outputId="830c66c6-568a-4f77-81a0-30bbbccb5d02" executionInfo={"status": "ok", "timestamp": 1568308923403, "user_tz": 360, "elapsed": 358, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 215}
df = pd.merge(merged, population)
print(df.shape)
df.head()
# + id="0iz91cRNe1Vv" colab_type="code" outputId="6209ae2b-3390-4c40-ebd6-54306612672e" executionInfo={"status": "ok", "timestamp": 1568308927088, "user_tz": 360, "elapsed": 359, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 235}
df = pd.merge(df,
entities[['country', 'name', 'world_4region', 'world_6region']],
left_on='geo', right_on='country')
print(df.shape)
df.head()
# + id="VSDMu3nQgAyf" colab_type="code" outputId="37d811a8-4d04-424a-efe2-24cd6369eaf5" executionInfo={"status": "ok", "timestamp": 1568309008661, "user_tz": 360, "elapsed": 309, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
df = df.rename(columns = {
'country': 'country_code',
'time': 'year',
'income_per_person_gdppercapita_ppp_inflation_adjusted': 'income',
'life_expectancy_years': 'lifespan',
'population_total': 'population',
'name': 'country',
'world_6region': '6region',
'world_4region': '4region'
})
df.head()
# + [markdown] colab_type="text" id="4OdEr5IFVdF5"
# ## Explore data
# + colab_type="code" id="4IzXea0T64x4" outputId="26bd85e3-cd02-4fea-fa00-19ac862d5804" executionInfo={"status": "ok", "timestamp": 1568309066827, "user_tz": 360, "elapsed": 306, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 190}
df.dtypes
# + id="V_VTEzJ5gYUm" colab_type="code" outputId="c5301de0-9ff1-4c5d-9124-fc52ff0f71da" executionInfo={"status": "ok", "timestamp": 1568309089290, "user_tz": 360, "elapsed": 321, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 288}
df.describe()
# + id="ZV9ooFazgjOG" colab_type="code" outputId="e16c2480-060f-446f-bbfb-3eaa04283d0c" executionInfo={"status": "ok", "timestamp": 1568309153908, "user_tz": 360, "elapsed": 290, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 168}
df.describe(exclude='number')
# + id="OfNDNciAg74g" colab_type="code" outputId="65be0d2a-358f-4b9f-d37a-fa0651dbddca" executionInfo={"status": "ok", "timestamp": 1568309245439, "user_tz": 360, "elapsed": 264, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
usa = df[df.country == 'United States']
usa.head()
# + id="36uVmuYFhBbv" colab_type="code" outputId="a588ba2e-166c-4f0d-beb1-239bd3d413a0" executionInfo={"status": "ok", "timestamp": 1568309258689, "user_tz": 360, "elapsed": 282, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 288}
usa.describe()
# + id="WTOz_KjBhQAq" colab_type="code" outputId="4878b765-4dc1-4cc3-b1dc-006dc0119750" executionInfo={"status": "ok", "timestamp": 1568309316318, "user_tz": 360, "elapsed": 293, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 138}
usa[usa.year.isin([1818,1918,2018])]
# + id="hBmh_cUjhbC-" colab_type="code" outputId="bf47cfc3-1ffe-46d5-dac1-b3fd35fce226" executionInfo={"status": "ok", "timestamp": 1568309378328, "user_tz": 360, "elapsed": 336, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 138}
china = df[df.country=='China']
china[china.year.isin([1818, 1918, 2018])]
# + [markdown] colab_type="text" id="hecscpimY6Oz"
# ## Plot visualization
# + colab_type="code" id="_o8RmX2M67ai" outputId="f5a3900b-ca35-48b7-f3fe-e326a6663695" executionInfo={"status": "ok", "timestamp": 1568310437780, "user_tz": 360, "elapsed": 334, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
import seaborn as sns
now = df[df.year == 2018]
then = df[df.year == 1918]
now.head()
# + id="61gnM3d9mReJ" colab_type="code" outputId="66732e2c-19dc-4899-ec9b-e0ab115b4e4b" executionInfo={"status": "ok", "timestamp": 1568310854765, "user_tz": 360, "elapsed": 2040, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 844}
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30, 400), alpha=0.7, data=then)
plt.xscale('log')
plt.title("The World in 1918")
plt.ylim(0,85)
plt.xlim(0,100000)
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30, 400), alpha=0.7, data=now)
plt.xscale('log')
plt.title("the World in 2018")
plt.ylim(0,85)
plt.xlim(0,100000)
# + id="imhI0uXSi_-w" colab_type="code" outputId="1ecc3e91-920e-4c41-aab5-d6c36ccd4ad1" executionInfo={"status": "ok", "timestamp": 1568310997364, "user_tz": 360, "elapsed": 1870, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# We *can* still use the figure/axes syntax from matplotlib with seaborn
# but it's much more common to use it when working with multiple subplots
# if we're working with just a single graph, then we typically use
# plt.blahblahblah (pyplot) syntax with seaborn
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16,8))
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30, 400), alpha=0.7, data=then, ax=ax[0]);
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30, 400), alpha=0.7, data=now, ax=ax[1]);
# + [markdown] colab_type="text" id="8OFxenCdhocj"
# ## Analyze outliers
# + colab_type="code" id="D59bn-7k6-Io" outputId="25d3f08c-3b55-47e7-b71c-66b06740e738" executionInfo={"status": "ok", "timestamp": 1568311058158, "user_tz": 360, "elapsed": 296, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Qatar is the richest country in 2018
now.sort_values('income', ascending=False)
# + id="o_1GBUqVoEnH" colab_type="code" outputId="41ec3b83-1e45-44b2-dac5-802637446094" executionInfo={"status": "ok", "timestamp": 1568311115684, "user_tz": 360, "elapsed": 288, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 78}
now_qatar = now[now.country=='Qatar']
now_qatar.head()
# + id="vA0g44feoKQU" colab_type="code" outputId="464709ab-def8-4d0d-846f-3d7e02ed0b8e" executionInfo={"status": "ok", "timestamp": 1568311377711, "user_tz": 360, "elapsed": 1126, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 386}
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30,400), data=now)
plt.xscale('log')
plt.ylim(0,90)
plt.title("Qatar is Really Rich")
plt.text(x=now_qatar.income-5000, y= now_qatar.lifespan+1, s='Qatar')
plt.show()
# + [markdown] colab_type="text" id="DNTMMBkVhrGk"
# ## Plot multiple years
# + colab_type="code" id="JkTUmYGF7BQt" outputId="88ff8af5-877b-4b2c-cdfa-cfd953bb3434" executionInfo={"status": "ok", "timestamp": 1568311801201, "user_tz": 360, "elapsed": 340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 198}
years = [1818, 1918, 2018]
centuries = df[df.year.isin(years)]
centuries.head()
# + id="gIzCi_WPp1eQ" colab_type="code" outputId="82917ed4-4e96-4f2a-805d-a539e4abfbf1" executionInfo={"status": "ok", "timestamp": 1568311966981, "user_tz": 360, "elapsed": 1676, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
fig = sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30,400), col='year', data=centuries)
axes = fig.axes.flatten()
axes[0].set_title('Poor and Sick in 1818')
axes[1].set_title('Healthier, but still poor in 1918')
axes[2].set_title('Healthier and Richer in 2018');
# + [markdown] colab_type="text" id="BB1Ki0v6hxCA"
# ## Point out a story
# + id="QlH4QDmPrcai" colab_type="code" colab={}
years = [1918, 1938, 1978, 1998, 2018]
# + id="2pXkpSAtrhTa" colab_type="code" outputId="4d598488-a4c5-417f-8a35-38bdd9a797f1" executionInfo={"status": "ok", "timestamp": 1568312294548, "user_tz": 360, "elapsed": 4670, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCz28Gz6CiIvYUZ2OONEmULLjSO02OJQDHnO_-Mrw=s64", "userId": "09124097936673074355"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
for year in years:
sns.relplot(x='income', y='lifespan', hue='6region', size='population',
sizes=(30,400), data=df[df.year==year])
plt.xscale('log')
plt.xlim((150, 150000))
plt.ylim((0, 90))
plt.title('Countries Above the Poverty Line in ' + str(year))
plt.axvline(x=1000, color='grey')
# + id="O54cCkAQr3n3" colab_type="code" colab={}
| module4-sequence-your-narrative/Drive_LS_DS8_124_Sequence_your_narrative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Planet Pieces
# ### A quick dive into what it takes to train and put together a neural network in two afternoons.
# ---
# ### Team members:
# *Nga (<NAME> - Louisiana State University
# <NAME> - Ceres Imaging
# <NAME> - University of Washington
# <NAME> - Utah State University
# <NAME> - Environmental Science Associates
# <NAME> - University of Washington
# <NAME> - University of Washington*
# ### Purpose:
#
# To explore the world of convolutional neural networks and train a model to classify land features in a high resolution context
# ### Data:
# 82 [PlanetScope Analytic Ortho Tile images](https://www.planet.com/products/satellite-imagery/planetscope-analytic-ortho-tile/)
# Collected in 2017 and 2019 between June and September
# - 2017-07-25
# - 2017-08-22
# - 2017-09-28
# - 2019-06-02
# - 2019-07-25
# - 2019-08-04
# - 2019-08-27
#
# Delivered pre-corrected for terrain effects (using Shuttle Radar Topography Mission, Intermap, and other local elevation datasets), sensor geometry, and projected
#
# **Spatial resolution**: 3m
# **Radiometric resolution**: 16 bit (top of atmosphere radiance radiometric range of 0-65536) 12 bit camera range (0 - 4096)
# **Spectral resolution**: Four bands - blue, green, red and near infrared
# **Spatial extent (per image)**: 25km x 25km (8000 pixel x 8000 pixel arrays)
# ### Packages used:
# numpy
# matplotlib
# rasterio
# tensorflow - keras
# `pandas`
# `xarray`
# ### What is a Convolutional Neural Network?
# ### What is image segmentation?
# ### General Approach
# *Establish git workflow*
#
# **Preliminary steps**
# - Download data (82 images)
# - Distribute data
#
# **Processing steps**
# 1. Convert data from top-of-atmosphere radiance to top-of-atmosphere reflectance (Michelle)
# 2. Create training dataset by labelling samples (Nga, Matt, and Shashank)
# 3. Build, modify, and train model on training dataset (Claire and Joshua)
# 4. Classify images (model + human help)
| contributors/jmhu/PlanetPieces_Intro_nopng.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
# import matplotlib.pyplot as plt
# import pandas as pd
# import numpy as np
# import requests
# import time
# from scipy.stats import linregress
# import citypy
# Workaround to fix Citipy issue. Performed a pip install citipy and still would not work with original code.
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from datetime import datetime
from scipy.stats import linregress
import scipy.stats as st
from pandas import DataFrame
from requests import get
# Import API key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file_cities = "output/cities.csv"
# Range of latitudes and longitudes
lat_range = (0, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
#len(cities)
#print(cities)
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
# Use url from class, ensure API Key is working, and convert temps to imperial (Fahrenheit)
url = 'http://api.openweathermap.org/data/2.5/weather?&units=imperial&' # To convert to Fahrenheit, I needed to have a workaround where I placed the & after imperial. Otherwise, the temps are not converted correctly
#url = 'http://api.openweathermap.org/data/2.5/weather?'
#units = 'imperial'
api_key = '28c4ccd34ec2c4e49331c9c55008fd8b' # My API key
# Test url to determine it works
#http://api.openweathermap.org/data/2.5/weather?&APPID=28c4ccd34ec2c4e49331c9c55008fd8b&units=imperial&q=chicago
# Create query_url
#query_url = url + "&appid=" + api_key + "&units=" + units + "&q=" + city
query_url = url + "appid=" + api_key + "&q=" + city
#print(query_url)
#type(query_url) # Check type
# +
# Set up a test of cities in list to iterate in the For Loop
#cities = ['London', 'notA_city', 'Chicago', 'Tokyo', 'Toronto', 'Orlando', 'Miami', 'Moscow', 'Hong Kong', 'Shanghai', 'Seoul', 'Paris', 'New York City']
# Initiate a list (columns) to hold reponses in a df
temp = []
humidity = []
cloudiness = []
wind_speed = []
lngs = []
lats = []
city_name = []
# +
# Use a For Loop to iterate through cities
for city in cities:
try:
query_url = url + "appid=" + api_key + "&q=" + city
weather_dict = get(query_url).json()
if 'coord' in weather_dict:
print('found', city)
else:
print(city, 'not found')
lats.append(weather_dict['coord']['lat']) # Append to list for each key/item found in weather_dict
lngs.append(weather_dict['coord']['lon'])
humidity.append(weather_dict['main']['humidity'])
cloudiness.append(weather_dict['clouds']['all'])
wind_speed.append(weather_dict['wind']['speed'])
temp.append(weather_dict['main']['temp'])
city_name.append(city)
except Exception as e:
print('Something broke...', e)
finally:
print('Finished trying to get', city)
city_name.to_csv("./output/cities.csv", index=False)
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# Set up a df by using dict. They are defined in the list above.
weather_df = DataFrame({
'City_Name': city_name,
'Temp': temp,
'Latitude': lats,
'Longitude': lngs,
'Humidity': humidity,
'Wind_Speed': wind_speed,
'Cloudiness': cloudiness,
})
weather_df
# type(weather_df) # Check it is a df of type
# Export the city data into a .csv file in the output folder
weather_df.to_csv("./output/weather_data.csv", index=False)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# Get the indices of cities that have humidity over 100%.
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
plt.scatter(weather_df['Latitude'], weather_df['Temp'])
plt.title(f"Latitude of City vs Current Temperature")
plt.grid(True)
plt.xlabel('Latitude')
plt.ylabel('Current Temperature')
plt.savefig("./output/lat_vs_temp.png", bbox_inches="tight")
plt.show()
# Analysis: There is a direct correlation of temperature relative to the equator (Latitude = 0).
# The closer the city is to the equator, the higher the temperature.
# -
# ## Latitude vs. Humidity Plot
# +
plt.scatter(weather_df['Latitude'], weather_df['Humidity'])
plt.title(f"Latitude of City vs Current Humidity")
plt.grid(True)
plt.xlabel('Latitude')
plt.ylabel('Current Humidity')
plt.savefig("./output/lat_vs_humidity.png", bbox_inches="tight")
plt.show()
# Analysis: In analyzing humidity relative to latitude, it is fairly distributed.
# That means high humidity can be found in cities near or far from the equator (Latitude = 0).
# -
# ## Latitude vs. Cloudiness Plot
# +
plt.scatter(weather_df['Latitude'], weather_df['Cloudiness'])
plt.title(f"Latitude of City vs Current Cloudiness")
plt.grid(True)
plt.xlabel('Latitude')
plt.ylabel('Current Cloudiness')
plt.savefig("./output/lat_vs_cloud.png", bbox_inches="tight")
plt.show()
# Analysis: In the analysis of cloudiness relative to latitude, it is distributed.
# That means cloudiness found in cities is not related to the distance of the cities to equator (Latitude = 0).
# -
# ## Latitude vs. Wind Speed Plot
# +
plt.scatter(weather_df['Humidity'], weather_df['Wind_Speed'])
plt.title(f"Latitude of City vs Current Wind Speed")
plt.grid(True)
plt.xlabel('Latitude')
plt.ylabel('Current Wind Speed')
plt.savefig("./output/lat_vs_wind.png", bbox_inches="tight")
plt.show()
# Analysis: In the analysis of cloudiness relative to latitude, there appears to be a correlation of higher wind speeds
# to the further away the city is from the equator (Latitude = 0).
# -
# ## Linear Regression
# Define the criterias for Northern and Southern Hemisphere. Use .loc method in a list. Create new dataframes.
northern_hemp_df = weather_df.loc[weather_df['Latitude'] >= 0]
southern_hemp_df = weather_df.loc[weather_df['Latitude'] < 0]
# +
# Create a scatter plot for latitude vs max temp (northern hemisphere)
#x_values = northern_hemp_df['Latitude']
#y_values = northern_hemp_df['Temp']
#plt.show()
# -
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
| WeatherPy/WeatherPY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
import sys
import os
sys.path.append(os.path.abspath("/media/sf_Project2/Code"))
from IO import Input
from IO import Output
import pandas as pd
class cd:
"""Context manager for changing the current working directory"""
def __init__(self, newPath):
self.newPath = os.path.expanduser(newPath)
def __enter__(self):
self.savedPath = os.getcwd()
os.chdir(self.newPath)
def __exit__(self, etype, value, traceback):
os.chdir(self.savedPath)
# -
with cd("/media/sf_Project2/Code"):
train_dataset = np.array(Input.load_trainset_caffefeatures(featureSelectionMethod='RF',Percentile = 100)).astype('float32')
train_labels = np.array(Input.load_trainset_labels()).astype('float32')
valid_dataset = np.array(Input.load_validationset_caffefeatures(featureSelectionMethod='RF',Percentile = 100)).astype('float32')
valid_labels = np.array(Input.load_validationset_labels()).astype('float32')
# +
num_labels=10
train_labels = np.squeeze((np.arange(num_labels) == train_labels[:,None]).astype(np.float32))
valid_labels = np.squeeze((np.arange(num_labels) == valid_labels[:,None]).astype(np.float32))
train_labels0 = train_labels[:,1]
train_labels0 = train_labels0.reshape((train_labels.shape[0],1))
print(train_labels0)
#print(train_dataset)
#train_labels = train_labels.reshape((train_labels.shape[0],1))
#valid_labels = valid_labels.reshape((valid_labels.shape[0],1))
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
feature_size = train_dataset.shape[1]
print(train_labels)
# -
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels,1))
/ predictions.shape[0])
# # Network:
# +
batch_size = 64
hlSize0 = 516
beta = 0.007
#decay_steps = 200
#decay_rate = 0.90
#learningStart=0.0007
decay_steps = 180
decay_rate = 0.96
learningStart=0.00012
stdv = 0.03
#patch_size = 5
#depth = 16
#num_hidden = 64
graph = tf.Graph()
with graph.as_default():
global_step = tf.Variable(0) # count the number of steps taken.
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, feature_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size,num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
#tf_test_dataset = tf.constant(test_dataset)
# Variables.
input_weights = tf.Variable(tf.truncated_normal(
[feature_size,hlSize0],
stddev=stdv))
input_biases = tf.Variable(tf.zeros([hlSize0]))
layer1_weights = tf.Variable(tf.truncated_normal(
[hlSize0,num_labels],
stddev=stdv))
layer1_biases = tf.Variable(tf.constant(0.0, shape=[num_labels]))
# Model.
def model(data):
layer1 = tf.nn.relu(tf.matmul(data, input_weights) + input_biases)
layer2 = tf.matmul(layer1, layer1_weights) + layer1_biases
return layer2
# Training computation.
logits = model(tf_train_dataset)
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
loss = loss + beta * tf.nn.l2_loss(input_weights) + \
beta * tf.nn.l2_loss(layer1_weights)
# beta * tf.nn.l2_loss(layer2_weights) + \
# beta * tf.nn.l2_loss(layer3_weights) + \
# beta * tf.nn.l2_loss(output_weights)
# Optimizer.
learning_rate = tf.train.exponential_decay(learningStart, global_step, decay_steps, decay_rate)
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss, global_step=global_step)
#optimizer = tf.train.GradientDescentOptimizer(0.00005).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset))
#test_prediction = tf.nn.softmax(model(tf_test_dataset))
# +
num_steps = 6001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size)]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 100 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
#print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
print("finished!")
input_weights_val = input_weights.eval()
input_biases_val = input_biases.eval()
layer1_weights_val = layer1_weights.eval()
layer1_biases_val = layer1_biases.eval()
valid_prediction_val = valid_prediction.eval()
# -
validData = pd.DataFrame(valid_prediction_val)
Output.to_outputfile(validData,1,'NNSTRUCTURE8valid')
with cd("/media/sf_Project2/Code"):
test_data = np.array(Input.load_testdata_caffefeatures(True,range(30000),'RF',100)).astype('float32')
with tf.Session() as session:
layer1 = tf.nn.relu(tf.matmul(test_data, input_weights_val) + input_biases_val)
layer2 = tf.matmul(layer1, layer1_weights_val) + layer1_biases_val
firstHalfTest = tf.nn.softmax(layer2).eval()
print(firstHalfTest)
with cd("/media/sf_Project2/Code"):
test_data = np.array(Input.load_testdata_caffefeatures(True,range(30000,60000),'RF',100)).astype('float32')
with tf.Session() as session:
layer1 = tf.nn.relu(tf.matmul(test_data, input_weights_val) + input_biases_val)
layer2 = tf.matmul(layer1, layer1_weights_val) + layer1_biases_val
secondHalfTest = tf.nn.softmax(layer2).eval()
print(secondHalfTest)
with cd("/media/sf_Project2/Code"):
test_data = np.array(Input.load_testdata_caffefeatures(True,range(60000,80000),'RF',100)).astype('float32')
with tf.Session() as session:
layer1 = tf.nn.relu(tf.matmul(test_data, input_weights_val) + input_biases_val)
layer2 = tf.matmul(layer1, layer1_weights_val) + layer1_biases_val
thirdHalfTest = tf.nn.softmax(layer2).eval()
print(thirdHalfTest)
testClass = np.concatenate([firstHalfTest,secondHalfTest, thirdHalfTest],0)
testClass = pd.DataFrame(testClass)
Output.to_outputfile(testClass,1,"NNSTRUCTURE8testset",validation=False)
| Project2/Code/Neural Networks/Untitled-1.79.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pickle
from itertools import chain
from collections import OrderedDict
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
import matplotlib.pylab as plt
from copy import deepcopy
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings(action="ignore", module="scipy", message="^internal gelsd")
import sys, os
sys.path.append(os.path.join(os.path.dirname("__file__"), '..', '..'))
from mela.util import plot_matrices, make_dir, get_struct_str, get_args, Early_Stopping, record_data, manifold_embedding
from mela.settings.filepath import variational_model_PATH, dataset_PATH
from mela.pytorch.net import Net
from mela.pytorch.util_pytorch import Loss_with_uncertainty
from mela.variational.util_variational import get_torch_tasks
from mela.variational.variational_meta_learning import Master_Model, Statistics_Net, Generative_Net, load_model_dict, get_regulated_statistics
from mela.variational.variational_meta_learning import VAE_Loss, sample_Gaussian, clone_net, get_nets, get_tasks, evaluate, get_reg, load_trained_models
from mela.variational.variational_meta_learning import plot_task_ensembles, plot_individual_tasks, plot_statistics_vs_z, plot_data_record, get_corrcoef
from mela.variational.variational_meta_learning import plot_few_shot_loss, plot_individual_tasks_bounce, plot_quick_learn_performance
from mela.variational.variational_meta_learning import get_latent_model_data, get_polynomial_class, get_Legendre_class, get_master_function
seed = 1
np.random.seed(seed)
torch.manual_seed(seed)
is_cuda = torch.cuda.is_available()
# -
# ## Training:
# +
task_id_list = [
# "latent-linear",
# "polynomial-3",
# "Legendre-3",
# "M-sawtooth",
# "M-sin",
# "M-Gaussian",
# "M-tanh",
# "M-softplus",
# "C-sin",
"C-tanh",
# "bounce-states",
# "bounce-images",
]
num_shots = 10
exp_id = "C-May8"
exp_mode = "meta"
input_size = 1
is_VAE = False
is_uncertainty_net = False
is_regulated_net = False
is_load_data = False
VAE_beta = 0.2
if task_id_list[0] == "C-sin":
statistics_output_neurons = 2
elif task_id_list[0] == "C-tanh":
statistics_output_neurons = 4
elif task_id_list[0] in ["bounce-states", "bounce-images"]:
statistics_output_neurons = 8
output_size = 1
lr = 5e-5
num_train_tasks = 50
num_test_tasks = 50
batch_size_task = min(50, num_train_tasks)
num_backwards = 1
num_iter = 10000
pre_pooling_neurons = 200
num_context_neurons = 0
statistics_pooling = "max"
main_hidden_neurons = (40, 40)
patience = 200
reg_amp = 1e-6
activation_gen = "leakyRelu"
activation_model = "leakyRelu"
optim_mode = "indi"
loss_core = "huber"
array_id = "new"
exp_id = get_args(exp_id, 1)
exp_mode = get_args(exp_mode, 2)
task_id_list = get_args(task_id_list, 3, type = "tuple")
statistics_output_neurons = get_args(statistics_output_neurons, 4, type = "int")
is_VAE = get_args(is_VAE, 5, type = "bool")
VAE_beta = get_args(VAE_beta, 6, type = "float")
lr = get_args(lr, 7, type = "float")
batch_size_task = get_args(batch_size_task, 8, type = "int")
pre_pooling_neurons = get_args(pre_pooling_neurons, 9, type = "int")
num_context_neurons = get_args(num_context_neurons, 10, type = "int")
statistics_pooling = get_args(statistics_pooling, 11)
main_hidden_neurons = get_args(main_hidden_neurons, 12, "tuple")
reg_amp = get_args(reg_amp, 13, type = "float")
activation_gen = get_args(activation_gen, 14)
activation_model = get_args(activation_model, 15)
optim_mode = get_args(optim_mode, 16)
is_uncertainty_net = get_args(is_uncertainty_net, 17, "bool")
loss_core = get_args(loss_core, 18)
array_id = get_args(array_id, 19)
try:
# %matplotlib inline
isplot = True
except:
isplot = False
# Settings:
reg_dict = {"statistics_Net": {"weight": reg_amp, "bias": reg_amp},
"generative_Net": {"weight": reg_amp, "bias": reg_amp, "W_gen": reg_amp, "b_gen": reg_amp}}
task_settings = {
"xlim": (-5, 5),
"num_examples": 20,
"test_size": 0.5,
}
struct_param_pre = [
[60, "Simple_Layer", {}],
# [60, "Simple_Layer", {}],
[60, "Simple_Layer", {}],
[pre_pooling_neurons, "Simple_Layer", {"activation": "linear"}],
]
struct_param_post = None
struct_param_gen_base = [
[60, "Simple_Layer", {}],
# [60, "Simple_Layer", {}],
[60, "Simple_Layer", {}],
]
isParallel = False
inspect_interval = 50
save_interval = 100
filename = variational_model_PATH + "/trained_models/{0}/Net_{1}_{2}_input_{3}_({4},{5})_stat_{6}_pre_{7}_pool_{8}_context_{9}_hid_{10}_batch_{11}_back_{12}_VAE_{13}_{14}_uncer_{15}_lr_{16}_reg_{17}_actgen_{18}_actmodel_{19}_struct_{20}_{21}_core_{22}_{23}_".format(
exp_id, exp_mode, task_id_list, input_size, num_train_tasks, num_test_tasks, statistics_output_neurons, pre_pooling_neurons, statistics_pooling, num_context_neurons, main_hidden_neurons, batch_size_task, num_backwards, is_VAE, VAE_beta, is_uncertainty_net, lr, reg_amp, activation_gen, activation_model, get_struct_str(struct_param_gen_base), optim_mode, loss_core, exp_id)
make_dir(filename)
print(filename)
# Obtain tasks:
assert len(task_id_list) == 1
dataset_filename = dataset_PATH + task_id_list[0] + "_{0}-shot.p".format(num_shots)
tasks = pickle.load(open(dataset_filename, "rb"))
tasks_train = get_torch_tasks(tasks["tasks_train"], task_id_list[0], is_cuda = is_cuda)
tasks_test = get_torch_tasks(tasks["tasks_test"], task_id_list[0], num_tasks = num_test_tasks, is_cuda = is_cuda)
# Obtain nets:
statistics_Net, generative_Net, generative_Net_logstd = get_nets(input_size = input_size, output_size = output_size, main_hidden_neurons = main_hidden_neurons,
pre_pooling_neurons = pre_pooling_neurons, statistics_output_neurons = statistics_output_neurons, num_context_neurons = num_context_neurons,
struct_param_pre = struct_param_pre,
struct_param_gen_base = struct_param_gen_base,
activation_statistics = activation_gen,
activation_generative = activation_gen,
activation_model = activation_model,
statistics_pooling = statistics_pooling,
isParallel = isParallel,
is_VAE = is_VAE,
is_uncertainty_net = is_uncertainty_net,
is_cuda = is_cuda,
)
if is_regulated_net:
struct_param_regulated_Net = [
[40, "Simple_Layer", {}],
[40, "Simple_Layer", {}],
[1, "Simple_Layer", {"activation": "linear"}],
]
generative_Net = Net(input_size = input_size, struct_param = struct_param_regulated_Net, settings = {"activation": activation_model})
master_model = Master_Model(statistics_Net, generative_Net, generative_Net_logstd, is_cuda = is_cuda)
# Setting up optimizer and loss functions:
if is_uncertainty_net:
optimizer = optim.Adam(chain.from_iterable([statistics_Net.parameters(), generative_Net.parameters(), generative_Net_logstd.parameters()]), lr = lr)
else:
optimizer = optim.Adam(chain.from_iterable([statistics_Net.parameters(), generative_Net.parameters()]), lr = lr)
if loss_core == "mse":
loss_fun_core = nn.MSELoss(size_average = True)
elif loss_core == "huber":
loss_fun_core = nn.SmoothL1Loss(size_average = True)
else:
raise
if is_VAE:
criterion = VAE_Loss(criterion = loss_fun_core, prior = "Gaussian", beta = VAE_beta)
else:
if is_uncertainty_net:
criterion = Loss_with_uncertainty(core = loss_core)
else:
criterion = loss_fun_core
early_stopping = Early_Stopping(patience = patience)
# Setting up recordings:
all_keys = list(tasks_train.keys()) + list(tasks_test.keys())
data_record = {"loss": {key: [] for key in all_keys}, "loss_sampled": {key: [] for key in all_keys}, "mse": {key: [] for key in all_keys},
"reg": {key: [] for key in all_keys}, "KLD": {key: [] for key in all_keys}}
info_dict = {"array_id": array_id}
info_dict["data_record"] = data_record
info_dict["model_dict"] = []
record_data(data_record, [exp_id, tasks_train, tasks_test, task_id_list, task_settings, reg_dict, is_uncertainty_net, lr, pre_pooling_neurons, num_backwards, batch_size_task,
struct_param_gen_base, struct_param_pre, struct_param_post, statistics_pooling, activation_gen, activation_model],
["exp_id", "tasks_train", "tasks_test", "task_id_list", "task_settings", "reg_dict", "is_uncertainty_net", "lr", "pre_pooling_neurons", "num_backwards", "batch_size_task",
"struct_param_gen_base", "struct_param_pre", "struct_param_post", "statistics_pooling", "activation_gen", "activation_model"])
# Training:
for i in range(num_iter + 1):
chosen_task_keys = np.random.choice(list(tasks_train.keys()), batch_size_task, replace = False).tolist()
if optim_mode == "indi":
if is_VAE:
KLD_total = Variable(torch.FloatTensor([0]), requires_grad = False)
if is_cuda:
KLD_total = KLD_total.cuda()
for task_key, task in tasks_train.items():
if task_key not in chosen_task_keys:
continue
((X_train, y_train), (X_test, y_test)), _ = task
for k in range(num_backwards):
optimizer.zero_grad()
if is_VAE:
statistics_mu, statistics_logvar = statistics_Net(torch.cat([X_train, y_train], 1))
statistics = sample_Gaussian(statistics_mu, statistics_logvar)
if is_regulated_net:
statistics = get_regulated_statistics(generative_Net, statistics)
y_pred = generative_Net(X_test, statistics)
loss, KLD = criterion(y_pred, y_test, mu = statistics_mu, logvar = statistics_logvar)
KLD_total = KLD_total + KLD
else:
if is_uncertainty_net:
statistics_mu, statistics_logvar = statistics_Net(torch.cat([X_train, y_train], 1))
y_pred = generative_Net(X_test, statistics_mu)
y_pred_logstd = generative_Net_logstd(X_test, statistics_logvar)
loss = criterion(y_pred, y_test, log_std = y_pred_logstd)
else:
statistics = statistics_Net(torch.cat([X_train, y_train], 1))
if is_regulated_net:
statistics = get_regulated_statistics(generative_Net, statistics)
y_pred = generative_Net(X_test, statistics)
loss = criterion(y_pred, y_test)
reg = get_reg(reg_dict, statistics_Net = statistics_Net, generative_Net = generative_Net, is_cuda = is_cuda)
loss = loss + reg
loss.backward(retain_graph = True)
optimizer.step()
# Perform gradient on the KL-divergence:
if is_VAE:
KLD_total = KLD_total / batch_size_task
optimizer.zero_grad()
KLD_total.backward()
optimizer.step()
record_data(data_record, [KLD_total], ["KLD_total"])
elif optim_mode == "sum":
optimizer.zero_grad()
loss_total = Variable(torch.FloatTensor([0]), requires_grad = False)
if is_cuda:
loss_total = loss_total.cuda()
for task_key, task in tasks_train.items():
if task_key not in chosen_task_keys:
continue
((X_train, y_train), (X_test, y_test)), _ = task
if is_VAE:
statistics_mu, statistics_logvar = statistics_Net(torch.cat([X_train, y_train], 1))
statistics = sample_Gaussian(statistics_mu, statistics_logvar)
y_pred = generative_Net(X_test, statistics)
loss, KLD = criterion(y_pred, y_test, mu = statistics_mu, logvar = statistics_logvar)
loss = loss + KLD
else:
if is_uncertainty_net:
statistics_mu, statistics_logvar = statistics_Net(torch.cat([X_train, y_train], 1))
y_pred = generative_Net(X_test, statistics_mu)
y_pred_logstd = generative_Net_logstd(X_test, statistics_logvar)
loss = criterion(y_pred, y_test, log_std = y_pred_logstd)
else:
statistics = statistics_Net(torch.cat([X_train, y_train], 1))
y_pred = generative_Net(X_test, statistics)
loss = criterion(y_pred, y_test)
reg = get_reg(reg_dict, statistics_Net = statistics_Net, generative_Net = generative_Net, is_cuda = is_cuda)
loss_total = loss_total + loss + reg
loss_total.backward()
optimizer.step()
else:
raise Exception("optim_mode {0} not recognized!".format(optim_mode))
loss_test_record = []
for task_key, task in tasks_test.items():
loss_test, _, _, _ = evaluate(task, statistics_Net, generative_Net, generative_Net_logstd = generative_Net_logstd, criterion = criterion, is_VAE = is_VAE, is_regulated_net = is_regulated_net)
loss_test_record.append(loss_test)
to_stop = early_stopping.monitor(np.mean(loss_test_record))
# Validation and visualization:
if i % inspect_interval == 0 or to_stop:
print("=" * 50)
print("training tasks:")
for task_key, task in tasks_train.items():
loss_test, loss_test_sampled, mse, KLD_test = evaluate(task, statistics_Net, generative_Net, generative_Net_logstd = generative_Net_logstd, criterion = criterion, is_VAE = is_VAE, is_regulated_net = is_regulated_net)
reg = get_reg(reg_dict, statistics_Net = statistics_Net, generative_Net = generative_Net, is_cuda = is_cuda).data[0]
data_record["loss"][task_key].append(loss_test)
data_record["loss_sampled"][task_key].append(loss_test_sampled)
data_record["mse"][task_key].append(mse)
data_record["reg"][task_key].append(reg)
data_record["KLD"][task_key].append(KLD_test)
print('{0}\ttrain\t{1} \tloss: {2:.5f}\tloss_sampled:{3:.5f} \tmse:{4:.5f}\tKLD:{5:.6f}\treg:{6:.6f}'.format(i, task_key, loss_test, loss_test_sampled, mse, KLD_test, reg))
for task_key, task in tasks_test.items():
loss_test, loss_test_sampled, mse, KLD_test = evaluate(task, statistics_Net, generative_Net, generative_Net_logstd = generative_Net_logstd, criterion = criterion, is_VAE = is_VAE, is_regulated_net = is_regulated_net)
reg = get_reg(reg_dict, statistics_Net = statistics_Net, generative_Net = generative_Net, is_cuda = is_cuda).data[0]
data_record["loss"][task_key].append(loss_test)
data_record["loss_sampled"][task_key].append(loss_test_sampled)
data_record["mse"][task_key].append(mse)
data_record["reg"][task_key].append(reg)
data_record["KLD"][task_key].append(KLD_test)
print('{0}\ttrain\t{1} \tloss: {2:.5f}\tloss_sampled:{3:.5f} \tmse:{4:.5f}\tKLD:{5:.6f}\treg:{6:.6f}'.format(i, task_key, loss_test, loss_test_sampled, mse, KLD_test, reg))
loss_train_list = [data_record["loss"][task_key][-1] for task_key in tasks_train]
loss_test_list = [data_record["loss"][task_key][-1] for task_key in tasks_test]
loss_train_sampled_list = [data_record["loss_sampled"][task_key][-1] for task_key in tasks_train]
loss_test_sampled_list = [data_record["loss_sampled"][task_key][-1] for task_key in tasks_test]
mse_train_list = [data_record["mse"][task_key][-1] for task_key in tasks_train]
mse_test_list = [data_record["mse"][task_key][-1] for task_key in tasks_test]
reg_train_list = [data_record["reg"][task_key][-1] for task_key in tasks_train]
reg_test_list = [data_record["reg"][task_key][-1] for task_key in tasks_test]
mse_few_shot = plot_few_shot_loss(master_model, tasks_test, isplot = isplot)
plot_quick_learn_performance(master_model, tasks_test)
record_data(data_record,
[np.mean(loss_train_list), np.median(loss_train_list), np.mean(reg_train_list), i,
np.mean(loss_test_list), np.median(loss_test_list), np.mean(reg_test_list),
np.mean(loss_train_sampled_list), np.median(loss_train_sampled_list),
np.mean(loss_test_sampled_list), np.median(loss_test_sampled_list),
np.mean(mse_train_list), np.median(mse_train_list),
np.mean(mse_test_list), np.median(mse_test_list),
mse_few_shot,
],
["loss_mean_train", "loss_median_train", "reg_mean_train", "iter",
"loss_mean_test", "loss_median_test", "reg_mean_test",
"loss_sampled_mean_train", "loss_sampled_median_train",
"loss_sampled_mean_test", "loss_sampled_median_test",
"mse_mean_train", "mse_median_train", "mse_mean_test", "mse_median_test",
"mse_few_shot",
])
if isplot:
plot_data_record(data_record, idx = -1, is_VAE = is_VAE)
print("Summary:")
print('\n{0}\ttrain\tloss_mean: {1:.5f}\tloss_median: {2:.5f}\tmse_mean: {3:.6f}\tmse_median: {4:.6f}\treg: {5:.6f}'.format(i, data_record["loss_mean_train"][-1], data_record["loss_median_train"][-1], data_record["mse_mean_train"][-1], data_record["mse_median_train"][-1], data_record["reg_mean_train"][-1]))
print('{0}\ttest\tloss_mean: {1:.5f}\tloss_median: {2:.5f}\tmse_mean: {3:.6f}\tmse_median: {4:.6f}\treg: {5:.6f}'.format(i, data_record["loss_mean_test"][-1], data_record["loss_median_test"][-1], data_record["mse_mean_test"][-1], data_record["mse_median_test"][-1], data_record["reg_mean_test"][-1]))
if is_VAE and "KLD_total" in locals():
print("KLD_total: {0:.5f}".format(KLD_total.data[0]))
if isplot:
plot_data_record(data_record, is_VAE = is_VAE)
# Plotting y_pred vs. y_target:
statistics_list_train, z_list_train = plot_task_ensembles(tasks_train, statistics_Net, generative_Net, is_VAE = is_VAE, is_regulated_net = is_regulated_net, title = "y_pred_train vs. y_train", isplot = isplot)
statistics_list_test, z_list_test = plot_task_ensembles(tasks_test, statistics_Net, generative_Net, is_VAE = is_VAE, is_regulated_net = is_regulated_net, title = "y_pred_test vs. y_test", isplot = isplot)
record_data(data_record, [np.array(z_list_train), np.array(z_list_test), np.array(statistics_list_train), np.array(statistics_list_test)],
["z_list_train_list", "z_list_test_list", "statistics_list_train_list", "statistics_list_test_list"])
if isplot:
print("train statistics vs. z:")
plot_statistics_vs_z(z_list_train, statistics_list_train)
print("test statistics vs. z:")
plot_statistics_vs_z(z_list_test, statistics_list_test)
# Plotting individual test data:
if "bounce" in task_id_list[0]:
plot_individual_tasks_bounce(tasks_test, num_examples_show = 40, num_tasks_show = 6, master_model = master_model, num_shots = 200)
else:
print("train tasks:")
plot_individual_tasks(tasks_train, statistics_Net, generative_Net, generative_Net_logstd = generative_Net_logstd, is_VAE = is_VAE, is_regulated_net = is_regulated_net, xlim = task_settings["xlim"])
print("test tasks:")
plot_individual_tasks(tasks_test, statistics_Net, generative_Net, generative_Net_logstd = generative_Net_logstd, is_VAE = is_VAE, is_regulated_net = is_regulated_net, xlim = task_settings["xlim"])
print("=" * 50 + "\n\n")
try:
sys.stdout.flush()
except:
pass
if i % save_interval == 0 or to_stop:
record_data(info_dict, [master_model.model_dict, i], ["model_dict", "iter"])
pickle.dump(info_dict, open(filename + "info.p", "wb"))
if to_stop:
print("The training loss stops decreasing for {0} steps. Early stopping at {1}.".format(patience, i))
break
# Plotting:
if isplot:
for task_key in tasks_train:
plt.semilogy(data_record["loss"][task_key], alpha = 0.6)
plt.show()
for task_key in tasks_test:
plt.semilogy(data_record["loss"][task_key], alpha = 0.6)
plt.show()
print("completed")
sys.stdout.flush()
# -
# ## Testing:
# +
lr = 1e-3
print(dataset_filename)
tasks = pickle.load(open(dataset_filename, "rb"))
tasks_test = get_torch_tasks(tasks["tasks_test"], task_id_list[0], is_cuda = is_cuda)
task_keys_all = list(tasks_test.keys())
mse_list_all = []
for i in range(int(len(tasks_test) / 100)):
print("{0}:".format(i))
task_keys_iter = task_keys_all[i * 100: (i + 1) * 100]
tasks_test_iter = {task_key: tasks_test[task_key] for task_key in task_keys_iter}
mse = plot_quick_learn_performance(master_model, tasks_test_iter, lr = lr, epochs = 20)['model_0'].mean(0)
mse_list_all.append(mse)
# +
plt.figure(figsize = (8,6))
mse_list_all = np.array(mse_list_all)
mse_mean = mse_list_all.mean(0)
mse_std = mse_list_all.std(0)
plt.fill_between(range(len(mse_mean)), mse_mean - mse_std * 1.96 / np.sqrt(int(len(tasks_test) / 100)), mse_mean + mse_std * 1.96 / np.sqrt(int(len(tasks_test) / 100)), alpha = 0.3)
plt.plot(range(len(mse_mean)), mse_mean)
plt.title("Tanh, 10-shot regression", fontsize = 20)
plt.xlabel("Number of gradient steps", fontsize = 18)
plt.ylabel("Mean Squared Error", fontsize = 18)
plt.show()
| variational/variational_meta_learning_regulated_exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
### rewards ###
import csv
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
import matplotlib.pyplot as plt
path_MLP = 'ToySyntheticRecordData-1510725920'
path_TINY = 'ToySyntheticRecordData-1510730017'
path_DEEP = 'ToySyntheticRecordData-1510734513'
iteration_MLP = np.arange(0,128000,500)
iteration_TINY = np.arange(0,155500,500)
iteration_DEEP = np.arange(0,88000,500)
t_MLP = np.linspace(0,60,len(iteration_MLP))
t_TINY = np.linspace(0,60,len(iteration_TINY))
t_DEEP = np.linspace(0,60,len(iteration_DEEP))
totals_MLP = []
for it_MLP in iteration_MLP:
data_MLP = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_MLP,it_MLP))
data_rewards_MLP = data_MLP['rewards']
rewards_MLP = np.zeros(len(data_rewards_MLP))
total_rewards_MLP = 0
for i1 in range(len(data_rewards_MLP)):
total_rewards_MLP += data_rewards_MLP[i1]
rewards_MLP[i1] = total_rewards_MLP
totals_MLP.append(total_rewards_MLP)
totals_TINY = []
for it_TINY in iteration_TINY:
data_TINY = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_TINY,it_TINY))
data_rewards_TINY = data_TINY['rewards']
rewards_TINY = np.zeros(len(data_rewards_TINY))
total_rewards_TINY = 0
for i2 in range(len(data_rewards_TINY)):
total_rewards_TINY += data_rewards_TINY[i2]
rewards_TINY[i2] = total_rewards_TINY
totals_TINY.append(total_rewards_TINY)
totals_DEEP = []
for it_DEEP in iteration_DEEP:
data_DEEP = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_DEEP,it_DEEP))
data_rewards_DEEP = data_DEEP['rewards']
rewards_DEEP = np.zeros(len(data_rewards_DEEP))
total_rewards_DEEP = 0
for i3 in range(len(data_rewards_DEEP)):
total_rewards_DEEP += data_rewards_DEEP[i3]
rewards_TINY[i3] = total_rewards_DEEP
totals_DEEP.append(total_rewards_DEEP)
## Normalization
totals_MLP = (totals_MLP - np.min(totals_MLP))/(np.max(totals_MLP) - np.min(totals_MLP))
totals_TINY = (totals_TINY - np.min(totals_TINY))/(np.max(totals_TINY) - np.min(totals_TINY))
totals_DEEP = (totals_DEEP - np.min(totals_DEEP))/(np.max(totals_DEEP) - np.min(totals_DEEP))
fig2, ax2 = plt.subplots()
ax2.plot(t_MLP,totals_MLP,label='MLP NN')
ax2.plot(t_TINY,totals_TINY,label='TINY NN')
ax2.plot(t_DEEP,totals_DEEP,label='DEEP NN')
ax2.set_title('3 Different Networks Training Results')
ax2.set_xlabel('Time(m)')
ax2.set_ylabel('Normalized Rewards')
ax2.legend()
ax2.set_xlim([-1, 61])
ax2.grid()
fig2.savefig('data/NN_Type/NN_TYPE_300_normalized.png')
print('Done')
# +
### original rewards ###
import csv
import pandas as pd
import numpy as np
from sklearn.utils import shuffle
import matplotlib.pyplot as plt
path_MLP = 'ToySyntheticRecordData-1510725920'
path_TINY = 'ToySyntheticRecordData-1510730017'
path_DEEP = 'ToySyntheticRecordData-1510734513'
iteration_MLP = np.arange(0,128000,500)
iteration_TINY = np.arange(0,155500,500)
iteration_DEEP = np.arange(0,88000,500)
t_MLP = np.linspace(0,60,len(iteration_MLP))
t_TINY = np.linspace(0,60,len(iteration_TINY))
t_DEEP = np.linspace(0,60,len(iteration_DEEP))
totals_MLP = []
for it_MLP in iteration_MLP:
data_MLP = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_MLP,it_MLP))
data_rewards_MLP = data_MLP['original_rewards']
rewards_MLP = np.zeros(len(data_rewards_MLP))
total_rewards_MLP = 0
for i1 in range(len(data_rewards_MLP)):
total_rewards_MLP += data_rewards_MLP[i1]
rewards_MLP[i1] = total_rewards_MLP
totals_MLP.append(total_rewards_MLP)
totals_TINY = []
for it_TINY in iteration_TINY:
data_TINY = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_TINY,it_TINY))
data_rewards_TINY = data_TINY['original_rewards']
rewards_TINY = np.zeros(len(data_rewards_TINY))
total_rewards_TINY = 0
for i2 in range(len(data_rewards_TINY)):
total_rewards_TINY += data_rewards_TINY[i2]
rewards_TINY[i2] = total_rewards_TINY
totals_TINY.append(total_rewards_TINY)
totals_DEEP = []
for it_DEEP in iteration_DEEP:
data_DEEP = pd.read_csv('data/NN_Type/{:s}/run_{:d}_0.csv'.format(path_DEEP,it_DEEP))
data_rewards_DEEP = data_DEEP['original_rewards']
rewards_DEEP = np.zeros(len(data_rewards_DEEP))
total_rewards_DEEP = 0
for i3 in range(len(data_rewards_DEEP)):
total_rewards_DEEP += data_rewards_DEEP[i3]
rewards_TINY[i3] = total_rewards_DEEP
totals_DEEP.append(total_rewards_DEEP)
## Normalization
totals_MLP = (totals_MLP - np.min(totals_MLP))/(np.max(totals_MLP) - np.min(totals_MLP))
totals_TINY = (totals_TINY - np.min(totals_TINY))/(np.max(totals_TINY) - np.min(totals_TINY))
totals_DEEP = (totals_DEEP - np.min(totals_DEEP))/(np.max(totals_DEEP) - np.min(totals_DEEP))
fig2, ax2 = plt.subplots()
ax2.plot(t_MLP,totals_MLP,label='MLP NN')
ax2.plot(t_TINY,totals_TINY,label='TINY NN')
ax2.plot(t_DEEP,totals_DEEP,label='DEEP NN')
ax2.set_title('3 Different Networks Training Results')
ax2.set_xlabel('Time(m)')
ax2.set_ylabel('Rewards')
ax2.legend()
ax2.set_xlim([-1, 61])
ax2.grid()
fig2.savefig('data/NN_Type/NN_TYPE_300_normalized_original.png')
print('Done')
| Report/Data/Plots/Codes/NN_300_labels_total.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GeoMMpax/Ethics/blob/master/V6_R_E_sig_ANN_FairLearnFrenchData.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YqAzy49D0Qrt"
# # MetaData
# + [markdown] id="GtM7Qas00WX7"
# Author <NAME> <EMAIL>
#
# Coauthor <NAME> <EMAIL>
#
# This is built for Google's cloud-based Colab. A libraries are !pip installed for each run.
#
# Data stored in Google Drive.
#
# Copy of V2 on 02Nov2021
#
# Copy of V3 on 16NOV2021
#
# Changes include scaling data and getting dummies for categorical variables before splitting into Train/Test
#
# Copy of V4 on 29NOV2021
#
# Changes include sigmoid in final layer for all models and updating the
# experiment so the activiation function was changed in the hidden layer.
#
# Copy of V5
#
# Added random seed control. Reduced activations fucntions from all 9 to 5.
# Softmax removed because it is for multi-class.
# Sigmoid removed because it is for binary (already in final layer)
#
# V6
# Stable. Experimental activation function "E" in both hidden layers.
# This copy made to test different dataset.
# + [markdown] id="3Qzt5H2PTHNR"
# ## Google Drive File
# + id="rvOh3OwyWelW" colab={"base_uri": "https://localhost:8080/"} outputId="4abad3fb-1525-4d96-9e28-cba3c8556f4d"
#https://pypi.org/project/PyDrive/
import time
start = time.time()
# !pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
end = time.time()
print("The time of execution of above program is :", end-start)
# + id="siFOuBXWWh7v" colab={"base_uri": "https://localhost:8080/"} outputId="e227d3e3-289f-4539-ba35-ef6df463a738"
start = time.time() #to meausure how long it takes
#authenticate users to have acces to google Drive. Click the link to get the code.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
end = time.time()
print("The time of execution of above program is :", end-start)
# + [markdown] id="C5y6FdYsS-xa"
# ## Library Loading
# + id="0JMF6jYsVwsR" colab={"base_uri": "https://localhost:8080/"} outputId="28f75a38-dd17-4d71-835d-9d28a5408dcd"
import pandas as pd
from matplotlib import pyplot as plt
import numpy as np
#It is a function that renders the figure in a notebook (instead of displaying a dump of the figure object).
# %matplotlib inline
# To remove the scientific notation from numpy arrays
#np.set_printoptions(suppress=True)
#Setting Seed for reproducable results (important to have tensorflow random seed set as well)
#https://datascience.stackexchange.com/questions/13314/causes-of-inconsistent-results-with-neural-network
np.random.seed(1)
# package for descriptive statistics
# !pip install researchpy
import researchpy as rp
# + id="8TEqf6-GWtAe"
#Get the file
#data from Kaggle: https://www.kaggle.com/blastchar/telco-customer-churn Accessed 2021Sep20
##data wrangled outside this Python notebook.
