code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# For loops can be used to repeatedly execute the same code
odds = [1, 3, 5, 7]
print(odds[0])
print(odds[1])
print(odds[2])
print(odds[3])
# Problem: What happens when we add/cut the elements? Better: iterate over the elements of the list
# The for loop also needs indented lines after the for to mark the executable part of
# the code
for number in odds:
print(number)
# The general form is
# for variable in collection:
# +
# A variable can be used in each iteration of the loop
length = 0
types = ['boolean', 'integer', 'string']
for name in types:
length = length + 1
print('There are', length, 'types in the list.')
print('We can also find this via the len() function: ', len(types))
# +
# A variable maintains its last value from the loop
name = 'bool'
for name in ['bool', 'integer', 'string']:
print(name)
print('after the loop, name is', name)
# +
# To create a list containing the numbers from 0 to N we can use the range function
for num in range(6):
print(num)
print()
# The range function is an important tool in python, it can also take more than one
# argument to specify start, stop and increment
for num in range(3,10,2):
print(num)
# -
# Quick Task: What happens here?
word = 'string'
for char in word:
print(char)
# +
# Task: Write code which calculates and prints the sum of all elements of a list
numbers = [124, 402, 36]
summed = 0
for num in numbers:
summed = summed + num
print(summed)
# +
# Task: Calculate the sum of all multiples of 3 up to 1000
sum = 0
for num in range(3,1000,3):
sum += num
print(sum)
# -
# If conditionals are used to execute pieces of code based upon a bool statement (True or False)
num = 37
if num > 100:
print('greater than 100')
else:
print('not greater than 100')
print('done')
# +
# The 'elif' statement can be used to chain together if statements (else if)
num = -3
if num > 0:
print(num, 'is positive')
elif num == 0:
print(num, 'is zero')
else:
print(num, 'is negative')
# +
# Note on comparison operators, we have already seen before the '==' as the comparing equal sign
# The other commonly used comparing operators are:
1 == 2
#> : greater than
#< : less than
#== : equal to
#!= : does not equal
#>= : greater than or equal to
#<= : less than or equal to
# +
# We can combine booleans using 'and' or 'or'
# For 'and' both boolean need to be true
if (1 > 0) and (-1 == 0):
print('both parts are true')
else:
print('at least one part is false')
# While for 'or' only one of the two needs to be true
if (1 < 0) or (-1 == 0):
print('at least one test is true')
# +
# Task: Write a piece of code which returns the absolute value of a number
a = 5
b = 10
diff = a-b
if diff < 0:
diff = diff*(-1)
# Task: Write a condition that prints True if a is within 10% of b, False otherwise, a,b,from Real+
if diff < 0.1 * b:
print('True')
else:
print('False')
print(abs(a - b) < 0.1 * abs(b))
# +
# Task: Sort a List of numbers into a list of all numbers <10 and >10
# It might be helpfull to use the append function to add elements to the end of an existing list
input = [12,15,16,19,24,25,27]
evens = []
odds = []
for num in input:
if(num>10):
evens.append(num)
else:
odds.append(num)
print(evens)
print(odds)
# +
# Functions
# -
# Expanding upon the use of function we remember the conversion example
mass_in_kg = 40
mass_in_g = mass_in_kg*1000
print("kg: ", mass_in_kg, " g: ", mass_in_g)
# +
# Here we want to define a new function called 'g_to_kg' which converts masses
# The def-Keyword designates a new function definition.
# Within the '()' We can specify the function arguments which are expected.
# In python, everything belonging to the function has to be indented
def g_to_kg(mass_in_g):
return mass_in_g/1000
print(g_to_kg(52))
# We can call functions within functions, we do not need a return statement
def say_hello():
print("Hello")
# +
# We can save the output of functions in variables and use the functions in calculations:
mass_in_g = 1000
mass_in_kg = g_to_kg(mass_in_g)
print(mass_in_kg)
mass_in_lb = g_to_kg(mass_in_g)/2.2
print(mass_in_lb)
# +
# Variable defined in functions go out of scopes when the function exits
def g_to_lb(mass_in_g):
temp_mass = g_to_kg(mass_in_g)
return temp_mass/2.2
# This will throw an error, result is not known outside the function
g_to_lb(3)
# #? print(temp_mass)
print(g_to_lb(5))
# +
# Task: Write a function which takes 2 integers and calculates the power of the first to the second
def power(base, exponent):
result = 1;
for ii in range(exponent):
result *= base
return result
print(power(4,4))
# -
# Task: Write a function which takes a number and a strings and converts from gram to either kg or lb
def convert_from_g(value, unit_to):
if(unit_to == 'kg'):
return g_to_kg(value)
elif(unit_to == 'lb'):
return g_to_lb(value)
else:
print("Invalid unit in input")
# +
# Task: Find the position of the smallest element in a given list
value = [3,4,5,6,7,8,2]
def smallest_element(input_list):
smallest_idx = 0;
for i in range(len(input_list)):
if input_list[i] < input_list[smallest_idx]:
smallest_idx = i;
return smallest_idx
print(smallest_element(value))
value.index(min(value))
# +
# If there ist time:
# A function argument can be set to a default in the definition using '='
# If no other value is given this default will be assumed
def display(a=1, b=2, c=3):
print('a:', a, 'b:', b, 'c:', c)
print('no parameters:')
display()
print('one parameter:')
display(55)
print('two parameters:')
display(55, 66)
# We can subvert the usual left to right filling of parameters by explicitely setting them
print('only setting the value of c')
display(c=77)
# Many library functions have default parameters in most arguments
# +
# Task: What do you expect to be printed?
def numbers(one, two=2, three, four=4):
n = str(one) + str(two) + str(three) + str(four)
return n
#print(numbers(1, three=3))
# +
# Help messages for custom functions
def rescale(input_array):
"""Takes an array as input, and returns a corresponding array scaled so
that 0 corresponds to the minimum and 1 to the maximum value of the input array.
Examples:
>>> rescale(range(0,10))
array([ 0. , 0.11111111, 0.22222222, 0.33333333, 0.44444444,
0.55555556, 0.66666667, 0.77777778, 0.88888889, 1. ])
"""
output_array = []
L = min(input_array)
H = max(input_array)
for i in input_array:
output_array.append((i - L) / (H - L))
return output_array
print(rescale([1,2.4,6.5,3.3]))
# We can now obtain our own custom help message
help(rescale)
# +
# google
# https://docs.python.org/3/
# stackoverflow
# +
# GET THE DATA! Download and unpack
| keywords_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as seaborn
import matplotlib.pyplot as plt
df = pd.read_csv('agora.csv')
df.head()
x = df.drop('Profit',axis=1) # asix=1 means col, axis = 0 means rows
x.head()
y = df['Profit']
y.head()
# # Manual
x['Marketing Spend'] = x['Marketing Spend'] / x['Marketing Spend'].max()
x.head()
# # Min Max Scaler
from sklearn.preprocessing import MinMaxScaler
minmax = MinMaxScaler(feature_range=(0, 3)) #default range 0,1
minmax
df['Marketing Spend'] = minmax.fit_transform(x[['Marketing Spend']])
df.head()
df['Administration'] = minmax.fit_transform(x[['Administration']])
df['Transport'] = minmax.fit_transform(x[['Transport']])
df.head()
# # Standard Scaler
df2 = pd.read_csv('agora.csv')
df2.head()
from sklearn.preprocessing import StandardScaler
std = StandardScaler()
# # Robust Scaler
df3 = pd.read_csv('agora.csv')
df3.head()
from sklearn.preprocessing import RobustScaler
rob = RobustScaler()
# # Max Absolute Scaler
df4 = pd.read_csv('agora.csv')
df4.head()
from sklearn.preprocessing import MaxAbsScaler
rob = MaxAbsScaler()
| class-5/Class 05 - Feature Transformations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# ### To import csv file use pd.read_csv
df = pd.read_csv('datasets/new.csv')
df
df = pd.read_csv('datasets/new.csv', skiprows = 1)
df
# ## When headers aren't provided
df = pd.read_csv('datasets/new2.csv')
df
# ### take header=none and provide list of columns to names
headers = ['tickers', 'eps', 'revenue', 'price', 'people']
df = pd.read_csv('datasets/new2.csv', header=None,names=headers )
df
# ### Reading only few rows:
df = pd.read_csv('datasets/new2.csv', header=None, names=headers, nrows=3)
df
# ### Treating na values and writing to csv
name= ['tickers','eps','revenue','price','people']
df = pd.read_csv('datasets/new2.csv',header=None,names=name,na_values=-1)
df
name= ['tickers','eps','revenue','price','people']
df = pd.read_csv('datasets/new2.csv',header=None,names=name,na_values={
'eps':['not available'],
'revenue':[-1],
'price':['n.a.'],
'people':['n.a.']
})
df
df.to_csv('datasets/final.csv', index=False, header=False, columns=['tickers','eps'])
# ### To import excel use pd.read_excel
| Pandas/Pandas2-read,write csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p36)
# language: python
# name: conda_pytorch_p36
# ---
# +
# %load_ext autoreload
# %autoreload 2
import torch
import matplotlib.pyplot as plt
import scipy, scipy.optimize
import numpy as np
import astro_dynamo
from astro_dynamo.snap import ParticleType
import mwtools.nemo
import galpy.potential
# %aimport astro_dynamo.analytic_potentials
# -
# Check we reproduce the same answer as galpy for some random potential
# +
q=0.5
#Getting units correct is painful. with ro=1 vo=1 and turn_physical_off then everything should be just G=1
galpy_pot = galpy.potential.TwoPowerTriaxialPotential(c=q,ro=1,vo=1)
galpy_pot.turn_physical_off()
pot = astro_dynamo.analytic_potentials.SpheroidalPotential(lambda m: galpy_pot._amp*galpy_pot._mdens(m),q=q)
# -
x=np.linspace(0,10,100)
plt.semilogy(x,list(map(lambda x: -galpy_pot.Rforce(x,1),x)),'r',label='galpy FR')
plt.semilogy(x,-pot.f_r_cyl(x,np.array([1.])),'--k')
plt.semilogy(x,list(map(lambda x: -galpy_pot.zforce(x,1),x)),'y',label='galpy Fz')
plt.semilogy(x,-pot.f_z(x,np.array([1.])),'--k',label='astro-dynamo')
plt.legend()
plt.ylabel('Force')
plt.xlabel('R')
x=np.linspace(0,10,100)
plt.plot(x,list(map(lambda x: galpy_pot.vcirc(x,0),x)),'r',label='galpy FR')
plt.plot(x,torch.sqrt(pot.vc2(x,np.array([0.]))),'--k',label='astro-dynamo')
plt.ylabel('$V_c$')
plt.xlabel('$R$')
# Try replacing the dark matter particles in a snapshot by an analytic profile
snap=astro_dynamo.snap.SnapShot('../inputmodels/M85_0.gz',
particle_type_mapping={0:ParticleType.DarkMatter,1:ParticleType.Star})
q,qerr = astro_dynamo.analytic_potentials.fit_q_to_snapshot(snap,plot=True,r_bins=50)
print(f'q={q:.3f}+-{qerr:.3f}')
# Define and test a spheriodal potential based on this fit
# +
def ein(m,rhor0,m0,alpha):
rho0 = rhor0 / (np.exp(-(2 / alpha) * ((8.2 / m0) ** alpha - 1)))
return rho0 * np.exp(-(2 / alpha) * ((m / m0) ** alpha - 1))
pot = astro_dynamo.analytic_potentials.fit_potential_to_snap(snap.dm,ein,init_parms=[1e-3,8.0,0.7],plot=True)
# +
r,dm_vc2 = mwtools.nemo.rotationcurve(snap.dm.as_numpy_array(),rrange=(0, 40))
r,stellar_vc2 = mwtools.nemo.rotationcurve(snap.stars.as_numpy_array(),rrange=(0, 40))
i = (np.abs(snap.dm.positions[:,0]) < 10) & \
(np.abs(snap.dm.positions[:,1]) < 10) & \
(np.abs(snap.dm.positions[:,2]) < 10)
r,dm_vc2_trunc = mwtools.nemo.rotationcurve(snap.dm[i].as_numpy_array(),rrange=(0, 40))
i = (np.abs(snap.stars.positions[:,0]) < 10) & \
(np.abs(snap.stars.positions[:,1]) < 10) & \
(np.abs(snap.stars.positions[:,2]) < 10)
r,stellar_vc2_trunc = mwtools.nemo.rotationcurve(snap.stars[i].as_numpy_array(),rrange=(0, 40))
# +
f,ax = plt.subplots(1,1)
ax.plot(r,np.sqrt(dm_vc2),label = 'DM Particles')
ax.plot(r,np.sqrt(stellar_vc2),label = 'Stellar Particles')
ax.plot(r,np.sqrt(dm_vc2_trunc),label = 'DM Particles in 10kpc box')
x=np.linspace(0.,40,100)
ax.plot(x,np.sqrt(pot.vc2(x,torch.tensor(0.0,dtype=torch.float64))),label = 'Einasto Fit')
r=r.copy()
ax.plot(r,np.sqrt(stellar_vc2+pot.vc2(r,torch.tensor(0.0,dtype=torch.float64)).numpy()),label = 'Total Vc: Einasto Fit')
ax.plot(r,np.sqrt(stellar_vc2+dm_vc2),label = 'Total Vc: Particles')
ax.set_xlim((0,20))
ax.set_ylabel('$V_c$')
ax.set_xlabel('$R$')
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# -
# Test the gridding of the potential
# +
pot.grid_acc()
maxi=1000
positions = snap.stars.positions
r_cyl = snap.stars.rcyl
z = snap.stars.positions[..., 2]
f_r_cyl,f_z = pot.get_accelerations_cyl(positions[:maxi,:]).t()
f,ax = plt.subplots(1,2, figsize = (8,4), sharey = 'row')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[0].semilogy()
ax[0].legend()
ax[0].set_ylabel('Fractional Difference')
ax[0].set_xlabel('R')
ax[1].plot(z[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[1].plot(z[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[1].semilogy()
ax[1].legend()
ax[1].set_xlabel('z')
# +
maxi=1000
positions = snap.stars.positions
r_cyl = snap.stars.rcyl
z = snap.stars.positions[..., 2]
acc = pot.get_accelerations(positions)
f_r_cyl = -torch.sqrt( acc[..., 0]**2 + acc[..., 1]**2 )
f_z = acc[..., 2]
f_r_cyl=f_r_cyl[:maxi]
f_z=f_z[:maxi]
f,ax = plt.subplots(1,2, figsize = (8,4), sharey = 'row')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[0].semilogy()
ax[0].legend()
ax[0].set_ylabel('Fractional Difference')
ax[0].set_xlabel('R')
ax[1].plot(z[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[1].plot(z[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[1].semilogy()
ax[1].legend()
ax[1].set_xlabel('z')
# +
gpu_pot = pot.to('cuda')
acc = gpu_pot.get_accelerations(positions)
f_r_cyl = -torch.sqrt( acc[..., 0]**2 + acc[..., 1]**2 )
f_z = acc[..., 2]
f_r_cyl=f_r_cyl[:maxi]
f_z=f_z[:maxi]
f,ax = plt.subplots(1,2, figsize = (8,4), sharey = 'row')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[0].plot(r_cyl[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[0].semilogy()
ax[0].legend()
ax[0].set_ylabel('Fractional Difference')
ax[0].set_xlabel('R')
ax[1].plot(z[:maxi],np.abs((pot.f_r_cyl(r_cyl[:maxi],z[:maxi])-f_r_cyl)/f_r_cyl),'.',label='$F_r$')
ax[1].plot(z[:maxi],np.abs((pot.f_z(r_cyl[:maxi],z[:maxi])-f_z)/f_z),'.',label='$F_z$')
ax[1].semilogy()
ax[1].legend()
ax[1].set_xlabel('z')
# -
| notebooks/EllipsoidalPotential.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.007608, "end_time": "2021-10-14T14:56:41.302876", "exception": false, "start_time": "2021-10-14T14:56:41.295268", "status": "completed"} tags=[]
# To run this example locally, execute: `ploomber examples -n python-api`.
#
# To start a free, hosted JupyterLab: [](https://mybinder.org/v2/gh/ploomber/binder-env/main?urlpath=git-pull%3Frepo%3Dhttps%253A%252F%252Fgithub.com%252Fploomber%252Fprojects%26urlpath%3Dlab%252Ftree%252Fprojects%252Fpython-api%252FREADME.ipynb%26branch%3Dmaster)
#
# Found an issue? [Let us know.](https://github.com/ploomber/projects/issues/new?title=python-api%20issue)
#
# Have questions? [Ask us anything on Slack.](http://community.ploomber.io/)
#
# + [markdown] papermill={"duration": 0.009189, "end_time": "2021-10-14T14:56:41.319122", "exception": false, "start_time": "2021-10-14T14:56:41.309933", "status": "completed"} tags=[]
# # Python API
#
# Pipeline project using the Python API.
#
# If you're new to the Python API, check out [python-api-examples/](../python-api-examples) directory, containing tutorials and more examples.
#
# ## Description
#
# This pipeline has three tasks:
#
# 1. Load task (Python function): CSV file
# 2. Clean task (Python script): Jupyter notebook and another CSV file
# 3. Plot task (Python scripts): Jupyter notebook
#
# ## Build
# + papermill={"duration": 13.144219, "end_time": "2021-10-14T14:56:54.470475", "exception": false, "start_time": "2021-10-14T14:56:41.326256", "status": "completed"} tags=[] language="bash"
# ploomber build
| python-api/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import datetime
from extract import get_full_data
def mean(arr):
cleaned = arr[arr > 0]
return np.mean(cleaned)
def std(arr):
cleaned = arr[arr > 0]
return np.std(cleaned)
def lwmean(arr):
return np.mean(arr[-7:])
def lwmax(arr):
return np.max(arr[-7:])
def blwmax(arr):
return np.max(arr[-14:-7])
def blwmean(arr):
return np.mean(arr[-14:-7])
def bblwmax(arr):
return np.max(arr[-21:-14])
def bblwmean(arr):
return np.mean(arr[-21:-14])
def bbblwmax(arr):
return np.max(arr[-28:-21])
def bbblwmean(arr):
return np.mean(arr[-28:-21])
def max_full(arr):
return np.max(arr)
def summary_save(source, label, threshold=None):
full_df = all_countries[all_countries['iso_code'].isin(source['iso_code'].unique())]
full_df = full_df[full_df['date'] == today]
full_df['case_per_millon'] = (1000000 * full_df['total_cases']) / full_df['population']
full_df['tests_per_millon'] = (1000000 * full_df['total_tests']) / full_df['population']
filtered = full_df[['iso_code', 'location', 'date', 'total_cases', 'new_cases', 'case_per_millon', 'tests_per_millon', 'total_tests', 'population']]
if threshold is not None:
filtered = filtered[filtered['case_per_millon'] > threshold]
filtered.reset_index(inplace=True, drop=True)
cap_mean = filtered['case_per_millon'].mean()
cap_std = filtered['case_per_millon'].std()
cap_max = filtered['case_per_millon'].max()
cap_min = filtered['case_per_millon'].min()
print(f"mean: {cap_mean}")
print(f"std: {cap_std}")
print(f"max: {cap_max}")
print(f"min: {cap_min}")
filtered.to_csv(f'data/{label}.csv', index=False)
filtered.to_csv(f'data/date/{label}{today}.csv', index=False)
return filtered
def get_summary(data, threshold=None):
if threshold is not None:
filtered = data[data['population'] > threshold]
filtered = filtered[['iso_code', 'new_cases']]
summary = filtered.groupby('iso_code').agg([mean, std, lwmean, blwmean, bblwmean, bbblwmean])
summary.columns = summary.columns.droplevel(0)
summary['descending'] = summary['mean'] > summary['lwmean']
summary['closing'] = (summary['mean'] - summary['std'] / 2) > summary['lwmean']
summary.reset_index(inplace=True)
return summary
# +
today_dt = datetime.date.today()
today = today_dt.strftime('%Y-%m-%d')
all_countries = get_full_data(force=True)
all_countries['cases_per_million'] = all_countries['total_cases'] / all_countries['population'] * 1000000
exclude_small = all_countries[all_countries['population'] > 500000]
small = all_countries[all_countries['population'] < 1000000]
small.head()
# -
# ### Cuales estan descendiendo o cerrando su curva?
summary = get_summary(all_countries, threshold=500000)
summary
# +
BOL = all_countries[all_countries['iso_code'] == 'BOL'][all_countries['date'] == today]['total_cases'].values[0]
bol_case_per_million = BOL / 11.63
print(f"BOL: {BOL}")
print(f"bol_case_per_million: {bol_case_per_million}")
# -
if not summary[summary['iso_code'] == 'BOL']['descending'].values[0]:
factor = 2
else:
factor =1
worse = exclude_small[exclude_small['cases_per_million'] > (bol_case_per_million * factor)][exclude_small['date'] == today]
worse = worse[worse['cases_per_million'] < 10000]
worse = worse.sort_values(['cases_per_million'], ascending=False)
worse = worse[['iso_code', 'location', 'date', 'total_cases', 'new_cases', 'cases_per_million', 'population']]
worse.reset_index(inplace=True, drop=True)
worst = worse.head(25)
worst.reset_index(inplace=True, drop=True)
worst.head(50)
summary_save(worst, 'worst', threshold=bol_case_per_million)
# ### Worse but small
worse_small = small[small['cases_per_million'] > (bol_case_per_million * factor)][small['date'] == today]
worse_small = worse_small.sort_values(['cases_per_million'], ascending=False)
worse_small = worse_small[['iso_code', 'location', 'date', 'total_cases', 'new_cases', 'cases_per_million', 'population']]
worse_small.reset_index(inplace=True, drop=True)
worse_small
summary_save(worse_small, 'worse_small')
# ### Que paises estan cerrrando o en decenso de su curva de contagio?
summary_save(summary[summary['descending']][summary['closing']], 'closing', threshold=1000)
summary[summary['descending']]
summary_save(summary[summary['descending']], 'descending', threshold=1000)
| capacity_calculator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# This notebook demonstrates pymatgen's functionality in terms of creating and editing molecules, as well as its integration with OpenBabel. For the latter, please note that you will need to have openbabel with python bindings installed. Please refer to pymatgen's documentation for installation details.
#
# Written using:
# - pymatgen==2018.3.13
# ## Molecules
from pymatgen import Molecule
# Create a methane molecule.
coords = [[0.000000, 0.000000, 0.000000],
[0.000000, 0.000000, 1.089000],
[1.026719, 0.000000, -0.363000],
[-0.513360, -0.889165, -0.363000],
[-0.513360, 0.889165, -0.363000]]
mol = Molecule(["C", "H", "H", "H", "H"], coords)
print(mol)
# A Molecule is simply a list of Sites.
print(mol[0])
print(mol[1])
# Break a Molecule into two by breaking a bond.
for frag in mol.break_bond(0, 1):
print(frag)
# Getting neighbors that are within 3 angstroms from C atom.
print(mol.get_neighbors(mol[0], 3))
#Detecting bonds
print(mol.get_covalent_bonds())
# If you need to run the molecule in a box with a periodic boundary condition
# code, you can generate the boxed structure as follows (in a 10Ax10Ax10A box)
structure = mol.get_boxed_structure(10, 10, 10)
print(structure)
# Writing to XYZ files (easy to open with many molecule file viewers)
from pymatgen.io.xyz import XYZ
xyz = XYZ(mol)
xyz.write_file("methane.xyz")
# ## Openbabel interface
# This section demonstrates pymatgen's integration with openbabel.
from pymatgen.io.babel import BabelMolAdaptor
import pybel as pb
a = BabelMolAdaptor(mol)
# Create a pybel.Molecule, which simplifies a lot of access
pm = pb.Molecule(a.openbabel_mol)
# Print canonical SMILES representation (unique and comparable).
print("Canonical SMILES = {}".format(pm.write("can")))
# Print Inchi representation
print("Inchi= {}".format(pm.write("inchi")))
# pb.outformats provides a listing of available formats.
# Let's do a write to the commonly used PDB file.
pm.write("pdb", filename="methane.pdb", overwrite=True)
# Generating ethylene carbonate (SMILES obtained from Wikipedia)
# And displaying the svg.
ec = pb.readstring("smi", "C1COC(=O)O1")
ec.make3D()
from IPython.core.display import SVG, display_svg
svg = SVG(ec.write("svg"))
display_svg(svg)
# ## Input/Output
# Pymatgen has built-in support for the XYZ and Gaussian, NWchem file formats. It also has support for most other file formats if you have openbabel with Python bindings installed.
# +
print(mol.to(fmt="xyz"))
print(mol.to(fmt="g09"))
print(mol.to(fmt="pdb")) #Needs Openbabel.
mol.to(filename="methane.xyz")
mol.to(filename="methane.pdb") #Needs Openbabel.
print(Molecule.from_file("methane.pdb"))
# -
# For more fine-grained control over output, you can use the underlying IO classes Gaussian and Nwchem, two commonly used computational chemistry programs.
from pymatgen.io.gaussian import GaussianInput
gau = GaussianInput(mol, charge=0, spin_multiplicity=1, title="methane",
functional="B3LYP", basis_set="6-31G(d)",
route_parameters={'Opt': "", "SCF": "Tight"},
link0_parameters={"%mem": "1000MW"})
print(gau)
# A standard relaxation + SCF energy nwchem calculation input file for methane.
from pymatgen.io.nwchem import NwTask, NwInput
tasks = [
NwTask.dft_task(mol, operation="optimize", xc="b3lyp",
basis_set="6-31G"),
NwTask.dft_task(mol, operation="freq", xc="b3lyp",
basis_set="6-31G"),
NwTask.dft_task(mol, operation="energy", xc="b3lyp",
basis_set="6-311G"),
]
nwi = NwInput(mol, tasks, geometry_options=["units", "angstroms"])
print(nwi)
# ## This concludes the demo on pymatgen's basic capabilities for molecules.
| notebooks/2013-01-01-Molecule.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="5b825984-72f9-589b-f4a1-d3fbad1d9598" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 30, "hidden": false, "row": 0, "width": 12}, "report_default": {"hidden": false}}}}
# # **An Interactive Data Science Tutorial**
#
#
# *[Based on the Titanic competition on Kaggle](https://www.kaggle.com/c/titanic)*
#
# *by <NAME> & <NAME>*
#
# *January 2017*
#
# ---
#
# ## Content
#
#
# 1. Business Understanding (5 min)
# * Objective
# * Description
# 2. Data Understanding (15 min)
# * Import Libraries
# * Load data
# * Statistical summaries and visualisations
# * Excersises
# 3. Data Preparation (5 min)
# * Missing values imputation
# * Feature Engineering
# 4. Modeling (5 min)
# * Build the model
# 5. Evaluation (25 min)
# * Model performance
# * Feature importance
# * Who gets the best performing model?
# 6. Deployment (5 min)
# * Submit result to Kaggle leaderboard
#
# [*Adopted from Cross Industry Standard Process for Data Mining (CRISP-DM)*](http://www.sv-europe.com/crisp-dm-methodology/)
#
# 
# + [markdown] _cell_guid="6f9380bf-1835-9f4f-b728-48bdb84e7cab" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 15, "hidden": false, "row": 30, "width": 12}, "report_default": {"hidden": false}}}}
# # 1. Business Understanding
#
# ## 1.1 Objective
# Predict survival on the Titanic
#
# ## 1.2 Description
# The sinking of the RMS Titanic is one of the most infamous shipwrecks in history. On April 15, 1912, during her maiden voyage, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 passengers and crew. This sensational tragedy shocked the international community and led to better safety regulations for ships.
#
# One of the reasons that the shipwreck led to such loss of life was that there were not enough lifeboats for the passengers and crew. Although there was some element of luck involved in surviving the sinking, some groups of people were more likely to survive than others, such as women, children, and the upper-class.
#
# In this challenge, we ask you to complete the analysis of what sorts of people were likely to survive. In particular, we ask you to apply the tools of machine learning to predict which passengers survived the tragedy.
#
# **Before going further, what do you think is the most important reasons passangers survived the Titanic sinking?**
#
# [Description from Kaggle](https://www.kaggle.com/c/titanic)
# + [markdown] _cell_guid="af225757-e074-9b64-ba8f-137252ea90e0" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 6, "hidden": false, "row": 45, "width": 12}, "report_default": {"hidden": false}}}}
# # 2. Data Understanding
#
# ## 2.1 Import Libraries
# First of some preparation. We need to import python libraries containing the necessary functionality we will need.
#
# *Simply run the cell below by selecting it and pressing the play button.*
# + _cell_guid="d3cb1c42-90ba-9674-0e72-0b4ee496fa42" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# Handle table-like data and matrices
import numpy as np
import pandas as pd
# Modelling Algorithms
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier , GradientBoostingClassifier
# Modelling Helpers
from sklearn.preprocessing import Imputer , Normalizer , scale
from sklearn.cross_validation import train_test_split , StratifiedKFold
from sklearn.feature_selection import RFECV
# Visualisation
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
# Configure visualisations
# %matplotlib inline
mpl.style.use( 'ggplot' )
sns.set_style( 'white' )
pylab.rcParams[ 'figure.figsize' ] = 8 , 6
# + [markdown] _cell_guid="cb44cb03-a5be-653d-34bf-f1e7eba133a4" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 51, "width": 12}, "report_default": {"hidden": false}}}}
# ## 2.2 Setup helper Functions
# There is no need to understand this code. Just run it to simplify the code later in the tutorial.
#
# *Simply run the cell below by selecting it and pressing the play button.*
# + _cell_guid="0bb8cf49-d080-46a7-2c66-fa967ad4db97" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
def plot_histograms( df , variables , n_rows , n_cols ):
fig = plt.figure( figsize = ( 16 , 12 ) )
for i, var_name in enumerate( variables ):
ax=fig.add_subplot( n_rows , n_cols , i+1 )
df[ var_name ].hist( bins=10 , ax=ax )
ax.set_title( 'Skew: ' + str( round( float( df[ var_name ].skew() ) , ) ) ) # + ' ' + var_name ) #var_name+" Distribution")
ax.set_xticklabels( [] , visible=False )
ax.set_yticklabels( [] , visible=False )
fig.tight_layout() # Improves appearance a bit.
plt.show()
def plot_distribution( df , var , target , **kwargs ):
row = kwargs.get( 'row' , None )
col = kwargs.get( 'col' , None )
facet = sns.FacetGrid( df , hue=target , aspect=4 , row = row , col = col )
facet.map( sns.kdeplot , var , shade= True )
facet.set( xlim=( 0 , df[ var ].max() ) )
facet.add_legend()
def plot_categories( df , cat , target , **kwargs ):
row = kwargs.get( 'row' , None )
col = kwargs.get( 'col' , None )
facet = sns.FacetGrid( df , row = row , col = col )
facet.map( sns.barplot , cat , target )
facet.add_legend()
def plot_correlation_map( df ):
corr = titanic.corr()
_ , ax = plt.subplots( figsize =( 12 , 10 ) )
cmap = sns.diverging_palette( 220 , 10 , as_cmap = True )
_ = sns.heatmap(
corr,
cmap = cmap,
square=True,
cbar_kws={ 'shrink' : .9 },
ax=ax,
annot = True,
annot_kws = { 'fontsize' : 12 }
)
def describe_more( df ):
var = [] ; l = [] ; t = []
for x in df:
var.append( x )
l.append( len( pd.value_counts( df[ x ] ) ) )
t.append( df[ x ].dtypes )
levels = pd.DataFrame( { 'Variable' : var , 'Levels' : l , 'Datatype' : t } )
levels.sort_values( by = 'Levels' , inplace = True )
return levels
def plot_variable_importance( X , y ):
tree = DecisionTreeClassifier( random_state = 99 )
tree.fit( X , y )
plot_model_var_imp( tree , X , y )
def plot_model_var_imp( model , X , y ):
imp = pd.DataFrame(
model.feature_importances_ ,
columns = [ 'Importance' ] ,
index = X.columns
)
imp = imp.sort_values( [ 'Importance' ] , ascending = True )
imp[ : 10 ].plot( kind = 'barh' )
print (model.score( X , y ))
# + [markdown] _cell_guid="ea8b0e99-e512-f1f5-ed3d-e7df876b9bed" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 56, "width": 12}, "report_default": {"hidden": false}}}}
# ## 2.3 Load data
# Now that our packages are loaded, let's read in and take a peek at the data.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="ee2677e2-b78b-250b-a908-816109e3ab91" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 61, "width": 4}, "report_default": {"hidden": false}}}}
# get titanic & test csv files as a DataFrame
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
full = train.append( test , ignore_index = True )
titanic = full[ :891 ]
del train , test
print ('Datasets:' , 'full:' , full.shape , 'titanic:' , titanic.shape)
# + [markdown] _cell_guid="76852c59-23bf-55b6-5b6d-f672c97114ae" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 6, "hidden": false, "row": 65, "width": 12}, "report_default": {"hidden": false}}}}
# ## 2.4 Statistical summaries and visualisations
#
# To understand the data we are now going to consider some key facts about various variables including their relationship with the target variable, i.e. survival.
#
# We start by looking at a few lines of the data
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="60d5efad-7649-c1e8-3cae-59bf562e8457" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 14, "hidden": false, "row": 71, "width": 8}, "report_default": {"hidden": false}}}}
# Run the code to see the variables, then read the variable description below to understand them.
titanic.head()
# + [markdown] _cell_guid="1f3708e1-b5ca-7540-39fc-da7453d0fb80" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 14, "hidden": false, "row": 85, "width": 12}, "report_default": {"hidden": false}}}}
# **VARIABLE DESCRIPTIONS:**
#
# We've got a sense of our variables, their class type, and the first few observations of each. We know we're working with 1309 observations of 12 variables. To make things a bit more explicit since a couple of the variable names aren't 100% illuminating, here's what we've got to deal with:
#
#
# **Variable Description**
#
# - Survived: Survived (1) or died (0)
# - Pclass: Passenger's class
# - Name: Passenger's name
# - Sex: Passenger's sex
# - Age: Passenger's age
# - SibSp: Number of siblings/spouses aboard
# - Parch: Number of parents/children aboard
# - Ticket: Ticket number
# - Fare: Fare
# - Cabin: Cabin
# - Embarked: Port of embarkation
#
# [More information on the Kaggle site](https://www.kaggle.com/c/titanic/data)
# + [markdown] _cell_guid="bcc371f8-8fbb-a582-5944-8537c152c6c9" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 99, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.1 Next have a look at some key information about the variables
# An numeric variable is one with values of integers or real numbers while a categorical variable is a variable that can take on one of a limited, and usually fixed, number of possible values, such as blood type.
#
# Notice especially what type of variable each is, how many observations there are and some of the variable values.
#
# An interesting observation could for example be the minimum age 0.42, do you know why this is?
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="88c8b958-0973-d27e-d463-58fada41900e" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 9, "hidden": false, "row": 107, "width": 7}, "report_default": {"hidden": false}}}}
titanic.describe()
# + [markdown] _cell_guid="c006f42b-cba3-7109-92fb-79676f726afb" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 116, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.2 A heat map of correlation may give us a understanding of which variables are important
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="84418f3e-32ab-7a6c-a60e-6d45760ee666" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 11, "hidden": false, "row": 71, "width": 4}, "report_default": {"hidden": false}}}}
plot_correlation_map( titanic )
# + [markdown] _cell_guid="952846c2-63a9-1923-2501-d3036056855b" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 120, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.3 Let's further explore the relationship between the features and survival of passengers
# We start by looking at the relationship between age and survival.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="617703c7-4ab2-186c-f4b1-40d5ffa37850" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 7, "height": 7, "hidden": false, "row": 107, "width": 4}, "report_default": {"hidden": false}}}}
# Plot distributions of Age of passangers who survived or did not survive
plot_distribution( titanic , var = 'Age' , target = 'Survived' , row = 'Sex' )
# + [markdown] _cell_guid="834c62d5-23e7-7e0c-d2d4-bc960d09a409" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 125, "width": 12}, "report_default": {"hidden": false}}}}
# Consider the graphs above. Differences between survival for different values is what will be used to separate the target variable (survival in this case) in the model. If the two lines had been about the same, then it would not have been a good variable for our predictive model.
#
# Consider some key questions such as; what age does males/females have a higher or lower probability of survival?
# + [markdown] _cell_guid="f8d077bf-e70c-29df-253e-5bd22ff3f06e" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 129, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.3 Excersise 1: Investigating numeric variables
# It's time to get your hands dirty and do some coding! Try to plot the distributions of Fare of passangers who survived or did not survive. Then consider if this could be a good predictive variable.
#
# *Hint: use the code from the previous cell as a starting point.*
# + _cell_guid="6fcddc9f-19be-c474-a639-79c6b2d2a41a" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Excersise 1
# Plot distributions of Fare of passangers who survived or did not survive
plot_distribution( titanic , var = 'Fare' , target = 'Survived' )
# + [markdown] _cell_guid="e9dffe91-ee29-9cca-2860-29a32a44e2af" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 6, "hidden": false, "row": 134, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.4 Embarked
# We can also look at categorical variables like Embarked and their relationship with survival.
#
# - C = Cherbourg
# - Q = Queenstown
# - S = Southampton
# + _cell_guid="b5beadda-fe40-d017-9bf8-29d372401c10" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 140, "width": 4}, "report_default": {"hidden": false}}}}
# Plot survival rate by Embarked
plot_categories( titanic , cat = 'Embarked' , target = 'Survived' )
# + [markdown] _cell_guid="33e7d091-af47-9a77-c907-81e710200c5f" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 6, "hidden": false, "row": 148, "width": 12}, "report_default": {"hidden": false}}}}
# ### 2.4.4 Excersise 2 - 5: Investigating categorical variables
# Even more coding practice! Try to plot the survival rate of Sex, Pclass, SibSp and Parch below.
#
# *Hint: use the code from the previous cell as a starting point.*
#
# After considering these graphs, which variables do you expect to be good predictors of survival?
# + _cell_guid="25022b4c-a631-45fa-ec2b-e7bd7ed11987" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Excersise 2
# Plot survival rate by Sex
plot_categories( titanic , cat = 'Sex' , target = 'Survived' )
# + _cell_guid="8c7af046-677c-07a7-d4bf-b28d10092f73" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Excersise 3
# Plot survival rate by Pclass
plot_categories( titanic , cat = 'Pclass' , target = 'Survived' )
# + _cell_guid="f5b4252e-4800-d674-f86f-01eba244230d" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Excersise 4
# Plot survival rate by SibSp
plot_categories( titanic , cat = 'SibSp' , target = 'Survived' )
# + _cell_guid="4dc79695-082e-84c0-0f0d-6fca5ad414dd" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Excersise 5
# Plot survival rate by Parch
plot_categories( titanic , cat = 'Parch' , target = 'Survived' )
# + [markdown] _cell_guid="c9888e98-091a-d98c-6ccf-b733a4f0d941" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 4, "hidden": false, "row": 61, "width": 4}, "report_default": {"hidden": false}}}}
# # 3. Data Preparation
# + [markdown] _cell_guid="e280f761-4b30-9776-09e3-fcaecf932a01" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 7, "hidden": false, "row": 154, "width": 12}, "report_default": {"hidden": false}}}}
# ## 3.1 Categorical variables need to be transformed to numeric variables
# The variables *Embarked*, *Pclass* and *Sex* are treated as categorical variables. Some of our model algorithms can only handle numeric values and so we need to create a new variable (dummy variable) for every unique value of the categorical variables.
#
# This variable will have a value 1 if the row has a particular value and a value 0 if not. *Sex* is a dichotomy (old school gender theory) and will be encoded as one binary variable (0 or 1).
#
# *Select the cells below and run it by pressing the play button.*
# + _cell_guid="75023b46-ded0-94ab-3ea1-7da1eec945fe" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# Transform Sex into binary values 0 and 1
sex = pd.Series( np.where( full.Sex == 'male' , 1 , 0 ) , name = 'Sex' )
# + _cell_guid="cf816542-c40f-abe0-b662-a91211882bff" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 7, "hidden": false, "row": 140, "width": 4}, "report_default": {"hidden": false}}}}
# Create a new variable for every unique value of Embarked
embarked = pd.get_dummies( full.Embarked , prefix='Embarked' )
embarked.head()
# + _cell_guid="01758920-20cc-e200-90aa-e4404c41a8d7" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 7, "hidden": false, "row": 140, "width": 4}, "report_default": {"hidden": false}}}}
# Create a new variable for every unique value of Embarked
pclass = pd.get_dummies( full.Pclass , prefix='Pclass' )
pclass.head()
# + [markdown] _cell_guid="d1269afc-b929-d519-9646-146f8a91b472" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 161, "width": 12}, "report_default": {"hidden": false}}}}
# ## 3.2 Fill missing values in variables
# Most machine learning alghorims require all variables to have values in order to use it for training the model. The simplest method is to fill missing values with the average of the variable across all observations in the training set.
#
# *Select the cells below and run it by pressing the play button.*
# + _cell_guid="04044a9c-603d-1963-34f4-a85efb8c9166" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 7, "hidden": false, "row": 166, "width": 4}, "report_default": {"hidden": false}}}}
# Create dataset
imputed = pd.DataFrame()
# Fill missing values of Age with the average of Age (mean)
imputed[ 'Age' ] = full.Age.fillna( full.Age.mean() )
# Fill missing values of Fare with the average of Fare (mean)
imputed[ 'Fare' ] = full.Fare.fillna( full.Fare.mean() )
imputed.head()
# + [markdown] _cell_guid="4f2ce8f7-8bf6-69f8-e2ae-5593b5cf98ac" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 173, "width": 12}, "report_default": {"hidden": false}}}}
# ## 3.3 Feature Engineering – Creating new variables
# Credit: http://ahmedbesbes.com/how-to-score-08134-in-titanic-kaggle-challenge.html
# + [markdown] _cell_guid="bd8efd0f-4af4-d709-cfeb-1907f885e341" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 177, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.3.1 Extract titles from passenger names
# Titles reflect social status and may predict survival probability
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="32772d9e-8c05-4eaf-5cd3-8425ae5837cd" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 7, "hidden": false, "row": 166, "width": 4}, "report_default": {"hidden": false}}}}
title = pd.DataFrame()
# we extract the title from each name
title[ 'Title' ] = full[ 'Name' ].map( lambda name: name.split( ',' )[1].split( '.' )[0].strip() )
# a map of more aggregated titles
Title_Dictionary = {
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir" : "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess":"Royalty",
"Dona": "Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr" : "Mr",
"Mrs" : "Mrs",
"Miss" : "Miss",
"Master" : "Master",
"Lady" : "Royalty"
}
# we map each title
title[ 'Title' ] = title.Title.map( Title_Dictionary )
title = pd.get_dummies( title.Title )
#title = pd.concat( [ title , titles_dummies ] , axis = 1 )
title.head()
# + [markdown] _cell_guid="a56838a1-5d47-194f-7ac6-06c397b60482" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 182, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.3.2 Extract Cabin category information from the Cabin number
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="87ed124f-d659-bf03-7dbf-accee0ddfe75" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 7, "hidden": false, "row": 186, "width": 7}, "report_default": {"hidden": false}}}}
cabin = pd.DataFrame()
# replacing missing cabins with U (for Uknown)
cabin[ 'Cabin' ] = full.Cabin.fillna( 'U' )
# mapping each Cabin value with the cabin letter
cabin[ 'Cabin' ] = cabin[ 'Cabin' ].map( lambda c : c[0] )
# dummy encoding ...
cabin = pd.get_dummies( cabin['Cabin'] , prefix = 'Cabin' )
cabin.head()
# + [markdown] _cell_guid="d2dba1d1-e7cd-b9c8-fcc7-fae22cca38da" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 193, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.3.3 Extract ticket class from ticket number
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="5319db41-01dd-3a48-83d8-56572272d966" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 197, "width": 12}, "report_default": {"hidden": false}}}}
# a function that extracts each prefix of the ticket, returns 'XXX' if no prefix (i.e the ticket is a digit)
def cleanTicket( ticket ):
ticket = ticket.replace( '.' , '' )
ticket = ticket.replace( '/' , '' )
ticket = ticket.split()
ticket = map( lambda t : t.strip() , ticket )
ticket = list(filter( lambda t : not t.isdigit() , ticket ))
if len( ticket ) > 0:
return ticket[0]
else:
return 'XXX'
ticket = pd.DataFrame()
# Extracting dummy variables from tickets:
ticket[ 'Ticket' ] = full[ 'Ticket' ].map( cleanTicket )
ticket = pd.get_dummies( ticket[ 'Ticket' ] , prefix = 'Ticket' )
ticket.shape
ticket.head()
# + [markdown] _cell_guid="7e812a29-3885-30a9-3d51-beda531016d9" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 205, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.3.4 Create family size and category for family size
# The two variables *Parch* and *SibSp* are used to create the famiy size variable
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="d885986a-77ea-b35c-5ddc-c16f54357232" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 7, "hidden": false, "row": 166, "width": 4}, "report_default": {"hidden": false}}}}
family = pd.DataFrame()
# introducing a new feature : the size of families (including the passenger)
family[ 'FamilySize' ] = full[ 'Parch' ] + full[ 'SibSp' ] + 1
# introducing other features based on the family size
family[ 'Family_Single' ] = family[ 'FamilySize' ].map( lambda s : 1 if s == 1 else 0 )
family[ 'Family_Small' ] = family[ 'FamilySize' ].map( lambda s : 1 if 2 <= s <= 4 else 0 )
family[ 'Family_Large' ] = family[ 'FamilySize' ].map( lambda s : 1 if 5 <= s else 0 )
family.head()
# + [markdown] _cell_guid="c20b06e8-7126-a80e-762a-8a383ffba9e1" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 210, "width": 12}, "report_default": {"hidden": false}}}}
# ## 3.4 Assemble final datasets for modelling
#
# Split dataset by rows into test and train in order to have a holdout set to do model evaluation on. The dataset is also split by columns in a matrix (X) containing the input data and a vector (y) containing the target (or labels).
# + [markdown] _cell_guid="29663deb-5bbf-b621-abe5-37cfa5e3d29f" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 10, "hidden": false, "row": 214, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.4.1 Variable selection
# Select which features/variables to inculde in the dataset from the list below:
#
# - imputed
# - embarked
# - pclass
# - sex
# - family
# - cabin
# - ticket
#
# *Include the variables you would like to use in the function below seperated by comma, then run the cell*
# + _cell_guid="4b3459bd-752c-0a96-22dc-8672900a0bb9" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 7, "hidden": false, "row": 224, "width": 11}, "report_default": {"hidden": false}}}}
# Select which features/variables to include in the dataset from the list below:
# imputed , embarked , pclass , sex , family , cabin , ticket
full_X = pd.concat( [ imputed , embarked , cabin , sex ] , axis=1 )
full_X.head()
# + [markdown] _cell_guid="cb10186c-81b3-f9cd-250a-fab46b0001f5" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 231, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.4.2 Create datasets
# Below we will seperate the data into training and test datasets.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="7ca114b3-f22c-b8bf-4e61-9c9bcd82fa63" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 4, "hidden": false, "row": 61, "width": 4}, "report_default": {"hidden": false}}}}
# Create all datasets that are necessary to train, validate and test models
train_valid_X = full_X[ 0:891 ]
train_valid_y = titanic.Survived
test_X = full_X[ 891: ]
train_X , valid_X , train_y , valid_y = train_test_split( train_valid_X , train_valid_y , train_size = .7 )
print (full_X.shape , train_X.shape , valid_X.shape , train_y.shape , valid_y.shape , test_X.shape)
# + [markdown] _cell_guid="f269d9b9-73fc-6d34-b46a-61fd6f1357ae" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 236, "width": 12}, "report_default": {"hidden": false}}}}
# ### 3.4.3 Feature importance
# Selecting the optimal features in the model is important.
# We will now try to evaluate what the most important variables are for the model to make the prediction.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="17c2f033-c43d-93f8-a697-bf32eec3b550" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 10, "hidden": false, "row": 241, "width": 4}, "report_default": {"hidden": false}}}}
plot_variable_importance(train_X, train_y)
# + [markdown] _cell_guid="84f76826-572a-e65d-7dd7-cd3791bf5237" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 251, "width": 12}, "report_default": {"hidden": false}}}}
# # 4. Modeling
# We will now select a model we would like to try then use the training dataset to train this model and thereby check the performance of the model using the test set.
#
# ## 4.1 Model Selection
# Then there are several options to choose from when it comes to models. A good starting point is logisic regression.
#
# **Select ONLY the model you would like to try below and run the corresponding cell by pressing the play button.**
# + [markdown] _cell_guid="bd56cc59-9abd-fb1f-e5e1-211c48776863" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 7, "height": 4, "hidden": false, "row": 186, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.1 Random Forests Model
# Try a random forest model by running the cell below.
# + _cell_guid="29893a22-aa92-b35d-881e-7ce907f3b3b2" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = RandomForestClassifier(n_estimators=100)
# + [markdown] _cell_guid="777e1893-585a-c545-250a-f6ea12c10fb9" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 4, "hidden": false, "row": 241, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.2 Support Vector Machines
# Try a Support Vector Machines model by running the cell below.
# + _cell_guid="c34e2a74-23b9-2916-683d-f0d6956ad5e6" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = SVC()
# + [markdown] _cell_guid="bc024820-b3db-4b2f-01aa-f9b35e413be1" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 4, "hidden": false, "row": 241, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.3 Gradient Boosting Classifier
# Try a Gradient Boosting Classifier model by running the cell below.
# + _cell_guid="120f00c8-b568-f3af-97fd-df9762d4aefb" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = GradientBoostingClassifier()
# + [markdown] _cell_guid="452de3e7-c672-0aeb-b49e-5bb6c400de75" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 4, "hidden": false, "row": 245, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.4 K-nearest neighbors
# Try a k-nearest neighbors model by running the cell below.
# + _cell_guid="862cf425-ab7c-6abb-3f86-fe6a63f4a790" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = KNeighborsClassifier(n_neighbors = 3)
# + [markdown] _cell_guid="34eb3eed-6d89-5c9e-31e7-40eb7e01639e" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 4, "hidden": false, "row": 245, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.5 Gaussian Naive Bayes
# Try a Gaussian Naive Bayes model by running the cell below.
# + _cell_guid="e6b97cb8-56c1-3b01-ca52-c2e9685b68d0" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = GaussianNB()
# + [markdown] _cell_guid="c3c89f76-4d2e-ae80-8b61-13feb6f33831" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 259, "width": 4}, "report_default": {"hidden": false}}}}
# ### 4.1.6 Logistic Regression
# Try a Logistic Regression model by running the cell below.
# + _cell_guid="f3573fce-2045-aa1d-e010-dc28139b5a16" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
model = LogisticRegression()
# + [markdown] _cell_guid="7e8d2a8d-a9f6-d416-74e7-4f5b711dcd98" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 263, "width": 12}, "report_default": {"hidden": false}}}}
# ## 4.2 Train the selected model
# When you have selected a dataset with the features you want and a model you would like to try it is now time to train the model. After all our preparation model training is simply done with the one line below.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="02a54d08-b02a-a4df-6d74-540be7d243d8" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 268, "width": 4}, "report_default": {"hidden": false}}}}
model.fit( train_X , train_y )
# + [markdown] _cell_guid="3868397c-30ed-a870-9c61-30b5d5ebdce7" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 10, "hidden": false, "row": 273, "width": 12}, "report_default": {"hidden": false}}}}
# # 5. Evaluation
# Now we are going to evaluate model performance and the feature importance.
#
# ## 5.1 Model performance
# We can evaluate the accuracy of the model by using the validation set where we know the actual outcome. This data set have not been used for training the model, so it's completely new to the model.
#
# We then compare this accuracy score with the accuracy when using the model on the training data. If the difference between these are significant this is an indication of overfitting. We try to avoid this because it means the model will not generalize well to new data and is expected to perform poorly.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="c4e9a0a3-f637-5568-e92c-07c0548cd89d" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 4, "hidden": false, "row": 259, "width": 4}, "report_default": {"hidden": false}}}}
# Score the model
print (model.score( train_X , train_y ) , model.score( valid_X , valid_y ))
# + [markdown] _cell_guid="cd75f2ca-dcb1-e2af-ebf8-e1dea0cf44a5" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 283, "width": 12}, "report_default": {"hidden": false}}}}
# ## 5.2 Feature importance - selecting the optimal features in the model
# We will now try to evaluate what the most important variables are for the model to make the prediction. The function below will only work for decision trees, so if that's the model you chose you can uncomment the code below (remove # in the beginning) and see the feature importance.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="f9412dd9-11ce-74ba-7fd6-148b2ae3bce3" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
#plot_model_var_imp(model, train_X, train_y)
# + [markdown] _cell_guid="e7d3ad7a-92eb-f0e0-4028-3eec53c65581" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 5, "hidden": false, "row": 288, "width": 12}, "report_default": {"hidden": false}}}}
# ### 5.2.1 Automagic
# It's also possible to automatically select the optimal number of features and visualize this. This is uncommented and can be tried in the competition part of the tutorial.
#
# *Select the cell below and run it by pressing the play button.*
# + _cell_guid="2e290c27-d9d2-4ebf-32bc-8ff56c68f763" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 293, "width": 4}, "report_default": {"hidden": false}}}}
rfecv = RFECV( estimator = model , step = 1 , cv = StratifiedKFold( train_y , 2 ) , scoring = 'accuracy' )
rfecv.fit( train_X , train_y )
#print (rfecv.score( train_X , train_y ) , rfecv.score( valid_X , valid_y ))
#print( "Optimal number of features : %d" % rfecv.n_features_ )
# Plot number of features VS. cross-validation scores
#plt.figure()
#plt.xlabel( "Number of features selected" )
#plt.ylabel( "Cross validation score (nb of correct classifications)" )
#plt.plot( range( 1 , len( rfecv.grid_scores_ ) + 1 ) , rfecv.grid_scores_ )
#plt.show()
# + [markdown] _cell_guid="61945225-1dcd-870d-bc49-851d5c91d1d5" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 8, "hidden": false, "row": 301, "width": 12}, "report_default": {"hidden": false}}}}
# ## 5.3 Competition time!
# It's now time for you to get your hands even dirtier and go at it all by yourself in a `challenge`!
#
# 1. Try to the other models in step 4.1 and compare their result
# * Do this by uncommenting the code and running the cell you want to try
# 2. Try adding new features in step 3.4.1
# * Do this by adding them in to the function in the feature section.
#
#
# **The winner is the one to get the highest scoring model for the validation set**
# + [markdown] _cell_guid="598cfdec-2804-0312-e69e-2ea779f045bc" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 7, "hidden": false, "row": 309, "width": 12}, "report_default": {"hidden": false}}}}
# # 6. Deployment
#
# Deployment in this context means publishing the resulting prediction from the model to the Kaggle leaderboard. To do this do the following:
#
# 1. select the cell below and run it by pressing the play button.
# 2. Press the `Publish` button in top right corner.
# 3. Select `Output` on the notebook menubar
# 4. Select the result dataset and press `Submit to Competition` button
# + _cell_guid="25cb37ca-225b-2917-00a8-5c08708ccf1d" extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
test_Y = model.predict( test_X )
passenger_id = full[891:].PassengerId
test = pd.DataFrame( { 'PassengerId': passenger_id , 'Survived': test_Y } )
test.shape
test.head()
test.to_csv( 'titanic_pred.csv' , index = False )
| Kaggle/KaggleTitanic/kaggleTitanicExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ada
# language: python
# name: ada
# ---
# +
from importlib import reload
import script_util
reload(script_util)
from script_util import root, rcParams, cmap
from typing import *
import matplotlib.pyplot as plt
from pathlib import Path
import json
import numpy as np
plt.rcParams.update(rcParams)
unit = 1
cmap
# -
B = 4
master = {
"v3-mirror-auto4-noaug":{
"paths":[root/"training116/00013-v3-mirror-auto4-noaug"],
"color": cmap(0*B+0),
"linestyle": "solid",
"label": "背景白で腰上の立ち絵,2万枚,augなし",
},
"v3-mirror-auto4-bg":{
"paths": [root/"training115/00009-v3-mirror-auto4-bg", root/"training115/00010-v3-mirror-auto4-bg-resumecustom", root/"training116/00014-v3-mirror-auto4-bg-resumecustom"],
"color": cmap(0*B+3),
"linestyle": "dashed",
"label": "背景白で腰上の立ち絵,2万枚,augはbitblit+geometry",
},
"v4-mirror-auto4-noaug":{
"paths":[root/"training116/00015-v4-mirror-auto4-noaug"],
"color": cmap(1*B+0),
"linestyle": "solid",
"label": "背景白で腰上の立ち絵,8万枚,augなし",
},
"v4-mirror-auto4-bg":{
"paths":[root/"training116/00016-v4-mirror-auto4-bg"],
"color": cmap(1*B+3),
"linestyle": "dashed",
"label": "背景白で腰上の立ち絵,8万枚,augはbitblit+geometry",
},
"portraits-mirror-auto4-noaug":{
"paths":[root/"training116/00009-portraits-mirror-auto4-noaug"],
"color": cmap(2*B+0),
"linestyle": "solid",
"label": "顔アップ,30万枚,augなし",
},
"portraits-mirror-auto4":{
"paths":[root/"training116/00010-portraits-mirror-auto4"],
"color": cmap(2*B+2),
"linestyle": "dashed",
"label": "顔アップ,30万枚,augはbitblit+geometry+color",
},
"portraits-mirror-auto4-target0.8-bg":{
"paths":[root/"training115/00014-portraits-mirror-auto4-target0.8-bg"],
"color": cmap(2*B+3),
"linestyle": "dashdot",
"label": "顔アップ,30万枚,augはbg,target=0.8",
},
"white_yc05_yw04-mirror-auto4-noaug":{
"paths":[root/"training116/00018-white_yc05_yw04-mirror-auto4-noaug"],
"color": cmap(3*B+0),
"linestyle": "solid",
# "label": "顔アップ(v4),8万枚,augなし",
},
"white_yc05_yw04-mirror-auto4-target0.7-bg":{
"paths":[root/"training116/00020-white_yc05_yw04-mirror-auto4-target0.7-bg"],
"color": cmap(3*B+3),
"linestyle": "dashdot",
# "label": "顔アップ(v4),8万枚,augなし",
},
}
# +
class Result(NamedTuple):
path: Path
metric: Dict[str, Any]
class Results:
def __init__(self, paths, metric="fid50k_full"):
self.results = list()
self.metric = metric
for i, path in enumerate(paths):
for j, line in enumerate((path/f"metric-{self.metric}.jsonl").read_text().strip().split("\n")):
if i != 0 and j == 0: continue
metric = json.loads(line)
self.results.append(Result(path, metric))
def __len__(self):
return len(self.results)
def metric_list(self):
return [metric["results"][self.metric] for path, metric in self.results]
def image(self, i):
path, metric = self.results[i]
num = metric["snapshot_pkl"].split("-")[-1].split(".")[0]
return path/f"fakes{num}.png"
# -
def myplot(names: List[str], best=1, metric="fid50k_full"):
fig, ax = plt.subplots()
maxX = 0
maxY = 0
minY = float("inf")
for name in names:
results = Results(master[name]["paths"], metric=metric)
Y = results.metric_list()
X = [unit*i for i in range(len(Y))]
maxX = max(maxX, max(X))
maxY = max(maxY, max(Y))
minY = min(minY, min(Y))
YX = list(zip(Y, X))
YX.sort()
bests = set()
for i in range(best):
y, x = YX[i]
bests.add(y)
# ax.text(x, 40*y-1200, f"{y:.1f}\n{str(results.image(x))}", color=master[name]["color"])
# ax.text(x, y-50, f"{y:.1f}\n{x}", color=master[name]["color"])
ax.text(x, y-minY, f"{y:.0f}", color=master[name]["color"])
plt.scatter([x], [y], lw=1, marker="x", color=master[name]["color"])
plt.plot(
X, Y,
lw=1, marker="",
label=master[name].get("label", name),
color=master[name].get("color", cmap(0)),
linestyle=master[name].get("linestyle", "solid")
)
ax.set_xticks([x for x in range(int(maxX)) if x%10 == 0])
ax.set_xticks([x for x in range(int(maxX)) if x%2 == 0], minor=True)
ax.set_yticks([y for y in range(int(maxY)) if y%10 == 0], minor=True)
ax.grid(which="major", alpha=0.6)
ax.grid(which="minor", alpha=0.2)
plt.xlabel("*200kimg")
plt.ylabel(metric)
plt.legend()
myplot([
"v3-mirror-auto4-noaug",
"v3-mirror-auto4-bg",
"v4-mirror-auto4-noaug",
"v4-mirror-auto4-bg",
])
myplot([
"portraits-mirror-auto4-noaug",
"portraits-mirror-auto4",
"portraits-mirror-auto4-target0.8-bg",
"white_yc05_yw04-mirror-auto4-noaug",
"white_yc05_yw04-mirror-auto4-target0.7-bg",
])
myplot(["v3-mirror-auto4-noaug", "v4-mirror-auto4-noaug"], metric="ppl2_wend", best=0)
# + tags=[]
myplot(["v3-mirror-auto4-bg","v3-mirror-auto4-noaug", "v4-mirror-auto4-noaug"], metric="kid50k_full", best=0)
| notebook_comparision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="vJ7lKzXZ-yUl"
import os
import time
import random
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.utils.data import DataLoader, Dataset
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.metrics import f1_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import confusion_matrix
# + id="yGeNhGm5o7jh"
random.seed(0)
np.random.seed(0)
# + colab={"base_uri": "https://localhost:8080/"} id="1uMjtpRDr-Da" outputId="935b9d2c-276f-456f-8645-34b436b1cdaf"
# Change this to run locally
is_local = False
if is_local:
data_loc = "../process/project_processed_data/"
else:
from google.colab import drive
drive.mount('/content/drive')
data_loc = "./drive/MyDrive/log_data/"
# + [markdown] id="wsxvHdJI-yUt"
# ## Load and Prepare Data
# + id="Uoav9WT1-yUu"
# Train sets
train_data = np.load('{}x_train_tf-idf.npy'.format(data_loc))
read_train_labels = pd.read_csv('{}y_train_tf-idf.csv'.format(data_loc))
train_labels = read_train_labels['Label'] == 'Anomaly'
train_labels = train_labels.astype(int)
# Test sets
test_data = np.load('{}x_test_tf-idf.npy'.format(data_loc))
read_test_labels = pd.read_csv('{}y_test_tf-idf.csv'.format(data_loc))
test_labels = read_test_labels['Label'] == 'Anomaly'
test_labels = test_labels.astype(int)
# + id="X09WrmMF-yUu"
train_data, val_data, train_labels, val_labels = train_test_split(train_data, train_labels, test_size=0.2, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="ta7khuA6F3e7" outputId="9c4c6267-7448-4cc9-f200-86828c7136f6"
train_labels = train_labels.reset_index(drop=True)
val_labels = val_labels.reset_index(drop=True)
print("train data shape")
print(train_data.shape)
print(train_labels.shape)
print("val data shape")
print(val_data.shape)
print(val_labels.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="LNcDmplg4GOp" outputId="be2d4298-3ba0-4dde-ee2d-f63743d24765"
print(len(test_labels))
anom = test_labels[test_labels > 0]
norm = test_labels[test_labels == 0]
print(anom.shape)
print(norm.shape)
print(len(anom)/len(test_labels))
# + colab={"base_uri": "https://localhost:8080/"} id="PL1F69-SsShW" outputId="d8c2f473-baa5-4237-dc05-034147aaab71"
print(len(train_labels))
anom = train_labels[train_labels > 0]
norm = train_labels[train_labels == 0]
print(anom.shape)
print(norm.shape)
print(len(anom)/len(train_labels))
# + id="sPraFUqn-yUx"
class logDataset(Dataset):
"""Log Anomaly Features Dataset"""
def __init__(self, data_vec, labels=None):
self.X = data_vec
self.y = labels
def __len__(self):
return len(self.X)
def __getitem__(self, idx):
data_matrix = self.X[idx]
if not self.y is None:
return(data_matrix, self.y[idx])
else:
return data_matrix
# + id="Ol2-sxy5-yUy"
# add 1 1 1 1 for padding
train_data = torch.tensor(train_data, dtype=torch.float32)
train_data = F.pad(input=train_data, pad=(1, 1, 1, 1), mode='constant', value=0) # pad all sides with 0s
train_data = np.expand_dims(train_data, axis=1)
test_data = torch.tensor(test_data, dtype=torch.float32)
test_data = F.pad(input=test_data, pad=(1, 1, 1, 1), mode='constant', value=0)
test_data = np.expand_dims(test_data, axis=1)
val_data = torch.tensor(val_data, dtype=torch.float32)
val_data = F.pad(input=val_data, pad=(1, 1, 1, 1), mode='constant', value=0)
val_data = np.expand_dims(val_data, axis=1)
# + id="ShW0RdNY-yU0"
# Hyperparameters
RANDOM_SEED = 1
LEARNING_RATE = 0.001
BATCH_SIZE = 128
NUM_EPOCHS = 10
NUM_CLASSES = 2
# Other
if torch.cuda.is_available():
DEVICE = "cuda:0"
else:
DEVICE = "cpu"
# + id="I7_3tMRl-yU0"
# pass datasets into the custom dataclass
train_dataset = logDataset(train_data, labels = train_labels)
test_dataset = logDataset(test_data, labels = test_labels)
val_dataset = logDataset(val_data, labels = val_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="zWwa2PG_-yU1" outputId="11b29c5f-192a-4d58-c289-ead77ad24692"
# use DataLoader class
train_loader = DataLoader(dataset=train_dataset,
batch_size=BATCH_SIZE,
num_workers=0, # couldn't use workers https://github.com/fastai/fastbook/issues/85
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=BATCH_SIZE,
num_workers=0,
shuffle=False)
val_loader = DataLoader(dataset=val_dataset,
batch_size=BATCH_SIZE,
num_workers=0,
shuffle=False)
# Checking the dataset
for data, labels in train_loader:
print('Matrix batch dimensions:', data.shape)
print('Matrix label dimensions:', labels.shape)
break
# Checking the dataset
for data, labels in test_loader:
print('Matrix batch dimensions:', data.shape)
print('Matrix label dimensions:', labels.shape)
break
# Checking the dataset
for data, labels in val_loader:
print('Matrix batch dimensions:', data.shape)
print('Matrix label dimensions:', labels.shape)
break
# + [markdown] id="hbvFNaRk-yU2"
# ## Setup Model
# + colab={"base_uri": "https://localhost:8080/"} id="KFxoCAHg-yU2" outputId="6e18dbfd-0af3-4534-f60d-9a97f2867f1d"
# Check that the train loader works correctly
device = torch.device(DEVICE)
torch.manual_seed(0)
for epoch in range(2):
for batch_idx, (x, y) in enumerate(train_loader):
print('Epoch:', epoch+1, end='')
print(' | Batch index:', batch_idx, end='')
print(' | Batch size:', y.size()[0])
x = x.to(device)
y = y.to(device)
break
# + id="doCsnPrG-yU2"
##########################
### MODEL
##########################
class logCNN(nn.Module):
def __init__(self, num_classes):
super(logCNN, self).__init__()
self.num_classes = num_classes
self.features = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2),
nn.Conv2d(16, 32, kernel_size=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2)
)
self.classifier = nn.Sequential(
nn.Linear(1056, 120),
nn.ReLU(),
nn.Linear(120, 84),
nn.ReLU(),
nn.Linear(84, num_classes),
)
def forward(self, x):
x = self.features(x)
x = torch.flatten(x, 1)
logits = self.classifier(x)
probas = F.softmax(logits, dim=1)
return logits, probas
# + id="EZLIAStI-yU3"
torch.manual_seed(RANDOM_SEED)
model = logCNN(NUM_CLASSES)
model.to(DEVICE)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
# + [markdown] id="Dr-b4WVw-yU3"
# # Train Model
# + tags=["outputPrepend"] colab={"base_uri": "https://localhost:8080/"} id="SeNobFr9-yU4" outputId="ba791175-ee54-4b45-a9c3-ed655e7ce432"
def compute_accuracy(model, data_loader, device):
correct_pred, num_examples = 0, 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(DEVICE)
targets = targets.to(DEVICE, dtype=torch.long) # had to pass in to torch.long based on the 1, 0 int labels
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
# This needs to be vectorized
def compute_f1(model, data_loader, device):
y_hats = []
y_acts = []
for i, (inputs, targets) in enumerate(data_loader):
yhat = model(inputs)[-1].cpu().detach().numpy().round()
yhat = np.argmax(yhat, axis=1)
y_hats.append(yhat)
y_acts.append(list(targets.cpu().detach().numpy()))
y_hats = [item for sublist in y_hats for item in sublist]
y_acts = [item for sublist in y_acts for item in sublist]
return f1_score(y_acts, y_hats)
start_time = time.time()
minibatch_cost = []
epoch_train_performance = []
epoch_val_performance = []
for epoch in range(NUM_EPOCHS):
model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(DEVICE)
targets = targets.to(DEVICE, dtype=torch.long) # another had to use torch.long
### FORWARD AND BACK PROP
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
minibatch_cost.append(cost)
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %04d/%04d | Cost: %.4f'
%(epoch+1, NUM_EPOCHS, batch_idx,
len(train_loader), cost, ))
model.eval()
with torch.set_grad_enabled(False): # save memory during inference
train_performance = compute_f1(model, train_loader, device=DEVICE)
val_performance = compute_f1(model, val_loader, device=DEVICE)
epoch_train_performance.append(train_performance)
epoch_val_performance.append(val_performance)
print('Epoch: %03d/%03d | Train: %.3f%% | Val: %.3f%%' % (
epoch+1, NUM_EPOCHS, train_performance, val_performance))
print('Time elapsed: %.2f min' % ((time.time() - start_time)/60))
print('Total Training Time: %.2f min' % ((time.time() - start_time)/60))
# + colab={"base_uri": "https://localhost:8080/", "height": 541} id="EGRqtOSy-yU5" outputId="00476482-36e0-4f96-af9a-3b21a0751188"
# plot cost functions
minibatch_cost_cpu = [i.cpu().detach().numpy() for i in minibatch_cost]
plt.plot(range(len(minibatch_cost_cpu)), minibatch_cost_cpu)
plt.ylabel('Cost Function Label')
plt.xlabel('Minibatch')
plt.show()
plt.plot(range(len(epoch_train_performance)), epoch_train_performance, label="train f1 scores")
plt.plot(range(len(epoch_val_performance)), epoch_val_performance, label="val f1 scores")
plt.ylabel('F1 Score')
plt.xlabel('Epoch')
plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left', borderaxespad=0.)
plt.show()
# + [markdown] id="obJPMecX-yU6"
# ## Evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="03nz_tXO-yU7" outputId="2cc3ffb8-79ab-4230-cfb7-a95e5f31f592"
# Evaluate metrics on the test set
y_hats = []
y_acts = []
counter = 0
for i, (inputs, targets) in enumerate(test_loader):
yhat = model(inputs)[-1].cpu().detach().numpy().round()
yhat = np.argmax(yhat, axis=1)
y_hats.append(yhat)
y_acts.append(list(targets.cpu().detach().numpy()))
counter += 1
y_hats = [item for sublist in y_hats for item in sublist]
y_acts = [item for sublist in y_acts for item in sublist]
print("TEST SET METRICS:")
f1 = f1_score(y_acts, y_hats)
print("f1 score : ", f1)
precision = precision_score(y_acts, y_hats)
print("precision", precision)
recall = recall_score(y_acts, y_hats)
print("recall", recall)
# + colab={"base_uri": "https://localhost:8080/"} id="h3_a-OxwsJwp" outputId="c38f2f47-abc1-4d24-b29a-7eaf5580db72"
test_ys = pd.DataFrame(list(zip(y_acts, y_hats)), columns=["y_true", "y_pred"])
print("TEST SET:\n")
print("anomalous:\n")
test_anomalous = test_ys[test_ys["y_true"]==1]
print("number of anomalies in the test set:", len(test_anomalous))
correct_anomalous = test_anomalous[test_anomalous["y_true"] == test_anomalous["y_pred"]]
print("number of anomalies correctly identified", len(correct_anomalous))
incorrect_anomalous = test_anomalous[test_anomalous["y_true"] != test_anomalous["y_pred"]]
print("number of anomalies incorrectly identified", len(incorrect_anomalous))
print("\nnormal:\n")
test_normals = test_ys[test_ys["y_true"]==0]
print("number of normals in the test set:", len(test_normals))
correct_normal = test_normals[test_normals["y_true"] == test_normals["y_pred"]]
print("number of normals correctly identified", len(correct_normal))
incorrect_normal = test_normals[test_normals["y_true"] != test_normals["y_pred"]]
print("number of normals incorrectly identified", len(incorrect_normal))
# + colab={"base_uri": "https://localhost:8080/"} id="IW4Z3-S5pPsR" outputId="a8e3228b-53f0-455b-9add-038356a0e3b0"
# Evaluate metrics on train set
y_hats = []
y_acts = []
counter = 0
for i, (inputs, targets) in enumerate(train_loader):
yhat = model(inputs)[-1].cpu().detach().numpy().round()
yhat = np.argmax(yhat, axis=1)
y_hats.append(yhat)
y_acts.append(list(targets.cpu().detach().numpy()))
counter += 1
y_hats = [item for sublist in y_hats for item in sublist]
y_acts = [item for sublist in y_acts for item in sublist]
print("TRAIN SET METRICS:")
f1 = f1_score(y_acts, y_hats)
print("f1 score : ", f1)
precision = precision_score(y_acts, y_hats)
print("precision", precision)
recall = recall_score(y_acts, y_hats)
print("recall", recall)
# + colab={"base_uri": "https://localhost:8080/"} id="aqZLZHRRpv-3" outputId="f733b39d-f4ec-4544-887d-1ea8a093885e"
train_ys = pd.DataFrame(list(zip(y_acts, y_hats)), columns=["y_true", "y_pred"])
print("TRAIN SET:\n")
print("anomalous:\n")
train_anomalous = train_ys[train_ys["y_true"]==1]
print("number of anomalies in the train set:", len(train_anomalous))
correct_anomalous = train_anomalous[train_anomalous["y_true"] == train_anomalous["y_pred"]]
print("number of anomalies correctly identified", len(correct_anomalous))
incorrect_anomalous = train_anomalous[train_anomalous["y_true"] != train_anomalous["y_pred"]]
print("number of anomalies incorrectly identified", len(incorrect_anomalous))
print("\nnormal:\n")
train_normals = train_ys[train_ys["y_true"]==0]
print("number of normals in the train set:", len(train_normals))
correct_normal = train_normals[train_normals["y_true"] == train_normals["y_pred"]]
print("number of normals correctly identified", len(correct_normal))
incorrect_normal = train_normals[train_normals["y_true"] != train_normals["y_pred"]]
print("number of normals incorrectly identified", len(incorrect_normal))
# + id="SN9sqHAiu3LG"
| model/anomaly_nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Explanatory Analysis ##
#
# Contuning with the question - i.e. How can our model helps support and augment health workers where hospital bed demand exists - Multivariate analysis did show a positive relationship between increase in bed availability across the poverty level.
#
# To explain this findings, I will use the Cluster analysis technique.
#
# - Cluster analysis, starting with top 5 states in poverty to help the model analyze the type of hospitals that have available bed capacity and their utlization.
# + slideshow={"slide_type": "subslide"}
# Import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as mtick
from matplotlib.ticker import PercentFormatter
import seaborn as sb
import os
import wget
# %matplotlib inline
# + slideshow={"slide_type": "subslide"}
# Load the dataset
pot_beds_df = pd.read_csv('poverty-beds-df.csv')
# + slideshow={"slide_type": "slide"}
# Lets get the top 5 states in poverty.
pot_beds_df.sort_values('PCT_POVALL_2018').drop_duplicates('STATE_NAME').nlargest(5, 'PCT_POVALL_2018')[['STATE_NAME', 'HQ_STATE']]
# + slideshow={"slide_type": "skip"}
# Build a dataset for the top 5 state in poverty.
top_povery_states = pot_beds_df.sort_values('PCT_POVALL_2018').drop_duplicates('HQ_STATE').nlargest(5, 'PCT_POVALL_2018')[['HQ_STATE']].HQ_STATE
top_povery_stbeds_df = pot_beds_df[pot_beds_df['HQ_STATE'].isin(top_povery_states)]
top_povery_stbeds_df.reset_index(drop=True, inplace=True)
top_povery_stbeds_df.sample(n=5)
# + slideshow={"slide_type": "slide"}
#Get a breakdown on available bed availability by hospital type given the high poverty level states.
plt.figure(figsize=[16,10])
g = sb.swarmplot(x="HQ_STATE", y="PCT_POTNL_INC_BED_CAPC", hue="HOSPITAL_TYPE",data=top_povery_stbeds_df,
palette="Set2", dodge=True, size=10)
plt.legend(loc = 6, bbox_to_anchor = (1.0, 0.5), fontsize=18)
plt.xlabel('State')
plt.ylabel('Potential Increase In Bed Availability')
g.set_title('States with Increase in Bed Availability in context of Hospital Types', fontdict={'fontsize':18}, pad=16)
plt.gca().yaxis.set_major_formatter(mtick.PercentFormatter())
plt.show()
# + slideshow={"slide_type": "slide"}
# What percentage increase in bed availability do we see across states and hospital types?
fig,ax = plt.subplots(figsize = [16,10])
cat_counts = top_povery_stbeds_df.groupby(['HOSPITAL_TYPE', 'HQ_STATE']).count()['PCT_POTNL_INC_BED_CAPC']
cat_counts = cat_counts.reset_index(name = 'num_var2_avg')
cat_counts = cat_counts.pivot(index = 'HQ_STATE', columns = 'HOSPITAL_TYPE',
values = 'num_var2_avg')
cmap = sb.cubehelix_palette(8)
g= sb.heatmap(cat_counts, annot = True,
vmin=0, vmax=100, linewidths=1, linecolor='grey', square=True, cmap=cmap, annot_kws={"fontsize":16},
cbar_kws = {'label' : '%(Potential Increase in Bed Availability)'})
g.set_facecolor('lightblue')
g.set_title('Precentage Increase in Bed Availability in context of Hospital Types', fontdict={'fontsize':18}, pad=16)
for t in ax.texts: t.set_text(t.get_text() + "%")
plt.xlabel('Hospital Type')
plt.ylabel('State')
plt.show()
# + slideshow={"slide_type": "slide"}
# What percentage of bed utilization do we see for the given availability of beds for people across the poverty level?
fig,ax = plt.subplots(figsize = [16,10])
plt.scatter(data=top_povery_stbeds_df, x='PCT_POVALL_2018', y='PCT_POTNL_INC_BED_CAPC',
c='PCT_BED_UTILIZATION',
cmap = 'viridis',
)
# plt.xlim(2,22)
# plt.ylim(0,100)
cbar = plt.colorbar()
cbar.ax.set_ylabel('Bed Utilization (%)')
plt.xlabel('Poverty Level')
plt.ylabel('Potential Increase In Bed Availability')
plt.gca().xaxis.set_major_formatter(mtick.PercentFormatter())
plt.gca().yaxis.set_major_formatter(mtick.PercentFormatter())
plt.title('Poverty Level to Bed Availabilty in the context of Bed Utilization')
plt.show()
# + slideshow={"slide_type": "subslide"}
# What is the median percentage of Bed utilization by Hospital type?
top_povery_stbeds_df.groupby(['HOSPITAL_TYPE'])['PCT_BED_UTILIZATION'].agg(['median']).sort_values(['median']).rename(
columns={'median' : 'PCT_BED_UTILIZATION'})
# + slideshow={"slide_type": "subslide"}
top_povery_stbeds_df.groupby(['HQ_STATE','HOSPITAL_TYPE'])['PCT_BED_UTILIZATION'].agg(['median']).sort_values(['HOSPITAL_TYPE']).reset_index().rename(
columns={'median' : 'PCT_BED_UTILIZATION'})
# + slideshow={"slide_type": "slide"}
# Categorical view of bed utilization by hospital type and state. This provides a view into bed utilization by categorical columns.
cat_sorted_df = top_povery_stbeds_df.groupby(['HOSPITAL_TYPE','HQ_STATE'])['PCT_BED_UTILIZATION'].agg(['median']).sort_values(['median']).reset_index().rename(
columns={'median' : 'PCT_BED_UTILIZATION'})
g = sb.catplot(x = "HQ_STATE", y = "PCT_BED_UTILIZATION", col="HOSPITAL_TYPE", col_wrap=3, kind="bar", data=cat_sorted_df, ci=None, height=8, color="c", linewidth=2)
g.set_axis_labels("STATE", "Hospital Bed Utilization %")
plt.subplots_adjust(hspace = 0.1)
plt.gca().yaxis.set_major_formatter(mtick.PercentFormatter())
plt.rcParams["axes.labelsize"] = 15
g.fig.subplots_adjust(top=0.9)
g.fig.suptitle('Bed Utilization by Hospital type in the context of top States in poverty', fontsize=16)
plt.show()
# + slideshow={"slide_type": "slide"}
#Pointplot to with join lines to show the change in utilization.
fig,ax = plt.subplots(figsize = [16,10])
ax = sb.pointplot(data = cat_sorted_df, x = 'HQ_STATE', y = 'PCT_BED_UTILIZATION', hue = 'HOSPITAL_TYPE',
palette = 'Blues', capsize=.2, dodge = 0.1, join=True, size=10, linewidth=2)
plt.title('Median Bed utilization across top 5 states in poverty')
plt.xlabel('State')
plt.ylabel('Median Bed utilization')
plt.gca().yaxis.set_major_formatter(mtick.PercentFormatter())
ax.set_yticklabels([],minor = True)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multivariate Analysis Findings ##
#
# ### Analysis on Hospital Bed Availability with reference to top 5 States in Poverty Level and hospital type ###
#
# - Short term acute care hospital are shown to have high bed availability across all 5 states. Louisiana and Arizona states have ** 49% ** and ** 31% ** of bed availability respectively.
# - ** District of Columbia ** has shown to be trailing in bed availability across all hospital types. A callout on this state being the top poverty state.
# - Overall ** Louisiana state ** is shown to have high bed availability across all hospital types.
# - ** Arizona state ** being the second highest poverty state stands second in high bed availability across all hospital types.
#
# ### Analysis on Bed Utilization with reference to top 5 States in Poverty Level and Hospital type ###
#
# - Psychiatric hospital type are the top utilized hospitals at ** 85% **. This utilization is distributed between Delaware, Arizona and Louisiana states.
# - Short Term Acute Care hospital type (shown high bed availability) utilization is at ** 49% ** across all 5 states. ** District of Columbia ** topping the utilization.
# - Childrens Hospital, Long Term Acute Care Hospital and Rehabilitation Hospital respectively have high bed utilization percentages.
# - New Mexico and Arizona states have only shown the Median Bed utilization for Critical Access Hospital at ** 33%**.
# + [markdown] slideshow={"slide_type": "slide"}
# # Conclusion: Findings and Next Steps #
#
# To better understand how socioeconomic status - such as populations living in federal poverty area impact hospital bed utilization and to help optimize scenario planning for when staff can be shifted around to serve those living in federal poverty areas.
#
# Historical Bed Utilization Rate, Poverty Level, Potential available in Bed capacity are key features that can help forecast the staffing needs of health workers (doctors, nurses etc). The staffing needs to a specific hospital type can be scored based on the availability enabling counties in poverty to be better served.
#
# As next steps, building a machine learning model that helps support and augment health workers where demand exists by integrating with existing staff scheduling system allowing managers to view the model recommendations, allowing healthcare stakeholders deliver better patient care and improving productivity.
| explanatory-povertylevel-bedutilization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.chdir(os.path.dirname("../"))
# +
import warnings
warnings.filterwarnings("ignore")
# -
# # deepOF data exploration
# Given a dataset, this notebook allows the user to
#
# * Load and process the dataset using deepof.data
# * Visualize data quality with interactive plots
# * Visualize training instances as multi-timepoint scatter plots with interactive configurations
# * Visualize training instances as video clips with interactive configurations
# +
import deepof.data
import deepof.utils
import numpy as np
import pandas as pd
import ruptures as rpt
import tensorflow as tf
from ipywidgets import interact, widgets
from IPython import display
from matplotlib.animation import FuncAnimation
from scipy.signal import savgol_filter
import matplotlib.pyplot as plt
import seaborn as sns
# -
# ### 1. Define and run project
exclude_bodyparts = tuple([""])
# Use deepof to load a project
proj = deepof.data.project(
path="../../Desktop/deepoftesttemp/",
arena_dims=[380],
arena_detection="rule-based",
exclude_bodyparts=exclude_bodyparts,
interpolate_outliers=True,
smooth_alpha=9,
).run()
# ### 2. Inspect dataset quality
all_quality = pd.concat([tab for tab in proj.get_quality().values()])
all_quality.boxplot(rot=45)
plt.ylim(0.99985, 1.00001)
plt.show()
@interact(quality_top=(0.0, 1.0, 0.01))
def low_quality_tags(quality_top):
pd.DataFrame(
pd.melt(all_quality)
.groupby("bodyparts")
.value.apply(lambda y: sum(y < quality_top) / len(y) * 100)
).sort_values(by="value", ascending=False).plot.bar(rot=45)
plt.xlabel("body part")
plt.ylabel("Tags with quality under {} (%)".format(quality_top * 100))
plt.tight_layout()
plt.legend([])
plt.show()
# In the cell above, you see the percentage of labels per body part which have a quality lower than the selected value (0.50 by default) **before** preprocessing. The values are taken directly from DeepLabCut.
# ### 3. Get coordinates, distances and angles
# And get speed, acceleration and jerk for each
# Get coordinates, speeds, accelerations and jerks for positions
position_coords = proj.get_coords(center="Center", align="Spine_1", align_inplace=True)
position_speeds = proj.get_coords(center="Center", speed=1)
position_accels = proj.get_coords(center="Center", speed=2)
position_jerks = proj.get_coords(center="Center", speed=3)
# Get coordinates, speeds, accelerations and jerks for distances
distance_coords = proj.get_distances()
distance_speeds = proj.get_distances(speed=1)
distance_accels = proj.get_distances(speed=2)
distance_jerks = proj.get_distances(speed=3)
# Get coordinates, speeds, accelerations and jerks for angles
angle_coords = proj.get_angles()
angle_speeds = proj.get_angles(speed=1)
angle_accels = proj.get_angles(speed=2)
angle_jerks = proj.get_angles(speed=3)
# ### 4. Analyse smoothing
@interact(test=position_coords.keys(),
bpart = set([i[0] for i in list(position_coords.values())[0].columns]),
max_obs=(100, 1000),
smooth_degree=widgets.IntSlider(min=2, max=11, value=9),
smooth_wlen=widgets.IntSlider(min=11, max=101, step=2, value=11))
def plot_smoothing(test, bpart, max_obs, smooth_degree, smooth_wlen):
fig = plt.figure(figsize=(12, 8))
no_smooth = deepof.data.project(
path="../../Desktop/deepoftesttemp/",
arena_dims=[380],
arena_detection="rule-based",
exclude_bodyparts=exclude_bodyparts,
interpolate_outliers=True,
smooth_alpha=None,
).run(verbose=0).get_coords(center="Center", align="Spine_1", align_inplace=True)
no_smooth_coords = no_smooth[test].iloc[:max_obs, :].loc[:, bpart]
smooth_coords = savgol_filter(no_smooth_coords, smooth_wlen, smooth_degree, axis=0)
plt.plot(no_smooth_coords, linestyle="--", label="no smoothing")
plt.plot(smooth_coords, color="red", label="savgol, poly={}, wlen={}".format(smooth_degree, smooth_wlen))
plt.title("SavGol smoothing exploration")
plt.xlabel("Time")
plt.ylabel("Coordinate value")
plt.legend()
plt.tight_layout()
plt.show()
# ### 5. Display training instances
# +
random_exp = np.random.choice(list(position_coords.keys()), 1)[0]
@interact(time_slider=(0.0, 15000, 25), length_slider=(10, 100, 5))
def plot_mice_across_time(time_slider, length_slider):
plt.figure(figsize=(10, 10))
for bpart in position_coords[random_exp].columns.levels[0]:
if bpart != "Center":
sns.scatterplot(
data=position_coords[random_exp].loc[
time_slider : time_slider + length_slider - 1, bpart
],
x="x",
y="y",
label=bpart,
palette=sns.color_palette("tab10"),
)
plt.title("Positions across time for centered data")
plt.legend(
fontsize=15,
bbox_to_anchor=(1.5, 1),
title="Body part",
title_fontsize=18,
shadow=False,
facecolor="white",
)
plt.ylim(-100, 60)
plt.xlim(-60, 60)
plt.show()
# -
# The figure above is a multi time-point scatter plot. The time_slider allows you to scroll across the video, and the length_slider selects the number of time-points to include. The idea is to intuitively visualize the data that goes into a training instance for a given preprocessing setting.
# +
# Auxiliary animation functions
def plot_mouse_graph(instant_x, instant_y, ax, edges):
"""Generates a graph plot of the mouse"""
plots = []
for edge in edges:
(temp_plot,) = ax.plot(
[float(instant_x[edge[0]]), float(instant_x[edge[1]])],
[float(instant_y[edge[0]]), float(instant_y[edge[1]])],
color="#006699",
linewidth=2.0,
)
plots.append(temp_plot)
return plots
def update_mouse_graph(x, y, plots, edges):
"""Updates the graph plot to enable animation"""
for plot, edge in zip(plots, edges):
plot.set_data(
[float(x[edge[0]]), float(x[edge[1]])],
[float(y[edge[0]]), float(y[edge[1]])],
)
# +
random_exp = np.random.choice(list(position_coords.keys()), 1)[0]
print(random_exp)
@interact(time_slider=(0.0, 15000, 25), length_slider=(10, 100, 5))
def animate_mice_across_time(time_slider, length_slider):
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
edges = deepof.utils.connect_mouse_topview()
for bpart in exclude_bodyparts:
if bpart:
edges.remove_node(bpart)
for limb in ["Left_fhip", "Right_fhip", "Left_bhip", "Right_bhip"]:
edges.remove_edge("Center", limb)
if ("Tail_base", limb) in list(edges.edges()):
edges.remove_edge("Tail_base", limb)
edges = edges.edges()
data = position_coords[random_exp].loc[
time_slider : time_slider + length_slider - 1, :
]
data["Center", "x"] = 0
data["Center", "y"] = 0
init_x = data.xs("x", level=1, axis=1, drop_level=False).iloc[0, :]
init_y = data.xs("y", level=1, axis=1, drop_level=False).iloc[0, :]
plots = plot_mouse_graph(init_x, init_y, ax, edges)
scatter = ax.scatter(x=np.array(init_x), y=np.array(init_y), color="#006699",)
# Update data in main plot
def animation_frame(i):
# Update scatter plot
x = data.xs("x", level=1, axis=1, drop_level=False).iloc[i, :]
y = data.xs("y", level=1, axis=1, drop_level=False).iloc[i, :]
scatter.set_offsets(np.c_[np.array(x), np.array(y)])
update_mouse_graph(x, y, plots, edges)
return scatter
animation = FuncAnimation(
fig, func=animation_frame, frames=length_slider, interval=75,
)
ax.set_title("Positions across time for centered data")
ax.set_ylim(-90, 60)
ax.set_xlim(-60, 60)
ax.set_xlabel("x")
ax.set_ylabel("y")
video = animation.to_html5_video()
html = display.HTML(video)
display.display(html)
plt.close()
# -
# The figure above displays exactly the same data as the multi time-point scatter plot, but in the form of a video (one training instance at the time).
# ### 6. Detect changepoints
test_rupt = np.array(position_coords["Test 1_s11"])
test_rupt.shape
algo = rpt.Pelt(model="rbf", min_size=10, jump=1).fit(test_rupt)
result = algo.predict(pen=3)
len(result)
cut = 1000
cut = result[np.argmin(np.abs(np.array(result) - cut))]
print("cut:", cut)
rpt.display(test_rupt[:cut], [i for i in result if i <= cut])
plt.show()
| supplementary_notebooks/deepof_data_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
#skip
! [ -e /content ] && pip install -Uqq fastpapers
# +
# default_exp core
# -
# # Core
#
# > Basic functions and classes usful to reproduce research papers.
#hide
from nbdev.showdoc import *
#export
import gc
import requests
from fastcore.all import *
from fastai.basics import *
from fastai.data.all import *
from fastai.vision.all import *
from fastai.vision.gan import *
from fastai.callback.hook import *
from fastai.test_utils import *
import pandas as pd
import seaborn as sns
# ## Debugging
#
# These are function that are useful for debugging.
#exports
def explode_types(o):
'''Like fastcore explode_types, but only shows __name__ of type.'''
if not is_listy(o): return type(o).__name__
return {type(o).__name__: [explode_types(o_) for o_ in o]}
#exports
def explode_lens(o):
if is_listy(o):
if all(is_listy(o_) for o_ in o):
return [explode_lens(o_) for o_ in o]
else: return len(o)
test_eq(explode_lens([[1,4], [[5,6,7], [1]]]), [2, [3, 1]])
#exports
def explode_shapes(o):
if not is_listy(o): return tuple(bind(getattr, arg0, 'shape')(o))
return [explode_shapes(o_) for o_ in o]
test_eq(explode_shapes([tensor([1,4]), [tensor([[4,5],[7,8]]), tensor([6])]]), [(2,), [(2,2), (1,)]])
#exports
def explode_ranges(o):
if not is_listy(o): return (float(o.min()), float(o.max()))
return [explode_ranges(o_) for o_ in o]
explode_ranges([tensor([1,4]), [tensor([[4,5],[7,8]]), tensor([6])]])
#exports
def pexpt(o): print(explode_types(o))
#exports
def pexpl(o): print(explode_lens(o))
#exports
def pexps(o): print(explode_shapes(o))
#export
def get_cudas():
'''Returns the number of tensors in cuda device.'''
n = 0
for o in gc.get_objects():
o = maybe_attr(o, 'data')
if torch.is_tensor(o):
if o.is_cuda: n += 1
return n
#export
def receptive_fields(model, nf, imsize, bs=64):
'''returns the size of the receptive field for each feature output.'''
# init parameters
for p in model.named_parameters():
if 'weight' in p[0]:
n = p[1].shape[1] if len(p[1].shape)==4 else 1
nn.init.constant_(p[1], 1./n)
elif 'bias' in p[0]:
nn.init.constant_(p[1], 0)
x = dummy_eval(model, imsize).detach()
outsz = x.shape[-2:]
with torch.no_grad():
rfs = []
model.eval()
model = model.cuda()
t = torch.eye(imsize[0]**2).reshape(imsize[0]*imsize[1], 1, imsize[0], imsize[1])
for i,batch in enumerate(chunked(t, bs)):
new = torch.cat(batch, dim=0).unsqueeze(1)
new = torch.cat([new for _ in range(nf)], dim=1).cuda()
rfs.append(model(new).cpu())
rfs = torch.cat(rfs, dim=0).squeeze()
rfs = rfs.reshape(imsize[0], imsize[1], outsz[0], outsz[1])
rfs = (rfs>0.99).sum(axis=(0,1)).float().sqrt()
return rfs
# ## N Images Classes
#
# These are classes to handle many images to many images generation
#export
class ImageNTuple(fastuple):
@classmethod
def create(cls, fns): return cls(tuple(PILImage.create(f) for f in tuplify(fns)))
def show(self, ctx=None, **kwargs):
all_tensors = all([isinstance(t, Tensor) for t in self])
same_shape = all([self[0].shape==t.shape for t in self[1:]])
if not all_tensors or not same_shape: return ctx
line = self[0].new_zeros(self[0].shape[0], self[0].shape[1], 10)
imgs = sum(L(zip(self, [line]*len(self))).map(list),[])[:-1]
return show_image(torch.cat(imgs, dim=2), ctx=ctx, **kwargs)
def requires_grad_(self, value):
for item in self: item.requires_grad_(value)
return self
@property
def shape(self):
all_tensors = all([isinstance(t, Tensor) for t in self])
same_shape = all([self[0].shape==t.shape for t in self[1:]])
if not all_tensors or not same_shape: raise AttributeError
return self[0].shape
#def detach(self):
# for item in self: item.detach()
# return self
path = untar_data(URLs.PETS)
files = get_image_files(path/"images")
# ImageNTuple will only show tha images if all of them are of same size, and if all of them are tensors.
imt2 = ImageNTuple.create((files[0], files[1]))
explode_types(imt2)
imt2 = ImageNTuple.create((files[0], files[1]))
imt2 = Resize(224)(imt2)
imt2 = ToTensor()(imt2)
imt2.show();
imt3 = ImageNTuple.create((files[0], files[1], files[2]))
imt3 = Resize(224)(imt3)
imt3 = ToTensor()(imt3)
ax = imt3.show()
test_eq(len(imt2), 2)
test_eq(len(imt3), 3)
# export
def ImageTupleBlock():
'''Like fastai tutoria siemese transform, but uses ImageNTuple.'''
return TransformBlock(type_tfms=ImageNTuple.create, batch_tfms=[IntToFloatTensor])
# ## GAN Models
# ### Conditional Generator -
#export
class ConditionalGenerator(nn.Module):
'''Wraper around a GAN generator that returns the generated image and the inp.'''
def __init__(self, gen):
super().__init__()
self.gen = gen
def forward(self, x):
if is_listy(x):
inp = torch.cat(x, axis=1)
else:
inp = x
return ImageNTuple(x, TensorImage(self.gen(inp)))
# Test that the ConditionalGenerator can generate images from one image
gen_base = basic_critic(32, 3)
gen_base = nn.Sequential(*list(gen_base.children())[:-2])
unet = DynamicUnet(gen_base, 3, (32, 32))
gen = ConditionalGenerator(unet)
out = gen(torch.rand(1, 3, 32, 32))
test_eq(explode_shapes(out), [(1, 3, 32, 32), (1, 3, 32, 32)])
test_eq(type(out), ImageNTuple)
# Test that the ConditionalGenerator can generate an image from two images
gen_base = basic_critic(32, 6)
gen_base = nn.Sequential(*list(gen_base.children())[:-2])
unet = DynamicUnet(gen_base, 3, (32, 32))
gen = ConditionalGenerator(unet)
dl = DataLoader(dataset=([imt2]), bs=1, after_item=IntToFloatTensor())
b = first(dl)
out = gen(b)
test_eq(type(out), ImageNTuple)
test_eq(len(out), 3)
# ### Siamese Critic -
#export
class SiameseCritic(Module):
def __init__(self, critic): self.critic = critic
def forward(self, x): return self.critic(torch.cat(x, dim=1))
critic = gan_critic(n_channels=6, nf=64)
scritic = SiameseCritic(critic)
# ## GAN Metrics
#
# These are Metrics that work with GANLearner
#export
class GenMetric(AvgMetric):
def accumulate(self, learn):
if learn.model.gen_mode:
bs = find_bs(learn.yb)
self.total += to_detach(self.func(learn, learn.pred, *learn.yb))*bs
self.count += bs
#export
class CriticMetric(AvgMetric):
def accumulate(self, learn):
if not learn.model.gen_mode:
bs = find_bs(learn.yb)
self.total += to_detach(self.func(learn, learn.pred, *learn.yb))*bs
self.count += bs
#export
def _l1(learn, output, target): return nn.L1Loss()(output[-1], target[-1])
l1 = GenMetric(_l1)
# ## Export GANLearner
# To export a `GANLearner` we need to set the learner into `gen_mode` and before recreating the optimization function we should call `set_freeze_model` to unfreeze all the parameters in the model so the old opt state can be loaded properly.
