code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ANOVA
# ANOVA is a method used to compare the means of more than two populations. So far, we
# have considered only a single population or at the most two populations. A one-way ANOVA uses one independent variable, while a two-way ANOVA uses two independent variables. The statistical distribution used in ANOVA is the F distribution, whose characteristics are as follows:
#
# 1. The F-distribution has a single tail (toward the right) and contains only positive values
#
# 
#
# 2. The F-statistic, which is the critical statistic in ANOVA, is the ratio of variation between the sample means to the variation within the sample. The formula is as follows.
# $$F = \frac{variation\ between\ sample\ means}{variation\ within\ the\ samples}$$
#
#
# 3. The different populations are referred to as treatments.
# 4. A high value of the F statistic implies that the variation between samples is considerable compared to variation within the samples. In other words, the populations or treatments from which the samples are drawn are actually different from one another.
# 5. Random variations between treatments are more likely to occur when the variation within the sample is considerable.
#
# Use a one-way ANOVA when you have collected data about one categorical independent variable and one quantitative dependent variable. The independent variable should have at least three levels (i.e. at least three different groups or categories).
#
# ANOVA tells you if the dependent variable changes according to the level of the independent variable. For example:
#
# + Your independent variable is social media use, and you assign groups to low, medium, and high levels of social media use to find out if there is a difference in hours of sleep per night.
# + Your independent variable is brand of soda, and you collect data on Coke, Pepsi, Sprite, and Fanta to find out if there is a difference in the price per 100ml.
#
# ANOVA determines whether the groups created by the levels of the independent variable are statistically different by calculating whether the means of the treatment levels are different from the overall mean of the dependent variable.
#
# If any of the group means is significantly different from the overall mean, then the null hypothesis is rejected.
#
# ANOVA uses the F-test for statistical significance. This allows for comparison of multiple means at once, because the error is calculated for the whole set of comparisons rather than for each individual two-way comparison (which would happen with a t-test).
#
# The F-test compares the variance in each group mean from the overall group variance. If the variance within groups is smaller than the variance between groups, the F-test will find a higher F-value, and therefore a higher likelihood that the difference observed is real and not due to chance.
#
# The assumptions of the ANOVA test are the same as the general assumptions for any parametric test:
#
# + **Independence of observations:** the data were collected using statistically-valid methods, and there are no hidden relationships among observations. If your data fail to meet this assumption because you have a confounding variable that you need to control for statistically, use an ANOVA with blocking variables.
# + **Normally-distributed response variable:** The values of the dependent variable follow a normal distribution.
# + **Homogeneity of variance:** The variation within each group being compared is similar for every group. If the variances are different among the groups, then ANOVA probably isn’t the right fit for the data.
# ## One-Way-ANOVA
#
# A few agricultural research scientists have planted a new variety of cotton called “AB
# cotton.” They have used three different fertilizers – A, B, and C – for three separate
# plots of this variety. The researchers want to find out if the yield varies with the type of
# fertilizer used. Yields in bushels per acre are mentioned in the below table. Conduct an
# ANOVA test at a 5% level of significance to see if the researchers can conclude that there
# is a difference in yields.
#
# | Fertilizer A | Fertilizer b | Fertilizer c |
# |--------------|--------------|--------------|
# | 40 | 45 | 55 |
# | 30 | 35 | 40 |
# | 35 | 55 | 30 |
# | 45 | 25 | 20 |
#
# Null hypothesis: $H_0 : \mu_1 = \mu_2 = \mu_3$
# Alternative hypothesis: $H_1 : \mu_1 ! = \mu_2 ! = \mu_3$
#
# the level of significance: $\alpha$=0.05
# +
import scipy.stats as stats
a=[40,30,35,45]
b=[45,35,55,25]
c=[55,40,30,20]
stats.f_oneway(a,b,c)
# -
# Since the calculated p-value (0.904)>0.05, we fail to reject the null hypothesis.There is no significant difference between the three treatments, at a 5% significance level.
# ## Two-way-ANOVA
#
# A botanist wants to know whether or not plant growth is influenced by sunlight exposure and watering frequency. She plants 30 seeds and lets them grow for two months under different conditions for sunlight exposure and watering frequency. After two months, she records the height of each plant, in inches.
# +
import numpy as np
import pandas as pd
#create data
df = pd.DataFrame({'water': np.repeat(['daily', 'weekly'], 15),
'sun': np.tile(np.repeat(['low', 'med', 'high'], 5), 2),
'height': [6, 6, 6, 5, 6, 5, 5, 6, 4, 5,
6, 6, 7, 8, 7, 3, 4, 4, 4, 5,
4, 4, 4, 4, 4, 5, 6, 6, 7, 8]})
# -
df[:10]
# +
import statsmodels.api as sm
from statsmodels.formula.api import ols
#perform two-way ANOVA
model = ols('height ~ C(water) + C(sun) + C(water):C(sun)', data=df).fit()
sm.stats.anova_lm(model, typ=2)
# -
# We can see the following p-values for each of the factors in the table:
#
# **water:** p-value = .000527
# **sun:** p-value = .0000002
# **water*sun:** p-value = .120667
#
# Since the p-values for water and sun are both less than .05, this means that both factors have a statistically significant effect on plant height.
#
# And since the p-value for the interaction effect (.120667) is not less than .05, this tells us that there is no significant interaction effect between sunlight exposure and watering frequency.
| 14. ANOVA - Analysis of Variance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Hello World program in Python
import json
import urllib.request # the lib that handles the url stuff
import pandas
from datetime import datetime, timedelta
import dateutil
pandas.options.display.float_format = '{:,.0f}'.format
url = "https://api.covidtracking.com/v1/us/current.json"
url = "https://api.covidtracking.com/v1/states/current.json"
url = "https://api.covidtracking.com/v1/states/daily.json"
#the_str = urllib.request.urlopen(url)
#df = pandas.read_json(the_str)
#df.to_json("covid_save.json")
df = pandas.read_json("covid_save.json")
#print(df)
states = ["AL","CT","FL","IL","NY","VA", "NJ"]
populations = { # per https://en.wikipedia.org/wiki/List_of_states_and_territories_of_the_United_States_by_population
"AL": 4903185,
"CT": 3565287,
"FL": 21477737,
"IL": 12671821,
"NY": 19453561,
"VA": 8535519,
"NJ": 9241900,
'USA': 328200000
}
mask_usage = { # as of 10/27/2020
"AL": .64,
"CT": .70,
"FL": .65,
"IL": .67,
"NY": .75,
"VA": .68
}
usa_deaths = { # From https://www.cdc.gov/nchs/fastats/leading-causes-of-death.htm
'Heart disease': 655381,
'Cancer': 599274,
'Accidents': 167127,
'Influenza+Pneumonia': 60000
}
cols = ["date","state","death","positive","dataQualityGrade"]
end_date = dateutil.parser.parse("12/23/2020") #datetime.now()
start_date = end_date - timedelta(days=30)
#print(now.strftime(f"%Y%m%d")+"-"+then.strftime(f"%Y%m%d"))
#today = 20201001
#print(today)
dates = [end_date.strftime(f"%Y%m%d"),start_date.strftime(f"%Y%m%d")]
print(dates)
df2 = df[df.state.isin(states)][df.date.isin(dates)][cols]
#print(df2)
#print(df2['death'].diff(periods=(len(states)*-1)))
df2['delta_d'] = df2['death'].diff(periods=(len(states)*-1))
df2['delta_p'] = df2['positive'].diff(periods=(len(states)*-1))
df2 = df2[df.date.isin([dates[0]])]
df2['case_mortality'] = df2['delta_d'] / df2['delta_p']
df2['population'] = df2['state'].map(populations)
df2['cases/100k'] = df2['delta_p'] / df2['population'] * 100000
df2['deaths/100k'] = df2['delta_d'] / df2['population'] * 100000
df2['deaths/100k per year'] = df2['deaths/100k'] * 12
df3 = df2.copy()
df3 = df3.set_index('state')
df3 = df3.drop(labels=['date','dataQualityGrade','death','positive','population','delta_p','delta_d','deaths/100k per year'],axis=1)
#print(df2.transpose())
formatters2 = {
'case_mortality':'{:.2%}'.format,
'cases/100k':'{:.2f}'.format,
'deaths/100k':'{:.2f}'.format,
'deaths/100k per year':'{:,.2f}'.format,
'population':lambda x : '{:,.1f}'.format(x/1000000)+'m'
}
df2 = df2.drop(labels=['date','dataQualityGrade'],axis=1)
df2 = df2.rename(columns=
{'death':'total deaths',
'positive':'total positive',
'delta_d':'death delta',
'delta_p':'positive delta',
})
#df2.style.format(formatters2)
for key, value in formatters2.items():
df2[key] = df2[key].apply(formatters2[key])
#print(df2.to_string(formatters=formatters2, float_format='{:,.0f}'.format))
print(df2.transpose())
flu_mortality = 17.1 # per 100k per https://www.cdc.gov/nchs/data/nvsr/nvsr68/nvsr68_09-508.pdf (row 8 on page 6) = 0.0171%
# If mortality is ~2% of cases, then case risk of 0.855%.
# -
df3
# +
# libraries
import matplotlib.pyplot as plt
import numpy as np
# create data
values=np.cumsum(np.random.randn(1000,1))
# use the plot function
#plt.plot(values)
plt.figure()
df3['case_mortality'].plot(kind='bar')
# -
df3['deaths/100k'].plot(kind='bar')
df3['cases/100k'].plot(kind='bar')
#date_time_obj = datetime.strptime("20201231", f"%Y%m%d")
#print('Date:', date_time_obj.date())
#print('Time:', date_time_obj.time())
#print('Date-time:', date_time_obj)
df['date-obj'] = pandas.to_datetime(df['date'], format=f"%Y%m%d")
df['date-obj']
# +
rolling_avg = 28
start_date = datetime.today() - timedelta(28*6)
#print(start_date)
#df_by_date = df[df['date-obj'] >= start_date]
df_by_date = df.set_index('date-obj')
df_by_date['population'] = df_by_date['state'].map(populations) / 100000
for state in states:
#ax = df_by_date[df_by_date.state.isin([state])]['death'].diff(periods=-30).plot(y=state,legend=True) #rolling(30).mean().plot()
ax = (df_by_date[df_by_date.state.isin([state])]['death'] / df_by_date[df_by_date.state.isin([state])]['population'] / rolling_avg)\
.diff(periods=-1*rolling_avg).plot(y=state,legend=True)
ax.legend(states)
#df_by_date[df_by_date.state.isin(['FL'])]['death'].diff(periods=-30).plot() #rolling(30).mean().plot()
#df_by_date['Influenza'] = 0.05
#ax = df_by_date['Influenza'].plot(y=state,legend=True)
for cause in usa_deaths:
df_by_date[cause] = usa_deaths[cause] / populations['USA'] * 100000 / 365
ax = df_by_date[cause].plot(y=state,legend=True,style="--",figsize=(10,5),title=f"Daily Deaths per 100k ({rolling_avg} day avg)")
ax.set_xlim(xmin=start_date)
ax.set_ylim(ymax=2.0)
#delim = ''
#title = ''
#cause_sum = 0
#for cause in usa_deaths:
# title += delim+cause
# delim = '+'
# cause_sum += usa_deaths[cause]
#df_by_date[title] = cause_sum / populations['USA'] * 100000 / 365
#print(cause_sum / populations['USA'] * 100000)
#ax = df_by_date[title].plot(y=state,legend=True)
# -
rolling_avg = 28
df_by_date = df.set_index('date-obj')
df_by_date['population'] = df_by_date['state'].map(populations) / 100000
for state in states:
ax = (df_by_date[df_by_date.state.isin([state])]['positive'] / df_by_date[df_by_date.state.isin([state])]['population'] / rolling_avg)\
.diff(periods=-28).plot(y=state,legend=True) #rolling(30).mean().plot()
ax.legend(states)
df_by_date = df.set_index('date-obj')
for state in states:
ax = (df_by_date[df_by_date.state.isin([state])]['death'].diff(periods=-28) / df_by_date[df_by_date.state.isin([state])]['positive'].diff(periods=-28)).plot(y=state,legend=True) #rolling(30).mean().plot()
ax.legend(states)
| covid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/mlelarge/dataflowr/blob/master/CEA_EDF_INRIA/AE_empty_colab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Unsupervised learning with Autoencoder
#
# Some piece of codes taken from https://github.com/kevinzakka/vae-pytorch
#
# <img src='img/Autoencoder_structure.png'>
#
# Description given by [Wikipedia](https://en.wikipedia.org/wiki/Autoencoder)
# +
import os
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
# %matplotlib inline
# -
# ## Loading MNIST
# +
# %mkdir data
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('data', train=True, download=True, transform=transforms.ToTensor()),
batch_size=256, shuffle=True)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('data', train=False, download=True, transform=transforms.ToTensor()),
batch_size=10, shuffle=False)
# -
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print('Using gpu: %s ' % torch.cuda.is_available())
# ## Helper Functions
# +
def to_img(x):
x = x.cpu().data.numpy()
x = 0.5 * (x + 1)
x = np.clip(x, 0, 1)
x = x.reshape([-1, 28, 28])
return x
def plot_reconstructions(model, conv=False):
"""
Plot 10 reconstructions from the test set. The top row is the original
digits, the bottom is the decoder reconstruction.
The middle row is the encoded vector.
"""
# encode then decode
data, _ = next(iter(test_loader))
if not conv:
data = data.view([-1, 784])
data.requires_grad = False
data = data.to(device)
true_imgs = data
encoded_imgs = model.encoder(data)
decoded_imgs = model.decoder(encoded_imgs)
true_imgs = to_img(true_imgs)
decoded_imgs = to_img(decoded_imgs)
encoded_imgs = encoded_imgs.cpu().data.numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(3, n, i + 1)
plt.imshow(true_imgs[i])
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.imshow(encoded_imgs[i].reshape(-1,4))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(3, n, i + 1 + n + n)
plt.imshow(decoded_imgs[i])
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# -
# ## Simple Auto-Encoder
#
# We'll start with the simplest autoencoder: a single, fully-connected layer as the encoder and decoder.
class AutoEncoder(nn.Module):
def __init__(self, input_dim, encoding_dim):
super(AutoEncoder, self).__init__()
self.encoder = nn.Linear(input_dim, encoding_dim)
self.decoder = nn.Linear(encoding_dim, input_dim)
def forward(self, x):
encoded = F.relu(self.encoder(x))
decoded = self.decoder(encoded)
return decoded
# +
input_dim = 784
encoding_dim = 32
model = AutoEncoder(input_dim, encoding_dim)
model = model.to(device)
optimizer = optim.Adam(model.parameters())
loss_fn = torch.nn.MSELoss()
# -
# Why did we take 784 as input dimension? What is the learning rate?
def train_model(model,loss_fn,data_loader=None,epochs=1,optimizer=None):
model.train()
for epoch in range(epochs):
for batch_idx, (data, _) in enumerate(train_loader):
data = data.view([-1, 784]).to(device)
optimizer.zero_grad()
output = model(data)
loss = loss_fn(output, data)
loss.backward()
optimizer.step()
if batch_idx % 100 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(data_loader.dataset),
100. * batch_idx / len(data_loader), loss.data.item()))
train_model(model, loss_fn,data_loader=train_loader,epochs=10,optimizer=optimizer)
plot_reconstructions(model)
# If you remove the non-linearity, what are you doing?
# ## Stacked Auto-Encoder
class DeepAutoEncoder(nn.Module):
def __init__(self, input_dim, encoding_dim):
super(DeepAutoEncoder, self).__init__()
self.encoder = nn.Sequential(
nn.Linear(input_dim, 128),
nn.ReLU(True),
nn.Linear(128, 64),
nn.ReLU(True),
nn.Linear(64, encoding_dim),
nn.ReLU(True),
)
self.decoder = nn.Sequential(
nn.Linear(encoding_dim, 64),
nn.ReLU(True),
nn.Linear(64, 128),
nn.ReLU(True),
nn.Linear(128, input_dim),
)
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
return x
# +
input_dim = 784
encoding_dim = 32
model = DeepAutoEncoder(input_dim, encoding_dim)
model = model.to(device)
optimizer = optim.Adam(model.parameters())
loss_fn = torch.nn.MSELoss()
# -
model.encoder
model.decoder
train_model(model, loss_fn,data_loader=train_loader,epochs=10,optimizer=optimizer)
plot_reconstructions(model)
# # Exercise
#
# - Change the loss to a nn.BCEWithLogits() loss.
#
# - Implement a weight sharing AE, for which the decoder weight matrix is just the transpose of the encoder weight matrix.
#
# - You may use F.linear(x, weight_matrix)
#
# A rapid google search gives:
#
# https://discuss.pytorch.org/t/how-to-create-and-train-a-tied-autoencoder/2585
# ## Convolutional Auto-Encoder
#
# Deconvolution are creating checkboard artefacts see [Odena et al.](https://distill.pub/2016/deconv-checkerboard/)
class ConvolutionalAutoEncoder(nn.Module):
def __init__(self):
super(ConvolutionalAutoEncoder, self).__init__()
self.encoder = nn.Sequential(
nn.Conv2d(in_channels=1, out_channels=16, kernel_size=3, stride=3, padding=1),
nn.ReLU(True),
nn.MaxPool2d(2, stride=2), # b, 16, 5, 5
nn.Conv2d(in_channels=16, out_channels=8, kernel_size=3, stride=2, padding=1),
nn.ReLU(True),
nn.MaxPool2d(2, stride=1)
)
self.decoder = nn.Sequential(
nn.ConvTranspose2d(in_channels=8, out_channels=16, kernel_size=3, stride=2), # b, 16, 5, 5
nn.ReLU(True),
nn.ConvTranspose2d(in_channels=16, out_channels=8, kernel_size=5, stride=3, padding=1), # b, 8, 15, 15
nn.ReLU(True),
nn.ConvTranspose2d(in_channels=8, out_channels=1, kernel_size=2, stride=2, padding=1), # b, 1, 28, 28
)
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
return x
model = ConvolutionalAutoEncoder()
model = model.to(device)
optimizer = optim.Adam(model.parameters())
loss_fn = torch.nn.BCEWithLogitsLoss()
# Why is
#
# `train_model(model,loss_fn,data_loader=train_loader,epochs=10,optimizer=optimizer)`
#
# not working? Make the necessary modification.
def train_convmodel(model,loss_fn,data_loader=None,epochs=1,optimizer=None):
model.train()
for epoch in range(epochs):
for batch_idx, (data, _) in enumerate(train_loader):
#
# your code here
#
if batch_idx % 100 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(data_loader.dataset),
100. * batch_idx / len(data_loader), loss.data.item()))
train_convmodel(model, loss_fn,data_loader=train_loader,epochs=10,optimizer=optimizer)
plot_reconstructions(model, conv=True)
# # Exercise
#
# Implement a denoising AE:
#
# 
#
# Use previous code and with minimal modifications, transform your AE in a denoising AE.
def train_denoising(model,loss_fn,data_loader=None,epochs=1,optimizer=None, noise=0.1):
model.train()
for epoch in range(epochs):
for batch_idx, (data, _) in enumerate(train_loader):
#
# your code here
#
if batch_idx % 100 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(data_loader.dataset),
100. * batch_idx / len(data_loader), loss.data.item()))
def plot_denoising(model, conv=False, noise=0.1):
"""
Plot 10 reconstructions from the test set. The top row is the original
digits, the bottom is the decoder reconstruction.
The middle row is the encoded vector.
"""
# encode then decode
data, _ = next(iter(test_loader))
data = data.to(device)
#
# your code here
#
if not conv:
data = data.view([-1, 784])
data.requires_grad = False
true_imgs = data
encoded_imgs = model.encoder(data)
decoded_imgs = model.decoder(encoded_imgs)
true_imgs = to_img(true_imgs)
decoded_imgs = to_img(decoded_imgs)
encoded_imgs = encoded_imgs.data.numpy()
n = 10
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(3, n, i + 1)
plt.imshow(true_imgs[i])
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax = plt.subplot(3, n, i + 1 + n)
plt.imshow(encoded_imgs[i].reshape(-1,4))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(3, n, i + 1 + n + n)
plt.imshow(decoded_imgs[i])
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
model = ConvolutionalAutoEncoder()
model = model.to(device)
optimizer = optim.Adam(model.parameters())
loss_fn = torch.nn.BCEWithLogitsLoss()
train_denoising(model, loss_fn,data_loader=train_loader,epochs=10,optimizer=optimizer, noise=0.2)
plot_denoising(model, conv=True, noise=0.2)
# You should obtain results like this:
#
# 
| CEA_EDF_INRIA/AE_empty_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Colaboratoryで実行する場合のみ実行" data-toc-modified-id="Colaboratoryで実行する場合のみ実行-1"><span class="toc-item-num">1 </span>Colaboratoryで実行する場合のみ実行</a></span></li><li><span><a href="#前処理" data-toc-modified-id="前処理-2"><span class="toc-item-num">2 </span>前処理</a></span></li><li><span><a href="#ニューラルネットワークの訓練" data-toc-modified-id="ニューラルネットワークの訓練-3"><span class="toc-item-num">3 </span>ニューラルネットワークの訓練</a></span></li><li><span><a href="#ニューラルネットワークの評価" data-toc-modified-id="ニューラルネットワークの評価-4"><span class="toc-item-num">4 </span>ニューラルネットワークの評価</a></span><ul class="toc-item"><li><span><a href="#学習の進捗" data-toc-modified-id="学習の進捗-4.1"><span class="toc-item-num">4.1 </span>学習の進捗</a></span></li><li><span><a href="#データ量とテストデータでの評価項目の関連" data-toc-modified-id="データ量とテストデータでの評価項目の関連-4.2"><span class="toc-item-num">4.2 </span>データ量とテストデータでの評価項目の関連</a></span></li><li><span><a href="#ROC曲線" data-toc-modified-id="ROC曲線-4.3"><span class="toc-item-num">4.3 </span>ROC曲線</a></span></li><li><span><a href="#Grad-CAM" data-toc-modified-id="Grad-CAM-4.4"><span class="toc-item-num">4.4 </span>Grad-CAM</a></span></li></ul></li></ul></div>
# -
# # Colaboratoryで実行する場合のみ実行
# - [Googleドライブの読み込みと保存](https://colab.research.google.com/notebooks/io.ipynb)
# +
import glob
import tarfile
from google.colab import drive
drive.mount('/content/gdrive')
# !mkdir images
# !mkdir logs
# !cp gdrive/My\ Drive/x-ray/* .
# !cp gdrive/My\ Drive/x-ray/images/* images
# +
gz_list = glob.glob("images/*.tar.gz")
for gz_file in gz_list:
tar = tarfile.open(gz_file)
tar.extractall()
tar.close()
# !pip install Keras==2.2.4
# !pip install Keras-Applications==1.0.6
# !pip install Keras-Preprocessing==1.0.5
# -
# # 前処理
# +
import os
import glob
import shutil
import numpy as np
import pandas as pd
import random
import pathlib
import keras
from keras_preprocessing.image import (array_to_img, img_to_array, load_img)
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from IPython.display import display
#auto relord modules
# %load_ext autoreload
# %autoreload 2
# +
random.seed(2018)
df = pd.read_csv('Data_Entry_2017.csv', encoding='us-ascii')
# Keep df where the file exist in images folder.
file_list = glob.glob("images/*")
file_list = [os.path.basename(file) for file in file_list]
file_list = pd.DataFrame(file_list, columns=['Image Index'])
df = pd.merge(df, file_list, on='Image Index', how='inner')
Findings = df["Finding Labels"].str.split('|')
Findings_list = [word for word_inner in Findings for word in word_inner]
Findings_list = pd.DataFrame(Findings_list, columns=['text'])
print(Findings_list['text'].value_counts())
Findings_list = list(Findings_list.drop_duplicates()['text'])
Findings_list.remove('No Finding')
for finding in Findings_list:
df[finding] = Findings.apply(lambda x: 1 if finding in x else 0)
# +
# Create train_list.txt and validation_list.txt
random.seed(2018)
train_val_list = set([])
with open('train_val_list.txt', 'r') as f:
for row in f:
train_val_list.add(row.rsplit()[0])
train_list = random.sample(train_val_list, int(0.8 * len(train_val_list)))
train_list = sorted(train_list)
validation_list = train_val_list - set(train_list)
validation_list = sorted(validation_list)
with open('train_list.txt', 'w') as f:
f.write("\n".join(train_list))
with open('validation_list.txt', 'w') as f:
f.write("\n".join(validation_list))
# +
for tt in ['train', 'validation', 'test']:
tmp_list = []
with open(tt + '_list.txt', 'r') as f:
for row in f:
tmp_list.append(row.rsplit()[0])
df[tt] = df['Image Index'].isin(tmp_list).apply(
lambda x: 'Y' if x == True else 'N')
def data_split(x):
if x.test == "Y":
return "test"
elif x.train == "Y":
return "train"
else:
return "validation"
df['split'] = df.apply(lambda x: data_split(x), axis=1)
# -
# # ニューラルネットワークの訓練
# +
import myTrainer
y_col = ['Effusion']
# +
# trainingx = myTrainer.trainer(
# trainer_name='densenet_p3',
# dataframe=df,
# data_proportion=10**-2,
# y_col=y_col,
# lr=10**-7,
# batch_size=32,
# initial_epoch=0,
# epochs=50)
# trainingx.lr_finder('densenet_p3')
# trainingx = myTrainer.trainer(
# trainer_name='densenet_p2',
# dataframe=df,
# data_proportion=10**-2,
# y_col=y_col,
# lr=10**-7,
# batch_size=32,
# initial_epoch=0,
# epochs=50)
# trainingx.lr_finder('densenet_p2')
# trainingx = myTrainer.trainer(
# trainer_name='densenet_p1',
# dataframe=df,
# data_proportion=10**-1,
# y_col=y_col,
# lr=10**-7,
# batch_size=32,
# initial_epoch=0,
# epochs=50)
# trainingx.lr_finder('p1')
# trainingx = myTrainer.trainer(
# trainer_name='densenet_p0',
# dataframe=df,
# data_proportion=10**-0,
# y_col=y_col,
# lr=10**-7,
# batch_size=32,
# initial_epoch=0,
# epochs=50)
# trainingx.lr_finder('densenet_p0')
# +
# import myTrainer
# training1 = myTrainer.trainer(
# trainer_name='densenet_lr7_p3',
# dataframe=df,
# data_proportion=10**-3,
# y_col=y_col,
# lr=10**-7,
# batch_size=32,
# initial_epoch=0,
# epochs=10)
# training1.training()
# training1.resume_training('densenet_lr7_p3_1')
# -
training3 = myTrainer.trainer(
trainer_name='densenet_lr3_p3',
dataframe=df,
data_proportion=10**-3,
y_col=y_col,
lr=10**-3,
batch_size=32,
initial_epoch=0,
epochs=50)
training3.training()
training3.evaluating()
training2 = myTrainer.trainer(
trainer_name='densenet_lr3_p2',
dataframe=df,
data_proportion=10**-2,
y_col=y_col,
lr=10**-3,
batch_size=32,
initial_epoch=0,
epochs=50)
training2.training()
training2.evaluating()
training1 = myTrainer.trainer(
trainer_name='densenet_lr3_p1',
dataframe=df,
data_proportion=10**-1,
y_col=y_col,
lr=10**-3,
batch_size=32,
initial_epoch=0,
epochs=50)
training1.training()
training1.evaluating()
training0 = myTrainer.trainer(
trainer_name='densenet_lr3_p0',
dataframe=df,
data_proportion=10**-0,
y_col=y_col,
lr=10**-3,
batch_size=32,
initial_epoch=0,
epochs=50)
training0.training()
training0.evaluating()
# # ニューラルネットワークの評価
# ## 学習の進捗
# +
df_training_log = pd.DataFrame()
for _ in range(4):
tmp = pd.read_csv(
'./logs/training_log_densenet_lr3_p' + str(_) + '.csv', sep='\t')
tmp['power'] = _
df_training_log = pd.concat([df_training_log, tmp])
vars = ['acc', 'loss', 'auc', 'fscore', 'precision', 'recall']
epoch = df_training_log['epoch'].unique() + 1
sns.set()
cmap = plt.get_cmap("Blues")
for var in vars:
for power in range(4):
training = df_training_log.query('power==' + str(power))[var]
val = df_training_log.query('power==' + str(power))['val_' + var]
plt.plot(
epoch,
training,
'bo',
label=str(10**-power) + ': Training ' + var,
color=cmap((4 - power) / 4))
plt.plot(
epoch,
val,
'b',
label=str(10**-power) + ': Validation ' + var,
color=cmap((4 - power) / 4))
plt.title('Training and validation ' + var)
plt.legend()
plt.savefig("logs/Effusion_Train_Val_p0-p3_" + var + ".svg")
plt.figure()
plt.show()
# +
for var in vars:
for power in [0]:
training = df_training_log.query('power==' + str(power))[var]
val = df_training_log.query('power==' + str(power))['val_' + var]
plt.plot(
epoch,
training,
'bo',
label='Training ' + var,
color=cmap((4 - power) / 4))
plt.plot(
epoch,
val,
'b',
label='Validation ' + var,
color=cmap((4 - power) / 4))
plt.title('Training and validation ' + var)
plt.legend()
plt.savefig("logs/Effusion_Train_Val_p0_" + var + ".svg")
plt.figure()
plt.show()
# -
# ## データ量とテストデータでの評価項目の関連
# +
df_test_log = pd.DataFrame()
for _ in range(4):
tmp = pd.read_csv(
'./logs/test_result_densenet_lr3_p' + str(_) + '.csv', sep='\t')
tmp['power'] = _
tmp['frac'] = 10**(-_)
df_test_log = pd.concat([df_test_log, tmp])
vars = ['acc', 'loss', 'auc', 'fscore', 'precision', 'recall']
sns.set()
cmap = plt.get_cmap("Blues")
#cmap = sns.cubehelix_palette(4, as_cmap=True)
for var in vars:
y = df_test_log[var]
x = df_test_log['frac']
plt.plot(x, y, 'b', color=cmap(1.0))
plt.title('Test ' + var)
plt.legend()
plt.xscale("log")
plt.savefig("logs/Effusion_Test_" + var + ".svg")
plt.figure()
plt.show()
# -
# ## ROC曲線
#http://scikit-learn.org/stable/auto_examples/model_selection/plot_roc.html
roc_auc_list, fpr_list, tpr_list = training0.auc_test()
plt.figure()
for i, finding in enumerate(y_col):
fpr = fpr_list[i]
tpr = tpr_list[i]
roc_auc = roc_auc_list[i]
plt.plot(
fpr,
tpr,
#color='darkorange',
lw=2,
label=finding + ' (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig("logs/Effusion_Test_ROC.svg")
plt.show()
# ## Grad-CAM
training0.Grad_CAM('00021381_013.png')
training0.Grad_CAM('00001437_012.png')
training0.Grad_CAM('00001558_016.png')
training0.Grad_CAM('00029894_000.png')
training0.Grad_CAM('00013337_000.png')
training0.Grad_CAM('00021181_002.png')
training0.Grad_CAM('00012045_009.png')
'''
00002395_007.png
00027028_017.png
00010007_168.png
00028974_016.png
00016291_002.png
00023058_004.png
00020277_001.png
00030634_000.png
00012834_034.png
00018427_011.png
00007034_016.png
00012834_122.png
00016972_025.png
00023283_019.png
00013285_026.png
00017714_006.png
00027631_000.png
00020751_003.png
'''
| CNN_x-ray.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from influxdb import InfluxDBClient
import random
from time import sleep
i = 0
for i in range(0,3600):
# クライアントのインスタンス作成
user = 'root'
password = '<PASSWORD>'
dbname = 'superdry'
host='influxdb'
port=8086
client = InfluxDBClient(host, port, user, password, dbname)
# データベース(なければ)作成
client.create_database(dbname)
# 送信内容
# json_body = [
# {
# "measurement": "sensordata",
# "tags": {
# "user": "umetsu",
# "id": "0"},
# "fields": {
# "temperature": 1.0,
# "humidity":2.0,
# "co2":3.0,
# "wetness":"4.0",
# "tvoc":5.0
# }
# }
# ]
json_body = [
{
"measurement": "sensordata",
"tags": {
"user": "umetsu",
"id": "0"},
"fields": {
"temperature": random.uniform(0.0,100.0),
"humidity":random.uniform(0.0,100.0),
"co2":random.uniform(0.0,100.0),
"wetness":random.uniform(0.0,100.0),
"tvoc":random.uniform(0.0,100.0)
}
}
]
# 内容確認
print(json_body)
# 送信
client.write_points(json_body)
i = i+1
sleep(5)
# -
| servers/jupyter/notebooks/send.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LogLoss
#
# Just run the right model. (or calibrate others, look at what is it calibration, not clear yet).
#
# ## Accuracy
#
# Fit any metrics, and tune threshold.
#
# ## Quadratic Weighted Kappa. (Cohen's Kappa)
#
# 1. Optimize MSE
#
# 2. Find right thresholds
# - Bad: np.round(predictions)
# - Better: optmize threshold.
#
# 3. Use soft kappa loss.
| old_files_unsorted_archive/classification_metrics_optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: airflow_jupyter
# language: python
# name: airflow_jupyter
# ---
# # this should be transformed to little library later on
# to ensure every task saves to thed same directory
# +
import random
import logging
from result_saver import ResultSaver
log = logging.getLogger("jupyter_code")
value = random.randint(10,20)
log.info(f'I have drawn {value} seconds for the next task!')
result = {
'sleeping_time': value
}
ResultSaver().save_result(result)
| docker/task2/code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ensemble (voting)
from si.data import Dataset, summary
from si.util import CrossValidationScore
import os
DIR = os.path.dirname(os.path.realpath('.'))
filename = os.path.join(DIR, 'datasets/breast-bin.data')
dataset = Dataset.from_data(filename)
summary(dataset)
dataset.toDataframe()
# Use accuracy as scorring function
from si.util import accuracy_score
# ### Decision Tree
from si.supervised import DecisionTree
dt = DecisionTree()
cv = CrossValidationScore(dt,dataset,score=accuracy_score)
cv.run()
cv.toDataframe()
# ### Logistic regression
from si.supervised import LogisticRegression
logreg = LogisticRegression()
cv = CrossValidationScore(logreg,dataset,score=accuracy_score)
cv.run()
cv.toDataframe()
# ### KNN
from si.supervised import KNN
knn = KNN(7)
cv = CrossValidationScore(knn,dataset,score=accuracy_score)
cv.run()
cv.toDataframe()
# ## Ensemble
from si.supervised import Ensemble, majority
en = Ensemble([dt,logreg,knn], majority, accuracy_score)
cv = CrossValidationScore(en,dataset,score=accuracy_score)
cv.run()
cv.toDataframe()
| scripts/eval3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fitting interaction paramaters for mixtures
#
#
# This notebook has the purpose of showing examples of how to fit interaction parameters for binary mixtures using experimental equilbrium data.
import numpy as np
from phasepy import component, mixture, prsveos
# +
#Vapor Liquid equilibria data obtanied from Rieder, <NAME>. y A. <NAME> (1949).