#Records with null for total charges removed
#"no internet service" and "no phone service" changed to "no"
# yes = 1 and no = 0
#gender female = 1 and male = 0
#customer ID removed
#recordID added
#stratified Sample 70/30 for based on "Gender" variable
#For Ver4 scaling data and getting dummies (i.e., one hot encoding) for categorical variables before splitting into Train/Test
# Stratied Sample 1 Telco Data
"""
downloaded = drive.CreateFile({'id':'1se_OSf2TmEBerxcDUU5mwjS-55iN-csF'}) # replace the id with id of file you want to access
downloaded.GetContentFile('WA_Fn-UseC_-Telco-Customer-Churn_WrangleTrain2.csv')
downloaded = drive.CreateFile({'id':'1wpB8kKSfqCEZXq0LUUlRVJFZ64vr6i0l'}) # replace the id with id of file you want to access
downloaded.GetContentFile('WA_Fn-UseC_-Telco-Customer-Churn_WrangleTEST2.csv')
"""
#Stratefied random split datafiles have "2" suffix Telco Data
# second stratefied random sample for this dataset, has "dec" suffix
"""
downloaded = drive.CreateFile({'id':'19hS4zGimbeFm2a1qMxyaDCuTLd1a0sLf'}) # replace the id with id of file you want to access
downloaded.GetContentFile('WA_Fn-UseC_-Telco-Customer-Churn_WrangleTrainDEC.csv')
downloaded = drive.CreateFile({'id':'1CHhJIFG0vYPl5as6sXfCUT3Jeibw0dJa'}) # replace the id with id of file you want to access
downloaded.GetContentFile('WA_Fn-UseC_-Telco-Customer-Churn_WrangleTESTdec.csv')
"""
# https://data.world/jfreex/e-commerce-users-of-a-french-c2c-fashion-store
# Target Variable made for transactions (items bought or sold)
# outliers removed at 3 standard deviations above the mean
#
#French Data Stratified Random Sample Jan 05, 2022 Random SEED == 47
"""
downloaded = drive.CreateFile({'id':'1BRlzi6jETpw66sDGB5Dhf53CkLp1zPtu'}) # replace the id with id of file you want to access
downloaded.GetContentFile('FrenchC2Train1.csv')
downloaded = drive.CreateFile({'id':'1P-Z4qCWjuOyEOmoXRhZI6TpTnzNaPukL'}) # replace the id with id of file you want to access
downloaded.GetContentFile('FrenchC2CTEST1.csv')
"""
#French Data Stratified Random Sample Jan 10, 2022 Random SEED == 67
downloaded = drive.CreateFile({'id':'1mOjfbmr7Vk3YqwpdbKlvfnXfMp1A8gJb'}) # replace the id with id of file you want to access
downloaded.GetContentFile('FrenchC2Train2.csv')
downloaded = drive.CreateFile({'id':'1jVMKvdJVswMJj9S9wqSdWdoYw9IF_-XH'}) # replace the id with id of file you want to access
downloaded.GetContentFile('FrenchC2CTEST2.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="f5mbIzCpXRg0" outputId="8769e99c-ba66-4e4e-f3fd-c96998a3adf8"
## read the training file
#dfTrain = pd.read_csv("WA_Fn-UseC_-Telco-Customer-Churn_WrangleTrain2.csv") # first stratified training dataset
#dfTrain = pd.read_csv("WA_Fn-UseC_-Telco-Customer-Churn_WrangleTrainDEC.csv") # second stratified training dataset
dfTrain = pd.read_csv('FrenchC2Train2.csv')
print("Training Data: ",dfTrain.shape)
## read the TEST file
# dfTEST = pd.read_csv("WA_Fn-UseC_-Telco-Customer-Churn_WrangleTEST2.csv") # first stratified testing dataset
#dfTEST = pd.read_csv("WA_Fn-UseC_-Telco-Customer-Churn_WrangleTESTdec.csv") # second stratified testing dataset
dfTEST = pd.read_csv("FrenchC2CTEST2.csv")
print("Testing Data: ",dfTEST.shape)
## show a sample of n records
#dfTrain.sample(10)
#dfTEST.sample(10)
del dfTrain['RecordID'] #"RecordID" included in dataset but removed because not variable in the analysis
del dfTEST['RecordID'] #"RecordID" included in dataset but removed because not variable in the analysis
## show the top 5 for Train
print("\n Training Data \n")
print(dfTrain.head())
#Displace the varaible types for Train
print(dfTrain.dtypes)
print(dfTrain.shape)
## show the top 5 for Testing
print("\n Testing Data \n")
print(dfTEST.head())
#Displace the varaible types for Testing
print(dfTEST.dtypes)
print(dfTEST.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="AO96LUMjbHNc" outputId="8eb64aac-fae3-4f6c-d9bd-eaba4450262a"
#install fairlearn
#if installing on a local IDE (e.g., Jupyter), do not use the exclamation point
# !pip install fairlearn
#install modeling packages
# !pip install tensorflow
# !pip install keras
# + [markdown] id="9xQVEQVbTTSV"
# ## Data File Wrangle
# + id="5Na3Wv079C3h"
# Separate Target Variable and Predictor Variables
# define variable to class: Churn
#"transact" is a dictomous variable used to identify customers who stop using the company's services
TargetVariable=["transact"]
#it is VERY important that the list of variables matches the left to right order in the CSVs!
PredictorVariables=["daysSinceLastLogin","productsListed","productsPassRate", "productsWished", "seniorityAsMonths",
"socialNbFollowers","socialNbFollows","socialProductsLiked","language_de", "language_en",
"language_es", "language_fr","language_it", "hasAnyApp","hasAndroidApp","hasIosApp", "hasProfilePicture","Gender_Byte"]
#NumericalPredictorVariables=["tenure","MonthlyCharges","TotalCharges"]
# + [markdown] id="XfV9759On6A5"
# ## Descriptive Statistics
# + colab={"base_uri": "https://localhost:8080/", "height": 668} id="KspXUV998uWV" outputId="bb3174fc-0b76-40bc-d273-ff77ae93982e"
print("dfTrain Predictor Variables")
rp.summary_cont(dfTrain[PredictorVariables])
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="A93rGcaxqGnI" outputId="dc96b79d-0200-4fca-9759-fb58db2f4f50"
print("dfTrain Target Variable")
rp.summary_cont(dfTrain[TargetVariable])
# + colab={"base_uri": "https://localhost:8080/", "height": 668} id="LIs_80o78gSX" outputId="e28e11f2-72db-4bf3-fab3-c33a0f71b87f"
print("dfTEST Predictor Variables")
rp.summary_cont(dfTEST[PredictorVariables])
# + colab={"base_uri": "https://localhost:8080/", "height": 135} id="xxk8AFwMpV-9" outputId="cce4f19d-9b69-462a-f8a2-84fef3772a98"
print("dfTEST Target Variable")
rp.summary_cont(dfTEST[TargetVariable])
# + colab={"base_uri": "https://localhost:8080/"} id="Qo68CbqeEkAr" outputId="5b4c6480-61fd-490c-d019-f504b1550d36"
"""Normally, random samples using 'from sklearn.model_selection import train_test_split'
identify training and testing data but, for repeatablility,
a static random stratified sample was made."""
#Training Data for X and y
X_train = dfTrain.drop('transact',axis='columns')
print("X_Train Shape:\n",X_train.shape, "\n")
print("X_train head:\n")
print(X_train.head())
y_train = dfTrain['transact']
print("y_train shape: \n",y_train.shape, "\n")
print("y_train head:\n")
print(y_train.head())
# Testing Data for X and y
X_test = dfTEST.drop('transact',axis='columns')
print("\nX_test shape: \n",X_test.shape,"\n")
print("X_test head:\n")
print(X_test.head())
y_test = dfTEST['transact']
print("y_test Shape:",y_test.shape,"\n")
print("y_test Head:\n")
print(y_test.head())
# Check shapes of training and testing datasets
assert(X_train.shape==(68976, 18))
assert(X_test.shape == (29562, 18))
print(X_train.shape)
print(X_test.shape)
#training data are rank one arrays that need to be reshaped
y_train=y_train.values.reshape((68976,1))
y_test=y_test.values.reshape((29562,1))
print(y_train.shape)
print(y_test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 325} id="us6CL1lTz09M" outputId="4812e7f4-508f-4336-dc53-355e21498f41"
print("\n X_test head: \n")
X_test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 325} id="5saHxAe27CsD" outputId="e40fa513-63e6-474a-964f-e2587596dd10"
print("\n X_train head: \n")
X_train.head()
# + [markdown] id="ezvTm7-YoDoM"
# # Activation Function Bias Analysis with FairLearn
# The Prupose of this code is to assess the technical bias resulting from the
# activation function used in the **hidden** layer of a Neural Network
# + [markdown] id="RY5OvhNtBPYi"
# # Modeling
# + id="l48i--sEBMXr"
#Modeling
import tensorflow as tf
#
# https://www.tensorflow.org/api_docs/python/tf/random/set_seed
# Set the random seed for repeatable results (important to have numpy random seed set as well)
tf.random.set_seed(1)
from tensorflow import keras
from keras.wrappers.scikit_learn import KerasClassifier
from keras.models import Sequential
from keras.layers import Dense
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import accuracy_score,precision_score,recall_score
from sklearn.model_selection import cross_val_score
from fairlearn.metrics import MetricFrame
from fairlearn.metrics import selection_rate,false_positive_rate,true_positive_rate,count
#Identifying the initializer used in each layer; seed set to 4 to a Python integer.
# "An initializer created with a given seed will always produce the same random tensor for a given shape and dtype."
# https://keras.io/api/layers/initializers/#randomnormal-class
initializer = tf.keras.initializers.RandomNormal(mean=0., stddev=1., seed=4)
# + [markdown] id="LNLh9cAiRx29"
# ## Relu Activation Function
# rectified linear unit activation = ReLU
#
# https://keras.io/api/layers/activations/#relu-function
# + id="AZVKwof-MZwN" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="62b25385-f471-4632-8c88-653afb21e15b"
# create ANN model https://thinkingneuron.com/how-to-use-artificial-neural-networks-for-classification-in-python/
### relu Activation function in all layers and sigmoid in output layer ###
model_relu = Sequential()
# Defining the Input layer and FIRST hidden layer, both are same!
model_relu.add(Dense(19, input_shape=(18,), kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# Defining the Second layer of the model
# after the first layer we don't have to specify input_dim as keras configure it automatically
model_relu.add(Dense(units=5, kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# The output neuron is a single fully connected node
#
model_relu.add(Dense(1, kernel_initializer=initializer,activation='sigmoid'))
# Compiling the model
model_relu.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fitting the ANN to the Training set
model_relu.fit(X_train, y_train, batch_size = 256, epochs = 30, verbose=1)
# Generating Predictions on testing data
Predictions_relu=model_relu.predict(X_test)
TestingData=pd.DataFrame(data=X_test.values, columns=PredictorVariables)
#TestingData.insert(len(df.columns), 'Churn', y_test.values)
#TestingData.assign(Churn=y_test)
TestingData["transact"]=y_test
TestingData['PredictedtransactRelu']=Predictions_relu
#transform Predicted transact from decimal to 0 or 1 for Confusion Matrix & insert into Fairlearn
TestingData["PredictedtransactReluClass"]= np.where(TestingData['PredictedtransactRelu']>=0.5, 1, 0)
#print(y_test[:10])
print(y_test.dtype)
print(TestingData.sample(11))
model_relu.evaluate(X_test, y_test)
#Fairlearn https://fairlearn.org/v0.7.0/quickstart.html
gm_relu = MetricFrame(metrics=accuracy_score, y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactReluClass'], sensitive_features=TestingData['Gender_Byte'])
print("\n")
print("Model score: ")
print(gm_relu.overall)
print("\n" "Model score by category: ")
print(gm_relu.by_group)
print(' Remember, from data wrangling, IF [Gender_Byte]="female" THEN "1" ELSE "0" ENDIF" \n')
#FairLearn Metrics
sr_relu = MetricFrame(metrics=selection_rate,y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactReluClass'], sensitive_features=TestingData['Gender_Byte'])
sr_relu.overall
print("1.0==female, 0.0==male")
sr_relu.by_group
#transform Sensative Varible from 0 or 1 to "male" or "female" for plotting
TestingData["GenderClass"]= np.where(TestingData['Gender_Byte']==1, "female","male")
#print(TestingData.head)
metrics = {
'accuracy': accuracy_score,
'precision': precision_score,
'recall': recall_score,
'false positive rate': false_positive_rate,
'true positive rate': true_positive_rate,
'selection rate': selection_rate,
'count': count}
metric_frame_relu = MetricFrame(metrics=metrics,
y_true=TestingData['transact'],
y_pred=TestingData['PredictedtransactReluClass'],
sensitive_features=TestingData["GenderClass"])
metric_frame_relu.by_group.plot.bar(
subplots=True,
layout=[3, 3],
legend=False,
figsize=[12, 8],
title="Show all relu metrics",
)
# + colab={"base_uri": "https://localhost:8080/", "height": 325} id="uwpCTh8VBdfh" outputId="e822bc30-7fa0-4086-fd12-6d0224748060"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="IZLxrL8LR5SZ"
# ## Tanh Activation Function
#
# Hyperbolic tangent activation function.
#
# https://keras.io/api/layers/activations/#tanh-function
# + id="ybATBIihsDgT" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="906b71c4-09b4-49bc-b790-0fbd6962a9eb"
# create ANN model https://thinkingneuron.com/how-to-use-artificial-neural-networks-for-classification-in-python/
### tanh Activation function in all layers and sigmoid in output layer ###
model_tanh = Sequential()
# Defining the Input layer and FIRST hidden layer, both are same!
model_tanh.add(Dense(19, input_shape=(18,), kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# Defining the Second layer of the model
# after the first layer we don't have to specify input_dim as keras configure it automatically
model_tanh.add(Dense(units=5, kernel_initializer=initializer, bias_initializer="zeros",activation='tanh'))
# The output neuron is a single fully connected node
#
model_tanh.add(Dense(1, kernel_initializer=initializer,activation='sigmoid'))
# Compiling the model
model_tanh.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fitting the ANN to the Training set
model_tanh.fit(X_train, y_train ,batch_size = 256, epochs = 30, verbose=1)
# Generating Predictions on testing data
Predictions_tanh=model_tanh.predict(X_test)
#TestingData=pd.DataFrame(data=X_test.values, columns=PredictorVariables)
#TestingData['transact']=y_test.values
TestingData['PredictedtransactTanh']=Predictions_tanh
#transform Predicted transact from decimal to 0 or 1 for Confusion Matrix & insert into Fairlearn
TestingData["PredictedtransactTanhClass"]= np.where(TestingData['PredictedtransactTanh']>=0.5, 1, 0)
print(y_test.dtype)
print(TestingData.sample(11))
model_tanh.evaluate(X_test, y_test)
#Fairlearn https://fairlearn.org/v0.7.0/quickstart.html
gm_tanh = MetricFrame(metrics=accuracy_score, y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactTanhClass'], sensitive_features=TestingData['Gender_Byte'])
print("\n")
print("Model score: ")
print(gm_tanh.overall)
print("\n" "Model score by category: ")
print(gm_tanh.by_group)
print(' Remember, from data wrangling, IF [Gender_Byte]="female" THEN "1" ELSE "0" ENDIF" \n')
#FairLearn Metrics
sr_tanh = MetricFrame(metrics=selection_rate,y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactTanhClass'], sensitive_features=TestingData['Gender_Byte'])
sr_tanh.overall
print("1.0==female, 0.0==male")
sr_tanh.by_group
#transform Sensative Varible from 0 or 1 to "male" or "female" for plotting
#TestingData["GenderClass"]= np.where(TestingData['gender']>=1, "female","male")
#print(TestingData.head)
metrics = {
'accuracy': accuracy_score,
'precision': precision_score,
'recall': recall_score,
'false positive rate': false_positive_rate,
'true positive rate': true_positive_rate,
'selection rate': selection_rate,
'count': count}
metric_frame_tanh = MetricFrame(metrics=metrics,
y_true=TestingData['transact'],
y_pred=TestingData['PredictedtransactTanhClass'],
sensitive_features=TestingData["GenderClass"])
metric_frame_tanh.by_group.plot.bar(
subplots=True,
layout=[3, 3],
legend=False,
figsize=[12, 8],
title="Show all tanh metrics",
)
# + id="sWYRSgbVFpjz" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="2ffc078f-92f9-4762-9f3e-f5f41a114578"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="i1RyYbiZSdQ4"
# ## Selu Activation Function
#
# Scaled Exponential Linear Unit (SELU).
#
# https://keras.io/api/layers/activations/#selu-function
# + id="cWfUGDVxsOmQ" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="de4418a2-f017-4807-9f6e-ac0e5915c416"
# create ANN model https://thinkingneuron.com/how-to-use-artificial-neural-networks-for-classification-in-python/
### Selu Activation function in all layers and sigmoid in output layer ###
model_selu = Sequential()
# Defining the Input layer and FIRST hidden layer, both are same!
model_selu.add(Dense(19, input_shape=(18,), kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# Defining the Second layer of the model
# after the first layer we don't have to specify input_dim as keras configure it automatically
model_selu.add(Dense(units=5, kernel_initializer=initializer, bias_initializer="zeros", activation='selu'))
# The output neuron is a single fully connected node
#
model_selu.add(Dense(1, kernel_initializer=initializer,activation='sigmoid'))
# Compiling the model
model_selu.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fitting the ANN to the Training set
model_selu.fit(X_train, y_train ,batch_size = 256, epochs = 30, verbose=1)
# Generating Predictions on testing data
Predictions_selu=model_selu.predict(X_test)
#TestingData=pd.DataFrame(data=X_test.values, columns=PredictorVariables)
#TestingData['transact']=y_test.values
TestingData['PredictedtransactSelu']=Predictions_selu
#transform Predicted transact from decimal to 0 or 1 for Confusion Matrix & insert into Fairlearn
TestingData["PredictedtransactSeluClass"]= np.where(TestingData['PredictedtransactSelu']>=0.5, 1, 0)
print(y_test.dtype)
print(TestingData.sample(11))
model_selu.evaluate(X_test, y_test)
#Fairlearn https://fairlearn.org/v0.7.0/quickstart.html
gm_selu = MetricFrame(metrics=accuracy_score, y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactSeluClass'], sensitive_features=TestingData['Gender_Byte'])
print("\n")
print("Model score: ")
print(gm_selu.overall)
print("\n" "Model score by category: ")
print(gm_selu.by_group)
print(' Remember, from data wrangling, IF [Gender_Byte]="female" THEN "1" ELSE "0" ENDIF" \n')
#FairLearn Metrics
sr_selu = MetricFrame(metrics=selection_rate,y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactSeluClass'], sensitive_features=TestingData['Gender_Byte'])
sr_selu.overall
print("1.0==female, 0.0==male")
sr_selu.by_group
#transform Sensative Varible from 0 or 1 to "male" or "female" for plotting
#TestingData["GenderClass"]= np.where(TestingData['Gender_Byte']>=1, "female","male")
#print(TestingData.head)
metrics = {
'accuracy': accuracy_score,
'precision': precision_score,
'recall': recall_score,
'false positive rate': false_positive_rate,
'true positive rate': true_positive_rate,
'selection rate': selection_rate,
'count': count}
metric_frame_selu = MetricFrame(metrics=metrics,
y_true=TestingData['transact'],
y_pred=TestingData['PredictedtransactSeluClass'],
sensitive_features=TestingData["GenderClass"])
metric_frame_selu.by_group.plot.bar(
subplots=True,
layout=[3, 3],
legend=False,
figsize=[12, 8],
title="Show all selu metrics",
)
# + id="xL5Z3qGhFt7z" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="c7e56a07-8aaf-4af6-a53c-26059634106a"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="X7tekLGXSjQn"
# ## Elu Activation Function
#
# Exponential Linear Unit.
#
# https://keras.io/api/layers/activations/#elu-function
# + id="4NH51L12sbFP" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="df6662ec-bc27-42a3-c37e-c4afc69e6854"
# create ANN model https://thinkingneuron.com/how-to-use-artificial-neural-networks-for-classification-in-python/
### elu Activation function in all layers and sigmoid in output layer ###
model_elu = Sequential()
# Defining the Input layer and FIRST hidden layer, both are same!
model_elu.add(Dense(19, input_shape=(18,), kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# Defining the Second layer of the model
# after the first layer we don't have to specify input_dim as keras configure it automatically
model_elu.add(Dense(units=5, kernel_initializer=initializer, bias_initializer="zeros", activation='elu'))
# The output neuron is a single fully connected node
#
model_elu.add(Dense(1, kernel_initializer=initializer, activation='sigmoid'))
# Compiling the model
model_elu.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fitting the ANN to the Training set
model_elu.fit(X_train, y_train ,batch_size = 256, epochs = 30, verbose=1)
# Generating Predictions on testing data
Predictions_elu=model_elu.predict(X_test)
#TestingData=pd.DataFrame(data=X_test.values, columns=PredictorVariables)
#TestingData['transact']=y_test.values
TestingData['PredictedtransactElu']=Predictions_elu
#transform Predicted transact from decimal to 0 or 1 for Confusion Matrix & insert into Fairlearn
TestingData["PredictedtransactEluClass"]= np.where(TestingData['PredictedtransactElu']>=0.5, 1, 0)
print(y_test.dtype)
print(TestingData.sample(11))
model_elu.evaluate(X_test, y_test)
#Fairlearn https://fairlearn.org/v0.7.0/quickstart.html
gm_elu = MetricFrame(metrics=accuracy_score, y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactEluClass'], sensitive_features=TestingData['Gender_Byte'])
print("\n")
print("Model score: ")
print(gm_elu.overall)
print("\n" "Model score by category: ")
print(gm_elu.by_group)
print(' Remember, from data wrangling, IF [Gender_Byte]="female" THEN "1" ELSE "0" ENDIF" \n')
#FairLearn Metrics
sr_elu = MetricFrame(metrics=selection_rate,y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactEluClass'], sensitive_features=TestingData['Gender_Byte'])
sr_elu.overall
print("1.0==female, 0.0==male")
sr_elu.by_group
#transform Sensative Varible from 0 or 1 to "male" or "female" for plotting
#TestingData["GenderClass"]= np.where(TestingData['Gender_Byte']>=1, "female","male")
#print(TestingData.head)
metrics = {
'accuracy': accuracy_score,
'precision': precision_score,
'recall': recall_score,
'false positive rate': false_positive_rate,
'true positive rate': true_positive_rate,
'selection rate': selection_rate,
'count': count}
metric_frame_elu = MetricFrame(metrics=metrics,
y_true=TestingData['transact'],
y_pred=TestingData['PredictedtransactEluClass'],
sensitive_features=TestingData["GenderClass"])
metric_frame_elu.by_group.plot.bar(
subplots=True,
layout=[3, 3],
legend=False,
figsize=[12, 8],
title="Show all elu metrics",
)
# + id="ZKTwruYIF3EQ" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="3c9611c4-2ca0-4011-96cf-0cede4614a35"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="Oj775-hISpuw"
# ## Exponential Activation Function
#
# https://keras.io/api/layers/activations/#exponential-function
# + id="SlurB-A1sgAB" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="cc0b230d-3111-425c-ec61-a5c7f4162b24"
# create ANN model https://thinkingneuron.com/how-to-use-artificial-neural-networks-for-classification-in-python/
### Sligtht variation from the rest...exponential Activation function in all layers did not work so relu is in first layer, exponential in hidden layer & sigmoid in output layer ###
model_exponential = Sequential()
# Defining the Input layer and FIRST hidden layer, both are same!
# SPECIAL for exponential model, the relu activation was used in the first hidden layer because the loss was "nan" for each epoch.
model_exponential.add(Dense(19, input_shape=(18,), kernel_initializer=initializer, bias_initializer="zeros", activation='relu'))
# Defining the Second layer of the model
# after the first layer we don't have to specify input_dim as keras configure it automatically
model_exponential.add(Dense(units=5, kernel_initializer=initializer, bias_initializer="zeros", activation='exponential'))
# The output neuron is a single fully connected node
#
model_exponential.add(Dense(1, kernel_initializer=initializer,activation='sigmoid'))
# Compiling the model
model_exponential.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Fitting the ANN to the Training set
model_exponential.fit(X_train, y_train ,batch_size = 256, epochs = 30, verbose=1)
# Generating Predictions on testing data
Predictions_exponential=model_exponential.predict(X_test)
#TestingData=pd.DataFrame(data=X_test.values, columns=PredictorVariables)
#TestingData['transact']=y_test.values
TestingData['PredictedtransactExponential']=Predictions_exponential
#transform Predicted transact from decimal to 0 or 1 for Confusion Matrix & insert into Fairlearn
TestingData["PredictedtransactExponentialClass"]= np.where(TestingData['PredictedtransactExponential']>=0.5, 1, 0)
print(y_test.dtype)
print(TestingData.sample(11))
model_exponential.evaluate(X_test, y_test)
#Fairlearn https://fairlearn.org/v0.7.0/quickstart.html
gm_exponential = MetricFrame(metrics=accuracy_score, y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactExponentialClass'], sensitive_features=TestingData['Gender_Byte'])
print("\n")
print("Model score: ")
print(gm_exponential.overall)
print("\n" "Model score by category: ")
print(gm_exponential.by_group)
print(' Remember, from data wrangling, IF [Gender_Byte]="female" THEN "1" ELSE "0" ENDIF" \n')
#FairLearn Metrics
sr_exponential = MetricFrame(metrics=selection_rate,y_true=TestingData['transact'], y_pred=TestingData['PredictedtransactExponentialClass'], sensitive_features=TestingData['Gender_Byte'])
sr_exponential.overall
print("1.0==female, 0.0==male")
sr_exponential.by_group
#transform Sensative Varible from 0 or 1 to "male" or "female" for plotting
#TestingData["GenderClass"]= np.where(TestingData['Gender_Byte']>=1, "female","male")
#print(TestingData.head)
metrics = {
'accuracy': accuracy_score,
'precision': precision_score,
'recall': recall_score,
'false positive rate': false_positive_rate,
'true positive rate': true_positive_rate,
'selection rate': selection_rate,
'count': count}
metric_frame_exponential = MetricFrame(metrics=metrics,
y_true=TestingData['transact'],
y_pred=TestingData['PredictedtransactExponentialClass'],
sensitive_features=TestingData["GenderClass"])
metric_frame_exponential.by_group.plot.bar(
subplots=True,
layout=[3, 3],
legend=False,
figsize=[12, 8],
title="Show all exponential metrics",
)
# + id="TBlqPcXMF51q" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="e6ceabb8-e11b-47b4-a019-a2e00375d6e4"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="UVD8J5nMnqVT"
# # Fairlearn Metrics
# + [markdown] id="rseY_tMOSxHp"
# ## Overall Model Metrics
# + id="RV10lI91Dy_4" colab={"base_uri": "https://localhost:8080/"} outputId="81c35317-3125-454a-987d-319b944caed4"
print ("Relu\n", metric_frame_relu.overall,
"\n\nTanh\n", metric_frame_tanh.overall,
"\n\nSelu\n", metric_frame_selu.overall,
"\n\nElu\n", metric_frame_elu.overall,
"\n\nExponential\n", metric_frame_exponential.overall,
"\n")
# + [markdown] id="y8F3FclPS4Ji"
# ## By Group Metrics
# + id="wJaQGxVkXXAE" colab={"base_uri": "https://localhost:8080/"} outputId="74bcb9c1-8ad4-4fce-add9-6987259acf8e"
print("Relu =", metric_frame_relu.by_group.to_dict(),"\n")
#print("Sigmoid =", metric_frame_sigmoid.by_group.to_dict(),"\n")
#print("Softmax =", metric_frame_softmax.by_group.to_dict(),"\n")
#print("Softplus =", metric_frame_softplus.by_group.to_dict(),"\n")
#print("Softsign =", metric_frame_softsign.by_group.to_dict(),"\n")
print("Tanh =", metric_frame_tanh.by_group.to_dict(),"\n")
print("Selu =", metric_frame_selu.by_group.to_dict(),"\n")
print("Elu =", metric_frame_elu.by_group.to_dict(),"\n")
print("Exponential=", metric_frame_exponential.by_group.to_dict(),"\n")
# + id="aQgS0toTQJw_" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="f6fd5843-de3a-46f3-924b-33ea075f417c"
print("\n X_test head: \n")
TestingData.head()
# + [markdown] id="0sf2VqkOZcQe"
# # Gender variable DoubleCheck
# Compare to make sure Gender Variable is properly identified. The count should mat counted.
# + id="gux-f8Wz9Xvb" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="cb509bc2-7cec-4566-aa8e-ec0cba243e40"
rp.summary_cont(TestingData["Gender_Byte"])
# + id="Nt1lTAvfOmLC" colab={"base_uri": "https://localhost:8080/", "height": 112} outputId="22141c40-e842-4b8e-cd21-4ec30520a1a0"
rp.summary_cat(TestingData["Gender_Byte"])
# + id="4g_jpsLGFetE" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="d633e338-3e94-4893-fb15-9900b6dc5e00"
print("\n X_test head: \n")
TestingData.head()
| V6_R_E_sig_ANN_FairLearnFrenchData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from time import sleep
# load the selenium driver from your machine
driver = webdriver.Chrome(
"/Users/jappanjeetsingh/Downloads/drivers/chromedriver")
driver.get("https://www.worldometers.info/coronavirus/")
sleep(5)
# finding the xpath to the table
table = driver.find_element_by_xpath(
"//*[@id=\"main_table_countries_today\"]/tbody[1]")
# finding the country
country = table.find_element_by_xpath("//td[contains(., 'India')]")
# we find the parent element of the country element and store it into row because we want all data in the row
row = country.find_element_by_xpath("./..")
data = row.text.split(" ")
print("Country: " + country.text)
print("Total cases: " + data[2])
print("New cases: " + data[3])
print("Total deaths: " + data[4])
print("New deaths: " + data[5])
print("Active cases: " + data[6])
print("Total recovered: " + data[7])
print("Serious, critical cases: " + data[8])
| Web Scraping (Beautiful Soup, Scrapy, Selenium)/webScraping_Day31/Scrape-Covid-India-Scenario/.ipynb_checkpoints/solution-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (lsl_eeg)
# language: python
# name: lsl_eeg
# ---
import os
# +
files = os.listdir("./data/raw/msw/ExperimentData")
for f in files: # iterate over files
# remove hidden/config files and folders
if not f.endswith(".raw"):
files.remove(f)
uids_exp = [f.split("-")[2].split(".")[0] for f in files]
# +
files = os.listdir("./data/raw/msw/EventData")
for f in files: # iterate over files
# remove hidden/config files and folders
if not f.endswith(".bin"):
files.remove(f)
uids_eve = [f.split("-")[2].split(".")[0] for f in files]
# -
print(len(uids_exp))
print(len(uids_eve))
exp = set(uids_exp)
eve = set(uids_eve)
diff_a = exp - eve
diff_a
diff_b = eve - exp
diff_b
for f in files:
uid = f.split("-")[2].split(".")[0]
if uid in diff_b:
os.remove(f"./data/raw/msw/EventData/{f}")
print(f"Removed: {f}")
| checks/raycast_prep/files_check_msw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computation on Arrays: Broadcasting
# We saw in the previous section how NumPy's universal functions can be used to *vectorize* operations and thereby remove slow Python loops.
# Another means of vectorizing operations is to use NumPy's *broadcasting* functionality.
# Broadcasting is simply a set of rules for applying binary ufuncs (e.g., addition, subtraction, multiplication, etc.) on arrays of different sizes.
# ## Introducing Broadcasting
#
# Recall that for arrays of the same size, binary operations are performed on an element-by-element basis:
import numpy as np
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
a + b
# Broadcasting allows these types of binary operations to be performed on arrays of different sizes–for example, we can just as easily add a scalar (think of it as a zero-dimensional array) to an array:
a + 5
# We can think of this as an operation that stretches or duplicates the value ``5`` into the array ``[5, 5, 5]``, and adds the results.
# The advantage of NumPy's broadcasting is that this duplication of values does not actually take place, but it is a useful mental model as we think about broadcasting.
#
# We can similarly extend this to arrays of higher dimension. Observe the result when we add a one-dimensional array to a two-dimensional array:
M = np.ones((3, 3))
M
M + a
# Here the one-dimensional array ``a`` is stretched, or broadcast across the second dimension in order to match the shape of ``M``.
#
# While these examples are relatively easy to understand, more complicated cases can involve broadcasting of both arrays. Consider the following example:
# +
a = np.arange(3)
b = np.arange(3)[:, np.newaxis]
print(a)
print(b)
# -
a + b
# Just as before we stretched or broadcasted one value to match the shape of the other, here we've stretched *both* ``a`` and ``b`` to match a common shape, and the result is a two-dimensional array!
# The geometry of these examples is visualized in the following figure.
# 
# The light boxes represent the broadcasted values: again, this extra memory is not actually allocated in the course of the operation, but it can be useful conceptually to imagine that it is.
# ## Rules of Broadcasting
#
# Broadcasting in NumPy follows a strict set of rules to determine the interaction between the two arrays:
#
# - Rule 1: If the two arrays differ in their number of dimensions, the shape of the one with fewer dimensions is *padded* with ones on its leading (left) side.
# - Rule 2: If the shape of the two arrays does not match in any dimension, the array with shape equal to 1 in that dimension is stretched to match the other shape.
# - Rule 3: If in any dimension the sizes disagree and neither is equal to 1, an error is raised.
#
# To make these rules clear, let's consider a few examples in detail.
# ### Broadcasting example 1
#
# Let's look at adding a two-dimensional array to a one-dimensional array:
M = np.ones((2, 3))
a = np.arange(3)
# Let's consider an operation on these two arrays. The shape of the arrays are
#
# - ``M.shape = (2, 3)``
# - ``a.shape = (3,)``
#
# We see by rule 1 that the array ``a`` has fewer dimensions, so we pad it on the left with ones:
#
# - ``M.shape -> (2, 3)``
# - ``a.shape -> (1, 3)``
#
# By rule 2, we now see that the first dimension disagrees, so we stretch this dimension to match:
#
# - ``M.shape -> (2, 3)``
# - ``a.shape -> (2, 3)``
#
# The shapes match, and we see that the final shape will be ``(2, 3)``:
M + a
# ### Broadcasting example 2
#
# Let's take a look at an example where both arrays need to be broadcast:
a = np.arange(3).reshape((3, 1))
b = np.arange(3)
# Again, we'll start by writing out the shape of the arrays:
#
# - ``a.shape = (3, 1)``
# - ``b.shape = (3,)``
#
# Rule 1 says we must pad the shape of ``b`` with ones:
#
# - ``a.shape -> (3, 1)``
# - ``b.shape -> (1, 3)``
#
# And rule 2 tells us that we upgrade each of these ones to match the corresponding size of the other array:
#
# - ``a.shape -> (3, 3)``
# - ``b.shape -> (3, 3)``
#
# Because the result matches, these shapes are compatible. We can see this here:
a + b
# ### Broadcasting example 3
#
# Now let's take a look at an example in which the two arrays are not compatible:
M = np.ones((3, 2))
a = np.arange(3)
# This is just a slightly different situation than in the first example: the matrix ``M`` is transposed.
# How does this affect the calculation? The shape of the arrays are
#
# - ``M.shape = (3, 2)``
# - ``a.shape = (3,)``
#
# Again, rule 1 tells us that we must pad the shape of ``a`` with ones:
#
# - ``M.shape -> (3, 2)``
# - ``a.shape -> (1, 3)``
#
# By rule 2, the first dimension of ``a`` is stretched to match that of ``M``:
#
# - ``M.shape -> (3, 2)``
# - ``a.shape -> (3, 3)``
#
# Now we hit rule 3–the final shapes do not match, so these two arrays are incompatible, as we can observe by attempting this operation:
M + a
# Note the potential confusion here: you could imagine making ``a`` and ``M`` compatible by, say, padding ``a``'s shape with ones on the right rather than the left.
# But this is not how the broadcasting rules work!
# That sort of flexibility might be useful in some cases, but it would lead to potential areas of ambiguity.
# If right-side padding is what you'd like, you can do this explicitly by reshaping the array (we'll use the ``np.newaxis`` keyword introduced in The Basics of NumPy Arrays):
a[:, np.newaxis].shape
M + a[:, np.newaxis]
# Also note that while we've been focusing on the ``+`` operator here, these broadcasting rules apply to *any* binary ``ufunc``.
# For example, here is the ``logaddexp(a, b)`` function, which computes ``log(exp(a) + exp(b))`` with more precision than the naive approach:
np.logaddexp(M, a[:, np.newaxis])
# For more information on the many available universal functions, refer to Computation on NumPy Arrays: Universal Functions.
# ## Broadcasting in Practice
# Broadcasting operations form the core of many examples we'll see throughout this book.
# We'll now take a look at a couple simple examples of where they can be useful.
# ### Centering an array
# In the previous section, we saw that ufuncs allow a NumPy user to remove the need to explicitly write slow Python loops. Broadcasting extends this ability.
# One commonly seen example is when centering an array of data.
# Imagine you have an array of 10 observations, each of which consists of 3 values.
# Using the standard convention, we'll store this in a $10 \times 3$ array:
X = np.random.random((10, 3))
# We can compute the mean of each feature using the ``mean`` aggregate across the first dimension:
Xmean = X.mean(0)
Xmean
# And now we can center the ``X`` array by subtracting the mean (this is a broadcasting operation):
X_centered = X - Xmean
# To double-check that we've done this correctly, we can check that the centered array has near zero mean:
X_centered.mean(0)
# To within machine precision, the mean is now zero.
# ### Plotting a two-dimensional function
# One place that broadcasting is very useful is in displaying images based on two-dimensional functions.
# If we want to define a function $z = f(x, y)$, broadcasting can be used to compute the function across the grid:
# +
# x and y have 50 steps from 0 to 5
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)[:, np.newaxis]
z = np.sin(x) ** 10 + np.cos(10 + y * x) * np.cos(x)
# -
# We'll use Matplotlib to plot this two-dimensional array (these tools will be discussed in full in Density and Contour Plots):
# %matplotlib inline
import matplotlib.pyplot as plt
plt.imshow(z, origin='lower', extent=[0, 5, 0, 5],
cmap='viridis')
plt.colorbar();
# The result is a compelling visualization of the two-dimensional function.
| notebooks/D1_L4_NumPy/05-Computation-on-arrays-broadcasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!DOCTYPE html>
# <html>
# <body>
# <div align="center">
# <h3>Prepared by <NAME></h3>
#
# <h1>Data Visualization With Matplotlib</h1>
#
# <h3>Follow Me on - <a href="https://www.linkedin.com/in/asif-bhat/">LinkedIn</a> <a href="https://mobile.twitter.com/_asifbhat_">Twitter</a> <a href="https://www.instagram.com/datasciencescoop/?hl=en">Instagram</a> <a href="https://www.facebook.com/datasciencescoop/">Facebook</a></h3>
# </div>
#
# </div>
# </body>
# </html>
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
#Graph Styling
# https://tonysyu.github.io/raw_content/matplotlib-style-gallery/gallery.html
plt.style.use('seaborn-darkgrid')
# # Bar Graphs
#Simple Bar Chart
id1 = np.arange(1,10)
score = np.arange(20,110,10)
plt.bar(id1,score)
plt.xlabel('Student ID')
plt.ylabel('Score')
plt.show()
# Changing color of the bar chart
id1 = np.arange(1,10)
score = np.arange(20,110,10)
plt.figure(figsize=(8,5)) # Setting the figure size
ax = plt.axes()
ax.set_facecolor("#ECF0F1") # Setting the background color by specifying the HEX Code
plt.bar(id1,score,color = '#FFA726')
plt.xlabel(r'$Student $ $ ID$')
plt.ylabel(r'$Score$')
plt.show()
#Plotting multiple sets of data
x1= [1,3,5,7]
x2=[2,4,6,8]
y1 = [7,7,7,7]
y2= [17,18,29,40]
plt.figure(figsize=(8,6))
ax = plt.axes()
ax.set_facecolor("white")
plt.bar(x1,y1,label = "First",color = '#42B300') # First set of data
plt.bar(x2,y2,label = "Second",color = '#94E413') # Second set of data
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# Horizontal Bar Chart
Age = [28,33,43,45,57]
Name = ["Asif", "Steve", 'John', "Ravi", "Basit"]
plt.barh(Name,Age, color ="yellowgreen")
plt.show()
# Changing the width of Bars
num1 = np.array([1,3,5,7,9])
num2 = np.array([2,4,6,8,10])
plt.figure(figsize=(8,4))
plt.bar(num1, num1**2, width=0.2 , color = '#FF6F00')
plt.bar(num2, num2**2, width=0.2 , color = '#FFB300')
plt.plot()
# Displaying values at the top of vertical bars
num1 = np.array([1,3,5,7,9])
num2 = np.array([2,4,6,8,10])
plt.figure(figsize=(10,6))
plt.bar(num1, num1**2, width=0.3 , color = '#FF6F00')
plt.bar(num2, num2**2, width=0.3 , color = '#FFB300')
for x,y in zip(num1,num1**2):
plt.text(x, y+0.05, '%d' % y, ha='center' , va= 'bottom')
for x,y in zip(num2,num2**2):
plt.text(x, y+0.05, '%d' % y, ha='center' , va= 'bottom')
plt.plot()
# +
x = np.arange(1,21)
plt.figure(figsize=(16,8))
y1 = np.random.uniform(0.1,0.7,20)
y2 = np.random.uniform(0.1,0.7,20)
plt.bar(x, +y1, facecolor='#C0CA33', edgecolor='white') #specify edgecolor by name
plt.bar(x, -y2, facecolor='#FF9800', edgecolor='white')
for x,y in zip(x,y1):
plt.text(x, y+0.05, '%.2f' % y, ha='center' , va= 'bottom', fontsize = 10)
plt.xlim(0,21)
plt.ylim(-1.25,+1.25)
plt.show()
# -
# ### Stacked Vertical Bar
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= np.array([17,18,29,40])
y2 =np.array([20,21,22,23])
y3 =np.array([5,9,11,12])
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.bar(x1,y3,label = "Cancelled Tickets",width = 0.5 ,bottom = y1+y2 , color = '#F7DC6F')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# ### Grouped Bar Chart
# +
# Grouped Bar Chart
plt.figure(figsize=(7,9))
# set width of bar
barWidth = 0.25
# set height of bar
y1= np.array([17,18,29,40])
y2 =np.array([20,21,22,23])
y3 =np.array([5,9,11,12])
# Set position of bar on X axis
pos1 = np.arange(len(y1))
pos2 = [x + barWidth for x in pos1]
pos3 = [x + barWidth for x in pos2]
# Make the plot
plt.bar(pos1, y1, color='#FBC02D', width=barWidth, label='Open')
plt.bar(pos2, y2, color='#F57F17', width=barWidth, label='Closed')
plt.bar(pos3, y3, color='#E65100', width=barWidth, label='Cancelled')
# Add xticks on the middle of the group bars
plt.xlabel('Assignee', fontweight='bold')
plt.ylabel('Number of Tickets', fontweight='bold')
plt.xticks([i + barWidth for i in range(len(y1))], ['Asif', 'Basit', 'Ravi', 'Minil'])
# Create legend & Show graphic
plt.legend()
plt.show()
np.arange(len(y1))
# -
# ### Stacked Vertical Bar
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(8,5))
plt.barh(x1,y1,label = "Open Tickets",color = '#FF6F00')
plt.barh(x1,y2,label = "Closed Tickets", left = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# ### Displaying values in Bar Charts
# +
# Displaying values in the stacked vertical bars using plt.text()
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(5,7))
plt.bar(x1,y1,label = "Open Tickets",width = 0.5,color = '#FF6F00')
plt.bar(x1,y2,label = "Closed Tickets",width = 0.5 ,bottom = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
plt.title ('$Bar $ $ Chart$')
for x,y in zip(x1,y1):
plt.text(x, y-10, '%d' % y, ha='center' , va= 'bottom')
for x,y,z in zip(x1,y2,y1):
plt.text(x, y+z-10, '%d' % y, ha='center' , va= 'bottom')
plt.legend()
plt.show()
# +
# Displaying values in the stacked horizontal bars using plt.text()
plt.style.use('seaborn-darkgrid')
x1= ['Asif','Basit','Ravi','Minil']
y1= [17,18,29,40]
y2 = [20,21,22,23]
plt.figure(figsize=(8,5))
plt.barh(x1,y1,label = "Open Tickets",color = '#FF6F00')
plt.barh(x1,y2,label = "Closed Tickets", left = y1 , color = '#FFB300')
plt.xlabel('$X$')
plt.ylabel('$Y$')
for x,y in zip(x1,y1):
plt.text(y-10, x, '%d' % y, ha='center' , va= 'bottom')
for x,y,z in zip(x1,y2,y1):
plt.text(y+z-10, x, '%d' % y, ha='center' , va= 'bottom')
plt.title ('$Bar $ $ Chart$')
plt.legend()
plt.show()
# +
# Displaying values at the top of the Grouped Bar Chart using plt.text()
plt.figure(figsize=(7,9))
# set width of bar
barWidth = 0.25
# set height of bar
y1= np.array([17,18,29,40])
y2 =np.array([20,21,22,23])
y3 =np.array([5,9,11,12])
# Set position of bar on X axis
pos1 = np.arange(len(y1))
pos2 = [x + barWidth for x in pos1]
pos3 = [x + barWidth for x in pos2]
# Make the plot
plt.bar(pos1, y1, color='#FBC02D', width=barWidth, label='Open')
plt.bar(pos2, y2, color='#F57F17', width=barWidth, label='Closed')
plt.bar(pos3, y3, color='#E65100', width=barWidth, label='Cancelled')
# Add xticks on the middle of the group bars
plt.xlabel('Assignee', fontweight='bold')
plt.ylabel('Number of Tickets', fontweight='bold')
plt.xticks([i + barWidth for i in range(len(y1))], ['Asif', 'Basit', 'Ravi', 'Minil'])
for x,y in zip(pos1,y1):
plt.text(x, y, '%d' % y, ha='center' , va= 'bottom')
for x,y in zip(pos2,y2):
plt.text(x, y, '%d' % y, ha='center' , va= 'bottom')
for x,y in zip(pos3,y3):
plt.text(x, y, '%d' % y, ha='center' , va= 'bottom')
plt.title ('$Grouped $ $ Bar $ $ Chart$')
# Create legend & Show graphic
plt.legend()
plt.show()
| Matplotlib/2. Bar Charts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Regular Expression
def search(pattern, text):
"""Return true if pattern appears anywhere in text."""
if pattern[0] == "^":
return match(pattern[1:], text)
else:
return match(".*"+pattern, text)
def match1(p, text):
"""Return true if first character of text matches pattern character p."""
if not text: return False
return p == '.' or p == text[0]
def match_star(p, pattern, text):
"""Return true if any number of char p, followed by pattern, matching text."""
return (match1(p, text) and match_star(p, pattern, text[1:])) or match(pattern, text)
def match(pattern, text):
"""Return True if pattern appears at the start of text."""
if pattern == "":
return True
elif pattern == "$":
return (text == "")
elif len(pattern) > 1 and pattern[1] in "*?":
p, op, pat = pattern[0], pattern[1], pattern[2:]
if op == "*":
return match_star(p, pat, text)
elif op == "?":
if match1(p, text) and match(pat, text[1:]):
return True
else:
return match(pat, text)
else:
return match1(pattern[0], text) and match(pattern[1:], text[1:])
def test():
assert search('baa*!', 'Sheep said baaaa!')
assert search('baa*!', 'Sheep said baaaa humbug') == False
assert match('baa*!', 'Sheep said baaaa!') == False
print "All test pass!"
test()
# # Language Processor
# +
def lit(string): return ('lit', string)
def seq(x, y): return ('seq', x, y)
def alt(x, y): return ('alt', x, y)
def star(x): return ('star', x)
def plus(x): return ('seq', x, star(x))
def opt(x): return alt(lit(''), x) #opt(x) means that x is optional
def oneof(chars): return ('oneof', tuple(chars))
dot = ('dot',)
eol = ('eol',)
def test():
assert lit('abc') == ('lit', 'abc')
assert seq(('lit', 'a'),
('lit', 'b')) == ('seq', ('lit', 'a'), ('lit', 'b'))
assert alt(('lit', 'a'),
('lit', 'b')) == ('alt', ('lit', 'a'), ('lit', 'b'))
assert star(('lit', 'a')) == ('star', ('lit', 'a'))
assert plus(('lit', 'c')) == ('seq', ('lit', 'c'),
('star', ('lit', 'c')))
assert opt(('lit', 'x')) == ('alt', ('lit', ''), ('lit', 'x'))
assert oneof('abc') == ('oneof', ('a', 'b', 'c'))
return 'tests pass'
print test()
# +
def matchset(pattern, text):
"Match pattern at start of text; return a set of remainders of text."
op, x, y = components(pattern)
if 'lit' == op:
return set([text[len(x):]]) if text.startswith(x) else null
elif 'seq' == op:
return set(t2 for t1 in matchset(x, text) for t2 in matchset(y, t1))
elif 'alt' == op:
return matchset(x, text) | matchset(y, text)
elif 'dot' == op:
return set([text[1:]]) if text else null
elif 'oneof' == op:
return set([text[1:]]) if text.startswith(x) else null
elif 'eol' == op:
return set(['']) if text == '' else null
elif 'star' == op:
return (set([text]) |
set(t2 for t1 in matchset(x, text)
for t2 in matchset(pattern, t1) if t1 != text))
else:
raise ValueError('unknown pattern: %s' % pattern)
null = frozenset()
def components(pattern):
"Return the op, x, and y arguments; x and y are None if missing."
x = pattern[1] if len(pattern) > 1 else None
y = pattern[2] if len(pattern) > 2 else None
return pattern[0], x, y
def test():
assert matchset(('lit', 'abc'), 'abcdef') == set(['def'])
assert matchset(('seq', ('lit', 'hi '),
('lit', 'there ')),
'hi there nice to meet you') == set(['nice to meet you'])
assert matchset(('alt', ('lit', 'dog'),
('lit', 'cat')), 'dog and cat') == set([' and cat'])
assert matchset(('dot',), 'am i missing something?') == set(['m i missing something?'])
assert matchset(('oneof', 'a'), 'aabc123') == set(['abc123'])
assert matchset(('eol',),'') == set([''])
assert matchset(('eol',),'not end of line') == frozenset([])
assert matchset(('star', ('lit', 'hey')), 'heyhey!') == set(['!', 'heyhey!', 'hey!'])
return 'tests pass'
print test()
# +
def search(pattern, text):
"Match pattern anywhere in text; return longest earliest match or None."
for i in range(len(text)):
m = match(pattern, text[i:])
if m is not None:
return m
def match(pattern, text):
"Match pattern against start of text; return longest match found or None."
remainders = matchset(pattern, text)
if remainders:
shortest = min(remainders, key=len)
return text[:len(text)-len(shortest)]
def test():
assert match(('star', ('lit', 'a')),'aaabcd') == 'aaa'
assert match(('alt', ('lit', 'b'), ('lit', 'c')), 'ab') == None
assert match(('alt', ('lit', 'b'), ('lit', 'a')), 'ab') == 'a'
assert match(seq(star(lit('a')), star(lit('b'))), 'abbv') == 'abb'
assert search(('alt', ('lit', 'b'), ('lit', 'c')), 'ab') == 'b'
return 'tests pass'
print test()
# +
def n_ary(f):
"""Given binary function f(x, y), return an n_ary function such
that f(x, y, z) = f(x, f(y,z)), etc. Also allow f(x) = x."""
def n_ary_f(x, *args):
return x if not args else f(x, n_ary_f(*args))
return n_ary_f
def test():
f = lambda x, y: ('seq', x, y)
g = n_ary(f)
assert g(2,3,4) == ('seq', 2, ('seq', 3, 4))
assert g(2) == 2
assert g(2,3) == ('seq', 2, 3)
return "tests pass"
print test()
# -
# # Decorator
# +
from functools import update_wrapper
def n_ary(f):
"""Given binary function f(x, y), return an n_ary function such
that f(x, y, z) = f(x, f(y,z)), etc. Also allow f(x) = x."""
def n_ary_f(x, *args):
return x if not args else f(x, n_ary_f(*args))
update_wrapper(n_ary_f, f) # update the function name and doc
return n_ary_f
@n_ary
def seq(x, y): return ('seq', x, y)
help(seq)
# +
def decorator(d):
def _d(fn):
return update_wrapper(d(fn), fn)
return update_wrapper(_d, d)
@decorator
def n_ary(f):
"""Given binary function f(x, y), return an n_ary function such
that f(x, y, z) = f(x, f(y,z)), etc. Also allow f(x) = x."""
def n_ary_f(x, *args):
return x if not args else f(x, n_ary_f(*args))
return n_ary_f
@n_ary
def seq(x, y): return ('seq', x, y)
help(seq)
help(n_ary)
print seq(2,3,4)
# -
# # Problem Set 1
# +
from functools import update_wrapper
from string import split
import re
def grammar(description, whitespace=r'\s*'):
"""Convert a description to a grammar. Each line is a rule for a
non-terminal symbol; it looks like this:
Symbol => A1 A2 ... | B1 B2 ... | C1 C2 ...
where the right-hand side is one or more alternatives, separated by
the '|' sign. Each alternative is a sequence of atoms, separated by
spaces. An atom is either a symbol on some left-hand side, or it is
a regular expression that will be passed to re.match to match a token.
Notation for *, +, or ? not allowed in a rule alternative (but ok
within a token). Use '\' to continue long lines. You must include spaces
or tabs around '=>' and '|'. That's within the grammar description itself.
The grammar that gets defined allows whitespace between tokens by default;
specify '' as the second argument to grammar() to disallow this (or supply
any regular expression to describe allowable whitespace between tokens)."""