#export
@patch
def export(self:GANLearner, fname='export.pkl', pickle_protocol=2):
"Export the content of `self` without the items and the optimizer state for inference"
if rank_distrib(): return # don't export if child proc
self.gan_trainer.switch(gen_mode=True)
self._end_cleanup()
old_dbunch = self.dls
self.dls = self.dls.new_empty()
state = self.opt.state_dict() if self.opt is not None else None
self.opt = None
with warnings.catch_warnings():
#To avoid the warning that come from PyTorch about model not being checked
warnings.simplefilter("ignore")
torch.save(self, self.path/fname, pickle_protocol=pickle_protocol)
set_freeze_model(self.model, True)
self.create_opt()
if state is not None: self.opt.load_state_dict(state)
self.dls = old_dbunch
self.gan_trainer.switch(gen_mode=True)
# ## ProgressImage Callback -
#export
class ProgressImage(Callback):
'''Shows a sample of the generator after every epoch. It is a good idea to keep a human in the loop.'''
run_after = GANTrainer
@delegates(show_image)
def __init__(self, out_widget, save_img=False, folder='pred_imgs', conditional=False, **kwargs):
self.out_widget = out_widget
self.kwargs = kwargs
self.save_img = save_img
self.folder = folder
self.conditional = conditional
if self.conditional:
self.title = 'Input-Real-Fake'
else:
self.title = 'Generated'
Path(self.folder).mkdir(exist_ok=True)
self.ax = None
def before_batch(self):
if self.gan_trainer.gen_mode and self.training:
self.last_gen_target = self.learn.to_detach(self.learn.yb)#[0][-1]
def after_train(self):
"Show a sample image."
if not hasattr(self.learn.gan_trainer, 'last_gen'): return
b = self.learn.gan_trainer.last_gen
gt = self.last_gen_target
b = self.learn.dls.decode((b,))
gt = self.learn.dls.decode(gt)
gt, imt = batch_to_samples((gt, b), max_n=1)[0]
gt, imt = gt[0][-1], imt[0]
if self.conditional:
imt = ToTensor()(ImageNTuple.create((*imt[:-1], gt, imt[-1])))
self.out_widget.clear_output(wait=True)
with self.out_widget:
if self.ax: self.ax.clear()
self.ax = imt.show(ax=self.ax, title=self.title, **self.kwargs)
display(self.ax.figure)
if self.save_img: self.ax.figure.savefig(self.path / f'{self.folder}/pred_epoch_{self.epoch}.png')
def after_fit(self):
if self.ax: plt.close(self.ax.figure)
# ## Show Results for TupleNImage
#export
@typedispatch
def show_results(x:TensorImage, y:ImageNTuple, samples, outs, ctxs=None, max_n=6, nrows=None, ncols=2, figsize=None, **kwargs):
max_n = min(x.shape[0], max_n)
if max_n<ncols: ncols = max_n
if figsize is None: figsize = (ncols*6, max_n//ncols * 3)
if ctxs is None: ctxs = get_grid(min(x[0].shape[0], max_n), nrows=None, ncols=ncols, figsize=figsize)
for i,ctx in enumerate(ctxs):
title = 'Input-Real-Fake'
ImageNTuple(x[i], y[1][i], outs[i][0][1]).show(ctx=ctx, title=title)
# ## Tuple in Ys Learner Fix -
#
# Learner.show_results and Learner.predict though an error if the ys are tuples. Let's fix that
#export
@patch
def show_results(self:GANLearner, ds_idx=1, dl=None, max_n=9, shuffle=True, **kwargs):
if dl is None: dl = self.dls[ds_idx].new(shuffle=shuffle)
b = dl.one_batch()
_,_,preds = self.get_preds(dl=[b], with_decoded=True)
preds = (preds,)
self.dls.show_results(b, preds, max_n=max_n, **kwargs)
# +
# #exort
# @patch
# def predict(self:GANLearner, item, rm_type_tfms=None, with_input=False):
# dl = self.dls.test_dl([item], rm_type_tfms=rm_type_tfms, num_workers=0)
# inp,preds,_,dec_preds = self.get_preds(dl=dl, with_input=True, with_decoded=True)
# i = getattr(self.dls, 'n_inp', -1)
# inp = (inp,) if i==1 else tuplify(inp)
# n_out = len(self.dls.tls) - i
# dec_preds = (dec_preds,) if n_out==1 else tuplify(dec_preds)
# dec = self.dls.decode_batch(inp + dec_preds)[0]
# dec_inp,dec_targ = map(detuplify, [dec[:i],dec[i:]])
# res = dec_targ,dec_preds[0],preds[0]
# if with_input: res = (dec_inp,) + res
# return res
# +
# @patch
# def tuplify(o:ImageNTuple, *kwargs): return (o,)
# -
# ## Datasets
#export
URLs.FACADES = 'http://efrosgans.eecs.berkeley.edu/pix2pix/datasets/facades.tar.gz'
URLs.FACADES_BASE = 'http://cmp.felk.cvut.cz/~tylecr1/facade/CMP_facade_DB_base.zip'
URLs.FACADES_EXTENDED = 'http://cmp.felk.cvut.cz/~tylecr1/facade/CMP_facade_DB_extended.zip'
URLs.CELEBA = '0B7EVK8r0v71pZjFTYXZWM3FlRnM'
URLs.COCO_VAL2017 = 'http://images.cocodataset.org/zips/val2017.zip'
URLs.COCO_TEST2017 = 'http://images.cocodataset.org/zips/test2017.zip'
URLs.COCO_TRAIN2017 = 'http://images.cocodataset.org/zips/train2017.zip'
URLs.COCO_TRAINVAL_ANN = 'http://images.cocodataset.org/annotations/annotations_trainval2017.zip'
URLs.COCO_TEST_ANN = 'http://images.cocodataset.org/annotations/image_info_test2017.zip'
# +
#export
def download_file_from_google_drive(file_id, destination, folder_name=None):
if folder_name:
dst = Config()['data'] / folder_name
if dst.exists():
return dst
else:
dst = Config()['data']
arch_dst = Config()['archive'] / destination
if not arch_dst.exists():
URL = "https://docs.google.com/uc?export=download"
session = requests.Session()
response = session.get(URL, params = { 'id' : file_id }, stream = True)
token = first([(k,v) for k,v in response.cookies.items() if k.startswith('download_warning')])[1]
if token:
params = { 'id' : file_id, 'confirm' : token }
response = session.get(URL, params = params, stream = True)
save_response_content(response, Config()['archive'] / destination)
file_extract(Config()['archive'] / destination, Config()['data'])
return dst
def save_response_content(response, destination):
CHUNK_SIZE = 32768
with open(destination, "wb") as f:
for chunk in response.iter_content(CHUNK_SIZE):
if chunk: # filter out keep-alive new chunks
f.write(chunk)
# -
# Original Inception's weights for FID by mseitzer
#export
FID_WEIGHTS_URL = 'https://github.com/mseitzer/pytorch-fid/releases/download/fid_weights/pt_inception-2015-12-05-6726825d.pth'
#export
def download_coco(force_download=False):
dest = Config()['data'] / 'coco'
paths = {'base': dest}
paths['train'] = untar_data(URLs.COCO_TRAIN2017, dest=dest, force_download=force_download)
paths['val'] = untar_data(URLs.COCO_VAL2017, dest=dest, force_download=force_download)
paths['test'] = untar_data(URLs.COCO_TEST2017, dest=dest, force_download=force_download)
paths['trainval_ann'] = untar_data(URLs.COCO_TRAINVAL_ANN, dest=dest, force_download=force_download)
paths['train_ann'] = paths['trainval_ann'] / 'instances_train2017.json'
paths['val_ann'] = paths['trainval_ann'] / 'instances_val2017.json'
return paths
# ### Renormalization stats
# from -1,1 to `imagenet_stats`
#exports
renorm_stats = (2*torch.tensor(imagenet_stats[0])-1).tolist(), (2*torch.tensor(imagenet_stats[1])).tolist()
x = TensorImage(torch.rand(1, 3, 32, 32))
xn = Normalize(0.5, 0.5)(x)
xin = Normalize.from_stats(*imagenet_stats, cuda=False)(x)
xn2in = Normalize.from_stats(*renorm_stats, cuda=False)(xn)
test(x, xn, nequals)
test(x, xin, nequals)
test(xin, xn, nequals)
test(xin, xn2in, all_equal)
# ## DataLoaders
#export
@patch
def is_relative_to(self:Path, *other):
"""Return True if the path is relative to another path or False.
"""
try:
self.relative_to(*other)
return True
except ValueError:
return False
#export
def _parent_idxs(items, name):
def _inner(items, name): return mask2idxs(Path(o).parent.name == name for o in items)
return [i for n in L(name) for i in _inner(items,n)]
#export
@delegates(get_image_files)
def get_tuple_files_by_stem(paths, folders=None, **kwargs):
if not is_listy(paths): paths = [paths]
files = []
for path in paths: files.extend(get_image_files(path, folders=folders))
out = L(groupby(files, attrgetter('stem')).values())
return out
#export
def ParentsSplitter(train_name='train', valid_name='valid'):
"Split `items` from the grand parent folder names (`train_name` and `valid_name`)."
def _inner(o):
tindex = _parent_idxs(L(o).itemgot(-1), train_name)
vindex = _parent_idxs(L(o).itemgot(-1), valid_name)
return tindex, vindex
return _inner
#export
class FilterRelToPath:
def __init__(self, path): self.path = path
def __call__(self, o): return L(o).filter(Self.is_relative_to(self.path))
#export
class CGANDataLoaders(DataLoaders):
"Basic wrapper around several `DataLoader`s with factory methods for CGAN problems"
@classmethod
@delegates(DataLoaders.from_dblock)
def from_paths(cls, input_path, target_path, train='train', valid='valid', valid_pct=None, seed=None, vocab=None, item_tfms=None,
batch_tfms=None, n_inp=1, add_normalize=True, **kwargs):
"Create from imagenet style dataset in `path` with `train` and `valid` subfolders (or provide `valid_pct`)"
splitter = ParentsSplitter(train_name=train, valid_name=valid) if valid_pct is None else RandomSplitter(valid_pct, seed=seed)
get_items = get_tuple_files_by_stem if valid_pct else partial(get_tuple_files_by_stem, folders=[train, valid])
if not Path(input_path).is_absolute(): input_path = Config()['data'] / input_path
if not Path(target_path).is_absolute(): target_path = Config()['data'] / target_path
if add_normalize:
if batch_tfms is None:
batch_tfms = [Normalize.from_stats([0.5]*3, [0.5]*3)]
else:
batch_tfms += [Normalize.from_stats([0.5]*3, [0.5]*3)]
dblock = DataBlock(blocks=(ImageTupleBlock, ImageTupleBlock),
get_items=get_items,
splitter=splitter,
get_x=FilterRelToPath(input_path),
get_y=noop,
item_tfms=item_tfms,
batch_tfms=batch_tfms)
return cls.from_dblock(dblock, [input_path, target_path], **kwargs)
@classmethod
@delegates(DataLoaders.from_dblock)
def from_path_ext(cls, path, folders, input_ext='.png', output_ext='.jpg', valid_pct=0.2, seed=None, item_tfms=None,
batch_tfms=None, **kwargs):
"Create from list of `fnames` in `path`s with `label_func`"
get_itmes = partial(get_tuple_files_by_stem, folders=folders)
files = get_itmes(path)
dblock = DataBlock(blocks=(ImageBlock, ImageTupleBlock),
get_items=get_itmes,
splitter=RandomSplitter(valid_pct, seed=seed),
get_x=lambda o: L(o).filter(lambda x: x.suffix==input_ext)[0],
get_y=lambda o: L(o).sorted(key=lambda x: {input_ext:0, output_ext:1}[x.suffix]),
item_tfms=item_tfms,
batch_tfms=batch_tfms)
return cls.from_dblock(dblock, path, **kwargs)
# +
#slow
path_base = untar_data(URLs.FACADES_BASE)
item_tfms=Resize(286, ResizeMethod.Squish, resamples=(Image.NEAREST, Image.NEAREST)),
batch_tfms=[Normalize.from_stats(0.5*torch.ones(3), 0.5*torch.ones(3)),
*aug_transforms(size=256, mult=0.0, max_lighting=0, p_lighting=0, mode='nearest')]
dls = CGANDataLoaders.from_path_ext(path_base.parent, ['base', 'extended'], item_tfms=item_tfms, batch_tfms=batch_tfms, bs=1)
# -
#slow
dls.show_batch()
#slow
item_tfms=Resize(286),
dls = CGANDataLoaders.from_paths('coco_under_water', 'coco', train='test2017', valid='val2017',
item_tfms=item_tfms, bs=16, num_workers=16, n_inp=-1)
#slow
dls.show_batch(max_n=2)
# ## Gather Logs Callback -
# +
# #export
# def basic_nested_repr(flds=None):
# if isinstance(flds, str): flds = re.split(', *', flds)
# flds = list(flds or [])
# def _f(self):
# sig = ', '.join(f'{o}={maybe_attr(nested_attr(self,o), "__name__")}' for o in flds)
# return f'{self.__class__.__name__}({sig})'
# return _f
# -
#export
def basic_name(flds=None):
if isinstance(flds, str): flds = re.split(', *', flds)
flds = list(flds or [])
def _f(self):
return '_'.join(f'{maybe_attr(nested_attr(self,o), "__name__")}' for o in flds)
return _f
#export
class GatherLogs(Callback):
'''Gather logs from one or more experiments.'''
run_after=Recorder
def __init__(self, experiments='logs', save_after_fit=True):
self.experiments = experiments
self.file = Path(self.experiments + '.pkl')
self.save_after_fit = save_after_fit
self.experiment = None
self.df = None
self.experiment_count = defaultdict(int)
def before_fit(self):
self.values = L()
if not self.experiment: self.set_experiment_name()
def after_epoch(self): self.values.append(self.recorder.log)
def set_experiment_name(self, name=None):
self.experiment = ifnone(name, basic_name('model.__class__,loss_func.__class__')(self))
self.experiment_count[name] += 1
def after_fit(self):
df = pd.DataFrame(self.values, columns=self.recorder.metric_names)
df['experiment'] = self.experiment
df['experiment_count'] = self.experiment_count[self.experiment]
df['time'] = pd.to_timedelta(df['time'].map(lambda x: '00:'+x if x.count(':')==1 else x))
df['time'] = df['time'].map(methodcaller('total_seconds'))
df = self.to_tidy(df)
self.df = pd.concat([self.df, df]).reset_index(drop=True)
if self.save_after_fit: self.save()
def to_tidy(self, df):
'''Creates tidy dataframe from metrics dataframe.'''
df = df.set_index(['epoch', 'time', 'experiment', 'experiment_count'])
df = df.stack().reset_index(level=4)
df['stage'] = df['level_4'].str.split('_').map(itemgetter(0))
df['metric'] = df['level_4'].str.split('_', n=1).map(itemgetter(1))
df = df.drop('level_4', axis=1)
df = df.set_index(['stage', 'metric'], append=True)
df[0] = df[0].astype(float)
df = df.unstack()
df.columns = df.columns.get_level_values(1)
df.columns.name = None
return df.reset_index()
@delegates(sns.relplot)
def plot_metric(self, y='loss', x='epoch' , col='stage', hue='experiment', kind='line', **kwargs):
return sns.relplot(data=self.df, x=x, y=y, col=col, hue=hue, kind=kind)
@delegates(sns.barplot)
def plot_time(self, x='experiment', y='time', **kwargs):
return sns.barplot(data=self.df[self.df['stage']=='train'], x=x, y=y, **kwargs)
def save(self):
self.learn = None
with self.file.open('bw') as f: pickle.dump(self, f)
learn = synth_learner()
gl = GatherLogs()
with learn.added_cbs(gl):
#gl.set_experiment_name('test1')
learn.fit(2)
gl.df
with learn.added_cbs(gl):
gl.set_experiment_name('test2')
learn.fit(2)
gl.df
gl.plot_metric();
gl.plot_time();
# ## RunNBatches Callback
#export
class RunNBatches(Callback):
run_after=Recorder
def __init__(self, n=2, no_valid=True): store_attr()
def after_batch(self):
if self.iter >= self.n:
if self.no_valid or not self.training: raise CancelFitException
raise CancelTrainException
def after_cancel_train(self): self.recorder.cancel_train = False
def after_cancel_validate(self): self.recorder.cancel_valid = False
learn = synth_learner()
with learn.added_cbs(RunNBatches()):
learn.fit(5)
with learn.added_cbs(RunNBatches(n=1, no_valid=False)):
learn.fit(5)
# ## Export Lib -
#hide
from nbdev.export import *
notebook2script()
| 00_core.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JamesJardinella/PRML/blob/master/Jardinella_Assignment_1_ch01_Introduction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ceRmVgOq6XmY"
# # 1. Introduction
# + [markdown] id="Wj_RTULpriir"
# <table class="tfo-notebook-buttons" align="left">
#
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/pantelis/PRML/blob/master/notebooks/ch01_Introduction.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# </table>
# + id="TMxmVuag7eyA" colab={"base_uri": "https://localhost:8080/"} outputId="26d0b3b3-bdac-4069-ba86-3cd6e65acf6b"
from google.colab import drive
drive.mount('/content/drive')
# + id="F3rYRMAD6ynO" colab={"base_uri": "https://localhost:8080/"} outputId="d8f88235-05e1-4db7-9860-739331516747"
# You need to adjust the directory names below for your own account
# e.g. you may elect to create ms-notebooks dir or not
# Execute this cell once
# 1. Download the repo and set it as the current directory
# %cd /content/drive/My Drive/Colab Notebooks/ml-notebooks
# !git clone https://github.com/pantelis/PRML
# %cd /content/drive/My Drive/Colab Notebooks/ml-notebooks/PRML
# 2. install the project/module
# !python setup.py install
# + id="iv9ADzLqiNsU" colab={"base_uri": "https://localhost:8080/"} outputId="1090e514-764d-4aba-f2bd-3e601369e01b"
# 3. Add the project directory to the path
# %cd /content/drive/My Drive/Colab Notebooks/ml-notebooks/PRML
import os, sys
sys.path.append(os.getcwd())
# + id="qwxjFZSR_vuX"
# Import seaborn
import seaborn as sns
# Apply the default theme
sns.set_theme()
# + id="pxBLdL3r6XmZ"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from prml.preprocess import PolynomialFeature
from prml.linear import (
LinearRegression,
RidgeRegression,
BayesianRegression
)
np.random.seed(1234)
# + [markdown] id="TWjYKLNc6Xmd"
# ## 1.1. Example: Polynomial Curve Fitting
# + [markdown] id="6XPE-VC_-OYK"
# The cell below defines $p_{data}(y|x)$ and generates the $\hat p_{data}(y|x)$
# + id="Tj4RTV3X6Xmd" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="8ff3be27-2ecd-40f4-a703-55c54c4030ca"
def create_toy_data(func, sample_size, std):
x = np.linspace(0, 1, sample_size) # p(x)
y = func(x) + np.random.normal(scale=std, size=x.shape)
return x, y
def func(x):
return np.sin(2 * np.pi * x)
x_train, y_train = create_toy_data(func, 10, 0.25)
x_test = np.linspace(0, 1, 100)
y_test = func(x_test)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.legend()
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
# + id="diJlsNGo6Xmg" colab={"base_uri": "https://localhost:8080/", "height": 661} outputId="b4333fec-8e99-453f-e6cd-d9003e3e1f90"
plt.subplots(figsize=(20, 10))
for i, degree in enumerate([0, 1, 3, 9]):
plt.subplot(2, 2, i + 1).set_title('M = {}'.format(degree))
####this makes 4+ subplots. Added title to make it clear which one I am comparing. Took me a while to figure out.
feature = PolynomialFeature(degree) ##### degree is M???
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegression()
model.fit(X_train, y_train)
print("w's for M = ", degree, model.w)
y = model.predict(X_test)
#print(degree,feature,y)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="hypothesis")
plt.ylim(-1.5, 1.5)
plt.annotate("M={}".format(degree), xy=(-0.15, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.legend(bbox_to_anchor=(1.05, 0.64), loc=2, borderaxespad=0.)
plt.show()
# + id="nfpg434z6Xmj" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="b1f83f57-3da5-4b0b-9b2c-cee1b4eede07"
def rmse(a, b):
return np.sqrt(np.mean(np.square(a - b)))
training_errors = []
test_errors = []
for i in range(10):
feature = PolynomialFeature(i) ##### is this the degree?
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = LinearRegression()
model.fit(X_train, y_train)
y = model.predict(X_test)
# print(model.coef_)
training_errors.append(rmse(model.predict(X_train), y_train))
test_errors.append(rmse(model.predict(X_test), y_test + np.random.normal(scale=0.25, size=len(y_test))))
plt.plot(training_errors, 'o-', mfc="none", mec="b", ms=10, c="b", label="Training")
plt.plot(test_errors, 'o-', mfc="none", mec="r", ms=10, c="r", label="Test")
plt.legend()
plt.xlabel("model capacity (degree)")
plt.ylabel("RMSE")
plt.show()
# + [markdown] id="nP67stVjbGRK"
# **Add a new text cell and explain why the loss function is called Root Mean Squared Error (RMSE) and what the square root offers, if anything, to finding a better solution.**
#
# RMSE--Root Mean Squared Error quantifies the mean distance of the residual (or error) between the expected valuea y and the values estimated by the model, $\hat y$. Because $\hat y$ for each point may be either above or below the actual y value, the value for the residual may be positive or negative. To avoid these values cancelling each other out when we find the total sum, we square each value and hence deal in positive values. Finding the root of this value brings the units of the function back to the original units in the data (they are no longer squared). The goal is to minimize this function within reason to avoid overfitting. In reality you can define an error function to suit your purposes--e.g. the squaring of this tends to overstate outlier points so you may want to add something to penalize those points. I have seen that folks often multiply the whole function by 1/2 to make it easier to take a derivative (the square root being the 1/2 power).
#
#
#
#
#
#
# + [markdown] id="lsuT0z1KimmB"
# **Model complexity (15 points)**
#
# **Add a text cell after the corresponding figure explaining the behavior of the test error for M=9 vs M=3**.
#
# The RMSE of the test is ~0.3 at M= 3 wheras it increases to nearly 0.4 at M=9. We see this result despite the near 0 RMSE we see with he training set at M=9. This is because adding degrees beyond 3 will fit the training data perfectly but will suffer from overfitting. You can see this if you look at the 4 subplots above where M=0 and M= 1 underfit as they are straight lines, whereas M=3 approximates the points nicely (but not exactly, leaving some room), and M=9 passes directly through all the training points exactly but is such a contorted function that it doesn't really track the general shape of the data--so it cannot predict well for data not in the training set which it has overfit.(actually it looks in the graph above like M=5 5, which is slightly different than in class)
#
#
#
#
# + [markdown] id="PTOLlihm6Xml"
# #### Regularization
# + id="18aoGaUg6Xml" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="5c7b0b1b-844a-4e09-8e19-b5d44b03c603"
feature = PolynomialFeature(9)
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = RidgeRegression(alpha=1e-3)
model.fit(X_train, y_train)
y = model.predict(X_test)
#y = model.predict(X_test)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="hypothesis")
plt.ylim(-1.5, 1.5)
plt.legend()
plt.annotate("M=9", xy=(-0.15, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.show()
# + [markdown] id="WoSp8MlZPVyn"
# **Regularization (20 points)**
#
# **Read the Linear Regression notes in the course site and add a text paragraph after the regularization cell explaining the output. Create a plot of the RMSE vs model complexity with regularization.**
# (note plot in 2 cells below)
#
# The MSE portion of the cost function tries to capture both the bias and the variance of the target variables and if the model complexity is too high it drives the bias down as the expense of variance. As variance increases, so does the generaliztion error which is where the model is overfitted. To combat this we add a regularization term reducesd the impact
#
# Regularization for linear models is usually accomplished via some sort of mechanism to constrain the model weights. In this case we are using the Ridge regression model which adds a regularization term to the error calculation/cost function during training to fit the data and minimize the weights. The additional term is $\alpha$\$\frac{1}{2}\\Sigma^n_{i=1}\theta^2_i$ where the hyperparameter controls the amount of regularization.
#
# This additional term can also be expressed as $\frac{1}{2}(\|w\|_2)^2 $ where $\|w\|_2$ represents the $l_2$ norm of the vector of weights [$\theta_1...\theta_n$].
#
# In the code we have $\alpha$ set to .001
#
# We can see the impact that regularization has if we compare the fit of the model in the LinearRegression with a polynomial of 9--which is highly overfitted, to the fit of the results of the Ridge regression with the same polynomial--which is fitted similar to the Linear regression with a lower polynomial with better generalization. We can also see this difference in the RMSE vs. Model Capacity graph below.
#
# Note: This is from Geron 133-134, I wanted to copy so I could remember and so I could play with Latex, which I find to be painful. Note the notes uses $\lambda$ insteat of $\alpha$ in the discussion of the 𝑙2 norm
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="QFsJxUe4sJFc" outputId="ab3b07e8-09b8-40ac-f858-b0d6e587b50d"
def rmse(a, b):
return np.sqrt(np.mean(np.square(a - b)))
training_errors = []
test_errors = []
for i in range(10):
feature = PolynomialFeature(i) ##### is this the degree?
X_train = feature.transform(x_train)
X_test = feature.transform(x_test)
model = RidgeRegression(alpha=1e-3)
model.fit(X_train, y_train)
y = model.predict(X_test)
# print(model.coef_)
training_errors.append(rmse(model.predict(X_train), y_train))
test_errors.append(rmse(model.predict(X_test), y_test + np.random.normal(scale=0.25, size=len(y_test))))
plt.plot(training_errors, 'o-', mfc="none", mec="b", ms=10, c="b", label="Training")
plt.plot(test_errors, 'o-', mfc="none", mec="r", ms=10, c="r", label="Test")
plt.legend()
plt.xlabel("model capacity (degree)")
plt.ylabel("RMSE")
plt.show()
# + [markdown] id="kfmJy1-96Xmo"
# ### 1.2.6 Bayesian curve fitting
# + id="GFCXxwiz6Xmo" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="3df98ea8-4143-482b-d728-280d4b749cd6"
model = BayesianRegression(alpha=2e-3, beta=2)
model.fit(X_train, y_train)
y, y_err = model.predict(X_test, return_std=True)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", s=50, label="training data")
plt.plot(x_test, y_test, c="g", label="$\sin(2\pi x)$")
plt.plot(x_test, y, c="r", label="mean")
plt.fill_between(x_test, y - y_err, y + y_err, color="pink", label="std.", alpha=0.5)
plt.xlim(-0.1, 1.1)
plt.ylim(-1.5, 1.5)
plt.annotate("M=9", xy=(0.8, 1))
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.legend(bbox_to_anchor=(1.05, 1.), loc=2, borderaxespad=0.)
plt.show()
# + id="i4VIskNS6Xmt"
| Jardinella_Assignment_1_ch01_Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Set 104b
#
# Complete the problems described below.
#
# Submit your work via email. A special email address has been setup for this purpose, and you can attach whatever files are appropriate to any mail sent to the address below. So I can tell which files below to whom, please **title all submitted files as**:
#
# LASTNAME firstname.py
#
# Submit this file to the class file submission email:
#
# <EMAIL>
#
# For one or more of the problems described below, a drawing or series of drawings may be required. These may be completed digitally (in Illustrator or similar) or by hand (in pencil or similar, then scanned). Please submit a single **multi-page** PDF (one page per problem) to the problem set submission email address.
# +
from decodes.core import *
from decodes.io.jupyter_out import JupyterOut
out = JupyterOut.two_pi( )
from math import *
import pprint
pp = pprint.PrettyPrinter(indent=4)
# -
# ## Python Dictionaries
# ### Dict Construction
# ### 104b.01
#
# The [Google Places API](https://developers.google.com/places/) provides software applications with access to geo-located search results. Given a search in a particular area, results may be returned in XML format, which stores nested key-value pairs that closely resemble and might also be structured as a Python Dictionary. While Python Dicts use the sytnax `key:value` the anagolous XML syntax would be `<key>value</key>`.
#
# Below is a simplified response from the Google Places API in XML format detailing the first few results from a search for restaurants near <NAME>. Reformat this information as a Python Dict.
#
# Note that since the keys of a Python Dict must be unique, this restriction does not hold for XML. As such, some changes have been made to the following results. In your reformatting, any numeric values should be expressed as number literals, and dll other values should be expressed as Strings.
#
# <?xml version="1.0" encoding="UTF-8"?>
# <PlaceSearchResponse>
# <status>OK</status>
# <result 1>
# <name><NAME></name>
# <vicinity>2600 Durant Avenue, Berkeley</vicinity>
# <type>restaurant</type>
# <rating>4.2</rating>
# </result 1>
# <result 2>
# <name><NAME></name>
# <vicinity>2399 Telegraph Avenue, Berkeley</vicinity>
# <type>store</type>
# <geometry>
# <location>37.8670352,-122.2587249</location>
# </geometry>
# <rating>4.5</rating>
# <price_level>1</price_level>
# </result 2>
# <result 3>
# <name>International House Cafe</name>
# <vicinity>2299 Piedmont Avenue, Berkeley</vicinity>
# <type>cafe</type>
# <geometry>
# <location>37.8696799,-122.2520091</location>
# </geometry>
# <rating>3.4</rating>
# <price_level>1</price_level>
# </result 3>
# <result 4>
# <name>Chipotle Mexican Grill</name>
# <vicinity>2311 Telegraph Avenue, Berkeley</vicinity>
# <type>restaurant</type>
# <rating>3.8</rating>
# </result 4>
# <result 5>
# <name><NAME></name>
# <vicinity>2522 Bancroft Way, Berkeley</vicinity>
# <type>cafe</type>
# <rating>4.2</rating>
# </result 5>
# </PlaceSearchResponse>
# +
#### START YOUR WORK HERE ####
api_response = {
'status':'OK',
'results':[
???,
???,
???,
???,
# ???
]
}
#### END YOUR WORK HERE ####
print(api_response)
# -
# ### Dict Access and Manipulation
# ### 104b.02
#
# Using the Dict constructed above, and following the samples found in the text concerning Dict access and manipulation, write code that constructs each of the following variables and/or manipulations:
#
# * Set the variable `how_much_chipotle_sux` to a number of the rating for Chipotle Mexican Grill.
# * Set the variable `bool_rice_and_bones` to a boolean value that tells us if the new Rice and Bones restaurant is present in the `place_search_response` dict.
# * Set the variable `lst_restaurants` to a List of all results where "type" is equal to "restaurant".
# * Remove the second result from the response.
# +
# some things to get us started
results = api_response['results']
idx_of_chipotle_result = -1
bool_rice_and_bones = False
lst_restaurants = []
#### START YOUR WORK HERE ####
# a loop over each result
for n, result in enumerate( results ):
# ??
# ??
# ??
# this should be set after the loop has finished
how_much_chipotle_sux = ??
# for removing the second response
# ??
#### END YOUR WORK HERE ####
print(how_much_chipotle_sux)
print(bool_rice_and_bones)
print(lst_restaurants)
# -
# ## Multi-Dimensionality
# ### Multiple Paths to Higher Dimensions
# ### 104b.03
#
# Populating the given collection `collection` only with the given variable `o`, construct a very simple multi-dimensional collection for which the overall **rank** is impossible to determine.
#
# The collection should be very simple (perhaps the simplest possible that meets this mandate) and should be constructed without the use of loops or other control structures.
# +
o = "object"
#### START YOUR WORK HERE ####
collection = []
#### END YOUR WORK HERE ####
# -
# ### Nests in Python
# ### 104b.04
#
# A rectangular grid of Points `pt_grid` is given, which might be comprised of any number of Points in the x and y directions. Write code to plot line Segments between each pair of adjacent Points in both horizontal and vertical directions. Store the Segments created in the `segs` List provided.
# +
x_cnt = random.choice([5,6,7])
y_cnt = random.choice([3,4,5])
rect_grid = [ [Point(x,y) for x in Interval(-4,4).divide(x_cnt-1,True)] for y in Interval(-2,2).divide(y_cnt-1,True)]
print("plotting a {} x {} rectangular grid".format(x_cnt, y_cnt))
out.put(rect_grid)
segs = []
#### START YOUR WORK HERE ####
for yi in range(y_cnt):
for xi in range(x_cnt):
# ??
#### END YOUR WORK HERE ####
out.put(segs)
out.draw()
out.clear()
# -
# ### 104b.05
# +
x_cnt, y_cnt = random.choice([8,9,10]), random.choice([5,6,7])
x_ival, y_ival = Interval(-3,4), Interval(-1,2)
tri_grid = []
vec_t = Vec((x_ival.eval(1/x_cnt) - x_ival.a)/2,0)
for yi in range(y_cnt):
row = [Point(x_ival.eval(xi/x_cnt),y_ival.eval(yi/y_cnt)) for xi in range(x_cnt)]
if yi%2 != 0: row = [pt+vec_t for pt in row[:-1]]
tri_grid.append(row)
print("plotting a {} x {} triangular grid".format(x_cnt, y_cnt))
out.put(tri_grid)
segs = []
#### START YOUR WORK HERE ####
for yi in range(y_cnt):
for xi in range(x_cnt):
# ??
#### END YOUR WORK HERE ####
out.put(segs)
out.draw()
out.clear()
# -
# ### Trees in Python
# ### 104b.06
#
# In the code below, the collection `tree` is constructed from two objects: `s`, a String, and `p`, a Point.
#
# Using Boolean or Float values, define the five variables at the end of the code with the appropriate values that describe the collection `tree`. If a property cannot be determined, define the variable as `False`.
# +
s = "string"
p = Point()
tree = [
s,
[s,s,s],
[p,p,p]
]
#### START YOUR WORK HERE ####
tree_rank = ??
tree_is_rectangular = ??
tree_is_jagged = ??
tree_is_homogeneous = ??
tree_is_heterogeneous = ??
#### END YOUR WORK HERE ####
# -
# ## Iterative Structures of Control
# ### Enumeration
# ### 104b.07
# Three collections of equal size are given: a List of Points `pts`, a List of Colors `clrs`, and a List of weight values `wgts`.
#
# Using the `enumerate` function, construct a loop that iterates over each of the given Points (and its index), assigns the cooresponding Color and weight value to the Point, and plots it.
# +
cnt = 100
pts = [Point.random(Interval(-3.5,3.5)) for n in range(cnt)]
clrs = [Color(random.random(),random.random(),0.5) for n in range(cnt)]
wgts = [random.uniform(2,10) for n in range(cnt)]
#### START YOUR WORK HERE ####
# ??
#### END YOUR WORK HERE ####
out.draw()
out.clear()
# -
# ### List Comprehesion
# ### 104b.08
#
# A collection of Points `pts` and a single Point `pt_attr` are given.
#
# Using a ***single line of code*** construct a new List `vecs` that contains a collection of Vecs that describe the direction and distance **from** `pt_attr` **to** each of the Points in `pts`. You will require List comprehnesion syntax for this.
# +
pt_attr = Point(4,4)
pts = [Point.random(Interval(-3.5,3.5)) for n in range(20)]
#### START YOUR WORK HERE ####
# ??
#### END YOUR WORK HERE ####
# -
# ### Iterators and Itertools
# ### 104b.09
#
# Two collections are given: one of Points and another of Vecs. ***Using the `product` function of the itertools module***, complete the code below such that each given Point is translated by each of the given Vecs, with a Segment drawn between the initial Point and the translated Point.
# +
cnt = 10
pts = [Point.random(Interval(-3.5,3.5)) for n in range(cnt)]
vecs = [Vec(0.5,0),Vec(0.5,0.5),Vec(0,0.5),Vec(-0.5,0.5) ]
#### START YOUR WORK HERE ####
# ??
#### END YOUR WORK HERE ####
out.put(pts)
out.draw()
out.clear()
# -
# ### Interval Objects in Decod.es
# ### 104b.10
#
# Two Points are given, `pt_a` and `pt_b`, at random positions and holding random weights and colors.
#
# Complete the code below to define a third Point, `pt_c` at a location halfway between the given Points, with a weight halfway between the weights of the given Points, and with a color halfway between the colors of the given Points. Interval objects will assist you in this.
# +
pt_a, pt_b = Point.random(Interval(-3,3)), Point.random(Interval(-3,3))
clr_a, clr_b = Color(random.random(),0,random.random()), Color(random.random(),random.random(),0)
wgt_a, wgt_b = random.uniform(2,10), random.uniform(2,10)
pt_a.set_color(clr_a)
pt_b.set_color(clr_b)
pt_a.set_weight(wgt_a)
pt_b.set_weight(wgt_b)
#### START YOUR WORK HERE ####
pt_c = ??
pt_c.set_weight(??)
pt_c.set_color(??)
#### END YOUR WORK HERE ####
out.put([pt_a,pt_b,pt_c])
out.draw()
out.clear()
# -
| problem sets/Problem Set 104b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''quantum'': conda)'
# language: python
# name: python_defaultSpec_1597662856438
# ---
# # Complex Arithmetic
#
# This is a tutorial designed to introduce you to complex arithmetic.
# This topic isn't particularly expansive, but it's important to understand it to be able to work with quantum computing.
#
# This tutorial covers the following topics:
#
# * Imaginary and complex numbers
# * Basic complex arithmetic
# * Complex plane
# * Modulus operator
# * Imaginary exponents
# * Polar representation
#
# If you need to look up some formulas quickly, you can find them in [this cheatsheet](https://github.com/microsoft/QuantumKatas/blob/main/quickref/qsharp-quick-reference.pdf).
#
# If you are curious to learn more, you can find more information at [Wikipedia](https://en.wikipedia.org/wiki/Complex_number).
# This notebook has several tasks that require you to write Python code to test your understanding of the concepts. If you are not familiar with Python, [here](https://docs.python.org/3/tutorial/index.html) is a good introductory tutorial for it.
#
# Let's start by importing some useful mathematical functions and constants, and setting up a few things necessary for testing the exercises. **Do not skip this step**.
#
# Click the cell with code below this block of text and press `Ctrl+Enter` (`⌘+Enter` on Mac).
# + tags=[]
# Run this cell using Ctrl+Enter (⌘+Enter on Mac).
from testing import exercise
from typing import Tuple
import math
Complex = Tuple[float, float]
Polar = Tuple[float, float]
# -
# # Algebraic Perspective
#
# ## Imaginary numbers
#
# For some purposes, real numbers aren't enough. Probably the most famous example is this equation:
#
# $$x^{2} = -1$$
#
# which has no solution for $x$ among real numbers. If, however, we abandon that constraint, we can do something interesting - we can define our own number. Let's say there exists some number that solves that equation. Let's call that number $i$.
#
# $$i^{2} = -1$$
#
# As we said before, $i$ can't be a real number. In that case, we'll call it an **imaginary unit**. However, there is no reason for us to define it as acting any different from any other number, other than the fact that $i^2 = -1$:
#
# $$i + i = 2i \\
# i - i = 0 \\
# -1 \cdot i = -i \\
# (-i)^{2} = -1$$
#
# We'll call the number $i$ and its real multiples **imaginary numbers**.
#
# > A good video introduction on imaginary numbers can be found [here](https://youtu.be/SP-YJe7Vldo).
# ### <span style="color:blue">Exercise 1</span>: Powers of $i$.
#
# **Input:** An even integer $n$.
#
# **Goal:** Return the $n$th power of $i$, or $i^n$.
#
# > Fill in the missing code (denoted by `...`) and run the cell below to test your work.
# + tags=[]
@exercise
def imaginary_power(n : int) -> int:
# If n is divisible by 4
if n % 4 == 0:
return 1
else:
return -1
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-1:-Powers-of-$i$.).*
# ## Complex Numbers
#
# Adding imaginary numbers to each other is quite simple, but what happens when we add a real number to an imaginary number? The result of that addition will be partly real and partly imaginary, otherwise known as a **complex number**. A complex number is simply the real part and the imaginary part being treated as a single number. Complex numbers are generally written as the sum of their two parts: $a + bi$, where both $a$ and $b$ are real numbers. For example, $3 + 4i$, or $-5 - 7i$ are valid complex numbers. Note that purely real or purely imaginary numbers can also be written as complex numbers: $2$ is $2 + 0i$, and $-3i$ is $0 - 3i$.
#
# When performing operations on complex numbers, it is often helpful to treat them as polynomials in terms of $i$.
# ### <span style="color:blue">Exercise 2</span>: Complex addition.
#
# **Inputs:**
#
# 1. A complex number $x = a + bi$, represented as a tuple `(a, b)`.
# 2. A complex number $y = c + di$, represented as a tuple `(c, d)`.
#
# **Goal:** Return the sum of these two numbers $x + y = z = g + hi$, represented as a tuple `(g, h)`.
#
# > A tuple is a pair of numbers.
# > You can make a tuple by putting two numbers in parentheses like this: `(3, 4)`.
# > * You can access the $n$th element of tuple `x` like so: `x[n]`
# > * For this tutorial, complex numbers are represented as tuples where the first element is the real part, and the second element is the real coefficient of the imaginary part
# > * For example, $1 + 2i$ would be represented by a tuple `(1, 2)`, and $7 - 5i$ would be represented by `(7, -5)`.
# >
# > You can find more details about Python's tuple data type in the [official documentation](https://docs.python.org/3/library/stdtypes.html#tuples).
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Remember, adding complex numbers is just like adding polynomials. Add components of the same type - add the real part to the real part, add the complex part to the complex part. <br>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=SfbjqVyQljk">here</a>.
# </details>
# + tags=[]
@exercise
def complex_add(x : Complex, y : Complex) -> Complex:
# You can extract elements from a tuple like this
a = x[0]
b = x[1]
c = y[0]
d = y[1]
# This creates a new variable and stores the real component into it
real = a + c
# Replace the ... with code to calculate the imaginary component
imaginary = b + d
# You can create a tuple like this
ans = (real, imaginary)
return ans
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-2:-Complex-addition.).*
# ### <span style="color:blue">Exercise 3</span>: Complex multiplication.
#
# **Inputs:**
#
# 1. A complex number $x = a + bi$, represented as a tuple `(a, b)`.
# 2. A complex number $y = c + di$, represented as a tuple `(c, d)`.
#
# **Goal:** Return the product of these two numbers $x \cdot y = z = g + hi$, represented as a tuple `(g, h)`.
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Remember, multiplying complex numbers is just like multiplying polynomials. Distribute one of the complex numbers:
#
# $$(a + bi)(c + di) = a(c + di) + bi(c + di)$$
#
# Then multiply through, and group the real and imaginary terms together.
# <br/>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=cWn6g8Qqvs4">here</a>.
# </details>
# + tags=[]
@exercise
def complex_mult(x : Complex, y : Complex) -> Complex:
# Fill in your own code
return ((x[0] * y[0] - x[1] * y[1]), (x[0] * y[1] + x[1] * y[0]))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-3:-Complex-multiplication.).*
# ## Complex Conjugate
#
# Before we discuss any other complex operations, we have to cover the **complex conjugate**. The conjugate is a simple operation: given a complex number $x = a + bi$, its complex conjugate is $\overline{x} = a - bi$.
#
# The conjugate allows us to do some interesting things. The first and probably most important is multiplying a complex number by its conjugate:
#
# $$x \cdot \overline{x} = (a + bi)(a - bi)$$
#
# Notice that the second expression is a difference of squares:
#
# $$(a + bi)(a - bi) = a^2 - (bi)^2 = a^2 - b^2i^2 = a^2 + b^2$$
#
# This means that a complex number multiplied by its conjugate always produces a non-negative real number.
#
# Another property of the conjugate is that it distributes over both complex addition and complex multiplication:
#
# $$\overline{x + y} = \overline{x} + \overline{y} \\
# \overline{x \cdot y} = \overline{x} \cdot \overline{y}$$
# ### <span style="color:blue">Exercise 4</span>: Complex conjugate.
#
# **Input:** A complex number $x = a + bi$, represented as a tuple `(a, b)`.
#
# **Goal:** Return $\overline{x} = g + hi$, the complex conjugate of $x$, represented as a tuple `(g, h)`.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=BZxZ_eEuJBM">here</a>.
# </details>
# + tags=[]
@exercise
def conjugate(x : Complex) -> Complex:
return (x[0], -x[1])
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-4:-Complex-conjugate.).*
# ## Complex Division
#
# The next use for the conjugate is complex division. Let's take two complex numbers: $x = a + bi$ and $y = c + di \neq 0$ (not even complex numbers let you divide by $0$). What does $\frac{x}{y}$ mean?
#
# Let's expand $x$ and $y$ into their component forms:
#
# $$\frac{x}{y} = \frac{a + bi}{c + di}$$
#
# Unfortunately, it isn't very clear what it means to divide by a complex number. We need some way to move either all real parts or all imaginary parts into the numerator. And thanks to the conjugate, we can do just that. Using the fact that any number (except $0$) divided by itself equals $1$, and any number multiplied by $1$ equals itself, we get:
#
# $$\frac{x}{y} = \frac{x}{y} \cdot 1 = \frac{x}{y} \cdot \frac{\overline{y}}{\overline{y}} = \frac{x\overline{y}}{y\overline{y}} = \frac{(a + bi)(c - di)}{(c + di)(c - di)} = \frac{(a + bi)(c - di)}{c^2 + d^2}$$
#
# By doing this, we re-wrote our division problem to have a complex multiplication expression in the numerator, and a real number in the denominator. We already know how to multiply complex numbers, and dividing a complex number by a real number is as simple as dividing both parts of the complex number separately:
#
# $$\frac{a + bi}{r} = \frac{a}{r} + \frac{b}{r}i$$
# ### <span style="color:blue">Exercise 5</span>: Complex division.
#
# **Inputs:**
#
# 1. A complex number $x = a + bi$, represented as a tuple `(a, b)`.
# 2. A complex number $y = c + di \neq 0$, represented as a tuple `(c, d)`.
#
# **Goal:** Return the result of the division $\frac{x}{y} = \frac{a + bi}{c + di} = g + hi$, represented as a tuple `(g, h)`.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=Z8j5RDOibV4">here</a>.
# </details>
# + tags=[]
@exercise
def complex_div(x : Complex, y : Complex) -> Complex:
return ((x[0] * y[0] + x[1] * y[1]) / (y[0]**2 + y[1]**2), (-x[0] * y[1] + x[1] * y[0]) / (y[0]**2 + y[1]**2))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-5:-Complex-division.).*
# # Geometric Perspective
#
# ## The Complex Plane
#
# You may recall that real numbers can be represented geometrically using the [number line](https://en.wikipedia.org/wiki/Number_line) - a line on which each point represents a real number. We can extend this representation to include imaginary and complex numbers, which gives rise to an entirely different number line: the imaginary number line, which only intersects with the real number line at $0$.
#
# A complex number has two components - a real component and an imaginary component. As you no doubt noticed from the exercises, these can be represented by two real numbers - the real component, and the real coefficient of the imaginary component. This allows us to map complex numbers onto a two-dimensional plane - the **complex plane**. The most common mapping is the obvious one: $a + bi$ can be represented by the point $(a, b)$ in the **Cartesian coordinate system**.
#
# 
#
# This mapping allows us to apply complex arithmetic to geometry, and, more importantly, apply geometric concepts to complex numbers. Many properties of complex numbers become easier to understand when viewed through a geometric lens.
# ## Modulus
#
# One such property is the **modulus** operator. This operator generalizes the **absolute value** operator on real numbers to the complex plane. Just like the absolute value of a number is its distance from $0$, the modulus of a complex number is its distance from $0 + 0i$. Using the distance formula, if $x = a + bi$, then:
#
# $$|x| = \sqrt{a^2 + b^2}$$
#
# There is also a slightly different, but algebraically equivalent definition:
#
# $$|x| = \sqrt{x \cdot \overline{x}}$$
#
# Like the conjugate, the modulus distributes over multiplication.
#
# $$|x \cdot y| = |x| \cdot |y|$$
#
# Unlike the conjugate, however, the modulus doesn't distribute over addition. Instead, the interaction of the two comes from the triangle inequality:
#
# $$|x + y| \leq |x| + |y|$$
# ### <span style="color:blue">Exercise 6</span>: Modulus.
#
# **Input:** A complex number $x = a + bi$, represented as a tuple `(a, b)`.
#
# **Goal:** Return the modulus of this number, $|x|$.
#
# > Python's exponentiation operator is `**`, so $2^3$ is `2 ** 3` in Python.
# >
# > You will probably need some mathematical functions to solve the next few tasks. They are available in Python's math library. You can find the full list and detailed information in the [official documentation](https://docs.python.org/3/library/math.html).
#
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# In particular, you might be interested in <a href=https://docs.python.org/3/library/math.html#math.sqrt>Python's square root function.</a><br>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=FwuPXchH2rA">here</a>.
# </details>
# + tags=[]
@exercise
def modulus(x : Complex) -> float:
return math.sqrt(x[0]**2 + x[1]**2)
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-6:-Modulus.).*
# ## Imaginary Exponents
#
# The next complex operation we're going to need is exponentiation. Raising an imaginary number to an integer power is a fairly simple task, but raising a number to an imaginary power, or raising an imaginary (or complex) number to a real power isn't quite as simple.
#
# Let's start with raising real numbers to imaginary powers. Specifically, let's start with a rather special real number - Euler's constant, $e$:
#
# $$e^{i\theta} = \cos \theta + i\sin \theta$$
#
# (Here and later in this tutorial $\theta$ is measured in radians.)
#
# Explaining why that happens is somewhat beyond the scope of this tutorial, as it requires some calculus, so we won't do that here. If you are curious, you can see [this video](https://youtu.be/v0YEaeIClKY) for a beautiful intuitive explanation, or [the Wikipedia article](https://en.wikipedia.org/wiki/Euler%27s_formula#Proofs) for a more mathematically rigorous proof.
#
# Here are some examples of this formula in action:
#
# $$e^{i\pi/4} = \frac{1}{\sqrt{2}} + \frac{i}{\sqrt{2}} \\
# e^{i\pi/2} = i \\
# e^{i\pi} = -1 \\
# e^{2i\pi} = 1$$
#
# > One interesting consequence of this is Euler's Identity:
# >
# > $$e^{i\pi} + 1 = 0$$
# >
# > While this doesn't have any notable uses, it is still an interesting identity to consider, as it combines 5 fundamental constants of algebra into one expression.
#
# We can also calculate complex powers of $e$ as follows:
#
# $$e^{a + bi} = e^a \cdot e^{bi}$$
#
# Finally, using logarithms to express the base of the exponent as $r = e^{\ln r}$, we can use this to find complex powers of any positive real number.
# ### <span style="color:blue">Exercise 7</span>: Complex exponents.
#
# **Input:** A complex number $x = a + bi$, represented as a tuple `(a, b)`.
#
# **Goal:** Return the complex number $e^x = e^{a + bi} = g + hi$, represented as a tuple `(g, h)`.
#
# > Euler's constant $e$ is available in the [math library](https://docs.python.org/3/library/math.html#math.e),
# > as are [Python's trigonometric functions](https://docs.python.org/3/library/math.html#trigonometric-functions).
# + tags=[]
@exercise
def complex_exp(x : Complex) -> Complex:
return (math.e**x[0] * math.cos(x[1]), math.e**x[0] * math.sin(x[1]))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-7:-Complex-exponents.).*
# ### <span style="color:blue">Exercise 8</span>*: Complex powers of real numbers.
#
# **Inputs:**
#
# 1. A non-negative real number $r$.
# 2. A complex number $x = a + bi$, represented as a tuple `(a, b)`.
#
# **Goal:** Return the complex number $r^x = r^{a + bi} = g + hi$, represented as a tuple `(g, h)`.
#
# > Remember, you can use functions you have defined previously
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# You can use the fact that $r = e^{\ln r}$ to convert exponent bases. Remember though, $\ln r$ is only defined for positive numbers - make sure to check for $r = 0$ separately!
# </details>
# + tags=[]
@exercise
def complex_exp_real(r : float, x : Complex) -> Complex:
if r == 0:
return (0.0, 0.0)
else:
return complex_exp((math.log(r) * x[0], math.log(r) * x[1]))
# -
# <i>Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-8*:-Complex-powers-of-real-numbers.).</i>
# ## Polar coordinates
#
# Consider the expression $e^{i\theta} = \cos\theta + i\sin\theta$. Notice that if we map this number onto the complex plane, it will land on a **unit circle** around $0 + 0i$. This means that its modulus is always $1$. You can also verify this algebraically: $\cos^2\theta + \sin^2\theta = 1$.
#
# Using this fact we can represent complex numbers using **polar coordinates**. In a polar coordinate system, a point is represented by two numbers: its direction from origin, represented by an angle from the $x$ axis, and how far away it is in that direction.
#
# Another way to think about this is that we're taking a point that is $1$ unit away (which is on the unit circle) in the specified direction, and multiplying it by the desired distance. And to get the point on the unit circle, we can use $e^{i\theta}$.
#
# A complex number of the format $r \cdot e^{i\theta}$ will be represented by a point which is $r$ units away from the origin, in the direction specified by the angle $\theta$.
#
# 
#
# Sometimes $\theta$ will be referred to as the number's **phase**.
# ### <span style="color:blue">Exercise 9</span>: Cartesian to polar conversion.
#
# **Input:** A complex number $x = a + bi$, represented as a tuple `(a, b)`.
#
# **Goal:** Return the polar representation of $x = re^{i\theta}$, i.e., the distance from origin $r$ and phase $\theta$ as a tuple `(r, θ)`.
#
# * $r$ should be non-negative: $r \geq 0$
# * $\theta$ should be between $-\pi$ and $\pi$: $-\pi < \theta \leq \pi$
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# <a href=https://docs.python.org/3/library/math.html#math.atan2>Python has a separate function</a> for calculating $\theta$ for this purpose.<br>
# A video explanation can be found <a href="https://www.youtube.com/watch?v=8RasCV_Lggg">here</a>.
# </details>
# + tags=[]
@exercise
def polar_convert(x : Complex) -> Polar:
# r = math.sqrt(x[0]**2 + x[1]**2)
#
# if r == 0:
# return (0.0, 0.0)
# else:
# theta = math.atan(x[1] / x[0])
# if x[0] >= 0:
# return (r, theta)
# elif x[1] >= 0:
# return (r, theta + math.pi)
# else:
# return (r, theta - math.pi)
return (math.sqrt(x[0]**2 + x[1]**2), math.atan2(x[1], x[0]))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-9:-Cartesian-to-polar-conversion.).*
# ### <span style="color:blue">Exercise 10</span>: Polar to Cartesian conversion.
#
# **Input:** A complex number $x = re^{i\theta}$, represented in polar form as a tuple `(r, θ)`.
#
# **Goal:** Return the Cartesian representation of $x = a + bi$, represented as a tuple `(a, b)`.
# + tags=[]
@exercise
def cartesian_convert(x : Polar) -> Complex:
return (x[0] * math.cos(x[1]), x[0] * math.sin(x[1]))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-10:-Polar-to-Cartesian-conversion.).*
# ### <span style="color:blue">Exercise 11</span>: Polar multiplication.
#
# **Inputs:**
#
# 1. A complex number $x = r_{1}e^{i\theta_1}$ represented in polar form as a tuple `(r1, θ1)`.
# 2. A complex number $y = r_{2}e^{i\theta_2}$ represented in polar form as a tuple `(r2, θ2)`.
#
# **Goal:** Return the result of the multiplication $x \cdot y = z = r_3e^{i\theta_3}$, represented in polar form as a tuple `(r3, θ3)`.
#
# * $r_3$ should be non-negative: $r_3 \geq 0$
# * $\theta_3$ should be between $-\pi$ and $\pi$: $-\pi < \theta_3 \leq \pi$
# * Try to avoid converting the numbers into Cartesian form.
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Remember, a number written in polar form already involves multiplication. What is $r_1e^{i\theta_1} \cdot r_2e^{i\theta_2}$?
# </details><details>
# <summary><strong>Need another hint? Click here</strong></summary>
# Is your θ not coming out correctly? Remember you might have to check your boundaries and adjust it to be in the range requested.
# </details>
# + tags=[]
@exercise
def polar_mult(x : Polar, y : Polar) -> Polar:
if x[1] + y[1] >= math.pi:
return (x[0] * y[0], (x[1] + y[1]) - 2 * math.pi)
elif x[1] + y[1] <= -math.pi:
return (x[0] * y[0], (x[1] + y[1]) + 2 * math.pi)
else:
return (x[0] * y[0], x[1] + y[1])
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-11:-Polar-multiplication.).*
# ### <span style="color:blue">Exercise 12</span>**: Arbitrary complex exponents.
#
# You now know enough about complex numbers to figure out how to raise a complex number to a complex power.
#
# **Inputs:**
#
# 1. A complex number $x = a + bi$, represented as a tuple `(a, b)`.
# 2. A complex number $y = c + di$, represented as a tuple `(c, d)`.
#
# **Goal:** Return the result of raising $x$ to the power of $y$: $x^y = (a + bi)^{c + di} = z = g + hi$, represented as a tuple `(g, h)`.
#
# <br/>
# <details>
# <summary><strong>Need a hint? Click here</strong></summary>
# Convert $x$ to polar form, and raise the result to the power of $y$.
# </details>
# + tags=[]
@exercise
def complex_exp_arbitrary(x : Complex, y : Complex) -> Complex:
x_polar = polar_convert(x)
if x_polar[0] == 0:
return (0, 0)
else:
x_polar_exponent = (math.log(x_polar[0]), x_polar[1])
multiplication = complex_mult(x_polar_exponent, (y[0], y[1]))
return cartesian_convert((math.exp(multiplication[0]), multiplication[1]))
# -
# *Can't come up with a solution? See the explained solution in the [Complex Arithmetic Workbook](./Workbook_ComplexArithmetic.ipynb#Exercise-12**:-Arbitrary-complex-exponents.).*
# ## Conclusion
#
# Congratulations! You should now know enough complex arithmetic to get started with quantum computing. When you are ready, you can move on to the next tutorial in this series, covering [linear algebra](../LinearAlgebra/LinearAlgebra.ipynb).
| tutorials/ComplexArithmetic/ComplexArithmetic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="nr04oer6VOHk"
#Description: this program detects if someone has diabetes using machine learning and python!
#Import the libraries
import pandas as pd
import numpy as np
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
import joblib
#Get the data for the model
df = pd.read_csv('https://raw.githubusercontent.com/ferris77/ml-web-app/main/diabetes.csv')
#We need to preprocess the data replacing zero values with suitable values (means)
df['Glucose'] = np.where(df['Glucose']==0,df['Glucose'].mean(),df['Glucose'])
df['BloodPressure'] = np.where(df['BloodPressure']==0,df['BloodPressure'].mean(),df['BloodPressure'])
df['SkinThickness'] = np.where(df['SkinThickness']==0,df['SkinThickness'].median(),df['SkinThickness'])
df['Insulin'] = np.where(df['Insulin']==0,df['Insulin'].median(),df['Insulin'])
df['BMI'] = np.where(df['BMI']==0,df['BMI'].mean(),df['BMI'])
# + colab={"base_uri": "https://localhost:8080/"} id="9Di0j-qeeAvO" outputId="acc6d852-1ade-49b3-afe3-b83fd3b716bd"
#Split the data into independentent 'X' and dependente 'Y' variables
X = df.iloc[:, 0:8].values #we want the array, not the df
Y = df.iloc[:, -1].values
#Split the data into 75% training and 25% testing
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.20, random_state=0)
#Create and train the model
model = RandomForestClassifier()
model.fit(X_train, Y_train)
#Let's check our models accuracy
predictions = model.predict(X_test)
print(f'Our first RFC model has an accuracy of: {accuracy_score(Y_test, predictions)*100:.2f}%')
# + colab={"base_uri": "https://localhost:8080/"} id="2qby4qB6f7Yd" outputId="b1eb2151-8569-45d8-d985-98fd63982214"
# Manual Hyperparameter Tuning
manual_tuned_model = RandomForestClassifier(n_estimators=100,criterion='gini',
max_features='sqrt',
min_samples_leaf=5,random_state=0)
manual_tuned_model.fit(X_train, Y_train)
manual_tuned_predictions = manual_tuned_model.predict(X_test)
print(f'Our manual tunned RFC model has an accuracy of: {accuracy_score(Y_test, manual_tuned_predictions)*100:.2f}%')
# + colab={"base_uri": "https://localhost:8080/"} id="3AceDCGAbxCG" outputId="51671c7c-d271-4dd9-ad55-489b34e79854"
#Hypertuning
from sklearn.model_selection import RandomizedSearchCV
import numpy as np
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 100, stop = 1200, num = 12)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt','log2']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 1000,10)]
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10,14]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4,6,8]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'criterion':['entropy','gini']}
print(random_grid)
auto_tuned_model = RandomForestClassifier()
RFC_randomcv = RandomizedSearchCV(estimator = auto_tuned_model,
param_distributions = random_grid,
n_iter = 20,
cv = 3,
verbose = 2,
random_state = 100,
n_jobs = -1)
### fit the randomized model
RFC_randomcv.fit(X_train,Y_train)
RFC_randomcv.best_params_
best_random_grid = RFC_randomcv.best_estimator_
auto_tuned_predictions = best_random_grid.predict(X_test)
print(f'Our manual tunned RFC model has an accuracy of: {accuracy_score(Y_test, auto_tuned_predictions)*100:.2f}%')
# + colab={"base_uri": "https://localhost:8080/"} id="B1BCfK1HcwBl" outputId="b5b6e4c5-ac98-4fd4-ded8-0fd0031716ce"
print(f'model accuracy: {accuracy_score(Y_test, predictions)*100:.2f}%')
print(f'manual_tuned_model accuracy: {accuracy_score(Y_test, manual_tuned_predictions)*100:.2f}%')
print(f'auto_tuned_model accuracy: {accuracy_score(Y_test, auto_tuned_predictions)*100:.2f}%')
# + colab={"base_uri": "https://localhost:8080/"} id="HBruX17GbYON" outputId="68f0e1d8-5bad-49f4-9972-860a418e9383"
#We reached higher accuracy with manual-tunned model, we will select this model
#to serialize and save to disk
joblib.dump(manual_tuned_model, 'model.pkl')
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <h1 align = 'center'> Neural Networks Demystified </h1>
# <h2 align = 'center'> Part 3: Gradient Descent </h2>
#
#
# <h4 align = 'center' > @stephencwelch </h4>
from IPython.display import YouTubeVideo
YouTubeVideo('5u0jaA3qAGk')
# <h3 align = 'center'> Variables </h3>
#
# |Code Symbol | Math Symbol | Definition | Dimensions
# | :-: | :-: | :-: | :-: |
# |X|$$X$$|Input Data, each row in an example| (numExamples, inputLayerSize)|
# |y |$$y$$|target data|(numExamples, outputLayerSize)|
# |W1 | $$W^{(1)}$$ | Layer 1 weights | (inputLayerSize, hiddenLayerSize) |
# |W2 | $$W^{(2)}$$ | Layer 2 weights | (hiddenLayerSize, outputLayerSize) |
# |z2 | $$z^{(2)}$$ | Layer 2 activation | (numExamples, hiddenLayerSize) |
# |a2 | $$a^{(2)}$$ | Layer 2 activity | (numExamples, hiddenLayerSize) |
# |z3 | $$z^{(3)}$$ | Layer 3 activation | (numExamples, outputLayerSize) |
# |J | $$J$$ | Cost | (1, outputLayerSize) |
# Last time we built a neural network in python that made really bad predictions of your score on a test based on how many hours you slept and how many hours you studied the night before. This time we'll focus on the theory of making those predictions better.
# We can initialize the network we built last time and pass in our normalized data, X, using our forward method, and have a look at our estimate of y, yHat.
# +
# %pylab inline
#Import code from last time:
from partTwo import *
# -
NN = Neural_Network()
yHat = NN.forward(X)
yHat
y
#Compare estimate, yHat, to actually score
bar([0,1,2], y, width = 0.35, alpha=0.8)
bar([0.35,1.35,2.35],yHat, width = 0.35, color='r', alpha=0.8)
grid(1)
legend(['y', 'yHat'])
# Right now our predictions are pretty inaccurate. To improve our model, we first need to quantify exactly how wrong our predictions are. We'll do this with a cost function. A cost function allows us to express exactly how wrong or "costly" our models is, given our examples.
# One way to compute an overall cost is to take each error value, square it, and add these values together. Multiplying by one half will make things simpler down the road. Now that we have a cost, or job is to minimize it. When someone says they’re training a network, what they really mean is that they're minimizing a cost function.
# $$
# J = \sum \frac{1}{2}(y-\hat{y})^2 \tag{5}
# $$
# OUR cost is a function of two things, our examples, and the weights on our synapses. We don't have much control of our data, so we'll minimize our cost by changing the weights.
# Conceptually, this is pretty simple concept. We have a collection of 9 individual weights, and we're saying that there is some combination of w's that will make our cost, J, as small as possible. When I first saw this problem in machine learning, I thought, I'll just try ALL THE WEIGHTS UNTIL I FIND THE BEST ONE! After all I have a computer!
# Enter the CURSE OF DIMENSIONALITY. Here's the problem. Let's pretend for a second that we only have 1 weight, instead of 9. To find the ideal value of our weight that will minimize our cost, we need to try a bunch of values for W, let's say we test 1000 values. That doesn't seem so bad, after all, my computer is pretty fast.
# +
import time
weightsToTry = np.linspace(-5,5,1000)
costs = np.zeros(1000)
startTime = time.clock()
for i in range(1000):
NN.W1[0,0] = weightsToTry[i]
yHat = NN.forward(X)
costs[i] = 0.5*sum((y-yHat)**2)
endTime = time.clock()
# -
timeElapsed = endTime-startTime
timeElapsed
# It takes about 0.04 seconds to check 1000 different weight values for our neural network. Since we’ve computed the cost for a wide range values of W, we can just pick the one with the smallest cost, let that be our weight, and we’ve trained our network.
plot(weightsToTry, costs)
grid(1)
ylabel('Cost')
xlabel('Weight')
# So you may be thinking that 0.04 seconds to train a network is not so bad, and we haven't even optimized anything yet. Plus, there are other, way faster languages than python our there.
# Before we optimize through, let's consider the full complexity of the problem. Remember the 0.04 seconds required is only for one weight, and we have 9 total! Let's next consider 2 weights for a moment. To maintain the same precision we now need to check 1000 times 1000, or one million values. This is a lot of work, even for a fast computer.
# +
weightsToTry = np.linspace(-5,5,1000)
costs = np.zeros((1000, 1000))
startTime = time.clock()
for i in range(1000):
for j in range(1000):
NN.W1[0,0] = weightsToTry[i]
NN.W1[0,1] = weightsToTry[j]
yHat = NN.forward(X)
costs[i, j] = 0.5*sum((y-yHat)**2)
endTime = time.clock()
# -
timeElapsed = endTime-startTime
timeElapsed
# After our 1 million evaluations we’ve found our solution, but it took an agonizing 40 seconds! The real curse of dimensionality kicks in as we continue to add dimensions. Searching through three weights would take a billion evaluations, or 11 hours! Searching through all 9 weights we need for our simple network would take 1,268,391,679,350,583.5 years. (Over a quardrillion years). So for that reason, the "just try everything" or brute force optimization method is clearly not going to work.
0.04*(1000**(9-1))/(3600*24*365)
# Let's return to the 1-dimensional case and see if we can be more clever. Let's evaluate our cost function for a specific value of w. If w is 1.1 for example, we can run our cost function, and see that J is 2.8. Now we haven't learned much yet, but let's try to add a little information to what we already know. What if we could figure out which way was downhill? If we could, we would know whether to make W smaller or larger do decrease the cost. We could test the cost function immediately to the left and right of our test point and see which is smaller. This is called numerical estimation, and is sometimes a good approach, but for us, there's a faster way. Let's look at our equations so far.
# $$
# z^{(2)} = XW^{(1)} \tag{1}\\
# $$
# $$
# a^{(2)} = f(z^{(2)}) \tag{2}\\
# $$
# $$
# z^{(3)} = a^{(2)}W^{(2)} \tag{3}\\
# $$
# $$
# \hat{y} = f(z^{(3)}) \tag{4}\\
# $$
# $$
# J = \sum \frac{1}{2}(y-\hat{y})^2 \tag{5}\\
# $$
# We have 5 equations, but we can really think of them as one big equation.
#
# And since we have one big equation that uniquely determines our cost, J, from X, y, W1, and W2, we can use our good friend calculus to find what we're looking for. We want to know "which way is downhill", that is, what is the rate of change of J with respect to W, also known as the derivative. And in this case, since we’re just considering one weight at a time, the partial derivative.
# We can derive an expression for dJdW, that will give us the rate of change of J with respect to W, for any value of W! If dJdW is positive, then the cost function is going uphill. If dJdW is negative the cost function is going downhill.
# Now we can really speed things up. Since we know in which direction the cost decreases, we can save all that time we would have spent searching in the wrong direction. We can save even more computational time by iteratively taking steps downhill and stopping when the cost stops getting smaller.
# This method is known as gradient descent, and although it may not seem so impressive in one dimension, it is capable of incredible speedups in higher dimensions. In fact, in our final video, we’ll show that what would have taken 10^27 function evaluations with our brute force method will take less than 100 evaluations with gradient descent. Gradient descent allows us to find needles in very very very large haystacks.
# Now before we celebrate too much here, there is a restriction. What if our cost function doesn't always go in the same direction? What if it goes up, then back down? The mathematical name for this is non-convex, and it could really throw off our gradient descent algorithm by getting it stuck in a local minima instead of our ideal global minima. One of the reasons we chose our cost function to be the sum of squared errors was to exploit the convex nature of quadratic equations.
# We know that the graph of y equals x squared is a nice convex parabola and it turns out that higher dimensional versions are too!
# Another piece of the puzzle here is that depending on how we use our data, it might not matter if or cost function is convex or not. If we use our examples one at a time instead of all at once, sometimes it won't matter if our cost function is convex, we will still find a good solution. This is called stochastic gradient descent. So maybe we shouldn't be afraid of non-convex cost functions, as Neural Network wizard <NAME> says in his excellent talk "Who is afraid on non-convex loss funtions?"
# Link to Yann's Talk:
# http://videolectures.net/eml07_lecun_wia/
# The details of gradient descent are a deep topic for another day, for now we're going to do our gradient descent "batch" style, where we use all our example at once, and the way we've setup our cost function will keep things nice and convex. Next time we'll compute and code up our gradients!
| Python/ML_DL/DL/Neural-Networks-Demystified-master/Part 3 Gradient Descent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import logging
import argparse
import numpy as np
import svgwrite
import sys
import drawing
import lyrics
from rnn import rnn
import pandas as pd
from wand.api import library
import wand.color
from wand.image import Image
import cv2
import re
class Hand(object):
def __init__(self):
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
self.nn = rnn(
log_dir='logs',
checkpoint_dir='checkpoints',
prediction_dir='predictions',
learning_rates=[.0001, .00005, .00002],
batch_sizes=[32, 64, 64],
patiences=[1500, 1000, 500],
beta1_decays=[.9, .9, .9],
validation_batch_size=32,
optimizer='rms',
num_training_steps=100000,
warm_start_init_step=17900,
regularization_constant=0.0,
keep_prob=1.0,
enable_parameter_averaging=False,
min_steps_to_checkpoint=2000,
log_interval=20,
logging_level=logging.CRITICAL,
grad_clip=10,
lstm_size=400,
output_mixture_components=20,
attention_mixture_components=10
)
self.nn.restore()
def write(self, filename, lines, biases=None, styles=None, stroke_colors=None, stroke_widths=None):
valid_char_set = set(drawing.alphabet)
#valid_char_set = {'.', ')', 'm', '1', 'M', 'S', 'B', 'U', 'p', 'Y', 'w', ':', '9', 'Z', '7', 'c', 'a', 'E', 'h', '\x00', 'e', '5', 'O', 'u', '3', 'v', 'A', 's', 'j', 'o', 'z', "'", 't', ',', 'P', 'g', ';', 'F', 'r', 'X', '6', 'T', 'N', '8', 'k', 'J', 'l', 'G', 'W', 'd', 'n', '4', 'q', '-', '0', '(', 'y', 'R', 'K', 'L', ' ', 'C', 'I', 'b', '/', 'D', 'H', 'Q', 'x', 'f', 'V', '2', 'i'}
#valid_char_set = {'\x00', 'W', 'N', 'u', 'Y', 'b', '0', 'B', 'J', 'c', 'n', '-', 'D', 'K', 'g', '7', '8', 'V', '9', ':', 'O', 'P', 'S', 'd', 'v', 'Z', 'M', 'f', ',', 'R', 'U', ')', '"', 'e', 'j', '2', ';', '5', '!', 's', 'z', '4', 'G', 'y', '3', 'k', 'm', 'i', "'", ' ', 'h', 'H', '.', '1', 'C', 'E', 'I', 'A', 'p', 'w', '(', 'l', 'r', 'o', 'L', '#', 'a', 't', 'x', 'T', 'q', 'F', '6'}
#print("set",valid_char_set)
for line_num, line in enumerate(lines):
if len(line) > 75:
raise ValueError(
(
"Each line must be at most 75 characters. "
"Line {} contains {}"
).format(line_num, len(line))
)
for char in line:
if char not in valid_char_set:
raise ValueError(
(
"Invalid character {} detected in line {}. "
"Valid character set is {}"
).format(char, line_num, valid_char_set)
)
strokes = self._sample(lines, biases=biases, styles=styles)
self._draw(strokes, lines, filename, line_num=line_num,stroke_colors=stroke_colors, stroke_widths=stroke_widths)
def _sample(self, lines, biases=None, styles=None):
num_samples = len(lines)
max_tsteps = 40*max([len(i) for i in lines])
biases = biases if biases is not None else [0.5]*num_samples
x_prime = np.zeros([num_samples, 1200, 3])
x_prime_len = np.zeros([num_samples])
chars = np.zeros([num_samples, 120])
chars_len = np.zeros([num_samples])
if styles is not None:
for i, (cs, style) in enumerate(zip(lines, styles)):
x_p = np.load('styles/style-{}-strokes.npy'.format(style))
c_p = np.load('styles/style-{}-chars.npy'.format(style)).tostring().decode('utf-8')
c_p = str(c_p) + " " + cs
c_p = drawing.encode_ascii(c_p)
c_p = np.array(c_p)
x_prime[i, :len(x_p), :] = x_p
x_prime_len[i] = len(x_p)
chars[i, :len(c_p)] = c_p
chars_len[i] = len(c_p)
else:
for i in range(num_samples):
encoded = drawing.encode_ascii(lines[i])
chars[i, :len(encoded)] = encoded
chars_len[i] = len(encoded)
[samples] = self.nn.session.run(
[self.nn.sampled_sequence],
feed_dict={
self.nn.prime: styles is not None,
self.nn.x_prime: x_prime,
self.nn.x_prime_len: x_prime_len,
self.nn.num_samples: num_samples,
self.nn.sample_tsteps: max_tsteps,
self.nn.c: chars,
self.nn.c_len: chars_len,
self.nn.bias: biases
}
)
samples = [sample[~np.all(sample == 0.0, axis=1)] for sample in samples]
return samples
def _draw(self, strokes, lines, filename, line_num,stroke_colors=None, stroke_widths=None):
stroke_colors = stroke_colors or ['black']*4
stroke_widths = stroke_widths or [2]*4
i=1
#i+=1
img_large = cv2.imread("templates/large1.jpg")
l_img = cv2.imread("large1.jpg",0)
for offsets, line, color, width in zip(strokes, lines, stroke_colors, stroke_widths):
svgfil = 'p' + str(i) + '.svg'
if i == 1:
line_height = 30
view_width = 200
dwg1 = svgwrite.Drawing(filename=svgfil)
dwg = dwg1
if i == 2:
line_height = 30
view_width = 300
dwg2 = svgwrite.Drawing(filename=svgfil)
dwg = dwg2
if i == 3:
line_height = 60
view_width = 450
dwg3 = svgwrite.Drawing(filename=svgfil)
dwg = dwg3
if i == 4:
line_height = 60
view_width = 450
dwg4 = svgwrite.Drawing(filename=svgfil)
dwg = dwg4
#line_height = 50
#view_width = 300
view_height = line_height*2
dwg.viewbox(width=view_width, height=view_height)
dwg.add(dwg.rect(insert=(0, 0), size=(view_width, view_height), fill='white'))
initial_coord = np.array([0, -(3*line_height / 4)])
if not line:
initial_coord[1] -= line_height
continue
offsets[:, :2] *= 1.1
strokes = drawing.offsets_to_coords(offsets)
strokes = drawing.denoise(strokes)
strokes[:, :2] = drawing.align(strokes[:, :2])
strokes[:, 1] *= -1
strokes[:, :2] -= strokes[:, :2].min() + initial_coord
strokes[:, 0] += (view_width - strokes[:, 0].max()) / 2
prev_eos = 1.0
p = "M{},{} ".format(0, 0)
for x, y, eos in zip(*strokes.T):
p += '{}{},{} '.format('M' if prev_eos == 1.0 else 'L', x, y)
prev_eos = eos
if i==1:
path1 = svgwrite.path.Path(p)
path1 = path1.stroke(color=color, width=width, linecap='round').fill("none")
dwg.add(path1)
if i==2:
path2 = svgwrite.path.Path(p)
path2 = path2.stroke(color=color, width=width, linecap='round').fill("none")
dwg.add(path2)
if i==3:
path3 = svgwrite.path.Path(p)
path3 = path3.stroke(color=color, width=width, linecap='round').fill("none")
dwg.add(path3)
if i==4:
path4 = svgwrite.path.Path(p)
path4 = path4.stroke(color=color, width=width, linecap='round').fill("none")
dwg.add(path4)
#initial_coord[1] -= line_height
dwg.save()
with Image(filename=svgfil, format='svg') as img:
img.format='png'
img.save(filename="s.png")
s_img = cv2.imread("s.png",0)
#control placement of text on the form
if i == 1:
x_offset=135
y_offset=50
if i==2:
x_offset=155
y_offset=110
if i==3:
x_offset=150
y_offset=300
if i==4:
x_offset=150
y_offset=390
l_img[y_offset:y_offset+s_img.shape[0], x_offset:x_offset+s_img.shape[1]] = s_img
outfile = "output/" + filename
i=i+1
cv2.imwrite(outfile,l_img)
s_img = np.ones(s_img.shape) *int(255)
cv2.imwrite("s.png",s_img)
# +
df = pd.read_csv('final100.csv')
# -
medlines = df['Drug Name']
# +
medlines.replace('_',' ',regex=True,inplace=True)
medlines.replace('Z','z',regex=True,inplace=True)
medlines.replace('X','x',regex=True,inplace=True)
medlines.replace('Q','q',regex=True,inplace=True)
medlines.replace('%',' ',regex=True,inplace=True)
medlines.replace('#',' ',regex=True,inplace=True)
medlines.replace('&',' ',regex=True,inplace=True)
medlines.replace('@',' ',regex=True,inplace=True)
names = df['first_name'] + " " + df['last_name']
# -
filenams = "Prescription" + df['Seq'].astype(int).astype(str) + ".png"
# +
hand = Hand()
for idx,drugs in enumerate(medlines):
lines = []
lines.append(names[idx])
lines.append(df['Address'][idx])
lines.append(drugs[:40])
lines.append(drugs[40:])
biases = [df['Biases'][idx] for i in lines]
styles = [df['Styles'][idx] for i in lines]
#Call write function to generate hanwritings.
hand.write(
filename=filenams[idx],
lines=lines,
biases=biases,
styles=styles)
# -
| demo-hanwriting-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jupyter Notebooks Inkscape Development.
import inkex
inkex.Effect
# +
# inkex.EffectExtension?
# -
inkex.Effect()
inkex.Effect()
e = inkex.Effect()
e.add_option
e.add_arguments
| inkex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Example B.1 Level Breadth-first Search
#
# Examples come from http://people.eecs.berkeley.edu/~aydin/GraphBLAS_API_C_v13.pdf
import numpy as np
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
import graphblas as gb
from graphblas import lib, ffi, Matrix, Vector, Scalar
from graphblas.base import NULL
from graphblas import dtypes
from graphblas import descriptor
from graphblas import unary, binary, monoid, semiring
from graphblas import io as gio
# Create initial data objects
edges = [
[3, 0, 3, 5, 6, 0, 6, 1, 6, 2, 4, 1],
[0, 1, 2, 2, 2, 3, 3, 4, 4, 5, 5, 6],
]
A = Matrix.from_values(edges[0], edges[1], [True for _ in edges[0]])
s = 1
A
gio.draw(A)
# ### level breadth-first search (BFS) in GraphBLAS
# ```
# 1 #include <stdlib.h>
# 2 #include <stdio.h>
# 3 #include <stdint.h>
# 4 #include <stdbool.h>
# 5 #include ”GraphBLAS.h”
# 6
# 7 /*
# 8 * Given a boolean n x n adjacency matrix A and a source vertex s, performs a BFS traversal
# 9 * of the graph and sets v[i] to the level in which vertex i is visited ( v[s] == 1 ).
# 10 * If i is not reachable from s, then v[i] = 0. ( Vector v should be empty on input. )
# 11 */
# 12 GrB_Info BFS( GrB Vector *v , GrB Matrix A, GrB Index s )
# 13 {
# 14 GrB_Index n;
# 15 GrB_Matrix_nrows(&n, A); // n = # of rows of A
# 16
# 17 GrB_Vector_new(v, GrB_INT32, n); // Vector<int32_t> v(n)
# 18
# 19 GrB_Vector q; // vertices visited in each level
# 20 GrB_Vector_new(&q ,GrB_BOOL, n); // Vector<bool> q (n )
# 21 GrB_Vector_setElement(q , ( bool ) true , s ) ; // q[s] = true , false everywhere else
# 22
# 23 /*
# 24 * BFS traversal and label the vertices.
# 25 */
# 26 int32_t d = 0 ; // d = level in BFS traversal
# 27 bool succ = false ; // succ == true when some successor found
# 28 do {
# 29 ++d ; // next level ( startwith 1)
# 30 GrB_assign (*v, q, GrB_NULL, d, GrB_ALL, n, GrB_NULL ) ; // v[q] = d
# 31 GrB_vxm(q, *v, GrB_NULL, GrB_LOR_LAND_SEMIRING_BOOL,
# 32 q, A, GrB_DESC_RC); // q[!v] = q ||.&& A ; finds all the
# 33 // unvisited successors from current q
# 34 GrB_reduce(&succ, GrB_NULL, GrB_LOR_MONOID_BOOL,
# 35 q, GrB_NULL ) ; // succ = || ( q )
# 36 } while ( succ ) ; // if there is no successor in q, we are done.
# 37
# 38 GrB_free (&q) ; // q vector no longer needed
# 39
# 40 return GrB_SUCCESS ;
# 41 }
# ```
# ## Python implementation
n = A.nrows
v = Vector(dtypes.INT32, n)
q = Vector(bool, n)
q[s] << True
succ = Scalar(bool)
d = 0 # level in BFS traversal
while True:
d += 1
# For the frontier, assign the depth level
v[:](mask=q.V) << d
# Compute the next frontier, masking out anything already assigned
q(~v.S, replace=True) << q.vxm(A, semiring.lor_land)
# If next frontier is empty, we're done
succ << q.reduce(monoid.lor, allow_empty=False)
if not succ:
break
v
# Let's Step thru each loop to watch the action unfold
# Only run this cell once -- it initializes things
v.clear()
q.clear()
q[s] << True
d = 0
d += 1
# For the frontier, assign the depth level
v[:](mask=q.V) << d
v
# Compute the next frontier, masking out anything already assigned
q(~v.S, replace=True) << q.vxm(A, semiring.lor_land)
q
# These are the next layer of the BFS, prep'd for the next iteration
succ << q.reduce(monoid.lor, allow_empty=False)
print("Continue" if succ else "Done")
| notebooks/Example B.1 -- Level BFS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/m-murphy/sketch-to-diagram/blob/master/generate_model.ipynb)
# + id="fDwtKlSwmUUz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ac08d45a-dd37-4297-fd7e-2a1b3848b6ec"
import os
import glob
import numpy as np
from tensorflow.keras import layers
from tensorflow import keras
import tensorflow as tf
import urllib.request
# !rm -r data
# !mkdir data
classes = ['circle', 'square', 'line', 'star', 'triangle', 'hexagon']
# + id="mlGjGp6ynuc7" colab_type="code" colab={}
def download():
base = 'https://storage.googleapis.com/quickdraw_dataset/full/numpy_bitmap/'
for c in classes:
url = f"{base}{c}.npy"
print(url)
urllib.request.urlretrieve(url, f'data/{c}.npy')
def load_data(root, vfold_ratio=0.2, max_items_per_class=1000000):
all_files = glob.glob(os.path.join(root, "*.npy"))
x = np.empty([0, 784])
y = np.empty([0])
class_names = []
for idx, file in enumerate(all_files):
data = np.load(file)
data = data[0: max_items_per_class, :]
labels = np.full(data.shape[0], idx)
x = np.concatenate((x, data), axis=0)
y = np.append(y, labels)
class_name, ext = os.path.splitext(os.path.basename(file))
class_names.append(class_name)
data = None
labels = None
permutation = np.random.permutation(y.shape[0])
x = x[permutation, :]
y = y[permutation]
vfold_size = int(x.shape[0]/100 * (vfold_ratio * 100))
x_test = x[0 : vfold_size, :]
y_test = y[0 : vfold_size]
x_train = x[vfold_size : x.shape[0], :]
y_train = y[vfold_size : y.shape[0]]
return x_train, y_train, x_test, y_test, class_names
# + id="ZyCdGwv0nlwO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="12828c34-9d71-4745-b7f6-196bfd148fc5"
download()
# + id="6IlPxWInrKh4" colab_type="code" colab={}
x_train, y_train, x_test, y_test, class_names = load_data('data')
num_classes = len(class_names)
image_size = 28
# + id="d4x7DeWcrji8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="712346b5-3e25-44d8-ba5b-6557260133f1"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, len(x_train))
plt.imshow(x_train[idx].reshape(28, 28))
print(class_names[int(y_train[idx].item())])
# + id="xc2zoLfxsoIS" colab_type="code" colab={}
x_train = x_train.reshape(x_train.shape[0], image_size, image_size, 1).astype('float32')
x_test = x_test.reshape(x_test.shape[0], image_size, image_size, 1).astype('float32')
x_train /= 255.0
x_test /= 255.0
# Convert class vectors to class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + id="DJDT7FjfsrVc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="6bed32a7-3a36-4fee-ee45-c8da73ddc742"
model = keras.Sequential()
model.add(layers.Convolution2D(32, (3, 3),
padding='same',
input_shape=x_train.shape[1:], activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
# model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Dropout(0.25))
# model.add(layers.Flatten())
# model.add(layers.Dense(128, activation='relu'))
# model.add(layers.Dropout(0.5))
# model.add(layers.Dense(num_classes, activation='softmax'))
model.add(layers.Convolution2D(64, (3, 3), padding='same', activation= 'relu'))
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Convolution2D(128, (3, 3), padding='same', activation= 'relu'))
model.add(layers.MaxPooling2D(pool_size =(2,2)))
model.add(layers.Dropout(0.25))
model.add(layers.Flatten())
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense(num_classes, activation='softmax'))
# Train model
adam = keras.optimizers.Adam()
model.compile(loss='categorical_crossentropy',
optimizer=adam,
metrics=['accuracy'])
print(model.summary())
tf.test.gpu_device_name()
# + id="g4IbduuMsyMT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="3e222061-a870-4064-8e7c-894f30a9c481"
model.fit(x = x_train, y = y_train, validation_split=0.1, batch_size = 128, verbose=2, epochs=10)
# + id="7kb6TFxutizt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4a2ec4d4-e3ae-49b6-cd75-7a613f7bc184"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test accuarcy: {:0.2f}%'.format(score[1] * 100))
# + id="--UCcr-_t_Az" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="516ddba8-d899-4787-b259-a04ed552eeb8"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, len(x_test))
img = x_test[idx]
plt.imshow(img.squeeze())
pred = model.predict(np.expand_dims(img, axis=0))[0]
print((-pred).argsort()[:6])
ind = (-pred).argsort()[:6]
latex = [class_names[x] for x in ind]
print(latex)
# + id="rSK595ovuF1N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1802ecda-e035-4d58-d409-a7c0caf3611d"
import json
with open('class_names.json', 'w') as file_handler:
class_name_dict = dict(enumerate(class_names))
print(class_name_dict)
file_handler.write(json.dumps(class_name_dict))
# for item in class_names:
# file_handler.write("{}\n".format(item))
# + id="gw9yZV1luI3N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 578} outputId="dd669e5d-2a59-46dc-b63a-418328c6c9bd"
# !pip install tensorflowjs
# + id="JOXpoKo0uPpk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="345f5e56-393b-4589-eda9-e9a25c35bd86"
model.save('keras.h5')
# !rm -r model
# !mkdir model
# !tensorflowjs_converter --input_format keras keras.h5 model/
# !cp class_names.json model/class_names.json
# !cp keras.h5 model/keras.h5
# !zip -r model.zip model
# + id="mPdH7932uoYh" colab_type="code" colab={}
from google.colab import files
files.download('model.zip')
| generate_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from scipy.stats import randint as sp_randint
# sklearn import
from sklearn.pipeline import Pipeline
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor, AdaBoostRegressor
from sklearn.model_selection import RandomizedSearchCV
from sklearn.metrics import make_scorer, r2_score
# my module imports
from optimalcodon.projects.rnastability.dataprocessing import get_data, general_preprocesing_pipeline
from optimalcodon.projects.rnastability import modelevaluation
# -
(train_x, train_y), (test_x, test_y) = get_data("../19-04-30-EDA/results_data/")
# +
# pre-process Pipeline
preprocessing = general_preprocesing_pipeline(train_x)
preprocessing.fit(train_x)
train_x_transformed = preprocessing.transform(train_x)
# -
# ***
# ## Decision Tree Regressor
np.arange(10, 25)
# +
tree_reg = DecisionTreeRegressor()
tree_grid = {
'min_samples_split': np.linspace(0.001, .03, 10),
'max_features': [None],
'splitter': ['best'],
'max_depth': np.arange(10, 25)
}
tree_search = modelevaluation.gridsearch(tree_reg, tree_grid, train_x_transformed, train_y, cores=15)
# -
# I will use the best parameter of the decision tree to train a random forest.
#
# ***
#
# ## Random Forest
# +
rf_reg = RandomForestRegressor(max_depth=14, n_jobs=8)
rf_grid = {
'n_estimators': np.arange(1700, 2000, 50)
}
rf_search = modelevaluation.gridsearch(rf_reg, rf_grid, train_x_transformed, train_y, cores=3)
# -
modelevaluation.eval_models({'rf': rf_search.best_estimator_}, preprocessing, test_x, test_y).to_csv("res3.csv")
# ***
#
# ## ADA BOOST
ada_reg = AdaBoostRegressor(DecisionTreeRegressor(max_depth=4))
ada_grid = {
'n_estimators': np.arange(10, 300, 100)
}
ada_search = modelevaluation.gridsearch(ada_reg, ada_grid, train_x_transformed, train_y, cores=17)
models = {
'decision tree': tree_search.best_estimator_,
'AdaBoost': ada_search.best_estimator_,
'random forest': rf_search.best_estimator_.set_params(n_jobs=2) # set params for cross validation
}
modelevaluation.eval_models(models, preprocessing, test_x, test_y).to_csv("results_data/val_non-Treemodels.csv")
modelevaluation.crossvalidation(models, train_x_transformed, train_y).to_csv('results_data/cv_Tree-models.csv', index=False)
| results/19-04-30-PredictiveModelDecayAllSpecies/19-05-01-TrainModels/03-TreeModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''PythonData'': conda)'
# language: python
# name: python37564bitpythondatacondaadf2dc53d8344d2f91c5b97fe5b73276
# ---
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
import copy
# +
chinese_data = "Data/chinese.csv"
indian_data = "Data/indian.csv"
french_data = "Data/french.csv"
italian_data = "Data/italian.csv"
mexican_data = "Data/mexican.csv"
southern_data = "Data/southern.csv"
greek_data = "Data/greek.csv"
nordic_data = "Data/nordic.csv"
caribbean_data = "Data/caribbean.csv"
middle_eastern_data = "Data/middle eastern.csv"
korean_data = "Data/korean.csv"
chinese_df = pd.read_csv(chinese_data)
indian_df = pd.read_csv(indian_data)
french_df = pd.read_csv(french_data)
italian_df = pd.read_csv(italian_data)
mexican_df = pd.read_csv(mexican_data)
southern_df = pd.read_csv(southern_data)
greek_df = pd.read_csv(greek_data)
nordic_df = pd.read_csv(nordic_data)
caribbean_df = pd.read_csv(caribbean_data)
middle_eastern_df = pd.read_csv(middle_eastern_data)
korean_df = pd.read_csv(korean_data)
# -
chinese_df = chinese_df.append(indian_df, ignore_index=True)
chinese_df = chinese_df.append(french_df, ignore_index=True)
chinese_df = chinese_df.append(italian_df, ignore_index=True)
chinese_df = chinese_df.append(mexican_df, ignore_index=True)
chinese_df = chinese_df.append(southern_df, ignore_index=True)
chinese_df = chinese_df.append(greek_df, ignore_index=True)
chinese_df = chinese_df.append(nordic_df, ignore_index=True)
chinese_df = chinese_df.append(caribbean_df, ignore_index=True)
chinese_df = chinese_df.append(middle_eastern_df, ignore_index=True)
chinese_df = chinese_df.append(korean_df, ignore_index=True)
chinese_df['instructions'].replace('', np.nan, inplace=True)
chinese_df.dropna(subset=['instructions'], inplace=True)
chinese_df
chinese_df.to_csv("Data/merged.csv", index=False, header=True)
#Import data into mongoDB
import pymongo
import csv
import json
from pymongo import MongoClient
client = MongoClient()
client = MongoClient('localhost', 27017)
db=client['foodtopia']
colmanager=db['food']
colmanager.drop()
colmanager.insert_many(chinese_df.to_dict('records'))
| foodtopia/templates/Merge_data_import_mongoDB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import time
from pybel.struct import (
get_subgraph_by_induction, expand_upstream_causal, get_upstream_causal_subgraph, has_protein_modification,
)
import hbp_knowledge
from pybel.dsl import Protein
from pybel_jupyter import to_jupyter
from pybel.constants import RELATION, HAS_VARIANT
# -
hbp_knowledge.VERSION
print(sys.version)
print(time.asctime())
graph = hbp_knowledge.get_graph()
graph.summarize()
# List all nodes that have to do with ``tau``.
nodes = []
for node in graph.nodes():
if 'name' in node and 'tau' in node.name.lower():
print(node)
nodes.append(node)
# Get post-translational modifications of the human Tau protein
mapt = Protein('HGNC', 'MAPT')
nodes = [
v
for u, v, key, data in graph.edges(keys=True, data=True)
if u == mapt and data[RELATION] == HAS_VARIANT and has_protein_modification(v)
]
nodes
# Induce a subgraph upstream of all PTMs of the human Tau protein.
mapt_ptm_controller_graph = get_upstream_causal_subgraph(graph, nodes)
mapt_ptm_controller_graph.name = 'MAPT Post-Translational Modification Controllers'
mapt_ptm_controller_graph.version = hbp_knowledge.VERSION
mapt_ptm_controller_graph.summarize()
to_jupyter(x)
# Get a list of controller proteins.
# +
controllers = [
node.name
for node in mapt_ptm_controller_graph
if isinstance(node, Protein) and 'MAPT' != node.name and 'HGNC'== node.namespace.upper()
]
print(controllers)
# -
# # TODO
#
#
# ## INDRA Enrichment
#
# 1. Generate query to find statements that have MAPT as an object and use a conversion statement
# 2. Get bigger list of
#
# ## Visualization of Relevant Chemical Space
#
# 1. Load ExCAPE-DB
# 2. Extract relationships from ExCAPE-DB related to these proteins
# 3. Calculate chemical descriptors
# 4. Show a t-SNE plot or related 2D depiction of chemical space
| notebooks/Get Controllers of Tau Modication.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Overview of Visualization for Deep Learning
# ## ComputeFest
# ### Winter, 2020
# <br><br>
# <img src="fig/logos.jpg" style="height:150px;">
# + slideshow={"slide_type": "skip"}
# Includes the necessary libraries
import numpy as np
import matplotlib.pylab as plt
import os
# for implementing neural network models
from keras import backend as K
from keras import layers
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense
from keras.callbacks import Callback, ModelCheckpoint, History
from keras.wrappers.scikit_learn import KerasClassifier
from keras.utils import np_utils
from keras.optimizers import SGD, Adam
# for data preprocessing and non-neural network machine learning models
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_moons, make_classification, make_circles
from sklearn.preprocessing import LabelEncoder, MinMaxScaler, scale
from sklearn.metrics import confusion_matrix, roc_auc_score
# %matplotlib inline
random_state = 0
random = np.random.RandomState(random_state)
# + slideshow={"slide_type": "skip"}
def scatter_plot_data(x, y, ax):
'''
scatter_plot_data scatter plots the patient data. A point in the plot is colored 'red' if positive outcome
and blue otherwise.
input:
x - a numpy array of size N x 2, each row is a patient, each column is a biomarker
y - a numpy array of length N, each entry is either 0 (no cancer) or 1 (cancerous)
ax - axis to plot on
returns:
ax - the axis with the scatter plot
'''
ax.scatter(x[y == 1, 0], x[y == 1, 1], alpha=0.9, c='red', label='class 1')
ax.scatter(x[y == 0, 0], x[y == 0, 1], alpha=0.9, c='blue', label='class 0')
ax.set_xlim((min(x[:, 0]) - 0.5, max(x[:, 0]) + 0.5))
ax.set_ylim((min(x[:, 1]) - 0.5, max(x[:, 1]) + 0.5))
ax.set_xlabel('marker 1')
ax.set_ylabel('marker 2')
ax.legend(loc='best')
return ax
def plot_decision_boundary(x, y, model, ax, poly_degree=1):
'''
plot_decision_boundary plots the training data and the decision boundary of the classifier.
input:
x - a numpy array of size N x 2, each row is a patient, each column is a biomarker
y - a numpy array of length N, each entry is either 0 (negative outcome) or 1 (positive outcome)
model - the 'sklearn' classification model
ax - axis to plot on
poly_degree - the degree of polynomial features used to fit the model
returns:
ax - the axis with the scatter plot
'''
# Plot data
ax.scatter(x[y == 1, 0], x[y == 1, 1], alpha=0.2, c='red', label='class 1')
ax.scatter(x[y == 0, 0], x[y == 0, 1], alpha=0.2, c='blue', label='class 0')
# Create mesh
x0_min = min(x[:, 0])
x0_max = max(x[:, 0])
x1_min = min(x[:, 1])
x1_max = max(x[:, 1])
interval = np.arange(min(x0_min, x1_min), max(x0_max, x1_max), 0.1)
n = np.size(interval)
x1, x2 = np.meshgrid(interval, interval)
x1 = x1.reshape(-1, 1)
x2 = x2.reshape(-1, 1)
xx = np.concatenate((x1, x2), axis=1)
# Predict on mesh points
if(poly_degree > 1):
polynomial_features = PolynomialFeatures(degree=poly_degree)
xx = polynomial_features.fit_transform(xx)
yy = model.predict(xx)
yy = yy.reshape((n, n))
# Plot decision surface
x1 = x1.reshape(n, n)
x2 = x2.reshape(n, n)
ax.contourf(x1, x2, yy, alpha=0.1, cmap='bwr')
ax.contour(x1, x2, yy, colors='black', linewidths=0.2)
ax.set_xlim((x0_min, x0_max))
ax.set_ylim((x1_min, x1_max))
ax.set_xlabel('marker 1')
ax.set_ylabel('marker 2')
ax.legend(loc='best')
return ax
# + [markdown] slideshow={"slide_type": "slide"}
# ## Outline
# 1. What is this workshop about?
# 2. A Motivating Real-world Example
# 3. Taxonomy of Deep Learning Visualization Literature
# 4. Hands-on Exercise: Saliency Maps for Model Diagnostic
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is This Workshop About?