# «Vapor-Liquid Equilibria Measured by a GillespieStill - Ethyl Alcohol - Water System».
#Ind. Eng. Chem. 41.12, 2905-2908.
#Saturation Pressure in bar
Pexp = np.array([1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013,
1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013,
1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013,
1.013, 1.013, 1.013, 1.013, 1.013, 1.013, 1.013])
#Saturation temeprature in Kelvin
Texp = np.array([372.45, 370.05, 369.15, 369.15, 368.75, 367.95, 366.95, 366.65,
366.05, 363.65, 363.65, 362.55, 361.55, 361.75, 360.35, 358.55,
357.65, 357.15, 356.55, 356.15, 355.45, 355.15, 354.55, 354.65,
354.35, 354.05, 353.65, 353.35, 353.15, 352.65, 351.95, 351.65,
351.55, 351.45])
#Liquid fraction mole array
Xexp = np.array([[0.0028, 0.0118, 0.0137, 0.0144, 0.0176, 0.0222, 0.0246, 0.0302,
0.0331, 0.0519, 0.053 , 0.0625, 0.0673, 0.0715, 0.0871, 0.126 ,
0.143 , 0.172 , 0.206 , 0.21 , 0.255 , 0.284 , 0.321 , 0.324 ,
0.345 , 0.405 , 0.43 , 0.449 , 0.506 , 0.545 , 0.663 , 0.735 ,
0.804 , 0.917 ],
[0.9972, 0.9882, 0.9863, 0.9856, 0.9824, 0.9778, 0.9754, 0.9698,
0.9669, 0.9481, 0.947 , 0.9375, 0.9327, 0.9285, 0.9129, 0.874 ,
0.857 , 0.828 , 0.794 , 0.79 , 0.745 , 0.716 , 0.679 , 0.676 ,
0.655 , 0.595 , 0.57 , 0.551 , 0.494 , 0.455 , 0.337 , 0.265 ,
0.196 , 0.083 ]])
#Vapor fraction mole array
Yexp = np.array([[0.032, 0.113, 0.157, 0.135, 0.156, 0.186, 0.212, 0.231, 0.248,
0.318, 0.314, 0.339, 0.37 , 0.362, 0.406, 0.468, 0.487, 0.505,
0.53 , 0.527, 0.552, 0.567, 0.586, 0.586, 0.591, 0.614, 0.626,
0.633, 0.661, 0.673, 0.733, 0.776, 0.815, 0.906],
[0.968, 0.887, 0.843, 0.865, 0.844, 0.814, 0.788, 0.769, 0.752,
0.682, 0.686, 0.661, 0.63 , 0.638, 0.594, 0.532, 0.513, 0.495,
0.47 , 0.473, 0.448, 0.433, 0.414, 0.414, 0.409, 0.386, 0.374,
0.367, 0.339, 0.327, 0.267, 0.224, 0.185, 0.094]])
datavle = (Xexp, Yexp, Texp, Pexp)
# +
water = component(name = 'Water', Tc = 647.13, Pc = 220.55, Zc = 0.229, Vc = 55.948, w = 0.344861,
ksv = [ 0.87292043, -0.06844994],
Ant = [ 11.72091059, 3852.20302815, -44.10441047],
cii = [ 1.16776082e-25, -4.76738739e-23, 1.79640647e-20],
GC = {'H2O':1})
ethanol = component(name = 'Ethanol', Tc = 514.0, Pc = 61.37, Zc = 0.241, Vc = 168.0, w = 0.643558,
ksv = [1.27092923, 0.0440421 ],
Ant = [ 12.26474221, 3851.89284329, -36.99114863],
cii = [ 2.35206942e-24, -1.32498074e-21, 2.31193555e-19],
GC = {'CH3':1, 'CH2':1, 'OH(P)':1})
mix = mixture(ethanol, water)
# -
# ## Fitting QMR mixing rule
#
# As an scalar is been fitted, scipy recommends to give a certain interval where the minimum could be found, the function ```fit_kij``` handles this optimization.
# +
from phasepy.fit import fit_kij
mixkij = mix.copy()
fit_kij((-0.15, -0.05), prsveos, mixkij, datavle)
# -
# ## Fitting NRTL interaction parameters
#
# As an array is been fitted, multidimentional optimization alogirthms are used, the function ```fit_nrtl``` handles this optimization with several options available. If a fixed value of the aleatory factor is used the initial guess has the following form:
#
# nrtl0 = np.array([A12, A21])
#
# If the aleatory factor needs to be optimized it can be included setting alpha_fixed to False, in this case the initial guess has the following form:
#
# nrtl0 = np.array([A12, A21, alpha])
#
# Temperature dependent parameters can be fitted setting the option Tdep = True in ```fit_nrtl```, when this option is used the parameters are computed as:
#
# $$
# A12 = A12_0 + A12_1 T \\
# A21 = A21_0 + A21_1 T
# $$
#
# The initial guess passed to the fit function has the following form:
#
# nrtl0 = np.array([A12_0, A21_0, A12_1, A21_1, alpha])
#
# or, if alpha fixed is used.
#
# nrtl0 = np.array([A12_0, A21_0, A12_1, A21_1])
# +
from phasepy.fit import fit_nrtl
mixnrtl = mix.copy()
#Initial guess of A12, A21
nrtl0 = np.array([-80., 650.])
fit_nrtl(nrtl0, mixnrtl, datavle, alpha_fixed = True)
#optimized values
[-84.77530335, 648.78439102]
#Initial guess of A12, A21
nrtl0 = np.array([-80., 650., 0.2])
fit_nrtl(nrtl0, mixnrtl, datavle, alpha_fixed = False)
#optimized values for A12, A21, alpha
[-5.53112687e+01, 6.72701992e+02, 3.19740734e-01]
# -
# By default Tsonopoulos virial correlation is calculated for vapor phase, if desired ideal gas or Abbott correlation can be used.
# +
#Initial guess of A12, A21
nrtl0 = np.array([-80., 650.])
fit_nrtl(nrtl0, mixnrtl, datavle, alpha_fixed = True, virialmodel = 'ideal_gas')
#optimized values
[-86.22483806, 647.6320968 ]
#Initial guess of A12, A21
nrtl0 = np.array([-80., 650.])
fit_nrtl(nrtl0, mixnrtl, datavle, alpha_fixed = True, virialmodel = 'Abbott')
#optimized values
[-84.81672981, 648.75311712]
# -
# ## Fitting Wilson interaction parameters
#
# As an array is been fitted, multidimentional optimization alogirthms are used, the function ```fit_wilson``` handles this optimization.
# +
from phasepy.fit import fit_wilson
mixwilson = mix.copy()
#Initial guess of A12, A21
wilson0 = np.array([-80., 650.])
fit_wilson(wilson0, mixwilson, datavle)
# -
# Similarly as when fitting nrtl parameters, Tsonopoulos virial correlation is used by default. Ideal gas or Abbott correlation can be used.
fit_wilson(wilson0, mixwilson, datavle, virialmodel = 'ideal_gas')
#optimized value
#[105.42279401, 517.2221969 ]
# ## Fitting Redlich-Kister interaction parameters
#
# As an array is been fitted, multidimentional optimization alogirthms are used, the function ```fit_rk``` handles this optimization. Redlich-Kister expansion is programmed for n terms of the expansion, this fitting function will optimize considering the lenght of the array passed as an initial guess.
#
# If rk0 is an scalar it reduced to Porter model, if it is array of size 2 it reduces to Margules Model.
#
# Temperature dependent parameters can be fitted in which case the initial guess will be splitted into two array.
#
# c, c1 = np.split(rk0, 2)
#
# Finally the parameters are computed as:
#
# G = c + c1/T
from phasepy.fit import fit_rk
mixrk = mix.copy()
rk0 = np.array([0, 0])
fit_rk(rk0, mixrk, datavle, Tdep = False)
fit_rk(rk0, mixrk, datavle, Tdep = False, virialmodel = 'ideal_gas')
# After the optimizations have been carried out, fitted data can be compared against experimental data.
# +
from phasepy import virialgamma
from phasepy.equilibrium import bubbleTy
prkij = prsveos(mixkij)
virialnrtl = virialgamma(mixnrtl, actmodel = 'nrtl')
virialwilson = virialgamma(mixwilson, actmodel = 'wilson')
virialrk = virialgamma(mixrk, actmodel = 'rk')
Ykij = np.zeros_like(Yexp)
Tkij = np.zeros_like(Pexp)
Ynrtl = np.zeros_like(Yexp)
Tnrtl = np.zeros_like(Pexp)
Ywilson = np.zeros_like(Yexp)
Twilson = np.zeros_like(Pexp)
Yrk = np.zeros_like(Yexp)
Trk = np.zeros_like(Pexp)
n = len(Pexp)
for i in range(n):
Ykij[:,i],Tkij[i] = bubbleTy(Yexp[:,i],Texp[i],Xexp[:,i],Pexp[i],prkij)
Ynrtl[:,i],Tnrtl[i] = bubbleTy(Yexp[:,i],Texp[i],Xexp[:,i],Pexp[i],virialnrtl)
Ywilson[:,i],Twilson[i] = bubbleTy(Yexp[:,i],Texp[i],Xexp[:,i],Pexp[i],virialwilson)
Yrk[:,i],Trk[i] = bubbleTy(Yexp[:,i],Texp[i],Xexp[:,i],Pexp[i],virialrk)
# +
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10,8))
ax=fig.add_subplot(221)
ax.plot(Xexp[0], Texp,'.', Yexp[0], Texp,'.')
ax.plot(Xexp[0], Tkij, Ykij[0], Tkij)
ax.set_xlabel('x,y')
ax.set_ylabel('T/K')
ax.text(0.5, 370, 'QMR')
ax2 = fig.add_subplot(222)
ax2.plot(Xexp[0], Texp,'.', Yexp[0], Texp,'.')
ax2.plot(Xexp[0], Tnrtl, Ynrtl[0], Tnrtl)
ax2.set_xlabel('x,y')
ax2.set_ylabel('T/K')
ax2.text(0.5, 370, 'NRTL')
ax3 = fig.add_subplot(223)
ax3.plot(Xexp[0], Texp,'.', Yexp[0], Texp,'.')
ax3.plot(Xexp[0], Trk, Yrk[0], Trk)
ax3.set_xlabel('x,y')
ax3.set_ylabel('T/K')
ax3.text(0.5, 370, 'Redlich-Kister')
ax4 = fig.add_subplot(224)
ax4.plot(Xexp[0], Texp,'.', Yexp[0], Texp,'.')
ax4.plot(Xexp[0], Twilson, Ywilson[0], Twilson)
ax4.set_xlabel('x,y')
ax4.set_ylabel('T/K')
ax4.text(0.5, 370, 'Wilson')
fig.show()
# -
| examples/Fitting Interaction Parameters for Mixtures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
def map_income(income):
if income<=9036.8:
return 'Low'
else:
return 'High'
average_number = pd.read_csv('../../TABLAS LATEX/languages.csv', sep=';', decimal=',')
average_number = average_number[['Language', 'Average']]
average_number
# # LOAD BIG MAC INDEX
big_mac_index = pd.read_csv('../data/big-mac-2021-01-01.csv')
big_mac_index = big_mac_index[['iso_a3', 'dollar_price']]
big_mac_index = big_mac_index.rename(columns={'dollar_price': 'big_mac_dollar_price', 'iso_a3': 'nationality'})
big_mac_index.head()
# +
import pycountry
stored_countries = {}
def map_iso2_to_iso3(iso2):
try:
if iso2 not in stored_countries:
iso3 = pycountry.countries.get(alpha_2=iso2).alpha_3
stored_countries[iso2] = iso3
return iso3
else:
return stored_countries[iso2]
except:
print(iso2)
return None
# -
# # Preply
preply = pd.read_csv('../data/results/final_dataframes/preply.csv', index_col=0)
preply = preply.drop_duplicates(subset=['user_name', 'language'])
preply['income_level'] = preply['income_level'].apply(map_income)
preply['nationality'] = preply['nationality'].apply(map_iso2_to_iso3)
preply = pd.merge(preply, big_mac_index, on='nationality', how='left')
preply.head()
preply['price_to_big_mac'] = preply.price/preply.big_mac_dollar_price
preply.head()
preply.columns
# +
from scipy.stats import ks_2samp
import numpy as np
def hypothesis_test(group1, group2, alpha=0.1):
st, p_value = ks_2samp(group1, group2)
if p_value<alpha:
return st, p_value
else:
return st, p_value
def compute_aggregated_feature_top_k(df, top_k, language_col, aggregation_col, target_cols, group1, group2):
count_group1 = 'count_{}'.format(group1)
count_group2 = 'count_{}'.format(group2)
variance_group1 = 'variance_{}'.format(group1)
variance_group2 = 'variance_{}'.format(group2)
mean_group1 = 'mean_{}'.format(group1)
mean_group2 = 'mean_{}'.format(group2)
median_group1 = 'median_{}'.format(group1)
median_group2 = 'median_{}'.format(group2)
results = pd.DataFrame(columns=['language', 'top_k', 'target_col', 'aggregation_col', mean_group1, median_group1, mean_group2, median_group2, count_group1, count_group2, variance_group1, variance_group2, 'statistic', 'p_value'])
for lang in df[language_col].unique():
temp = df[df[language_col]==lang]
temp = temp.sort_values(by='position', ascending=True)
for target in target_cols:
temp = temp.dropna(subset=[target])
if top_k is not None:
temp = temp.head(top_k)
temp[target] = pd.to_numeric(temp[target], errors='coerce')
g1 = temp[temp[aggregation_col]==group1][target].values
g2 = temp[temp[aggregation_col]==group2][target].values
g1_count = len(g1)
g2_count = len(g2)
g1_mean = np.nanmean(g1) if g1_count else None
g2_mean = np.nanmean(g2) if g2_count else None
g1_median = np.nanmedian(g1) if g1_count else None
g2_median = np.nanmedian(g2) if g2_count else None
g1_var = g1.var() if g1_count else None
g2_var = g2.var() if g2_count else None
#Hypothesis testing
if len(g1)>0 and len(g2)>0:
st, p_value = hypothesis_test(g1, g2)
else:
st, p_value = None, None
results = results.append({'language': lang, 'top_k': len(temp), 'target_col': target, 'aggregation_col': aggregation_col,
mean_group1: g1_mean, median_group1: g1_median, mean_group2: g2_mean, median_group2: g2_median, count_group1: g1_count, count_group2: g2_count,
variance_group1: g1_var, variance_group2: g2_var, 'statistic': st, 'p_value': p_value}, ignore_index=True)
return results
# -
preply_results_40 = compute_aggregated_feature_top_k(preply, 40, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
preply_results_all = compute_aggregated_feature_top_k(preply, None, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
preply_results = pd.concat([preply_results_40, preply_results_all])
preply_results = pd.merge(preply_results, average_number, how='left', left_on='language', right_on='Language')
preply_results = preply_results.rename(columns={'Average': 'average_num_teachers'})
preply_results.head()
preply_results.to_csv('../data/results/features_analysis/income/preply_big_mac.csv')
# # Italki
italki = pd.read_csv('../data/results/final_dataframes/italki.csv', index_col=0)
italki = italki.drop_duplicates(subset=['user_id', 'language'])
italki['income_level'] = italki['income_level'].apply(map_income)
italki['nationality'] = italki['nationality'].apply(map_iso2_to_iso3)
italki = pd.merge(italki, big_mac_index, on='nationality', how='left')
italki.head()
italki['price_to_big_mac'] = italki.price/italki.big_mac_dollar_price/100
italki.head()
italki.head()
italki.columns
italki_results_40 = compute_aggregated_feature_top_k(italki, 40, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
italki_results_all = compute_aggregated_feature_top_k(italki, None, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
italki_results = pd.concat([italki_results_40, italki_results_all])
italki_results = pd.merge(italki_results, average_number, how='left', left_on='language', right_on='Language')
italki_results = italki_results.rename(columns={'Average': 'average_num_teachers'})
italki_results.head()
italki_results.to_csv('../data/results/features_analysis/income/italki_big_mac.csv')
italki_results[italki_results['p_value']<0.1]
# # Verbling
verbling = pd.read_csv('../data/results/final_dataframes/verbling.csv', index_col=0)
verbling = verbling.drop_duplicates(subset=['first_name', 'last_name', 'language'])
verbling['income_level'] = verbling['income_level'].apply(map_income)
verbling['nationality'] = verbling['nationality'].apply(map_iso2_to_iso3)
verbling = pd.merge(verbling, big_mac_index, on='nationality', how='left')
verbling.head()
verbling['price_to_big_mac'] = verbling.price/verbling.big_mac_dollar_price
verbling.head()
verbling.columns
verbling_results_40 = compute_aggregated_feature_top_k(verbling, 40, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
verbling_results_all = compute_aggregated_feature_top_k(verbling, None, 'language', 'income_level', ['price_to_big_mac'], 'High', 'Low')
verbling_results = pd.concat([verbling_results_40, verbling_results_all])
verbling_results = pd.merge(verbling_results, average_number, how='left', left_on='language', right_on='Language')
verbling_results = verbling_results.rename(columns={'Average': 'average_num_teachers'})
verbling_results.head()
verbling_results.to_csv('../data/results/features_analysis/income/verbling_big_mac.csv')
verbling_results[verbling_results['p_value']<0.1]
| analysis/statistical_price_analysis/Features analysis wrt nationality BIG MAC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ipywidgets as widgets
# ## Button event
#
# https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20Events.html#
print(widgets.Button.on_click.__doc__)
# +
button = widgets.Button(description="Click Me!", button_style="primary")
output = widgets.Output()
def on_button_clicked(b):
with output:
print("Button clicked.")
button.on_click(on_button_clicked)
widgets.VBox([button, output])
# -
# ## Other widget events
#
# https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20Events.html#Traitlet-events
print(widgets.Widget.observe.__doc__)
# ### First example
# +
buttons = widgets.ToggleButtons(
value=None,
options=["Show", "Hide", "Close"],
button_style="primary",
)
buttons.style.button_width = "80px"
html = widgets.HTML(
value='<img src="https://earthengine.google.com/static/images/earth-engine-logo.png" width="100" height="100">'
)
vbox = widgets.VBox([buttons, html])
vbox
# +
def handle_btn_click(change):
if change['new'] == 'Show':
vbox.children = [buttons, html]
elif change['new'] == 'Hide':
vbox.children = [buttons]
elif change['new'] == 'Close':
buttons.close()
html.close()
vbox.close()
buttons.observe(handle_btn_click, "value")
# -
# ### Second example
# +
dropdown = widgets.Dropdown(
options=["Landsat", "Sentinel", "MODIS"],
value=None,
description="Satellite:",
style={"description_width": "initial"},
layout=widgets.Layout(width="250px")
)
btns = widgets.ToggleButtons(
value=None,
options=["Apply", "Reset", "Close"],
button_style="primary",
)
btns.style.button_width = "80px"
output = widgets.Output()
box = widgets.VBox([dropdown, btns, output])
box
# +
def dropdown_change(change):
if change['new']:
with output:
output.clear_output()
print(change['new'])
dropdown.observe(dropdown_change, "value")
# +
def button_click(change):
with output:
output.clear_output()
if change['new'] == "Apply":
if dropdown.value is None:
print("Please select a satellie from the dropdown list.")
else:
print(f"You selected {dropdown.value}")
elif change['new'] == 'Reset':
dropdown.value = None
else:
box.close()
btns.observe(button_click, "value")
# -
# ### Third example
#
# - https://geemap.org/notebooks/39_timelapse
# - https://geemap.org/notebooks/71_timelapse
# - https://gishub.org/gee-ngrok
#
# 
| examples/ipywidgets_events.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from glob import glob
import pickle
import numpy as np
import pandas as pd
import re
from IPython.display import display
# -
class Aggregator:
def __init__(self, path, task, logname="log.log", flagname='log_flags.pkl', verbose=False):
self.path = path
self.task = task
self.flagfiles = [file for path, subdir, files in os.walk(os.path.join(path, task))
for file in glob(os.path.join(path, flagname))]
if verbose:
print(self.flagfiles)
# logs = [file for path, subdir, files in os.walk(os.path.join(path, task))
# for file in glob(os.path.join(path, logname))]
self.logfiles = [f.replace(flagname, logname) for f in self.flagfiles]
self.rename = {'binarize_MNIST': 'binarized',
'normalise_fb': 'Nfb',
'num_glimpses': 'glimpses',
'num_classes_kn': 'KK',
'num_uk_test': 'UU',
'num_uk_test_used': 'UU used',
'num_uk_train': 'KU',
'scale_sizes': 'scales',
'size_z': 'z size',
'uk_cycling': 'cycl',
'z_B_center': 'z center (B)',
'z_dist': 'z dist',
'z_B_kl': 'eq',
'rl_reward': 'R',
'pre_train_epochs': 'pre',
'pre_train_policy': 'pre policy',
}
def _rename(self, columns):
return [self.rename[c] if (c in self.rename.keys()) else c for c in columns]
def _parse_results(self, file, keyword='TEST: '):
results = {}
with open(file, 'r') as f:
text = f.read()
test_log = re.findall('(?<={}).*(?=\n)'.format(keyword), text)
if test_log:
final = test_log[-1].replace(':', '').split(' ')
for i in range(len(final) // 2):
name = final[2*i].split('/')[-1]
value = final[2*i + 1]
results[name] = value
return results
def get_overview(self, param_cols=None, metrics=None, groupby='glimpses', sortby=None, incl_last_valid=False):
rs = []
for log, flag in zip(self.logfiles, self.flagfiles):
with open(flag, 'rb') as f:
params = pickle.load(f)
results = self._parse_results(log)
if incl_last_valid:
results_valid = self._parse_results(log, keyword='VALID: ')
results.update({'val_' + k: v for k, v in results_valid.items()})
exp_name = log.split('\\')[-2]
if results:
results.update(params)
results['exp_name'] = exp_name
rs.append(results)
if not rs:
return
df = pd.DataFrame(rs)
# df = df.set_index('name')
df.columns = self._rename(df.columns)
df['scales'] = df['scales'].apply(lambda v: '{}x{}'.format(len(v), v[0]))
self.available_columns = sorted(df.columns)
if param_cols is not None:
df = df.set_index(param_cols)
if metrics is not None:
if incl_last_valid:
metrics += ['val_' + m for m in metrics]
df = df[metrics + [groupby]]
df = df.pivot(columns=groupby).swaplevel(axis=1).sort_index(axis=1, level=0, sort_remaining=False)
if sortby is not None:
df = df.sort_values(sortby, ascending=False)
return df
PATH = os.path.join('logs')
print(os.listdir(PATH))
# # MNIST
# +
params = ['planner', 'scales', 'pre', 'pre policy', 'z size', 'z dist', 'eq', 'z center (B)', 'Nfb']
metrics = ['acc', 'f1', 'loss', 'T']
mnist = Aggregator(PATH, 'MNIST/rl3', verbose=False)
df = mnist.get_overview(params, metrics, sortby=(7, 'f1'))
df
# +
params = ['planner', 'scales', 'pre', 'pre policy', 'R', 'z size', 'z dist', 'eq', 'z center (B)', 'Nfb']
metrics = ['acc', 'f1', 'loss', 'T']
mnist = Aggregator(PATH, 'MNIST/AI8', verbose=False)
df = mnist.get_overview(params, metrics, sortby=(5, 'f1'))
df
# +
params = ['planner', 'scales', 'pre', 'pre policy', 'R', 'z size', 'z dist', 'eq', 'z center (B)', 'Nfb']
metrics = ['acc', 'f1', 'loss', 'T']
mnist = Aggregator(PATH, 'MNIST/AI_pixel8', verbose=False)
df = mnist.get_overview(params, metrics, sortby=(5, 'f1'))
df
# -
# # MNIST_UK
# +
params = ['planner', 'scales', 'pre', 'pre policy', 'z size', 'z dist', 'z center (B)', 'Nfb',
'KK', 'KU', 'UU', 'UU used', 'cycl']
metrics = ['f1', 'acc', 'acc_kn', 'acc_uk', 'loss', 'T', 'pct_noDecision']
mnist_uk = Aggregator(PATH, 'MNIST_UK', verbose=False)
df = mnist_uk.get_overview(params, metrics, sortby=(7, 'f1'))
df
# -
# # MNIST_OMNI_notMNIST
# +
params = ['planner', 'scales', 'pre', 'pre policy', 'z size', 'z dist', 'z center (B)', 'Nfb',
'uk_pct', 'KK', 'KU', 'UU', 'binarized']
metrics = ['f1', 'acc', 'acc_kn', 'acc_uk']
mnist_omni_notmnist = Aggregator(PATH, 'MNIST_OMNI_notMNIST/rl')
df = mnist_omni_notmnist.get_overview(params, metrics, sortby=(7, 'f1'), incl_last_valid=True)
df
| notebooks/metrics_overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python3
from collections import defaultdict
import matplotlib
matplotlib.use("PGF")
import matplotlib.pyplot as plt
import numpy as np
import re
from scipy.optimize import minimize
import sys
sys.path.append("../../enrichment/src/")
from analyser import Analyser, Plotter, atom_to_mass_frac, mass_to_atom_frac
from multi_isotope_calculator import Multi_isotope
import plotsettings as ps
plt.style.use("seaborn")
plt.rcParams.update(ps.tex_fonts())
# -
# Global variables
NUC_ID = (922320000, 922330000, 922340000, 922350000, 922360000, 922380000)
SEPARATION_FACTOR = 1.35
def simulation_tails(fname, uranium_type="Natural"):
"""Get the reprocessed and depleted uranium tails as dict"""
if uranium_type=="Natural":
sink = "DepletedNaturalUSink"
elif uranium_type=="Reprocessed":
sink = "DepletedReprocessedUSink"
else:
msg = "'uranium_type' has to be either 'Natural' or 'Reprocessed'"
raise ValueError(msg)
a = Analyser(fname)
sim_end = a.query(selection='EndTime', table='Finish')[0][0]
results = a.query(selection='NucId, Quantity',
table='ExplicitInventory',
condition='Time==? AND AgentId==?',
vals=(sim_end, a.names[sink]))
comp = dict([(key, 0) for key in range(232, 239) if key!=237] )
quantity = 0
for key, value in results:
key = key/10000 - 92*1000
comp[key] = value
quantity += value
for key, value in comp.items():
comp[key] = value / quantity
return comp, quantity
# + slideshow={"slide_type": "-"}
def enrichment_feed_and_tails(origin, burnup):
"""Prepare data: feed used and expected tails composition"""
if origin=="Natural":
#Mass fractions of natU enrichment tails taken from the Cyclus
# output file
natU_comp = {'234': 0.0054, '235': (0.7204, 2, 0.3)}
seu_tails = np.array([0., 0., 1.5440247618063e-05,
0.00290322658192604, 0.,
0.997081333170456])
heu_tails = np.array([0., 0., 1.27218682709261e-05,
0.00285479562964945, 0.,
0.99713248250208])
return natU_comp, (seu_tails, heu_tails)
elif origin=="Reprocessed":
# Load, filter and format feed data
data = np.load("../data/SERPENT_outputs_NatU_percentages.npy").item()
feed_composition = {}
normalisation = 0
for iso in [i for i in range(234, 239) if i!=237]:
value = data[burnup][f"U{iso}"]
feed_composition[str(iso)] = value * 100
normalisation += value
for key, val in feed_composition.items():
feed_composition[key] = val/normalisation
feed_composition['232'] = 0.
feed_composition['233'] = 0
# The U238 content is calculated by the enrichment module
del feed_composition['238']
# Get SEU and HEU tails
if burnup=="0.5MWd":
seu_tails = np.array([0., 0., 1.35406410557832e-05,
0.00269133511129306, 4.13592084547905e-05,
0.997253765039196])
elif burnup=="2MWd":
seu_tails = np.array([0., 0., 1.56662456546925e-05,
0.00269248329581373, 0.000163308471630726,
0.997128541986901])
else:
raise ValueError("'burnup' has to be '0.5MWd' or '2MWd'")
concentration = feed_composition
concentration['235'] = (feed_composition['235'], 90., 0.3)
m = Multi_isotope(concentration, feed=1, process='centrifuge',
alpha235=SEPARATION_FACTOR, downblend=True)
m.calculate_staging()
heu_tails = m.xt
return feed_composition, (seu_tails, heu_tails)
else:
raise ValueError("'origin' has to be 'Natural' or 'Reprocessed'")
# -
def mix_compositions(comp1, comp2, mass1, mass2):
return (mass1*comp1 + mass2*comp2) / (mass1+mass2)
def mixing_ratios(sim_tails_comp, tails_comp):
"""Calculate how much of comp1 is added to comp2 using mix_comp
Here, the mixing with the following compositions is calculated:
mix_comp = (a*comp1 + b*comp2) / (a+b)
b is set to 1 such that this function calculates how much of
comp1 is added to comp2 per unit of comp2. In other words, a is
given in units of comp2.