G = {' ': whitespace}
description = description.replace('\t', ' ') # no tabs!
for line in split(description, '\n'):
lhs, rhs = split(line, ' => ', 1)
alternatives = split(rhs, ' | ')
G[lhs] = tuple(map(split, alternatives))
return G
def decorator(d):
"Make function d a decorator: d wraps a function fn."
def _d(fn):
return update_wrapper(d(fn), fn)
update_wrapper(_d, d)
return _d
@decorator
def memo(f):
"""Decorator that caches the return value for each call to f(args).
Then when called again with same args, we can just look it up."""
cache = {}
def _f(*args):
try:
return cache[args]
except KeyError:
cache[args] = result = f(*args)
return result
except TypeError:
# some element of args can't be a dict key
return f(args)
return _f
def parse(start_symbol, text, grammar):
"""Example call: parse('Exp', '3*x + b', G).
Returns a (tree, remainder) pair. If remainder is '', it parsed the whole
string. Failure iff remainder is None. This is a deterministic PEG parser,
so rule order (left-to-right) matters. Do 'E => T op E | T', putting the
longest parse first; don't do 'E => T | T op E'
Also, no left recursion allowed: don't do 'E => E op T'"""
tokenizer = grammar[' '] + '(%s)'
def parse_sequence(sequence, text):
result = []
for atom in sequence:
tree, text = parse_atom(atom, text)
if text is None: return Fail
result.append(tree)
return result, text
@memo
def parse_atom(atom, text):
if atom in grammar: # Non-Terminal: tuple of alternatives
for alternative in grammar[atom]:
tree, rem = parse_sequence(alternative, text)
if rem is not None: return [atom]+tree, rem
return Fail
else: # Terminal: match characters against start of text
m = re.match(tokenizer % atom, text)
return Fail if (not m) else (m.group(1), text[m.end():])
# Body of parse:
return parse_atom(start_symbol, text)
Fail = (None, None)
JSON = grammar("""value => number | array | string | object | true | false | null
object => { } | { members }
members => pair , members | pair
pair => string : value
array => [[] []] | [[] elements []]
elements => value , elements | value
string => "[^"]*"
number => int frac exp | int frac | int exp | int
int => [-+]?[1-9][0-9]*
frac => [.][0-9]+
exp => [eE][-+][0-9]+""", whitespace='\s*')
def json_parse(text):
return parse('value', text, JSON)
def test():
assert json_parse('["testing", 1, 2, 3]') == (
['value', ['array', '[', ['elements', ['value',
['string', '"testing"']], ',', ['elements', ['value', ['number',
['int', '1']]], ',', ['elements', ['value', ['number',
['int', '2']]], ',', ['elements', ['value', ['number',
['int', '3']]]]]]], ']']], '')
assert json_parse('-123.456e+789') == (
['value', ['number', ['int', '-123'], ['frac', '.456'], ['exp', 'e+789']]], '')
assert json_parse('{"age": 21, "state":"CO","occupation":"rides the rodeo"}') == (
['value', ['object', '{', ['members', ['pair', ['string', '"age"'],
':', ['value', ['number', ['int', '21']]]], ',', ['members',
['pair', ['string', '"state"'], ':', ['value', ['string', '"CO"']]],
',', ['members', ['pair', ['string', '"occupation"'], ':',
['value', ['string', '"rides the rodeo"']]]]]], '}']], '')
return 'tests pass'
print test()
# -
# # Problem Set 3
# +
import re
def findtags(text):
parms = '(?:\w+\s*=\s*"[^"]*"\s*)*'
tags = '(<\s*\w+\s*' + parms + '\s*/?>)'
return re.findall(tags, text)
testtext1 = """
My favorite website in the world is probably
<a href="www.udacity.com">Udacity</a>. If you want
that link to open in a <b>new tab</b> by default, you should
write <a href="www.udacity.com"target="_blank">Udacity</a>
instead!
"""
testtext2 = """
Okay, so you passed the first test case. <let's see> how you
handle this one. Did you know that 2 < 3 should return True?
So should 3 > 2. But 2 > 3 is always False.
"""
testtext3 = """
It's not common, but we can put a LOT of whitespace into
our HTML tags. For example, we can make something bold by
doing < b > this < /b >, Though I
don't know why you would ever want to.
"""
def test():
assert findtags(testtext1) == ['<a href="www.udacity.com">',
'<b>',
'<a href="www.udacity.com"target="_blank">']
assert findtags(testtext2) == []
assert findtags(testtext3) == ['< b >']
return 'tests pass'
print test()
# -
| Making Tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/novoic/ml-challenge/blob/master/text_challenge.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="NOHLZHtggQtk"
# <a href="https://novoic.com"><img src="https://assets.novoic.com/logo_320px.png" alt="Novoic logo" width="160"/></a>
#
# # Novoic ML challenge – text data
# + [markdown] colab_type="text" id="zlll6t_8M5fy"
# ## Introduction
# Welcome to the Novoic ML challenge!
#
# This is an open-ended ML challenge to help us identify exceptional researchers and engineers. The guidance below describes an open-source dataset that you can use to demonstrate your research skills, creativity, coding ability, scientific communication or anything else you think is important to the role.
#
# Before starting the challenge, go ahead and read our CEO's [Medium post](https://medium.com/@emil_45669/the-doctor-is-ready-to-see-you-tube-videos-716b12367feb) on what we're looking for in our Research Scientists, Research Engineers and ML Interns. We recommend you spend around three hours on this (more or less if you wish), which you do not have to do in one go. Please make use of any resources you like.
#
# This is the text version of the challenge. Also available are audio and image versions. You can access all three from [this GitHub repo](https://github.com/novoic/ml-challenge).
#
# Best of luck – we're looking forward to seeing what you can do!
# + [markdown] colab_type="text" id="GUJZyzMB_2TA"
# ## Prepare the data
# Copy the dataset to a local directory – this should take a few seconds.
# + colab={} colab_type="code" id="AaRNBDz4nN1t"
# !mkdir -p data
# !gsutil -m cp -r gs://novoic-ml-challenge-text-data/* ./data
# + [markdown] colab_type="text" id="BpJ7NOCXkCKs"
# ## Data description
#
# The data comprises 5,574 SMS messages. Each message is labelled as either 'ham' (legitimate) or spam.
#
# Each line in `data.txt` corresponds to one message. The first word is the data label (either `ham` or `spam`), followed by a tab (`\t`) character and then the message.
# + colab={} colab_type="code" id="muZyXMc8rbAK"
with open('data/data.txt', 'r') as f:
msgs = f.read().splitlines()
# + colab={} colab_type="code" id="6Ey4KqN1uBqN"
print(msgs[10])
print(msgs[11])
# + [markdown] colab_type="text" id="UeYkLol_rZoT"
#
# For more information about the dataset, see its `README.md`.
#
# Directory structure:
# ```
# data/
# ├── data.txt
# ├── LICENSE
# └── README.md
# ```
#
#
#
#
#
# + [markdown] colab_type="text" id="K7x5PwDaFDdy"
# ## The challenge
# This is an open-ended challenge and we want to witness your creativity. Some suggestions:
# - Data exploration/visualization
# - Binary classification
# - Unsupervised clustering
# - Model explainability
#
# You're welcome to explore one or more of these topics, or do something entirely different.
#
# Create, iterate on, and validate your work in this notebook, using any packages of your choosing.
#
# **You can access a GPU via `Runtime -> Change runtime type` in the toolbar.**
#
# ## Submission instructions
# Once you're done, send this `.ipynb` notebook (or a link to it hosted on Google Drive/GitHub with appropriate permissions) to <EMAIL>, ensuring that outputs from cells (text, plots etc) are preserved.
#
# If you haven't applied already, make sure you submit an application first through our [job board](https://novoic.com/careers/).
# + [markdown] colab_type="text" id="VXJdZxNrK008"
# ## Your submission
# The below sets up TensorFlow as an example but feel free to use any framework you like.
# + colab={} colab_type="code" id="t7A3vYU3LRz_"
# The default TensorFlow version on Colab is 1.x. Uncomment the below to use TensorFlow 2.x instead.
# # %tensorflow_version 2.x
# + colab={} colab_type="code" id="d_-0bsdzK6sy"
import tensorflow as tf
tf.__version__
# + [markdown] colab_type="text" id="Y8PoDC7xLkU4"
# Take the wheel!
| .ipynb_checkpoints/text_challenge-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# 
#
# # ExempliPy
# by [<NAME>](www.tonysaad.net) <br/>
# Assistant Professor of [Chemical Engineering](www.che.utah.edu) <br/>
# [University of Utah](www.utah.edu)
#
#
# A collection of example of problems solved numerically with Python. Applications span physics, chemical, mechanical, civil, and electrical engineering.
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#ExempliPy" data-toc-modified-id="ExempliPy-1"><span class="toc-item-num">1 </span>ExempliPy</a></div><div class="lev1 toc-item"><a href="#Free-Fall:-ODE-Time-Integration" data-toc-modified-id="Free-Fall:-ODE-Time-Integration-2"><span class="toc-item-num">2 </span>Free Fall: ODE Time Integration</a></div><div class="lev2 toc-item"><a href="#Method-1:-Using-Lists" data-toc-modified-id="Method-1:-Using-Lists-21"><span class="toc-item-num">2.1 </span>Method 1: Using Lists</a></div><div class="lev2 toc-item"><a href="#Method-2:-Using-Numpy-Arrays" data-toc-modified-id="Method-2:-Using-Numpy-Arrays-22"><span class="toc-item-num">2.2 </span>Method 2: Using Numpy Arrays</a></div><div class="lev1 toc-item"><a href="#Interpolation" data-toc-modified-id="Interpolation-3"><span class="toc-item-num">3 </span>Interpolation</a></div>
# + [markdown] slideshow={"slide_type": "slide"}
# # Free Fall: ODE Time Integration
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider the free fall of an astronaut subject to drag. The governing equation is according to Newton's second law is:
# $$m \frac{\text{d}u}{\text{d} t} = m g - c u$$
# or
# $$\frac{\text{d}u}{\text{d} t} = g - \frac{c}{m} u$$
# where $u$(m/s) is the (downward) speed of the astronaut, $g$(m/s/s) is the acceleration of gravity, and $c$(kg/s) is the drag coefficient. Here, the drag force acts in the direction opposite to the fall of the astronaut and is given by $F_\text{D} = cu\mathbf{j}$
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Method 1: Using Lists
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
dt = 0.5 # step size in s
u=[0.0] # create a list for the velocity array. This contains the initial condition
t=[0.0] # create a list for the time array. starts at t = 0.0
tend = 30.0 # set end time
c = 12.5 # drag coefficientkg/s
m = 60 # object's mass, kg
g = 9.81 # gravitational acceleration m/s/s
# t[-1] returns the last element in the list
while t[-1] < tend:
unp1 = u[-1] + dt * (g - c/m*u[-1]) # time advance
u.append(unp1)
t.append(t[-1] + dt)
# tplot = np.linspace(t0,tend,len(u)) # create an equally space array for time. This will be used for plotting.
plt.plot(t,u)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Method 2: Using Numpy Arrays
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
dt = 0.5 # step size in s
t0 = 0.0 # set initial time
tend = 30.0 # set end time
ndt = int( (tend-t0)/dt ) # number of timesteps that we will take
t = np.linspace(t0,tend,ndt) # create an equally space array for time. This will be used for plotting.
u= np.zeros(ndt) # allocate a numpy array of the same size as the number of timesteps
c = 12.5 # drag coefficientkg/s
m = 60 # object's mass, kg
g = 9.81 # gravitational acceleration m/s/s
n = 0 # just a counter
while n < ndt-1:
u[n+1] = u[n] + dt *(g - c/m*u[n]) # time advance
n += 1
plt.plot(t,u)
# + [markdown] slideshow={"slide_type": "slide"}
# # Interpolation
# + [markdown] slideshow={"slide_type": "fragment"}
# Use linear, polynomial, and cubic spline interpolants to interpolate the function
# $$ f(x) = e^{-x^2/\sigma^2}$$
# on the interval $[-1,1]$. Start with $n=10$ samples and experiment with different values of the standard deviation, $\sigma$.
# + slideshow={"slide_type": "subslide"}
import numpy as np
from numpy import interp
from numpy import polyfit, polyval, poly1d
from scipy.interpolate import CubicSpline
# %matplotlib inline
import matplotlib.pyplot as plt
n = 10 # sampling points - we will use this many samples
# we want to interpolate this gaussian data
σ = 0.4
y = lambda x: np.exp(-x**2/σ**2)
# exact solution
xe = np.linspace(-1,1,200) # create equally spaced points between -1 and 1
ye = y(xe)
plt.figure(figsize=(8, 6))
# sampling points
xi = np.linspace(-1,1,n)
yi = y(xi)
# plot sample point locations
plt.plot(xi,yi,'o',markersize=10)
plt.plot(xe,ye,'k-',label='Exact')
# linear interpolation. Interpolate to to xe using sampling points xi
ylin = interp(xe, xi, yi)
plt.plot(xe,ylin,'r-',label='Linear')
# polynomial interpolation. Interpolate to to xe using sampling points xi
p = np.polyfit(xi, yi, n-1)
ypoly =polyval(p,xe)
plt.plot(xe,ypoly,'b-', label='Polynomial')
# cubic spline interpolation. Interpolate to to xe using sampling points xi
cs = CubicSpline(xi,yi)
ycs = cs(xe)
plt.plot(xe,ycs,'g-', label='Cubic Spline')
# finalize plot
plt.legend()
plt.draw()
# + [markdown] slideshow={"slide_type": "slide"}
# More examples coming soon!
| exemplipy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# (file-types:notebooks)=
# # Jupyter Notebook files
#
# You can create content with Jupyter notebooks.
# For example, the content for the current page is contained in {download}`this notebook file <./notebooks.ipynb>`.
#
# ```{margin}
# If you'd like to write in plain-text files, but still keep a notebook structure, you can write
# Jupyter notebooks with MyST Markdown, which are then automatically converted to notebooks.
# See [](./myst-notebooks.md) for more details.
# ```
#
# Jupyter Book supports all Markdown that is supported by Jupyter Notebook.
# This is mostly a flavour of Markdown called [CommonMark Markdown](https://commonmark.org/) with minor modifications.
# For more information about writing Jupyter-flavoured Markdown in Jupyter Book, see [](./markdown.md).
#
# ## Code blocks and image outputs
#
# Jupyter Book will also embed your code blocks and output in your book.
# For example, here's some sample Matplotlib code:
from matplotlib import rcParams, cycler
import matplotlib.pyplot as plt
import numpy as np
plt.ion()
# + tags=["remove-stdout"]
# Fixing random state for reproducibility
np.random.seed(19680801)
N = 10
data = [np.logspace(0, 1, 100) + np.random.randn(100) + ii for ii in range(N)]
data = np.array(data).T
cmap = plt.cm.coolwarm
rcParams['axes.prop_cycle'] = cycler(color=cmap(np.linspace(0, 1, N)))
from matplotlib.lines import Line2D
custom_lines = [Line2D([0], [0], color=cmap(0.), lw=4),
Line2D([0], [0], color=cmap(.5), lw=4),
Line2D([0], [0], color=cmap(1.), lw=4)]
fig, ax = plt.subplots(figsize=(10, 5))
lines = ax.plot(data)
ax.legend(custom_lines, ['Cold', 'Medium', 'Hot']);
# -
# Note that the image above is captured and displayed in your site.
# + tags=["popout", "remove-input", "remove-stdout"]
# Fixing random state for reproducibility
np.random.seed(19680801)
N = 10
data = [np.logspace(0, 1, 100) + .1*np.random.randn(100) + ii for ii in range(N)]
data = np.array(data).T
cmap = plt.cm.coolwarm
rcParams['axes.prop_cycle'] = cycler(color=cmap(np.linspace(0, 1, N)))
from matplotlib.lines import Line2D
custom_lines = [Line2D([0], [0], color=cmap(0.), lw=4),
Line2D([0], [0], color=cmap(.5), lw=4),
Line2D([0], [0], color=cmap(1.), lw=4)]
fig, ax = plt.subplots(figsize=(10, 5))
lines = ax.plot(data)
ax.legend(custom_lines, ['Cold', 'Medium', 'Hot'])
ax.set(title="Smoother linez")
# + [markdown] tags=["popout"]
# ```{margin} **You can also pop out content to the side!**
# For more information on how to do this,
# check out the {ref}`layout/sidebar` section.
# ```
# -
# ## Removing content before publishing
#
# You can also remove some content before publishing your book to the web.
# For reference, {download}`you can download the notebook content for this page <notebooks.ipynb>`.
# + tags=["remove-cell"]
thisvariable = "none of this should show up in the textbook"
fig, ax = plt.subplots()
x = np.random.randn(100)
y = np.random.randn(100)
ax.scatter(x, y, s=np.abs(x*100), c=x, cmap=plt.cm.coolwarm)
ax.text(0, .5, thisvariable, fontsize=20, transform=ax.transAxes)
ax.set_axis_off()
# -
# You can **remove only the code** so that images and other output still show up.
# + tags=["hide-input"]
thisvariable = "this plot *will* show up in the textbook."
fig, ax = plt.subplots()
x = np.random.randn(100)
y = np.random.randn(100)
ax.scatter(x, y, s=np.abs(x*100), c=x, cmap=plt.cm.coolwarm)
ax.text(0, .5, thisvariable, fontsize=20, transform=ax.transAxes)
ax.set_axis_off()
# -
# Which works well if you'd like to quickly display cell output without cluttering your content with code.
# This works for any cell output, like a Pandas DataFrame.
# + tags=["hide-input"]
import pandas as pd
pd.DataFrame([['hi', 'there'], ['this', 'is'], ['a', 'DataFrame']], columns=['Word A', 'Word B'])
# -
# See {ref}`hiding/remove-content` for more information about hiding and removing content.
# ## Interactive outputs
#
# We can do the same for *interactive* material. Below we'll display a map
# using [folium](https://python-visualization.github.io/folium/). When your book is built,
# the code for creating the interactive map is retained.
#
# ```{margin}
# **This will only work for some packages.** They need to be able to output standalone
# HTML/Javascript, and not
# depend on an underlying Python kernel to work.
# ```
# +
import folium
m = folium.Map(
location=[45.372, -121.6972],
zoom_start=12,
tiles='Stamen Terrain'
)
folium.Marker(
location=[45.3288, -121.6625],
popup='Mt. Hood Meadows',
icon=folium.Icon(icon='cloud')
).add_to(m)
folium.Marker(
location=[45.3311, -121.7113],
popup='Timberline Lodge',
icon=folium.Icon(color='green')
).add_to(m)
folium.Marker(
location=[45.3300, -121.6823],
popup='Some Other Location',
icon=folium.Icon(color='red', icon='info-sign')
).add_to(m)
m
# -
# ## Rich outputs from notebook cells
# Because notebooks have rich text outputs, you can store these in
# your Jupyter Book as well! For example, here is the command line help
# menu, see how it is nicely formatted.
# !jupyter-book build --help
# And here is an error. You can mark notebook cells as "expected to error" by adding a
# `raises-exception` tag to them.
# + tags=["raises-exception"]
this_will_error
# -
# ## More features with Jupyter notebooks
#
# There are many other features of Jupyter notebooks to take advantage of,
# such as automatically generating Binder links for notebooks or connecting your content with a kernel in the cloud.
# For more information browse the pages in this site, and [](content:code-outputs) in particular.
| docs/file-types/notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Continuous Control
#
# ---
#
# In this notebook, you will learn how to use the Unity ML-Agents environment for the second project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.
#
# ### 1. Start the Environment
#
# We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
from unityagents import UnityEnvironment
import numpy as np
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/Reacher.app"`
# - **Windows** (x86): `"path/to/Reacher_Windows_x86/Reacher.exe"`
# - **Windows** (x86_64): `"path/to/Reacher_Windows_x86_64/Reacher.exe"`
# - **Linux** (x86): `"path/to/Reacher_Linux/Reacher.x86"`
# - **Linux** (x86_64): `"path/to/Reacher_Linux/Reacher.x86_64"`
# - **Linux** (x86, headless): `"path/to/Reacher_Linux_NoVis/Reacher.x86"`
# - **Linux** (x86_64, headless): `"path/to/Reacher_Linux_NoVis/Reacher.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `Reacher.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="Reacher.app")
# ```
env = UnityEnvironment(file_name='Reacher.app')
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 2. Examine the State and Action Spaces
#
# In this environment, a double-jointed arm can move to target locations. A reward of `+0.1` is provided for each step that the agent's hand is in the goal location. Thus, the goal of your agent is to maintain its position at the target location for as many time steps as possible.
#
# The observation space consists of `33` variables corresponding to position, rotation, velocity, and angular velocities of the arm. Each action is a vector with four numbers, corresponding to torque applicable to two joints. Every entry in the action vector must be a number between `-1` and `1`.
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
# -
# ### 4. It's Your Turn!
#
# Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
# ```python
# env_info = env.reset(train_mode=True)[brain_name]
# ```
# #### Algorithm to Use: DDPG
# We have just learned Deep Deterministic Policy Gradients (DDPG), a actor-critic algorithm or a DQN method for continuous action space. The actor approximates the optimal policy deterministically and output the best believed action for any given state. The critic learns the optimal value function by using the actors best believed action. There are two main features of DDPG:
#
# - experience reply buffer which refers to learn from the previous experience (memory), as in DQN. We set a Reply buffer with fixed size (BUFFER_SIZE) and stored the experiences in the buffer. When updating the parameters, we sample a few (batch_size) memory from the buffer. In this way, we can break the sequential nature of experiences and stabilize the learning algorithm.
# - soft updates to the target networks
#
# Note: there are 4 neural networks:
# - two regular networks (like the local/evaluate network in DQN whose parameters update all the time): one for the actor and one for the critic
# - two target networks (like the target network in DQN whose parameters update after a certain steps): one for the actor and one for the critic
# The target networks update using a soft updates strategy, e.g, tau = 0.01 which means when updating the target network $\theta_{target}$,
# $$\theta_{target} = \tau \cdot \theta_{local} + (1 - \tau)\cdot \theta_{target} $$
# Here, $\tau \leq 1$ and $\theta_{target}$ can be for either actor target network or critic target network.
# #### Extended ides
#
# When solving the first project: Banana, I have tried to use prioritized experience reply with different DQN based methods. The results showed that with prioritized experience reply, less episodes are needed for solving a problem. I would like also to integrate prioritized experience reply with DDPG and compare it to DDPG without prioritized experience reply.
#
# Note: with prioritized experience reply in DQN, we selects the sample according to the priorities (https://arxiv.org/pdf/1511.05952.pdf). Samples drawn from the buffer are fed into the DQN algorithm before getting priorities based on the magnitude of TD error. There are two additional hyperparameters $\alpha$ and $\beta$ for controlling how much prioritized experience replay affects the sampling distribution and network parameter updates.
#
# #### Define DDPG run pipeline
# +
# import
from collections import deque
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
# from ddpg_agent import Agent
from ddpg_agent import Agent
from torchsummary import summary
import time
plt.ion()
# -
def ddpg_singe_agent(n_episodes=2000, max_t = 1000, window_size=100, score_threshold=30.0,
print_interval=10, epochs=1000):
scores_deque = deque(maxlen=window_size)
scores = []
best_average_score = -np.inf
print("Training on {} started...".format(agent.device))
for i_episode in range(1, epochs+1):
env_info = env.reset(train_mode=True)[brain_name]
states = env_info.vector_observations
agent.reset()
episode_scores = np.zeros(num_agents)
for t in range(max_t):
actions = agent.act(states)
env_info = env.step(actions)[brain_name]
next_states = env_info.vector_observations
rewards = env_info.rewards
dones = env_info.local_done
agent.step(states=states, actions=actions, rewards=rewards, next_states=next_states, dones=dones)
episode_scores += np.array(rewards)
states = next_states
if np.any(dones):
break
episode_score = np.mean(episode_scores) # Summary of scores for this episode
scores_deque.append(episode_score)
scores.append(episode_score)
average_score = np.mean(scores_deque)
print('\rEpisode: {}\tAverage Score: {:.2f}\tCurrent Score: {:.2f}'.format(i_episode, average_score, episode_score), end="")
if i_episode % print_interval == 0:
print('\rEpisode: {}\tAverage Score: {:.2f}\tCurrent Score: {:.2f}'.format(i_episode, average_score, episode_score))
if average_score >= score_threshold:
print('\nEnvironment solved in {} episodes!\tAverage Score: {:.2f}'.format(i_episode-window_size, average_score))
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
break
np.save('scores.npy', scores)
return scores
# ### Watch an trained agnet
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))
# When finished, close the environment.
env.close()
| p2_continuous-control/Continuous_Control_Single_Agent_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="CazISR8X_HUG"
# # Multiple Linear Regression
# + [markdown] colab_type="text" id="pOyqYHTk_Q57"
# ## Importing the libraries
# -
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] colab_type="text" id="vgC61-ah_WIz"
# ## Importing the dataset
# -
dataset = pd.read_csv('50_Startups.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
print(X[:10])
print(y[:10])
# + [markdown] colab_type="text" id="VadrvE7s_lS9"
# ## Encoding categorical data
# -
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
ct = ColumnTransformer(transformers=[('encoder', OneHotEncoder(), [3])] , remainder='passthrough')
X = np.array(ct.fit_transform(X))
print(X[:10])
# + [markdown] colab_type="text" id="WemVnqgeA70k"
# ## Splitting the dataset into the Training set and Test set
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] colab_type="text" id="k-McZVsQBINc"
# ## Training the Multiple Linear Regression model on the Training set
# -
# sklearn.linear_model handles:
# - the dummy variable trap (dropping the last/dependent dummy variable for each set)
# - the selection of the most statisticly significant features (stepwise regression)
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
# + [markdown] colab_type="text" id="xNkXL1YQBiBT"
# ## Predicting the Test set results
# -
y_pred = regressor.predict(X_test)
np.set_printoptions(precision=2)
# evaluate performance by comparing the predicted profits and the real profits
print(np.concatenate((y_pred.reshape(len(y_pred), 1), y_test.reshape(len(y_test), 1)), axis=1))
# ## Making a single prediction (for example the profit of a startup with R&D Spend = 160000, Administration Spend = 130000, Marketing Spend = 300000 and State = 'California')
print(regressor.predict([[1, 0, 0, 160000, 130000, 300000]]))
# Therefore, our model predicts that the profit of a Californian startup which spent 160000 in R&D, 130000 in Administration and 300000 in Marketing is $ 181566,92.
#
# **Important note 1:** Notice that the values of the features were all input in a double pair of square brackets. That's because the "predict" method always expects a 2D array as the format of its inputs. And putting our values into a double pair of square brackets makes the input exactly a 2D array. Simply put:
#
# $1, 0, 0, 160000, 130000, 300000 \rightarrow \textrm{scalars}$
#
# $[1, 0, 0, 160000, 130000, 300000] \rightarrow \textrm{1D array}$
#
# $[[1, 0, 0, 160000, 130000, 300000]] \rightarrow \textrm{2D array}$
#
# **Important note 2:** Notice also that the "California" state was not input as a string in the last column but as "1, 0, 0" in the first three columns. That's because of course the predict method expects the one-hot-encoded values of the state, and as we see in the second row of the matrix of features X, "California" was encoded as "1, 0, 0". And be careful to include these values in the first three columns, not the last three ones, because the dummy variables are always created in the first columns.
# ## Getting the final linear regression equation with the values of the coefficients
print(regressor.coef_)
print(regressor.intercept_)
# Therefore, the equation of our multiple linear regression model is:
#
# $$\textrm{Profit} = 86.6 \times \textrm{Dummy State 1} - 873 \times \textrm{Dummy State 2} + 786 \times \textrm{Dummy State 3} + 0.773 \times \textrm{R&D Spend} + 0.0329 \times \textrm{Administration} + 0.0366 \times \textrm{Marketing Spend} + 42467.53$$
#
# **Important Note:** To get these coefficients we called the "coef_" and "intercept_" attributes from our regressor object. Attributes in Python are different than methods and usually return a simple value or an array of values.
| regression/multiple_linear_regression/my_multiple_linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
# # Maximising the utility of an Open Address
#
# <NAME> (GeoLytics), <NAME> (UU), <NAME> (UU), <NAME> (UU), <NAME> (Beare Essentials)
#
#
# 
#
#
# Go down for licence and other metadata about this presentation
# + [markdown] nbpresent={"id": "d4771fb7-04a1-4096-854f-0997ebe4dd8b"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "470f5f91-3b52-4171-a749-c2e589cfc338"} slideshow={"slide_type": "slide"}
# # The view of addressing from United Utilities
#
# Unless states otherwise all content is under a CC-BY licence
#
# 
#
#
# You can access this presentation on github:
#
# [https://github.com/AntArch/20150305_AddressDay.git](https://github.com/AntArch/20150305_AddressDay.git)
#
#
# 
#
# + [markdown] nbpresent={"id": "d7b93b9e-0316-40d2-9ad5-adab3fb37747"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "1057f676-f0de-4297-b8db-ad2d4eb9c665"} slideshow={"slide_type": "subslide"}
# ## Using Ipython for presentations
#
# A short video showing how to use Ipython for presentations
# + nbpresent={"id": "8fc4a806-4783-4034-b936-88d1b4f87ee5"}
from IPython.display import YouTubeVideo
YouTubeVideo('F4rFuIb1Ie4')
# + nbpresent={"id": "23f3b29b-34f8-4c52-a35b-bfb62ada785f"}
## PDF output using pandoc
import os
### Export this notebook as markdown
commandLineSyntax = 'ipython nbconvert --to markdown 201609_UtilityAddresses_Presentation.ipynb'
print (commandLineSyntax)
os.system(commandLineSyntax)
### Export this notebook and the document header as PDF using Pandoc
commandLineSyntax = 'pandoc -f markdown -t latex -N -V geometry:margin=1in DocumentHeader.md 201609_UtilityAddresses_Presentation.md --filter pandoc-citeproc --latex-engine=xelatex --toc -o interim.pdf '
os.system(commandLineSyntax)
### Remove cruft from the pdf
commandLineSyntax = 'pdftk interim.pdf cat 1-5 18-end output 201609_UtilityAddresses_Presentation.pdf'
os.system(commandLineSyntax)
### Remove the interim pdf
commandLineSyntax = 'rm interim.pdf'
os.system(commandLineSyntax)
# + [markdown] nbpresent={"id": "fc648d92-f376-4423-82df-f1f70cff02ef"} slideshow={"slide_type": "subslide"}
# ## The environment
#
# In order to replicate my environment you need to know what I have installed!
# + [markdown] nbpresent={"id": "fd6428a6-3f0c-4df4-a019-ca745b03f5ae"} slideshow={"slide_type": "skip"}
# ### Set up watermark
#
# This describes the versions of software used during the creation.
#
# Please note that critical libraries can also be watermarked as follows:
#
# ```python
# # # %watermark -v -m -p numpy,scipy
# ```
# + nbpresent={"id": "f28ecdcb-c05c-4492-8e61-dbf913814ab9"} slideshow={"slide_type": "skip"}
# %install_ext https://raw.githubusercontent.com/rasbt/python_reference/master/ipython_magic/watermark.py
# %load_ext watermark
# + nbpresent={"id": "2bac0c6b-93cc-40bf-8a5f-0ebe2cd901b9"}
# %watermark -a "<NAME>" -d -v -m -g
# + nbpresent={"id": "28c39680-76cc-4ea5-8654-d596ed6f7872"}
#List of installed conda packages
# !conda list
# + nbpresent={"id": "664d13f2-be01-4a90-8e70-90f6f991cf23"}
#List of installed pip packages
# !pip list
# + [markdown] nbpresent={"id": "30ba2b86-2580-4f33-a5cf-9c0a67c346b6"} slideshow={"slide_type": "subslide"}
# ## Running dynamic presentations
#
# You need to install the [RISE Ipython Library](https://github.com/damianavila/RISE) from [<NAME>](https://github.com/damianavila) for dynamic presentations
# + [markdown] nbpresent={"id": "4aaf6362-e9db-42b4-b981-0188500a2308"} slideshow={"slide_type": "slide"}
# To convert and run this as a static presentation run the following command:
# + nbpresent={"id": "6d38b76f-f2ed-40f3-8b06-a77f0d3d950b"}
# Notes don't show in a python3 environment
# !jupyter nbconvert 201609_UtilityAddresses_Presentation.ipynb --to slides --post serve
# + [markdown] nbpresent={"id": "54794ad2-9588-4ac2-9a8e-2747d7038cc7"}
# To close this instances press *control 'c'* in the *ipython notebook* terminal console
#
# Static presentations allow the presenter to see *speakers notes* (use the 's' key)
#
# If running dynamically run the scripts below
# + [markdown] nbpresent={"id": "bf640cff-9590-4cad-a769-6b2f153276c5"} slideshow={"slide_type": "subslide"}
# ## Pre load some useful libraries
# + nbpresent={"id": "39d75d5d-063a-427a-982e-6d86bd7285d4"} slideshow={"slide_type": "-"}
#Future proof python 2
from __future__ import print_function #For python3 print syntax
from __future__ import division
# def
import IPython.core.display
# A function to collect user input - ipynb_input(varname='username', prompt='What is your username')
def ipynb_input(varname, prompt=''):
"""Prompt user for input and assign string val to given variable name."""
js_code = ("""
var value = prompt("{prompt}","");
var py_code = "{varname} = '" + value + "'";
IPython.notebook.kernel.execute(py_code);
""").format(prompt=prompt, varname=varname)
return IPython.core.display.Javascript(js_code)
# inline
# %pylab inline
# + [markdown] nbpresent={"id": "04d308c3-3f5b-417b-969d-af88640e6934"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "628a2fab-6117-451c-86bf-112808eb53c9"} slideshow={"slide_type": "slide"}
# ## About me
#
#
#
# 
#
# * Honorary Research Fellow, University of Nottingham: [orcid](http://orcid.org/0000-0002-2991-811X)
# * Director, Geolytics Limited - A spatial data analytics consultancy
#
# ## About this presentation
#
# * [Available on GitHub](https://github.com/AntArch/Presentations_Github/tree/master/20151008_OpenGeo_Reuse_under_licence) - https://github.com/AntArch/Presentations_Github/
# * [Fully referenced PDF](https://github.com/AntArch/Presentations_Github/blob/master/201609_UtilityAddresses_Presentation/201609_UtilityAddresses_Presentation.pdf)
#
# + [markdown] nbpresent={"id": "2887ec79-f276-42ec-844e-1724a84a363a"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # Addresses support:
#
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## everyday life
#
# 
#
# They are part of the fabric of everyday life
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Economy and commerce
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Governance
#
# 
#
# * Without an address, it is harder for individuals to register as legal residents.
# * They are *not citizens* and are excluded from:
# * public services
# * formal institutions.
# * This impacts on democracy.
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Urban Development
#
# 
#
# * Key to managing *the explosion* of rural to urban migration.
# * Informal settlements housing the urban poor.
# * Poor infrastructure services.
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Legal and Social integration
#
# 
#
# * Formal versus Informal
# * Barring individuals and businesses from systems:
# * financial
# * legal
# * government
# * ....
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Sustainability and risk management
#
# 
#
# * Addresses, geodemographics and spatial infrastructure support
# * sustainability
# * resilience
# * disaster management
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Global Wellbeing
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Addresses bridge gaps
#
# Addresses provide the link between ***people*** and ***place***
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # Utility Addresses
#
#
# ## In the beginning ...... was the ledger
#
# 
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Bespoke digital addresses
#
# * Digitisation and data entry to create a bespoke Address Database -
# * Fit for UU's operational purpose
# * Making utilities a key *owner* of address data
# * Subject to IP against PAF matching
#
# 
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Policy mandates
#
# Open Water - A shared view of addresses requiring a new addressing paradigm - Address Base Premium?
#
# 
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
#
# # Utility addressing:
#
#
#
# * Postal delivery (Billing)
# * Services and Billing to properties within the extent of the UU operational area
# * Billing to customers outside the extent of UU operational area
# * Asset/Facilities Management (infrastructure)
# * Premises
# * But utilties manage different assets to Local Authorities
# * is an address the best way to manage a geo-located asset?
# * Bill calculation
# * Cross referencing Vaulation Office and other detals.
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
#
#
# . . . .
#
# **It's not just postal addressing**
#
# . . . .
#
# **Address credibility is critical**
#
# . . . .
#
# Utilities see the full life-cycle of an address - especially the birth and death
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## asset management
#
# * UU manage assets and facilities
#
# > According to ABP a Waste Water facility is neither a postal address or a non-postal address.
#
# Really? Is it having an existential crisis?
#
# 
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## A connected spatial network
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Serving customers who operate **somewhere**
#
# 
#
# * UU serve customers located in
# * Buildings
# * Factories
# * Houses
# * Fields
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Serving customers who operate **anywhere**
#
# 
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
# # Utility addressing issues
#
#
# * Addresses are a pain
# * Assets as locations
# * Services as locatons
# * People at locations
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
# # Issues: When most people in the UK think of addresses they think of a postal address.
#
# * Is *Postal* a constraining legacy?
# * Is *address* a useful term?
#
# 
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# # Issues: Do formal *addresses* actually help utilities?
#
# * External addresses (ABP for example) are another product(s) to manage
# * which may not fit the real business need
# * which may not have full customer or geographic coverage
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
#
# # What is an address?
#
# ## Address abstraction
#
# * Address did not spring fully formed into existance.
# * They are used globally
# * but developed nationally
# * and for different reasons
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Royal Mail - postal delivery
#
# 
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# In a postal system:
#
# * a *Delivery Point* (DP) is a single mailbox or other place at which mail is delivered.
# * a single DP may be associated with multiple addresses
# * An *Access Point* provides logistical detail.
#
# The postal challenge is to solve the last 100 meters. In such a scenario the *post person* is critical.
#
# DPs were collected by the Royal Mail for their operational activities and sold under licence as the *Postal Address File* (PAF). PAF is built around the 8-character *Unique Delivery Point Reference Number* (UDPRN). The problem with PAF is that the spatial context is not incorporated into the product. Delivery points are decoupled from their spatial context - a delivery point with a spatial context should provide the clear location of the point of delivery (a door in a house, a post-room at an office etc.).
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## LLPG - asset management
#
# ](https://dl.dropboxusercontent.com/u/393477/ImageBank/ForAddresses/LBH-LLPG-System-flow-Diagram.png)
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# An LLPG (Local Land and Property Gazetteer) is a collection of address and location data created by a local authority.
#
# It is an Asset/Facilities Management tool to support public service delivery:
#
# * Local Authority
# * Police
# * Fire
# * Ambulance
#
# It incorporates:
#
# * Non postal addresses (i.e. something that the Royal Mail wouldn't deliver post to)
#
# * a 12-digit Unique Property Reference Number for every building and plot of land
# * National Street Gazetteer
#
# Prior to the initialization of the LLPGs, local authorities would have different address data held across different departments and the purpose of the Local Land and Property Gazetteers was to rationalize the data, so that a property or a particular plot of land is referred to as the same thing, even if they do have different names.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses as assets?
#
# ](http://joncruddas.org.uk/sites/joncruddas.org.uk/files/styles/large/public/field/image/post-box.jpg?itok=ECnzLyhZ)
#
# * So what makes the following 'non-postal' *facilities* addresses:
# * Chimney
# * Post box - which is clearly having a letter delivered ;-)
# * Electricity sub-station
# * Public Telephone
# * Tennis Courts
# * Context is critical
# * So why is a waste-water facility not an address in ABP?
# * Because it is not *of interest* to a council and the Royal Mail have never been asked to deliver mail to it.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Korea: The Jibeon system - taxation
#
# 
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# * Until recently, the Republic of Korea (Korea) has used land parcel numbers ( jibeon) to identify unique locations.
# * These parcel numbers were assigned chronologically according to date of construction and without reference to the street where they were located.
# * This meant that adjacent buildings did not necessarily follow a sequential numbering system.
# * This system was initially used to identify land for census purposes and to levy taxes.
# * In addition, until the launch of the new addressing system, the jibeon was also used to identify locations (i.e. a physical address).
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## World Bank - social improvement
#
# 
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# The World Bank has taken a *street addressing* view-point (@_addressing_2012, p.57). This requires up-to-date mapping and bureacracy (to deliver a street gazetteer and to provide the street infrastructure (furniture)). However, (@_addressing_2012, p.44) demonstrates that this is a cumbersome process with a number of issues, not the least:
#
# * Urban bias
# * Cost of infrastucture development
# * Lack of community involvment
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Denmark: An addressing commons with impact
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Denmark: An addressing commons with impact
#
# * Geocoded address infrastructure
# * Defined the semantics of purpose
# * what is an address
# * Open data
# * an address commons
# * The re-use statistics are staggering:
# * 70% of deliveries are to the private sector,
# * 20% are to central government
# * 10% are to municipalities.
# * Benefits:
# * Efficiencies
# * No duplication
# * Improved confidence
# * Known quality
#
# A credible service providing a mutlitude of efficiencies (@_addressing_2012, pp.50 - 54)
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
#
# # UK Addressing
#
# ## Geoplace - Formal
#
# ](https://www.geoplace.co.uk/documents/10181/67776/NAG+infographic/835d83a5-e2d8-4a26-bc95-c857b315490a?t=1434370410424)
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# * GeoPlace is a limited liability partnership owned equally by the [Local Government Association](http://www.local.gov.uk/) and [Ordnance Survey](http://www.ordnancesurvey.co.uk/).
# * It has built a synchronised database containing spatial address data from
# * 348 local authorities in England and Wales (the *Local Land and Property Gazetteers* (LLPG) which cumulatively create the *National Land and Property Gazetteer* (NLPG)),
# * Royal Mail,
# * Valuation Office Agency and
# * Ordnance Survey datasets.
# * The NAG Hub database is owned by GeoPlace and is the authoritative single source of government-owned national spatial address information, containing over 225 million data records relating to about 34 million address features. GeoPlace is a production organisation with no product sales or supply operations.
# * The NAG is made available to public and private sector customers through Ordnance Survey’s [AddressBase](http://www.ordnancesurvey.co.uk/business-and-government/products/addressbase.html) products.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## The AddressBase Family
#
# 
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# * The National Address Gazetteer Hub database is owned by GeoPlace and is claimed to be *the authoritative single source of government-owned national spatial address information*, containing over 225 million data records relating to about 34 million address features.
# * Each address has its own *Unique Property Reference Number* (UPRN). The AddressBase suite have been designed to integrate into the [Ordnance Survey MasterMap suite of products](http://www.ordnancesurvey.co.uk/business-and-government/products/mastermap-products.html).
#
# AddressBase is available at three levels of granularity (lite, plus and premium).
#
# * AB+ merges two address datasets together (PAF and Local Authority) to provide the best available view of addresses currently defined by Local Authorities, giving many advantages over AddressBase.
# * AB+ lets you link additional information to a single address, place it on a map, and carry out spatial analysis that enables improved business practices.
# * Geoplace argue that further value comes from additional information in the product which includes:
# * A more detailed classification – allowing a better understanding of the type (e.g. Domestic, Commercial or Mixed) and function of a property (e.g. Bank or Restaurant)
# * Local Authority addresses not contained within PAF – giving a more complete picture of the current addresses and properties (assuming they are in scope (see below))
# * Cross reference to OS MasterMap TOIDs – allowing simple matching to OS MasterMap Address Layer 2, Integrated Transport Network or Topography Layer
# * Spatial coordinates
# * Unique Property Reference Number (UPRN) – which provides the ability to cross reference data with other organisations, and maintain data integrity.
# * Premium includes the address lifecycle
#
#
# AddressBase supports the UK Location Strategy concept of a 'core reference geography', including the key principles of the European Union INSPIRE directive, that data should only be collected once and kept where it can be maintained most effectively (see [AddressBase products user guide](http://www.ordnancesurvey.co.uk/docs/user-guides/addressbase-products-user-guide.pdf)). *It's probably worthwhile mentioning that this is not an open address layer - however, a [2104 feasibility study sponsored by the department of Business, Innovation and Skills](https://www.gov.uk/government/publications/an-open-national-address-gazetteer) included a recommendation that AddressBase lite is made openly available*.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Address lifecycle
#
# ](https://dl.dropboxusercontent.com/u/393477/ImageBank/ForAddresses/ABP_Lifecycle.png)
#
#
# + [markdown] slideshow={"slide_type": "notes"}
# * This ability to maintain an overview of the lifecycle of address and property status means the AddressBase Premium has introduced new potential use cases.
# * This has seen companies incorporating AddressBase Premium into their business systems to replace PAF or bespoke addressing frameworks - in theory the ability to authoritatively access the address lifecycle provides greater certainty for a number of business operations.
#
# * At *United Utilites* (UU) AddressBase Premium is replacing a multitude of bespoke and PAF based addressing products.
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## [Open National Address Gazetteer](https://www.gov.uk/government/publications/an-open-national-address-gazetteer) - *informal?*
#
# The *Department for Business, Innovation & Skills* (BIS) on the need for an [Open National Address Gazetteer](https://www.gov.uk/government/publications/an-open-national-address-gazetteer) commissioned a review of *open addressing* which was published January 2014.
#
# . . . . .
#
# It recommended:
#
# * the UK Government commission an 'open' addressing product based on a variation of the 'freemium' model
# * data owners elect to release a basic ('Lite') product as Open Data that leaves higher value products to be licensed
#
# . . . . .
#
# AddressBase Lite was proposed with an annual release cycle. Critically this contains the UPRN which could be be key for product interoperability.
# * This would allow the creation of a shared interoperable address spine along the lines of the Denmark model
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Open NAG - [*'Responses received'*](https://www.gov.uk/government/publications/an-open-national-address-gazetteer) April 2014
#
# With the exception of the PAF advisory board and Royal Mail there was support for the BIS review across the respondants with some notable calls for the *Totally Open* option (particularly from those organisations who are not part of the Public Sector Mapping Agreement) and that the UPRN should be released under an open data licence (as a core reference data set that encourages product interoperability).
#
# . . . . .
#
# A number of quotes have been selected below:
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses as an Open Core Reference
#
# >....Address data and specific locations attached to them **are part of Core Reference data sets recognised by government as a key component of our National Information Infrastructure** (as long argued by APPSI). The report published by BIS gives us **a chance to democratise access to addressing data** and meet many of the Government’s avowed intentions. We urge acceptance of Option 6 *(freemium)* or 7 *(an independent open data product)*.
#
# **<NAME> *Chair of the Advisory Panel on Public Sector Information* **
#
# >....**Freely available data are much more likely to be adopted** by users and embedded in operational systems. **A national register, free at the point of delivery will undoubtedly help in joining up services, increasing efficiency and reducing duplication**.
#
# **Office of National Statistics**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Monopoly rent exploitation
#
# >... we expressed concern that, for almost all other potential customers (non-public sector), **the prices are prohibitive**, and appear designed to protect OS’s existing policy of setting high prices for a small captive market, **extracting monopoly rent**.
#
# **<NAME> *Director, DUG* **
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## The benefit of current credible addresses
#
# >**The problem of out-of-date addresses is a very significant commercial cost** for the whole of the UK and is also arguably underplayed in the report.
#
# **Individual Respondent 3**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Licences
#
# >Whatever licence the data is available under, **it must permit the data to be combined with other open data and then re-published**. ... The [Open Government Licence](http://www.nationalarchives.gov.uk/doc/open-government-licence/version/2/) fulfils this criteria, but it should be noted that the [OS OpenData Licence](http://www.ordnancesurvey.co.uk/docs/licences/os-opendata-licence.pdf) (enforced by OS on it's OS OpenData products, and via the PSMA) does not. The use of the latter would represent a significant restriction on down-stream data use, and so should be avoided.
#
# **Individual Respondent 6**
#
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
# # Taking Stock
#
# ## Addresses are heterogeneous
#
# 
#
# In terms of:
#
# * What they mean
# * What they are used for
# * Who uses them
# * How they are accessed
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Assets can have addresses
#
# So - anything can have an address (the *Internet of Things*)
#
# ](http://joncruddas.org.uk/sites/joncruddas.org.uk/files/styles/large/public/field/image/post-box.jpg?itok=ECnzLyhZ)
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## National data silos
#
# 
#
# They have been created to solve national issues.
#
# No unifying semantics
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ##
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses are bureaucratic and costly
#
# 
#
# Severely protracted when formal/informal issues are encountered.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses can be opaque
#
# 
#
# **transparent and reproducible?**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses are of global significance
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses are ripe for disruption
#
# 
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
#
# # Address Disruption
#
# ## Formal versus informal
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Technology
#
# Streets are so last century.....
#
# 
#
# * Ubiquitous GPS/GNSS
# * Structured crowdsourced geo-enabled content (wikipedia, OSM)
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Interoperability
#
# 
#
# * Will the semantic web provide address interoperabilty?
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Globalisation
#
# 
#
# * Addressing is a **core reference geography**
# * Global brands will demand (or invoke) consistent global addressing
# * How will licences impact on this?
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# ## [Core reference geographies](http://www.slideshare.net/geocommunitylive/bob-barr-what-are-core-reference-geographies)
#
# <NAME> has described [core reference geographies](http://www.slideshare.net/geocommunitylive/bob-barr-what-are-core-reference-geographies) as geographic data which:
#
# * Are definitive
# * Should be collected and maintained once and used many times
# * Are Natural monopolies (which addresses are)
# * Have variable value in different applications
# * Have highly elastic demand
#
# **Global addresses are a core reference geography.**
#
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
# # A new global address paradigm?
#
# * [Amazon drone delivery in the UK requires](https://www.theguardian.com/technology/2016/jul/25/amazon-to-test-drone-delivery-uk-government)
# * A new view over addressing complements streets and buildings but is geo-coded at source
# * and supports accurate delivery throughout the delivery chain using a global referencing system.
#
# Is there a universal approach which allows all avenues to be satisfied?
#
# 
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## How might this look?
#
# .
# .
#
# Requirements for a Global Address Framework
#
# .
# .
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## WGS84 algorithmic address minting
#
# 
#
# **A global addressing framework needs to be transparent and reproducible.**
#
# **A global addressing framework should be based on a spatial reference system.**
#
# **A global addressing framework needs to be lightweight and cheap so it can be implemented in a timely manner.**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Small footprint
#
# 
#
# **Ubiquitous access across platforms.**
#
# **No dependency on internet access.**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Short/memorable
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Self checking
#
# 
#
# **Improving validity and credibility of downstream business processes.**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Unlimited spatial recording
#
# 
#
# * What are the spatial requirements for the range of addressing options?
# * [Manila has a population density of 42,857 people per square km](http://en.wikipedia.org/wiki/List_of_cities_proper_by_population_density).
# * [Map Kibera](http://mapkibera.org/) has revolutionised services in Kibera (Kenya). Address Kibera could do the same thing for citizenship.
#
# **A global addressing framework should meet the needs of the rural, urban, formal and informal communities equally.**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Open and interoperable
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Open and interoperable
#
# > the lack of a consistent and transparent legal and policy framework for sharing spatial data continues to be an additional roadblock.
#
# @pomfret_spatial_2010
#
# **A global addressing framework should be open or available with as few barriers as possible.**
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Indoor use and 3D
#
# 
#
# Incorporating wifi-triangulation - *individual room* addressing and navigation.
#
# Seamless integration with BIM and CityGML.
#
# *Addressing isn't only about buildings - think about the Internet of Things*
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Inherent geo-statistical aggregation (spatially scalable)
#
# 
#
# GIS free multi-scale analysis and reporting during disaster scenarios.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Technology benchmarking
#
# BCS examples (in alphabetical order):
#
# * [GeoHash](http://en.wikipedia.org/wiki/Geohash)
# * gcpvj1r2vnbp
# * [Maidenhead Locator System](http://en.wikipedia.org/wiki/Maidenhead_Locator_System)
# * IO91wm (it has a very large footprint)
# * [MapCode](http://www.mapcode.com/)
# * GBR JD.VJ
# * [Natural Area Code](http://nactag.info/map.asp)
# * 8KDB PGFD
# * [Pyxis](http://www.pyxisinnovation.com/)
# * [What3Words](http://what3words.com/)
# * move slam stress
#
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # Utility address concepts
#
#
# * A means of communicating location to third parties in a way **they** understand.
# * Delivery
# * Contract engineer
# * Incident reporting
# * Hence, addresses are all about sharing
# * We need to *buy into* disambiguating stakeholder semantics
# * Democratise the infrastructure
# * Democratise re-use
# * Everything is mediated by a human in the information exchange
# * Everyone has their own semantics
# * Formal and vernacular geographies
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Addresses mediate space
#
# In business systems addresses are bridge a between technology stacks and social systems.
#
#
#
# 
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Addresses mediate space
#
# In business systems addresses are bridge a between technology stacks and social systems.
#
#
#
# 
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# * Most people in the UK think of an address as a *postal address*
# * This is a mindset we should be trying to break
# * A delivery address is only one facet to an address
# * What do addresses enable
# * Services
# * Postal services
# * Utility services
# * etc
# * Routing
# * Vehicle navigation
# * People navigation
# * Asset/Infrastructure management
# * Information integration
# * Lifecycle
# * Geodemographics
# * Hence, addressing information covers a range of requirements:
# * Semantic
# * GIS
# * Database
# * Challenges
# * find an unambiguous way to encode these different address types across the enterprise (and/or as part of an open initiative)
# * find ways to dynamically transform these address so that each end-user community get the most appropriate address be they:
# * formal addresses
# * vernacular (informal) addresses
# * Postal address
# * Asset location
#
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# In terms of assets two things spring to mind -
#
# 1. we no longer need streets and buildings to provide an address.
# * GNSS already does this globally.
# * The challenge is to translate GNSS into something appropriate for other services
# 1. The Access point/Delivery point metaphor used by Royal Mail may be important for traction
# * solving the last 100m problem (or location of local drone delivery depot)
#
# 
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "slide"}
#
# # Current utility addressing?
#
# ## A shared view over addressing?
#
# 
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## A shared view over addressing?