#
# - **What:** those of you coming from data intensive disciplines are fluent in the techniques of data visualization - this is an well-established, deep and interdisciplinary field. Today we focus on visualizations that probe the properties of machine learning models built for data sets. <br><br>
#
# - **Who:** this workshop is intended for folks who have working familiarity with some machine models (neural network based or not). This introductory lecture will give a brief guided tour of the literature. We will define what is a neural networks model and given motivatations for model visualization.
#
# The following workshop will guide you through implementation of a number of these visualization techniques.<br><br>
#
# - **How:** notes for this lecture are online and are best used as a conceptual foundation for the workshop, as well as a starting point for exploring this body of literature.
# + [markdown] slideshow={"slide_type": "slide"}
# # A Motivating Real-world Example
# + [markdown] slideshow={"slide_type": "slide"}
# ## Predicting Positive Outcomes for IVF Patients
#
# <img src="fig/ivf.png" style="height:300px;">
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Simple Model: Logistic Regression
# We build a simple predictive model: **logistic regression**. We model the probability of an patient $\mathbf{x} \in \mathbb{R}^{\text{input}}$ having a positive outcome (encoded by $y=1$) as a function of its distance from a hyperplane parametrized by $\mathbf{w}$ that separates the outcome groups in the input space.
# <img src="./fig/fig0.png" style='height:300px;'>
#
# That is, we model $p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}(\mathbf{w}^\top \mathbf{x})$. Where $g(\mathbf{x}) = \mathbf{w}^\top \mathbf{x}=0$ is the equation of the decision boundary.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualizing the Top Predictors
#
# We sort the weights of the logistic regression model
#
# $$p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}(\mathbf{w}^\top \mathbf{x})$$
#
# and visualize the weights and the corresponding data attribute. Why is this a good idea?
#
# <img src="./fig/features.png" style='height:500px;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interpreting the Top Predictors
#
# We visualize the distribution of the predictor `E2_max` for both postive and negative outcome groups. What can we conclude about this predictor?
# <img src="./fig/feature_1.png" style='height:300px;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interpreting the Top Predictors
# We visualize the distribution of the predictor `Trigger_Med_nan` for both postive and negative outcome groups. What can we conclude about this predictor?
#
# <img src="./fig/feature_2.png" style='height:300px;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Lessons for Visualization
#
# Choosing/designing machine learning visualization requires that we think about:
#
# 1. ***Why and for whom*** **to visualize**: for example
# - are we visualizing to diagnose problems with our models?
# - are we visualizing to interpret our models with clinical meaningfulness?<br><br>
#
# 2. ***What and how*** **to visualize**: for example
# - do we visualize decision boundaries, weights of our model, and or distributional differences in the data?
#
# **Note:** what is possible to visualize depends very much on the internal mechanics of our model and the data!
# + [markdown] slideshow={"slide_type": "slide"}
# ## What If the Decision Boundary is Not Linear?
#
# How would you parametrize a ellipitical decision boundary?
#
# <img src="./fig/fig1.png" style='height:300px;'>
#
# We can say that the decision boundary is given by a ***quadratic function*** of the input:
#
# $$
# g(\mathbf{x}) = w_1x^2_1 + w_2x^2_2 + w_3 = 0
# $$
#
# We can fit such a decision boundary using logistic regression with degree 2 polynomial features.
# + [markdown] slideshow={"slide_type": "slide"}
# ## How would you parametrize an arbitrary complex decision boundary?
#
# <img src="./fig/fig2.png" style='height:300px;'>
#
# It's not easy to think of a function $g(\mathbf{x})$ can capture this decision boundary. Also, assuming an exact form for $g$ is restrictive.
#
# **GOAL:** Build increasingly good approximations, $\widehat{g}$, of the "true deicision boundary" $g$ by composing simple functions. This $\widehat{g}$ is essentially a **neural network**.
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is a Neural Network?
#
# **Goal:** build a complex function $\widehat{g}$ by composing simple functions.
#
# For example, let the following picture represents the approximation $\widehat{g}(\mathbf{x}) = f\left(\sum_{i}w_ix_i\right)$, where $f$ is a non-linear transform:
#
# <img src="./fig/fig4.png" style='height:200px;'>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Neural Networks as Function Approximators
#
# Then we can define the function $\widehat{g}$ with a graphical schema representing a complex series of compositions and sums of the form, $f\left(\sum_{i}w_ix_i\right)$
#
# <img src="./fig/fig5.png" style='height:300px;'>
#
# This is a ***neural network***. We denote the weights of the neural network collectively by $\mathbf{W}$.
# The non-linear function $f$ is called the ***activation function***. Nodes in this diagram that are not representing input or output are called ***hidden nodes***. This diagram, along with the choice of activation function $f$, that defines $\widehat{g}$ is called the ***architecture***.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Training Neural Networks
#
# We ***train*** a neural network classifier,
#
# $$p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}\left(\widehat{g}_{\mathbf{W}}(\mathbf{x})\right)$$
#
# by finding weights $\mathbf{W}$ that parametrizes the function $\widehat{g}_{\mathbf{W}}$ that "best fits the data" - e.g. best separate the classes.
#
# Typically, we optimize the fit of $\widehat{g}_{\mathbf{W}}$ to the data **incrementally** and **greedily** in a process called ***gradient descent***.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What to Visualize for Neural Network Models?
# For logistic regression, $p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}(\mathbf{w}^\top \mathbf{x})$, we were able to interrogate the model by printing out the weights of the model.
#
# For a neural network classifier, $p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}\left(\widehat{g}_{\mathbf{W}}(\mathbf{x})\right)$, would it be helpful to print out all the weights?
#
# <img src="./fig/fig5.png" style='height:300px;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Weight Space Versus Function Space
#
# While it's convienient to build up a complex function by composing simple ones, understanding the impact of each weight on the outcome is now much more difficult.
#
# In fact, the relationship between the space of weights of a neural network and the function the network represents is extremely complicated:
# 1. the same function may be represented by two very different set of weights for the same architecture<br><br>
# 2. the architecture may be overly expressive - it can express the function $\widehat{g}$ using a subset of the weights and hidden nodes (i.e. the trained model can have weights that are zero or nodes that contribute little to the computation).
#
# **Question:** are there more global/heuristic properties of these models that we can visualize?
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Non-linear Classification Problem
# + [markdown] slideshow={"slide_type": "skip"}
# #### Make a toy data set that cannot be classified by a linear decision boundary.
#
# We make use of the `.make_*()` functions provided by the `sklearn` libary, specifying the noise level (how much the two classes overlap) and the number of samples in our data set. By default, the classes generated will be balanced (i.e. there will be roughly an equal number of instances in both classes).
# + slideshow={"slide_type": "skip"}
# generate a toy classification data set with non-linear decision boundary
X, Y = make_moons(noise=0.1, random_state=random_state, n_samples=1000)
# we scale the data so that the magnitude along each input dimension are roughly equal
X = scale(X)
# split the data set into testing and training, with 30% for test
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=.3)
# + [markdown] slideshow={"slide_type": "skip"}
# #### Fit a logistic regression model with a linear decision boundary
# We fit an instance of `sklearn`'s `LogisticRegression` model to our data. Recall that 'fitting' the model means finding the coefficients, $\mathbf{w}$, of the linear decision boundary that best fit the data. Here the criteria of fitness will be the likelihood of the training data given the model $\mathbf{w}$. `LogisticRegression` requires that you choose a method for maximizing the likelihood over $\mathbf{w}$ (e.g. `lbfgs`).
# + slideshow={"slide_type": "skip"}
# instantiate a sklearn logistic regression model
logistic = LogisticRegression(solver='lbfgs')
# fit the model to the training data
logistic.fit(X_train, Y_train)
# + [markdown] slideshow={"slide_type": "skip"}
# #### Evaluating the fitted model
#
# To evaluate the mode that we've fitted to the data, we can compute a number of metrics like accuracy (or a confusion matrix, balanced accuracy, F1 etc if the classes are unbalanced). These metrics tell us how well our model does but not why it is able to achieve such performance. When possible, it is still preferable to visualize the data along with the classifier's decision boundary (or surface in higher dimensions) - this gives a nuanced picture that explains the models performance on various metrics.
# + slideshow={"slide_type": "skip"}
# evaluate the accuracy of our classifier on the training and testing data
print('Training accuracy of linear logistic regression:', logistic.score(X_train, Y_train))
print('Testing accuracy of linear logistic regression:', logistic.score(X_test, Y_test))
# + slideshow={"slide_type": "fragment"}
degree_of_polynomial = 1
fig, ax = plt.subplots(1, 2, figsize=(15, 5))
scatter_plot_data(X_train, Y_train, ax[0])
ax[0].set_title('Training Data')
plot_decision_boundary(X_train, Y_train, logistic, ax[1], degree_of_polynomial)
ax[1].set_title('Decision Boundary')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Neural Network Model for Classification
# We train a neural network classifier (with relu activation): $p(y=1 | \mathbf{w}, \mathbf{x}) = \mathrm{sigmoid}\left(\widehat{g}_{\mathbf{W}}(\mathbf{x})\right)$
#
#
# <img src="fig/architecture.jpeg" style="height:350px;">
#
#
# *Generated with: [`http://alexlenail.me/NN-SVG/index.html`](http://alexlenail.me/NN-SVG/index.html)*
# + [markdown] slideshow={"slide_type": "skip"}
# #### Implement the neural network classifier in `keras`
#
# The `python` library `keras` provides convenient and intuitive api for quickly building up deep learning models. Essentially, if you have a graphical representation of the neural network architecture you can translate this graph, layer by layer, into `keras` objects.
# + slideshow={"slide_type": "skip"}
def create_model(input_dim, optimizer):
'''
create_model creates a neural network model for classification in keras.
input:
input_dim - the number of data attributes
optimizer - the method of optimization for training the neural network
returns:
model - the keras neural network model
'''
# create sequential multi-layer perceptron in keras
model = Sequential()
#layer 0 to 1: input to 100 hidden nodes
model.add(Dense(100, input_dim=input_dim, activation='relu',
kernel_initializer='random_uniform',
bias_initializer='zeros'))
#layer 1 to 2: 100 hidden nodes to 2 hidden nodes
model.add(Dense(2, activation='relu',
kernel_initializer='random_uniform',
bias_initializer='zeros'))
#binary classification, one output: 2 hidden nodes to one output node
#since we're doing classification, we apply the sigmoid activation to the
#linear combination of the input coming from the previous hidden layer
model.add(Dense(1, activation='sigmoid',
kernel_initializer='random_uniform',
bias_initializer='zeros'))
# configure the model
model.compile(optimizer=optimizer,
loss='binary_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] slideshow={"slide_type": "skip"}
# #### Fit a neural network classifier
# We fit the neural network model to our data. Recall that 'fitting' the model means finding the network weights, $\mathbf{W}$, parametrizing a non-linear decision boundary that best fit the data. Again, the criteria of fitness here will be the likelihood of the training data given the model $\mathbf{w}$. `keras` requires that you choose a method for iteratively maximizing the likelihood over $\mathbf{w}$ - there are many flavors of gradient descent (e.g. `sgd`, `adam`), these different methods may learn very different models.
# + slideshow={"slide_type": "skip"}
input_dim = X_train.shape[1]
model = create_model(input_dim, 'sgd')
# fit the model to the data and save value of the objective function at each step of training
history = model.fit(X_train, Y_train, batch_size=20, shuffle=True, epochs=500, verbose=0)
# + [markdown] slideshow={"slide_type": "skip"}
# #### Diagnosing potential problems during training
#
# A way to diagnose potential basic issues during training is visualizing the objective function (model fitness) during gradient descent. Since we typically frame maximizing model fitness as minimizing a ***loss function***, we expect to see the loss to go down over iterations of gradient descent.
# + slideshow={"slide_type": "skip"}
# plot the loss function and the evaluation metric over the course of training
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
ax.plot(np.array(history.history['accuracy']), color='blue', label='training accuracy')
ax.plot(np.array(history.history['loss']), color='red', label='training loss')
ax.set_title('Loss and Accuracy During Training')
ax.legend(loc='upper right')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## How Well Does Our Neural Network Classifier Do?
# + slideshow={"slide_type": "fragment"}
# Evaluate the training and testing performance of your model
# Note: you should check both the loss function and your evaluation metric
score = model.evaluate(X_train, Y_train, verbose=0)
print('Train accuracy:', score[1], '\n\n')
# + [markdown] slideshow={"slide_type": "skip"}
# So the neural network classifier is much much more accurate than the logistic regression model with a linear decision boundary. But how exactly is it able to achieve this performance. Again, we want a much more nuanced understanding of the model performance than that which is given by numeric summary metrics.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why is a Neural Network Classifier So Effective?
#
# Visualizing the decision boundary:
# + slideshow={"slide_type": "fragment"}
degree_of_polynomial = 1
fig, ax = plt.subplots(1, 2, figsize=(15, 5))
scatter_plot_data(X_train, Y_train, ax[0])
ax[0].set_title('Training Data')
plot_decision_boundary(X_train, Y_train, model, ax[1], degree_of_polynomial)
ax[1].set_title('Decision Boundary')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why is a Neural Network Classifier So Effective?
#
# Visualizing the output of the last hidden layer.
# + [markdown] slideshow={"slide_type": "skip"}
# Before neural network models became wildly popular in machine learning, a common method for building non-linear classifiers is to first map the data, in the input space $\mathbb{R}^{\text{input}}$, into a 'feature' space $\mathbb{R}^{\text{feature}}$, such that the classes are well-separated in the feature space. Then, a linear classifier can be fitted to the transformed data.
#
# If we ignore the output node of our neural network classifier, we are left with a function, $\mathbb{R}^{2} \to \mathbb{R}^2$, mapping the data from the input space to a 2-dimensional feature space. The transformed data (and in general, the output from a hidden layer in a neural network) is called a ***representation*** of the data.
#
# <img src="fig/architecture.jpeg" style="height:350px;">
#
# Visualizing these representations can often shed light on how and what neural network models learns from the data.
# + slideshow={"slide_type": "skip"}
# get the class probabilities predicted by our MLP on the training set
Y_train_pred = model.predict(X_train)
Y_train_pred = Y_train_pred.reshape((Y_train_pred.shape[0], ))
# get the activations for the last hidden layer in the network
last_hidden_layer = -2
latent_representation = K.function([model.input, K.learning_phase()], [model.layers[last_hidden_layer].output])
activations = latent_representation([X_train, 1.])[0]
# + slideshow={"slide_type": "fragment"}
# plot the latent representation of our training data at the first hidden layer
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
ax.scatter(activations[Y_train_pred >= 0.5, 0], activations[Y_train_pred >= 0.5, 1], color='red', label='Class 1')
ax.scatter(activations[Y_train_pred < 0.5, 0], activations[Y_train_pred < 0.5, 1], color='blue', label='Class 0')
ax.set_title('Toy Classification Dataset Transformed by the NN')
ax.legend()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Two Interpretations of a Neural Network Classifier:
# <table>
# <tr><td><font size="3">A Complex Decision Boundary $g\quad\quad\quad\quad\quad$</font></td>
# <td><font size="3">A Transformation $g_0$ and a linear model $g_1\quad\quad\quad\quad$</font></td>
# <tr><td><img src="fig/decision.png" style="height:350px;"></td>
# <td><img src="fig/architecture2.png" style="height:400px;"></td></tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Is the Model Right for the Right Reasons?
#
# Now that we see that the boosted performance of neural network models come from the fact that they can express complex functions (either capturing decision boundaries or transformations of the data).
#
# Do these visualization help us answer the same questions we asked of the logistic regression model?
#
# - which attributes are most predictive of a positive outcome?<br><br>
# - is the relationship between these attributes and the outcome clinically meaningful?
# + [markdown] slideshow={"slide_type": "slide"}
# ## Lessons for Visualization
# Choosing/designing machine learning visualization requires that we think about:
#
# 1. ***Why and for whom*** **to visualize**: for example
# - are we visualizing to diagnose problems with our models?
# - are we visualizing to interpret our models with clinical meaningfulness?<br><br>
#
# 2. ***What and how*** **to visualize**: for example
# - do we visualize decision boundaries, weights of our model, and or distributional differences in the data?
#
# **Note:** what is possible to visualize depends very much on the internal mechanics of our model and the data!<br><br>
#
# 3. **Deep models present unique challenges for visualization**: we can answer the same questions about the model, but our method of interrogation must change!
# + [markdown] slideshow={"slide_type": "slide"}
# # Taxonomy of Deep Learning Visualization Literature
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why and for Whom
#
# 1. **Interpretability & Explainability:** understand how deep learning models make decisions and what representations they have learned.<br><br>
# 2. **Debugging & Improving Models:** help model developers build and debug their models, with the hope of expediting the iterative experimentation process to ultimately improve performance.<br><br>
# 3. **Teaching Deep Learning Concepts:** educate non-expert users about AI.
#
# *From: [Visual Analytics in Deep Learning: An Interrogative Survey for the Next Frontiers](https://arxiv.org/pdf/1801.06889.pdf)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## TensorBoard
#
# TensorBoard provides a wide array of visualization tools for model diagnostics.
#
# <img src="./fig/tensorboard.png" style='height:500px;'>
#
# *From: [Tensorboard](https://www.tensorflow.org/tensorboard)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Explaining Classifier Decisions in Medical Imaging
#
# Visualization of the evidence for the correct classification of an MRI as positive of presence of disease.
#
# <img src="./fig/medical.png" style='height:450px;'>
#
# *From: [Visualizing Deep Neural Network Decisions: Prediction Difference Analysis](https://arxiv.org/pdf/1702.04595.pdf)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## How and What
#
# **What technical components of neural networks could be visualized?**
# - Computational Graph & Network Architecture
# - Learned Model Parameters: weights, filters
# - Individual Computational Units: activations, gradients
# - Aggregate information: performance metrics
#
# **How can they be insightfully visualized?**
#
# How depends on the type of data and model as well as our specific investigative goal.
# + [markdown] slideshow={"slide_type": "slide"}
# # All Data Types
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activations and Weights
# By visualizing the network weights and activations (the output of hidden nodes) as we train, we can diagnose issues that ultimately impact model performance.
#
# The following visualizes the distribution of activations in two hidden layers over the course of training. What problems do we see?
#
# <img src="./fig/tensor_board.png" style='height:350px;'>
#
# *From: [Tensorboard](https://www.tensorflow.org/tensorboard)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualizing Top Predictors by Input Gradient
# Since the input gradient of an objective function for a trained model indicates which input dimensions has the greatest effect on the model decision at an input $\mathbf{x}$, we can visualize the "top predictors" of outcome for a particular input $\mathbf{x}$.
#
# We can think of this as approximating our neural network model with a linear model locally at an input $\mathbf{x}$ and then interpreting the weights of this linear approximation.
#
# <img src="./fig/features.png" style='height:300px;'>
#
# *From: [How to Explain Individual Classification Decisions](http://www.jmlr.org/papers/volume11/baehrens10a/baehrens10a.pdf)*
# + [markdown] slideshow={"slide_type": "slide"}
# # Image Data
# + [markdown] slideshow={"slide_type": "slide"}
# ## Saliency Maps
#
# When each input dimension is a pixel, we can visualize the input gradients for a particular image as a heat-map, called ***saliency map***. Higher gradient regions representing portions of the image that is most impactful for the model's decision.
#
# <img src="./fig/saliency.png" style='height:450px;'>
#
# *From: [Top-down Visual Saliency Guided by Captions](https://www.groundai.com/project/top-down-visual-saliency-guided-by-captions/2)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Class Maximization
# We can also visualize the image exemplar of a class, according to a trained model. That is, we find image $\mathbf{x}^*$ that maximizes the "chances" of classifying that image as class $c$:
# $
# \mathbf{x}^* = \mathrm{argmax}_\mathbf{x}\; \mathrm{score}_c(\mathbf{x})
# $
#
# <img src="./fig/class_max.png" style='height:450px;'>
#
# *From: [Deep Inside Convolutional Networks](https://arxiv.org/pdf/1312.6034.pdf), [Multifaceted Feature Visualization](https://arxiv.org/pdf/1602.03616.pdf)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Working with Image Data is Challenging
#
# In applications involving images, the first task is often to parse an image into a set of 'features' that are relevant for the task at hand. That is, we prefer not to work with images as a set of pixels.
#
# <img src="./fig/fig8.jpg" style='height:300px;'>
#
# Formally, we want to learn a function $h$ mapping an image $X$ to a set of $K$ ***features*** $[F_1, F_2, \ldots, F_K]$, where each $F_k$ is an image represented as an array. We want to learn a neural network, called a **convolutional neural network**, to represent such a function $h$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Convolutional Layers
# A ***convolutional neural network*** typically consists of feature extracting layers and condensing layers.
#
# The feature extracting layers are called ***convolutional layers***, each node in these layers uses a small fixed set of weights to transform the image in the following way:
# <img src="./fig/fig9.gif" style="width: 500px;" align="center"/>
# This set of fixed weights for each node in the convolutional layer is often called a ***filter***.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Connections to Classical Image Processing
# The term "filter" comes from image processing where one has standard ways to transforms raw images:
#
# <img src="./fig/fig10.png" style="width: 400px;" align="center"/>
# + [markdown] slideshow={"slide_type": "skip"}
# ## What Do Filters Do?
# For example, to blur an image, we can pass an $n\times n$ filter over the image, replacing each pixel with the average value of its neighbours in the $n\times n$ window. The larger the window, the more intense the blurring effect. This corresponds to the Box Blur filter, e.g. $\frac{1}{9}\left(\begin{array}{ccc}1 & 1 & 1\\ 1 & 1 & 1 \\1 & 1 & 1\end{array}\right)$:
#
# <img src="./fig/fig11.png" style="width: 600px;" align="center"/>
#
# In an Gaussian blur, for each pixel, closer neighbors have a stronger effect on the value of the pixel (i.e. we take a weighted average of neighboring pixel values).
#
# + [markdown] slideshow={"slide_type": "skip"}
# ## Convolutional Networks for Image Classification in `keras`
#
# #### Defining image size, filter size etc:
#
# ``` python
# # image shape
# image_shape = (64, 64)
# # Stride size
# stride_size = (2, 2)
# # Pool size
# pool_size = (2, 2)
# # Number of filters
# filters = 2
# # Kernel size
# kernel_size = (5, 5)
# ```
#
# #### The model:
#
# ``` python
# cnn_model = Sequential()
# # feature extraction layer 0: convolution
# cnn_model.add(Conv2D(filters, kernel_size=kernel_size, padding='same',
# activation='tanh',
# input_shape=(image_shape[0], image_shape[1], 1)))
# # feature extraction layer 1: max pooling
# cnn_model.add(MaxPooling2D(pool_size=pool_size, strides=stride_size))
#
# # input to classification layers: flattening
# cnn_model.add(Flatten())
#
# # classification layer 0: dense non-linear transformation
# cnn_model.add(Dense(10, activation='tanh'))
# # classification layer 3: output label probability
# cnn_model.add(Dense(1, activation='sigmoid'))
#
# # Compile model
# cnn_model.compile(optimizer='Adam',
# loss='binary_crossentropy',
# metrics=['accuracy'])
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ## Feature Extraction for Classification
#
# Rather than processing image data with a pre-determined set of filters, we want to learn the filters of a CNN for feature extraction. Our goal is to extract features that best helps us to perform our downstream task (e.g. classification).
#
# **Idea:** We train a CNN for feature extraction and a model (e.g. MLP, decision tree, logistic regression) for classification, *simultaneously* and *end-to-end*.
#
# <img src="./fig/fig13.png" style="width: 800px;" align="center"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## What to Visualize for CNNs?
# The first things to try are:
# 1. visualize the result of applying a learned filter to an image
# 2. visualize the filters themselves:
#
# <table>
# <tr><td><img src="fig/bird.png" style="height:250px;"></td>
# <td><img src="fig/bird_features.png" style="height:250px;"></td>
# <td><img src="fig/bird_filters.png" style="height:250px;"></td></tr>
# </table>
#
# Unfortunately, these simple visualizations don't shed much light on what the model has learned.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activation Maximization: Generating Exemplars
# Rather than visualizing a particular filter or the representation of a particular image at a hidden layer, we can visualize the image that maximize the output and hence impact of that filter or layer. Such an image is an exemplar of the filter or feature that the model has learned.
#
# That is, we find image $\mathbf{x}^*$ that maximizes activation of a filter or a hidden layer (representation) while holding the network weights fixed:
#
# $$
# \mathbf{x}^* = \mathrm{argmax}_\mathbf{x}\; \mathrm{activation}_{\text{$f$ or $l$}}(\mathbf{x}).
# $$
#
# <img src="./fig/activation_max.png" style="width:1000px;" align="center"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualizing Convolutional Features By Activation Maximization
#
#
# <img src="./fig/feature_viz.png" style="width:1000px;" align="center"/>
#
# *From: [Feature Visualization](https://distill.pub/2017/feature-visualization/)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualizing Convolutional Filters By Activation Maximization
#
# <table>
# <tr><td><img src="fig/filters.jpg" style="height:350px;"></td>
# <td><img src="fig/filters2.jpg" style="height:350px;"></td>
# <td><img src="fig/filters3.jpg" style="height:350px;"></td>
# </tr>
# </table>
#
# *From: [DeepViz](https://github.com/yosinski/deep-visualization-toolbox)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interpretation with a Grain of Salt
#
# - Some widely deployed saliency methods are independent of both the data the model was trained on, and the model parameters!
#
# - A transformation with no effect on the model can cause numerous saliency methods to incorrectly attribute!
#
# <img src="fig/saliency_problems.png" style="height:350px;">
#
# *From: [Sanity Checks for Saliency Maps](https://papers.nips.cc/paper/8160-sanity-checks-for-saliency-maps.pdf), [THE (UN)RELIABILITY OF SALIENCY METHODS](https://arxiv.org/pdf/1711.00867.pdf)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example: Saliency Maps for Model Diagnostic
#
# Here is a guided example of using saliency maps to diagnose problems with a neural work classifier (this has not yet been converted to `keras`!).
#
# https://tinyurl.com/w5h54vg
# + [markdown] slideshow={"slide_type": "skip"}
# # Text Data
# + [markdown] slideshow={"slide_type": "skip"}
# ## Representing Textual Data
# Comparing the content of the following two sentences is easy for an English speaking human (clearly both are discussing the same topic, but with different emotional undertone):
#
# 1. Linear R3gr3ssion is very very cool!
# 2. What don’t I like it a single bit? Linear regressing!
#
# But a computer doesn’t understand
# - which words are nouns, verbs etc (grammar)
# - how to find the topic (word ordering)
# - feeling expressed in each sentence (sentiment)
# We need to represent the sentences in formats that a computer can easily process and manipulate.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Preprocessing
#
# If we’re interested in the topics/content of text, we may find many components of English sentences to be uninformative.
#
# 1. Word ordering
# 2. Punctuation
# 3. Conjugation of verbs (go vs going), declension of nouns (chair vs chairs)
# 4. Capitalization
# 5. Words with mostly grammatical functions: prepositions (before, under), articles (the, a, an) etc
# 6. Pronouns?
#
# These uninformative features of text will only confuse and distract a machine and should be removed.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Representing Documents: Bag Of Words
#
# After preprocessing our sentences:
#
# 1. (**S1**) linear regression is very very cool
# 2. (**S2**) what don’t like single bit linear regression
#
# We represent text in the format that is most accessible to a computer: numeric.
#
# We simply make a vector of the counts of the words in each sentence.
#
# <img src="./fig/fig15.png" style='height:100px;'>
#
#
# Turning a piece of text into a vector of word counts is called ***Bag of Words***.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Solving Machine Learning Tasks Using Bag of Words
#
# We can apply any existing machine learning model for classification or regression to the numerical representations of text data! In particular, we can apply a neural network classifier to BoW representations of text to perform document classification.
#
# <img src="./fig/fig16.png" style='height:300px;'>
# + [markdown] slideshow={"slide_type": "skip"}
# ## What Do the Hidden Layers in the Network Mean?
# The hidden layers of the document classifying are representations of words are **low-dimensional** real-valued vectors. These vectors are called ***word embeddings***. Sometimes, these representation captures semantic information and are much more compressed than the BoW representations!
#
# Such embeddings are often extracted and used in future tasks in the place of BoW representations.
#
# <img src="./fig/fig17.png" style='height:400px;'>
#
# Commonly used word embeddings:
# 1. **Word2Vec:** embeddings (representations of words extracted from a hidden layers of a neural network model) obtained by training a neural network to predict a target word using context, or to predict a target context using a word.<br><br>
#
# 2. **GloVe:** Ratios of word-word co-occurrence probabilities can encode meaning. Learn word vectors (embeddings) such that their dot product equals the logarithm of the words’ probability of co-occurrence (matrix factorization). This associates the log ratios of co-occurrence probabilities with vector differences in the word vector space.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Visualizing Word Embeddings
#
# There are a number of tools that allows users to interactively visualizes embeddings by rendering them in two or three dimensions.
#
# <img src="./fig/embedding.gif" style='height:400px;'>
#
# *From: [Embedding Projector](http://projector.tensorflow.org)*
# + [markdown] slideshow={"slide_type": "skip"}
# ## Visualizing Bias
# By visualizing word embeddings, we can often discover conceptual biases underlying the way we use language today - machine learning models often pick up these biases and propagate them unknowingly.
#
# For example, word embeddings often capture semantic relationship between words - the vector for 'apple' maybe close in Euclidean distance to the vector for 'pear'. However, given the training data (documents generated by human beings) word embeddings will also learn to associate 'man' with 'engineer' and 'woman' with 'homemaker'.
#
# <img src="./fig/bias.png" style='height:400px;'>
#
# *From: [wordbias](http://wordbias.umiacs.umd.edu)*
# + [markdown] slideshow={"slide_type": "slide"}
# ## Additional Resources
#
# 1. [Visual Analytics in Deep Learning: An Interrogative Survey for the Next Frontiers](https://arxiv.org/pdf/1801.06889.pdf)
#
# 2. [Understanding Neural Networks via Feature Visualization: A survey](https://arxiv.org/pdf/1904.08939.pdf)
#
# 3. [Visualizing and Understanding Recurrent Networks](http://vision.stanford.edu/pdf/KarpathyICLR2016.pdf)
#
# 4. [Visualizing memorization in RNNs](https://distill.pub/2019/memorization-in-rnns/)
#
# 5. [distill.pub](https://distill.pub)
#
# 6. [Google PAIR](https://research.google/teams/brain/pair/)
| content/lectures/lecture8/notebook/.ipynb_checkpoints/lecture8-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="aIMn8J64Dym-"
# # Tokyo Olympic 2020
# Data from Arjunprasadsarkhel in Kaggle.
# + [markdown] id="CL04gmw2EOy_"
# ## Import Data and Libraries
# + colab={"base_uri": "https://localhost:8080/"} id="cz7ZlxYP12bm" outputId="bdd8659a-8cf8-4b07-ec82-3d7a8ec566de"
# !pip install --upgrade plotly
# + colab={"base_uri": "https://localhost:8080/"} id="o5OA63B32Kh9" outputId="dd82c3b3-55a9-4b86-93b9-3faea546b166"
# !mkdiry ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !chmod 600 ~/.kaggle/kaggle.json
# !kaggle datasets download -d arjunprasadsarkhel/2021-olympics-in-tokyo
# !unzip 2021-olympics-in-tokyo.zip
# + colab={"base_uri": "https://localhost:8080/"} id="m5Gu8Yq366Kv" outputId="69ca2132-f2ee-42ea-8c9e-e5c7c30014ab"
# !pip install pyspark
# + colab={"base_uri": "https://localhost:8080/", "height": 214} id="0KtsB2Ne7W6B" outputId="52cbc76f-a833-4158-c5d3-3b552c9269cd"
import pyspark
spark = pyspark.sql.SparkSession(pyspark.SparkContext())
spark
# + id="vRRC-Lmz7ooj"
import pandas as pd
def excel_to_csv(name):
pd.read_excel(name + '.xlsx').to_csv(name + '.csv', sep=',', index=False)
excel_to_csv('Athletes')
excel_to_csv('Coaches')
excel_to_csv('EntriesGender')
excel_to_csv('Medals')
excel_to_csv('Teams')
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="c29rD7lK8hCb" outputId="93c31a72-1bdd-4a29-91f0-096bcd907ff0"
athletes = pd.read_csv('Athletes.csv')
coaches = pd.read_csv('Coaches.csv')
entriesGender = pd.read_csv('EntriesGender.csv')
medals = pd.read_csv('Medals.csv')
teams = pd.read_csv('Teams.csv')
athletes.head()
# + [markdown] id="efPkFAmhEXM5"
# ## Medals
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="DdJ6ky5P9MWN" outputId="36d59b0b-c0e8-4ba7-d744-cdfd4fcaeba4"
medals.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="9swY35vy9UZL" outputId="77a098f1-86aa-4104-fb8f-5101ab4c1c44"
medals.sort_values('Rank by Total', inplace=True)
medals.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="PH1uz0bD9hSL" outputId="301e6ca4-5b5d-46aa-cbed-a95e77d16d1a"
import plotly.express as px
fig = px.bar(medals, x='Team/NOC', y=['Gold', 'Silver', 'Bronze'], color_discrete_sequence=['Gold', 'Silver', 'Brown'], title='Number of Medals of Country')
fig.show()
# + [markdown] id="7Z-J94ndEfoi"
# ## Coaches
# + colab={"base_uri": "https://localhost:8080/", "height": 665} id="cDmBpwsQBdqC" outputId="9d61ad77-1c9e-4cb7-84d1-0b7ff734452a"
coaches.head(20)
# + colab={"base_uri": "https://localhost:8080/"} id="TiRYjNabCGbu" outputId="dc37db15-0ab4-447c-bf6c-6fbdef29364b"
spark.read.csv('Coaches.csv', header=True, inferSchema=True).createOrReplaceTempView('Coaches')
spark.sql('Select * From Coaches').show(5)
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="F2yAEkfTC-Ws" outputId="4fff0ce8-8b46-48a3-d58e-ab140d89836c"
pd_coaches_query = spark.sql('Select NOC, Count(NOC) as Count From Coaches Group By NOC Order By NOC').toPandas()
pd_coaches_query.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="MXG3VwVCER3d" outputId="5fd6c0a3-c7bb-42ee-b09d-d8e781282be4"
fig = px.bar(pd_coaches_query, x='NOC', y='Count', color='Count', title='Number of coaches from each country')
fig.show()
# + [markdown] id="Ywptlz-EEmJz"
# ## Athletes
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="r6VgqOpPEw2D" outputId="6f46d25a-fced-4244-fd90-eb4f6f05f1ac"
athletes.head()
# + colab={"base_uri": "https://localhost:8080/"} id="DqcJ9f4NE1JB" outputId="7a9b1b22-1418-4ce4-e569-8d2f61cc50d4"
spark.read.csv('Athletes.csv', header=True, inferSchema=True).createOrReplaceTempView('Athletes')
spark.sql('Select * From athletes Order By NOC, Discipline').show(20)
# + colab={"base_uri": "https://localhost:8080/"} id="p9MUOMrlFhnO" outputId="eaf8b63f-4c3e-4b03-f211-d015afcd0d0c"
spark.sql('''Select first(NOC) as NOC, first(Discipline) as Discipline,
count(*) as Count from athletes Group By NOC, Discipline
order by NOC, Discipline''').show(20)
# + id="U1xi13ZiGrUA"
pd_athletes_query = spark.sql('''Select first(NOC) as NOC, first(Discipline) as Discipline,
count(*) as Count from athletes Group By NOC, Discipline
order by NOC, Discipline''').toPandas()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="2Uqu4YeIGmOh" outputId="800dfd52-5ff7-40ca-dd9b-9141b7fe760f"
fig = px.bar(pd_athletes_query, x='NOC', y='Count', color='Discipline', title='Number of athletes from each country per discipline')
fig.show()
# + colab={"base_uri": "https://localhost:8080/"} id="TCdB7OstHYoo" outputId="3608e074-7d6c-45e2-8707-7dab93e54817"
athletes['Discipline'].value_counts().head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="6vjbaJfdIEwW" outputId="867ae468-934c-4d72-e6e6-5f0c70d16d68"
pd_athletes_query = pd_athletes_query.loc[pd_athletes_query['Discipline'].isin(['Athletics', 'Swimming', 'Football'])]
pd_athletes_query.Discipline.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="iGb17i1xJBs0" outputId="9c77fac0-ca33-4559-faec-6ab0c15d8b89"
fig = px.bar(pd_athletes_query, x='NOC', y='Count', color='Discipline', title='Number of athletes from each country per highest discipline')
fig.show()
# + [markdown] id="KGhH5oWyEwVE"
# ## Gender
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="6z59B03JJmIV" outputId="dd7930f7-da85-42ff-8482-0f08bb10e72a"
entriesGender.head(5)
# + colab={"base_uri": "https://localhost:8080/"} id="2m5bzaubKJUz" outputId="29b820a0-df7c-4f02-88cd-aacca089ab4d"
len(entriesGender)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="fpEnzV-CJ3Q2" outputId="82131fcf-36fc-497d-8f51-7094e889032b"
from plotly.subplots import make_subplots
import plotly.graph_objects as go
specs = [[{'type':'domain'}, {'type':'domain'}]] *23
fig = make_subplots(rows=23, cols=2,
subplot_titles=entriesGender['Discipline'], specs=specs)
sub_figs = []
for index, row in entriesGender.iterrows():
sub_fig = go.Pie(labels=['Female', 'Male'], values=[row['Female'],
row['Male']])
sub_figs.append(sub_fig)
k = 0
for i in range(1, 24):
for j in range(1,3):
fig.add_trace(sub_figs[k], i, j)
k += 1
fig.update_layout(showlegend=False, height=10000, width=800, title_text="Distribution of Gender amongst each game")
fig.update_traces(textposition='inside', textinfo='label+percent', hoverinfo='label+value+percent')
fig = go.Figure(fig)
fig.show()
# + [markdown] id="1x-RE549E2Qa"
# ## Teams
# + colab={"base_uri": "https://localhost:8080/", "height": 665} id="CRkLlcyhKuU0" outputId="3208a416-7460-4d1f-e3a2-9b24eb3442da"
teams.head(20)
# + colab={"base_uri": "https://localhost:8080/"} id="O2SDKWLrR-hW" outputId="6449d53d-df5a-4136-e807-b00c52815ea3"
spark.read.csv('Teams.csv', header=True, inferSchema=True).createOrReplaceTempView('Teams')
spark.sql('Select * From Teams Order By NOC, Event').show(20)
# + colab={"base_uri": "https://localhost:8080/"} id="A-V4hsgcQll6" outputId="394a4679-1d51-4837-d8df-bd216bd4e866"
spark.sql('''Select first(NOC) as NOC, first(Event) as Event,
count(*) as Count from teams Group By NOC, Event
order by NOC, Count''').show(5)
| NewTokyoEDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="jcaItn7dGHr_" colab_type="code" outputId="d606d7c7-5f8b-405d-f8f5-63a0ad2b0223" colab={"base_uri": "https://localhost:8080/", "height": 34}
dict = {}
type(dict)
# + id="0JKJe4hVZ6I3" colab_type="code" outputId="7d5f68e6-b44d-403d-aef2-bb4c6f25b856" colab={"base_uri": "https://localhost:8080/", "height": 104}
# %time dic ={'m1': 5,'m2': 3.5,'m3': 2.5,'m4': 3}
# %time dict(m1='iron man',m2='infinty war',m3='raganrok')
# + id="wIbuImCFay3n" colab_type="code" outputId="8da24c9c-c7a5-4163-a5b4-75ed0f6eeb13" colab={"base_uri": "https://localhost:8080/", "height": 34}
movies = ['ABCD','ABCD2']
actors = ['Canish', 'bcanish']
abc = {mov:actors for mov in movies }#for act in actors}
abc
# + id="BcHrfKBTdctw" colab_type="code" outputId="a03cff32-297b-44da-e8a9-64ce21be2171" colab={"base_uri": "https://localhost:8080/", "height": 69}
abc['gajni'] = "ganish"
abc
# + id="0ZZj4YWueRHb" colab_type="code" outputId="5aef6fbd-cf5c-4e58-e838-ed6cec24d6d2" colab={"base_uri": "https://localhost:8080/", "height": 69}
abc['gajni'] = ["ganish",'ccanish']
abc
# + id="s-B3XfitfaTy" colab_type="code" outputId="ee528ecb-dfde-44dd-e898-200c8c07b730" colab={"base_uri": "https://localhost:8080/", "height": 347}
abc['ABCD'].append('lsh')
abc
# + id="-XhuwYmXfu_u" colab_type="code" outputId="294d902a-0d2b-4158-c2a9-5b0be2fbd2fb" colab={"base_uri": "https://localhost:8080/", "height": 121}
for k,v in abc.items():
print(k,":",v)
actors.pop()
abc
# + id="GdrHMI-HhPey" colab_type="code" colab={}
| ml_course/ipynbfiles/Dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="cDgpEHaUETGA"
# ##### Copyright 2019 Google LLC.
#
#
# + cellView="form" colab={} colab_type="code" id="esgwBPvbEVsD"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="YMDobAW8mfWo"
# # Non-rigid surface deformation
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/non_rigid_deformation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/graphics/blob/master/tensorflow_graphics/notebooks/non_rigid_deformation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="L9EFL7fhiHbD"
# Non-rigid surface deformation is a technique that, among other things, can be used to interactively manipulate meshes or to deform a template mesh to fit to a point-cloud. When manipulating meshes, this can for instance allow users to move the hand of a character, and have the rest of the arm deform in a realistic manner. It is interesting to note that the deformation can also be performed over the scale of parts or the entire mesh.
#
#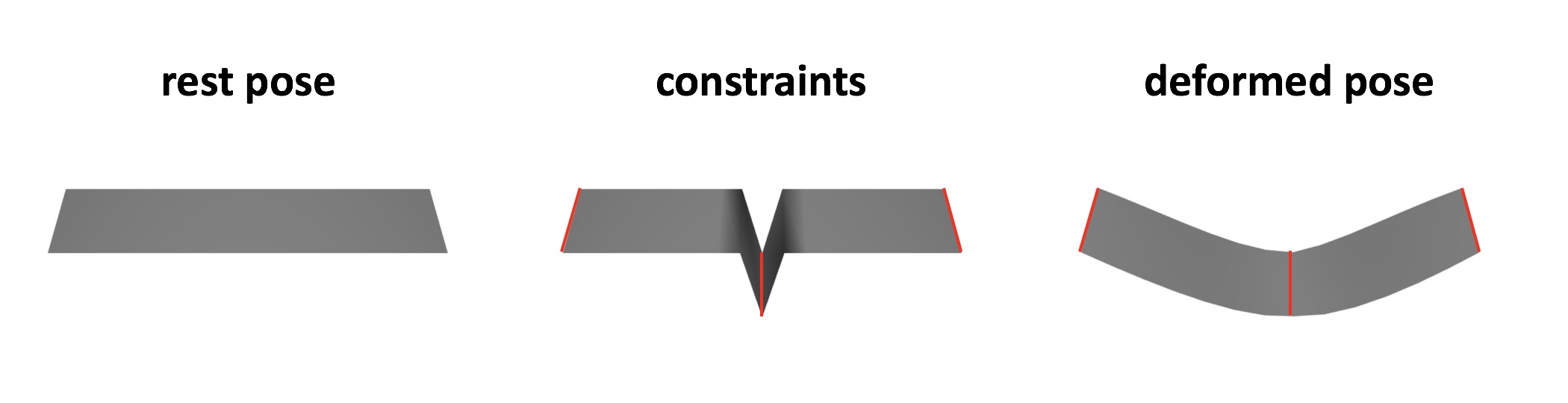 
#
# This notebook illustrates how to use [Tensorflow Graphics](https://github.com/tensorflow/graphics) to perform deformations similiar to the one contained in the above image.
# + [markdown] colab_type="text" id="BSMKlF6nAEPE"
# ## Setup & Imports
#
# If Tensorflow Graphics is not installed on your system, the following cell can install the Tensorflow Graphics package for you.
# + colab={} colab_type="code" id="gpmGP3DgAKRe"
# !pip install tensorflow_graphics
# + [markdown] colab_type="text" id="-YxmozRVANeD"
# Now that Tensorflow Graphics is installed, let's import everything needed to run the demo contained in this notebook.
# + colab={} colab_type="code" id="-OAKdOoTf_-f"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import numpy as np
import tensorflow as tf
from tensorflow_graphics.geometry.deformation_energy import as_conformal_as_possible
from tensorflow_graphics.geometry.representation.mesh import utils as mesh_utils
from tensorflow_graphics.geometry.transformation import quaternion
from tensorflow_graphics.math.optimizer import levenberg_marquardt
from tensorflow_graphics.notebooks import threejs_visualization
from tensorflow_graphics.notebooks.resources import triangulated_stripe
tf.compat.v1.enable_eager_execution()
# + [markdown] colab_type="text" id="1jLD48VvMquE"
# In this example, we build a mesh that corresponds to a flat and rectangular surface. Using the sliders, you can control the position of the deformation constraints applied to that surface, which respectively correspond to all the points along the left boundary, center, and right boundary of the mesh.
# + colab={} colab_type="code" id="IpztINWQ4Q4H"
mesh_rest_pose = triangulated_stripe.mesh
connectivity = mesh_utils.extract_unique_edges_from_triangular_mesh(triangulated_stripe.mesh['faces'])
camera = threejs_visualization.build_perspective_camera(
field_of_view=40.0, position=(0.0, -5.0, 5.0))
width = 500
height = 500
_ = threejs_visualization.triangular_mesh_renderer([mesh_rest_pose],
width=width,
height=height,
camera=camera)
# + cellView="form" colab={} colab_type="code" id="JVSAgVRXypak"
###############
# UI controls #
###############
#@title Constraints on the deformed pose { vertical-output: false, run: "auto" }
constraint_1_z = 0 #@param { type: "slider", min: -1, max: 1 , step: 0.05 }
constraint_2_z = -1 #@param { type: "slider", min: -1, max: 1 , step: 0.05 }
constraint_3_z = 0 #@param { type: "slider", min: -1, max: 1 , step: 0.05 }
vertices_rest_pose = tf.Variable(mesh_rest_pose['vertices'])
vertices_deformed_pose = np.copy(mesh_rest_pose['vertices'])
num_vertices = vertices_deformed_pose.shape[0]
# Adds the user-defined constraints
vertices_deformed_pose[0, 2] = constraint_1_z
vertices_deformed_pose[num_vertices // 2, 2] = constraint_1_z
vertices_deformed_pose[num_vertices // 4, 2] = constraint_2_z
vertices_deformed_pose[num_vertices // 2 + num_vertices // 4, 2] = constraint_2_z
vertices_deformed_pose[num_vertices // 2 - 1, 2] = constraint_3_z
vertices_deformed_pose[-1, 2] = constraint_3_z
mesh_deformed_pose = {
'vertices': vertices_deformed_pose,
'faces': mesh_rest_pose['faces']
}
vertices_deformed_pose = tf.Variable(vertices_deformed_pose)
# Builds a camera and render the mesh.
camera = threejs_visualization.build_perspective_camera(
field_of_view=40.0, position=(0.0, -5.0, 5.0))
_ = threejs_visualization.triangular_mesh_renderer([mesh_rest_pose],
width=width,
height=height,
camera=camera)
_ = threejs_visualization.triangular_mesh_renderer([mesh_deformed_pose],
width=width,
height=height,
camera=camera)
geometries = threejs_visualization.triangular_mesh_renderer(
[mesh_deformed_pose], width=width, height=height, camera=camera)
################
# Optimization #
################
def update_viewer_callback(iteration, objective_value, variables):
"""Callback to be called at each step of the optimization."""
geometries[0].getAttribute('position').copyArray(
variables[0].numpy().ravel().tolist())
geometries[0].getAttribute('position').needsUpdate = True
geometries[0].computeVertexNormals()
def deformation_energy(vertices_deformed_pose, rotation):
"""As conformal as possible deformation energy."""
return as_conformal_as_possible.energy(
vertices_rest_pose,
vertices_deformed_pose,
rotation,
connectivity,
aggregate_loss=False)
def soft_constraints(vertices_deformed_pose):
"""Soft constrains forcing results to obey the user-defined constraints."""
weight = 10.0
return (
weight * (vertices_deformed_pose[0, 2] - constraint_1_z),
weight * (vertices_deformed_pose[num_vertices // 2, 2] - constraint_1_z),
weight * (vertices_deformed_pose[num_vertices // 4, 2] - constraint_2_z),
weight * (vertices_deformed_pose[num_vertices // 2 + num_vertices // 4, 2] -
constraint_2_z),
weight *
(vertices_deformed_pose[num_vertices // 2 - 1, 2] - constraint_3_z),
weight * (vertices_deformed_pose[-1, 2] - constraint_3_z),
)
def fitting_energy(vertices_deformed_pose, rotation):
deformation = deformation_energy(vertices_deformed_pose, rotation)
constraints = soft_constraints(vertices_deformed_pose)
return tf.concat((deformation, constraints), axis=0)
rotations = tf.Variable(quaternion.from_euler(np.zeros((num_vertices, 3))))
max_iterations = 15 #@param { isTemplate: true, type: "integer" }
_ = levenberg_marquardt.minimize(
residuals=fitting_energy,
variables=(vertices_deformed_pose, rotations),
max_iterations=int(max_iterations),
callback=update_viewer_callback)
| tensorflow_graphics/notebooks/non_rigid_deformation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp mymodel01
# -
# # mymodel01
# > API details.
#hide
from nbdev.showdoc import *
# # Introduction
# !pip install nbdev==1.1.23
# ### In this notebook kernel, I'm going to predictions customers are eligible for the loan and check whether what are the missing criteria to know why customer not getting loan to make there own house.
#
#
# <div class="text-success "><h4> We will learning about, Data Analysis Preprocess such as, </h4></div>
#
# ---
#
# > ### Steps are:
#
#
# 1. [Gathering Data](#1)
# - [Exploratory Data Analysis](#2)
# - [Data Visualizations](#3)
# - [Machine Learning Model Decision.](#4)
# - [Traing the ML Model](#5)
# - [Predict Model](#6)
# - [Deploy Model](#7)
#
#
#
#
# **Hope** you guys ****Love It**** and get a better **learning experience**. 🙏
#
# ---
#
# <div class="text-danger" >
# <h4>Let's Say, You are the owner of the <b>Housing Finance Company</b> and you want to build your own model to predict the customers are applying for the home loan and company want to check and validate the customer are eligible for the home loan.
# </h4>
# </div>
#
# # <div class="text-primary"> The Problem is, </div>
#
# ### In a Simple Term, Company wants to make automate the Loan Eligibility Process in a real time scenario related to customer's detail provided while applying application for home loan forms.
#
#
# You will use the training set to build your model, and the test set to validate it. Both the files are stored on the web as CSV files; their URLs are already available as character strings in the sample code.
#
#
# First of all, we need to importing the necessary packages to work with the data to solve our problem
# # Import Packages
#export
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# +
# Show the Dataset Path to get detaset
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# -
# You can load this data with the `read_csv()` method from `pandas` package. It converts the data set to a python dataframe.
#
# ## Dataset Key Information.
#
# ---
#
# > - `Loan_ID`--------------> Unique Loan ID.
# - `Gender` --------------> Male/ Female
# - `Married` --------------> Applicant married (Y/N)
# - `Dependents` ------------> Number of dependents
# - `Education` -------------> Applicant Education (Graduate/ Under Graduate)
# - `Self_Employed` ---------> Self-employed (Y/N)
# - `ApplicantIncome` -------> Applicant income
# - `CoapplicantIncome` -----> Coapplicant income
# - `LoanAmount` -----------> Loan amount in thousands
# - `Loan_Amount_Term` ------> Term of a loan in months
# - `Credit_History` --------> Credit history meets guidelines
# - `Property_Area` ---------> Urban/ Semi-Urban/ Rural
# - `Loan_Status` -----------> Loan approved (Y/N)
# <a id="1"></a><br>
# # 1. Gathering Data
# +
#export
# Create New Variable and stores the dataset values as Data Frame
# loan_train = pd.read_csv('/kaggle/input/loan-eligible-dataset/loan-train.csv')
# loan_test = pd.read_csv('/kaggle/input/loan-eligible-dataset/loan-test.csv')
traindata_url="https://gist.githubusercontent.com/erwangranger/c1534647ecd6efc6188a92d69657f3bc/raw/eb6da3554971d9c3ab8f0506b37f285fb6453642/loans_train.csv"
testdata_url="https://gist.githubusercontent.com/erwangranger/43749b3d049b572ae83572719519cbbc/raw/47277db78d37994f8d9347265078be6619df234d/loans_test.csv"
loan_train = pd.read_csv(traindata_url)
loan_test = pd.read_csv(testdata_url)
# -
# - Lets display the some few information from our large datasets
#
# Here, We shows the first five rows from datasets
loan_train.head()
# - As we can see in the above output, there are too many columns, ( columns known as features as well. )
#
# We can also use `loan_train` to show few rows from the first five and last five record from the dataset
loan_train
# > ### Here, we can see there are many rows and many columns, To know how many records and columns are available in our dataset, we can use the `shape` attribute or we can use `len()` to know how many records and how many features available in the dataset.
print("Rows: ", len(loan_train))
# Pandas has inbuild attribute to get all column from the dataset, With the help of this feature we can get the how many column available we have.
print("Columns: ", len(loan_train.columns))
# Also we can get the shape of the dataset using `shape` attribute
print("Shape : ", loan_train.shape)
# > ### *After we collecting the data, Next step we need to understand what kind of data we have.*
# ### Also we can get the column as an list(array) from dataset
#
# > **Note: DataFrame.columns returns the total columns of the dataset,
# > Store the number of columns in variable `loan_train_columns`**
#export
loan_train_columns = loan_train.columns # assign to a variable
loan_train_columns # print the list of columns
# ### Now, Understanding the Data
# - First of all we use the `loan_train.describe()` method to shows the important information from the dataset
# - It provides the `count`, `mean`, `standard deviation (std)`, `min`, `quartiles` and `max` in its output.
loan_train.describe()
# #### As I said the above cell, this the information of all the methamatical details from dataset. Like `count`, `mean`, `standard deviation (std)`, `min`, `quartiles(25%, 50%, 75%)` and `max`.
# > ### Another method is `info()`, This method show us the information about the dataset, Like
#
# 1. What's the type of culumn have?
# - How many rows available in the dataset?
# - What are the features are there?
# - How many null values available in the dataset?
# - Ans so on...
loan_train.info()
# As we can see in the output.
#
# 1. There are `614` entries
# - There are total 13 features (0 to 12)
# - There are three types of datatype `dtypes: float64(4), int64(1), object(8)`
# - It's Memory usage that is, `memory usage: 62.5+ KB`
# - Also, We can check how many missing values available in the `Non-Null Count` column
# <a id="2"></a><br>
# # 2. Exploratory Data Analysis
#
# In this section, We learn about extra information about data and it's characteristics.
#
# - First of all, We explore object type of data
# So let's make a function to know how many types of values available in the column
def explore_object_type(df ,feature_name):
"""
To know, How many values available in object('categorical') type of features
And Return Categorical values with Count.
"""
if df[feature_name].dtype == 'object':
print(df[feature_name].value_counts())
# - After defined a function, Let's call it. and check what's the output of our created function.
# Now, Test and Call a function for gender only
explore_object_type(loan_train, 'Gender')
# - Here's one little issue occurred, Suppose in your datasets there are lots of feature to defined like this above code.
# +
# Solution is, Do you remember we have variable with name of `loan_train_columns`, Right, let's use it
# 'Loan_ID', 'Gender', 'Married', 'Dependents', 'Education', 'Self_Employed', 'Property_Area', 'Loan_Status'
for featureName in loan_train_columns:
if loan_train[featureName].dtype == 'object':
print('\n"' + str(featureName) + '\'s" Values with count are :')
explore_object_type(loan_train, str(featureName))
# -
# > ## *Note: Your output maybe shorter or longer, It's totally depend upon your dataset's columns*
#
#
# - We need to fill null values with `mean` and `median` using `missingno` package
# +
#export
# !pip install missingno
import missingno as msno
# +
# list of how many percentage values are missing
loan_train
loan_train.isna().sum()
# round((loan_train.isna().sum() / len(loan_train)) * 100, 2)
# -
msno.bar(loan_train)
msno.matrix(loan_train )
# - As we can see here, there are too many columns missing with small amount of null values so we use `mean` amd `mode` to replace with `NaN` values.
# +
#export
loan_train['Credit_History'].fillna(loan_train['Credit_History'].mode(), inplace=True) # Mode
loan_test['Credit_History'].fillna(loan_test['Credit_History'].mode(), inplace=True) # Mode
loan_train['LoanAmount'].fillna(loan_train['LoanAmount'].mean(), inplace=True) # Mean
loan_test['LoanAmount'].fillna(loan_test['LoanAmount'].mean(), inplace=True) # Mean
# -
# ### # convert Categorical variable with Numerical values.
# `Loan_Status` feature boolean values, So we replace `Y` values with `1` and `N` values with `0`
# and same for other `Boolean` types of columns
# +
#export
loan_train.Loan_Status = loan_train.Loan_Status.replace({"Y": 1, "N" : 0})
# loan_test.Loan_Status = loan_test.Loan_Status.replace({"Y": 1, "N" : 0})
loan_train.Gender = loan_train.Gender.replace({"Male": 1, "Female" : 0})
loan_test.Gender = loan_test.Gender.replace({"Male": 1, "Female" : 0})
loan_train.Married = loan_train.Married.replace({"Yes": 1, "No" : 0})
loan_test.Married = loan_test.Married.replace({"Yes": 1, "No" : 0})
loan_train.Self_Employed = loan_train.Self_Employed.replace({"Yes": 1, "No" : 0})
loan_test.Self_Employed = loan_test.Self_Employed.replace({"Yes": 1, "No" : 0})
# +
#export
loan_train['Gender'].fillna(loan_train['Gender'].mode()[0], inplace=True)
loan_test['Gender'].fillna(loan_test['Gender'].mode()[0], inplace=True)
loan_train['Dependents'].fillna(loan_train['Dependents'].mode()[0], inplace=True)
loan_test['Dependents'].fillna(loan_test['Dependents'].mode()[0], inplace=True)
loan_train['Married'].fillna(loan_train['Married'].mode()[0], inplace=True)
loan_test['Married'].fillna(loan_test['Married'].mode()[0], inplace=True)
loan_train['Credit_History'].fillna(loan_train['Credit_History'].mean(), inplace=True)
loan_test['Credit_History'].fillna(loan_test['Credit_History'].mean(), inplace=True)
# -
# * Here, `Property_Area`, `Dependents` and `Education` has multiple values so now we can use `LabelEncoder` from `sklearn` package
# +
#export
from sklearn.preprocessing import LabelEncoder
feature_col = ['Property_Area','Education', 'Dependents']
le = LabelEncoder()
for col in feature_col:
loan_train[col] = le.fit_transform(loan_train[col])
loan_test[col] = le.fit_transform(loan_test[col])
# -
# > ### Finally, We have all the features with numerical values,
# <a id="3"></a><br>
# # 3. Data Visualizations
#
#
# In this section, We are showing the visual information from the dataset, For that we need some pakages that are `matplotlib` and `seaborn`
#
#
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style('dark')
# -
loan_train
# +
loan_train.plot(figsize=(18, 8))
plt.show()
# +
plt.figure(figsize=(18, 6))
plt.subplot(1, 2, 1)
loan_train['ApplicantIncome'].hist(bins=10)
plt.title("Loan Application Amount ")
plt.subplot(1, 2, 2)
plt.grid()
plt.hist(np.log(loan_train['LoanAmount']))
plt.title("Log Loan Application Amount ")
plt.show()
# +
plt.figure(figsize=(18, 6))
plt.title("Relation Between Applicatoin Income vs Loan Amount ")
plt.grid()
plt.scatter(loan_train['ApplicantIncome'] , loan_train['LoanAmount'], c='k', marker='x')
plt.xlabel("Applicant Income")
plt.ylabel("Loan Amount")
plt.show()
# -
plt.figure(figsize=(12, 6))
plt.plot(loan_train['Loan_Status'], loan_train['LoanAmount'])
plt.title("Loan Application Amount ")
plt.show()
plt.figure(figsize=(12,8))
sns.heatmap(loan_train.corr(), cmap='coolwarm', annot=True, fmt='.1f', linewidths=.1)
plt.show()
# In this heatmap, we can clearly seen the relation between two variables
# <a id="4"></a><br>
# # 4. Choose ML Model.
# * In this step, We have a lots of Machine Learning Model from sklearn package, and we need to decide which model is give us the better performance. then we use that model in final stage and send to the production level.
# +
#export
# import ml model from sklearn pacakge
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# -
# First of all, we are use `LogisticRegression` from `sklearn.linear_model` package. Here is the little information about `LogisticRegression`.
#
# `Logistic Regression` is a **classification algorithm**. It is used to predict a binary outcome (`1 / 0`, `Yes / No`, and `True / False`) given a set of independent variables. To represent binary / categorical outcome, we use dummy variables. You can also think of logistic regression as a special case of linear regression when the outcome variable is categorical, where we are using log of odds as the dependent variable.
#
# 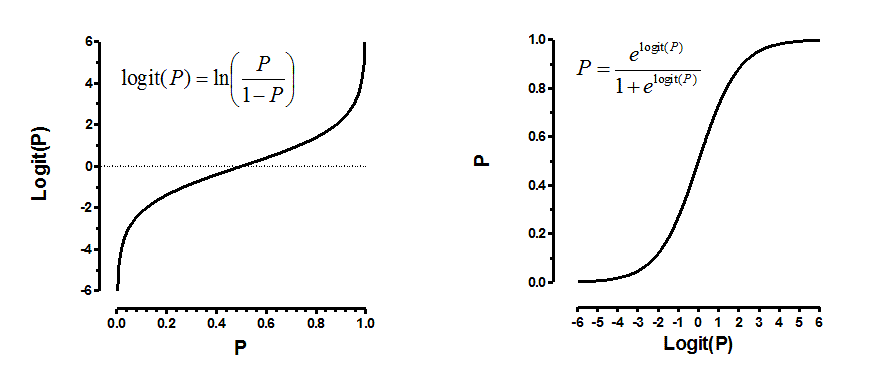
# * Let's build the model
#export
logistic_model = LogisticRegression()
# <a id="5"></a><br>
# # 5. Traing the ML Model
# > ### **Before fitting the model, We need to decide how many feature are available for testing and training, then after complete this step. fitt the model**
#
# Currently, we are using `Credit_History', 'Education', 'Gender` features for training so let's create train and test variables
# +
#export
train_features = ['Credit_History', 'Education', 'Gender']
x_train = loan_train[train_features].values
y_train = loan_train['Loan_Status'].values
x_test = loan_test[train_features].values
# -
#export
logistic_model.fit(x_train, y_train)
# <a id="6"></a><br>
# # 6. Predict Model
# +
#export
# Predict the model for testin data
predicted = logistic_model.predict(x_test)
# -
# check the coefficeints of the trained model
print('Coefficient of model :', logistic_model.coef_)
# check the intercept of the model
print('Intercept of model',logistic_model.intercept_)
# Accuray Score on train dataset
# accuracy_train = accuracy_score(x_test, predicted)
score = logistic_model.score(x_train, y_train)
print('accuracy_score overall :', score)
print('accuracy_score percent :', round(score*100,2))
# +
# predict the target on the test dataset
predict_test = logistic_model.predict(x_test)
print('Target on test data',predict_test)
# -
# <a id="7"></a><br>
# # 7. Deploy Model
#
# - Finally, we are done so far. The last step is to deploy our model in production map. So we need to export our model and bind with web application API.
#
# Using pickle we can export our model and store in to `logistic_model.pkl` file, so we can ealy access this file and calculate customize prediction using Web App API.
#
#
# #### A little bit information about pickle:
#
# `Pickle` is the standard way of serializing objects in Python. You can use the pickle operation to serialize your machine learning algorithms and save the serialized format to a file. Later you can load this file to deserialize your model and use it to make new predictions
#
#
# >> Here is example of the Pickle export model
#
#
#
# ```
# model.fit(X_train, Y_train)
# # save the model to disk
# filename = 'finalized_model.sav'
# pickle.dump(model, open(filename, 'wb'))
#
# # some time later...
#
# # load the model from disk
# loaded_model = pickle.load(open(filename, 'rb'))
# result = loaded_model.score(X_test, Y_test)
# print(result)
# ```
#export
import pickle as pkl
#export
filename = 'logistic_model_01.pkl'
pkl.dump(logistic_model, open(filename, 'wb')) # wb means write as binary
# ### Now, You can check your current directory. You can see the file with named "logistic_model.pkl"
# - To read model from file
#
# ```
# # load the model from disk
# loaded_model = pkl.load(open(filename, 'rb')) # rb means read as binary
# result = loaded_model.score(X_test, Y_test)
#
# ```
#
# +
#loaded_model = pkl.load(open(filename, 'rb')) # rb means read as binary
#result = loaded_model.score(x_test, y_test)
# -
from nbdev.export import *
notebook2script()
| create_model_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Magic from here onwards, do not touch
# %load_ext autoreload
# %autoreload 2
# +
import math
import wandb
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
from tqdm import tqdm_notebook as tqdm
matplotlib.rc('font', **{'size': 16})
# -
api = wandb.Api()
sweep = api.sweep("ko3n1g/equivariant-attention/t1u6qh1s")
sweep_data = []
for run in tqdm(sweep.runs):
history = run.history()
df = history.dropna(subset=['Test loss'], axis=0)
break
df
df = pd.DataFrame(sweep_data)
# +
plt.figure(figsize=(15,6))
plt.subplot(1, 3, 1)
plt.plot(df["_step"], df['Test loss'], linewidth=3, label='Test loss')
plt.tight_layout()
plt.legend(loc='upper right')
plt.ylabel('Loss')
plt.xlabel('Step')
plt.grid(b=True, which='major', axis='y')
plt.subplot(1, 3, 2)
plt.plot(df["_step"], df['Test pos_mse'], linewidth=3, label='Test pos_mse')
plt.tight_layout()
plt.legend(loc='upper right')
plt.ylabel('Loss')
plt.xlabel('Step')
plt.grid(b=True, which='major', axis='y')
plt.subplot(1, 3, 3)
plt.plot(df["_step"], df['Test vel_mse'], linewidth=3, label='Test vel_mse')
plt.tight_layout()
plt.legend(loc='upper right')
plt.ylabel('Loss')
plt.xlabel('Step')
plt.grid(b=True, which='major', axis='y')
plt.savefig(f'Final model vs epoch.pdf', bbox_inches='tight')
plt.savefig(f'Final model vs epoch.png', bbox_inches='tight')
plt.show()
# -
| experiments/nbody/Model comparison - Final runs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Tce3stUlHN0L"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="qFdPvlXBOdUN"
# # Decoding API
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/text/guide/decoding_api"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/text/blob/master/docs/guide/decoding_api.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/text/blob/master/docs/guide/decoding_api.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/text/docs/guide/decoding_api.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="xHxb-dlhMIzW"
# ## Overview
# In the recent past, there has been a lot of research in language generation with auto-regressive models. In auto-regressive language generation, the probability distribution of token at time step *K* is dependent on the model's token-predictions till step *K-1*. For these models, decoding strategies such as Beam search, Greedy, Top-p, and Top-k are critical components of the model and largely influence the style/nature of the generated output token at a given time step *K*.
#
# For example, **Beam search** reduces the risk of missing hidden high probability tokens by
# keeping the most likely num_beams of hypotheses at each time step and eventually
# choosing the hypothesis that has the overall highest probability.
# [Murray et al. (2018)](https://arxiv.org/abs/1808.10006) and
# [Yang et al. (2018)](https://arxiv.org/abs/1808.09582) show that beam search
# works well in Machine Translation tasks.
# Both **Beam search** & **Greedy** strategies have a possibility of generating
# repeating tokens.
#
# [Fan et. al (2018)](https://arxiv.org/pdf/1805.04833.pdf) introduced **Top-K
# sampling**, in which the K most likely tokens are filtered and the probability mass
# is redistributed among only those K tokens.
#
# [<NAME> et. al (2019)](https://arxiv.org/pdf/1904.09751.pdf) introduced
# **Top-p sampling**, which chooses from the smallest possible set of tokens with
# cumulative probability that adds upto the probability *p*. The probability mass is then
# redistributed among this set. This way, the size of the set of tokens can
# dynamically increase and decrease.
# **Top-p, Top-k** are generally used in tasks such as story-generation.
#
# The Decoding API provides an interface to experiment with different decoding strategies on auto-regressive models.
#
# 1. The following sampling strategies are provided in sampling_module.py, which inherits from the base Decoding class:
# * [top_p](https://arxiv.org/abs/1904.09751) : [github](https://github.com/tensorflow/models/blob/master/official/nlp/modeling/ops/sampling_module.py#L65)
#
# * [top_k](https://arxiv.org/pdf/1805.04833.pdf) : [github](https://github.com/tensorflow/models/blob/master/official/nlp/modeling/ops/sampling_module.py#L48)
#
# * Greedy : [github](https://github.com/tensorflow/models/blob/master/official/nlp/modeling/ops/sampling_module.py#L26)
#
# 2. Beam search is provided in beam_search.py. [github](https://github.com/tensorflow/models/blob/master/official/nlp/modeling/ops/beam_search.py)
# + [markdown] id="MUXex9ctTuDB"
# ## Setup
# + id="60_D9NLa9KhJ"
# !pip install -q -U tensorflow-text==2.7.3
# + id="FJV1ttb8dZyQ"
# !pip install -q tf-models-official==2.7.0
# + id="IqR2PQG4ZaZ0"
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from official import nlp
from official.nlp.modeling.ops import sampling_module
from official.nlp.modeling.ops import beam_search
# + [markdown] id="j9r8-Q_CekYK"
# ## Initialize Sampling Module in TF-NLP.
#
#
# * **symbols_to_logits_fn** : Use this closure to call the model to predict the logits for the `index+1` step. Inputs and outputs for this closure are as follows:
# ```
# Args:
# 1] ids : Current decoded sequences. int tensor with shape (batch_size, index + 1 or 1 if padded_decode is True)],
# 2] index [scalar] : current decoded step,
# 3] cache [nested dictionary of tensors] : Only used for faster decoding to store pre-computed attention hidden states for keys and values. More explanation in the cell below.
# Returns:
# 1] tensor for next-step logits [batch_size, vocab]
# 2] the updated_cache [nested dictionary of tensors].
# ```
# The cache is used for faster decoding.
# Here is a [reference](https://github.com/tensorflow/models/blob/master/official/nlp/modeling/ops/beam_search_test.py#L88) implementation for the above closure.
#
#
# * **length_normalization_fn** : Use this closure for returning length normalization parameter.
# ```
# Args:
# 1] length : scalar for decoded step index.
# 2] dtype : data-type of output tensor
# Returns:
# 1] value of length normalization factor.
# ```
#
# * **vocab_size** : Output vocabulary size.
#
# * **max_decode_length** : Scalar for total number of decoding steps.
#
# * **eos_id** : Decoding will stop if all output decoded ids in the batch have this eos_id.
#
# * **padded_decode** : Set this to True if running on TPU. Tensors are padded to max_decoding_length if this is True.
#
# * **top_k** : top_k is enabled if this value is > 1.
#
# * **top_p** : top_p is enabled if this value is > 0 and < 1.0
#
# * **sampling_temperature** : This is used to re-estimate the softmax output. Temperature skews the distribution towards high probability tokens and lowers the mass in tail distribution. Value has to be positive. Low temperature is equivalent to greedy and makes the distribution sharper, while high temperature makes it more flat.
#
# * **enable_greedy** : By default, this is True and greedy decoding is enabled. To experiment with other strategies, please set this to False.
# + [markdown] id="xqpGECmAeu7Q"
# ## Initialize the Model hyperparameters
# + id="KtylpxOmceaC"
params = {}
params['num_heads'] = 2
params['num_layers'] = 2
params['batch_size'] = 2
params['n_dims'] = 256
params['max_decode_length'] = 4
# + [markdown] id="pwdM2pl3RSPb"
# In auto-regressive architectures like Transformer based [Encoder-Decoder](https://arxiv.org/abs/1706.03762) models,
# Cache is used for fast sequential decoding.
# It is a nested dictionary storing pre-computed hidden-states (key and values in the self-attention blocks and in the cross-attention blocks) for every layer.
# + [markdown] id="A_xX-fbze8_S"
# ## Initialize cache.
# + id="mMOeXVmbdilM"
cache = {
'layer_%d' % layer: {
'k': tf.zeros([params['batch_size'], params['max_decode_length'], params['num_heads'], int(params['n_dims']/params['num_heads'])], dtype=tf.float32),
'v': tf.zeros([params['batch_size'], params['max_decode_length'], params['num_heads'], int(params['n_dims']/params['num_heads'])], dtype=tf.float32)
} for layer in range(params['num_layers'])
}
print("cache key shape for layer 1 :", cache['layer_1']['k'].shape)
# + [markdown] id="-9BGL3gOe-2K"
# ## Define closure for length normalization if needed.
# This is used for normalizing the final scores of generated sequences and is optional
#
# + id="U82B_tH2d294"
def length_norm(length, dtype):
"""Return length normalization factor."""
return tf.pow(((5. + tf.cast(length, dtype)) / 6.), 0.0)
# + [markdown] id="TJdqBNBbS78n"
# ## Create model_fn
# In practice, this will be replaced by an actual model implementation such as [here](https://github.com/tensorflow/models/blob/master/official/nlp/transformer/transformer.py#L236)
# ```
# Args:
# i : Step that is being decoded.
# Returns:
# logit probabilities of size [batch_size, 1, vocab_size]
# ```
# + id="xVeis7YZfaQM"
probabilities = tf.constant([[[0.3, 0.4, 0.3], [0.3, 0.3, 0.4],
[0.1, 0.1, 0.8], [0.1, 0.1, 0.8]],
[[0.2, 0.5, 0.3], [0.2, 0.7, 0.1],
[0.1, 0.1, 0.8], [0.1, 0.1, 0.8]]])
def model_fn(i):
return probabilities[:, i, :]
# + [markdown] id="_G2rkaCPfcka"
# ## Initialize symbols_to_logits_fn
#
# + id="1B6T3629fdKJ"
def _symbols_to_logits_fn():
"""Calculates logits of the next tokens."""
def symbols_to_logits_fn(ids, i, temp_cache):
del ids
logits = tf.cast(tf.math.log(model_fn(i)), tf.float32)
return logits, temp_cache
return symbols_to_logits_fn
# + [markdown] id="rhosGmvZffke"
# ## Greedy
# Greedy decoding selects the token id with the highest probability as its next id: $id_t = argmax_{w}P(id | id_{1:t-1})$ at each timestep $t$. The following sketch shows greedy decoding.
# + id="JZ-p0JdbfyJ7"
greedy_obj = sampling_module.SamplingModule(
length_normalization_fn=None,
dtype=tf.float32,
symbols_to_logits_fn=_symbols_to_logits_fn(),
vocab_size=3,
max_decode_length=params['max_decode_length'],
eos_id=10,
padded_decode=False)
ids, _ = greedy_obj.generate(
initial_ids=tf.constant([9, 1]), initial_cache=cache)
print("Greedy Decoded Ids:", ids)
# + [markdown] id="pOmG0IE6ff40"
# ## top_k sampling
# In *Top-K* sampling, the *K* most likely next token ids are filtered and the probability mass is redistributed among only those *K* ids.
# + id="qkIDv7VZfuzr"
top_k_obj = sampling_module.SamplingModule(
length_normalization_fn=length_norm,
dtype=tf.float32,
symbols_to_logits_fn=_symbols_to_logits_fn(),
vocab_size=3,
max_decode_length=params['max_decode_length'],
eos_id=10,
sample_temperature=tf.constant(1.0),
top_k=tf.constant(3),
padded_decode=False,
enable_greedy=False)
ids, _ = top_k_obj.generate(
initial_ids=tf.constant([9, 1]), initial_cache=cache)
print("top-k sampled Ids:", ids)
# + [markdown] id="PaEv2c_cflsE"
# ## top_p sampling
# Instead of sampling only from the most likely *K* token ids, in *Top-p* sampling chooses from the smallest possible set of ids whose cumulative probability exceeds the probability *p*.
# + id="WzHslibyfs6K"
top_p_obj = sampling_module.SamplingModule(
length_normalization_fn=length_norm,
dtype=tf.float32,
symbols_to_logits_fn=_symbols_to_logits_fn(),
vocab_size=3,
max_decode_length=params['max_decode_length'],
eos_id=10,
sample_temperature=tf.constant(1.0),
top_p=tf.constant(0.9),
padded_decode=False,
enable_greedy=False)
ids, _ = top_p_obj.generate(
initial_ids=tf.constant([9, 1]), initial_cache=cache)
print("top-p sampled Ids:", ids)
# + [markdown] id="hTSdHdTjfoPV"
# ## Beam search decoding
# Beam search reduces the risk of missing hidden high probability token ids by keeping the most likely num_beams of hypotheses at each time step and eventually choosing the hypothesis that has the overall highest probability.