"""
# Assure correct formatting
sim_tails_comp = np.array(list(sim_tails_comp.values()))
# special case: mix comp contains no comp2: return a large number
if np.all(sim_tails_comp - tails_comp[0] < 1e-10):
print(f"Only SEU tails, no HEU tails produced!")
return 1e200
mass_ratio = ((tails_comp[1]-sim_tails_comp)
/ (sim_tails_comp-tails_comp[0]))
if np.std(mass_ratio[~np.isnan(mass_ratio)]) > 1e-10:
print()
msg = (f"Values differ from each other!\n"
+ f"mass_ratio:\n{mass_ratio}\n"
+ f"Composition 1:\n{tails_comp[0]}\n"
+ f"Composition 2:\n{tails_comp[1]}\n"
+ f"Mixed final composition:\n{sim_tails_comp}")
raise RuntimeError(msg)
# Remove possible nans from isotopes
mass_ratio = np.mean(mass_ratio[~np.isnan(mass_ratio)])
return mass_ratio
def tails_per_product_qty(concentrations, enrichment_level):
m = Multi_isotope(concentrations, max_swu=np.inf, feed=np.inf,
product=1, downblend=True, process='centrifuge',
alpha235=SEPARATION_FACTOR)
m.set_product_enrichment(enrichment_level)
m.calculate_staging()
tails = m.t
product = m.p
if abs(product-1) > 1e-10:
raise RuntimeError("Something fishy going on here")
return tails / product
def tails_qty(origin, burnup):
"""Calculate the amount of HEU and SEU produced"""
print(f"\n{origin} uranium, burnup of {burnup}")
# Get tails composition in depleted U sink from simulation
fname_burnup = re.sub("\.", "", burnup)
fname = (f"../data/run_two_repositories_{fname_burnup}_0/"
+ f"run_two_repositories_{fname_burnup}.sqlite")
sim_tails_comp, sim_tails_qty = simulation_tails(fname,
uranium_type=origin)
# Get feed and predicted tails compositions
feed_comp, tails_comp = enrichment_feed_and_tails(origin, burnup)
seu_per_heu_tails = mixing_ratios(sim_tails_comp, tails_comp)
seu_tails_qty = (sim_tails_qty * seu_per_heu_tails
/ (1.+seu_per_heu_tails))
heu_tails_qty = sim_tails_qty / (1.+seu_per_heu_tails)
print(f"Total qty: {sim_tails_qty:9.0f} kg\n"
+ f"SEU tails: {seu_tails_qty:9.0f} kg\n"
+ f"HEU tails: {heu_tails_qty:9.0f} kg\n")
enrichment_lvl = (1.1, 90.)
label = ("SEU", "HEU")
tails = (seu_tails_qty, heu_tails_qty)
for xp, name, tail in zip(enrichment_lvl, label, tails):
t_per_p = tails_per_product_qty(feed_comp, xp)
product = tail / t_per_p
print(f"Produced {name}: {product:9.1f} kg")
return
def main():
burnup = ("0.5MWd", "2MWd")
origin = ("Natural", "Reprocessed")
for bu in burnup:
for orig in origin:
tails_qty(orig, bu)
return
| analysis/notebook_leonia_tails.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.io)
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2019.</p>
| 25_despliegue_en_aws.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Requirements/Libraries
import requests
from owlready2 import *
from rdflib import Graph
from SPARQLWrapper import SPARQLWrapper, JSON
# ## Load an ontology and print 10 classes
# +
#Load ontology
urionto="http://www.cs.ox.ac.uk/isg/ontologies/dbpedia.owl"
onto = get_ontology(urionto).load()
print("Classes in DBpedia ontology: " + str(len(list(onto.classes()))))
#Print 10 classes
i=0
for cls in onto.classes():
if "http://dbpedia.org/ontology/" in cls.iri:
i=i+1
print("\t"+cls.iri)
if i==9:
break
# -
# ## Loads and queries a local RDF Knowledge Graph
# +
file="../files/nobelprize_kg.nt"
g = Graph()
g.parse(file, format="nt")
print("\n\n" + file + " has '" + str(len(g)) + "' triples.")
#SPARQL query
nobelprize_query = """
SELECT DISTINCT ?x WHERE {
?laur <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> <http://data.nobelprize.org/terms/Laureate> . ?laur <http://www.w3.org/2000/01/rdf-schema#label> ?x .
?laur <http://xmlns.com/foaf/0.1/gender> \"female\" .
}"""
qres = g.query(nobelprize_query)
print("Female laureates:")
for row in qres:
print("%s" % row)
# -
# ## Queries remote Knowledge Graph via its Endpoint
# +
dbpedia_endpoint = "http://dbpedia.org/sparql"
dbpedia_query = """
SELECT DISTINCT ?x WHERE {
<http://dbpedia.org/resource/Chicago_Bulls> <http://www.w3.org/1999/02/22-rdf-syntax-ns#type> ?x .
}"""
sparqlw = SPARQLWrapper(dbpedia_endpoint)
sparqlw.setReturnFormat(JSON)
sparqlw.setQuery(dbpedia_query)
results = sparqlw.query().convert()
#Prints JSON file with results
#print(results)
print("\nQuerying DBPedia Knowledge Graph (types of Chicago Bulls)")
for result in results["results"]["bindings"]:
#Prints individual results
print(result["x"]["value"])
# -
# ## Gets pre-computed vector embedding for a KG entity
# +
print("\nVector embedding for the KG resource 'Chicago Bulls':")
#http://dbpedia.org/resource/Chicago_Bulls
kg_entity = "Chicago_Bulls"
r = requests.get('http://www.kgvec2go.org/rest/get-vector/dbpedia/' + kg_entity)
print(r.text)
# -
| lab1/lab1-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# The Centre for Research on the Epidemiology of Disasters' Data set about the American Typhoons (2000-2022)
# 1. Determine the typhoon(s) from 2000-2022 that brought the greatest number of casualties to the different locations in America.
#
# +
import pandas as pd
# casualties of storms in America based in EMDAT datasets
data = pd.read_excel(r'Data sets\2000-2022-emdat_public_2022_04_24_query_uid-XuKaJG.xlsx', sheet_name="Filtered Data Level 2")
df = pd.DataFrame(data) #convert dataset into dataframe
df = df.fillna(0) # replace NaN with zero value
#*************************NEW DATAFRAME***************************************
#selecting all needed and specific columns from original dataframe/dataset and creating new dataframe named new_df
new_df = df.iloc[:,[0,1,6,7,9,10, 12, 34, 35, 36, 37,38, 40, 41, 42, 43, 44]].copy()
#changing data types of specific columns using dictionary
convert_datatypes = {"Total Deaths":int,
"No Injured":int,
"No Affected":int,
"No Homeless":int,
"Total Affected":int,
"Reconstruction Costs, Adjusted ('000 US$)": int,
"Insured Damages ('000 US$)": int,
"Insured Damages, Adjusted ('000 US$)": int,
"Total Damages ('000 US$)":int,
"Total Damages, Adjusted ('000 US$)": int}
new_df= new_df.astype(convert_datatypes) #converting columns datatypes
#***********************SUB DATAFRAME (YEAR 2000)***************************
#creating subdataframe name=typ_y2000 and selecting rows where year=2000
typ_y2000 = new_df.loc[new_df['Year']==2000].copy()
#selecting columns for sorting based on column "Total Affected"
typ_y2000_af = typ_y2000.iloc[:,[0,1,3,4,5,11]]
typ_y2000_af = typ_y2000_af.sort_values(by = 'Total Affected', ascending = False) #sorting of data in descending order
print('\nYear:2000 \n(Sorted by column: Total Affected)')
display(typ_y2000_af) #displaying of selected data
# +
#selecting columns for sorting based on column "Total Deaths"
typ_y2000_d = typ_y2000.iloc[:,[0,1,3,4,5,7]]
typ_y2000_d = typ_y2000_d.sort_values(by = 'Total Deaths', ascending = False) #sorting of data in descending order
print('\n(Sorted by column: Total Deaths)')
display(typ_y2000_d) #displaying of selected data
# +
#selecting columns for sorting based on column "Total Damages, Adjusted"
typ_y2000_DA = typ_y2000.iloc[:,[0,1,3,4,5,16]]
typ_y2000_DA = typ_y2000_DA.sort_values(by = "Total Damages, Adjusted ('000 US$)", ascending = False) #sorting of data in descending order
print("\n(Sorted by column: Total Damages, Adjusted ('000 US$))")
display(typ_y2000_DA) #displaying of selected data
# -
display(new_df)
#
| EDA to Typhoon Mitigation and Response Framework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="kdOgSV5Io03P"
# `Add csv files on to colab..`
# + [markdown] id="pY8n2JfPo8QZ"
# Install RDKit library to start working with python...
# + colab={"base_uri": "https://localhost:8080/"} id="Q-dgRhRkwAb0" outputId="9ad7654c-990b-4872-ad34-f4e4f05e2df0"
# Install RDKit.
# !pip install kora
import kora.install.rdkit
# + id="O4II53mRrPp7"
import kora.install.rdkit
# + id="RXg_4dQFwKPg"
from rdkit import Chem
from rdkit.Chem import Draw
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem import rdMolDescriptors
from rdkit.Chem import Descriptors
from rdkit.Chem import AllChem
from rdkit import DataStructs
import numpy as np
# + [markdown] id="ARbisBI1pE27"
# `Reading Sider data... (Smiles + ADR Labels)`
# + colab={"base_uri": "https://localhost:8080/", "height": 343} id="PN3fTVmmV7bj" outputId="ec526384-fc5d-49d2-da9a-13c21f4f1d51"
import pandas as pd
sider = pd.read_csv('sider.csv')
sider.head()
# + colab={"base_uri": "https://localhost:8080/"} id="-rKQY6ta6CDG" outputId="c5a47e41-acaf-4844-becb-e4771f033272"
sider.shape
# + [markdown] id="L_Q0DCpWdkgi"
# ### `Dropping Unnecessary Columns`
#
# As per the research,w.k.t 'Product Issues', 'Investigations', and
# 'Social Circumstances' was not useful for predicting..
# + colab={"base_uri": "https://localhost:8080/"} id="6o2Z7yxndl8X" outputId="ae0d6f82-eb77-4273-b3db-1b1802276b97"
'Investigations' in sider.columns
# + colab={"base_uri": "https://localhost:8080/"} id="naTC2fmXdu9T" outputId="9fe5f6ae-abfa-453f-c607-12ce55524cb5"
'Product issues' in sider.columns
# + colab={"base_uri": "https://localhost:8080/"} id="2rmwjWGJd5zr" outputId="17b305b5-7f28-4570-94ec-3f8c2051c81f"
'Social circumstances' in sider.columns
# + id="0nc3Zzdm1zXH"
sider.drop(['Investigations','Product issues' ,'Social circumstances'],axis=1,inplace=True)
# + [markdown] id="fkzb2ybD8Y_f"
# **Reading offside_socs_modified data**
# + colab={"base_uri": "https://localhost:8080/", "height": 326} id="JxkOgXN2zqAl" outputId="8e6d748a-a733-4b5e-c620-0a4b9bc68209"
offside_socs_modified = pd.read_csv('offside_socs_modified.csv')
offside_socs_modified.head()
# + colab={"base_uri": "https://localhost:8080/"} id="t72Nm1cR6EUu" outputId="e05d4c2f-6e44-47d9-edd1-054b7e974b3a"
offside_socs_modified.shape
# + [markdown] id="rNBGQSCw8s3R"
# **Checking for the common smile used in offside_socs_modified and sider**
#
# + colab={"base_uri": "https://localhost:8080/"} id="ZxM6Y9MX1R37" outputId="e6aaa669-e47c-493c-e6d8-f97ee90a9507"
df3 = pd.merge(offside_socs_modified, sider, on='smiles',how='outer', indicator='Exist')
df3 = df3.loc[df3['Exist'] =='both']
print('Number of Common smile in both data farmes ',df3.shape)
# + [markdown] id="OVXBchHRjB8z"
# ### Concatenating the Dataframes
# + id="N6evZUjl5CkT"
dfs = [offside_socs_modified, sider]
sider = pd.concat(dfs)
# + colab={"base_uri": "https://localhost:8080/"} id="aU9BtkNl6UWt" outputId="6a73e5f5-d67c-47c3-e373-d8a6a318f0c0"
print('Number of Columns in merged sider', sider.shape[0])
# + [markdown] id="dDLmm7TwjO5G"
# # Molecular Descriptors
# + [markdown] id="o9Wv6uAhA5XK"
# **Smiles to Molecules...**
#
#
# + id="mT4P1rX1WDaq" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="39150d3d-73cf-47a2-ce0d-1a2022d1d23b"
mol_list = []
for smile in sider.iloc[:5,0]:
mol = Chem.MolFromSmiles(smile) # Converting to Molecules images from smiles
mol_list.append(mol)
img = Draw.MolsToGridImage(mol_list, molsPerRow=5)
img
# + [markdown] id="ripR6z_kawD5"
# ### `Molecular Weights`
#
# Getting Molecular Weights from smiles
# + id="XGZbMGYNXjUD" colab={"base_uri": "https://localhost:8080/"} outputId="2068527d-4f9b-40f5-83e6-8d908e2b08b2"
mol_weights = []
mol_str=[]
for smile in sider.iloc[:,0]:
mol = Chem.MolFromSmiles(smile)
mol_str.append(mol)
mol_weights.append(Descriptors.MolWt(mol))
# + colab={"base_uri": "https://localhost:8080/"} id="p3LpfwQ2sB5J" outputId="03c10d5a-20d3-462e-9222-e0a16fc80559"
print(mol_weights)
# + [markdown] id="qkL8KQJCa2cy"
# Concatenating weights to sider..
# + id="i_INqiTw7FLP"
sider['Molecular_Weights']=mol_weights
# + colab={"base_uri": "https://localhost:8080/", "height": 326} id="hDaJLWRjanHS" outputId="a1047c36-727d-4ffc-fe0b-00ee792f42e1"
sider[:5]
# + [markdown] id="XKKrfbDRcGLT"
# ### `Radical Electrons`
#
# Getting no.of radical electrons from smiles
# + colab={"base_uri": "https://localhost:8080/"} id="DK6h6785aoPK" outputId="d8f3a259-798f-4e67-fd8a-0574fa54bb44"
radical_electrons = []
for smile in sider.iloc[:,0]:
mol = Chem.MolFromSmiles(smile)
radical_electrons.append(Descriptors.NumRadicalElectrons(mol))
# + id="TP91z4uQ7o38"
sider['Radical Electrons'] =radical_electrons
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="uAbxUZd_cmCy" outputId="8536fdc4-353b-4c56-ac05-3a07edc540e0"
sider[:4]
# + [markdown] id="Yv3jn5oNcsth"
# ### `Valence Electrons`
# + colab={"base_uri": "https://localhost:8080/"} id="8b9JT865cysf" outputId="fd3f5ddd-d48d-4f55-fd52-6ee0a867f4ce"
valence_electrons = []
for smile in sider.iloc[:,0]:
mol = Chem.MolFromSmiles(smile)
valence_electrons.append(Descriptors.NumValenceElectrons(mol))
# + id="ui6XEZhbdIVH"
sider['Valence Electrons'] = valence_electrons
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="r5zMABRfdMie" outputId="6bcb3314-7f63-4074-d853-c0e0d5cec905"
sider[:2]
# + id="TyfmSJzIfoZD"
# + [markdown] id="dxGC8kLwT5LI"
# ### Finger Print
# + id="diGbj2jogrb-" colab={"base_uri": "https://localhost:8080/"} outputId="d6a5c397-3e41-4399-c603-d8cb903432a8"
fingerprint = []
bi={}
for smile in sider.iloc[:,0]:
mol = Chem.MolFromSmiles(smile)
fingerprint.append(rdMolDescriptors.GetMorganFingerprintAsBitVect(mol, radius=2, bitInfo=bi))
# + id="6FjdeDnGsa28"
sider['fingerprint']=fingerprint
# + colab={"base_uri": "https://localhost:8080/", "height": 338} id="KtAzlGloh84g" outputId="c031dc84-a87c-4b2d-8f25-3078b49e7b82"
sider.head(2)
# + id="XR7N620Hsa9r"
# + id="5JCMIUI5sbBG"
# + id="ifbvtr89sbFL"
# + id="Nd-v8kj8sbI4"
# + id="nWFo8t5bsbP3"
# + id="thvGo-YpsbS9"
# + id="sI66TnokCVwL"
| CreatingFeatures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
__author__ = "__fbbianco"
'''
CUSP UCSL 2016
shows that the unmber of floors in Mannhattan building
are distributed in a Poissonian way (more or less)
'''
import pandas as pd
import pylab as pl
matplotlib.style.use('ggplot')
# %pylab inline
# I downloaded a database of building heights in Manhattan here http://chriswhong.github.io/plutoplus/#
pdh = pd.read_csv("pluto.csv")
# +
fig = pl.figure(figsize = (20,5))
#plot a histogram of the number of floors in Manhattan buildings
fig.add_subplot(1,2,1)
pdh.numfloors.hist(bins = np.arange(0,50,2.5), alpha = 0.7, label="Manhattan lots (Pluto)")
pl.legend()
pl.ylabel("Number of lots")
pl.xlabel("Number of floors")
xlim(0,50)
#plot a histogram of the number of floors in Manhattan buildings and overplot a Poisson distribution
fig.add_subplot(1,2,2)
pdh.numfloors.hist(bins = np.arange(0,50,2.5), alpha = 0.7, label="Manhattan lots (Pluto)")
pl.hist(np.random.poisson(5, size=40000), alpha = 0.7, color='k',
bins = np.arange(0,50,2.5), label = r"Poisson distribution $\lambda=5$")
pl.legend()
pl.ylabel("Number of lots")
pl.xlabel("Number of floors")
| notebooks/05-00 Statistics/manfloors_poisson.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.4
# language: julia
# name: julia-1.5
# ---
# # Functions and Modeling Applications
# Contents:
#
# - [Functions and Modeling Applications](#Functions-and-Modeling-Applications)
# - [Creating Functions](#Creating-Functions)
# - [Loops](#Loops)
# - [Linear Algebra](#Linear-Algebra)
# - [Finite Markov Chains](#Finite-Markov-Chains)
#
#
# This lab covers:
#
# (1) User-defined functions;
#
# (2) Loops;
#
# (3) Linear algebra applications;
#
# (4) Modeling finite-state Markov chains.
#
# ----
# ## Creating Functions
# In this section we will cover the basics of creating custom functions.
# A function is essentially an object that takes a set of inputs, applies some sort of procedure to said inputs, and spits out a result.
#
# Functions can be handy for organizing code that is likely to be routinely re-used in the future with varying inputs.
# Let's define a function named `add` that takes two variables, `x` and `y`, as inputs, and returns their sum.
function add(x, y)
z = x + y
return z
end
# To check, we call on `add()` with the inputs `x=2` and `y=3` -- obviously the output is supposed to be 5.
add(2,3)
# Success!
#
# Recall that we stored the output into a variable `z` that was defined inside of the function.
#
# If we try to call on `z` in the global environment (outside of the function), we will get an error since `z` only lives within the function:
z
# Now let's define a function called `all_operations` that takes takes two variables, `x` and `y`, as inputs, and returns their sum, difference, product, and quotient.
function all_operations(x, y)
sum = x + y
difference = x - y
product = x * y
quotient = x / y
result = (sum, difference, product, quotient)
return result
end
# To check whether `all_operations` works, we call on it with the inputs `x=1` and `y=2`:
all_operations(1, 2)
# Notice that the output of `all_operations()` is a tuple with four entires.
#
# Tuples are useful as function output objects because we can easily store their entries as separate variables:
# +
# Store output of `all_operations(1,2)` as separate variables
xy_sum, xy_difference, xy_product, xy_quotient = all_operations(1,2)
# Print all collected variables
@show xy_sum
@show xy_difference
@show xy_product
@show xy_quotient;
# -
# An alternative simpler way of defining the `all_operations` function by creating an equivalent `all_operations_v2`:
function all_operations_v2(x,y)
(x + y, x - y, x * y, x / y)
end
# What did we do differently?
# - We defined and stored all operation results in an unassigned tuple ;
# - We didn't use `return` at the end of the function to return the output.
#
# When we called on the function, Julia noticed the lack of a `return` command and chose the last thing it saw as the output -- in our case this was the unassigned tuple.
#
# Is this alternative way of defining `all_operations()` better? Not necessarily -- it depends on the context. For example, you might find that shorter code isn't *necessarily* easier to read.
#
# Let's just apply `all_operations()` and `all_operations_v2()` to the same inputs and check whether the outputs match using a custom-defined function `check()`:
# +
# Create var. `condition` that tests whether outputs are equivalent
condition = all_operations(1,2) == all_operations_v2(1,2)
# Create fun. `check` w/ input `condition`
function check(condition)
if condition == true
result = "The two functions are the same!"
end
if condition != true
result = "The two functions are not the same!"
end
return result
end
# Run `check` on `condition`
check(condition)
# -
# The `check()` function, as defined in the previous cell, is pretty clunky -- let's simplify it:
# +
# Re-define function `check()`
function check(condition)
if condition == true
return "The two functions are the same!"
else
return "The two functions are not the same!"
end
end
# Call on `check()`
check(condition)
# -
# Or alternatively:
# +
# Re-define function `check()
function check(condition)
if condition == true
return "The two functions are the same!"
end
"The two functions are not the same!"
end
# Call on `check()`
check(condition)
# -
# Again -- in the case of simple functions such as the ones shown above, being super efficient is not necessary.
#
# But clunky code can make larger scripts hard to read, and potentially even run slow!
# Now let's talk math.
#
# Defining mathematical functions in Julia is easy.
#
# Let's define the polynomial mapping $f:\mathbb{R} \rightarrow \mathbb{R}$ such that $f(x) = x^2 - 3x + 2$:
f(x) = x^2 - 3x + 2
# Suppose we're interested in knowing the value of $f(\pi)$:
f(pi)
# Alternatively:
f(π)
# There are a lot of details on user-defined functions that we haven't been able to cover here, but I think the above should be enough to get you started.
#
# Visit the official Julia manual section on [functions](https://docs.julialang.org/en/v1/manual/functions/) to learn more.
# ---
# ## Loops
# Let's print every integer between 1 and 5 using a while loop:
# +
# Initial value
i = 1
# While loop:
while i <= 5 # Run until i = 5
println(i) # Print i
i = i + 1 # Add 1 to i for the next iteration of the loop
end
# -
# The above can be accomplished more easily using a for loop:
for i in 1:5
println(i)
end
# We can pass any kind of sequence to a for loop.
#
# For example, we can print the set of odd numbers between 1 and 5 by defining a sequence called `sequence` and then iterating the values of said sequence:
# +
sequence = [1.0,3.0,5.0]
for i in sequence
println(i)
end
# -
# What if we want to instead print the index associated with the entries of `sequence` instead of the entry values themselves?
for i in eachindex(sequence)
println(i)
end
# Suppose we want to square all values of `sequence` and store it as a separate array called `seq_out`.
#
# We can accomplish this using a for loop that goes through each entry of `sequence`, squares it, and stores it as the corresponding entry of `seq_out`:
# +
# Declare `seq_out` as a vector
# w/ the same number of entries as `sequence`
seq_out = zeros(length(sequence))
# Run a for-loop that goes through
# the indexes of `sequence`
for i in eachindex(sequence)
seq_out[i] = sequence[i]^2
end
# Print `seq_out`
seq_out
# -
# Alternatively, we can use a for loop in a **comprehension**:
seq_out = [sequence[i]^2 for i in eachindex(sequence)]
# Even better -- we can **broadcast** (remember this from last lab?) `^2` across `sequence`.
seq_out = sequence.^2
# ---
# ## Linear Algebra
# First, we load up the `LinearAlgebra` package.
using LinearAlgebra
# Let's assume we have vectors $a_1 = (1, 2, 3)'$ and $a_2 = (4, 5, 6)'$.
#
# We start by defining these two column vectors:
a_1 = [1; 2; 3]
a_2 = [4, 5, 6];
# Recall that whether we use `;`'s or `,`'s to separate entries in single-entry arrays, we still get column vectors.
# Obviously $a_1$ and $a_2$ do not span each other, but let's just check to be sure:
A = [a_1 a_2]
b = A \ zeros(3)
# Since the zero vector is the only solution for $x$ in $A \, x = b$ where $A = [ a_1 a_2 ]$, then $a_1$ and $a_2$ must be linearly independent.
# Find the dot product of $a_1$ and $a_2$:
a_1' * a_2
# Alternatively:
dot(a_1, a_2)
# Now let's find $a_1 a_2'$:
a_1 * a_2'
# Now let's add the two vectors:
a_1 + a_2
# Subtract $a_2$ from $a_1$:
a_1 - a_2
# Let's scale vector $a_1$ by 3:
3a_1
# Equivalently:
3 * a_1
# Equivalently:
3 .* a_1
# The norm of vector $a_1$:
norm(a_1)
# Since $a_1$ and $a_2$ cannot span $\mathbb{R}^3$, we can find another orthogonal vector $a_3$.
#
# Is $a_3 = a_1 + a_2$ orthogonal? (Obviously not, by definition, but let's practice checking)
a_3 = a_1 + a_2
b = A \ a_3
# Since there exists a non-trivial solution to $x$ in $[a_1 \, a_2] \, x = A \, x = a_3$, then $a_3$ is not linearly independent.
# We can find a linearly independent $a_3$ by guessing some initial vector $b_3$, projecting it onto the columns of $A$ to obtain the projection $\hat{b}_3$, and then extracting the orthogonal $a_3 = b_3 - \hat{b}_3$:
b_3 = [2, 3, 4]
a_3 = b_3 - (A * inv(A'A) * A' * b_3)
a_3
# Now let's check whether $a_3$ is actually linearly independent by redefining $A$ as $A = [a_1 \, a_2 \, a_3]$ and solving for $x$ in $A \, x = 0$:
A = [a_1 a_2 a_3]
A \ zeros(3)
# Since the only solution for $x$ in $A \, x = 0$ is the trivial solution, then $A$ must be full-rank.
# Now let's check the eigenvalues and eigenvectors of $A$:
eigenv_A, eigenvec_A = eigen(A);
eigenv_A
eigenvec_A
# ---
# ## Finite Markov Chains
# Suppose that for a Markov process with three states we are given an initial state distribution and a stochastic matrix.