#
# Not really....
#
# * ABP isn’t a silver bullet
# * Subset of required ‘formal - delivery’ addresses
# * Mis-match in terms of assets
# * Why does a sewage works not have an address when a post-box does?
# * Not plug and play
# * Lag in the system - the lifecycle feedback does not have credibility for time critical applications.
# * The co-ordinating spine is not freely available (under a permissive licence)
# * Inset areas - an aglomoration of 'addresses'
# * VOA link is a cludge
#
# 
#
# ## Addresses should mediate systems
#
# * Bridge the gap between a building focussed 2d view of the world and the 3d world in which we inhabit.
# * Harmonise the edge cases relationships between UPRNs and VOAs
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## Issues about ABP
#
# * Users over a barrel
# * Needed to buy ABP as AL2 was being removed
# * Data model change
# * a hostage to someone else's data model
# * Lifecycle benefit not being realised (at least not for utilities)
# * Altough utilities have a significant value add
# * Update frequency
# * Different view of property hierarchy
# * 2d and 3d metaphors against VOA
# * a better 2.5 view of the world would be appreciated.
#
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "subslide"}
# ## This begs the question
#
# As a key creator of addresses should utilities replace a functional bespoke address system with an address framework (ABP) that does not meet all their business requirements?
#
# This creates a paradox when products like AddressBase are stipulated in Government policy documents (such as OpenWater)
#
# How can this gap be bridged?
#
# **Addresses need to be fit-for-purpose for the end user**
#
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # Future Addressing
#
#
#
# ## What do Utilities need from an Open Address infrastructure
#
# > <NAME> will talk about how addresses are employed within United Utilities: from bespoke addressing, to the current implementation of Geoplace’s Address Base. The current approach to addressing hinders effective market activities so consideration is given to how Open approaches can disrupt the addressing landscape and improve utility services.
#
# * Should this simply emulate Address Base Premium?
# * No
# * Like Denmark should it exploit technological developments to be:
# * More robust
# * Improve use case
# * More flexible
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Future Addressing
#
#
#
# ## What do Utilities need from an Open Address infrastructure
#
# * Should it embrace technological development to make operational activities more efficient
# * Use disruptive technologies to facilitate geo-coded addressing of assets in a flexible and credible manner
# * How can such an infrastructure interoperate with other formal and informal sources to provide benefits
# * What licence would a service be under.
# * OS? -No
# * The point is to encourage:
# * adoption
# * engagement
# * re-use
#
# > We would like to see any *open addressing infrastructure* in the OK **not simply aim to emulate ABP** but instead **provide a platform for 21st Century addressing on a global platform**
#
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## What can Utilities bring to Open Addresses
#
# * A credible publisher of addressing updates under open licences providing:
# * additional content
# * improved lifecycle information
# * Critical lifecycle data updates
# * potentially faster than local government.
#
# The address lifecycle element helps UU provide operational capacity for new builds and provides greater confidence when changing asset GIS and client details when a property is demolished.
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] nbpresent={"id": "b2ca43ef-2e99-479c-b47d-9e7a8ede9572"} slideshow={"slide_type": "subslide"}
# ## What can Open Addresses bring to Utilities
#
# * Fill the gap of formal addresses
# * But share a common reference Spine
# * UPRN?
# * But what about the 3d world
# * Add value
# * Embed a complementary geoaddressing paradigm
# * Linked data?
# * Property life-cycle?
# * Spatially consistent
# * Crowd enhanced
# * Service innovation
# * enhanced business intelligence from shared knowledge
# * geo-demographics protecting the disenfranchised
# * who are our sensitive customers - what are their needs?
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # Final thoughts
#
# Should an Open Address infrastructure emulate current models or should it be the foundation of a new addressing paradigm fit for 21st century challenges
#
# Utilities have the potential to be:
#
# * Key consumers of open addressing data
# * Key providers of open addressing content
#
# **United Utilities would like to help frame this debate and be part of any solution.**
# + [markdown] slideshow={"slide_type": "skip"}
# \newpage
# + [markdown] slideshow={"slide_type": "slide"}
# # References
# -
| 20190306_BCS_Scotland_Presentation/.ipynb_checkpoints/201609_UtilityAddresses_Presentation_Pre_Cull-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# TensorFlow GPU support document
#
# - GPU Guide: https://www.tensorflow.org/guide/gpu
# - Compatibility Matrix: https://www.tensorflow.org/install/source#gpu
# Get GPU status with Nvidia System Management Interface (nvidia-smi)
# Check driver version and CUDA version are compatible with TensorFlow
# !nvidia-smi
# Check the cuda toolkit version
# ! nvcc -V
# Get TensorFlow version and the number of GPUs visible to TensorFlow runtime
import tensorflow as tf
print("TensorFlow Version:", tf.__version__)
print("Num GPUs Available: ", len(tf.config.list_physical_devices('GPU')))
# +
# Run 'MatMul' with GPU
tf.debugging.set_log_device_placement(True)
# Create some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
| examples/notebook-examples/data/tensorflow2/1_advanced/tensorflow2_gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython import display
import os
def show_app(app, port = 9999,
width = 700,
height = 350,
offline = False,
in_binder = None):
in_binder ='JUPYTERHUB_SERVICE_PREFIX' in os.environ if in_binder is None else in_binder
if in_binder:
base_prefix = '{}proxy/{}/'.format(os.environ['JUPYTERHUB_SERVICE_PREFIX'], port)
url = 'https://hub.mybinder.org{}'.format(base_prefix)
app.config.requests_pathname_prefix = base_prefix
else:
url = 'http://localhost:%d' % port
iframe = '<a href="{url}" target="_new">Open in new window</a><hr><iframe src="{url}" width={width} height={height}></iframe>'.format(url = url,
width = width,
height = height)
display.display_html(iframe, raw = True)
if offline:
app.css.config.serve_locally = True
app.scripts.config.serve_locally = True
return app.run_server(debug=False, # needs to be false in Jupyter
host = '0.0.0.0',
port=port)
import dash
import dash_core_components as dcc
import dash_html_components as html
from plotly import graph_objs as go
from dash.dependencies import Input, Output
test_dash_app = dash.Dash(__name__, url_base_pathname='/', csrf_protect=False)
test_dash_app.layout = html.Div([dcc.RadioItems(id='item_list',
options = [dict(label = k, value = k) for k in ['Hey', 'Bob']]),
dcc.RadioItems(id='subitem_list', value = [])])
@test_dash_app.callback(
Output(component_id='subitem_list', component_property='options'),
[Input(component_id='item_list', component_property='value')]
)
def update_lesion_list(selected_idx):
return [{'label': '<img src="https://dummyimage.com/%i.jpg">hey</img>' % (100+i), 'value': i} for i, lab_name in enumerate('abcde')]
show_app(test_dash_app)
test_dash_app = dash.Dash(__name__, url_base_pathname='/', csrf_protect=False)
test_dash_app.layout = html.Div([dcc.RadioItems(id='item_list',
options = [dict(label = k, value = k) for k in ['Hey', 'Bob']]),
html.Div(id='button_list')])
@test_dash_app.callback(
Output(component_id='button_list', component_property='children'),
[Input(component_id='item_list', component_property='value')]
)
def update_button_list(selected_idx):
if selected_idx is not None:
return [html.Button('Hey %04d' % (i), id = 'id_%s_%04d' % (selected_idx, i)) for i in range(4)]
show_app(test_dash_app)
# +
test_dash_app = dash.Dash(__name__, url_base_pathname='/', csrf_protect=False)
test_dash_app.layout = html.Div([dcc.RadioItems(id='item_list',
options = [dict(label = k, value = k) for k in ['Hey', 'Bob']]),
html.Div(id='button_list')])
def fancy_button_adder(*args):
return [html.P(y) for arg in args]
@test_dash_app.callback(
Output(component_id='button_list', component_property='children'),
[Input(component_id='item_list', component_property='value')]
)
def update_button_list(selected_idx):
if selected_idx is not None:
out_id = 'div_%s' % selected_idx
out_obj_list = [html.Div('ClickOutputs',id = out_id)]
out_dep_obj = Output(component_id= 'click_msg', component_property='children')
in_dep_obj = []
for i in range(4):
c_id = 'id_%s_%04d' % (selected_idx, i)
out_obj_list += [html.Button('Hey %04d' % (i), id = c_id) ]
in_dep_obj += [Input(component_id=c_id, component_property='n_clicks')]
test_dash_app.callback(out_dep_obj, in_dep_obj)(fancy_button_adder)
return out_obj_list
#test_dash_app.config['suppress_callback_exceptions']=True
# -
show_app(test_dash_app)
# +
test_dash_app = dash.Dash(__name__, url_base_pathname='/', csrf_protect=False)
test_dash_app.layout = html.Div([dcc.RadioItems(id='item_list',
options = [dict(label = k, value = k) for k in ['Hey', 'Bob']]),
html.Div(id='button_list'),
dcc.Location(id='url', refresh=False),
html.Div('No Clicks', id = 'click_msg')])
@test_dash_app.callback(
Output(component_id='button_list', component_property='children'),
[Input(component_id='item_list', component_property='value')]
)
def update_button_list(selected_idx):
if selected_idx is not None:
return [html.Div([html.Br(),
dcc.Link('Hey %04d' % (i),
href = 'id_%s_%04d' % (selected_idx, i),
style = {'color': '#1EAEDB', 'text-decoration': 'underline',
'cursor': 'pointer'})]) for i in range(4)]
@test_dash_app.callback(
Output(component_id='click_msg', component_property='children'),
[Input(component_id='url', component_property='pathname')]
)
def update_click_msg(in_url):
if in_url != '/':
p_url = in_url.split('/')[-1]
return p_url
else:
return 'No Clicks'
# -
show_app(test_dash_app)
| notebooks/FancyRadioItems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this assignment, you'll implement an L-layered deep neural network and train it on the MNIST dataset. The MNIST dataset contains scanned images of handwritten digits, along with their correct classification labels (between 0-9). MNIST's name comes from the fact that it is a modified subset of two data sets collected by NIST, the United States' National Institute of Standards and Technology.<br>
# ## Data Preparation
# +
import numpy as np
import pickle
import gzip
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import h5py
import sklearn
import sklearn.datasets
import scipy
from PIL import Image
from scipy import ndimage
# %matplotlib inline
# -
# The MNIST dataset we use here is 'mnist.pkl.gz' which is divided into training, validation and test data. The following function <i> load_data() </i> unpacks the file and extracts the training, validation and test data.
def load_data():
f = gzip.open('mnist.pkl.gz', 'rb')
f.seek(0)
training_data, validation_data, test_data = pickle.load(f, encoding='latin1')
f.close()
return (training_data, validation_data, test_data)
# Let's see how the data looks:
training_data, validation_data, test_data = load_data()
training_data
# shape of data
print(training_data[0].shape)
print(training_data[1].shape)
print("The feature dataset is:" + str(training_data[0]))
print("The target dataset is:" + str(training_data[1]))
print("The number of examples in the training dataset is:" + str(len(training_data[0])))
print("The number of points in a single input is:" + str(len(training_data[0][1])))
# Now, as discussed earlier in the lectures, the target variable is converted to a one hot matrix. We use the function <i> one_hot </i> to convert the target dataset to one hot encoding.
def one_hot(j):
# input is the target dataset of shape (m,) where m is the number of data points
# returns a 2 dimensional array of shape (10, m) where each target value is converted to a one hot encoding
# Look at the next block of code for a better understanding of one hot encoding
n = j.shape[0]
new_array = np.zeros((10, n))
index = 0
for res in j:
new_array[res][index] = 1.0
index = index + 1
return new_array
data = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
print(data.shape)
one_hot(data)
# The following function data_wrapper() will convert the dataset into the desired shape and also convert the ground truth labels to one_hot matrix.
def data_wrapper():
tr_d, va_d, te_d = load_data()
training_inputs = np.array(tr_d[0][:]).T
training_results = np.array(tr_d[1][:])
train_set_y = one_hot(training_results)
validation_inputs = np.array(va_d[0][:]).T
validation_results = np.array(va_d[1][:])
validation_set_y = one_hot(validation_results)
test_inputs = np.array(te_d[0][:]).T
test_results = np.array(te_d[1][:])
test_set_y = one_hot(test_results)
return (training_inputs, train_set_y, test_inputs, test_set_y)
train_set_x, train_set_y, test_set_x, test_set_y = data_wrapper()
print ("train_set_x shape: " + str(train_set_x.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x shape: " + str(test_set_x.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
# We can see that the data_wrapper has converted the training and validation data into numpy array of desired shapes. Let's convert the actual labels into a dataframe to see if the one hot conversions are correct.
y = pd.DataFrame(train_set_y)
print("The target dataset is:" + str(training_data[1]))
print("The one hot encoding dataset is:")
y
# Now let us visualise the dataset. Feel free to change the index to see if the training data has been correctly tagged.
index = 1000
k = train_set_x[:,index]
k = k.reshape((28, 28))
plt.title('Label is {label}'.format(label= training_data[1][index]))
plt.imshow(k, cmap='gray')
# # Feedforward
# ### sigmoid
# This is one of the activation functions. It takes the cumulative input to the layer, the matrix **Z**, as the input. Upon application of the **`sigmoid`** function, the output matrix **H** is calculated. Also, **Z** is stored as the variable **sigmoid_memory** since it will be later used in backpropagation.You use _[np.exp()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.exp.html)_ here in the following way. The exponential gets applied to all the elements of Z.
def sigmoid(Z):
# Z is numpy array of shape (n, m) where n is number of neurons in the layer and m is the number of samples
# sigmoid_memory is stored as it is used later on in backpropagation
H = 1/(1+np.exp(-Z))
sigmoid_memory = Z
return H, sigmoid_memory
Z = np.arange(8).reshape(4,2)
print ("sigmoid(Z) = " + str(sigmoid(Z)))
# ### relu
# This is one of the activation functions. It takes the cumulative input to the layer, matrix **Z** as the input. Upon application of the **`relu`** function, matrix **H** which is the output matrix is calculated. Also, **Z** is stored as **relu_memory** which will be later used in backpropagation. You use _[np.maximum()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.maximum.html)_ here in the following way.
def relu(Z):
# Z is numpy array of shape (n, m) where n is number of neurons in the layer and m is the number of samples
# relu_memory is stored as it is used later on in backpropagation
H = np.maximum(0,Z)
assert(H.shape == Z.shape)
relu_memory = Z
return H, relu_memory
Z = np.array([1, 3, -1, -4, -5, 7, 9, 18]).reshape(4,2)
print ("relu(Z) = " + str(relu(Z)))
# ### softmax
# This is the activation of the last layer. It takes the cumulative input to the layer, matrix **Z** as the input. Upon application of the **`softmax`** function, the output matrix **H** is calculated. Also, **Z** is stored as **softmax_memory** which will be later used in backpropagation. You use _[np.exp()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.exp.html)_ and _[np.sum()](https://docs.scipy.org/doc/numpy-1.10.0/reference/generated/numpy.sum.html)_ here in the following way. The exponential gets applied to all the elements of Z.
def softmax(Z):
# Z is numpy array of shape (n, m) where n is number of neurons in the layer and m is the number of samples
# softmax_memory is stored as it is used later on in backpropagation
Z_exp = np.exp(Z)
Z_sum = np.sum(Z_exp,axis = 0, keepdims = True)
H = Z_exp/Z_sum #normalising step
softmax_memory = Z
return H, softmax_memory
Z = np.array([[11,19,10], [12, 21, 23]])
#Z = np.array(np.arange(30)).reshape(10,3)
H, softmax_memory = softmax(Z)
print(H)
print(softmax_memory)
# ### initialize_parameters
# Let's now create a function **`initialize_parameters`** which initializes the weights and biases of the various layers. One way to initialise is to set all the parameters to 0. This is not a considered a good strategy as all the neurons will behave the same way and it'll defeat the purpose of deep networks. Hence, we initialize the weights randomly to very small values but not zeros. The biases are initialized to 0. Note that the **`initialize_parameters`** function initializes the parameters for all the layers in one `for` loop.
#
# The inputs to this function is a list named `dimensions`. The length of the list is the number layers in the network + 1 (the plus one is for the input layer, rest are hidden + output). The first element of this list is the dimensionality or length of the input (784 for the MNIST dataset). The rest of the list contains the number of neurons in the corresponding (hidden and output) layers.
#
# For example `dimensions = [784, 3, 7, 10]` specifies a network for the MNIST dataset with two hidden layers and a 10-dimensional softmax output.
#
# Also, notice that the parameters are returned in a dictionary. This will help you in implementing the feedforward through the layer and the backprop throught the layer at once.
def initialize_parameters(dimensions):
# dimensions is a list containing the number of neuron in each layer in the network
# It returns parameters which is a python dictionary containing the parameters "W1", "b1", ..., "WL", "bL":
np.random.seed(2)
parameters = {}
L = len(dimensions) # number of layers in the network + 1
for l in range(1, L):
parameters['W' + str(l)] = np.random.randn(dimensions[l], dimensions[l-1]) * 0.1
parameters['b' + str(l)] = np.zeros((dimensions[l], 1))
assert(parameters['W' + str(l)].shape == (dimensions[l], dimensions[l-1]))
assert(parameters['b' + str(l)].shape == (dimensions[l], 1))
return parameters
dimensions = [784, 3,7,10]
parameters = initialize_parameters(dimensions)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
# print("W3 = " + str(parameters["W3"]))
# print("b3 = " + str(parameters["b3"]))
# ### layer_forward
#
# The function **`layer_forward`** implements the forward propagation for a certain layer 'l'. It calculates the cumulative input into the layer **Z** and uses it to calculate the output of the layer **H**. It takes **H_prev, W, b and the activation function** as inputs and stores the **linear_memory, activation_memory** in the variable **memory** which will be used later in backpropagation.
#
# <br> You have to first calculate the **Z**(using the forward propagation equation), **linear_memory**(H_prev, W, b) and then calculate **H, activation_memory**(Z) by applying activation functions - **`sigmoid`**, **`relu`** and **`softmax`** on **Z**.
#
# <br> Note that $$H^{L-1}$$ is referred here as H_prev. You might want to use _[np.dot()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)_ to carry out the matrix multiplication.
# +
#Graded
def layer_forward(H_prev, W, b, activation = 'relu'):
# H_prev is of shape (size of previous layer, number of examples)
# W is weights matrix of shape (size of current layer, size of previous layer)
# b is bias vector of shape (size of the current layer, 1)
# activation is the activation to be used for forward propagation : "softmax", "relu", "sigmoid"
# H is the output of the activation function
# memory is a python dictionary containing "linear_memory" and "activation_memory"
if activation == "sigmoid":
Z = np.dot(W, H_prev) + b #write your code here W * H_prev + b
linear_memory = (H_prev, W, b)
H, activation_memory = sigmoid(Z) #write your code here
elif activation == "softmax":
Z = np.dot(W, H_prev) + b #write your code here
linear_memory = (H_prev, W, b)
H, activation_memory = softmax(Z) #write your code here
elif activation == "relu":
Z = np.dot(W, H_prev) + b #write your code here
linear_memory = (H_prev, W, b)
H, activation_memory = relu(Z) #write your code here
assert (H.shape == (W.shape[0], H_prev.shape[1]))
memory = (linear_memory, activation_memory)
return H, memory
# +
# verify
# l-1 has two neurons, l has three, m = 5
# H_prev is (l-1, m)
# W is (l, l-1)
# b is (l, 1)
# H should be (l, m)
H_prev = np.array([[1,0, 5, 10, 2], [2, 5, 3, 10, 2]])
W_sample = np.array([[10, 5], [2, 0], [1, 0]])
b_sample = np.array([10, 5, 0]).reshape((3, 1))
H = layer_forward(H_prev, W_sample, b_sample, activation="sigmoid")[0]
H
# -
# You should get:<br>
# array([[1. , 1. , 1. , 1. , 1. ],<br>
# [0.99908895, 0.99330715, 0.99999969, 1. , 0.99987661],<br>
# [0.73105858, 0.5 , 0.99330715, 0.9999546 , 0.88079708]])
#
# ### L_layer_forward
# **`L_layer_forward`** performs one forward pass through the whole network for all the training samples (note that we are feeding all training examples in one single batch). Use the **`layer_forward`** you have created above here to perform the feedforward for layers 1 to 'L-1' in the for loop with the activation **`relu`**. The last layer having a different activation **`softmax`** is calculated outside the loop. Notice that the **memory** is appended to **memories** for all the layers. These will be used in the backward order during backpropagation.
# +
#Graded
def L_layer_forward(X, parameters):
# X is input data of shape (input size, number of examples)
# parameters is output of initialize_parameters()
# HL is the last layer's post-activation value
# memories is the list of memory containing (for a relu activation, for example):
# - every memory of relu forward (there are L-1 of them, indexed from 1 to L-1),
# - the memory of softmax forward (there is one, indexed L)
memories = []
H = X
L = len(parameters) // 2 # number of layers in the neural network
# Implement relu layer (L-1) times as the Lth layer is the softmax layer
for l in range(1, L):
H_prev = H #write your code here
H, memory = layer_forward(H_prev, parameters['W' + str(l)], parameters['b' + str(l)], 'relu') #write your code here
memories.append(memory)
# Implement the final softmax layer
# HL here is the final prediction P as specified in the lectures
HL, memory = layer_forward(H, parameters['W' + str(L)], parameters['b' + str(L)], 'softmax')#write your code here
memories.append(memory)
assert(HL.shape == (10, X.shape[1]))
return HL, memories
# -
# verify
# X is (784, 10)
# parameters is a dict
# HL should be (10, 10)
x_sample = train_set_x[:, 10:20]
print(x_sample.shape)
HL = L_layer_forward(x_sample, parameters=parameters)[0]
print(HL[:, :5])
# You should get:
#
# (784, 10)<br>
# [[0.10106734 0.10045152 0.09927757 0.10216656 0.1 ]<br>
# [0.10567625 0.10230873 0.10170271 0.11250099 0.1 ]<br>
# [0.09824287 0.0992886 0.09967128 0.09609693 0.1 ]<br>
# [0.10028288 0.10013048 0.09998149 0.10046076 0.1 ]<br>
# [0.09883601 0.09953443 0.09931419 0.097355 0.1 ]<br>
# [0.10668575 0.10270912 0.10180736 0.11483609 0.1 ]<br>
# [0.09832513 0.09932275 0.09954792 0.09627089 0.1 ]<br>
# [0.09747092 0.09896735 0.0995387 0.09447277 0.1 ]<br>
# [0.09489069 0.09788255 0.09929998 0.08915178 0.1 ]<br>
# [0.09852217 0.09940447 0.09985881 0.09668824 0.1 ]]
# # Loss
#
# ### compute_loss
# The next step is to compute the loss function after every forward pass to keep checking whether it is decreasing with training.<br> **`compute_loss`** here calculates the cross-entropy loss. You may want to use _[np.log()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.log.html)_, _[np.sum()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.log.html)_, _[np.multiply()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.multiply.html)_ here. Do not forget that it is the average loss across all the data points in the batch. It takes the output of the last layer **HL** and the ground truth label **Y** as input and returns the **loss**.
# +
#Graded
def compute_loss(HL, Y):
# HL is probability matrix of shape (10, number of examples)
# Y is true "label" vector shape (10, number of examples)
# loss is the cross-entropy loss
m = Y.shape[1]
loss = (-1./m) * np.sum(np.multiply(Y, np.log(HL))) #write your code here, use (1./m) and not (1/m)
loss = np.squeeze(loss) # To make sure that the loss's shape is what we expect (e.g. this turns [[17]] into 17).
assert(loss.shape == ())
return loss
# +
# sample
# HL is (10, 5), Y is (10, 5)
np.random.seed(2)
HL_sample = np.random.rand(10,5)
Y_sample = train_set_y[:, 10:15]
print(HL_sample)
print(Y_sample)
print(compute_loss(HL_sample, Y_sample))
# -
# You should get:<br>
#
# [[0.4359949 0.02592623 0.54966248 0.43532239 0.4203678 ]<br>
# [0.33033482 0.20464863 0.61927097 0.29965467 0.26682728]<br>
# [0.62113383 0.52914209 0.13457995 0.51357812 0.18443987]<br>
# [0.78533515 0.85397529 0.49423684 0.84656149 0.07964548]<br>
# [0.50524609 0.0652865 0.42812233 0.09653092 0.12715997]<br>
# [0.59674531 0.226012 0.10694568 0.22030621 0.34982629]<br>
# [0.46778748 0.20174323 0.64040673 0.48306984 0.50523672]<br>
# [0.38689265 0.79363745 0.58000418 0.1622986 0.70075235]<br>
# [0.96455108 0.50000836 0.88952006 0.34161365 0.56714413]<br>
# [0.42754596 0.43674726 0.77655918 0.53560417 0.95374223]]<br>
# [[0. 0. 0. 0. 0.]<br>
# [0. 0. 0. 0. 1.]<br>
# [0. 0. 0. 0. 0.]<br>
# [1. 0. 1. 0. 0.]<br>
# [0. 0. 0. 0. 0.]<br>
# [0. 1. 0. 0. 0.]<br>
# [0. 0. 0. 1. 0.]<br>
# [0. 0. 0. 0. 0.]<br>
# [0. 0. 0. 0. 0.]<br>
# [0. 0. 0. 0. 0.]]<br>
# 0.8964600261334037
# # Backpropagation
# Let's now get to the next step - backpropagation. Let's start with sigmoid_backward.
#
# ### sigmoid-backward
# You might remember that we had created **`sigmoid`** function that calculated the activation for forward propagation. Now, we need the activation backward, which helps in calculating **dZ** from **dH**. Notice that it takes input **dH** and **sigmoid_memory** as input. **sigmoid_memory** is the **Z** which we had calculated during forward propagation. You use _[np.exp()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.exp.html)_ here the following way.
def sigmoid_backward(dH, sigmoid_memory):
# Implement the backpropagation of a sigmoid function
# dH is gradient of the sigmoid activated activation of shape same as H or Z in the same layer
# sigmoid_memory is the memory stored in the sigmoid(Z) calculation
Z = sigmoid_memory
H = 1/(1+np.exp(-Z))
dZ = dH * H * (1-H)
assert (dZ.shape == Z.shape)
return dZ
# ### relu-backward
# You might remember that we had created **`relu`** function that calculated the activation for forward propagation. Now, we need the activation backward, which helps in calculating **dZ** from **dH**. Notice that it takes input **dH** and **relu_memory** as input. **relu_memory** is the **Z** which we calculated uring forward propagation.
def relu_backward(dH, relu_memory):
# Implement the backpropagation of a relu function
# dH is gradient of the relu activated activation of shape same as H or Z in the same layer
# relu_memory is the memory stored in the sigmoid(Z) calculation
Z = relu_memory
dZ = np.array(dH, copy=True) # dZ will be the same as dA wherever the elements of A weren't 0
dZ[Z <= 0] = 0
assert (dZ.shape == Z.shape)
return dZ
# ### layer_backward
#
# **`layer_backward`** is a complimentary function of **`layer_forward`**. Like **`layer_forward`** calculates **H** using **W**, **H_prev** and **b**, **`layer_backward`** uses **dH** to calculate **dW**, **dH_prev** and **db**. You have already studied the formulae in backpropogation. To calculate **dZ**, use the **`sigmoid_backward`** and **`relu_backward`** function. You might need to use _[np.dot()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)_, _[np.sum()](https://docs.scipy.org/doc/numpy/reference/generated/numpy.sum.html)_ for the rest. Remember to choose the axis correctly in db.
# +
#Graded
def layer_backward(dH, memory, activation = 'relu'):
# takes dH and the memory calculated in layer_forward and activation as input to calculate the dH_prev, dW, db
# performs the backprop depending upon the activation function
linear_memory, activation_memory = memory
if activation == "relu":
dZ = relu_backward(dH, activation_memory) #write your code here
H_prev, W, b = linear_memory
m = H_prev.shape[1]
dW = (1./m) * np.dot(dZ, H_prev.T) #write your code here, use (1./m) and not (1/m)
db = (1./m) * np.sum(dZ, axis = -1, keepdims=True) #write your code here, use (1./m) and not (1/m)
dH_prev = np.dot(W.T, dZ) #write your code here
elif activation == "sigmoid":
dZ = sigmoid_backward(dH, activation_memory) #write your code here
H_prev, W, b = linear_memory
m = H_prev.shape[1]
dW = (1./m) * np.dot(dZ, H_prev.T) #write your code here, use (1./m) and not (1/m)
db = (1./m) * np.sum(dZ, axis = -1, keepdims=True) #write your code here, use (1./m) and not (1/m)
dH_prev = np.dot(W.T, dZ) #write your code here
return dH_prev, dW, db
# +
# verify
# l-1 has two neurons, l has three, m = 5
# H_prev is (l-1, m)
# W is (l, l-1)
# b is (l, 1)
# H should be (l, m)
H_prev = np.array([[1,0, 5, 10, 2], [2, 5, 3, 10, 2]])
W_sample = np.array([[10, 5], [2, 0], [1, 0]])
b_sample = np.array([10, 5, 0]).reshape((3, 1))
H, memory = layer_forward(H_prev, W_sample, b_sample, activation="relu")
np.random.seed(2)
dH = np.random.rand(3,5)
dH_prev, dW, db = layer_backward(dH, memory, activation = 'relu')
print('dH_prev is \n' , dH_prev)
print('dW is \n' ,dW)
print('db is \n', db)
# -
# You should get:<br>
# dH_prev is <br>
# [[5.6417525 0.66855959 6.86974666 5.46611139 4.92177244]<br>
# [2.17997451 0.12963116 2.74831239 2.17661196 2.10183901]]<br>
# dW is <br>
# [[1.67565336 1.56891359]<br>
# [1.39137819 1.4143854 ]<br>
# [1.3597389 1.43013369]]<br>
# db is <br>
# [[0.37345476]<br>
# [0.34414727]<br>
# [0.29074635]]<br>
#
# ### L_layer_backward
#
# **`L_layer_backward`** performs backpropagation for the whole network. Recall that the backpropagation for the last layer, i.e. the softmax layer, is different from the rest, hence it is outside the reversed `for` loop. You need to use the function **`layer_backward`** here in the loop with the activation function as **`relu`**.
# +
#Graded
def L_layer_backward(HL, Y, memories):
# Takes the predicted value HL and the true target value Y and the
# memories calculated by L_layer_forward as input
# returns the gradients calulated for all the layers as a dict
gradients = {}
L = len(memories) # the number of layers
m = HL.shape[1]
Y = Y.reshape(HL.shape) # after this line, Y is the same shape as AL
# Perform the backprop for the last layer that is the softmax layer
current_memory = memories[-1]
linear_memory, activation_memory = current_memory
dZ = HL - Y
H_prev, W, b = linear_memory
# Use the expressions you have used in 'layer_backward'
gradients["dH" + str(L-1)] = np.dot(W.T, dZ) #write your code here
gradients["dW" + str(L)] = (1./m) * np.dot(dZ, H_prev.T) #write your code here, use (1./m) and not (1/m)
gradients["db" + str(L)] = (1./m) * np.sum(dZ, axis = -1, keepdims=True) #write your code here, use (1./m) and not (1/m)
# Perform the backpropagation l-1 times
for l in reversed(range(L-1)):
# Lth layer gradients: "gradients["dH" + str(l + 1)] ", gradients["dW" + str(l + 2)] , gradients["db" + str(l + 2)]
current_memory = memories[l]
dH_prev_temp, dW_temp, db_temp = layer_backward(gradients["dH" + str(l+1)], current_memory, activation='relu') #write your code here
gradients["dH" + str(l)] = dH_prev_temp #write your code here
gradients["dW" + str(l + 1)] = dW_temp #write your code here
gradients["db" + str(l + 1)] = db_temp #write your code here
return gradients
# +
# verify
# X is (784, 10)
# parameters is a dict
# HL should be (10, 10)
x_sample = train_set_x[:, 10:20]
y_sample = train_set_y[:, 10:20]
HL, memories = L_layer_forward(x_sample, parameters=parameters)
gradients = L_layer_backward(HL, y_sample, memories)
print('dW3 is \n', gradients['dW3'])
print('db3 is \n', gradients['db3'])
print('dW2 is \n', gradients['dW2'])
print('db2 is \n', gradients['db2'])
# -
# You should get:<br>
#
# dW3 is <br>
# [[ 0.02003701 0.0019043 0.01011729 0.0145757 0.00146444 0.00059863 0. ]<br>
# [ 0.02154547 0.00203519 0.01085648 0.01567075 0.00156469 0.00060533 0. ]<br>
# [-0.01718407 -0.00273711 -0.00499101 -0.00912135 -0.00207365 0.00059996 0. ]<br>
# [-0.01141498 -0.00158622 -0.00607049 -0.00924709 -0.00119619 0.00060381 0. ]<br>
# [ 0.01943173 0.0018421 0.00984543 0.01416368 0.00141676 0.00059682 0. ]<br>
# [ 0.01045447 0.00063974 0.00637621 0.00863306 0.00050118 0.00060441 0. ]<br>
# [-0.06338911 -0.00747251 -0.0242169 -0.03835708 -0.00581131 0.0006034 0. ]<br>
# [ 0.01911373 0.001805 0.00703101 0.0120636 0.00138836 -0.00140535 0. ]<br>
# [-0.01801603 0.0017357 -0.01489228 -0.02026076 0.00133528 0.00060264 0. ]<br>
# [ 0.0194218 0.00183381 0.00594427 0.01187949 0.00141043 -0.00340965 0. ]]<br>
# db3 is <br>
# [[ 0.10031756]<br>
# [ 0.00460183]<br>
# [-0.00142942]<br>
# [-0.0997827 ]<br>
# [ 0.09872663]<br>
# [ 0.00536378]<br>
# [-0.10124784]<br>
# [-0.00191121]<br>
# [-0.00359044]<br>
# [-0.00104818]]<br>
# dW2 is <br>
# [[ 4.94428956e-05 1.13215514e-02 5.44180380e-02]<br>
# [-4.81267081e-05 -2.96999448e-05 -1.81899582e-02]<br>
# [ 5.63424333e-05 4.77190073e-03 4.04810232e-02]<br>
# [ 1.49767478e-04 -1.89780927e-03 -7.91231369e-03]<br>
# [ 1.97866094e-04 1.22107085e-04 2.64140566e-02]<br>
# [ 0.00000000e+00 -3.75805770e-04 1.63906102e-05]<br>
# [ 0.00000000e+00 0.00000000e+00 0.00000000e+00]]<br>
# db2 is <br>
# [[ 0.013979 ]<br>
# [-0.01329383]<br>
# [ 0.01275707]<br>
# [-0.01052957]<br>
# [ 0.03179224]<br>
# [-0.00039877]<br>
# [ 0. ]]<br>
# # Parameter Updates
#
# Now that we have calculated the gradients. let's do the last step which is updating the weights and biases.
# +
#Graded
def update_parameters(parameters, gradients, learning_rate):
# parameters is the python dictionary containing the parameters W and b for all the layers
# gradients is the python dictionary containing your gradients, output of L_model_backward
# returns updated weights after applying the gradient descent update
L = len(parameters) // 2 # number of layers in the neural network
for l in range(L):
parameters["W" + str(l+1)] = parameters["W" + str(l+1)] - learning_rate * gradients["dW" + str(l+1)] #write your code here
parameters["b" + str(l+1)] = parameters["b" + str(l+1)] - learning_rate * gradients["db" + str(l+1)]#write your code here
return parameters
# -
# Having defined the bits and pieces of the feedforward and the backpropagation, let's now combine all that to form a model. The list `dimensions` has the number of neurons in each layer specified in it. For a neural network with 1 hidden layer with 45 neurons, you would specify the dimensions as follows:
dimensions = [784, 45, 10] # three-layer model
# # Model
#
# ### L_layer_model
#
# This is a composite function which takes the training data as input **X**, ground truth label **Y**, the **dimensions** as stated above, **learning_rate**, the number of iterations **num_iterations** and if you want to print the loss, **print_loss**. You need to use the final functions we have written for feedforward, computing the loss, backpropagation and updating the parameters.
# +
#Graded
def L_layer_model(X, Y, dimensions, learning_rate = 0.0075, num_iterations = 3000, print_loss=False):
# X and Y are the input training datasets
# learning_rate, num_iterations are gradient descent optimization parameters
# returns updated parameters
np.random.seed(2)
losses = [] # keep track of loss
# Parameters initialization
parameters = initialize_parameters(dimensions) #write your code here
for i in range(0, num_iterations):
# Forward propagation
HL, memories = L_layer_forward(X, parameters) #write your code here
# Compute loss
loss = compute_loss(HL, Y) #write your code here
# Backward propagation
gradients = L_layer_backward(HL, Y, memories) #write your code here
# Update parameters.
parameters = update_parameters(parameters, gradients, learning_rate) #write your code here
# Printing the loss every 100 training example
if print_loss and i % 100 == 0:
print ("Loss after iteration %i: %f" %(i, loss))
losses.append(loss)
# plotting the loss
plt.plot(np.squeeze(losses))
plt.ylabel('loss')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
# -
# Since, it'll take a lot of time to train the model on 50,000 data points, we take a subset of 5,000 images.
train_set_x_new = train_set_x[:,0:5000]
train_set_y_new = train_set_y[:,0:5000]
train_set_x_new.shape
# Now, let's call the function L_layer_model on the dataset we have created.This will take 10-20 mins to run.
parameters = L_layer_model(train_set_x_new, train_set_y_new, dimensions, num_iterations = 2000, print_loss = True)
def predict(X, y, parameters):
# Performs forward propogation using the trained parameters and calculates the accuracy
m = X.shape[1]
n = len(parameters) // 2 # number of layers in the neural network
# Forward propagation
probas, caches = L_layer_forward(X, parameters)
p = np.argmax(probas, axis = 0)
act = np.argmax(y, axis = 0)
print("Accuracy: " + str(np.sum((p == act)/m)))
return p
# Let's see the accuray we get on the training data.
pred_train = predict(train_set_x_new, train_set_y_new, parameters)
# We get ~ 88% accuracy on the training data. Let's see the accuray on the test data.
pred_test = predict(test_set_x, test_set_y, parameters)
# It is ~87%. You can train the model even longer and get better result. You can also try to change the network structure.