# + id="U1jIPF_qfqcO"
beam_size = 2
params['batch_size'] = 1
beam_cache = {
'layer_%d' % layer: {
'k': tf.zeros([params['batch_size'], params['max_decode_length'], params['num_heads'], params['n_dims']], dtype=tf.float32),
'v': tf.zeros([params['batch_size'], params['max_decode_length'], params['num_heads'], params['n_dims']], dtype=tf.float32)
} for layer in range(params['num_layers'])
}
print("cache key shape for layer 1 :", beam_cache['layer_1']['k'].shape)
ids, _ = beam_search.sequence_beam_search(
symbols_to_logits_fn=_symbols_to_logits_fn(),
initial_ids=tf.constant([9], tf.int32),
initial_cache=beam_cache,
vocab_size=3,
beam_size=beam_size,
alpha=0.6,
max_decode_length=params['max_decode_length'],
eos_id=10,
padded_decode=False,
dtype=tf.float32)
print("Beam search ids:", ids)
| docs/guide/decoding_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # 예제 9-2 미리 학습한 모델을 사용하는 방법
# +
# # %load /home/sjkim/.jupyter/head.py
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from importlib import reload
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
#os.environ["CUDA_VISIBLE_DEVICES"]="0"
# seaborn
#import seaborn as sns
#sns.set( style = 'white', font_scale = 1.7)
#sns.set_style('ticks')
#plt.rcParams['savefig.dpi'] = 200
# font for matplotlib
#import matplotlib
#import matplotlib.font_manager as fm
#fm.get_fontconfig_fonts()
#font_location = '/usr/share/fonts/truetype/nanum/NanumGothicBold.ttf'
#font_name = fm.FontProperties(fname=font_location).get_name()
#matplotlib.rc('font', family=font_name)
# -
import ex9_2_pretarined_method as example
example.main()
| nb_ex9_2_pretrained_method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Normalized Consumption Visualizations
#
# <NAME> - <EMAIL>
import pandas as pd
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime
import timeit
import os
from datetime import datetime
from __future__ import division
from pylab import *
import matplotlib.dates as mdates
from matplotlib import ticker
import datetime
import matplotlib
sns.set_style("whitegrid")
# %matplotlib inline
repos_path = "/Users/Clayton/temporal-features-for-buildings/"
meta = pd.read_csv(os.path.join(repos_path,"data/raw/meta_open.csv"), index_col='uid', parse_dates=["datastart","dataend"], dayfirst=True)
temp = pd.read_csv((os.path.join(repos_path,"data/interim/temp_open_utc_complete.csv")), index_col='timestamp', parse_dates=True).tz_localize('utc')
# # Individual visualization
# +
def get_individual_data(temp, meta, building):
timezone = meta.T[building].timezone
start = meta.T[building].datastart
end = meta.T[building].dataend
return pd.DataFrame(temp[building].tz_convert(timezone).truncate(before=start,after=end))
def get_individual_data_notz(temp, meta, building):
start = meta.T[building].datastart
end = meta.T[building].dataend
return pd.DataFrame(temp[building].truncate(before=start,after=end))
# -
temp_loadratio = pd.DataFrame()
for building in meta.index:
df = get_individual_data(temp, meta, building)
df_min = df.resample('D').min()
df_max = df.resample('D').max()
df_loadratio = df_min / df_max
temp_loadratio = pd.merge(temp_loadratio, df_loadratio.tz_localize(None), right_index=True, left_index=True, how='outer')
temp_loadratio.tail()
building = "Office_Ellie"
start = '2012-12-01'
end = '2012-12-30'
df = get_individual_data(temp, meta, building).truncate(before=start,after=end)
df_loadratio = get_individual_data_notz(temp_loadratio, meta, building).truncate(before=start,after=end).dropna()
df_loadratio.head()
def plot_line_example(df_1, df_2, color):
sns.set(rc={"figure.figsize": (12,4)})
sns.set_style('whitegrid')
fig = plt.figure()
fig.autofmt_xdate()
fig.subplots_adjust(hspace=.5)
gs = GridSpec(100,100,bottom=0.18,left=0.18,right=0.88)
df_1.columns = ["Actual kWh"]
#df_predicted.columns = ["Predicted kWh"]
ax1 = fig.add_subplot(gs[1:60,:])
df_1.plot(ax = ax1)
ax1.xaxis.set_visible(False)
ax1.set_title("Hourly kWh")
ax2 = fig.add_subplot(gs[68:,:])
df_2 = df_2.tz_localize(None)
x = mdates.drange(df_2.index[0], df_2.index[-1] + datetime.timedelta(days=1), datetime.timedelta(days=1))
y = np.linspace(0, len(df_2.columns), len(df_2.columns)+1)
data = np.array(df_2.T)
datam = np.ma.array(data, mask=np.isnan(data))
cmap = matplotlib.cm.get_cmap(color)
qmesh = ax2.pcolormesh(x, y, datam, cmap=cmap)
ax2.set_title("Normalized Hourly Consumption [kWh/m2]")
#leftspacing,
cbaxes = fig.add_axes([0.18, 0.08, 0.7, 0.02])
cbar = fig.colorbar(qmesh, ax=ax2, orientation='horizontal', cax=cbaxes)
ax2.axis('tight')
ax2.xaxis_date()
ax2.yaxis.set_visible(False)
myFmt = mdates.DateFormatter('%b %d')
ax2.xaxis.set_major_formatter(myFmt)
plot_line_example(df, df_loadratio, "PuBu")
plt.savefig(os.path.join(repos_path,"reports/figures/featureoverviews/loadratio_example.pdf"));
# +
#sns.palplot(sns.light_palette("viridis"))
# -
# # Overview heatmap
# +
def heatmap_data(df):
x = mdates.drange(df.index[0], df.index[-1] + datetime.timedelta(days=1), datetime.timedelta(days=1))
y = np.linspace(0, len(df.columns), len(df.columns)+1)
return x,y
#This function changes the year for all of the buildings so that they can be overlapped for plotting
# THE RESULTING DATAFRAME IS ONLY FOR "GENERALIZED" PLOTTING TO GET THE DATA SETS TO OVERLAP -- DON'T PERFORM CALCULATIONS ON IT
def changeto2015(x):
try:
return x.replace(year=2015)
except:
return np.nan
def is_leap_and_29Feb(s):
return (s.index.year % 4 == 0) & ((s.index.year % 100 != 0) | (s.index.year % 400 == 0)) & (s.index.month == 2) & (s.index.day == 29)
def generalize_timestamp_forplotting(meta, temp):
temp_generalized = pd.DataFrame()
buildinglist = list(meta.index)
for building in buildinglist:
#Change to timezone
timezone = meta.T[building].timezone
start = meta.T[building].datastart
end = meta.T[building].dataend
building_data = pd.DataFrame(temp[building].tz_convert(timezone).truncate(before=start,after=end))
#Remove leap year day for 2012
mask = is_leap_and_29Feb(building_data)
building_data = building_data.loc[~mask]
#Change to 2015 -- we arbitrarily use this year
building_data.index = building_data.index.map(lambda t: changeto2015(t))
#Remove tz awareness, resample to normalize the timestamps and ffill to get rid of the gaps
building_data = building_data.tz_localize(None)
building_data = building_data.resample('D').mean().sort_index()
building_data = building_data.ffill()
#Merge into dataset
temp_generalized = pd.merge(temp_generalized, building_data, right_index=True, left_index=True, how='outer')
temp_generalized = temp_generalized.resample('D').mean().sort_index()
#print "finished building: "+building
return temp_generalized
def generalize_timestamp_forplotting_notz(meta, temp):
temp_generalized = pd.DataFrame()
buildinglist = list(meta.index)
for building in buildinglist:
#Change to timezone
start = meta.T[building].datastart
end = meta.T[building].dataend
building_data = pd.DataFrame(temp[building].truncate(before=start,after=end))
#Remove leap year day for 2012
mask = is_leap_and_29Feb(building_data)
building_data = building_data.loc[~mask]
#Change to 2015 -- we arbitrarily use this year
building_data.index = building_data.index.map(lambda t: changeto2015(t))
#Remove tz awareness, resample to normalize the timestamps and ffill to get rid of the gaps
building_data = building_data.tz_localize(None)
building_data = building_data.resample('D').mean().sort_index()
building_data = building_data.ffill()
#Merge into dataset
temp_generalized = pd.merge(temp_generalized, building_data, right_index=True, left_index=True, how='outer')
temp_generalized = temp_generalized.resample('D').mean().sort_index()
#print "finished building: "+building
return temp_generalized
def plotmap(df, color, cbarlabel, xaxislabel, yaxislabel, graphiclabel, filelabel):
import matplotlib.dates as mdates
from matplotlib import ticker
import datetime
import matplotlib
import seaborn as sns
# Set up the size/style
sns.set(rc={"figure.figsize": (5,11)})
sns.set_style("whitegrid")
numberofplots = 1
fig = plt.figure()
x = mdates.drange(df.index[0], df.index[-1] + datetime.timedelta(days=1), datetime.timedelta(days=1))
y = np.linspace(0, len(df.columns), len(df.columns)+1)
ax = fig.add_subplot(numberofplots, 1, 1)
data = np.array(df.T)
datam = np.ma.array(data, mask=np.isnan(data))
cmap = matplotlib.cm.get_cmap(color)
qmesh = ax.pcolormesh(x, y, datam, cmap=cmap)
cbaxes = fig.add_axes([0.15, 0.15, 0.7, 0.02])
cbar = fig.colorbar(qmesh, ax=ax, orientation='horizontal', cax=cbaxes)
cbar.ax.tick_params(length = 0)
cbar.set_label(cbarlabel)
ax.axis('tight')
ax.xaxis_date()
fig.autofmt_xdate()
fig.subplots_adjust(hspace=.5)
ax.set_xlabel(xaxislabel)
ax.set_ylabel(yaxislabel)
ax.set_title(graphiclabel)
ax.set_yticklabels(df.columns)
tick_locator = ticker.MaxNLocator(nbins=110)
loc = ticker.MultipleLocator(base=1.0) # this locator puts ticks at regular intervals
ax.locator_params(axis='y', nbins=100)
myFmt = mdates.DateFormatter('%b')
ax.xaxis.set_major_formatter(myFmt)
plt.subplots_adjust(bottom=0.2)
def plotmap_subplots(df, meta, color, cbarlabel, xaxislabel, yaxislabel, graphiclabel, filelabel, z_min, z_max):
import matplotlib.dates as mdates
from matplotlib import ticker
import datetime
import matplotlib
import seaborn as sns
# Set up the size/style
sns.set(rc={"figure.figsize": (5,11)})
sns.set_style("whitegrid")
cmap = matplotlib.cm.get_cmap(color)
fig = plt.figure()
fig.autofmt_xdate()
fig.subplots_adjust(hspace=.5)
gs = GridSpec(100,100,bottom=0.18,left=0.18,right=0.88)
# Plot Offices
df_office = df[df.columns[df.columns.str.contains("Office")]]
office_height = int(len(df_office.T)/len(df.T)*100-2)
ax1 = fig.add_subplot(gs[1:office_height,:])
x,y = heatmap_data(df_office)
data = np.array(df_office.T)
datam = np.ma.array(data, mask=np.isnan(df_office))
qmesh = ax1.pcolormesh(x, y, datam, cmap=cmap, vmin=z_min, vmax=z_max)
ax1.axis('tight')
ax1.xaxis_date()
ax1.set_title("Offices")
ax1.xaxis.set_visible(False)
# Plot UnivLab
df_lab = df[df.columns[df.columns.str.contains("UnivLab")]]
lab_height = int(len(df_lab.T)/len(df.T)*100-2)
ax2 = fig.add_subplot(gs[office_height+2:office_height+lab_height,:])
x,y = heatmap_data(df_lab)
data = np.array(df_lab.T)
datam = np.ma.array(data, mask=np.isnan(df_lab))
qmesh = ax2.pcolormesh(x, y, datam, cmap=cmap, vmin=z_min, vmax=z_max)
ax2.axis('tight')
ax2.xaxis_date()
ax2.set_title("University Labs")
ax2.xaxis.set_visible(False)
# Plot UnivClass
df_uniclass = df[df.columns[df.columns.str.contains("UnivClass")]]
uniclass_height = int(len(df_uniclass.T)/len(df.T)*100-2)
ax3 = fig.add_subplot(gs[office_height+lab_height+2:office_height+lab_height+uniclass_height,:])
x,y = heatmap_data(df_uniclass)
data = np.array(df_uniclass.T)
datam = np.ma.array(data, mask=np.isnan(df_uniclass))
qmesh = ax3.pcolormesh(x, y, datam, cmap=cmap, vmin=z_min, vmax=z_max)
ax3.axis('tight')
ax3.xaxis_date()
ax3.set_title("University Classrooms")
ax3.xaxis.set_visible(False)
# Plot PrimClass
df_primclass = df[df.columns[df.columns.str.contains("PrimClass")]]
primclass_height = int(len(df_primclass.T)/len(df.T)*100-2)
ax4 = fig.add_subplot(gs[office_height+lab_height+uniclass_height+2:office_height+lab_height+uniclass_height+primclass_height,:])
x,y = heatmap_data(df_primclass)
data = np.array(df_primclass.T)
datam = np.ma.array(data, mask=np.isnan(df_primclass))
qmesh = ax4.pcolormesh(x, y, datam, cmap=cmap, vmin=z_min, vmax=z_max)
ax4.axis('tight')
ax4.xaxis_date()
ax4.set_title("Primary/Secondary Classroom")
ax4.xaxis.set_visible(False)
# Plot Univ Dorms
df_unidorm = df[df.columns[df.columns.str.contains("UnivDorm")]]
unidorm_height = int(len(df_unidorm.T)/len(df.T)*100-2)
ax5 = fig.add_subplot(gs[office_height+lab_height+uniclass_height+primclass_height+2:office_height+lab_height+uniclass_height+primclass_height+unidorm_height,:])
x,y = heatmap_data(df_unidorm)
data = np.array(df_unidorm.T)
datam = np.ma.array(data, mask=np.isnan(df_unidorm))
qmesh = ax5.pcolormesh(x, y, datam, cmap=cmap, vmin=z_min, vmax=z_max)
ax5.axis('tight')
ax5.xaxis_date()
ax5.set_title("University Dorms")
cbaxes = fig.add_axes([0.18, 0.23, 0.7, 0.02])
cbar = fig.colorbar(qmesh, orientation='horizontal', cax=cbaxes) #ax=ax,
cbar.ax.tick_params(length = 0)
cbar.set_label(cbarlabel)
myFmt = mdates.DateFormatter('%b')
ax5.xaxis.set_major_formatter(myFmt)
fig.text(0.14, 0.5, yaxislabel, va='center', rotation='vertical')
#plt.subplots_adjust(bottom=0.2)
#plt.savefig(filelabel+".png", dpi=500)maps import *
# +
#temp_residuals_normalized.describe().T["count"].value_counts()
# +
#temp_loadratio
# -
temp_loadratio_generalized = generalize_timestamp_forplotting_notz(meta, temp_loadratio)
# +
#temp_loadratio_generalized.head()
# -
temp_loadratio_generalized_sorted = temp_loadratio_generalized[list(temp_loadratio_generalized.sum().sort_values().index)]
temp_loadratio_generalized_sorted.head()#.describe().T.describe()
plotmap_subplots(temp_loadratio_generalized_sorted, meta, 'PuBu', "Daily Load Ratio",
"Timeline", "", "kWh Difference", "Daily Load Ratio", 0, 1)
plt.savefig(os.path.join(repos_path,"reports/figures/featureoverviews/loadratio_heatmap.pdf"));
# # Loop through and make ratio-based features
#
# - Daily
# - Weekends
# - Weekdays
#
# Ratios Tested
# - min vs. max
# - mean vs. max
# - median vs. max
# - min vs 95 perc
# - mean vs 95 perc
# - median vs 95 perc
# +
#df.groupby(df.index.date).quantile(0)
# -
df.quantile(0)
def make_temporalfeatures(df, label, overall_label):
features = pd.DataFrame()
features[overall_label+"_"+label+"_mean"] = df.mean()
features[overall_label+"_"+label+"_min"] = df.min()
features[overall_label+"_"+label+"_max"] = df.max()
features[overall_label+"_"+label+"_std"] = df.std()
return features
def get_ratio(df, quan_num, quan_denom):
df_num = df.groupby(df.index.date).quantile(quan_num)
df_denom = df.groupby(df.index.date).quantile(quan_denom)
ratio = df_num / df_denom
return ratio
runs = {
'minvsmax':[0,1],
'meanvsmax':[0.5,1],
'minvs95':[0,0.95],
'meanvs95':[0.5,0.95]
}
runtypes = {
'all':[0,6],
'weekdays':[0,4],
'weekend':[5,6]
}
# +
#tempfeatures
# -
df[(df.index.weekday >= 5) & (df.index.weekday <= 6)].head()
# +
overall_start_time = timeit.default_timer()
ratios = pd.DataFrame()
for runtype in runtypes:
for run in runs:
temp_loadratio = pd.DataFrame()
for building in meta.index:
df = get_individual_data(temp, meta, building)
df = df[(df.index.weekday >= runtypes[runtype][0]) & (df.index.weekday <= runtypes[runtype][1])]
df_loadratio = get_ratio(df, runs[run][0], runs[run][1])
temp_loadratio = pd.merge(temp_loadratio, df_loadratio, right_index=True, left_index=True, how='outer')
tempfeatures = make_temporalfeatures(temp_loadratio, run, runtype)
ratios = pd.merge(ratios, tempfeatures, right_index=True, left_index=True, how='outer')
print "Calculated "+run+" and "+runtype+" in "+str(timeit.default_timer() - overall_start_time)+" seconds"
print "Calculated everything in "+str(timeit.default_timer() - overall_start_time)+" seconds"
# -
# ## Make ratios
ratios = pd.read_csv(os.path.join(repos_path,"data/processed/feature_ratios.csv"), index_col='building_name')
ratios.head()
ratios.columns = "BG_"+ratios.columns
ratios.info()
ratios.index.name = "building_name"
ratios.columns.name = "feature_name"
ratios.to_csv(os.path.join(repos_path,"data/processed/feature_ratios.csv"))
ratios.describe().T.describe()
| 03_LoadRatio Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CPAS Exploration
# Connecting to CPAS with cx_Oracle
# cx_Oracle is a Python extension module that enables access to Oracle Database
import cx_Oracle
import sqlalchemy
import os
import urllib
import json
import pandas as pd
# +
USERNAME = os.environ.get('CPAS_USERNAME')
PASSWORD = os.environ.get('CPAS_PASSWORD')
HOSTNAME = '10.32.196.224'
PORT = '1332'
SERVICE_NAME = 'visispd1.lacity.org'
dsn = cx_Oracle.makedsn(HOSTNAME, PORT, service_name=SERVICE_NAME)
connection = cx_Oracle.connect(user=USERNAME, password=PASSWORD, dsn=dsn)
# -
engine = sqlalchemy.create_engine(
f"oracle+cx_oracle://{USERNAME}:{PASSWORD}@{HOSTNAME}:{PORT}/"
f"?service_name={SERVICE_NAME}"
)
# # Accessing Grant Applications
def get_grnt_app(year):
USER_APP = f"GRNT{year}"
yr_app = year
return pd.read_sql(
f"""SELECT * FROM
(SELECT
apl.grnt_aplctn_id,
apl.prjct_ttl,
apl.fnd_yr,
(SELECT
field_value
FROM
{USER_APP}.grnt_l_stat,
{USER_APP}.grnt_stat
WHERE
grnt_stat.grnt_l_stat = grnt_l_stat.field_id
AND apl.grnt_aplctn_id = grnt_stat.grnt_aplctn_id
) AS aplctn_status,
(SELECT
field_value
FROM
grnt.grnt_l_dept,
{USER_APP}.grnt_cntct
WHERE
grnt_cntct.grnt_l_dept = grnt_l_dept.field_id
AND cty_dept_yn = 1
AND apl.grnt_city_dept_cntct_id = grnt_cntct_id
) AS dept_nm
FROM
{USER_APP}.grnt_aplctn apl
WHERE
1 = 1
)
WHERE
1 = 1
AND fnd_yr = {yr_app}
ORDER BY
prjct_ttl,
dept_nm,
aplctn_status""", engine)
get_grnt_app(46).head(5)
# ## Creating a csv file for grant years 33-46
import csv
with open('df_app.csv',mode = 'w') as f:
f = csv.writer(f)
for year in range(37,47):
df_app = pd.DataFrame(get_grnt_app(year))
df_app.to_csv('df_app.csv', mode = 'a', header = True, index = False)
# ## List of departments submitting applications between grant years 33-46
import pandas as pd
import itertools
from collections import Counter
adn = df_app.dept_nm.unique()
print(pd.value_counts(adn))
list(i for i,_ in itertools.groupby(sorted(adn)))
# ## List of project titles
import pandas as pd
import itertools
pt = df_app.prjct_ttl.unique()
list(i for i,_ in itertools.groupby(sorted(pt)))
# ## Script to create json files for individual grant years
grnt41_app = pd.DataFrame(get_grnt_app(41))
grnt41_app.to_json('grnt41_app.json', orient = 'records')
# ## json files covering multiple grant years
def grnt_app(year):
user_num = get_grnt_app(year)
yr_app = year
return f'grnt{user_num}_app = pd.DataFrame{yr_app}'
for user_num in range(33,47):
f"grnt{user_num}_app.to_json(r'grnt{user_num}_app.json', orient = 'records')"
grnt_app(42)
user_num = (grnt_app(43))
f"grnt{user_num}_app = pd.DataFrame{user_num}"
f"grnt{user_num}_app.to_json(r'grnt{user_num}_app.json', orient = 'records')"
# +
# shape of the dataframe
df_app_tot = pd.read_csv('../data/df_app.csv')
print(df_app_tot.shape, '\n')
# data type of each column
print(df_app_tot.dtypes, '\n')
# number of null values
print(df_app_tot.info())
# +
# number of unique values of column dept
print(df_app_tot.dept_nm.nunique(),'\n')
# unique values of column dept
print(df_app_tot.dept_nm.unique())
# -
# # Accessing GRP Information
def get_grnt_grp(year):
USER_grp = f"GRNT{year}"
return pd.read_sql(
f"""SELECT DISTINCT
{USER_grp}.grnt_gpr.yr,
{USER_grp}.grnt_gpr.grnt_gpr_id,
{USER_grp}.grnt_gpr.dept,
{USER_grp}.grnt_gpr.pid,
{USER_grp}.grnt_gpr.actv_nbr,
{USER_grp}.grnt_gpr.proj_nm,
{USER_grp}.grnt_gpr.actv_nm,
{USER_grp}.grnt_gpr.proj_addr,
{USER_grp}.grnt_gpr.proj_desc,
{USER_grp}.grnt_gpr.natl_obj,
{USER_grp}.grnt_gpr.hud_cd,
{USER_grp}.grnt_hud_cd.grnt_hud_cd_id,
{USER_grp}.grnt_hud_cd.ttl,
{USER_grp}.grnt_hud_cd.regulation_cit,
{USER_grp}.grnt_gpr.grnt_l_accmplsh,
{USER_grp}.grnt_gpr.obj_cnt,
{USER_grp}.grnt_gpr.otcm_cnt,
{USER_grp}.grnt_gpr.accmplsh_actl_units,
{USER_grp}.grnt_gpr.accmplsh_narrtv,
{USER_grp}.grnt_gpr.fund_amt,
{USER_grp}.grnt_gpr.drn_thru_amt,
{USER_grp}.grnt_gpr.tot_accmplsh,
{USER_grp}.grnt_gpr.tot_hsg,
{USER_grp}.grnt_gpr.accmplsh_narrtv_updt,
{USER_grp}.grnt_gpr.aprv_anlst_email,
{USER_grp}.grnt_gpr.aprv_anlst_tel,
{USER_grp}.grnt_gpr.aprv_anlst_sig_dt,
{USER_grp}.grnt_gpr.aprv_supv_nm,
{USER_grp}.grnt_gpr.aprv_supv_email,
{USER_grp}.grnt_gpr.aprv_anlst_dept_nm,
{USER_grp}.grnt_gpr.gpr_subm_dt,
{USER_grp}.grnt_gpr.grnt_l_gpr_actv_stts,
{USER_grp}.grnt_gpr.ent_in_idis_dt
FROM
{USER_grp}.grnt_gpr
INNER JOIN {USER_grp}.grnt_hud_cd ON {USER_grp}.grnt_gpr.hud_cd = {USER_grp}.grnt_hud_cd.hud_cd
WHERE
{USER_grp}.grnt_gpr.yr > '2009'
ORDER BY
{USER_grp}.grnt_gpr.yr,
{USER_grp}.grnt_gpr.proj_nm""", engine)
import csv
with open('df_grp.csv',mode = 'w') as f:
f = csv.writer(f)
for year in range(35,45):
df_grp = pd.DataFrame(get_grnt_grp(year))
df_grp.to_csv('df_grp.csv', mode = 'a', header = True, index = False)
# +
# shape of the dataframe
df_grp_tot = pd.read_csv('df_grp.csv')
print(df_grp_tot.shape, '\n')
# data type of each column
print(df_grp_tot.dtypes, '\n')
# number of null values
print(df_grp_tot.info())
# +
# number of unique values of column dept
print(df_grp_tot.dept.nunique(),'\n')
# unique values of column dept
print(df_grp_tot.dept.unique())
# -
grnt44_grp = pd.DataFrame(get_grnt_grp(44))
grnt44_grp.to_json('grnt44_grp.json', orient = 'records')
def grnt_grp(year):
user_num = get_grnt_grp(year)
yr_grp = year
return f'grnt{user_num}_grp = pd.DataFrame{yr_grp}'
for user_num in range(33,46):
f"grnt{user_num}_grp.to_json(r'grnt{user_num}_grp.json', orient = 'records')"
# +
# grnt_grp(44)
# -
grnt44_grp = pd.DataFrame(get_grnt_grp(44))
grnt44_grp.to_json('grnt_grp.json', orient = 'records')
# ## Accessing PEP Information
def get_grnt_pep(year):
USER_pep = f"GRNT{year}"
yr_pep = year
return pd.read_sql(
f"""SELECT
{yr_pep} AS pgm_year,
grnt_pep_id,
pep_agcy_nm,
pep_proj_nm,
citywide_cncl_dist_yn,
grnt_l_pep_stts,
to_char(pep_vrftn_dt, 'mm/dd/yyyy') AS pep_vrftn_dt,
to_char(pep_to_mgmt_dt, 'mm/dd/yyyy') AS pep_to_mgmt_dt,
to_char(pep_send_out_dt, 'mm/dd/yyyy') AS pep_send_out_dt,
grnt_l_send_out_mthd,
to_char(cdbo_send_vrftn_dt, 'mm/dd/yyyy') AS cdbo_send_vrftn_dt,
to_char(pep_rcv_dt, 'mm/dd/yyyy') AS pep_rcv_dt,
pep_rcv_dt AS pep_rcv_dt_2,
(
SELECT
field_value
FROM
{USER_pep}.grnt_l_pep_typ
WHERE
field_id = gp.grnt_l_pep_typ
) AS grnt_l_pep_typ_value,
(
SELECT
prjct_ttl
FROM
{USER_pep}.grnt_aplctn
WHERE
grnt_aplctn_id = gp.grnt_aplctn_id
) AS prjct_ttl,
cdbg_fnd_amt,
(
SELECT
proj_id
FROM
{USER_pep}.grnt_prpsd_proj
WHERE
grnt_aplctn_id = gp.grnt_aplctn_id
) AS proj_id,
(
SELECT
pep_note
FROM
{USER_pep}.grnt_pep_note
WHERE
grnt_l_pep_note_typ = 470
AND grnt_pep_id = gp.grnt_pep_id
) AS trackingcomment,
(
SELECT
pep_note
FROM
{USER_pep}.grnt_pep_note
WHERE
grnt_l_pep_note_typ = 485
AND grnt_pep_id = gp.grnt_pep_id
) AS pep_vrftn_comment,
(
SELECT
pep_note
FROM
{USER_pep}.grnt_pep_note
WHERE
grnt_l_pep_note_typ = 487
AND grnt_pep_id = gp.grnt_pep_id
) AS pep_to_mgmt_comment,
(
SELECT
pep_note
FROM
grnt43.grnt_pep_note
WHERE
grnt_l_pep_note_typ = 488
AND grnt_pep_id = gp.grnt_pep_id
) AS cdbo_send_vrftn_comment,
(
SELECT
field_value
FROM
{USER_pep}.grnt_l_pep_stts
WHERE
field_id = gp.grnt_l_pep_stts
) AS grnt_l_pep_stts_value,
(
SELECT
field_value
FROM
{USER_pep}.grnt_l_send_out_mthd
WHERE
field_id = gp.grnt_l_send_out_mthd
) AS grnt_l_send_out_mthd_value,
(
SELECT
ldp.field_value
FROM
{USER_pep}.grnt_aplctn apl,
{USER_pep}.grnt_cntct cicon,
grnt.grnt_l_dept ldp
WHERE
apl.grnt_aplctn_id = gp.grnt_aplctn_id
AND cicon.cty_dept_yn = 1
AND apl.grnt_city_dept_cntct_id = cicon.grnt_cntct_id
AND cicon.grnt_l_dept = ldp.field_id
) AS department,
grnt_l_rec_color,
(
SELECT
field_value
FROM
{USER_pep}.grnt_l_rec_color
WHERE
field_id = gp.grnt_l_rec_color
) AS grnt_l_rec_color_value,
(
SELECT
grnt_user.grnt_utility.value_list('
select
field_value
from
{USER_pep}.GRNT_PEP_CNCL_DIST A,
{USER_pep}.GRNT_L_CNCL_DIST B
where B.FIELD_ID=A.GRNT_L_CNCL_DIST
and
grnt_pep_id = '
|| gp.grnt_pep_id)
FROM
dual
) AS council_district,
temp.status_date
FROM
{USER_pep}.grnt_pep gp,
(
SELECT
MAX(mod_dt) AS status_date,
pk_id
FROM
{USER_pep}.grnt_hist
WHERE
table_nm = 'GRNT_PEP'
AND pk_nm = 'GRNT_PEP_ID'
AND field_nm = 'GRNT_L_PEP_STTS'
GROUP BY
pk_id
) temp
WHERE
gp.grnt_pep_id = temp.pk_id (+)
AND grnt_pep_id IN (
SELECT
grnt_pep_id
FROM
{USER_pep}.grnt_pep
)
ORDER BY
proj_id,
pep_agcy_nm""", engine)
# +
#get_grnt_pep(45) # script works for grnt38 - 45
# -
import csv
import json
with open('df_pep.csv',mode = 'w') as f:
f = csv.writer(f)
for year in range(38,46):
df_pep = pd.DataFrame(get_grnt_pep(year))
df_pep.to_csv('df_pep.csv', mode = 'a', header =True, index = False)
def grnt_pep(year):
user_num = get_grnt_pep(year)
yr_pep = year
return f'grnt{user_num}_pep = pd.DataFrame{yr_pep}'
for user_num in range(33,46):
f"grnt{user_num}_pep.to_json(r'grnt{user_num}_pep.json', orient = 'records')"
grnt41_pep = pd.DataFrame(get_grnt_pep(41))
grnt41_pep.to_json('grnt41_pep.json', orient = 'records')
# +
# shape of the dataframe
df_pep_tot = pd.read_csv('df_pep.csv')
print(df_pep_tot.shape, '\n')
# data type of each column
print(df_pep_tot.dtypes, '\n')
# number of null values
print(df_pep_tot.info())
# +
# number of unique values of column dept
print(df_pep_tot.department.nunique(),'\n')
# unique values of column dept
print(df_pep_tot.department.unique())
# -
# ## Basic Visualizations
# Fund Amt by Council District for Grant year 44
import matplotlib
df_pep = pd.read_json("grnt41_pep.json")
df_pep.boxplot('cdbg_fnd_amt', 'council_district', rot = 30, figsize=(15,10))
# +
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import colors
from matplotlib.ticker import PercentFormatter
np.random.seed(1968001)
# +
df_grp = pd.read_csv("df_grp.csv")
y = df_grp['hud_cd']
x = df_grp['fund_amt']
fig, ax = plt.subplots(tight_layout = True, figsize=(15,10) )
ax.scatter(x,y)
# +
## Need to check for duplications over multiple years
df_grp = pd.read_csv("df_grp.csv")
fig, axs = plt.subplots(sharey = True, tight_layout = True, figsize=(25,10))
plt.yscale('linear')
plt.title('Linear')
axs.bar(df_grp['hud_cd'], df_grp['drn_thru_amt'])
plt.ylim(0, 1.4e7)
# +
df_grp = pd.read_csv("df_grp.csv")
np.random.seed(1968001)
x = df_grp['hud_cd']
y = df_grp['fund_amt']
plt.subplots(sharey = True, tight_layout = True, figsize=(30,10))
plt.bar(x,y)
plt.yscale('linear')
plt.title('Linear', fontsize=20)
plt.xticks(fontsize=20)
plt.yticks(fontsize=20)
plt.grid()
plt.show()
# +
import seaborn as sns
import pandas.util.testing as tm
df_grp_json = pd.read_json("grnt44_grp.json")
sns.set(style = 'ticks', color_codes = True)
sns.catplot(x = "regulation_cit", y = "fund_amt", data = df_grp_json,
height = 7, aspect = 2)
# +
import seaborn as sns
import pandas.util.testing as tm
df_grp_json = pd.read_json("grnt44_grp.json")
sns.set(style = 'ticks', color_codes = True)
sns.catplot(x = "yr", y = "dept", data = df_grp_json, height = 7, aspect = 2)
# -
sns.catplot(x = "yr", y = "ttl", data = df_grp_json,
height = 10, aspect = 1.5)
# +
import seaborn as sns
import pandas.util.testing as tm
df_pep_json = pd.read_json("grnt41_pep.json")
sns.set(style = 'ticks', color_codes = True)
sns.catplot(x = "cdbg_fnd_amt", y = "department", data = df_pep_json,
height = 5, aspect = 1)
# -
| notebooks/grnt_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('bmh')
plt.rcParams['figure.dpi'] = 300
plt.rcParams['font.size'] = 8
plt.rcParams['legend.fontsize'] = 7
plt.rcParams['legend.loc'] = 'lower right'
plt.rcParams['figure.facecolor'] = '#FFFFFF'
# + tags=[]
env_names = ['Hopper-v2', 'Walker2d-v2', 'HalfCheetah-v2', 'Ant-v2', 'Humanoid-v2']
nenvs = 5
nrow = 2
ncol = (nenvs + nrow - 1) // nrow
fig, axs = plt.subplots(nrow, ncol, figsize=(4 * ncol, 3 * nrow))
log_dirs = [
('logs', 'sac'),
]
for log_dir, label in log_dirs:
for i in range(ncol * nrow):
row = i // ncol
col = i % ncol
ax = axs[row, col]
if i < len(env_names):
env_dir = os.path.join(log_dir, env_names[i])
if os.path.isdir(env_dir):
all_results = []
min_len = 1e6
for result in os.listdir(env_dir):
if 'txt' in result:
result_file = os.path.join(env_dir, result)
results = np.loadtxt(result_file)
all_results.append(results)
min_len = min(min_len, results.shape[0])
all_results = [result[:min_len] for result in all_results]
all_results = np.stack(all_results, axis=0)
xs = all_results[0, :, 0]
mean = np.mean(all_results[:, :, 1], axis=0)
std = np.std(all_results[:, :, 1], axis=0)
ax.plot(xs, mean, label=label, linewidth=1)
ax.fill_between(xs, mean - std, mean + std, alpha=0.25)
ax.set_xlabel('Environment Steps ($\\times 10^6%$)')
ax.set_ylabel('Episode Return')
ax.set_title(env_names[i])
if i == 0:
ax.legend()
else:
ax.axis('off')
plt.tight_layout()
plt.show()
# -
| learning_curves/plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Supervised learning with scikit-learn
#
# In this notebook, we review scikit-learn's API for training a model.
# +
from sklearn.datasets import fetch_openml
blood = fetch_openml('blood-transfusion-service-center', as_frame=True)
# -
blood.frame.head()
X, y = blood.data, blood.target
X.head()
y.head()
y.value_counts(normalize=True)
# ## Split Data
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=0
)
# -
X_train.head()
y_train.value_counts(normalize=True)
y_test.value_counts(normalize=True)
# ### Stratify!
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=0, stratify=y
)
y_train.value_counts(normalize=True)
y_test.value_counts(normalize=True)
# ## scikit-learn API
from sklearn.linear_model import Perceptron
percept = Perceptron()
percept.fit(X_train, y_train)
percept.predict(X_train)
y_train
percept.score(X_train, y_train)
percept.score(X_test, y_test)
# ## Another estimator
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
rf.score(X_train, y_train)
rf.score(X_test, y_test)
# ## Are these results any good?
from sklearn.dummy import DummyClassifier
dc = DummyClassifier(strategy='prior')
dc.fit(X_train, y_train)
dc.score(X_test, y_test)
# ## Exercise 1
#
# 1. Import and evaluate the performance of `sklearn.linear_model.LogisticRegression` on the above dataset
# 2. How does the test performance compare to the ones we already looked at?
# +
# # %load solutions/02-ex1-solution.py
# -
# ## Exercise 2
#
# 1. Load the wine dataset from `sklearn.datasets` module using the `load_wine` dataset.
# 2. Split it into a training and test set using `train_test_split`.
# 3. Train and evalute `sklearn.neighbors.KNeighborsClassifer`, `sklearn.ensemble.RandomForestClassifier` and `sklearn.linear_model.LogisticRegression` on the wine dataset.
# 4. How do they perform on the training and test set?
# 5. Which one is best on the training set and which one is best on the test set?
# +
# # %load solutions/02-ex2-solution.py
| notebooks/02-supervised-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
final_df = pd.read_csv('./final_df.csv',low_memory=False)
final_df = final_df.applymap(lambda s:s.lower() if type(s) == str else s)
# +
def search(df = final_df, text = None,columns = ['Continent', 'Country', 'Region', 'Cultural Group', 'Local name', 'Keywords for printouts', 'Class for printouts', 'Processes for printouts', 'Materials for printouts', 'Date made', 'Field Collector', 'When collected', 'PRM Source', 'Acquired'], incomplete_columns=['PRM Source', 'Description', 'Research notes', 'Publications history', 'Primary documentation']):
text = re.findall(r'(\w+)',text)
c=0
for i in columns:
# print(i)
if c==0:
if len(df[df[i].str.contains('|'.join(text))==True])>0:
# print(len(df[df[i].str.contains('|'.join(text))==True]))
appa = df[df[i].str.contains('|'.join(text))==True]
c+=1
else:
pass
else:
appa.append(df[df[i].str.contains('|'.join(text))==True])
appa['ALL']= ''
for i in incomplete_columns:
appa['ALL']= appa['ALL'] + appa[i]
appa['ALL'] = appa['ALL'].str.len()*1
appa['ALL'].fillna(0, inplace=True)
appa = appa.sort_values('ALL')
appa.drop_duplicates(keep='first',inplace = True)
return appa
def year(df= final_df,Before = False, value=1940):
if Before == True:
return df[df['Accession_Number_Year']<=value]
else:
return df[df['Accession_Number_Year']>=value]
# -
search(text='africa')
| Search/search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Exercise #1
#
# Date: 2021년 3월 22일
#
# ## Practice 1 - `pandas`
#
# 아래의 긴 `str ` 값을 `pandas.DataFrame`로 만드세요.
#
# * 조건 1: 파일로 생성해서 읽으면 안되요.
#
# ```python
# new_str = """name age email
# jang 40 <EMAIL>
# chris 30 <EMAIL>
# jinie 20 <EMAIL>
# simon 30 <EMAIL>
# sonia 20 <EMAIL>
# sooji 20 <EMAIL>
# """
# ```
#
# 잘 되었을 경우 아래와 같이 나올거에요.
#
# 
#
# ### Answer
#
# ```python
# import pandas as pd
# from io import StringIO
#
# df = pd.read_csv(StringIO(new_str), sep = "\s+")
# print(df)
# ```
#
#
#
# ## Practice 2 - `sort`
#
# Practice 1에서 사용했던 6개의 이메일 주소에 대해 알파벳 오름차순으로 정렬하세요.
#
# 다만, 먼저 성으로 정렬하고, 그 후 이름으로 정렬하세요.
#
# 잘 되었을 경우 아래와 같이 나올 거에요.
#
# * 힌트 1: `sort` (또는 `sorted`)를 2번 할 필요는 없어요.
#
# ```python
# answer = [
# '<EMAIL>',
# '<EMAIL>',
# '<EMAIL>',
# '<EMAIL>',
# '<EMAIL>',
# '<EMAIL>']
# ```
#
# ### Answer
#
# #### `re.split` 을 사용
#
# ```python
# import re
#
# l = list(df["email"])
#
# def get_name(addr):
# first, last, *_ = re.split("[\.@]", addr)
# return last, first
#
# sorted(l, key=get_name)
# ```
#
# ### `str.split` 을 사용
#
# ```python
# l = list(df["email"])
#
# def get_name(addr):
# name, company = addr.split("@")
# first, last = name.split(".")
# return last, first
#
# sorted(l, key=get_name)
# ```
#
#
#
| Python Exercise 1 (Answer).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # How to remove large intermediate files without breaking signatures
# + [markdown] kernel="SoS"
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
# * **Key points**:
# * `zapped` files have only their signatures
#
# + [markdown] kernel="SoS"
# ### Removal of large intermediate files
# + [markdown] kernel="SoS"
# SoS keep tracks of all intermediate files and will rerun steps only if any of the tracked files are removed or changed. However, it is often desired to remove some of the large non-essential intemediate files to reduce diskspace used by completed workflows, while allowing the workflow to be re-executed without these files. SoS provides a command
#
# ```
# sos remove files --zap
# ```
#
# to zap specified file, or for example
#
# ```
# sos remove . --size +5G --zap
# ```
# to zap all files larger than 5G. This command removes specified files but keeps a special `{file}.zapped` file with essential information (e.g. md5 signature, and size). SoS would consider a file exist when a `.zapped` file is present and will only regenerate the file if the actual file is needed for a later step.
# + [markdown] kernel="SoS"
# For example, let us execute a workflow with output `temp/result.txt`, and `temp/size.txt`.
# + kernel="SoS"
# %run -s force
[10]
output: "temp/result.txt"
# added comment
sh: expand=True
dd if=/dev/urandom of={_output} count=2000
[20]
output: 'temp/size.txt'
with open(_output[0], 'w') as sz:
sz.write(f"Modified {_input}: {os.path.getsize(_input[0])}\n")
# + [markdown] kernel="SoS"
# and let us zap the intermediate file `temp/result.txt`,
# + kernel="SoS"
!sos remove temp/result.txt --zap -y
!ls temp
# + [markdown] kernel="SoS"
# As you can see, `temp/result.txt` is replaced with `temp/result.txt.zapped`. Now if you rerun the workflow
# + kernel="SoS"
# %rerun -s default -v2
# + [markdown] kernel="SoS"
# ## Further reading
#
# *
| src/user_guide/zap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import json
import os
import requests
import sys
from gensim.utils import simple_preprocess
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Dense, Dropout, LSTM
from keras.utils import np_utils
from keras.callbacks import ModelCheckpoint
# -
path = './Lyrics_JSON'
# +
# Aggregating separate JSON lyric files
def gather_data(path_to_data):
data = []
for f in os.listdir(path):
if os.path.isdir(f) == False:
if f[-4:] == 'json':
with open(os.path.join(path, f)) as t:
#for lyrics in t['Lyrics']:
text = t.read().strip('\n')
data.append(str(text))
return data
# -
lyrics = gather_data(path)
# +
# def tokenize(text):
# return [token for token in simple_preprocess(text)]
# +
# with open('lyrics_aggregated.txt', 'w') as filehandle:
# filehandle.writelines('%s\n' % line for line in lyrics)
# -
text = (open('lyrics_aggregated.txt').read())
text = text.lower()
# +
# Simple pass at cleaning text
# text = text.replace('\n', '')
text = text.replace('{', '')
text = text.replace('}', '')
# text = text.replace("\", '')
text = text.replace('[', '')
text = text.replace(']', '')
text = text.replace('lyrics', '')
text = text.replace('title', '')
text = text.replace('"', '')
text = text.replace('genius', '')
# +
# Creating character/word mappings
# All unique characters/words are mapped to a number
characters = sorted(list(set(text)))
chars_to_int = dict((c, i) for i, c in enumerate(characters))
# -
n_chars = len(text)
n_vocab = len(characters)
print(f'total characters: {n_chars}, total vocab: {n_vocab}')
# +
# Training and target array for LSTM model
X = []
y = []
seq_length = 100
for i in range(0, n_chars - seq_length, 1):
sequence = text[i:i + seq_length]
label = text[i + seq_length]
X.append([chars_to_int[char] for char in sequence])
y.append(chars_to_int[label])
n_patterns = len(X)
print(f'total patterns: {n_patterns}')
# +
# Modifying array shapes for LSTM, transform y into one-hot encoded
X_modified = np.reshape(X, (n_patterns, seq_length, 1))
#normalize
X_modified = X_modified / float(n_vocab)
y_modified = np_utils.to_categorical(y)
# +
# Sequential model with two LSTM layers with 400 units each
# Dropoout layer to check for over-fitting
model = Sequential()
model.add(LSTM(256, input_shape=(X_modified.shape[1], X_modified.shape[2])))
model.add(Dropout(0.2))
model.add(Dense(y_modified.shape[1], activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam')
# +
# %%time
# Baseline model
model.fit(X_modified, y_modified, epochs=3, batch_size=128)
# model.save_weights('text_generator_400_0.2_400_0.2_baseline.h5')
# -
int_to_char = dict((i, c) for i, c in enumerate(characters))
# +
# Pick a random seed from text data
start = np.random.randint(0, len(X) - 1)
pattern = X[start]
print('Seed:')
print("\"", ''.join([int_to_char[value] for value in pattern])), "\""
# Generate characters
for i in range(500):
x = np.reshape(pattern, (1, len(pattern), 1))
x = x / float(n_vocab)
prediction = model.predict(x, verbose=0)
index = np.random.choice(len(prediction[0]), p=prediction[0])
result = int_to_char[index]
seq_in = [int_to_char[value] for value in pattern]
sys.stdout.write(result)
pattern.append(index)
pattern = pattern[1:len(pattern)]
# -
prediction
| notebooks/billie2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 1 Day 5
# +
#Write a program to identify sublist is there in the given list in the same order,
#if yes print "it's a match" if no print "it's gone" in function.
# +
list1=[1,3,5,7,2,4,6,8,9]
list2=[1,1,5]
i=0
while(i<=8):
if(len(list2)>0 and list1[i]==list2[0]):
list2.pop(0)
else:
break
i+=1
if(len(list2)==0):
print("it's a match")
else:
print("it's gone")
# +
list1=[1,1,5]
list2=[1,1,5]
i=0
while(i<=8):
if(len(list2)>0 and list1[i]==list2[0]):
list2.pop(0)
else:
break
i+=1
if(len(list2)==0):
print("it's a match")
else:
print("it's gone")
# -
# # Assignment 2 Day 5
# +
#Make a Function for Prime Numbers and use Filter to filter out all the prime numbers from 1-2500
# -
def prime(n):
for i in range(2,n):
if n % i == 0:
break
else:
return n
primeNo = filter(prime, range(1, 2500+1))
print("Prime numbers between 1-2500:",list(primeNo))
# # Assignment 3 Day 5
# +
#Make a lambda function for capitalizing the whole sentence passed using arguments
#And map all the sentences in the List,with tha lambda function
# -
#capitalizing whole sentence
lst = ["hey this is vighnatha","i am from hyderabad"]
print("Original list: ",lst)
result = list(map(lambda words: " ".join([word.upper() for word in words.split( )]) ,lst))
print("capitalized list:",result)
#capitalizing only the first letter of every word in a sentence
lst = ["<NAME> is vighnatha","i am from hyderabad"]
print("Original list: ",lst)
result = list(map(lambda words: " ".join([word.capitalize() for word in words.split( )]) ,lst))
print("capitalized list:",result)
| day 5 assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# ### Creating Data Frames
# documentation: http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html
#
# DataFrame is a 2-dimensional labeled data structure with columns of potentially different types. You can think of it
# like a spreadsheet or SQL table, or a dict of Series objects.
#
# You can create a data frame using:
# - Dict of 1D ndarrays, lists, dicts, or Series
# - 2-D numpy.ndarray
# - Structured or record ndarray
# - A Series
# - Another DataFrame
#
# ### Data Frame attributes
# | T | Transpose index and columns | |
# |---------|-------------------------------------------------------------------------------------------------------------------|---|
# | at | Fast label-based scalar accessor | |
# | axes | Return a list with the row axis labels and column axis labels as the only members. | |
# | blocks | Internal property, property synonym for as_blocks() | |
# | dtypes | Return the dtypes in this object. | |
# | empty | True if NDFrame is entirely empty [no items], meaning any of the axes are of length 0. | |
# | ftypes | Return the ftypes (indication of sparse/dense and dtype) in this object. | |
# | iat | Fast integer location scalar accessor. | |
# | iloc | Purely integer-location based indexing for selection by position. | |
# | is_copy | | |
# | ix | A primarily label-location based indexer, with integer position fallback. | |
# | loc | Purely label-location based indexer for selection by label. | |
# | ndim | Number of axes / array dimensions | |
# | shape | Return a tuple representing the dimensionality of the DataFrame. | |
# | size | number of elements in the NDFrame | |
# | style | Property returning a Styler object containing methods for building a styled HTML representation fo the DataFrame. | |
# | values | Numpy representation of NDFrame | |
import pandas as pd
import numpy as np
# ### Creating data frames from various data types
# documentation: http://pandas.pydata.org/pandas-docs/stable/dsintro.html
#
# cookbook: http://pandas.pydata.org/pandas-docs/stable/cookbook.html
# ##### create data frame from Python dictionary
my_dictionary = {'a' : 45., 'b' : -19.5, 'c' : 4444}
print(my_dictionary.keys())
print(my_dictionary.values())
# ##### constructor without explicit index
cookbook_df = pd.DataFrame({'AAA' : [4,5,6,7], 'BBB' : [10,20,30,40],'CCC' : [100,50,-30,-50]})
cookbook_df
# ##### constructor contains dictionary with Series as values
series_dict = {'one' : pd.Series([1., 2., 3.], index=['a', 'b', 'c']),
'two' : pd.Series([1., 2., 3., 4.], index=['a', 'b', 'c', 'd'])}
series_df = pd.DataFrame(series_dict)
series_df
# ##### dictionary of lists
produce_dict = {'veggies': ['potatoes', 'onions', 'peppers', 'carrots'],
'fruits': ['apples', 'bananas', 'pineapple', 'berries']}
produce_dict
# ##### list of dictionaries
data2 = [{'a': 1, 'b': 2}, {'a': 5, 'b': 10, 'c': 20}]
pd.DataFrame(data2)
# ##### dictionary of tuples, with multi index
pd.DataFrame({('a', 'b'): {('A', 'B'): 1, ('A', 'C'): 2},
('a', 'a'): {('A', 'C'): 3, ('A', 'B'): 4},
('a', 'c'): {('A', 'B'): 5, ('A', 'C'): 6},
('b', 'a'): {('A', 'C'): 7, ('A', 'B'): 8},
('b', 'b'): {('A', 'D'): 9, ('A', 'B'): 10}})
| python-scripts/data_analytics_learn/link_pandas/Ex_Files_Pandas_Data/Exercise Files/04_01/Begin/.ipynb_checkpoints/Create-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zXLKwrRBBYAD" colab_type="text"
# #Trabalho de Avaliação e Desempenho
#
# Grupo:<br>
# <NAME> - 115150099 <br>
# <NAME> - 116150432 <br>
# <NAME> - 116033119 <br>
# <NAME> - 114148170 <br>
# <NAME> - 116140788 <br>
# <NAME> - 116165607 <br>
#
# Professor(a): <NAME> <br>
# Período: 2019.2
# + [markdown] id="o2-iD4puBnl_" colab_type="text"
# ##Questão 4
# + [markdown] id="lc1OiC7mgvSl" colab_type="text"
# ### Código-base
# + id="fz754CCxBW0r" colab_type="code" colab={}
import numpy as np
import scipy as sp
import scipy.stats
import math
import time
import random
from prettytable import PrettyTable
import matplotlib.pyplot as plt
from decimal import *
class Eventos:
def __init__(self, tipo, fimEvento, duracaoServico):
self.tipo = tipo
self.fimEvento = fimEvento #Do ponto de vista do lambda é a chegada na fila e do ponto de vista de mi é o tempo de sair do servidor.
self.duracaoServico = duracaoServico
def simulaMG1 (λ1, λ2, µ1, µ2, nEventos, X = random.expovariate, tempoTotalSimulacao = 5000): # Simula uma fila M/G/1, retornando o E(N)
linhaDoTempo = 0
#tempoUltimoEvento = 0
area = 0
N0 = 0
T0 = 0
numeroClienteT1 = 0
numeroClienteT2 = 0
numeroTotalCliente = 0
tempTotalServico = 0
fila = [] #Fila. OBS: A primeira posição representa o servidor, logo o cliente que ocupar tal posição estar no servidor.
eventos = [] #Fila de eventos
atualEvento = 0
#Primeira ocorrência de eventos (λ1 e λ2)
temp = random.expovariate(λ1)
eventos.append(Eventos("Chegada1",temp , temp))
if(λ2 != 0):
temp = random.expovariate(λ2)
eventos.append(Eventos("Chegada2", temp, temp))
while(linhaDoTempo <= tempoTotalSimulacao):
#eventos = sorted(eventos,key=attrgetter('fimEvento'))
eventos.sort(key=lambda eventos: eventos.fimEvento)
#print(eventos)
atualEvento = eventos[0] #Pega o primeiro evento
eventos = eventos[1:] #Tira o primeiro evento da fila de eventos
linhaDoTempo = atualEvento.fimEvento #Avança a linha do tempo
filaAux = fila[1:] #Não podemos considerar a posição 0 na ordenação, uma vez que a mesma repesenta o servidor.
filaAux.sort()
for i in range(0,len(filaAux)):
fila[i+1] = filaAux[i]
if(atualEvento.tipo == "Chegada1"):
numeroClienteT1 += 1
fila.append(1)
temp = random.expovariate(λ1)
eventos.append(Eventos("Chegada1", linhaDoTempo + temp, temp))
if(len(fila) == 1):
temp = X(µ1)
eventos.append(Eventos("Servico1", linhaDoTempo + temp, temp))
numeroClienteT1 -= 1
elif(atualEvento.tipo == "Chegada2"):
numeroClienteT2 += 1
fila.append(2)
temp = random.expovariate(λ2)
eventos.append(Eventos("Chegada2", linhaDoTempo +temp, temp))
#temp = random.expovariate(λ1)
#eventos.append(Eventos("Chegada1", linhaDoTempo + temp, temp))
if(len(fila) == 1):
temp = X(µ2)
eventos.append(Eventos("Servico2", linhaDoTempo + temp, temp))
numeroClienteT2 -= 1
elif(atualEvento.tipo == "Servico1"):
fila = fila[1:]
numeroTotalCliente += 1
tempTotalServico += atualEvento.duracaoServico
if(len(fila) != 0 and fila[0] == 1):
temp = X(µ1)
eventos.append(Eventos("Servico1", linhaDoTempo + temp, temp))
elif(µ2 != 0 and len(fila) != 0 and fila[0] == 2 ):
temp = X(µ2)
eventos.append(Eventos("Servico2", linhaDoTempo + temp, temp))
elif(atualEvento.tipo == "Servico2"):
fila = fila[1:]
numeroTotalCliente += 1
tempTotalServico += atualEvento.duracaoServico
if(len(fila) != 0 and fila[0] == 1):
temp = X(µ1)
eventos.append(Eventos("Servico1", linhaDoTempo + temp, temp))
elif(µ2 != 0 and len(fila) != 0 and fila[0] == 2 ):
temp = X(µ2)
eventos.append(Eventos("Servico2", linhaDoTempo + temp, temp))
if(len(fila) != N0):
if(N0 > 0 ):
area = area + (N0 - 1)*(linhaDoTempo - T0)
N0 = len(fila)
T0 = linhaDoTempo
return area/linhaDoTempo, numeroTotalCliente, (tempTotalServico/numeroTotalCliente), (tempTotalServico/linhaDoTempo) #Retorna [Nq], N, [X], ρ
def simulaGeral(λ1, λ2, µ1, µ2, nSimulacoes, X = random.expovariate):
# Array para o resultado de cada uma das simulações
Nq_barras = []
W_barras = []
X_barras = []
T_barras = []
N_s = []
ρ_s = []
# Médias das simulações
media_simus_Nq = 0
media_simus_W = 0
media_simus_X = 0
media_simus_N = 0
media_simus_ρ = 0
media_simus_T = 0
desvio_simus_Nq = 0
desvio_simus_W = 0
desvio_simus_X = 0
desvio_simus_N = 0
desvio_simus_ρ = 0
desvio_simus_T = 0
for i in range(nSimulacoes): #Realiza n Simulacões.
NQ_barra, N, X_barra, ρ = simulaMG1(λ1, λ2, µ1, µ2, 1000, X) #Simulações com 1000 eventos cada
Nq_barras.append(NQ_barra)
N_s.append(N)
X_barras.append(X_barra)
W_barras.append(Nq_barras[i]/(λ1+λ2))
ρ_s.append(ρ)
T_barras.append(W_barras[i] + X_barras[i])
#Média Para Nq_barra de todas as simulações
media_simus_Nq = np.array(Nq_barras).mean()
desvio_simus_Nq = np.std(np.array(Nq_barras))
#intervaloConf_Nq = sp.stats.norm.interval(0.95, loc=media_simus_Nq, scale=desvio_simus_Nq)
intervaloConf_Nq = []
intervaloConf_Nq.append(media_simus_Nq - 1.96*(desvio_simus_Nq/math.sqrt(len(Nq_barras))))
intervaloConf_Nq.append(media_simus_Nq + 1.96*(desvio_simus_Nq/math.sqrt(len(Nq_barras))))
#Média Para W_barra de todas as simulações
media_simus_W = np.array(W_barras).mean()
desvio_simus_W = np.std(np.array(W_barras))
#intervaloConf_W = sp.stats.norm.interval(0.95, loc=media_simus_W, scale=desvio_simus_W)
intervaloConf_W = []
intervaloConf_W.append(media_simus_W - 1.96*(desvio_simus_W/math.sqrt(len(W_barras))))
intervaloConf_W.append(media_simus_W + 1.96*(desvio_simus_W/math.sqrt(len(W_barras))))
#Média para N (Número de clientes) de todas as simulações
media_simus_N = np.array(N_s).mean()
desvio_simus_N = np.std(np.array(N_s))
#intervaloConf_N = sp.stats.norm.interval(0.95, loc=media_simus_N, scale=desvio_simus_N)
intervaloConf_N = []
intervaloConf_N.append(media_simus_N - 1.96*(desvio_simus_N/math.sqrt(len(N_s))))
intervaloConf_N.append(media_simus_N + 1.96*(desvio_simus_N/math.sqrt(len(N_s))))
#Média para X_barra (Tempo de serviço médio) de todas as simulações
media_simus_X = np.array(X_barras).mean()
desvio_simus_X = np.std(np.array(X_barras))
#intervaloConf_X = sp.stats.norm.interval(0.95, loc=media_simus_X, scale=desvio_simus_X)
intervaloConf_X = []
intervaloConf_X.append(media_simus_X - 1.96*(desvio_simus_X/math.sqrt(len(X_barras))))
intervaloConf_X.append(media_simus_X + 1.96*(desvio_simus_X/math.sqrt(len(X_barras))))
#Média para ρ (Utilização) de todas as simulações
media_simus_ρ = np.array(ρ_s).mean()
desvio_simus_ρ = np.std(np.array(ρ_s))
#intervaloConf_ρ = sp.stats.norm.interval(0.95, loc=media_simus_ρ, scale=desvio_simus_ρ)
intervaloConf_ρ = []
intervaloConf_ρ.append(media_simus_ρ - 1.96*(desvio_simus_ρ/math.sqrt(len(ρ_s))))
intervaloConf_ρ.append(media_simus_ρ + 1.96*(desvio_simus_ρ/math.sqrt(len(ρ_s))))
#Média para T_barra(Tempo médio no sistema) de todas as simulações
media_simus_T = np.array(T_barras).mean()
desvio_simus_T = np.std(np.array(T_barras))
#intervaloConf_T = sp.stats.norm.interval(0.95, loc=media_simus_T, scale=desvio_simus_T)
intervaloConf_T = []
intervaloConf_T.append(media_simus_T - 1.96*(desvio_simus_T/math.sqrt(len(T_barras))))
intervaloConf_T.append(media_simus_T + 1.96*(desvio_simus_T/math.sqrt(len(T_barras))))
return [intervaloConf_Nq, media_simus_Nq, intervaloConf_W, media_simus_W, media_simus_N, media_simus_X, media_simus_ρ, media_simus_T, intervaloConf_N, intervaloConf_X, intervaloConf_ρ, intervaloConf_T]
# + id="B2FwSY0Du1Ic" colab_type="code" outputId="8f9b6074-a47e-4a84-b3ea-8f9dc769e6e8" executionInfo={"status": "ok", "timestamp": 1577141470800, "user_tz": 180, "elapsed": 1817, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
def Ns_barra(λ, X_barra):
return λ*X_barra
def X_barra(µ = 0, a = 0, b = 0, tipoServico = 'e'):
if tipoServico == 'e':
return 1/µ
elif tipoServico == 'u':
return (a+b)/2
else:
return 1/µ
def X2_barra(µ = 0, a = 0, b = 0, tipoServico = 'e'):
if tipoServico == 'e':
return 2/(pow(µ,2))
elif tipoServico == 'u':
return ( pow(a, 2) + a*b + pow(b,2) )/3
else:
return 1/(pow(µ,2))
#def Xr_barra(λ, µ, a, b, tipoServico='e'):
# return (2* X2_barra(µ, tipoServico) ) / X_barra(µ, a, b, tipoServico)
def Xr_barra(λ1, µ1, λ2, µ2, a1 = 0, b1 = 0, a2 = 0, b2 = 0, tipoServico='e'):
return (1/2)*( λ1* X2_barra(µ1, a1, b1, tipoServico) + λ2* X2_barra(µ2, a2, b2, tipoServico) )
def P(tipo, λ1, λ2):
if (tipo == 1):
return λ1/ (λ1 + λ2)
else:
return λ2/ (λ1 + λ2)
def Nq_barra(λ = 0, µ = 0, tipoServico = 'e'):
if (tipoServico == 'e'):
ρ = λ / µ
return pow(ρ,2) / (1-ρ)
else:
return 0
def ρ_geral(λ1, µ1, λ2, µ2):
return (λ1/µ1) + (λ2/µ2)
def W_barra(cliente, λ1, µ1, λ2, µ2, a1 = 0, b1 = 0, a2 = 0, b2 = 0, tipoServico = 'e'):
ρ1 = λ1*X_barra(µ1,a1,b1,tipoServico)
ρ2 = λ2*X_barra(µ2,a2,b2,tipoServico)
if(cliente == 1):
return Xr_barra(λ1, µ1, λ1, µ1, a1, b1, a2, b2, tipoServico)/(1-ρ1)
else:
if (ρ1+ρ2 != 1):
ρ = ρ1+ρ2
else:
ρ = 0.999
return Xr_barra(λ1, µ1, λ1, µ1, a1, b1, a2, b2, tipoServico)/( (1-ρ1)*(1-ρ) )
def calculoAnalitico(λ1 = 0, λ2 = 0, µ1 = 0, µ2 = 0, a1 = 0, b1 = 0, a2 = 0, b2 = 0, tipoServico = 'e'):
W1 = W_barra(1, λ1, µ1, λ2, µ2, a1, b1, a2, b2, tipoServico)
W2 = W_barra(2, λ1, µ1, λ2, µ2, a1, b1, a2, b2,tipoServico)
W = W1*P(1, λ1, λ2) + W2*P(2, λ1, λ2)
Nq1 = λ1*W1
Nq2 = λ2*W2
#Nq2 = (λ1 + λ2)*W
Nq = Nq1 + Nq2
return [Nq, W]
print(calculoAnalitico(0.05, 0.2, 1, 0.5, 0, 0, 'e'))
# + id="8VpFYqwhCX6B" colab_type="code" colab={}
#Imprimir os gráficos
def printGraficoQ3(cenario, result, maxXsticks):
ultimoNqCadeia = -1
ultimoNqSimu = result[1][len(result[1])-1]
ultimoWCadeia = -1
ultimoWSimu = result[3][len(result[3])-1]
plt.figure(figsize= [20, 20])
plt.subplot(211)
plt.title ("λ1 x número de clientes no cenário " + str(cenario) )
plt.bar(result[0], result[1], width = 0.01, color = 'blue', yerr = result[2], capsize=7, edgecolor = 'black', align='edge', label='Simulação')
#if(cenario != 4):
# resultAnalitico = result[4][:len(result[4])-1]
# resultAnalitico.append(result[1][len(result[1])-1])
#else:
# resultAnalitico = result[4]
plt.bar([ x+0.01 for x in result[0] ], result[4] , width = 0.01, color = 'cyan', edgecolor = 'black', align='edge', label='Analítico')
if(cenario != 3):
plt.bar([ x+0.02 for x in result[0] ], result[len(result)-2], width = 0.01, color = 'red', edgecolor = 'black', align='edge', label='Cadeia de Markov')
ultimoNqCadeia = result[len(result)-2][len(result[len(result)-2])-1]
if(cenario != 4):
plt.xticks( np.array(range(5, maxXsticks, 5)) * 0.01 )
else:
plt.xticks( [0.08])
plt.xlabel('λ1', fontsize=15)
plt.ylabel('E(Nq)', fontsize=15)
axesNq = plt.gca()
axesNq.set_ylim([0,max(ultimoNqCadeia, ultimoNqSimu)+10])
plt.legend()
plt.subplot(212)
plt.title ("λ1 x tempo médio de clientes na fila de espera do cenário " + str(cenario) )
plt.bar(result[0], result[3], width = 0.01, color = 'blue', yerr = result[5], capsize=7, edgecolor = 'black', align='edge', label='Simulação')
#if(cenario != 1):
# if(cenario == 2 or cenario == 3):
# resultAnalitico = result[14][:len(result[14])-1]
# resultAnalitico.append(result[3][len(result[3])-1])
# else:
# resultAnalitico = result[4]
plt.bar([ x+0.01 for x in result[0] ], result[14] , width = 0.01, color = 'cyan', edgecolor = 'black', align='edge', label='Analítico')
if(cenario != 3):
plt.bar([ x+0.02 for x in result[0] ], result[len(result)-1], width = 0.01, color = 'red', edgecolor = 'black', align='edge', label='Cadeia de Markov')
ultimoWCadeia = result[len(result)-1][len(result[len(result)-1])-1]
if(cenario != 4):
plt.xticks( np.array(range(5, maxXsticks, 5)) * 0.01 )
else:
plt.xticks( [0.08])
#plt.bar(result[0], result[3], width = 0.03, , edgecolor = 'black')
plt.xlabel('λ1', fontsize=15)
plt.ylabel('E(W)', fontsize=15)
axesNq = plt.gca()
axesNq.set_ylim([0,max(ultimoWCadeia, ultimoWSimu)+10])
plt.legend()
plt.show()
# + [markdown] id="tRIu0GFREqp_" colab_type="text"
# ### Cenário 2
# + [markdown] id="bZS_AL10EzN6" colab_type="text"
# #### Simulação e solução analítica
# + id="gtvR23sWE_lB" colab_type="code" outputId="3d9e666b-85ec-4dca-d44b-2f6c382ca9fa" executionInfo={"status": "ok", "timestamp": 1577141696703, "user_tz": 180, "elapsed": 227692, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 311}
from prettytable import PrettyTable
def simulaCenario2():
medias_Nq = []
medias_NqAnalitico = []
medias_WAnalitico = []
medias_W = []
medias_N = []
medias_X = []
medias_T = []
ρs = []
λs = []
confsNq = []
confsW = []
confsN = []
confsX = []
confsρ = []
confsT = []
resultCenario2 = []
for i in range(5, 65, 5): # Para cada λ1 do cenário 1
λ = 0.01 * i # Gera o λ1
λs.append(λ) # Guarda os λ num array
result = simulaGeral(λ , 0.2, 1, 0.5, 1000) # Faz a simulação para cada λ
resultAnal = calculoAnalitico(λ, 0.2, 1, 0.5, 'e')
medias_Nq.append(result[1]) # Guarda o valor do E(Nq) para cada λ
medias_W.append(result[3]) # Guarda o valor do E(W) para cada λ
medias_N.append(result[4])
medias_X.append(result[5])
ρs.append(result[6])
medias_T.append(result[7])
medias_NqAnalitico.append(resultAnal[0]) # Guarda o valor do E(Nq) usando Little para cada λ
medias_WAnalitico.append(resultAnal[1])
confsNq.append(result[1] - result[0][0]) # Guarda o valor da diferença entre a média e um dos extremos do intervalo de confiança para cada λ
confsW.append(result[3] - result[2][0])
confsN.append(result[4] - result[8][0])
confsX.append(result[5] - result[9][0])
confsρ.append(result[6] - result[10][0])
confsT.append(result[7] - result[11][0])
# Salva tudo num array geral
resultCenario2.append(λs)
resultCenario2.append(medias_Nq)
resultCenario2.append(confsNq)
resultCenario2.append(medias_W)
resultCenario2.append(medias_NqAnalitico)
resultCenario2.append(confsW)
resultCenario2.append(medias_N)
resultCenario2.append(medias_X)
resultCenario2.append(ρs)
resultCenario2.append(medias_T)
resultCenario2.append(confsN)
resultCenario2.append(confsX)
resultCenario2.append(confsρ)
resultCenario2.append(confsT)
resultCenario2.append(medias_WAnalitico)
return resultCenario2
resultCenario2 = simulaCenario2()
#print(resultCenario1[6])
tabelaCenario2 = PrettyTable()
tabelaCenario2.header = True
nomes_colunas = ['λ', 'Nº de clientes; (Intervalo de confiança)', 'E[X]; (Intervalo de confiança)', 'E[W]; (Intervalo de confiança)', 'E[T]; (Intervalo de confiança)', 'ρ; (Intervalo de confiança)', 'E[Nq]; (Intervalo de confiança)']
tabelaCenario2.add_column(nomes_colunas[0], [ round(lambd,2) for lambd in resultCenario2[0] ], align='c')
tabelaCenario2.add_column(nomes_colunas[1], [ f"{round(resultCenario2[6][i],5)}; ({round(resultCenario2[6][i]-resultCenario2[10][i],5)}, {round(resultCenario2[6][i]+resultCenario2[10][i],5)})" for i in range(len(resultCenario2[6])) ], align='c' )
tabelaCenario2.add_column(nomes_colunas[2], [ f"{round(resultCenario2[7][i],5)}; ({round(resultCenario2[7][i]-resultCenario2[11][i],5)}, {round(resultCenario2[7][i]+resultCenario2[11][i],5)})" for i in range(len(resultCenario2[7])) ], align='c' )
tabelaCenario2.add_column(nomes_colunas[3], [ f"{round(resultCenario2[3][i],5)}; ({round(resultCenario2[3][i]-resultCenario2[5][i],5)}, {round(resultCenario2[3][i]+resultCenario2[5][i],5)})" for i in range(len(resultCenario2[3])) ], align='c' )
tabelaCenario2.add_column(nomes_colunas[4], [ f"{round(resultCenario2[9][i],5)}; ({round(resultCenario2[9][i]-resultCenario2[13][i],5)}, {round(resultCenario2[9][i]+resultCenario2[13][i],5)})" for i in range(len(resultCenario2[9])) ], align='c' )
tabelaCenario2.add_column(nomes_colunas[5], [ f"{round(resultCenario2[8][i],5)}; ({round(resultCenario2[8][i]-resultCenario2[12][i],5)}, {round(resultCenario2[8][i]+resultCenario2[12][i],5)})" for i in range(len(resultCenario2[8])) ], align='c' )
tabelaCenario2.add_column(nomes_colunas[6], [ f"{round(resultCenario2[1][i],5)}; ({round(resultCenario2[1][i]-resultCenario2[2][i],5)}, {round(resultCenario2[1][i]+resultCenario2[2][i],5)})" for i in range(len(resultCenario2[1])) ], align='c' )
print(tabelaCenario2.get_string(title="Resultados do Cenário 2 - Simulação"))
#tabela2Cenario2 = PrettyTable()
#nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]']
#tabela2Cenario2.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario2[0] ])
#tabela2Cenario2.add_column(nomes_colunas2[1], [ f"{round(resultCenario2[1][i],5)}; ({round(resultCenario2[1][i]-resultCenario2[2][i],5)}, {round(resultCenario2[1][i]+resultCenario2[2][i],5)})" for i in range(len(resultCenario2[1])) ])
#tabela2Cenario2.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario2[4] ])
#print(tabela2Cenario2.get_string(title="Resultados do Cenário 2 - Simulação e solução analítica"))
#tabela3Cenario2 = PrettyTable()
#nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]']
#tabela3Cenario2.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario2[0] ])
#tabela3Cenario2.add_column(nomes_colunas3[1], [ f"{round(resultCenario2[3][i],5)}; ({round(resultCenario2[3][i]-resultCenario2[5][i],5)}, {round(resultCenario2[3][i]+resultCenario2[5][i],5)})" for i in range(len(resultCenario2[3])) ], align='c')
#tabela3Cenario2.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario2[14] ])
#print(tabela3Cenario2.get_string(title="Resultados do Cenário 2 - Simulação e solução analítica"))
# + [markdown] id="KRYxsWYPE4J7" colab_type="text"
# #### <NAME>
# FILA<br>
# <img src = "https://drive.google.com/uc?export=view&id=1sZVDH7J9uje0wVbw0TRpsmBJa1CRmXVI"/>
#
# <br>
# CADEIA <br>
# <img src = "https://drive.google.com/uc?export=view&id=1NpVq1SO804c8Fhcz7FzC_Mio7PhgfPFJ"/>
# + id="Kk7kGsAkFAKF" colab_type="code" outputId="1cfc3336-9363-48a4-dfd6-94852246e2ec" executionInfo={"status": "ok", "timestamp": 1577141732275, "user_tz": 180, "elapsed": 263251, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/"}
import pandas as pd
import numpy as np
from numpy.linalg import matrix_power
from numpy.linalg import norm
import numpy.linalg as lin
def check_state(f1, f2, serving):
if f1 == 0 and f2 == 0 and serving != 0:
raise Exception("invalido: se sistema está vazio, não pode ter ninguém sendo servido")
if f1 > 0 and f2 > 0 and serving != 1 and serving != 2:
raise Exception("invalido: se sistema está ocupado, tem que estar sevindo alguém")
if f1 > 0 and f2 == 0 and serving != 1:
raise Exception("invalido: se só há clientes do tipo 1, então tem que estar servindo o tipo 1")
if f1 == 0 and f2 > 0 and serving != 2:
raise Exception("invalido: se só há clientes do tipo 2, então tem que estar servindo o tipo 2")
def get_state_name(f1, f2, serving):
check_state(f1, f2, serving)
s1 = "*" if serving == 1 else " "
s2 = "*" if serving == 2 else " "
return f"{f1:5}{s1}/{f2:5}{s2}"
def get_next_states(f1, f2, serving, λ1, λ2, µ1, µ2):
check_state(f1, f2, serving)
r = {}
# Chegada de um cliente do tipo 1:
if serving == 0:
r[get_state_name(f1+1,f2,1)] = λ1
else:
r[get_state_name(f1+1,f2,serving)] = λ1
# Chegada de um cliente do tipo 2:
if f1 == 0:
r[get_state_name(0,f2+1,2)] = λ2
else:
r[get_state_name(f1,f2+1,serving)] = λ2
# Saida de um cliente do tipo 1:
if serving == 1:
if f1 > 1:
r[get_state_name(f1-1,f2,1)] = µ1
elif f2 > 0:
r[get_state_name(0,f2,2)] = µ1
else:
r[get_state_name(0,0,0)] = µ1
# Saida de um cliente do tipo 2:
if serving == 2:
if f1 > 0:
r[get_state_name(f1,f2-1,1)] = µ2
elif f2 > 1:
r[get_state_name(0,f2-1,2)] = µ2
else:
r[get_state_name(0,0,0)] = µ2
return r
def create_ctmc_states(size, λ1, λ2, µ1, µ2):
states = {}
# adding state " 0 / 0 "
states[get_state_name(0,0,0)] = get_next_states(0,0,0, λ1, λ2, µ1, µ2)
for f1 in range(0,size):
for f2 in range(0,size):
if f1 == 0 and f2 == 0:
serve_list = []
elif f1 == 0:
serve_list = [2]
elif f2 == 0:
serve_list = [1]
else:
serve_list = [1,2]
for serving in serve_list:
states[get_state_name(f1,f2,serving)] = get_next_states(f1,f2,serving, λ1, λ2, µ1, µ2)
return states
def create_ctmc_matrix(size, λ1, λ2, µ1, µ2):
states = create_ctmc_states(size, λ1, λ2, µ1, µ2)
df = pd.DataFrame(states)
df = df.reindex(sorted(df.columns), axis=1)
df = df.sort_index()
df = df.transpose()
df = df[df.index]
columns = df.columns
np.fill_diagonal(df.values, -df.sum(axis=1))
df = df.fillna(0)
return df.to_numpy(), columns
# Calcula os π
def calculaCadeiaMarkovCenario2(λ1, λ2, µ1, µ2):
result = create_ctmc_matrix(40, λ1, λ2, µ1, µ2)
Q = result[0]
columns = result[1]
# Trecho a seguir calcula os π
sz = Q.shape[0]
Qt = Q.transpose()
Qt[sz-1,:] = 1
b = np.zeros((sz,1))
b[sz-1] = 1
x = lin.solve(Qt.T.dot(Qt), Qt.T.dot(b))
πi = {}
# Agrupa os πi que apresentam o mesmo número de pessoas no sistema para um único π
for i in range(len(columns)):
txt = columns[i].split(",")
for k in range(len(txt)):
txt2 = txt[k].split("/")
t1 = txt2[0].replace("*","")
t2 = txt2[1].replace("*","")
txt2 = (t1, t2)
numberCustumerSystem = int(txt2[0])+int(txt2[1])
if numberCustumerSystem in πi:
πi[numberCustumerSystem] = x[i] + πi[numberCustumerSystem]
else:
πi[numberCustumerSystem] = x[i]
return list(πi.values())
def cadeiaMarkovCenario2():
Nqs = []
W = []
Nq = 0
for i in range(5, 65, 5):
λ1 = 0.01 * i
λ2 = 0.2
µ1 = 1
µ2 = 0.5
πk = (calculaCadeiaMarkovCenario2(λ1, λ2, µ1, µ2))
Nq = 0
# Faz os cálculos dos Nq e W
for j in range(1,len(πk)):
#print(len(πk))
if(πk[j] < 0):
Nq += 0
else:
Nq += (j-1)*πk[j]
Nqs.append(Nq[0])
W.append(Nq[0]/(λ1+λ2))
return [Nqs, W]
resultCMCenario2 = cadeiaMarkovCenario2()
print(resultCMCenario2[0])
print(resultCMCenario2[1])
# + [markdown] id="32G1x5HsE5_M" colab_type="text"
# #### Comparação
# + colab_type="code" id="dXNkWdIQOiNa" outputId="2bb12832-0b20-4538-95cf-be9f952bf0a4" executionInfo={"status": "ok", "timestamp": 1577141732281, "user_tz": 180, "elapsed": 263239, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 605}
# Tabela 2
tabela2Cenario2 = PrettyTable()
nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]', 'Cadeia de Markov: E[Nq]']
tabela2Cenario2.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario2[0] ])
tabela2Cenario2.add_column(nomes_colunas2[1], [ f"{round(resultCenario2[1][i],5)}; ({round(resultCenario2[1][i]-resultCenario2[2][i],5)}, {round(resultCenario2[1][i]+resultCenario2[2][i],5)})" for i in range(len(resultCenario2[1])) ])
tabela2Cenario2.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario2[4] ])
tabela2Cenario2.add_column(nomes_colunas2[3], [ round(item,5) for item in resultCMCenario2[0] ] )
print(tabela2Cenario2.get_string(title="Resultados do Cenário 2 - Simulação, solução analítica e cadeia"))
# Tabela 3
tabela3Cenario2 = PrettyTable()
nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]', 'Cadeia de Markov: E[W]']
tabela3Cenario2.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario2[0] ])
tabela3Cenario2.add_column(nomes_colunas3[1], [ f"{round(resultCenario2[3][i],5)}; ({round(resultCenario2[3][i]-resultCenario2[5][i],5)}, {round(resultCenario2[3][i]+resultCenario2[5][i],5)})" for i in range(len(resultCenario2[3])) ])
tabela3Cenario2.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario2[14] ])
tabela3Cenario2.add_column(nomes_colunas3[3], [ round(item,5) for item in resultCMCenario2[1] ])
print(tabela3Cenario2.get_string(title="Resultados do Cenário 2 - Simulação, solução analítica e cadeia"))
# + id="b9T4Bf5kFAzH" colab_type="code" outputId="b2ce2cb8-9634-4917-9590-da3694b77ae7" executionInfo={"status": "ok", "timestamp": 1577141733504, "user_tz": 180, "elapsed": 264438, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
resultCenario2.append(resultCMCenario2[0])
resultCenario2.append(resultCMCenario2[1])
printGraficoQ3(2, resultCenario2, 65)
# + [markdown] id="5-8Gxp_cLuh_" colab_type="text"
# ### Cenário 3
# + [markdown] id="iNlu9fLVLyNr" colab_type="text"
# #### Simulação e solução analítica
# + id="9KvRyTU4L6fl" colab_type="code" outputId="5d422fa8-b803-48e6-bd6d-289a048c3e81" executionInfo={"status": "ok", "timestamp": 1577141935508, "user_tz": 180, "elapsed": 466424, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 311}
def deterministica(L):
return 1/L
def simulaCenario3():
medias_Nq = []
medias_NqAnalitico = []
medias_WAnalitico = []
medias_W = []
medias_N = []
medias_X = []
medias_T = []
ρs = []
λs = []
confsNq = []
confsW = []
confsN = []
confsX = []
confsρ = []
confsT = []
resultCenario3 = []
for i in range(5, 65, 5): # Para cada λ1 do cenário 1
λ = 0.01 * i # Gera o λ1
λs.append(λ) # Guarda os λ num array
result = simulaGeral(λ , 0.2, 1, 0.5, 1000, deterministica) # Faz a simulação para cada λ
resultAnal = calculoAnalitico(λ,0.2,1,0.5,0,0,'d')
medias_Nq.append(result[1]) # Guarda o valor do E(Nq) para cada λ
medias_W.append(result[3]) # Guarda o valor do E(W) para cada λ
medias_N.append(result[4])
medias_X.append(result[5])
ρs.append(result[6])
medias_T.append(result[7])
medias_NqAnalitico.append(resultAnal[0]) # Guarda o valor do E(Nq) usando Little para cada λ
confsNq.append(result[1] - result[0][0]) # Guarda o valor da diferença entre a média e um dos extremos do intervalo de confiança para cada λ
confsW.append(result[3] - result[2][0])
confsN.append(result[4] - result[8][0])
confsX.append(result[5] - result[9][0])
confsρ.append(result[6] - result[10][0])
confsT.append(result[7] - result[11][0])
medias_WAnalitico.append(resultAnal[1])
# Salva tudo num array geral
resultCenario3.append(λs)
resultCenario3.append(medias_Nq)
resultCenario3.append(confsNq)
resultCenario3.append(medias_W)
resultCenario3.append(medias_NqAnalitico)
resultCenario3.append(confsW)
resultCenario3.append(medias_N)
resultCenario3.append(medias_X)
resultCenario3.append(ρs)
resultCenario3.append(medias_T)
resultCenario3.append(confsN)
resultCenario3.append(confsX)
resultCenario3.append(confsρ)
resultCenario3.append(confsT)
resultCenario3.append(medias_WAnalitico)
return resultCenario3
resultCenario3 = simulaCenario3()
tabelaCenario3 = PrettyTable()
nomes_colunas = ['λ', 'Nº de clientes; (Intervalo de confiança)', 'E[X]; (Intervalo de confiança)', 'E[W]; (Intervalo de confiança)', 'E[T]; (Intervalo de confiança)', 'ρ; (Intervalo de confiança)', 'E[Nq]; (Intervalo de confiança)']
tabelaCenario3.add_column(nomes_colunas[0], [ round(lambd,2) for lambd in resultCenario3[0] ], align='c')
tabelaCenario3.add_column(nomes_colunas[1], [ f"{round(resultCenario3[6][i],5)}; ({round(resultCenario3[6][i]-resultCenario3[10][i],5)}, {round(resultCenario3[6][i]+resultCenario3[10][i],5)})" for i in range(len(resultCenario3[6])) ], align='c' )
tabelaCenario3.add_column(nomes_colunas[2], [ f"{round(resultCenario3[7][i],5)}; ({round(resultCenario3[7][i]-resultCenario3[11][i],5)}, {round(resultCenario3[7][i]+resultCenario3[11][i],5)})" for i in range(len(resultCenario3[7])) ], align='c' )
tabelaCenario3.add_column(nomes_colunas[3], [ f"{round(resultCenario3[3][i],5)}; ({round(resultCenario3[3][i]-resultCenario3[5][i],5)}, {round(resultCenario3[3][i]+resultCenario3[5][i],5)})" for i in range(len(resultCenario3[3])) ], align='c' )
tabelaCenario3.add_column(nomes_colunas[4], [ f"{round(resultCenario3[9][i],5)}; ({round(resultCenario3[9][i]-resultCenario3[13][i],5)}, {round(resultCenario3[9][i]+resultCenario3[13][i],5)})" for i in range(len(resultCenario3[9])) ], align='c' )
tabelaCenario3.add_column(nomes_colunas[5], [ f"{round(resultCenario3[8][i],5)}; ({round(resultCenario3[8][i]-resultCenario3[12][i],5)}, {round(resultCenario3[8][i]+resultCenario3[12][i],5)})" for i in range(len(resultCenario3[8])) ], align='c' )
tabelaCenario3.add_column(nomes_colunas[6], [ f"{round(resultCenario3[1][i],5)}; ({round(resultCenario3[1][i]-resultCenario3[2][i],5)}, {round(resultCenario3[1][i]+resultCenario3[2][i],5)})" for i in range(len(resultCenario3[1])) ], align='c' )
print(tabelaCenario3.get_string(title="Resultados do Cenário 3 - Simulação"))
#tabela2Cenario3 = PrettyTable()
#nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]']
#tabela2Cenario3.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario3[0] ])
#tabela2Cenario3.add_column(nomes_colunas2[1], [ f"{round(resultCenario3[1][i],5)}; ({round(resultCenario3[1][i]-resultCenario3[2][i],5)}, {round(resultCenario3[1][i]+resultCenario3[2][i],5)})" for i in range(len(resultCenario3[1])) ])
#tabela2Cenario3.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario3[4] ])
#print(tabela2Cenario3.get_string(title="Resultados do Cenário 3 - Simulação e solução analítica"))
#tabela3Cenario3 = PrettyTable()
#nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]']
#tabela3Cenario3.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario3[0] ])
#tabela3Cenario3.add_column(nomes_colunas3[1], [ f"{round(resultCenario3[3][i],5)}; ({round(resultCenario3[3][i]-resultCenario3[5][i],5)}, {round(resultCenario3[3][i]+resultCenario3[5][i],5)})" for i in range(len(resultCenario3[3])) ], align='c')
#tabela3Cenario3.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario3[14] ])
#print(tabela3Cenario3.get_string(title="Resultados do Cenário 2 - Simulação e solução analítica"))
# + [markdown] id="IIZtQUYdL0_y" colab_type="text"
# ####Cadeia de Markov
# + id="t1sMCgAnL7RK" colab_type="code" colab={}
# + [markdown] id="-tfftf7kL2Mw" colab_type="text"
# #### Comparação
# + colab_type="code" id="s3tD8b_2Q31V" outputId="70863927-cd8f-421a-d50f-d12de493b9ac" executionInfo={"status": "ok", "timestamp": 1577141935511, "user_tz": 180, "elapsed": 466398, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 605}
# Tabela 2
tabela2Cenario3 = PrettyTable()
#nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]', 'Cadeia de Markov: E[Nq]']
nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]']
tabela2Cenario3.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario3[0] ])
tabela2Cenario3.add_column(nomes_colunas2[1], [ f"{round(resultCenario3[1][i],5)}; ({round(resultCenario3[1][i]-resultCenario3[2][i],5)}, {round(resultCenario3[1][i]+resultCenario3[2][i],5)})" for i in range(len(resultCenario3[1])) ])
tabela2Cenario3.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario3[4] ])
#tabela2Cenario3.add_column(nomes_colunas2[3], [ round(item,5) for item in resultCMCenario3[0] ] )
print(tabela2Cenario3.get_string(title="Resultados do Cenário 3 - Simulação, solução analítica e cadeia"))
# Tabela 3
tabela3Cenario3 = PrettyTable()
#nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]', 'Cadeia de Markov: E[W]']
nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]']
tabela3Cenario3.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario3[0] ])
tabela3Cenario3.add_column(nomes_colunas3[1], [ f"{round(resultCenario3[3][i],5)}; ({round(resultCenario3[3][i]-resultCenario3[5][i],5)}, {round(resultCenario3[3][i]+resultCenario3[5][i],5)})" for i in range(len(resultCenario3[3])) ])
tabela3Cenario3.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario3[14] ])
#tabela3Cenario3.add_column(nomes_colunas3[3], [ round(item,5) for item in resultCMCenario3[1] ])
print(tabela3Cenario3.get_string(title="Resultados do Cenário 3 - Simulação, solução analítica e cadeia"))
# + id="YqzFwIWKL8NY" colab_type="code" outputId="bf52a056-faf6-4552-e4fd-b6061f45ba63" executionInfo={"status": "ok", "timestamp": 1577141936781, "user_tz": 180, "elapsed": 467651, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
printGraficoQ3(3, resultCenario3, 65)
# + [markdown] id="x_CfpPXkMACi" colab_type="text"
# ###Cenário 4
# + [markdown] id="Q6V8salSMCUG" colab_type="text"
# ####Simulação e solução analítica
# + id="umujXfqtMFPw" colab_type="code" outputId="e6f84a36-f5ea-4c74-e08c-a992a1945881" executionInfo={"status": "ok", "timestamp": 1577141941587, "user_tz": 180, "elapsed": 472445, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/"}
def uniforme(L):
return random.uniform(L[0], L[1])
def simulaCenario4():
medias_Nq = []
medias_NqAnalitico = []
medias_WAnalitico = []
medias_W = []
medias_N = []
medias_X = []
medias_T = []
ρs = []
λs = [0.08]
confsNq = []
confsW = []
confsN = []
confsX = []
confsρ = []
confsT = []
resultCenario4 = []
result = simulaGeral(0.08 , 0.05, [5,15], [1,2], 1000, uniforme) # Faz a simulação para cada λ
resultAnal = calculoAnalitico(0.08,0.05,0,0, 5,15,1,3,'u')
medias_Nq.append(result[1]) # Guarda o valor do E(Nq) para cada λ
medias_W.append(result[3]) # Guarda o valor do E(W) para cada λ
medias_N.append(result[4])
medias_X.append(result[5])
ρs.append(result[6])
medias_T.append(result[7])
medias_NqAnalitico.append(resultAnal[0]) # Guarda o valor do E(Nq) usando Little para cada λ
medias_WAnalitico.append(resultAnal[1])
confsNq.append(result[1] - result[0][0]) # Guarda o valor da diferença entre a média e um dos extremos do intervalo de confiança para cada λ
confsW.append(result[3] - result[2][0])
confsN.append(result[4] - result[8][0])
confsX.append(result[5] - result[9][0])
confsρ.append(result[6] - result[10][0])
confsT.append(result[7] - result[11][0])
# Salva tudo num array geral
resultCenario4.append(λs)
resultCenario4.append(medias_Nq)
resultCenario4.append(confsNq)
resultCenario4.append(medias_W)
resultCenario4.append(medias_NqAnalitico)
resultCenario4.append(confsW)
resultCenario4.append(medias_N)
resultCenario4.append(medias_X)
resultCenario4.append(ρs)
resultCenario4.append(medias_T)
resultCenario4.append(confsN)
resultCenario4.append(confsX)
resultCenario4.append(confsρ)
resultCenario4.append(confsT)
resultCenario4.append(medias_WAnalitico)
return resultCenario4
resultCenario4 = simulaCenario4()
tabelaCenario4 = PrettyTable()
nomes_colunas = ['λ', 'Nº de clientes; (Intervalo de confiança)', 'E[X]; (Intervalo de confiança)', 'E[W]; (Intervalo de confiança)', 'E[T]; (Intervalo de confiança)', 'ρ; (Intervalo de confiança)', 'E[Nq]; (Intervalo de confiança)']
tabelaCenario4.add_column(nomes_colunas[0], [ round(lambd,2) for lambd in resultCenario4[0] ], align='c')
tabelaCenario4.add_column(nomes_colunas[1], [ f"{round(resultCenario4[6][i],5)}; ({round(resultCenario4[6][i]-resultCenario4[10][i],5)}, {round(resultCenario4[6][i]+resultCenario4[10][i],5)})" for i in range(len(resultCenario4[6])) ], align='c' )
tabelaCenario4.add_column(nomes_colunas[2], [ f"{round(resultCenario4[7][i],5)}; ({round(resultCenario4[7][i]-resultCenario4[11][i],5)}, {round(resultCenario4[7][i]+resultCenario4[11][i],5)})" for i in range(len(resultCenario4[7])) ], align='c' )
tabelaCenario4.add_column(nomes_colunas[3], [ f"{round(resultCenario4[3][i],5)}; ({round(resultCenario4[3][i]-resultCenario4[5][i],5)}, {round(resultCenario4[3][i]+resultCenario4[5][i],5)})" for i in range(len(resultCenario4[3])) ], align='c' )
tabelaCenario4.add_column(nomes_colunas[4], [ f"{round(resultCenario4[9][i],5)}; ({round(resultCenario4[9][i]-resultCenario4[13][i],5)}, {round(resultCenario4[9][i]+resultCenario4[13][i],5)})" for i in range(len(resultCenario4[9])) ], align='c' )
tabelaCenario4.add_column(nomes_colunas[5], [ f"{round(resultCenario4[8][i],5)}; ({round(resultCenario4[8][i]-resultCenario4[12][i],5)}, {round(resultCenario4[8][i]+resultCenario4[12][i],5)})" for i in range(len(resultCenario4[8])) ], align='c' )
tabelaCenario4.add_column(nomes_colunas[6], [ f"{round(resultCenario4[1][i],5)}; ({round(resultCenario4[1][i]-resultCenario4[2][i],5)}, {round(resultCenario4[1][i]+resultCenario4[2][i],5)})" for i in range(len(resultCenario4[1])) ], align='c' )
print(tabelaCenario4.get_string(title="Resultados do Cenário 4 - Simulação"))
tabela2Cenario4 = PrettyTable()
nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]']
tabela2Cenario4.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario4[0] ])
tabela2Cenario4.add_column(nomes_colunas2[1], [ f"{round(resultCenario4[1][i],5)}; ({round(resultCenario4[1][i]-resultCenario4[2][i],5)}, {round(resultCenario4[1][i]+resultCenario4[2][i],5)})" for i in range(len(resultCenario4[1])) ])
tabela2Cenario4.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario4[4] ])
print(tabela2Cenario4.get_string(title="Resultados do Cenário 4 - Simulação e solução analítica"))
tabela3Cenario4 = PrettyTable()
nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]']
tabela3Cenario4.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario4[0] ])
tabela3Cenario4.add_column(nomes_colunas3[1], [ f"{round(resultCenario4[3][i],5)}; ({round(resultCenario4[3][i]-resultCenario4[5][i],5)}, {round(resultCenario4[3][i]+resultCenario4[5][i],5)})" for i in range(len(resultCenario4[3])) ], align='c')
tabela3Cenario4.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario4[14] ])
print(tabela3Cenario4.get_string(title="Resultados do Cenário 2 - Simulação e solução analítica"))
# + [markdown] id="jjax48AZMHQr" colab_type="text"
# ####<NAME>
# + id="pBnwjE0nMMdz" colab_type="code" outputId="2c95b5d1-b273-4520-880b-fae27ea2e1da" executionInfo={"status": "ok", "timestamp": 1577141944828, "user_tz": 180, "elapsed": 475678, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/"}
import pandas as pd
import numpy as np
from numpy.linalg import matrix_power
from numpy.linalg import norm
import numpy.linalg as lin
def check_state(f1, f2, serving):
if f1 == 0 and f2 == 0 and serving != 0:
raise Exception("invalido: se sistema está vazio, não pode ter ninguém sendo servido")
if f1 > 0 and f2 > 0 and serving != 1 and serving != 2:
raise Exception("invalido: se sistema está ocupado, tem que estar sevindo alguém")
if f1 > 0 and f2 == 0 and serving != 1:
raise Exception("invalido: se só há clientes do tipo 1, então tem que estar servindo o tipo 1")
if f1 == 0 and f2 > 0 and serving != 2:
raise Exception("invalido: se só há clientes do tipo 2, então tem que estar servindo o tipo 2")
def get_state_name(f1, f2, serving):
check_state(f1, f2, serving)
s1 = "*" if serving == 1 else " "
s2 = "*" if serving == 2 else " "
return f"{f1:5}{s1}/{f2:5}{s2}"
def get_next_states(f1, f2, serving, λ1, λ2, µ1, µ2):
check_state(f1, f2, serving)
r = {}
# Chegada de um cliente do tipo 1:
if serving == 0:
r[get_state_name(f1+1,f2,1)] = λ1
else:
r[get_state_name(f1+1,f2,serving)] = λ1
# Chegada de um cliente do tipo 2:
if f1 == 0:
r[get_state_name(0,f2+1,2)] = λ2
else:
r[get_state_name(f1,f2+1,serving)] = λ2
# Saida de um cliente do tipo 1:
if serving == 1:
if f1 > 1:
r[get_state_name(f1-1,f2,1)] = µ1
elif f2 > 0:
r[get_state_name(0,f2,2)] = µ1
else:
r[get_state_name(0,0,0)] = µ1
# Saida de um cliente do tipo 2:
if serving == 2:
if f1 > 0:
r[get_state_name(f1,f2-1,1)] = µ2
elif f2 > 1:
r[get_state_name(0,f2-1,2)] = µ2
else:
r[get_state_name(0,0,0)] = µ2
return r
def create_ctmc_states(size, λ1, λ2, µ1, µ2):
states = {}
# adding state " 0 / 0 "
states[get_state_name(0,0,0)] = get_next_states(0,0,0, λ1, λ2, µ1, µ2)
for f1 in range(0,size):
for f2 in range(0,size):
if f1 == 0 and f2 == 0:
serve_list = []
elif f1 == 0:
serve_list = [2]
elif f2 == 0:
serve_list = [1]
else:
serve_list = [1,2]
for serving in serve_list:
states[get_state_name(f1,f2,serving)] = get_next_states(f1,f2,serving, λ1, λ2, µ1, µ2)
return states
def create_ctmc_matrix(size, λ1, λ2, µ1, µ2):
states = create_ctmc_states(size, λ1, λ2, µ1, µ2)
df = pd.DataFrame(states)
df = df.reindex(sorted(df.columns), axis=1)
df = df.sort_index()
df = df.transpose()
df = df[df.index]
columns = df.columns
np.fill_diagonal(df.values, -df.sum(axis=1))
df = df.fillna(0)
return df.to_numpy(), columns
# Calcula os π
def calculaCadeiaMarkovCenario4(λ1, λ2, µ1, µ2):
result = create_ctmc_matrix(40, λ1, λ2, µ1, µ2)
Q = result[0]
columns = result[1]
# Trecho a seguir calcula os π
sz = Q.shape[0]
Qt = Q.transpose()
Qt[sz-1,:] = 1
b = np.zeros((sz,1))
b[sz-1] = 1
x = lin.solve(Qt.T.dot(Qt), Qt.T.dot(b))
πi = {}
# Agrupa os πi que apresentam o mesmo número de pessoas no sistema para um único π
for i in range(len(columns)):
txt = columns[i].split(",")
for k in range(len(txt)):
txt2 = txt[k].split("/")
t1 = txt2[0].replace("*","")
t2 = txt2[1].replace("*","")
txt2 = (t1, t2)
numberCustumerSystem = int(txt2[0])+int(txt2[1])
if numberCustumerSystem in πi:
πi[numberCustumerSystem] = x[i] + πi[numberCustumerSystem]
else:
πi[numberCustumerSystem] = x[i]
return list(πi.values())
def cadeiaMarkovCenario4():
Nqs = []
W = []
Nq = 0
λ1 = 0.08
λ2 = 0.05
µ1 = (5 + 15)/2
µ2 = (1 + 3)/2
πk = (calculaCadeiaMarkovCenario4(λ1, λ2, µ1, µ2))
Nq = 0
# Faz os cálculos dos Nq e W
for j in range(1,len(πk)):
if(πk[j] < 0):
Nq += 0
else:
Nq += (j-1)*πk[j]
Nqs.append(Nq[0])
W.append(Nq[0]/(λ1+λ2))
return [Nqs, W]
resultCMCenario4 = cadeiaMarkovCenario4()
print("Nq:",resultCMCenario4[0])
print("W:",resultCMCenario4[1])
# + [markdown] id="fma_7Ix9MNI3" colab_type="text"
# ####Comparação
# + colab_type="code" id="j6kUh2ObV8Gy" outputId="b6711fc0-ba99-4aa1-d0fc-dad44057b0ec" executionInfo={"status": "ok", "timestamp": 1577141944831, "user_tz": 180, "elapsed": 475673, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/"}
# Tabela 2
tabela2Cenario4 = PrettyTable()
nomes_colunas2 = ['λ', 'Simulação: E[Nq]; (Intervalo de confiança)', 'Analítico: E[Nq]', 'Cadeia de Markov: E[Nq]']
tabela2Cenario4.add_column(nomes_colunas2[0], [ round(lambd,2) for lambd in resultCenario4[0] ])
tabela2Cenario4.add_column(nomes_colunas2[1], [ f"{round(resultCenario4[1][i],5)}; ({round(resultCenario4[1][i]-resultCenario4[2][i],5)}, {round(resultCenario4[1][i]+resultCenario4[2][i],5)})" for i in range(len(resultCenario4[1])) ])
tabela2Cenario4.add_column(nomes_colunas2[2], [ round(item,5) for item in resultCenario4[4] ])
tabela2Cenario4.add_column(nomes_colunas2[3], [ round(item,5) for item in resultCMCenario4[0] ] )
print(tabela2Cenario4.get_string(title="Resultados do Cenário 4 - Simulação, solução analítica e cadeia"))
# Tabela 3
tabela3Cenario4 = PrettyTable()
nomes_colunas3 = ['λ', 'Simulação: E[W]; (Intervalo de confiança)', 'Analítico: E[W]', 'Cadeia de Markov: E[W]']
tabela3Cenario4.add_column(nomes_colunas3[0], [ round(lambd,2) for lambd in resultCenario4[0] ])
tabela3Cenario4.add_column(nomes_colunas3[1], [ f"{round(resultCenario4[3][i],5)}; ({round(resultCenario4[3][i]-resultCenario4[5][i],5)}, {round(resultCenario4[3][i]+resultCenario4[5][i],5)})" for i in range(len(resultCenario4[3])) ])
tabela3Cenario4.add_column(nomes_colunas3[2], [ round(item,5) for item in resultCenario4[14] ])
tabela3Cenario4.add_column(nomes_colunas3[3], [ round(item,5) for item in resultCMCenario4[1] ])
print(tabela3Cenario4.get_string(title="Resultados do Cenário 4 - Simulação, solução analítica e cadeia"))
# + id="P7azPGXXMPOx" colab_type="code" outputId="84adf317-4423-45b3-d7f1-0ae23f88322c" executionInfo={"status": "ok", "timestamp": 1577141945688, "user_tz": 180, "elapsed": 476522, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "06722449926182729976"}} colab={"base_uri": "https://localhost:8080/"}
resultCenario4.append(resultCMCenario4[0])
resultCenario4.append(resultCMCenario4[1])
printGraficoQ3(4, resultCenario4, 0)
| Trabalho_AD/TrabalhoAD_Questao4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Author: <NAME>
# #### Program: Prediction of House Value using California State dataset.
# #### Last Modified: Feb 25, 2018
# +
from __future__ import division, print_function, unicode_literals
import os
import numpy as np
np.random.seed(42)
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Function to save the figures.
PROJECT_ROOT_DIR = "."
PROGRAM = "california_state_data"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", PROGRAM)
def save_fig (fig_id, tight_layout=True, fig_extension='png',resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path,format=fig_extension, dpi=resolution)
# -
# ### Get the Data
# +
import os
import tarfile
from six.moves import urllib
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml/master/"
HOUSING_PATH = os.path.join("datasets", "housing")
HOUSING_URL = DOWNLOAD_ROOT + "datasets/housing/housing.tgz"
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
if not os.path.isdir(housing_path):
os.makedirs(housing_path)
tgz_path = os.path.join(housing_path,"housing.tgz")
urllib.request.urlretrieve(housing_url,tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
# -
fetch_housing_data()
# +
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
# -
housing = load_housing_data()
housing.head()
housing.info()
housing["ocean_proximity"].value_counts()
housing.describe()
# %matplotlib inline
import matplotlib.pyplot as plt
housing.hist(bins=50, figsize=(20,15))
save_fig("attribute_histogram_plots")
plt.show()
| House Value Prediction - Califonia Dataset..ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yukinaga/ai_programming/blob/main/lecture_09/03_morphological_analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="T5Y9J5ZXseMI"
# # 形態素解析
# 形態素解析とは、自然言語を形態素にまで分割することです。
# 形態素とは、言葉が意味を持つまとまりの単語の最小単位のことです。
# 今回は、形態素解析を用いて単語に分割します。
# + [markdown] id="x8PoTnfx8JfL"
# ## ライブラリのインストール
# Janomeは日本語の形態素解析が可能なツールです。
# + id="CRJJYYW-8R9d"
# !pip install janome
# + [markdown] id="TCSMvucAseML"
# Janomeを使って形態素解析を行いましょう。
# Tokenizerをインポートします。
# + id="Fz3h1d6WseML"
from janome.tokenizer import Tokenizer
t = Tokenizer()
s = "すもももももももものうち"
for token in t.tokenize(s):
print(token)
# + [markdown] id="0uRsDzb1seMN"
# ## 分かち書き
# Janomeを使って分かち書きを行います。
# 分かち書きとは、文章を単語ごとに分割することです。
# `tokenize`の際に引数を`wakati=True`にすることで、各単語に分割できます。
# `tokenize`関数はgeneratorの形で分割された単語を返すので、単語をリストに格納したい場合は`list()`によりリストに変換します。
# + id="QexY60s0seMN"
from janome.tokenizer import Tokenizer
t = Tokenizer()
s = "すもももももももものうち"
word_list = t.tokenize(s, wakati=True)
word_list = list(word_list) # generatorをリストに変換
print(word_list)
# + [markdown] id="q1ndrQINseMN"
# ## コーパスを分かち書き
# 前回前処理を行った「我輩は猫である」に対して、分かち書きを行います。
# + id="3diiEHcPseMO"
from janome.tokenizer import Tokenizer
import pickle
t = Tokenizer()
with open('wagahai_list.pickle', mode='rb') as f:
wagahai_list = pickle.load(f)
for sentence in wagahai_list:
print(list(t.tokenize(sentence, wakati=True)))
# + [markdown] id="5X1e41y8seMO"
# collectionsを使うことで、各単語の出現回数をカウントすることができます。
# + id="XMvAC3t7seMO"
import collections
t = Tokenizer()
words = []
for sentence in wagahai_list:
words += list(t.tokenize(sentence, wakati=True)) # リストwordsに全ての単語を入れる
c = collections.Counter(words)
print(c)
# + [markdown] id="kwwGr1ANseMP"
# ## 課題:
# 前回の課題で前処理した「銀河鉄道の夜」で各単語数をカウントしてみましょう。
# + id="jPPJdA9ZseMP"
| lecture_09/03_morphological_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Run ADAM
#
# The model ADAM (Annual Danish Aggregate Model)is a model of the Danish
# economy maintained and developed by Danmarks Statistik. A Link to the model: https://www.dst.dk/pubomtale/18836. It has 4624 equations
#
# Danmarks Statistik has kindly provided the model version JUN17X and an associated databank for this demo.
#
# The model and databank is not the current model and forecast, but serves as an example.
#
# This is a basic demo. However you can use all the features of ModelFlow to manipulate the model.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Import ModelFlow and Pandas
# + slideshow={"slide_type": "-"}
# %load_ext autoreload
# %autoreload 2
import pandas as pd
from ipywidgets import interact,Dropdown
from IPython.display import display, clear_output
import sys
sys.path.append('modelflow/')
from modelsandbox import newmodel
from modelclass import model
from modeljupyter import inputwidget
# + [markdown] slideshow={"slide_type": "slide"}
# ## Read model and databank
# + slideshow={"slide_type": "-"}
fadam = open('adam/jul17x.txt','rt').read()
bank = pd.read_pickle('adam/lang100_2017.pc')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Make a model instance
# -
turbo = 0 # if you set turbo=1 the model will be compiled, it takes time but specd the running up
madam = newmodel(fadam,modelname='ADAM')
madam.use_preorder = True # for speedup
# + [markdown] slideshow={"slide_type": "slide"}
# ## Run the Baseline
# -
basedf = madam(bank,2018,2030,conv=['YR','UL'],antal=100,alfa=0.4,ljit=turbo,chunk=30,relconv = 0.0000001,silent=True,debug=0)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Define a interface to experiments
#
# The next cell defines interactive widget.
#
# You can add additional variables using by adding extra entries to the slidedef dictionary
# +
# ADAM Variable descriptions
vtrans = {'ENL' :'Balance of Payment, surplus','UL':'Unemployment','FY':'Real GDP',
'TFN_O':'Public surplus','TFN_OY':'Public surplus, Percent of GDP'}
slidedef = {'Value added tax rate': {'var':'TG','value':0.0,'min':-0.1, 'max':0.1,'op':'+'},
'Extra labour supply, 1000 ': {'var':'UQ','value':0.0,'min':-8, 'max':8, 'op':'+','step':1},
}
input = inputwidget(madam,basedf,slidedef=slidedef
,showout=True,varpat='FY ENL UL TFN_O TFN_OY',modelopt={'silent':True},trans=vtrans)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Now make the experiment
# -
display(input)
# ## Attributions to the results from changed variables
display(madam.get_att_gui(desdic=vtrans))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Attributions to an equation (here FY) from its input.
# You can decide how many levels back in the dependency tree you want to calculate the attributions
# -
@interact
def explain(Variable=Dropdown(options = sorted(madam.endogene),value='FY'),
Levels=[0,1,2,3]):
print(madam.allvar[Variable]['frml'])
_ = madam.explain(Variable,up=Levels,svg=1,dec=2)
# + [markdown] slideshow={"slide_type": "skip"}
# ## Uploade and downloade to the virtual machine.
# In general you should be able to both upload files from your local machine to the virtual machine, and and to download files from the virtual machine to the local machine.