#
# We are told to compute the state density in 10 periods (at $t=10$).
#
# The given initial distribution is $P_0 = (1/3, 1/3, 1/3)'$, while the stochastic matrix is
#
# $$M = \begin{bmatrix} 0.95 & 0.05 & 0 \\ 0.15 & 0.75 & 0.1 \\ 0 & 0.5 & 0.5 \end{bmatrix} \, .$$
# +
# Define initial state distr.
P0 = [1/3, 1/3, 1/3]
# Define stochastic matrix
M = [0.95 0.05 0 ; 0.15 0.75 0.1 ; 0.0 0.5 0.5]
# Compute state distr. at t = 10
P10 = (P0' * M^(10))'
# -
# More generally, what if we're interested in computing the state density for a variety of $n \in \mathbb{N}$ periods?
#
# This is when writing custom functions comes in handy!
#
# Let's create a function named `markov_chain` with inputs for $P_0$, $M$, and $n$:
# +
function markov_chain(P0, M, n)
# Start by creating a copy of P0
# to later feed into the loop
P = copy(P0)
# Run a loop that computes
# P n-steps ahead
for i in 1:n
new_P = (P' * M)'
P = new_P
end
# Return the final state distr.
return P
end
# -
# We can now try our new function out using the previously-defined `P0` and `M` with `n` set to 10:
markov_chain(P0, M, 10)
# Does this match `P10`?
markov_chain(P0, M, 10) ≈ P10
# It does!
#
# We can now use this function to do a whole bunch of useful things.
#
# For example, given the same $P_0$ and $M$, we can now create a list of $P_n$ for $n = 1,2,\ldots,20$:
probabilities = [markov_chain(P0, M, n) for n in 1:20];
# We may visualize the probability of each state across time by gathering and plotting the corresponding series for all three states:
# +
state1 = zeros(20)
state2 = zeros(20)
state3 = zeros(20)
for i in 1:20
state1[i] = probabilities[i][1]
state2[i] = probabilities[i][2]
state3[i] = probabilities[i][3]
end
# -
# Let's load the `Plots` package to make a couple of plots.
using Plots
# First, let's make a simple plot that contains all series:
time = 1:20
plot(time, state1) # Plot state 1 density
plot!(time, state2) # Plot state 2 density
plot!(time, state3) # Plot state 3 density
# Notice that with time, being in state 1 becomes more likely, while states 2 and 3 becomes less likely.
# We may also create a separate plot for each series, but include them in a single composition:
p1 = plot(time, state1, title = "Normal Growth") # Plot state 1 probability
p2 = plot(time, state2, title = "Recession") # Plot state 2 probability
p3 = plot(time, state3, title = "Deep Recession") # Plot state 3 probability
plot(p1, p2, p3, layout = (3,1))
# Now let's write a function that computes the probability of a given sequence of outcomes.
#
# (You should have seen something like this on Assignment 1.)
# +
function outcome_prob(outcome, P0, M)
# Make sure `outcome` contains integers
# to allow for indexing
outcome = floor.(Int64, outcome)
# Store probability of initial state
probability = P0[outcome[1]]
# Compute probability of `outcome` sequence
for i in 2:length(outcome)
probability = probability * M[outcome[i-1], outcome[i]]
end
# Return `probability` -- prob. of sequence `outcome`
return probability
end
# -
# Let's keep using our previously-defined initial distribution $P_0$ and stochastic matrix $M$.
#
# We can now feed the function the following sequence of states to check its functionality: $11$ -- the probability of this sequence of outcomes should obviously be (1/3)(19/20) according to our defined $P_0$ and $M$.
outcome_prob(ones(2), P0, M)
# What if we want to find out the probability of having one of the following outcomes: (1) $11$, and (2) $22$ ?
#
# We can use `outcome_prob()` to compute the probability of each outcome, and then find their sum.
outcome_prob([1,1], P0, M) + outcome_prob([2,2], P0, M)
# ---
| week02/.ipynb_checkpoints/lab02-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from hw_demo_estimation import etl, graph_manipulation as gm, data_viz as dv, data_manipulation as dm
from networkx.algorithms.assortativity import average_neighbor_degree
from networkx.algorithms.cluster import clustering
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# # Demography Estimation Homework
#
# This is a markdown cell in a jupyter notebook where I can write analysis about the charts and the statistics that I calculated
nodes, edges = etl.load_and_select_profiles_and_edges()
G=gm.create_graph_from_nodes_and_edges(nodes, edges)
dv.plot_degree_distribution(G)
# descriptive analytics of the nodes
# age distribution by gender
dv.plot_age_distribution_by_gender(nodes)
edges_w_features = gm.add_node_features_to_edges(nodes, edges)
dv.plot_age_relations_heatmap(edges_w_features)
# THE END
directed = dm.compute_directed_edges(edges_w_features)
predict = dm.add_nbrs_by_gender(nodes, directed)
predict.loc[predict["TRAIN_TEST"]=="TEST", "gender"] = 1.0
predict.loc[(predict["TRAIN_TEST"]=="TEST") & (predict["0_nbrs"]>predict["1_nbrs"]), "gender"] = 0.0
predict[predict["TRAIN_TEST"]=="TEST"]["gender"]
dv.plot_node_statistic_by_gender(nodes, dict(G.degree))
dv.plot_node_statistic_by_gender(nodes, average_neighbor_degree(G))
dv.plot_node_statistic_by_gender(nodes, clustering(G))
directed = directed.dropna()
# +
directed["weight"] = 1/directed["smaller_id"].map(dict(G.degree()))
directed[["gender_x", "gender_y"]] = directed[["gender_x", "gender_y"]].replace({0:-1})
directed["y_axis"] = directed["gender_y"]*(directed["AGE_y"]-15)
to_plot = directed.groupby(["AGE_x", "gender_x", "y_axis", "smaller_id"]).sum()["weight"]
to_plot = to_plot.groupby(["AGE_x", "gender_x", "y_axis"]).mean()
df = to_plot[to_plot.index.get_level_values("gender_x").isin([1.0])].groupby(["AGE_x", "y_axis"]).sum()
df = df.reset_index().pivot("y_axis", "AGE_x", "weight")
from seaborn import heatmap
heatmap(df, cmap="coolwarm")
# +
df = to_plot[to_plot.index.get_level_values("gender_x").isin([-1.0])].groupby(["AGE_x", "y_axis"]).sum()
df = df.reset_index().pivot("y_axis", "AGE_x", "weight")
heatmap(df, cmap="coolwarm")
# -
for indexer in [
[True]*len(counter), # no filtering
(counter["gender_x"]==0.0) & (counter["gender_y"]==0.0),
(counter["gender_x"]==1.0) & (counter["gender_y"]==1.0),
((counter["gender_x"]==1.0) & (counter["gender_y"]==0.0)) | ((counter["gender_x"]==0.0) & (counter["gender_y"]==1.0))
]:
plt.figure()
counter["temp"] = 1.0
arr = counter[indexer].groupby(["AGE_x", "AGE_y"]).sum()["temp"].reset_index().pivot("AGE_x", "AGE_y", "temp").fillna(0).to_numpy()
arr = arr + arr.transpose()
heatmap(arr, cmap="coolwarm").invert_yaxis()
Ez az ábra azért más, mint a tanulmányban lévő, mert a gráfunkban súlyozatlan élek vannak
# +
import pandas
directed = dm.compute_directed_edges(edges_w_features).dropna()
directed["weight"] = 1/directed["smaller_id"].map(dict(G.degree()))
directed["age_diff"] = directed["AGE_y"]-directed["AGE_x"]
df = pandas.DataFrame()
df["AGE_x"]=directed["AGE_x"].unique()
df = df.set_index("AGE_x")
gender = {0.0: "M", 1.0: "F"}
plot_idx = [[], []]
for g1 in [0, 1]:
for g2 in [0.0, 1.0]:
for lower, upper in [(-5, 5), (20, 30), (-30, -20)]:
filtered = directed[(directed["gender_x"] == g1) & (directed["gender_y"] == g2) & (directed["age_diff"] > lower) & (directed["age_diff"] < upper) ]
to_plot = filtered.groupby(["smaller_id", "AGE_x"]).sum()["weight"]
to_plot = to_plot.groupby("AGE_x").mean().reset_index()
to_plot = to_plot.set_index("AGE_x")
name = " ".join([str(it) for it in [gender[g1], gender[g2], lower, upper]])
plot_idx[g1].append(name)
df[name] = to_plot["weight"]
df = df.fillna(0).sort_index()
df[plot_idx[0]].plot(figsize=(10, 10));
plt.figure()
df[plot_idx[1]].plot(figsize=(10, 10));
# -
# itt úgy tűnik, mintha a középső tartományból valamiért elvesztek volna az adatok
# # 3.feladat
from networkx import find_cliques
tris = [c for c in find_cliques(G) if len(c)==3]
gender_tris = [[n.loc[it, "gender"] for it in tri ] for tri in tris]
keys = [(3, 0), (2, 1), (1, 2), (0, 3)]
distr = dict()
for key in keys:
distr[key] = 0
for tri in gender_tris:
num_0 = tri.count(0)
num_1 = tri.count(1)
if (num_0, num_1) in keys:
distr[(num_0, num_1)] +=1
N = sum(distr.values())
for key in keys:
distr[key] /= N
distr
# A distr változó tartalmazza az ismert nemű csúcspontok alapján becsült eloszlását az egyes típusú háromszögeknek, ahol a típus alatt azt értjük, milyen nemből mennyit tartalmaz a háromszög. Feltételezve, hogy ezen eloszlás a teljes gráfra igaz maximum likelihood becslést végezhetünk az ismeretlen csúcsok nemére az alapján, hogy milyen típusú háromszögek jönnek létre, ha valamilyen neműnek vesszük az egyes csúcspontokat. Bár egyértelműen nem függetlenek egymástól a háromszögek típusa, ennek ellenére egy közelítő ML becslésként vehetjük a likelihoodot a létrejövő háromszögek valószínűségének a szorzatának.
| POKEC Network Analysis Stub for Students.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Text Using Markdown
#
# **If you double click on this cell**, you will see the text change so that all of the formatting is removed. This allows you to edit this block of text. This block of text is written using [Markdown](http://daringfireball.net/projects/markdown/syntax), which is a way to format text using headers, links, italics, and many other options. Hit _shift_ + _enter_ or _shift_ + _return_ on your keyboard to show the formatted text again. This is called "running" the cell, and you can also do it using the run button in the toolbar.
# # Code cells
#
# One great advantage of IPython notebooks is that you can show your Python code alongside the results, add comments to the code, or even add blocks of text using Markdown. These notebooks allow you to collaborate with others and share your work. The following cell is a code cell.
# +
# Hit shift + enter or use the run button to run this cell and see the results
print 'hello world'
# +
# The last line of every code cell will be displayed by default,
# even if you don't print it. Run this cell to see how this works.
2 + 2 # The result of this line will not be displayed
3 + 3 # The result of this line will be displayed, because it is the last line of the cell
# -
# # Nicely formatted results
#
# IPython notebooks allow you to display nicely formatted results, such as plots and tables, directly in
# the notebook. You'll learn how to use the following libraries later on in this course, but for now here's a
# preview of what IPython notebook can do.
# +
# If you run this cell, you should see the values displayed as a table.
# Pandas is a software library for data manipulation and analysis. You'll learn to use it later in this course.
import pandas as pd
df = pd.DataFrame({'a': [2, 4, 6, 8], 'b': [1, 3, 5, 7]})
df
# +
# If you run this cell, you should see a scatter plot of the function y = x^2
# %pylab inline
import matplotlib.pyplot as plt
xs = range(-30, 31)
ys = [x ** 2 for x in xs]
plt.scatter(xs, ys)
# -
# # Creating cells
#
# To create a new **code cell**, click "Insert > Insert Cell [Above or Below]". A code cell will automatically be created.
#
# To create a new **markdown cell**, first follow the process above to create a code cell, then change the type from "Code" to "Markdown" using the dropdown next to the run, stop, and restart buttons.
# # Re-running cells
#
# If you find a bug in your code, you can always update the cell and re-run it. However, any cells that come afterward won't be automatically updated. Try it out below. First run each of the three cells. The first two don't have any output, but you will be able to tell they've run because a number will appear next to them, for example, "In [5]". The third cell should output the message "Intro to Data Analysis is awesome!"
class_name = "Intro to Data Analysis"
message = class_name + " is awesome!"
message
# Once you've run all three cells, try modifying the first one to set `class_name` to your name, rather than "Intro to Data Analysis", so you can print that you are awesome. Then rerun the first and third cells without rerunning the second.
#
# You should have seen that the third cell still printed "Intro to Data Analysis is awesome!" That's because you didn't rerun the second cell, so even though the `class_name` variable was updated, the `message` variable was not. Now try rerunning the second cell, and then the third.
#
# You should have seen the output change to "*your name* is awesome!" Often, after changing a cell, you'll want to rerun all the cells below it. You can do that quickly by clicking "Cell > Run All Below".
#
# One final thing to remember: if you shut down the kernel after saving your notebook, the cells' output will still show up as you left it at the end of your session when you start the notebook back up. However, the state of the kernel will be reset. If you are actively working on a notebook, remember to re-run your cells to set up your working environment to really pick up where you last left off.
| .ipynb_checkpoints/ipython_notebook_tutorial-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
#creating single dimension array
a = np.array([1,3,23,45,2])
b = np.array([34,45,5,6,7])
a
b
#creating a 2D array
x = np.array([[1,34],[3,45],[23,5],[45,6],[2,7]])
x
x[0][1]
x[0:2]
x+3
x*2
x**3
x.shape
y = x.reshape(2,5)
y
#only columns
y[:,1]
z = np.zeros((5,5))
z
z = np.ones((5,5), dtype="int32")
z
np.add(x,x)
x/25
| Mathops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings("ignore")
df = pd.read_csv("../data/user_detail.csv")
df.review = df.review.apply(lambda x: x.replace("\r", " "))
df=df[['score', 'review']]
df.score = df.score.apply(lambda x: 0 if x<6 else 1)
# -
# ### train_test split
# +
from sklearn.model_selection import train_test_split
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords
train, test = train_test_split(df)
# -
def except_stopwords(words):
stopWords = set(stopwords.words('english'))
wordsFiltered = []
for w in words:
if w not in stopWords:
wordsFiltered.append(w)
return wordsFiltered
# ### tokenize + stopwords check
# %%time
train.review = train.review.apply(lambda x: except_stopwords(word_tokenize(x)))
test.review = test.review.apply(lambda x: except_stopwords(word_tokenize(x)))
# ### pos tagging
# +
from nltk.tag import pos_tag
def postagger(doc):
return ["/".join(p) for p in pos_tag(doc)]
# -
# %%time
train_docs = [(postagger(row[1]), row[0]) for row in train.values]
test_docs = [(postagger(row[1]), row[0]) for row in test.values]
# +
import pickle
with open('train_docs', 'wb') as f:
pickle.dump(train_docs, f)
with open('test_docs', 'wb') as f:
pickle.dump(test_docs, f)
# -
with open('train_docs', 'rb') as f:
train_docs = pickle.load(f)
with open('test_docs', 'rb') as f:
test_docs = pickle.load(f)
# ---
tokens = [t for d in train_docs for t in d[0]]
print(len(tokens))
# +
import nltk
from pprint import pprint
text = nltk.Text(tokens, name='NMSC')
print(len(text.tokens)) # returns number of tokens
print(len(set(text.tokens))) # returns number of unique tokens
pprint(text.vocab().most_common(10)) # returns frequency distribution
# -
# %matplotlib inline
text.plot(30)
# ---
# ### doc2vec
# +
from collections import namedtuple
TaggedDocument = namedtuple('TaggedDocument', 'words tags')
tagged_train_docs = [TaggedDocument(d, [c]) for d, c in train_docs]
tagged_test_docs = [TaggedDocument(d, [c]) for d, c in test_docs]
# -
from gensim.models import doc2vec
# 사전 구축
doc_vectorizer = doc2vec.Doc2Vec(vector_size=300, alpha=0.025, min_alpha=0.025, seed=0)
doc_vectorizer.build_vocab(tagged_train_docs)
doc_vectorizer.train(tagged_train_docs, total_words=200, epochs=10)
pprint(doc_vectorizer.wv.most_similar('super/VBP'))
pprint(doc_vectorizer.wv.most_similar('unfortunately/RB'))
# %%time
train_x = [doc_vectorizer.infer_vector(doc.words) for doc in tagged_train_docs]
train_y = [doc.tags[0] for doc in tagged_train_docs]
len(train_x)
# %%time
test_x = [doc_vectorizer.infer_vector(doc.words) for doc in tagged_test_docs]
test_y = [doc.tags[0] for doc in tagged_test_docs]
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state=0)
classifier.fit(train_x, train_y)
classifier.score(test_x, test_y)
# +
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import *
y_pred = classifier.predict(test_x)
print("acc:", accuracy_score(test_y, y_pred))
print(confusion_matrix(test_y, y_pred))
print(classification_report(test_y, y_pred))
# -
# Naive bayes와 비교
# - doc2vec와는 궁합이 잘 안맞는가...
# +
from sklearn.preprocessing import minmax_scale
train_x1 = minmax_scale(train_x)
test_x1 = minmax_scale(test_x)
from sklearn.naive_bayes import MultinomialNB
MNB = MultinomialNB(alpha=0.5)
MNB.fit(train_x1, train_y)
MNB.score(test_x1, test_y)
# +
y_pred = MNB.predict(test_x)
print("acc:", accuracy_score(test_y, y_pred))
print(confusion_matrix(test_y, y_pred))
print(classification_report(test_y, y_pred))
# -
from sklearn.linear_model import SGDClassifier
clf = SGDClassifier(loss='log', random_state=1, max_iter=1)
clf.fit(train_x, train_y)
clf.score(test_x, test_y)
# +
y_pred = clf.predict(test_x)
print("acc:", accuracy_score(test_y, y_pred))
print(confusion_matrix(test_y, y_pred))
print(classification_report(test_y, y_pred))
# -
# ### todo
# - gridsearch
| note/03.NLP_doc2vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Dq0kkUAc5O5n" colab_type="text"
# ## Dependencies
# + id="dQO2Qp272_ho" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="8d65d47c-2942-4c40-d561-84fecf7cb25d"
# !pip uninstall -y tensorflow numpy && pip install tensorflow-gpu==1.14.0 tf_sentencepiece
# + id="xRYbl_We431V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="5f755702-9233-4bbd-8c54-e12d7b0f9a94"
# !curl -o one_to_many.zip https://dl.dropboxusercontent.com/s/bssvce8t00zjwxj/one_to_many.zip?dl=0
# + id="b1kWSRVb5F-U" colab_type="code" colab={}
# !unzip -q one_to_many.zip
# + id="UA7Nd3qo5KM-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="fc7d8ae5-6e15-4377-dfeb-baed5fb6a77f"
# !curl -o mlt.zip https://codeload.github.com/suyash/mlt/zip/master
# + id="vg2sGM4N5Ln9" colab_type="code" colab={}
# !unzip -q mlt.zip && mv mlt-master/mlt ./
# + [markdown] id="SuEV4CUW5ROy" colab_type="text"
# ## One to Many Translation
# + id="Z1oy_H3o5MvI" colab_type="code" colab={}
import numpy as np
import tensorflow as tf
import tf_sentencepiece as tfs
from mlt.evaluation import predict
from mlt.layers import Attention, ConditionalNormalization, Gelu, MultiplyConstant, PaddingAndLookaheadMask, PaddingMask, PositionalEncoding
# + id="NXVjFAP55Y9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6216fe02-6d49-4e1b-84db-c258ba07fd63"
tf.__version__
# + id="1UrtqoA85aLG" colab_type="code" colab={}
sess = tf.InteractiveSession()
# + id="JAfwOm9x5baX" colab_type="code" colab={}
en_model_file = "final/sentencepiece/para_crawl/ende_plain_text/models/unigram/8192/a.model"
de_model_file = "final/sentencepiece/para_crawl/ende_plain_text/models/unigram/8192/b.model"
fr_model_file = "final/sentencepiece/para_crawl/enfr_plain_text/models/unigram/8192/b.model"
es_model_file = "final/sentencepiece/para_crawl/enes_plain_text/models/unigram/8192/b.model"
it_model_file = "final/sentencepiece/para_crawl/enit_plain_text/models/unigram/8192/b.model"
# + id="tvxp2Swp5d2M" colab_type="code" colab={}
en_offset = 0
fr_offset = 8192
de_offset = fr_offset + 8192
es_offset = de_offset + 8192
it_offset = es_offset + 8192
# + id="sbrg8m1U5fIW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="70786a53-ce1e-44aa-866b-9d7b6f33ea21"
with sess.as_default():
model = tf.keras.experimental.load_from_saved_model("final/model114", custom_objects={
"MultiplyConstant": MultiplyConstant,
"PositionalEncoding": PositionalEncoding,
"PaddingMask": PaddingMask,
"PaddingAndLookaheadMask": PaddingAndLookaheadMask,
"Attention": Attention,
"ConditionalNormalization": ConditionalNormalization,
"Gelu": Gelu,
})
# + [markdown] id="FgKgIm625jRP" colab_type="text"
# ### English to German
# + id="XH1XFVga5gZ5" colab_type="code" colab={}
a = tfs.encode(["This is a problem that we have to solve."], model_file=en_model_file, add_bos=True, add_eos=True)[0] + en_offset
# + id="cN0jhGry5odk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="9abaf8fb-707f-408e-dbdd-654e6fa71709"
ids, probs = predict(
model=model,
inputs=a,
inpf=tf.constant([1.0]),
tarf=tf.constant([1.0, 0.0, 0.0, 0.0]),
bos_id=de_offset + 1,
eos_id=de_offset + 2,
beam_size=5,
vocab_size=40960,
alpha=1.0,
)
# + id="8MsOxURV5qH6" colab_type="code" colab={}
mask = tf.cast(tf.not_equal(ids, 0), tf.int32)
seq_len = tf.reduce_sum(mask, axis=-1)
ids = ids + mask * -de_offset
probs = tf.math.exp(probs)
# + id="66oTPUsR5req" colab_type="code" colab={}
ids_, probs_, seq_len_ = sess.run([ids, probs, seq_len])
# + id="o883Aol85xNy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="40ad7b52-b497-41e8-f89d-fd1fec9b6cf1"
[x.decode("utf-8") for x in sess.run(tfs.decode(ids_[0], seq_len_[0], model_file=de_model_file))], probs_[0]
# + [markdown] id="UpaALHvr56x0" colab_type="text"
# ### English to French
# + id="QgjkgisG5yV1" colab_type="code" colab={}
a = tfs.encode(["This is a problem that we have to solve"], model_file=en_model_file, add_bos=True, add_eos=True)[0] + en_offset
# + id="tXYizgI06C_O" colab_type="code" colab={}
ids, probs = predict(
model=model,
inputs=a,
inpf=tf.constant([1.0]),
tarf=tf.constant([0.0, 1.0, 0.0, 0.0]),
bos_id=fr_offset + 1,
eos_id=fr_offset + 2,
beam_size=5,
vocab_size=40960,
alpha=1.0,
)
# + id="en6IYi_M6D5-" colab_type="code" colab={}
mask = tf.cast(tf.not_equal(ids, 0), tf.int32)
seq_len = tf.reduce_sum(mask, axis=-1)
ids = ids + mask * -fr_offset
probs = tf.math.exp(probs)
# + id="nmlqX_lw6FI6" colab_type="code" colab={}
ids_, probs_, seq_len_ = sess.run([ids, probs, seq_len])
# + id="c1dl-hAE6GLG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="eaeeb08d-d7fd-4fe3-8a7d-2ff91fcbf406"
[x.decode("utf-8") for x in sess.run(tfs.decode(ids_[0], seq_len_[0], model_file=fr_model_file))], probs_[0]
# + [markdown] id="Gt3aez4k6Lzv" colab_type="text"
# ### English to Spanish
# + id="TDy9PKeP6HbX" colab_type="code" colab={}
a = tfs.encode(["This is a problem that we have to solve"], model_file=en_model_file, add_bos=True, add_eos=True)[0] + en_offset
# + id="ULV7SQu76S7f" colab_type="code" colab={}
ids, probs = predict(
model=model,
inputs=a,
inpf=tf.constant([1.0]),
tarf=tf.constant([0.0, 0.0, 1.0, 0.0]),
bos_id=es_offset + 1,
eos_id=es_offset + 2,
beam_size=5,
vocab_size=40960,
alpha=1.0,
)
# + id="yBceuuwB6Uaz" colab_type="code" colab={}
mask = tf.cast(tf.not_equal(ids, 0), tf.int32)
seq_len = tf.reduce_sum(mask, axis=-1)
ids = ids + mask * -es_offset
probs = tf.math.exp(probs)
# + id="Waq32kLs6Vyy" colab_type="code" colab={}
ids_, probs_, seq_len_ = sess.run([ids, probs, seq_len])
# + id="DXXxpcLk6XBL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="b83d028b-b818-4aca-fbe9-3a80cee03b71"
[x.decode("utf-8") for x in sess.run(tfs.decode(ids_[0], seq_len_[0], model_file=es_model_file))], probs_[0]
# + [markdown] id="LgWiupN-6b-N" colab_type="text"
# ### English to Italian
# + id="FWkLJP9Y6YX_" colab_type="code" colab={}
a = tfs.encode(["This is a problem that we have to solve"], model_file=en_model_file, add_bos=True, add_eos=True)[0] + en_offset
# + id="k33TP3wZ6e7v" colab_type="code" colab={}
ids, probs = predict(
model=model,
inputs=a,
inpf=tf.constant([1.0]),
tarf=tf.constant([0.0, 0.0, 0.0, 1.0]),
bos_id=it_offset + 1,
eos_id=it_offset + 2,
beam_size=5,
vocab_size=40960,
alpha=1.0,
)
# + id="F2eNZu0K6f9Y" colab_type="code" colab={}
mask = tf.cast(tf.not_equal(ids, 0), tf.int32)
seq_len = tf.reduce_sum(mask, axis=-1)
ids = ids + mask * -it_offset
probs = tf.math.exp(probs)
# + id="_YeRSjk36hL0" colab_type="code" colab={}
ids_, probs_, seq_len_ = sess.run([ids, probs, seq_len])
# + id="2Armf6bX6icz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="9ba96879-e3f4-47a1-f47b-9316f19e3f45"
[x.decode("utf-8") for x in sess.run(tfs.decode(ids_[0], seq_len_[0], model_file=it_model_file))], probs_[0]
# + id="2WKbUxHh6jlB" colab_type="code" colab={}
sess.close()
| one_to_many_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "3"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. Meet Professor <NAME>
# <p>An investment may make sense if we expect it to return more money than it costs. But returns are only part of the story because they are risky - there may be a range of possible outcomes. How does one compare different investments that may deliver similar results on average, but exhibit different levels of risks?</p>
# <p><img style="float: left ; margin: 5px 20px 5px 1px;" width="200" src="https://s3.amazonaws.com/assets.datacamp.com/production/project_66/img/sharpe.jpeg"></p>
# <p>Enter <NAME>. He introduced the <a href="https://web.stanford.edu/~wfsharpe/art/sr/sr.htm"><em>reward-to-variability ratio</em></a> in 1966 that soon came to be called the Sharpe Ratio. It compares the expected returns for two investment opportunities and calculates the additional return per unit of risk an investor could obtain by choosing one over the other. In particular, it looks at the difference in returns for two investments and compares the average difference to the standard deviation (as a measure of risk) of this difference. A higher Sharpe ratio means that the reward will be higher for a given amount of risk. It is common to compare a specific opportunity against a benchmark that represents an entire category of investments.</p>
# <p>The Sharpe ratio has been one of the most popular risk/return measures in finance, not least because it's so simple to use. It also helped that Professor Sharpe won a Nobel Memorial Prize in Economics in 1990 for his work on the capital asset pricing model (CAPM).</p>
# <p>Let's learn about the Sharpe ratio by calculating it for the stocks of the two tech giants Facebook and Amazon. As a benchmark, we'll use the S&P 500 that measures the performance of the 500 largest stocks in the US.</p>
# + dc={"key": "3"} tags=["sample_code"]
# Importing required modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Settings to produce nice plots in a Jupyter notebook
plt.style.use('fivethirtyeight')
# %matplotlib inline
# Reading in the data
stock_data = pd.read_csv('datasets/stock_data.csv', parse_dates=['Date'], index_col=['Date']).dropna()
benchmark_data = pd.read_csv('datasets/benchmark_data.csv', parse_dates=['Date'], index_col=['Date']).dropna()
# + dc={"key": "11"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. A first glance at the data
# <p>Let's take a look the data to find out how many observations and variables we have at our disposal.</p>
# + dc={"key": "11"} tags=["sample_code"]
# Display summary for stock_data
print('Stocks\n')
stock_data.info()
print(stock_data.head())
# Display summary for benchmark_data
print('\nBenchmarks\n')
benchmark_data.info()
print(benchmark_data.head())
# + dc={"key": "18"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Plot & summarize daily prices for Amazon and Facebook
# <p>Before we compare an investment in either Facebook or Amazon with the index of the 500 largest companies in the US, let's visualize the data, so we better understand what we're dealing with.</p>
# + dc={"key": "18"} tags=["sample_code"]
# visualize the stock_data
stock_data.plot(title= 'Stock Data', subplots=True)
# summarize the stock_data
stock_data.describe()
# + dc={"key": "25"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Visualize & summarize daily values for the S&P 500
# <p>Let's also take a closer look at the value of the S&P 500, our benchmark.</p>
# + dc={"key": "25"} tags=["sample_code"]
# plot the benchmark_data
benchmark_data.plot(title='S&P 500')
# summarize the benchmark_data
benchmark_data.describe()
# + dc={"key": "32"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. The inputs for the Sharpe Ratio: Starting with Daily Stock Returns
# <p>The Sharpe Ratio uses the difference in returns between the two investment opportunities under consideration.</p>
# <p>However, our data show the historical value of each investment, not the return. To calculate the return, we need to calculate the percentage change in value from one day to the next. We'll also take a look at the summary statistics because these will become our inputs as we calculate the Sharpe Ratio. Can you already guess the result?</p>
# + dc={"key": "32"} tags=["sample_code"]
# calculate daily stock_data returns
stock_returns = stock_data.pct_change()
# plot the daily returns
stock_returns.plot()
# summarize the daily returns
stock_returns.describe()
# + dc={"key": "39"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. Daily S&P 500 returns
# <p>For the S&P 500, calculating daily returns works just the same way, we just need to make sure we select it as a <code>Series</code> using single brackets <code>[]</code> and not as a <code>DataFrame</code> to facilitate the calculations in the next step.</p>
# + dc={"key": "39"} tags=["sample_code"]
# calculate daily benchmark_data returns
sp_returns = benchmark_data['S&P 500'].pct_change()
# plot the daily returns
sp_returns.plot()
# summarize the daily returns
sp_returns.describe()
# + dc={"key": "46"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. Calculating Excess Returns for Amazon and Facebook vs. S&P 500
# <p>Next, we need to calculate the relative performance of stocks vs. the S&P 500 benchmark. This is calculated as the difference in returns between <code>stock_returns</code> and <code>sp_returns</code> for each day.</p>
# + dc={"key": "46"} tags=["sample_code"]
# calculate the difference in daily returns
excess_returns = stock_returns.sub(sp_returns, axis=0)
# plot the excess_returns
excess_returns.plot()
# summarize the excess_returns
excess_returns.describe()
# + dc={"key": "53"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. The Sharpe Ratio, Step 1: The Average Difference in Daily Returns Stocks vs S&P 500
# <p>Now we can finally start computing the Sharpe Ratio. First we need to calculate the average of the <code>excess_returns</code>. This tells us how much more or less the investment yields per day compared to the benchmark.</p>
# + dc={"key": "53"} tags=["sample_code"]
# calculate the mean of excess_returns
avg_excess_return = excess_returns.mean()
# plot avg_excess_returns
avg_excess_return.plot.bar(title='Mean of the Return Difference')
# + dc={"key": "60"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. The Sharpe Ratio, Step 2: Standard Deviation of the Return Difference
# <p>It looks like there was quite a bit of a difference between average daily returns for Amazon and Facebook.</p>
# <p>Next, we calculate the standard deviation of the <code>excess_returns</code>. This shows us the amount of risk an investment in the stocks implies as compared to an investment in the S&P 500.</p>
# + dc={"key": "60"} tags=["sample_code"]
# calculate the standard deviations
sd_excess_return = excess_returns.std()
# plot the standard deviations
sd_excess_return.plot.bar(title='Standard Deviation of the Return Difference')
# + dc={"key": "67"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 10. Putting it all together
# <p>Now we just need to compute the ratio of <code>avg_excess_returns</code> and <code>sd_excess_returns</code>. The result is now finally the <em>Sharpe ratio</em> and indicates how much more (or less) return the investment opportunity under consideration yields per unit of risk.</p>
# <p>The Sharpe Ratio is often <em>annualized</em> by multiplying it by the square root of the number of periods. We have used daily data as input, so we'll use the square root of the number of trading days (5 days, 52 weeks, minus a few holidays): √252</p>
# + dc={"key": "67"} tags=["sample_code"]
# calculate the daily sharpe ratio
daily_sharpe_ratio = avg_excess_return.div(sd_excess_return)
# annualize the sharpe ratio
annual_factor = np.sqrt(252)
annual_sharpe_ratio = daily_sharpe_ratio.mul(annual_factor)
# plot the annualized sharpe ratio
annual_sharpe_ratio.plot.bar(title='Annualized Sharpe Ration: Stocks vs S&P 500')
# + dc={"key": "74"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 11. Conclusion
# <p>Given the two Sharpe ratios, which investment should we go for? In 2016, Amazon had a Sharpe ratio twice as high as Facebook. This means that an investment in Amazon returned twice as much compared to the S&P 500 for each unit of risk an investor would have assumed. In other words, in risk-adjusted terms, the investment in Amazon would have been more attractive.</p>
# <p>This difference was mostly driven by differences in return rather than risk between Amazon and Facebook. The risk of choosing Amazon over FB (as measured by the standard deviation) was only slightly higher so that the higher Sharpe ratio for Amazon ends up higher mainly due to the higher average daily returns for Amazon. </p>
# <p>When faced with investment alternatives that offer both different returns and risks, the Sharpe Ratio helps to make a decision by adjusting the returns by the differences in risk and allows an investor to compare investment opportunities on equal terms, that is, on an 'apples-to-apples' basis.</p>
# + dc={"key": "74"} tags=["sample_code"]
# Uncomment your choice.
buy_amazon = True
buy_facebook = False
| notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alastra32/DS-Unit-2-Applied-Modeling/blob/master/module4/assignment_applied_modeling_4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nCc3XZEyG3XV" colab_type="text"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Applied Modeling, Module 4
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] Continue to iterate on your project: data cleaning, exploratory visualization, feature engineering, modeling.