# <br>Below, you can see which all numbers are incorrectly identified by the neural network by changing the index.
index = 3474
k = test_set_x[:,index]
k = k.reshape((28, 28))
plt.title('Label is {label}'.format(label=(pred_test[index], np.argmax(test_set_y, axis = 0)[index])))
plt.imshow(k, cmap='gray')
| src/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="MvrZGmWA8LQG" executionInfo={"status": "ok", "timestamp": 1634410491306, "user_tz": 180, "elapsed": 30675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
# %%capture
# !pip install pandas
# !pip install numpy
# !pip install tensorflow
# !pip install keras
# !pip install sklearn
# !pip install matplotlib
# !pip install seaborn
# !pip install unidecode
# !pip install -U imbalanced-learn
# !pip3 install pickle5
# + colab={"base_uri": "https://localhost:8080/"} id="jbkU3YVN8Pwd" executionInfo={"status": "ok", "timestamp": 1634410494555, "user_tz": 180, "elapsed": 3261, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="d9a60138-91ed-40ad-d99d-96a16d5a803b"
import tensorflow as tf
import pandas as pd
import warnings
import unidecode
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pickle5 as pickle
import random
import os
import re
import time
import nltk
nltk.download('stopwords')
sns.set_style('darkgrid')
from imblearn.over_sampling import RandomOverSampler
from nltk.corpus import stopwords
from sklearn.model_selection import train_test_split
from tensorflow import keras
from keras.models import Sequential, Model
from keras.layers import Reshape, Dense, Dropout, Flatten, Input, MaxPooling2D, Convolution2D, Embedding, Concatenate
from keras.regularizers import l2
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from sklearn.metrics import f1_score, confusion_matrix
sw = set(stopwords.words('english'))
os.environ['PYTHONHASHSEED']=str(23)
tf.random.set_seed(23)
random.seed(23)
warnings.filterwarnings('ignore')
np.random.seed(23)
# + [markdown] id="fbIsBrCP1iMA"
# ## Preprocessing
# + id="DVGCip5q1hla" executionInfo={"status": "ok", "timestamp": 1634410494562, "user_tz": 180, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def remove_username(text):
text = re.sub(r'\@[^\s]+', ' ', text)
return text
def remove_newline(text):
text = text.replace('\n', ' ')
return text
def only_letters(text):
text = re.sub(r'[^a-záâàãéêèẽíìîĩóòõôúùũû\s]+', ' ', text)
return text
def remove_link(text):
text = re.sub(r'www\.?[^\s]+', ' ', text)
return text
def remove_hyperlink(text):
text = re.sub(r'\<.?\>', ' ', text)
return text
def remove_accent(text):
text = unidecode.unidecode(text)
return text
def adjustment_text(text):
text = re.sub(r'\s+', ' ', text)
text = text.strip()
return text
def remove_stopwords(text):
text = [word for word in text.split() if word not in sw]
text = ' '.join(text)
return text
def remove_spam(text):
text = re.sub(r'\&', ' ', text)
text = re.sub(r'\<', ' ', text)
text = re.sub(r'\>', ' ', text)
text = re.sub(r'\#follow|\#followme|\#like|\#f4f|\#photooftheday', ' ', text)
return text
def remove_slangs(text):
text = re.sub(r' b4 ', ' before ', text)
text = re.sub(r' 2b ', ' to be ', text)
text = re.sub(r' 2morrow ', ' tomorrow ', text)
text = re.sub(r' rn ', ' right now ', text)
text = re.sub(r' brb ', ' be right back ', text)
text = re.sub(r' mb ', ' my bad ', text)
text = re.sub(r' luv ', ' love ', text)
text = re.sub(r' b ', ' be ', text)
text = re.sub(r' r ', ' are ', text)
text = re.sub(r' u ', ' you ', text)
text = re.sub(r' y ', ' why ', text)
text = re.sub(r' ur ', ' your ', text)
text = re.sub(r' hbd ', ' happy birthday ', text)
text = re.sub(r' bday ', ' birthday ', text)
text = re.sub(r' bihday ', ' birthday ', text)
text = re.sub(r' omg ', ' oh my god ', text)
text = re.sub(r' lol ', ' laughing out loud ', text)
return text
def remove_abbreviations(text):
text = re.sub(r" can\'t ", " can not ", text)
text = re.sub(r" i\'m ", " i am ", text)
text = re.sub(r" i\'ll ", " i will ", text)
text = re.sub(r" i\'d ", " i would ", text)
text = re.sub(r" i\'ve ", " i have ", text)
text = re.sub(r" ain\'t ", " am not ", text)
text = re.sub(r" haven\'t ", " have not ", text)
text = re.sub(r" hasn\'t ", " has not ", text)
text = re.sub(r" can\'t ", " can not ", text)
text = re.sub(r" won\'t ", " will not ", text)
text = re.sub(r" you\'re ", " you are ", text)
text = re.sub(r" we\'re ", " we are ", text)
text = re.sub(r" they\'re ", " they are ", text)
text = re.sub(r" he\'s ", " he is ", text)
text = re.sub(r" she\'s ", " she is ", text)
text = re.sub(r" it\'s ", " it is ", text)
text = re.sub(r" don\'t ", " do not ", text)
text = re.sub(r" doesn\'t ", " does not ", text)
text = re.sub(r" wouldn\'t ", " would not ", text)
text = re.sub(r" couldn\'t ", " could not ", text)
text = re.sub(r" shouldn\'t ", " should not ", text)
return text
def remove_one_len_word(text):
text = re.sub(r'\b[a-z]\b', ' ', text)
return text
def preprocessing(data):
data['cleaned_tweet'] = data['tweet'].apply(str)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(lambda x: x.lower())
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_newline)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_hyperlink)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_spam)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_link)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_username)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_abbreviations)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(only_letters)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_accent)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_slangs)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_stopwords)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(remove_one_len_word)
data['cleaned_tweet'] = data['cleaned_tweet'].apply(adjustment_text)
return data
# + [markdown] id="wM2uoJUZ8oVr"
# from google.colab import files
# uploaded = files.upload()
# + [markdown] id="FRxNPczhV8wy"
# # # !unzip tokenizer_RNN_seed23
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="0PRNAa9_-qIM" executionInfo={"status": "ok", "timestamp": 1634410494564, "user_tz": 180, "elapsed": 40, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="939ac868-132e-419c-b17a-9cfcb2359b8b"
normal_data = pd.read_csv('Data/train.csv')
normal_data = normal_data.drop(columns=['id'])
normal_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="iv3WqTC3kKPP" executionInfo={"status": "ok", "timestamp": 1634410494568, "user_tz": 180, "elapsed": 36, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="ec3e93dd-272b-4d18-f5cb-7e5a991880ec"
normal_data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="2sOor4hzRfB2" executionInfo={"status": "ok", "timestamp": 1634410495353, "user_tz": 180, "elapsed": 809, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="491974a7-033c-4aa1-ac3c-6b039a7e90e4"
plt.figure(figsize=(8, 6))
sns.countplot(data=normal_data, x='label', color='cornflowerblue')
plt.xlabel('Classe', fontsize=14)
plt.xticks(fontsize=13)
plt.ylabel('Quantidade de mensagens', fontsize=14)
plt.yticks(fontsize=13)
plt.savefig('Images/data_unbalanced.png')
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="YWsDE9A2kD_u" executionInfo={"status": "ok", "timestamp": 1634410495355, "user_tz": 180, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="9089828f-4709-4567-88e5-2d3ce486b7d9"
ros = RandomOverSampler(random_state=23, sampling_strategy='minority')
X_resampled, y_resampled = ros.fit_resample(normal_data[['tweet']], normal_data['label'])
data_augmentation = pd.concat([X_resampled, y_resampled], axis=1)
data_augmentation.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="SVjsRUIWPYST" executionInfo={"status": "ok", "timestamp": 1634410495357, "user_tz": 180, "elapsed": 34, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="e20f3f13-d57e-4a86-b497-394cf2d634fa"
plt.figure(figsize=(8, 6))
sns.countplot(data=data_augmentation, x='label', color='cornflowerblue')
plt.xlabel('Classe', fontsize=14)
plt.xticks(fontsize=13)
plt.ylabel('Quantidade de mensagens', fontsize=14)
plt.yticks(fontsize=13)
plt.savefig('Images/data_balanced.png')
# + colab={"base_uri": "https://localhost:8080/"} id="szfpHt7rzTIb" executionInfo={"status": "ok", "timestamp": 1634410495359, "user_tz": 180, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="be54cc05-3d52-44a1-bd8a-dd01bc619a12"
data_augmentation.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="2Y4K9Bt11Zeu" executionInfo={"status": "ok", "timestamp": 1634410498243, "user_tz": 180, "elapsed": 2907, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="cbbec8b7-b3b5-4748-dd9d-25e0e9a4e510"
preprocessed_data = normal_data.copy()
preprocessed_data = preprocessing(preprocessed_data)
preprocessed_data = preprocessed_data.replace('None', pd.NA)
preprocessed_data = preprocessed_data.dropna()
preprocessed_data = preprocessed_data.drop_duplicates()
preprocessed_data = preprocessed_data.drop(columns=['tweet'])
preprocessed_data = preprocessed_data.rename(columns={'cleaned_tweet': 'tweet'})
preprocessed_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="pTkLD9L4zM0Z" executionInfo={"status": "ok", "timestamp": 1634410498245, "user_tz": 180, "elapsed": 36, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="377931d1-4a8a-4f68-a27f-c10b7c807928"
preprocessed_data.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="hebFvYTaBpAG" executionInfo={"status": "ok", "timestamp": 1634410498247, "user_tz": 180, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="2c9339bf-825d-4337-cc91-3f63d1d63e10"
ros = RandomOverSampler(random_state=23, sampling_strategy='minority')
X_resampled, y_resampled = ros.fit_resample(preprocessed_data[['tweet']], preprocessed_data['label'])
data_preprocessing_augmentation = pd.concat([X_resampled, y_resampled], axis=1)
data_preprocessing_augmentation.head()
# + colab={"base_uri": "https://localhost:8080/"} id="Q1gSgHkFzPOt" executionInfo={"status": "ok", "timestamp": 1634410498249, "user_tz": 180, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="d2cb1651-1cf1-4315-ee74-8f63023743f8"
data_preprocessing_augmentation.shape
# + [markdown] id="SBHqM8OyIt1P"
# ## Tokenizer
# + [markdown] id="jo2peEd9ItHt"
# with open(r"Tokenizer/tokenizer_non_static.pickle", "rb") as output_file:
# tokenizer_non_static = pickle.load(output_file)
# + [markdown] id="DCMsEXk9JKw5"
# with open(r"Tokenizer/tokenizer_non_static_augmentantion.pickle", "rb") as output_file:
# tokenizer_non_static_augmentation = pickle.load(output_file)
# + [markdown] id="4j9quWvyJTvz"
# with open(r"Tokenizer/tokenizer_non_static_preprocessing.pickle", "rb") as output_file:
# tokenizer_non_static_preprocessing = pickle.load(output_file)
# + [markdown] id="zsH-Oa7zJT1b"
# with open(r"Tokenizer/tokenizer_non_static_preprocessing_augmentantion.pickle", "rb") as output_file:
# tokenizer_non_static_preprocessing_augmentation = pickle.load(output_file)
# + [markdown] id="Y5UyBoIuJfhN"
# with open(r"Tokenizer/tokenizer_rand.pickle", "rb") as output_file:
# tokenizer_rand = pickle.load(output_file)
# + [markdown] id="2ptmhy4eJfmK"
# with open(r"Tokenizer/tokenizer_rand_augmentantion.pickle", "rb") as output_file:
# tokenizer_rand_augmentation = pickle.load(output_file)
# + [markdown] id="R0FkNJIcJfrj"
# with open(r"Tokenizer/tokenizer_rand_preprocessing.pickle", "rb") as output_file:
# tokenizer_rand_preprocessing = pickle.load(output_file)
# + [markdown] id="f2oRqYedJfyi"
# with open(r"Tokenizer/tokenizer_rand_preprocessing_augmentantion.pickle", "rb") as output_file:
# tokenizer_rand_preprocessing_augmentation = pickle.load(output_file)
# + id="VdWFvuxLVsmh" executionInfo={"status": "ok", "timestamp": 1634410498251, "user_tz": 180, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
with open(r"content/Tokenizer/tokenizer.pickle", "rb") as output_file:
tokenizer_lstm = pickle.load(output_file)
# + id="iVncBz2wcUiM" executionInfo={"status": "ok", "timestamp": 1634410501994, "user_tz": 180, "elapsed": 3761, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
all_messages = pd.concat([data_preprocessing_augmentation,
preprocessed_data,
normal_data,
data_augmentation], axis=0)
all_messages = all_messages.reset_index(drop=True)
tokenizer = Tokenizer()
tokenizer.fit_on_texts(all_messages['tweet'].values)
# + id="Ukix7VqDVlZW" executionInfo={"status": "ok", "timestamp": 1634410502005, "user_tz": 180, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
with open('Tokenizer/tokenizer.pickle', 'wb') as handle:
pickle.dump(tokenizer, handle, protocol=pickle.HIGHEST_PROTOCOL)
# + colab={"base_uri": "https://localhost:8080/", "height": 196} id="5EAASAM8WVG6" executionInfo={"status": "ok", "timestamp": 1634410502006, "user_tz": 180, "elapsed": 34, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="7167056e-3606-4dce-fd77-b198c0e47b5a"
teste = pd.DataFrame({'tweet': ["the next school year is the year for exams.ð¯ can't think about that ð #school #exams #hate #imagine #actorslife #revolutionschool #girl",
"we won!!! love the land!!! #allin #cavs #champions #cleveland #clevelandcavaliers ⦠",
"it was a hard monday due to cloudy weather. disabling oxygen production for today. #goodnight #badmonday "]})
teste['tokenized_cnn'] = tokenizer.texts_to_sequences(teste['tweet'].values)
teste['tokenized_lstm'] = tokenizer_lstm.texts_to_sequences(teste['tweet'].values)
teste.head()
# + colab={"base_uri": "https://localhost:8080/"} id="dqc18LbdXDvC" executionInfo={"status": "ok", "timestamp": 1634410502009, "user_tz": 180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="4be7fba7-2365-40b5-f9c2-c89a64dd7b7b"
teste['tokenized_lstm'].equals(teste['tokenized_cnn'])
# + [markdown] id="iCpQSNEE7wlo"
# ## Word2vec
# + [markdown] id="HmVKbyWJ7y1W"
# with open(r"Data/word2vec.pickle", "rb") as output_file:
# word2vec_embedding = pickle.load(output_file)
# + [markdown] id="Df2gQAxIsteF"
# with open(r"Data/word2vec_preprocessing.pickle", "rb") as output_file:
# word2vec_preprocessing_embedding = pickle.load(output_file)
# + id="yl5H1vv5cX8k" executionInfo={"status": "ok", "timestamp": 1634410503922, "user_tz": 180, "elapsed": 1930, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
with open(r"Data/word2vec_total.pickle", "rb") as output_file:
word2vec = pickle.load(output_file)
# + [markdown] id="2tnMZB6L9Cwf"
# ## Parameters
# + id="RLPjaO0_9HRS" executionInfo={"status": "ok", "timestamp": 1634410503932, "user_tz": 180, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
non_linearity_function = 'relu'
kernel_size = [3, 4, 5]
filters = 100
dropout_rate = 0.5
l2_constraint = 3
epochs = 10
batch_size = 100
embedding_dim = 300
length_size = 30
# + [markdown] id="xgf0e9zIM_Ee"
# ## Tokenization + padding + splitting data step
# + id="wQ6cy3KnF4s_" executionInfo={"status": "ok", "timestamp": 1634410503933, "user_tz": 180, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def preprocessing_step(tokenizer, data, model, preprocessing, augmentation):
# tokenizer = Tokenizer()
# tokenizer.fit_on_texts(data['tweet'].values)
data['tokenized'] = tokenizer.texts_to_sequences(data['tweet'].values)
vocab_size = len(tokenizer.word_index) + 1
X = pad_sequences(sequences = data['tokenized'],
maxlen = length_size,
padding = 'post')
y = data['label']
X_train, X_validation, y_train, y_validation = train_test_split(X, y, test_size=0.15, random_state=23)
return vocab_size, tokenizer, X_train, X_validation, y_train, y_validation
# + id="VyA3hpt6W9XD" executionInfo={"status": "ok", "timestamp": 1634410503935, "user_tz": 180, "elapsed": 26, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def generate_cm(model, X_validation, y_validation, prep, augmentation, nome):
plt.figure(figsize = (10, 7))
predicted_validation = (model.predict(X_validation) > 0.5).astype("int32")
matrix = confusion_matrix(y_validation, predicted_validation, labels=[0, 1])
sns.set(font_scale=1.4)
sns.heatmap(matrix, annot=True, cmap="Blues", fmt='d', annot_kws={"size": 16})
plt.xlabel('Classe prevista')
plt.ylabel('Classe real')
if prep:
if augmentation:
plt.savefig('Images/matriz_confusao_' + nome + '_preprocessing_augmentation.jpg')
else:
plt.savefig('Images/matriz_confusao_' + nome + '_preprocessing.jpg')
else:
if augmentation:
plt.savefig('Images/matriz_confusao_' + nome + '_augmentation.jpg')
else:
plt.savefig('Images/matriz_confusao_' + nome + '.jpg')
# + [markdown] id="Raq3KN353mxO"
# ## Predict
# + id="dSZq35fD3lTY" executionInfo={"status": "ok", "timestamp": 1634410503938, "user_tz": 180, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def predict(tokenizer, model, prep, augmentation, nome):
test = pd.read_csv('Data/test.csv')
if prep:
test = preprocessing(test)
test['tokenized'] = tokenizer.texts_to_sequences(test['cleaned_tweet'].values)
else:
test['tokenized'] = tokenizer.texts_to_sequences(test['tweet'].values)
X_test = pad_sequences(sequences = test['tokenized'],
maxlen = length_size,
padding = 'post')
predicted = (model.predict(X_test) > 0.5).astype("int32")
prediction = pd.DataFrame()
prediction['id'] = test['id']
prediction['label'] = predicted
if prep:
if augmentation:
prediction.to_csv('Submission/' + nome + '_preprocessing_augmentation.csv', index=False)
else:
prediction.to_csv('Submission/' + nome + '_preprocessing.csv', index=False)
else:
if augmentation:
prediction.to_csv('Submission/' + nome + '_augmentation.csv', index=False)
else:
prediction.to_csv('Submission/' + nome + '.csv', index=False)
# + [markdown] id="KmfpDUot3oyv"
# ## Save Models
# + id="_ou2_2Zq3o7f" executionInfo={"status": "ok", "timestamp": 1634410503941, "user_tz": 180, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def save_model(modelo, nome_modelo, preprocessing, augmentation):
file_name = 'model_'
if preprocessing:
if augmentation:
file_name = file_name + nome_modelo + '_preprocessing_augmentantion'
else:
file_name = file_name + nome_modelo + '_preprocessing'
else:
if augmentation:
file_name = file_name + nome_modelo + '_augmentantion'
else:
file_name = file_name + nome_modelo
modelo.save('Model/' + file_name + '.h5')
# + [markdown] id="PWYmZv062f_D"
# ## Save embeddings
# + id="xtMYplLE2gPS" executionInfo={"status": "ok", "timestamp": 1634410503943, "user_tz": 180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def save_embedding(modelo, tokenizer, nome_modelo, preprocessing, augmentation):
embeddings = modelo.get_layer('embedding').get_weights()[0]
w2v_my = {}
for word, index in tokenizer.word_index.items():
w2v_my[word] = embeddings[index]
file_name = 'embedding_'
if preprocessing:
if augmentation:
file_name = file_name + nome_modelo + '_preprocessing_augmentantion.pickle'
else:
file_name = file_name + nome_modelo + '_preprocessing.pickle'
else:
if augmentation:
file_name = file_name + nome_modelo + '_augmentantion.pickle'
else:
file_name = file_name + nome_modelo + '.pickle'
with open('Model/' + file_name + '.h5', 'wb') as handle:
pickle.dump(w2v_my, handle, protocol=pickle.HIGHEST_PROTOCOL)
# + [markdown] id="Xn-bmRXK8fwV"
# ## CNN-rand
# + id="9A-7AYJW8fMI" executionInfo={"status": "ok", "timestamp": 1634410503945, "user_tz": 180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def cnn_rand(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, preprocessing, augmentation):
#model input
input = Input(shape=(length_size, ))
#embedding layer
embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
input_length=length_size,
name='embedding')(input)
reshape = Reshape((length_size, embedding_dim, 1))(embedding)
#convolution layer
convs = []
for size in kernel_size:
conv = Convolution2D(filters=filters,
kernel_size=(size, embedding_dim),
activation=non_linearity_function,
kernel_regularizer=l2(l2_constraint))(reshape)
pool = MaxPooling2D(strides=(1, 1),
pool_size=(2, 1),
padding='valid')(conv)
convs.append(pool)
#concatenate convs layers
concatenated = Concatenate(axis=1)(convs)
#flatten layer
flatten = Flatten()(concatenated)
#droupout layer
dropout = Dropout(0.5)(flatten)
#output layer
output = Dense(units=1, activation='sigmoid')(dropout)
model_random = Model(inputs=input, outputs=output)
model_random.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
#model_random.summary()
history_random = model_random.fit(X_train,
y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(X_validation, y_validation))
#save_embedding(model_random, tokenizer, 'CNN-rand', preprocessing, augmentation)
save_model(model_random, 'CNN-rand', preprocessing, augmentation)
predicted_validation = (model_random.predict(X_validation) > 0.5).astype("int32")
score = f1_score(y_validation, predicted_validation, average='weighted')
score = round(score, 4)
generate_cm(model_random, X_validation, y_validation, preprocessing, augmentation, 'CNN-rand')
predict(tokenizer, model_random, preprocessing, augmentation, 'CNN-rand')
return model_random, history_random, score
# + [markdown] id="CI9jb9a40sWZ"
# ## CNN-static
# + id="JDx6XooT0ulS" executionInfo={"status": "ok", "timestamp": 1634410503948, "user_tz": 180, "elapsed": 31, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def cnn_static(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, preprocessing, augmentation):
#model input
input = Input(shape=(length_size, ))
#embedding layer
embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
input_length=length_size,
weights=[word2vec],
trainable=False,
name='embedding')(input)
reshape = Reshape((length_size, embedding_dim, 1))(embedding)
#convolution layer
convs = []
for size in kernel_size:
conv = Convolution2D(filters=filters,
kernel_size=(size, embedding_dim),
activation=non_linearity_function,
kernel_regularizer=l2(l2_constraint))(reshape)
pool = MaxPooling2D(strides=(1, 1),
pool_size=(2, 1),
padding='valid')(conv)
convs.append(pool)
#concatenate convs layers
concatenated = Concatenate(axis=1)(convs)
#flatten layer
flatten = Flatten()(concatenated)
#droupout layer
dropout = Dropout(0.5)(flatten)
#output layer
output = Dense(units=1, activation='sigmoid')(dropout)
model_static = Model(inputs=input, outputs=output)
model_static.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
#model_static.summary()
history_static = model_static.fit(X_train,
y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(X_validation, y_validation))
# save_embedding(model_static, tokenizer, 'CNN-static', preprocessing, augmentation)
save_model(model_static, 'CNN-static', preprocessing, augmentation)
predicted_validation = (model_static.predict(X_validation) > 0.5).astype("int32")
score = f1_score(y_validation, predicted_validation, average='weighted')
score = round(score, 4)
generate_cm(model_static, X_validation, y_validation, preprocessing, augmentation, 'CNN-static')
predict(tokenizer, model_static, preprocessing, augmentation, 'CNN-static')
return model_static, history_static, score
# + [markdown] id="O3f-PZSIBoaS"
# ## CNN non-static
# + id="KC_AHFhvBq7h" executionInfo={"status": "ok", "timestamp": 1634410503950, "user_tz": 180, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
def cnn_non_static(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, preprocessing, augmentation):
#model input
input = Input(shape=(length_size, ))
#embedding layer
embedding = Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
input_length=length_size,
weights=[word2vec],
trainable=True,
name='embedding')(input)
reshape = Reshape((length_size, embedding_dim, 1))(embedding)
#convolution layer
convs = []
for size in kernel_size:
conv = Convolution2D(filters=filters,
kernel_size=(size, embedding_dim),
activation=non_linearity_function,
kernel_regularizer=l2(l2_constraint))(reshape)
pool = MaxPooling2D(strides=(1, 1),
pool_size=(2, 1),
padding='valid')(conv)
convs.append(pool)
#concatenate convs layers
concatenated = Concatenate(axis=1)(convs)
#flatten layer
flatten = Flatten()(concatenated)
#droupout layer
dropout = Dropout(0.5)(flatten)
#output layer
output = Dense(units=1, activation='sigmoid')(dropout)
model_non_static = Model(inputs=input, outputs=output)
model_non_static.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
#model_non_static.summary()
history_non_static = model_non_static.fit(X_train,
y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(X_validation, y_validation))
#save_embedding(model_non_static, tokenizer, 'CNN-non-static', preprocessing, augmentation)
save_model(model_non_static, 'CNN-non-static', preprocessing, augmentation)
predicted_validation = (model_non_static.predict(X_validation) > 0.5).astype("int32")
score = f1_score(y_validation, predicted_validation, average='weighted')
score = round(score, 4)
generate_cm(model_non_static, X_validation, y_validation, preprocessing, augmentation, 'CNN-non-static')
predict(tokenizer, model_non_static, preprocessing, augmentation, 'CNN-non-static')
return model_non_static, history_non_static, score
# + [markdown] id="Cg6enS3gDizi"
# ## Main
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="xDxJkScXDlFI" executionInfo={"status": "ok", "timestamp": 1634422120863, "user_tz": 180, "elapsed": 11616944, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="0f1a52bb-2431-4e08-f04e-1f3d5248f01f"
use_augmentation = [False, True]
use_preprocessing = [False, True]
models_used = []
preprocessing_used = []
augmentation_used = []
scores_validation = []
tempos = []
models = ['rand', 'non_static']
for model in models:
for aug in use_augmentation:
for prep in use_preprocessing:
if aug and prep:
vocab_size, tokenizer, X_train, X_validation, y_train, y_validation = preprocessing_step(tokenizer, data_preprocessing_augmentation, model, prep, aug)
elif aug and not prep:
vocab_size, tokenizer, X_train, X_validation, y_train, y_validation = preprocessing_step(tokenizer, data_augmentation, model, prep, aug)
elif prep and not aug:
vocab_size, tokenizer, X_train, X_validation, y_train, y_validation = preprocessing_step(tokenizer, preprocessed_data, model, prep, aug)
else:
vocab_size, tokenizer, X_train, X_validation, y_train, y_validation = preprocessing_step(tokenizer, normal_data, model, prep, aug)
print('Modelo: {}\nPré-processamento: {}\nBalanceamento: {}\n'.format(model, prep, aug))
if model == 'rand':
ini = time.time()
m, history, validation_score = cnn_rand(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, prep, aug)
fim = time.time()
tempo = fim - ini
elif model == 'static':
ini = time.time()
m, history, validation_score = cnn_static(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, prep, aug)
fim = time.time()
tempo = fim - ini
elif model == 'non_static':
ini = time.time()
m, history, validation_score = cnn_non_static(vocab_size, tokenizer, X_train, X_validation, y_train, y_validation, prep, aug)
fim = time.time()
tempo = fim - ini
else:
print('Modelo não encontrado')
break
print('Modelo finalizado!\n')
models_used.append(model)
tempos.append(tempo)
preprocessing_used.append(prep)
augmentation_used.append(aug)
scores_validation.append(validation_score)
# + id="4PJDAY2JDlIm" executionInfo={"status": "ok", "timestamp": 1634422121282, "user_tz": 180, "elapsed": 31, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
results = pd.DataFrame()
results['modelo'] = models_used
results['tempo'] = tempos
results['pré_processamento'] = preprocessing_used
results['balanceamento'] = augmentation_used
results['score_validação'] = scores_validation
# + id="7ko0dEU9Dxne" colab={"base_uri": "https://localhost:8080/", "height": 300} executionInfo={"status": "ok", "timestamp": 1634422121285, "user_tz": 180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="4b4b3564-fee4-44ec-fc9b-d4e551e637f4"
results.head(20)
# + id="C0wXkrN4rMUd" executionInfo={"status": "ok", "timestamp": 1634422121288, "user_tz": 180, "elapsed": 27, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}}
results.to_csv('tempos_textCNN2D_GPU.csv', index=False)
# + id="54rzghrL3jNS" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1634422199463, "user_tz": 180, "elapsed": 78199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="0cab1654-52aa-47ee-d17d-8b1d3746ccf7"
# !zip -r /content/model.zip /content/Model
# + id="Mdhry_XBybwY" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1634422199467, "user_tz": 180, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="2fedcc36-fd18-4061-ee75-eb8807b5e8a2"
# !zip -r /content/submission.zip /content/Submission/
# + id="IyrPHyonN0R0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1634422199469, "user_tz": 180, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="be3683c2-1fc9-4c63-a941-fd8671d3f1b0"
# !zip -r /content/images_cnn.zip /content/Images/
# + id="nHyl8viViEEp" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1634422200145, "user_tz": 180, "elapsed": 694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "13108790060757877730"}} outputId="7eb3d54b-1b92-4557-d721-5671ac5c6ab1"
# !zip -r /content/tokenizer.zip /content/Tokenizer
# + id="WxHcIEk8ffXQ"
while True:
pass
# + id="qScJ-UopHSNO"
1+1
# + [markdown] id="u8d45Vi10XSX"
# ## Referências
# + [markdown] id="N3-LaAxV0WJw"
# https://www.kaggle.com/hamishdickson/cnn-for-sentence-classification-by-yoon-kim
#
# https://github.com/pinkeshbadjatiya/twitter-hatespeech/blob/master/cnn.py
#
# https://github.com/alexander-rakhlin/CNN-for-Sentence-Classification-in-Keras/blob/master/sentiment_cnn.py
#
# https://github.com/satya-thirumani/Python/blob/master/Sentiment%20Analysis/AV_practice_problem_Twitter_Sentiment_Analysis.ipynb
#
# https://github.com/yoonkim/CNN_sentence/blob/23e0e1f7355705bb083043fda05c031b15acb38c/conv_net_classes.py#L340
#
# https://github.com/Jverma/cnn-text-classification-keras/blob/master/text_cnn.py
#
# https://github.com/bentrevett/pytorch-sentiment-analysis/blob/master/4%20-%20Convolutional%20Sentiment%20Analysis.ipynb
#
# https://github.com/dennybritz/cnn-text-classification-tf/blob/master/text_cnn.py
| Code/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Linear Regression - Analysis
# ============
# ***
#
# We're going to pick up where we left off at the end of the exploration and define a linear model with two independent variables determining the dependent variable, Interest Rate.
#
# Our investigation is now defined as:
#
# _Investigate FICO Score and Loan Amount as predictors of Interest Rate for the Lending Club sample of 2,500 loans._
#
# We use Multivariate Linear Regression to model Interest Rate variance with FICO Score and Loan Amount using:
#
# $$InterestRate = a_0 + a_1 * FICOScore + a_2 * LoanAmount$$
#
# We're going to use modeling software to generate the model coefficients $a_0$, $a_1$ and $a_2$ and then some error estimates that we'll only touch upon lightly at this point.
#
# +
# %pylab inline
import pylab as pl
import numpy as np
#from sklearn import datasets, linear_model
import pandas as pd
import statsmodels.api as sm
# import the cleaned up dataset
df = pd.read_csv('../datasets/loanf.csv')
intrate = df['Interest.Rate']
loanamt = df['Loan.Amount']
fico = df['FICO.Score']
# reshape the data from a pandas Series to columns
# the dependent variable
y = np.matrix(intrate).transpose()
# the independent variables shaped as columns
x1 = np.matrix(fico).transpose()
x2 = np.matrix(loanamt).transpose()
# put the two columns together to create an input matrix
# if we had n independent variables we would have n columns here
x = np.column_stack([x1,x2])
# create a linear model and fit it to the data
X = sm.add_constant(x)
model = sm.OLS(y,X)
f = model.fit()
print 'Coefficients: ', f.params[0:2]
print 'Intercept: ', f.params[2]
print 'P-Values: ', f.pvalues
print 'R-Squared: ', f.rsquared
# -
# So we have a lot of numbers here and we're going to understand some of them.
#
# Coefficients: contains $a_1$ and $a_2$ respectively.
# Intercept: is the $a_0$.
#
# How good are these numbers, how reliable? We need to have some idea. After all we are estimating. We're going to learn a very simple pragmatic way to use a couple of these.
#
# Let's look at the second two numbers.
# We are going to talk loosely here so as to give some flavor of why these are important.
# But this is by no means a formal explanation.
#
# P-Values are probabilities. Informally, each number represents a probability that the respective coefficient we have is a really bad one. To be fairly confident we want this probability to be close to zero. The convention is it needs to be 0.05 or less.
# For now suffice it to say that if we have this true for each of our coefficients then we have good confidence in the model. If one or other of the coefficients is equal to or greater than 0.05 then we have less confidence in that particular dimension being useful in modeling and predicting.
#
# $R$-$squared$ or $R^2$ is a measure of how much of the variance in the data is captured by the model. What does this mean? For now let's understand this as a measure of how well the model captures the **spread** of the observed values not just the average trend.
#
# R is a coefficient of correlation between the independent variables and the dependent variable - i.e. how much the Y depends on the separate X's. R lies between -1 and 1, so $R^2$ lies between 0 and 1.
#
# A high $R^2$ would be close to 1.0 a low one close to 0. The value we have, 0.65, is a reasonably good one. It suggests an R with absolute value in the neighborhood of 0.8.
# The details of these error estimates deserve a separate discussion which we defer until another time.
#
# In summary we have a linear multivariate regression model for Interest Rate based on FICO score and Loan Amount which is well described by the parameters above.
#
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| notebooks/A3. Linear Regression - Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from lifelines.datasets import load_rossi
rossi = load_rossi()
rossi.head()
# let's b-spline age
cph = CoxPHFitter().fit(rossi, "week", "arrest", formula="fin + bs(age, df=4) + wexp + mar + paro + prio")
# +
# now we need to "extend" our data to plot it
# we'll plot age over it's observed range
age_range = np.linspace(rossi['age'].min(), rossi['age'].max(), 50)
# need to create a matrix of variables at their means, _except_ for age.
x_bar = cph._central_values
df_varying_age = pd.concat([x_bar] * 50).reset_index(drop=True)
df_varying_age['age'] = age_range
df_varying_age.head()
# -
cph.predict_log_partial_hazard(df_varying_age).plot()
# +
# compare to _not_ bspline-ing:
cph = CoxPHFitter().fit(rossi, "week", "arrest", formula="fin + age + wexp + mar + paro + prio")
age_range = np.linspace(rossi['age'].min(), rossi['age'].max(), 50)
# need to create a matrix of variables at their means, _except_ for age.
x_bar = cph._central_values
df_varying_age = pd.concat([x_bar] * 50).reset_index(drop=True)
df_varying_age['age'] = age_range
cph.predict_log_partial_hazard(df_varying_age).plot()
# -
| examples/B-splines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''3.9.2'': pyenv)'
# name: python3
# ---
# + cell_id="00000-991a5153-c0b7-4743-858f-db509d0163a9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1621269381567 source_hash="1d682613" tags=[]
import json
with open('../src/lib/vendors.json', 'r') as f:
v = json.load(f)
# + cell_id="00001-7b63542c-015e-4c40-b22b-8a71eed0a271" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1621269415517 source_hash="5b2f3127" tags=[]
vendors = [x['slug'] for x in v['vendors']]
# + cell_id="00002-27660675-5bcc-40f8-a975-7866ed3e78a0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1621270160411 source_hash="482f8a72" tags=[]
urls = [
'/',
'/features/versioning',
'/features/realtime-collaboration',
'/features/comments'
]
# + cell_id="00002-ae4b6a7b-7a4b-4cb8-997a-2d88832246f1" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1621270163009 source_hash="6a4f61a5" tags=[]
for v in vendors:
url = f'/alternatives/{v}'
urls.append(url)
print(f"'{url}',")
# + cell_id="00002-c01071a5-d48d-47cd-84f3-ca40fbfaff9f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=38 execution_start=1621270166450 source_hash="bb62da1c" tags=[]
import itertools
for v1, v2 in itertools.combinations(vendors, 2):
url = f'/compare/{v1}/{v2}'
urls.append(url)
print(f"'{url}',")
print(f"'{url}',")
# + cell_id="00005-e6367096-4ffb-46d4-b885-1b4900be5165" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1621270173414 source_hash="dc03d704" tags=[]
urls
# + cell_id="00004-9b64a1de-7f6b-4e0b-a454-cd94c0f8ca34" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1621270304422 source_hash="f8662b3e" tags=[]
import xml.etree.cElementTree as ET
import datetime
root = ET.Element('urlset')
root.attrib['xmlns:xsi']="http://www.w3.org/2001/XMLSchema-instance"
root.attrib['xsi:schemaLocation']="http://www.sitemaps.org/schemas/sitemap/0.9 http://www.sitemaps.org/schemas/sitemap/0.9/sitemap.xsd"
root.attrib['xmlns']="http://www.sitemaps.org/schemas/sitemap/0.9"
for url in urls:
dt = datetime.datetime.now().strftime ("%Y-%m-%d")
doc = ET.SubElement(root, "url")
ET.SubElement(doc, "loc").text = "https://datasciencenotebook.org"+url
ET.SubElement(doc, "lastmod").text = dt
ET.SubElement(doc, "changefreq").text = "weekly"
ET.SubElement(doc, "priority").text = "1.0"
tree = ET.ElementTree(root)
tree.write('../static/sitemap.xml', encoding='utf-8', xml_declaration=True)
# + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
# <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=d391d205-19c3-4542-b8d3-26ed01b7ba99' target="_blank">
# <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB<KEY> > </img>
# Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebooks/sitemap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import mahotas as mh
from mahotas.features import surf
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import *
from sklearn.cluster import MiniBatchKMeans
import glob
# +
all_instance_filenames = []
all_instance_targets = []
number_images = 1000
data_path = './data/dog-v-cat/train/{}.{}.jpg'
for sp in ['cat', 'dog']:
for n in range(1, number_images+1):
target = 1 if sp == 'cat' else 0
path = data_path.format(sp, n)
all_instance_filenames.append(path)
all_instance_targets.append(target)
surf_features = []
counter = 0
for f in all_instance_filenames:
if counter % 100 == 0:
print "Read {} images".format(counter)
counter += 1
image = mh.imread(f, as_grey=True)
surf_features.append(surf.surf(image)[:, 5:])
print '*Finished reading images*'
# -
train_len = int(len(all_instance_filenames) * .60)
X_train_surf_features = np.concatenate(surf_features[:train_len])
X_test_surf_features = np.concatenate(surf_features[train_len:])
y_train = all_instance_targets[:train_len]
y_test = all_instance_targets[train_len:]
# +
n_clusters = 300
print 'Clustering', len(X_train_surf_features), 'features'
estimator = MiniBatchKMeans(n_clusters=n_clusters)
estimator.fit_transform(X_train_surf_features)
# find the cluster associated with each of the extracted SURF descriptors and count.
X_train = []
for instance in surf_features[:train_len]:
clusters = estimator.predict(instance)
features = np.bincount(clusters)
if len(features) < n_clusters:
features = np.append(features, np.zeros((1, n_clusters - len(features))))
X_train.append(features)
# +
X_test = []
for instance in surf_features[train_len:]:
clusters = estimator.predict(instance)
features = np.bincount(clusters)
if len(features) < n_clusters:
features = np.append(features, np.zeros((1, n_clusters - len(features))))
X_test.append(features)
# +
clf = LogisticRegression(C=0.001, penalty='l2')
clf.fit_transform(X_train, y_train)
predictions = clf.predict(X_test)
print classification_report(y_test, predictions)
print 'Precision:', precision_score(y_test, predictions)
print 'Recall:', recall_score(y_test, predictions)
print 'Accuracy:', accuracy_score(y_test, predictions)
# -
| .ipynb_checkpoints/Dog V Cat-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Crop Areas and Yields foo.005 http://mapspam.info
# foo.005 http://mapspam.info
# SRC: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DHXBJX
#
# Highlight difference between area, production and yield... yield is a rate, production is a total. All three are important.
#
# 4 datasets for 4 crops, 6 layers per Crop (Wheat, Rice, Soybean, Maize)
# - Irrigated
# -- harv area
# -- production
# -- yield
# - Rainfed
# -- harv area
# -- production
# -- yield
#
# OR 2 datasets for Irrigated / Rainfed technologies, 3 layers per crop for 12 layers (harv area wheat, production wheat, yield wheat, etc)
#
# OR 3 datasets for Harvested Area / Production / Yield statistics, 2 layers per crop for 8 layers (irrigated wheat, rainfed wheat, etc)
#
# <u>Available files</u>
# * cell5m_allockey_xy.csv
# primary key for cell grid, xy centroid
# * <b>spam2005V3r1_global_harv_area.geotiff.zip</b>
# * spam2005V3r1_global_phys_area.geotiff.zip
# * <b>spam2005V3r1_global_prod.geotiff.zip</b>
# * spam2005V3r1_global_val_prod_agg.geotiff.zip
# * * includes measures of value of food crops vs. non-food crops
# * <b>spam2005V3r1_global_yield.geotiff.zip</b>
#
# <u>Crops (for now, all food crops)</u>
# * wheat (whea)
# * rice (rice)
# * maize (maiz)
# * soybean (soyb)
#
# <u>Technologies</u>
# * A (all technologies)
# * H (rainfed high inputs)
# * I (irrigated high inputs)
# * L (rainfed low inputs)
# * R (all rainfed technologies)
# * S (rainfed subsistence)
#
#
#
# Import libraries
# +
# Libraries for downloading data from remote server (may be ftp)
import requests
from urllib.request import urlopen
from contextlib import closing
import shutil
# Library for uploading/downloading data to/from S3
import boto3
# Libraries for handling data
import rasterio as rio
import numpy as np
# from netCDF4 import Dataset
# import pandas as pd
# import scipy
# Libraries for various helper functions
# from datetime import datetime
import os
import threading
import sys
from glob import glob
# -
# s3 tools
# +
s3_upload = boto3.client("s3")
s3_download = boto3.resource("s3")
s3_bucket = "wri-public-data"
s3_folder = "resourcewatch/raster/*"
s3_file = "*.tif"
s3_key_orig = s3_folder + s3_file
s3_key_edit = s3_key_orig[0:-4] + "_edit.tif"
class ProgressPercentage(object):
def __init__(self, filename):
self._filename = filename
self._size = float(os.path.getsize(filename))
self._seen_so_far = 0
self._lock = threading.Lock()
def __call__(self, bytes_amount):
# To simplify we'll assume this is hooked up
# to a single filename.
with self._lock:
self._seen_so_far += bytes_amount
percentage = (self._seen_so_far / self._size) * 100
sys.stdout.write("\r%s %s / %s (%.2f%%)"%(
self._filename, self._seen_so_far, self._size,
percentage))
sys.stdout.flush()
# -
# Potentially useful for online directories
# +
# View files in a ftp, see cli.015
remote_path = "ftp://cidportal.jrc.ec.europa.eu/jrc-opendata/EDGAR/datasets/v431_v2"
remote_path_BC = remote_path + "/BC/TOTALS/"
file = req.urlopen(remote_path_BC).read().splitlines()
# Copy from https or ftp
with(closing(urlopen(online_folder + most_recent))) as r:
with(open(local_orig, 'wb')) as f:
shutil.copyfileobj(r, f)
# -
# Define local file locations
# +
local_folder = "/Users/nathansuberi/Desktop/RW_Data/Rasters/*"
file_name = "*"]
local_orig = local_folder + file_name
orig_extension_length = 4 #4 for each char in .tif
local_edit = local_orig[:-orig_extension_length] + "edit.tif"
# -
# Use rasterio to reproject and compress
# +
# Note - this is the core of Vizz's netcdf2tif function
with rio.open(local_orig, 'r') as src:
# This assumes data is readable by rasterio
# May need to open instead with netcdf4.Dataset, for example
data = src.read()[0]
rows = data.shape[0]
columns = data.shape[1]
print(rows)
print(columns)
# Latitude bounds
south_lat = -90
north_lat = 90
# Longitude bounds
west_lon = -180
east_lon = 180
transform = rasterio.transform.from_bounds(west_lon, south_lat, east_lon, north_lat, columns, rows)
# Profile
no_data_val = *
target_projection = 'EPSG:4326'
target_data_type = np.float64
profile = {
'driver':'GTiff',
'height':rows,
'width':columns,
'count':1,
'dtype':target_data_type,
'crs':target_projection,
'transform':transform,
'compress':'lzw',
'nodata': no_data_val
}
with rio.open(local_edit, "w", **profile) as dst:
dst.write(data.astype(profile["dtype"]), 1)
# -
# Upload orig and edit files to s3
# +
# Original
s3_upload.upload_file(local_orig, s3_bucket, s3_key_orig,
Callback=ProgressPercentage(local_orig))
# Edit
s3_upload.upload_file(local_edit, s3_bucket, s3_key_edit,
Callback=ProgressPercentage(local_edit))
| ResourceWatchCode/Raster Dataset Processing/Raster Prep Notebooks/foo.005 (update metadata - matches with area, production, and yield, need to update crops and technologies).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This notebook makes co-occurance searches based on the outputs of T1.1.1.
# It makes bipartite and tripartite coincidences at the single sentence and single block level
# Correlation plots are produced to indicate co-occurence frequencies.
# If data is missing, it is download from google drive.
# initial commit by <NAME> (<EMAIL>) for the CoronaWhy project.
import numpy as np
import pylab
import pandas as pd
import json
import os
import requests
# -
def DownloadFiller(file, url):
if not os.path.isfile(file):
myfile = requests.get(url)
open(file, 'wb').write(myfile.content)
DownloadFiller("./TitleAbstractMatches_therapies.csv", "https://docs.google.com/uc?export=download&id=1zcEfIGYgbqQrsS_DVm-e8IsJRM3dhc5w")
DownloadFiller("./TitleAbstractMatches_drugs.csv", "https://docs.google.com/uc?export=download&id=1dQsWY5gyKtaAYlJprkeWq-nF697uFymK")
DownloadFiller("./TitleAbstractMatches_exps.csv", "https://docs.google.com/uc?export=download&id=1zcEfIGYgbqQrsS_DVm-e8IsJRM3dhc5w")
DownloadFiller("./TitleAbstractMatches_virusnames.csv", "https://docs.google.com/uc?export=download&id=1e-52CS4zX8qXk9euUE0fzVQAWITSmWoS")
# Load CSV files
dat_therapies=pd.read_csv("./TitleAbstractMatches_therapies.csv")
dat_drugs= pd.read_csv("./TitleAbstractMatches_drugs.csv")
dat_viruses= pd.read_csv("./TitleAbstractMatches_virusnames.csv")
dat_exps= pd.read_csv("./TitleAbstractMatches_exptypes.csv")
# Drop pointless column
dat_drugs=dat_drugs.drop('Unnamed: 0',axis=1).set_index('block')
dat_therapies=dat_therapies.drop('Unnamed: 0',axis=1).set_index('block')
dat_viruses=dat_viruses.drop('Unnamed: 0',axis=1).set_index('block')
dat_exps=dat_exps.drop('Unnamed: 0',axis=1).set_index('block')
# With the new extended drug lexicon, we'll need to restrict to a
# subset for good visuals
drugsubset=["naproxen","clarithromycin","chloroquine","kaletra","Favipiravir","Avigan",'hydroxychloroquine','baricitinib']
# +
# We'll use this function later to see if two words are in the same sentence
# within the block
def SameSentenceCheck(block,pos1,pos2):
if(pos1<pos2):
Interstring=block[int(pos1):int(pos2)]
else:
Interstring=block[int(pos2):int(pos1)]
SentenceEnders=[".",";","?","!"]
for s in SentenceEnders:
if s in Interstring:
return 0
return 1
# +
def Make2DPlot(dat_joined, factor1, factor2, single_sentence_plots=False):
if(single_sentence_plots):
grouped = dat_joined[dat_joined.same_sentence==True].groupby(['word_'+factor1,'word_'+factor2])
else:
grouped = dat_joined.groupby(['word_'+factor1,'word_'+factor2])
Values = grouped.count().values[:,0]
Index=grouped.count().index
Index1=[]
Index2=[]
for i in Index:
Index1.append(i[0])
Index2.append(i[1])
Uniq1=np.unique(Index1)
Uniq2=np.unique(Index2)
for i in range(0,len(Index1)):
Index1[i]=np.where(Index1[i]==Uniq1)[0][0]
Index2[i]=np.where(Index2[i]==Uniq2)[0][0]
pylab.figure(figsize=(5,5),dpi=200)
hist=pylab.hist2d(Index1,Index2, (range(0,len(Uniq1)+1),range(0,len(Uniq2)+1)), weights=Values,cmap='Blues')
pylab.xticks(np.arange(0,len(Uniq1))+0.5, Uniq1,rotation=90)
pylab.yticks(np.arange(0,len(Uniq2))+0.5, Uniq2)
pylab.clim(0,np.max(hist[0])*1.5)
for i in range(0,len(Uniq1)):
for j in range(0,len(Uniq2)):
pylab.text(i+0.5,j+0.5,int(hist[0][i][j]),ha='center',va='center')
pylab.colorbar()
if(single_sentence_plots):
pylab.title(factor1+" and " +factor2+" in One Sentence")
pylab.tight_layout()
pylab.savefig("Overlap"+factor1+"_Vs_"+factor2+"_2D_sentence.png",bbox_inches='tight',dpi=200)
else:
pylab.title(factor1+" and " +factor2+" in One Block")
pylab.tight_layout()
pylab.savefig("Overlap"+factor1+"_Vs_"+factor2+"_2D_block.png",bbox_inches='tight',dpi=200)
# -
# # Virus / Therapy word coincidences
# +
# Prune and join, and extract overlap counts
dat_joined_vt=dat_therapies.join(dat_viruses, rsuffix='_virus',lsuffix="_therapy")
dat_joined_vt=dat_joined_vt[dat_joined_vt.notna().word_therapy & dat_joined_vt.notna().word_virus]
#Make single sentence index
dat_joined_vt=dat_joined_vt.drop(["sha_therapy","blockid_therapy","sec_therapy"],axis=1).reset_index().rename(columns={"sha_virus":"sha","blockid_virus":"blockid","sec_virus":"sec"})
SingleSentence=[]
for i in dat_joined_vt.index:
SingleSentence.append(SameSentenceCheck(dat_joined_vt.block[i],dat_joined_vt.pos_virus[i],dat_joined_vt.pos_therapy[i]))
dat_joined_vt.insert(len(dat_joined_vt.columns),'same_sentence',SingleSentence)
dat_joined_vt.to_csv("Overlaps_Virus_Therapy.csv")
# -
Make2DPlot(dat_joined_vt,"virus","therapy")
Make2DPlot(dat_joined_vt,"virus","therapy",single_sentence_plots=True)
# # Virus / Drug coincidences
#
# +
# Prune and join, and extract overlap counts
dat_joined_vd=dat_drugs.join(dat_viruses, rsuffix='_virus',lsuffix="_drug")
dat_joined_vd=dat_joined_vd[dat_joined_vd.notna().word_drug & dat_joined_vd.notna().word_virus]
dat_joined_vd=dat_joined_vd.drop(["sha_drug","blockid_drug","sec_drug"],axis=1).reset_index().rename(columns={"sha_virus":"sha","blockid_virus":"blockid","sec_virus":"sec"})
SingleSentence=[]
for i in dat_joined_vd.index:
SingleSentence.append(SameSentenceCheck(dat_joined_vd.block[i],dat_joined_vd.pos_drug[i],dat_joined_vd.pos_drug[i]))
dat_joined_vd.insert(len(dat_joined_vd.columns),'same_sentence',SingleSentence)
dat_joined_vd.to_csv("Overlaps_Virus_Drug.csv")
# -
Make2DPlot(dat_joined_vd[dat_joined_vd.word_drug.isin(drugsubset)],"virus","drug")
Make2DPlot(dat_joined_vd[dat_joined_vd.word_drug.isin(drugsubset)],"virus","drug",single_sentence_plots=True)
# # Drug / Therapy Coincidences
# +
# Prune and join, and extract overlap counts
dat_joined_dt=dat_drugs.join(dat_therapies, rsuffix='_therapy',lsuffix="_drug")
dat_joined_dt=dat_joined_dt[dat_joined_dt.notna().word_drug & dat_joined_dt.notna().word_therapy]
dat_joined_dt=dat_joined_dt.drop(["sha_drug","blockid_drug","sec_drug"],axis=1).reset_index().rename(columns={"sha_therapy":"sha","blockid_therapy":"blockid","sec_therapy":"sec"})
SingleSentence=[]
for i in dat_joined_dt.index:
SingleSentence.append(SameSentenceCheck(dat_joined_dt.block[i],dat_joined_dt.pos_drug[i],dat_joined_dt.pos_therapy[i]))
dat_joined_dt.insert(len(dat_joined_dt.columns),'same_sentence',SingleSentence)
dat_joined_dt.to_csv("Overlaps_Drug_Therapy.csv")
# -
Make2DPlot(dat_joined_dt[dat_joined_dt.word_drug.isin(drugsubset)],"drug","therapy")
Make2DPlot(dat_joined_dt[dat_joined_dt.word_drug.isin(drugsubset)],"drug","therapy",single_sentence_plots=True)
# # Experiment Type and Drug
# +
# Prune and join, and extract overlap counts
dat_joined_de=dat_drugs.join(dat_exps, rsuffix='_exp',lsuffix="_drug")
dat_joined_de=dat_joined_de[dat_joined_de.notna().word_drug & dat_joined_de.notna().word_exp]
dat_joined_de=dat_joined_de.drop(["sha_drug","blockid_drug","sec_drug"],axis=1).reset_index().rename(columns={"sha_exp":"sha","blockid_exp":"blockid","sec_exp":"sec"})
SingleSentence=[]
for i in dat_joined_de.index:
SingleSentence.append(SameSentenceCheck(dat_joined_de.block[i],dat_joined_de.pos_drug[i],dat_joined_de.pos_exp[i]))
dat_joined_de.insert(len(dat_joined_de.columns),'same_sentence',SingleSentence)
dat_joined_de.to_csv("Overlaps_Drug_Experiment.csv")
# -
Make2DPlot(dat_joined_de[dat_joined_de.word_drug.isin(drugsubset)],"drug","exp")
Make2DPlot(dat_joined_de[dat_joined_de.word_drug.isin(drugsubset)],"drug","exp",single_sentence_plots=True)
# # Tripartite Coincidences
dat_joined_vtd=dat_therapies.join(dat_viruses, rsuffix='_virus',lsuffix="_therapy").join(dat_drugs)
dat_joined_vtd=dat_joined_vtd[dat_joined_vtd.notna().word_therapy & dat_joined_vtd.notna().word_virus & dat_joined_vtd.notna().word]
grouped_vtd=dat_joined_vtd.groupby(['word_therapy','word_virus','word'])
grouped_vtd.count().sha_therapy
dat_joined_vtd=dat_joined_vtd.reset_index().drop(['sha_therapy','blockid_therapy','sec_therapy','sha_virus','blockid_virus','sec_virus'],axis=1).rename(columns={'word':'word_drug','pos':'pos_drug'}).set_index('sha')
dat_joined_vtd=dat_joined_vtd[["block","sec","blockid","word_therapy","pos_therapy","word_virus", "pos_virus","word_drug","pos_drug"]]
dat_joined_vtd.to_csv("Overlaps_Drug_Therapy_Virus.csv")
| drug_treatment_extraction/notebooks/Task_1T.1.2.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3 (4 threads)
# language: julia
# name: julia-1.0k
# ---
# ## CÁLCULOS COM MATEMÁTICA SIMBÓLICA
#
# A computação simbólica trabalha com o cálculo de objetos matemáticos simbolicamente, no qual é possível manipular expressões matemáticas e realizar cálculos numéricos. Dessa forma podemos realizar operações matemáticas, como por exemplo encontrar as raízes de uma expressão $ax^2 + bx -c = 0$ de forma exata. A abordagem simbólica é de domínio dos Sistemas de Álgebra Computacional (CAS em ingles) e é trabalhado através de programas como o Mathematica®, o Maple® e o Maxima. Julia possui o pacote `SymPy.jl` que permite utilizar o SymPy do Python via pacote `Pycall.jl` para realizar calculos de matemática simbólica. Dessa forma, é possível fatorar números inteiros e polinômios, resolver sistemas lineares e não lineares, operar com números complexos, simplificar expressões, calcular limites, integrais e diferenciar funções, resolver EDOs de primeira ordem, resolver grande parte das EDOs lineares de segunda ordem, além outras operações e funções. Julia também suporta uma série de funções especiais e é capaz de criar gráficos via gnu-plot, possui métodos para resolver equações polinômiais e manipular matrizes (por exemplo, escalonar e calcular autovalores e autovetores).
# Uma ou várias variáveis simbólicas são definidas de 3 formas diferentes:
# ```julia
# @vars x y
# @syms x y
# x, y = Sym("x, y")
# ```
# Uma variável simbólica não possui valor pré-definido e dessa forma permite manipulações algébricas.
# ### CARREGANDO PACOTE E DEFININDO VARIÁVEIS###
# +
# Carregar o pacote SymPy
# +
# Definir as variáveis simbólicas x e y
# +
# verificar tipo das variáveis
# -
# ### TESTANDO O CÁLCULO SIMBÓLICO###
# +
# operação com variaveis simbólicos
# -
# ### CALCULOS COM EXPRESSÕES SIMBÓLICAS
# +
# Carregar o pacote SymPy e criar variáveis x e y
# -
# **Expansão dos termos de uma expressões**
# +
# atribuir a exp1 a expressão (x + 5)^2*(x^2 + x)
# +
# comando expand
# -
# **Fatorização**
# +
# atribuir a exp2 a expressão simbólica x^7 + x^6 + x^5 + x^4 + x^3 + x^2 + x + 1
# +
# comando factor
# -
# **Frações parciais**
# +
# atribuir a exp3 a expressão racional y3 (2*x^2 + 3*x)/(x^2 - 3*x +2)
# -
# comando apart
# **Simplificação de expressões**
# +
#comando simplify
# -
# ### CALCULO DIFERENCIAL E INTEGRAL
# +
# carregar sympy e as variáveis x e y
# +
# Definir a função f(x) = x^2 + x - 2 / x - 1
# +
# limit x-> 0
# +
# derivada primeira de x^2 - 2*x em x
# +
# derivada segunda
# +
# derivada parcial de x*y -exp(x) - y em relação a x
# +
# integral indefinida de x^2 - 2*x
# +
# integral definida de x^2 - 2*x entre x1 = 0 e x2 = 1
# +
# integração dupla definida de x*y - exp(x) - y entre y1=0 e y2=1 e x1=0 e x2=1
# -
# ### EQUAÇÕES DIFERENCIAIS
# +
using SymPy
@syms x
@symfuns y
# +
# definir edo1 sendo y'(x) + x
# +
# sol_edo recebe a Solução edo1
# +
# sol_edo_vi = Solução EDO com valores inciais y(0) = 1
# +
# ou somente a função "y" de sol_edo_vi
# +
# edo2 segunda ordem y''(x) - 4*y(x) -x
# +
# sol_edo2 recebe a Solução edo2 para y(0) = 1 e y'(0) = 0
# -
# # %%%Fim Matemática Simbólica%%%
| 4-CALCULO-MATEMATICA-SIMBOLICA-EXE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yiranamejia/AnalisisyVisualizacion/blob/master/tutorial_mitigar_bias_en_word_embeddings_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="TylXHnmHHPSm"
# # **Diagnóstico y mitigación de sesgo de género en embeddings de palabras**
#
# ## basado en el workshop https://learn.responsibly.ai/word-embedding
#
# y sobre el toolkit [`responsibly`](https://docs.responsibly.ai/) - para auditar y mitigar sesgo y obtener equidad en los sistemas de aprendizaje automático.
#
# # Descargos
#
# En este ejemplo nos enfocamos en sesgo de género simplificándolo a un fenómeno binario, pero entendemos que se trata de una sobresimplificación, una primera aproximación a la familia de soluciones de mitigación que requiere de una mayor complejidad para tratar los fenómenos de sesgo como construcciones sociales.
#
# Este material es un ejercicio puntual, no una perspectiva completa sobre sesgo en aprendizaje automático, equidad o inteligencia artificial responsable.
# + [markdown] id="r4nFGQ75HPSn"
# # Configuración
# + [markdown] id="DHn7LJEFHPSn"
# ## Instalar `responsibly`
# + id="7PNDdTweHPSn" colab={"base_uri": "https://localhost:8080/"} outputId="10a895bb-2de4-47a4-d99b-2dbb9aa6348a"
# !pip install --user responsibly
# + [markdown] id="haXHrM7eHPSn"
# ## Validar la instalación de `responsibly`
# + id="Bx-k9YV9HPSn"
import responsibly
# deberían obtener '0.1.2'
responsibly.__version__
# + [markdown] id="hwbS87m7HPSn"
# ---
#
# Si están trabajando en Colab, es normal que después de la instalación tengan el error **`ModuleNotFoundError: No module named 'responsibly'`**.
# <br/> <br/>
# Reinicien el Kernel/Runtime (usen el menú de arriba o el botón en la notebook), salteen la celda de instalación (`!pip install --user responsibly`) y ejecuten la celda previa de vuelta.
# + [markdown] id="2aN-ZeM5HPSo"
# # Jugar con el embedding de Word2Vec
#
# Con el paquete [`responsibly`](http://docs.responsibly.ai) viene la función [`responsibly.we.load_w2v_small`]() que devuelve un objeto [`KeyedVectors`](https://radimrehurek.com/gensim/models/keyedvectors.html#gensim.models.keyedvectors.KeyedVectors) de [`gensim`](https://radimrehurek.com/gensim/). Este modelo fue entrenado con Google News - 100B tokens, vocabulario de 3 millones, vectores de 300 dimensiones, sólo nos quedamos con el vocabulario en minúscula.
#
# Para más información: [Word2Vec](https://code.google.com/archive/p/word2vec/) -
#
# ## Propiedades Básicas
# + id="3MbuyNyFHPSo"
# ignorar warnings
# en general no queremos hacerlo pero ahora nos queremos enfocar en otra cosa
import warnings
warnings.filterwarnings('ignore')
# + id="o2yChPuMHPSo"
from responsibly.we import load_w2v_small
w2v_small = load_w2v_small()
# + id="MWa0pheRHPSo"
# tamanio del vocabulario
len(w2v_small.vocab)
# + id="tilWEJIDHPSo"
# obtener el vector de la palabra "home"
print('home =', w2v_small['home'])
# + id="X6DWNcVsHPSo"
# la dimensión del embedding de la palabra, en este caso, es 300
len(w2v_small['home'])
# + id="g5i6Nd3EHPSo"
# todas las palabras están normalizadas (=tienen una norma igual a uno como vectores)
from numpy.linalg import norm
norm(w2v_small['home'])
# + id="gfkBhxYuHPSo"
# asegurémonos que todos los vectores están normalizados
from numpy.testing import assert_almost_equal
length_vectors = norm(w2v_small.vectors, axis=1)
assert_almost_equal(actual=length_vectors,
desired=1,
decimal=5)
# + [markdown] id="HhCem0fsHPSo"
# ## Medir la similitud entre palabras
#
# Usaremos el [coseno](https://es.wikipedia.org/wiki/Similitud_coseno) como medida de similitud (o distancia) entre palabras.
# - Mide el coseno del ángulo entre dos vectores.
# - Rango entre 1 (vectores idénticos) y -1 (vectores opuestos).
# - En Python, para vectores normalizados (arrays de Numpy), usar el operador `@`
# + id="d-7nbBcsHPSo"
w2v_small['cat'] @ w2v_small['cat']
# + id="wwBriungHPSo"
w2v_small['cat'] @ w2v_small['cats']
# + id="LU2_chyBHPSo"
from math import acos, degrees
degrees(acos(w2v_small['cat'] @ w2v_small['cats']))
# + id="JmmYgVTyHPSo"
w2v_small['cat'] @ w2v_small['dog']
# + id="pZ4QGX3QHPSo"
degrees(acos(w2v_small['cat'] @ w2v_small['dog']))
# + id="_vAGtxd4HPSo"
w2v_small['cat'] @ w2v_small['cow']
# + id="uO9Ymx-CHPSo"
degrees(acos(w2v_small['cat'] @ w2v_small['cow']))
# + id="9EXNejAxHPSo"
w2v_small['cat'] @ w2v_small['graduated']
# + id="wLR5i4QIHPSo"
degrees(acos(w2v_small['cat'] @ w2v_small['graduated']))
# + [markdown] id="UqHoqJnQHPSo"
# ## Visualización del Word Embedding usando T-SNE
#
# <small>fuente: [Google's Seedbank](https://research.google.com/seedbank/seed/pretrained_word_embeddings)</small>
# + id="HSaC_nyXHPSo"
from sklearn.manifold import TSNE
from matplotlib import pylab as plt
# take the most common words in the corpus between 200 and 600
words = [word for word in w2v_small.index2word[200:600]]
# convert the words to vectors
embeddings = [w2v_small[word] for word in words]
# perform T-SNE
words_embedded = TSNE(n_components=2).fit_transform(embeddings)
# ... and visualize!
plt.figure(figsize=(20, 20))
for i, label in enumerate(words):
x, y = words_embedded[i, :]
plt.scatter(x, y)
plt.annotate(label, xy=(x, y), xytext=(5, 2), textcoords='offset points',
ha='right', va='bottom', size=11)
plt.show()
# + [markdown] id="2laNOxSAHPSo"
# ### Extra: [Tensorflow Embedding Projector](http://projector.tensorflow.org)
# + [markdown] id="I0orOF4LHPSo"
# ## Palabras más semejantes
#
# Cuáles son las palabras más semejantes a una determinada palabra?
# + id="Kr8o2o_THPSo"
w2v_small.most_similar('cat')
# + [markdown] id="tQLpS8IlHPSp"
# ### EXTRA: Cuál es la palabra que desentona?
#
# Dada una lista de palabras, cuál desentona? Es decir, cuál es la que está más lejos de la media de palabras.
# + id="tr3hQvF0HPSp"
w2v_small.doesnt_match('breakfast cereal dinner lunch'.split())
# + [markdown] id="vT_RDdLFHPSp"
# ## Suma de palabras
#
# 
#
# <small>fuente: [Wikipedia](https://commons.wikimedia.org/wiki/File:Vector_add_scale.svg)</small>
# + id="safkv9RCHPSp"
# nature + science = ?
w2v_small.most_similar(positive=['nature', 'science'])
# + [markdown] id="QgRMunP4HPSp"
# ## Analogía de vectores
# + [markdown] id="Zo1smCA6HPSp"
# 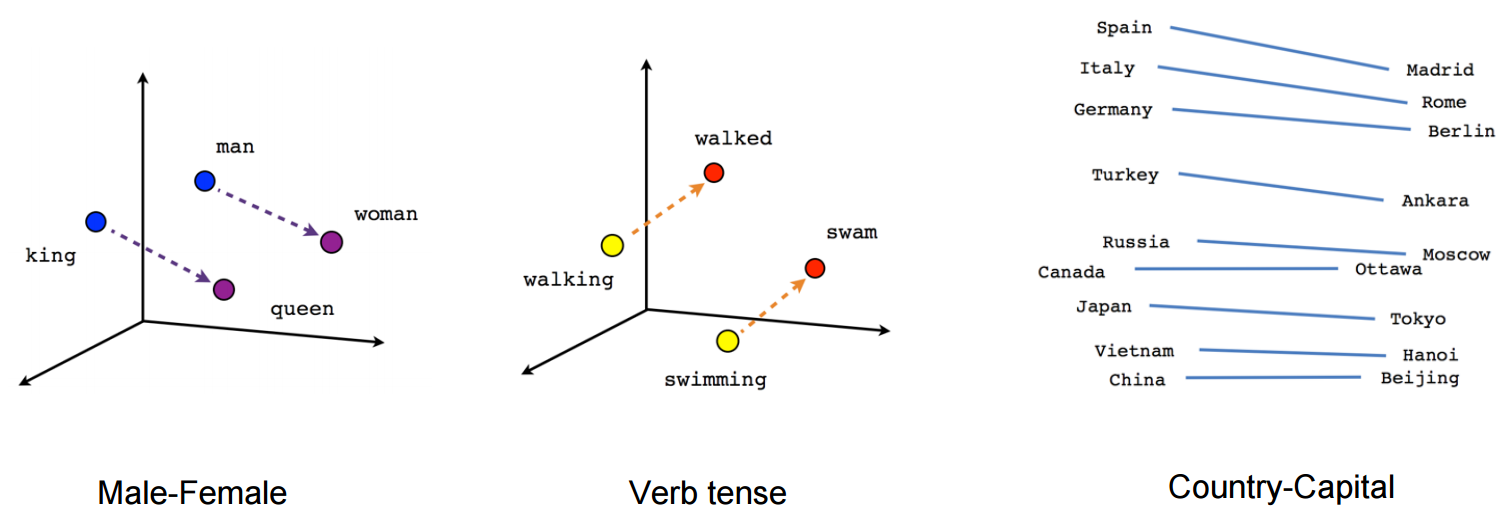
# <small>fuente: [Documentación de Tensorflow](https://www.tensorflow.org/tutorials/representation/word2vec)</small>
# + id="VMEgOEhgHPSp"
# man:king :: woman:?
# king - man + woman = ?
w2v_small.most_similar(positive=['king', 'woman'],
negative=['man'])
# + id="DudawGFKHPSp"
w2v_small.most_similar(positive=['big', 'smaller'],
negative=['small'])
# + [markdown] id="ja00u1VDHPSp"
# ## La dirección de un embedding puede verse como una relación
#
# # $\overrightarrow{she} - \overrightarrow{he}$
# # $\overrightarrow{smaller} - \overrightarrow{small}$
# # $\overrightarrow{Spain} - \overrightarrow{Madrid}$
#
# + [markdown] id="n9OBIpv2HPSp"
# # Diagnosticamos sesgo de género en embeddings
#
# <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>. [Man is to computer programmer as woman is to homemaker? debiasing word embeddings](https://arxiv.org/abs/1607.06520). NIPS 2016.