#
# So even when though the virtual machines disappear you can still get the output to our local machine. And you can modify this notebook and download it for later use.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Now try turbo = 1
# Edit the cell above where turbo is set to 0 and change it to 1
#
# Then run the notebook again. This will trigger a just in time compilation by the NUMBA library. The compilation will take time, but the simulation will be much faster.
| ADAM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# > **Copyright (c) 2020 <NAME>**<br><br>
# > **Copyright (c) 2021 Skymind Education Group Sdn. Bhd.**<br>
# <br>
# Licensed under the Apache License, Version 2.0 (the \"License\");
# <br>you may not use this file except in compliance with the License.
# <br>You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0/
# <br>
# <br>Unless required by applicable law or agreed to in writing, software
# <br>distributed under the License is distributed on an \"AS IS\" BASIS,
# <br>WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# <br>See the License for the specific language governing permissions and
# <br>limitations under the License.
# <br>
# <br>
# **SPDX-License-Identifier: Apache-2.0**
# <br>
# # Introduction
#
# In this notebook, you will be introduced the concept of Object Oriented Programming and the implementation of OOP in Python
# # Notebook Content
#
# * [Object Oriented Programming Python](#Object-Oriented-Programming-Python)
#
# * [What is Object Oriented Programming (OOP)?](#What-is-Object-Oriented-Programming-(OOP)?)
# * [What are the advantages of OOP?](#What-are-the-advantages-of-OOP?)
# * [4 Main Concepts of OOP](#4-Main-Concepts-of-OOP)
#
#
# * [Implementation of OOP](#Implementation-of-OOP)
# * [Classes](#Classes)
# * [Objects](#Objects)
# * [Private Instance Variables & Methods](#Private-Instance-Variables-&-Methods)
# * [Inheritance](#Inheritance)
# * [Polymorphism](#Polymorphism)
# * [Advice](#Advice:)
#
#
# * [Advanced OOP Python](#Advanced-OOP-Python)
# + [markdown] id="K4M4QUPHqqEW"
# # Object Oriented Programming Python
# -
# ### What is Object Oriented Programming (OOP)?
#
# OOP is a method of structuring program based on the concept of "**objects**". An object consists of properties and behaviors which are unique and specific.
# + [markdown] id="Q9ol_R7VqqEa"
# ### What are the advantages of OOP?
#
# </br>
#
# 1. Code usability
# 2. Data redundancy
# 3. Code maintenance
# 4. Better code security
# 5. Easy to implement
# 6. Better productivity
# 7. Easy troubleshooting
# 8. Polymophism flexibility
# 9. Problem solvings
#
# </br>
#
# Further reading: https://www.educba.com/advantages-of-oop/
# + [markdown] id="cbyKZoYWqqEb"
# ### 4 Main Concepts of OOP
# </br>
#
# <h4>
#
# | No. | Concepts | Definitions
# |:---:|:-------------:|:------------------------------------------------------------------------|
# |1. | Encapsulation | Restricting the direct access to some components of an object. |
# |2. | Abstraction | Reducing complexity of program by hiding unnecessary data of an object. |
# |3. | Inheritance | Declaring hierarchy of classes that share common attributes and methods.|
# |4. | Polymorphism | Defining object with multiple forms. |
#
# </h4>
# </br>
#
# Further readings:
# - https://www.sumologic.com/glossary/encapsulation/
# - https://stackify.com/oop-concept-inheritance/
# - https://www.upgrad.com/blog/polymorphism-in-oops/
# + [markdown] id="9yINyQ8TqqEc"
# # Implementation of OOP
# + [markdown] id="TtgCLVLIqqEd"
# ### Classes
# </br>
#
# A class is a blueprint of creating an object with consists of properties (**instance variables**) and behaviours (**methods**)
# + id="a8ffYWgyqqEe"
class Transportation:
# Class attribute
function = "To carry people from place to place."
# Constructor
def __init__(self, colour, tyres, doors):
# Instance variables
self.colour = colour
self.tyres = tyres
self.doors = doors
# Instance methods
def startEngine(self):
print("Engine start...")
# + [markdown] id="8waJMmV1qqEg"
# ### Objects
# </br>
#
# An object is an instance of a class that has unique properties and behaviours
# + id="cUQBMxtDqqEj" outputId="13540ba0-b2c1-45b5-b2d9-eb81e5f9a70f"
# Creating transportation object
transport = Transportation("red", 4, 4)
# Accessing instance variable
print(f"This transport has {transport.tyres} tyres.")
# Accessing instance method
transport.startEngine()
# + [markdown] id="E5wF-Z4-qqEl"
# ### Private Instance Variables & Methods
# </br>
#
# In actual terms (practically), python doesn’t have anything called private member variable in Python.
#
# However, to make a variable and method becomes **private**, you just have to put **__** (double underscore) in front of variable and method.
#
# **Note**: Private variables and methods cannot be accessed except inside an object.
# + id="4O5rZOb0qqEm"
class Ferry:
def __init__(self, location):
self.location = location
# Private instance variable
self.__captain = "John"
# Private method
def __repair(self):
print("Repairing ship...")
# Public method
def startRepair(self):
self.__repair()
# + id="Tmn7UFKUqqEn" outputId="0ef17ed3-cdc2-4714-a1be-d91adaafe12f"
# Initialize object
ferry = Ferry("Penang")
# Access private variable -> Return AttributeError
try:
print(ferry.__captain)
except AttributeError as e:
print(e)
# Access private method -> Return AttributeError
try:
ferry.__repair()
except AttributeError as e:
print(e)
ferry.startRepair()
# + [markdown] id="C3QzaZ5tqqEn"
# ### Inheritance
# </br>
#
# Reusability of code that inherits all the methods and properties from another class.
#
# **Parent**: Class being inherited from, also called base class. <br> </br>
# **Child** : Class that inherits from another class, also called derived class.
# + id="NEWa5QrNqqEo"
# Derived from Transportation class
class Car(Transportation):
def __init__(self, colour, tyres, doors, size):
# initialize superclass
super().__init__(colour, tyres, doors)
self.size = size
def getDimension(self):
print("{} m (width) x {} m (height)".format(*self.size))
def description(self):
print("This car is in {} colour, has {} tyres and {} doors with size of {}m x {}m".format(self.colour, self.tyres, self.doors, *self.size))
# + id="HiN3nKitqqEo" outputId="94adaa0a-6e58-4b65-deef-a7cc0dda6dd1"
# Initialize car object
car = Car("blue", 4, 4, size=(5, 15))
# Calling parent class method
car.startEngine()
# Calling instance method
car.getDimension()
car.description()
# + [markdown] id="GNfZxH2fqqEp"
# ### Polymorphism
# </br>
#
# **Method Overriding**: Reimplementing a method inherited from the parent class in the child class.
# + id="rXmmBRf5qqEp"
class Bike(Transportation):
def __init__(self, colour, tyres, doors, price):
super().__init__(colour, tyres, doors)
self.price = price
# Override the inherited method
def startEngine(self):
print("Bike has no engine...")
# + id="9ee9G6TrqqEp" outputId="79ff964a-99de-44cc-9fc9-11c041890c30"
# Initialize bike object
bike = Bike("black", 2, 0, price=587.00)
# Calling overrided method
bike.startEngine()
# + [markdown] id="o9aM725oqqEp"
# ### Advice:
# </br>
#
# **Two leading underscores method** avoids method to be overridden by a subclass.
# + id="rqb4rCVBqqEu"
class Parent:
def __whoAmI(self):
print("I am parent")
def describe(self):
self.__whoAmI()
# Inherit parent class
class Child(Parent):
def __init__(self):
super().__init__()
# Try to override the private method of parent class
def __whoAmI(self):
print("I am children")
# + id="Ns2-U_3bqqEu" outputId="cf29f505-c3cb-4b14-be8d-403bb3cc86f6"
parent = Parent()
parent.describe()
child = Child()
# __whoAmI method in subclass is not overrided
child.describe()
# + [markdown] id="YpGeSSDZqqEv"
# # Advanced OOP Python
# + [markdown] id="0hPEEyXnqqEv"
# <h3> Dunder / Magic Method </h3>
# </br>
#
# Dunder method can override functionality for built-in functions for custom classes which in the form of `__<methodname>__`
# </br>
#
# List of all dunder methods in python:
#
# **Basic Customizations**
#
# `__new__(self)` return a new object (an instance of that class). It is called before `__init__` method.
#
# `__init__(self)` is called when the object is initialized. It is the constructor of a class.
#
# `__del__(self)` for del() function. Called when the object is to be destroyed. Can be used to commit unsaved data or close connections.
#
# `__repr__(self)` for repr() function. It returns a string to print the object. Intended for developers to debug. Must be implemented in any class.
#
# `__str__(self)` for str() function. Return a string to print the object. Intended for users to see a pretty and useful output. If not implemented, `__repr__` will be used as a fallback.
#
# `__bytes__(self)` for bytes() function. Return a byte object which is the byte string representation of the object.
#
# `__format__(self)` for format() function. Evaluate formatted string literals like % for percentage format and ‘b’ for binary.
#
# `__lt__(self, anotherObj)` for < operator.
#
# `__le__(self, anotherObj)` for <= operator.
#
# `__eq__(self, anotherObj)` for == operator.
#
# `__ne__(self, anotherObj)` for != operator.
#
# `__gt__(self, anotherObj)` for > operator.
#
# `__ge__(self, anotherObj)` for >= operator.
#
# **Arithmetic Operators**
#
# `__add__(self, anotherObj)` for + operator.
#
# `__sub__(self, anotherObj)` for – operation on object.
#
# `__mul__(self, anotherObj)` for * operation on object.
#
# `__matmul__(self, anotherObj)` for @ operator (numpy matrix multiplication).
#
# `__truediv__(self, anotherObj)` for simple / division operation on object.
#
# `__floordiv__(self, anotherObj)` for // floor division operation on object.
#
# **Type Conversion**
#
# `__abs__(self)` make support for abs() function. Return absolute value.
#
# `__int__(self)` support for int() function. Returns the integer value of the object.
#
# `__float__(self)` for float() function support. Returns float equivalent of the object.
#
# `__complex__(self)` for complex() function support. Return complex value representation of the object.
#
# `__round__(self, nDigits)` for round() function. Round off float type to 2 digits and return it.
#
# `__trunc__(self)` for trunc() function of math module. Returns the real value of the object.
#
# `__ceil__(self)` for ceil() function of math module. The ceil function Return ceiling value of the object.
#
# `__floor__(self)` for floor() function of math module. Return floor value of the object.
#
# **Emulating Container Types**
#
# `__len__(self)` for len() function. Returns the total number in any container.
#
# `__getitem__(self, key)` to support indexing. Like `container[index]` calls `container.__getitem(key)` explicitly.
#
# `__setitem__(self, key, value)` makes item mutable (items can be changed by index), like `container[index] = otherElement`.
#
# `__delitem__(self, key)` for del() function. Delete the value at the index key.
#
# `__iter__(self)` returns an iterator when required that iterates all values in the container.
#
# <br></br>
#
# Further reading:
# https://holycoders.com/python-dunder-special-methods/
# + id="BmEOPYPyqqEx"
class Coordinate:
def __init__(self, x, y):
self.x = x
self.y = y
def __str__(self):
return "(x: {}, y: {})".format(self.x, self.y)
def __add__(self, coord):
assert isinstance(coord, Coordinate)
return Coordinate(self.x + coord.x, self.y + coord.y)
def __sub__(self, coord):
assert isinstance(coord, Coordinate)
return Coordinate(self.x - coord.x, self.y - coord.y)
def __len__(self):
return 2
def distance(self, coord):
assert isinstance(coord, Coordinate)
return (self.y - coord.y) / (self.x - coord.x)
# + id="yXvl93XPqqEx" outputId="8de21cc8-2680-45d1-a729-0c829108f426"
# Initialize 2 coordinates
coord1 = Coordinate(5, 10)
coord2 = Coordinate(9, 17)
print("Coordinate 1:", coord1)
print("Coordinate 2:", coord2)
# Add two coordinates
ans = coord1 + coord2
print("Sum of two coordinates =", ans)
# Substract two coordinates
ans = coord1 - coord2
print("Difference of two coordinates =", ans)
# Distance between two coordinates
ans = coord1.distance(coord2)
print("Distance between two coordinates =", ans)
# -
# # Contributors
#
# **Author**
# <br><NAME>
# # References
#
# 1. [Python Documentation](https://docs.python.org/3/)
# 2. [Object Oriented Programming Python 3](https://realpython.com/python3-object-oriented-programming/)
# 3. [Python OOP](https://www.programiz.com/python-programming/object-oriented-programming)
| nlp-labs/Day_01/Python_Basic/06_OOP_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Nesting List
# +
c = ['ccc','ddd']
b = ['bb', c, 'ee', 'ff']
h = ['hh', 'ii']
x = ['a', b, 'g', h, 'j']
# -
print( x)
# # Coding Notes
# The isinstance() function returns True if the specified object is of the specified type otherwise False.
#
# ## Syntax
# isinstance(object, type)
#
# ## Parameter Values
#
# | Parameter | Description |
# | ----------- | ----------- |
# | object | Required. An object. |
# | type | A type or a class, or a tuple of types and/or classes |
#
#
#
#
isinstance("A", str)
# # Coding Example
#
# Converting Nesting List to Python List.
#
# The first IF and FOR statement checks the object for string and puts string value in the Python List.
#
# The ELIF goes down the one level when non string value is found.
#
# The RETURN is only used after the entire Nest List converted to Python List and Python List is printed out.
#
class Nesting:
def __init__(self):
self.final_list = []
def toList(self, nest):
for element in nest:
if isinstance(element, str):
self.final_list.append(element);
elif isinstance(element, list):
self.toList(element);
return self.final_list;
n=Nesting()
print( n.toList(x) )
| TypeDataStructures/Nesting List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import pandas as pd
import panel as pn
import plotly.express as px
import plotly.io as pio
import plotly.graph_objects as go
pn.extension("plotly", sizing_mode="stretch_width")
# -
# ## Styling Plotly for Panel
#
# In this example we will show how to style Plotly plots with Panel for both the `default` and the `dark` theme.
#
# 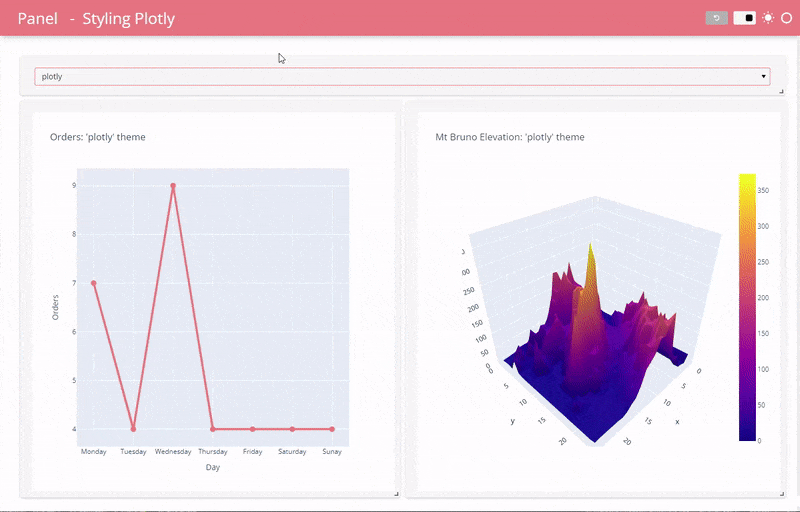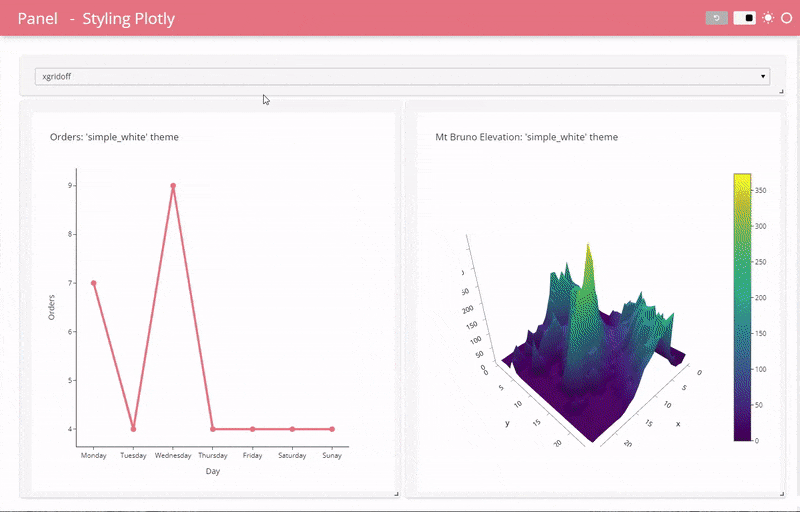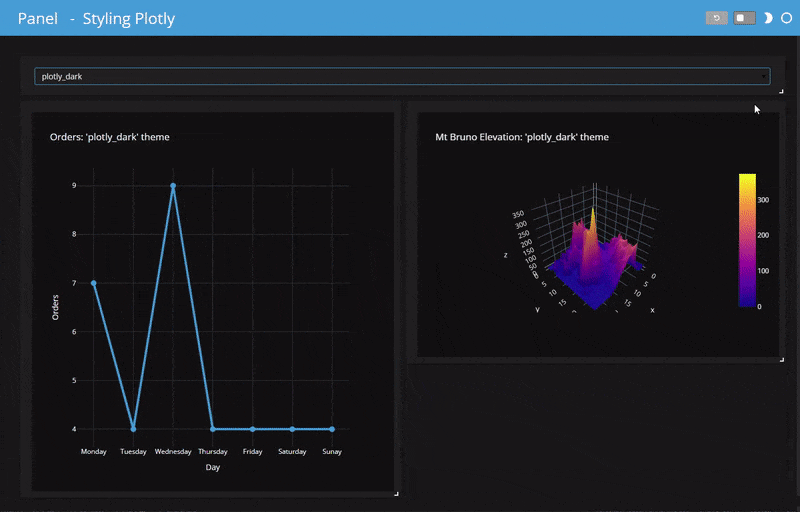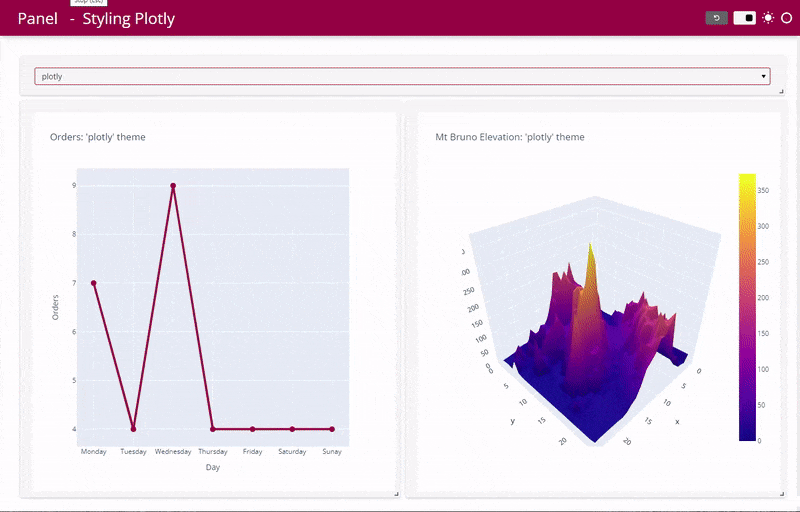
# ## Get or set the theme
#
# When we use the Fast templates the `theme` can be found in the `session_args`.
def get_theme():
return pn.state.session_args.get("theme", [b'default'])[0].decode()
theme=get_theme()
theme
# ## Select a nice accent color
#
# Below we create some functionality to *cycle through* a list of nice accent colors. You would probably just set the `accent_color` and `color` for your specific use case.
nice_accent_colors = [
("#00A170", "white"), # Mint
("#DAA520", "white"), # Golden Rod
("#F08080", "white"), # Light Coral
("#4099da", "white"), # Summery Sky
("#2F4F4F", "white"), # Dark Slate Grey
("#A01346", "white"), # Fast
]
def get_nice_accent_color():
"""Returns the 'next' nice accent color"""
if not "color_index" in pn.state.cache:
pn.state.cache["color_index"]=0
elif pn.state.cache["color_index"]==len(nice_accent_colors)-1:
pn.state.cache["color_index"]=0
else:
pn.state.cache["color_index"]+=1
return nice_accent_colors[pn.state.cache["color_index"]]
accent_color, color = get_nice_accent_color()
pn.pane.Markdown(f"# Color: {accent_color}", background=accent_color, height=70, margin=0, style={"color": color, "padding": "10px"})
# ## Plotly
#
# Plotly provides a list of built in templates in `plotly.io.templates`. See the [Plotly Templates Guide](https://plotly.com/python/templates/).
#
# Let's define a [`Select`](https://panel.holoviz.org/reference/widgets/Select.html) widget to explore the templates. We will set the default value to `plotly` or `plotly_dark` depending on the theme.
plotly_template = pn.widgets.Select(options=sorted(pio.templates))
if theme=="dark":
plotly_template.value="plotly_dark"
else:
plotly_template.value="plotly"
# ## Plotly Express
#
# Plotly Express provides a `template` argument. Let's try to use it.
data = pd.DataFrame(
[
("Monday", 7),
("Tuesday", 4),
("Wednesday", 9),
("Thursday", 4),
("Friday", 4),
("Saturday", 4),
("Sunay", 4),
],
columns=["Day", "Orders"],
)
def get_express_plot(template=plotly_template.value, accent_color=accent_color):
fig = px.line(
data,
x="Day",
y="Orders",
template=template,
color_discrete_sequence=(accent_color,),
title=f"Orders: '{template}' theme"
)
fig.update_traces(mode="lines+markers", marker=dict(size=10), line=dict(width=4))
fig.layout.autosize = True
return fig
# Let's [bind](https://panel.holoviz.org/user_guide/APIs.html#reactive-functions) `get_express_plot` to the `plotly_template` widget and lay out the two in a `Column`.
get_express_plot=pn.bind(get_express_plot, template=plotly_template)
express_plot=pn.pane.panel(get_express_plot, config={"responsive": True}, sizing_mode="stretch_both", name="EXPRESS")
pn.Column(plotly_template, express_plot, sizing_mode="stretch_both")
# ## Plotly Graph Objects Figure
#
# You can set the theme of a Plotly Graph Objects Figure via the `update_layout` method.
z_data = pd.read_csv("https://raw.githubusercontent.com/plotly/datasets/master/api_docs/mt_bruno_elevation.csv")
def get_go_plot(template=plotly_template.value, accent_color=accent_color):
figure = go.Figure(
data=go.Surface(z=z_data.values),
layout=go.Layout(
title="Mt Bruno Elevation",
))
figure.layout.autosize = True
figure.update_layout(template=template, title="Mt Bruno Elevation: '%s' theme" % template)
return figure
# Letss [bind](https://panel.holoviz.org/user_guide/APIs.html#reactive-functions) `get_go_plot` to the `plotly_template` widget and lay everything using `Tabs`and `Column`.
get_go_plot=pn.bind(get_go_plot, template=plotly_template)
go_plot=pn.pane.panel(get_go_plot, config={"responsive": True}, sizing_mode="stretch_both", name="GRAPH OBJECTS")
pn.Column(plotly_template, go_plot, min_height=600)
# ## Wrap it up in a nice template
#
# Here we use the [`FastGridTemplate`](https://panel.holoviz.org/reference/templates/FastListTemplate.html#templates-gallery-fastgridtemplate)
template = pn.template.FastGridTemplate(
site="Panel",
title="Styling Plotly",
sidebar=[plotly_template],
accent_base_color=accent_color,
header_background=accent_color,
header_color=color,
row_height=70,
save_layout=True, prevent_collision=True,
)
template.main[0:1,:]=plotly_template
template.main[1:10,0:6]=express_plot
template.main[1:10,6:12]=go_plot
template.servable();
# You can serve the app via `panel serve PlotlyStyle.ipynb` and find it at `http://localhost:5006/PlotlyStyle`. You should add the `--autoreload` flag while developing for *hot reloading*.
| examples/gallery/styles/PlotlyStyle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
os.environ['FR_Endpoint'] = "<YOUR_FORM_RECOGNIZER_ENDPOINT>"
os.environ['FR_Key'] = '<YOUR_FORM_RECOGNIZER_KEY>'
os.environ['FR_Model_ID'] = '<YOUR_CUSTOM_FORM_RECOGNIZER_MODEL_ID>'
os.environ['Form_Uri'] = '<YOUR_FORM_URI>'
# +
import os
import json
import numpy as np
import pandas as pd
from azure.core.exceptions import ResourceNotFoundError
from azure.ai.formrecognizer import FormRecognizerClient, FormTrainingClient
from azure.ai.formrecognizer import FormTrainingClient
from azure.core.credentials import AzureKeyCredential
#Get form recognizer endpoint/key, custom model ID, and form URI (blob uri w/ SAS Key)
endpoint = os.environ.get('FR_Endpoint')
key = os.environ.get('FR_Key')
model_id = os.environ.get('FR_Model_ID')
formUri = os.environ.get('Form_Uri')
#Create Form Recognizer Client
form_recognizer_client = FormRecognizerClient(endpoint, AzureKeyCredential(key))
# +
#Recognize text fields from custom model
poller = form_recognizer_client.begin_recognize_custom_forms_from_url(
model_id=model_id, form_url=formUri)
result = poller.result()
#Print detected fields
for recognized_form in result:
print("Form type: {}".format(recognized_form.form_type))
print()
for name, field in recognized_form.fields.items():
print("{} - {}".format(field.label_data.text if field.label_data else name,
field.value))
| Consuming_Custom_FR_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.12 64-bit (''dl_env'': venv)'
# language: python
# name: python3
# ---
# + [markdown] id="vio35IzlY124"
# # HOMEWORK 3 NEURAL NETWORKS AND DEEP LEARNING
#
# ---
# A.A. 2021/22 (6 CFU) - Dr. <NAME>, Dr. <NAME>
# ---
# Student: <NAME>
# ---
# id: 2020374
# + [markdown] id="neMVrqt0Y127"
# # Reinforced Learning
# + [markdown] id="ymxY-f60Y129"
# ### General overview
# In this homework you will learn how to implement and test neural network models for
# solving reinforcement learning problems. The basic tasks for the homework will require to implement some
# extensions to the code that you have seen in the Lab. The advanced tasks will require to train and test your
# learning agent on a different type of input (image pixels) or Gym environment. You can just choose one of
# the advanced tasks to get the maximum grade. If you are interested in improving your skills, feel free to try
# both advanced tasks. Given the higher computational complexity of RL, in this homework you don’t need to
# tune learning hyperparameters using search procedures and cross-validation; however, you are encouraged
# to play with model hyperparameters to find a satisfactory configuration.
#
#
# - 3 pt: use the notebook of Lab 07 to study how the exploration profile (either using eps-greedy or
# softmax) impacts the learning curve. Tune a bit the model hyperparameters or tweak the reward
# function to speed-up learning convergence (i.e., reach the same accuracy with fewer training episodes).
#
# YOU CAN DO JUST 1 OR BOTH
# - 5 pt: extend the notebook used in Lab 07, in order to learn to control the CartPole environment using
# directly the screen pixels, rather than the compact state representation used during the Lab (cart
# position, cart velocity, pole angle, pole angular velocity). NB: this will require to change the
# “observation_space” and to look for smart ways of encoding the pixels in a compact way to reduce
# computational complexity (e.g., crop the image around the pole, use difference of consecutive frames
# as input to consider temporal context, etc.).
#
# OR
#
# - 5 pt: train a deep RL agent on a different Gym environment. You are free to choose whatever Gym
# environment you like from the available list, or even explore other simulation platforms.
# + id="g15CvplwY12-"
# Boolean value to check if we are in colab or not
colab = False
show_video = False
show_render = False
load_model = True
# + colab={"base_uri": "https://localhost:8080/"} id="jBelL8nBZVsn" outputId="b1822e26-682a-49f2-ac0b-86b0618c8e0c"
if colab:
# !pip install gym
else:
# %pip install gym
# + id="a7EkHQ0VsNnJ"
import random
import torch
import numpy as np
import gym
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
from torch import nn
from collections import deque # this python module implements exactly what we need for the replay memeory
#use gpu if available
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# + colab={"base_uri": "https://localhost:8080/"} id="E0oXbzI2ZAfv" outputId="3fde884a-75dd-40a6-8d14-42c3f62763d0"
if colab:
# !apt update
# !apt-get install python-opengl -y
# !apt install xvfb -y
# !pip install pyvirtualdisplay
# !pip install piglet
import glob
import io
import base64
import os
from IPython.display import HTML
from IPython import display as ipythondisplay
from pyvirtualdisplay import Display
from gym.wrappers import Monitor
display = Display(visible=0, size=(1400, 900))
display.start()
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY"))==0:
# !bash ../xvfb start
# %env DISPLAY=:1
"""
Utility functions to enable video recording of gym environment and displaying it
To enable video, just do "env = wrap_env(env)""
"""
def show_videos():
mp4list = glob.glob('video/*.mp4')
mp4list.sort()
for mp4 in mp4list:
print(f"\nSHOWING VIDEO {mp4}")
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
ipythondisplay.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
def wrap_env(env, video_callable=None):
env = Monitor(env, './video', force=True, video_callable=video_callable)
return env
# + id="1axKWEBl8uik"
class ReplayMemory(object):
def __init__(self, capacity, state_size=2):
#capacity, state, action, reward, next_state, mask
self.memory = torch.zeros((capacity, 2*state_size+1+1+1), dtype=torch.float32, device=device) #capacity, action, reward, 2 states
self.state_size = state_size
self.memory_index = 0
self.capacity = capacity
self.full = False
def push(self, state, action, next_state, reward):
if state is None or action is None or reward is None:
print("WARNING: None value passed to ReplayMemory.push()")
return
# TODO: Add the tuple (state, action, next_state, reward) to the queue
s = torch.from_numpy(state).float().to(device)
a = torch.tensor([action], dtype=torch.float32, device=device)
r = torch.tensor([reward], dtype=torch.float32, device=device)
if next_state is not None:
next_s = torch.from_numpy(next_state).float().to(device)
mask = torch.tensor([1], dtype=torch.float32, device=device)
else:
next_s = torch.zeros((self.state_size), dtype=torch.float32, device=device)
mask = torch.tensor([0], dtype=torch.float32, device=device) # mask is false
to_add = torch.cat((s, a, r, next_s, mask), dim=0 )
#assert to_add.shape == (1, 2*self.state_size+1+1+1), f"to_add: {to_add.shape}, \n {to_add}"
self.memory[self.memory_index, :] = to_add
#increase the index, and wrap it around if it is bigger than the capacity
if self.memory_index == self.capacity-1:
self.full = True
self.memory_index = (self.memory_index + 1) % self.memory.shape[0]
def sample(self, batch_size):
batch_size = min(batch_size, len(self)) # Get all the samples if the requested batch_size is higher than the number of sample currently in the memory
# select indeces
pool = self.capacity if self.full else self.memory_index-1
indeces = np.random.choice(pool, batch_size, replace=False)
# return the samples
ret = self.memory[indeces, :]
states = ret[:, 0:self.state_size].unsqueeze(1)
actions = ret[:, self.state_size].long()
rewards = ret[:, self.state_size+1].float()
non_final_mask = ret[:, self.state_size+2].bool()
non_final_next_states = ret[:, self.state_size+3:].unsqueeze(1)
non_final_next_states = non_final_next_states[non_final_mask]
#print(f"states: {states.shape}, actions: {actions.shape}, rewards: {rewards.shape}, non_final_next_states: {non_final_next_states.shape}, non_final_mask: {non_final_mask.shape}")
return states, actions, rewards, non_final_next_states, non_final_mask
def __len__(self):
if self.full:
return self.capacity
else:
return self.memory_index
# return len(self.memory) # Return the number of samples currently stored in the memory
# + id="W41rXekb8x7K"
class DQN(nn.Module):
def __init__(self, state_space_dim, action_space_dim):
super().__init__()
n_hid = 128
self.linear = nn.Sequential(
#inpt layer
nn.Linear(state_space_dim, n_hid),
nn.Tanh(),
#nn.Dropout(0.2), #bad
#hidden layer
nn.Linear(n_hid, n_hid),
nn.Tanh(),
#nn.Dropout(0.2), #bad
#outpt layer
nn.Linear(n_hid, action_space_dim),
)
def forward(self, x):
return self.linear(x)
# + [markdown] id="ynfEXqjGfbpl"
# ## Exploration Policy
# + id="ZXfh_Ub1fv4c"
def choose_action_epsilon_greedy(net, state, epsilon):
if epsilon > 1 or epsilon < 0:
raise Exception('The epsilon value must be between 0 and 1')
# Evaluate the network output from the current state
with torch.no_grad():
net.eval()
state = torch.tensor(state, dtype=torch.float32,device=device) # Convert the state to tensor
net_out = net(state)
# Get the best action (argmax of the network output)
best_action = int(net_out.argmax())
# Get the number of possible actions
action_space_dim = net_out.shape[-1]
# Select a non optimal action with probability epsilon, otherwise choose the best action
if random.random() < epsilon:
# List of non-optimal actions
non_optimal_actions = [a for a in range(action_space_dim) if a != best_action]
# Select randomly
action = random.choice(non_optimal_actions)
else:
# Select best action
action = best_action
return action, net_out.cpu().numpy()
# + id="taW_cjBsf4sW"
def choose_action_softmax(net, state, temperature):
if temperature < 0:
raise Exception('The temperature value must be greater than or equal to 0 ')
# If the temperature is 0, just select the best action using the eps-greedy policy with epsilon = 0
if temperature == 0:
return choose_action_epsilon_greedy(net, state, 0)
# Evaluate the network output from the current state
with torch.no_grad():
net.eval()
state = torch.tensor(state, dtype=torch.float32, device=device)
net_out = net(state)
# Apply softmax with temp
temperature = max(temperature, 1e-8) # set a minimum to the temperature for numerical stability
softmax_out = nn.functional.softmax(net_out / temperature, dim=0).cpu().numpy()
# Sample the action using softmax output as mass pdf
all_possible_actions = np.arange(0, softmax_out.shape[-1])
action = np.random.choice(all_possible_actions, p=softmax_out) # this samples a random element from "all_possible_actions" with the probability distribution p (softmax_out in this case)
return action, net_out.cpu().numpy()
# + [markdown] id="QMXhJfZcpqIA"
# ### Exploration profile
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="G-f78CC3ptt2" outputId="fd64b464-4063-495b-8477-6532613451a0"
### Define exploration profile
def get_exploration_profile(initial_value=0.5, num_iterations=1000, speed = 3): # speed = speed of convergence, bigger is faster to reach 0
if initial_value < 0:
raise Exception('The initial value must be > 0')
if initial_value > 1:
exp_decay = np.exp(-np.log(initial_value) / num_iterations * speed) # We compute the exponential decay in such a way the shape of the exploration profile does not depend on the number of iterations
else :
exp_decay = np.exp(np.log(initial_value) / num_iterations * speed)
exploration_profile = [initial_value * (exp_decay ** i) for i in range(num_iterations)]
return exploration_profile
exploration_profile = get_exploration_profile()
### Plot exploration profile
plt.figure(figsize=(12,3))
plt.plot(exploration_profile)
plt.grid()
plt.xlabel('Iteration')
plt.ylabel('Exploration profile (Softmax temperature)')
# + [markdown] id="zjKmdFhowRbj"
# # Gym Environment ('MountainCar-v0')
# + colab={"base_uri": "https://localhost:8080/"} id="Ar0yNaNrCnjn" outputId="3fd78bd9-e4cf-4b14-d5d5-a30f0f4aaa88"
### Create environment
chosen_env = 'MountainCar-v0'
env = gym.make(chosen_env) # Initialize the Gym environment
env.seed(0) # Set a random seed for the environment (reproducible results)
# Get the shapes of the state space (observation_space) and action space (action_space)
state_space_dim = env.observation_space.shape[0]
action_space_dim = env.action_space.n
print(f"state_space_dim: {state_space_dim}, action_space_dim: {action_space_dim}")
if colab:
env = wrap_env(env, video_callable=lambda episode_id: True)
# + [markdown] id="9KXpjzf2vdeL"
# # Network update
# + id="lVpk-g0i9d-B"
# Set random seeds
torch.manual_seed(0)
np.random.seed(0)
random.seed(0)
### PARAMETERS
gamma = 0.97 #0.97 gamma parameter for the long term reward
replay_memory_capacity = 20000 #10000 Replay memory capacity
lr = 2e-2 #1e-2 Optimizer learning rate
target_net_update_steps = 5 #10 Number of episodes to wait before updating the target network
batch_size = 4096 #128 Number of samples to take from the replay memory for each update
bad_state_penalty = 0 #0 Penalty to the reward when we are in a bad state (in this case when the pole falls down)
min_samples_for_training = 5000 #1000 Minimum samples in the replay memory to enable the training
use_epsilon_greedy = False #False Use epsilon greedy exploration or not
#set up exploration profile for softmax
initial_value = 4 #5 Initial temperature/epsilon value for the exploration profile
num_iterations = 300 # Number of episodes 1000,
convergence_speed = 6 #6 Speed of convergence of the exploration profile
exploration_profile = get_exploration_profile(initial_value, num_iterations, speed = convergence_speed)
# + [markdown] id="pvSmo5w7Y13M"
# lowering target_net_update_steps from 10 to 8 had a huge impact, learning rate could be set higher too and gamma could be lowered.
# + id="2k3ZQuh8xHo7"
### Initialize the replay memory
replay_mem = ReplayMemory(replay_memory_capacity)
### Initialize the policy network
policy_net = DQN(state_space_dim, action_space_dim).to(device)
### Initialize the target network with the same weights of the policy network
target_net = DQN(state_space_dim, action_space_dim).to(device)
target_net.load_state_dict(policy_net.state_dict()) # This will copy the weights of the policy network to the target network
### Initialize the optimizer
optimizer = torch.optim.Adam(policy_net.parameters(), lr=lr) # The optimizer will update ONLY the parameters of the policy network
### Initialize the loss function (Huber loss)
loss_fn = nn.SmoothL1Loss()
#list of losses and scores
losses, scores = [],[]
# + id="Sj1hEvPOvkBX"
def update_step(policy_net, target_net, replay_mem, gamma, optimizer, loss_fn, batch_size):
# Sample the data from the replay memory
states, actions, rewards, non_final_next_states, non_final_mask = replay_mem.sample(batch_size)
#print(f"states: {states.shape}, actions: {actions.shape}, rewards: {rewards.shape}, non_final_next_states: {non_final_next_states.shape}, non_final_mask: {non_final_mask.shape}")
# Compute all the Q values (forward pass)
policy_net.train()
q_values = policy_net(states).squeeze(1)
# Select the proper Q value for the corresponding action taken Q(s_t, a)
#print(f"q_values: {q_values.shape}, actions: {actions}")
state_action_values = q_values.gather(1, actions.unsqueeze(1))
# Compute the value function of the next states using the target network V(s_{t+1}) = max_a( Q_target(s_{t+1}, a)) )
with torch.no_grad():
target_net.eval()
q_values_target = target_net(non_final_next_states).squeeze(1)
next_state_max_q_values = torch.zeros(batch_size, device=device)
#print(f"q_values_target: {q_values_target.shape}, non_final_mask: {non_final_mask.shape}, next_state_max_q_values: {next_state_max_q_values.shape}")
next_state_max_q_values[non_final_mask] = q_values_target.max(dim=1)[0]
# Compute the expected Q values
expected_state_action_values = rewards + (next_state_max_q_values * gamma)
expected_state_action_values = expected_state_action_values.unsqueeze(1) # Set the required tensor shape
# Compute the Huber loss
loss = loss_fn(state_action_values, expected_state_action_values)
# Optimize the model
optimizer.zero_grad()
loss.backward()
# Apply gradient clipping (clip all the gradients greater than 2 for training stability)
nn.utils.clip_grad_norm_(policy_net.parameters(), 2)
optimizer.step()
return loss.item()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["8e2fbf81537546668e2899f16d9b3be9", "7d265e41e381415bb671f818264dd590", "f3bb82f8b7734f90924832e2638ec039", "1442803c5a92408fa87ec3b33b897ecd", "d2c1223cb93c499198fb938ce12140af", "001de6b3e2794971bd82aeaf8e0a3f9f", "<KEY>", "<KEY>", "816903b033194420b016330111b9daeb", "c9e354f4a14e498ca31841c56e5f5b80", "ef024dc7ca8b409fa723b63c85e68df4"]} id="WF6Zf53FoRDZ" outputId="b466b533-110d-4095-8799-94d6295c37a0"
if not load_model:
# Initialize the Gym environment
env = gym.make(chosen_env)
env.seed(0) # Set a random seed for the environment (reproducible results)
# This is for creating the output video in Colab, not required outside Colab
if colab:
env = wrap_env(env, video_callable=lambda episode_id: episode_id % 100 == 0) # Save a video every 100 episodes
for episode_num, tau in enumerate(tqdm(exploration_profile)):
if use_epsilon_greedy:
epsilon = tau
# Reset the environment and get the initial state
state = env.reset()
# Reset the score, score will be the maximum position reached
position = -0.5
max_speed = 0
done = False
score = -1
episode_loss = []
tot_reward= 0
while not done:
# Choose the action following the policy
action, q_values = choose_action_epsilon_greedy(policy_net, state, epsilon) if use_epsilon_greedy else choose_action_softmax(policy_net, state, temperature=tau)
# Apply the action and get the next state, the reward and a flag "done" that is True if the game is ended
next_state, reward, done, info = env.step(action)
## SCORE UPDATE AND REWARD UPDATE
curr_position = -1
if done: # more than 200 steps or car arrived at the end (>= 0.5)
next_state = None
curr_position = -1
curr_speed = -1
score = max(score, state[0]+1.2)
else:
curr_position = next_state[0]
curr_speed = next_state[1]
# big reward if it achieves the goal
if reward == 0:
reward += 200
else:
reward = -0.1
#reward if it beats it's previous score
if curr_position > position:
reward += 1 + 2*(curr_position - position)
position = curr_position
#reward if it achieves a higher speed
if curr_speed > max_speed:
reward += 1 + 1*(curr_speed - max_speed)
max_speed = curr_speed
# Update the replay memory
replay_mem.push(state, action, next_state, reward)
# Update the network
if len(replay_mem) > min_samples_for_training: # we enable the training only if we have enough samples in the replay memory, otherwise the training will use the same samples too often
loss = update_step(policy_net, target_net, replay_mem, gamma, optimizer, loss_fn, batch_size)
episode_loss.append(loss)
# Visually render the environment (disable to speed up the training)
if not colab and show_render:
env.render()
# Set the current state for the next iteration
state = next_state
tot_reward += reward
#end of while loop
#get episode loss
average_episode_loss = np.mean(episode_loss)
losses.append(average_episode_loss)
#get episode score
scores.append(score)
# Update the target network every target_net_update_steps episodes
if episode_num % target_net_update_steps == 0:
#print('Updating target network...')
target_net.load_state_dict(policy_net.state_dict()) # This will copy the weights of the policy network to the target network
# Print the final score
print(f"EPISODE: {episode_num + 1} - FINAL SCORE: {score:.2f} - Temperature/epsilon: {tau:.4f}, total reward: {tot_reward:.2f}")
if episode_num > num_iterations*0.9:
break
env.close()
else:
# Load the model
policy_net.load_state_dict(torch.load(f'good_models/hill_policy__net.pt'))
target_net.load_state_dict(torch.load(f'good_models/hill_target_net.pt'))
# load scores and losses
losses = np.load(f'good_models/hill_losses.npy').tolist()
scores = np.load(f'good_models/hill_scores.npy').tolist()
print('Model loaded')
# + [markdown] id="FLZ_6SacY13O"
# Reward function is key, in this case the idea is that the agent get rewards only when it beats himself
# + id="7FQjjVTiP2T6"
# Display the videos, not required outside Colab
if colab and show_video:
show_videos()
# + colab={"base_uri": "https://localhost:8080/", "height": 336} id="d29HBc8HY13P" outputId="310f23dc-a19c-42c8-d4ef-b64d96733e28"
#plot scores and losses
fig = plt.figure(figsize=(12,3))
plt.subplot(1,2,1)
plt.plot(scores, label='Scores')
plt.title('Scores')
plt.plot(exploration_profile, label='Exploration profile')
plt.ylim(0, 1.8)
plt.grid()
plt.legend()
plt.subplot(1,2,2)
plt.plot(losses, label='Losses')
plt.title('Losses and exploration profile')
plt.plot(exploration_profile, label='Exploration profile')
plt.ylim(0,0.2)
plt.legend()
plt.grid()
plt.show()
fig.savefig('good_models/hill_scores_losses_exploration.eps', format='eps', dpi=1000)
# + [markdown] id="1ix8XN_sY13P"
# Adding an exploration profile makes the loss go down, however the score goes down with the loss too
# + [markdown] id="JkG9iDZTIhzc"
# # Final test
# + colab={"base_uri": "https://localhost:8080/"} id="vJKgnu3_IjWE" outputId="e0b2eedb-dfc5-4439-8649-bcdc30ef2ad1"
# Initialize the Gym environment
env = gym.make(chosen_env)
env.seed(1) # Set a random seed for the environment (reproducible results)
# This is for creating the output video in Colab, not required outside Colab
if colab:
env = wrap_env(env, video_callable=lambda episode_id: True) # Save a video every episode
# Let's try for a total of 10 episodes
scores = []
for num_episode in range(50):
# Reset the environment and get the initial state
state = env.reset()
# Reset the score. The final score will be the total amount of steps before the pole falls
score = -0.5
done = False
# Go on until the pole falls off or the score reach 490
while not done:
# Choose the best action (temperature 0)
action, q_values = choose_action_epsilon_greedy(policy_net, state, epsilon=0) if use_epsilon_greedy else choose_action_softmax(policy_net, state, temperature=0)
# Apply the action and get the next state, the reward and a flag "done" that is True if the game is ended
next_state, reward, done, info = env.step(action)
# Visually render the environment
if not colab and show_render:
env.render()
# Update the final score (+1 for each step)
score = max(score, next_state[0]+1.2)
# Set the current state for the next iteration
state = next_state
# Print the final score
#print(f"EPISODE {num_episode + 1} - FINAL SCORE: {score}")
scores.append(score)
env.close()
#print the average score
print(f"AVERAGE SCORE: {np.mean(scores):.4f}, environment is always solved for 1.7")
# + id="pIz9tDlBY13Q"
#let an agent play for a few episodes
#doesn't work in colab, but works in local
if not colab:
env = gym.make(chosen_env)
#env.seed(42)
show_render = True
scores = []
# Reset the environment and get the initial state
state = env.reset()
# Reset the score. The final score will be the total amount of steps before the pole falls
# Go on
for i in range(1000):
# Choose the best action (temperature 0)
action, q_values = choose_action_epsilon_greedy(policy_net, state, epsilon=0) if use_epsilon_greedy else choose_action_softmax(policy_net, state, temperature=0)
# Apply the action and get the next state, the reward and a flag "done" that is True if the game is ended
next_state, reward, done, info = env.step(action)
# Visually render the environment
if not colab and show_render:
env.render()
# Set the current state for the next iteration
state = next_state
env.close()
# -
if not load_model:
#save the model
torch.save(policy_net.state_dict(), f'good_models/hill_policy__net.pt')
torch.save(target_net.state_dict(), f'good_models/hill_target_net.pt')
#save losses
np.save(f'good_models/hill_losses.npy', losses)
#save scores
np.save(f'good_models/hill_scores.npy', scores)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="mxdtGJutLqlw" outputId="346fb19e-28c9-47e9-ec1e-790fbf63d51e"
# Display the videos, not required outside Colab
if colab:
show_videos()
| hw3/hw3_hill_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # STEP 5: ETL the data from 3NF tables to Facts & Dimension Tables
# **IMPORTANT:** The following exercise depends on first having successing completed Exercise 1: Step 4.
#
# Start by running the code in the cell below to connect to the database. If you are coming back to this exercise, then uncomment and run the first cell to recreate the database. If you recently completed steps 1 through 4, then skip to the second cell.
# +
# # !PGPASSWORD=student createdb -h 127.0.0.1 -U student pagila
# # !PGPASSWORD=student psql -q -h 127.0.0.1 -U student -d pagila -f Data/pagila-schema.sql
# # !PGPASSWORD=student psql -q -h 127.0.0.1 -U student -d pagila -f Data/pagila-data.sql
# +
# %load_ext sql
DB_ENDPOINT = "127.0.0.1"
DB = 'pagila'
DB_USER = 'student'
DB_PASSWORD = '<PASSWORD>'
DB_PORT = '5432'
# postgresql://username:password@host:port/database
conn_string = "postgresql://{}:{}@{}:{}/{}" \
.format(DB_USER, DB_PASSWORD, DB_ENDPOINT, DB_PORT, DB)
print(conn_string)
# %sql $conn_string
# -
# ### Introducing SQL to SQL ETL
# When writing SQL to SQL ETL, you first create a table then use the INSERT and SELECT statements together to populate the table. Here's a simple example.
# First, you create a table called test_table.
# + language="sql"
# CREATE TABLE test_table
# (
# date timestamp,
# revenue decimal(5,2)
# );
# -
# Then you use the INSERT and SELECT statements to populate the table. In this case, the SELECT statement extracts data from the `payment` table and INSERTs it INTO the `test_table`.
# + language="sql"
# INSERT INTO test_table (date, revenue)
# SELECT payment_date AS date,
# amount AS revenue
# FROM payment;
# -
# Then you can use a SELECT statement to take a look at your new table.
# %sql SELECT * FROM test_table LIMIT 5;
# If you need to delete the table and start over, use the DROP TABLE command, like below.
# %sql DROP TABLE test_table
# Great! Now you'll do the same thing below to create the dimension and fact tables for the Star Schema using the data in the 3NF database.
#
# ## ETL from 3NF to Star Schema
# ### 3NF - Entity Relationship Diagram
#
# <img src="./pagila-3nf.png" width="50%"/>
#
# ### Star Schema - Entity Relationship Diagram
#
# <img src="pagila-star.png" width="50%"/>
# In this section, you'll populate the tables in the Star schema. You'll `extract` data from the normalized database, `transform` it, and `load` it into the new tables.
#
# To serve as an example, below is the query that populates the `dimDate` table with data from the `payment` table.
# * NOTE 1: The EXTRACT function extracts date parts from the payment_date variable.
# * NOTE 2: If you get an error that says that the `dimDate` table doesn't exist, then go back to Exercise 1: Step 4 and recreate the tables.
# + language="sql"
# INSERT INTO dimDate (date_key, date, year, quarter, month, day, week, is_weekend)
# SELECT DISTINCT(TO_CHAR(payment_date :: DATE, 'yyyyMMDD')::integer) AS date_key,
# date(payment_date) AS date,
# EXTRACT(year FROM payment_date) AS year,
# EXTRACT(quarter FROM payment_date) AS quarter,
# EXTRACT(month FROM payment_date) AS month,
# EXTRACT(day FROM payment_date) AS day,
# EXTRACT(week FROM payment_date) AS week,
# CASE WHEN EXTRACT(ISODOW FROM payment_date) IN (6, 7) THEN true ELSE false END AS is_weekend
# FROM payment;
# -
# TODO: Now it's your turn. Populate the `dimCustomer` table with data from the `customer`, `address`, `city`, and `country` tables. Use the starter code as a guide.
# + language="sql"
# INSERT INTO dimCustomer (customer_key, customer_id, first_name, last_name, email, address,
# address2, district, city, country, postal_code, phone, active,
# create_date, start_date, end_date)
# SELECT c.customer_id as customer_key,
# c.customer_id,
# c.first_name,
# c.last_name,
# c.email,
# a.address,
# a.address2,
# a.district,
# ci.city,
# co.country,
# a.postal_code,
# a.phone,
# c.active,
# c.create_date,
# now() AS start_date,
# now() AS end_date
# FROM customer c
# JOIN address a ON (c.address_id = a.address_id)
# JOIN city ci ON (a.city_id = ci.city_id)
# JOIN country co ON (ci.country_id = co.country_id);
# -
# TODO: Populate the `dimMovie` table with data from the `film` and `language` tables. Use the starter code as a guide.
# + language="sql"
# INSERT INTO dimMovie (movie_key,film_id,title,description,release_year,language,
# original_language,rental_duration,length,rating,special_features)
# SELECT distinct f.film_id as movie_key,
# f.film_id,
# f.title,
# f.description,
# f.release_year,
# l.name as language,
# orig_lang.name AS original_language,
# f.rental_duration,
# f.length,
# f.rating,
# f.special_features
# FROM film f
# JOIN language l ON (f.language_id=l.language_id)
# LEFT JOIN language orig_lang ON (f.original_language_id = orig_lang.language_id);
# -
# TODO: Populate the `dimStore` table with data from the `store`, `staff`, `address`, `city`, and `country` tables. This time, there's no guide. You should write the query from scratch. Use the previous queries as a reference.
# + language="sql"
# INSERT INTO dimStore (store_key,store_id,address,address2, district, city, country,
# postal_code,manager_first_name,manager_last_name,start_date,end_date)
# SELECT distinct s.store_id as store_key,
# s.store_id,
# a.address,
# a.address2,
# a.district,
# ci.city,
# co.country,
# a.postal_code,
# st.first_name as manager_first_name,
# st.last_name as manager_last_name,
# now() as start_date,
# now() as end_date
# FROM store s
# JOIN staff st ON (s.store_id = st.store_id)
# JOIN address a ON (st.address_id = a.address_id)
# JOIN city ci ON (a.city_id = ci.city_id)
# JOIN country co ON (ci.country_id = co.country_id);
#
#
#
#
#
# -
# TODO: Populate the `factSales` table with data from the `payment`, `rental`, and `inventory` tables. This time, there's no guide. You should write the query from scratch. Use the previous queries as a reference.
# + language="sql"
# INSERT INTO factSales (sales_key, date_key,customer_key,movie_key,store_key,sales_amount)
# SELECT Distinct p.payment_id as sales_key,
# (TO_CHAR(p.payment_date :: DATE, 'yyyyMMDD')::integer) AS date_key,
# r.customer_id as customer_key,
# i.film_id as movie_key,
# i.store_id as store_key,
# p.amount as sales_amount
# FROM payment p
# JOIN rental r ON (p.rental_id = r.rental_id)
# JOIN inventory i ON (r.inventory_id = i.inventory_id)
#
#
#
#
#
# -
| Cloud_DWH/L1 E1 - Step 5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Data from the Web
# The internet is full of useful (as well as useless) information, so sometimes it might be very helpful to get data from it and process it locally.
#
# There are different ways to get data from the web, the most used are:
# - **Web Scraping**: download the html content from the page and then look from it to extract information
# - **Using Web API**: APIs ([Application Programming Interface](https://en.wikipedia.org/wiki/API)) is code that is meant to be called from other code instead of displayed visually to a user. We'll be looking at the most common type: [REST API](https://en.wikipedia.org/wiki/Representational_state_transfer)
#
# Often Web API need authentication, but ofter you can get a API key after a quick free signup.
# [Here's a non-exaustive list](https://github.com/public-apis/public-apis) of open APIs.
# ## External Libraries
# We'll be using these two external libraries:
# - [requests](https://requests.readthedocs.io/en/master/): a more user-friendly alternative to the built-in library `urllib.request`
# - [beautifulsoup4](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) a html parser, which is a type of software that builds a data structure from given inputs (usually of text kind).
#
# which you can install by runnin the cell below
# !pip install requests
# !pip install beautifulsoup4
# let's import from them to check they were installed correctly
import requests
from bs4 import BeautifulSoup
# ## Web Scraping
r = requests.get("https://www.metaweather.com/21125/")
r
# + tags=[]
html_doc = r.text
html_doc[:200]
# + [markdown] jupyter={"outputs_hidden": true} tags=[]
# if you want you can try render the html within the notebook by using
# ```python
# from IPython.core.display import HTML
# HTML(r.text)
# ```
# -
# We could try an do string manipulation (or using [regular expression syntax](https://www.w3schools.com/python/python_regex.asp)), but using a pre-built parser is usually less painful :)
soup = BeautifulSoup(html_doc, 'html.parser')
# + tags=[]
weather_data = []
for item in soup.findAll('div'):
if item.has_attr('data-date'):
weather_date = item['data-date']
weather_description = item.find("span").get_text()
weather_day_data = {
"date": weather_date,
"description": weather_description,
}
weather_data.append(weather_day_data)
# -
weather_data
# ## Web API
# APIs are usually much more stable and nicer to work with, but you usually need to read through some documentation to learn what you can do and which particular urls (called "endpoints") you need to use.
#
# In our case, [MetaWeather API documentation](https://www.metaweather.com/api/) tells us that to get a similar output as our scraped data, we need to use the endpoint `/api/location/(woeid)/` where `woeid` is the identifier of the location we want the weather from.
r = requests.get("https://www.metaweather.com/api/location/21125?api=hjagr0r3hg03hrghg0g3rah0")
r
api_weather_data = r.json()
# + tags=[]
api_weather_data
# -
# # Pandas to the rescue
# !pip install lxml
# +
import pandas as pd
list_of_dfs = pd.read_html('https://en.wikipedia.org/wiki/S%26P_500')
first_df = list_of_dfs[1].iloc[:-4] # last four columns are statistics
df = first_df.set_index('Year')
change_values_as_strings = df['Change in Index'].str.replace('−','-').str.replace('%','') # cleaning up wierd characters
change_values_as_numbers = pd.to_numeric(change_values_as_strings)
change_values_as_numbers.plot(grid=True)
# -
# # HomeWork
#
# For a more flexible application we could get the `woeid` from a city name or lat/lon coordinates by using other provided endpoints: `/api/location/search/?query=(query)` and `/api/location/search/?lattlong=(latt),(long)`
#
# Create two functions:
# ```python
# def get_woeid_from_city_name(city_name):
# ...
#
# def get_woeid_from_latlon(lat, lon):
# ...
# ```
#
# which will return the `woeid` for the given input. Then try to combine this with the previous code to produce a function that gets the weather for the next days.
# + tags=[]
...
# -
# ### Possible solution
# + jupyter={"source_hidden": true} tags=[]
def _get_woeid_from_city_name(city_name):
json_data = requests.get(f"https://www.metaweather.com/api/location/search/?query={city_name}").json()
if not json_data:
raise ValueError(f"No city found with name: {city_name}")
first_result_woeid = json_data[0]['woeid']
return first_result_woeid
def _get_woeid_from_latlon(lat, lon):
json_data = requests.get(f"https://www.metaweather.com/api/location/search/?lattlong={lat},{lon}").json()
if not json_data:
raise ValueError(f"No location found with latitude {lat} and longitude {lon}")
first_result_woeid = json_data[0]['woeid']
return first_result_woeid
# which can be tested with
_get_woeid_from_city_name("Glasgow")
_get_woeid_from_latlon(55.864200, -4.251800)
def _get_weather_from_woeid(woeid):
json_data = requests.get(f"https://www.metaweather.com/api/location/{woeid}").json()
json_data['consolidated_weather']
return json_data
def print_weather(city_name=None, lat=None, lon=None):
if city_name is None:
woeid = _get_woeid_from_latlon(lat, lon)
else:
woeid = _get_woeid_from_city_name(city_name)
weather_data = _get_weather_from_woeid(woeid)
for i in weather_data['consolidated_weather']:
print(i['applicable_date'], i['weather_state_name'])
# which can be tested with
print_weather("Glasgow")
# -
| 2020-21_semester2/09_Data_from_Web.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Javascript (Node.js)
// language: javascript
// name: javascript
// ---
// + [markdown] deletable=false
// # ibm_db.beginTransaction()
// + [markdown] deletable=false
// ## Purpose:
// + [markdown] deletable=false
// Begin a transaction.
// + [markdown] deletable=false
// ## Syntax:
// + [markdown] deletable=false
// conn.beginTransaction(callback)
// + [markdown] deletable=false
// ## Parameters:
// + [markdown] deletable=false
//
// * __*callback :*__ `callback (err)`
//
// + [markdown] deletable=false
// ## Return values:
// + [markdown] deletable=false
// * If __successful__, begins the transaction.
// * If __unsuccessful__, returns `Error`
// + [markdown] deletable=false
// ## Description:
// + [markdown] deletable=false
// The __ibm_db.beginTransaction()__ API is used to begin the transaction in an IBM Db2 server or database.<p>
// + [markdown] deletable=false
// ## Example:
//
// + deletable=false
/*
#----------------------------------------------------------------------------------------------#
# NAME: ibm_db-beginTransaction.js #
# #
# PURPOSE: This program is designed to illustrate how to use the ibm_db.beginTransaction() #
# API to begin the transaction in an remote Db2 server. #
# #
# Additional APIs used: #
# ibm_db.open() # #
# ibm_db.commitTransaction() # #
# ibm_db.querySync() #
# ibm_db.closeSync() #
# #
# #
#----------------------------------------------------------------------------------------------#
# DISCLAIMER OF WARRANTIES AND LIMITATION OF LIABILITY #
# #
# (C) COPYRIGHT International Business Machines Corp. 2018 All Rights Reserved #
# Licensed Materials - Property of IBM #
# #
# US Government Users Restricted Rights - Use, duplication or disclosure restricted by GSA #
# ADP Schedule Contract with IBM Corp. #
# #
# The following source code ("Sample") is owned by International Business Machines #
# Corporation ("IBM") or one of its subsidiaries and is copyrighted and licensed, not sold. #
# You may use, copy, modify, and distribute the Sample in any form without payment to IBM, #
# for the purpose of assisting you in the creation of Python applications using the ibm_db #
# library. #
# #
# The Sample code is provided to you on an "AS IS" basis, without warranty of any kind. IBM #
# HEREBY EXPRESSLY DISCLAIMS ALL WARRANTIES, EITHER EXPRESS OR IMPLIED, INCLUDING, BUT NOT #
# LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE. #
# Some jurisdictions do not allow for the exclusion or limitation of implied warranties, so #
# the above limitations or exclusions may not apply to you. IBM shall not be liable for any #
# damages you suffer as a result of using, copying, modifying or distributing the Sample, #
# even if IBM has been advised of the possibility of such damages. #
#----------------------------------------------------------------------------------------------#
*/
var ibmdb = require("ibm_db")
, cn = "DATABASE=dbName;HOSTNAME=myhost;PORT=dbport;PROTOCOL=TCPIP;UID=username;PWD=password";
ibmdb.open(cn, function (err, conn) {
if (conn) {
if (conn.connected) {
console.log("\n A database connection has been created successfully.\n");
}
else if (err) {
console.log(JSON.stringify(err));
return;
}
}
conn.beginTransaction(function (err) {
conn.querySync("create table customer(customercode varchar(10))");
var result = conn.querySync("insert into customer (customerCode) values ('stevedave')");
console.log("The transaction has begin.\n")
conn.commitTransaction(function (err) {
if (err) {
//error during commit
console.log(err);
return conn.closeSync();
}
console.log(conn.querySync("select * from customer where customerCode = 'stevedave'"));
conn.querySync("drop table customer");
//Close the connection
conn.closeSync();
});
});
});
// -
| Jupyter_Notebooks/ibm_db-beginTransaction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interactive Hyper-parameter Configuration
# This is a use case provided by RLzoo to support an interactive hyper-parameter configuration process. It is built with *ipywidgets* package, so make sure you have the package installed:
#
# ```! pip3 install ipywidgets==7.5.1```
#
# You just need to **run** each cell (Shift+Enter) and **select** the sliders or dropdown lists to configure the hyper-parameters for the learning process, for whichever algorithm and environment supported in RLzoo.
#
# It follows four steps:
# 1. Environment Configuration
# 2. Environment Information Display and Algorithm Configuration
# 3. Algorithm Parameters Display and Learning Parameters Configuration
# 4. Launch Learning with Visualization
#
# Tips:
# To stop the learning process and start a new one, you needs to restart the kernel (always work) or interrupt the kernel (not always work).
#
# Have fun!
# +
"""
1. Environment Configuration
-----------------------------
Run a environment selector and select a environment you like.
Tips: no need to rerun after selection, directly go to next cell.
"""
from rlzoo.interactive.common import *
from rlzoo.interactive.components import *
from rlzoo.algorithms import *
from rlzoo.common.env_wrappers import build_env, close_env
env_sel = EnvironmentSelector()
display(env_sel)
# +
"""
2. Environment Information Display and Algorithm Configuration
--------------------------------------------------------------
Run this code to create the enivronment instance.
Tips: need to rerun every time you want to create a new environment with above cell, \
because this cell builds the environment.
"""
try:
close_env(env) # close the previous environment
except:
pass
env = build_env(**env_sel.value)
print('Environment created!')
display(EnvInfoViewer(env))
# run a algorithm selector and select a RL algorithm
alog_sel = AlgorithmSelector(env)
display(alog_sel)
# +
"""
3. Algorithm Parameters Display and Learning Parameters Configuration
----------------------------------------------------------------------
Call the default parameters of the selected algorithm in our environment and display them, \
then select learning parameters.
Tips: need to rerun after you created a different algorithm or environment.
"""
EnvType, AlgName = env_sel.value['env_type'], alog_sel.value
alg_params, learn_params = call_default_params(env, EnvType, AlgName)
print('Default parameters loaded!')
# see the networks, optimizers and adjust other parameters
algiv = AlgoInfoViewer(alog_sel, alg_params, learn_params)
display(algiv)
# run this to generate the algorithm instance with the algorithm parameter settings above
alg_params = algiv.alg_params
alg = eval(AlgName+'(**alg_params)')
print('Algorithm instance created!')
# +
"""
4. Launch Learning with Visualization
---------------------------------------
Run the cell to train the algorithm with the configurations above.
"""
learn_params = algiv.learn_params
om = OutputMonitor(learn_params, smooth_factor=algiv.smooth_factor)
display(om)
with om.print_out:
alg.learn(env=env, plot_func=om.plot_func, **learn_params)
# -
# whenever leaving the page, please close the environment by the way
close_env(env)
print('Environment closed')
| rlzoo/interactive/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Mutation-Based Fuzzing
#
# Most [randomly generated inputs](Fuzzer.ipynb) are syntactically _invalid_ and thus are quickly rejected by the processing program. To exercise functionality beyond input processing, we must increase chances to obtain valid inputs. One such way is so-called *mutational fuzzing* – that is, introducing small changes to existing inputs that may still keep the input valid, yet exercise new behavior. We show how to create such mutations, and how to guide them towards yet uncovered code, applying central concepts from the popular AFL fuzzer.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# **Prerequisites**
#
# * You should know how basic fuzzing works; for instance, from the ["Fuzzing"](Fuzzer.ipynb) chapter.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Fuzzing a URL Parser
#
# Many programs expect their inputs to come in a very specific format before they would actually process them. As an example, think of a program that accepts a URL (a Web address). The URL has to be in a valid format (i.e., the URL format) such that the program can deal with it. When fuzzing with random inputs, what are our chances to actually produce a valid URL?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# To get deeper into the problem, let us explore what URLs are made of. A URL consists of a number of elements:
#
# scheme://netloc/path?query#fragment
#
# where
#
# * `scheme` is the protocol to be used, including `http`, `https`, `ftp`, `file`...
# * `netloc` is the name of the host to connect to, such as `www.google.com`
# * `path` is the path on that very host, such as `search`
# * `query` is a list of key/value pairs, such as `q=fuzzing`
# * `fragment` is a marker for a location in the retrieved document, such as `#result`
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# In Python, we can use the `urlparse()` function to parse and decompose a URL into its parts.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import fuzzingbook_utils
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
try:
from urlparse import urlparse # Python 2
except ImportError:
from urllib.parse import urlparse # Python 3
urlparse("http://www.google.com/search?q=fuzzing")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# We see how the result encodes the individual parts of the URL in different attributes.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let us now assume we have a program that takes a URL as input. To simplify things, we won't let it do very much; we simply have it check the passed URL for validity. If the URL is valid, it returns True; otherwise, it raises an exception.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
def http_program(url):
supported_schemes = ["http", "https"]
result = urlparse(url)
if result.scheme not in supported_schemes:
raise ValueError("Scheme must be one of " + repr(supported_schemes))
if result.netloc == '':
raise ValueError("Host must be non-empty")
# Do something with the URL
return True
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let us now go and fuzz `http_program()`. To fuzz, we use the full range of printable ASCII characters, such that `:`, `/`, and lowercase letters are included.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Fuzzer import fuzzer
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
fuzzer(char_start=32, char_range=96)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let's try to fuzz with 1000 random inputs and see whether we have some success.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
for i in range(1000):
try:
url = fuzzer()
result = http_program(url)
print("Success!")
except ValueError:
pass
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# What are the chances of actually getting a valid URL? We need our string to start with `"http://"` or `"https://"`. Let's take the `"http://"` case first. These are seven very specific characters we need to start with. The chance of producing these seven characters randomly (with a character range of 96 different characters) is $1 : 96^7$, or
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
96 ** 7
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# The odds of producing a `"https://"` prefix are even worse, at $1 : 96^8$:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
96 ** 8
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# which gives us a total chance of
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
likelihood = 1 / (96 ** 7) + 1 / (96 ** 8)
likelihood
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# And this is the number of runs (on average) we'd need to produce a valid URL scheme:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
1 / likelihood
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let's measure how long one run of `http_program()` takes:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Timer import Timer
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
trials = 1000
with Timer() as t:
for i in range(trials):
try:
url = fuzzer()
result = http_program(url)
print("Success!")
except ValueError:
pass
duration_per_run_in_seconds = t.elapsed_time() / trials
duration_per_run_in_seconds
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# That's pretty fast, isn't it? Unfortunately, we have a lot of runs to cover.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
seconds_until_success = duration_per_run_in_seconds * (1 / likelihood)
seconds_until_success
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# which translates into
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
hours_until_success = seconds_until_success / 3600
days_until_success = hours_until_success / 24
years_until_success = days_until_success / 365.25
years_until_success
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# Even if we parallelize things a lot, we're still in for months to years of waiting. And that's for getting _one_ successful run that will get deeper into `http_program()`.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# What basic fuzzing will do well is to test `urlparse()`, and if there is an error in this parsing function, it has good chances of uncovering it. But as long as we cannot produce a valid input, we are out of luck in reaching any deeper functionality.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Mutating Inputs
#
# The alternative to generating random strings from scratch is to start with a given _valid_ input, and then to subsequently _mutate_ it. A _mutation_ in this context is a simple string manipulation - say, inserting a (random) character, deleting a character, or flipping a bit in a character representation. This is called *mutational fuzzing* – in contrast to the _generational fuzzing_ techniques discussed earlier.
#
# Here are some mutations to get you started:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import random
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
def delete_random_character(s):
"""Returns s with a random character deleted"""
if s == "":
return s
pos = random.randint(0, len(s) - 1)
# print("Deleting", repr(s[pos]), "at", pos)
return s[:pos] + s[pos + 1:]
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
seed_input = "A quick brown fox"
for i in range(10):
x = delete_random_character(seed_input)
print(repr(x))
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
def insert_random_character(s):
"""Returns s with a random character inserted"""
pos = random.randint(0, len(s))
random_character = chr(random.randrange(32, 127))
# print("Inserting", repr(random_character), "at", pos)
return s[:pos] + random_character + s[pos:]
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
for i in range(10):
print(repr(insert_random_character(seed_input)))
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
def flip_random_character(s):
"""Returns s with a random bit flipped in a random position"""
if s == "":
return s
pos = random.randint(0, len(s) - 1)
c = s[pos]
bit = 1 << random.randint(0, 6)
new_c = chr(ord(c) ^ bit)
# print("Flipping", bit, "in", repr(c) + ", giving", repr(new_c))
return s[:pos] + new_c + s[pos + 1:]
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
for i in range(10):
print(repr(flip_random_character(seed_input)))
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let us now create a random mutator that randomly chooses which mutation to apply:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
def mutate(s):
"""Return s with a random mutation applied"""
mutators = [
delete_random_character,
insert_random_character,
flip_random_character
]
mutator = random.choice(mutators)
# print(mutator)
return mutator(s)
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
for i in range(10):
print(repr(mutate("A quick brown fox")))
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# The idea is now that _if_ we have some valid input(s) to begin with, we may create more input candidates by applying one of the above mutations. To see how this works, let's get back to URLs.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Mutating URLs
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# Let us now get back to our URL parsing problem. Let us create a function `is_valid_url()` that checks whether `http_program()` accepts the input.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
def is_valid_url(url):
try:
result = http_program(url)
return True
except ValueError:
return False
# + slideshow={"slide_type": "fragment"}
assert is_valid_url("http://www.google.com/search?q=fuzzing")
assert not is_valid_url("xyzzy")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let us now apply the `mutate()` function on a given URL and see how many valid inputs we obtain.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
seed_input = "http://www.google.com/search?q=fuzzing"
valid_inputs = set()
trials = 20
for i in range(trials):
inp = mutate(seed_input)
if is_valid_url(inp):
valid_inputs.add(inp)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# We can now observe that by _mutating_ the original input, we get a high proportion of valid inputs:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
len(valid_inputs) / trials
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# What are the odds of also producing a `https:` prefix by mutating a `http:` sample seed input? We have to insert ($1 : 3$) the right character `'s'` ($1 : 96$) into the correct position ($1 : l$), where $l$ is the length of our seed input. This means that on average, we need this many runs:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
trials = 3 * 96 * len(seed_input)
trials
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# We can actually afford this. Let's try:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Timer import Timer
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
trials = 0
with Timer() as t:
while True:
trials += 1
inp = mutate(seed_input)
if inp.startswith("https://"):
print(
"Success after",
trials,
"trials in",
t.elapsed_time(),
"seconds")
break
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# Of course, if we wanted to get, say, an `"ftp://"` prefix, we would need more mutations and more runs – most important, though, we would need to apply _multiple_ mutations.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multiple Mutations
#
# So far, we have only applied one single mutation on a sample string. However, we can also apply _multiple_ mutations, further changing it. What happens, for instance, if we apply, say, 20 mutations on our sample string?
# + slideshow={"slide_type": "subslide"}
seed_input = "http://www.google.com/search?q=fuzzing"
mutations = 50
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
inp = seed_input
for i in range(mutations):
if i % 5 == 0:
print(i, "mutations:", repr(inp))
inp = mutate(inp)
# + [markdown] slideshow={"slide_type": "fragment"}
# As you see, the original seed input is hardly recognizable anymore. By mutating the input again and again, we get a higher variety in the input.
# + [markdown] slideshow={"slide_type": "subslide"}
# To implement such multiple mutations in a single package, let us introduce a `MutationFuzzer` class. It takes a seed (a list of strings) as well as a minimum and a maximum number of mutations.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Fuzzer import Fuzzer
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class MutationFuzzer(Fuzzer):
def __init__(self, seed, min_mutations=2, max_mutations=10):
self.seed = seed
self.min_mutations = min_mutations
self.max_mutations = max_mutations
self.reset()
def reset(self):
self.population = self.seed
self.seed_index = 0
# + [markdown] slideshow={"slide_type": "subslide"}
# In the following, let us develop `MutationFuzzer` further by adding more methods to it. The Python language requires us to define an entire class with all methods as a single, continuous unit; however, we would like to introduce one method after another. To avoid this problem, we use a special hack: Whenever we want to introduce a new method to some class `C`, we use the construct
#
# ```python
# class C(C):
# def new_method(self, args):
# pass
# ```
#
# This seems to define `C` as a subclass of itself, which would make no sense – but actually, it introduces a new `C` class as a subclass of the _old_ `C` class, and then shadowing the old `C` definition. What this gets us is a `C` class with `new_method()` as a method, which is just what we want. (`C` objects defined earlier will retain the earlier `C` definition, though, and thus must be rebuilt.)
# + [markdown] slideshow={"slide_type": "subslide"}
# Using this hack, we can now add a `mutate()` method that actually invokes the above `mutate()` function. Having `mutate()` as a method is useful when we want to extend a `MutationFuzzer` later.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class MutationFuzzer(MutationFuzzer):
def mutate(self, inp):
return mutate(inp)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let's get back to our strategy, maximizing _diversity in coverage_ in our population. First, let us create a method `create_candidate()`, which randomly picks some input from our current population (`self.population`), and then applies between `min_mutations` and `max_mutations` mutation steps, returning the final result:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class MutationFuzzer(MutationFuzzer):
def create_candidate(self):
candidate = random.choice(self.population)
trials = random.randint(self.min_mutations, self.max_mutations)
for i in range(trials):
candidate = self.mutate(candidate)
return candidate
# + [markdown] slideshow={"slide_type": "subslide"}
# The `fuzz()` method is set to first pick the seeds; when these are gone, we mutate:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class MutationFuzzer(MutationFuzzer):
def fuzz(self):
if self.seed_index < len(self.seed):
# Still seeding
self.inp = self.seed[self.seed_index]
self.seed_index += 1
else:
# Mutating
self.inp = self.create_candidate()
return self.inp
# + slideshow={"slide_type": "subslide"}
seed_input = "http://www.google.com/search?q=fuzzing"
mutation_fuzzer = MutationFuzzer(seed=[seed_input])
mutation_fuzzer.fuzz()
# + slideshow={"slide_type": "fragment"}
mutation_fuzzer.fuzz()
# + slideshow={"slide_type": "fragment"}
mutation_fuzzer.fuzz()
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# With every new invocation of `fuzz()`, we get another variant with multiple mutations applied. The higher variety in inputs, though, increases the risk of having an invalid input. The key to success lies in the idea of _guiding_ these mutations – that is, _keeping those that are especially valuable._
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Guiding by Coverage
#
# To cover as much functionality as possible, one can rely on either _specified_ or _implemented_ functionality, as discussed in the ["Coverage"](Coverage.ipynb) chapter. For now, we will not assume that there is a specification of program behavior (although it _definitely_ would be good to have one!). We _will_ assume, though, that the program to be tested exists – and that we can leverage its structure to guide test generation.
#
# Since testing always executes the program at hand, one can always gather information about its execution – the least is the information needed to decide whether a test passes or fails. Since coverage is frequently measured as well to determine test quality, let us also assume we can retrieve coverage of a test run. The question is then: _How can we leverage coverage to guide test generation?_
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# One particularly successful idea is implemented in the popular fuzzer named [American fuzzy lop](http://lcamtuf.coredump.cx/afl/), or *AFL* for short. Just like our examples above, AFL evolves test cases that have been successful – but for AFL, "success" means _finding a new path through the program execution_. This way, AFL can keep on mutating inputs that so far have found new paths; and if an input finds another path, it will be retained as well.
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us build such a strategy. We start with introducing a `Runner` class that captures the coverage for a given function. First, a `FunctionRunner` class:
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Fuzzer import Runner
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class FunctionRunner(Runner):
def __init__(self, function):
"""Initialize. `function` is a function to be executed"""
self.function = function
def run_function(self, inp):
return self.function(inp)
def run(self, inp):
try:
result = self.run_function(inp)
outcome = self.PASS
except Exception:
result = None
outcome = self.FAIL
return result, outcome
# + slideshow={"slide_type": "fragment"}
http_runner = FunctionRunner(http_program)
http_runner.run("https://foo.bar/")
# + [markdown] slideshow={"slide_type": "subslide"}
# We can now extend the `FunctionRunner` class such that it also measures coverage. After invoking `run()`, the `coverage()` method returns the coverage achieved in the last run.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
from Coverage import Coverage, population_coverage
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class FunctionCoverageRunner(FunctionRunner):
def run_function(self, inp):
with Coverage() as cov:
try:
result = super().run_function(inp)
except Exception as exc:
self._coverage = cov.coverage()
raise exc
self._coverage = cov.coverage()
return result
def coverage(self):
return self._coverage
# + slideshow={"slide_type": "fragment"}
http_runner = FunctionCoverageRunner(http_program)
http_runner.run("https://foo.bar/")
# + [markdown] slideshow={"slide_type": "subslide"}
# Here are the first five locations covered:
# + slideshow={"slide_type": "fragment"}
print(list(http_runner.coverage())[:5])
# + [markdown] run_control={} slideshow={"slide_type": "subslide"}
# Now for the main class. We maintain the population and a set of coverages already achieved (`coverages_seen`). The `fuzz()` helper function takes an input and runs the given `function()` on it. If its coverage is new (i.e. not in `coverages_seen`), the input is added to `population` and the coverage to `coverages_seen`.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
class MutationCoverageFuzzer(MutationFuzzer):
def reset(self):
super().reset()
self.coverages_seen = set()
# Now empty; we fill this with seed in the first fuzz runs
self.population = []
def run(self, runner):
"""Run function(inp) while tracking coverage.
If we reach new coverage,
add inp to population and its coverage to population_coverage
"""
result, outcome = super().run(runner)
new_coverage = frozenset(runner.coverage())
if outcome == Runner.PASS and new_coverage not in self.coverages_seen:
# We have new coverage
self.population.append(self.inp)
self.coverages_seen.add(new_coverage)
return result
# + [markdown] run_control={} slideshow={"slide_type": "subslide"}
# Let us now put this to use:
# + slideshow={"slide_type": "fragment"}
seed_input = "http://www.google.com/search?q=fuzzing"
mutation_fuzzer = MutationCoverageFuzzer(seed=[seed_input])
mutation_fuzzer.runs(http_runner, trials=10000)
mutation_fuzzer.population
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Success! In our population, _each and every input_ now is valid and has a different coverage, coming from various combinations of schemes, paths, queries, and fragments.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
all_coverage, cumulative_coverage = population_coverage(
mutation_fuzzer.population, http_program)
import matplotlib.pyplot as plt
plt.plot(cumulative_coverage)
plt.title('Coverage of urlparse() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('lines covered');
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# The nice thing about this strategy is that, applied to larger programs, it will happily explore one path after the other – covering functionality after functionality. All that is needed is a means to capture the coverage.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * Randomly generated inputs are frequently invalid – and thus exercise mostly input processing functionality.
# * Mutations from existing valid inputs have much higher chances to be valid, and thus to exercise functionality beyond input processing.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# Our aim is still to sufficiently cover functionality, such that we can trigger as many bugs as possible. To this end, we focus on two classes of techniques:
#
# 1. Try to cover as much _specified_ functionality as possible. Here, we would need a _specification of the input format,_ distinguishing between individual input elements such as (in our case) numbers, operators, comments, and strings – and attempting to cover as many of these as possible. We will explore this as it comes to [grammar-based testing](GrammarFuzzer.ipynb), and especially in [grammar-based mutations](Parser.ipynb).
#
# 2. Try to cover as much _implemented_ functionality as possible. The concept of a "population" that is systematically "evolved" through "mutations" will be explored in depth when discussing [search-based testing](SearchBasedFuzzer.ipynb). Furthermore, [symbolic testing](SymbolicFuzzer.ipynb) introduces how to systematically reach program locations by solving the conditions that lie on their paths.
#
# These two techniques make up the gist of the book; and, of course, they can also be combined with each other. As usual, we provide runnable code for all. Enjoy!
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 1: Fuzzing CGI decode with Mutations
#
# Apply the above _guided_ mutation-based fuzzing technique on `cgi_decode()` from the ["Coverage"](Coverage.ipynb) chapter. How many trials do you need until you cover all variations of `+`, `%` (valid and invalid), and regular characters?
# + run_control={} slideshow={"slide_type": "skip"}
from Coverage import cgi_decode
# + run_control={} slideshow={"slide_type": "fragment"}
seed = ["Hello World"]
cgi_runner = FunctionCoverageRunner(cgi_decode)
m = MutationCoverageFuzzer(seed)
results = m.runs(cgi_runner, 10000)
# + slideshow={"slide_type": "fragment"}
m.population
# + slideshow={"slide_type": "subslide"}
cgi_runner.coverage()
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
all_coverage, cumulative_coverage = population_coverage(
m.population, cgi_decode)
import matplotlib.pyplot as plt
plt.plot(cumulative_coverage)
plt.title('Coverage of cgi_decode() with random inputs')
plt.xlabel('# of inputs')
plt.ylabel('lines covered');
# + [markdown] slideshow={"slide_type": "fragment"}
# After 10,000 runs, we have managed to synthesize a `+` character and a valid `%xx` form. We can still do better.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# ### Exercise 2: Fuzzing bc with Mutations
#
# Apply the above mutation-based fuzzing technique on `bc`, as in the chapter ["Introduction to Fuzzing"](Fuzzer.ipynb).
#
# #### Part 1: Non-Guided Mutations
#
# Start with non-guided mutations. How many of the inputs are valid?
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** This is just a matter of tying a `ProgramRunner` to a `MutationFuzzer`:
# + slideshow={"slide_type": "skip"} solution2="hidden"
from Fuzzer import ProgramRunner
# + slideshow={"slide_type": "skip"} solution2="hidden"
seed = ["1 + 1"]
bc = ProgramRunner(program="bc")
m = MutationFuzzer(seed)
outcomes = m.runs(bc, trials=100)
# + slideshow={"slide_type": "skip"} solution2="hidden"
outcomes[:3]
# + slideshow={"slide_type": "skip"} solution2="hidden"
sum(1 for completed_process, outcome in outcomes if completed_process.stderr == "")
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 2: Guided Mutations
#
# Continue with _guided_ mutations. To this end, you will have to find a way to extract coverage from a C program such as `bc`. Proceed in these steps:
#
# First, get [GNU bc](https://www.gnu.org/software/bc/); download, say, `bc-1.07.1.tar.gz` and unpack it:
# + slideshow={"slide_type": "fragment"}
# !curl -O http://ftp.gnu.org/gnu/bc/bc-1.07.1.tar.gz
# + slideshow={"slide_type": "fragment"}
# !tar xfz bc-1.07.1.tar.gz
# + [markdown] slideshow={"slide_type": "subslide"}
# Second, configure the package:
# + slideshow={"slide_type": "subslide"}
# !cd bc-1.07.1; ./configure
# + [markdown] slideshow={"slide_type": "subslide"}
# Third, compile the package with special flags:
# + slideshow={"slide_type": "subslide"}
# !cd bc-1.07.1; make CFLAGS="--coverage"
# + [markdown] slideshow={"slide_type": "subslide"}
# The file `bc/bc` should now be executable...
# + slideshow={"slide_type": "fragment"}
# !cd bc-1.07.1/bc; echo 2 + 2 | ./bc
# + [markdown] slideshow={"slide_type": "fragment"}
# ...and you should be able to run the `gcov` program to retrieve coverage information.
# + slideshow={"slide_type": "fragment"}
# !cd bc-1.07.1/bc; gcov main.c
# + [markdown] slideshow={"slide_type": "subslide"}
# As sketched in the ["Coverage" chapter](Coverage.ipynb), the file [bc-1.07.1/bc/main.c.gcov](bc-1.07.1/bc/main.c.gcov) now holds the coverage information for `bc.c`. Each line is prefixed with the number of times it was executed. `#####` means zero times; `-` means non-executable line.
# + [markdown] slideshow={"slide_type": "fragment"}
# Parse the GCOV file for `bc` and create a `coverage` set, as in `FunctionCoverageRunner`. Make this a `ProgramCoverageRunner` class that would be constructed with a list of source files (`bc.c`, `main.c`, `load.c`) to run `gcov` on.
# + [markdown] slideshow={"slide_type": "fragment"}
# When you're done, don't forget to clean up:
# + slideshow={"slide_type": "fragment"}
# !rm -fr bc-1.07.1 bc-1.07.1.tar.gz
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 3
#
# In this [blog post](https://lcamtuf.blogspot.com/2014/08/binary-fuzzing-strategies-what-works.html), the author of _American Fuzzy Lop_ (AFL), a very popular mutation-based fuzzer discusses the efficiency of various mutation operators. Implement four of them and evaluate their efficiency as in the examples above.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 4
#
# When adding a new element to the list of candidates, AFL does actually not compare the _coverage_, but adds an element if it exercises a new _branch_. Using branch coverage from the exercises of the ["Coverage"](Coverage.ipynb) chapter, implement this "branch" strategy and compare it against the "coverage" strategy, above.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 5
#
# Design and implement a system that will gather a population of URLs from the Web. Can you achieve a higher coverage with these samples? What if you use them as initial population for further mutation?
| docs/notebooks/MutationFuzzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 1-1.1 Intro Python
# ## Getting started with Python in Jupyter Notebooks
# - **Python 3 in Jupyter notebooks**
# - **`print()`**
# - **comments**
# - data types basics
# - variables
# - addition with Strings and Integers
# - Errors
# - character art
#
# -----
#
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - **use Python 3 in Jupyter notebooks**
# - **write working code using `print()` and `#` comments**
# - combine Strings using string addition (`+`)
# - add numbers in code (`+`)
# - troubleshoot errors
# - create character art
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concept</B></font>
# ## Hello World! - python `print()` statement
# Using code to write "Hello World!" on the screen is the traditional first program when learning a new language in computer science
#
# Python has a very simple implementation:
# ```python
# print("Hello World!")
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ## "Hello World!"
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/6f5784c6-eece-4dfe-a14e-9dcf6ee81a7f/Unit1_Section1.1-Hello_World.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/6f5784c6-eece-4dfe-a14e-9dcf6ee81a7f/Unit1_Section1.1-Hello_World.vtt","srclang":"en","kind":"subtitles","label":"english"}])
#
# Our "Hello World!" program worked because this notebook hosts a python interpreter that can run python code cells.
#
# Try showing
# ```python
# "Hello programmer!"
# ```
# enter new text inside the quotations in the cell above. Click on the cell to edit the code.
#
# What happens if any part of `print` is capitalized or what happens there are no quotation marks around the greeting?
# ## Methods for running the code in a cell
# 1. **Click in the cell below** and **press "Ctrl+Enter"** to run the code
# or
# 2. **Click in the cell below** and **press "Shift+Enter"** to run the code and move to the next cell
#
# 3. **Menu: Cell**...
# a. **> Run Cells** runs the highlighted cell(s)
# b. **> Run All Above** runs the highlighted cell and above
# c. **> Run All Below** runs the highlighted cell and below
# -
# <font size="4" color="#00A0B2" face="verdana"> <B>Example</B></font>
# + slideshow={"slide_type": "subslide"}
# [ ] Review the code, run the code
print("Hello World!")
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concept</B></font>
# ## Comments
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/34e2afb1-d07a-44ca-8860-bba1a5476caa/Unit1_Section1.1-Comments.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/34e2afb1-d07a-44ca-8860-bba1a5476caa/Unit1_Section1.1-Comments.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# When coding, programmers include comments for explanation of how code works for reminders and to help others who encounter the code
# ### comment start with the `#` symbol
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
# +
# this is how a comment looks in python code
# every comment line starts with the # symbol
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
# ## Program: "Hello World!" with comment
# - add a comment describing the code purpose
# - create an original "Hello World" style message
# +
# Add a comment "Hello world"
# -
print("Hello World!")
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Notebooks and Libraries
# Jupyter Notebooks provide a balance of jotting down important summary information along with proving a live code development environment where we can write and run python code. This course uses cloud hosted Jupyter [Notebooks](https://notebooks.azure.com) on Microsoft Azure and we will walk through the basics and some best practices for notebook use.
#
# ## add a notebook library
# - New: https://notebooks.azure.com/library > New Library
# - Link: from a shared Azure Notebook library link > open link, sign in> clone and Run
# - Add: open library > Add Notebooks > from computer > navigate to file(s)
#
# ## working in notebook cells
# - **Markdown cells** display text in a web page format. Markdown is code that formats the way the cell displays (*this cell is Markdown*)
#
# - **Code cells** contain python code and can be interpreted and run from a cell. Code cells display code and output.
#
# - **in edit** or **previously run:** cells can display in editing mode or cells can display results of *code* having been run
#
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/6b9134fc-c7d7-4d25-b0a7-bdb79d3e1a5b/Unit1_Section1.1-EditRunSave.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/6b9134fc-c7d7-4d25-b0a7-bdb79d3e1a5b/Unit1_Section1.1-EditRunSave.vtt","srclang":"en","kind":"subtitles","label":"english"}])
#
# ### edit mode
# - **text** cells in editing mode show markdown code
# - Markdown cells keep editing mode appearance until the cell is run
# - **code** (python 3) cells in editing look the same after editing, but may show different run output
# - clicking another cell moves the green highlight that indicates which cell has active editing focus
#
# ### cells need to be saved
# - the notebook will frequently auto save
# - **best practice** is to manually save after editing a cell using **"Ctrl + S"** or alternatively, **Menu: File > Save and Checkpoint**
#
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Altering Notebook Structure
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/cb195105-eee8-4068-9007-64b2392cd9ff/Unit1_Section1.1-Language_Cells.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/cb195105-eee8-4068-9007-64b2392cd9ff/Unit1_Section1.1-Language_Cells.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### add a cell
# - Highlight any cell and then... add a new cell using **Menu: Insert > Insert Cell Below** or **Insert Cell Above**
# - Add with Keyboard Shortcut: **"ESC + A"** to insert above or **"ESC + B"** to insert below
#
# ### choose cell type
# - Format cells as Markdown or Code via the toolbar dropdown or **Menu: Cell > Cell Type > Code** or **Markdown**
# - Cells default to Code when created but can be reformatted from code to Markdown and vice versa
#
# ### change notebook page language
# - The course uses Python 3 but Jupyter Notebooks can be in Python 2 or 3 (and a language called R)
# - To change a notebook to Python 3 go to **"Menu: Kernel > Change Kernel> Python 3"**
#
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
# ## Insert a new cell
# - Insert a new Code cell below with a comment describing the task
# - edit cell: add print() with the message "after edit, save!"
# - run the cell
# +
# display "after edit save"
# -
print("after edit save!")