# - [ ] Make a Shapley force plot to explain at least 1 individual prediction.
# - [ ] Share at least 1 visualization on Slack.
#
# (If you haven't completed an initial model yet for your portfolio project, then do today's assignment using your Tanzania Waterpumps model.)
#
# ## Stretch Goals
# - [ ] Make Shapley force plots to explain at least 4 individual predictions.
# - If your project is Binary Classification, you can do a True Positive, True Negative, False Positive, False Negative.
# - If your project is Regression, you can do a high prediction with low error, a low prediction with low error, a high prediction with high error, and a low prediction with high error.
# - [ ] Use Shapley values to display verbal explanations of individual predictions.
# - [ ] Use the SHAP library for other visualization types.
#
# The [SHAP repo](https://github.com/slundberg/shap) has examples for many visualization types, including:
#
# - Force Plot, individual predictions
# - Force Plot, multiple predictions
# - Dependence Plot
# - Summary Plot
# - Summary Plot, Bar
# - Interaction Values
# - Decision Plots
#
# We just did the first type during the lesson. The [Kaggle microcourse](https://www.kaggle.com/dansbecker/advanced-uses-of-shap-values) shows two more. Experiment and see what you can learn!
#
#
# ## Links
# - [Kaggle / <NAME>: Machine Learning Explainability — SHAP Values](https://www.kaggle.com/learn/machine-learning-explainability)
# - [<NAME>: Interpretable Machine Learning — Shapley Values](https://christophm.github.io/interpretable-ml-book/shapley.html)
# - [SHAP repo](https://github.com/slundberg/shap) & [docs](https://shap.readthedocs.io/en/latest/)
# + [markdown] id="x5ZcA_ZL-8JG" colab_type="text"
# ## Setup
# + id="DM6fbgb4jSec" colab_type="code" outputId="f1533b65-1e75-4461-b54c-2a6d117c7925" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# If you're in Colab...
import os, sys
in_colab = 'google.colab' in sys.modules
if in_colab:
# Install required python packages:
# category_encoders, version >= 2.0
# pandas-profiling, version >= 2.0
# plotly, version >= 4.0
# !pip install --upgrade category_encoders pandas-profiling plotly
# + id="o50qKqSIiafP" colab_type="code" colab={}
import pandas as pd
from sklearn.model_selection import train_test_split
# merge train_features.csv & train_labels.csv
trainandval = pd.merge(pd.read_csv('https://raw.githubusercontent.com/alastra32/DS-Unit-2-Kaggle-Challenge/master/data/tanzania/train_features.csv'),
pd.read_csv('https://raw.githubusercontent.com/alastra32/DS-Unit-2-Kaggle-Challenge/master/data/tanzania/train_labels.csv'))
# read test_features.csv & sample_submission.csv
test = pd.read_csv('https://raw.githubusercontent.com/alastra32/DS-Unit-2-Kaggle-Challenge/master/data/tanzania/test_features.csv')
sample_submission = pd.read_csv('https://raw.githubusercontent.com/alastra32/DS-Unit-2-Kaggle-Challenge/master/data/tanzania/sample_submission.csv')
# + id="CWZ9qhvHiafU" colab_type="code" colab={}
# import block
pd.set_option('display.max_columns', None)
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score
import category_encoders as ce
from xgboost import XGBClassifier
# + id="9N1--TLUiafY" colab_type="code" outputId="4b88e403-09b5-4ba0-965a-a331d247c07c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# train validation split
train, val = train_test_split(trainandval, train_size=0.95, test_size=0.05,
stratify=trainandval['status_group'], random_state=42)
train.shape, val.shape, test.shape
# + [markdown] id="vyKNJx_B_FIP" colab_type="text"
# ## Manual Mode
#
#
# + id="PRXUtcKx4Dn0" colab_type="code" colab={}
# We need a function that returns the mode of a given series for the imputer function.
def manual_mode(feature):
try:
return feature.mode()[0]
except:
pass
# + [markdown] id="ukWfLSXF_Kau" colab_type="text"
# ## Imputer
# + colab_type="code" id="oFwcq_XJ5Db8" colab={}
# imputes by the lowest non-null region measure
def fill_nulls(df, feature, method):
#attempt to fill nulls by method in succesively larger geographic scopes
df = df.copy()# avoid settingwithcopy warning
geo_scopes = ['ward', 'lga', 'region', 'basin']
if method == 'mode':
method = manual_mode
for scope in geo_scopes:
if df[feature].isnull().sum() == 0:
break
df[feature] = df[feature].fillna(df.groupby(scope)[feature].transform(method))
return df[feature]
def impute(df, features, method):
#imputation of given features by given method (mean/median/mode)
df = df.copy()
for feature in features:
df[feature] = fill_nulls(df, feature, method)
return df
# + [markdown] id="GHdYiB3w_Uyw" colab_type="text"
# ## Wrangler
# + colab_type="code" id="todzeNu6Edkq" colab={}
def flag_missing_values(df):
'''add "<FEATURE>_MISSING" flag feature for all columns with nulls'''
df.copy()
columns_with_nulls = df.columns[df.isna().any()]
for col in columns_with_nulls:
df[col+'_MISSING'] = df[col].isna()
return df
def convert_dummy_nulls(df):
'''Convert 0 to NaN's'''
df = df.copy()
# replace near-zero latitudes with zero
df['latitude'] = df['latitude'].replace(-2e-08, 0)
zero_columns = ['longitude', 'latitude', 'construction_year', 'gps_height',
'population']
for col in zero_columns:
df[col] = df[col].replace(0, np.nan)
return df
def clean_text_columns(df):
'''convert text to lowercase, remove non-alphanumerics, unknowns to NaN'''
df = df.copy()
text_columns = df[df.columns[(df.applymap(type) == str).all(0)]]
unknowns = ['unknown', 'notknown', 'none', 'nan', '']
for col in text_columns:
df[col] = df[col].str.lower().str.replace('\W', '')
df[col] = df[col].replace(unknowns, np.nan)
return df
def get_distances_to_population_centers(df):
'''create a distance feature for population centers'''
df = df.copy()
population_centers = {'dar': (6.7924, 39.2083),
'mwanza': (2.5164, 32.9175),
'dodoma': (6.1630, 35.7516)}
for city, loc in population_centers.items():
df[city+'_distance'] = ((((df['latitude']-loc[0])**2)
+ ((df['longitude']-loc[1])**2))**0.5)
return df
def engineer_date_features(df):
df = df.copy()
# change date_recorded to datetime format
df['date_recorded'] = pd.to_datetime(df.date_recorded,
infer_datetime_format=True)
# extract components from date_recorded
df['year_recorded'] = df['date_recorded'].dt.year
df['month_recorded'] = df['date_recorded'].dt.month
df['day_recorded'] = df['date_recorded'].dt.day
df['inspection_interval'] = df['year_recorded'] - df['construction_year']
return df
def wrangle(df):
'''cleaning/engineering function'''
df = df.copy()
df = convert_dummy_nulls(df)
df = clean_text_columns(df)
df = get_distances_to_population_centers(df)
df = engineer_date_features(df)
df = flag_missing_values(df)
drop_features = ['recorded_by', 'id', 'date_recorded']
df = df.drop(columns=drop_features)
# Apply imputation
numeric_columns = df.select_dtypes(include = 'number').columns
nonnumeric_columns = df.select_dtypes(exclude = 'number').columns
df = impute(df, numeric_columns, 'median')
df = impute(df, nonnumeric_columns, 'mode')
return df
# + [markdown] id="8fdKICh-_kpF" colab_type="text"
# ## Engineer, Pipe, and Train
# + id="SGxbu3mRiafw" colab_type="code" colab={}
# clean and engineer all datasets
train_wrangled = wrangle(train)
val_wrangled = wrangle(val)
test_wrangled = wrangle(test)
# + colab_type="code" id="eF-B9S3HAkNy" colab={}
# arrange data into X features matrix and y target vector
target = 'status_group'
X_train = train_wrangled.drop(columns=target)
y_train = train_wrangled[target]
X_val = val_wrangled.drop(columns=target)
y_val = val_wrangled[target]
X_test = test_wrangled
# + id="_f7uhECciaf6" colab_type="code" outputId="1ac27d09-e154-45e5-ea0e-4760d1acbcf5" colab={"base_uri": "https://localhost:8080/", "height": 141}
# Use Ordinal Encoder, outside of a pipeline
encoder = ce.OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded= encoder.fit_transform(X_val)
model = RandomForestClassifier(n_estimators=129, max_depth=29, min_samples_leaf=2,
random_state=42, min_impurity_decrease=2.22037e-16, n_jobs=-1)
model.fit(X_train_encoded, y_train)
# + id="uBcCbN7Biaf-" colab_type="code" outputId="f0a80d7b-21dd-453e-e5e3-2f38687a7058" colab={"base_uri": "https://localhost:8080/", "height": 34}
#score
model.score(X_val_encoded,y_val)
# + id="SUzd7xBab1lY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 97} outputId="fd5c1656-967a-4980-bb67-13895b3b52bc"
row = X_test.iloc[[3232]]
row
# + id="NAWl2rI64zNQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 585} outputId="d3dd2cc6-a67b-473e-c161-969e045b6c6b"
# !pip install shap
# + id="CDS7q4ov3w7v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 213} outputId="95ceb4d6-4916-46cc-c5ac-3caae68dc6b1"
import shap
# processor = pipeline[:-1]
explainer = shap.TreeExplainer(model)
row_process = encoder.transform(row)
shap_values = explainer.shap_values(row_process)
shap.initjs()
shap.force_plot(
base_value=explainer.expected_value[0],
shap_values=shap_values[0],
features=row
)
# + id="PiKT7Bm74GKw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c13ee0a1-aca4-432b-be20-f59b1bcaa419"
feature_names = row.columns
feature_values = row.values[0]
shaps = pd.Series(shap_values[0][0], zip(feature_names,feature_values))
shaps.sort_values().plot.barh(color='grey', figsize=(15,20));
# + id="cbfkbude4GY7" colab_type="code" colab={}
| module4/assignment_applied_modeling_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IPython Magic Commands
# The previous two sections showed how IPython lets you use and explore Python efficiently and interactively.
# Here we'll begin discussing some of the enhancements that IPython adds on top of the normal Python syntax.
# These are known in IPython as *magic commands*, and are prefixed by the ``%`` character.
# These magic commands are designed to succinctly solve various common problems in standard data analysis.
# Magic commands come in two flavors: *line magics*, which are denoted by a single ``%`` prefix and operate on a single line of input, and *cell magics*, which are denoted by a double ``%%`` prefix and operate on multiple lines of input.
# We'll demonstrate and discuss a few brief examples here, and come back to more focused discussion of several useful magic commands later in the chapter.
# ## Pasting Code Blocks: ``%paste`` and ``%cpaste``
#
# When working in the IPython interpreter, one common gotcha is that pasting multi-line code blocks can lead to unexpected errors, especially when indentation and interpreter markers are involved.
# A common case is that you find some example code on a website and want to paste it into your interpreter.
# Consider the following simple function:
#
# ``` python
# >>> def donothing(x):
# ... return x
#
# ```
# The code is formatted as it would appear in the Python interpreter, and if you copy and paste this directly into IPython you get an error:
#
# ```ipython
# In [2]: >>> def donothing(x):
# ...: ... return x
# ...:
# File "<ipython-input-20-5a66c8964687>", line 2
# ... return x
# ^
# SyntaxError: invalid syntax
# ```
#
# In the direct paste, the interpreter is confused by the additional prompt characters.
# But never fear–IPython's ``%paste`` magic function is designed to handle this exact type of multi-line, marked-up input:
#
# ```ipython
# In [3]: %paste
# >>> def donothing(x):
# ... return x
#
# ## -- End pasted text --
# ```
#
# The ``%paste`` command both enters and executes the code, so now the function is ready to be used:
#
# ```ipython
# In [4]: donothing(10)
# Out[4]: 10
# ```
#
# A command with a similar intent is ``%cpaste``, which opens up an interactive multiline prompt in which you can paste one or more chunks of code to be executed in a batch:
#
# ```ipython
# In [5]: %cpaste
# Pasting code; enter '--' alone on the line to stop or use Ctrl-D.
# :>>> def donothing(x):
# :... return x
# :--
# ```
#
# These magic commands, like others we'll see, make available functionality that would be difficult or impossible in a standard Python interpreter.
# ## Running External Code: ``%run``
# As you begin developing more extensive code, you will likely find yourself working in both IPython for interactive exploration, as well as a text editor to store code that you want to reuse.
# Rather than running this code in a new window, it can be convenient to run it within your IPython session.
# This can be done with the ``%run`` magic.
#
# For example, imagine you've created a ``myscript.py`` file with the following contents:
#
# ```python
# #-------------------------------------
# # file: myscript.py
#
# def square(x):
# """square a number"""
# return x ** 2
#
# for N in range(1, 4):
# print(N, "squared is", square(N))
# ```
#
# You can execute this from your IPython session as follows:
#
# ```ipython
# In [6]: %run myscript.py
# 1 squared is 1
# 2 squared is 4
# 3 squared is 9
# ```
#
# Note also that after you've run this script, any functions defined within it are available for use in your IPython session:
#
# ```ipython
# In [7]: square(5)
# Out[7]: 25
# ```
#
# There are several options to fine-tune how your code is run; you can see the documentation in the normal way, by typing **``%run?``** in the IPython interpreter.
# ## Timing Code Execution: ``%timeit``
# Another example of a useful magic function is ``%timeit``, which will automatically determine the execution time of the single-line Python statement that follows it.
# For example, we may want to check the performance of a list comprehension:
#
# ```ipython
# In [8]: %timeit L = [n ** 2 for n in range(1000)]
# 1000 loops, best of 3: 325 µs per loop
# ```
#
# The benefit of ``%timeit`` is that for short commands it will automatically perform multiple runs in order to attain more robust results.
# For multi line statements, adding a second ``%`` sign will turn this into a cell magic that can handle multiple lines of input.
# For example, here's the equivalent construction with a ``for``-loop:
#
# ```ipython
# In [9]: %%timeit
# ...: L = []
# ...: for n in range(1000):
# ...: L.append(n ** 2)
# ...:
# 1000 loops, best of 3: 373 µs per loop
# ```
#
# We can immediately see that list comprehensions are about 10% faster than the equivalent ``for``-loop construction in this case.
# We'll explore ``%timeit`` and other approaches to timing and profiling code in [Profiling and Timing Code](01.07-Timing-and-Profiling.ipynb).
# ## Help on Magic Functions: ``?``, ``%magic``, and ``%lsmagic``
#
# Like normal Python functions, IPython magic functions have docstrings, and this useful
# documentation can be accessed in the standard manner.
# So, for example, to read the documentation of the ``%timeit`` magic simply type this:
#
# ```ipython
# In [10]: %timeit?
# ```
#
# Documentation for other functions can be accessed similarly.
# To access a general description of available magic functions, including some examples, you can type this:
#
# ```ipython
# In [11]: %magic
# ```
#
# For a quick and simple list of all available magic functions, type this:
#
# ```ipython
# In [12]: %lsmagic
# ```
#
# Finally, I'll mention that it is quite straightforward to define your own magic functions if you wish.
# We won't discuss it here, but if you are interested, see the references listed in [More IPython Resources](01.08-More-IPython-Resources.ipynb).
| notebooks/Python-in-2-days/D1_L2_IPython/04-Magic-Commands.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas
import numpy as np
import pandas as pd
labels = ["a","b","c"]
mydata = [10,20,30]
arr = np.array(mydata)
d = {"a":10,"b":20,"c":30}
pd.Series(data = mydata)
pd.Series(data = mydata,index = labels)
pd.Series(mydata,labels)
pd.Series(arr,labels)
pd.Series(d)
pd.Series(data=[sum,print])
ser1 = pd.Series([1,2,3],["USA","GERMANY","ALGERIA"])
ser1
ser2 = pd.Series([4,5,6],["USA","GERMANY","ALGERIA"])
ser2
ser1["USA"]
ser3 = pd.Series(data= labels)
ser3
ser3[0]
ser1
ser2
ser1 + ser2
| Pandas/.ipynb_checkpoints/Panda_Series-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# More process building tools!
# ----------------------------
# Use multiple feature extractors (on the same data), concatenate results.
from sklearn.pipeline import make_union, make_pipeline
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.svm import LinearSVC
from sklearn.grid_search import GridSearchCV
import numpy as np
from sklearn.datasets import fetch_20newsgroups
news = fetch_20newsgroups()
data, y = news.data, news.target
np.bincount(y)
from sklearn.cross_validation import train_test_split
data_train, data_test, y_train, y_test = train_test_split(data, y)
# +
char_and_word = make_union(CountVectorizer(analyzer="char"),
CountVectorizer(analyzer="word"))
text_pipe = make_pipeline(char_and_word, LinearSVC(dual=False))
param_grid = {'linearsvc__C': 10. ** np.arange(-3, 3)}
grid = GridSearchCV(text_pipe, param_grid=param_grid, cv=5, verbose=10)
# -
grid.fit(data_train, y_train)
param_grid = {'featureunion__countvectorizer-1__ngram_range': [(1, 3), (1, 5), (2, 5)],
'featureunion__countvectorizer-2__ngram_range': [(1, 1), (1, 2), (2, 2)],
'linearsvc__C': 10. ** np.arange(-3, 3)}
| 07 - Feature Union.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
# # Python 연산과 Numpy 연산 차이점
t = [1, 2, 3]
# ## 1. Python 연산
# numpy가 아니라 list라서 아래는 오류 난다.
t + 5
t * 5 # 파이썬의 * 연산은 반복된다
# ## 2.Numpy 연산
np.int32(t) + 5
np.int32(t) * 5
# ### a. Numpy Array
a = np.arange(10)
a
a + 3 # Broadcasting 지원
a + a
a[[2, 4, 1]] # index array
# ### b. Slicing
a[:3]
# ## 3. Fancy Indexing
b = a.reshape(2, 5)
b
b[-1, -1] # fancy indexing 마지막 행, 마지막열
b[[1,0], -1]
b[:, -1]
b[:, [2, -1, 0]]
# # TensorFlow 와 Numpy reshape 차이점
a= np.arange(11, 19).reshape(-1, 1, 1, 2)
# +
# 앞에 -1을 넣은것은 전체 갯수를 차원으로 나눈것
# -
a
| _pages/AI/TensorFlow/src/NCIA-CNN/Day_01_02_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def enc_keyless2(message): # ROW TRANSPOSITION encryption in which we encrytp the plain by transpose of matrix
text=""
i=0
while(i<len(message)):
if(message[i]!=' '):
text=text+message[i]
i=i+1
result=""
i=0
while((i*i)<len(text)):
i=i+1
j=0
count=0
while(j<i and count<len(text)):
k=0
l=j
while(k<i and count<len(text)):
result=result+text[l]
count=count+1
k=k+1
l=l+4
j=j+1
return result
# result contains the cipher text
def enc_keyless1(message): # RAIL FENCE keyless transoposition for encryption where we simple use two rows and generate the cipher text
text=""
i=0
while(i<len(message)):
if(message[i]!=' '):
text=text+message[i]
i=i+1
result=""
i=0
while(i<len(text)):
result=result+text[i]
i=i+2 #appending even text
i=1
while(i<len(text)):
result=result+text[i]
i=i+2 #appending odd text
return result
#result has the cipher text which is made by rail fence
def dec_keyless1(cipher,a,b): # RAIL FENCE keyless decryption using the cipher text we got, a and b contains odd and even characters
i=len(a)
j=len(b)
result=""
print("decryption of rail fence cipehr text is \t\t",end="") #here we print the decypted text
if i>j:
for k in range(i-1):
print(a[k],end="")
print(b[k],end="")
print(a[k+1])
else:
for k in range(j):
print(a[k],end="")
print(b[k],end="")
def dec_keyless2(final): # ROW TRANSPOSITION decryption in which we decrypt the cipher by reverse transpose of matrix
lis1=[]
k=-1
count=0
while(k<4 and count<len(final)):
k=k+1
i=k
while(i<len(final) and count<len(final)):
lis1.append(final[i])
i=i+4
count=count+1
str1=''.join([str(elem) for elem in lis1])
return str1
# str1 contains the decrypted text
# +
def main():
message=input("Enter the message\n")
message=message.upper()
cipher1=enc_keyless1(message) # encrypt the RAIL FENCE transpositon plain text
list1=[]
list2=[]
for i in range(len(message)):
if i%2==0:
list1.append(message[i]) #list1 contains even length
else:
list2.append(message[i]) #list2 contains odd length
print("rail fence cipher text is\t\t\t\t"+cipher1) #print the encrypted text
dec_keyless1(cipher1,list1,list2) #decrypt the RAIL FENCE cipher text
cipher2=enc_keyless2(message) #encrypt the ROW TRANSPOSITION plain text
print("matrix transposition cipher text is\t\t\t"+cipher2) #print the encrypted text
count=0
final=[]
s=[]
k=-1
for i in message:
s.append(i)
while(k<4 and count<len(message)):
k=k+1
i=k
while(i<len(message) and count<len(message)):
final.append(s[i])
i=i+4
count=count+1
plain2=dec_keyless2(final) #decrypt the ROW TRANSPOSITION cipher text
print("decryption of matrix transposition cipher is\t\t"+message) #print the decrypted text
if __name__=='__main__':
main()
# -
#
| NTC Assignment/Ciphers/Transposition Cipher (Keyless)/B180441CS_BHUKYA_05a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
# http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html
from sklearn.decomposition import PCA
# http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html
# <h2>Random Data with PCA</h2>
# <h4>Demonstrate how PCA works with completely random dataset</h4>
# <br>
# Features: 10 random columns<br>
# Output: PCA Components<br>
#
# Objective: <br><quote>There should not be much reduction in dimensions when using all random colums</quote>
# 1000 rows x 10 columns
np.random.seed(5)
random_data = np.random.rand(1000,10)
random_data.shape
df = pd.DataFrame(random_data)
df.head()
df.corr()
df.dtypes
# +
# Test PCA
# +
# Two modes to test with PCA
# How many components we need in final output or how much variance do we need to capture as a percentage
pca = PCA(n_components=0.9) # percentage of variance to capture
#pca = PCA(n_components=2) # number of components
# -
pca.fit(df)
# number of components PCA came up with
pca.n_components_
df.head()
pca.transform(df)
def transform_with_pca(pca, df, columns):
transformed_data = pca.transform(df[columns])
tcols = []
for i in range(pca.n_components_):
tcols.append('component_' + str(i))
print ('components:',tcols)
df_transformed = pd.DataFrame(transformed_data, columns=tcols)
for col in df_transformed.columns:
df[col] = df_transformed[col]
df.drop(columns, inplace=True, axis=1)
return tcols
transform_with_pca(pca,df, df.columns)
df.head()
# ## Summary
#
# 1. With random datasets, not much reduction is possible
# 2. We are capturing 90% variance and PCA came up 9 components
# 3. In the next demo, let’s look a dataset that has related features.
#
| pca/ExplorePCA/random_data_preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Start from importing dataset loading module and numpy for some processing
from mnist import MNIST
import numpy as np
# initialize a dataset loader and use gzip files
mnist_data = MNIST(path='./samples/', gz=True)
train_data = mnist_data.load_training()
len(train_data)
# ### The training set contains 60,000 examples
len(train_data[0])
len(train_data[1])
# ### train_data[0] is the input arrays (images)
len(train_data[0][0])
len(train_data[1][0]) # Since train_data[1] is a list of integers, calling len() on integers causes an error
print(type(train_data[0]), type(train_data[1]))
# ### load_training() returns a tuple with two lists: [1] lists of values for data points (60000 lists, each with 784 entries), [2] a list of labels as integers (60000 integers)
X_train = np.array(train_data[0])
y_train = np.array(train_data[1]).reshape((-1, 1))
X_train.shape
y_train.shape
X_train.max()
X_train.min()
y_train.max()
y_train.min()
# ## Import matplotlib for visualization
import matplotlib.pyplot as plt
# %matplotlib inline
# # Let's pick 5 random images from the training set!
fig, axes = plt.subplots(nrows=1, ncols=5, figsize=(20, 20), dpi=100)
for n, ax in zip(np.random.permutation(len(X_train))[:5], axes):
ax.set_xticks([])
ax.set_yticks([])
ax.set_title("True Label : " + str(y_train[n]))
ax.imshow(X_train[n].reshape(28, 28), cmap="Greys")