#
# ¿Cómo afecta el sesgo de género en embeddings en el contexto de aplicaciones downstream?
#
# 
#
# <small>fuente: <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., ... & <NAME>. (2019). [Mitigating Gender Bias in Natural Language Processing: Literature Review](https://arxiv.org/pdf/1906.08976.pdf). arXiv preprint arXiv:1906.08976.</small>
#
# + [markdown] id="FPSnOYQIHPSp"
# ## Probemos algunas propiedades con expresiones que sabemos que están fuertemente marcadas por el género
#
#
# + id="AEDgqRlFHPSp"
# she:sister :: he:?
# sister - she + he = ?
w2v_small.most_similar(positive=['sister', 'he'],
negative=['she'])
# + [markdown] id="pLRbogzEHPSp"
# ```
# queen-king
# waitress-waiter
# sister-brother
# mother-father
# ovarian_cancer-prostate_cancer
# convent-monastery
# ```
# + id="AlErzh68HPSp"
w2v_small.most_similar(positive=['nurse', 'he'],
negative=['she'])
# + [markdown] id="K74Iz-14HPSp"
# ```
# sewing-carpentry
# nurse-doctor
# blond-burly
# giggle-chuckle
# sassy-snappy
# volleyball-football
# register_nurse-physician
# interior_designer-architect
# feminism-conservatism
# vocalist-guitarist
# diva-superstar
# cupcakes-pizzas
# housewife-shopkeeper
# softball-baseball
# cosmetics-pharmaceuticals
# petite-lanky
# charming-affable
# hairdresser-barber
# ```
# + [markdown] id="zvfPt1T-HPSp"
# Parece que el método de generar analogías no es la forma más adecuada de observar sesgo en los embeddings, por la paradoja del observador: introduce sesgo, fuerza la producción de estereotipos de género!
#
# <NAME>., <NAME>., <NAME>. (2019). [Fair is Better than Sensational: Man is to Doctor as Woman is to Doctor](https://arxiv.org/abs/1905.09866).
#
# + [markdown] id="X2F_l7OeHPSp"
# ## Qué sí nos da la analogía? La dirección del género!
#
# # $\overrightarrow{she} - \overrightarrow{he}$
# + id="zGjRzqqbHPSp"
gender_direction = w2v_small['she'] - w2v_small['he']
gender_direction /= norm(gender_direction)
# + id="Nw3Xed-kHPSp"
gender_direction @ w2v_small['architect']
# + id="V4Ip0nTKHPSp"
gender_direction @ w2v_small['interior_designer']
# + [markdown] id="6RuU3hbEHPSp"
# Con todos los recaudos de saber que estamos sobresimplificando el fenómeno, podemos ver que la palabra *architect* aparece en más contextos con *he* que con *she*, y viceversa para *interior designer*.**
# + [markdown] id="j8uZ6qLQHPSp"
# Basándonos en esta propiedad, podemos calcular la dirección del género usano varios pares de palabras que sabemos que están fuertemente marcadas para género.:
#
# - woman - man
# - girl - boy
# - she - he
# - mother - father
# - daughter - son
# - gal - guy
# - female - male
# - her - his
# - herself - himself
# - Mary - John
# + [markdown] id="hYl3D_ATHPSp"
# ## Prueben con algunas palabras
# Reflexión: ¿están haciendo análisis exploratorio o evaluación sistemática?
# + id="BCGf0oHBHPSp"
gender_direction @ w2v_small['word']
# + [markdown] id="KshpmrI9HPSp"
# Proyecciones
# + id="PhfOAPsTHPSp"
from responsibly.we import GenderBiasWE
w2v_small_gender_bias = GenderBiasWE(w2v_small, only_lower=True)
# + id="y7OjvenQHPSp"
w2v_small_gender_bias.positive_end, w2v_small_gender_bias.negative_end
# + id="BBSW9vuSHPSq"
# dirección del género
w2v_small_gender_bias.direction[:10]
# + id="9WxA6tfvHPSq"
from responsibly.we.data import BOLUKBASI_DATA
neutral_profession_names = BOLUKBASI_DATA['gender']['neutral_profession_names']
# + id="5lSOGF5BHPSq"
neutral_profession_names[:8]
# + [markdown] id="jzjYnRJ8HPSq"
# Nota: `actor` está en la lista de nombres de profesión neutros, y no`actress` porque parece que el uso de la palabra ha cambiado con el tiempo y ahora es más neutro, en comparación por ejemplo con waiter-waitress (ver [Wikipedia - The term Actress](https://en.wikipedia.org/wiki/Actor#The_term_actress))
# + id="iI93U0JlHPSq"
len(neutral_profession_names)
# + id="1JS7UZjyHPSs"
# the same of using the @ operator on the bias direction
w2v_small_gender_bias.project_on_direction(neutral_profession_names[0])
# + [markdown] id="HDPo28WyHPSs"
# Visualicemos las proyecciones de las profesiones (neutras y específicas) en la dirección del género.
# + id="zEnirv55HPSt"
import matplotlib.pylab as plt
f, ax = plt.subplots(1, figsize=(10, 10))
w2v_small_gender_bias.plot_projection_scores(n_extreme=20, ax=ax);
# + [markdown] id="_z_VCT-eHPSt"
# EXTRA: Demo - Visualizando sesgo de género con [Nubes de palabras](http://wordbias.umiacs.umd.edu/)
# + [markdown] id="EtcH2sX_HPSt"
# Las proyecciones en la dirección de género de las palabras de profesiones se corresponden con datos de ocupación desglosados por género, según se puede ver en el porcentaje de mujeres en diversas profesiones según la encuesta de población de 2017 del Labor Force Statistics: https://arxiv.org/abs/1804.06876
# + id="BZoIxVJ4HPSt"
from operator import itemgetter # 🛠️ For idiomatic sorting in Python
from responsibly.we.data import OCCUPATION_FEMALE_PRECENTAGE
sorted(OCCUPATION_FEMALE_PRECENTAGE.items(), key=itemgetter(1))
# + id="vSrQIXF7HPSt"
f, ax = plt.subplots(1, figsize=(10, 8))
w2v_small_gender_bias.plot_factual_association(ax=ax);
# + [markdown] id="nQw3uMaDHPSt"
# <NAME>., <NAME>., <NAME>., & <NAME>. (2018). [Word embeddings quantify 100 years of gender and ethnic stereotypes](https://www.pnas.org/content/pnas/115/16/E3635.full.pdf). Proceedings of the National Academy of Sciences, 115(16), E3635-E3644.
#
# 
#
# <small>Data: Google Books/Corpus of Historical American English (COHA)</small>
# + [markdown] id="Kve5HaIUHPSt"
# ## Medición directa del sesgo
#
# 1. Proyectamos cada uno de los nombres de profesión neutros en la dirección de género
# 2. Calculamos el valor absoluto de cada proyección
# 3. Lo promediamos
# + id="OEN5PVPsHPSu"
# función de alto nivel en responsibly
w2v_small_gender_bias.calc_direct_bias()
# + id="TgtF2eUqHPSu"
# qué hace responsibly internamente:
neutral_profession_projections = [w2v_small[word] @ w2v_small_gender_bias.direction
for word in neutral_profession_names]
abs_neutral_profession_projections = [abs(proj) for proj in neutral_profession_projections]
sum(abs_neutral_profession_projections) / len(abs_neutral_profession_projections)
# + [markdown] id="hZoPjpEeHPSu"
# **Atención** la medición directa de sesgo está haciendo asunciones fuertes sobre las palabras neutras.
# + [markdown] id="uwoxH_RgHPSu"
# ## 5.10 - [EXTRA] Medición indirecta del sesgo
# Semejanza por proyección en la misma "dirección de género".
# + id="Pamry1hmHPSv"
w2v_small_gender_bias.generate_closest_words_indirect_bias('softball',
'football')
# + [markdown] id="NlS0ydglHPSv"
# # Mitigar sesgo
#
# > We intentionally do not reference the resulting embeddings as "debiased" or free from all gender bias, and
# prefer the term "mitigating bias" rather that "debiasing," to guard against the misconception that the resulting
# embeddings are entirely "safe" and need not be critically evaluated for bias in downstream tasks. <small><NAME>., & <NAME>. (2019). [Probabilistic Bias Mitigation in Word Embeddings](https://arxiv.org/pdf/1910.14497.pdf). arXiv preprint arXiv:1910.14497.</small>
#
#
# ## Neutralizar
#
# Si neutralizamos, vamos a eliminar la proyección de género de todas las palabras excepto las de género neutro, y después normalizamos.
#
# **Atención** un prerequisito fuerte es tener la lista de palabras fuertemente marcadas para género.
# + id="FGGDnQHTHPSv"
w2v_small_gender_debias = w2v_small_gender_bias.debias(method='neutralize', inplace=False)
# + id="pVoPZ9qvHPSv"
print('home:',
'before =', w2v_small_gender_bias.model['home'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['home'] @ w2v_small_gender_debias.direction)
# + id="pum8xVStHPSv"
print('man:',
'before =', w2v_small_gender_bias.model['man'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['man'] @ w2v_small_gender_debias.direction)
# + id="kiN3so-cHPSv"
print('woman:',
'before =', w2v_small_gender_bias.model['woman'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['woman'] @ w2v_small_gender_debias.direction)
# + id="hSbuQ_0nHPSv"
w2v_small_gender_debias.calc_direct_bias()
# + id="nviSfKo0HPSv"
f, ax = plt.subplots(1, figsize=(10, 10))
w2v_small_gender_debias.plot_projection_scores(n_extreme=20, ax=ax);
# + id="ytG7tYD6HPSv"
f, ax = plt.subplots(1, figsize=(10, 8))
w2v_small_gender_debias.plot_factual_association(ax=ax);
# + [markdown] id="D72h6h_-HPSv"
# ## [EXTRA] Ecualizar
#
# Las palabras en la lista de palabras marcadas para género (como por ejemplo `man` y `woman`) pueden tener una proyección diferente en la dirección de género. Eso puede resultar en una similitud diferente a palabras neutras, como `kitchen`.
# + id="9NpdcrJVHPSv"
w2v_small_gender_debias.model['man'] @ w2v_small_gender_debias.model['kitchen']
# + id="7jSEfiZPHPSv"
w2v_small_gender_debias.model['woman'] @ w2v_small_gender_debias.model['kitchen']
# + id="jEwqwZXoHPSv"
BOLUKBASI_DATA['gender']['equalize_pairs'][:10]
# + [markdown] id="4vELxkXLHPSv"
# ## Eliminación de sesgo dura: Neutralizar y Ecualizar
# + id="Hm9XmdjiHPSw"
w2v_small_gender_debias = w2v_small_gender_bias.debias(method='hard', inplace=False)
# + id="ScM9OhHwHPSw"
print('home:',
'before =', w2v_small_gender_bias.model['home'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['home'] @ w2v_small_gender_debias.direction)
# + id="VTLS5X6WHPSx"
print('man:',
'before =', w2v_small_gender_bias.model['man'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['man'] @ w2v_small_gender_debias.direction)
# + id="1YxQoXzOHPSx"
print('woman:',
'before =', w2v_small_gender_bias.model['woman'] @ w2v_small_gender_bias.direction,
'after = ', w2v_small_gender_debias.model['woman'] @ w2v_small_gender_debias.direction)
# + id="fv_ioBbkHPSx"
w2v_small_gender_debias.calc_direct_bias()
# + id="f7TlMzRUHPSx"
w2v_small_gender_debias.model['man'] @ w2v_small_gender_debias.model['kitchen']
# + id="mMtftVhxHPSx"
w2v_small_gender_debias.model['woman'] @ w2v_small_gender_debias.model['kitchen']
# + id="vLwPjbgIHPSx"
f, ax = plt.subplots(1, figsize=(10, 10))
w2v_small_gender_debias.plot_projection_scores(n_extreme=20, ax=ax);
# + [markdown] id="gYfDqt5aHPSx"
# Después de mitigar el sesgo, el rendimiento del embedding resultante en benchmarks estándares no se ve fuertemente afectado.
# + id="y3f3WQhgHPSx"
w2v_small_gender_bias.evaluate_word_embedding()
# + id="ZB_b9z16HPSx"
w2v_small_gender_debias.evaluate_word_embedding()
# + [markdown] id="TL5krvKnHPS1"
# # Explorar otros tipos de sesgo en word embeddings
# + [markdown] id="hblQa1vKHPS1"
# ### Sesgo racial
#
# Usaremos la clase [`responsibly.we.BiasWordEmbedding`](http://docs.responsibly.ai/word-embedding-bias.html#ethically.we.bias.BiasWordEmbedding). `GenderBiasWE` es una subclase de `BiasWordEmbedding`.
# + id="IXOGwjCgHPS1"
from responsibly.we import BiasWordEmbedding
w2v_small_racial_bias = BiasWordEmbedding(w2v_small, only_lower=True)
# + [markdown] id="V_m-sCQPHPS2"
# Identificar la dirección racial usando el método `sum`
# + id="-NgemvQPHPS2"
white_common_names = ['Emily', 'Anne', 'Jill', 'Allison', 'Laurie', 'Sarah', 'Meredith', 'Carrie',
'Kristen', 'Todd', 'Neil', 'Geoffrey', 'Brett', 'Brendan', 'Greg', 'Matthew',
'Jay', 'Brad']
black_common_names = ['Aisha', 'Keisha', 'Tamika', 'Lakisha', 'Tanisha', 'Latoya', 'Kenya', 'Latonya',
'Ebony', 'Rasheed', 'Tremayne', 'Kareem', 'Darnell', 'Tyrone', 'Hakim', 'Jamal',
'Leroy', 'Jermaine']
w2v_small_racial_bias._identify_direction('Whites', 'Blacks',
definitional=(white_common_names, black_common_names),
method='sum')
# + [markdown] id="BDuYsHv7HPS2"
# Usar los nombres de profesión neutros para medir el sesgo racial.
# + id="OQxxl7LIHPS2"
neutral_profession_names = BOLUKBASI_DATA['gender']['neutral_profession_names']
# + id="ZSVyfTklHPS2"
neutral_profession_names[:10]
# + id="zGMT4tNfHPS2"
f, ax = plt.subplots(1, figsize=(10, 10))
w2v_small_racial_bias.plot_projection_scores(neutral_profession_names, n_extreme=20, ax=ax);
# + [markdown] id="JPn1vjxZHPS2"
# Calcular la medida directa de sesgo
# + id="_2zrnHd6HPS2"
# Your Code Here...
# + [markdown] id="yJFRjlJ1HPS2"
# Sigan explorando el sesgo racial
# + id="toREKl5YHPS2"
# Your Code Here...
# + [markdown] id="BuF8ZOFWHPS3"
# # Recursos
#
# ## [Doing Data Science Responsibly - Resources](https://handbook.responsibly.ai/appendices/resources.html)
#
# In particular:
#
# - CVPR 2020 - [FATE Tutorial](https://youtu.be/-xGvcDzvi7Q) [Video]
#
# - fast.ai - [Algorithmic Bias (NLP video 16)](https://youtu.be/pThqge9QDn8) [Video]
#
# - <NAME>, <NAME>, <NAME> - [Fairness and machine learning - Limitations and Opportunities](https://fairmlbook.org/) [Textbook]
#
#
#
# ## Non-Technical Overview with More Downstream Application Examples
# - [Google - Text Embedding Models Contain Bias. Here's Why That Matters.](https://developers.googleblog.com/2018/04/text-embedding-models-contain-bias.html)
# - [<NAME> (UCLA) - What It Takes to Control Societal Bias in Natural Language Processing](https://www.youtube.com/watch?v=RgcXD_1Cu18)
# - <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., ... & <NAME>. (2019). [Mitigating Gender Bias in Natural Language Processing: Literature Review](https://arxiv.org/pdf/1906.08976.pdf). arXiv preprint arXiv:1906.08976.
#
# ## Additional Related Work
#
# - **Understanding Bias**
# - <NAME>., <NAME>., & <NAME>. (2019, July). [Understanding Undesirable Word Embedding Associations](https://arxiv.org/pdf/1908.06361.pdf). In Proceedings of the 57th Annual Meeting of the Association for Computational Linguistics (pp. 1696-1705). - **Including critical analysis of the current metrics and debiasing methods (quite technical)**
#
# - <NAME>., <NAME>., <NAME>., & <NAME>. (2019, May). [Understanding the Origins of Bias in Word Embeddings](https://arxiv.org/pdf/1810.03611.pdf). In International Conference on Machine Learning (pp. 803-811).
#
#
# - **Discovering Biases**
# - <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2019, January). [What are the biases in my word embedding?](https://arxiv.org/pdf/1812.08769.pdf). In Proceedings of the 2019 AAAI/ACM Conference on AI, Ethics, and Society (pp. 305-311). ACM.
# Measuring Gender Bias in Word Embeddings across Domains and Discovering New Gender Bias Word Categories
#
# - <NAME>., & <NAME>. (2019, August). [Measuring Gender Bias in Word Embeddings across Domains and Discovering New Gender Bias Word Categories](https://www.aclweb.org/anthology/W19-3804). In Proceedings of the First Workshop on Gender Bias in Natural Language Processing (pp. 25-32).
#
#
# - **Fairness in Classification**
# - <NAME>., <NAME>., & <NAME>. (2019, August). [Debiasing Embeddings for Reduced Gender Bias in Text Classification](https://arxiv.org/pdf/1908.02810.pdf). In Proceedings of the First Workshop on Gender Bias in Natural Language Processing (pp. 69-75).
#
# - <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., ... & <NAME>. (2019, June). [What's in a Name? Reducing Bias in Bios without Access to Protected Attributes](https://arxiv.org/pdf/1904.05233.pdf). In Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers) (pp. 4187-4195).
#
#
# - **Other**
#
# - <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2019, June). [Gender Bias in Contextualized Word Embeddings](https://arxiv.org/pdf/1904.03310.pdf). In Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers) (pp. 629-634). [slides](https://jyzhao.net/files/naacl19.pdf)
#
# - <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. [Analyzing and Mitigating Gender Bias in Languages with Grammatical Gender and Bilingual Word Embeddings](https://aiforsocialgood.github.io/icml2019/accepted/track1/pdfs/47_aisg_icml2019.pdf). ICML 2019 - AI for Social Good. [Poster](https://aiforsocialgood.github.io/icml2019/accepted/track1/posters/47_aisg_icml2019.pdf)
#
# - <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. [Gender Bias in Multilingual Embeddings](https://www.researchgate.net/profile/Subhabrata_Mukherjee/publication/340660062_Gender_Bias_in_Multilingual_Embeddings/links/5e97428692851c2f52a6200a/Gender-Bias-in-Multilingual-Embeddings.pdf).
#
#
# ##### Complete example of using `responsibly` with Word2Vec, GloVe and fastText: http://docs.responsibly.ai/notebooks/demo-gender-bias-words-embedding.html
#
#
# ## Bias in NLP
#
# Around dozen of papers on this field until 2019, but nowdays plenty of work is done. Two venues from back then:
# - [1st ACL Workshop on Gender Bias for Natural Language Processing](https://genderbiasnlp.talp.cat/)
# - [NAACL 2019](https://naacl2019.org/)
#
| tutorial_mitigar_bias_en_word_embeddings_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # ZapZap Awards!
#
# ### Bem vindo ao ZapZap Awards! Use as setinhas ← ↑ → ↓ pra navegar pelas paginas
# + slideshow={"slide_type": "skip"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sklearn as sk
from collections import defaultdict
import random
import requests
import json, string
import settings
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
pd.set_option('display.max_colwidth', -1)
# -
# + slideshow={"slide_type": "skip"}
df = pd.read_csv(f'{settings.GROUP_NAME}_messages.csv')
df.datetime = df.datetime.str.replace(' ', 'T')
df.loc[df.text.isnull(), 'text'] = ''
# + slideshow={"slide_type": "skip"}
df.head()
# + [markdown] slideshow={"slide_type": "skip"}
# # Normalizing dataframe
#
# + slideshow={"slide_type": "skip"}
import re
import string
def sanitize(text):
text = text.lower()
text = re.sub(r'\bkkk+', 'kkkk', text)
text = re.sub(r'\b(h[aeui]*){3,}', 'hahaha', text)
return re.sub('[{}]'.format(string.punctuation), '', text)
def shortify_links(text):
return re.sub(r'(https?://.*?/.{10})(.*?)( |$)', r'\1[...]', text)
print(sanitize('Todo dia, sièges. hahahahahaha haehaehaehea kkkkkkkk VAMO Q VAMO! 😀')) # TODO: enrich examples
df['stext'] = df.text.apply(sanitize)
df['short_links_text'] = df.text.apply(shortify_links)
df['datetime'] = pd.to_datetime(df.datetime)
df['date'] = df.datetime.dt.date
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# # <center> Estatisticas por Pessoa </center>
#
# Antes de comecarmos com os awards, vamos levantar umas estatisticas interessantes
# -
from zapzap.stats import Stats
stats = Stats(df, STOP_WORDS)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Horas favoritas
#
# Aqui temos o numero de mensagens q cada um mandou ao longo do ano, ao longo das 24h de um dia
#
# Tem gente q nao mandou uma unica mensagem as 5 da manha (no meu caso, foi shifted, para as 7)!
#
# (Aperte ↓ pra entrar nesse topico)
# -
stats.favorite_hours()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Palavras caracteristicas
#
# Quais as palavras que voce falou e que ninguem mais usa?
#
# Omiti os nomes das pessoas nesse ae. Quero ver se vcs conseguem adivinhar quem eh quem (pelo menos os seus proprios).
# + slideshow={"slide_type": "subslide"}
stats.characteristic_person()
# + [markdown] slideshow={"slide_type": "slide"}
# # <center> Estatisticas do Surfe </center>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Palavras favoritas
#
# As palavras mais usadas no surfe, por mes.
#
# Primeiro, filtradas por palavras [comuns](https://raw.githubusercontent.com/stopwords-iso/stopwords-pt/master/stopwords-pt.txt)
#
# Depois, sem filtros
# + slideshow={"slide_type": "subslide"}
stats.group_most_common_words(print_word_count=False)
# + [markdown] slideshow={"slide_type": "slide"}
# Agora, sem mais delongas...
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Single Awards
#
# Vamos a eles... os awards individuais! Ô familia, quarta feira abencoada...
#
# ## Troféu Metalinguistico - SAIU!
# Quem falou mais as palavras 'surf' e 'night'
# ## Troféu Early riser - SAIU
# Quem manda a primeira mensagem do dia (>6am)
# ## Troféu Wanderley - SAIU
# Quem grita mais? Quem usou mais caps lock em 2017
# ## Troféu Machado de Assis
# Quem tem o maior vocabulario? distinct words / words
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Troféu Pagerank
# O mais citado
# ## Troféu Feliz
# Quem ri mais?
# ## Troféu Fanfarrao
# Quem mudou mais o subject do grupo
# ## Troféu Audiófilo
# Quem mandou mais audio
# ## Troféu AlfiNET
# Quem mandou mais imagens/videos
# ## Troféu Vacuo
# Quem mandou a ultima mensagem da conversa (uma conversa) = 3h
# + [markdown] slideshow={"slide_type": "subslide"}
# # Segundo round
#
# ## Troféu NDP (sponsor Dibob)
# Autoexplicativo
# ## Troféu Popstar
# Quem foi mais chamado pelo nome próprio/apelidos
# ## Troféu Atention seeker
# Mais chamou alguém por nome proprio/apelidos
# ## Troféu Amizade
# Quem mais xingou os amiguinhos
# ## Troféu Chulo (sponsor Chulé)
# Quem mais falou palavrão
# ## Trofeu Mais Que Mil Palavras
# Quem mais usou emoji
# -
from zapzap.awards import Awards
awards = Awards(df, group_name='surf na night')
# + [markdown] slideshow={"slide_type": "slide"}
# # Troféu Metalinguistico 🏆
#
# Quem falou mais as palavras 'surf', 'night' e variantes
# -
import logging
logging.root.setLevel(logging.INFO)
awards.metalinguistic()
# + [markdown] slideshow={"slide_type": "fragment"}
# Com 19 citacoes, o ouro vai pro **BODE**! Prata pra mim e Bronze pro Chule!
#
# Eis ai um verdadeiro surfista noturno, um grande abracador da causa, sempre esta na atividade, estronda, farpacao... Um abraco mano debo! Comecando os trabalhos!
# + [markdown] slideshow={"slide_type": "slide"}
# # Troféu Early riser 🏆
# Quem mandou mais vezes a primeira mensagem do dia (>6am)
#
# + slideshow={"slide_type": "skip"}
awards.early_riser()
# + [markdown] slideshow={"slide_type": "fragment"}
# Mas que momento! Temos claramente a familia caprina vencendo no quesito quem cedo madruga. <NAME> vai levar uma medalha de bronze pra casa tambem!
#
# Ceia, uma criatura naturalmente noturna, conseguiu empatar com o nosso trio ternura do sumico. Luke, David e o lendario Guilha.
#
# Gostaria de enfatizar que ganhei do Law e do Kim. Q minha fama de dorminhoco seja revista!
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## <NAME> 🏆
# Quem grita mais? Quem usou mais caps lock em 2017
# + slideshow={"slide_type": "subslide"}
awards.shouts()
# + [markdown] slideshow={"slide_type": "fragment"}
# Cara! Achei esse resultado mt curioso!
#
# Kimzola ganhou em primeiro lugar DISPARADO!
#
# Em segundo lugar o mano CEIA, quem diria cara. Ele fala pouco, mas fala gritando.
#
# E o Dibob, provavelmente o favorito, conseguiu chegar no top 3.
#
# Parabens aos envolvidos!
# + [markdown] slideshow={"slide_type": "slide"}
#
# # To be developed...
# <img src="https://gcn.com/~/media/GIG/GCN/Redesign/Generic/AgileDevelopment.png">
# + slideshow={"slide_type": "skip"}
############################################################################################### APELIDAS #######
#display(df[df.stext.str.contains(r'\b(?:{})'.format(r'fre'), regex=True)])
df.stext.str.extract(r'.*\b({})'.format(r'fr.*?\b'), expand=False).dropna().unique()
# + slideshow={"slide_type": "skip"}
df.stext.str.contains('.*raf', regex=True)
# + slideshow={"slide_type": "skip"}
df[df.stext.str.match(r'.*\b({})'.format(r'.*o cao\b'))].head(100)
# + [markdown] slideshow={"slide_type": "skip"}
# # Single Stats
#
# ## Favorite Hours
# Normalizado por total de mensagens
# ## Palavras caracteristicas
# As palavras que so vc fala
#
#
# # Surf Stats
#
# ## Palavras favoritas
#
# ## Mensagens por hora/weekday
#
#
# # Surf Awards
#
# ## <NAME>
# Dia com maior numero de mensagens
# ## Combo
# Dias consecutivos com mais de n mensagens
# ## AntiCombo
# Dias consecutivos com 0 mensagens
# + [markdown] slideshow={"slide_type": "skip"}
# # Surf Stats
#
# ## Palavras favoritas
#
# ## Mensagens por hora/weekday
# + language="html"
# <style>
# .chat {
# display: flex;
# flex-direction: column;
# }
# .box-right {
# display: flex;
# justify-content: flex-end;
#
# }
# .box-left {
# display: flex;
# justify-content: flex-start;
# }
# .box-right-content {
# min-height: 30px;
# max-width: 40%;
# color: white;
# background: #ff974b;
# margin-right: 7px;
# padding: 10px;
# box-shadow: 1px 1px 3px 0px #CCC;
# border-radius: 10px;
# margin-bottom: 5px;
# }
# .box-left-content {
# min-height: 30px;
# max-width: 40%;
# background-color: #EDEDED;
# color: #4A4A4A;
# margin-right: 7px;
# padding: 10px;
# box-shadow: 1px 1px 3px 0px #CCC;
# border-radius: 10px;
# margin-bottom: 5px;
# }
# .chat-text {
# margin:0;
# }
# .chat-time {
# opacity: 0.6;
# font-size: 0.917em;
# line-height: 1.167em;
# margin:0 !important;
# }
# .box-info {
# display: flex;
# justify-content: center;
# }
# .box-info-content {
# background-color: #333;
# opacity: 0.5;
# padding: 5px;
# min-width: 125px;
# box-shadow: 1px 1px 3px 0px #CCC;
# border-radius: 10px;
# color: #FFF;
# text-align: center;
# font-size: 0.917em;
#
#
# }
# </style>
# -
def owner_msg(text, time):
return f'''<div class="box-right">
<div class="box-right-content"> <p class="chat-text">{text}</p> <p class="chat-time">{time}</p></div>
</div>'''
def other_msg(author, text, time):
return f'''<div class="box-left">
<div class="box-left-content"> <p class="chat-text">{author}: {text}</p> <p class="chat-time">{time}</p></div>
</div>'''
def info_msg(text):
return f'''<div class="box-info">
<div class="box-info-content"> {text} </div>
</div>'''
# +
def display_chat(df, owner):
html = ''
html += info_msg('A very nice chat!')
for idx, author, text, datetime in df[['name', 'text', 'datetime']][:50].to_records():
html += other_msg(author, text, datetime) if author != owner else owner_msg(text, datetime)
from IPython.display import display, HTML
display(HTML(html))
display_chat(df, df.name[0])
# -
| 3_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Unsupervised State-Space Modelling Using Reproducing Kernels
# ### by <NAME>
#
# The following is a demonstration of **pykssm**, a simple library for unsupervised kernel state-space modelling (KSSM), which allows the user to estimate the transition function of a nonlinear state-space model without supervision, i.e. only by looking at the output observations of the system, with no access to the internal state.
#
# The system is assumed to be of the form
# $$
# \begin{aligned}
# x_t &= f(x_{t-1}) + \eta \\
# y_t &= h(x_t) + \mu,
# \end{aligned}
# $$
# where $\eta$ and $\mu$ are random variables (usually gaussian).
#
# Then, $f(x)$ is estimated in a Reproducing Kernel Hilbert Space defined by the user, i.e. as the parametric $\hat f(x) = \sum_{i=1}^N a_i K(x, s^i), \; a_i \in \mathbb{R}$.
#
# Three systems are explored: static offline estimation, where there exists a fixed transition function $f$; time-varying online estimation, where the transition function $f_t$ changes with time; and frequency estimation, which uses real data and is estimated using more than one time step (nonlinear autoregressive model).
#
# Based on the work of <NAME>, <NAME> and <NAME>, ["Unsupervised State-Space Modeling Using Reproducing Kernels." IEEE Transactions on Signal Processing 63.19 (2015): 5210-5221.](http://ieeexplore.ieee.org/document/7130658/). If you've found the code useful for your own work, please cite the paper.
# +
# Headers
# %matplotlib inline
import sys
sys.path.append("..")
import numpy as np
import pykssm
from matplotlib import rcParams
import matplotlib.pyplot as mtp
from mpl_toolkits.mplot3d import Axes3D
rcParams['axes.labelsize'] = 18
rcParams['xtick.labelsize'] = 18
rcParams['ytick.labelsize'] = 18
rcParams['legend.fontsize'] = 18
rcParams['axes.titlesize'] = 18
rcParams['axes.labelsize'] = 18
rcParams['font.family'] = 'sans-serif'
rcParams['font.serif'] = ['Computer Modern Roman']
rcParams['font.sans-serif'] = ['Helvetica']
rcParams['mathtext.fontset'] = 'stix'
rcParams['text.usetex'] = True
rcParams['lines.markeredgewidth'] = 1.5
rcParams['lines.linewidth'] = 1.5
# %load_ext autoreload
# %autoreload 2
# #%matplotlib notebook
# -
# # Offline Modelling
#
# In the first scenario, a fixed transition function $f(x) = 10 \,\text{sinc}(\frac{x}{7})$ is used, as well as the trivial identity sensor model $h(x) = x$. KSSM is implemented to estimate $f(\cdot)$ using 40 observations.
#
# The model can be written as
# $$
# \begin{aligned}
# x_t &= f(x_{t-1}) + \eta &&= 10 \,\text{sinc}(\frac{x_{t-1}}{7}) + \mathcal{N}(0, 2^2) \\
# y_t &= h(x_t) + \mu &&= x_t + \mathcal{N}(0, 2^2).
# \end{aligned}
# $$
#
# The first state is drawn from $\mathcal{N}(0, 10)$.
# +
# Simple model
f = lambda x: 10 * np.sinc(x / 7)
h = lambda x: x
sigmax0 = np.sqrt(10)
sigmax = 1#2
sigmay = 1#2
size = 40
x0 = 0 + np.random.randn() * sigmax0
(x, y) = pykssm.filter(f, h, sigmax, sigmay, x0, size)
# Let's see the data we'll be working with,
# the output of the filter
mtp.figure(figsize=(10, 2.5))
mtp.plot(list(range(1, size + 1)), y, "o", markerfacecolor="none", markeredgecolor="b", clip_on=False)
mtp.title("Observations $y_{1:" + str(size) + "}$")
mtp.xlabel("Time [samples]")
mtp.ylabel("$y_t$")
yrange = np.max(y) - np.min(y)
mtp.axis([1, size, np.min(y) - 0.2 * yrange, np.max(y) + 0.2 * yrange])
mtp.show()
# +
# %%time
# KSSM, the core of this notebook
(samples, like, svectors, kernel) = (
pykssm.offline(observations = y[np.newaxis].T,
#svectors = np.array([[0], [10]]),
hsensor = lambda x: x,
invhsensor = lambda y: y,
kernel = pykssm.GaussianKernel(np.sqrt(10)),
nsamples = 1200,
sigmax = sigmax,
sigmay = sigmay,
smcprior = lambda: np.array([0 + np.random.randn() * sigmax0]), #prior x0
verbose = True))
# kernel (output) describes the used kernel reproducing hilbert space, with all parameters set (in this case sigma is
# set to sqrt(10) but if the argument is ommited from the input kernel argument, it would be deduced from the data);
#
# svectors are the support vectors for the representation of the transition function, i.e. the centres of the kernels
# that will be mixed together to form the estimate;
#
# samples is a list of samples from the distribution p(f_t | y_{1:T}), where f_t is represented by a list {a_i}
# of mixing parameters that weight each support vector.
# So, each sampled transition function can be calculated using the expression
# f(x) = sum(a_i * kernel(svectors[i], x)).
#
# like are the likelihoods of each sample.
#
# For this example, a gaussian kernel has been used (usually good for its universality properties),
# but it may be replaced with any other (and it's very simple to create new kernels).
# Supervised alternative (Kernel regression)
supervisedf = pykssm.kls(x[:-1][np.newaxis].T, x[1:][np.newaxis].T, svectors, kernel, 1.0)
# +
supervisedf = pykssm.kls(x[:-1][np.newaxis].T, f(x[:-1][np.newaxis].T), svectors, kernel, 0.001)
supervisedf = pykssm.kls(y[:-1][np.newaxis].T, (y[1:][np.newaxis].T), svectors, kernel, 0.000)
# +
# State transition function estimation
limit = 5.0 * np.ceil(1/5.0 * 2.0 * max(abs(min(x)), abs(max(x))))
grid = np.arange(-limit, limit, 0.025)
# Discard the first 200 samples as the MCMC converges to the desired distribution
smean = np.mean(np.array(samples[200:]), 0)
svar = np.var (np.array(samples[200:]), 0)
# Real transition function
real = [f(i) for i in grid]
# Mean estimate and its marginal deviation (note that
# since support vectors are constants and the mixture
# is a linear combination, the variance just requires
# evaluating the mixture with the weight variances)
estmean = np.array([kernel.mixture_eval(smean, svectors, [i])[0] for i in grid])
estvar = np.array([kernel.mixture_eval(svar, svectors, [i])[0] for i in grid])
eststd = np.sqrt(estvar)
estsupervised = np.array([kernel.mixture_eval(supervisedf, svectors, [i])[0] for i in grid])
mtp.figure(figsize = (8, 5))
# Observed transitions
mtp.plot(x[:-1], x[1:], "or", clip_on=False, label="State samples", markeredgewidth=0.0)
mtp.plot(y[:-1], y[1:], "og", clip_on=False, label="State samples", markeredgewidth=0.0)
mtp.plot(np.array(svectors).T[0], np.zeros(len(svectors)),
"ob", clip_on=False, label="Support vector centers", markeredgewidth=0.0)
mtp.fill_between(grid, estmean - eststd, estmean + eststd, color="b", alpha=0.4, linewidth=0.0)
mtp.plot(grid, estmean, "-b", label = "KSSM estimate")
mtp.plot(grid, estsupervised, "-g", label = "Supervised estimate")
mtp.plot(grid, real, "-k", label = "True function")
mtp.legend(loc='center left', bbox_to_anchor=(1, 0.5))
mtp.title("State transition function estimation")
mtp.xlabel("$x_t$")
mtp.ylabel("$x_{t+1}$")
mtp.xlim(np.min(grid), np.max(grid))
mtp.show()
# +
# Posterior evolution
# Supervised alternative
# supervisedlike = []
#for i in range(0, len(samples)):
# sfilter = pykssm.SMC(observations = y[np.newaxis].T,
# prior = lambda: np.array([0 + np.random.randn() * sigmax0]),
# ftransition = lambda x: kernel.mixture_eval(supervisedf, svectors, x) + np.random.randn() * sigmax,
# hsensor = lambda x, y: 1.0 / (np.sqrt(2.0 * np.pi)*sigmay) * np.exp(-0.5 * np.dot(x - y, x - y) / sigmay**2),
# nsamples = 200)
# supervisedlike.append(sfilter.get_likelihood())
sfilter = pykssm.SMC(observations = y[np.newaxis].T,
prior = lambda: np.array([0 + np.random.randn() * sigmax0]),
ftransition = lambda x: kernel.mixture_eval(supervisedf, svectors, x) + np.random.randn() * sigmax,
hsensor = lambda x, y: 1.0 / (np.sqrt(2.0 * np.pi)*sigmay) * np.exp(-0.5 * np.dot(x - y, x - y) / sigmay**2),
nsamples = 200)
supervisedlike = sfilter.get_likelihood() * np.ones(len(samples))
mtp.figure(figsize = (20, 6))
mtp.semilogy(like, "-b", label="MCMC samples")
mtp.semilogy(supervisedlike, "-r", label="Supervised solution")
mtp.legend(loc='center right')
mtp.title("Posterior evolution")
mtp.xlabel("Time [samples]")
mtp.ylabel("Posterior probability")
mtp.show()
# -
mtp.figure(figsize=(12,6))
samples.shape
supervisedf.shape
mtp.plot(samples[:,:,0],samples[:,:,1])
mtp.scatter(supervisedf[:,0],supervisedf[:,1],color='r')
# # Online Modelling
#
# In the second scenario, the transition function varies in time, following the equation
# $$
# f(x) = \left\{
# \begin{array}{ll}
# \frac{x}{2} + 25 \frac{x}{1 + x^2} & \mbox{if } x < 30 \\
# \frac{60 - t}{30} \cdot (\frac{x}{2} + 25 \frac{x}{1 + x^2}) + \frac{t - 30}{30} \cdot 10 \, \text{sinc}(\frac{x}{7}) & \mbox{if } 30 \leq x \leq 60 \\
# 10 \, \text{sinc}(\frac{x}{7}) & \mbox{if } x > 60.
# \end{array}
# \right.
# $$
#
# The sensor model is a linear function of the state, $h(x) = \frac{x}{2} + 5$. The system is run for 90 iterations.
#
# The model can be written as
# $$
# \begin{aligned}
# x_t &= f_t(x_{t-1}) + \eta &&= f_t(x_{t-1}) + \mathcal{N}(0, 2^2) \\
# y_t &= h(x_t) + \mu &&= \frac{x}{2} + 5 + \mathcal{N}(0, 2^2).
# \end{aligned}
# $$
#
# The first state is drawn from $\mathcal{N}(0, 1^2)$.
# +
# Time-varying model
def ft(t, x):
# time-invariant stable
def flow(x):
return x / 2 + 25 * x / (1 + x * x)
# time-invariant unstable (two accumulation points)
def fhigh(x):
return 10 * np.sinc(x / 7)
# linear interpolation between the previous two
def fmid(t, x):
return (60 - t) / 30 * flow(x) + (t - 30) / 30 * fhigh(x)
if t < 30:
return flow(x)
elif t > 60:
return fhigh(x)
else:
return fmid(t, x)
ht = lambda x: x / 2 + 5
sigmaxt0 = 1
sigmaxt = 1
sigmayt = np.sqrt(0.5)
sizet = 90
xt0 = 0 + np.random.randn() * sigmaxt0
(xt, yt) = pykssm.filter_t(ft, ht, sigmaxt, sigmayt, xt0, sizet)
# Output of the filter
mtp.figure(figsize=(10, 2.5))
mtp.plot(list(range(1, sizet + 1)), yt, "o", markerfacecolor="none", markeredgecolor="b", clip_on=False)
mtp.title("Observations $y_{1:" + str(sizet) + "}$")
mtp.xlabel("Time [samples]")
mtp.ylabel("$y_t$")
yrange = np.max(yt) - np.min(yt)
mtp.axis([1, sizet, np.min(yt) - 0.2 * yrange, np.max(yt) + 0.2 * yrange])
mtp.show()
# +
# Filter state transition function in time
limit = 10.0 * np.ceil(1/10.0 *2.0 * max(abs(min(xt)), abs(max(xt))))
grid = np.arange(-limit, limit, 0.025)
mtp.figure(figsize=(8, 5))
for t in range(30, 61, 3):
labelindex = str(t)
if t == 30:
labelindex = "\le 30"
elif t == 60:
labelindex = "\ge 60"
mtp.plot(grid, [ft(t, i) for i in grid], "-",
color=(0, 0.15 + 0.85 * (1 - (t - 30) / 30), 0.15 + 0.85 * (t - 30) / 30),
label="$f_{" + labelindex + "}$")
mtp.legend(loc='center left', bbox_to_anchor=(1, 0.5))
mtp.title("State transition function in time")
mtp.xlabel("$x_t$")
mtp.ylabel("$x_{t+1}$")
mtp.show()
# +
# %%time
# #%%prun -D profile.prof -q
# Time-varying KSSM, the core of this notebook
# (state transition function) transition standard deviation
sigmaf = 0.2
estimate = pykssm.online(observations = yt[np.newaxis].T,
hsensor = lambda x: (x / 2 + 5),
invhsensor = lambda y: 2 * (y - 5),
theta = lambda f1, f2: 1.0 / ((2 * np.pi)**len(f1) * sigmaf) *
np.exp(-0.5 * np.sum((f1 - f2) * (f1 - f2)) / sigmaf**2),
kernel = pykssm.GaussianKernel(),
nsamples = 400,
sigmax = sigmaxt,
sigmay = sigmayt,
smcprior = lambda: np.array([0 + np.random.randn() * sigmaxt0]),
verbose = True)
# estimate is an array of tuples of the form (samples, likelihoods, svectors, kernel), each one corresponding
# to a time step and similar to the offline case.
# +
# Time-varying state transition function estimation
limit = 10.0 * np.ceil(1/10.0 * 2.0 * max(abs(min(xt)), abs(max(xt))))
grid = np.arange(-limit, limit, 0.025)
# We'll observe the estimate at two moments: 30 and 90 seconds,
# which are the end points for both time-invariant zones.
# The estimates' first index correspond to the samples, of which
# the first 200 are discarded while the MCMC converges to the desired distribution
smean30 = np.mean(np.array(estimate[30 - 1][0][200:]), 0)
svar30 = np.var (np.array(estimate[30 - 1][0][200:]), 0)
smean90 = np.mean(np.array(estimate[89 - 1][0][200:]), 0)
svar90 = np.var (np.array(estimate[89 - 1][0][200:]), 0)
# The estimates' second index corresponds to the support vectors at that time
svectors30 = estimate[30 - 1][2]
svectors90 = estimate[89 - 1][2]
# The estimates' third index corresponds to the kernel descriptor
kernel30 = estimate[30 - 1][3]
kernel90 = estimate[89 - 1][3]
# Real transition functions
real30 = [ft(30, i) for i in grid]
real90 = [ft(89, i) for i in grid]
# Mean estimate and its marginal deviation
estmean30 = np.array([kernel30.mixture_eval(smean30, svectors30, [i])[0] for i in grid])
estvar30 = np.array([kernel30.mixture_eval(svar30, svectors30, [i])[0] for i in grid])
eststd30 = np.sqrt(estvar30)
estmean90 = np.array([kernel90.mixture_eval(smean90, svectors90, [i])[0] for i in grid])
estvar90 = np.array([kernel90.mixture_eval(svar90, svectors90, [i])[0] for i in grid])
eststd90 = np.sqrt(estvar90)
mtp.figure(figsize = (8, 5))
# First time-invariant zone estimate
mtp.fill_between(grid, estmean30 - eststd30, estmean30 + eststd30, color="g", alpha=0.4, linewidth=0.0)
mtp.plot(grid, estmean30, "-g", label = "$\hat f_{30}$, KSSM estimate")
mtp.plot(grid, real30, "-k", label = "$f_{30}$, True function")
#mtp.plot(2 * (yt[0:30] - 5), 2 * (yt[1:31] - 5), "og", clip_on=False, label="$s_{30}$, State samples")
mtp.plot(xt[0:30], xt[1:31], "og", clip_on=False, label="$s_{30}$, State samples")
#mtp.plot(np.array(svectors90).T[0], np.zeros(len(svectors90)),
# "ob", clip_on=False, label="Support vector centers", markeredgewidth=0.0)
# Second time-invariant zone estimate
mtp.fill_between(grid, estmean90 - eststd90, estmean90 + eststd90, color="b", alpha=0.4, linewidth=0.0)
mtp.plot(grid, estmean90, "-b", label = "$\hat f_{90}$, KSSM estimate")
mtp.plot(grid, real90, "--k", label = "$f_{90}$, True function")
#mtp.plot(2 * (yt[60:89] - 5), 2 * (yt[61:90] - 5), "ob", clip_on=False, label="$s_{90}$, State samples")
mtp.plot(xt[60:89], xt[61:90], "ob", clip_on=False, label="$s_{90}$, State samples")
mtp.legend(loc='center left', bbox_to_anchor=(1, 0.5))
mtp.title("State transition function estimation")
mtp.xlabel("$x_t$")
mtp.ylabel("$x_{t+1}$")
mtp.show()
# +
# Internal state of the filter
mtp.figure(figsize=(10, 2.5))
mtp.plot(list(range(1, sizet + 1)), xt, "-b")
mtp.title("State signal $x_{1:" + str(sizet) + "}$")
mtp.xlabel("Time [samples]")
mtp.ylabel("$x_t$")
xrange = np.max(xt) - np.min(xt)
mtp.axis([1, sizet, np.min(xt) - 0.2 * xrange, np.max(xt) + 0.2 * xrange])
mtp.show()
# -
# # Frequency Analysis
#
# In the third scenario, 200 data points are extracted from a real stream of frequency measurements of the [UK national grid for the day 17 July 2014](http://www.gridwatch.templar.co.uk/), which has been normalized into the $[-8, 8]$ region.
#
# To be able to measure the performance of the algorithm, these measurements have been used as groundtruth state, and measurements have been synthesized on top of them by adding additive gaussian noise from $\mathcal{N}(0, 0.25^2)$.
#
# The state transition noise has been assumed normal from $\mathcal{N}(0, 0.25^2)$.
#
# To estimate and predict the frequency just using the previous time step doesn't have much power, so this experiment will use two time steps to predict. This way there's both information about the frequency and its derivative (albeit noisy). This means that the transition function becomes multivariate, of the form
# $$
# F_t(X_t) =
# \begin{pmatrix}f_t(x_t, x_{t-1}) \\
# x_t \end{pmatrix}.
# $$
#
# The sensor model is the identity function (for the first state coordinate) with the additional noise indicated above.
#
# The full model can be written as
# $$
# \begin{aligned}
# \begin{pmatrix}x_t\\
# x_{t-1} \end{pmatrix} &= F_t\begin{pmatrix}x_{t-1}\\
# x_{t-2}\end{pmatrix} + \eta &&= \begin{pmatrix}f_t(x_t, x_{t-1}) \\
# x_t \end{pmatrix} + \mathcal{N}(0,
# \begin{pmatrix}0.25^2 & 0 \\
# 0 & 0 \end{pmatrix}) \\
# y_t &= h(x_t) + \mu &&= \begin{pmatrix}1 & 0\end{pmatrix} \begin{pmatrix}x_t \\ x_{t-1}\end{pmatrix} + \mathcal{N}(0, 0.25^2),
# \end{aligned}
# $$
#
# where $f_t(\cdot)$ is estimated using kernels, just like the previous experiments.
#
# The first state is drawn from $\mathcal{N}(0, 0.25^2)$.
#
# Note that the details of the nonlinear autoregressive model are not explicitly coded (including heuristics on the undetermined sensor function inverse). Instead the library automatically expands everything as necessary; the only required input is the number of delays to consider.