# ### Insert another new cell
# - Insert a new Code cell below
# - edit cell: add print() with the message showing the keyboard Shortcut to save **Ctrl + s**
# - run the cell
print("CTRL + s")
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Absolute Beginner/Module_1_1.1_Absolute_Beginner_START_HERE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Tests with kernel_size 4
# +
mask_type = 'V'
kernel_size=(4, 4)
padding = keras.layers.ZeroPadding2D(padding=((1,0),0))
conv = MaskedConv2D(mask_type=mask_type,
filters=1,
kernel_size=kernel_size,
padding='same',
kernel_initializer='ones',
bias_initializer='zeros')
cropping = keras.layers.Cropping2D(cropping=((0, 1), 0))
x = padding(test_ones_2d)
x = conv(x)
result = cropping(x)
print('MASK')
print(conv.mask.numpy().squeeze())
print('')
print('OUTPUT')
print(result.numpy().squeeze())
# +
mask_type = 'A'
kernel_size=(1, 4)
conv = MaskedConv2D(mask_type=mask_type,
filters=1,
kernel_size=kernel_size,
padding='same',
kernel_initializer='ones',
bias_initializer='zeros')
result = conv(test_ones_2d)
print('MASK')
print(conv.mask.numpy().squeeze())
print('')
print('OUTPUT')
print(result.numpy().squeeze())
# +
mask_type = 'B'
kernel_size=(1, 4)
conv = MaskedConv2D(mask_type=mask_type,
filters=1,
kernel_size=kernel_size,
padding='same',
kernel_initializer='ones',
bias_initializer='zeros')
result = conv(test_ones_2d)
print('MASK')
print(conv.mask.numpy().squeeze())
print('')
print('OUTPUT')
print(result.numpy().squeeze())
| WIP/6-gated_pixelcnn_cropped/masked_vs_cropped_even_filter_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python-tutorial]
# language: python
# name: conda-env-python-tutorial-py
# ---
#Use miniconda-analysis environment
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
import co2_timeseries_tools as co2tt
from scipy import stats
from scipy import signal
from ctsm_py import utils
import pandas as pd
# ### Obs Data
path='/glade/work/dll/CTSM_py/notebooks/'
brw={'name':'Barrow', 'acronym': 'brw', 'lat': 71.3, 'lon':360-156.61, 'z': 11.0}
mlo={'name':'<NAME>', 'acronym': 'mlo', 'lat': 19.5, 'lon':360-155.6, 'z':3397.0}
alt={'name':'Alert', 'acronym': 'alt', 'lat': 82.5, 'lon':360-62.5, 'z':200.0}
azr={'name': 'Azores', 'acronym':'azr','lat':38.8, 'lon':360-27.4, 'z':19.0}
cba={'name': '<NAME>', 'acronym':'cba', 'lat':55.2, 'lon':360-162.7, 'z':21.3}
kum={'name':'Kumukahi', 'acronym':'kum', 'lat':19.7, 'lon':360-155.0, 'z':0.3}
ESRL=[brw, mlo, alt, azr, cba, kum]
lats=np.array([71.3,19.5,82.5,38.8,55.2,19.7])
lons=np.array([360-156.61,360-155.6,360-62.5,360-27.4,360-162.7,360-155.0])
#note that the 'lev' variable only goes from 0 to 1000, not sure how to translate from 'alt' to 'lev'
alt=np.array([11.0,3397.0,200.0,19.0,21.3,0.3])
cesm1levs=np.array([25,20,25,25,25,25])
cesm2levs=np.array([31,22,31,31,31,31])
# +
minYear = 1981 # minimum year for an 'early' trend, used later for plotting
for site in ESRL:
# print(site)
filename=path+'co2_'+site['acronym']+'_surface-flask_1_ccgg_month.txt'
#import glob
#filename=glob.glob(partialname+ '*co2')
with open(filename, 'r') as fid:
first_line=fid.readline()
nheader=first_line[-3:-1]
nheader=np.int(np.float(nheader))
data=np.loadtxt(filename, usecols=(1,2,3), skiprows=nheader)
time=data[:,0]+data[:,1]/12
co2=data[:,2]
month=data[:,1]
year=data[:,0]
site['year']=year
site['month']=month
site['co2']=co2
#for y in range(len(site['year'])):
# site['min'][y] = site['co2'].min()
# site['max'][y] = site['co2'].max()
# -
# ### Problem 1
# I would like to calculate max - min for each year but can't figure out how to loop over the years.
# The code below calculates absolute min and max across all years
#
# ### Solution 1
# Ideally, I'd do this with groupby, or assign 'name' as the index for year & CO2, but this isn't working as I'd expect
#
# Instead I'll just create data arrays to hold output I'm expecting, and throw out years with < 12 month of data
# +
#convert to pandas dataframe
df = pd.DataFrame(ESRL,columns = ['name','year','co2'])
# create arrays to hold results (here xr)
# nobs will be used to count obs for each year & mask values when < 12
years = range(int(ESRL[0].get('year').min()),int(ESRL[0].get('year').max()))
minCO2 = xr.DataArray(np.nan, coords={'name': df.name, 'year':years}, dims=['name','year'])
maxCO2 = xr.DataArray(np.nan, coords={'name': df.name, 'year':years}, dims=['name','year'])
nobs = xr.DataArray(np.nan, coords={'name': df.name, 'year':years}, dims=['name','year'])
# loop through each site & year
for i in range(len(df.name)):
for j in range(len(years)):
temp = np.where(ESRL[i]['year']==years[j],ESRL[i]['co2'],np.nan)
maxCO2[i,j] = np.nanmax(temp)
minCO2[i,j] = np.nanmin(temp)
nobs[i,j] = np.isnan(temp)[np.isnan(temp) == False].size
ampCO2 = maxCO2 - minCO2
ampCO2 = ampCO2.where(nobs==12)
ampCO2.plot.line(hue='name',x='year');
#print()
# -
# not sure how you want to handle missing data...?
# ### Model Data
# +
simyrs = "185001-201412"
var = "CO2"
datadir = "/glade/p/cesm/lmwg_dev/dll/"
subdir = "/atm/proc/tseries/month_1/"
Mod1dir = "CESM2_Coupled_NoCrop/"
sim = "b.e21.BHIST_BPRP.f09_g17.CMIP6-esm-hist.001"
sim2 = "b40.20th.1deg.coup.001"
# -
data1 = utils.time_set_mid(xr.open_dataset(datadir+Mod1dir+sim+".cam.h0."+var+"."+simyrs+".nc", decode_times=True), 'time')
data2 = utils.time_set_mid(xr.open_dataset(datadir+Mod1dir+sim2+".cam2.h0."+var+".185001-200512.nc", decode_times=True), 'time')
# ### Converting CO2 units to ppm
#conversion of CESM CO2 to ppm
convert = 10.0**6 * 28.966/44.0
# +
# %%time
## Are you intendings to swap CESM1 with data2 and CESM2 with data1?
CESM1ppm = data2.CO2.sel(time=slice('1950','2014')) * convert
CESM2ppm = data1.CO2.sel(time=slice('1950','2014')) * convert
CESM1ppm.attrs['units'] = 'ppm'
CESM2ppm.attrs['units'] = 'ppm'
# -
# ### Selecting sites for comparison to observations
# +
#initialize list using '[]' and dictionary using '{}'
CESM1points = {}
CESM2points = {}
#lat and lon are actual values to pull out, level refers to a specific index, not a value, so requires 'isel'
for x in range(6):
CESM1pointloop = CESM1ppm.sel(lat=lats[x], lon=lons[x], method="nearest")
CESM2pointloop = CESM2ppm.sel(lat=lats[x], lon=lons[x], method="nearest")
CESM1pointloop = CESM1pointloop.isel(lev=cesm1levs[x])
CESM2pointloop = CESM2pointloop.isel(lev=cesm2levs[x])
CESM1points[x] = CESM1pointloop
CESM2points[x] = CESM2pointloop
# -
temp = CESM2points[0].sel(time=slice('1980','1985'))
#temp.plot()
xr.DataArray(signal.detrend(temp),coords={'time':temp.time}, dims='time').groupby("time.month").mean().plot();
# ### Calculating detrended annual cycle for early and late time periods
# +
CESM1_ann_early = {}
CESM1_ann_late = {}
CESM2_ann_early = {}
CESM2_ann_late = {}
CESM1_ann_early_detrend = {}
CESM1_ann_late_detrend = {}
CESM2_ann_early_detrend = {}
CESM2_ann_late_detrend = {}
for site, data in CESM1points.items():
CESM1late = data.sel(time=slice('2000','2005'))
CESM1early = data.sel(time=slice('1980','1985'))
CESM1_ann_late[site] = CESM1late.groupby("time.month").mean()
CESM1_ann_early[site] = CESM1early.groupby("time.month").mean()
CESM1_ann_early_detrend[site] = xr.DataArray(signal.detrend(CESM1early),coords={'time':CESM1early.time}, dims='time').groupby("time.month").mean()
CESM1_ann_late_detrend[site] = xr.DataArray(signal.detrend(CESM1late),coords={'time':CESM1late.time}, dims='time').groupby("time.month").mean()
print(CESM1_ann_early_detrend[1])
print(type(CESM1_ann_early_detrend))
for site, data in CESM2points.items():
CESM2late = data.sel(time=slice('2000','2005'))
CESM2early = data.sel(time=slice('1980','1985'))
CESM2_ann_late[site] = CESM2late.groupby("time.month").mean()
CESM2_ann_early[site] = CESM2early.groupby("time.month").mean()
CESM2_ann_early_detrend[site] = xr.DataArray(signal.detrend(CESM2early),coords={'time':CESM2early.time}, dims='time').groupby("time.month").mean()
CESM2_ann_late_detrend[site] = xr.DataArray(signal.detrend(CESM2late),coords={'time':CESM2late.time}, dims='time').groupby("time.month").mean()
print(CESM1_ann_early_detrend[1])
# -
# ## Problem #2
# ### Detrended plot has oddly high late-season values
# Bottom plot is detrended. Should I be using a different function?
#
# ## solution
# Above, I changed when you're taking the mean of the detrended data. Now the data are detrended, then the monthly averages are taken
# ### Absolute value plot
CESM1_ann_early[0].plot();
# ### Detrended Plot
plt.plot(CESM1_ann_early_detrend[0])
plt.plot(CESM1_ann_late_detrend[0]);
| notebooks/CO2Amp_Problems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# uncomment this on first run
# # !wget 'https://archive.ics.uci.edu/ml/machine-learning-databases/00331/sentiment%20labelled%20sentences.zip'
# -
# !ls
# +
# uncomment this on first run
# # !unzip sentiment\ labelled\ sentences.zip
# -
# !ls
# !ls sentiment\ labelled\ sentences
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import scipy
import sklearn
import matplotlib.pyplot as plt
import seaborn as sns
# Grab and process the raw data.
data_path = ("sentiment labelled sentences/amazon_cells_labelled.txt")
sms_raw = pd.read_csv(data_path, delimiter= '\t', header=None)
sms_raw.head()
# -
sms_raw.columns = ['message', 'pos_neg']
# +
# I think the keywords are:
keywords = ['good', 'bad', 'great', 'disappoint', 'negative', 'unsatisfactory', 'sweet', 'excellent']
for key in keywords:
# Note that we add spaces around the key so that we're getting the word,
# not just pattern matching.
sms_raw[str(key)] = sms_raw.message.str.contains(
' ' + str(key) + ' ',
case=False
)
sms_raw.head()
# +
# making a true false columm
sms_raw["pos_neg_bool"] = (sms_raw["pos_neg"] == 0)
sms_raw.head()
# +
# Our data is binary / boolean, so we're importing the Bernoulli classifier.
from sklearn.naive_bayes import BernoulliNB
# Instantiate our model and store it in a new variable.
bnb = BernoulliNB()
data = sms_raw[keywords]
target = sms_raw["pos_neg_bool"]
bnb.fit(data, target)
y_pred = bnb.predict(data)
# Display our results.
print("Number of mislabeled points out of a total {} points : {}".format(
data.shape[0],
(target != y_pred).sum()
))
# -
| feedback-analysis-with-naive-bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
def calculaPorDia(x):
#print(x[3]/x[6].days)
#print(x[3],'',x[7])
return x[3]/x[7].days
hotels = []
with open('hotels-in-Munich.txt') as f:
for line in f:
if len(line)>10:
hotels.append([str(n.replace('\xa0','').replace('"','').replace('[','').replace(']','')) for n in line.strip().split(';')])
#print(line)
import datetime
hotelDict = []
for i in hotels:
#print(i[0][3:5])
dtIni = datetime.datetime(int(i[0][6:10]), int(i[0][3:5]), int(i[0][0:2]), 0, 0, 0)
dtEnd = datetime.datetime(int(i[1][6:10]), int(i[1][3:5]), int(i[1][0:2]), 0, 0, 0)
hotelDict.append({'de':dtIni,'ate':dtEnd,'nome':i[2],'preco':i[3],'nota':i[4],'distancia':i[5],'url':i[6]})
df = pd.DataFrame(hotelDict)
df.drop_duplicates(inplace = True)
df = df[df['preco']>'-1']
df['preco'].replace(regex=True,inplace=True,to_replace=r'\D',value=r'')
df['preco'] = pd.to_numeric(df['preco'])
df['dias'] = (df['ate']-df['de']).replace(regex=True,to_replace=r'\D',value=r'')
df["precoPorNoite"] = df.apply(calculaPorDia,axis=1)
display(df.describe())
# +
datetimeFormat = '%Y-%m-%d %H:%M:%S.%f'
#df['diasteste'] = datetime.datetime.strptime(df['ate'],datetimeFormat) - datetime.datetime.strptime(df['de'],datetimeFormat)
df['dias'] = (df['ate']-df['de']).replace(regex=True,to_replace=r'\D',value=r'')
df["nota"] = pd.to_numeric(df["nota"])
df['distanciaMedida'] = pd.np.where(df.distancia.str.contains("km"),"km","m")
df["distanciaNumerica"] = pd.to_numeric(df['distancia'].replace(regex=True,to_replace=r'\D',value=r''))#, downcast='float')
df["distanciaMetros"] = pd.np.where(df.distancia.str.contains("km") & df.distancia.str.contains("."),df.distanciaNumerica*1000,df["distanciaNumerica"])
df["distanciaMetros"] = pd.np.where(df.distanciaMetros > 9000,(df.distanciaNumerica/10),df["distanciaNumerica"])
df["distanciaMetros"] = df["distanciaMetros"].apply(lambda x: x*1000 if x < 100 else x)
#df["precoPorNoite"] = df.apply(calculaPorDia,axis=1)
df = df[df['precoPorNoite']<800]
df = df[df['nota']>7.5]
df = df[df['distanciaMetros']<2000]
df = df[~df["nome"].str.contains('Hostel')]
del df['distanciaMedida']
del df['distanciaNumerica']
df.drop_duplicates(inplace = True)
display(df.describe())
display(df)
# -
import plotly.express as px
#fig = px.scatter(df, x="precoPorNoite", y="nota",color="nome")
#fig = px.scatter(df, x="preco", y="nota",color="nome")
#fig.show()
import plotly.express as px
fig = px.scatter(df, x="precoPorNoite", y="nota", size="distanciaMetros", color="nome",hover_name="nome",log_x=True)
fig.show()
# +
df['link'] ='<a href="http://'+df['url']+'">Clique</a>'
df= df.drop_duplicates(subset=['nome', 'preco'])
dfa = df.copy()
del dfa['url']
del dfa['de']
del dfa['ate']
del dfa['dias']
del dfa['distanciaMetros']
dfa = dfa.to_html()
print('de: ',df['de'].max())
print('até: ',df['ate'].max())
#display(dfa)
#display(df.drop_duplicates().sort_values(by=['precoPorNoite'],ascending=False))
from IPython.core.display import display, HTML
display(HTML(dfa))
# -
| BookingScraper-joao_v2/BookingScraper/bookingReadMunique.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import os
import sys
import fnmatch
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from collections import defaultdict
# -
plt.style.use('fivethirtyeight')
# +
WS_REAL = defaultdict(list)
valid_final_season = {}
FEAT = ['Age','WS']
for YR in range(1980,2020):
table = np.load('tables_{}.pkl'.format(YR),allow_pickle=True)
for team in table:
stats = table[team]['advanced'][FEAT]
for item in stats.itertuples():
try:
v = [0.0 if _=='' else _ for _ in item[1:]]
#hgt = table[team]['roster'].loc[item[0]]['Ht']
#hgt = [int(_) for _ in hgt.split('-')]
#hgt = 12*hgt[0] + hgt[1]
#v.append(hgt)
WS_REAL[item[0]].append(np.array(v).astype(np.float))
except:
print(item)
# -
d = []
for k in WS_REAL:
t = np.array(WS_REAL[k])
tm = t[:,1]#*t[:,3]
tm = tm.astype(np.float)
d.append(np.vstack([[k for _ in range(tm.shape[0])],tm,t[:,1]]).T)
table = np.load('tables_{}.pkl'.format(2019),allow_pickle=True)
for team in table:
stats = table[team]['advanced']
for item in stats.itertuples():
valid_final_season[item[0]] = 1
X = []
y = []
for name in WS_REAL:
stats = WS_REAL[name]
yrs = len(stats)
X += stats
for i in range(yrs-1):
y.append(0.0)
y.append(float(name not in valid_final_season))
from sklearn.linear_model import LogisticRegression, ElasticNet, Ridge,RidgeClassifier
from sklearn.svm import LinearSVC,SVC
from sklearn.preprocessing import PolynomialFeatures,StandardScaler
Xo = np.array(X)
yo = np.array(y)
X = Xo.copy()
y = yo.copy()
X[:,0] -= 18
X[:,0] *= X[:,0]
#X[:,-1] /= X[:,-1].max()
#X = np.hstack([X,(X[:,0] * X[:,1]).reshape((-1,1))])
# +
fexp = PolynomialFeatures(2,interaction_only=True)
scal = StandardScaler()
pX = X#fexp.fit_transform(X)#fexp.fit_transform(X)
#pX[:,0] = pX[:,0]**2
clf = LogisticRegression(C=1e6,solver='liblinear')#,class_weight='balanced')
clf.fit(pX,y)
clf.score(pX,y),(1-y.sum()/y.shape[0])
# -
_ = plt.hist(clf.predict_proba(pX)[:,1],50,density=True)
FEAT,clf.coef_,clf.intercept_
import statsmodels.api as sm
FEAT
# +
X2 = sm.add_constant(pX)
df_tmp = pd.DataFrame(X2,columns=['1'] + FEAT )
df_tmp.columns = ['(Age-18)^2' if _ == 'Age' else _ for _ in df_tmp.columns]
logit_mod = sm.Logit(y,df_tmp)#fexp.get_feature_names(FEAT)))
logit_res = logit_mod.fit(disp=0)
print()
np.mean((logit_res.predict() >0.5)== y.astype(np.bool))
logit_res.summary()
# +
xp = np.linspace(18,50)
p = lambda x: 1.0/(1.0+np.exp(-x))
#p = lambda x: np.log(np.exp(x)+1)
for WS in [-1,0,1,3,5,7]:
#plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*xp),label='WS0')
plt.plot(xp,p(logit_res.params[0] + logit_res.params[1]*(xp-18)**2 + logit_res.params[2]*WS ),label='WS {}'.format(WS))
plt.xlim(18,55)
plt.legend()
plt.xlabel('Age')
plt.ylabel('Prob of Retiring')
plt.title('NBA (Age-18)^2 + WS Model')
plt.ylim(0,1)
plt.xlim(18,50)
plt.tight_layout()
plt.grid(True)
plt.savefig('retire.png',facecolor='w',edgecolor='w')
# +
xp = np.linspace(18,60)
p = lambda x: 1.0/(1.0+np.exp(-x))
#p = lambda x: np.log(np.exp(x)+1)
for WS in [-1,0,1,3,5,7]:
#plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*xp),label='WS0')
plt.plot(xp,100*p(clf.intercept_ + clf.coef_[0,0]*(xp-18)**2 + clf.coef_[0,1]*WS ),label='WS {}'.format(WS))
plt.xlim(18,55)
plt.legend()
plt.xlabel('Age')
plt.ylabel('Prob of Retiring')
plt.title('New BBGM Model')
plt.ylim(0,100)
plt.xlim(35,60)
plt.tight_layout()
plt.grid(True)
plt.savefig('retire.png',facecolor='w',edgecolor='w')
# -
pX
df2 = pd.read_csv('beta_stats.csv')#big_stat
from matplotlib.colors import LogNorm
plt.style.use('fivethirtyeight')
dft = df2[df2.MP*df2.G > 300]
_ = plt.hexbin(dft.Ovr,dft['WS'],gridsize=40,norm=LogNorm())
#plt.ylim(-5,40)
#plt.xlim(30,85)
plt.xlabel('Ovr')
plt.ylabel('WS')
xp = np.linspace(30,90)
#plt.plot(xp,(1/64)*np.maximum(xp-40,0)**2,c='r')
plt.title('bbgm ovr -> WS')
plt.savefig('wsovr3.png',edgecolor='w',facecolor='w')
# +
WS_FAKE = defaultdict(list)
valid_final_season2 = {}
FEAT2 = ['Name','Age','WS','TS%','3PAr','Pot']
for item in df2[FEAT2].itertuples():
try:
v = [0.0 if _=='' else _ for _ in item[2:]]
#hgt = table[team]['roster'].loc[item[0]]['Ht']
#hgt = [int(_) for _ in hgt.split('-')]
#hgt = 12*hgt[0] + hgt[1]
#v.append(hgt)
WS_FAKE[item[1]].append(np.array(v).astype(np.float))
except:
raise#print(item)
# -
X2 = []
y2 = []
for name in WS_FAKE:
stats = WS_FAKE[name]
yrs = len(stats)
X2 += stats
for i in range(yrs-1):
y2.append(0.0)
y2.append(1.0)
if len(y2) != len(X2):
print('omg',len(y2),len(X2),name)
X2 = np.array(X2)
y2 = np.array(y2)
# +
clf2 = LogisticRegression(C=1e6,solver='liblinear')#,class_weight='balanced')
clf2.fit(X2,y2)
clf2.score(X2,y2),(1-y2.sum()/y2.shape[0])
# -
_ = plt.hist(clf2.predict_proba(X2)[:,1],50,density=True)
FEAT2,clf2.coef_,clf2.intercept_
X3 = sm.add_constant(X2)
X3 = pd.DataFrame(X3,columns=['1'] + FEAT2[1:])
logit_mod = sm.Logit(y2,X3)
logit_res = logit_mod.fit(disp=0)
print(logit_res.summary())
np.mean((logit_res.predict() >0.5)== y2.astype(np.bool))
xp = np.linspace(20,40)
p = lambda x: 1.0/(1.0+np.exp(-x))
for WS in [-1,0,1,3,5,7]:
#plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*xp),label='WS0')
plt.plot(xp,p(clf2.intercept_ + clf2.coef_[0,0]*xp + clf2.coef_[0,1]*WS+ clf2.coef_[0,2]*30+ clf2.coef_[0,3]*5 + clf2.coef_[0,4]*50 ),label='WS {}'.format(WS))
plt.xlim(20,46)
plt.legend()
plt.title('BBGM, TS%:30, 3PAr:5')
plt.xlabel('Age')
plt.ylabel('P(Retire | Age, WS)')
plt.tight_layout()
plt.savefig('retireFAKE.png',facecolor='w',edgecolor='w')
df2['OvrSm'] = (np.maximum(0,df2['Ovr']-37))**2
df3 = df2[df2.Age > 22]
clf_ws = sm.OLS(df3.WS,df3['OvrSm']).fit()
from scipy.stats import pearsonr
print(pearsonr(clf_ws.predict(),df3.WS)[0])
clf_ws.summary()
plt.scatter(df3.WS,clf_ws.predict(),alpha=0.5,s=8)
clf3 = ElasticNet(alpha=0,fit_intercept=False)
#df3 = df2[df2.Age > 22]
clf3.fit(np.array(df3.OvrSm).reshape((-1,1)),df3.WS)
clf3.score(np.array(df3.OvrSm).reshape((-1,1)),df3.WS)
# +
xp = np.linspace(18,60)
p = lambda x: 1.0/(1.0+np.exp(-x))
#p = lambda x: np.log(np.exp(x)+1)
for OVR in [50,55,60,65]:
#plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*xp),label='WS0')
plt.plot(xp,100*p(clf.intercept_ + clf.coef_[0,0]*(xp-18)**2 + clf.coef_[0,1]* clf3.coef_[0]*(OVR-37)**2 ),label='OVR {}'.format(OVR))
plt.xlim(18,55)
plt.legend()
plt.xlabel('Age')
plt.ylabel('Prob of Retiring')
plt.title('New BBGM Retirement')
plt.ylim(0,100)
plt.xlim(35,60)
plt.tight_layout()
plt.grid(True)
plt.savefig('retire3.png',facecolor='w',edgecolor='w')
# -
clf3.coef_,clf3.intercept_,clf3.coef_[0]
clf3.coef_[0]*50+clf3.intercept_
# +
xp = np.linspace(20,60)
p = lambda x: 1.0/(1.0+np.exp(-x))
for POT in [40,50,60,70]:
#plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*xp),label='WS0')
plt.plot(xp,p(clf.intercept_ + clf.coef_[0,0]*(xp-18)**2 + clf.coef_[0,1]* (clf3.coef_[0]*((POT-37)**2)) ),label='Pot {}'.format(POT))
plt.xlim(20,55)
plt.ylim(0,1)
plt.legend()
plt.xlabel('Age')
plt.ylabel('P(Retire | Age, Pot)')
plt.title('Overall model')
plt.tight_layout()
plt.savefig('retire2.png',facecolor='w',edgecolor='w')
# -
clf.intercept_ ,1/clf.coef_[0,0],clf.coef_[0,1]
# +
df2['retire'] = clf.intercept_ + clf.coef_[0,0]*(np.maximum(df2.Age,18)-18)**2 + clf.coef_[0,1]*np.maximum(df2.WS,clf3.coef_[0]*((np.maximum(37,df2.Pot)-37)**2))
df2['retire'] = 1/(1+np.exp(-df2.retire))
# -
1/clf3.coef_,clf3.intercept_
plt.hist(df2.retire,20)
retired = df2[df2.retire >0.5][['WS','Age','Pot','retire']]
retired
plt.style.use('fivethirtyeight')
plt.figure(figsize=(10,6))
for i,col in enumerate(retired.columns):
plt.subplot(2,2,1+i)
plt.hist(df2[df2.retire < 0.5][col],20,density=True,alpha=0.8,label='not retired')
plt.hist(df2[df2.retire > 0.5][col],8,density=True,alpha=0.8,label='retired')
plt.title('Retirement ' + col + '\n (weight: Balanced)')
if i == 0:
plt.xlim(-6,10)
plt.legend()
plt.tight_layout()
plt.hexbin(df2.Age,df2.retire,gridsize=17)
plt.xlabel('Age')
plt.ylabel('P(Retire)')
plt.title('Balanced')
plt.colorbar()
dft = df2[(df2.Ovr < 62) & (df2.WS > 16)]
dft
pd.set_option("display.precision", 2)
df2[(df2.Season == 2187) & (df2.Team == 'MIA')].sort_values('MP',0,False)[['Name','Pos','Age','Ovr','Salary','GS','MP','WS','PTS','TRB','AST','STL','Blk','TOV']]
from tabulate import tabulate
print(tabulate(df2[(df2.Season == 2187) & (df2.Team == 'MIA')].sort_values('MP',0,False)[['Name','Pos','Age','Salary','GS','MP','WS','PTS','TRB','AST','STL','Blk','TOV']]))
| retire.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Watch Me Code 2: Dictionary Methods
#
student = { 'Name' : 'Michael', 'GPA': 3.2, 'Ischool' : True }
print(student)
# KeyError, keys must match exactly
student['gpa']
try:
print(student['Age'])
except KeyError:
print('The key "Age" does not exist.')
list(student.values())
list(student.keys())
for key in student.keys():
print(student[key])
del student['GPA']
print(student)
| content/lessons/10/Watch-Me-Code/WMC2-Dict-Methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Define inputs for the script
z_blocks = [{'z_start':, 'N_zs':, 'z_space':, 'channel_each_z':[]},
{'z_start':, 'N_zs':, 'z_space':, 'channel_each_z':[]}]
shutter_file = r"test.xml"
# +
# Generate the z positions
z_positions = []
for z_b in z_blocks:
for i in range(z_b['N_zs']):
for j in range(len(z_b['channel_each_z'])):
z_positions.append(z_b['z_start'] + z_b['z_space'] * i)
print(','.join([f'{z:.1f}' for z in z_positions]))
# +
# Generate the shutter file
head = f'''<?xml version="1.0" encoding="ISO-8859-1"?>
<repeat>
<oversampling>1</oversampling>
<frames>{len(z_positions)}</frames>
'''
events = ''
end = '</repeat>'
count = 0
for z_b in z_blocks:
for i in range(z_b['N_zs']):
for j in range(len(z_b['channel_each_z'])):
channel = z_b['channel_each_z'][j]
if not (channel is None):
events += f''' <event>
<channel>{channel}</channel>
<power>1</power>
<on>{count:.1f}</on>
<off>{count + 1:.1f}</off>
<color>255,255,255</color>
</event>
'''
count += 1
events += '\n'
print(head + events + end)
with open(shutter_file, 'w') as f:
f.write(head + events + end)
| data_acquisition/imaging/generate_z_positions_and_shutter_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The goal of data cleaning notebook for `wiai-facility` is 3-fold:
#
# 1. Create symlinks from `raw/audio/*.wav` <- `processed/audio/*.wav`
# 2. Use `CaC_work_sheet.csv` to create `processed/annotations.csv`
# 3. Use `CaC_work_sheet.csv` to create `processed/attributes.csv`
# %load_ext autoreload
# %autoreload 2
# +
from os import makedirs, symlink, rmdir, listdir
from os.path import join, dirname, exists, isdir, basename, splitext
from shutil import rmtree
import math
from collections import defaultdict
import pandas as pd
import numpy as np
from glob import glob
from tqdm import tqdm
import librosa
from librosa import get_duration
import scipy.io.wavfile as wav
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
from termcolor import colored
from cac.utils.io import save_yml
from cac.utils.pandas import apply_filters
from cac.utils.file import get_audio_type, get_unique_id
# +
# directory where the data resides
data_root = '/data/wiai-facility/'
# src and destination directories
load_root = join(data_root, 'raw')
save_root = join(data_root, 'processed')
makedirs(save_root, exist_ok=True)
load_audio_dir = join(load_root, 'audio')
save_audio_dir = join(save_root, 'audio')
makedirs(save_audio_dir, exist_ok=True)
# -
data_sheet = pd.read_csv(join(load_audio_dir, 'CaC_work_sheet_nov23-anonymized.csv'))
data_sheet.shape
# #### Important check: find out files that are unreadable via `EOFError` (can't be discovered by `librosa`)
files = glob(join(load_audio_dir, '*/*/*.wav'))
len(files)
# #### Generate symlinks from `raw` <- `processed`
# +
user_dirs = [f for f in glob(join(load_audio_dir, '*')) if isdir(f)]
files = []
timestamps = []
users = []
for user_dir in tqdm(user_dirs):
user_files = glob(join(user_dir, '*/*.wav'))
for user_file in user_files:
users.append(basename(user_dir))
filename = basename(user_file)
if 'breathing' in filename:
filename = 'breathing'
elif 'cough_sound_recording_1' in filename:
filename = 'cough_1'
elif 'cough_sound_recording_2' in filename:
filename = 'cough_2'
elif 'cough_sound_recording_3' in filename:
filename = 'cough_3'
elif 'speech_recording' in filename:
filename = 'audio_1_to_10'
elif 'room_recording' in filename:
filename = 'room_sound'
elif 'aaaaa_recording' in filename:
filename = 'a_sound'
elif 'eeeee_recording' in filename:
filename = 'e_sound'
elif 'ooooo_recording' in filename:
filename = 'o_sound'
else:
import ipdb; ipdb.set_trace()
# strongly dependent on structure
timestamps.append(user_file.split('/')[-2])
save_filename = '_'.join([*user_file.split('/')[-3:-1], filename + '.wav'])
save_path = join(save_audio_dir, save_filename)
# ignore .wav
files.append(splitext(save_filename)[0])
if not exists(save_path):
symlink(user_file, save_path)
# -
# #### Creating `attributes.csv` and `annotations.csv`
sound_labels = {
'breathing': 'breathing',
'cough_1': 'cough',
'cough_2': 'cough',
'cough_3': 'cough',
'audio_1_to_10': 'audio_1_to_10',
'room_sound': 'room_sound',
'a_sound': 'a_sound',
'e_sound': 'e_sound',
'o_sound': 'o_sound'
}
unsup_label_keys = [
'enroll_patient_gender',
'patient_id',
'enroll_patient_age',
'enroll_state',
'enroll_facility',
'enroll_habits',
'enroll_travel_history',
'enroll_comorbidities',
'enroll_contact_with_confirmed_covid_case',
'enroll_fever',
'enroll_days_with_fever',
'enroll_cough',
'enroll_days_with_cough',
'enroll_shortness_of_breath',
'enroll_days_with_shortness_of_breath',
'enroll_patient_temperature',
'enroll_patient_respiratory_rate',
'enroll_cough_relief_measures',
'testresult_covid_test_result'
]
data_sheet[unsup_label_keys] = data_sheet[unsup_label_keys].fillna('NA')
# +
files = []
unsup_labels = []
clf_labels = []
users = []
for index in tqdm(range(len(data_sheet)), desc="Iterating over all patients"):
row = data_sheet.loc[index]
recording_dir = row['audio_folder']
user_timestamp = '_'.join(recording_dir.split('/')[-2:])
user = user_timestamp.split('/')[0]
disease_status = row['testresult_covid_test_result']
user_files = []
user_clf_labels = []
user_unsup_labels = []
user_ids = []
for key, value in sound_labels.items():
file = '/'.join([save_audio_dir, '_'.join([user_timestamp, f'{key}.wav'])])
if key != 'room_sound':
clf_label = [value, disease_status]
else:
clf_label = [value]
unsup_label = dict(row[unsup_label_keys])
unsup_label['dataset-name'] = 'wiai-facility'
if exists(file):
user_files.append(file)
user_clf_labels.append(clf_label)
user_ids.append(user)
user_unsup_labels.append(unsup_label)
files.extend(user_files)
clf_labels.extend(user_clf_labels)
unsup_labels.extend(user_unsup_labels)
users.extend(user_ids)
# -
# len(starts), len(ends), \
len(files), len(users), len(clf_labels), len(unsup_labels)
df = pd.DataFrame({'file': files, 'classification': clf_labels, 'unsupervised': unsup_labels, 'users': users})
df.shape
df['id'] = df['file'].apply(get_unique_id)
df['audio_type'] = df['file'].apply(get_audio_type)
df.head()
df.tail()
# save the dataframe
annotation_save_path = join(save_root, 'annotation.csv')
df.to_csv(annotation_save_path, index=False)
# save the dataframe
annotation_save_path = join(save_root, 'attributes.csv')
data_sheet.to_csv(annotation_save_path, index=False)
# Check the total duration of the dataset
durations = []
for filename in tqdm(files, desc='Durations'):
filepath = join(save_audio_dir, filename + '.wav')
if exists(filepath):
duration = get_duration(filename=filepath)
durations.append(duration)
sum(durations)
# #### Junk code
from joblib import Parallel, delayed
df = {'file': [], 'classification': [], 'users': [], 'start': [], 'end': []}
def update_df_by_user_files(index):
row = data_sheet.loc[index]
recording_dir = row['audio_folder']
user_timestamp = '_'.join(recording_dir.split('/')[-2:])
user = user_timestamp.split('/')[0]
disease_status = row['testresult_covid_test_result']
user_files = []
user_labels = []
user_filesecs = []
user_ids = []
user_fstarts = []
for key, value in sound_labels.items():
file = '/'.join([save_audio_dir, '_'.join([user_timestamp, f'{key}.wav'])])
if key != 'room_sound':
label = [value, disease_status]
else:
label = [value]
if exists(file):
user_files.append(file)
user_labels.append(label)
user_filesecs.append(get_duration(filename=file))
user_fstarts.append(0)
user_ids.append(user)
df['file'].extend(user_files)
df['classification'].extend(user_labels)
df['end'].extend(user_filesecs)
df['users'].extend(user_ids)
df['start'].extend(user_fstarts)
iterator = tqdm(range(len(data_sheet)), desc="Iterating over all patients")
Parallel(n_jobs=10, require='sharedmem')(delayed(update_df_by_user_files)(index) for index in iterator);
df = pd.DataFrame(df)
# +
non_existent = []
exceptions = []
for file in tqdm(files):
if exists(file):
try:
signal, rate = librosa.load(file)
except Exception as ex:
exceptions.append((type(ex).__name__. str(ex), file))
else:
non_existent.append(file)
# -
non_existent
exceptions
# !ls /data/wiai-facility/processed/audio
files = listdir(save_audio_dir)
len(files)
files[0]
# +
invalid_files = []
for file in tqdm(files, desc='Checking valid files'):
fpath = f'/data/wiai-crowdsourced/processed/audio/{file}'
try:
S, R = librosa.load(fpath)
except:
invalid_files.append(file)
# -
len(invalid_files)
| datasets/cleaning/wiai-facility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wikimedia data
#
# - [Wikimedia Downloads: Analytics Datasets](https://dumps.wikimedia.org/other/analytics/)
# Info about Pageviews, mediacounts and unique devices:
# - [Pageviews since may 2015](https://dumps.wikimedia.org/other/pageviews/):
# ```
# https://dumps.wikimedia.org/other/pageviews/[YEAR]/[YEAR]-[2-DIGIT-MONTH]/pageviews-YYYYMMDD-HHMMSS.GZ
# ```
#
# - [Siteviews interactive analysis](https://tools.wmflabs.org/siteviews/?platform=all-access&source=pageviews&agent=all-agents&start=2015-07&end=2017-09&sites=all-projects)
#
# ## Running this notebook:
#
# Dependencies:
# - Bokeh
# - Pandas
#
# Enable widgetsnbextension:
# ```
# $ jupyter nbextension enable --py --sys-prefix widgetsnbextension
# ```
#
# !pip install -e $PWD/awscosts
# dataframe-related imports
import wikimedia_scraper as ws
from datetime import datetime
import pandas as pd
import numpy as np
from datetime import timedelta
# +
# plotting-related imports
import matplotlib.pylab as plt
# %matplotlib inline
from bokeh.io import push_notebook, show, output_notebook
from bokeh.plotting import figure
from bokeh.models import DatetimeTickFormatter, NumeralTickFormatter, BasicTickFormatter
from bokeh.models.tickers import FixedTicker
# -
# ## Get data source
# +
# use New Wikipedia scraper and store in a dataframe
project = 'za' # 0.00
project = 'el' # 0.22
project = 'he' # 0.37
project = 'uk' # 0.43
project = 'es' # 8.10
project = 'fr' # 6.34
project = 'en' # 14.62
start_date = datetime(2016, 11, 1)
end_date = datetime(2017, 10, 30)
ws.output_notebook()
traffic_generator = ws.get_traffic_generator(start_date, end_date, projects=(project,))
df = pd.DataFrame(list(traffic_generator))
df.head()
# +
# set date as index
df = df.set_index(pd.DatetimeIndex(df['date']))
df = df.drop(['date'], axis=1)
df = df.loc[df['project']==project]
df.head()
# +
# z-score (not really meaningful for this study)
#df["col_zscore"] = (df['hits'] - df['hits'].mean())/df['hits'].std(ddof=0)
# Filtering between dates example (not used for now)
#mask = (df.index >= '2017-05-22 15:00:00') & (df.index <= '2017-05-23 5:00:00')
#filtered_df = df.loc[mask]
# -
# ## Yearly hits data normalization & plotting
# +
# need to convert types to avoid a INF value while computing mean value (too big number?)
df['hits'] = df['hits'].astype(float)
# rolling mean
df['normalized_hits'] = df['hits'].astype(float)/df['hits'].astype(int).sum()
df['rolling'] = df['normalized_hits'].rolling(window=24*7, min_periods=3).mean()
df.head()
# +
# plotting yearly data
# using BOKEH
year_plot = figure(title="wikipedia visits per hour", x_axis_type="datetime")
year_plot.yaxis.formatter = BasicTickFormatter(use_scientific=False)
year_plot.xaxis.formatter = DatetimeTickFormatter(
hours = [ '%R' ],
days = [ '%d %b' ],
months = [ '%b' ],
years = [ '%a %H' ],
)
year_zscore_data = year_plot.line(df.index, df['normalized_hits'], color="#2222aa", line_width=1)
year_rolling_data = year_plot.line(df.index, df['rolling'], color="red", line_width=1)
#output_notebook()
#show(year_plot, notebook_handle=True)
#push_notebook()
# using matplotlib
plt.plot(df['normalized_hits'], color='blue', label='hits')
plt.plot(df['rolling'], color='red',label='Original')
# -
# ## The "_Average week_" calculation
# +
# Add new columns based on date index
df['weekday'] = df.index.weekday_name
df['hour'] = df.index.hour
startdate = datetime(1970,1,5)
days = ('Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday')
df3 = pd.DataFrame()
for day in days:
dfx1 = pd.DataFrame(index=pd.date_range(start=startdate, periods=24, freq='H'), columns=['hits'])
hitmeans = df.loc[df['weekday']==day].groupby('hour')['hits'].mean()
dfx1['hits'] = np.array(hitmeans)
startdate += timedelta(days=1)
df3 = pd.concat([df3,dfx1])
df3['normalized_hits'] = df3['hits'].astype(float)/df3['hits'].astype(int).sum()
df3['rolling'] = df3['normalized_hits'].rolling(window=24, min_periods=3).mean()
# +
# plotting avg week
avg_week_plot = figure(title="wikipedia average week (normalized)", x_axis_type="datetime")
avg_week_plot.yaxis.formatter = BasicTickFormatter(use_scientific=False)
avg_week_plot.xaxis.formatter = DatetimeTickFormatter(
hours = [ '%R' ],
days = [ '%a' ],
months = [ '%a %H' ],
)
avg_week_plot.yaxis[0].formatter = NumeralTickFormatter(format='0.000a')
avg_week_data = avg_week_plot.line(df3.index, df3['normalized_hits'], color="#2222aa", line_width=1)
avg_week_rolling_data = avg_week_plot.line(df3.index, df3['rolling'], color="red", line_width=1)
#output_notebook()
#show(avg_week_plot, notebook_handle=True)
#push_notebook()
plt.plot(df3['normalized_hits'], color='blue', label='hits')
plt.plot(df3['rolling'], color='red',label='Original')
# -
# ## Construct a synthetic year
# FIXME: datetime.now() has to be replaced with the last monday at 0:00
dfy = pd.DataFrame(index=pd.date_range(start=datetime.now(), periods=52*7*24, freq='H'), columns=['normalized_hits'])
dfy['normalized_hits'] = list(df3['normalized_hits']) * 52
# +
# plotting yearly data
synthetic_year_plot = figure(title="wikipedia visits per hour", x_axis_type="datetime")
synthetic_year_plot.yaxis.formatter = BasicTickFormatter(use_scientific=False)
synthetic_year_plot.xaxis.formatter = DatetimeTickFormatter(
hours = [ '%R' ],
days = [ '%d %b' ],
months = [ '%b' ],
years = [ '%a %H' ],
)
year_zscore_data = synthetic_year_plot.line(dfy.index, dfy['normalized_hits'], color="#2222aa", line_width=1)
#year_rolling_data = year_plot.line(df.index, dfy['rolling'], color="red", line_width=1)
#output_notebook()
#show(synthetic_year_plot, notebook_handle=True)
#push_notebook()
plt.plot(dfy['normalized_hits'], color='blue', label='hits')
# -
# ## Growth function
# ### No growth
dfy['growth_factor'] = (1,) * len(dfy)
dfy.head()
# To be requested from the user
scale_factor = df3['hits'].astype(int).sum()
dfy['hits'] = dfy['normalized_hits'] * dfy['growth_factor'] * scale_factor
# ### Linear growth (10% over len(dfy))
# +
dfy['growth_factor'] = (np.nan,) * len(dfy)
dfy['growth_factor'][ 0] = 1
dfy['growth_factor'][-1] = 1.10
dfy.interpolate(inplace=True, method='linear')
plt.plot(dfy['growth_factor'])
# -
dfy['hits'] = dfy['normalized_hits'] * dfy['growth_factor'] * scale_factor
# +
# plotting yearly data
synthetic_year_plot = figure(title="wikipedia visits per hour", x_axis_type="datetime")
synthetic_year_plot.yaxis.formatter = BasicTickFormatter(use_scientific=False)
synthetic_year_plot.xaxis.formatter = DatetimeTickFormatter(
hours = [ '%R' ],
days = [ '%d %b' ],
months = [ '%b' ],
years = [ '%a %H' ],
)
year_zscore_data = synthetic_year_plot.line(dfy.index, dfy['hits'], color="#2222aa", line_width=1)
#year_rolling_data = year_plot.line(df.index, dfy['rolling'], color="red", line_width=1)
#output_notebook()
#show(synthetic_year_plot, notebook_handle=True)
#push_notebook()
plt.plot(dfy['hits'], color='#2222aa', label='hits')
# -
# ### Linear growth
# +
dfy['x'] = (np.nan,) * len(dfy)
dfy['x'][ 0] = 0
dfy['x'][-1] = 1
dfy=dfy.drop(['growth_factor'], axis=1)
dfy.interpolate(inplace=True, method='linear')
# -
# ### Exponential growth (275% over len(dfy))
# +
from math import log
dfy['growth_factor'] = np.exp(log(2)*dfy['x'])
# -
plt.plot(dfy['growth_factor'])
dfy['hits'] = dfy['normalized_hits'] * dfy['growth_factor'] * scale_factor
# +
# plotting yearly data
synthetic_year_plot = figure(title="wikipedia visits per hour", x_axis_type="datetime")
synthetic_year_plot.yaxis.formatter = BasicTickFormatter(use_scientific=False)
synthetic_year_plot.xaxis.formatter = DatetimeTickFormatter(
hours = [ '%R' ],
days = [ '%d %b' ],
months = [ '%b' ],
years = [ '%a %H' ],
)
year_zscore_data = synthetic_year_plot.line(dfy.index, dfy['hits'], color="#2222aa", line_width=1)
#year_rolling_data = year_plot.line(df.index, dfy['rolling'], color="red", line_width=1)
#output_notebook()
#show(synthetic_year_plot, notebook_handle=True)
#push_notebook()
plt.plot(dfy['hits'])
# +
dfy['date'] = dfy.index
import awscosts
MB_per_request = 128
ms_per_req=200
max_reqs_per_second = 1000
mylambda = awscosts.Lambda(MB_per_req=MB_per_request, ms_per_req=ms_per_req)
#myec2 = awscosts.EC2(instance_type='m4.4xlarge', MB_per_req=MB_per_request, ms_per_req=ms_per_req)
myec2 = awscosts.EC2(instance_type='m4.4xlarge', max_reqs_per_second=max_reqs_per_second)
dfy['lambda_cost'] = dfy.apply(lambda x: mylambda.get_cost(date = x['date'], reqs = x['hits']), axis=1)
dfy['ec2_cost'] = dfy.apply(lambda x: myec2.get_cost_and_num_instances(3600, reqs = x['hits'])[0], axis=1)
#dfy['ec2_cost2'] = dfy.apply(lambda x: myec2.get_cost_per_hour(reqs = x['hits']), axis=1)
dfy['instances'] = dfy.apply(lambda x: myec2.get_num_instances(reqs = x['hits']/3600), axis=1)
dfy.head()
# +
d2=pd.DataFrame(dfy.groupby(dfy.index.month)['lambda_cost'].sum())
d2['ec2_cost']=pd.DataFrame(dfy.groupby(dfy.index.month)['ec2_cost'].sum())
d2['delta']=d2['lambda_cost']-d2['ec2_cost']
d2['ratio λ/ec2']=d2['lambda_cost']/d2['ec2_cost']
d2['ratio λ/ec2'].mean()
# -
d2=d2.style.format({'lambda_cost': "$ {:.2f}", 'ec2_cost': '$ {:.2f}', 'ratio λ/ec2':'{:.2f}'})
d2
| web_traffic_growth_simulator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Buffered Text-to-Speech
#
# In this tutorial, we are going to build a state machine that controls a text-to-speech synthesis. The problem we solve is the following:
#
# - Speaking the text takes time, depending on how long the text is that the computer should speak.
# - Commands for speaking can arrive at any time, and we would like our state machine to process one of them at a time. So, even if we send three messages to it shortly after each other, it processes them one after the other.
#
# While solving this problem, we can learn more about the following concepts in STMPY state machines:
#
# - **Do-Activities**, which allow us to encapsulate the long-running text-to-speech function in a state machine.
# - **Deferred Events**, which allow us to ignore incoming messages until a later state, when we are ready again.
# # Text-to-Speech
# ## Mac
#
# On a Mac, this is a function to make your computer speak:
# +
from os import system
def text_to_speech(text):
system('say {}'.format(text))
# -
# Run the above cell so the function is available in the following, and then execute the following cell to test it:
text_to_speech("Hello. I am a computer.")
# ## Windows
#
# TODO: We should have some code to run text to speech on Windows, too!
# # State Machine 1
#
# With this function, we can create our first state machine that accepts a message and then speaks out some text. (Let's for now ignore how we get the text into the method, we will do that later.)
#
# 
#
# Unfortunately, this state machine has a problem. This is because the method `text_to_speech(text)` is taking a long time to complete. This means, for the entire time that it takes to speak the text, nothing else can happen in all the state machines that are part of the same driver!
# # State Machine 2
#
# ## Long-Running Actions
#
# The way this function is implented makes that it **blocks**. This means, the Python program is busy executing this function as long as the speech takes to pronouce the message. Longer message, longer blocking.
# You can test this by putting some debugging aroud the function, to see when the functions returns:
print('Before speaking.')
text_to_speech("Hello. I am a computer.")
print('After speaking.')
# You see that the string _"After speaking"_ is printed after the speaking is finished. During the execution, the program is blocked and does not do anything else.
#
# When our program should also do other stuff at the same time, either completely unrelated to speech or even just accepting new speech commands, this is not working! The driver is now completely blocked with executing the speech method, not being able to do anything else.
# ## Do-Activities
#
# Instead of executing the method as part of a transition, we execute it as part of a state. This is called a **Do-Activity**, and it is declared as part of a state. The do-activity is started when the state is entered. Once the activity is finished, the state machine receives the event `done`, which triggers it to switch into another state.
#
# 
#
# You may think now that the do-activity is similar to an entry action, as it is started when entering a state. However, a do-activity is started as part of its own thread, so that it does not block any other behavior from happening. Our state machine stays responsive, and so does any of the other state machines that may be assigned to the same driver. This happens in the background, STMPY is creating a new thread for a do-activity, starts it, and dispatches the `done` event once the do-activity finishes.
#
# When the do-activity finishes (in the case of the text-to-speech function, this means when the computer is finished talking), the state machine dispatches _automatically_ the event `done`, which brings the state machine into the next state.
#
# - A state with a do activity can therefore only declare one single outgoing transition that is triggered by the event `done`.
# - A state can have at most one do-activity.
# - A do-activity cannot be aborted. Instead, it should be programmed so that the function itself terminates, indicated for instance by the change of a variable.
#
# The following things are still possible in a state with a do-activity:
#
# - A state with a do-activity can have entry and exit actions. They are simply executed before or after the do activities.
# - A state with a do-activity can have internal transition, since they don't leave the state.
# +
from stmpy import Machine, Driver
from os import system
import logging
debug_level = logging.DEBUG
logger = logging.getLogger('stmpy')
logger.setLevel(debug_level)
ch = logging.StreamHandler()
ch.setLevel(debug_level)
formatter = logging.Formatter('%(asctime)s - %(name)-12s - %(levelname)-8s - %(message)s')
ch.setFormatter(formatter)
logger.addHandler(ch)
class Speaker:
def speak(self, string):
system('say {}'.format(string))
speaker = Speaker()
t0 = {'source': 'initial', 'target': 'ready'}
t1 = {'trigger': 'speak', 'source': 'ready', 'target': 'speaking'}
t2 = {'trigger': 'done', 'source': 'speaking', 'target': 'ready'}
s1 = {'name': 'speaking', 'do': 'speak(*)'}
stm = Machine(name='stm', transitions=[t0, t1, t2], states=[s1], obj=speaker)
speaker.stm = stm
driver = Driver()
driver.add_machine(stm)
driver.start()
driver.send('speak', 'stm', args=['My first sentence.'])
driver.send('speak', 'stm', args=['My second sentence.'])
driver.send('speak', 'stm', args=['My third sentence.'])
driver.send('speak', 'stm', args=['My fourth sentence.'])
driver.wait_until_finished()
# -
# The state machine 2 still has a problem, but this time another one: If we receive a new message with more text to speak _while_ we are in state `speaking`, this message is discarded. Our next state machine will fix this.
# # State Machine 3
#
# As you know, events arriving in a state that do not declare outgoing triggers with that event, are discarded (that means, thrown away). For our state machine 2 above this means that when we are in state `speaking` and a new message arrives, this message is discarded. However, what we ideally want is that this message is handled once the currently spoken text is finished. There are two ways of achieving this:
#
# 1. We could build a queue variable into our logic, and declare a transition that puts any arriving `speak` message into that queue. Whenever the currently spoken text finishes, we take another one from the queue until the queue is empty again. This has the drawback that we need to code the queue ourselves.
# 2. We use a mechanism called **deferred event**, which is part of the state machine mechanics. This is the one we are going to use below.
#
# ## Deferred Events
#
# A state can declare that it wants to **defer** an event, which simply means to not handle it. For our speech state machine it means that state `speaking` can declare that it defers event `speak`.
#
# 
#
# Any event that arrives in a state that defers it, is ignored by that state. It is as if it never arrived, or as if it is invisible in the incoming event queue. Only once we switch into a next state that does not defer it, it gets visible again, and then either consumed by a transition, or discarded if the state does not declare any transition triggered by it.
#
# +
s1 = {'name': 'speaking', 'do': 'speak(*)', 'speak': 'defer'}
stm = Machine(name='stm', transitions=[t0, t1, t2], states=[s1], obj=speaker)
speaker.stm = stm
driver = Driver()
driver.add_machine(stm)
driver.start()
driver.send('speak', 'stm', args=['My first sentence.'])
driver.send('speak', 'stm', args=['My second sentence.'])
driver.send('speak', 'stm', args=['My third sentence.'])
driver.send('speak', 'stm', args=['My fourth sentence.'])
driver.wait_until_finished()
# -
| Buffered Text-to-Speech.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring the UTx000 Extension Beacon Data
# (See the [GH repo](https://github.com/intelligent-environments-lab/utx000))
import warnings
warnings.filterwarnings('ignore')
# # Estimating Ventilation Rates
# Using CO$_2$, we can estimate the ventilation rates in the space using various methodologies.
# ## Package Import
# +
import os
import sys
sys.path.append('../')
from src.features import build_features
from src.visualization import visualize
import pandas as pd
pd.set_option('display.max_columns', 200)
import numpy as np
from datetime import datetime, timedelta
import math
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib.dates as mdates
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
# -
# # Data Import
#
# ## Beacon Data
# We are primarily concerned with the beacon data when participants are asleep since that is the only time we can be sure participants are home in their bedrooms. However, we also need all the processed beacon data for later.
processed_beacon = pd.read_csv('../data/processed/beacon-ux_s20.csv',
index_col="timestamp",parse_dates=True,infer_datetime_format=True)
processed_beacon.head()
beacon = pd.read_csv('../data/processed/beacon_by_night-ux_s20.csv',
index_col="timestamp",parse_dates=["timestamp","start_time","end_time"],infer_datetime_format=True)
beacon.head()
# ## Participant Information
# We need the participant beacon, beiwe, and fitbit IDs so we can cross-reference the various pieces of information.
# getting pt names
names = pd.read_excel('../data/raw/utx000/admin/id_crossover.xlsx',sheet_name='all')
names = names[['beiwe','first','last','sex']]
# getting beacon only pts with names
info = pd.read_excel('../data/raw/utx000/admin/id_crossover.xlsx',sheet_name='beacon')
info = info[['redcap','beiwe','beacon','lat','long','volume','roommates']] # keep their address locations
info = info.merge(left_on='beiwe',right=names,right_on='beiwe')
info.head()
# ## Fitbit Data
# We will need some information from each of the participants that Fitbit logs like body weight.
daily = pd.read_csv('../data/processed/fitbit-daily-ux_s20.csv',
index_col=0,parse_dates=True,infer_datetime_format=True)
daily.head()
# # Ventilation based on constant CO2 concentration
# We can estimate the ventilation rate of the participants' bedrooms based on a constant CO2 concentration:
#
# $$
# \lambda = \frac{E}{V \left(C - pC_0\right)}
# $$
#
# In our case we have to make quite a few assumptions:
# - $E$: Emission rate of CO2 can be estimated for humans to be approximately $30 g/hr$. We can further refine this value by considering the height, weight, sex, and activity level of the participant - all information that we have access to.
# - $V$: Arguably the most difficult value to track down and the one that could affect the answer the most. We can assume a volume of each of the participants bedrooms, but there is the possiblility of refining this value by cross-referencing the participants' addresses with any known apartment complexes. From there, we can estimate the floor plan based on the number of roommates.
# - $p$: Penetration factor of CO2 from outside into the indoor space - we can safely assume this to be 1.
# - $C_0$: Outdoor concentration of CO2 - another safe assumption of 400 ppm.
# ## Emission Rate
# Based on an article that Sangeetha sent me, we can determine the emission rate of CO2 from the following equation:
#
# $$
# V_{CO_2} = (BMR)(M)(T/P)(0.000179)
# $$
#
# where:
# - $V_{CO_2}$ is the volumetric emission rate of CO2 in L/s
# - $BMR$ is the body mass ratio determined from Persily and De Jonge 2016
# - $M$: Activity level in METs
# - $T$ is the temperature in Kelvin
# - $P$ is the pressure in kPa
# ### Body Mass Ratio
# The body mass ratio is given by one of the two equations since our participants are in the age range of 18-30
#
# $$
# BMR_{male} = 0.063m + 2.896\\
# BMR_{female} = 0.062m + 2.036
# $$
#
# where $m$ is the body mass in kilograms. We have the pariticpants body weights assuming they logged them into Fitbit. We can grab these data and put it alongside the participants beacon, beiwe, and fitbit IDs.
#
# We will take the body weight for each participant, average it over the total amount of time logged (just in case they changed it which I doubt anyone did) and store that value in our ```ventilation_info``` dataframe.
# +
weight_dict = {'beiwe':[],'mass':[]}
for pt in daily['beiwe'].unique():
daily_pt = daily[daily['beiwe'] == pt]
weight_dict['beiwe'].append(pt)
weight_dict['mass'].append(np.nanmean(daily_pt['weight'])*0.453592) # convert to kg
mass_info = info.merge(left_on='beiwe',right=pd.DataFrame(weight_dict),right_on='beiwe')
mass_info.sort_values("beacon",ascending=True)
# -
# In order to properly calculate the BMR of the participants, we need to know whether they are male or female. Participants had to mark whether or not they were male or female on the CRISIS baseline survey we sent via REDCap.
# getting sex from baseline survey
#crisis_baseline = pd.read_csv('../data/raw/utx000/surveys/CRISIS_E3_labels.csv',
#usecols=[0,3],names=['redcap','sex'],header=0)
#sex_info = mass_info.merge(left_on='redcap',right=crisis_baseline,right_on='redcap',how='outer')
#sex_info.dropna(subset=['beacon'],axis=0,inplace=True)
# adding sex manually
#sex_info.iloc[3,-1] = 'Male' # <NAME>
#sex_info.iloc[4,-1] = 'Female' # <NAME>
#sex_info.iloc[8,-1] = 'Female' # <NAME>
#sex_info.to_csv("~/Desktop/info.csv")
# <div class="alert alert-block alert-success">
#
# A `sex` column has beend added to the `id_crossover` file so the above cell and import of the CRISIS survey is no longer needed.
#
# </div>
# Now we calculate the BMR based on the mass and sex of the participants
def get_BMR(sex, mass):
'''
Calculates the BMR based on mass assuming an age range between 18 and 30
Inputs:
- sex: string in ['Male','Female']
- mass: double of the mass in kg
Returns BMR from Persily and De Jong 2016
'''
if sex.lower() == 'male':
return 0.063*mass + 2.896
elif sex.lower() == 'female':
return 0.062*mass + 2.036
else:
return 0
mass_info['bmr'] = mass_info.apply(lambda row: get_BMR(row['sex'],row['mass']),axis=1)
pt_info = mass_info.set_index('beiwe')
pt_info.head()
# ### Activity Level in METs
# The Persily and De Jong 2016 article gives values for activity level in METs dependingon the activity. We can assume that our participants are asleep since we are only using data when Fitbit detects sleeping individuals. The MET value for a sleeping individual, from the article, is **0.95**.
# ### Temperature
# The **temperature will vary depending on the night** that we choose to analyze and therefore will alter the emission rate of CO2. Thus we cannot establish a static emission rate.
# ### Pressure
# The pressure will more or less remain constant since we are at the same altitude for the most part in Austin. The National Weather Service has a [station at the airport](https://w1.weather.gov/data/obhistory/KAUS.html) in Austin that reports the atmopsheric pressure is around **102.5 kPa**.
def get_emission_rate(BMR, T):
'''
Calculates the CO2 emission rate
Inputs:
- BMR: double of body-mass-ratio
- T: double of Temperature in Kelvin
Returns CO2 emission rate in L/s
'''
# Assumed constants
M = 0.95 #METs
P = 102.5 #kPa
return BMR * M * (T / P) * 0.000179
# ## Volume
# We can always assume a value for the volume of the space, or we can go a bit more in-depth by looking at the addresses of the participants.
#
# Doing this presents their own set of problems since we now have to trust the answers of the participants and the addresses could be different than the addresess we sent the beacons to. Rather, we can simply use average numbers:
# - **Stand-Alone Home**: We can use the US average of 11x12 feet with 9 foot ceilings - **1188 feet**
# - **Apartment**: Another average value would be 10x12 with 9 foot ceilings - **1080 feet**
# <div class="alert-block alert alert-success">
# The volume values have already been added to the info.csv file for each participant based on the address used to ship the beacons to, <b>not</b> the response in the EE survey.
# <div>
# ## Outdoor CO2
# We can assume the outdoor CO2 concentration is 400 ppm, but we need to convert this value into g/m$^3$.
def convert_ppm_to_gm3(concentration, mm=44.0, mv=24.5):
'''
Converts the ppm of a gas to g/m3
Inputs:
- concentration: double specifying the concentration in ppm
- mm: double specifying the molar mass of the compound (default is CO2)
- mv: double of molar volume (default is at room temperature)
Returns concentration in g/m3
'''
return concentration / 10**6 * mm / mv * 1000
# **Testing**
#
# Compare to calculator [here](https://www.teesing.com/en/page/library/tools/ppm-mg3-converter)
convert_ppm_to_gm3(400)
# ## Constant CO2
# Now we need to find periods during the evening when the CO2 concentration is constant. We can do this by looking at the mean difference in CO2 concentration during a certain window. If the change is less than a certain threshold, we can then assume the concentration is stable.
#
# We can also cross-reference with the temperature to make sure that the AC isn't just cycling on and off quickly keeping the CO2 within a tight window.
# ### Processing Temperature
# We want to make sure to use the temperature sensors from the DGS sensors rather than the CO2 sensor. There might be times when one of the temperature sensors is working or they are both operating. In the latter case, we want to average the values and then combine the temperature values into one column.
#
# We will also need the temperature to better estimate the emission rate.
beacon_co2 = beacon[['redcap','beiwe','beacon','co2','temperature_c','rh','start_time','end_time']]
beacon_co2.head()
# ### CO2 Trend Periods
# The function below gets the periods of a certain trends for CO2: increasing, decreasing, or constant (within a threshold).
def get_co2_periods(df, window=12, co2_threshold=10, t_threshold=0.25, time_threshold=300, change='decrease'):
'''
Finds and keeps periods of CO2 change or consistency
Inputs:
- df: Dataframe holding the measured CO2 concentrations at 5-minute timestamps
- window: integer specifying how many timesteps the increase/decrease has to last
Returns dataframe with only increasing/decreasing periods greater in length than the window time periods
'''
# getting differences
df['change'] = df['co2'] - df['co2'].shift(1)
df['change'] = df['change'].shift(-1)
df['t_change'] = df['temperature_c'] - df['temperature_c'].shift(1)
df['t_change'] = df['t_change'].shift(-1)
df["time"] = df.index
df['dtime'] = df["time"] - df["time"].shift(1)
df['dtime'] = df['dtime'].shift(-1)
# find periods of increase/decrease and giving them unique labels
i = 0
periods = []
period = 1
if change == 'decrease':
while i < len(df):
while df['change'][i] < 0 and df['t_change'][i] <= 0 and df['dtime'][i].total_seconds() <= time_threshold:
periods.append(period)
i += 1
periods.append(0)
period += 1
i += 1
elif change == 'increase':
while i < len(df):
while df['change'][i] > 0 and abs(df['t_change'][i]) <= t_threshold and df['dtime'][i].total_seconds() <= time_threshold:
periods.append(period)
i += 1
periods.append(0)
period += 1
i += 1
else: # constant periods
while i < len(df):
while abs(df['change'][i]) < co2_threshold and df['t_change'][i] <= 0 and df['dtime'][i].total_seconds() <= time_threshold:
periods.append(period)
i += 1
periods.append(0)
period += 1
i += 1
# removing periods shorter than the window
df['period'] = periods
df = df[df['period'] > 0]
for period in df['period'].unique():
temp = df[df['period'] == period]
if len(temp) < window:
df = df[df['period'] != period]
return df
# ## Air Exchange Rate
# Now we have all the information we need for the equation, we can determine the air exchange rate in each of the participants' homes.
#
# We do need to find some extra information to convert our units:
# - **Density of CO2 ($\rho_{CO_2}$)**: The density of CO2 can be found [here](www.engineeringtoolbox.com/carbon-dioxide-density-specific-weight-temperature-pressure-d_2018.html). We want to look for nearly atmospheric pressure and room temperature.
# - **Conversions**: Like cubic feet to cubic meters, ppm to g/m$^3$, and the like.
def get_ach_from_constant_co2(E, V, C, C0=400.0, p=1.0):
'''
Calculates the air exchange rate for constant CO2 events
Inputs:
- E: double of emission rate in L/s
- V: double of volume in ft3
- C: double of room co2 concetration in ppm
- C0: double of outdoor co2 concentration in ppm (default is 400)
- p: double of penetration factor (default is 1)
Returns ach in 1/h
'''
# defining constants
rho = 1.8 # g/L
# converting units
E_gs = E * rho # L/s to g/s
V_m3 = V * 0.0283168 # ft3 to m3
C_gm3 = convert_ppm_to_gm3(C) # ppm to g/m3
C0_gm3 = convert_ppm_to_gm3(C0) # ppm to g/m3
return E_gs / (V_m3 * (C_gm3 - p*C0_gm3)) * 3600
# ## Calculating emission rates and ventilation rates
# Putting everything together now we loop through each night for each participant, calculate the emission rate, and then determine the ventilation rate.
def plot_constant_co2_period(df, ach, pt="", save=False):
"""plots the relevant variables from a constant co2 period"""
fig, ax = plt.subplots(figsize=(8,6))
ax.plot(df.index,df["co2"],color="seagreen")
ax.set_ylim([400,2000])
ax.set_ylabel("CO$_2$ (ppm)",fontsize=16)
plt.yticks(fontsize=14)
plt.xticks(fontsize=14,ha="left",rotation=-45)
ax2 = ax.twinx()
ax2.plot(df.index,df["temperature_c"],color="cornflowerblue")
ax2.spines['right'].set_color('cornflowerblue')
ax2.set_ylim([20,30])
plt.yticks(fontsize=14)
ax2.set_ylabel("Temperature ($^\circ$C)",fontsize=16,color="cornflowerblue")
ax2.xaxis.set_major_locator(mdates.MinuteLocator(interval=5))
ax2.xaxis.set_major_formatter(mdates.DateFormatter("%H:%M"))
plt.xticks(fontsize=14,ha="left",rotation=-45)
period = df['period'][0]
ax.set_title(f"ID: {pt} - Period: {period} - ACH: {round(ach,2)}")
if save:
plt.savefig(f"../reports/figures/beacon_summary/ventilation_estimates/method_0-{pt}-{period}.pdf",bbox_inches="tight")
plt.show()
plt.close()
ventilation_df = pd.DataFrame()
for pt in beacon_co2['beiwe'].unique(): # cycling through each of the participants
# setting up the dictionary to add pt values to
pt_dict = {'beiwe':[],'beacon':[],'start':[],'end':[],'co2_mean':[],'co2_delta':[],'temperature_c_mean':[],'temperature_c_delta':[],'e':[],'ach':[]}
# pt-specific dataframes
beacon_pt = beacon_co2[beacon_co2['beiwe'] == pt]
info_pt = pt_info[pt_info.index == pt]
for start, end in zip(beacon_pt['start_time'].unique(),beacon_pt['end_time'].unique()): # looping through sleep events
beacon_pt_night = beacon_pt[start:end] # masking for iaq data during sleep
if len(beacon_pt_night) > 36: # looking for nights when we have at least 3 hours (beacon data is at 5-min)
constant_periods = get_co2_periods(beacon_pt_night[['co2','temperature_c','rh']],window=12,change='constant')
n = len(constant_periods)
if n > 0:
for period in constant_periods['period'].unique():
constant_by_period = constant_periods[constant_periods['period'] == period]
C = np.nanmean(constant_by_period['co2'])
if C > 600:
dC = np.nanmean(constant_by_period['change'])
T = np.nanmean(constant_by_period['temperature_c'])
dT = np.nanmean(constant_by_period['t_change'])
pt_dict['beiwe'].append(pt)
pt_dict['beacon'].append(info_pt['beacon'].values[0])
pt_dict['start'].append(start)
pt_dict['end'].append(end)
pt_dict['co2_mean'].append(C)
pt_dict['co2_delta'].append(dC)
pt_dict['temperature_c_mean'].append(T)
pt_dict['temperature_c_delta'].append(dT)
E = get_emission_rate(pt_info.loc[pt,'bmr'],T+273)
pt_dict['e'].append(E)
V = info_pt['volume'].values[0]
ACH = get_ach_from_constant_co2(E,V,C)
pt_dict['ach'].append(ACH)
#plot_constant_co2_period(constant_by_period,ACH,pt,save=True)
ventilation_df = ventilation_df.append(pd.DataFrame(pt_dict))
ventilation_df = ventilation_df.groupby(["start","end","beiwe"]).mean().reset_index()
# # Volume based on CO2 growth
# While not a good estimate, we can use periods of uniterrupted CO2 concentration increases to estimate what the volume of the space might be. Under these conditions, we might assume the the contribution from outdoors is negligible and we simply have accumulation:
#
# $$
# V \frac{dC}{dt} = E
# $$
# ## Estimating volume
# We can rearrange the original equation to solve for the volume. We have to use our previously defined function to estimate the emission rate from a person though.
def get_volume(df,E,n_people=1):
'''
Estimates the volume based on the CO2 emission rate and negligible infiltration/exfiltration
Inputs:
- df: Dataframe indexed by time holding the CO2 and temperature values over an increasing period of CO2
- E: double of emission rate of CO2 in L/s
- n_people: integer specifying the number of CO2 emitters - default is 1 person
Returns the volume in m3 and ft3
'''
# defining constants
rho = 1.8 # g/L
# converting units
E_gs = E * rho # L/s to g/s
df['c'] = convert_ppm_to_gm3(df['co2']) # ppm to g/m3
# Calculating
V_si = (E_gs * (df.index[-1] - df.index[0]).total_seconds()) / (df['c'][-1] - df['c'][0])
V_ip = V_si / 0.0283168
return V_si, V_ip
# ## Calculating volumes for individuals
# +
v_df = pd.DataFrame()
for pt in beacon_co2['beiwe'].unique():
v_dict = {'beiwe':[],'beacon':[],'start':[],'end':[],'starting_co2':[],'co2_delta':[],'ending_co2':[],'R^2':[],'volume_est':[],'volume_gen':[]}
# getting pt-specific data
beacon_co2_pt = beacon_co2[beacon_co2['beiwe'] == pt]
info_pt = pt_info[pt_info.index == pt]
# getting co2 periods of increase
increasing_co2 = get_co2_periods(beacon_co2_pt,window=8,change='increase')
for period in increasing_co2['period'].unique():
increasing_period_pt = increasing_co2[increasing_co2['period'] == period]
T = np.nanmean(increasing_period_pt['temperature_c'])
E = get_emission_rate(pt_info.loc[pt,'bmr'],T+273)
V_gen = info_pt['volume'].values[0]
V_est = get_volume(increasing_period_pt,E)[1]
# Checking linearity
Y = increasing_period_pt['co2']
X = np.arange(0,len(increasing_period_pt)*5,5)
X = sm.add_constant(X)
model = sm.OLS(Y,X)
results = model.fit()
# adding information to dict
for key, value_to_add in zip(v_dict.keys(),[pt,info_pt['beacon'].values[0],
increasing_period_pt.index[0],increasing_period_pt.index[-1],
increasing_period_pt['co2'][0],
increasing_period_pt['co2'][-1]-increasing_period_pt['co2'][0],
increasing_period_pt['co2'][-1],
results.rsquared,V_est,V_gen]):
v_dict[key].append(value_to_add)
v_df = v_df.append(pd.DataFrame(v_dict))
# Removing bad values
v_df = v_df[v_df['R^2'] > 0.99]
v_df = v_df[v_df['volume_est'] < 10000]
# Averaging
v_df_averaged = v_df.groupby('beiwe').mean()
# -
v_df_averaged
# # Ventilation based on CO2 decay
# We can estimate the ventilation rate of the participants' bedrooms based on a decay in the CO2 concentration:
#
# $$
# C_t = C_{t=0}e^{-\lambda t} + \left(pC_0 + \frac{E}{\lambda V}\right)\left(1 - e^{-\lambda t}\right)
# $$
#
# The benefit of this approach is that the exchange rate we get might be agnostic to the accuracy of the sensor assuming the sensor is precise (just off from the true value by a constant difference).
#
# Many of the variables from the previous analysis show up here again which is nice since we already found those.
# - $C_t$: CO2 concentation at some time, $t$
# - $C_{t=0}$: CO2 concentration at the beginning of the analysis period
# - $E$: Emission rate of CO2 estimated from the Persily and Jonge (2016) paper which looks at the activity, sex, and BMR of the participant.
# - $V$: We opted for a more generic value rather than diving too deep.
# - $\lambda$: air exchange rate
# - $p$: Penetration factor of CO2 from outside into the indoor space - we can safely assume this to be 1.
# - $C_0$: Outdoor concentration of CO2 - another safe assumption of 400 ppm.
#
# There are two challenges with this approach:
# 1. What data do we need and from what time?
# 1. How do we identify periods of CO2 decay?
# 2. The solution involves an iterative approach since we cannot easily solve for the air exchange rate, $\lambda$
#
# ## Occupant still in the space
# If we do not assume that the participant left the space, we can directly apply the above equation. We had to go through a similar process we did when estimating the air exchange rate based on a constant CO2 concentration i.e. we need the emission rate and the volume of the space.
#
# ## Occupant leaves
# If the participant leaves the bedroom, we don't have any emission sources. Therefore, our equation simplifies to:
#
# $$
# C_t = C_{t=0}e^{-\lambda t} + pC_0\left(1 - e^{-\lambda t}\right)
# $$
#
# ## Occupant leaves and no outdoor penetration
# By not considering any outdoor penetration, we can further simplify our equation:
#
# $$
# C_t = C_{t=0}e^{-\lambda t}
# $$
# ## New Beacon Data - Wake up periods
# We need to grab a different set of beacon data to do this assessment. Instead of looking at CO2 data during the evening, we actually need the CO2 data for a few hours right AFTER the participants woke up. We should be able to easily do this by looking at the stop times from the filtered beacon data and parsing out timestamps for the few hours after this from the processed beacon data.
def get_morning_beacon_data(night_df,all_df,num_hours=3):
'''
Grabs beacon data for hours after the participant has woken up.
Inputs:
- night_df: dataframe holding the nightly measured beacon values
- all_df: dataframe holding all the beacon measured values
- num_hours: number of hours after waking up to consider - default is 3
Returns a dataframe of beacon values for the "morning"
'''
df = pd.DataFrame()
for pt in night_df['beiwe'].unique():
# pt-specific data
night_pt = night_df[night_df['beiwe'] == pt]
beacon_pt = all_df[all_df['beiwe'] == pt]
for wake_time in night_pt['end_time'].unique():
temp = beacon_pt[wake_time:pd.to_datetime(wake_time)+timedelta(hours=num_hours)]
temp['start_time'] = wake_time
df = df.append(temp)
return df
# Now we use the functioned we defined above, get combined T/RH columns, and remove unecessary columns.
morning_beacon = get_morning_beacon_data(beacon,processed_beacon)
morning_beacon_co2 = morning_beacon[['redcap','beiwe','beacon','co2','temperature_c','rh','start_time']]
morning_beacon_co2.head()
# ## Identifying periods of decay
# To determine periods of decay we can recycle a lot of the work we did for the growth periods, but just look at the reverse situation.
#
# The code is actually so similar, that all is required is a switch of the inequality. Therefore, we just need to call the function ```get_co2_periods()``` with ```change = 'decreasing'```.
decay_co2 = get_co2_periods(morning_beacon_co2, window=6, change='decrease')
decay_co2.head()
# ### Further refining the periods of decay
# We are looking for the air exchange rate, so we need to make sure the AC is running during these decay periods. We can do so by looking at the temperature. Therefore we can further restrict our dataframe.
def get_change_T_periods(df, window=6,change='decrease'):
'''
Finds and keeps periods of T change
Inputs:
- df: Dataframe holding the measured T concentrations at 5-minute timestamps
- window: integer specifying how many timesteps the increase/decrease has to last
Returns dataframe with only increasing/decreasing periods greater in length than the window time periods
'''
# getting difference
df['temperature_change'] = df['temperature_c'] - df['temperature_c'].shift(1)
df['temperature_change'] = df['temperature_change'].shift(-1)
# find periods of increase/decrease and giving them unique labels
i = 0
periods = []
period = 1
if change == 'decrease':
while i < len(df):
while df['temperature_change'][i] <= 0:
periods.append(period)
i += 1
periods.append(0)
period += 1
i += 1
else: #increase
while i < len(df):
while df['temperature_change'][i] >= 0:
periods.append(period)
i += 1
periods.append(0)
period += 1
i += 1
# removing periods shorter than the window
df['temperature_period'] = periods
df = df[df['temperature_period'] > 0]
for period in df['temperature_period'].unique():
temp = df[df['temperature_period'] == period]
if len(temp) < window:
df = df[df['temperature_period'] != period]
return df
# The above function helps us identify periods when the temperature is constant and/or increasing/decreasing. We can use the periods to further restrict our co2 decay dataset.
# ## Iteratively solving for ventilation rate
# Now that we can get the periods of decreasing CO2 concentration, we can use the dynamic solution and continuously vary the air exchange rate until we get the lowest error between the measured and estimated concentrations at $t > 0$.
def get_ach_from_dynamic_co2(df, E, V, C0=400.0, p=1.0, plot=False, pt="", period="", method="", save=False):
'''
Calculates the ACH based on a dynamic solution to the mass balance equation
Inputs:
- df: dataframe indexed by time with CO2 column for CO2 measurements in ppm
- E: double of emission rate in L/s
- V: double of volume in ft3
- C0: double of outdoor co2 concentration in ppm (default is 400)
- p: double of penetration factor (default is 1)
Returns ach in 1/h
'''
# defining constants
rho = 1.8 # g/L
# converting units
E_gh = E * rho *3600 # L/s to g/h
V_m3 = V * 0.0283168 # ft3 to m3
df['c'] = convert_ppm_to_gm3(df['co2']) # ppm to g/m3
C0_gm3 = convert_ppm_to_gm3(C0) # ppm to g/m3
C_t0 = df['c'][0]
min_rmsd = math.inf
ach = -1
C_to_plot = df['c'].values
for ell in np.arange(0,20.01,0.001):
Cs = []
for i in range(len(df)):
t = i*300/3600
Cs.append(C_t0 * math.exp(-ell*t) + (p*C0_gm3 - E_gh/(V_m3*ell))*(1 - math.exp(-ell*t)))
rmsd = 0
for C_est, C_meas in zip(Cs,df['c']):
rmsd += (C_est-C_meas)**2
rmsd = math.sqrt(rmsd/len(Cs))
if rmsd < min_rmsd:
min_rmsd = rmsd
ach = ell
C_to_plot = Cs
# Plotting to compare results if flag turned on
if plot:
fig, ax = plt.subplots(figsize=(8,6))
ax.plot(df.index,df['c'],color='seagreen',label='Measured')
ax.plot(df.index,C_to_plot,color='firebrick',label=f'ACH={round(ach,2)}; RMSD={round(rmsd,3)}')
for i in range(len(Cs)):
ax.annotate(str(round(df['c'].values[i],2)),(df.index[i],df['c'].values[i]),ha="left",fontsize=12)
ax.annotate(str(round(C_to_plot[i],2)),(df.index[i],C_to_plot[i]),ha="right",fontsize=12)
ax.set_ylabel("CO$_2$ (g/m$^3$)",fontsize=16)
plt.yticks(fontsize=14)
ax.legend(fontsize=14)
plt.xticks(fontsize=14,ha="left",rotation=-15)
ax2 = ax.twinx()
ax2.plot(df.index,df['temperature_c'],color='cornflowerblue',label='Temperature')
ax2.spines['right'].set_color('cornflowerblue')
ax2.set_ylim([20,30])
plt.yticks(fontsize=14)
ax2.set_ylabel("Temperature ($^\circ$C)",fontsize=16,color="cornflowerblue")
ax2.xaxis.set_major_locator(mdates.MinuteLocator(interval=5))
ax2.xaxis.set_major_formatter(mdates.DateFormatter("%H:%M"))
ax.set_title(f"ID: {pt} - Period: {period}")
if save:
plt.savefig(f"../reports/figures/beacon_summary/ventilation_estimates/method_{method}-{pt}-{period}.pdf",bbox_inches="tight")
plt.show()
plt.close()
return ach, min_rmsd, C_to_plot
# <div class="alert-block alert alert-warning">
# <p>The above code could use some upgrades in terms of computation time as well as the method used to check how close the solution is. However, it seems to do the trick as of right now.</p>
# </div>
# ## Calculating ventilation rate based on decay
# We can use the above equations to test and check what the ventilation rate must be under various conditions.
# Here we go through the three different equations (starting bottom to top) to understand how the ventilation rate changes based on our assumptions.
# some parameters for all three cases:
window = 12
min_co2 = 600
plotting = False
# ### Occupant leaves and no outdoor CO2 penetration
# The most simple case we have: basic decay equation.
decay_eq3_df = pd.DataFrame()
for pt in beacon_co2['beiwe'].unique():
decay_dict = {'beiwe':[],'beacon':[],'start':[],'end':[],'ending_co2_meas':[],'ending_co2_calculated':[],'rmsd':[],'ach':[]}
# getting pt-specific data
beacon_co2_pt = morning_beacon_co2[morning_beacon_co2['beiwe'] == pt]
info_pt = pt_info[pt_info.index == pt]
# getting
decreasing_co2_ac_pt = get_co2_periods(beacon_co2_pt,window=window,change='decrease')
#decreasing_co2_ac_pt = get_change_T_periods(decreasing_co2_pt)
for period in decreasing_co2_ac_pt['period'].unique():
decreasing_period_ac_pt = decreasing_co2_ac_pt[decreasing_co2_ac_pt['period'] == period]
if np.nanmin(decreasing_period_ac_pt['co2']) >= min_co2:
T = np.nanmean(decreasing_period_ac_pt['temperature_c'])
E = 0
V = info_pt['volume'].values[0]
#print(f'{pt}: period {period}')
ach, ss, C_est = get_ach_from_dynamic_co2(decreasing_period_ac_pt,E,V,p=0,plot=plotting,pt=pt,period=period,method=1,save=False)
# adding information to dict
for key, value_to_add in zip(decay_dict.keys(),[pt,info_pt['beacon'].values[0],
decreasing_period_ac_pt.index[0],decreasing_period_ac_pt.index[-1],
decreasing_period_ac_pt['c'][-1],C_est[-1],
ss,ach]):
decay_dict[key].append(value_to_add)
decay_eq3_df = decay_eq3_df.append(pd.DataFrame(decay_dict))
# ### Occupant leaves
# Now we add in some penetration from outdoors
decay_eq2_df = pd.DataFrame()
for pt in beacon_co2['beiwe'].unique():
decay_dict = {'beiwe':[],'beacon':[],'start':[],'end':[],'ending_co2_meas':[],'ending_co2_calculated':[],'rmsd':[],'ach':[]}
# getting pt-specific data
beacon_co2_pt = morning_beacon_co2[morning_beacon_co2['beiwe'] == pt]
info_pt = pt_info[pt_info.index == pt]
# getting
decreasing_co2_ac_pt = get_co2_periods(beacon_co2_pt,window=window,change='decrease')
#decreasing_co2_ac_pt = get_change_T_periods(decreasing_co2_pt)
for period in decreasing_co2_ac_pt['period'].unique():
decreasing_period_ac_pt = decreasing_co2_ac_pt[decreasing_co2_ac_pt['period'] == period]
if np.nanmin(decreasing_period_ac_pt['co2']) >= min_co2:
T = np.nanmean(decreasing_period_ac_pt['temperature_c'])
E = 0
V = info_pt['volume'].values[0]
ach, ss, C_est = get_ach_from_dynamic_co2(decreasing_period_ac_pt,E,V,plot=plotting,pt=pt,period=period,method=2,save=True)
# adding information to dict
for key, value_to_add in zip(decay_dict.keys(),[pt,info_pt['beacon'].values[0],
decreasing_period_ac_pt.index[0],decreasing_period_ac_pt.index[-1],
decreasing_period_ac_pt['c'][-1],C_est[-1],
ss,ach]):
decay_dict[key].append(value_to_add)
decay_eq2_df = decay_eq2_df.append(pd.DataFrame(decay_dict))
# ### Occupant stays
# Lastly, we add in an emission source (the participant stays in their bedroom).
decay_df = pd.DataFrame()
for pt in beacon_co2['beiwe'].unique():
decay_dict = {'beiwe':[],'beacon':[],'start':[],'end':[],'ending_co2_meas':[],'ending_co2_calculated':[],'rmsd':[],'ach':[]}
# getting pt-specific data
beacon_co2_pt = morning_beacon_co2[morning_beacon_co2['beiwe'] == pt]
info_pt = pt_info[pt_info.index == pt]
# getting
decreasing_co2_ac_pt = get_co2_periods(beacon_co2_pt,window=window,change='decrease')
#decreasing_co2_ac_pt = get_change_T_periods(decreasing_co2_pt)
for period in decreasing_co2_ac_pt['period'].unique():
decreasing_period_ac_pt = decreasing_co2_ac_pt[decreasing_co2_ac_pt['period'] == period]
if np.nanmin(decreasing_period_ac_pt['co2']) >= min_co2:
T = np.nanmean(decreasing_period_ac_pt['temperature_c'])
E = get_emission_rate(pt_info.loc[pt,'bmr'],T+273)
V = info_pt['volume'].values[0]
ach, ss, C_est = get_ach_from_dynamic_co2(decreasing_period_ac_pt,E,V,plot=plotting)
#print(f'{pt}: period {period}')
# adding information to dict
for key, value_to_add in zip(decay_dict.keys(),[pt,info_pt['beacon'].values[0],
decreasing_period_ac_pt.index[0],decreasing_period_ac_pt.index[-1],
decreasing_period_ac_pt['c'][-1],C_est[-1],
ss,ach]):
decay_dict[key].append(value_to_add)
decay_df = decay_df.append(pd.DataFrame(decay_dict))
decay_df["ACH_EQ1"] = decay_df["ach"].values
decay_df["RMSD_EQ1"] = decay_df["rmsd"].values
decay_df['ACH_EQ3'] = decay_eq3_df['ach'].values
decay_df['RMSD_EQ3'] = decay_eq3_df['rmsd'].values
decay_df['ACH_EQ2'] = decay_eq2_df['ach'].values
decay_df['RMSD_EQ2'] = decay_eq2_df['rmsd'].values
decay_df["ach"] = decay_df['ACH_EQ2']
decay_df['method'] = 'decay'
decay_df['beacon'].unique()
ventilation_df['method'] = 'constant'
ventilation_df['beacon'].unique()
ventilation_estimates = decay_df[['beacon','beiwe','start','end','ach','method']].append(ventilation_df[['beacon','beiwe','start','end','ach','method']])
fig, ax = plt.subplots(figsize=(12,8))
#sns.stripplot(x='Beacon',y='ACH',color='#333f48',data=decay_df,s=15,jitter=0,marker='d',label='Decay')
#sns.stripplot(x='Beacon',y='ACH',color='#bf5700',data=ventilation_df,s=12,jitter=0,label='Constant')
ax.scatter(decay_df['beacon'],decay_df['ach'],s=220,color='#333f48',marker='d',label='Decay')
ax.scatter(ventilation_df['beacon'],ventilation_df['ach'],s=175,color='#bf5700',label='Constant')
ax.legend(frameon=False)
ax.set_ylim([0,1.6])
ax.set_xlim([0,40])
ax.set_xticks(ventilation_estimates['beacon'].unique())
#plt.savefig('../reports/conferences/ASHRAESummer_2020/ventilation_estimates.pdf')
plt.show()
plt.close()
# ### Saving
decay_df.to_csv("../data/processed/beacon-ventilation_from_decay.csv",index=False)
decay_df.head()
ventilation_df.to_csv("../data/processed/beacon-ventilation_from_ss.csv",index=False)
ventilation_df.head()
ventilation_estimates.to_csv("../data/processed/beacon-ventilation.csv",index=False)
ventilation_estimates.head()
| notebooks/4.1.4-hef-beacon-ventilation_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Introduction to programming
# ## Legend
# <div class = "alert alert-block alert-info">
# In <b> blue</b>, the <b> instructions </b> and <b> goals </b> are highlighted.
# </div>
# <div class = "alert alert-block alert-success">
# In <b> green</b>, the <b> information </b> is highlighted.
# </div>
# <div class = "alert alert-block alert-warning">
# In <b> yellow</b>, the <b> exercises </b> are highlighted.
# </div>
# <div class = "alert alert-block alert-danger">
# In <b> red</b>, the <b> error </b> and <b> alert messages </b> are highlighted.
# </div>
# ## Instructions
# <div class = "alert alert-block alert-info">
# <ul>
# <li> Click "Run" on each cell to go through the code in each cell. This will take you through the cell and print out the results. </li>
# <li> If you wish to see all the outputs at once in the whole notebook, just click Cell and then Run All. </li>
# </ul>
# </div>
# ## Goal
# <div class = "alert alert-block alert-info">
# After this workshop, the student should get more familiar with the following topics: <br>
# <ul>
# <li> printing basic statements and commands in Jupyter Notebook</li>
# <li> performing basic arithmetic calculations in Python</li>
# <li> improving an existent model of the code</li>
# <li> recognizing and checking variable types in Python </li>
# </ul>
#
# <b> These objectives are in agreement with the National 3 Scottish Curriculum for high-school students. </b> <br> <br> <b> Note: </b> For most of the workshop, the student will be given some coding examples. In some cases, the student will have to code himself/herself. The coding part is optional, but highly recommended to approach.
#
# </div>
# ## Explore
# ### Variable types
# <div class = "alert alert-block alert-success">
# Let us talk about one important aspect in computing. So far, we have put straight instructions to calculate sums, differences, products and so on. However, we can work with what we call <b> variables. </b> The best example is shown below:
# </div>
# +
a = 1
b = 2
print("Variable a has the following value: " + str(a))
print("Variable b has the following value: " + str(b))
print("The sum of the variables is the following: " + str(a+b))
print("The difference of the variables is the following: " + str(a-b))
# -
# <div class = "alert alert-block alert-success">
# At the beginning, when we calculated directly print(2+3), the number within the brackets is called a variable, as $ 2+3 = 5$. Now, we assigned a and b variables two values. The expression $ a=1 $ in Python stands for the following: <b> create a variable a and assign it value 1 </b>. Let us learn something about the variables: they can have various types. How can that be?
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> Run through the following cells and investigate the outputs. Discuss the outputs of the code with your colleagues and teacher. Can you suspect what does the type function do?
# </div>
a = 3
type(a)
b = 3.4
type(b)
c = 'a'
type(c)
d = "a"
type(d)
e = True
type(e)
# <div class = "alert alert-block alert-success">
#
# There are four main data types in Python:
#
# <ul>
#
# <li> data type <b> int </b>, where the variable is an integer: $ 0 $, $ \pm 1 $, $ \pm 2 $, $ \pm 3 $,... </li>
#
# <li> data type <b> float </b>, where the variable is any type of decimal (rational or irrational numbers): $1.5$, $ \pi $, $3.00$, $2.347563786578325678$, ... </li>
#
# <li> data type <b> string </b>, where the variable can be a single character - a,b,A,C,z,Z - or any arrangement of more characters - "it is nice", "feggk", "wGsX" </li>
#
# <li> data type <b> bool </b>, where the variable is a logical truth value - either <b> True </b> or <b> False </b> </li>
#
# </ul>
#
# </div>
# <div class = "alert alert-block alert-warning">
# What type of varibale is $ f = -2 $? Use the acquired information first, then check on your Notebook.
# +
# Type the answer here
# -
# We have assigned plenty of variables: a,b,c,d,e,f... Cannot we do better?
a = -2
a
# <div class = "alert alert-block alert-success">
# Before a was 3, and now we assigned it to 2... <b> Variables in Python can be given other values at any time! </b> This is one of the great advantages of programming!!
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Predict: </b> Can you figure out what will be the final value of a in the bottom code cell?
# </div>
# First think about the output of the code, then run the cell
a=3
a=2
a=-1
a=0.4
a=0
print(a)
# All the previous values of a - $ (3,2,-1, 0.4) $ are now lost.
# <div class = "alert alert-block alert-warning">
# <b> Exercise: </b> Can you run again through the same values of a, and print each value, so that each time the variable is printed?
# +
# Type your answer here
# -
# <div class = "alert alert-block alert-warning">
# <b> Predict: </b> Take again the cell below and analyse it for a bit: what happens if you remove the str part. Talk about it in pairs or threes, do not remove anything from the cell, just process it through your mind. After you had some debate, delete the str type and the brackets ():
# </div>
# +
a = 1
b = 2
print("Variable a has the following value: " + str(a))
# -
# <div class = "alert alert-block alert-danger">
# When taken together with a string, the variable a needs to be converted to a string as well, otherwise in the print sequence there will be combination of strings and integers, and this does not work for the print statement. Hence, by adding the str() command, a becomes from integer a string. <b> Now go back to the question at the beginning: </b> What happens if I just type the following command? Again, talk about it in pairs before you type the cell:
# </div>
print(Hello!! Welcome to your first programming workshop. I am here to help you)
# <div class = "alert alert-block alert-danger">
# <b> Answer: </b> The whole sequence is treated as a pile of integers!! This is clearly not the case, as hello is a word in our case, not a number. Of course, the situation looks a lot different in the following scenario:
# </div>
# In this scenario, let us take a variable hello which is an integer of value 2. Afterwards, we will print the sentence hello, as well as another statment about the hello variable. There will be some subtleties which need to be discussed in groups.
# print solution here
hello = 2
print("hello!!")
print("The hello variable takes value: " + str(hello))
# <div class = "alert alert-block alert-warning">
# <b> Investigate: </b> How do hello and "hello" differ?
# </div>
# <div class = "alert alert-block alert-success">
# In one case, hello is a string, while in the other case, hello is an integer.
# </div>
# <div class = "alert alert-block alert-warning">
# <b> Investigate again: </b> Compare the following without running them initially:
# </div>
print(2+3)
print("2+3")
# #### Difference between = and ==
a = 5
a = 4
print(a)
print(5 == 4)
print(5 = 4)
# <div class = "alert alert-block alert-success">
# So the main idea is the following: <b> = </b> expression <b> assigns </b> a value to the variable, while <b> == </b> expresion <b> stands for the equality sign. </b> This is the one you are very familiar with: is 1 equal to 3, 4 equal to 4 and so on...
# </div>
# ## Take-away
# <div class = "alert alert-block alert-success">
# This is it for today, and well done for managing to go through the material!! <br> <br> After this session, you should be more familiar with how simple sentences, numbers and conditional statements can be printed in Python. Also, feel free to work more on this notebook using any commands you would like. <br> <br>
# <b> <Note:> </b> Always keep a back-up of the notebook, in case the original one is altered.
# </div>
# For today's session, this should be enough! See you later!!
print("Bye bye! :D")
# 
| GeneralExemplars/Coding Activities for Schools/National 3/Intro_Nat3_Part_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Install the PIC-SURE-HPDS client library
# +
import sys
# #!{sys.executable} -m pip uninstall git+https://github.com/hms-dbmi/pic-sure-python-client.git
# #!{sys.executable} -m pip uninstall git+https://github.com/hms-dbmi/pic-sure-python-adapter-hpds.git
# !{sys.executable} -m pip install --upgrade pip
# !{sys.executable} -m pip install git+https://github.com/hms-dbmi/pic-sure-python-client.git
# !{sys.executable} -m pip install git+https://github.com/hms-dbmi/pic-sure-python-adapter-hpds.git
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install matplotlib
# -
# # Connect directly to the HPDS resource, bypassing all security. This should never be possible in an instance holding private data.
#
# # An HPDS instance should always be hosted behind a PIC-SURE API and PIC-SURE Auth Micro-App instance if it has any privacy concern.
from pandas import pandas
import PicSureHpdsLib
adapter = PicSureHpdsLib.BypassAdapter("http://pic-sure-hpds-nhanes:8080/PIC-SURE")
resource = adapter.useResource()
# # Retrieve the data dictionary
data_dictionary = resource.dictionary().find()
all_concepts = data_dictionary.DataFrame()
# # See how many concepts were loaded in total
all_concepts.shape[0]
# # Check the expected counts and ranges for each of your variables. This will require comparing these values to your source data.