# ## Now that we checked that the dataset is good, let's load test dataset as well!
X_test, y_test = mnist_data.load_testing()
X_test = np.array(X_test)
y_test = np.array(y_test).reshape(-1, 1)
# ## Import torch for neural networks
import torch
import torch.nn as nn
import torch.nn.functional as F
# ### Use CUDA device if available. Use CPU otherwise
device = "cuda" if torch.cuda.is_available() else "cpu"
# ### Input Normalization
# * currently this doesn't seem to provide any significant improvement in training, so it is defined but not used for now
# +
def normalize(input_tensor, dim=None):
if dim is None:
mean = input_tensor.data.mean()
std = input_tensor.data.std()
else:
mean = input_tensor.data.mean(dim=dim)
std = input_tensor.data.std(dim=dim)
input_tensor.data = input_tensor.data - mean # mean of 0
input_tensor.data = input_tensor.data / std.max(torch.tensor(1e-12, dtype=std.dtype, device=std.device)) # standard deviation of 1, prevent division by zero
def convert_and_normalize_input(X, dtype=None, device=None, dim=None):
X_minmaxed = torch.tensor(X / 255, dtype=dtype, device=device)
normalize(X_minmaxed, dim=dim)
X_normalized = X_minmaxed
return X_normalized
# -
# ## Do minmax scaling and transform into torch tensors
# X_train = convert_and_normalize_input(X_train, dtype=torch.float64, device=device)
# X_test = convert_and_normalize_input(X_test, dtype=torch.float64, device=device)
X_train = torch.tensor(X_train / 255, dtype=torch.float64, device=device)
X_test = torch.tensor(X_test / 255, dtype=torch.float64, device=device)
X_train
X_train.shape
X_test.shape
# ### For classification, we also need to turn the labels to one-hot vectors (vectors each having value 1 for corresponding class and 0 for everything else)
def labels_to_onehot(Y, device=None):
return torch.tensor(Y, dtype=torch.uint8, device=device).eq(torch.arange(0, 10, dtype=torch.uint8, device=device))
Y_train = labels_to_onehot(y_train, device=device)
Y_train.data = Y_train.data.type(torch.float64)
Y_test = labels_to_onehot(y_test, device=device)
Y_test.data = Y_test.data.type(torch.float64)
# ## For hyperparameter tuning in the future, the training set is randomly devided into 50,000 training examples and 10,000 validation examples
# Take out some examples from the training set as the validation set (50000 for training, 10000 for validation)
randidx = torch.randperm(len(X_train))
X_val = X_train[randidx[:10000]]
X_train = X_train[randidx[10000:]]
Y_val = Y_train[randidx[:10000]]
Y_train = Y_train[randidx[10000:]]
print(len(X_train))
print(len(X_val))
print(len(Y_train))
print(len(Y_val))
# ## Initialize weights and biases
# Two layers - layer1 : (input to hidden) -> layer2 : (hidden to output)
# +
epsilon = 0.01
w1 = torch.tensor(torch.rand(100, X_train.shape[1], dtype=torch.float64) * 2 * epsilon - epsilon, device=device, requires_grad=True)
b1 = torch.tensor(torch.rand(100, 1, dtype=torch.float64) * 2 * epsilon - epsilon, device=device, requires_grad=True)
w2 = torch.tensor(torch.rand(10, w1.shape[0], dtype=torch.float64) * 2 * epsilon - epsilon, device=device, requires_grad=True)
b2 = torch.tensor(torch.rand(10, 1, dtype=torch.float64) * 2 * epsilon - epsilon, device=device, requires_grad=True)
# -
w1
# ## Useful and necessary functions
# - re-initializing parameters (4 different methods are available : uniform, standard normal (std can be adjusted), He, and Xavier)
# - mini-batch sampling (Generater function that yields mini-batch samples every iteration)
# - feedforward
# - prediction (Turns raw probability values into prediction labels; integer values)
# - binary cross entropy loss
# - L2 regularization loss
# - loss function (cross entropy + L2 regularization)
# - accuracy evaluation
# +
def reset(tensors, method='uniform', e=0.01):
"""
re-initialize parameters using the given method
tensors : 'list' of tensors
method : {'uniform'|'std'|'he'|'xavier'}, default:'uniform'
e : maximum absolute value for 'uniform' | standard deviation for 'std' | ignored otherwise
"""
method = method.lower()
if method not in ('uniform', 'std', 'he', 'xavier'):
raise ValueError("method must be one of these options : {'uniform'|'std'|'he'|'xavier'}")
if method == 'uniform':
for tensor in tensors:
tensor.data = torch.tensor(torch.rand(tensor.size(), dtype=tensor.dtype, device=tensor.device) * 2 * e - e)
elif method == 'std':
for tensor in tensors:
tensor.data = torch.tensor(torch.randn(tensor.size(), dtype=tensor.dtype, device=tensor.device) * e)
elif method == 'he':
for tensor in tensors:
tensor.data = torch.tensor(torch.randn(tensor.size(), dtype=tensor.dtype, device=tensor.device) * torch.sqrt(torch.tensor(2 / tensor.size()[1], dtype=tensor.dtype, device=tensor.device)))
else:
for tensor in tensors:
tensor.data = torch.tensor(torch.randn(tensor.size(), dtype=tensor.dtype, device=tensor.device) * torch.sqrt(torch.tensor(2 / sum(tensor.size()), dtype=tensor.dtype, device=tensor.device)))
def mini_batch(X, Y=None, batch_size=100, shuffle=True):
assert Y is None or len(X) == len(Y), "length of the first dimensions in both matrices must match"
assert isinstance(batch_size, int), "the batch size must be an integer"
if batch_size > len(X) or batch_size <= 0:
batch_size = len(X)
n_batches = len(X) // batch_size
if shuffle is True:
rand_idx = torch.randperm(len(X))
if Y is None:
for batch_idx in range(n_batches):
yield X[rand_idx[batch_size * batch_idx:batch_size * (batch_idx + 1)]]
else:
for batch_idx in range(n_batches):
yield (X[rand_idx[batch_size * batch_idx:batch_size * (batch_idx + 1)]],
Y[rand_idx[batch_size * batch_idx:batch_size * (batch_idx + 1)]])
else:
if Y is None:
for batch_idx in range(n_batches):
yield X[batch_size * batch_idx:batch_size * (batch_idx + 1)]
else:
for batch_idx in range(n_batches):
yield (X[batch_size * batch_idx:batch_size * (batch_idx + 1)],
Y[batch_size * batch_idx:batch_size * (batch_idx + 1)])
def feedforward(input_data, weight_matrices, biase_matrices=None):
assert biase_matrices is None or len(weight_matrices) == len(biase_matrices)
output = input_data.clone()
for n, weights in enumerate(weight_matrices):
output = output.mm(weights.t())
if biase_matrices is not None:
output = output + biase_matrices[n].t()
output = torch.sigmoid(output)
return output
def predict(probs, one_hot=False):
preds = torch.tensor(probs.argmax(dim=1).reshape((-1, 1)), dtype=torch.uint8, device=probs.device)
if one_hot is True:
preds.data = preds.eq(torch.arange(0, 10, dtype=torch.uint8, device=probs.device))
return preds
def binary_class_cross_entropy_loss(pred, true, per_example=False):
assert len(pred) == len(true)
loss = torch.sum(-true * torch.log(pred) - (1 - true) * torch.log(1 - pred), dim=1, dtype=pred.dtype)
if per_example is False:
loss = loss.sum() / len(true)
if loss.item() != loss.item(): # if the result is nan (probably due to 0 value inside log functions)
loss = torch.sum(-true * torch.log(pred + 1e-12) - (1 - true) * torch.log(1 - pred + 1e-12), dtype=pred.dtype)
return loss
def regularization_loss(weight_matrices, regular_coef, n_examples):
loss = (regular_coef / (2 * n_examples)) * sum([weights.pow(2).sum() for weights in weight_matrices])
return loss
def loss_function(prob, true, weight_matrices, regular_coef, n_examples):
return binary_class_cross_entropy_loss(prob, true) + regularization_loss(weight_matrices, regular_coef, n_examples)
def accuracy_score(pred, true):
assert len(pred) == len(true)
pred = pred.cuda().argmax(dim=1)
true = true.cuda().argmax(dim=1)
accuracy = pred.eq(true).sum(dtype=pred.dtype).item() / float(len(true))
return accuracy
# -
# Bundle the weights and biases into two separate lists so that I can use them for my functions
weights = [w1, w2]
biases = [b1, b2]
# ### Here, re-initialize the parameters using He et al initialization
reset(weights + biases, method='he')
# ## Setting up logs
# These will help tracking the progress
# Set up logs
train_loss_history = []
train_acc_history = []
val_loss_history = []
val_acc_history = []
# ## Finding optimal learning rate
# From [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/abs/1506.01186)
# +
# Finding the best learning rate
def lr_find(X, Y, weight_matrices, bias_matrices, batch_size=100, start_lr=1e-4, end_lr=1.0, n_steps=100, reg_coef=0, reset_method='uniform', e=0.01):
assert start_lr <= end_lr, "'end_lr' must be greater than or equal to 'start_lr'"
assert isinstance(n_steps, int) and n_steps >= 0, "'n_steps' must be an integer value greater than or equal to 0"
# Use the clones so that we won't end up messing up with our original weights...
weight_matrices = [weight.clone() for weight in weight_matrices]
bias_matrices = [bias.clone() for bias in bias_matrices]
params = weight_matrices + bias_matrices
reset(params, method=reset_method, e=e)
# Since cloned tensors are "edges", they do not retain gradients by default.
# We have to manually set it
for param in params:
param.retain_grad()
learning_rate = start_lr
lr_history = []
acc_history = []
lr_diff = end_lr - start_lr
lr_step_change = lr_diff / n_steps
report_every_n_steps = n_steps // 10
step_count = 0
while step_count < n_steps:
for X_batch, Y_batch in mini_batch(X, Y, batch_size=batch_size):
output = feedforward(X, weight_matrices, bias_matrices)
loss = loss_function(output, Y, weight_matrices, reg_coef, len(X))
loss.backward()
for param in params:
param.data -= learning_rate * param.grad.data
param.grad.data.zero_()
lr_history.append(learning_rate)
acc_history.append(accuracy_score(predict(output, one_hot=True), Y))
learning_rate += lr_step_change
step_count += 1
if step_count % report_every_n_steps == 0:
print(f"{step_count} step(s) of {n_steps} ...")
if step_count >= n_steps:
break
return (lr_history, acc_history)
def lr_plot(lr_list, acc_list, log_scale=False):
if log_scale:
plt.xscale('log')
plt.locator_params(axis='y', nbins=10)
plt.xlabel('learning rate' + ('', '(in log scale)')[bool(log_scale)])
plt.ylabel('accuracy')
plt.grid()
plt.plot(lr_list, acc_list)
# -
logs = lr_find(X_train, Y_train, weights, biases, batch_size=-1, start_lr=1e-1, end_lr=3.0, n_steps=100, reg_coef=1, reset_method='he')
print(len(logs[0]))
# The entire plot
lr_plot(logs[0], logs[1])
# First quarter
lr_plot(logs[0][:len(logs[0])//4], logs[1][:len(logs[0])//4])
# Second quarter
lr_plot(logs[0][len(logs[0])//4:len(logs[0])//2], logs[1][len(logs[0])//4:len(logs[0])//2])
# Third quarter
lr_plot(logs[0][len(logs[0])//2:(len(logs[0])*3)//4], logs[1][len(logs[0])//2:(len(logs[0])*3)//4])
# Forth quarter
lr_plot(logs[0][(len(logs[0])*3)//4:], logs[1][(len(logs[0])*3)//4:])
# ## Let's now set hyperparameters and start training!
# Hyperparameters
epoch = 100
lambda_term = 1
learning_rate = 1.0
batch_size = -1 # the entire training set
# +
tol_loss = 1e-3
tol_step = 1e-8
reset_progress = True
initialization = 'he'
if reset_progress is True:
reset(weights, method=initialization)
reset(biases, method=initialization)
train_loss_history = []
train_acc_history = []
val_loss_history = []
val_acc_history = []
exit_reason = ""
backups = [{'parameters':[param.clone() for param in (weights + biases)],
'index':len(train_loss_history),
'train_loss':999 if train_loss_history == [] else train_loss_history[-1]}]
backup_during_training = True
n_checkpoint = 10 # backup the parameters and iteration number n_checkpoint times during a full set of iterations
checkpoint_interval = epoch // n_checkpoint
for iteration in range(epoch):
batch_losses = []
batch_accs = []
for X_batch, Y_batch in mini_batch(X_train, Y_train, batch_size=batch_size):
output = feedforward(X_batch, weights, biases)
loss = loss_function(output, Y_batch, weights, lambda_term, len(X_batch))
loss.backward()
for param in (weights + biases):
param.data -= learning_rate * param.grad.data
param.grad.data.zero_()
train_pred = predict(output, one_hot=True)
batch_losses.append(loss.item())
batch_accs.append(accuracy_score(train_pred, Y_batch))
avg_loss = sum(batch_losses) / len(batch_losses)
avg_acc = sum(batch_accs) / len(batch_accs)
train_loss_history.append(avg_loss)
train_acc_history.append(avg_acc)
val_prob = feedforward(X_val, weights, biases)
val_pred = predict(val_prob, one_hot=True)
val_loss_history.append(binary_class_cross_entropy_loss(val_prob, Y_val).item())
val_acc_history.append(accuracy_score(val_pred, Y_val))
if (iteration + 1) % 10 == 0 or iteration == 0:
print(f"\nIteration {iteration + 1}")
print(f"[Training Loss = {train_loss_history[-1]:.6f}\t"
f"Training Accuracy = {train_acc_history[-1]:.6f}]\n"
f"[Validation Loss = {val_loss_history[-1]:.6f}\t\t"
f"Validation Accuracy = {val_acc_history[-1]:.6f}]")
if backup_during_training is True and iteration % checkpoint_interval == 0:
backups.append({'parameters':[param.clone() for param in (weights + biases)],
'index':len(train_loss_history),
'train_loss':train_loss_history[-1]})
if train_loss_history[-1] < tol_loss:
exit_reason = "Loss value less than tol_loss value"
break
elif len(train_loss_history) > 1 and abs(train_loss_history[-1] - train_loss_history[-2]) < tol_step:
exit_reason = "Loss value change less than tol_step value"
break
elif iteration == (epoch - 1):
exit_reason = "Iteration finished"
test_accuracy = accuracy_score(feedforward(X_test, weights, biases), Y_test)
print(f"\n{exit_reason}")
print(f"Training finished in {iteration + 1} iterations")
print(f"\nFinal training loss = {train_loss_history[-1]}")
print(f"Final training accuracy = {train_acc_history[-1]}")
print(f"\nFinal validation loss = {val_loss_history[-1]}")
print(f"Final validation accuracy = {val_acc_history[-1]}")
print(f"\nTest accuracy = {test_accuracy}")
# -
# ## This part of code is for restoring backed up parameters in case things go crazy
list(enumerate([backup['train_loss'] for backup in backups]))
# +
are_you_sure_you_want_to_revert = False # Are you really really sure??
i = 0 # Which backup
if are_you_sure_you_want_to_revert is True:
for n, param in enumerate(weights + biases):
param.data = backups[i]['parameters'][n].data
train_loss_history = train_loss_history[:backups[i]['index']]
train_acc_history = train_acc_history[:backups[i]['index']]
val_loss_history = val_loss_history[:backups[i]['index']]
val_acc_history = val_acc_history[:backups[i]['index']]
# -
# ---
# ## Plot the training loss over iterations
plt.plot(range(1, len(train_loss_history) + 1), train_loss_history)
plt.grid()
plt.title("Train Loss Over Iteration")
plt.xlabel("Iteration")
plt.ylabel("Loss")
plt.xlim(0, None)
plt.ylim(0, None);
len(train_loss_history)
# ### Looks good!
# ## Let's see the examples that it performs well and those that it does poorly
loss_sorted = binary_class_cross_entropy_loss(feedforward(X_train, weights, biases), Y_train, per_example=True).sort(dim=0)
top5_highloss = (loss_sorted[0][-5:], loss_sorted[1][-5:])
top5_lowloss = (loss_sorted[0][:5], loss_sorted[1][:5])
fig, axes = plt.subplots(nrows=1, ncols=len(top5_highloss[1]), figsize=(20, 20), dpi=100)
for loss, idx, ax in zip(top5_highloss[0], top5_highloss[1], axes):
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(f"true : {Y_train[idx].argmax()}")
ax.set_xlabel(f"predicted : {predict(feedforward(X_train[idx].unsqueeze(0), weights, biases)).item()}\n"
f"loss : {loss}")
ax.imshow(X_train[idx].reshape(28, 28), cmap="Greys")
fig, axes = plt.subplots(nrows=1, ncols=len(top5_lowloss[1]), figsize=(20, 20), dpi=100)
for loss, idx, ax in zip(top5_lowloss[0], top5_lowloss[1], axes):
ax.set_xticks([])
ax.set_yticks([])
ax.set_title(f"true : {Y_train[idx].argmax()}")
ax.set_xlabel(f"predicted : {predict(feedforward(X_train[idx].unsqueeze(0), weights, biases)).item()}\n"
f"loss : {loss}")
ax.imshow(X_train[idx].reshape(28, 28), cmap="Greys")
# ### Interestingly, it got really good at classifying few specific digits!
# ---
# ## Making the training process into a function so that it can be used in different contexts
# * (WIP)!
def train(X_train, Y_train, weight_matrices, bias_matrices=None, X_val=None, Y_val=None, X_test=None, Y_test=None,
epoch=100, learning_rate=1.0, reg_coef=0,
report_interval=10, reset_progress=False,
tol_loss=1e-3, tol_step = 1e-8,
logs=[[],[],[],[]], backups=[], n_backups=0):
assert bias_matrices is None or len(weight_matrices) == len(bias_matrices), "The number of weights and biases must match"
assert isinstance(report_interval, int), "The reporting interval must be an integer value, but {type(report_interval)} received"
assert isinstance(n_backups, int), f"The number of backups must be an integer value, but {type(n_backups)} received"
train_loss_history = logs[0]
train_acc_history = logs[1]
val_loss_history = logs[2]
val_acc_history = logs[3]
params = weight_matrices + bias_matrices if bias_matrices is not None else weight_matrices
if reset_progress is True:
reset(weight_matrices)
if bias_matrices is not None:
reset(bias_matrices)
train_loss_history.clear()
train_acc_history.clear()
val_loss_history.clear()
val_acc_history.clear()
exit_reason = ""
if report_interval <= 0:
report_interval = epoch + 1
backup_during_training = True if n_backups > 0 else False
if backup_during_training:
backup_interval = epoch // n_backups # backup the parameters, iteration number(starting from 1), and train loss for 'n_backups' times during a full set of iterations
backups.append({'weights':[weight.data.clone() for weight in weight_matrices],
'biases':[bias.data.clone() for bias in (bias_matrices if bias_matrices is not None else [])],
'index':len(train_loss_history),
'train_loss':999 if train_loss_history == [] else train_loss_history[-1]})
for iteration in range(epoch):
output = feedforward(X_train, weight_matrices, bias_matrices)
loss = loss_function(output, Y_train, weight_matrices, reg_coef, len(X_train))
loss.backward()
for param in ():
param.data -= learning_rate * param.grad.data
param.grad.data.zero_()
train_pred = predict(output, one_hot=True)
train_loss_history.append(loss.item())
train_acc_history.append(accuracy_score(train_pred, Y_train))
if X_val is not None and Y_val is not None:
val_prob = feedforward(X_val, weights, biases)
val_pred = predict(val_prob, one_hot=True)
val_loss_history.append(binary_class_cross_entropy_loss(val_prob, Y_val).item())
val_acc_history.append(accuracy_score(val_pred, Y_val))
if (iteration + 1) % report_interval == 0:
print(f"\nIteration {iteration + 1}")
print(f"[Training Loss = {train_loss_history[-1]:.6f}\t"
f"Training Accuracy = {train_acc_history[-1]:.6f}]")
if X_val is not None and Y_val is not None:
print(f"[Validation Loss = {val_loss_history[-1]:.6f}\t\t"
f"Validation Accuracy = {val_acc_history[-1]:.6f}]")
if backup_during_training is True and iteration % checkpoint_interval == 0:
backups.append({'parameters':[param.clone() for param in (weights + biases)],
'index':len(train_loss_history),
'train_loss':train_loss_history[-1]})
if train_loss_history[-1] < tol_loss:
exit_reason = "Loss value less than tol_loss value"
break
elif len(train_loss_history) > 1 and abs(train_loss_history[-1] - train_loss_history[-2]) < tol_step:
exit_reason = "Loss value change less than tol_step value"
break
elif iteration == (epoch - 1):
exit_reason = "Iteration finished"
print(f"\n{exit_reason}")
print(f"Training finished in {iteration + 1} iterations")
print(f"\nFinal training loss = {train_loss_history[-1]}")
print(f"Final training accuracy = {train_acc_history[-1]}")
if X_val is not None and Y_val is not None:
print(f"\nFinal validation loss = {val_loss_history[-1]}")
print(f"Final validation accuracy = {val_loss_history[-1]}")
if X_test is not None and Y_test is not None:
test_accuracy = accuracy_score(feedforward(X_test, weight_matrices, bias_matrices), Y_test)
print(f"\nTest accuracy = {test_accuracy}")
| simple_nn_for_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple usage of a set of MPI engines
# This example assumes you've started a cluster of N engines (4 in this example) as part
# of an MPI world.
#
# Our documentation describes [how to create an MPI profile](https://ipyparallel.readthedocs.io/en/stable/process.html#using-ipcluster-in-mpiexec-mpirun-mode)
# and explains [basic MPI usage of the IPython cluster](https://ipyparallel.readthedocs.io/en/stable/mpi.html).
#
#
# For the simplest possible way to start 4 engines that belong to the same MPI world,
# you can run this in a terminal:
#
# <pre>
# ipcluster start --engines=MPI -n 4
# </pre>
#
# or start an MPI cluster from the cluster tab if you have one configured.
#
# Once the cluster is running, we can connect to it and open a view into it:
import ipyparallel as ipp
rc = ipp.Client()
view = rc[:]
# Let's define a simple function that gets the MPI rank from each engine.
@view.remote(block=True)
def mpi_rank():
from mpi4py import MPI
comm = MPI.COMM_WORLD
return comm.Get_rank()
mpi_rank()
# To get a mapping of IPython IDs and MPI rank (these do not always match),
# you can use the get_dict method on AsyncResults.
mpi_rank.block = False
ar = mpi_rank()
ar.get_dict()
# With %%px cell magic, the next cell will actually execute *entirely on each engine*:
# +
# %%px
from mpi4py import MPI
comm = MPI.COMM_WORLD
size = comm.Get_size()
rank = comm.Get_rank()
if rank == 0:
data = [(i+1)**2 for i in range(size)]
else:
data = None
data = comm.scatter(data, root=0)
assert data == (rank+1)**2, 'data=%s, rank=%s' % (data, rank)
{
'data': data,
'rank': rank,
}
| examples/Using MPI with IPython Parallel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 函数
#
# - 函数可以用来定义可重复代码,组织和简化
# - 一般来说一个函数在实际开发中为一个小功能
# - 一个类为一个大功能
# - 同样函数的长度不要超过一屏
def fun_name():
print('天才')
fun_name() #()表示函数调用
# ## 定义一个函数
#
# def function_name(list of parameters):
#
# do something
# 
# - 以前使用的random 或者range 或者print.. 其实都是函数或者类
# ## 调用一个函数
# - functionName()
# - "()" 就代表调用
def panduanjiou(): #这个必须先运行一下哦,然后就可以调用了哦
num1 = eval(input('>>'))
if num1 % 2 == 0:
print ('偶数')
else :
print ('奇数')
panduanjiou()
def panduansushu():
count = 0
num2 = int(input('>>'))
for s in range(2,num2):
if num2 % s == 0:
count += 1
if count == 1:
print ('是素数')
else :
print ('不是')
panduansushu()
# 
# ## 带返回值和不带返回值的函数
# - return 返回的内容
# - return 返回多个值
# - 一般情况下,在多个函数协同完成一个功能的时候,那么将会有返回值
# 
#
# - 当然也可以自定义返回None
# ## EP:
# 
# ## 类型和关键字参数
# - 普通参数
# - 多个参数
# - 默认值参数
# - 不定长参数
import os
def kuajiang(name):
os.system('say {}你真是一个天才'.format(name))
kuajiang('嘎') #名字是可以随意添加的
kuajiang(name = '阿达') #带参数名的传递方式
def input_():
num = eval(input('>>'))
res3 = san(num)
res2 = er(num)
print (res3 - res2)
def san(num): #定义可以放在任意位置
return num ** 3
def er(num):
return num ** 2
input_()
x =2
# ## 普通参数
# ## 多个参数
# ## 默认值参数
import os
#如果参数含有默认值,一定要放到最后面!!!!!!!!!
def kuajiang(name1,name2,name3):
print('say {}{}{}你们都是小帅哥'.format(name1,name2,name3))
kuajiang(name1='z',name2='f',name3='h')
# +
#def login(is_ok_and_login = False):
# acount == str(input('账号或密码不正确,请再次输入账号:'))
# password == int(input('请再次输入密码:'))
# -
acount = '<EMAIL>' #str(input('输入账号:'))
password = '<PASSWORD>' #int(input('输入密码:'))
is_ok_and_qitian = False
def login(account_login,password_login):
if account_login == acount and password_login == password:
print('登陆成功')
else:
print('账号或密码错误')
# +
def qitian():
global is_ok_and_qitian
#如果你想要为一个定义在函数外的变量赋值,那么你就得告诉Python这个变量名不是局部的,而是 全局 的。
#我们使用global语句完成这一功能。没有global语句,是不可能为定义在函数外的变量赋值的。
#你可以使用定义在函数外的变量的值(假设在函数内没有同名的变量)。然而,我并不鼓励你这样做,
#并且你应该尽量避免这样做,因为这使得程序的读者会不清楚这个变量是在哪里定义的。使用global语句可以清楚地表明变量是在外面的块定义的。
if is_ok_and_qitian == False:
print('是否七天免登陆?y/n')
res = input('>>')
account_login = input('输入账号:')
password_login = input('输入密码:')
if res == 'y':
login(account_login,password_login)
is_ok_and_qitian = True
else:
login(account_login,password_login)
else:
print('登陆成功!')
# -
qitian()
# ## 强制命名
# +
#定义时,前面会加一个*,再后面输入是必须添加定义参数
# -
# ## 不定长参数
# - \*args
# > - 不定长,来多少装多少,不装也是可以的
# - 返回的数据类型是元组
# - args 名字是可以修改的,只是我们约定俗成的是args
# - \**kwargs
# > - 返回的字典
# - 输入的一定要是表达式(键值对)
# - name,\*args,name2,\**kwargs 使用参数名
# *args是下水道,来多少装多少
# **kwargs是加强版的下水道
def test1(*args):
print(args)
test1(1,2,3,4,5,6,7,8,9) # 元组 (可迭代)
def test2(**kwages): #函数的传参
print(kwages)
test2(a=1,b=2,c=3) #这个输出的是“字典” 有键名和键值,‘’里面是键名
def test3(*args,**kwargs):
print(args)
print(kwargs)
test3(1,2,3,a=4,b=5,c=6) #都是 可传可不传的
# ## 变量的作用域
# - 局部变量 local
# - 全局变量 global
# - globals 函数返回一个全局变量的字典,包括所有导入的变量
# - locals() 函数会以字典类型返回当前位置的全部局部变量。
a =100
b =200
c =True
d =[]
def test4():
a = 1000
b = 2000
c = False
d.append(999)
print(a,b,c)
print(a,b,c)
test4()
print(a,b,c,d)
locals()
# ## 注意:
# - global :在进行赋值操作的时候需要声明
# - 官方解释:This is because when you make an assignment to a variable in a scope, that variable becomes local to that scope and shadows any similarly named variable in the outer scope.
# - 
# # Homework
# - 1
# 
def homework1():
n = 1
count = 0
a = int(input('>>'))
for s in range(1,a):
if n < a:
c = n * (3 * n - 1) / 2
n += 1
print(int(c),end=' ')
count += 1
if(count%10==0):
print(end='\n')
homework1()
# - 2
# 
a = int(input("您要键入的数字:"))
#变成字符串了str~~~
b = len(str(a)) #len是用来获取字符串长度
print(b)
# 三为数的太简单,无视并跳过!!!! 下面为 n 位数的计算方法函数~~~~~~~~~~~~~
def homework2():
# 输入任意数字并赋值给input_num
input_num = input('请输入数字:')
# 声明一个变量用于存放相加计算后的数字,初始值设定为0
sum_num = 0
# 判断输入的内容是否为数字
#if input_num.isdigit():
# 循环input_num字符长度次,其中range()为python中生成数字序列的函数,len()为python获取字符串长度的函数
# 此循环的意思等于JavaScript中的for(i = 0; i < input_num.length; i++){}
for i in range(len(input_num)):
# python中截取字符串中某一位的字符,方法为str[0:2](即:截取出名称为str的字符串的第0位到第2位之间的字符)
# 此处的意思是,截取输入的字符中第i个到第i+1个字符之间的字符,并赋值给cut_num
cut_num = input_num[i:i + 1]
# 重新给变量sum_num赋值为sum_num+截取到的数字的和
sum_num = sum_num + int(cut_num)
# 打印(即:在控制台输出)输入的数字中各个数位数字相加的总和
print('输入了' + str(len(input_num)) + '个数字\n数字的综合为:' + str(sum_num))
#else:
# print('你输入的不是数字,我不能计算,再见')
homework2()
# - 3
# 
def homework3():
a,b,c = eval(input('输入三个数:'))
if a>b>c :
a,b,c=a,b,c
elif a>c>b :
a,b,c=a,c,b
elif b>a>c :
a,b,c=b,a,c
elif b>c>a :
a,b,c=b,c,a
elif c>a>b :
a,b,c=c,a,b
elif c>b>a :
a,b,c=c,b,a
print('升序排列后为:' + str(c),str(b),str(a))
homework3()
# - 4
# 
def homework4():
money = int(input('输入投资额:'))
lilv = eval(input('输入百分比格式的年利率:'))
lixi = 0
print('Years Future Value')
for i in range(1,31):
lixi = lixi + money*lilv / 100
money = money + lixi
print(str(i)+ ' '+ str(money))
homework4()
# - 5
# 
def homework5(ch1,ch2,number):
ch1,ch2,number = input('输入两个字符,和每行打印的个数:')
# - 6
# 
def homework6():
year1 = int(input('输入启示年份:'))
year2 = int(input('输入末尾年份:'))
sum1 = 0
for i in range(0,(year2 - year1)):
if (year1 % 100 == 0 and year1 % 400 == 0) or (year1 % 100 != 0 and year1 % 4 == 0) :
sum1 += 366
year1 += 1
else :
year1 += 1
sum1 += 365
print(sum1)
homework6()
# - 7
# 
# - 8
# 
# - 9
# 
# 
# - 10
# 
# - 11
# ### 去网上寻找如何用Python代码发送邮件
| wk_test5_914.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# # PixelCNN
#
# **Author:** [ADMoreau](https://github.com/ADMoreau)<br>
# **Date created:** 2020/05/17<br>
# **Last modified:** 2020/05/23<br>
# **Description:** PixelCNN implemented in Keras.
# + [markdown] colab_type="text"
# ## Introduction
#
# PixelCNN is a generative model proposed in 2016 by <NAME> et al.
# (reference: [Conditional Image Generation with PixelCNN Decoders](https://arxiv.org/abs/1606.05328)).
# It is designed to generate images (or other data types) iteratively
# from an input vector where the probability distribution of prior elements dictates the
# probability distribution of later elements. In the following example, images are generated
# in this fashion, pixel-by-pixel, via a masked convolution kernel that only looks at data
# from previously generated pixels (origin at the top left) to generate later pixels.
# During inference, the output of the network is used as a probability distribution
# from which new pixel values are sampled to generate a new image
# (here, with MNIST, the pixel values range from white (0) to black (255).
#
# + colab_type="code"
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tqdm import tqdm
# + [markdown] colab_type="text"
# ## Getting the data
#
# + colab_type="code"
# Model / data parameters
num_classes = 10
input_shape = (28, 28, 1)
n_residual_blocks = 5
# The data, split between train and test sets
(x, _), (y, _) = keras.datasets.mnist.load_data()
# Concatenate all of the images together
data = np.concatenate((x, y), axis=0)
# Round all pixel values less than 33% of the max 256 value to 0
# anything above this value gets rounded up to 1 so that all values are either
# 0 or 1
data = np.where(data < (0.33 * 256), 0, 1)
data = data.astype(np.float32)
# + [markdown] colab_type="text"
# ## Create two classes for the requisite Layers for the model
#
# + colab_type="code"
# The first layer is the PixelCNN layer. This layer simply
# builds on the 2D convolutional layer, but includes masking.
class PixelConvLayer(layers.Layer):
def __init__(self, mask_type, **kwargs):
super(PixelConvLayer, self).__init__()
self.mask_type = mask_type
self.conv = layers.Conv2D(**kwargs)
def build(self, input_shape):
# Build the conv2d layer to initialize kernel variables
self.conv.build(input_shape)
# Use the initialized kernel to create the mask
kernel_shape = self.conv.kernel.get_shape()
self.mask = np.zeros(shape=kernel_shape)
self.mask[: kernel_shape[0] // 2, ...] = 1.0
self.mask[kernel_shape[0] // 2, : kernel_shape[1] // 2, ...] = 1.0
if self.mask_type == "B":
self.mask[kernel_shape[0] // 2, kernel_shape[1] // 2, ...] = 1.0
def call(self, inputs):
self.conv.kernel.assign(self.conv.kernel * self.mask)
return self.conv(inputs)
# Next, we build our residual block layer.
# This is just a normal residual block, but based on the PixelConvLayer.
class ResidualBlock(keras.layers.Layer):
def __init__(self, filters, **kwargs):
super(ResidualBlock, self).__init__(**kwargs)
self.conv1 = keras.layers.Conv2D(
filters=filters, kernel_size=1, activation="relu"
)
self.pixel_conv = PixelConvLayer(
mask_type="B",
filters=filters // 2,
kernel_size=3,
activation="relu",
padding="same",
)
self.conv2 = keras.layers.Conv2D(
filters=filters, kernel_size=1, activation="relu"
)
def call(self, inputs):
x = self.conv1(inputs)
x = self.pixel_conv(x)
x = self.conv2(x)
return keras.layers.add([inputs, x])
# + [markdown] colab_type="text"
# ## Build the model based on the original paper
#
# + colab_type="code"
inputs = keras.Input(shape=input_shape)
x = PixelConvLayer(
mask_type="A", filters=128, kernel_size=7, activation="relu", padding="same"
)(inputs)
for _ in range(n_residual_blocks):
x = ResidualBlock(filters=128)(x)
for _ in range(2):
x = PixelConvLayer(
mask_type="B",
filters=128,
kernel_size=1,
strides=1,
activation="relu",
padding="valid",
)(x)
out = keras.layers.Conv2D(
filters=1, kernel_size=1, strides=1, activation="sigmoid", padding="valid"
)(x)
pixel_cnn = keras.Model(inputs, out)
adam = keras.optimizers.Adam(learning_rate=0.0005)
pixel_cnn.compile(optimizer=adam, loss="binary_crossentropy")
pixel_cnn.summary()
pixel_cnn.fit(
x=data, y=data, batch_size=128, epochs=50, validation_split=0.1, verbose=2
)
# + [markdown] colab_type="text"
# ## Demonstration
#
# The PixelCNN cannot generate the full image at once. Instead, it must generate each pixel in
# order, append the last generated pixel to the current image, and feed the image back into the
# model to repeat the process.
#
# You can use the trained model hosted on [Hugging Face Hub](https://huggingface.co/keras-io/pixel-cnn-mnist) and try the demo on [Hugging Face Spaces](https://huggingface.co/spaces/keras-io/pixelcnn-mnist-image-generation).