# +
# UK national grid frequency data
frequency = np.load("frequency.npy")
frequency = 50 * (frequency - 50)
delays = 1
sigmaxf0 = 0.25
sigmaxf = 0.25
sigmayf = 0.25
xf = frequency[:200]
yf = np.array([x + sigmayf * np.random.randn() for x in xf])
sizefq = len(xf)
# Plot of the input data
mtp.figure(figsize=(10, 2.5))
mtp.plot(list(range(1, sizefq + 1)), xf, "-", markerfacecolor="none", markeredgecolor="b", clip_on=False)
mtp.title("Frequency $x_{1:" + str(sizefq) + "}$")
mtp.xlabel("Time [samples]")
mtp.ylabel("$x_t$")
xrange = np.max(frequency) - np.min(frequency)
mtp.axis([1, sizefq, np.min(frequency) - 0.2 * xrange, np.max(frequency) + 0.2 * xrange])
mtp.show()
# +
# %%time
# Online estimation of frequency transition function
sigmaff = 0.2
estimate = (
pykssm.autoregressive(observations = yf[0:10][np.newaxis].T,
delays = 1,
hsensor = lambda x: x,
invhsensor = lambda y: y,
theta = lambda f1, f2: 1.0 / ((2 * np.pi)**len(f1) * sigmaff) *
np.exp(-0.5 * np.sum((f1 - f2) * (f1 - f2)) / sigmaff**2),
kernel = pykssm.GaussianKernel(),
nsamples = 400,
sigmax = sigmaxf,
sigmay = sigmayf,
smcprior = lambda: np.array([xf[0] + np.random.randn() * sigmaxf0]),
verbose = True))
# estimate is an array of tuples of the form (samples, likelihoods, svectors, kernel), each one corresponding
# to a time step and similar to the offline case.
# +
# Time-varying state transition function estimation
highlimit = max(xf[:7])
lowlimit = min(xf[:7])
span = highlimit - lowlimit
highlimit = 1.0 * np.ceil ( 1/1.0 * 0.5 * span + highlimit)
lowlimit = 1.0 * np.floor(-1/1.0 * 0.5 * span + lowlimit)
gridx = np.linspace(lowlimit, highlimit, 64)
gridy = np.linspace(lowlimit, highlimit, 64)
# We'll observe the estimate at two moments: 30 and 90 seconds,
# which are the end points for both time-invariant zones.
# The estimates' first index correspond to the samples, of which
# the first 200 are discarded while the MCMC converges to the desired distribution
smean2 = np.mean(np.array(estimate[2 - 1][0][200:]), 0)
smean3 = np.mean(np.array(estimate[3 - 1][0][200:]), 0)
smean4 = np.mean(np.array(estimate[4 - 1][0][200:]), 0)
smean7 = np.mean(np.array(estimate[7 - 1][0][200:]), 0)
# The estimates' second index corresponds to the support vectors at that time
svectors2 = estimate[2 - 1][2]
svectors3 = estimate[3 - 1][2]
svectors4 = estimate[4 - 1][2]
svectors7 = estimate[7 - 1][2]
# The estimates' third index corresponds to the kernel descriptor
kernel2 = estimate[2 - 1][3]
kernel3 = estimate[3 - 1][3]
kernel4 = estimate[4 - 1][3]
kernel7 = estimate[7 - 1][3]
# Mean estimate and its marginal deviation
estmean2 = np.array([[kernel2.mixture_eval(smean2, svectors2, [i, k])[0] for k in gridy] for i in gridx])
estmean3 = np.array([[kernel3.mixture_eval(smean3, svectors3, [i, k])[0] for k in gridy] for i in gridx])
estmean4 = np.array([[kernel4.mixture_eval(smean4, svectors4, [i, k])[0] for k in gridy] for i in gridx])
estmean7 = np.array([[kernel7.mixture_eval(smean7, svectors7, [i, k])[0] for k in gridy] for i in gridx])
mgx, mgy = np.meshgrid(gridx, gridy)
sampledx = np.concatenate(([0], xf[:-2]))
sampledy = xf[:-1]
sampledz = xf[1:]
fig = mtp.figure(figsize = (16, 10))
ax = fig.add_subplot(2, 2, 1, projection="3d")
ax.plot_wireframe(mgx, mgy, estmean2, color=(0, 0, 0, 0.6), cstride=4, rstride=4, label="$\hat f_{2}$, KSSM estimate")
ax.scatter(sampledx[0:2], sampledy[0:2], sampledz[0:2], c="b", s=40,
clip_on=False, depthshade=False, label="$s_{1:2}$, State samples")
ax.set_title("Time $t=2$")
ax = fig.add_subplot(2, 2, 2, projection="3d")
ax.plot_wireframe(mgx, mgy, estmean3, color=(0, 0, 0, 0.6), cstride=4, rstride=4, label="$\hat f_{3}$, KSSM estimate")
ax.scatter(sampledx[0:3], sampledy[0:3], sampledz[0:3], c="b", s=40,
clip_on=False, depthshade=False, label="$s_{1:3}$, State samples")
ax.set_title("Time $t=3$")
ax = fig.add_subplot(2, 2, 3, projection="3d")
ax.plot_wireframe(mgx, mgy, estmean4, color=(0, 0, 0, 0.6), cstride=4, rstride=4, label="$\hat f_{4}$, KSSM estimate")
ax.scatter(sampledx[0:4], sampledy[0:4], sampledz[0:4], c="b", s=40,
clip_on=False, depthshade=False, label="$s_{1:4}$, State samples")
ax.set_title("Time $t=4$")
ax = fig.add_subplot(2, 2, 4, projection="3d")
ax.plot_wireframe(mgx, mgy, estmean7, color=(0, 0, 0, 0.6), cstride=4, rstride=4, label="$\hat f_{7}$, KSSM estimate")
ax.scatter(sampledx[0:7], sampledy[0:7], sampledz[0:7], c="b", s=40,
clip_on=False, depthshade=False, label="$s_{1:7}$, State samples")
ax.set_title("Time $t=7$")
fig.suptitle("Frequency transition estimation", fontsize=18)
mtp.show()
| demo/Kernel State-Space Modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from scrape_abc import utils
url = "http://www.abc.org.br/membro/isaac-roitman/"
name = "<NAME>"
abc_page = utils.AbcPage(url)
abc_page.get_info()
abc_page.write_wikipage(onmc=True, woman=False)
# +
print(f"Creating page for {name}")
print(f"https://author-disambiguator.toolforge.org/names_oauth.php?precise=0&name={name.replace(' ', '+')}&doit=Look+for+author&limit=500&filter=")
# -
qid = input("Enter Wikidata QID for " + name )
abc_page.print_qs(qid, woman=False)
print(f'{qid}|P166|Q3132815|P580|+1994-04-08T00:00:00Z/11|S854|"https://web.archive.org/web/20070213055821/http://www.mct.gov.br/index.php/content/view/11199.html?area=allAreas&categoria=allMembros"')
man_abc_onc_df = man_abc_onc_df.reset_index(drop=True).drop(0)
man_abc_onc_df.head()
man_abc_onc_df.to_csv("man_to_create.csv")
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('/home/aman-py/Desktop/HackerEarth/ZS Data Science Challange/dataset/yds_train2018.csv',index_col='S_No')
df.head()
df.shape
df.info()
df.describe()
x = df[['Year','Month','Product_ID','Country_col']]
y = df[['Sales']]
print(x.head(),'\n',y.head())
from sklearn import *
x_train,x_test,y_train,y_test = cross_validation.train_test_split(x,y,test_size = 0.25,random_state=1)
clf = random_projection.BaseRandomProjection()
clf.fit(X=x_train,y=y_train)
clf.score(x_test,y_test)
p =clf.predict(x_test)
import numpy as np
type(y_test)
p.shape
y_test.shape
# +
import numpy as np
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
# +
y_true = [3, -0.5, 2, 7]; y_pred = [2.5, -0.3, 2, 8]
m = np.array(y_true).reshape(-1,1)
y_pred=np.array(y_pred).reshape(-1,1)
mean_absolute_percentage_error(p,y_test)
# -
df.loc[df['Sales']==0,:]
from sklearn import linear_model
a=np.array(y_true)
a.reshape(-1,1)
y_train.head()
k = list(df['Country'].unique())
arr = []
for i in df['Country']:
arr.append(k.index(i))
df['Country_col'] = arr
from sklearn import model_selection
knn = model_selection.
| ZS Data Science Challange/Challenge Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from pypfopt.efficient_frontier import EfficientFrontier
from pypfopt import risk_models
from pypfopt import expected_returns
import numpy as np
# -
import os
dirname = os.getcwd()
parent_dirname = os.path.dirname(dirname)
# Read in price data
df = pd.read_csv(os.path.join(parent_dirname, 'data/raw/dji_adj_price_daily.csv'),
parse_dates=True,
index_col="Date")
# Calculate daily returns
# ret = (df - df.shift()) / df.shift()
ret = np.log(df) - np.log(df.shift())
# do not drop NA
# +
# remove STI
# ret = ret.drop('^STI', axis=1)
# -
# Calculate mean and covariance of daily log return
mu = ret.mean()
S = ret.cov()
# Plot the portfolios
plot_df = pd.DataFrame({'mu':mu, 'S':np.sqrt(np.diag(S))})
plot_df
plot_df.plot.scatter(x='S', y='mu')
# Calculate correlation
r = ret.corr()
# r
# Portfolio analysis
ef = EfficientFrontier(mu, S, weight_bounds=(0, 1))
# optimize wrt max sharpe ratio
raw_weights = ef.max_sharpe(risk_free_rate=0.02)
cleaned_weights = ef.clean_weights()
# View clean weights
cleaned_weights
ef.portfolio_performance(verbose=True)
# +
# how much of each stock to buy?
capital = 10000
from pypfopt.discrete_allocation import DiscreteAllocation, get_latest_prices
latest_prices = get_latest_prices(df)
da = DiscreteAllocation(raw_weights, latest_prices, total_portfolio_value=capital)
allocation, leftover = da.lp_portfolio()
print("Discrete allocation:", allocation)
print("Funds remaining: ${:.2f}".format(leftover))
# -
| notebooks/2.0-hwant-DJI-portfolio-optimization-daily.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# [](https://github.com/awslabs/aws-data-wrangler)
#
# # 25 - Redshift - Loading Parquet files with Spectrum
# ## Enter your bucket name:
import getpass
bucket = getpass.getpass()
PATH = f"s3://{bucket}/files/"
# ## Mocking some Parquet Files on S3
# +
import awswrangler as wr
import pandas as pd
df = pd.DataFrame({
"col0": [0, 1, 2, 3, 4, 5, 6, 7, 8, 9],
"col1": ["a", "b", "c", "d", "e", "f", "g", "h", "i", "j"],
})
df
# -
wr.s3.to_parquet(df, PATH, max_rows_by_file=2, dataset=True, mode="overwrite");
# ## Crawling the metadata and adding into Glue Catalog
wr.s3.store_parquet_metadata(
path=PATH,
database="aws_data_wrangler",
table="test",
dataset=True,
mode="overwrite"
)
# ## Running the CTAS query to load the data into Redshift storage
con = wr.redshift.connect(connection="aws-data-wrangler-redshift")
query = "CREATE TABLE public.test AS (SELECT * FROM aws_data_wrangler_external.test)"
with con.cursor() as cursor:
cursor.execute(query)
# ## Running an INSERT INTO query to load MORE data into Redshift storage
df = pd.DataFrame({
"col0": [10, 11],
"col1": ["k", "l"],
})
wr.s3.to_parquet(df, PATH, dataset=True, mode="overwrite");
query = "INSERT INTO public.test (SELECT * FROM aws_data_wrangler_external.test)"
with con.cursor() as cursor:
cursor.execute(query)
# ## Checking the result
query = "SELECT * FROM public.test"
wr.redshift.read_sql_table(con=con, schema="public", table="test")
con.close()
| tutorials/025 - Redshift - Loading Parquet files with Spectrum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geo_env
# language: python
# name: geo_env
# ---
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import plotly.express as px
from sklearn.model_selection import train_test_split
import statsmodels.api as sm
# -
df = pd.read_csv("../data/raw/data.csv")
df.head()
fig,axs = plt.subplots(1,2)
sm.qqplot(df.LoanAmount,line="q",ax=axs[0])
sm.qqplot(np.log(df.LoanAmount),line="q",ax=axs[1])
plt.show()
fig,axs = plt.subplots(1,2)
sm.qqplot(df.ApplicantIncome,line="q",ax=axs[0])
sm.qqplot(np.log(df.ApplicantIncome),line="q",ax=axs[1])
plt.show()
df.LoanAmount = np.log(df.LoanAmount)
df["TotalApplicantIncome"] = np.log(df.ApplicantIncome+df.CoapplicantIncome)
df.ApplicantIncome = np.log(df.ApplicantIncome)
df.drop(columns=['CoapplicantIncome','Loan_ID','Loan_Amount_Term'],inplace=True)
df.to_csv(r'../data/interim/2_feature_engineered/1_base_data.csv',index=False)
| notebooks/3_data_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 反向传播算法
#
# ## 激活函数导数
#
# ### Sigmoid函数导数
#
# Sigmoid函数表达式:$$\sigma(x) = \frac{1}{1 + e^{-x}}$$
# Sigmoid函数的导数表达式:$$\frac{d}{dx} \sigma(x) = \sigma(1-\sigma)$$
# +
# 导入 numpy 库
import numpy as np
from matplotlib import pyplot as plt
plt.rcParams['font.size'] = 16
plt.rcParams['font.family'] = ['STKaiti']
plt.rcParams['axes.unicode_minus'] = False
def set_plt_ax():
# get current axis 获得坐标轴对象
ax = plt.gca()
ax.spines['right'].set_color('none')
# 将右边 上边的两条边颜色设置为空 其实就相当于抹掉这两条边
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
# 指定下边的边作为 x 轴,指定左边的边为 y 轴
ax.yaxis.set_ticks_position('left')
# 指定 data 设置的bottom(也就是指定的x轴)绑定到y轴的0这个点上
ax.spines['bottom'].set_position(('data', 0))
ax.spines['left'].set_position(('data', 0))
def sigmoid(x):
# 实现 sigmoid 函数
return 1 / (1 + np.exp(-x))
def sigmoid_derivative(x):
# sigmoid 导数的计算
# sigmoid 函数的表达式由手动推导而得
return sigmoid(x)*(1-sigmoid(x))
# +
x = np.arange(-6.0, 6.0, 0.1)
sigmoid_y = sigmoid(x)
sigmoid_derivative_y = sigmoid_derivative(x)
set_plt_ax()
plt.plot(x, sigmoid_y, color='C9', label='Sigmoid')
plt.plot(x, sigmoid_derivative_y, color='C4', label='导数')
plt.xlim(-6, 6)
plt.ylim(0, 1)
plt.legend(loc=2)
plt.show()
# -
# ### ReLU 函数导数
# ReLU 函数的表达式:$$\text{ReLU}(x)=\max(0,x)$$
# ReLU 函数的导数表达式:$$\frac{d}{dx} \text{ReLU} = \left \{
# \begin{array}{cc}
# 1 \quad x \geqslant 0 \\
# 0 \quad x < 0
# \end{array} \right.$$
# +
def relu(x):
return np.maximum(0, x)
def relu_derivative(x): # ReLU 函数的导数
d = np.array(x, copy=True) # 用于保存梯度的张量
d[x < 0] = 0 # 元素为负的导数为 0
d[x >= 0] = 1 # 元素为正的导数为 1
return d
# +
x = np.arange(-6.0, 6.0, 0.1)
relu_y = relu(x)
relu_derivative_y = relu_derivative(x)
set_plt_ax()
plt.plot(x, relu_y, color='C9', label='ReLU')
plt.plot(x, relu_derivative_y, color='C4', label='导数')
plt.xlim(-6, 6)
plt.ylim(0, 6)
plt.legend(loc=2)
plt.show()
# -
# ### LeakyReLU函数导数
#
# LeakyReLU 函数的表达式:$$\text{LeakyReLU} = \left\{ \begin{array}{cc}
# x \quad x \geqslant 0 \\
# px \quad x < 0
# \end{array} \right.$$
#
# LeakyReLU的函数导数表达式:$$\frac{d}{dx} \text{LeakyReLU} = \left\{ \begin{array}{cc}
# 1 \quad x \geqslant 0 \\
# p \quad x < 0
# \end{array} \right.$$
# +
def leakyrelu(x, p):
y = np.copy(x)
y[y < 0] = p * y[y < 0]
return y
# 其中 p 为 LeakyReLU 的负半段斜率,为超参数
def leakyrelu_derivative(x, p):
dx = np.ones_like(x) # 创建梯度张量,全部初始化为 1
dx[x < 0] = p # 元素为负的导数为 p
return dx
# +
x = np.arange(-6.0, 6.0, 0.1)
p = 0.1
leakyrelu_y = leakyrelu(x, p)
leakyrelu_derivative_y = leakyrelu_derivative(x, p)
set_plt_ax()
plt.plot(x, leakyrelu_y, color='C9', label='LeakyReLU')
plt.plot(x, leakyrelu_derivative_y, color='C4', label='导数')
plt.xlim(-6, 6)
plt.yticks(np.arange(-1, 7))
plt.legend(loc=2)
plt.show()
# -
# ### Tanh 函数梯度
#
# tanh函数的表达式:$$\tanh(x)=\frac{e^x-e^{-x}}{e^x + e^{-x}}= 2 \cdot \text{sigmoid}(2x) - 1$$
# tanh函数的导数表达式:$$
# \begin{aligned}
# \frac{\mathrm{d}}{\mathrm{d} x} \tanh (x) &=\frac{\left(e^{x}+e^{-x}\right)\left(e^{x}+e^{-x}\right)-\left(e^{x}-e^{-x}\right)\left(e^{x}-e^{-x}\right)}{\left(e^{x}+e^{-x}\right)^{2}} \\
# &=1-\frac{\left(e^{x}-e^{-x}\right)^{2}}{\left(e^{x}+e^{-x}\right)^{2}}=1-\tanh ^{2}(x)
# \end{aligned}
# $$
def sigmoid(x): # sigmoid 函数实现
return 1 / (1 + np.exp(-x))
def tanh(x): # tanh 函数实现
return 2*sigmoid(2*x) - 1
def tanh_derivative(x): # tanh 导数实现
return 1-tanh(x)**2
# +
x = np.arange(-6.0, 6.0, 0.1)
tanh_y = tanh(x)
tanh_derivative_y = tanh_derivative(x)
set_plt_ax()
plt.plot(x, tanh_y, color='C9', label='Tanh')
plt.plot(x, tanh_derivative_y, color='C4', label='导数')
plt.xlim(-6, 6)
plt.ylim(-1.5, 1.5)
plt.legend(loc=2)
plt.show()
# -
# ## 链式法则
# +
import tensorflow as tf
# 构建待优化变量
x = tf.constant(1.)
w1 = tf.constant(2.)
b1 = tf.constant(1.)
w2 = tf.constant(2.)
b2 = tf.constant(1.)
# 构建梯度记录器
with tf.GradientTape(persistent=True) as tape:
# 非 tf.Variable 类型的张量需要人为设置记录梯度信息
tape.watch([w1, b1, w2, b2])
# 构建 2 层线性网络
y1 = x * w1 + b1
y2 = y1 * w2 + b2
# 独立求解出各个偏导数
dy2_dy1 = tape.gradient(y2, [y1])[0]
dy1_dw1 = tape.gradient(y1, [w1])[0]
dy2_dw1 = tape.gradient(y2, [w1])[0]
# 验证链式法则, 2 个输出应相等
print(dy2_dy1 * dy1_dw1)
print(dy2_dw1)
# -
# ## Himmelblau 函数优化实战
#
# Himmelblau 函数是用来测试优化算法的常用样例函数之一,它包含了两个自变量$x$和$y$,数学表达式是:$$
# f(x, y)=\left(x^{2}+y-11\right)^{2}+\left(x+y^{2}-7\right)^{2}
# $$
# +
from mpl_toolkits.mplot3d import Axes3D
def himmelblau(x):
# himmelblau 函数实现,传入参数 x 为 2 个元素的 List
return (x[0] ** 2 + x[1] - 11) ** 2 + (x[0] + x[1] ** 2 - 7) ** 2
# -
x = np.arange(-6, 6, 0.1) # 可视化的 x 坐标范围为-6~6
y = np.arange(-6, 6, 0.1) # 可视化的 y 坐标范围为-6~6
print('x,y range:', x.shape, y.shape)
# 生成 x-y 平面采样网格点,方便可视化
X, Y = np.meshgrid(x, y)
print('X,Y maps:', X.shape, Y.shape)
Z = himmelblau([X, Y]) # 计算网格点上的函数值
# 绘制 himmelblau 函数曲面
fig = plt.figure('himmelblau')
ax = fig.gca(projection='3d') # 设置 3D 坐标轴
ax.plot_surface(X, Y, Z, cmap = plt.cm.rainbow ) # 3D 曲面图
ax.view_init(60, -30)
ax.set_xlabel('x')
ax.set_ylabel('y')
plt.show()
# +
# 参数的初始化值对优化的影响不容忽视,可以通过尝试不同的初始化值,
# 检验函数优化的极小值情况
# [1., 0.], [-4, 0.], [4, 0.]
# 初始化参数
x = tf.constant([4., 0.])
for step in range(200):# 循环优化 200 次
with tf.GradientTape() as tape: #梯度跟踪
tape.watch([x]) # 加入梯度跟踪列表
y = himmelblau(x) # 前向传播
# 反向传播
grads = tape.gradient(y, [x])[0]
# 更新参数,0.01 为学习率
x -= 0.01*grads
# 打印优化的极小值
if step % 20 == 19:
print ('step {}: x = {}, f(x) = {}'.format(step, x.numpy(), y.numpy()))
# -
# ## 反向传播算法实战
# +
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.datasets import make_moons
from sklearn.model_selection import train_test_split
plt.rcParams['font.size'] = 16
plt.rcParams['font.family'] = ['STKaiti']
plt.rcParams['axes.unicode_minus'] = False
# -
def load_dataset():
# 采样点数
N_SAMPLES = 2000
# 测试数量比率
TEST_SIZE = 0.3
# 利用工具函数直接生成数据集
X, y = make_moons(n_samples=N_SAMPLES, noise=0.2, random_state=100)
# 将 2000 个点按着 7:3 分割为训练集和测试集
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=TEST_SIZE, random_state=42)
return X, y, X_train, X_test, y_train, y_test
def make_plot(X, y, plot_name, XX=None, YY=None, preds=None, dark=False):
# 绘制数据集的分布, X 为 2D 坐标, y 为数据点的标签
if (dark):
plt.style.use('dark_background')
else:
sns.set_style("whitegrid")
plt.figure(figsize=(16, 12))
axes = plt.gca()
axes.set(xlabel="$x_1$", ylabel="$x_2$")
plt.title(plot_name, fontsize=30)
plt.subplots_adjust(left=0.20)
plt.subplots_adjust(right=0.80)
if XX is not None and YY is not None and preds is not None:
plt.contourf(XX, YY, preds.reshape(XX.shape), 25, alpha=1, cmap=plt.cm.Spectral)
plt.contour(XX, YY, preds.reshape(XX.shape), levels=[.5], cmap="Greys", vmin=0, vmax=.6)
# 绘制散点图,根据标签区分颜色
plt.scatter(X[:, 0], X[:, 1], c=y.ravel(), s=40, cmap=plt.cm.Spectral, edgecolors='none')
plt.show()
X, y, X_train, X_test, y_train, y_test = load_dataset()
# 调用 make_plot 函数绘制数据的分布,其中 X 为 2D 坐标, y 为标签
make_plot(X, y, "Classification Dataset Visualization ")
class Layer:
# 全连接网络层
def __init__(self, n_input, n_neurons, activation=None, weights=None,
bias=None):
"""
:param int n_input: 输入节点数
:param int n_neurons: 输出节点数
:param str activation: 激活函数类型
:param weights: 权值张量,默认类内部生成
:param bias: 偏置,默认类内部生成
"""
# 通过正态分布初始化网络权值,初始化非常重要,不合适的初始化将导致网络不收敛
self.weights = weights if weights is not None else np.random.randn(n_input, n_neurons) * np.sqrt(1 / n_neurons)
self.bias = bias if bias is not None else np.random.rand(n_neurons) * 0.1
self.activation = activation # 激活函数类型,如’sigmoid’
self.last_activation = None # 激活函数的输出值o
self.error = None # 用于计算当前层的delta 变量的中间变量
self.delta = None # 记录当前层的delta 变量,用于计算梯度
# 网络层的前向传播函数实现如下,其中last_activation 变量用于保存当前层的输出值:
def activate(self, x):
# 前向传播函数
r = np.dot(x, self.weights) + self.bias # X@W+b
# 通过激活函数,得到全连接层的输出o
self.last_activation = self._apply_activation(r)
return self.last_activation
# 上述代码中的self._apply_activation 函数实现了不同类型的激活函数的前向计算过程,
# 尽管此处我们只使用Sigmoid 激活函数一种。代码如下:
def _apply_activation(self, r):
# 计算激活函数的输出
if self.activation is None:
return r # 无激活函数,直接返回
# ReLU 激活函数
elif self.activation == 'relu':
return np.maximum(r, 0)
# tanh 激活函数
elif self.activation == 'tanh':
return np.tanh(r)
# sigmoid 激活函数
elif self.activation == 'sigmoid':
return 1 / (1 + np.exp(-r))
return r
# 针对于不同类型的激活函数,它们的导数计算实现如下:
def apply_activation_derivative(self, r):
# 计算激活函数的导数
# 无激活函数,导数为1
if self.activation is None:
return np.ones_like(r)
# ReLU 函数的导数实现
elif self.activation == 'relu':
grad = np.array(r, copy=True)
grad[r > 0] = 1.
grad[r <= 0] = 0.
return grad
# tanh 函数的导数实现
elif self.activation == 'tanh':
return 1 - r ** 2
# Sigmoid 函数的导数实现
elif self.activation == 'sigmoid':
return r * (1 - r)
return r
# 神经网络模型
class NeuralNetwork:
def __init__(self):
self._layers = [] # 网络层对象列表
def add_layer(self, layer):
# 追加网络层
self._layers.append(layer)
# 网络的前向传播只需要循环调各个网络层对象的前向计算函数即可,代码如下:
# 前向传播
def feed_forward(self, X):
for layer in self._layers:
# 依次通过各个网络层
X = layer.activate(X)
return X
def backpropagation(self, X, y, learning_rate):
# 反向传播算法实现
# 前向计算,得到输出值
output = self.feed_forward(X)
for i in reversed(range(len(self._layers))): # 反向循环
layer = self._layers[i] # 得到当前层对象
# 如果是输出层
if layer == self._layers[-1]: # 对于输出层
layer.error = y - output # 计算2 分类任务的均方差的导数
# 关键步骤:计算最后一层的delta,参考输出层的梯度公式
layer.delta = layer.error * layer.apply_activation_derivative(output)
else: # 如果是隐藏层
next_layer = self._layers[i + 1] # 得到下一层对象
layer.error = np.dot(next_layer.weights, next_layer.delta)
# 关键步骤:计算隐藏层的delta,参考隐藏层的梯度公式
layer.delta = layer.error * layer.apply_activation_derivative(layer.last_activation)
# 循环更新权值
for i in range(len(self._layers)):
layer = self._layers[i]
# o_i 为上一网络层的输出
o_i = np.atleast_2d(X if i == 0 else self._layers[i - 1].last_activation)
# 梯度下降算法,delta 是公式中的负数,故这里用加号
layer.weights += layer.delta * o_i.T * learning_rate
def train(self, X_train, X_test, y_train, y_test, learning_rate, max_epochs):
# 网络训练函数
# one-hot 编码
y_onehot = np.zeros((y_train.shape[0], 2))
y_onehot[np.arange(y_train.shape[0]), y_train] = 1
# 将One-hot 编码后的真实标签与网络的输出计算均方误差,并调用反向传播函数更新网络参数,循环迭代训练集1000 遍即可
mses = []
accuracys = []
for i in range(max_epochs + 1): # 训练1000 个epoch
for j in range(len(X_train)): # 一次训练一个样本
self.backpropagation(X_train[j], y_onehot[j], learning_rate)
if i % 10 == 0:
# 打印出MSE Loss
mse = np.mean(np.square(y_onehot - self.feed_forward(X_train)))
mses.append(mse)
accuracy = self.accuracy(self.predict(X_test), y_test.flatten())
accuracys.append(accuracy)
print('Epoch: #%s, MSE: %f' % (i, float(mse)))
# 统计并打印准确率
print('Accuracy: %.2f%%' % (accuracy * 100))
return mses, accuracys
def predict(self, X):
return self.feed_forward(X)
def accuracy(self, X, y):
return np.sum(np.equal(np.argmax(X, axis=1), y)) / y.shape[0]
nn = NeuralNetwork() # 实例化网络类
nn.add_layer(Layer(2, 25, 'sigmoid')) # 隐藏层 1, 2=>25
nn.add_layer(Layer(25, 50, 'sigmoid')) # 隐藏层 2, 25=>50
nn.add_layer(Layer(50, 25, 'sigmoid')) # 隐藏层 3, 50=>25
nn.add_layer(Layer(25, 2, 'sigmoid')) # 输出层, 25=>2
mses, accuracys = nn.train(X_train, X_test, y_train, y_test, 0.01, 1000)
# +
x = [i for i in range(0, 101, 10)]
# 绘制MES曲线
plt.title("MES Loss")
plt.plot(x, mses[:11], color='blue')
plt.xlabel('Epoch')
plt.ylabel('MSE')
plt.show()
# -
# 绘制Accuracy曲线
plt.title("Accuracy")
plt.plot(x, accuracys[:11], color='blue')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.show()
| tensorflow_v2/dragen1860/ch07/ch07-反向传播算法.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importando librerías necesarias
import requests
import os
from bs4 import BeautifulSoup
from humanfriendly import format_timespan
import time
begin_time = time.time()
# Pegar la URL de cada módulo aquí: notar que la url va entre comillas simples '
module0_url = 'https://delftxdownloads.tudelft.nl/EnerTran1x_Energy_Markets_of_Today/Module_0/'
module1_url = 'https://delftxdownloads.tudelft.nl/EnerTran1x_Energy_Markets_of_Today/Module_1/'
module2_url = "https://delftxdownloads.tudelft.nl/EnerTran1x_Energy_Markets_of_Today/Module_2/"
module3_url = 'https://delftxdownloads.tudelft.nl/EnerTran1x_Energy_Markets_of_Today/Module_3/'
module4_url = 'https://delftxdownloads.tudelft.nl/EnerTran1x_Energy_Markets_of_Today/Module_4/'
all_urls = [module0_url, module1_url, module2_url, module3_url, module4_url]
print('---> DESCARGANDO EL CURSO "ENERGY MARKETS OF TODAY" DE TUDELFT <---')
course_folder = 'Energy Markets of Today - TUDelft'
# Creando la carpeta general de curso
try:
os.mkdir(course_folder)
print("")
except FileExistsError:
print("")
# Generando licencia de software
with open('LICENCE.txt', "w") as file:
file.write('MIT License\n\nCopyright (c) 2020 <NAME>\n\nPermission is hereby granted, free of charge, to any person obtaining a copy\nof this software and associated documentation files (the "Software"), to deal\nin the Software without restriction, including without limitation the rights\nto use, copy, modify, merge, publish, distribute, sublicense, and/or sell\ncopies of the Software, and to permit persons to whom the Software is\nfurnished to do so, subject to the following conditions:\n\nThe above copyright notice and this permission notice shall be included in all\ncopies or substantial portions of the Software.\n\nTHE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR\nIMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,\nFITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE\nAUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER\nLIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,\nOUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE\nSOFTWARE.')
# Generando licencia de curso
with open('Energy Markets of Today - TUDelft\TUDelft Copyright.txt', "w") as file:
file.write('Unless otherwise specified the Course Materials of this course are Copyright Delft University of Technology and are licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.\n\nFor further information: https://creativecommons.org/licenses/by-nc-sa/4.0/')
# ----------------------------------------------------------------------
# Descarga archivos para cada url (módulo)
for module_url in all_urls:
# Creando carpetas para módulos
import os
dirName = module_url[-9:-1].upper()
dirName = dirName.replace("O", "Ó")
dirName = dirName.replace("E", "O")
dirName = dirName.replace("_", " ")
try:
os.mkdir('Energy Markets of Today - TUDelft/{}'.format(dirName))
print("----------------------------------------------")
print("LA CARPETA DEL '" + dirName.upper() + "' HA SIDO CREADA")
except FileExistsError:
print("----------------------------------------------")
print("LA CARPETA DEL'" + dirName.upper() + "' YA EXISTÍA")
# ----------------------------------------------------------------------
def get_file_links():
# Crear objeto de respuesta de los módulos del curso
r = requests.get(module_url)
# Crear objeto beautiful-soup
soup = BeautifulSoup(r.content,'html5lib')
# Encuentra links dentro de la página web
links = soup.findAll('a')
# Seleccionando archivos con las siguientes terminaciones: 720.mp4, .srt y slides.pdf
file_links = [module_url + link['href'] for link in links
if link['href'].endswith('360.mp4') or link['href'].endswith('.srt') or link['href'].endswith('slides.pdf')]
return file_links
# ----------------------------------------------------------------------
def download_file_series(file_links):
for link in file_links:
'''itera a través de todos los enlaces en file_links
y descarga uno por uno'''
# Obteniendo el nombre de cada archivo dividiendo la URL con / y obteniendo la última parte
file_name = link.split('/')[-1]
# Texto que se muestra en la descarga
if file_name[-3:]=='srt':
print("Descargando subtítulo: " + file_name)
elif file_name[-3:]=='pdf':
print("Descargando diapositiva: " + file_name)
elif file_name[-3:]=='mp4':
print("Descargando video: " + file_name)
# Crear objeto de respuesta de cada archivo del curso
r = requests.get(link, stream = True)
# Empieza la descarga
with open('{}/{}/{}'.format(course_folder, dirName, file_name), 'wb') as f:
for chunk in r.iter_content(chunk_size = 1024*1024):
if chunk:
f.write(chunk)
print("Terminado")
print("EL "+ dirName.upper()+ " HA SIDO DESCARGADO\n")
return
# ----------------------------------------------------------------------
if __name__ == "__main__":
# Seleccionando los archivos
file_links = get_file_links()
# Descargando todos los archivos
download_file_series(file_links)
print('______________________________________________\n')
print('---> ¡CURSO "ENERGY MARKETS OF TODAY" DESCARGADO! <---\n')
# muestra el tiempo de ejecución
end_time = time.time() - begin_time
print("Tiempo total de descarga: ", format_timespan(end_time))
print(input("\nPresione ENTER para salir del programa: "))
| DescargarCursoEDX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# =========================================================
# Principal components analysis (PCA)
# =========================================================
#
# These figures aid in illustrating how a point cloud
# can be very flat in one direction--which is where PCA
# comes in to choose a direction that is not flat.
#
#
#
# +
print(__doc__)
# Authors: <NAME>
# <NAME>
# <NAME>
# License: BSD 3 clause
from sklearn.decomposition import PCA
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
# #############################################################################
# Create the data
e = np.exp(1)
np.random.seed(4)
def pdf(x):
return 0.5 * (stats.norm(scale=0.25 / e).pdf(x)
+ stats.norm(scale=4 / e).pdf(x))
y = np.random.normal(scale=0.5, size=(30000))
x = np.random.normal(scale=0.5, size=(30000))
z = np.random.normal(scale=0.1, size=len(x))
density = pdf(x) * pdf(y)
pdf_z = pdf(5 * z)
density *= pdf_z
a = x + y
b = 2 * y
c = a - b + z
norm = np.sqrt(a.var() + b.var())
a /= norm
b /= norm
# #############################################################################
# Plot the figures
def plot_figs(fig_num, elev, azim):
fig = plt.figure(fig_num, figsize=(4, 3))
plt.clf()
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=elev, azim=azim)
ax.scatter(a[::10], b[::10], c[::10], c=density[::10], marker='+', alpha=.4)
Y = np.c_[a, b, c]
# Using SciPy's SVD, this would be:
# _, pca_score, V = scipy.linalg.svd(Y, full_matrices=False)
pca = PCA(n_components=3)
pca.fit(Y)
pca_score = pca.explained_variance_ratio_
V = pca.components_
x_pca_axis, y_pca_axis, z_pca_axis = V.T * pca_score / pca_score.min()
x_pca_axis, y_pca_axis, z_pca_axis = 3 * V.T
x_pca_plane = np.r_[x_pca_axis[:2], - x_pca_axis[1::-1]]
y_pca_plane = np.r_[y_pca_axis[:2], - y_pca_axis[1::-1]]
z_pca_plane = np.r_[z_pca_axis[:2], - z_pca_axis[1::-1]]
x_pca_plane.shape = (2, 2)
y_pca_plane.shape = (2, 2)
z_pca_plane.shape = (2, 2)
ax.plot_surface(x_pca_plane, y_pca_plane, z_pca_plane)
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
elev = -40
azim = -80
plot_figs(1, elev, azim)
elev = 30
azim = 20
plot_figs(2, elev, azim)
plt.show()
| lab10/decomposition/plot_pca_3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
# %load_ext autoreload
# %autoreload 1
from ercollect import molecule as mol
from ercollect.molecule import molecule
from ercollect import rxn_syst
from ercollect.rxn_syst import reaction, get_RS
import numpy as np
import random
import os
import requests
from rdkit.Chem import Draw
from rdkit.Chem import AllChem as Chem
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem.Draw.MolDrawing import MolDrawing, DrawingOptions
from IPython.display import clear_output
from ercollect import SABIO_IO
# Author: <NAME>
#
# Date Created: 08 Dec 2018
#
# Distributed under the terms of the MIT License.
# # Notebook to clean up collected KEGG entries
# This notebook contains some fixes for a bug in the code that did not translate the roles of molecules properly when using an already collected KEGG molecule
# # PROBLEM 1:
# ## 08/12/18
# - found a bug in KEGG_IO that overwrote the role of a given component when translating to an existing molecule using the KEGG ID translator
# ## Step 1:
# - For each RS, check components roles compared to that downloaded from KEGG.
# - if they differ, then rewrite role
# - set RS properties to None
# - IMPORTANT!
# - in the cases where the same molecule exists as a reactant and product then there is no trivial way to make sure this fix is done properly. So those RS are deleted to be recollected
directory = '/home/atarzia/psp/screening_results/'
directory += 'new_reactions_kegg_atlas/'
# directory += 'new_reactions_sabio_wcharge/'
# directory += 'biomin_search_sabio_wcharge/'
# + code_folding=[2]
for i, rs in enumerate(rxn_syst.yield_rxn_syst_filelist(output_dir=directory,
filelist=directory+'file_list_1.txt')):
delete = False
print(rs.pkl)
if rs.components is None:
continue
# get Reaction information from KEGG API
URL = 'http://rest.kegg.jp/get/reaction:'+rs.DB_ID
request = requests.post(URL)
if request.text != '':
request.raise_for_status()
# because of the formatting of KEGG text - this is trivial
equations_string = request.text.split('EQUATION ')[1].split('\n')[0].rstrip()
if '<=>' in equations_string:
# implies it is reversible
reactants, products = equations_string.split("<=>")
# print("reaction is reversible")
else:
reactants, products = equations_string.split("=")
reactants = [i.lstrip().rstrip() for i in reactants.split("+")]
products = [i.lstrip().rstrip() for i in products.split("+")]
# collect KEGG Compound/Glycan ID
# check if reactant or product are compound or glycan
# remove stoichiometry for now
comp_list = []
for r in reactants:
if 'G' in r:
# is glycan
KID = 'G'+r.split('G')[1].rstrip()
elif 'C' in r:
# is compound
KID = 'C'+r.split('C')[1].rstrip()
elif 'D' in r:
# is drug
KID = 'D'+r.split('D')[1].rstrip()
comp_list.append((KID, 'reactant'))
for r in products:
if 'G' in r:
# is glycan
KID = 'G'+r.split('G')[1].rstrip()
elif 'C' in r:
# is compound
KID = 'C'+r.split('C')[1].rstrip()
elif 'D' in r:
# is drug
KID = 'D'+r.split('D')[1].rstrip()
comp_list.append((KID, 'product'))
for m in rs.components:
count_matched = 0
for comp in comp_list:
if comp[0] == m.KEGG_ID:
m.role = comp[1]
count_matched += 1
if count_matched > 1:
print(rs.pkl, '-- has same molecule on both sides')
delete = True
continue
if delete is True:
os.system('rm '+directory+rs.pkl)
elif delete is False:
rs.save_object(directory+rs.pkl)
print(i, 'done')
# -
# ## Step 2:
# - set RS properties to None
for i, rs in enumerate(rxn_syst.yield_rxn_syst(output_dir=directory)):
rs.delta_comp = None
rs.delta_sa = None
rs.max_XlogP = rs.max_comp_size = rs.max_logP = rs.min_logP = rs.min_XlogP = None
rs.p_max_comp = rs.r_max_comp = rs.p_max_sa = rs.r_max_sa = None
print(i, 'done')
rs.save_object(directory+rs.pkl)
| archived_code/notebooks/clean_up_KEGG_RS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # STORE SALES PREDICTIONS
# # 1. BUSINESS UNDERSTANDING
# ### 1.1. Business objectives
# Store managers are tasked with **predicting their daily sales for up to six weeks in advance**.
# ### 1.2. Assess situation
# Store sales are influenced by many factors, including promotions, competition, school and state holidays, seasonality, and locality. With thousands of individual managers predicting sales based on their unique circumstances, the accuracy of results can be quite varied.
# ### 1.3. Project goals
# This project intends to build a regression model to predict daily sales up to 6 weeks ahead using machine learning algorithms.
# # 2. DATA UNDERSTANDING
# ### 2.1. Import libraries and helper functions
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import inflection
import pickle
import xgboost as xgb
from sklearn.preprocessing import StandardScaler, MinMaxScaler, RobustScaler, LabelEncoder
from sklearn.linear_model import LinearRegression, Lasso
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error, mean_absolute_percentage_error, mean_squared_error
from sklearn.model_selection import TimeSeriesSplit
from sklearn.model_selection import RandomizedSearchCV
from boruta import BorutaPy
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.rcParams['figure.figsize']=[20,10]
plt.rcParams['font.size']=10
sns.set()
jupyter_settings()
# -
# ### 2.2. Collect initial data
# +
# Loading data
df_raw_store = pd.read_csv('../data/store.csv', low_memory=False)
df_raw_train = pd.read_csv('../data/train.csv', low_memory=False)
# Merging
df = pd.merge(df_raw_train, df_raw_store, how='left', on='Store')
# Snakecase pattern
columns_old = list(df.columns)
snakecase = lambda x: inflection.underscore(x)
columns_new = map(snakecase, columns_old)
df.columns = columns_new
# Dataset
df1 = df.copy()
# -
# ### 2.3. Describe data
# +
# Visualizing dataset
display(df1.head())
display(df1.tail())
# Data info
display(df1.info())
# -
# ### 2.4. Explore data
# ### 2.4.1. Numerical attributes
# Numerical attributes
num_attributes = df1.select_dtypes( include = ['int64','float64'] )
display(num_attributes.describe().T)
num_attributes.hist(bins=50);
# ### 2.4.2. Datetime attribute
# +
# Datetime attributes
df1['date'] = pd.to_datetime( df1['date'])
# Lowest date
print('Lowest date: {}'.format(df1['date'].min() ) )
# Biggest date
print('Biggest date: {}'.format(df1['date'].max() ) )
#Visualizing date by sales
aux = df1[['date', 'sales']].groupby('date').sum().reset_index()
sns.scatterplot(x='date', y='sales', data=aux)
# -
# ### 2.4.3. Categorical attributes
# +
# Categorical attributes
cat_attributes = df1.select_dtypes( exclude = ['int64','float64', 'datetime'] )
display( cat_attributes.apply( lambda x: x.unique().shape[0] ) )
# Visualizing categorical by sales
aux = df1[ df1['sales'] > 0 ]
plt.subplot(2, 2, 1)
sns.boxplot(x='state_holiday', y= 'sales', data= aux)
plt.subplot(2, 2, 2)
sns.boxplot(x='store_type', y= 'sales', data= aux)
plt.subplot(2, 2, 3)
sns.boxplot(x='assortment', y= 'sales', data= aux)
plt.subplot(2, 2, 4)
sns.boxplot(x='promo_interval', y= 'sales', data= aux)
# -
# ### 2.5. Verify data quality
# Checking NA
df1.isna().sum()
# # 3. DATA PREPARATION
# ### 3.1. Select data
# +
# Dataset
df2 = df1.copy()
# Dropping 'customers' -> It won't be available on the moment of predictions (test dataset)
df2.drop('customers', axis=1, inplace=True)
# Excluding rows where 'open'== 0 and 'sales' == 0 -> the store wasn't working or there are no sales
df2 = df2[ (df2['open'] != 0) & (df2['sales'] > 0)]
# Dropping 'open' -> It is not necessary anymore
df2.drop('open', axis=1, inplace=True)
# -
# ### 3.2. Clean data
# +
# Fillout NAs
# ['competition_distance'] -> NA can means there is no competitor store near.
# Filling with the highest value (75860)
df2['competition_distance'] = df2['competition_distance'].apply( lambda x: 75860 if math.isnan(x) else x )
# ['competition_open_since_(month/year)']
# Filling with the month/year of sale's date ('date'.month / 'date'.year)
df2['competition_open_since_month'] = df2.apply( lambda x: x['date'].month if math.isnan(x['competition_open_since_month']) else x['competition_open_since_month'], axis=1)
df2['competition_open_since_year'] = df2.apply( lambda x: x['date'].year if math.isnan(x['competition_open_since_year']) else x['competition_open_since_year'], axis=1)
# ['promo2_since_(year/week)'] -> NA occur when store is not participating to 'promo2' (promo2 == 0)
# Filling with the year/week of sale's date ('date'.year / 'date'.week)
df2['promo2_since_year'] = df2.apply( lambda x: x['date'].year if math.isnan(x['promo2_since_year']) else x['promo2_since_year'], axis=1)
df2['promo2_since_week'] = df2.apply( lambda x: x['date'].week if math.isnan(x['promo2_since_week']) else x['promo2_since_week'], axis=1)
# ['promo_interval'] -> NA occur when store is not participating to 'promo2' (promo2 == 0)
# Filling with zero
df2['promo_interval'].fillna(0, inplace=True)
# -
# ### 3.3. Construct data
# +
# Dataset
df3 = df2.copy()
# Deriving 'date' of sale
df3['year'] = df3['date'].apply( lambda x: x.year )
df3['month'] = df3['date'].apply( lambda x: x.month )
df3['day'] = df3['date'].apply( lambda x: x.day )
# Deriving 'competition_open_since_(month/year)' to indicate the time of competition related to date of sale
df3['competition_open_since'] = df3.apply(lambda x: datetime.datetime(
day=1,
month= int(x['competition_open_since_month']),
year= int(x['competition_open_since_year'])), axis=1 )
df3['competition_time'] = df3['competition_open_since'] - df3['date']
# Deriving 'promo2_since_(week/year)' to indicate the time of participation in the promo2 related to date of sale
df3['promo2_since'] = df3.apply( lambda x: str(int(x['promo2_since_year'])) + '-' + str(int(x['promo2_since_week'])), axis=1 )
df3['promo2_since'] = df3.apply( lambda x: datetime.datetime.strptime( x['promo2_since']+'-1', '%Y-%W-%w' ), axis=1 )
df3['promo2_time'] = df3['promo2_since'] - df3['date']
# Deriving 'promo_interval' to indicate if promo2 was activated on date of sale
dict_promo_interval = {'Jan,Apr,Jul,Oct': [1, 4, 7, 10],
'Feb,May,Aug,Nov': [2, 5, 8, 11],
'Mar,Jun,Sept,Dec': [3, 6, 9, 12],
0: [0]}
df3['promo_interval'] = df3['promo_interval'].map(dict_promo_interval)
df3['promo2_activated'] = df3.apply( lambda x: 1 if ( x['month'] in x['promo_interval'] ) else 0, axis=1 )
# Dropping attributes that have been replaced by derived variables
cols_to_drop = ['competition_open_since_month','competition_open_since_year', 'competition_open_since',
'promo2_since_week', 'promo2_since_year', 'promo2_since', 'promo_interval']
df3.drop(cols_to_drop, axis=1, inplace=True)
# -
# ### 3.4. Format data
# +
# Reorganizing dataset
df4 = df3.copy()
df4 = df4[['store', 'sales', 'date', 'year', 'month', 'day', 'day_of_week', 'state_holiday',
'school_holiday', 'store_type', 'assortment', 'competition_distance', 'competition_time',
'promo', 'promo2', 'promo2_time', 'promo2_activated']]
#Verifying types
df4.dtypes
# -
# Changing data type - converting timedelta to integer
df4['competition_time'] = df4['competition_time'].apply( lambda x: int(x.days) )
df4['promo2_time'] = df4['promo2_time'].apply( lambda x: int(x.days) )
# ### 3.5. Rescaling
# +
# Dataset
df5 = df4.copy()
# Robust Scaler
robust_scaler = RobustScaler()
df5[['competition_distance', 'competition_time', 'promo2_time']] = robust_scaler.fit_transform( df5[['competition_distance', 'competition_time', 'promo2_time']] )
# Min Max Scaler
minmax_scaler = MinMaxScaler()
df5['year'] = minmax_scaler.fit_transform( df5['year'].values.reshape(-1,1) )
# -
# ### 3.6. Encoding
# +
# 'state_holiday' - one hot encoding
df5 = pd.get_dummies( df5, prefix=['state_holiday'], columns=['state_holiday'])
# 'store_type' - label encoding
df5['store_type'] = LabelEncoder().fit_transform( df5['store_type'] )
# 'assortment' - ordinal encoding
assortment_dict = {'a':1, 'b':2, 'c':3}
df5['assortment'] = df5['assortment'].map(assortment_dict)
# -
# ### 3.7. Transformations
# Logarithmic Transformation
df5['sales'] = np.log1p( df5['sales'] )
# +
# Cyclic Transformation
df5['month_sin'] = df5['month'].apply( lambda x: np.sin( x * ( 2. * np.pi/12) ) )
df5['month_cos'] = df5['month'].apply( lambda x: np.cos( x * ( 2. * np.pi/12) ) )
df5['day_sin'] = df5['day'].apply( lambda x: np.sin( x * ( 2. * np.pi/31) ) )
df5['day_cos'] = df5['day'].apply( lambda x: np.cos( x * ( 2. * np.pi/31) ) )
df5['day_of_week_sin'] = df5['day_of_week'].apply( lambda x: np.sin( x * ( 2. * np.pi/7) ) )
df5['day_of_week_cos'] = df5['day_of_week'].apply( lambda x: np.cos( x * ( 2. * np.pi/7) ) )
# Dropping cyclically transformed attributes
df5.drop(['month', 'day', 'day_of_week'], axis=1, inplace=True)
# -
# # 4. MODELING
# ### 4.1. Split dataset into train and test
# +
# Sort dataset by date in ascending order
df6 = df5.copy()
df6 = df6.sort_values(by='date', ascending=True).reset_index(drop=True)
# Separate the data from the last 6 weeks for test dataset and the others for training dataset
date_start_last_6_weeks = df6['date'].max() - datetime.timedelta(days=6*7)
df6_train = df6[ df6['date'] < date_start_last_6_weeks ]
df6_test = df6[ df6['date'] >= date_start_last_6_weeks ]
X_train = df6_train.drop(['sales', 'date'], axis=1)
y_train = df6_train['sales']
X_test = df6_test.drop(['sales', 'date'], axis=1)
y_test = df6_test['sales']
# -
# ### 4.2. Feature Selector - Boruta
# +
## Training and test dataset for Boruta
#X_train_boruta = X_train.values
#y_train_boruta = y_train.values.ravel()
#
## Defining Random Forest Regressor
#rf = RandomForestRegressor( n_jobs=-1 )
#
## Defining Boruta
#boruta = BorutaPy( rf, n_estimators='auto', verbose=2, random_state=42 ).fit( X_train_boruta, y_train_boruta )
# +
## Best features
#cols_selected = boruta.support_.tolist()
#
#cols_selected_boruta = X_train.iloc[:, cols_selected].columns.tolist()
#
## Features not selected
#cols_not_selected_boruta = list( np.setdiff1d( X_train.columns, cols_selected_boruta) )
# +
#cols_selected_boruta
# +
cols_selected_boruta = ['store',
'store_type',
'assortment',
'competition_distance',
'competition_time',
'promo',
'promo2',
'promo2_time',
'month_cos',
'day_sin',
'day_cos',
'day_of_week_sin',
'day_of_week_cos']
X_train = X_train[ cols_selected_boruta ]
X_test = X_test[ cols_selected_boruta ]
# -
# ### 4.3. Build models
# Training 5 models, with and without cross validation:
#
# 1. Average (baseline model)
# 2. Linear Regression
# 3. Linear Regression Regularized
# 4. Random Forest Regressor
# 5. XGBoost Regressor
# +
# Model's Performance Function
def ml_error(model_name,y,yhat):
mae = mean_absolute_error(y,yhat)
mape = mean_absolute_percentage_error(y,yhat)
rmse = np.sqrt(mean_squared_error(y,yhat))
return pd.DataFrame({'Model Name': model_name,
'MAE': mae,
'MAPE': mape,
'RMSE': rmse}, index=[0])
# Cross Validation Function with Time Series Split
def cross_validation(x_training, y_training, kfold, model_name, model, verbose=False):
mae_list = []
mape_list = []
rmse_list = []
tscv = TimeSeriesSplit(n_splits=kfold)
for fold, (train_index, test_index) in enumerate(tscv.split(x_training)):
print('FOLD:', fold)
print('TRAIN:', train_index)
print('TEST:', test_index)
x_training_cv = x_training.iloc[train_index]
y_training_cv = y_training.iloc[train_index]
x_validation_cv = x_training.iloc[test_index]
y_validation_cv = y_training.iloc[test_index]
# model
m = model.fit(x_training_cv, y_training_cv)
# predict
yhat = m.predict(x_validation_cv)
# performance
m_result = ml_error(model_name, np.expm1(y_validation_cv), np.expm1(yhat))
# store performance of each Kfolds iteration
mae_list.append(m_result['MAE'])
mape_list.append(m_result['MAPE'])
rmse_list.append(m_result['RMSE'])
return pd.DataFrame({'Model Name': model_name,
'MAE CV': np.round(np.mean(mae_list),2).astype(str) + '+/-' + np.round(np.std(mae_list),2).astype(str),
'MAPE CV': np.round(np.mean(mape_list),2).astype(str) + '+/-' + np.round(np.std(mape_list),2).astype(str),
'RMSE CV': np.round(np.mean(rmse_list),2).astype(str) + '+/-' + np.round(np.std(rmse_list),2).astype(str)},index=[0])
# -
# ### 4.3.1. Average
# +
# Model
aux = df6_test[['store','sales']].groupby('store').mean().reset_index().rename(columns={'sales':'predictions'})
aux = pd.merge(df6_test, aux, how='left', on='store')
# Prediction
yhat_baseline = aux['predictions']
# Performance
baseline_result = ml_error('Average Model',np.expm1(y_test),np.expm1(yhat_baseline))
baseline_result
# -
# ### 4.3.2. Linear Regression
# +
# Model
lr = LinearRegression().fit( X_train, y_train )
# Prediction
yhat_lr = lr.predict( X_test )
# Performance
lr_result = ml_error('Linear Regression',np.expm1(y_test),np.expm1(yhat_lr))
lr_result
# -
# Performance with Cross Validation
lr_result_cv = cross_validation(X_train, y_train, 5, 'Linear Regression', lr, verbose=False)
lr_result_cv
# ### 4.3.3. Linear Regression Regularized
# +
# Model
lrr = Lasso(alpha=0.01).fit( X_train, y_train )
# Prediction
yhat_lrr = lrr.predict( X_test )
# Performance
lrr_result = ml_error('Linear Regression - Lasso', np.expm1(y_test), np.expm1(yhat_lrr))
lrr_result
# -
# Performance with Cross Validation
lrr_result_cv = cross_validation(X_train, y_train, 5, 'Linear Regression - Lasso', lrr, verbose=False)
lrr_result_cv
# ### 4.3.4. Random Forest Regressor
# +
# Model
rf = RandomForestRegressor( n_estimators=100, n_jobs=-1, random_state=42 ).fit( X_train, y_train )
# Prediction
yhat_rf = rf.predict( X_test )
# Performance
rf_result = ml_error('Random Forest Regressor', np.expm1(y_test), np.expm1(yhat_rf))
rf_result
# -
# Performance with Cross Validation
rf_result_cv = cross_validation(X_train, y_train, 5, 'Random Forest Regressor', rf, verbose=False)
rf_result_cv
# ### 4.3.5. XGBoost Regressor
# +
# Model
model_xgb = xgb.XGBRegressor().fit(X_train, y_train)
# Prediction
yhat_xgb = model_xgb.predict( X_test )
# Performance
xgb_result = ml_error('XGBoost Regressor', np.expm1(y_test), np.expm1(yhat_xgb))
xgb_result
# -
# Performance with Cross Validation
xgb_result_cv = cross_validation(X_train, y_train, 5, 'XGBoost Regressor', model_xgb, verbose=False)
xgb_result_cv
# ### 4.4. Assess model
# ### 4.4.1. Compare Single Models Performance
modelling_result = pd.concat([baseline_result,lr_result,lrr_result,rf_result,xgb_result])
modelling_result.sort_values('RMSE')
# ### 4.4.2. Compare Cross Validated Models Performance
modelling_result_cv = pd.concat([lr_result_cv,lrr_result_cv,rf_result_cv,xgb_result_cv])
modelling_result_cv.sort_values('RMSE CV')