# +
sorted_concepts = all_concepts.sort_index(axis = 0)
sorted_concepts
# -
# # Because the source file is sorted by CONCEPT_PATH and PATIENT_NUM it is fairly straight forward to confirm that the rows match. First we retrieve all concepts for all patients.
query = resource.query()
query.select().add(data_dictionary.keys())
all_concepts_dataframe = query.getResultsDataFrame()
# # We unpivot the data back into a single line per patient and concept like the original file sorted in the same order as the original file for performance.
melted = pandas.melt(all_concepts_dataframe, col_level=0, id_vars=['Patient ID'])
loaded_data = melted.sort_values(by=['variable', 'Patient ID'])
# # We then load the original source file
original_data = pandas.read_csv('allConcepts.csv')
# # ... and join the NVAL_NUM and TVAL_CHAR entries on our loaded dataset
joined_data = pandas.merge(original_data, loaded_data, left_on=['PATIENT_NUM','CONCEPT_PATH'], right_on=['Patient ID','variable']).drop(columns=['Patient ID', 'variable'])
# # We can see here that most of the data seems ok, but we can do better than that.
joined_data
# # We filter out all rows where the value in our loaded dataset matches either the NVAL_NUM or TVAL_CHAR value
unmatched_rows = joined_data[(joined_data['value'] != joined_data['TVAL_CHAR']) | (joined_data['value'] != joined_data['NVAL_NUM'])]
unmatched_rows
# # We see that some numeric values loaded are failing to match because they are interpreted as text, so we parse them and filter the matches out of the dataset.
unmatched_numeric = unmatched_rows
unmatched_numeric[['value']] = unmatched_numeric[['value']].apply(pandas.to_numeric)
still_unmatched = unmatched_numeric[(unmatched_numeric['value'] != unmatched_numeric['NVAL_NUM'])]
# # As it turns out, some numeric values are in the TVAL_CHAR column, we also parse them and filter out their matches as HPDS automatically converts these for us on load.
still_unmatched
unmatched_char = still_unmatched
unmatched_char = unmatched_char.loc[unmatched_char['TVAL_CHAR'] != 'E']
unmatched_char[['TVAL_CHAR']] = unmatched_char[['TVAL_CHAR']].apply(pandas.to_numeric)
unmatched_char
still_unmatched_char = unmatched_char[(unmatched_char['value'] != unmatched_char['TVAL_CHAR'])]
# # We can now see that all of the discrepancies are explained by the above parsing oddities. This means all our data is in the HPDS datastore.
still_unmatched_char
| jupyter-notebooks/Data Load Statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="LcZmS4jqK-uD" outputId="0b9f9bd6-4718-410f-c158-9923cbbdfd83" colab={"base_uri": "https://localhost:8080/"}
pip install imageai --upgrade
# + id="L45np-FfK-uI" outputId="205735d4-2516-42e9-aeeb-92f2661681d3" colab={"base_uri": "https://localhost:8080/", "height": 375}
from imageai.Detection import ObjectDetection
import os
execution_path = os.getcwd()
detector = ObjectDetection()
detector.setModelTypeAsRetinaNet()
detector.setModelPath( os.path.join(execution_path , "resnet50_coco_best_v2.1.0.h5"))
detector.loadModel()
detections = detector.detectObjectsFromImage(input_image=os.path.join(execution_path , "image.jpg"),
output_image_path=os.path.join(execution_path , "imagenew.jpg"))
for eachObject in detections:
print(eachObject["name"] , " : " , eachObject["percentage_probability"] )
# + id="sNEXBWNjK-uK" outputId="8944a1ec-6aa3-4b6f-a79f-d29409fc606c" colab={"base_uri": "https://localhost:8080/", "height": 351}
from imageai.Detection.Custom import CustomObjectDetection
detector = CustomObjectDetection()
detector.setModelTypeAsYOLOv3()
detector.setModelPath("detection_model-ex-010--loss-0044.074.h5")
detector.setJsonPath("./custom/puzzle/json/detection_config.json")
detector.loadModel()
detections = detector.detectObjectsFromImage(input_image="image.jpg", output_image_path="image_detected.jpg")
for detection in detections:
print(detection["name"], " : ", detection["percentage_probability"], " : ", detection["box_points"])
# + id="lsTYrWsH1T_z" outputId="e0caa61a-7e6b-4f35-d21c-7459e4994295" colab={"base_uri": "https://localhost:8080/"}
# ls
# + id="pDtGOWyM1UjQ"
| OpenAI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Lesson 3: New Activities and Intents
# * 싸이그래머 / 인지모델링 - 파트 1 : 안드로이드
# * 김무성
# # Contents
# * Introduction to Lesson 3
# * What to do next?
# * Sunshine App UX Mocks
# * List Item Click Listener
# * ItemClickListener and Toast
# * Create New Activity
# * Match Our Code
# * Intents Framework
# * Intents as Envelopes
# * Launch DetailActivity
# * Display Content in DetailActivity
# * It works!
# * Settings UX
# * Preferences
# * Create SettingsActivity
# * Launch SettingsActivity
# * Location Setting XML
# * Modify SettingsActivity
# * Use SharedPreferences
# * Update Data on Activity Start
# * Temperature Units Setting
# * Debug Breakpoints
# * Launching Implicit Intents
# * Add Map Location Intent
# * Intent Resolution
# * Share Intent is Awesome
# * Broadcast Intents
# * Intent Filters
# * Lesson 3 Recap
# * Storytime: Android Distribution Platform
# # Introduction to Lesson 3
# # What to do next?
# # Sunshine App UX Mocks
# <img src="figures/cap3.1.png" width=600 />
# <img src="figures/cap3.2.png" width=600 />
# # List Item Click Listener
# <img src="figures/cap3.3.png" width=600 />
# <img src="figures/cap3.4.png" width=600 />
# # ItemClickListener and Toast
# <img src="figures/cap3.5.png" width=600 />
# <img src="figures/cap3.6.png" width=600 />
# <img src="figures/cap3.7.png" width=600 />
# <img src="figures/cap3.8.png" width=600 />
# <img src="figures/cap3.9.png" width=600 />
# <img src="figures/cap3.10.png" width=600 />
# # Create New Activity
# <img src="figures/cap3.11.png" width=600 />
# <img src="figures/cap3.12.png" width=600 />
# <img src="figures/cap3.13.png" width=600 />
# <img src="figures/cap3.14.png" width=600 />
# <img src="figures/cap3.15.png" width=600 />
# <img src="figures/cap3.16.png" width=600 />
# <img src="figures/cap3.17.png" width=600 />
# <img src="figures/cap3.18.png" width=600 />
# # Match Our Code
# # Intents Framework
# <img src="figures/cap3.19.png" width=600 />
# <img src="figures/cap3.20.png" width=600 />
# <img src="figures/cap3.21.png" width=600 />
# <img src="figures/cap3.22.png" width=600 />
# <img src="figures/cap3.23.png" width=600 />
# <img src="figures/cap3.24.png" width=600 />
# <img src="figures/cap3.25.png" width=600 />
# # Intents as Envelopes
# <img src="figures/cap3.26.png" width=600 />
# <img src="figures/cap3.27.png" width=600 />
# <img src="figures/cap3.28.png" width=600 />
# <img src="figures/cap3.29.png" width=600 />
# <img src="figures/cap3.30.png" width=600 />
# <img src="figures/cap3.31.png" width=600 />
# <img src="figures/cap3.32.png" width=600 />
# <img src="figures/cap3.33.png" width=600 />
# <img src="figures/cap3.34.png" width=600 />
# <img src="figures/cap3.35.png" width=600 />
# <img src="figures/cap3.36.png" width=600 />
# <img src="figures/cap3.37.png" width=600 />
# <img src="figures/cap3.38.png" width=600 />
# <img src="figures/cap3.39.png" width=600 />
# <img src="figures/cap3.40.png" width=600 />
# <img src="figures/cap3.41.png" width=600 />
# <img src="figures/cap3.42.png" width=600 />
# # Launch DetailActivity
# <img src="figures/cap3.43.png" width=600 />
# <img src="figures/cap3.44.png" width=600 />
# <img src="figures/cap3.45.png" width=600 />
# <img src="figures/cap3.46.png" width=600 />
# <img src="figures/cap3.47.png" width=600 />
# <img src="figures/cap3.48.png" width=600 />
# <img src="figures/cap3.49.png" width=600 />
# <img src="figures/cap3.50.png" width=600 />
# # Display Content in DetailActivity
# <img src="figures/cap3.51.png" width=600 />
# <img src="figures/cap3.52.png" width=600 />
# <img src="figures/cap3.53.png" width=600 />
# <img src="figures/cap3.54.png" width=600 />
# <img src="figures/cap3.55.png" width=600 />
# <img src="figures/cap3.56.png" width=600 />
# <img src="figures/cap3.57.png" width=600 />
# <img src="figures/cap3.58.png" width=600 />
# <img src="figures/cap3.59.png" width=600 />
# # It works!
# <img src="figures/cap3.60.png" width=600 />
# <img src="figures/cap3.61.png" width=600 />
# # Settings UX
# <img src="figures/cap3.62.png" width=600 />
# <img src="figures/cap3.63.png" width=600 />
# <img src="figures/cap3.64.png" width=600 />
# <img src="figures/cap3.65.png" width=600 />
# <img src="figures/cap3.66.png" width=600 />
# <img src="figures/cap3.67.png" width=600 />
# <img src="figures/cap3.68.png" width=600 />
# <img src="figures/cap3.69.png" width=600 />
# <img src="figures/cap3.70.png" width=600 />
# <img src="figures/cap3.71.png" width=600 />
# <img src="figures/cap3.72.png" width=600 />
# # Preferences
# <img src="figures/cap3.73.png" width=600 />
# # Create SettingsActivity
# <img src="figures/cap3.74.png" width=600 />
# <img src="figures/cap3.75.png" width=600 />
# <img src="figures/cap3.76.png" width=600 />
# <img src="figures/cap3.77.png" width=600 />
# <img src="figures/cap3.78.png" width=600 />
# <img src="figures/cap3.79.png" width=600 />
# # Launch SettingsActivity
# <img src="figures/cap3.80.png" width=600 />
# <img src="figures/cap3.81.png" width=600 />
# <img src="figures/cap3.82.png" width=600 />
# <img src="figures/cap3.83.png" width=600 />
# <img src="figures/cap3.84.png" width=600 />
# <img src="figures/cap3.85.png" width=600 />
# <img src="figures/cap3.86.png" width=600 />
# <img src="figures/cap3.87.png" width=600 />
# <img src="figures/cap3.88.png" width=600 />
# <img src="figures/cap3.89.png" width=600 />
# <img src="figures/cap3.90.png" width=600 />
# <img src="figures/cap3.91.png" width=600 />
# <img src="figures/cap3.92.png" width=600 />
# <img src="figures/cap3.93.png" width=600 />
# <img src="figures/cap3.94.png" width=600 />
# <img src="figures/cap3.95.png" width=600 />
# # Location Setting XML
# <img src="figures/cap3.96.png" width=600 />
# <img src="figures/cap3.97.png" width=600 />
# <img src="figures/cap3.98.png" width=600 />
# <img src="figures/cap3.99.png" width=600 />
# <img src="figures/cap3.100.png" width=600 />
# <img src="figures/cap3.101.png" width=600 />
# <img src="figures/cap3.102.png" width=600 />
# <img src="figures/cap3.103.png" width=600 />
# <img src="figures/cap3.104.png" width=600 />
# <img src="figures/cap3.105.png" width=600 />
# <img src="figures/cap3.106.png" width=600 />
# <img src="figures/cap3.107.png" width=600 />
# <img src="figures/cap3.108.png" width=600 />
# <img src="figures/cap3.109.png" width=600 />
# <img src="figures/cap3.110.png" width=600 />
# <img src="figures/cap3.111.png" width=600 />
# <img src="figures/cap3.112.png" width=600 />
# <img src="figures/cap3.113.png" width=600 />
# <img src="figures/cap3.114.png" width=600 />
# <img src="figures/cap3.115.png" width=600 />
# <img src="figures/cap3.116.png" width=600 />
# <img src="figures/cap3.117.png" width=600 />
# <img src="figures/cap3.118.png" width=600 />
# <img src="figures/cap3.119.png" width=600 />
# <img src="figures/cap3.120.png" width=600 />
# <img src="figures/cap3.121.png" width=600 />
# # Modify SettingsActivity
# <img src="figures/cap3.122.png" width=600 />
# <img src="figures/cap3.123.png" width=600 />
# <img src="figures/cap3.124.png" width=600 />
# <img src="figures/cap3.125.png" width=600 />
# <img src="figures/cap3.126.png" width=600 />
# <img src="figures/cap3.127.png" width=600 />
# <img src="figures/cap3.128.png" width=600 />
# <img src="figures/cap3.129.png" width=600 />
# <img src="figures/cap3.130.png" width=600 />
# <img src="figures/cap3.131.png" width=600 />
# <img src="figures/cap3.132.png" width=600 />
# <img src="figures/cap3.133.png" width=600 />
# <img src="figures/cap3.134.png" width=600 />
# <img src="figures/cap3.135.png" width=600 />
# <img src="figures/cap3.136.png" width=600 />
# # Use SharedPreferences
# <img src="figures/cap3.137.png" width=600 />
# <img src="figures/cap3.138.png" width=600 />
# <img src="figures/cap3.139.png" width=600 />
# <img src="figures/cap3.140.png" width=600 />
# <img src="figures/cap3.141.png" width=600 />
# <img src="figures/cap3.142.png" width=600 />
# <img src="figures/cap3.143.png" width=600 />
# <img src="figures/cap3.144.png" width=600 />
# # Update Data on Activity Start
# <img src="figures/cap3.145.png" width=600 />
# <img src="figures/cap3.146.png" width=600 />
# <img src="figures/cap3.147.png" width=600 />
# <img src="figures/cap3.148.png" width=600 />
# <img src="figures/cap3.149.png" width=600 />
# <img src="figures/cap3.149.png" width=600 />
# <img src="figures/cap3.150.png" width=600 />
# <img src="figures/cap3.151.png" width=600 />
# <img src="figures/cap3.152.png" width=600 />
# <img src="figures/cap3.153.png" width=600 />
# <img src="figures/cap3.154.png" width=600 />
# <img src="figures/cap3.155.png" width=600 />
# <img src="figures/cap3.156.png" width=600 />
# <img src="figures/cap3.157.png" width=600 />
# <img src="figures/cap3.158.png" width=600 />
# # Temperature Units Setting
# <img src="figures/cap3.159.png" width=600 />
# <img src="figures/cap3.160.png" width=600 />
# <img src="figures/cap3.161.png" width=600 />
# <img src="figures/cap3.162.png" width=600 />
# <img src="figures/cap3.163.png" width=600 />
# <img src="figures/cap3.164.png" width=600 />
# # Debug Breakpoints
# # Launching Implicit Intents
# <img src="figures/cap3.165.png" width=600 />
# # Add Map Location Intent
# <img src="figures/cap3.166.png" width=600 />
# <img src="figures/cap3.167.png" width=600 />
# <img src="figures/cap3.168.png" width=600 />
# <img src="figures/cap3.169.png" width=600 />
# <img src="figures/cap3.170.png" width=600 />
# <img src="figures/cap3.171.png" width=600 />
# <img src="figures/cap3.172.png" width=600 />
# <img src="figures/cap3.173.png" width=600 />
# <img src="figures/cap3.174.png" width=600 />
# <img src="figures/cap3.175.png" width=600 />
# <img src="figures/cap3.176.png" width=600 />
# <img src="figures/cap3.179.png" width=600 />
# <img src="figures/cap3.180.png" width=600 />
# <img src="figures/cap3.181.png" width=600 />
# <img src="figures/cap3.182.png" width=600 />
# <img src="figures/cap3.183.png" width=600 />
# <img src="figures/cap3.184.png" width=600 />
# <img src="figures/cap3.185.png" width=600 />
# # Intent Resolution
# <img src="figures/cap3.186.png" width=600 />
# <img src="figures/cap3.187.png" width=600 />
# <img src="figures/cap3.188.png" width=600 />
# <img src="figures/cap3.189.png" width=600 />
# <img src="figures/cap3.190.png" width=600 />
# <img src="figures/cap3.191.png" width=600 />
# # Share Intent is Awesome
# <img src="figures/cap3.192.png" width=600 />
# <img src="figures/cap3.193.png" width=600 />
# <img src="figures/cap3.194.png" width=600 />
# <img src="figures/cap3.195.png" width=600 />
# <img src="figures/cap3.196.png" width=600 />
# <img src="figures/cap3.197.png" width=600 />
# <img src="figures/cap3.200.png" width=600 />
# <img src="figures/cap3.201.png" width=600 />
# <img src="figures/cap3.202.png" width=600 />
# <img src="figures/cap3.203.png" width=600 />
# <img src="figures/cap3.204.png" width=600 />
# <img src="figures/cap3.205.png" width=600 />
# <img src="figures/cap3.206.png" width=600 />
# <img src="figures/cap3.207.png" width=600 />
# <img src="figures/cap3.208.png" width=600 />
# <img src="figures/cap3.209.png" width=600 />
# <img src="figures/cap3.210.png" width=600 />
# <img src="figures/cap3.211.png" width=600 />
# <img src="figures/cap3.212.png" width=600 />
# <img src="figures/cap3.213.png" width=600 />
# <img src="figures/cap3.214.png" width=600 />
# <img src="figures/cap3.215.png" width=600 />
# <img src="figures/cap3.216.png" width=600 />
# <img src="figures/cap3.217.png" width=600 />
# <img src="figures/cap3.218.png" width=600 />
# <img src="figures/cap3.219.png" width=600 />
# <img src="figures/cap3.220.png" width=600 />
# <img src="figures/cap3.221.png" width=600 />
# <img src="figures/cap3.222.png" width=600 />
# <img src="figures/cap3.223.png" width=600 />
# <img src="figures/cap3.224.png" width=600 />
# <img src="figures/cap3.225.png" width=600 />
# <img src="figures/cap3.226.png" width=600 />
# <img src="figures/cap3.227.png" width=600 />
# <img src="figures/cap3.228.png" width=600 />
# <img src="figures/cap3.229.png" width=600 />
# <img src="figures/cap3.230.png" width=600 />
# <img src="figures/cap3.231.png" width=600 />
# <img src="figures/cap3.232.png" width=600 />
# <img src="figures/cap3.233.png" width=600 />
#
# <img src="figures/cap3.234.png" width=600 />
# <img src="figures/cap3.235.png" width=600 />
# # Broadcast Intents
# <img src="figures/cap3.236.png" width=600 />
# <img src="figures/cap3.237.png" width=600 />
# <img src="figures/cap3.238.png" width=600 />
# # Intent Filters
# <img src="figures/cap3.239.png" width=600 />
# <img src="figures/cap3.240.png" width=600 />
# <img src="figures/cap3.241.png" width=600 />
# <img src="figures/cap3.242.png" width=600 />
# <img src="figures/cap3.245.png" width=600 />
# <img src="figures/cap3.246.png" width=600 />
# <img src="figures/cap3.247.png" width=600 />
# <img src="figures/cap3.248.png" width=600 />
# <img src="figures/cap3.249.png" width=600 />
# <img src="figures/cap3.250.png" width=600 />
# <img src="figures/cap3.251.png" width=600 />
# <img src="figures/cap3.252.png" width=600 />
# # Lesson 3 Recap
# # Storytime: Android Distribution Platform
# # 참고자료
| part2/android/sunshine03/03_New_Activities_and_Intents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.13 ('ml')
# language: python
# name: python3
# ---
# ### Implement decision tree
# Use decision tree to solve the Balance Scale Weight & Distance Database problem.<br>
# Following the example from: <br>
# https://www.geeksforgeeks.org/decision-tree-implementation-python/ <br>
# Link to the dataset: <br>
# https://archive.ics.uci.edu/ml/machine-learning-databases/balance-scale/
#
import numpy as np
import pandas as pd
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
import sklearn.tree
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
# #### Data preparation
# +
# Download and loda data
balance_data = pd.read_csv(
'https://archive.ics.uci.edu/ml/machine-learning-'+
'databases/balance-scale/balance-scale.data',
sep= ',', header = None)
# Printing the dataset shape
print ("Dataset Length: ", len(balance_data))
print ("Dataset Shape: ", balance_data.shape)
# Printing the dataset obseravtions
print ("Dataset: ",balance_data.head())
# +
# train-test split
# Separating the target variable
X = balance_data.values[:, 1:5]
Y = balance_data.values[:, 0]
# Splitting the dataset into train and test
X_train, X_test, y_train, y_test = train_test_split(
X, Y, test_size = 0.3, random_state = 100)
# -
# #### Train model
#
# There are two algorithms to train the model (choose features for the nodes):
# - Gini impurity
# - Information gain (entropy based)
#
# We will perform both of them.
# Function to perform training with Gini index (impurity)
def train_using_gini(X_train, X_test, y_train):
# Creating the classifier object
clf_gini = DecisionTreeClassifier(criterion = "gini",
random_state = 100,max_depth=3, min_samples_leaf=5)
# Performing training
clf_gini.fit(X_train, y_train)
return clf_gini
# Function to perform training with entropy.
def tarin_using_entropy(X_train, X_test, y_train):
# Decision tree with entropy
clf_entropy = DecisionTreeClassifier(
criterion = "entropy", random_state = 100,
max_depth = 3, min_samples_leaf = 5)
# Performing training
clf_entropy.fit(X_train, y_train)
return clf_entropy
# train model using gini index
clf_gini = train_using_gini(X_train, X_test, y_train)
# train model using information gain
clf_entropy = tarin_using_entropy(X_train, X_test, y_train)
# plot decision tree using gini index
sklearn.tree.plot_tree(clf_gini)
# plot decision tree using information gain
sklearn.tree.plot_tree(clf_entropy)
# #### Model evaluation
# Function to make predictions
def prediction(X_test, clf_object):
# Predicton on test with giniIndex
y_pred = clf_object.predict(X_test)
print("Predicted values:")
print(y_pred)
return y_pred
# Function to calculate accuracy
def cal_accuracy(y_test, y_pred):
print("Confusion Matrix: ",
confusion_matrix(y_test, y_pred))
print ("Accuracy : ",
accuracy_score(y_test,y_pred)*100)
print("Report : ",
classification_report(y_test, y_pred))
print("Results Using Gini Index:")
# Prediction using gini
y_pred_gini = prediction(X_test, clf_gini)
cal_accuracy(y_test, y_pred_gini)
print("Results Using Entropy:")
# Prediction using entropy
y_pred_entropy = prediction(X_test, clf_entropy)
cal_accuracy(y_test, y_pred_entropy)
| classic/decision_tree/decision_tree_sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img style="float: center; width: 100%" src="https://raw.githubusercontent.com/andrejkk/TalksImgs/master/FrontSlideUpperBan.png">
# <p style="margin-bottom:2cm;"></p>
#
# <center>
# <H1> 1. Uvod v Optimizacijo (OP) v telekomunikacijah (TK) OP v TK </H1>
#
#
#
# <br><br>
# <H3> <NAME>, Lucami, FE </H3>
# <H4> Kontakt: prof. dr. <NAME>, <EMAIL>, skype=akosir_sid </H4>
# </center>
#
#
# <p style="margin-bottom:2cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 1 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> 1. Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> Cilji, ponudba </div>
# </div>
#
#
# ## Cilji, ponudba, izpiti
#
# - Cilji predmeta: Prepoznati probleme optimizacije v realni praksi in skrajšati čas do njihove rešitve:
# - seznaniti s postopki optimizacije
# - usposobiti za samostojne preproste analize, uporaba računalnika
# - usposobiti za sodelovanje s strokovnjaki pri zahtevnejših analizah
# - seznaniti z možnostmi optimizacije
#
# - Spletna ponudba
# - Spletna stran EFE
#
# - Izpitni red
# - Projekti vaj (30%)+ projekti (30%)+ ustni izpit (40%), udeležba vseh treh prispevkov je obvezna
# - Izpitni red FE
#
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 2 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# # 1.1. Umestitev optimizacije v TK
#
# ■ Osnovni pojmi in primeri
#
# ■ Sodobne telekomunikacije in optimizacija
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 3 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
# ## ■ Osnovni pojmi in primeri
#
#
# - Izvor pojma?
# - Kaj so operacijske raziskave
# - Uporaba v logistiki, prometu, industrijski procesi, ...
#
#
# - Kaj je optimizacija
# - Optimizacijska naloga
# - Primer optimizacije:
# 1. optimalna oblika posode
# 2. Problem najkrajše (najcenejše) poti;
#
#
# - Pomembni pojmi
# - Optimizacijski problemi
# - Optimizacijska naloga
# - Algoritem
# - Podatkoven strukture (graf, vrsta, sklad)
# - Problemi: optimalno razvrščanje, optimalna alokacija virov, ...
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 4 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
# ## ■ Sodobne telekomunikacije in optimizacija
#
# - Optimizacija na OSI nivojih? Ne - uporabnik daje kriterijski funkcijo
# - TEDx Ljubljana govor o Socialni Inteligenci https://www.youtube.com/watch?v=YtmAcupMYK8&t=235s
#
# <img style="float: right; width: 400px;" src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/OSI_slo.png">
# <img style="float: left; width: 300px;" src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/ORinTC_slo.png">
#
#
# <br>
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 5 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Sodobne telekomunikacije in optimizacija
#
# - Uporabnik je v središču: upravljalski vidik
# - Vodstva velikih TK podjetij: „ne upravlja več uprava, ampak uporabniki“:
# - Izbor storitev
# - Neposreden vpliv na uspešnost podjetij
# - Odvisnost od socialnih omrežij
#
# - Danes ne plačujemo več “impulzov in Kb”, ampak pakete, znotraj katerih uporabniki sami določamo uporabo
#
# - Skupni moto gurujev: „User data is the oil of the future“
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 6 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (1)
#
#
# - <NAME> se definicija
# - Komunikacija: prenos informacij s simboli
# - Tele - : s pomočjo električnih sistemov
# - Spreminjajo civilizacijo
#
# - Zgodovina – komunikacijska tehnologija:
# - -4. stoletje: grški hidravlični sistemi semafor;
# - 1792: <NAME>, zgradil prvi stalni vizualni radiotelegraf;
# - 1839: komercialni električni telegraf;
# - 1858: telegrafski kabel Atlantik
# - 1878: telefon
# - 1900: začetek telegrafije , radiokomunikacij
# - 1920: začetek radiokomunikacij
# - 1940: frekvenčno multipleksiranje
# - 1960: satelitske komunikacije
# - 1980: optično vlakno
# - 2000: Gigabiten prenos po vlaknu
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 7 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (2)
#
# - Zgodovina – komunikacijska tehnologija (nad.):
# - Eneida, Didona, Kartagina, volovska koža, poskusi s pasom, krog
# - Platon (Republika) – osnovna delitev dela – človek, ki počne le en posel mora postati v tem izvrsten
# - Tiran v Sirakuzi poveri obrambo Arhimedu
# - <NAME>
# - 17 stol Fermat, <NAME>; verjetnost
# - 1758 <NAME> - ekonomske tablice
# - 1764 <NAME> - parni stroj
# - 1776 Monge - problem vpliva vzdrževanja poti na ceno prevozov
# - 1776 <NAME> (Wealth of Nations) - o prednostih delitve dela
# - 1802 Ampere - začetki teorije iger
# - 1824 Fourier - sistemi linearnih neenačb - LP
# - 1832 <NAME> (On Economy of Machines and Manufactures)
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 8 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (3)
#
# Zgodovina z OR povezanih dogodkov
#
# - 18?? Hamilton - hamiltonov problem v teoriji grafov
# - bencinski motor
# - elektromotor
# - 1874 Walras - ravnotežje v ekonomiki
# - 18-- Pareto – večkriterijalnost
# - 18-- Kirchoff - vezja
# - 1911 <NAME> - znanstveno upravljanje proizvodnje (usposabljanje, časovne analize, standardi);
# - Standardizacijo so uporabljali že v beneških ladjedelnicah
# - 1913 <NAME> - razporejanje poslov po strojih, <NAME> - tekoči trak
# - 1914 <NAME>, enačbe za napovedovanje izhoda bitk glede na števičnost in ognjeno moč nasprotnikov
# - 1915 <NAME>, osnovni obrazec za velikost naročila v teoriji zalog
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 9 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (4)
#
# - 19-- <NAME>, Markov
# - 1925 Leontijev - ravnotežje narodnega gospodarstva SZ
# - 1921-27 Borel - teorija iger
# - 1928 Janos (John) <NAME> - osnovni izrek teorije iger izrek o minimaksu
# - 1930 <NAME>, povezanost reklame in prodaje ter dohodka in prebivališča kupcev
# - 193- <NAME>, poudarek na človeških dejavnikih, ublažitev tehničnega pristopa
# - 1936 Konig - teorija grafov
# - 1938 prva skupina OR v Angliji pri RAFu
# - 1939 P.M.S. Blackettova skupina
# - 1939 Kantorovič - metoda za reševanje LP
#
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 10 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (5)
#
# - 1939-45 <NAME>, <NAME> - teorija
# - ok. 1940 Tolstoj, Kantorovič, Hitchcock - transportni problem; računalniki
# - 194- Metropolis, Ulam - Monte Carlo
# - 1947 <NAME> - LP
# - 195- <NAME> - omrežja
# - 195- <NAME> - nelinearno programiranje
# - 195- <NAME> - kazenske metode
# - 195- Bellman - dinamično programiranje
# - 195- POLARIS - CPM, PERT
# - 1950 dalje: statistične/verjetnostne metode za nadzor kakovosti, modeliranje – računalniki
# - 1950 ustanovljeno (British) Operational Research Society
# - 1952 ustanovljeno Operations Research Society of America
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 11 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1. 1. Umestitev optimizacije v TK </div>
# </div>
#
#
# ## ■ Kaj so telekomunikacije, zgodovina (6)
#
# - 1953 ustanovljen The institute of Management Science
# - 1959 ustanovljeno International Federation of ORS
# - 1960 univerzitetni študij
# - od 1960 teorija odločanja, ciljno programiranje, večkriterijska optimizacija
# - od 1970 povezave z informacijskimi sistemi – sistemi za podporo odločanja;
# - umetna inteligenca
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 12 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# # 1.2. Primeri in posebnosti OR v TK
#
# ■ Logistični primeri
#
# ■ Specifike OP v TK (1): Specifika prometa
#
# ■ Specifike OP v TK (2): Kompleksnost sistemov
#
# ■ Specifike OP v TK (3): Kriterijska funkcija in merjenje uporabnikov
#
# ■ Značilen potek reševanja problema
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 13 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
#
# ## ■ Logistični primeri
#
# - Problem: minimalni stroški prevozov hitre pošte
# - Optimizacijski prostor: model cestnega omrežja področja, ki ga pokrivamo
# - Kriterijska funkcija: stroški, ki so sestavljeni iz
# - Amortizacija vozil, servis
# - Stroški dela
# - Poraba goriva
# - Omejitve:
# - Fizične omejitve vozil in mreže
# - Zakonske omejitve
# - Ali je rešljivo?
#
#
# <img style="float:right; width:300px; margin:-200px 0 0 0;" src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/CityMap.jpg">
#
#
#
# <p style="margin-bottom:1cm;"></p><br><br><br>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 14 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
#
# ## ■ Specifike OP v TK (1): specifika prometa
#
# - Statistične lastnosti prometa
# - Poissonov model ne velja
# - Samopodobnost
# - Hurstov parameter
#
# <p>
# - Kdaj in zakaj?
# - Ko imamo večje število uporabnikov, ki brska po spletnih straneh
#
#
#
# <img style="float: right; width: 300px; margin:-150px 0 0 0;" src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/SelfSimilarTr_slo.png">
#
#
#
# <p style="margin-bottom:1cm;"><br><br><br><br><br><br><br><br><br><br></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 15 </div>
# </div><img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# ## ■ Specifike OP v TK (2): Kompleksnost sistemov
#
# - Uporabnik je človek
# - QoE
# - MOS: srednja vrednost mnenja
# - Statistična evalvacija vprašalnikov
#
# <p>
# - Sistem s povratno vezavo
# - Odmetavanje paketov
#
# <img style="float: right; width: 500px; margin:-80px 20px 10px 0;" src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/ComplexNetSys_ang.png">
#
#
# <p style="margin-bottom:1cm;"></p><br><br><br><br><br>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 16 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# ## ■ Specifike OP v TK (3): Kriterijska funkcija in merjenje uporabnikov
#
#
# - Kriterijska funkcija ni matematična funkcija, ampak učinek storitve iz stališča uporabnika
# - učinkovitost in učinki storitve so
# - slabo definirani
# - težko merljivi
# - zahtevajo zapleten uporabniške eksperimente
#
#
# - posebno poglavje
#
#
# <p style="margin-bottom:1cm;"></p><br><br><br><br><br>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 17</div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# ## ■ Značilen potek reševanja problema
#
# - Bistveno je poznavanje področja <p>
# (ang. domain knowledge)
# - Več iteracij
# - Človeški vidik: implementacija <p>
# rezultatov optimizacije je izredno naporna
#
# <img style="float: right; width: 250px; margin:-160px 20px 10px 0;"
# src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/OptimizProc_slo.png">
#
#
# <p style="margin-bottom:1cm;"><br><br><br><br><br></p><br><br><br><br><br><br><br><br><br><br>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 18 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# ## ■ Wikipedia povezave (1)
#
# <table style="width:70%" align="left">
# <tr>
# <th>The algorithm and the computational complexity (algorithm, computational complexity).
# </th>
# <th>
# http://en.wikipedia.org/wiki/Algorithm
# http://en.wikipedia.org/wiki/Computational_complexity
# http://en.wikipedia.org/wiki/Analysis_of_algorithms
# http://en.wikipedia.org/wiki/Numerical_error </th>
# </tr>
# <tr>
# <td>Graph theory (description, operations on graphs, basic graph algorithms, selected properties of graphs) and data structures.
# </td>
# <td>
# http://en.wikipedia.org/wiki/Graph_theory
# http://en.wikipedia.org/wiki/Data_structures
# </td>
# </tr>
# <tr>
# <td>Introduction to operations research and optimization. Optimization task (formulation, objective function, types of solutions).
# </td>
# <td>http://en.wikipedia.org/wiki/Operations_research
# http://en.wikipedia.org/wiki/Optimization </td>
# </tr>
# <tr>
# <td>Combinatorial optimization, linear programming and integer programming (simplex method, travel salesman problem and a knapsack problem).
# </td>
# <td>http://en.wikipedia.org/wiki/Combinatorial_optimization
# http://en.wikipedia.org/wiki/Linear_programming
# http://en.wikipedia.org/wiki/Simplex_algorithm
# http://en.wikipedia.org/wiki/Traveling_salesman_problem
# </tr>
# <tr>
# <td>Network analysis (maximal flow, minimum cost, shortest path, optimal labeling).
# </td>
# <td>http://en.wikipedia.org/wiki/Maximal_flow
# http://en.wikipedia.org/wiki/Minimum_cost_flow_problem
# http://en.wikipedia.org/wiki/Shortest_path
# </tr>
# </table>
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 19 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="display:flex;font-weight:bold;font-size:0.9em;">
# <div style="flex:1;width:50%;"> Uvod v Optimizacijo v telekomunikacijah </div>
# <div style="flex:1;width:50%;text-align:right;"> 1.2. Primeri in posebnosti OR v TK </div>
# </div>
#
#
# ## ■ Wikipedia povezave (2)
#
#
# <table style="width:70%" align="left">
# <tr>
# <th>
# Nonlinear optimization (gradient and Newton methods, constrained optimization).
# </th>
# <th>
# http://en.wikipedia.org/wiki/Nonlinear_programming
# http://en.wikipedia.org/wiki/Constraint_optimization
# http://en.wikipedia.org/wiki/Nonlinear_conjugate_gradient_method
# </th>
# </tr>
# <tr>
# <th>
# Dynamic programming and game theory.
# </th>
# <th>
# http://en.wikipedia.org/wiki/Dynamic_programming
# http://en.wikipedia.org/wiki/Game_theory
# </th>
# </tr>
# <tr>
# <th>
# Markov chains (classification of states, ergodicity).
# </th>
# <th>
# http://en.wikipedia.org/wiki/Markov_chain
# </th>
# </tr>
# <tr>
# <th>
# Time series
# </th>
# <th>
# http://en.wikipedia.org/wiki/Time_series
# http://en.wikipedia.org/wiki/Traffic_generation_model
# http://en.wikipedia.org/wiki/Autoregressive_fractionally_integrated_moving_average
# http://en.wikipedia.org/wiki/Self-similar_process
# </th>
# </tr>
# <tr>
# <th>
# Decission theory.
# </th>
# <th>
# http://en.wikipedia.org/wiki/Decision_theory
# </th>
# </tr>
# <tr>
# <th>
# Heuristic optimization methods
# </th>
# <th>
# http://en.wikipedia.org/wiki/Heuristic_algorithm
# </th>
# </tr>
# <tr>
# <th>
# Important applications in telecommunications (topology control, optimal resource allocation, optimal routing, optimal recovery after an error).
# </th>
# <th>
# http://en.wikipedia.org/wiki/Routing
# http://en.wikipedia.org/wiki/Scheduling
# </th>
# </tr>
# </table>
#
#
#
# <p style="margin-bottom:1cm;"></p>
# <div style="width:100%;text-align:right;font-weight:bold;font-size:1.2em;"> 20 </div>
# <img src="https://raw.githubusercontent.com/andrejkk/ORvTK_SlidesImgs/master/footer_full.jpg">
# -
| AKosir-OPvTK-Lec01_Intro_Slides_SLO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib as mpl
#mpl.use('pdf')
import matplotlib.pyplot as plt
import numpy as np
plt.rc('font', family='serif', serif='Times')
plt.rc('text', usetex=True)
plt.rc('xtick', labelsize=6)
plt.rc('ytick', labelsize=6)
plt.rc('axes', labelsize=6)
#axes.linewidth : 0.5
plt.rc('axes', linewidth=0.5)
#ytick.major.width : 0.5
plt.rc('ytick.major', width=0.5)
plt.rcParams['xtick.direction'] = 'in'
plt.rcParams['ytick.direction'] = 'in'
plt.rc('ytick.minor', visible=True)
#plt.style.use(r"..\..\styles\infocom.mplstyle") # Insert your save location here
# width as measured in inkscape
fig_width = 3.487
#height = width / 1.618 / 2
fig_height = fig_width / 1.618 * 2
# -
import matplotlib
matplotlib.__version__
# +
small_link_data = np.full((5, 21), 0.1355019)
small_node_data = np.full((5, 21), 0.16319444)
#print(small_link_data)
with open("failure/small-link-utilization.csv", "r") as f:
f1 = f.readlines()
for index in range(1, 11):
line = f1[index]
line = line.split(",")
for i in range(0, 5):
small_link_data[i, index-1] = line[i]
#print(line)
#print(type(f1))
with open("failure/small-node_utilization.csv", "r") as f:
f1 = f.readlines()
for index in range(1, 11):
line = f1[index]
line = line.split(",")
for i in range(0, 5):
small_node_data[i, index-1] = line[i]
#print(line)
#print(type(f1))
print(small_link_data)
print(small_node_data)
print(type(small_link_data[0,0]))
#x = np.arange(0.0, 3*np.pi , 0.1)
#plt.plot(x, np.sin(x))
#plt.show()
small_link_data = 100 * small_link_data
small_node_data = 100 * small_node_data
# +
medium_link_data = np.full((5, 21), 0.1355018958)
medium_node_data = np.full((5, 21), 0.1631944444)
#mesh3data = np.zeros((5, 11))
with open("failure20stages/Medium-Failure-20stage/no-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
#print(start_line)
#print(len(f1))
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[0, index] = float(line[3])
medium_link_data[0, index] = float(line[4])
#mesh3data[0, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/Link-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[1, index] = float(line[3])
medium_link_data[1, index] = float(line[4])
#mesh3data[1, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/LimitedReconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
start_line1 = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
start_line1 = start_line + 16
for index in range(start_line+16, len(f1)):
if f1[index].find("%Stage") >= 0:
break
else:
start_line1 = start_line1 + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[2, index] = float(line[3])
medium_link_data[2, index] = float(line[4])
#mesh3data[2, index] = int(line[1])
for index in range(0, 16):
line = f1[index+start_line1]
line = line.split(",")
medium_node_data[3, index] = float(line[3])
medium_link_data[3, index] = float(line[4])
#mesh3data[3, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/Any-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[4, index] = float(line[3])
medium_link_data[4, index] = float(line[4])
medium_link_data = 100 * medium_link_data
medium_node_data = 100 * medium_node_data
print(medium_node_data)
print(medium_link_data)
# +
large_link_data = np.zeros((5, 21))
large_node_data = np.zeros((5, 21))
with open("failure20stages/Big-Failure-20stage/no-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
#print(start_line)
#print(len(f1))
for index in range(0, 21):
line = f1[index+start_line]
line = line.split(",")
large_node_data[0, index] = float(line[3])
large_link_data[0, index] = float(line[4])
#mesh3data[0, index] = int(line[1])
with open("failure20stages/Big-Failure-20stage/Link-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 21):
line = f1[index+start_line]
line = line.split(",")
large_node_data[1, index] = float(line[3])
large_link_data[1, index] = float(line[4])
#mesh3data[1, index] = int(line[1])
with open("failure20stages/Big-Failure-20stage/LimitedReconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
start_line1 = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
start_line1 = start_line + 21
for index in range(start_line+21, len(f1)):
if f1[index].find("%Stage") >= 0:
break
else:
start_line1 = start_line1 + 1
for index in range(0, 21):
line = f1[index+start_line]
line = line.split(",")
large_node_data[2, index] = float(line[3])
large_link_data[2, index] = float(line[4])
for index in range(0, 21):
line = f1[index+start_line1]
line = line.split(",")
large_node_data[3, index] = float(line[3])
large_link_data[3, index] = float(line[4])
with open("failure20stages/Big-Failure-20stage/Any-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 21):
line = f1[index+start_line]
line = line.split(",")
large_node_data[4, index] = float(line[3])
large_link_data[4, index] = float(line[4])
large_link_data = 100 * large_link_data
large_node_data = 100 * large_node_data
print(large_node_data)
print(large_link_data)
# +
fig, ax = plt.subplots(nrows=3, ncols=2)
stage_small = np.arange(21)
stage_big = np.arange(21)
print(stage_big)
print(ax)
marker_list = ["p", "v", "x", ".", "*"]
label_list = ['No-rec', 'Link-rec', 'Lim-rec(5,0)', 'Lim-rec(5,2)', 'Any-rec']
for i in range(5):
ax[0,0].plot(stage_small, small_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[0,1].plot(stage_small, small_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,0].plot(stage_big, large_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,1].plot(stage_big, large_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
# +
fig, ax = plt.subplots(nrows=3, ncols=2)
stage = np.arange(21)
print(stage)
print(ax)
marker_list = ["p", "v", "x", ".", "*"]
label_list = ['No-rec', 'Link-rec', 'Lim-rec(5,0)', 'Lim-rec(5,2)', 'Any-rec']
for i in range(5):
ax[0,0].plot(stage, small_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[0,1].plot(stage, small_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[1,0].plot(stage, medium_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[1,1].plot(stage, medium_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,0].plot(stage, large_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,1].plot(stage, large_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(0, 2):
ax[0, i].set_xlabel('Recovery stages with 20\% substrate failures')
for i in range(0, 2):
ax[1, i].set_xlabel('Recovery stages with 30\% substrate failures')
for i in range(0, 2):
ax[2, i].set_xlabel('Recovery stages with 40\% substrate failures')
for i in range(0, 3):
ax[i,0].set_ylabel("\% of node utilization")
for i in range(0, 3):
ax[i,1].set_ylabel("\% of link utilization")
#for i in range(0, 3):
ax[0,0].legend(loc='upper center', bbox_to_anchor=(1.15, 1.3),
ncol=5, prop={'size': 5})
ax[0,0].yaxis.set_label_coords(-0.16, 0.5)
ax[0,1].yaxis.set_label_coords(-0.16, 0.5)
for i in range(1, 3):
for j in range(0, 2):
ax[i,j].yaxis.set_label_coords(-0.164, 0.5)
for i in range(0, 3):
for j in range(0, 2):
ax[i,j].xaxis.set_label_coords(0.5, -0.19)
#x1.yaxis.set_label_coords(-0.17,0.5)
ax[0, 0].set_title('(a) 20\% substrate failures', y=-0.55, fontsize=7)
ax[0, 1].set_title('(b) 20\% substrate failures', y=-0.55, fontsize=7)
ax[1, 0].set_title('(c) 30\% substrate failures', y=-0.55, fontsize=7)
ax[1, 1].set_title('(d) 30\% substrate failures', y=-0.55, fontsize=7)
ax[2, 0].set_title('(e) 40\% substrate failures', y=-0.55, fontsize=7)
ax[2, 1].set_title('(f) 40\% substrate failures', y=-0.55, fontsize=7)
mpl.pyplot.subplots_adjust(wspace = 0.3, hspace=0.6)
fig.subplots_adjust(left=.10, bottom=.11, right=.97, top=.93)
plt.show()
# +
fig.set_size_inches(fig_width, fig_height)
plt.show()
fig.savefig('heuristic-utilization1.pdf')
# +
small_link_data = np.full((5, 11), 0.1355019)
small_node_data = np.full((5, 11), 0.16319444)
#print(small_link_data)
with open("failure/small-link-utilization.csv", "r") as f:
f1 = f.readlines()
for index in range(1, 11):
line = f1[index]
line = line.split(",")
for i in range(0, 5):
small_link_data[i, index-1] = line[i]
#print(line)
#print(type(f1))
with open("failure/small-node_utilization.csv", "r") as f:
f1 = f.readlines()
for index in range(1, 11):
line = f1[index]
line = line.split(",")
for i in range(0, 5):
small_node_data[i, index-1] = line[i]
#print(line)
#print(type(f1))
print(small_link_data)
print(small_node_data)
print(type(small_link_data[0,0]))
#x = np.arange(0.0, 3*np.pi , 0.1)
#plt.plot(x, np.sin(x))
#plt.show()
small_link_data = 100 * small_link_data
small_node_data = 100 * small_node_data
# +
medium_link_data = np.full((5, 16), 0.1355018958)
medium_node_data = np.full((5, 16), 0.1631944444)
#mesh3data = np.zeros((5, 11))
with open("failure20stages/Medium-Failure-20stage/no-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
#print(start_line)
#print(len(f1))
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[0, index] = float(line[3])
medium_link_data[0, index] = float(line[4])
#mesh3data[0, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/Link-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[1, index] = float(line[3])
medium_link_data[1, index] = float(line[4])
#mesh3data[1, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/LimitedReconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
start_line1 = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
start_line1 = start_line + 16
for index in range(start_line+16, len(f1)):
if f1[index].find("%Stage") >= 0:
break
else:
start_line1 = start_line1 + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[2, index] = float(line[3])
medium_link_data[2, index] = float(line[4])
#mesh3data[2, index] = int(line[1])
for index in range(0, 16):
line = f1[index+start_line1]
line = line.split(",")
medium_node_data[3, index] = float(line[3])
medium_link_data[3, index] = float(line[4])
#mesh3data[3, index] = int(line[1])
with open("failure20stages/Medium-Failure-20stage/Any-reconfig120.csv", "r") as f:
f1 = f.readlines()
start_line = 0
for line in f1:
if line.find("%Stage") >= 0:
break
else:
start_line = start_line + 1
for index in range(0, 16):
line = f1[index+start_line]
line = line.split(",")
medium_node_data[4, index] = float(line[3])
medium_link_data[4, index] = float(line[4])
medium_link_data = 100 * medium_link_data
medium_node_data = 100 * medium_node_data
print(medium_node_data)
print(medium_link_data)
# +
fig, ax = plt.subplots(nrows=3, ncols=2)
stage = np.arange(21)
stage_small = np.arange(11)
stage_medium = np.arange(16)
stage_large = np.arange(21)
print(stage)
print(ax)
marker_list = ["p", "v", "x", ".", "*"]
label_list = ['No-rec', 'Link-rec', 'Lim-rec(5,0)', 'Lim-rec(5,2)', 'Any-rec']
for i in range(5):
ax[0,0].plot(stage_small, small_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[0,1].plot(stage_small, small_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[1,0].plot(stage_medium, medium_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[1,1].plot(stage_medium, medium_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,0].plot(stage_large, large_node_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(5):
ax[2,1].plot(stage_large, large_link_data[i], linewidth=1,
marker=marker_list[i], markersize=3, markeredgewidth=0.5,
label=label_list[i])
for i in range(0, 2):
ax[0, i].set_xlabel('Recovery stages with 20\% substrate failures')
for i in range(0, 2):
ax[1, i].set_xlabel('Recovery stages with 30\% substrate failures')
for i in range(0, 2):
ax[2, i].set_xlabel('Recovery stages with 40\% substrate failures')
for i in range(0, 3):
ax[i,0].set_ylabel("\% of node utilization")
for i in range(0, 3):
ax[i,1].set_ylabel("\% of link utilization")
#for i in range(0, 3):
ax[0,0].legend(loc='upper center', bbox_to_anchor=(1.15, 1.3),
ncol=5, prop={'size': 5})
ax[0,0].yaxis.set_label_coords(-0.16, 0.5)
ax[0,1].yaxis.set_label_coords(-0.16, 0.5)
for i in range(1, 3):
for j in range(0, 2):
ax[i,j].yaxis.set_label_coords(-0.164, 0.5)
for i in range(0, 3):
for j in range(0, 2):
ax[i,j].xaxis.set_label_coords(0.5, -0.19)
#x1.yaxis.set_label_coords(-0.17,0.5)
ax[0, 0].set_title('(a) 20\% substrate failures', y=-0.55, fontsize=7)
ax[0, 1].set_title('(b) 20\% substrate failures', y=-0.55, fontsize=7)
ax[1, 0].set_title('(c) 30\% substrate failures', y=-0.55, fontsize=7)
ax[1, 1].set_title('(d) 30\% substrate failures', y=-0.55, fontsize=7)
ax[2, 0].set_title('(e) 40\% substrate failures', y=-0.55, fontsize=7)
ax[2, 1].set_title('(f) 40\% substrate failures', y=-0.55, fontsize=7)
mpl.pyplot.subplots_adjust(wspace = 0.3, hspace=0.6)
fig.subplots_adjust(left=.10, bottom=.11, right=.97, top=.93)
for i in range(0, 3):
for j in range(0, 2):
ax[i, j].grid(lw = 0.25)
plt.show()
# +
fig.set_size_inches(fig_width, fig_height)
plt.show()
fig.savefig('heuristic-utilization1.pdf')
| python-plot/slice-restoration/.ipynb_checkpoints/compare-heuristic-utilization-20stages-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Error and Exception Handling
5/0
print("hellow")
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
try:
c = a/b
print(c)
except ZeroDivisionError:
print("Exception is caught ZeroDivisionError")
print('Hello Error')
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
print(c)
print('Hello World')
# -
a = [1,2,3]
try:
print(a[8])
except IndexError:
print('Yeah! I Have handled that error')
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
print(c)
print(d[2])
except ZeroDivisionError:
print('Exception is caught ZeroDivisionError')
print('Hello World')
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
try:
c = a/b
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print("Else " +str(c))
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
print(e)
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print(c)
# +
print("First")
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
# e = d[5]
except ZeroDivisionError:
print("Handled ZeroDivisionError")
else:
print(c)
print("last")
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
except ZeroDivisionError: #Type Z and press tab to get list of Exception
print("Exception is caught ZeroDivisionError")
except IndexError: #Type I and press tab to get list of Exception
print("Exception is caught is list index out of range")
else:
print(c)
print(e)
# +
d = []
try:
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
e = d[5]
except: # here we did not explain the type of exception. Genral exception will
# be used. means it will catch every kind of exception
print("koi si bhi Exception is caught")
else:
print(c)
# +
# a =int(input("Enter a number"))
# b =int(input("Enter a number"))
d = []
try:
'''built in message print karna hy exception ka to {except Exception as e:}
is me jistrah ki acception aegi wo uska message khud he utha k laega or print karega
is code ko khud se run kare or exception raise kar k dekhen'''
a =int(input("Enter a number"))
b =int(input("Enter a number"))
c = a/b
ee = d[5]
except Exception as e:
print("Handled Error == " +str(e))
else:
print(c)
print('HelloWorld')
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
"""yahan ek humne apni message likha hy or ek built in use kia hy"""
except ZeroDivisionError:
print("Handel ZeroDivisionError")
except Exception as e:
print("Handle Error == " +str(e))
else:
print(c)
# +
a =int(input("Enter a number"))
b =int(input("Enter a number"))
d = []
try:
c = a/b
e = d[5]
except ZeroDivisionError:
print("Handel ZeroDivisionError")
else:
print(c)
finally:
print("Finally will always run")
# -
# + active=""
# ###Rasing our own exceptions
#
# We can rasie exception by ourselves. ony any condition we defined like if a>10 raise exception
# like if user inputs value 10 raise exception bla bla bla bla......
# +
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise Exception("Age can not be greater then 80 and less then 16")
self.name = name
self.age = age
# +
age = int(input("Enter students age: "))
st = Student("ALi",age)
print(st.age)
# + active=""
# #Catching the exception raised by ourself###
# Exception catch karne ki liye wahi kam karenge k jahan se raise hone k dar ho usko try me
# dal den. or except me catch kara den. Go through the code......
# +
class Student():
def __init__(self,name,age):
if age > 80:
raise Exception("Age can not be greater then 80")
self.name = name
self.age = age
# +
'''ab humen try except laga diya hy code crash nahi hoga or izzat batch jaegi hahahahah'''
try:
age = int(input("Enter students age: "))
st = Student("Hello",age) # send different ages and see the result
except Exception as e:
print("Exception "+str(e))
else :
print(st.age)
# + active=""
#
# #####################################################
# #######Importantttttttttttttttttttttttttttttttt######
# #####################################################
# + active=""
# Making our class of exception::::::::
# Raising exception ourselves::::::
#
# Last code me we used python exception class
# Now we will make our own exception class and will used in the code.
# Dont panic guls and guys...
#
#
# +
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise StudentAgeException("Age can not be greater then 80 and less then 16")
# we used here own class of exceptio
self.name = name
self.age = age
'''All the exception raised are derived from a class Exception by default
We can make our own class of exception and can derived(inherit) it from parent
Exception class like this:'''
class StudentAgeException(Exception):
pass
# +
age = int(input("Enter students age: "))
st = Student("Hello",age)
# abhi it is creating exception that is not handled
print(st.age)
####Note###
'''exception raise karane k liye humne Exception python ki general wali use nahiiiiiii ki hy
abhi code crash hogaya hy kyu k humne abhi exception catch nahiiiiiiiiiiiiiiiiii ki hy'''
# + active=""
#
# #################################################################################
# Raising and catching our own exception [[[[defined in our own exception class]]]]
# Raising and catching our own exception [[[[defined in our own exception class]]]]
# Raising and catching our own exception [[[[defined in our own exception class]]]]
# Raising and catching our own exception [[[[defined in our own exception class]]]]
# Raising and catching our own exception [[[[defined in our own exception class]]]]
# #################################################################################
# +
class Student():
def __init__(self,name,age):
if age > 80 or age <16:
raise StudentAgeException("Age can not be greater then 80 and less then 16")
self.name = name
self.age = age
class StudentAgeException(Exception):
pass
try:
age = int(input("Enter students age: "))
st = Student("Hello",age)
except StudentAgeException as e:
print("StudentAgeException == "+str(e))
else :
print(st.age)
# -
| Q1/Notebooks files/exceptions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import json
import matplotlib.pyplot as plt
data = []
with open('DeltaDump2015All.txt') as f:
for line in f:
data.append(json.loads(line))
d=dict()
for item in data:
if(item['uid']!="none" and item['uid'] not in d.keys()):
ud=item['uid']
c=["No"]
t=list()
for time in data:
if(time['uid']==ud and'id' in time['edata']['eks'].keys()):
s=time['edata']['eks']['id']
if("suitcase" in s):
c[0]="Yes"
if(time['uid']==ud):
t.append(time['ts'])
sorted(t)
hr=t[0]
c.append(int(hr[11:13]))
d[ud]=c
td=d.values()
rewards=dict()
for tim in td:
if(tim[0]=="No"):
if (tim[1] not in rewards):
rewards[tim[1]]=[0, 1]
else:
rewards[tim[1]][1]+=1
elif(tim[0]=="Yes"):
if (tim[1] not in rewards):
rewards[tim[1]]=[1, 1]
else:
rewards[tim[1]][1]+=1
rewards[tim[1]][0]+=1
plt.title("Percentage of users that access rewards by hour")
plt.xlabel("Time of day")
plt.ylabel("Percentage of users")
plt.axis([0, 24, 0, 100])
p=list()
for x, y in rewards.values():
p.append((float(x)/float(y))*100)
plt.plot(rewards.keys(), p)
plt.show()
| platform-scripts/Takeoff/Take-Off_Userpercentage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (PySpark)
# language: python
# name: python3
# ---
# +
import os,sys,re
# sys.path.insert(0, "/home/wenhao71/.pylib/3")
# %load_ext autoreload
# %autoreload 2
# +
from typing import *
import wfdb
import scipy
import json
from io import StringIO
from scipy.io import loadmat
from glob import glob
from copy import deepcopy
from random import shuffle
from functools import reduce
from tqdm import tqdm
from scipy.signal import resample, resample_poly
import torch
from itertools import product
from easydict import EasyDict as ED
from data_reader import CPSC2020Reader as CR
from dataset import CPSC2020
from cfg import TrainCfg, ModelCfg, PreprocCfg
from utils import list_sum
from torch_ecg.torch_ecg.models.nets import (
BCEWithLogitsWithClassWeightLoss,
default_collate_fn as collate_fn,
)
from torch.utils.data import DataLoader
# -
pwd
db_dir = '/media/cfs/wenhao71/data/CPSC2020/TrainingSet/'
dr = CR(db_dir)
dr.all_records
# +
# for rec in dr.all_records:
# data = dr.load_data(rec, keep_dim=False)
# print(f"{rec} has max abs value {np.max(np.abs(data))} (max positive values {np.max(data)}, max negative values {np.min(data)})")
# +
# for rec in dr.all_records:
# ann = dr.load_ann(rec)
# spb = ann["SPB_indices"]
# pvc = ann["PVC_indices"]
# if len(np.diff(spb)) > 0:
# print(f"{rec}: min dist among SPB = {np.min(np.diff(spb))}")
# if len(np.diff(pvc)) > 0:
# print(f"{rec}: min dist among PVC = {np.min(np.diff(pvc))}")
# diff = [s-p for s,p in product(spb, pvc)]
# if len(diff) > 0:
# print(f"{rec}: min dist between SPB and PVC = {np.min(np.abs(diff))}")
# -
rec = dr.all_records[1]
dr.load_ann(rec)
dr.locate_premature_beats(rec, "pvc")
# +
# dr.plot(rec,sampfrom=1459472,ticks_granularity=2,sampto=1469472)
# -
dr.plot(rec,sampfrom=0,ticks_granularity=2,sampto=4000)
ModelCfg
dataset_cfg = ED(TrainCfg.copy())
dataset_cfg.model_name = "seq_lab"
dataset_cfg.classes = ModelCfg[dataset_cfg.model_name].classes
dataset_cfg.class_map = ModelCfg[dataset_cfg.model_name].class_map
PreprocCfg
ds = CPSC2020(dataset_cfg, training=True)
len(list_sum([v for v in ds.all_segments.values()]))
for rec, l_seg in ds.all_segments.items():
print(rec, len(l_seg))
# +
# with open(ds.segments_json, "w") as f:
# json.dump(ds.all_segments, f)
# -
ds._load_seg_seq_lab(ds.segments[0]).shape
ds.__DEBUG__=True
data, label = ds[14000]
np.where(label[:,1]>0.5)[0]*8/400
# +
# with open("/media/cfs/wenhao71/data/CPSC2020/TrainingSet/segments_backup2/crnn_segments.json", "r") as f:
# old_segs = json.load(f)
# for rec, l_seg in old_segs.items():
# print(rec, len(l_seg))
# -
train_loader = DataLoader(
dataset=ds,
batch_size=32,
shuffle=True,
num_workers=8,
pin_memory=True,
drop_last=False,
collate_fn=collate_fn,
)
for signals, labels in train_loader:
break
signals.shape
labels.shape
pred = torch.rand((32, 500, 2))
loss = torch.nn.BCEWithLogitsLoss()
loss(pred, labels)
ds.disable_data_augmentation()
ds.__DEBUG__=True
ds[1]
ds.segments[10004]
ds[10004]
np.std(ds[100095][0]), np.mean(ds[100095][0])
ds._get_seg_ampl(ds[10003])
# +
# dr.plot(rec="A02", sampfrom=169*seg_len, sampto=186*seg_len)
# +
# flat_segs = {rec:[] for rec in dr.all_records}
# valid_segs = {rec:[] for rec in dr.all_records}
# for i, rec in enumerate(ds.reader.all_records):
# for idx, seg in enumerate(ds.all_segments[rec]):
# seg_data = ds._load_seg_data(seg)
# if ds._get_seg_ampl(seg_data) < 0.1:
# flat_segs[rec].append(seg)
# else:
# valid_segs[rec].append(seg)
# print(f"{idx+1}/{len(ds.all_segments[rec])} @ {i+1}/{len(ds.reader.all_records)}", end="\r")
# +
# valid_segs
# +
# len(list_sum([v for v in valid_segs.values()]))
# +
# len(list_sum([v for v in flat_segs.values()]))
# +
# len(list_sum([v for v in ds.all_segments.values()]))
# +
# with open(ds.segments_json, "w") as f:
# json.dump(valid_segs, f)
# -
ds.__DEBUG__ = True
ds.disable_data_augmentation()
all_segs = list_sum([v for k,v in ds.all_segments.items()])
# +
mean分布, std分布 = [], []
for i, (rec, l_seg) in enumerate(ds.all_segments.items()):
for idx, seg in enumerate(l_seg):
data = ds._load_seg_data(seg)
mean分布.append(np.mean(data))
std分布.append(np.std(data))
# print(f"{idx+1}/{len(l_seg)} @ {i+1}/{len(ds.all_segments)}", end="\r", flush=True)
# -
len(mean分布), len(std分布)
plt.hist(mean分布)
np.median(mean分布), np.mean(mean分布)
plt.hist(std分布)
np.median(std分布), np.mean(std分布)
# +
# check = []
# for i, (rec, l_seg) in enumerate(ds.all_segments.items()):
# for idx, seg in enumerate(l_seg):
# data = ds._load_seg_data(seg)
# if np.std(data) < 0.002:
# check.append(seg)
# -
std分布_sorted = sorted(std分布)
std分布_sorted[:10]
np.sum(np.array(std分布)<0.03)
check = [all_segs[i] for i in np.where(np.array(std分布)<0.03)[0]]
no = 0
print(check[no])
ds.plot_seg(check[no],ticks_granularity=2)
no += 1
print(no)
ds.plot_seg(ds.all_segments["A09"][2911])
len(list_sum([v for v in valid_segs.values()]))
with open(ds.segments_json, "w") as f:
json.dump(valid_segs, f)
data = ds._load_seg_data("S08_0001043")
dr.plot("A01", data=data/np.std(data)*0.25, ticks_granularity=2)
ds.plot_seg("S08_0001043")
ecg_denoise(ds._load_seg_data(ds.segments[0]),400,{"ampl_min":0.15}) == [[0,4000]]
from scipy.ndimage import median_filter
data = ds._load_seg_data("S06_0005081")
from signal_processing.ecg_denoise import ecg_denoise
# ecg_denoise(data,400,{})
itvs = ecg_denoise(dr.load_data("A02"),400,{"ampl_min":0.15})
sum([itv[1]-itv[0] for itv in itvs])
ecg_denoise(ds._load_seg_data())
no = 370
ds._load_seg_data(ds.segments[no]).shape
plt.plot(ds._load_seg_data(ds.segments[no]))
ds._load_seg_label(ds.segments[no])
ds._load_seg_beat_ann(ds.segments[no])
for seg in ds.segments:
lb = ds._load_seg_label(seg)
if (lb[1:]>0).any():
print(seg)
ds.plot_seg("S01_0001781", ticks_granularity=2)
# +
# ds._slice_one_record(rec=rec,verbose=2)
# +
# ds._slice_data(verbose=2)
# -
# +
# ds._preprocess_one_record("A01",config=PreprocCfg)
# -
ds.all_segments
pd = loadmat(os.path.join(ds.preprocess_dir,"A01-bandpass.mat"))
pd_rpeaks = loadmat(os.path.join(ds.rpeaks_dir, "A01-bandpass.mat"))
pd['ecg'].squeeze()
pd_rpeaks['rpeaks'].squeeze()
rpeaks = pd_rpeaks['rpeaks'].squeeze()[np.where(pd_rpeaks['rpeaks'].squeeze()<400*60*3)]
# dr.plot("A01", data=pd['ecg'].squeeze()[:400*60*3], rpeak_inds=rpeaks,ticks_granularity=2)
dr.plot("A01", rpeak_inds=rpeaks,ticks_granularity=2)
dr.load_ann("A01")
ds._slice_one_record("A01", verbose=2)
from signal_processing.ecg_rpeaks_dl import seq_lab_net_detect
data = dr.load_data(rec,keep_dim=False)
data.shape
rpeaks = seq_lab_net_detect(data[:30*60*400], 400, verbose=2)
rpeaks
dr.plot(rec, data[:3*60*400], rpeak_inds=rpeaks, ticks_granularity=2)
np.ones((20,20))[0,:5]
data = dr.load_data(rec, sampfrom=0, sampto=4000, keep_dim=False)
data.shape
diff = np.max(data) - np.min(data)
bw = gen_baseline_wander(4000,400,[0.33, 0.1, 0.05, 0.01], diff*np.array([0.02, 0.04, 0.07, 0.1]), 0, diff*0.01)
bw.shape
fig,ax = plt.subplots(figsize=(20,6))
ax.plot(bw)
ax.plot(data)
ax.plot(data+bw)
hehe = np.ones((20,4))
hehe[np.array([2,3,4,5]),...]
data.shape
data_rsmp = scipy.signal.resample_poly(data, up=51, down=50)
fig,ax = plt.subplots(figsize=(20,6))
ax.plot(data_rsmp)
ax.plot(data)
# ?scipy.signal.resample
# ?np.append
hehe = np.ones((20,20))
for _ in range(10):
hehe = np.append(hehe, np.zeros((1,20)), axis=0)
# ?hehe.reshape
np.ones((10,)).reshape((1,-1)).shape
| inspect_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 7: Handy Haversacks
# You land at the regional airport in time for your next flight. In fact, it looks like you'll even have time to grab some food: all flights are currently delayed due to issues in luggage processing.
#
# Due to recent aviation regulations, many rules (your puzzle input) are being enforced about bags and their contents; bags must be color-coded and must contain specific quantities of other color-coded bags. Apparently, nobody responsible for these regulations considered how long they would take to enforce!
#
# For example, consider the following rules:
#
# * light red bags contain 1 bright white bag, 2 muted yellow bags.
# * dark orange bags contain 3 bright white bags, 4 muted yellow bags.
# * bright white bags contain 1 shiny gold bag.
# * muted yellow bags contain 2 shiny gold bags, 9 faded blue bags.
# * shiny gold bags contain 1 dark olive bag, 2 vibrant plum bags.
# * dark olive bags contain 3 faded blue bags, 4 dotted black bags.
# * vibrant plum bags contain 5 faded blue bags, 6 dotted black bags.
# * faded blue bags contain no other bags.
# * dotted black bags contain no other bags.
#
# These rules specify the required contents for 9 bag types. In this example, every faded blue bag is empty, every vibrant plum bag contains 11 bags (5 faded blue and 6 dotted black), and so on.
#
# You have a shiny gold bag. If you wanted to carry it in at least one other bag, how many different bag colors would be valid for the outermost bag? (In other words: how many colors can, eventually, contain at least one shiny gold bag?)
#
# In the above rules, the following options would be available to you:
#
# * A bright white bag, which can hold your shiny gold bag directly.
# * A muted yellow bag, which can hold your shiny gold bag directly, plus some other bags.
# * A dark orange bag, which can hold bright white and muted yellow bags, either of which could then hold your shiny gold bag.
# * A light red bag, which can hold bright white and muted yellow bags, either of which could then hold your shiny gold bag.
#
# So, in this example, the number of bag colors that can eventually contain at least one shiny gold bag is 4.
#
# How many bag colors can eventually contain at least one shiny gold bag? (The list of rules is quite long; make sure you get all of it.)
with open("input.txt","r") as f:
input_data = [x.split(" contain ") for x in f.read().split("\n")]
def remove_end(x):
if x.endswith("bags"):
return x[:-5]
elif x.endswith("bag"):
return x[:-4]
return None
def construct_bag_dict(input_data):
bag_dict = dict()
for input_row in input_data:
key = remove_end(input_row[0])
bag_dict[key] = dict()
values = [remove_end(v) for v in input_row[1][:-1].split(", ")]
for v in values:
value_ele = v.split(" ")
if value_ele[0] != "no":
number = int(value_ele[0])
color = " ".join(value_ele[1:])
bag_dict[key][color] = number
return bag_dict
bag_dict = construct_bag_dict(input_data)
def find_shiny_gold_in_x(bag_dict, x):
if len(bag_dict[x]) == 0:
return x == "shiny gold"
result = 0
for k, v in bag_dict[x].items():
if k == "shiny gold":
result = result + v
result = result + find_shiny_gold_in_x(bag_dict, k) * v
return result
sum([find_shiny_gold_in_x(bag_dict, x) > 0 for x in bag_dict])
# # Part Two
# It's getting pretty expensive to fly these days - not because of ticket prices, but because of the ridiculous number of bags you need to buy!
#
# Consider again your shiny gold bag and the rules from the above example:
#
# * faded blue bags contain 0 other bags.
# * dotted black bags contain 0 other bags.
# * vibrant plum bags contain 11 other bags: 5 faded blue bags and 6 dotted black bags.
# * dark olive bags contain 7 other bags: 3 faded blue bags and 4 dotted black bags.
#
# So, a single shiny gold bag must contain 1 dark olive bag (and the 7 bags within it) plus 2 vibrant plum bags (and the 11 bags within each of those): 1 + 1*7 + 2 + 2*11 = 32 bags!
#
# Of course, the actual rules have a small chance of going several levels deeper than this example; be sure to count all of the bags, even if the nesting becomes topologically impractical!
#
# Here's another example:
#
# * shiny gold bags contain 2 dark red bags.
# * dark red bags contain 2 dark orange bags.
# * dark orange bags contain 2 dark yellow bags.
# * dark yellow bags contain 2 dark green bags.
# * dark green bags contain 2 dark blue bags.
# * dark blue bags contain 2 dark violet bags.
# * dark violet bags contain no other bags.
#
# In this example, a single shiny gold bag must contain 126 other bags.
#
# How many individual bags are required inside your single shiny gold bag?
#
#
def find_bag_in_x(bag_dict, x):
if len(bag_dict[x]) == 0:
return 0
return sum([(find_bag_in_x(bag_dict, k) + 1) * v for k, v in bag_dict[x].items()])
find_bag_in_x(bag_dict, "shiny gold")
| 2020/Day07/Day7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fitting the distribution of heights data
# ## Instructions
#
# In this assessment you will write code to perform a steepest descent to fit a Gaussian model to the distribution of heights data that was first introduced in *Mathematics for Machine Learning: Linear Algebra*.
#
# The algorithm is the same as you encountered in *Gradient descent in a sandpit* but this time instead of descending a pre-defined function, we shall descend the $\chi^2$ (chi squared) function which is both a function of the parameters that we are to optimise, but also the data that the model is to fit to.
#
# ## How to submit
#
# Complete all the tasks you are asked for in the worksheet. When you have finished and are happy with your code, press the **Submit Assingment** button at the top of this notebook.
#
# ## Get started
# Run the cell below to load dependancies and generate the first figure in this worksheet.
# Run this cell first to load the dependancies for this assessment,
# and generate the first figure.
from readonly.HeightsModule import *
# ## Background
# If we have data for the heights of people in a population, it can be plotted as a histogram, i.e., a bar chart where each bar has a width representing a range of heights, and an area which is the probability of finding a person with a height in that range.
# We can look to model that data with a function, such as a Gaussian, which we can specify with two parameters, rather than holding all the data in the histogram.
#
# The Gaussian function is given as,
# $$f(\mathbf{x};\mu, \sigma) = \frac{1}{\sigma\sqrt{2\pi}}\exp\left(-\frac{(\mathbf{x} - \mu)^2}{2\sigma^2}\right)$$
#
# The figure above shows the data in orange, the model in magenta, and where they overlap in green.
# This particular model has not been fit well - there is not a strong overlap.
#
# Recall from the videos the definition of $\chi^2$ as the squared difference of the data and the model, i.e $\chi^2 = |\mathbf{y} - f(\mathbf{x};\mu, \sigma)|^2$. This is represented in the figure as the sum of the squares of the pink and orange bars.
#
# Don't forget that $\mathbf{x}$ an $\mathbf{y}$ are represented as vectors here, as these are lists of all of the data points, the |*abs-squared*|${}^2$ encodes squaring and summing of the residuals on each bar.
#
# To improve the fit, we will want to alter the parameters $\mu$ and $\sigma$, and ask how that changes the $\chi^2$.
# That is, we will need to calculate the Jacobian,
# $$ \mathbf{J} = \left[ \frac{\partial ( \chi^2 ) }{\partial \mu} , \frac{\partial ( \chi^2 ) }{\partial \sigma} \right]\;. $$
#
# Let's look at the first term, $\frac{\partial ( \chi^2 ) }{\partial \mu}$, using the multi-variate chain rule, this can be written as,
# $$ \frac{\partial ( \chi^2 ) }{\partial \mu} = -2 (\mathbf{y} - f(\mathbf{x};\mu, \sigma)) \cdot \frac{\partial f}{\partial \mu}(\mathbf{x};\mu, \sigma)$$
# With a similar expression for $\frac{\partial ( \chi^2 ) }{\partial \sigma}$; try and work out this expression for yourself.
#
# The Jacobians rely on the derivatives $\frac{\partial f}{\partial \mu}$ and $\frac{\partial f}{\partial \sigma}$.
# Write functions below for these.
# PACKAGE
import matplotlib.pyplot as plt
import numpy as np
# +
# GRADED FUNCTION
# This is the Gaussian function.
def f (x,mu,sig) :
return np.exp(-(x-mu)**2/(2*sig**2)) / np.sqrt(2*np.pi) / sig
# Next up, the derivative with respect to μ.
# If you wish, you may want to express this as f(x, mu, sig) multiplied by chain rule terms.
# === COMPLETE THIS FUNCTION ===
def dfdmu (x,mu,sig) :
return f(x, mu, sig) * (x-mu)/sig**2
# Finally in this cell, the derivative with respect to σ.
# === COMPLETE THIS FUNCTION ===
def dfdsig (x,mu,sig) :
return -f(x, mu, sig)/sig + f(x, mu, sig) * (x-mu)**2/sig**3
# -
# Next recall that steepest descent shall move around in parameter space proportional to the negative of the Jacobian,
# i.e., $\begin{bmatrix} \delta\mu \\ \delta\sigma \end{bmatrix} \propto -\mathbf{J} $, with the constant of proportionality being the *aggression* of the algorithm.
#
# Modify the function below to include the $\frac{\partial ( \chi^2 ) }{\partial \sigma}$ term of the Jacobian, the $\frac{\partial ( \chi^2 ) }{\partial \mu}$ term has been included for you.
# +
# GRADED FUNCTION
# Complete the expression for the Jacobian, the first term is done for you.
# Implement the second.
# === COMPLETE THIS FUNCTION ===
def steepest_step (x, y, mu, sig, aggression) :
J = np.array([
-2*(y - f(x,mu,sig)) @ dfdmu(x,mu,sig),
-2*(y - f(x,mu,sig)) @ dfdsig(x,mu,sig) # Replace the ??? with the second element of the Jacobian.
])
step = -J * aggression
return step
# -
# ## Test your code before submission
# To test the code you've written above, run all previous cells (select each cell, then press the play button [ ▶| ] or press shift-enter).
# You can then use the code below to test out your function.
# You don't need to submit these cells; you can edit and run them as much as you like.
# +
# First get the heights data, ranges and frequencies
x,y = heights_data()
# Next we'll assign trial values for these.
mu = 155 ; sig = 6
# We'll keep a track of these so we can plot their evolution.
p = np.array([[mu, sig]])
# Plot the histogram for our parameter guess
histogram(f, [mu, sig])
# Do a few rounds of steepest descent.
for i in range(50) :
dmu, dsig = steepest_step(x, y, mu, sig, 2000)
mu += dmu
sig += dsig
p = np.append(p, [[mu,sig]], axis=0)
# Plot the path through parameter space.
contour(f, p)
# Plot the final histogram.
histogram(f, [mu, sig])
# -
# Note that the path taken through parameter space is not necesarily the most direct path, as with steepest descent we always move perpendicular to the contours.
| Multivariate Calculus/(Week-6)Fitting+the+distribution+of+heights+data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import cv2
import numpy as np
import math
import matplotlib.pyplot as plt
import os
import sys
import itertools
from os import listdir
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.neighbors import KNeighborsClassifier
from sklearn.decomposition import PCA
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.svm import LinearSVC
import time
from sklearn.linear_model import LogisticRegression, Lasso
num_samples = 1000
# Function to load the images given the folder name
def load_path(path):
imgs = []
files = listdir('./MIO-TCD-Classification/train/{}/'.format(path))
img_counter = 0
for img in files:
img = cv2.cvtColor(cv2.imread('./MIO-TCD-Classification/train/{}/{}'.format(path,img)), cv2.COLOR_BGR2GRAY)
img = cv2.resize(img, (64,64))
imgs.append(img)
img_counter += 1
if(img_counter==num_samples): break
return imgs
articulated_truck = load_path('articulated_truck')
background = load_path('background')
bicycle = load_path('bicycle')
bus = load_path('bus')
car = load_path('car')
motorcycle = load_path('motorcycle')
non_motorized_vehicle = load_path('non-motorized_vehicle')
pedestrian = load_path('pedestrian')
pickup_truck = load_path('pickup_truck')
single_unit_truck = load_path('single_unit_truck')
work_van = load_path('work_van')
X = articulated_truck + background + bicycle + bus + car + motorcycle + non_motorized_vehicle + pedestrian + pickup_truck + single_unit_truck + work_van
y = ['articulated_truck']*num_samples + ['background']*num_samples + ['bicycle']*num_samples + ['bus']*num_samples + ['car']*num_samples + ['motorcycle']*num_samples + ['non_motorized_vehicle']*num_samples + ['pedestrian']*num_samples + ['pickup_truck']*num_samples + ['single_unit_truck']*num_samples + ['work_van']*num_samples
# Function to compute HoG features of a list of images
def compute_HoG(imgs, cell_size, block_size, nbins, h, w):
img_features = []
hog = cv2.HOGDescriptor(_winSize=((w // cell_size[1]) * cell_size[1],
(h // cell_size[0]) * cell_size[0]),
_blockSize=(block_size[1] * cell_size[1],
block_size[0] * cell_size[0]),
_blockStride=(cell_size[1], cell_size[0]),
_cellSize=(cell_size[1], cell_size[0]),
_nbins=nbins)
n_cells = (h // cell_size[0], w // cell_size[1])
for img in imgs:
hog_feats = hog.compute(img)\
.reshape(n_cells[1] - block_size[1] + 1,
n_cells[0] - block_size[0] + 1,
block_size[0], block_size[1], nbins) \
.transpose((1, 0, 2, 3, 4)) # index blocks by rows first
# computation for BlockNorm
gradients = np.full((n_cells[0], n_cells[1], nbins), 0, dtype=float)
cell_count = np.full((n_cells[0], n_cells[1], 1), 0, dtype=int)
for off_y in range(block_size[0]):
for off_x in range(block_size[1]):
gradients[off_y:n_cells[0] - block_size[0] + off_y + 1,
off_x:n_cells[1] - block_size[1] + off_x + 1] += \
hog_feats[:, :, off_y, off_x, :]
cell_count[off_y:n_cells[0] - block_size[0] + off_y + 1,
off_x:n_cells[1] - block_size[1] + off_x + 1] += 1
# Average gradients
gradients /= cell_count
img_features.append(gradients)
return img_features
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.15, random_state=69)
c_size = (4,4)
b_size = (4,4)
bins = 8
height = 64
width = 64
HoG_feat = compute_HoG(X_train, c_size, b_size, bins, height, width)
# -
bin = 4
test = HoG_feat[0]
plt.pcolor(test[:, :, bin])
plt.gca().invert_yaxis()
plt.gca().set_aspect('equal', adjustable='box')
plt.title("HOG bin = 4")
plt.colorbar()
plt.show()
# Part 4
then = time.clock()
clf = LinearSVC()
HoG_feat = np.array(HoG_feat)
clf.fit(HoG_feat.reshape(HoG_feat.shape[0],-1),y_train)
now = time.clock()
print(now - then)
then = time.clock()
HoG_feat_test = compute_HoG(X_test, c_size, b_size, bins, height, width)
HoG_feat_test = np.array(HoG_feat_test)
pred = clf.predict(HoG_feat_test.reshape(HoG_feat_test.shape[0],-1))
now = time.clock()
print(now - then)
print(classification_report(y_test,pred))
# +
# Adapted from sklearn COnfusion Matrix tutorial
# https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html#sphx-glr-auto-examples-model-selection-plot-confusion-matrix-py
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.figure(figsize = (12,12))
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
#print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, pred)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
# plt.figure()
# plot_confusion_matrix(cnf_matrix, classes = list(set(y_test)),
# title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes = list(set(y_test)), normalize=True,
title='Normalized confusion matrix')
plt.show()
# -
then = time.clock()
clf = LogisticRegression(random_state=0, solver='lbfgs',multi_class='multinomial')
clf.fit(HoG_feat.reshape(HoG_feat.shape[0],-1),y_train)
now = time.clock()
print(now - then)
then = time.clock()
HoG_feat_test = compute_HoG(X_test, c_size, b_size, bins, height, width)
HoG_feat_test = np.array(HoG_feat_test)
pred = clf.predict(HoG_feat_test.reshape(HoG_feat_test.shape[0],-1))
now = time.clock()
print(now - then)
print(classification_report(y_test,pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, pred)
np.set_printoptions(precision=2)
plot_confusion_matrix(cnf_matrix, classes = list(set(y_test)), normalize=True,
title='Normalized confusion matrix')
# +
from sklearn.linear_model import LogisticRegression
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
clf1 = LogisticRegression(solver='lbfgs', max_iter=1000, random_state=123)
clf2 = RandomForestClassifier(n_estimators=100, random_state=123)
clf3 = GaussianNB()
X = (HoG_feat.reshape(HoG_feat.shape[0],-1))
y = y_train
eclf = VotingClassifier(estimators=[('lr', clf1), ('rf', clf2), ('gnb', clf3)],
voting='soft',
weights=[6, 5, 1])
# predict class probabilities for all classifiers
probas = [c.fit(X, y).predict_proba(X) for c in (clf1, clf2, clf3, eclf)]
# get class probabilities for the first sample in the dataset
class1_1 = [pr[0, 0] for pr in probas]
class2_1 = [pr[0, 1] for pr in probas]
# plotting
N = 4 # number of groups
ind = np.arange(N) # group positions
width = 0.35 # bar width
fig, ax = plt.subplots()
# bars for classifier 1-3
p1 = ax.bar(ind, np.hstack(([class1_1[:-1], [0]])), width,
color='green', edgecolor='k')
p2 = ax.bar(ind + width, np.hstack(([class2_1[:-1], [0]])), width,
color='lightgreen', edgecolor='k')
# bars for VotingClassifier
p3 = ax.bar(ind, [0, 0, 0, class1_1[-1]], width,
color='blue', edgecolor='k')
p4 = ax.bar(ind + width, [0, 0, 0, class2_1[-1]], width,
color='steelblue', edgecolor='k')
# plot annotations
plt.axvline(2.8, color='k', linestyle='dashed')
ax.set_xticks(ind + width)
ax.set_xticklabels(['LogisticRegression\nweight 1',
'GaussianNB\nweight 1',
'RandomForestClassifier\nweight 5',
'VotingClassifier\n(average probabilities)'],
rotation=40,
ha='right')
plt.ylim([0, 1])
plt.title('Class probabilities for sample 1 by different classifiers')
plt.legend([p1[0], p2[0]], ['class 1', 'class 2'], loc='upper left')
plt.tight_layout()
plt.show()
# -
then = time.clock()
HoG_feat_test = compute_HoG(X_test, c_size, b_size, bins, height, width)
HoG_feat_test = np.array(HoG_feat_test)
pred = clf.predict(HoG_feat_test.reshape(HoG_feat_test.shape[0],-1))
now = time.clock()
print(now - then)
print(classification_report(y_test,pred))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, pred)
np.set_printoptions(precision=2)
plot_confusion_matrix(cnf_matrix, classes = list(set(y_test)), normalize=True,
title='Normalized confusion matrix')
| Final Project/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Cells with personal data as part of a string or input values have either been scrubbed and replaced with "ADDRESS"
## or deleted
# -
import json
import pandas as pd
from collections import Counter
from datetime import date, time, datetime
from matplotlib import pyplot as plt
day = date.today().strftime("%B %Y")
# months = ['JANUARY', 'FEBRUARY', 'MARCH', 'APRIL', 'MAY', 'JUNE', 'JULY', 'AUGUST',
# 'SEPTEMBER', 'OCTOBER', 'NOVEMBER', 'DECEMBER']
home_count=[]
work_count1=[]
work_count2=[]
full_year=[]
full_year=[]
for month in months:
f = open(f'C:/Users/jspit/OneDrive/Documents/Takeout/Location History/Semantic Location History/2020/2020_{month}.json')
data=json.load(f)
address_list=[]
for items in data.values():
for item in items:
for k in item.values():
try:
address_list.append(k['location']['address'].split('\n')[0])
except:
None
loc_count = dict(Counter(address_list))
print(loc_count)
full_year.append({"Month": month, "Locations": loc_count})
try:
home_count.append(loc_count['ADDRESS'])
work_count1.append(loc_count['ADDRESS'])
work_count2.append(loc_count['ADDRESS'])
except:
work_count1.append(0)
work_count2.append(0)
print(work_count1)
print(loc_count)
print(full_year)
# +
df = pd.json_normalize(full_year)
df.fillna(0, inplace=True)
df.reset_index()
df.set_index('Month', inplace=True)
for column, data in df.iteritems():
if not any(i > 2 for i in data.values):
del df[column]
df.astype(int)
for column, data in df.iteritems():
df.rename({column: column.strip('Locations.')}, inplace=True)
df
pd.set_option('display.max_columns',25)
df
# -
plt.bar(months, df_places['Vons'], label='Vons')
plt.bar(months, df_places['Friend House'], label="friend's house")
plt.bar(months, df_places['Trader Joes'], label = "Trader Joes")
plt.bar(months, df_places['Hiking trail'], label='Hiking Trail')
plt.grid(True)
plt.xticks(months, rotation=30)
plt.title('Frequently Visited Places - 2020')
plt.ylabel('Frequency Count')
plt.legend()
# +
df = pd.json_normalize(full_year)
dfx = df.loc[:,['Month', 'Locations.ADDRESS', 'Locations.ADDRESS','Locations.ADDRESS','Locations.ADDRESS']]
dfx.rename(columns={'Locations.ADDRESS':'Home','Locations.ADDRESS':'Work Location 1a', 'Locations.ADDRESS':'Work Location 1b',
'Locations.ADDRESS': 'Work Location 2' }, inplace=True)
dfx = dfx.fillna(0)
dfx = dfx.astype({'Work Location 1a': 'int64', 'Work Location 1b': 'int64'})
dfx['Work Location 1'] = dfx['Work Location 1a'] + dfx['Work Location 1b']
dfx
# +
dfx1 = pd.DataFrame(dfx)
del dfx1['Work Location 1a']
del dfx1['Work Location 1b']
# -
dfx1
dfx2 = dfx1[['Month', 'Work Location 1', 'Work Location 2']]
dfx2
plt.plot(months,home_count, label='Home')
plt.plot(months,dfx['Work Location 1'], label='Work Location 1')
plt.plot(months,dfx['Work Location 2'], label='Work Location 2')
plt.xlabel('Month')
plt.xticks(months, rotation=30)
plt.ylabel('Frequency of Arrivals')
plt.title('Quantity of Times Arriving Home and Work Per Month - 2020')
plt.legend(bbox_to_anchor=(1, 1))
plt.grid(True)
plt.rcParams['figure.figsize'] = (9,5)
plt.show()
plt.tight_layout()
# +
from vincenty import vincenty
dist_list=[]
dist_month = {}
for month in months:
f = open(f'C:/Users/jspit/OneDrive/Documents/Takeout/Location History/Semantic Location History/2020/2020_{month}.json')
data=json.load(f)
for items in data['timelineObjects']:
try:
item = items['activitySegment']
start_coord = (int(item['startLocation']['latitudeE7'])/1e7, int(item['startLocation']['longitudeE7'])/1e7)
end_coord = (int(item['endLocation']['latitudeE7'])/1e7, int(item['endLocation']['longitudeE7'])/1e7)
distance = vincenty(start_coord, end_coord, miles=True)
dist_month = {"Month": month, "Distance":distance}
except:
None
dist_list.append(dist_month)
df2 = pd.DataFrame(dist_list)
df2_grp = df2.groupby(['Month'])
dist_totals = df2_grp['Distance'].sum()
df3 = pd.DataFrame(dist_totals)
df3 = df3.rename(columns = {'Distance': 'Monthly Distance Sum'})
df3
# +
months = ['JANUARY', 'FEBRUARY', 'MARCH', 'APRIL', 'MAY', 'JUNE', 'JULY', 'AUGUST',
'SEPTEMBER', 'OCTOBER', 'NOVEMBER', 'DECEMBER']
val= []
for index, row in df3.iterrows():
val.append(row['Monthly Distance Sum'])
dfsort = pd.DataFrame({'Month': [i for i in months], 'num':[0,1,2,3,4,5,6,7,8,9,10,11]})
df4 = pd.merge(df3, dfsort, on='Month')
df4=df4.sort_values('num')
plt1 = df4.plot(kind='bar',x='Month', y = 'Monthly Distance Sum', ylabel='Monthly Distance (mi)', title ='Monthly Distance Totals', figsize=(8,8))
plt1.get_legend().remove()
# +
newlist=[]
for month in months:
f = open(f'C:/Users/jspit/OneDrive/Documents/Takeout/Location History/Semantic Location History/2020/2020_{month}.json')
data=json.load(f)
for items in data['timelineObjects']:
try:
item = items['activitySegment']
start_coord = (int(item['startLocation']['latitudeE7'])/1e7, int(item['startLocation']['longitudeE7'])/1e7)
end_coord = (int(item['endLocation']['latitudeE7'])/1e7, int(item['endLocation']['longitudeE7'])/1e7)
distance = vincenty(start_coord, end_coord, miles=True)
travtype = item['activityType']
dist_month = {"Month": month, "Distance":distance, "Activity Type": travtype}
except:
None
newlist.append(dist_month)
df5 = pd.DataFrame(newlist)
df5
df5_grp = df5.groupby(['Month', 'Activity Type'])
act_totals = df5_grp['Distance'].sum()
act_totals
df6 = pd.DataFrame(act_totals)
df5_grp2 = df6.groupby(['Month', 'Activity Type'])
activity_totals = df5_grp2['Distance'].sum()
df5 = pd.DataFrame(activity_totals)
df5
# -
df5_grp2 = df5.groupby(['Activity Type'])
activity_totals = df5_grp2['Distance'].sum()
df5 = pd.DataFrame(activity_totals)
df5
df5.reset_index(inplace=True)
df5.sort_values(by='Distance', ascending=False)
labels= ['', 'FLYING', '', 'IN_PASSENGER_VEHICLE', '', '', 'SKIING', 'WALKING']
x = [i for i in df5['Distance']]
explode=[0, 0.5, 0.1,0.4, 0.1, 0.1, 0.5, 0.7]
y = plt.pie(x, labels=labels, autopct='%1.1f%%', radius=2, explode=explode)
| Google_Location_Data_2020-Scrubbed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Dithering Telescope - random offsets
# In this notebook, one can set a sky position to look at (boresight position) and a source (with profile and position) and study the effect of possible telescope offsets (such as fiber positioner offset, boresight offsets) on the signal-to-noise ratio (SNR).
#
import astropy.units as u
import matplotlib.pyplot as plt
import numpy as np
import sys
import os
sys.path.append('../../py')
import dithering
#
# After the general packages and the special package "dithering" is imported, one start with setting the source and boresight positions. By default, "generate_source()" function generates a source like a QSO. For other types of sources, you can uncomment/comment the lines below.
#
# ### Constructing the object
#
# One can uncomment/comment a source below to study
dit = dithering.dithering("../../config/desi-noblur-nooffset.yaml")
# QSO
#source = dit.generate_source(disk_fraction=0., bulge_fraction=0.)
#source_type = 'qso'
# ELG
source = dit.generate_source(disk_fraction=1., bulge_fraction=0.)
source_type = 'elg'
# LRG
#source = dit.generate_source(disk_fraction=0., bulge_fraction=1.)
#source_type = 'lrg'
dit.set_source_position(20.*u.deg, 25.*u.deg)
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.run_simulation(source_type, *source, report=True)
# ### Rotating the positioner along $\Phi$ and $\Theta$
#
# If the fiber aperture is not covering the source, we may have to move the fiber around a bit to find the maximum
# The example below starts from -2 degrees and scans up to 2 degrees to find the maximum.
#dit.change_alt_az_bore_position(20.*u.deg, 24.5*u.deg)
nbins = 50
#dit.set_focal_plane_position()
rotations = np.linspace(-.5, .5, nbins)
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
thetas = []
phis = []
SNR_b = []
SNR_r = []
SNR_z = []
for i in range(nbins):
for j in range(nbins):
dit.set_positioner_phi(rotations[i]*u.deg)
dit.set_positioner_theta(rotations[j]*u.deg)
dit.make_positioner_rotation()
dit.run_simulation(source_type, *source, report=False)
phis.append((rotations[i]*u.deg).to(u.arcsec).value)
thetas.append((rotations[j]*u.deg).to(u.arcsec).value)
SNR_b.append(np.median(dit.SNR['b'][0]))
SNR_r.append(np.median(dit.SNR['r'][0]))
SNR_z.append(np.median(dit.SNR['z'][0]))
# camera b
plt.hist2d(phis, thetas, weights=SNR_b, bins=nbins)
plt.xlabel("$\Phi$ [arcsec]")
plt.ylabel("$\Theta$ [arcsec]")
plt.title("SNR [camera b]")
plt.colorbar()
plt.savefig("theta_phi_b-band.pdf")
plt.show()
# camera r
plt.hist2d(phis, thetas, weights=SNR_r, bins=nbins)
plt.xlabel("$\Phi$ [arcsec]")
plt.ylabel("$\Theta$ [arcsec]")
plt.title("SNR [camera r]")
plt.colorbar()
plt.savefig("theta_phi_r-band.pdf")
plt.show()
# camera z
plt.hist2d(phis, thetas, weights=SNR_z, bins=nbins)
plt.xlabel("$\Phi$ [arcsec]")
plt.ylabel("$\Theta$ [arcsec]")
plt.title("SNR [camera z]")
plt.colorbar()
plt.savefig("theta_phi_z-band.pdf")
plt.show()
# ### Testing the random offsets - simple case
# Here we report some of the conditions of the simulation before we start changing them. Then we add random fiber placement offsets.
print("Nominal results:")
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.run_simulation(source_type, *source)
# adding some random offset
dit.add_random_offset_fiber_position(var_x=5*u.um)
print("------------------------------")
print("With fiber positioner offsets:")
dit.run_simulation(source_type, *source)
# ### Testing the random offsets - random fiber positioner offsets
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.run_simulation(source_type, *source, report=False)
num_iterations = 1000
snr_b = []
snr_r = []
snr_z = []
for i in range(num_iterations):
# move the fiber to its initial point before randomly moving it
dit.set_focal_plane_position()
dit.add_random_offset_fiber_position()
dit.run_simulation(source_type, *source, report=False)
snr_b.append(np.median(dit.SNR['b'][0]))
snr_r.append(np.median(dit.SNR['r'][0]))
snr_z.append(np.median(dit.SNR['z'][0]))
plt.clf()
plt.hist(snr_b, bins = 20)
plt.show()
# ### Testing the random offsets - random fiber positioner offsets
# One difference between the previous cell and the one below is that we introduce 1 arcsec movement to the telescope in azimuthal direction
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg+1*u.arcsec)
dit.run_simulation(source_type, *source, report=False)
num_iterations = 1000
snr_b = []
snr_r = []
snr_z = []
for i in range(num_iterations):
# move the fiber to its initial point before randomly moving it
dit.set_focal_plane_position()
dit.add_random_offset_fiber_position()
dit.run_simulation(source_type, *source, report=False)
snr_b.append(np.median(dit.SNR['b'][0]))
snr_r.append(np.median(dit.SNR['r'][0]))
snr_z.append(np.median(dit.SNR['z'][0]))
plt.clf()
plt.hist(snr_b, bins = 20)
plt.show()
# ### Testing random offsets - random boresight altitude offsets
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.run_simulation(source_type, *source, report=False)
num_iterations = 1000
snr_b = []
snr_r = []
snr_z = []
for i in range(num_iterations):
# move the fiber to its initial point before randomly moving it
#dit.set_focal_plane_position()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.add_random_boresight_offset(var_az=0*u.arcsec)
dit.run_simulation(source_type, *source, report=False)
snr_b.append(np.median(dit.SNR['b'][0]))
snr_r.append(np.median(dit.SNR['r'][0]))
snr_z.append(np.median(dit.SNR['z'][0]))
plt.clf()
plt.hist(snr_b, bins = 20)
plt.show()
# ### Testing random offsets - random boresight azimuth offsets
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.run_simulation(source_type, *source, report=False)
num_iterations = 1000
snr_b = []
snr_r = []
snr_z = []
for i in range(num_iterations):
# move the fiber to its initial point before randomly moving it
#dit.set_focal_plane_position()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.add_random_boresight_offset(var_alt=0*u.arcsec)
dit.run_simulation(source_type, *source, report=False)
snr_b.append(np.median(dit.SNR['b'][0]))
snr_r.append(np.median(dit.SNR['r'][0]))
snr_z.append(np.median(dit.SNR['z'][0]))
plt.clf()
plt.hist(snr_b, bins = 20)
plt.show()
# ### Testing random offsets - random boresight azimuth and altitude offsets
dit.set_positioner_theta(0*u.deg)
dit.set_positioner_phi(0*u.deg)
dit.make_positioner_rotation()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.run_simulation(source_type, *source, report=False)
num_iterations = 1000
snr_b = []
snr_r = []
snr_z = []
for i in range(num_iterations):
# move the fiber to its initial point before randomly moving it
#dit.set_focal_plane_position()
dit.set_boresight_position(20.*u.deg, 24.5*u.deg)
dit.set_focal_plane_position()
dit.add_random_boresight_offset()
dit.run_simulation(source_type, *source, report=False)
snr_b.append(np.median(dit.SNR['b'][0]))
snr_r.append(np.median(dit.SNR['r'][0]))
snr_z.append(np.median(dit.SNR['z'][0]))
plt.clf()
plt.hist(snr_b, bins = 20)
plt.show()
| docs/nb/dithering_randoms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo 2 : Data Modeling, Creating a Table with PostgreSQL
# <img src="images/postgresql-logo.png" width="250" height="250">
# configuring Postgres, creating users, and creating databases using the psql utility
# https://www.codementor.io/@engineerapart/getting-started-with-postgresql-on-mac-osx-are8jcopb
# psycopg2 https://pynative.com/python-postgresql-tutorial/
# # PostgreSQL and AutoCommits
# ## Walk through the basics of PostgreSQL autocommits
## import postgreSQL adapter for the Python
import psycopg2
# ### Create a connection to the database
# 1. Connect to the local instance of PostgreSQL (*127.0.0.1*)
# 2. Use the database/schema from the instance.
# 3. The connection reaches out to the database (*dataenginneering*) and use the correct privilages to connect to the database (*user = postgres and password = <PASSWORD>*).
conn = psycopg2.connect("host=127.0.0.1 dbname=dataengineering user=postgres password=<PASSWORD>")
# ### Use the connection to get a cursor that will be used to execute queries.
# https://www.postgresql.org/docs/9.2/plpgsql-cursors.html
cur = conn.cursor()
# ### Create a database to work in
cur.execute("select * from users ")
# ### Error occurs, but it was to be expected because table has not been created as yet. To fix the error, create the table.¶
cur.execute("CREATE TABLE test (col1 int, col2 int, col3 int);")
# ### Error indicates we cannot execute this query. Since we have not committed the transaction and had an error in the transaction block, we are blocked until we restart the connection.
conn = psycopg2.connect("host=127.0.0.1 dbname=dataengineering user=postgres password=<PASSWORD>")
cur = conn.cursor()
# In our exercises instead of worrying about commiting each transaction or getting a strange error when we hit something unexpected, let's set autocommit to true. **This says after each call during the session commit that one action and do not hold open the transaction for any other actions. One action = one transaction.**
# In this demo we will use automatic commit so each action is commited without having to call `conn.commit()` after each command. **The ability to rollback and commit transactions are a feature of Relational Databases.**
conn.set_session(autocommit=True)
cur.execute("select * from test")
cur.execute("CREATE TABLE test (col1 int, col2 int, col3 int);")
# ### Once autocommit is set to true, we execute this code successfully. There were no issues with transaction blocks and we did not need to restart our connection.
cur.execute("select * from test")
cur.execute("select count(*) from test")
print(cur.fetchall())
# # Creating a Table with PostgreSQL
# ## Walk through the basics of PostgreSQL:
# <br><li>Creating a table <li>Inserting rows of data, <li>Running a simple SQL query to validate the information.
# ### Typically, we would use a python wrapper called *psycopg2* to run the PostgreSQL queries. This library should be preinstalled but in the future to install this library, run the following command in the notebook to install locally:
# `!pip3 install --user psycopg2`
# #### More documentation can be found here: http://initd.org/psycopg/
# #### Import the library
# Note: An error might popup after this command has executed. Read it carefully before proceeding.
import psycopg2
# ### Create a connection to the database
# 1. Connect to the local instance of PostgreSQL (*127.0.0.1*)
# 2. Use the database/schema from the instance.
# 3. The connection reaches out to the database (*dataenginneering*) and use the correct privilages to connect to the database (*user = postgres and password = <PASSWORD>*).