# + colab_type="code"
from IPython.display import Image, display
# Create an empty array of pixels.
batch = 4
pixels = np.zeros(shape=(batch,) + (pixel_cnn.input_shape)[1:])
batch, rows, cols, channels = pixels.shape
# Iterate over the pixels because generation has to be done sequentially pixel by pixel.
for row in tqdm(range(rows)):
for col in range(cols):
for channel in range(channels):
# Feed the whole array and retrieving the pixel value probabilities for the next
# pixel.
probs = pixel_cnn.predict(pixels)[:, row, col, channel]
# Use the probabilities to pick pixel values and append the values to the image
# frame.
pixels[:, row, col, channel] = tf.math.ceil(
probs - tf.random.uniform(probs.shape)
)
def deprocess_image(x):
# Stack the single channeled black and white image to RGB values.
x = np.stack((x, x, x), 2)
# Undo preprocessing
x *= 255.0
# Convert to uint8 and clip to the valid range [0, 255]
x = np.clip(x, 0, 255).astype("uint8")
return x
# Iterate over the generated images and plot them with matplotlib.
for i, pic in enumerate(pixels):
keras.preprocessing.image.save_img(
"generated_image_{}.png".format(i), deprocess_image(np.squeeze(pic, -1))
)
display(Image("generated_image_0.png"))
display(Image("generated_image_1.png"))
display(Image("generated_image_2.png"))
display(Image("generated_image_3.png"))
| examples/generative/ipynb/pixelcnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np, matplotlib.pyplot as plt, pandas as pd
dataset = pd.read_csv('50_startups.csv')
dataset.head()
X = dataset.iloc[:,:-1].values
Y = dataset.iloc[:,-1].values
# +
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
ct = ColumnTransformer([("State",OneHotEncoder(),[3])],remainder = 'passthrough')
X = ct.fit_transform(X)
# -
X = np.array(X[:,[0,3]])
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size = 0.2, random_state =0)
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train,Y_train)
y_pred = regressor.predict(X_test)
fig,ax = plt.subplots()
ax.plot(Y_test) #default color red
ax.plot(y_pred) #default color blue
Y_test
y_pred- array([103015.20159795, 132582.27760817, 132447.73845176, 71976.09851257,
178537.48221058, 116161.24230165, 67851.69209675, 98791.73374686,
113969.43533013, 167921.06569553])
y_pred_new - array([104793.79144158, 133950.03032448, 135315.09697767, 72317.50514982,
179170.2710366 , 109948.04206623, 65795.34503314, 100610.57995573,
111554.00908759, 169523.90691948])
Y_test- array([103282.38, 144259.4 , 146121.95, 77798.83, 191050.39, 105008.31,
81229.06, 97483.56, 110352.25, 166187.94])
| Regression/Multiple Linear Regression/Checking result of Backward Elimination .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import pickle as cPickle
import numpy as np
from sklearn import svm
import sklearn.utils
from scipy.sparse import csr_matrix
from sklearn.model_selection import GridSearchCV, cross_val_score
from sklearn.metrics import classification_report
import random
import matplotlib.pyplot as plt
from scipy.stats.stats import pearsonr
from collections import defaultdict
import math
from sklearn import preprocessing
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
from scipy.stats import spearmanr
from sklearn import linear_model
import re
import copy
import seaborn as sns
import pandas as pd
import scipy.stats
import statsmodels.stats.proportion
from sklearn.cross_validation import LeaveOneOut
from prediction_utils.show_examples import update, generate_snapshots, clean
from prediction_utils.features2vec import _get_term_features, _get_last_n_action_features, \
_get_action_features, _get_repeatition_features, _get_balance_features, documents2feature_vectors
# +
import matplotlib
from matplotlib.ticker import FuncFormatter
def to_percent(y, position):
# Ignore the passed in position. This has the effect of scaling the default
# tick locations.
s = str(int(100 * y))
# The percent symbol needs escaping in latex
if matplotlib.rcParams['text.usetex'] is True:
return s + r'$\%$'
else:
return s + '%'
# -
COLOR = ["#bb5f4c",
"#8e5db0",
"#729b57"]
# +
import matplotlib.pyplot as plt
import numpy as np
plt.rcdefaults()
fig, ax = plt.subplots(figsize=(8, 4))
# Example data
# conv+user C = 0.007
# BOW C= 0.00007
# Human 1 C = 0.0007
# FULL C = 0.0007
# User only C = 0.005
# conv only C = 0.005
methods = ('Human', 'Human Perception', 'BOW', \
'Conversational + Question', 'Conversational only', \
'Participant Features')
y_pos = np.arange(len(methods))
performance = ( 0.595, 0.551, 0.554, 0.578, 0.564, 0.530)
err = [0.017,0.011, 0.011, 0.011, 0.011, 0.01]
barwidth = 0.5
gap = 0.1
ax.barh(y_pos * (barwidth + gap), performance, barwidth, xerr=err, align='center',
color=[COLOR[c] for c in [0, 0, 1, 1, 1, 2]], ecolor='black')
ax.set_xlim(right=0.80)
upperbound = 0.759
fontsize=13
for i in range(len(methods)):
text = '%.1f'%(performance[i]* 100) + '%'
if methods[i] == 'Human Perception':
text += '$\dag$'
if methods[i] in ['BOW']:
text += '*'
if methods[i] == 'Participant Features':
text += '***'
ax.text( upperbound - 0.01, i * (barwidth+gap), text, fontsize=fontsize, horizontalalignment ='right')
ax.text( 0.01, i * (barwidth+gap),methods[i], horizontalalignment ='left', fontsize=fontsize, fontweight='bold', color='white')
ax.set_yticks([])
#ax.set_yticklabels(methods)
ax.invert_yaxis() # labels read top-to-bottom
ax.set_xlabel('Accuracy', fontsize=fontsize)
plt.axvline(x=0.5, color='k', linestyle='--')
plt.axvline(x=upperbound, color='k', linestyle='-.')
#plt.axhline(y=1.5, color='k', linestyle='-')
#plt.axhline(y=4.5, color='k', linestyle='-')
formatter = FuncFormatter(to_percent)
# Set the formatter
plt.gca().xaxis.set_major_formatter(formatter)
plt.xticks(fontsize=fontsize)
plt.show()
# -
def plot_profiles1(profiles, ASPECTS, experience=-1):
catergories = {'Min': 0, 'Max': 1, 'In the Middle': 2, 'Anonymous':3, 'New Comer':4, 'No Gap': 5, 'Bot': 6}
cats = ['min', 'max', 'in the middle', 'Anonymous', 'New Comer']
f, ax = plt.subplots(1, figsize=(13,6))
bar_width = 0.4
bar_l = [i for i in range(len(ASPECTS))]
tick_pos = [i+bar_width for i in bar_l]
colors = ['pink', 'mediumslateblue', 'steelblue', 'mediumaquamarine', 'darksalmon']
bads = [[[], [], [], [], [], [], []], [[], [], [], [], [], [], []]]
total = len(profiles[0])
alpha=[0.9, 0.3]
conv_label = ['Offender is ', 'Non-offender is ']
mins = [[], []]
cnts = [[[], [], [], [], [], [], []], [[], [], [], [], [], [], []]]
rects = []
for clss in [0, 1]:
for aspect in ASPECTS:
cur = []
for ind in range(len(catergories)):
bads[clss][ind].append(0)
cnts[clss][ind].append(0)
for p in profiles[clss]:
# if not('experience') in p or p['experience'] <= experience:
# continue
bads[clss][catergories[p[aspect]]][-1] += 1
cnts[clss][catergories[p[aspect]]][-1] += 1
if catergories[p[aspect]] == 0:
cur.append(1)
elif catergories[p[aspect]] < 3:
cur.append(0)
mins[clss].append(cur)
previous = [0 for a in ASPECTS]
first_three = [0 for a in ASPECTS]
for bad in bads[clss][:3]:
for ii, b in enumerate(bad):
first_three[ii] += b
for ind,bad in enumerate(bads[clss][:3]):
for ii, b in enumerate(bad):
if first_three[ii]: bad[ii] = bad[ii] / first_three[ii]
bads[clss][ind] = bad
rects = ax.bar(bar_l, bad, label=conv_label[clss] + cats[ind], bottom = previous, alpha=alpha[clss], \
color=colors[ind],width=bar_width,edgecolor='white')
for ind, rect in enumerate(rects):
ax.text(rect.get_x() + rect.get_width()/2., (bad[ind] / 3 + previous[ind]),
'%.1f' % (bad[ind]*100) + '%',
ha='center', va='bottom')
for ii, b in enumerate(bad):
previous[ii] += b
# ax.legend(loc="upper center", bbox_to_anchor=(1,1), fontsize='large')
ax.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc=3,
ncol=3, mode="expand", borderaxespad=0., fontsize='large')
bar_l = [b+bar_width for b in bar_l]
if clss:
print('Good Total:')
else:
print('Bad Total:')
for ii,aspect in enumerate(ASPECTS):
print(aspect, first_three[ii])
ax.set_ylabel("Percentage among All the Cases", fontsize='large')
# ax.set_xlabel("Aspect")
Xticks = ['Proportion replied',\
'Being replied latency', 'Reply latency', \
'Age', 'Status', \
'# edits on Wikipedia']
plt.xticks([t - bar_width / 2 for t in tick_pos], Xticks, fontsize='large')
# ax.set_xlabel("")
# rotate axis labels
plt.setp(plt.gca().get_xticklabels(), rotation=20, horizontalalignment='right')
# plt.title('Who\'s the Attacker')
# shot plot
plt.show()
# for aspect in ASPECTS:
# print(aspect, first_three[0], first_three[1])
print('Test 1')
for ind, aspect in enumerate(ASPECTS):
print(aspect)
print('Average in Ggap: ', np.mean(mins[1][ind]))
print('Average of Bgap: ', np.mean(mins[0][ind]))
if np.mean(mins[1][ind]) == 1 or np.mean(mins[1][ind]) == 0:
continue
print(scipy.stats.mannwhitneyu(mins[0][ind], mins[1][ind]))
print('\n')
print('Test 2')
clss = 0
for ind, aspect in enumerate(ASPECTS):
print(aspect, ':', scipy.stats.binom_test(cnts[clss][0][ind], cnts[clss][0][ind] + cnts[clss][1][ind]))
# print(cnts[clss][0][ind], cnts[clss][1][ind])
print('\n')
print('Test 3')
clss = 1
for ind, aspect in enumerate(ASPECTS):
print(aspect, ':', scipy.stats.binom_test(cnts[clss][0][ind], cnts[clss][0][ind] + cnts[clss][1][ind]))
| experimental/conversation_go_awry/Result Plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 (''.venv'': pipenv)'
# language: python
# name: python3
# ---
# # pip install matplotlib ipympl
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
X = np.linspace(0, 2*np.pi)
Y = np.sin(X)
fig, ax = plt.subplots()
ax.plot(X, Y)
| src/test/datascience/widgets/notebooks/matplotlib_widgets_inline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cognitive Age Prediction with MEG
# +
import mne
import numpy as np
import pandas as pd
from camcan.utils import (plot_pred, plot_learning_curve,
plot_barchart, run_ridge,
plot_error_age, plot_error_segments,
plot_boxplot)
# %matplotlib inline
# -
CV = 10
# store mae, std for the summary plot
meg_mae = {}
meg_mae_std = {}
meg_pred_diff = {}
# ## Read Data
subjects_data = pd.read_csv('../../data/participant_data.csv', index_col=0)
subjects_predictions = pd.DataFrame(subjects_data.age, index=subjects_data.index, dtype=float)
subjects_data.head()
FREQ_BANDS = ('alpha',
'beta_high',
'beta_low',
'delta',
'gamma_high',
'gamma_lo',
'gamma_mid',
'low',
'theta')
# +
MEG_SOURCE_SPACE_DATA = '../../data/meg_source_space_data.h5'
meg_data = pd.read_hdf(MEG_SOURCE_SPACE_DATA, key='meg')
columns_to_exclude = ('band', 'fmax', 'fmin', 'subject')
parcellation_labels = [c for c in meg_data.columns if c
not in columns_to_exclude]
band_data = [meg_data[meg_data.band == bb].set_index('subject')[
parcellation_labels] for bb in FREQ_BANDS]
meg_data = pd.concat(band_data, axis=1, join='inner', sort=False)
print(f'Found {len(meg_data)} subjects')
# -
# ## The Brain Age Prediction using MEG data in Source Space
# +
df_pred, arr_mae, arr_r2, train_sizes, train_scores, test_scores = \
run_ridge(meg_data, subjects_data, alphas=np.logspace(-3, 5, 100), cv=CV)
arr_mae = -arr_mae
mae = arr_mae.mean()
std = arr_mae.std()
print('MAE: %.2f' % mae)
print('MAE STD: %.2f' % std)
meg_mae['MEG'] = arr_mae
meg_mae_std['MEG'] = (mae, std)
subjects_predictions.loc[df_pred.index, 'MEG'] = df_pred['y']
plot_pred(subjects_predictions.loc[df_pred.index].age,
subjects_predictions.loc[df_pred.index, 'MEG'],
mae, title='Age Prediction from MEG with SPoC')
plot_learning_curve(train_sizes, train_scores, test_scores)
# -
plot_error_age(subjects_predictions, ylim=(-2, 55))
plot_error_segments(subjects_predictions, segment_len=10)
plot_boxplot(meg_mae, title='Age Prediction, MEG')
| notebook/age_prediction/meg_age_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://leetcode.com/problemset/top-interview-questions/
from typing import List
import json
# +
# Problem 101, https://leetcode.com/problems/symmetric-tree/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def isSymmetric(self, root: TreeNode) -> bool:
if root is None:
return True
pairs = [(root.left, root.right)]
while len(pairs) > 0:
l, r = pairs.pop()
if l is None:
if r is None:
continue
else:
return False
if r is None:
return False
if l.val != r.val:
return False
pairs.append((l.right, r.left))
pairs.append((l.left, r.right))
return True
def stringToTreeNode(input):
input = input.strip()
input = input[1:-1]
if not input:
return None
inputValues = [s.strip() for s in input.split(',')]
root = TreeNode(int(inputValues[0]))
nodeQueue = [root]
front = 0
index = 1
while index < len(inputValues):
node = nodeQueue[front]
front = front + 1
item = inputValues[index]
index = index + 1
if item != "null":
leftNumber = int(item)
node.left = TreeNode(leftNumber)
nodeQueue.append(node.left)
if index >= len(inputValues):
break
item = inputValues[index]
index = index + 1
if item != "null":
rightNumber = int(item)
node.right = TreeNode(rightNumber)
nodeQueue.append(node.right)
return root
def main():
import sys
import io
s = '''[1,2,2,3,4,4,3]
[1,2,2,null,3,null,3]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
root = stringToTreeNode(line);
ret = Solution().isSymmetric(root)
out = (ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 102, https://leetcode.com/problems/binary-tree-level-order-traversal/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def levelOrder(self, root: TreeNode) -> List[List[int]]:
if root is None:
return []
lo = []
t = [(0, root)]
while len(t) > 0:
d, n = t.pop()
if d == len(lo):
lo.append([])
lo[d].append(n.val)
if not n.right is None:
t.append((d+1, n.right))
if not n.left is None:
t.append((d+1, n.left))
return lo
def stringToTreeNode(input):
input = input.strip()
input = input[1:-1]
if not input:
return None
inputValues = [s.strip() for s in input.split(',')]
root = TreeNode(int(inputValues[0]))
nodeQueue = [root]
front = 0
index = 1
while index < len(inputValues):
node = nodeQueue[front]
front = front + 1
item = inputValues[index]
index = index + 1
if item != "null":
leftNumber = int(item)
node.left = TreeNode(leftNumber)
nodeQueue.append(node.left)
if index >= len(inputValues):
break
item = inputValues[index]
index = index + 1
if item != "null":
rightNumber = int(item)
node.right = TreeNode(rightNumber)
nodeQueue.append(node.right)
return root
def main():
import sys
import io
s = '''[3,9,20,null,null,15,7]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
root = stringToTreeNode(line);
ret = Solution().levelOrder(root)
out = (ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 103, https://leetcode.com/problems/binary-tree-zigzag-level-order-traversal/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def zigzagLevelOrder(self, root: TreeNode) -> List[List[int]]:
if root is None:
return []
zzlo, d = [], 0
zig, zag = [root], []
while len(zig) > 0:
while len(zig) > 0:
n = zig.pop()
if d == len(zzlo):
zzlo.append([])
zzlo[d].append(n.val)
if not n.left is None:
zag.append(n.left)
if not n.right is None:
zag.append(n.right)
print('zag', [za.val for za in zag])
print([zi.val for zi in zig], [za.val for za in zag])
d += 1
while len(zag) > 0:
n = zag.pop()
if d == len(zzlo):
zzlo.append([])
zzlo[d].append(n.val)
if not n.right is None:
zig.append(n.right)
if not n.left is None:
zig.append(n.left)
print('zig', [zi.val for zi in zig])
print([zi.val for zi in zig], [za.val for za in zag])
d += 1
return zzlo
def stringToTreeNode(input):
input = input.strip()
input = input[1:-1]
if not input:
return None
inputValues = [s.strip() for s in input.split(',')]
root = TreeNode(int(inputValues[0]))
nodeQueue = [root]
front = 0
index = 1
while index < len(inputValues):
node = nodeQueue[front]
front = front + 1
item = inputValues[index]
index = index + 1
if item != "null":
leftNumber = int(item)
node.left = TreeNode(leftNumber)
nodeQueue.append(node.left)
if index >= len(inputValues):
break
item = inputValues[index]
index = index + 1
if item != "null":
rightNumber = int(item)
node.right = TreeNode(rightNumber)
nodeQueue.append(node.right)
return root
def main():
import sys
import io
s = '''[3,9,20,null,null,15,7]
[1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
root = stringToTreeNode(line);
ret = Solution().zigzagLevelOrder(root)
out = (ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 104, https://leetcode.com/problems/maximum-depth-of-binary-tree/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def maxDepth(self, root: TreeNode) -> int:
if root is None:
return 0
ds = []
t = [(1, root)]
while len(t) > 0:
d, n = t.pop()
if n.left is None:
if n.right is None:
ds.append(d)
else:
t.append((d+1, n.right))
else:
t.append((d+1, n.left))
if not n.right is None:
t.append((d+1, n.right))
return max(ds)
def stringToTreeNode(input):
input = input.strip()
input = input[1:-1]
if not input:
return None
inputValues = [s.strip() for s in input.split(',')]
root = TreeNode(int(inputValues[0]))
nodeQueue = [root]
front = 0
index = 1
while index < len(inputValues):
node = nodeQueue[front]
front = front + 1
item = inputValues[index]
index = index + 1
if item != "null":
leftNumber = int(item)
node.left = TreeNode(leftNumber)
nodeQueue.append(node.left)
if index >= len(inputValues):
break
item = inputValues[index]
index = index + 1
if item != "null":
rightNumber = int(item)
node.right = TreeNode(rightNumber)
nodeQueue.append(node.right)
return root
def main():
import sys
import io
s = '''[3,9,20,null,null,15,7]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
root = stringToTreeNode(line);
ret = Solution().maxDepth(root)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 105, https://leetcode.com/problems/construct-binary-tree-from-preorder-and-inorder-traversal/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def buildTree(self, preorder: List[int], inorder: List[int]) -> TreeNode:
# print(preorder, inorder)
n = len(preorder)
if n == 0:
return None
node = TreeNode(preorder[0])
pos = inorder.index(preorder[0])
if pos > 0:
node.left = self.buildTree(preorder[1:pos+1], inorder[0:pos])
if pos < n-1:
node.right = self.buildTree(preorder[pos+1:n], inorder[pos+1:n])
return node
def stringToIntegerList(input):
return json.loads(input)
def treeNodeToString(root):
if not root:
return "[]"
output = ""
queue = [root]
current = 0
while current != len(queue):
node = queue[current]
current = current + 1
if not node:
output += "null, "
continue
output += str(node.val) + ", "
queue.append(node.left)
queue.append(node.right)
return "[" + output[:-2] + "]"
def main():
import sys
import io
s = '''[8,4,2,1,3,6,5,7,12,10,9,11,14,13,15]
[1,2,3,4,5,6,7,8,9,10,11,12,13,14,15]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
preorder = stringToIntegerList(line);
line = next(lines)
inorder = stringToIntegerList(line);
ret = Solution().buildTree(preorder, inorder)
out = treeNodeToString(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 108, https://leetcode.com/problems/convert-sorted-array-to-binary-search-tree/
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def sortedArrayToBST(self, nums: List[int]) -> TreeNode:
print(nums)
n = len(nums)
if n < 2:
return TreeNode(nums[0]) if n == 1 else None
mid = n // 2
node = TreeNode(nums[mid])
node.left = self.sortedArrayToBST(nums[0:mid])
node.right = self.sortedArrayToBST(nums[mid+1:n])
return node
def stringToIntegerList(input):
return json.loads(input)
def treeNodeToString(root):
if not root:
return "[]"
output = ""
queue = [root]
current = 0
while current != len(queue):
node = queue[current]
current = current + 1
if not node:
output += "null, "
continue
output += str(node.val) + ", "
queue.append(node.left)
queue.append(node.right)
return "[" + output[:-2] + "]"
def main():
import sys
import io
s = '''[-10,-3,0,5,9]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
nums = stringToIntegerList(line);
ret = Solution().sortedArrayToBST(nums)
out = treeNodeToString(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 116, https://leetcode.com/problems/populating-next-right-pointers-in-each-node/
# Definition for a Node.
class TreeNode:
def __init__(self, val: int = 0, left: 'Node' = None, right: 'Node' = None, next_: 'Node' = None):
self.val = val
self.left = left
self.right = right
self.next = next_
class Solution:
def connect(self, root: 'Node') -> 'Node':
if root is None:
return None
t = [root]
while len(t) > 0:
node = t.pop()
print(node.val)
if node.left:
node.left.next = node.right
if node.next:
node.right.next = node.next.left
t.append(node.left)
t.append(node.right)
return root
def stringToTreeNode(input):
input = input.strip()
input = input[1:-1]
if not input:
return None
inputValues = [s.strip() for s in input.split(',')]
root = TreeNode(int(inputValues[0]))
nodeQueue = [root]
front = 0
index = 1
while index < len(inputValues):
node = nodeQueue[front]
front = front + 1
item = inputValues[index]
index = index + 1
if item != "null":
leftNumber = int(item)
node.left = TreeNode(leftNumber)
nodeQueue.append(node.left)
if index >= len(inputValues):
break
item = inputValues[index]
index = index + 1
if item != "null":
rightNumber = int(item)
node.right = TreeNode(rightNumber)
nodeQueue.append(node.right)
return root
def treeNodeToString(root):
if not root:
return "[]"
output = ""
queue = [root]
current = 0
while current != len(queue):
node = queue[current]
current = current + 1
if not node:
output += "null, "
continue
next_val = None if node.next is None else node.next.val
output += str(node.val) + f'({next_val})'+ ", "
queue.append(node.left)
queue.append(node.right)
return "[" + output[:-2] + "]"
def main():
import sys
import io
s = '''[1,2,3,4,5,6,7]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
root = stringToTreeNode(line);
ret = Solution().connect(root)
out = treeNodeToString(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 118, https://leetcode.com/problems/pascals-triangle/
class Solution:
def generate(self, numRows: int) -> List[List[int]]:
dp = [[1] for _ in range(0, numRows)]
for i in range(1, numRows):
for j in range(1, i):
dp[i].append(dp[i-1][j-1] + dp[i-1][j])
dp[i].append(1)
return dp
def main():
import sys
import io
s = '''5
10'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
x = int(line);
ret = Solution().generate(x)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 121, https://leetcode.com/problems/best-time-to-buy-and-sell-stock/
class Solution:
def maxProfit(self, prices: List[int]) -> int:
n = len(prices)
if n < 2:
return 0
dp = [0 for i in range(0, n)] # max profit of selling on day
for i in range(1, n):
p = prices[i] - prices[i-1] # profit of single day
if dp[i-1] > 0:
dp[i] = dp[i-1] + p
else:
dp[i] = p
print(i, p, dp)
return max(dp)
def stringToIntegerList(input):
return json.loads(input)
def main():
import sys
import io
s = '''[7,1,5,3,6,4]
[7,6,4,3,1]
[7,2,5,3,6,4,1,2]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
prices = stringToIntegerList(line);
ret = Solution().maxProfit(prices)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 122, https://leetcode.com/problems/best-time-to-buy-and-sell-stock-ii/
class Solution:
def maxProfit(self, prices: List[int]) -> int:
mp = 0
for i in range(1, len(prices)):
p = prices[i] - prices[i-1]
if p > 0:
mp += p
return mp
def stringToIntegerList(input):
return json.loads(input)
def main():
import sys
import io
s = '''[7,1,5,3,6,4]
[1,2,3,4,5]
[7,6,4,3,1]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
prices = stringToIntegerList(line);
ret = Solution().maxProfit(prices)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 125, https://leetcode.com/problems/valid-palindrome/submissions/
class Solution:
def isPalindrome(self, s: str) -> bool:
if s == '':
return True
l = s.lower()
n = len(l)
p1, p2 = 0, n - 1
while p1 < p2:
while p1 < p2 and not l[p1].isalnum():
p1 += 1
while p1 < p2 and not l[p2].isalnum():
p2 -= 1
if l[p1] != l[p2]:
return False
p1 += 1
p2 -= 1
return True
def stringToString(input):
import json
return json.loads(input)
def main():
import sys
import io
s = '''"A man, a plan, a canal: Panama"
"race a car"
"0a"
""'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
s = stringToString(line);
ret = Solution().isPalindrome(s)
out = (ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 128, https://leetcode.com/problems/longest-consecutive-sequence/
class Solution:
def longestConsecutive(self, nums: List[int]) -> int:
if len(nums) == 0:
return 0
nums_sorted = sorted(list(set(nums)))
n = len(nums_sorted)
max_l, l = 1, 1
for i in range(1, n):
if nums_sorted[i] - nums_sorted[i-1] == 1:
l += 1
else:
max_l = max(max_l, l)
l = 1
return max(max_l, l)
def stringToIntegerList(input):
return json.loads(input)
def main():
import sys
import io
s ='''[100, 4, 200, 1, 3, 2]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
nums = stringToIntegerList(line);
ret = Solution().longestConsecutive(nums)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 141, https://leetcode.com/problems/linked-list-cycle/
# Definition for singly-linked list.
class ListNode:
def __init__(self, x):
self.val = x
self.next = None
class Solution:
def hasCycle(self, head: ListNode) -> bool:
if head is None or head.next is None:
return False
p, q = head, head.next
while True:
p = p.next
if q.next is None or q.next.next is None:
return False
q = q.next.next
if p.val == q.val:
return True
return True
def stringToIntegerList(input):
return json.loads(input)
def stringToListNode(input):
# Generate list from the input
numbers = stringToIntegerList(input)
# Now convert that list into linked list
dummyRoot = ListNode(0)
ptr = dummyRoot
for number in numbers:
ptr.next = ListNode(number)
ptr = ptr.next
ptr = dummyRoot.next
return ptr
def cycle(head, pos):
if pos < 0:
return
cycle_node = head
while pos > 0:
cycle_node = cycle_node.next
pos -= 1
tail = head
while tail.next != None:
tail = tail.next
tail.next = cycle_node
def main():
import sys
import io
s = '''[3,2,0,-4]
1
[1,2]
0
[1]
-1'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
head = stringToListNode(line);
line = next(lines)
pos = int(line);
cycle(head, pos)
ret = Solution().hasCycle(head)
out = (ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 143, https://leetcode.com/problems/reorder-list/
# Definition for singly-linked list.
class ListNode:
def __init__(self, val=0, next=None):
self.val = val
self.next = next
class Solution:
def reorderList(self, head: ListNode) -> None:
"""
Do not return anything, modify head in-place instead.