# ### 4.4.3. Chosen Model -> XGBoost Regressor
# Comparing the metrics above, the model that achieved the best performance was Random Forest Regressor.
# However the **XGBoost Regressor** has achieved similar performance and it is lighter than Random Forest to be carried.
# ### 4.5. Hyperparameter Fine Tuning - Random Search
# +
## Parameters Variability
#param = {'n_estimators':[1500,1700,2500,3000],
# 'eta':[0.01,0.03],
# 'max_depth':[3,5,7],
# 'subsample':[0.3,0.5,0.7],
# 'colsample_bytree':[0.3,0.5,0.7],
# 'min_child_weight':[3,5,7]}
#
## Random Search
#tscv = TimeSeriesSplit(n_splits=5)
#rscv = RandomizedSearchCV( estimator=model_xgb, param_distributions=param, n_iter=5, n_jobs=-1, cv=tscv, random_state=42, verbose=True)
#model_xgb_tuned = rscv.fit(X_train, y_train)
#model_xgb_tuned.best_params_
# -
# ### 4.6. Final Model
# +
param_tuned = {'n_estimators':3000,
'eta':0.03,
'max_depth':5,
'subsample':0.7,
'colsample_bytree':0.7,
'min_child_weight':3}
# model
model_xgb_tuned = xgb.XGBRegressor(objective='reg:squarederror',
n_estimators=param_tuned['n_estimators'],
eta=param_tuned['eta'],
max_depth=param_tuned['max_depth'],
subsample=param_tuned['subsample'],
colsample_bytree=param_tuned['colsample_bytree'],
min_child_weight =param_tuned['min_child_weight']).fit(X_train,y_train)
#prediction
yhat_xgb_tuned = model_xgb_tuned.predict(X_test)
#performance
xgb_result_tuned = ml_error('XGBoost Regressor Tuned',np.expm1(y_test),np.expm1(yhat_xgb_tuned))
xgb_result_tuned
# -
# Save trained model
pickle.dump( model_xgb_tuned, open( '../model/model_store_sales_prediction.pkl', 'wb') )
# # 5. EVALUATION
# +
# Dataset to evaluation
df7 = df6_test.copy()
df7['sales'] = np.expm1(df7['sales'])
df7['predictions'] = np.expm1(yhat_xgb_tuned)
# Sum of predictions
df7_aux = df7[['store','predictions']].groupby('store').sum().reset_index()
# MAE and MAPE
df7_aux1 = df7[['store','predictions','sales']].groupby('store').apply(lambda x: mean_absolute_error(x['sales'], x['predictions'])).reset_index().rename(columns={0:'MAE'})
df7_aux2 = df7[['store','predictions','sales']].groupby('store').apply(lambda x: mean_absolute_percentage_error(x['sales'],x['predictions'])).reset_index().rename(columns={0:'MAPE'})
# Merge
df7_aux3 = pd.merge(df7_aux1,df7_aux2,how='inner',on='store')
df7_aux4 = pd.merge(df7_aux,df7_aux3,how='inner',on='store')
# Scenarios
df7_aux4['worst_scenario'] = df7_aux4['predictions'] - df7_aux4['MAE']
df7_aux4['best_scenario'] = df7_aux4['predictions'] + df7_aux4['MAE']
# Order columns
df7_aux4 = df7_aux4[['store','predictions','worst_scenario','best_scenario','MAE','MAPE']]
df7_aux4
# -
plt.figure(figsize = (15,7))
sns.scatterplot(x = 'store',y = 'MAPE', data = df7_aux4);
# Total Business Impact
df7_aux5 = df7_aux4[['predictions','worst_scenario','best_scenario']].apply(lambda x: np.sum(x),axis=0).reset_index().rename(columns={'index': 'Scenarios',0:'Values'})
df7_aux5['Values'] = df7_aux5['Values'].map('R$ {:,.2f}'.format)
df7_aux5
# Machine Learning Performance
sns.lineplot(x='date',y='sales',data=df7,label='SALES');
sns.lineplot(x='date',y='predictions',data=df7,label='PREDICTIONS');
| notebooks/store_sales_predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import geopandas as gpd
import pandas as pd
import os
path = r"T:\DCProjects\Modeling\KateModel\requests\20220210 lane miles"
file = '2020_emme_links.shp'
df = pd.read_excel(os.path.join(path, 'CLMPO Lane Miles.xlsx'), skiprows=1)
df.rename(columns={'Unnamed: 0':'Functional Classification'}, inplace=True)
df_mapping = pd.DataFrame({'fed_class': df['Functional Classification'].values})
sort_mapping = df_mapping.reset_index().set_index('fed_class')
def aggLaneMiles(file):
gdf = gpd.read_file(os.path.join(path,'output',file))
aggdata = gdf[['miles', 'fed_class']].groupby('fed_class').agg('sum')
aggdata = {'fed_class':aggdata.index.values,'miles':aggdata.miles.values}
aggdata = pd.DataFrame(aggdata)
aggdata['fed_class'] = aggdata['fed_class'].str.replace('and','&')
aggdata['order'] = aggdata['fed_class'].map(sort_mapping['index'])
return aggdata.dropna().sort_values('order').miles.values
aggLaneMiles(file)
df['Base Year'] = aggLaneMiles('2020_emme_links.shp')
df['Future Year'] = aggLaneMiles('2045_emme_links.shp')
df
df.to_csv(os.path.join(path, 'output', 'CLMPO_Lane_Miles.csv'), index=False)
| data_requests/lane_miles/aggregate_lane_miles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import pandas as pd
import numpy as np
# this data is based on the output of 1.0.0_Data_Filtering
data = pd.read_csv('../data/filtered_reviews_in_Phonex.csv')
data.head(3)
def train_valid_test_split(data, m, n):
'''
construct rating matrix from data
the columns of which represent business_id
the rows of which represent user_id
the values of whose elements represent the according ratings
@ data: filterd_reviews
@ m: counts of ratings for validation
@ n: counts of ratings for test
'''
# to construct sparse matrix
# train
train_user_id = []
train_business_id = []
train_stars = []
# validation
valid_user_id = []
valid_business_id = []
valid_stars = []
# train + validation
train_valid_user_id = []
train_valid_business_id = []
train_valid_stars = []
# test
test_user_id = []
test_business_id = []
test_stars = []
user_id_lst = data['user_id'].unique().tolist() # rows of sparse matrix
busi_id_lst = data['business_id'].unique().tolist() # columns of sparse matrix
train_sparse_matrix = np.zeros(shape=(len(user_id_lst), len(busi_id_lst)))
valid_sparse_matrix = np.zeros(shape=(len(user_id_lst), len(busi_id_lst)))
train_valid_sparse_matrix = np.zeros(shape=(len(user_id_lst), len(busi_id_lst)))
test_sparse_matrix = np.zeros(shape=(len(user_id_lst), len(busi_id_lst)))
ranking_df = data[['user_id','business_id','stars','date']].groupby(['user_id'])
for group_name, group_df in ranking_df:
group_df = group_df.sort_values(by='date')
# if the len(group_df) > valid_m + test_n, split the group_df as
# training set : group_df.iloc[:len(group_df)-m-n, :]
# validation set : group_df.iloc[len(group_df)-m-n:len(group_df)-n, :]
# test set : group_df.iloc[len(group_df)-n:, :]
# otherwise, not split the group_df
# keep the group_df as training set
if len(group_df) > m+n:
training_set = group_df.iloc[:len(group_df)-m-n, :]
train_user_id.extend(training_set.loc[:,'user_id'].tolist())
train_business_id.extend(training_set.loc[:,'business_id'].tolist())
train_stars.extend(training_set.loc[:,'stars'].tolist())
validation_set = group_df.iloc[len(group_df)-m-n:len(group_df)-n, :]
valid_user_id.extend(validation_set.loc[:,'user_id'].tolist())
valid_business_id.extend(validation_set.loc[:,'business_id'].tolist())
valid_stars.extend(validation_set.loc[:,'stars'].tolist())
train_validation_set = group_df.iloc[:len(group_df)-n, :]
train_valid_user_id.extend(train_validation_set.loc[:,'user_id'].tolist())
train_valid_business_id.extend(train_validation_set.loc[:,'business_id'].tolist())
train_valid_stars.extend(train_validation_set.loc[:,'stars'].tolist())
testing_set = group_df.iloc[len(group_df)-n:, :]
test_user_id.extend(testing_set.loc[:,'user_id'].tolist())
test_business_id.extend(testing_set.loc[:,'business_id'].tolist())
test_stars.extend(testing_set.loc[:,'stars'].tolist())
else:
training_set = group_df
train_user_id.extend(training_set.loc[:,'user_id'].tolist())
train_business_id.extend(training_set.loc[:,'business_id'].tolist())
train_stars.extend(training_set.loc[:,'stars'].tolist())
train_df = pd.DataFrame({'user_id': train_user_id, 'business_id': train_business_id, 'stars': train_stars})
valid_df = pd.DataFrame({'user_id': valid_user_id, 'business_id': valid_business_id, 'stars': valid_stars})
train_valid_df = pd.DataFrame({'user_id': train_valid_user_id, 'business_id': train_valid_business_id, 'stars': train_valid_stars})
test_df = pd.DataFrame({'user_id': test_user_id, 'business_id': test_business_id, 'stars': test_stars})
for i in range(len(train_df)):
ratings = train_df.iloc[i, 2] # stars
row_index = user_id_lst.index(train_df.iloc[i, 0]) # user_id
column_index = busi_id_lst.index(train_df.iloc[i, 1]) # business_id
train_sparse_matrix[row_index, column_index] = ratings
for i in range(len(valid_df)):
ratings = valid_df.iloc[i, 2] # stars
row_index = user_id_lst.index(valid_df.iloc[i, 0]) # user_id
column_index = busi_id_lst.index(valid_df.iloc[i, 1]) # business_id
valid_sparse_matrix[row_index, column_index] = ratings
for i in range(len(train_valid_df)):
ratings = train_valid_df.iloc[i, 2] # stars
row_index = user_id_lst.index(train_valid_df.iloc[i, 0]) # user_id
column_index = busi_id_lst.index(train_valid_df.iloc[i, 1]) # business_id
train_valid_sparse_matrix[row_index, column_index] = ratings
for i in range(len(test_df)):
ratings = test_df.iloc[i, 2] # stars
row_index = user_id_lst.index(test_df.iloc[i, 0]) # user_id
column_index = busi_id_lst.index(test_df.iloc[i, 1]) # business_id
test_sparse_matrix[row_index, column_index] = ratings
# calculate sparstiy of the matrix
train_sparsity = 1 - np.count_nonzero(train_sparse_matrix)/ (train_sparse_matrix.shape[0] * train_sparse_matrix.shape[1])
valid_sparsity = 1 - np.count_nonzero(valid_sparse_matrix)/ (valid_sparse_matrix.shape[0] * valid_sparse_matrix.shape[1])
train_valid_sparsity = 1 - np.count_nonzero(train_valid_sparse_matrix)/ (train_valid_sparse_matrix.shape[0] * train_valid_sparse_matrix.shape[1])
test_sparsity = 1 - np.count_nonzero(test_sparse_matrix)/ (test_sparse_matrix.shape[0] * test_sparse_matrix.shape[1])
train_sparsity *= 100
valid_sparsity *=100
train_valid_sparse_matrix *= 100
test_sparsity *= 100
print (f'{len(user_id_lst)} users')
print (f'{len(busi_id_lst)} business')
print (f'Train_rating_matrix Sparsity: {round(train_sparsity,4)}%')
print (f'Valid_rating_matrix Sparsity: {round(valid_sparsity,4)}%')
print(f'Test_rating_matrix Sparsity: {round(test_sparsity,4)}%')
return train_sparse_matrix, valid_sparse_matrix, train_valid_sparse_matrix, test_sparse_matrix, \
train_df, valid_df, train_valid_df, test_df, \
user_id_lst, busi_id_lst
train_sparse_matrix, valid_sparse_matrix, train_valid_sparse_matrix, test_sparse_matrix, \
train_df, valid_df, train_valid_df, test_df, \
user_id_lst, busi_id_lst = train_valid_test_split(data=data, m=1, n=1)
# +
train_valid_sparse_matrix = np.vstack((train_sparse_matrix, valid_sparse_matrix))
train_valid_df = pd.concat((train_df, valid_df), axis=0)
np.save('train_sparse_matrix.npy', train_sparse_matrix)
np.save('valid_sparse_matrix.npy', valid_sparse_matrix)
np.save('test_sparse_matrix.npy', test_sparse_matrix)
np.save('train_valid_sparse_matrix.npy', train_valid_sparse_matrix)
train_df.to_pickle('../data/train_df.pkl')
valid_df.to_pickle('../data/valid_df.pkl')
test_df.to_pickle('../data/test_df.pkl')
train_valid_df.to_pickle('../data/train_valid_df.pkl')
# -
test_sparse_matrix.shape
# # long format to sparse
def long_format_to_sparse(data, pre_feature):
'''
@ pre_feature: the feature represents predict_ratings
'''
test_sparse_matrix = np.zeros(shape=(len(user_id_lst), len(busi_id_lst)))
for i in range(len(data)):
predict_col_index = data.columns.get_loc(pre_feature)
predict_ratings = data.iloc[i, predict_col_index]
row_index = user_id_lst.index(data.iloc[i, 0]) # user_id
column_index = busi_id_lst.index(data.iloc[i, 1]) # business_id
test_sparse_matrix[row_index, column_index] = predict_ratings
return test_sparse_matrix
# +
# Example
# -
nlp_long_format = pd.read_csv('Predictions_CB_bus.csv')
nlp_sparse = long_format_to_sparse(nlp_long_format, 'prediction_ratings')
nlp_sparse
nlp_sparse.shape
# # sparse to long format
def sparse_to_long_format(sparse_matrix):
user_loc_lst = np.nonzero(sparse_matrix)[0]
busi_loc_lst = np.nonzero(sparse_matrix)[1]
prediction = [nlp_sparse[loc] for loc in zip(user_loc_lst, busi_loc_lst)]
user_id = [user_id_lst[i] for i in user_loc_lst]
busi_id = [busi_id_lst[i] for i in busi_loc_lst]
long_format = pd.DataFrame({'user_id': user_id,
'busi_id': busi_id,
'prediction_ratings': prediction})
return long_format
# +
# Example
# -
long_format = sparse_to_long_format(nlp_sparse)
long_format.head(1)
| code/1.1_Train_Test_Split.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Cargamos datos
import Loading_data
from matplotlib import pyplot as plt
import warnings
warnings.filterwarnings('ignore')
df = Loading_data.Get_Nacion()
df.head()
# +
import pandas as pdg
momo = pd.read_csv('https://momo.isciii.es/public/momo/data')
# +
def get_momo():
return pd.read_csv('https://momo.isciii.es/public/momo/data')
def get_momo_by_year():
kk = get_momo()
# Enrich data
kk = kk[(kk['ambito']=='nacional') & (kk['nombre_gedad']=='todos') & (kk['nombre_sexo' ] =='todos') ]
kk['date'] =kk['fecha_defuncion']
kk['date'] = pd.to_datetime(kk['date'])
kk['year'], kk['month'] = kk['date'].dt.year, kk['date'].dt.month
kk["month"] = kk.month.map("{:02}".format)
kk['year-month'] = kk['year'].astype(str) + "-" + kk['month'].astype(str)
ss = kk[['defunciones_observadas','year-month']].groupby(['year-month'])['defunciones_observadas'].agg('sum').to_frame()
ss['month'] = ss.index.astype(str).str[5:7]
ss['year-month'] = ss.index
muertes_2018 = ss[(ss['year-month'] >= '2018-01') & (ss['year-month'] < '2019-01')][['defunciones_observadas','month']]
muertes_2019 = ss[(ss['year-month'] >= '2019-01') & (ss['year-month'] < '2020-01')][['defunciones_observadas','month']]
muertes_2020 = ss[(ss['year-month'] >= '2020-01') & (ss['year-month'] < '2021-01')][['defunciones_observadas','month']]
muertes_2018=muertes_2018.rename(columns = {'defunciones_observadas':'2018'})
muertes_2019=muertes_2019.rename(columns = {'defunciones_observadas':'2019'})
muertes_2020=muertes_2020.rename(columns = {'defunciones_observadas':'2020'})
muertes_2018 = muertes_2018.reset_index(drop=True)
muertes_2019 = muertes_2019.reset_index(drop=True)
muertes_2020 = muertes_2020.reset_index(drop=True)
muertes_temp = pd.merge( muertes_2019,muertes_2018, on="month", how='left')
muertes_temp
muertes_temp2 = pd.merge( muertes_2019,muertes_2020, on="month", how='left')
muertes_temp2
muertes_totales = pd.merge( muertes_temp, muertes_temp2)
muertes_totales.index=muertes_totales
muertes_totales.index =muertes_totales['month']
del muertes_totales['month']
return muertes_totales[['2018','2019','2020']]
df=get_momo_by_year()
# +
from matplotlib import pyplot as plt
from IPython.display import display, HTML
import pandas as pd
import numpy as np
fig = plt.figure(figsize=(8, 6), dpi=80)
for ca in df.columns:
plt.plot(df[ca])
plt.legend(df.columns)
fig.suptitle('Comparativa', fontsize=20)
plt.show()
df['resta 2020 y 2019'] = df['2020'] - df['2019']
df
# -
nacional = momo[
(momo['ambito' ] =='nacional') &
(momo['nombre_gedad'] =='todos') &
(momo['nombre_sexo' ] =='todos')]
nacional
| jupyter/Momo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing model performance with boxplots
# How to plot multiple boxplots in the same figure. Useful when we need to visualize the mean/median model performance, along with variability (standard deviation/interquantile range).
# Let us assume we have a number of trained models and we want to assess how they perform
# on the same intependent test set using a performance metric, such as accuracy, Dice, AUC, etc.
#
# ## Independent models
# When evaluating independent models it is useful to plot their performance as a
# boxplot. Boxplots are a convenient way to visualize the variability of a model's
# performance on a set of datapoints (that comprise the test set). [Box plots](https://en.wikipedia.org/wiki/Box_plot) Visualize the 25th and 75th percentiles of the data distribution as a box, while the median is marked as a horizontal line inside the box. The whiskers can have several different meanings, but the most common (and the one used by default in [matplotlib](https://matplotlib.org/3.1.0/api/_as_gen/matplotlib.pyplot.boxplot.html)) is encompassing all datapoints lying in at most +- 1.5 times the Inter-Quantile-Range (IQR). IQR corresponds to the distance between the 25th and 75th percentiles (i.e., the height of the box).
#
# If we prefer a cleaner look, we can only plot the medians, along with error-bars
# that correspond to the 25th and 75th percentile (showing the interquantile range)
#
# The performance of different models can be visualized via boxplots as follows:
# 
# **Figure 1.** Visualizing the performance of independent models via boxplots.
#
# 
# **Figure 2.** Visualizing the performance of independent models via median+error bars corresponding to the interquantile range (the 25th and 75th percentiles).
#
# ## Dependent models
# Sometimes we want to visualize a series of dependent models. E.g. how the performance
# of a model changes after training for additional epochs, or after adding new datapoints
# to the training set. In this case, it is convenient to visualize this continuity
# by connecting the boxplots. In this example, we connect the boxplots by drawing a
# line that connects the medians
#
# Similarly, we can connect the medians of the cleaner plot that only visualizes
# the median and the interquantile range.
#
# 
# **Figure 3.** Visualizing the performance of dependent models via boxplots.
#
# 
# **Figure 4.** Visualizing the performance of dependent models via median+error bars corresponding to the interquantile range (the 25th and 75th percentiles). This cleaner plot is preferable if we intend to plot a number of different lines (corresponding to different sets of dependent models) on top of each other.
#
# The plots are generated using artificial data. All plots are created in generate_plot.py.
#
# ## About
# Personal website: https://users.isc.tuc.gr/~nchlis/
#
# For tutorials on Machine Learning projects you can visit
# - https://nchlis.github.io/
# - https://github.com/nchlis/
#
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### How To Break Into the Field
#
# Now you have had a closer look at the data, and you saw how I approached looking at how the survey respondents think you should break into the field. Let's recreate those results, as well as take a look at another question.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import HowToBreakIntoTheField as t
# %matplotlib inline
df = pd.read_csv('./survey_results_public.csv')
schema = pd.read_csv('./survey_results_schema.csv')
df.head()
# -
# #### Question 1
#
# **1.** In order to understand how to break into the field, we will look at the **CousinEducation** field. Use the **schema** dataset to answer this question. Write a function called **get_description** that takes the **schema dataframe** and the **column** as a string, and returns a string of the description for that column.
# +
def get_description(column_name, schema=schema):
'''
INPUT - schema - pandas dataframe with the schema of the developers survey
column_name - string - the name of the column you would like to know about
OUTPUT -
desc - string - the description of the column
'''
desc = list(schema[schema['Column'] == column_name]['Question'])[0]
return desc
#test your code
#Check your function against solution - you shouldn't need to change any of the below code
get_description(df.columns[0]) # This should return a string of the first column description
# -
#Check your function against solution - you shouldn't need to change any of the below code
descrips = set(get_description(col) for col in df.columns)
t.check_description(descrips)
# The question we have been focused on has been around how to break into the field. Use your **get_description** function below to take a closer look at the **CousinEducation** column.
get_description('CousinEducation')
# #### Question 2
#
# **2.** Provide a pandas series of the different **CousinEducation** status values in the dataset. Store this pandas series in **cous_ed_vals**. If you are correct, you should see a bar chart of the proportion of individuals in each status. If it looks terrible, and you get no information from it, then you followed directions. However, we should clean this up!
# +
cous_ed_vals = df.CousinEducation.value_counts()#Provide a pandas series of the counts for each CousinEducation status
cous_ed_vals # assure this looks right
# +
# The below should be a bar chart of the proportion of individuals in your ed_vals
# if it is set up correctly.
(cous_ed_vals/df.shape[0]).plot(kind="bar");
plt.title("Formal Education");
# -
# We definitely need to clean this. Above is an example of what happens when you do not clean your data. Below I am using the same code you saw in the earlier video to take a look at the data after it has been cleaned.
# +
possible_vals = ["Take online courses", "Buy books and work through the exercises",
"None of these", "Part-time/evening courses", "Return to college",
"Contribute to open source", "Conferences/meet-ups", "Bootcamp",
"Get a job as a QA tester", "Participate in online coding competitions",
"Master's degree", "Participate in hackathons", "Other"]
def clean_and_plot(df, title='Method of Educating Suggested', plot=True):
'''
INPUT
df - a dataframe holding the CousinEducation column
title - string the title of your plot
axis - axis object
plot - bool providing whether or not you want a plot back
OUTPUT
study_df - a dataframe with the count of how many individuals
Displays a plot of pretty things related to the CousinEducation column.
'''
study = df['CousinEducation'].value_counts().reset_index()
study.rename(columns={'index': 'method', 'CousinEducation': 'count'}, inplace=True)
study_df = t.total_count(study, 'method', 'count', possible_vals)
study_df.set_index('method', inplace=True)
if plot:
(study_df/study_df.sum()).plot(kind='bar', legend=None);
plt.title(title);
plt.show()
props_study_df = study_df/study_df.sum()
return props_study_df
props_df = clean_and_plot(df)
# -
# #### Question 4
#
# **4.** I wonder if some of the individuals might have bias towards their own degrees. Complete the function below that will apply to the elements of the **FormalEducation** column in **df**.
# +
def higher_ed(formal_ed_str):
'''
INPUT
formal_ed_str - a string of one of the values from the Formal Education column
OUTPUT
return 1 if the string is in ("Master's degree", "Doctoral", "Professional degree")
return 0 otherwise
'''
if formal_ed_str in ("Master's degree", "Doctoral", "Professional degree"):
return 1
else:
return 0
df["FormalEducation"].apply(higher_ed)[:5] #Test your function to assure it provides 1 and 0 values for the df
# -
# Check your code here
df['HigherEd'] = df["FormalEducation"].apply(higher_ed)
higher_ed_perc = df['HigherEd'].mean()
t.higher_ed_test(higher_ed_perc)
# #### Question 5
#
# **5.** Now we would like to find out if the proportion of individuals who completed one of these three programs feel differently than those that did not. Store a dataframe of only the individual's who had **HigherEd** equal to 1 in **ed_1**. Similarly, store a dataframe of only the **HigherEd** equal to 0 values in **ed_0**.
#
# Notice, you have already created the **HigherEd** column using the check code portion above, so here you only need to subset the dataframe using this newly created column.
# +
ed_1 = df[df['HigherEd'] == 1] # Subset df to only those with HigherEd of 1
ed_0 = df[df['HigherEd'] == 0] # Subset df to only those with HigherEd of 0
print(ed_1['HigherEd'][:5]) #Assure it looks like what you would expect
print(ed_0['HigherEd'][:5]) #Assure it looks like what you would expect
# +
#Check your subset is correct - you should get a plot that was created using pandas styling
#which you can learn more about here: https://pandas.pydata.org/pandas-docs/stable/style.html
ed_1_perc = clean_and_plot(ed_1, 'Higher Formal Education', plot=False)
ed_0_perc = clean_and_plot(ed_0, 'Max of Bachelors Higher Ed', plot=False)
comp_df = pd.merge(ed_1_perc, ed_0_perc, left_index=True, right_index=True)
comp_df.columns = ['ed_1_perc', 'ed_0_perc']
comp_df['Diff_HigherEd_Vals'] = comp_df['ed_1_perc'] - comp_df['ed_0_perc']
comp_df.style.bar(subset=['Diff_HigherEd_Vals'], align='mid', color=['#d65f5f', '#5fba7d'])
# -
# #### Question 6
#
# **6.** What can you conclude from the above plot? Change the dictionary to mark **True** for the keys of any statements you can conclude, and **False** for any of the statements you cannot conclude.
# +
sol = {'Everyone should get a higher level of formal education': False,
'Regardless of formal education, online courses are the top suggested form of education': True,
'There is less than a 1% difference between suggestions of the two groups for all forms of education': False,
'Those with higher formal education suggest it more than those who do not have it': True}
t.conclusions(sol)
# -
# This concludes another look at the way we could compare education methods by those currently writing code in industry.
| lessons/CRISP_DM/.ipynb_checkpoints/How To Break Into the Field - Solution -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yohanesnuwara/numerical-method/blob/master/LMU_course_Igel/Week2_SecondDerivative.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] nbpresent={"id": "5d520f9b-b602-4c79-8a60-fe27a18ed013"} id="-8AxFJWKAlFf" colab_type="text"
# <div style='background-image: url("title01.png") ; padding: 0px ; background-size: cover ; border-radius: 5px ; height: 200px'>
# <div style="float: right ; margin: 50px ; padding: 20px ; background: rgba(255 , 255 , 255 , 0.7) ; width: 50% ; height: 150px">
# <div style="position: relative ; top: 50% ; transform: translatey(-50%)">
# <div style="font-size: xx-large ; font-weight: 900 ; color: rgba(0 , 0 , 0 , 0.8) ; line-height: 100%">Computers, Waves, Simulations</div>
# <div style="font-size: large ; padding-top: 20px ; color: rgba(0 , 0 , 0 , 0.5)">Finite-Difference Method - Second Derivative</div>
# </div>
# </div>
# </div>
# + [markdown] nbpresent={"id": "ef866370-5d28-4549-a6f8-e41bee86acdf"} id="KxL5SCnHAlFg" colab_type="text"
# #### This exercise covers the following aspects:
# * Initializing a Gaussian test function
# * Calculation of numerical second derivative with 3-point operator
# * Accuracy improvement of numerical derivative with 5-point operator
# + code_folding=[] nbpresent={"id": "218aaa8f-5740-4c7f-b7bc-c6c168ba2ecd"} id="5HO19oB-AlFh" colab_type="code" colab={}
# Import Libraries
import numpy as np
from math import *
import matplotlib.pyplot as plt
# + [markdown] nbpresent={"id": "78c4fa61-9316-4e30-9d70-76bf31a62797"} id="PVCJZpdbAlFk" colab_type="text"
# We initialize a Gaussian function
#
# \begin{equation}
# f(x)=\dfrac{1}{\sqrt{2 \pi a}}e^{-\dfrac{(x-x_0)^2}{2a}}
# \end{equation}
#
# Note that this specific definition is a $\delta-$generating function. This means that $\int{f(x) dx}=1$ and in the limit $a\rightarrow0$ the function f(x) converges to a $\delta-$function.
# + code_folding=[] nbpresent={"id": "ff03aeb5-ea0c-49ea-966b-41e32a43652a"} id="VXIXOQVJAlFl" colab_type="code" colab={}
# Initialization
xmax=10.0 # physical domain (m)
nx=100 # number of space samples
a=.25 # exponent of Gaussian function
dx=xmax/(nx-1) # Grid spacing dx (m)
x0 = xmax/2 # Center of Gaussian function x0 (m)
x=np.linspace(0,xmax,nx) # defining space variable
# Initialization of Gaussian function
f=(1./sqrt(2*pi*a))*np.exp(-(((x-x0)**2)/(2*a)))
# + code_folding=[] nbpresent={"id": "731499f2-5c1a-46a3-9dd7-0fe9e740e8d4"} id="CmTlObAgAlFn" colab_type="code" outputId="7a423efc-d7e7-4027-83e5-86fa7f9aca47" colab={}
# Plotting of gaussian
plt.figure(figsize=(10,6))
plt.plot(x, f)
plt.title('Gaussian function')
plt.xlabel('x, m')
plt.ylabel('Amplitude')
plt.xlim((0, xmax))
plt.grid()
plt.show()
# + [markdown] nbpresent={"id": "9dfc0b23-793e-44b8-bbf7-6211f6ba6d66"} id="ufb8O2NBAlFr" colab_type="text"
# Now let us calculate the second derivative using the finite-difference operator with three points
#
# \begin{equation}
# f^{\prime\prime}_{num}(x)=\dfrac{f(x+dx)-2 f(x)+f(x-dx)}{dx^2}
# \end{equation}
#
# and compare it with the analytical solution
# \begin{equation}
# f^{\prime\prime}(x)= \dfrac{1}{\sqrt{2\pi a}} ( \dfrac{(x-x_0)^2}{a^2}- \dfrac{1}{a} ) \ e^{-\dfrac{(x-x_0)^2}{2a}}
# \end{equation}
# + code_folding=[] nbpresent={"id": "1a162fb0-320a-4eef-81db-298723350bee"} id="12OTUoDkAlFr" colab_type="code" colab={}
# Second derivative with three-point operator
# Initiation of numerical and analytical derivatives
nder3=np.zeros(nx) # numerical derivative
ader=np.zeros(nx) # analytical derivative
# Numerical second derivative of the given function
for i in range (1, nx-1):
nder3[i]=(f[i+1] - 2*f[i] + f[i-1])/(dx**2)
# Analytical second derivative of the Gaissian function
ader=1./sqrt(2*pi*a)*((x-x0)**2/a**2 -1/a)*np.exp(-1/(2*a)*(x-x0)**2)
# Exclude boundaries
ader[0]=0.
ader[nx-1]=0.
# Calculate rms error of numerical derivative
rms = np.sqrt(np.mean((nder3-ader)**2))
# + code_folding=[] id="ybk1pEDgAlFt" colab_type="code" outputId="183fe212-3f39-44eb-bc98-b6372cf45e2f" colab={}
# Plotting
plt.figure(figsize=(10,6))
plt.plot (x, nder3,label="Numerical Derivative, 3 points", lw=2, color="violet")
plt.plot (x, ader, label="Analytical Derivative", lw=2, ls="--")
plt.plot (x, nder3-ader, label="Difference", lw=2, ls=":")
plt.title("Second derivative, Err (rms) = %.6f " % (rms) )
plt.xlabel('x, m')
plt.ylabel('Amplitude')
plt.legend(loc='lower left')
plt.grid()
plt.show()
# + [markdown] nbpresent={"id": "7661a51a-d5ee-4479-b8d4-78e9da50bfaa"} id="gazmf9-XAlFw" colab_type="text"
# In the cell below calculation of the first derivative with four points is provided with the following weights:
#
# \begin{equation}
# f^{\prime\prime}(x)=\dfrac{-\dfrac{1}{12}f(x-2dx)+\dfrac{4}{3}f(x-dx)-\dfrac{5}{2}f(x) +\dfrac{4}{3}f(x+dx)-\dfrac{1}{12}f(x+2dx)}{dx^2}
# \end{equation}
# + code_folding=[] nbpresent={"id": "b3dac541-c1db-46a5-b4d3-a0060005afba"} id="4y-vo_wJAlFw" colab_type="code" colab={}
# First derivative with four points
# Initialisation of derivative
nder5=np.zeros(nx)
# Calculation of 2nd derivative
for i in range (1, nx-2):
nder5[i] = (-1./12 * f[i - 2] + 4./3 * f[i - 1] - 5./2 * f[i] \
+4./3 * f[i + 1] - 1./12 * f[i + 2]) / dx ** 2
# Exclude boundaries
ader[1]=0.
ader[nx-2]=0.
# Calculate rms error of numerical derivative
rms=rms*0
rms = np.sqrt(np.mean((nder5-ader)**2))
# + code_folding=[] nbpresent={"id": "654f633a-08ce-4bec-9d1c-2b54af728587"} id="b8rn8Zb4AlFz" colab_type="code" outputId="771d7385-4d9b-4805-ab0a-04a306c2ab74" colab={}
# Plotting
plt.figure(figsize=(10,6))
plt.plot (x, nder5,label="Numerical Derivative, 5 points", lw=2, color="violet")
plt.plot (x, ader, label="Analytical Derivative", lw=2, ls="--")
plt.plot (x, nder5-ader, label="Difference", lw=2, ls=":")
plt.title("Second derivative, Err (rms) = %.6f " % (rms) )
plt.xlabel('x, m')
plt.ylabel('Amplitude')
plt.legend(loc='lower left')
plt.grid()
plt.show()
# + [markdown] id="qjPGOdKtAlF1" colab_type="text"
# ### Conclusions
#
# * 3-point finite-difference approximations can provide estimates of the 2nd derivative of a function
# * We can increase the accuracy of the approximation by using further functional values further
# * A 5-point operator leads to substantially more accurate results
| LMU_course_Igel/Week2_SecondDerivative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# We create logistic regression model (i.e. multi-layer perceptron with no hidden layer), and evaluate it on the canonical MNIST dataset.
# +
import os
import gzip
import numpy as np
import autodiff as ad
from autodiff import initializers
from autodiff import optimizers
random_state = np.random.RandomState(0)
# -
def read_mnist_labels(fn):
with gzip.open(fn, 'rb') as f:
content = f.read()
num_images = int.from_bytes(content[4:8], byteorder='big')
labels = np.zeros((num_images, 10), dtype=np.float32)
indices = np.fromstring(content[8:], dtype=np.uint8)
labels[range(num_images), indices] += 1
return labels
def read_mnist_images(fn):
with gzip.open(fn, 'rb') as f:
content = f.read()
num_images = int.from_bytes(content[4:8], byteorder='big')
height = int.from_bytes(content[8:12], byteorder='big')
width = int.from_bytes(content[12:16], byteorder='big')
images = np.fromstring(content[16:], dtype=np.uint8).reshape((num_images, height, width))
images = images.astype(np.float32) / 255.
return images
# Make sure you have the downloaded the following 4 files, and place them under the current directory.
train_images = read_mnist_images('train-images-idx3-ubyte.gz')
train_labels = read_mnist_labels('train-labels-idx1-ubyte.gz')
test_images = read_mnist_images('t10k-images-idx3-ubyte.gz')
test_labels = read_mnist_labels('t10k-labels-idx1-ubyte.gz')
# Build a logistic regression model with l2 regularization.
# +
reg = 1e-3
tni = initializers.TruncatedNormalInitializer(mean=0.0, stddev=0.01, seed=0)
zi = initializers.ZerosInitializer()
gd = optimizers.GradientDescentOptimizer(alpha=0.5)
inputs = ad.placeholder((None, 784))
labels = ad.placeholder((None, 10))
weight = ad.variable((784, 10), tni)
bias = ad.variable((10,), zi)
logits = ad.matmul(inputs, weight) + bias
loss = ad.reduce_mean(ad.softmax_cross_entropy_loss(labels, logits))
loss = loss + ad.l2norm(weight, reg)
# -
# setup the graph and runtime
# +
graph = ad.get_default_graph()
graph.initialize_variables()
runtime = ad.RunTime()
graph.set_runtime(runtime)
# -
# Training stage: run forward backward cycles on the computational graph.
batch_size = 100
for i in range(1000):
which = random_state.choice(train_images.shape[0], batch_size, False)
inputs_val = train_images[which].reshape((batch_size, -1))
labels_val = train_labels[which]
feed_dict = {inputs: inputs_val, labels: labels_val}
with runtime.forward_backward_cycle():
gd.optimize(loss, feed_dict)
if i % 100 == 0:
loss_val = loss.forward(feed_dict)
logits_val = logits.forward(feed_dict)
print('step: %d, loss: %f, accuracy: %f' % (i, loss_val, np.mean(np.argmax(logits_val, axis=1) == np.argmax(labels_val, axis=1))))
# At this point we are out of the scope of an active `RunTime`, so its attributes should all be empty.
assert not runtime._fwval
assert not runtime._bwval
assert not runtime._cache_data
# But `Variables` still hold their updated values. So we can save the logistic regression variable weights to a file.
graph.save_variables('lr_weights')
# And then restore from it.
# +
var_dict = np.load('lr_weights.npy', allow_pickle=True).item()
graph.initialize_variables(var_dict=var_dict)
# -
# Evaluate on test set using the restored variable weights.
# +
feed_dict = {inputs: test_images.reshape((-1, 784))}
with runtime.forward_backward_cycle():
logits_val = logits.forward(feed_dict)
print('accuracy', np.mean(np.argmax(logits_val, axis=1) == np.argmax(test_labels, axis=1)))
| demos/logistic_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''venv'': venv)'
# language: python
# name: python3
# ---
# # Vector Similarity
# ## Adding Vector Fields
# +
import redis
from redis.commands.search.field import VectorField
from redis.commands.search.query import Query
r = redis.Redis(host='localhost', port=36379)
schema = (VectorField("v", "HNSW", {"TYPE": "FLOAT32", "DIM": 2, "DISTANCE_METRIC": "L2"}),)
r.ft().create_index(schema)
# -
# ## Searching
# ### Querying vector fields
# + pycharm={"name": "#%%\n"}
r.hset("a", "v", "aaaaaaaa")
r.hset("b", "v", "aaaabaaa")
r.hset("c", "v", "aaaaabaa")
q = Query("*=>[KNN 2 @v $vec]").return_field("__v_score")
r.ft().search(q, query_params={"vec": "aaaaaaaa"})
| docs/examples/search_vector_similarity_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="FpHPaEEoVSUn" colab_type="text"
# # tensorflow transfer learning sample
#
# tensorflow で転移学習を行うサンプルです。
#
# - [Transfer learning with a pretrained ConvNet][tutorial]
#
# [tutorial]: https://www.tensorflow.org/tutorials/images/transfer_learning
# + [markdown] id="lcmZkCYOLQvk" colab_type="text"
# ## 環境の確認
# + id="LnMGmEyyLT2f" colab_type="code" outputId="4a1b01f1-5363-4b0e-ace3-aaf7f14bbf3c" colab={"base_uri": "https://localhost:8080/", "height": 53}
# !cat /etc/issue
# + id="UDMUXlaTLpG2" colab_type="code" outputId="c2d91e02-1e16-4c30-c485-8f7e66ff135b" colab={"base_uri": "https://localhost:8080/", "height": 71}
# !free -h
# + id="WHn3f12zLsTw" colab_type="code" outputId="e0e84be6-8fde-48a4-cc3b-4fa54e2b0ad0" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !cat /proc/cpuinfo
# + id="3FUZPKAcL4jO" colab_type="code" outputId="61f67b34-e47a-414e-a06c-76d54f38d9fa" colab={"base_uri": "https://localhost:8080/", "height": 323}
# !nvidia-smi
# + id="w2V3sVtNPYdT" colab_type="code" outputId="677ae93f-e772-42d5-e1ba-c939f94a9588" colab={"base_uri": "https://localhost:8080/", "height": 35}
# !python --version
# + id="Zyf3UfsENhOf" colab_type="code" colab={}
from logging import Logger
def get_logger() -> Logger:
import logging
logger = logging.getLogger(__name__)
fmt = "%(asctime)s %(levelname)s %(name)s :%(message)s"
logging.basicConfig(level=logging.INFO, format=fmt)
return logger
logger = get_logger()
# + id="EppKsLuHMYjR" colab_type="code" outputId="b1d6b7a3-48cd-4391-b929-4308c80363ca" colab={"base_uri": "https://localhost:8080/", "height": 35}
def check_tf_version() -> None:
import tensorflow as tf
logger.info(tf.__version__)
check_tf_version()
# + [markdown] id="SKyYk7oMY21S" colab_type="text"
# ## ソースコードの取得
# + id="PjgDKO_RV_U5" colab_type="code" outputId="cd3b6e75-37f8-4e2e-807b-0b48df311010" colab={"base_uri": "https://localhost:8080/", "height": 395}
# 対象のコードを取得
# !git clone -n https://github.com/iimuz/til.git
# %cd til
# !git checkout 71dc5ca
# %cd python/tensorflow-transfer-learning
# + [markdown] id="_vUlewGLYtT1" colab_type="text"
# ## 実行
# + [markdown] id="J4llcBuDKMus" colab_type="text"
# ### 事前準備
# + id="H8Ng5oAwIevW" colab_type="code" colab={}
import tensorflow.compat.v1 as tfv1
tfv1.enable_eager_execution()
# + [markdown] id="NDZFpP_IKKCr" colab_type="text"
# ### データセットの確認
# + id="8AwAXG4UHZ7V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6c8351ed-7b63-4b58-93d0-eac9ea73ef0e"
# %run -i datasets.py
# + id="nv1OyOv3I9f4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 145} outputId="59baa002-21a9-4ddf-9bab-819051b4cc0a"
import datasets
raw_train, raw_validation, _, metadata = datasets.get_batch_dataset(shuffle_seed=0)
# + [markdown] id="leX-NiQxKRHh" colab_type="text"
# ### 特徴抽出ネットワークの確認
# + id="sLKS5JeWHdNl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1f5364f7-a374-4116-922c-fa251a7edaa5"
# %run -i network_fe.py
# + [markdown] id="071mN33PKYZu" colab_type="text"
# ### 特徴抽出の実行
# + id="g17JAmxFHhSn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d7e45f4d-40e0-4349-b352-94b388d70987"
import network_fe
import utils
from tensorflow.python.data.ops.dataset_ops import DatasetV1Adapter
def train_fe(dataset_train: DatasetV1Adapter, dataset_validation: DatasetV1Adapter) -> None:
base_learning_rate = 0.0001
epochs = 10
model = network_fe.MobileNetV2FE()
model.compile(
optimizer=tf.keras.optimizers.RMSprop(lr=base_learning_rate),
loss="binary_crossentropy",
metrics=["accuracy"],
)
# initial model accuracy
loss0, accuracy0 = model.evaluate(dataset_validation, steps=20)
logger.info(f"initial loss: {loss0:.2f}, acc: {accuracy0:.2f}")
# training
checkpoint = utils.load_checkpoints(model, save_dir="_data/ckpt_feature")
history = model.fit(
dataset_train, epochs=epochs, validation_data=dataset_validation, callbacks=[checkpoint]
)
utils.plot_history(history)
train_fe(raw_train, raw_validation)
# + [markdown] id="oniEZcuqKa19" colab_type="text"
# ### Fine Tuning のネットワークの確認
# + id="Lm7MKzfcKnSp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a5f37a9-ea9c-4658-a7b9-4808b322be0a"
# %run -i network_ft.py
# + [markdown] id="4RL1Q_a1KskP" colab_type="text"
# ### Fine Tuning の実行
# + id="1pMqz9p5Hnpj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="78b9e47d-eb5b-4e4c-b174-2e94d7ced0e1"
import network_ft
def train_ft(dataset_train: DatasetV1Adapter, dataset_validation: DatasetV1Adapter) -> None:
base_learning_rate = 0.0001
epochs = 10
model = network_ft.MobileNetV2FT()
model.compile(
optimizer=tf.keras.optimizers.RMSprop(lr=base_learning_rate),
loss="binary_crossentropy",
metrics=["accuracy"],
)
# initial model accuracy
loss0, accuracy0 = model.evaluate(dataset_validation, steps=20)
logger.info(f"initial loss: {loss0:.2f}, acc: {accuracy0:.2f}")
# training
checkpoint = utils.load_checkpoints(model, save_dir="_data/ckpt_finetuning")
history = model.fit(
dataset_train, epochs=epochs, validation_data=dataset_validation, callbacks=[checkpoint]
)
utils.plot_history(history)
train_ft(raw_train, raw_validation)
| machine_learning/tf_transfer_learning/tensorflow_transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from distutils.version import LooseVersion
from scipy.stats import norm
from sklearn.neighbors import KernelDensity
# +
# Plot the progression of histograms to kernels
np.random.seed(1)
N = 20
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
bins = np.linspace(-5, 10, 10)
fig, ax = plt.subplots(2, 2, sharex=True, sharey=True)
fig.subplots_adjust(hspace=0.05, wspace=0.05)
# histogram 1
ax[0, 0].hist(X[:, 0], bins=bins, fc='#AAAAFF', **density_param)
ax[0, 0].text(-3.5, 0.31, "Histogram")
# histogram 2
ax[0, 1].hist(X[:, 0], bins=bins + 0.75, fc='#AAAAFF', **density_param)
ax[0, 1].text(-3.5, 0.31, "Histogram, bins shifted")
# tophat KDE
kde = KernelDensity(kernel='tophat', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 0].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 0].text(-3.5, 0.31, "Tophat Kernel Density")
# Gaussian KDE
kde = KernelDensity(kernel='gaussian', bandwidth=0.75).fit(X)
log_dens = kde.score_samples(X_plot)
ax[1, 1].fill(X_plot[:, 0], np.exp(log_dens), fc='#AAAAFF')
ax[1, 1].text(-3.5, 0.31, "Gaussian Kernel Density")
for axi in ax.ravel():
axi.plot(X[:, 0], np.full(X.shape[0], -0.01), '+k')
axi.set_xlim(-4, 9)
axi.set_ylim(-0.02, 0.34)
for axi in ax[:, 0]:
axi.set_ylabel('Normalized Density')
for axi in ax[1, :]:
axi.set_xlabel('x')
# ----------------------------------------------------------------------
# Plot all available kernels
X_plot = np.linspace(-6, 6, 1000)[:, None]
X_src = np.zeros((1, 1))
fig, ax = plt.subplots(2, 3, sharex=True, sharey=True)
fig.subplots_adjust(left=0.05, right=0.95, hspace=0.05, wspace=0.05)
def format_func(x, loc):
if x == 0:
return '0'
elif x == 1:
return 'h'
elif x == -1:
return '-h'
else:
return '%ih' % x
for i, kernel in enumerate(['gaussian', 'tophat', 'epanechnikov',
'exponential', 'linear', 'cosine']):
axi = ax.ravel()[i]
log_dens = KernelDensity(kernel=kernel).fit(X_src).score_samples(X_plot)
axi.fill(X_plot[:, 0], np.exp(log_dens), '-k', fc='#AAAAFF')
axi.text(-2.6, 0.95, kernel)
axi.xaxis.set_major_formatter(plt.FuncFormatter(format_func))
axi.xaxis.set_major_locator(plt.MultipleLocator(1))
axi.yaxis.set_major_locator(plt.NullLocator())
axi.set_ylim(0, 1.05)
axi.set_xlim(-2.9, 2.9)
ax[0, 1].set_title('Available Kernels')
# ----------------------------------------------------------------------
# Plot a 1D density example
N = 100
np.random.seed(1)
X = np.concatenate((np.random.normal(0, 1, int(0.3 * N)),
np.random.normal(5, 1, int(0.7 * N))))[:, np.newaxis]
X_plot = np.linspace(-5, 10, 1000)[:, np.newaxis]
true_dens = (0.3 * norm(0, 1).pdf(X_plot[:, 0])
+ 0.7 * norm(5, 1).pdf(X_plot[:, 0]))
fig, ax = plt.subplots()
ax.fill(X_plot[:, 0], true_dens, fc='black', alpha=0.2,
label='input distribution')
colors = ['navy', 'cornflowerblue', 'darkorange']
kernels = ['gaussian', 'tophat', 'epanechnikov']
lw = 2
for color, kernel in zip(colors, kernels):
kde = KernelDensity(kernel=kernel, bandwidth=0.5).fit(X)
log_dens = kde.score_samples(X_plot)
ax.plot(X_plot[:, 0], np.exp(log_dens), color=color, lw=lw,
linestyle='-', label="kernel = '{0}'".format(kernel))
ax.text(6, 0.38, "N={0} points".format(N))
ax.legend(loc='upper left')
ax.plot(X[:, 0], -0.005 - 0.01 * np.random.random(X.shape[0]), '+k')
ax.set_xlim(-4, 9)
ax.set_ylim(-0.02, 0.4)
plt.show()
# -
| Untitled1.ipynb |