#
# ### Note 1: This block of code will be standard in all notebooks.
# ### Note 2: Adding the `try except` will make sure errors are caught and understood
try:
conn = psycopg2.connect("host=127.0.0.1 dbname=dataengineering user=postgres password=<PASSWORD>")
except psycopg2.Error as e:
print("Error: Could not make connection to the Postgres database")
print(e)
# ### Use the connection to get a cursor that can be used to execute queries.
try:
cur = conn.cursor()
except psycopg2.Error as e:
print("Error: Could not get curser to the Database")
print(e)
# ### Use automactic commit so that each action is commited without having to call conn.commit() after each command. The ability to rollback and commit transactions is a feature of Relational Databases.
conn.set_session(autocommit=True)
# ### Test the Connection and Error Handling Code
# The try-except block should handle the error: We are trying to do a select * on a table but the table has not been created yet.
try:
cur.execute("select * from quran.surah")
except psycopg2.Error as e:
print(e)
# ### Create a database to work in
try:
cur.execute("create database quran")
except psycopg2.Error as e:
print(e)
# ### Close our connection to the default database, reconnect to the Quran database, and get a new cursor.
# +
try:
conn.close()
except psycopg2.Error as e:
print(e)
try:
conn = psycopg2.connect("host=127.0.0.1 dbname=quran user=postgres password=<PASSWORD>")
except psycopg2.Error as e:
print("Error: Could not make connection to the Postgres database")
print(e)
try:
cur = conn.cursor()
except psycopg2.Error as e:
print("Error: Could not get curser to the Database")
print(e)
conn.set_session(autocommit=True)
# -
# ### We will create AL-QURAN, each surah has a lot of information we could add to the AL-QURAN database. We will design english translation by create quran_index, sura, aya, text
#
# `Table Name: English Saheeh International
# column 1: Quran Index
# column 2: Surah
# column 3: Ayah
# column 4: Text Translation`
#
# ### Translate this information into a Create Table Statement.
#
# Review this document on PostgreSQL datatypes: https://www.postgresql.org/docs/9.5/datatype.html
#
try:
cur.execute("CREATE TABLE IF NOT EXISTS en_sahih (index int, surah int, ayah int, text varchar);")
except psycopg2.Error as e:
print("Error: Issue creating table")
print (e)
# ### No error was found, but lets check to ensure our table was created. `select count(*)` which should return 0 as no rows have been inserted in the table.
# +
try:
cur.execute("select count(*) from en_sahih")
except psycopg2.Error as e:
print("Error: Issue creating table")
print (e)
print(cur.fetchall())
# -
# ### Insert two rows
## 1
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(1, 1, 1, 'In the name of Allah, the Entirely Merciful, the Especially Merciful.'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
# +
## 2
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(2, 1, 2, '[All] praise is [due] to Allah, Lord of the worlds -'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
## 3
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(3, 1, 3, 'The Entirely Merciful, the Especially Merciful,'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
## 4
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(4, 1, 4, 'Sovereign of the Day of Recompense.'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
## 5
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(5, 1, 5, 'It is You we worship and You we ask for help.'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
## 6
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(6, 1, 6, 'Guide us to the straight path -'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
## 7
try:
cur.execute("INSERT INTO en_sahih (index, surah, ayah, text) \
VALUES (%s, %s, %s, %s)", \
(7, 1, 7, 'The path of those upon whom You have bestowed favor, not of those who have evoked [Your] anger or of those who are astray.'))
except psycopg2.Error as e:
print("Error: Inserting Rows")
print (e)
# -
# ### Validate your data was inserted into the table.
# The while loop is used for printing the results. If executing queries in the Postgres shell, this would not be required.
# ### Note: If you run the insert statement code more than once, you will see duplicates of your data. PostgreSQL allows for duplicates.
# +
try:
cur.execute("SELECT * FROM en_sahih;")
except psycopg2.Error as e:
print("Error: select *")
print (e)
row = cur.fetchone()
while row:
print(row)
row = cur.fetchone()
# -
# ### Drop the table to avoid duplicates and clean up
try:
cur.execute("DROP table en_sahih")
except psycopg2.Error as e:
print("Error: Dropping table")
print (e)
# ### Close the cursor and connection.
cur.close()
conn.close()
| 5-data-modeling/5-2-data-modeling_postgresql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [python3]
# language: python
# name: Python [python3]
# ---
# This notebook implements a sequential backward selection (SBS) algorithm for feature selection.
# Reference to [Sequential feature selection algorithms](https://github.com/rasbt/python-machine-learning-book/blob/master/code/ch04/ch04.ipynb)
#
#
# +
from sklearn.base import clone
from itertools import combinations
import numpy as np
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
class SBS():
def __init__(self, estimator, k_features, scoring=accuracy_score,
test_size=0.25, random_state=1):
'''
k_features: the desired number of features we want to return
estimator: model for classification on the feature subsets
scoring: metric to evaluate the performance of a model (accuracy_score by default)
'''
self.scoring = scoring
self.estimator = clone(estimator)
self.k_features = k_features
self.test_size = test_size
self.random_state = random_state
def fit(self, X, y):
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=self.test_size,
random_state=self.random_state)
dim = X_train.shape[1]
self.indices_ = tuple(range(dim))
self.subsets_ = [self.indices_]
score = self._calc_score(X_train, y_train,
X_test, y_test, self.indices_)
self.scores_ = [score]
# feature subset is created by the itertools.combinations until the feature subset
# has the desired dimensionality
# in each iteration the accuracy score of the best subset is collected in self.scores_
# the column indices of the final feature subset are assigned to self.indices_
while dim > self.k_features:
scores = []
subsets = []
for p in combinations(self.indices_, r=dim - 1):
score = self._calc_score(X_train, y_train,
X_test, y_test, p)
scores.append(score)
subsets.append(p)
best = np.argmax(scores)
self.indices_ = subsets[best]
self.subsets_.append(self.indices_)
dim -= 1
self.scores_.append(scores[best])
self.k_score_ = self.scores_[-1]
return self
# transform method returns a new data array with the selected feature columns
def transform(self, X):
return X[:, self.indices_]
def _calc_score(self, X_train, y_train, X_test, y_test, indices):
self.estimator.fit(X_train[:, indices], y_train)
y_pred = self.estimator.predict(X_test[:, indices])
score = self.scoring(y_test, y_pred)
return score
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import StandardScaler
stdsc = StandardScaler()
X_train_std = stdsc.fit_transform(X_train)
X_test_std = stdsc.transform(X_test)
knn = KNeighborsClassifier(n_neighbors=2)
# selecting features
sbs = SBS(knn, k_features=1)
sbs.fit(X_train_std, y_train)
# plotting performance of feature subsets
k_feat = [len(k) for k in sbs.subsets_]
plt.plot(k_feat, sbs.scores_, marker='o')
plt.ylim([0.7, 1.1])
plt.ylabel('Accuracy')
plt.xlabel('Number of features')
plt.grid()
plt.tight_layout()
# plt.savefig('./sbs.png', dpi=300)
plt.show()
# -
k5 = list(sbs.subsets_[8])
print(df_wine.columns[1:][k5])
knn.fit(X_train_std[:, k5], y_train)
print('Training accuracy:', knn.score(X_train_std[:, k5], y_train))
print('Test accuracy:', knn.score(X_test_std[:, k5], y_test))
# +
import pandas as pd
from sklearn.model_selection import train_test_split
df_wine = pd.read_csv('https://archive.ics.uci.edu/'
'ml/machine-learning-databases/wine/wine.data',
header=None)
df_wine.columns = ['Class label', 'Alcohol', 'Malic acid', 'Ash',
'Alcalinity of ash', 'Magnesium', 'Total phenols',
'Flavanoids', 'Nonflavanoid phenols', 'Proanthocyanins',
'Color intensity', 'Hue', 'OD280/OD315 of diluted wines',
'Proline']
X, y = df_wine.iloc[:, 1:].values, df_wine.iloc[:, 0].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# +
from sklearn.ensemble import RandomForestClassifier
import numpy as np
feat_labels = df_wine.columns[1:]
forest = RandomForestClassifier(n_estimators=10000,
random_state=0,
n_jobs=-1)
forest.fit(X_train, y_train)
importances = forest.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(X_train.shape[1]):
print("%2d) %-*s %f" % (f + 1, 30,
feat_labels[indices[f]],
importances[indices[f]]))
# +
import matplotlib.pyplot as plt
plt.title('Feature Importances')
plt.bar(range(X_train.shape[1]),
importances[indices],
color='lightblue',
align='center')
plt.xticks(range(X_train.shape[1]),
feat_labels[indices], rotation=90)
plt.xlim([-1, X_train.shape[1]])
plt.tight_layout()
#plt.savefig('./random_forest.png', dpi=300)
plt.show()
# -
| ch4/Implement_sequential_feature_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Dynamic size plot with bokeh
# Creating a plot with a dynamic width size can change with the web page size.
# +
# Data handing tools
import numpy as np
# Bokeh tools
from bokeh.plotting import figure, show, output_file
from bokeh.io import curdoc, output_notebook
# + [markdown] pycharm={"name": "#%% md\n"}
# Generating random values.
# + pycharm={"name": "#%%\n"}
x = np.random.randint(-10, 10, size=10)
y = np.random.randint(-10, 10, size=10)
# + [markdown] pycharm={"name": "#%% md\n"}
# Setting 'night_sky' for the plot.
# + pycharm={"name": "#%%\n"}
curdoc().theme = 'contrast'
# + [markdown] pycharm={"name": "#%% md\n"}
# Setting plot to show inside the notebook and on a html webpage.
# + pycharm={"name": "#%%\n"}
output_file('stretching_plot.html')
output_notebook()
# + [markdown] pycharm={"name": "#%% md\n"}
# Setting up a figure frame.
# + pycharm={"name": "#%%\n"}
p = figure(title='Dynamic Size Plot', sizing_mode='stretch_width', plot_height=500, x_axis_label='x-axis', y_axis_label='y-axis')
p.title.align = "center"
# + [markdown] pycharm={"name": "#%% md\n"}
# Creating a scatter plot inside the figure.
# + pycharm={"name": "#%%\n"}
p.circle(x, y, fill_color='green', size=20)
show(p)
| stretching_plot/stretching_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: OceanParcels
# language: python
# name: oceanparcels
# ---
# # Adding Back the Tides - CIOPS
# An effort to make the daily files more accurate as they are currently lacking the tidal pumping that is so important to the flow of the Salish Sea
import xarray as xr
from pathlib import Path
import numpy as np
import datetime as dt
float(np.nan)
# #### unlike what we did with SSC in this file make all add back tides days seperate to reduce space requirement
# #### just doing 2018 for now
# plus 30 days on the end for running purposes
startday = [dt.datetime(2017,12,31)+dt.timedelta(days=i) for i in range(int(399))]
print(startday[-1])
# +
# for i in range(131,len(startday)-1):
# print(i)
# -
folders = [dt.datetime(2017,12,31)+dt.timedelta(days=7*(i+1)) for i in range(int(57))]
print(folders[-1])
folders = np.repeat(folders,7)
# +
# start = dt.datetime(2015,11,22) #start day of your run
#dates for each run
# numdays = 2 #15 for all except the last run (7)
# date_list = [start + dt.timedelta(days=x) for x in range(numdays)]
# folderday = [start+ dt.timedelta(days=7),start+ dt.timedelta(days=7*2)]#,start+ dt.timedelta(days=7*3)]
# folderday = np.repeat(folderday,7)
# folderday = folderday[:-6]
date_list_daily = [startday[0],startday[1]]
folderday_daily = [folders[0], folders[1]]
# for whatever reason the naming it raelly really weird and the daily and the hourly files names for the SAME DAY aren't\
# on the same day.. must take hourly files a day ahead woooooooooooo
date_list_hourly = [startday[1],startday[2]]
folderday_hourly = [folders[1], folders[2]]
# -
"{:%Y%m%d}00/BC12_1h_grid_U_2D_{:%Y%m%d}_{:%Y%m%d}.nc".format(folderday_hourly[0], date_list_hourly[0], date_list_hourly[0])
# +
path = Path("/ocean/mdunphy/CIOPSW-BC12/")
drop_vars = (
"sbu", "tauuo", "time_counter_bounds", "time_instant_bounds", "uos", "time_instant",
)
files = [sorted(path.glob("{:%Y%m%d}00/BC12_1h_grid_U_2D_{:%Y%m%d}_{:%Y%m%d}.nc".format(folderday_hourly[i], date_list_hourly[i], date_list_hourly[i]))) for i in range(len(date_list_hourly))]
mydata = xr.open_mfdataset(files, drop_variables=drop_vars)
ut_h = mydata['ubar']
# -
files
ut_h
ut_h[0,646,267].values
# +
drop_vars = (
"sbv", "tauvo", "time_counter_bounds", "time_instant_bounds", "vos", "time_instant",
)
files = [sorted(path.glob("{:%Y%m%d}00/BC12_1h_grid_V_2D_{:%Y%m%d}_{:%Y%m%d}.nc".format(folderday_hourly[i], date_list_hourly[i], date_list_hourly[i]))) for i in range(len(date_list_hourly))]
mydata = xr.open_mfdataset(files, drop_variables=drop_vars)
vt_h = mydata['vbar']
# +
# 2D hourly outputs are alraedy in bartropic form so you dont need to do any conversion!
# +
# Now get the required data from the daily files
drop_vars = (
"depthu_bounds", "nav_lat", "nav_lon", 'time_counter_bounds', 'time_instant',
'time_instant_bounds',
)
files = [sorted(path.glob("{:%Y%m%d}00/BC12_1d_grid_U_{:%Y%m%d}_{:%Y%m%d}.nc".format(folderday_daily[i], date_list_daily[i], date_list_daily[i]))) for i in range(len(date_list_daily))]
mydata = xr.open_mfdataset(files, drop_variables=drop_vars)
u_d = mydata['uo']
drop_vars = (
"depthv_bounds", "nav_lat", "nav_lon", 'time_counter_bounds', 'time_instant',
'time_instant_bounds',
)
files = [sorted(path.glob("{:%Y%m%d}00/BC12_1d_grid_V_{:%Y%m%d}_{:%Y%m%d}.nc".format(folderday_daily[i], date_list_daily[i], date_list_daily[i]))) for i in range(len(date_list_daily))]
mydata = xr.open_mfdataset(files, drop_variables=drop_vars)
v_d = mydata['vo']
# -
u_d = u_d.where(u_d != 0)
u_d[0,:,267,646].values
u_d[0,:,240,646].values
u_d
# we WILL need to do some conversions here, so get e3t from the mesh_mask file
mydata = xr.open_dataset("/ocean/mdunphy/CIOPSW-BC12/grid/mesh_mask_Bathymetry_NEP36_714x1020_SRTM30v11_NOAA3sec_WCTSS_JdeFSalSea.nc")
e3t = mydata['e3t_0']
e3t
# +
# convert e3t to e3u and to e3v
e3t_xshift = e3t.shift(x=-1,fill_value=0)
e3u = e3t_xshift+e3t
e3u = e3u*0.5
e3u = e3u.rename({'z': 'depthu'})
e3t_yshift = e3t.shift(y=-1,fill_value=0)
e3v = e3t_yshift+e3t
e3v = e3v*0.5
e3v = e3v.rename({'z': 'depthv'})
# -
#calcuate bartropic component
ut_d = (u_d*e3u[0,:,:,:]).sum(dim='depthu')/e3u[0,:,:,:].sum(dim='depthu')
ut_d[0,646,267].values
#subtract from u to get baroclinic component
uc_d = u_d-ut_d #does this work even though their ut_d lacks the depth dimension?
# interpolate + resample uc_d to get it in an hourly format
uc_d[0,:,646,267].values
offset = dt.timedelta(hours=1) # daily avg timestamp as next day time 0, offset by -23 hours to line up with hourly data
uc_h_interp = uc_d.resample(time_counter="1H", loffset=offset).interpolate("linear")
#instead of taking 12hours off each side like when done with SSC, this method takes off 24hours on the end!
# , loffset=offset
uc_h_interp
uc_h_interp[0,0,646,267].values
u_new = ut_h + uc_h_interp
u_new = u_new.isel(time_counter = np.arange(0,24,1))
u_new = u_new.rename('vozocrtx')
u_new
u_new[0,646,267,0].values
encoding={
"vozocrtx": {"zlib": True, "complevel": 4, "_FillValue": np.nan}
}
encoding
# + jupyter={"source_hidden": true} tags=[]
# encoding = {
# "depthu": {
# "zlib": True, "complevel": 4,
# "chunksizes": {
# "depthu": 75,
# }
# },
# "vozocrtx": {
# "zlib": True, "complevel": 4,
# "chunksizes": {
# "time_counter": 1, "y": 1020, "x": 714, "depthu": 75,
# }
# }
# }
# encoding = {
# "depthu": {
# "zlib": True, "complevel": 4,
# "chunksizes": [75],
# },
# "time_counter": {
# "zlib": True, "complevel": 4,
# "chunksizes": [1],
# "units": "hours since 1970-01-01 00:00:00"
# },
# "nav_lat": {
# "zlib": True, "complevel": 4,
# "chunksizes": {
# "y": 1020, "x": 714,
# }
# },
# "nav_lon": {
# "zlib": True, "complevel": 4,
# "chunksizes": {
# "y": 1020, "x": 714,
# }
# },
# "vozocrtx": {
# "zlib": True, "complevel": 4,
# "chunksizes": {
# "time_counter": 1, "y": 1020, "x": 714, "depthu": 75,
# }
# }
# }
# -
path = "/ocean/rbeutel/data/"
u_new.to_netcdf(str(path)+'u_new_{:%d%b%y}_fill.nc'.format(date_list_hourly[0],date_list_hourly[0]), encoding=encoding)
print('u_new_{:%d%b%y}_{:%d%b%y}.nc'.format(date_list_hourly[0],date_list_hourly[0]))
#calcuate bartropic component
vt_d = (v_d*e3v[0,:,:,:]).sum(dim='depthv')/e3v[0,:,:,:].sum(dim='depthv')
#subtract from v to get baroclinic component
vc_d = v_d-vt_d
vc_h_interp = vc_d.resample(time_counter="1H", loffset=offset).interpolate("linear")
v_new = vt_h + vc_h_interp
v_new = v_new.isel(time_counter = np.arange(0,24,1))
v_new = v_new.rename('vomecrty')
v_new
v_new.to_netcdf(str(path)+'v_new_{:%d%b%y}_{:%d%b%y}.nc'.format(date_list[0],date_list[-1]))
| ArtificialTides/BaroclinicBarotropic-BC12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/martin-fabbri/colab-notebooks/blob/master/tree-based-models/tree_based_models_02.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="h41beIilC4SC" colab_type="text"
# # Decision Trees and Random Forest
# + id="XsY16nJaC4Dw" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
from sklearn.datasets import load_digits
from sklearn.datasets import make_blobs
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
sns.set()
warnings.filterwarnings('ignore')
# + [markdown] id="uNU6txCvJDr8" colab_type="text"
# Decision trees are extremely intuitive ways to classify or label objects. The whole concept is to simply ask a series of questions designed to zero-in on the classification.
#
# The tree's binary splitting makes them extremelly efficient. In a well-constructed tree, each question will cit the number of options by approximately half. The trick comes in deciding which question to ask at each step.
#
# The questions generally take the form of axis-aligned splits in data, where each node in the tree splits the data into two groups using a cutoff value within one of the features.
# + [markdown] id="_9XhJDd0Q7cn" colab_type="text"
# ## Tree Plotting Utilities (keep collapsed)
# + id="L2Bbx7fCQ7F_" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeClassifier
from ipywidgets import interact
def visualize_classifier(model, X, y, ax=None, cmap='rainbow'):
ax = ax or plt.gca()
# Plot the training points
ax.scatter(X[:, 0], X[:, 1], c=y, s=30, cmap=cmap,
clim=(y.min(), y.max()), zorder=3)
ax.axis('tight')
ax.axis('off')
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# fit the estimator
model.fit(X, y)
xx, yy = np.meshgrid(np.linspace(*xlim, num=200),
np.linspace(*ylim, num=200))
Z = model.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
# Create a color plot with the results
n_classes = len(np.unique(y))
contours = ax.contourf(xx, yy, Z, alpha=0.3,
levels=np.arange(n_classes + 1) - 0.5,
cmap=cmap, clim=(y.min(), y.max()),
zorder=1)
ax.set(xlim=xlim, ylim=ylim)
def visualize_tree(estimator, X, y, boundaries=True,
xlim=None, ylim=None, ax=None):
ax = ax or plt.gca()
# Plot the training points
ax.scatter(X[:, 0], X[:, 1], c=y, s=30, cmap='viridis',
clim=(y.min(), y.max()), zorder=3)
ax.axis('tight')
ax.axis('off')
if xlim is None:
xlim = ax.get_xlim()
if ylim is None:
ylim = ax.get_ylim()
# fit the estimator
estimator.fit(X, y)
xx, yy = np.meshgrid(np.linspace(*xlim, num=200),
np.linspace(*ylim, num=200))
Z = estimator.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
n_classes = len(np.unique(y))
Z = Z.reshape(xx.shape)
contours = ax.contourf(xx, yy, Z, alpha=0.3,
levels=np.arange(n_classes + 1) - 0.5,
cmap='viridis', clim=(y.min(), y.max()),
zorder=1)
ax.set(xlim=xlim, ylim=ylim)
# Plot the decision boundaries
def plot_boundaries(i, xlim, ylim):
if i >= 0:
tree = estimator.tree_
if tree.feature[i] == 0:
ax.plot([tree.threshold[i], tree.threshold[i]], ylim, '-k', zorder=2)
plot_boundaries(tree.children_left[i],
[xlim[0], tree.threshold[i]], ylim)
plot_boundaries(tree.children_right[i],
[tree.threshold[i], xlim[1]], ylim)
elif tree.feature[i] == 1:
ax.plot(xlim, [tree.threshold[i], tree.threshold[i]], '-k', zorder=2)
plot_boundaries(tree.children_left[i], xlim,
[ylim[0], tree.threshold[i]])
plot_boundaries(tree.children_right[i], xlim,
[tree.threshold[i], ylim[1]])
if boundaries:
plot_boundaries(0, xlim, ylim)
def plot_tree_interactive(X, y):
def interactive_tree(depth=5):
clf = DecisionTreeClassifier(max_depth=depth, random_state=0)
visualize_tree(clf, X, y)
return interact(interactive_tree, depth=[1, 5])
def randomized_tree_interactive(X, y):
N = int(0.75 * X.shape[0])
xlim = (X[:, 0].min(), X[:, 0].max())
ylim = (X[:, 1].min(), X[:, 1].max())
def fit_randomized_tree(random_state=0):
clf = DecisionTreeClassifier(max_depth=15)
i = np.arange(len(y))
rng = np.random.RandomState(random_state)
rng.shuffle(i)
visualize_tree(clf, X[i[:N]], y[i[:N]], boundaries=False,
xlim=xlim, ylim=ylim)
interact(fit_randomized_tree, random_state=[0, 100]);
# + [markdown] id="MkNPpC6vPL4O" colab_type="text"
# ## Decision Tree Example
# + id="nd4OEJ8BC37c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 361} outputId="fefb5a0b-35b2-427d-9b1b-085d8903cdcf"
fig = plt.figure(figsize=(10, 4))
ax = fig.add_axes([0, 0, 0.8, 1], frameon=False, xticks=[], yticks=[])
ax.set_title('Example Decision Tree: Animal Classification', size=24)
def text(ax, x, y, t, size=20, **kwargs):
ax.text(x, y, t,
ha='center', va='center', size=size,
bbox=dict(boxstyle='round', ec='k', fc='w'), **kwargs)
text(ax, 0.5, 0.9, "How big is\nthe animal?", 20)
text(ax, 0.3, 0.6, "Does the animal\nhave horns?", 18)
text(ax, 0.7, 0.6, "Does the animal\nhave two legs?", 18)
text(ax, 0.12, 0.3, "Are the horns\nlonger than 10cm?", 14)
text(ax, 0.38, 0.3, "Is the animal\nwearing a collar?", 14)
text(ax, 0.62, 0.3, "Does the animal\nhave wings?", 14)
text(ax, 0.88, 0.3, "Does the animal\nhave a tail?", 14)
text(ax, 0.4, 0.75, "> 1m", 12, alpha=0.4)
text(ax, 0.6, 0.75, "< 1m", 12, alpha=0.4)
text(ax, 0.21, 0.45, "yes", 12, alpha=0.4)
text(ax, 0.34, 0.45, "no", 12, alpha=0.4)
text(ax, 0.66, 0.45, "yes", 12, alpha=0.4)
text(ax, 0.79, 0.45, "no", 12, alpha=0.4)
ax.plot([0.3, 0.5, 0.7], [0.6, 0.9, 0.6], '-k')
ax.plot([0.12, 0.3, 0.38], [0.3, 0.6, 0.3], '-k')
ax.plot([0.62, 0.7, 0.88], [0.3, 0.6, 0.3], '-k')
ax.plot([0.0, 0.12, 0.20], [0.0, 0.3, 0.0], '--k')
ax.plot([0.28, 0.38, 0.48], [0.0, 0.3, 0.0], '--k')
ax.plot([0.52, 0.62, 0.72], [0.0, 0.3, 0.0], '--k')
ax.plot([0.8, 0.88, 1.0], [0.0, 0.3, 0.0], '--k')
ax.axis([0, 1, 0, 1])
# + [markdown] id="QjW5YxEXP8MJ" colab_type="text"
# ## Creating a decision tree
# + id="V4kgtPxkC4Ab" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 268} outputId="b85d6ae3-c8d9-4d7f-8168-ec8ddce66bc7"
X, y = make_blobs(n_samples=300, centers=4, random_state=0, cluster_std=1.0)
plt.scatter(X[:,0], X[:,1], c=y, s=50, cmap='viridis');
# + [markdown] id="jca0VWgXPlEd" colab_type="text"
# A simple decision tree build on this data will interactively split the data along one of the other axis according to some quantitative criterion.
# + id="K26xXt_vC33T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 210} outputId="945669ce-788f-4c14-cbb3-7b9ce03b0f82"
fig, ax = plt.subplots(1, 3, figsize=(10, 3))
fig.subplots_adjust(left=0.02, right=0.98, wspace=0.1)
for axi, depth in zip(ax, range(1, 4)):
model = DecisionTreeClassifier(max_depth=depth)
visualize_tree(model, X, y, ax=axi)
axi.set_title('depth = {0}'.format(depth));
# + id="gJCSg0BwC3zn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="9f0e6a63-bbc3-410e-ead4-62b1348f0a43"
# without max_depth restrictions
dt = DecisionTreeClassifier()
visualize_tree(dt, X, y)
# + [markdown] id="Lk-WsXtO7Ve3" colab_type="text"
# ## Decision Trees Overfitting
#
# Such overfitting turns out to be a general property of decision tres; it is very easy to go too deep in the tree, and thus end up fitting details particular to the analysed sample that might not general well with the overall distribution.
#
# Another way to see overfitting is to look at model trained with different subsets of the same dataset.
#
# + id="98Hf6Ag6C3wb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="480f4988-e5b5-4657-b4a4-9d6c69966f4a"
model = DecisionTreeClassifier()
fig, ax = plt.subplots(1, 2, figsize=(10, 6))
fig.subplots_adjust(left=0.0625, right=0.95, wspace=0.1)
visualize_tree(model, X[::2], y[::2], boundaries=False, ax=ax[0])
visualize_tree(model, X[1::2], y[1::2], boundaries=False, ax=ax[1])
# + id="kpBqEhPhC3tN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="a93af898-8997-437a-df10-cc9dc2995dd5"
model = DecisionTreeClassifier(max_depth=4)
fig, ax = plt.subplots(1, 2, figsize=(10, 6))
fig.subplots_adjust(left=0.0625, right=0.95, wspace=0.1)
visualize_tree(model, X[::2], y[::2], boundaries=False, ax=ax[0])
visualize_tree(model, X[1::2], y[1::2], boundaries=False, ax=ax[1])
# + [markdown] id="rjduKj1XES_L" colab_type="text"
# We can identify that our model consistently produces some similar regions, while there are some regions that are unique to an spefic model.
#
# **The key observation is that inconsistencies tend to happen where the classification is less certain. We might be able to conbine the results of different models, reduce those unceirtainties, and come up with a better model.**
#
# **The sum could be better than it's parts.**
# + [markdown] id="zFrJpTkaF5Fg" colab_type="text"
# ## Ensembles of Estimators: Random Forest
#
# This notions, that multiple oferfitting estimators can be combined to reduce the effect of overfitting, is that underlies an ensemble method called bagging. Bagging makes use of an ensemble (a grab bag, perhaps) of parallel estimators, each of which overfits the data combining the results to find a better model. ***An ensemble of randomized decision trees is know as random forest***.
#
# This type of bagging classification can be done manually using sklearn's ``BaggingClassifier``.
# + id="0dJQWIQJC3qA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="4b714141-ffa8-42ba-b5a4-8cb3a3b5ef39"
tree = DecisionTreeClassifier()
bag = BaggingClassifier(tree, n_estimators=100, max_samples=0.8, random_state=1)
bag.fit(X, y)
visualize_classifier(bag, X, y)
# + [markdown] id="GucuGjtoKJcx" colab_type="text"
# In the above example, we have randomized the data by fitting each estimator witha random subset of 80%. In practice, decision trees are more effective randomized by injecting some stochasticity in how the splits are chosen.
# + id="sj7DsJY6C3mx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="ec203e05-6d84-4f7a-99aa-b6c8a508b19a"
model = RandomForestClassifier(n_estimators=100, random_state=1)
visualize_classifier(model, X, y)
# + [markdown] id="KC9DVAL9LDIT" colab_type="text"
# ## Random Forest Regression
# + id="ULoytv46C3lG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 268} outputId="85e24bc1-c966-4038-bd87-d5bd123bef74"
rng = np.random.RandomState(42)
X = 10 * rng.rand(200)
def model(data, sigma=0.3):
fast_oscilation = np.sin(5 * data)
slow_oscilation = np.sin(0.5 * data)
noise = sigma * rng.randn(len(data))
return slow_oscilation + fast_oscilation + noise
y = model(X)
plt.errorbar(X, y, 0.3, fmt='o');
# + id="4iloq1fCC3hx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 268} outputId="9ab4735c-dc97-47ca-f184-a2868bac63a4"
forest = RandomForestRegressor(200)
forest.fit(x[:, None], y)
x_fit = np.linspace(0, 10, 1000)
y_fit = forest.predict(x_fit[:, None])
y_true = model(x_fit, sigma=0)
plt.errorbar(X, y, 0.3, fmt='o', alpha=0.5)
plt.plot(x_fit, y_fit, '-r')
plt.plot(x_fit, y_true, '-k', alpha=0.5);
# + [markdown] id="eBRKnERVWq64" colab_type="text"
# ## Classifiying MNIST
# + id="-m70coM8C3er" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fb2424d2-ac10-4ecd-f713-4cff4e90507b"
digits = load_digits()
digits.keys()
# + id="cC-s02HlC3bp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 463} outputId="c8c62a0a-39a7-45bb-c46c-af23520c15bf"
fig = plt.figure(figsize=(6, 6))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1, hspace=0.05, wspace=0.05)
for i in range(64):
ax = fig.add_subplot(8, 8, i + 1, xticks=[], yticks=[])
ax.imshow(digits.images[i], cmap=plt.cm.binary, interpolation='nearest')
ax.text(0, 7, str(digits.target[i]));
# + id="wlCt-E6cPS3U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="06f534c7-9627-410e-8a26-ba9af6a1daee"
X_train, X_test, y_train, y_test = train_test_split(digits.data,
digits.target,
random_state=0)
model = RandomForestClassifier(n_estimators=1000)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
# + id="wKt5pVhnPSzY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="fb2833b7-f<PASSWORD>-4f4e-9dce-<PASSWORD>"
mat = confusion_matrix(y_test, y_pred)
sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False)
plt.xlabel('true label')
plt.ylabel('predicted label');
# + id="_oAsG--wPSwK" colab_type="code" colab={}
# + id="zsEM0yW8C3Ym" colab_type="code" colab={}
# + id="rLWVZ7UnC3Vi" colab_type="code" colab={}
# + id="eMcRgMosC3SD" colab_type="code" colab={}
# + id="rgV7uI0WC3O1" colab_type="code" colab={}
# + id="fdKb_hA5C3L3" colab_type="code" colab={}
# + id="VxJMcEc7C3I9" colab_type="code" colab={}
# + id="9oD0aEHlC3F-" colab_type="code" colab={}
# + id="GHKjmNYIC3DE" colab_type="code" colab={}
# + id="2n8Wm0buC16o" colab_type="code" colab={}
# + id="NPgH1UfLPSp9" colab_type="code" colab={}
| tree-based-models/tree_based_models_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from libspiral import seqset
max_results = 20
sample = raw_input('Sample: ')
my_graph = seqset('/mnt/{0}.gbwt'.format(sample))
while True:
sequence = raw_input('Query: ')
if sequence == '':
break
new_ctx = my_graph.find(sequence)
if(new_ctx.valid):
count = new_ctx.end - new_ctx.begin
print 'Found {0} entries\n'.format(count)
if count > max_results:
print 'Showing the first {0} matches'.format(max_results)
count = max_results
for i in range(new_ctx.begin, new_ctx.begin + count):
print my_graph.entry(i).sequence
print
else:
print 'No entries found.'
# -
| python/jupyter/agbtdemo/query_seqset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Made on July 25th, 2019
@author: <NAME>
@contact: <EMAIL>
"""
x_Delta = np.log10(54) # In our time units, the time between SDSS and HSC
default_Delta_value = -0.0843431604042636
data_path_times = '/home/tpena01/AGN_variability_project/Simulations/light_curves/v_bend_in_times_10/results_v_bend_in_times_10_{}.bin'
data_path_divided = '/home/tpena01/AGN_variability_project/Simulations/light_curves/v_bend_in_divided_10/results_v_bend_in_divided_10_{}.bin'
data_path_gets_2 = '/home/tpena01/AGN_variability_project/Simulations/light_curves/a_low_in_gets_2/results_a_low_in_gets_2_{}.bin'
data_path_gets_0 = '/home/tpena01/AGN_variability_project/Simulations/light_curves/a_low_in_gets_0/results_a_low_in_gets_0_{}.bin'
data_path_fiducial = '/home/tpena01/AGN_variability_project/Simulations/light_curves/default_params/results_Default_curve_{}.bin'
# # Setup and imports
# +
import sys
print("sys version: {}".format(sys.version))
# This project is entirely in python 3.7
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib qt
# If you don't have an X server, line 7 might crash your kernel. Try '%matplotlib inline' instead.
import numpy as np
print("numpy version: {}".format(np.__version__))
from tqdm import tqdm
# This gives for loops progress bars.
import random
# This helps choosing random numbers from our arrays
random.seed() # Randomize seed
from IPython.core.display import display, HTML
# An alternate, cleaner take on the jupyter workspace
display(HTML("<style>.container { width:100% !important; }</style>"))
# + [markdown] heading_collapsed=true
# # Data extraction function
# + code_folding=[] hidden=true
def extract_data(brightest_percent_lower=1, brightest_percent_upper=0, num_random_points=1000, t_max=10000, length_curve=2**24, num_curves=20, path=data_path):
"""
Input: Parameters documented in the cell above
Output: Graph of delta Eddington ratio as a function of delta time.
"""
if brightest_percent_lower <= brightest_percent_upper:
sys.exit('Can\'t have an interval where the lower bound is greater than or equal to the upper bound. Remember, things are reversed. 100% is a reasonable lower bound, and brightest_percent_upper defaults to zero.')
# Load the data
default_curves = []
for i in tqdm(range(num_curves + 1)):
if i == 0:
continue # for some reason the results files start at 1 and not 0
_er_curve = np.zeros(length_curve, dtype=float)
_er_curve = np.fromfile(path.format(str(i)))
default_curves.append(_er_curve)
default_curves = np.array(default_curves)
default_curves = np.log10(default_curves) # Move everything into the log domain
# Cut out the last t_max points
cut_curves = np.zeros((np.array(list(default_curves.shape)) - np.array([0, t_max])))
for i in tqdm(range(num_curves)):
cut_curves[i, :] = default_curves[i, :-t_max]
##
# Select all points brighter than brightest_percent_lower%
num_brightest_lower = int(np.floor((cut_curves[0].shape[0] * (brightest_percent_lower/100))))
if brightest_percent_lower == 100:
num_brightest_lower = cut_curves[0].shape[0]
if brightest_percent_lower == 0:
sys.exit('Cannot use 0 as a lower bound.')
else:
indices_lower = []
for i in tqdm(range(num_curves)):
indices_lower.append(np.argpartition(cut_curves[i, :], -num_brightest_lower)[-num_brightest_lower:])
indices_lower = np.array(indices_lower)
# Select all points brighter than brightest_percent_upper%
num_brightest_upper = int(np.floor((cut_curves[0].shape[0] * (brightest_percent_upper/100))))
if brightest_percent_upper == 100:
num_brightest_upper = cut_curves[0].shape[0]
if brightest_percent_upper == 0:
indices_upper = []
for i in range(num_curves):
indices_upper.append(np.array([]))
else:
indices_upper = []
for i in tqdm(range(num_curves)):
indices_upper.append(np.argpartition(cut_curves[i, :], -num_brightest_upper)[-num_brightest_upper:])
indices_upper = np.array(indices_upper)
indices = []
for i in range(num_curves):
indices.append(np.setdiff1d(indices_lower[i], indices_upper[i], assume_unique=True))
##
# Randomly sample from the chosen indices
chosen_indices = []
for brightest_points_in_curve in tqdm(indices):
chosen_indices.append(random.sample(list(brightest_points_in_curve), num_random_points))
chosen_indices = np.array(chosen_indices, dtype=int)
# Find the smallest number that we've chosen (We print this out later)
small_points = []
for i in tqdm(range(num_curves)):
small_points.append(np.min(cut_curves[i][chosen_indices[i]]))
smallest_point = "Min log(Edd): " + str(np.min(small_points))[:6]
# Select all our points
t_examine = np.logspace(0, np.log10(t_max), np.log(t_max)*10 + 1).astype(int)
t_log = np.log10(t_examine) # Used later
t_array = np.tile(t_examine, (num_random_points, 1))
master_array = np.zeros(t_examine.shape, dtype=int)
for i in tqdm(range(num_curves)):
indices_array = np.tile(chosen_indices[i, :], (t_array.shape[1], 1)).T
indices_array = indices_array + t_array
master_array = np.vstack((default_curves[i][indices_array], master_array))
master_array = np.delete(master_array, -1, 0)
starting_vals = np.copy(master_array[:, 0])
for i in tqdm(range(master_array.shape[1])):
master_array[:, i] = master_array[:, i] - starting_vals
# Find our trends
means = []
stands = []
for i in tqdm(range(master_array.shape[1])):
means.append(np.mean(master_array[:, i]))
stands.append(np.std(master_array[:, i]))
means = np.array(means)
stands = np.array(stands)
# Get a line of best fit
best_fit = np.poly1d(np.poly1d(np.polyfit(t_log.astype(float)[1:], means.astype(float)[1:], 1)))
return (t_log, means, stands, best_fit, smallest_point)
# -
# # Main
# +
t_log05_times, means05_times, stands05_times, best_fit05_times, smallest_point05_times = extract_data(brightest_percent_lower=0.5, brightest_percent_upper=0.1, path=data_path_times)
t_log05_divided, means05_times_divided, stands05_divided, best_fit05_divided, smallest_point05_divided = extract_data(brightest_percent_lower=0.5, brightest_percent_upper=0.1, path=data_path_divided)
t_log05_gets2, means05_gets2, stands05_gets2, best_fit05_gets2, smallest_point05_gets2 = extract_data(brightest_percent_lower=0.5, brightest_percent_upper=0.1, path=data_path_gets_2)
t_log05_gets0, means05_gets0, stands05_gets0, best_fit05_gets0, smallest_point05_gets0 = extract_data(brightest_percent_lower=0.5, brightest_percent_upper=0.1, path=data_path_gets_0)
t_log05_fiducial, means05_fiducial, stands05_fiducial, best_fit05_fiducial, smallest_point05_fiducial = extract_data(brightest_percent_lower=0.5, brightest_percent_upper=0.1, path=data_path_fiducial)
# + [markdown] heading_collapsed=true
# # Delta value
# + hidden=true
Delta_value = means05[np.where(t_log05==x_Delta)][0]
print('For this set of parameters, Delta is ' + str(Delta_value - default_Delta_value) + '.')
print('Remember, a negative delta (approximately) means that the curve was steeper than the default plot of log Edd. Ratio as a function of time.')
# -
# # Graphs
# ## 10-panel plots
# + code_folding=[0] hide_input=false
# Delta Eddington ratio plots
with plt.style.context('seaborn-paper'):
fig, ax = plt.subplots(5, 2, figsize=(20, 10), sharex=True, sharey=True, gridspec_kw={'width_ratios': [1, 1], 'wspace':0, 'left':0.04, 'right':0.96, 'bottom':0.05, 'top':0.92})
fig.suptitle('Data from: ' + data_path)
ax[0,0].set_title('100%', fontsize=13)
ax[0,0].tick_params(direction='in', length=6, width=1.5)
ax[0,0].spines['top'].set_linewidth(1.5)
ax[0,0].spines['right'].set_linewidth(1.5)
ax[0,0].spines['bottom'].set_linewidth(1.5)
ax[0,0].spines['left'].set_linewidth(1.5)
ax[0,0].hlines(0, 0, t_log100[-1] + 0.2, linewidth=1, linestyle='--')
ax[0,0].errorbar(t_log100, means100, yerr=stands100, fmt='s', alpha=0.5, color=line_color)
ax[0,0].plot(t_log100[1:], best_fit100(t_log100[1:].astype(float)), ls='--', color='orange')
ax[0,0].text(0, -1, smallest_point100)
ax[0,1].set_title('50%', fontsize=13)
ax[0,1].tick_params(direction='in', length=6, width=1.5)
ax[0,1].spines['top'].set_linewidth(1.5)
ax[0,1].spines['right'].set_linewidth(1.5)
ax[0,1].spines['bottom'].set_linewidth(1.5)
ax[0,1].spines['left'].set_linewidth(1.5)
ax[0,1].hlines(0, 0, t_log50[-1] +0.2, linewidth=1, linestyle='--')
ax[0,1].errorbar(t_log50, means50, yerr=stands50, fmt='s', alpha=0.5, color=line_color)
ax[0,1].plot(t_log50[1:], best_fit50(t_log50[1:].astype(float)), ls='--', color='orange')
ax[0,1].text(0, -1, smallest_point50)
ax[1,0].set_title('10%', fontsize=13)
ax[1,0].tick_params(direction='in', length=6, width=1.5)
ax[1,0].spines['top'].set_linewidth(1.5)
ax[1,0].spines['right'].set_linewidth(1.5)
ax[1,0].spines['bottom'].set_linewidth(1.5)
ax[1,0].spines['left'].set_linewidth(1.5)
ax[1,0].hlines(0, 0, t_log10[-1] +0.2, linewidth=1, linestyle='--')
ax[1,0].errorbar(t_log10, means10, yerr=stands10, fmt='s', alpha=0.5, color=line_color)
ax[1,0].plot(t_log10[1:], best_fit10(t_log10[1:].astype(float)), ls='--', color='orange')
ax[1,0].text(0, -1, smallest_point10)
ax[1,1].set_title('5%', fontsize=13)
ax[1,1].tick_params(direction='in', length=6, width=1.5)
ax[1,1].spines['top'].set_linewidth(1.5)
ax[1,1].spines['right'].set_linewidth(1.5)
ax[1,1].spines['bottom'].set_linewidth(1.5)
ax[1,1].spines['left'].set_linewidth(1.5)
ax[1,1].hlines(0, 0, t_log5[-1] +0.2, linewidth=1, linestyle='--')
ax[1,1].errorbar(t_log5, means5, yerr=stands5, fmt='s', alpha=0.5, color=line_color)
ax[1,1].plot(t_log5[1:], best_fit5(t_log5[1:].astype(float)), ls='--', color='orange')
ax[1,1].text(0, -1, smallest_point5)
ax[2,0].set_title('1%', fontsize=13)
ax[2,0].tick_params(direction='in', length=6, width=1.5)
ax[2,0].spines['top'].set_linewidth(1.5)
ax[2,0].spines['right'].set_linewidth(1.5)
ax[2,0].spines['bottom'].set_linewidth(1.5)
ax[2,0].spines['left'].set_linewidth(1.5)
ax[2,0].hlines(0, 0, t_log1[-1] +0.2, linewidth=1, linestyle='--')
ax[2,0].errorbar(t_log1, means1, yerr=stands1, fmt='s', alpha=0.5, color=line_color)
ax[2,0].plot(t_log1[1:], best_fit1(t_log1[1:].astype(float)), ls='--', color='orange')
ax[2,0].text(0, -1, smallest_point1)
ax[2,1].set_title('0.5%', fontsize=13)
ax[2,1].tick_params(direction='in', length=6, width=1.5)
ax[2,1].spines['top'].set_linewidth(1.5)
ax[2,1].spines['right'].set_linewidth(1.5)
ax[2,1].spines['bottom'].set_linewidth(1.5)
ax[2,1].spines['left'].set_linewidth(1.5)
ax[2,1].hlines(0, 0, t_log05[-1] +0.2, linewidth=1, linestyle='--')
ax[2,1].errorbar(t_log05, means05, yerr=stands05, fmt='s', alpha=0.5, color=line_color)
ax[2,1].plot(t_log05[1:], best_fit05(t_log05[1:].astype(float)), ls='--', color='orange')
ax[2,1].text(0, -1, smallest_point05)
ax[3,0].set_title('0.1%', fontsize=13)
ax[3,0].tick_params(direction='in', length=6, width=1.5)
ax[3,0].spines['top'].set_linewidth(1.5)
ax[3,0].spines['right'].set_linewidth(1.5)
ax[3,0].spines['bottom'].set_linewidth(1.5)
ax[3,0].spines['left'].set_linewidth(1.5)
ax[3,0].hlines(0, 0, t_log01[-1] +0.2, linewidth=1, linestyle='--')
ax[3,0].errorbar(t_log01, means01, yerr=stands01, fmt='s', alpha=0.5, color=line_color)
ax[3,0].plot(t_log01[1:], best_fit01(t_log01[1:].astype(float)), ls='--', color='orange')
ax[3,0].text(0, -1, smallest_point01)
ax[3,1].set_title('0.05%', fontsize=13)
ax[3,1].tick_params(direction='in', length=6, width=1.5)
ax[3,1].spines['top'].set_linewidth(1.5)
ax[3,1].spines['right'].set_linewidth(1.5)
ax[3,1].spines['bottom'].set_linewidth(1.5)
ax[3,1].spines['left'].set_linewidth(1.5)
ax[3,1].hlines(0, 0, t_log005[-1] +0.2, linewidth=1, linestyle='--')
ax[3,1].errorbar(t_log005, means005, yerr=stands005, fmt='s', alpha=0.5, color=line_color)
ax[3,1].plot(t_log005[1:], best_fit005(t_log005[1:].astype(float)), ls='--', color='orange')
ax[3,1].text(0, -1, smallest_point005)
ax[4,0].set_title('0.02%', fontsize=13)
ax[4,0].tick_params(direction='in', length=6, width=1.5)
ax[4,0].spines['top'].set_linewidth(1.5)
ax[4,0].spines['right'].set_linewidth(1.5)
ax[4,0].spines['bottom'].set_linewidth(1.5)
ax[4,0].spines['left'].set_linewidth(1.5)
ax[4,0].set_xlabel('log(t/time units)', fontsize=13)
ax[4,0].set_ylabel('Mean $\Delta$log(Edd. Ratio)', fontsize=13)
ax[4,0].hlines(0, 0, t_log002[-1] +0.2, linewidth=1, linestyle='--')
ax[4,0].errorbar(t_log002, means002, yerr=stands002, fmt='s', alpha=0.5, color=line_color)
ax[4,0].plot(t_log002[1:], best_fit002(t_log002[1:].astype(float)), ls='--', color='orange')
ax[4,0].text(0, -1, smallest_point002)
ax[4,1].set_title('As small as possible (0.006%)', fontsize=13)
ax[4,1].tick_params(direction='in', length=6, width=1.5)
ax[4,1].spines['top'].set_linewidth(1.5)
ax[4,1].spines['right'].set_linewidth(1.5)
ax[4,1].spines['bottom'].set_linewidth(1.5)
ax[4,1].spines['left'].set_linewidth(1.5)
ax[4,1].set_xlabel('log(t/time units)', fontsize=13)
ax[4,1].hlines(0, 0, t_loginf[-1] +0.2, linewidth=1, linestyle='--')
ax[4,1].errorbar(t_loginf, meansinf, yerr=standsinf, fmt='s', alpha=0.5, color=line_color)
ax[4,1].plot(t_loginf[1:], best_fitinf(t_loginf[1:].astype(float)), ls='--', color='orange')
ax[4,1].text(0, -1, smallest_pointinf)
plt.savefig('10-panel_eddington_plot.pdf', bbox_inches='tight')
plt.show()
# + code_folding=[] hide_input=false
# Structure function plots
with plt.style.context('seaborn-paper'):
fig, ax = plt.subplots(1,
1,
figsize=(20, 10),
gridspec_kw={
'wspace': 0,
'left': 0.04,
'right': 0.96,
'bottom': 0.05,
'top': 0.92
})
ax.set_title('0.05% Structure function', fontsize=13)
ax.tick_params(direction='in', length=6, width=1.5)
ax.spines['top'].set_linewidth(1.5)
ax.spines['right'].set_linewidth(1.5)
ax.spines['bottom'].set_linewidth(1.5)
ax.spines['left'].set_linewidth(1.5)
ax.set_xlabel('log(t/time units)', fontsize=13)
ax.set_ylabel('log(SF)', fontsize=13)
ax.plot(t_log05_divided[1:],
np.log10(stands05_divided[1:]),
color='green',
label='v_bend_in divided by 10')
ax.plot(t_log05_times[1:],
np.log10(stands05_times[1:]),
color='purple',
label='v_bend_in times by 10')
ax.plot(t_log05_divided[1:],
np.log10(stands05_fiducial[1:]),
color='blue',
label='default parameters')
ax.plot(t_log05_gets0[1:],
np.log10(stands05_gets0[1:]),
color='orange',
label='a_low_in set to 0')
ax.plot(t_log05_gets2[1:],
np.log10(stands05_gets2[1:]),
color='aqua',
label='a_low_in set to 2')
plt.legend()
plt.savefig('Comparison_of_0.5percent_structure_functions.pdf',
bbox_inches='tight')
plt.show()
# +
default_curves = []
for i in tqdm(range(20 + 1)):
if i == 0:
continue # for some reason the results files start at 1 and not 0
_er_curve = np.zeros(2**24, dtype=float)
_er_curve = np.fromfile(data_path_gets_0.format(str(i)))
default_curves.append(_er_curve)
default_curves = np.array(default_curves)
default_curves = np.log10(default_curves) # Move everything into the log domain
# -
plt.plot(default_curves[0][5000:6100])
plt.show()
# + [markdown] heading_collapsed=true
# ## 4-Line plots
# + hidden=true
# PSD regime determination plot
with plt.style.context('seaborn-paper'):
fig, ax = plt.subplots(1, figsize=(20, 10))
ax.set_title('Mean change in log(Edd. Ratio) as a function of chosen percent. Data from: '+ data_path, fontsize=13)
ax.tick_params(direction='in', length=6, width=1.5)
ax.spines['top'].set_linewidth(1.5)
ax.spines['right'].set_linewidth(1.5)
ax.spines['bottom'].set_linewidth(1.5)
ax.spines['left'].set_linewidth(1.5)
ax.set_xlabel('log(% total data)', fontsize=18)
ax.set_xlim(x[0], x[-1])
ax.set_ylabel('Mean $\Delta$log(Edd. Ratio)', fontsize=18)
ax.plot(x, log_t_1, label='log(t/time units) = 1', marker='o', markersize=10)
ax.plot(x, log_t_2, label='log(t/time units) = 2', marker='o', markersize=10)
ax.plot(x, log_t_3, label='log(t/time units) = 3', marker='o', markersize=10)
ax.plot(x, log_t_4, label='log(t/time units) = 4', marker='o', markersize=10)
ax.legend(prop={'size':18})
plt.savefig('4-line_eddington_plot.pdf', bbox_inches='tight')
plt.show()
# + hidden=true
# Structure functions in a strange space
with plt.style.context('seaborn-paper'):
fig, ax = plt.subplots(1, figsize=(20, 10))
ax.set_title('log(SF) as a function of log(chosen percent). Data from: ' + data_path, fontsize=13)
ax.tick_params(direction='in', length=6, width=1.5)
ax.spines['top'].set_linewidth(1.5)
ax.spines['right'].set_linewidth(1.5)
ax.spines['bottom'].set_linewidth(1.5)
ax.spines['left'].set_linewidth(1.5)
ax.set_xlabel('log(% total data)', fontsize=18)
ax.set_xlim(x[0], x[-1])
ax.set_ylabel('log(SF)', fontsize=18)
ax.plot(x, log_t_1_stands, label='log(t/time units) = 1', marker='o', markersize=10)
ax.plot(x, log_t_2_stands, label='log(t/time units) = 2', marker='o', markersize=10)
ax.plot(x, log_t_3_stands, label='log(t/time units) = 3', marker='o', markersize=10)
ax.plot(x, log_t_4_stands, label='log(t/time units) = 4', marker='o', markersize=10)
ax.legend(prop={'size':18})
plt.savefig('4-line_SF_plot.pdf', bbox_inches='tight')
plt.show()
| Simulations/visualization/misc_graphs/Factory2.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from em_examples.DCWidget_Overburden_2_5D import *
from IPython.display import display
# %matplotlib inline
from matplotlib import rcParams
rcParams['font.size'] = 14
# # Effects of a highly Conductive surface layer
# # Purpose
#
# For a direct current resistivity (DCR) survey, currents are injected to the earth, and flow.
# Depending upon the conductivity contrast current flow in the earth will be distorted, and these changes
# can be measurable on the sufurface electrodes.
# Here, we focus on a bloc target embedded in a halfspace below a highly conductive surface layer, and investigate what are happening in the earth when static currents are injected. The conductive layer will also impact the illumination of the target (conductor or resistor).
# By investigating changes in currents, electric fields, potential, and charges upon different geometry, Tx and Rx location, we understand geometric effects of the conductive layer for DCR survey.
# # Setup
# <img src=https://github.com/geoscixyz/em_apps/blob/master/images/Dcapps_Overburden_draw.png?raw=true>
# # Question
#
# - How does the Target affect the apparent resistivity? Is there a difference if you add or remove the target?
# ## Overburden model
# - **survey**: Type of survey
# - **A**: Electrode A (+) location
# - **B**: Electrode B (-) location
# - **M**: Electrode A (+) location
# - **N**: Electrode B (-) location
# - **$\rho_{1}$**: Resistivity of the half-space
# - **$\rho_{2}$**: Resistivity of the overburden
# - **$\rho_{3}$**: Resistivity of the target
# - **Overburden_thick**: thickness of the overburden
# - **target_thick**: thickness of the target
# - **target_wide**: width of the target
# - **whichprimary**: which model to consider as primary: either uniform background or Overburden model
# - **ellips_a**: x radius of ellipse
# - **ellips_b**: z radius of ellipse
# - **xc**: x location of ellipse center
# - **zc**: z location of ellipse center
# - **Field**: Field to visualize
# - **Type**: which part of the field
# - **Scale**: Linear or Log Scale visualization
#
# ### **When typing modifications to values, do not forget to PRESS ENTER**
# ### **Do not forget to hit Run Interact to update the figure after you made modifications**
app = valley_app()
| notebooks/eletrorresistividade/DC_Overburden_2_5D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Is <NAME> finally better than <NAME>?
# + run_control={"frozen": false, "read_only": false}
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
sns.set_style('whitegrid')
# +
jordan = 'http://espn.go.com/nba/player/stats/_/id/1035/michael-jordan'
lebron = 'http://www.espn.com/nba/player/stats/_/id/1966/lebron-james'
tab1 = pd.read_html(jordan, header=1)
tab2 = pd.read_html(lebron, header=1)
# -
avgs_jordan = tab1[1][0:-1]
avgs_lebron = tab2[1][0:-1]
combined = pd.concat([avgs_jordan['FG%'], avgs_lebron['FG%']],axis='columns')
combined.columns = ['Jordan', 'Lebron']
combined[['Jordan', 'Lebron']] = combined[['Jordan', 'Lebron']].astype(float)
combined
# ### Since we care about how "efficient" a player is, season averages should be used
combined.describe()
# **There you have it - Lebron has a higher overal FG% than Jordan**
# **Let's chart the season avg FG%**
fig, ax = plt.subplots(figsize=(20, 10))
combined.plot(ax=ax, linewidth=4)
ax.set_title('Season Average Field Goal Percentages', fontsize=30, weight='bold')
ax.set_ylim(bottom=0, top=0.8)
ax.set_xlabel('Number of Seasons')
plt.show()
# It is incredible that Lebron has improved with age.
| jupyter_notebooks/pandas/read_html.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
from joblib import load, dump
trained = load("pipe.joblib")
# -
lr = trained.steps[1][1]
class EvilMetaEstimator:
def __init__(self, orig_model):
self.orig_model = orig_model
def predict(self, X):
print("fooled you!")
return self.orig_model.predict(X)
trained.steps[-1] = ("LogisticRegression", EvilMetaEstimator(lr))
trained.predict(["hello"])
dump(trained, "pipe-evil.joblib")
# +
from joblib import load
load("pipe-evil.joblib").predict(["hello"])
# -
# ## We're not out of the woods yet!
#
# Technically, the above attack "fails" when we have a new environment that does not have the `EvilMetaEstimator` defined.
#
# But that doesn't mean that there might be other tomfoolery!
class EvilThing:
def predict(self, X):
print("fooled you!")
return [1 for _ in X]
trained = load("pipe.joblib")
evil_pipe = EvilThing()
# +
from joblib import dump, load
dump(evil_pipe, "pipe-evil.joblib")
# +
from joblib import dump, load
pipe_loaded = load("pipe-evil.joblib")
pipe_loaded.predict(["hello"])
# +
import hashlib
def calc_checksum(path):
md5_hash = hashlib.md5()
with open(path, "rb") as f:
content = f.read()
md5_hash.update(content)
digest = md5_hash.hexdigest()
print(digest)
calc_checksum("pipe.joblib")
calc_checksum("pipe-evil.joblib")
| beware-the-evil.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # EXTRAS
# # Interactivity
# matplotlib has it's own set of widgets that you can use, but recently, Jupyter / Ipython gained the interact() function
# (see http://ipywidgets.readthedocs.io/en/latest/examples/Using%20Interact.html )
#
# Note: something changed in mpl 2.0 that we now need a `plt.show()` here. See:
# https://github.com/jupyter-widgets/ipywidgets/issues/1179
import matplotlib.pyplot as plt
import numpy as np
from ipywidgets import interact
plt.figure()
x = np.linspace(0,1,100)
def plotsin(f):
plt.plot(x, np.sin(2*np.pi*x*f))
plt.show()
interact(plotsin, f=(1,10,0.1))
# +
# interactive histogram
def hist(N, sigma):
r = sigma*np.random.randn(N)
plt.hist(r, density=True, bins=20)
x = np.linspace(-5,5,200)
plt.plot(x, np.exp(-x**2/(2*sigma**2))/(sigma*np.sqrt(2.0*np.pi)),
c="r", lw=2)
plt.xlabel("x")
plt.show()
interact(hist, N=(100,10000,10), sigma=(0.5,5,0.1))
# -
# # Final fun
# if you want to make things look hand-drawn in the style of xkcd, rerun these examples after doing
# plt.xkcd()
plt.xkcd()
| day2/13. matplotlib - extras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 1. Natural Language Toolkit
#
# ### What is Natural Language Toolkit?
#
# Natural Language Toolkit (or NLTK for short) is a group of libraries and programs used for symbolic and statistical natural language processing.
#
# As it has been mentioned previously, data for NLP model has to be preprocessed prior to the training procedure. Such preprocessing operations could include, converting string-type data to numerical data, performing semantical analysis, etc. All of these (and many more) operations can be simply implemented using the NLTK library.
#
# In the following section, we will look at the most relevant functions.
#Setting up
# !pip install nltk
# +
import nltk
nltk.download()
# -
# ### Tokenization
#
# 
#
# Prior to processing textual data, we should first tokenize it. In other words, we should split it into smaller parts (sentences to words, paragraphs to sentences), as it reduces further processing time.
#
# #### Sentence tokenization
#
# As the name might suggest, in the sentence tokenization we aim to split groups of sentences/paragraphs to shorter sentences.
# +
from nltk.tokenize import sent_tokenize
Text = "Natural language processing (NLP) refers to the branch of computer science. To be more specific, the branch of artificial intelligence. It is concerned with giving computers the ability to understand text and spoken words in much the same way human beings can."
sent_tokenize = sent_tokenize(Text)
sent_tokenize
# -
# #### Word tokenization
#
# In contrast to sentence tokenization, the goal of word tokenization is to divide textual data into individual words.
# +
word_tokenize = nltk.word_tokenize(Text)
word_tokenize
# -
# ### Stemming
#
# 
#
# Stemming is the process of producing morphological variants using the given base word. The usage of stemming algorithms allows multiple word variations to share a *same* meaning / attribute (for instance, 'fish', 'fishing' are variants of the base word 'fish').
#
# In many cases, stemming algorithm tends to cut the end of the word until the base word is found, as it works in the most cases.
#
# Let's look at one of the most common stemming tools implementation - **Porter Stemmer**.
# +
from nltk.stem.porter import *
stemmer = PorterStemmer()
words = ['fishing', 'believes', 'writes', 'loving', 'cats']
for word in words:
print(word + '-----' + stemmer.stem(word))
# -
# ### Lemmatization
#
# In contrast to stemming, lemmatization does not apply simple *'word end cutting'* and rather considers a full vocabulary to apply a morphological word analysis. The simple examples of such reduction could be interpretting 'was' as one of the 'be' or 'mice' as 'mouse'.
#
# As a result, lemmatization is considered to be more informative than simple stemming, but also, slower.
# +
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer();
words = ['believes', 'lives', 'mice']
for word in words:
print(word + '-----' + lemmatizer.lemmatize(word))
# -
# On the other hand, same words can be interpreted differently based on the part of speech. To correctly interpret such words, we can specify the ```pos``` argument in the lemmatizer function.
# +
#crossing as adjective
print(lemmatizer.lemmatize('crossing', pos = 'a'))
#crossing as verb
print(lemmatizer.lemmatize('crossing', pos = 'v'))
# -
# ### Stopwords
#
# Stopwords are the words in any language which do not add much meaning to a sentence, thus can be ignored without lossing accuracy. The examples of such words could be 'is', 'the', 'at', however, each NLP tool has a different list of stopwords.
# +
from nltk.corpus import stopwords
print(stopwords.words('english'))
# -
# As these words do not carry much information, they can be removed, however, this depends from case to case.
#
# Generally, NLP models for text classification, sentiment analysis, spam classification (and so on) would require the removal of stop words.
#
# On the other hand, if we are dealing with the translation models, stopwords should be left as they might provide some contextual information.
#
# Let's remove stopwords from the previously analyzed NLP definition.
# +
from nltk.corpus import stopwords
stopwords = set(stopwords.words('english'))
Text = "Natural language processing (NLP) refers to the branch of computer science. To be more specific, the branch of artificial intelligence. It is concerned with giving computers the ability to understand text and spoken words in much the same way human beings can."
words = nltk.word_tokenize(Text)
filtered_words = []
for word in words:
if word not in stopwords:
filtered_words.append(word)
print(filtered_words)
| Week-07/1_Natural Language Toolkit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="lux5l2hpRsPE"
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
import statsmodels.formula.api as smf
# + [markdown] id="oXxoFK5RXCxN"
# ### Functional analysis of transcriptomics data from SARS-CoV-2 infected cell lines
#
# Data is from [GSE147507](https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE147507)
#
# You need to upload the data files (from Google Drive link) to the *content* directory of your Colab enviroment. If you run this localy, you have to change the path to files according to your enviroment.
# + id="Gr6uQrl3Xupi"
data = pd.read_csv('/content/GSE147507_RawReadCounts_Human.csv',
sep=',', header=0, index_col=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="tiawhlvFz5w6" outputId="05ba5e5a-1a0a-4a28-c779-7c3589c90409"
data.head()
# + id="MoTd3MA10BQh"
meta = pd.read_csv('/content/meta.csv',
sep=',', header=0, index_col=0)
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="Vns8h9ij0JZ6" outputId="6b1d04c2-4999-4753-d57b-5de0ac01c2c9"
meta.head()
# + colab={"base_uri": "https://localhost:8080/"} id="HZA626gR0UwB" outputId="fc80ea12-d2c6-4002-cac0-789972f67bd5"
np.sum(meta.index != data.columns)
# + colab={"base_uri": "https://localhost:8080/"} id="IkAwpykl0i5Q" outputId="d68f86cf-5b43-4054-90a0-22d0f56b10b6"
meta['Cell'].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="dADWtg-m0t2u" outputId="da90d360-7184-43f2-ac8c-1acc5d93edd2"
meta['Treatment'].value_counts()
# + [markdown] id="VHlZ3CF0Xpdx"
# ### Basic exploratory analysis with PCA
# + id="5hSZ328uXQVl" colab={"base_uri": "https://localhost:8080/", "height": 333} outputId="af3de836-2ffc-4024-9b1a-a9fcd7c2a2ce"
plt.hist(data.loc['WASH7P'])
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="DPCrOnk51SQl" outputId="2fbece29-6ba0-4f36-be34-7c60b6cdabfa"
sns.scatterplot(data.loc['WASH7P'], data.loc['STAT1'])
# + colab={"base_uri": "https://localhost:8080/", "height": 333} id="nt6RhIpC1mIr" outputId="905f3f6b-17a7-4c6f-dfc3-3b2ce2d95cab"
plt.hist(data.loc['STAT1'])
# + id="eWedqXTd1y9O"
data_log = np.log2(data+1)
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="P7gr-AKx2Cp1" outputId="99cda2b9-9cbb-4a27-d5e7-585aaf246a2b"
plt.hist(data_log.loc['STAT1'])
# + colab={"base_uri": "https://localhost:8080/"} id="uXZab78l2Mob" outputId="3dd11aec-5475-4a99-f00a-3838d574e0cf"
data.shape
# + id="n-i5PT5O2hxK"
model = PCA(2)
# + id="IcxJcBK52qmG"
data_pca = model.fit_transform(data_log.T)
# + id="Tk_jkvpW24NI"
data_pca = pd.DataFrame(data_pca, index=data_log.columns, columns=['PC1', 'PC2'])
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="mGdrt-XP3K7P" outputId="bf545068-b078-409e-a565-a22a615f09a9"
data_pca.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="Lla2VIhZ3NpE" outputId="a827ad37-42db-4b57-e3bc-f72cde9b03a3"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Cell'])
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="6Wo6FA3s3zca" outputId="b9b26f8a-0ed7-40fc-d6df-ddfcc8fb24bb"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Treatment'])
# + id="oCilht5C3-A2"
meta['Total_count'] = data.sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="yLtuGDjE4L9p" outputId="83ff5491-545d-4722-a097-f8781ba296c4"
meta.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="ryQG-N9l4OWn" outputId="6bb760e0-1bfb-4880-fa7d-6d4aa0de365f"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Total_count'])
# + id="BYcBOpYj4hU1"
### normalizálás
data_norm = data / data.sum()
# + colab={"base_uri": "https://localhost:8080/"} id="Cyv75c9W4zPl" outputId="c8d9f4e3-e019-44c6-be5d-8bf57de5e162"
data_norm.sum()
# + id="Rvymnr0947Gt"
data_lognorm = np.log2(data_norm+1)
# + id="nrGlKlXU5BZX"
model = PCA(2)
data_pca = model.fit_transform(data_lognorm.T)
data_pca = pd.DataFrame(data_pca, index=data_log.columns, columns=['PC1', 'PC2'])
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="cX6jjo8h5OFq" outputId="642213c1-4522-47fe-aace-a0bec6ab792c"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Cell'])
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="UXGcsG665TCX" outputId="feeea9dd-f3f3-4065-f2ae-a69c758c08d4"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Treatment'])
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="ChBDGjKz5d9w" outputId="76f81d98-edc3-47ce-c9e6-925632390a9b"
sns.scatterplot(x=data_pca['PC1'], y=data_pca['PC2'], hue=meta['Total_count'])
# + id="7J3PwI2R5onY"
### mock / SARS-CoV-2
fil = np.in1d(meta['Treatment'], ['Mock', 'SARS-CoV-2'])
# + id="UCsHFRN25-Oq"
meta = meta[fil]
data_lognorm = data_lognorm[meta.index]
# + colab={"base_uri": "https://localhost:8080/"} id="Ej9hvt7F6MjN" outputId="2b6bf4aa-c788-4c9b-ccbb-eb530ad89af1"
meta.shape
# + [markdown] id="8hS4F5HNX1Xg"
# ### Basics of statistical modelling
# + id="fzvPxywOYtRF"
toy = pd.DataFrame(index=['A','B','C','D','E','F'], columns=['Meas', 'Group'])
toy['Group'] = ['A','A','A','B','B','B']
np.random.seed(310)
toy.loc[['A','B','C'], 'Meas'] = np.random.normal(0, 1, size=3)
toy.loc[['D','E','F'], 'Meas'] = np.random.normal(2, 1, size=3)
# + id="QkYQkbhy7zzy"
toy['Meas'] = toy['Meas'].astype(float)
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="yisyHbjV_YQM" outputId="fa473278-4e90-414a-d87c-4905e1e11692"
toy
# + id="WJfYRQUj7HKD"
from scipy.stats import ttest_ind
# + colab={"base_uri": "https://localhost:8080/"} id="nQSwfGPX73zt" outputId="a44d0843-9cee-4b31-a449-bd9d6ccda7b7"
ttest_ind(toy.loc[['A','B','C'], 'Meas'], toy.loc[['D','E','F'], 'Meas'])
# + id="qU8jelye733l"
model = smf.ols('Meas ~ Group', data=toy).fit()
# + colab={"base_uri": "https://localhost:8080/"} id="q3OwWOMv736M" outputId="424cc1cd-2faa-417c-810a-4c8272ee266c"
model.params
# + colab={"base_uri": "https://localhost:8080/"} id="De6rbecM_GoW" outputId="61b9327f-bf8a-4411-a592-4b72c914a89f"
model.pvalues
# + id="xjTKYbqDABCN"
### genek
# + id="6CrlFK8BAMfA"
### remove all 0s
fil = data_lognorm.std(1) != 0.0
data_lognorm = data_lognorm[fil]
# + id="MA0wIwOKAmTh"
### sorok a mintak
data_stat = pd.concat([data_lognorm.T, meta[['Cell','Treatment']]], 1)
# + id="soiYGhkPBEVz"
model = smf.ols('STAT1 ~ Treatment', data=data_stat).fit()
# + colab={"base_uri": "https://localhost:8080/"} id="0N_CeuOABW6N" outputId="3b3bcbb0-2f23-46db-a0eb-298395082daf"
model.params
# + colab={"base_uri": "https://localhost:8080/"} id="k9FyZcUtBW9D" outputId="f9bc86ad-3135-475c-f2ee-9f60d4bb937b"
model.pvalues
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="W48-bJfmBXA0" outputId="a71f7dcb-1f01-482e-b4a2-f1506fdb6082"
sns.boxplot(x=data_stat['Treatment'], y=data_stat['STAT1'])
# + id="dXt6uwalBXDt"
model = smf.ols('STAT1 ~ Treatment + Cell + Treatment:Cell', data=data_stat).fit()
# + colab={"base_uri": "https://localhost:8080/"} id="x0qGf0dwCJqU" outputId="bfb1d3ae-0814-4a81-c457-475b20bf5dcb"
model.pvalues
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="ad1hD6xYCRE9" outputId="cc1b4b83-a89d-4862-c02d-953de20e4f5f"
sns.boxplot(x=data_stat['Treatment'], y=data_stat['STAT1'], hue=meta['Cell'])
# + colab={"base_uri": "https://localhost:8080/", "height": 386} id="OIS3N5HRDDFd" outputId="576d8a83-e5d5-464e-b5d2-eb36a61ed07c"
data_stat.head()
# + id="v7ISjKvlELEy"
good_columns = [x for x in data_stat.columns if ('-' not in x)&('.' not in x)]
# + id="9h3zyNipEU-Z"
data_stat = data_stat[good_columns]
# + id="wycYN0tbDohJ"
results = pd.DataFrame(index=data_stat.columns[0:-2],
columns=['Pval', 'Param'])
# + id="IY8SMpyyC6x2"
### takes a few minutes
for gene in data_stat.columns[0:-2]:
model = smf.ols(gene + ' ~ Cell + Treatment', data=data_stat).fit()
results.loc[gene] = model.pvalues['Treatment[T.SARS-CoV-2]'], model.params['Treatment[T.SARS-CoV-2]']
# + colab={"base_uri": "https://localhost:8080/"} id="SGpjdm7eE1jc" outputId="514cb5e9-a088-4073-ec8d-992bf50cfa4c"
#### you can see some immune related genes in the most significantly diferentially expressed genes
results['Pval'].sort_values().head(20)
# + [markdown] id="bdSTGNh0YvaT"
# ### Homework
#
# * Calculate coefficients (for SARS-CoV-2 treatment) for Calu-3 and A549 cells, and plot them (scatter plot) against each other. This will show us how similar are the response of these cells to infection. You will have to filter for these data (have 2 DataFrames, containing Mock and SARS-CoV-2 infected samples, and either Calu-3 or A549 cell lines), and run a statistical model with only 'Treatment' factor (basically it is a t-test).
# * Please upload this notbook (your_name.ipynb) to the Week6 folder (you should have write access to this, if not please let me know)
# * install [DESeq2](https://bioconductor.org/packages/release/bioc/html/DESeq2.html) library in R.
# + id="amrEjG6CYwxP"
| code/Week6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy.stats as ss
import matplotlib.pyplot as plt
# ML from scratch
class hmm:
def __init__(self,n,A,p,mu,sig):
self.n = int(n)
self.A = np.asarray(A)
self.p = np.asarray(p)
self.mu = mu
self.sig = sig
self.h = [None]
self.x = [None]
self.steps = 0
def sample(self,steps):
self.steps = steps
self.h = np.random.choice(self.n, size=steps, p=list(self.p))
self.x = np.random.normal(loc=self.mu[self.h[0]],scale=self.sig[self.h[0]],size=steps)
for i in range(steps-1):
self.h[i+1] = np.random.choice(self.n,size=1,p=list(self.A[:,self.h[i]]))
self.x[i+1] = np.random.normal(loc=self.mu[self.h[i+1]],scale=self.sig[self.h[i+1]],size=1)
return self.h, self.x
def fit(self,x_obs,delta_tol,iter_max):
# initialized your marginals for hidden state
self.steps = len(x_obs)
self.x = x_obs
self.h_marginal = np.ones((self.n,self.steps))
delta_curr = np.ones(1)
iter_curr = 0
while delta_curr>delta_tol and iter_curr<iter_max:
mu_old = self.mu
# E-step:
self.h_marginal = self.E_step(self.x,self.A, self.p, self.mu, self.sig)
# M-step
self.A, self.p, self.mu, self.sig = self.M_step(self.x,self.h_marginal)
# check how much the estimated mean has moved increase counter
delta_curr = np.sum(np.abs(self.mu-mu_old))
iter_curr+=1
return self, iter_curr
def infer(self,observed_x_seq):
hidden = Viterbi()
return hidden
def forward(self,x,A,p,mu,sig):
alpha = np.zeros((self.n,self.steps))
for i in range(self.n):
alpha[i,0] = p[i]*ss.norm.pdf(x[0],mu[i],sig[i])
for t in range(self.steps-1):
for i in range(self.n):
for j in range(self.n):
alpha[i,t+1] += alpha[j,t]*A[i,j]*ss.norm.pdf(x[t+1],mu[i],sig[i])
return alpha
def backward(self,x,A,p,mu,sig):
beta = np.zeros((self.n,self.steps))
beta[:,-1] = 1
for t in range(self.steps-2,-1,-1):
for i in range(self.n):
for j in range(self.n):
beta[i,t] += beta[j,t+1]*A[j,i]*ss.norm.pdf(x[t+1],mu[j],sig[j])
return beta
def E_step(self,x,A_old,p_old,mu_old,sig_old):
alpha = self.forward(x,A_old,p_old,mu_old,sig_old) # this is an NxT matrix
beta = self.backward(x,A_old,p_old,mu_old,sig_old) # this is an NxT matrix
h_marginal = np.multiply(alpha,beta)
h_marginal = np.divide(h_marginal,np.tile(np.sum(h_marginal,axis=0),(self.n,1)))
return h_marginal
def M_step(self,x,h_marginal):
A = np.zeros((self.n,self.n))
for i in range(self.n):
for j in range(self.n):
for t in range(self.steps-1):
A[i,j] += h_marginal[j,t]*h_marginal[i,t+1]
# normalizethe columns
for i in range(self.n):
A[:,i] = A[:,i]/np.sum(A[:,i])
# pick initial
p = h_marginal[:,0]
mu = np.divide(np.matmul(h_marginal,x),np.sum(h_marginal,axis=1))
for i in range(self.n):
sig[i] = np.sqrt(np.matmul(h_marginal[i,:],(x-mu[i])**2)/np.sum(h_marginal[i,:]))
return A,p,mu,sig
# some generic helper funcitons
def normalpdf(x,mu,sig):
return 1/np.sqrt(2*np.pi*sig**2)*np.exp(-(x-mu)**2/2/sig**2)
# -
A = [[0.9,0.1],[0.1,0.9]]
p = [0.5,0.5]
mu = [0,1]
sig = [0.2,0.2]
T = 500;
myhmm = hmm(2,A,p,mu,sig)
h,x_obs = myhmm.sample(T)
plt.plot(x_obs)
plt.plot(h)
plt.show()
# now we try to learn this sequence of randomly generated observation
# intitial guesses
A0 = [[0.8,0.2],[0.2,0.8]]
p0 = [0.5,0.5]
mu0 = [0.1,0.9]
sig0 = [0.1,0.1]
tol_mu = 1e-4
iter_max = 1000
testhmm = hmm(2,A0,p0,mu0,sig0)
testhmm.sample(T)
h_marginals0= testhmm.E_step(x_obs,np.asarray(A0),np.asarray(p0),np.asarray(mu0),np.asarray(sig0))
A1,p1,mu1,sig1 = testhmm.M_step(x_obs,h_marginals0)
print(A1)
print(p1)
h_marginals1= testhmm.E_step(x_obs,np.asarray(A1),np.asarray(p1),np.asarray(mu1),np.asarray(sig1))
A2,p2,mu2,sig2 = testhmm.M_step(x_obs,h_marginals1)
print(A2)
plt.plot(h_marginals0.T)
plt.show()
#hmmfit,iters = testhmm.fit(x_obs,tol_mu,iter_max)
#print(iters)
#print(hmmfit.A)
#print(hmmfit.p)
#print(hmmfit.mu)
#print(hmmfit.sig)
| HMM/hmm_CS_dysfunctional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vivekwisdom/NLP/blob/develop/Sentiment%20Classification/Sentiment_Classification_CNN_LSTM_keras.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1uS-l6IMPj85" colab_type="text"
# [Source]('https://keras.io/examples/imdb_cnn_lstm/')
# + id="Ok0INx6KRAui" colab_type="code" colab={}
pip install seqeval version_information
# + id="NMLVps-6Pj9A" colab_type="code" outputId="b5651144-7c67-464b-b6f7-10233cc79ff9" colab={"base_uri": "https://localhost:8080/", "height": 280}
from __future__ import print_function
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import tensorflow as tf
import tensorflow_hub as hub
from keras import backend as K
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import LSTM, Embedding, Dense, TimeDistributed, Dropout, Bidirectional, Lambda, Activation, Conv1D, MaxPooling1D
# from seqeval.metrics import precision_score, recall_score, f1_score, classification_report
from keras.datasets import imdb
print("GPU is", "available" if tf.test.is_gpu_available() else "NOT AVAILABLE")
# %load_ext version_information
# %version_information pandas, numpy, keras, tensorflow, sklearn
# + id="uL9-DhpcPj9I" colab_type="code" colab={}
# Embedding
max_features = 20000
maxlen = 500
embedding_size = 256
# + id="eAY6K6RdPj9N" colab_type="code" colab={}
# Convolution
kernel_size = 2
filters = 64
pool_size = 2
# + id="BFbOxwnjPj9T" colab_type="code" colab={}
# LSTM
lstm_output_size = 25
# + id="Hefv_YtjPj9Y" colab_type="code" colab={}
# Training
batch_size = 100 #batch_size is highly sensitive. Only 2 epochs are needed as the dataset is very small.
epochs = 5
# + id="2duz5nKcPj9q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="b6738afc-878f-43dc-8dcd-5cfea3101cc7"
print('Loading Full data...')
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=max_features)
print(len(x_train), 'train sequences')
print(len(x_test), 'test sequences')
# + [markdown] id="F8B_np_A3gvS" colab_type="text"
# # Simple EDA on Data
# + id="eLSntYHw3mOm" colab_type="code" colab={}
X = np.concatenate((x_train, x_test), axis=0)
y = np.concatenate((y_train, y_test), axis=0)
# + id="U-eDXqs23wob" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="339f243b-d2a4-4ea8-da4b-2f9eb1338a90"
# summarize size
print("Training data: ")
print(X.shape)
print(y.shape)
# + id="RPUcN6rG3w00" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4a7adfb9-891c-4b57-f25e-31a6004ed16e"
# Summarize number of classes
print("Classes: ")
print(np.unique(y))
# + id="0GDJPpj63w3n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="ebb2a4b9-e994-4f9c-9bf3-4e4fdd20f2a1"
# Summarize number of words
print("Number of words: ")
print(len(np.unique(np.hstack(X))))
# + id="98iatNxx3w6i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="ca836929-9fa8-474c-c172-7b4e4ce55ea9"
# Summarize review length
print("Review length: ")
result = [len(x) for x in X]
print("Mean %.2f words (%f) and median is (%f)" % (np.mean(result), np.std(result), np.median(result)))
# plot review length
# plt.hist(result)
plt.boxplot(result)
plt.show()
# + id="34dUUJkxPj9s" colab_type="code" outputId="f1019f60-ae83-4910-c121-c5fcdb0a37ef" colab={"base_uri": "https://localhost:8080/", "height": 34}
print('Pad sequences (samples x time)')
x_train = sequence.pad_sequences(x_train, maxlen=maxlen)
x_test = sequence.pad_sequences(x_test, maxlen=maxlen)
# + id="opRB4rNfPj_1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e160554c-341c-4a01-8aab-49ecc20985e7"
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
# + [markdown] id="945GQiJF60j_" colab_type="text"
# ## Build Model
# + id="qvB8TYpw6SRr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="9b09d1ae-d8a3-4fe1-8504-d8df9f39eb31"
print('Build model...')
model = Sequential()
model.add(Embedding(max_features, embedding_size, input_length=maxlen))
model.add(Dropout(0.25))
model.add(Conv1D(filters,
kernel_size,
padding='valid',
activation='relu',
strides=1))
model.add(MaxPooling1D(pool_size=pool_size))
model.add(Bidirectional(LSTM(lstm_output_size)))
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.summary()
# + [markdown] id="I5aCpx457I4D" colab_type="text"
# ## Compile the model
# + id="kAT1uUhv7Lsh" colab_type="code" colab={}
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + id="6aRJ508p7MxJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="2f705756-859c-40e9-fa88-4256db2f6059"
print('Train...')
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test))
# + id="bc9Gbi4J8-a_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="96dae2be-0270-4334-92b3-9114293eec53"
score, acc = model.evaluate(x_test, y_test, batch_size=batch_size)
print('Test score:', score)
print('Test accuracy:', acc)
# + id="Srz-91qtF23l" colab_type="code" colab={}
#Will try agaun later to implement other papers in https://paperswithcode.com/sota/sentiment-analysis-on-imdb
# + id="WBmUZY6YouhA" colab_type="code" colab={}
| Sentiment Classification/Sentiment_Classification_CNN_LSTM_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ydCMp2Pf-wzo" colab_type="text"
# ### Install Dependencies
# + id="_1Ui5cxV-1D2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="48655ce4-1312-4c4c-b188-3403334e775b"
# !pip install kaggle contractions
# + [markdown] id="FSBcEaHWAdQC" colab_type="text"
# ### Import Dependencies
# + id="FZ2wOqcxAgKl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="1d6c6a1f-d336-4b4a-efc2-463d58926335"
import os
os.environ['KAGGLE_USERNAME'] = 'spyrosmouselinos'
os.environ['KAGGLE_KEY'] = 'a907fb69eab07900ccb6e1f2874fd343'
import re
import contractions
import numpy as np
import pandas as pd
import nltk
nltk.download('wordnet')
nltk.download('punkt')
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
from kaggle.api.kaggle_api_extended import KaggleApi
from zipfile import ZipFile
# + [markdown] id="GHbd7wGM_Bc6" colab_type="text"
# ### Connect to Kaggle API and download dataset
# + id="-LS71Vxh_Gq5" colab_type="code" colab={}
api = KaggleApi()
api.authenticate()
if not os.path.exists('train.csv'):
api.competition_download_file('twitter-sentiment-analysis2','train.csv')
zf = ZipFile('train.csv.zip', 'r')
zf.extractall('./')
zf.close()
os.remove('train.csv.zip')
data = pd.read_csv('train.csv', delimiter=',', encoding='latin-1')
data.drop(columns='ItemID', inplace=True)
# + [markdown] id="TxY7Jo3ZAzZC" colab_type="text"
# ### Text Preprocessing / Splitting
# + id="Gp9Gzefs-Np_" colab_type="code" colab={}
data['SentimentText'] = data['SentimentText'].str.lower()
# + id="_r-3oN8e-NqE" colab_type="code" colab={}
def convert_emojis(sentence):
# Converts known emojis to sentiment words
# :) --> happyface
# (: --> happyface
# :p --> happyface
# :P --> happyface
# ;p --> happyface
# :] --> happyface
# [: --> happyface
# :o --> surpriseface
# :O --> surpriseface
# :( --> sadface
# ): --> sadface
# :'( --> sadface
# :S --> sadface
# :\ --> sadface
# :[ --> sadface
# ]: ---> sadface
return
def preprocess(sentence):
# Convert to Lower Case
sentence = sentence.lower()
# Replace Contractions
sentence = contractions.fix(sentence, slang=True)
# Remove Links
sentence = re.sub(r'(http|https|www)\S+', ' ', sentence)
# Remove usernames
sentence = re.sub(r'@\w+', ' ', sentence)
# Remove non-word characters
sentence = re.sub(r'\W', ' ', sentence)
# Remove underscores
sentence = re.sub(r'[-_]', ' ', sentence)
# Remove numbers
sentence = re.sub(r'[0-9]', ' ', sentence)
# Remove all single characters
sentence = re.sub(r'\s+[a-zA-Z]\s+', ' ', sentence)
# Substituting multiple spaces with single space
sentence = re.sub(r'\s+', ' ', sentence, flags=re.I)
# Lemmatization
sentence = sentence.split()
sentence = [lemmatizer.lemmatize(word) for word in sentence]
sentence = ' '.join(sentence)
return sentence
# + id="UT9_njYN-NqJ" colab_type="code" colab={}
data['SentimentText'] = data['SentimentText'].apply(lambda x : preprocess(x))
# + id="k3eY0UCK-NqR" colab_type="code" colab={}
### Convert into lists and split
y = data['Sentiment'].values
x = data['SentimentText'].values
# + id="q47I6nyc-NqV" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3, stratify=y, shuffle=True)
# + [markdown] id="8CncrlPtIs2w" colab_type="text"
# ### Features
# * TF
# * TF-IDF
#
# ### Classifiers
# * Dummy
# * Logistic Regression
# * Logistic Regression SGD
# * Naive Bayes
# * KNN
# * SVM
# * MLP
# * Random Forest
# + id="fAULAJOmN3Be" colab_type="code" colab={}
from sklearn.pipeline import Pipeline
from sklearn.naive_bayes import MultinomialNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.decomposition import TruncatedSVD
from sklearn.feature_extraction.text import TfidfTransformer, CountVectorizer
from sklearn.dummy import DummyClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC as SVM
# + [markdown] id="WRNWwP1wLVkR" colab_type="text"
# ### Dummy Classifier (Input/Tuning is Irrelevant) ✔
# + id="hlrtsecXIyVP" colab_type="code" colab={}
# Make classifier
base = DummyClassifier(strategy='most_frequent')
# Fit on Train Data
base.fit(np.zeros_like(y_train), y_train)
# Make Predictions on Training Set
predictions = base.predict(np.zeros_like(y_train))
score = accuracy_score(y_train, predictions)
print("train accuracy: %.2f%%" % (score*100))
# Make Predictions on Test Set
predictions_test = base.predict(np.zeros_like(y_test))
score = accuracy_score(y_test, predictions_test)
print("test accuracy: %.2f%%" % (score*100))
print("Test data confusion matrix")
y_true = pd.Series(y_test, name='True')
y_pred = pd.Series(predictions_test, name='Predicted')
pd.crosstab(y_true, y_pred)
# + id="0tyo8KmsL8Hl" colab_type="code" colab={}
# + [markdown] id="QsmGbyOGNbo0" colab_type="text"
# ### Logistic Regression
# + id="NNMyFoQ-NjVs" colab_type="code" colab={}
pipeline = Pipeline([
('vect', CountVectorizer(min_df=10, max_features=5000, stop_words=stopwords.words('english'))),
('tfidf', TfidfTransformer()),
('dim_reduction', TruncatedSVD()),
('clf', LogisticRegression(class_weight='balanced'))
])
parameters = {
'vect__max_df': (0.6, 0.7),
'vect__ngram_range': ((1,2),(1,3)),
'dim_reduction__n_components': (100, 500, 1000),
'tfidf__use_idf': (True, False),
'tfidf__norm': ('l1', 'l2'),
'clf__C': (1.0, 0.1, 0.01),
}
clf = GridSearchCV(pipeline, parameters, scoring='f1', cv=3, n_jobs=-1)
clf.fit(x_train, y_train)
print("Best Score: ", clf.best_score_)
print("Best Params: ", clf.best_params_)
# + id="d6pyBWp7SA24" colab_type="code" colab={}
# Results of Grid Search
# Best Score: 0.762361919024294
# Best Params: {'clf__C': 1.0, 'tfidf__norm': 'l2', 'tfidf__use_idf': True, 'vect__max_df': 0.7, 'vect__min_df': 0.0, 'vect__ngram_range': (1, 3)}
# + [markdown] id="YRVh8rIfWpoc" colab_type="text"
# ### Logistic Regression SGD
# + id="to-PuGH7WrzM" colab_type="code" colab={}
pipeline = Pipeline([
('vect', CountVectorizer(min_df=10, max_features=5000,stop_words=stopwords.words('english'))),
('tfidf', TfidfTransformer()),
('dim_reduction', TruncatedSVD()),
('clf', SGDClassifier(loss="log", max_iter=1000, class_weight='balanced'))
])
parameters = {
'vect__max_df': (0.6, 0.7),
'vect__ngram_range': ((1,2), (1,3)),
'dim_reduction__n_components': (100,500,1000),
'tfidf__use_idf': (True, False),
'tfidf__norm': ('l1', 'l2'),
'clf__penalty':('l1','l2'),
'clf__alpha':(0.01, 0.001, 0.0001)
}
clf = GridSearchCV(pipeline, parameters, scoring='f1', cv=3, n_jobs=-1)
clf.fit(x_train, y_train)
print("Best Score: ", clf.best_score_)
print("Best Params: ", clf.best_params_)
# + id="IItP3pOVYHty" colab_type="code" colab={}
# Best Score: 0.7605481382673948
# Best Params: {'clf__alpha': 0.0001, 'clf__penalty': 'elasticnet', 'tfidf__norm': 'l2', 'tfidf__use_idf': True, 'vect__max_df': 1.0, 'vect__min_df': 0.0, 'vect__ngram_range': (1, 2)}
# + [markdown] id="zyGFFsE1YwB-" colab_type="text"
# ### Naive Bayes
# + id="djCuX7-HYYZi" colab_type="code" colab={}
pipeline = Pipeline([
('vect', CountVectorizer(min_df=10, max_features=5000, stop_words=stopwords.words('english'))),
('tfidf', TfidfTransformer()),
('dim_reduction', TruncatedSVD()),
('clf', MultinomialNB(fit_prior=True))
])
parameters = {
'vect__max_df': (0.6, 0.7),
'vect__ngram_range': ((1, 2), (1,3)),
'dim_reduction__n_components': (100,500,1000),
'tfidf__use_idf': (True, False),
'tfidf__norm': ('l1', 'l2'),
'clf__alpha': (1.0, 0.1, 0.01)
}
clf = GridSearchCV(pipeline, parameters, scoring='f1', cv=3, n_jobs=-1)
clf.fit(x_train, y_train)
print("Best Score: ", clf.best_score_)
print("Best Params: ", clf.best_params_)
# + id="Bwe-jaMwZixJ" colab_type="code" colab={}
# + [markdown] id="TPzqCsYwZjNE" colab_type="text"
# ### KNN
# + id="rMRG-LSoZqNC" colab_type="code" colab={}
pipeline = Pipeline([
('vect', CountVectorizer(min_df=10,max_features=5000,stop_words=stopwords.words('english'))),
('tfidf', TfidfTransformer()),
('dim_reduction', TruncatedSVD()),
('clf', KNeighborsClassifier())
])
parameters = {
'vect__max_df': (0.6, 0.7),
'vect__ngram_range': ((1, 2), (1,3)),
'dim_reduction__n_components': (100,500,1000),
'tfidf__use_idf': (True, False),
'tfidf__norm': ('l1', 'l2'),
'clf__n_neighbors': (3,5,7)
}
clf = GridSearchCV(pipeline, parameters, scoring='f1', cv=3, n_jobs=-1)
clf.fit(x_train, y_train)
print("Best Score: ", clf.best_score_)
print("Best Params: ", clf.best_params_)
# + id="TBQlESJabgjL" colab_type="code" colab={}
# + [markdown] id="ZEV5gdPubg-K" colab_type="text"
# ### SVM
# + id="XcZXkpN4b12l" colab_type="code" colab={}
pipeline = Pipeline([
('vect', CountVectorizer(min_df=10, max_features=5000, stop_words=stopwords.words('english'))),
('tfidf', TfidfTransformer()),
('dim_reduction', TruncatedSVD()),
('clf', SVM())
])
parameters = {
'vect__max_df': (0.6, 0.7),
'vect__ngram_range': ((1, 2), (1,3)),
'tfidf__use_idf': (True, False),
'dim_reduction__n_components': (100,500,1000),
'tfidf__norm': ('l1', 'l2'),
'clf__C':(1.0, 0.1, 0.01),
'clf__kernel':('linear','sigmoid')
}
clf = GridSearchCV(pipeline, parameters, scoring='f1', cv=3, n_jobs=-1)
clf.fit(x_train, y_train)
print("Best Score: ", clf.best_score_)
print("Best Params: ", clf.best_params_)
# + id="tuin2JW1dh-q" colab_type="code" colab={}
| text_analytics/Assignment2/Assignment2.ipynb |