"""
if head is None or head.next is None or head.next.next is None:
return
slow, fast = head, head
while fast and fast.next:
slow = slow.next
fast = fast.next.next
nodes_to_reverse, nodes_reversed = slow.next, None
slow.next = None
while nodes_to_reverse:
next_reverse = nodes_to_reverse.next
nodes_to_reverse.next = nodes_reversed
nodes_reversed = nodes_to_reverse
nodes_to_reverse = next_reverse
print('head', listNodeToString(head), 'tail', listNodeToString(nodes_reversed))
p, q = head, nodes_reversed
while q:
p_next, q_next = p.next, q.next
p.next = q
q.next = p_next
p = p_next
q = q_next
return
def stringToIntegerList(input):
return json.loads(input)
def stringToListNode(input):
# Generate list from the input
numbers = stringToIntegerList(input)
# Now convert that list into linked list
dummyRoot = ListNode(0)
ptr = dummyRoot
for number in numbers:
ptr.next = ListNode(number)
ptr = ptr.next
ptr = dummyRoot.next
return ptr
def listNodeToString(node):
if not node:
return "[]"
result = ""
while node:
result += str(node.val) + ", "
node = node.next
return "[" + result[:-2] + "]"
def main():
import sys
import io
s = '''[1,2,3]
[1,2,3,4]
[1,2,3,4,5]
[1,2,3,4,5,6]
[1,2,3,4,5,6,7]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
head = stringToListNode(line);
ret = Solution().reorderList(head)
out = listNodeToString(head)
if ret is not None:
print("Do not return anything, modify head in-place instead.")
else:
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 146, https://leetcode.com/problems/lru-cache/
class LRUCache:
def __init__(self, capacity: int):
self.cache = {}
self.capacity = capacity
self.indices = []
self.NOT_FOUND = -1
def get(self, key: int) -> int:
print('key', key, 'cache', self.cache, 'indices', self.indices)
if key in self.cache:
# update to latest
self.indices.remove(key)
self.indices.append(key)
return self.cache[key]
return self.NOT_FOUND
def put(self, key: int, value: int) -> None:
print('key', key, 'cache', self.cache, 'indices', self.indices)
if key in self.cache:
print('existing')
self.indices.remove(key)
self.indices.append(key)
elif len(self.indices) < self.capacity:
print('new')
self.indices.append(key)
else:
print('replace')
# remove earliest
earliest = self.indices[0]
del self.indices[0]
del self.cache[earliest]
self.indices.append(key)
self.cache[key] = value
cache = LRUCache(2);
cache.put(1, 1);
cache.put(2, 2);
print('get:', cache.get(1));
cache.put(3, 3);
print('get:', cache.get(2));
cache.put(4, 4);
print('get:', cache.get(1));
print('get:', cache.get(3));
print('get:', cache.get(4));
# +
# Problem 146, https://leetcode.com/problems/lru-cache/
class Node:
def __init__(self, key, val):
self.key = key
self.val = val
self.prev = None
self.next = None
class LRUCache:
def __init__(self, capacity: int):
self.capacity = capacity
self.NOT_FOUND = -1
self.cache = {}
self.used = 0
self.index_head = Node('head', None)
self.index_tail = Node('tail', None)
self.index_head.next = self.index_tail
self.index_tail.prev = self.index_head
def get(self, key: int) -> int:
head, p, tail, q = [], self.index_head.next, [], self.index_tail.prev
while p.key != 'tail':
head.append(p.val)
p = p.next
while q.key != 'head':
tail.append(q.val)
q = q.prev
print('head to tail', head, 'tail to head', tail)
if key in self.cache:
node = self.cache[key]
# remove
node_prev = node.prev
node_next = node.next
node_prev.next = node_next
node_next.prev = node_prev
# insert to tail
node.prev = self.index_tail.prev
node.next = self.index_tail
node.prev.next = node
node.next.prev = node
return node.val
return self.NOT_FOUND
def put(self, key: int, value: int) -> None:
head, p, tail, q = [], self.index_head.next, [], self.index_tail.prev
while p.key != 'tail':
head.append(p.val)
p = p.next
while q.key != 'head':
tail.append(q.val)
q = q.prev
print('head to tail', head, 'tail to head', tail)
if key in self.cache:
self.cache[key].val = value
self.get(key)
elif self.used < self.capacity:
print('new')
self.used += 1
# insert to tail
node = Node(key, value)
node.prev = self.index_tail.prev
node.next = self.index_tail
print(node.prev.key, node.next.key)
node.prev.next = node
node.next.prev = node
self.cache[key] = node
else:
print('replace')
# remove earliest
earliest = self.index_head.next
del self.cache[earliest.key]
earliest.next.prev = earliest.prev
earliest.prev.next = earliest.next
# insert to tail
node = Node(key, value)
node.prev = self.index_tail.prev
node.next = self.index_tail
node.prev.next = node
node.next.prev = node
self.cache[key] = node
cache = LRUCache(3);
cache.put(1, 1);
cache.put(2, 2);
cache.put(3, 3);
cache.put(4, 4);
print('get:', cache.get(4));
print('get:', cache.get(3));
print('get:', cache.get(2));
print('get:', cache.get(1));
cache.put(5, 5);
print('get:', cache.get(1));
print('get:', cache.get(2));
print('get:', cache.get(3));
print('get:', cache.get(4));
print('get:', cache.get(5));
# +
# Problem 148, https://leetcode.com/problems/sort-list/
# Definition for singly-linked list.
class ListNode:
def __init__(self, val=0, next=None):
self.val = val
self.next = next
class Solution:
def sortList(self, head: ListNode) -> ListNode:
if head is None:
# len: 0
return head
if head.next is None:
# len: 1
return head
if head.next.next is None:
# len: 2
if head.val < head.next.val:
return head
else:
new_head = head.next
new_head.next = head
head.next = None
return new_head
# find middle
p, q = head, head
while q.next and q.next.next:
p = p.next
q = q.next.next
# sort two parts
second_part = self.sortList(p.next)
p.next = None
first_part = self.sortList(head)
if first_part.val > second_part.val:
first_part, second_part = second_part, first_part
print('sorted first part:', listNodeToString(first_part))
print('sorted second part:', listNodeToString(second_part))
# merge
p, q, r = first_part, first_part.next, second_part
while q and r:
if q.val < r.val:
p.next = q
p = p.next
q = q.next
else:
p.next = r
p = p.next
r = r.next
# concat rest
if q is None:
p.next = r
elif r is None:
p.next = q
return first_part
def stringToIntegerList(input):
return json.loads(input)
def stringToListNode(input):
# Generate list from the input
numbers = stringToIntegerList(input)
# Now convert that list into linked list
dummyRoot = ListNode(0)
ptr = dummyRoot
for number in numbers:
ptr.next = ListNode(number)
ptr = ptr.next
ptr = dummyRoot.next
return ptr
def listNodeToString(node):
if not node:
return "[]"
result = ""
while node:
result += str(node.val) + ", "
node = node.next
return "[" + result[:-2] + "]"
def main():
import sys
import io
s = '''[4,2,1,3]
[-1,5,3,4,0]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
head = stringToListNode(line);
ret = Solution().sortList(head)
out = listNodeToString(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 189, https://leetcode.com/problems/rotate-array/
class Solution:
def rotate(self, nums: List[int], k: int) -> None:
"""
Do not return anything, modify nums in-place instead.
"""
n = len(nums)
if n < 2:
return
if k > n-1:
k %= n
print(k)
hcf, hcf_x = n, k # highest common factor
while hcf_x != 0:
t = hcf % hcf_x
hcf = hcf_x
hcf_x = t
print('hcf', hcf)
for i in range(0, hcf):
pos, val, tmp = i, nums[i], None
for j in range(0, (n // hcf) - 1):
next_pos = (pos+k) % n
tmp = nums[next_pos]
nums[next_pos] = val
print((i, j), (pos, next_pos), nums)
pos, val = next_pos, tmp
nums[i] = val
print(i, nums)
def stringToIntegerList(input):
return json.loads(input)
def integerListToString(nums, len_of_list=None):
if not len_of_list:
len_of_list = len(nums)
return json.dumps(nums[:len_of_list])
def main():
import sys
import io
s = '''[1,2,3,4,5,6,7]
3
[-1,-100,3,99]
2
[1,2,3,4,5,6,7,8,9,0]
4'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
nums = stringToIntegerList(line);
line = next(lines)
k = int(line);
ret = Solution().rotate(nums, k)
out = integerListToString(nums)
if ret is not None:
print("Do not return anything, modify nums in-place instead.")
else:
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# +
# Problem 198, https://leetcode.com/problems/house-robber/
class Solution:
def rob(self, nums: List[int]) -> int:
n = len(nums)
if n < 3:
if n == 0:
return 0
return max(nums)
dp = [0 for i in range(0, n+1)] # max when robbing house
dp[0], dp[1], dp[2] = 0, nums[0], nums[1]
for i in range(2, n):
dp[i+1] = max(dp[i-1], dp[i-2]) + nums[i]
print(dp)
return max(dp[n-1], dp[n])
def stringToIntegerList(input):
return json.loads(input)
def main():
import sys
import io
s = '''[1,2,3,1]
[2,7,9,3,1]
[1,9,1,1,9]'''
def readlines():
for line in s.splitlines():
yield line.strip('\n')
lines = readlines()
while True:
try:
line = next(lines)
nums = stringToIntegerList(line);
ret = Solution().rob(nums)
out = str(ret);
print(out)
except StopIteration:
break
if __name__ == '__main__':
main()
# -
| leetcode/InterviewII.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Part 3
#
# In this part, you will implement feature consistency analysis for model selection
import neuralflow
from neuralflow.utilities.FC_nonstationary import JS_divergence_tdp, FeatureComplexities, FeatureComplexity
import numpy as np
import matplotlib.pyplot as plt, matplotlib.gridspec as gridspec
# We generated 400 trials of data from the ramping dynamics, split it into to equal data samples and optimized a model on each of the datasample for 10000 GD iterations. Let us initialize an EnergyModel class instance and load the optimization results.
# +
EnergyModelParams = {'pde_solve_param':{'method':{'name': 'SEM', 'gridsize': {'Np': 8, 'Ne': 16}}},
'Nv': 111,
'peq_model':{'model': 'uniform', 'params': {}},
'p0_model': {'model': 'single_well', 'params': {'miu': 200, 'xmin': -0.3}},
'D0': 0.1,
'boundary_mode':'absorbing',
'num_neuron':1,
'firing_model':[{'model': 'linear','params': {'r_slope': 50, 'r_bias': 60}}],
'verbose':True
}
# An instance of EnergyModel class that will be used for FC analysis. Note that here we set peq to uniform to pretend
# that we do not know the correct model
em_fitting = neuralflow.EnergyModel(**EnergyModelParams)
# Load the optimization results
data1 = dict(np.load('data/Ex3_datasample1.npz',allow_pickle=True))
data2 = dict(np.load('data/Ex3_datasample2.npz',allow_pickle=True))
# Also calculate the ground-truth model for visual comparison
peq_gt = neuralflow.peq_models.linear_pot(em_fitting.x_d_,em_fitting.w_d_,slope=-5)
# -
# Before proceding to feature consistency analysis, let us visualize the results of the first optimization.
# Here we plot the potential function at selected iteration, and the negative loglikelihood as in the previous exercise. In addition to the training cost, we also plot validation cost (validation negative loglikelihood), which is calculated for each model produced by gradient descent on validation data (that was not used for training).
#
# You can see that both validation and training costs display long horizontal plateaus (after initial fast descent). Due to the nuances in the data, the minimum of the validation cost can be achieved anywhere on this plateau. However, as you can see in the picture, if the minimum is achieved on a late iteration, the corresponding potential function will be overfitted. Thus, we need an alternative method for model selection.
# +
lls=data1['logliks']
lls_CV=data1['logliksCV']
#Shift training and validated loglikelihoods such that they both start from 0 for visualisation purposes
lls= (lls-lls[0])
lls_CV=(lls_CV-lls_CV[0])
fig=plt.figure(figsize=(20,7))
gridspec.GridSpec(2,6)
Iterations =[1,13,50,1000,5000,10000]
Iterations_indices = np.argmin(np.abs(np.subtract.outer(data1['iter_num'],Iterations)),axis=0)
colors=[[0.0, 0.0, 1.0],
[0.2, 0.4, 0.8],
[0.4, 0.2, 0.6],
[0.6, 0.2, 0.4],
[0.8, 0.4, 0.2],
[1.0, 0.2, 0.0]]
# Plot negative loglikelihood vs. iteration number
plt.subplot2grid((2,6), (0,0), colspan=2, rowspan=2)
# Training negative loglikelihood
plt.plot(np.arange(1,lls.size+1),lls,color='black',linewidth=3,label='training')
# Validation negative loglikelihood
plt.plot(np.arange(1,lls_CV.size+1),lls_CV,color='red',linewidth=3, label='validation')
plt.xlabel('Iteration #', fontsize=18)
plt.ylabel(r'$-\log\mathcal{L}$', fontsize=18)
plt.legend()
#Plot potentials. Potential is calculated from peq by taking negative log: Phi = - log(peq).
for i,Iter in enumerate(Iterations):
plt.subplot2grid((2,6), (i//3,2+i%3))
plt.plot(em_fitting.x_d_,-np.log(data1['peqs'][...,Iterations_indices[i]]),color=colors[i],linewidth=3)
plt.plot(em_fitting.x_d_,-np.log(peq_gt),color='grey',linewidth=2)
plt.title(f'Iteration {Iter}')
# -
# Let us implement feature consistency analysis for model selection.
# +
#Only consider the models at selected iterations (arranged on a logspace between 0 and 10,000).
number_of_models = 100
iterations_selected=np.unique(np.concatenate((np.array([0]),np.logspace(0,4,number_of_models))).astype(int))
#Find the indices of the selected iterations in the data1/data2 arrays
iteration_indices = np.argmin(np.abs(np.subtract.outer(data1['iter_num'],iterations_selected)),axis=0)
# For each model calculate feature complexity, and the eigenvalues and eigenvectors of the operator H0
# using our utility function
FCs_array1, lQ_array1, Qx_array1=FeatureComplexities(data1,em_fitting,iteration_indices)
FCs_array2, lQ_array2, Qx_array2=FeatureComplexities(data2,em_fitting,iteration_indices)
# -
# Now calculate JS divergence at each level of feature complexity
# +
# This parameter will determine how many models from sequence 2 are compared with a model from sequence 1 for each
# level of feature complexity
FC_stride = 5
# Preallocate JS divergence array
JS = np.zeros(FCs_array1.size)
# For the second sequence, mid_inds2 will contain index of the model for each feature complexity in data2.
min_inds2 = np.zeros_like(JS)
for i in range(FCs_array1.size):
# We only consider 1 model from the sequence of models optimized on data sample 1.
FC1_ind = i
# Find the index of a model in the second sequence FCs_array2 that have the closest feature complexity
# to the model from the first sequence
ind_seq_2 = np.argmin(np.abs(FCs_array1[FC1_ind]-FCs_array2))
# Select the indices of 2*FC_stride-1 models from the second sequence of models around the index ind_seq_2
FC2_ind = np.array(np.arange(max(0,ind_seq_2-FC_stride),min(FCs_array2.size-1,ind_seq_2+1+FC_stride)))
#Compute JSD for each pair of models selected from the sequences 1 and 2.
JS_cur = np.zeros(FC2_ind.size)
for i2, ind2 in enumerate(FC2_ind):
peq_ind1=iteration_indices[FC1_ind]
peq_ind2=iteration_indices[ind2]
JS_cur[i2]=JS_divergence_tdp(data1['peqs'][...,peq_ind1],em_fitting.D_,em_fitting.p0_,
data2['peqs'][...,peq_ind2],em_fitting.D_,em_fitting.p0_,
em_fitting.w_d_, lQ_array1[FC1_ind,:], Qx_array1[FC1_ind,:],
lQ_array2[ind2,:], Qx_array2[ind2,:],1,10)
# Find the index of model2 in data2
m2 = np.argmin(JS_cur)
min_inds2[i] = iteration_indices[FC2_ind[m2]]
# JS[i] should be equal to the minimal value of JS_cur
######INSERT YOUR CODE HERE############
JS[i] = np.min(JS_cur)
#######################################
# For the first sequence of model, we only consider a single model for each feature complexity
min_inds1 = iteration_indices[np.arange(FCs_array1.size)]
# -
# Now threshold FC divergence and plot the results
# +
#JS divergence threshold
JS_thres=0.001
#Index of the optimal model
optimal_ind=np.where(JS>JS_thres)[0][0]-1
#Optimal FC
FC_opt = FCs_array1[optimal_ind]
#For illustration purposes, choose early and late FCs.
FC_late=FC_opt + 1
late_ind = np.where(FCs_array1>FC_late)[0][0]
FC_early=FC_opt - 1
early_ind = np.where(FCs_array1>FC_early)[0][0]
#Create a ground-truth model, calculate its Feature Complexity.
EnergyModelParams = {'pde_solve_param':{'method':{'name': 'SEM', 'gridsize': {'Np': 8, 'Ne': 16}}},
'Nv': 111,
'peq_model':{"model": "linear_pot", "params": {"slope": -5}},
'p0_model': {'model': 'single_well', 'params': {'miu': 200, 'xmin': -0.3}},
'D0': 0.1,
'boundary_mode':'absorbing',
'num_neuron':1,
'firing_model':[{'model': 'linear','params': {'r_slope': 50, 'r_bias': 60}}],
'verbose':True
}
em_gt = neuralflow.EnergyModel(**EnergyModelParams)
FC_gt = FeatureComplexity(em_gt)
#Visualization
fig=plt.figure(figsize=(20,7));
gs=gridspec.GridSpec(2,6,height_ratios=[3,2],hspace=0.5, wspace=0.5);
line_colors = [[239/255, 48/255, 84/255], [0, 127/255, 1], [0.5, 0.5, 0.5]]
dot_colors = [[0.6,0.6,0.6], [1, 169/255, 135/255], [147/255, 192/255, 164/255]]
ax = plt.subplot(gs[0,:3])
ax.set_title('Feature complexity grows with iteration number',fontsize=18)
ax.plot(data1['iter_num'][iteration_indices],FCs_array1,color=line_colors[0],linewidth=3,label='Data sample 1')
ax.plot(data2['iter_num'][iteration_indices],FCs_array2,color=line_colors[1],linewidth=3,label='Data sample 2')
ax.hlines(FC_gt,data1['iter_num'][iteration_indices][0],data1['iter_num'][iteration_indices][-1],color=line_colors[2],linewidth=2,label='Ground truth')
plt.xscale('log')
plt.xlabel('Iteration number', fontsize=14)
plt.ylabel('Feature complexity', fontsize=14)
ax=plt.subplot(gs[0,3:])
ax.set_title('JS divergence as a function of feature complexity',fontsize=18)
ax.plot(FCs_array1,JS, color = [0.47, 0.34, 0.66],linewidth=3)
ax.plot(FC_early,JS[np.argmin(np.abs(FCs_array1-FC_early))],'.',markersize=20,color=dot_colors[0])
ax.plot(FC_opt,JS[np.argmin(np.abs(FCs_array1-FC_opt))],'.',markersize=20,color=dot_colors[1])
ax.plot(FC_late,JS[np.argmin(np.abs(FCs_array1-FC_late))],'.',markersize=20,color=dot_colors[2])
ax.hlines(JS_thres,FCs_array1[0],FCs_array1[-1],linestyles='dashed',color=line_colors[2],linewidth=2,label='Ground truth')
plt.xlabel('Feature complexity',fontsize=14)
plt.ylabel('JS divergence', fontsize=14)
ax=plt.subplot(gs[1,:2])
ax.set_title(r'$\mathcal{M}<\mathcal{M}^*$', fontsize=18)
ax.plot(em_gt.x_d_,-np.log(data1['peqs'][...,int(min_inds1[early_ind])]),color=line_colors[0],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(data2['peqs'][...,int(min_inds2[early_ind])]),color=line_colors[1],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(em_gt.peq_),color=[0.5, 0.5, 0.5],linewidth=2)
plt.xlabel(r'Latent state, $x$', fontsize=14)
plt.ylabel(r'Potential, $\Phi(x)$', fontsize=14)
ax=plt.subplot(gs[1,2:4])
ax.set_title(r'$\mathcal{M}=\mathcal{M}^*$', fontsize=18)
ax.plot(em_gt.x_d_,-np.log(data1['peqs'][...,int(min_inds1[optimal_ind])]),color=line_colors[0],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(data2['peqs'][...,int(min_inds2[optimal_ind])]),color=line_colors[1],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(em_gt.peq_),color=[0.5, 0.5, 0.5],linewidth=2)
plt.xlabel('latent state, x', fontsize=14)
plt.ylabel(r'Potential, $\Phi(x)$', fontsize=14)
ax=plt.subplot(gs[1,4:])
ax.set_title(r'$\mathcal{M}>\mathcal{M}^*$', fontsize=18)
ax.plot(em_gt.x_d_,-np.log(data1['peqs'][...,int(min_inds1[late_ind])]),color=line_colors[0],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(data2['peqs'][...,int(min_inds2[late_ind])]),color=line_colors[1],linewidth=3)
ax.plot(em_gt.x_d_,-np.log(em_gt.peq_),color=[0.5, 0.5, 0.5],linewidth=2)
plt.xlabel('latent state, x', fontsize=14)
plt.ylabel(r'Potential, $\Phi(x)$', fontsize=14)
| Tutorial/CCN2021/Completed/Part3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Entropic Portfolio Optimization
#
# In this notebook we show how to use the exponential cone to model the perspective of the `log_sum_exp` function and its application in portfolio optimization.
#
# ## 1. Entropic Value at Risk Optimization
#
# ### 1.1 The Entropic Value at Risk
#
# The Entropic Value at Risk (EVaR) which is a new risk measure introduced by __[Ahmadi-Javid (2012)](https://link.springer.com/article/10.1007/s10957-011-9968-2?r=1&l=ri&fst=0&error=cookies_not_supported&code=ccfb8a5e-692b-43d1-b76e-ae596c7f0bed)__. It is the upper bound based on Chernoff Inequality of Value at Risk (VaR) and Conditional Value at Risk (CVaR), formally it is defined as:
#
# $$
# \text{EVaR}_{\alpha}(X) = \inf_{z>0} \left \{z\log \left ( \frac{1}{\alpha} M_{X} (\frac{1}{z}) \right ) \right \}
# $$
#
# Where $M_{X} (t) = \text{E} [e^{tX}]$ is the moment generating function and $\alpha \in [0,1]$ is the significance level.
#
# ### 1.2 EVaR Minimization
#
# To discretize the EVaR we need the perspective of the `log_sum_exp` function, we can do this using the exponential cone in CVXPY. The discipined convex programming (DCP) problem of EVaR minimization was proposed by __[Cajas (2021)](https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3792520)__ and it is posed as:
#
# $$
# \begin{equation}
# \begin{aligned}
# & \underset{x, \, z, \, t, \, u}{\text{min}} & & t + z \ln \left ( \frac{1}{\alpha T} \right ) \\
# & \text{s.t.} & & \mu x^{\tau} \geq \bar{\mu} \\
# & & & \sum_{i=1}^{N} x_i = 1 \\
# & & & z \geq \sum^{T}_{j=1} u_{j} \\
# & & & (-r_{j}x^{\tau}-t, z, u_{j}) \in K_{\text{exp}} \; \forall \; j=1, \ldots, T \\
# & & & x_i \geq 0 \; ; \; \forall \; i =1, \ldots, N \\
# \end{aligned}
# \end{equation}
# $$
#
# Where $t$ is an auxiliar variable that represents the perspectives of the `log_sum_exp` function, $z$ is the factor of perspective function, $u_{j}$ is an auxiliary variable, $x$ are the weights of assets, $\mu$ is the mean vector of expected returns, $\bar{\mu}$ the minimum expected return of portfolio, $K_{\text{exp}}$ is an exponential cone and $r$ is the matrix of observed returns.
# +
####################################
# Downloading Data
####################################
# !pip install --quiet yfinance
import numpy as np
import pandas as pd
import yfinance as yf
import warnings
warnings.filterwarnings("ignore")
yf.pdr_override()
pd.options.display.float_format = '{:.4%}'.format
# Date range
start = '2016-01-01'
end = '2019-12-30'
# Tickers of assets
assets = ['TGT', 'CMCSA', 'CPB', 'MO', 'T', 'BAX', 'BMY',
'MSFT', 'SEE', 'VZ', 'CNP', 'NI', 'GE', 'GOOG']
assets.sort()
# Downloading data
data = yf.download(assets, start = start, end = end)
data = data.loc[:,('Adj Close', slice(None))]
data.columns = assets
# Calculating returns
Y = data[assets].pct_change().dropna()
display(Y.head())
# +
####################################
# Finding the Min EVaR Portfolio
####################################
import cvxpy as cp
# Defining initial inputs
mu = Y.mean().to_numpy().reshape(1,-1)
returns = Y.to_numpy()
n = returns.shape[0]
# Defining initial variables
w = cp.Variable((mu.shape[1], 1))
alpha = 0.05
ret = mu @ w
X = returns @ w
# Entropic Value at Risk Model Variables
t = cp.Variable((1, 1))
z = cp.Variable((1, 1), nonneg=True)
ui = cp.Variable((n, 1))
constraints = [cp.sum(ui) <= z,
cp.constraints.ExpCone(-X - t, np.ones((n, 1)) @ z, ui)] # Exponential cone constraint
# Budget and weights constraints
constraints += [cp.sum(w) == 1,
w >= 0]
# Defining risk objective
risk = t + z * np.log(1 / (alpha * n))
objective = cp.Minimize(risk)
# Solving problem
prob = cp.Problem(objective, constraints)
prob.solve()
# Showing Optimal Weights
weights = pd.DataFrame(w.value, index=assets, columns=['Weights'])
display(weights)
# -
# ## 2. Entropic Drawdown at Risk Optimization
#
# ### 2.1 The Entropic Drawdown at Risk
#
# The Entropic Drawdown at Risk (EDaR) which is a new risk measure introduced by __[Cajas (2021)](https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3792520)__. It is the upper bound based on Chernoff Inequality of Drawdown at Risk (DaR) and Conditional Drawdown at Risk (CDaR), formally it is defined as:
#
# $$
# \begin{equation}
# \begin{aligned}
# \text{EDaR}_{\alpha}(X) & = \text{EVaR}_{\alpha}(\text{DD}(X)) \\
# \text{EDaR}_{\alpha}(X) & = \inf_{z>0} \left \{ z \ln \left (\frac{1}{\alpha}M_{\text{DD}(X)} \left (\frac{1}{z} \right ) \right ) \right \} \\
# \end{aligned}
# \end{equation}
# $$
#
# where $M_{X}(t)$ is the moment generating function of $t$, $\alpha \in [0,1]$ is the significance level and $\text{DD}(X)$ is the drawdown of $X$.
#
# ### 2.2 Maximization of Return/EDaR ratio
#
# This problem is a __[linear fractional programming](https://en.wikipedia.org/wiki/Linear-fractional_programming)__ problem and can be converted to a DCP problem using __Charnes and Cooper transformation__. The DCP problem of maximization of return EDaR ratio was proposed by __[Cajas (2021)](https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3792520)__ and it is posed as:
#
# $$
# \begin{equation}
# \begin{aligned}
# & \underset{y, \, k, \, z, \, t, \, u, \, d}{\text{min}} & & t + z \ln \left ( \frac{1}{\alpha T} \right )\\
# & \text{s.t.} & & \mu y^{\tau} - r_{f} k= 1 \\
# & & & \sum_{i=1}^{N} y_{i} = k \\
# & & & z \geq \sum^{T}_{j=1} u_{j} \\
# & & & (d_{j} - R_{j} y^{\tau} - t, z, u_{j}) \in K_{\text{exp}} \; \forall \; j =1, \ldots, T \\
# & & & d_{j} \geq R_{j} y^{\tau} \; \forall \; j=1, \ldots, T \\
# & & & d_{j} \geq d_{j-1} \; \forall \; j=1, \ldots, T \\
# & & & d_{j} \geq 0 \; \forall \; j=1, \ldots, T \\
# & & & d_{0} = 0 \\
# & & & k \geq 0 \\
# & & & y_{i} \geq 0 \; ; \; \forall \; i =1, \ldots, N \\
# \end{aligned}
# \end{equation}
# $$
#
# where $R_{j} x^{\tau} = \sum^{j}_{i=1} r_{i} x^{\tau}$, $d_{j}$ is a variable that represents the uncompounded cumulative return of the portfolio and $r_{f}$ is the risk free rate.
#
# Finally, the optimal portfolio is obtained making the transformation $x = y / k$.
# +
#######################################
# Finding the max return/EDaR Portfolio
#######################################
# Defining initial inputs
mu = Y.mean().to_numpy().reshape(1,-1)
returns = Y.to_numpy()
nav = Y.cumsum().to_numpy()
n = returns.shape[0]
# Defining initial variables
w = cp.Variable((mu.shape[1], 1))
k = cp.Variable((1, 1))
rf0 = 0
alpha = 0.05
ret = mu @ w
X1 = nav @ w
# Drawdown Variables
d = cp.Variable((nav.shape[0] + 1, 1))
constraints = [d[1:] >= X1,
d[1:] >= d[:-1],
d[1:] >= 0,
d[0] == 0]
# Entropic Drawdown at Risk Model Variables
t = cp.Variable((1, 1))
z = cp.Variable((1, 1), nonneg=True)
ui = cp.Variable((n, 1))
constraints += [cp.sum(ui) <= z,
cp.constraints.ExpCone(d[1:] - X1 - t, np.ones((n, 1)) @ z, ui)] # Exponential cone constraint
# Budget and weights constraints
constraints += [cp.sum(w) == k,
ret - rf0 * k == 1,
w >= 0,
k >= 0]
# Defining risk objective
risk = t + z * np.log(1 / (alpha * n))
objective = cp.Minimize(risk)
# Solving problem
prob = cp.Problem(objective, constraints)
prob.solve()
# Showing Optimal Weights
weights = pd.DataFrame(w.value/k.value, index=assets, columns=['Weights'])
display(weights)
# -
# For more portfolio optimization models and applications, you can see the CVXPY based library __[Riskfolio-Lib](https://github.com/dcajasn/Riskfolio-Lib)__.
| examples/notebooks/WWW/Entropic Portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ashesm/ISYS5002_PORTFOLIO1/blob/main/Business_Buzzword_Generator_STUDENTS.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="f8b21259-c1a0-4ecb-a02f-e35565841067"
# # Business Buzzword Generator
#
# Lets write a real-world business application using Python. Want to write a program to genrate business phrases. The program takes three lists of words, randomly picks one word from each list, combines the words into a phrase, and then prints out the phrase. Here is the psuedocode:
#
# importing the random module
# make three lists, one of buzzword, one of actions, and one of outcomes
# randomly choose one buzzword, action, and outcome from each list
# now build the phrase by "adding" the words together
# output the phrase
#
# A good use of pseudocode is each line becomes comments in the code. Each line of pseudo code has been pasted into a cell below. Try to implement each as python statement or statements.
# + id="6b9a2fc6-f8ae-4778-a5e7-9659136091fd"
# importing the random modul
import random
# + colab={"base_uri": "https://localhost:8080/"} id="52db4e24-c2e3-46f2-8a96-0fece1f437f2" outputId="9d430e86-dfaf-40ea-9552-6d64ce931198"
# make three lists, one of buzzwords, one of actions, and one of outcomes
verbs = ['actualize', 'administrate','aggregate']
adjectives = ['24/7, acurate', 'adaptive']
nouns = ['adoption', 'alignments','application']
verbs
# + colab={"base_uri": "https://localhost:8080/"} id="YkPcv1Ka8Kdv" outputId="9d896676-8a1b-452a-cf0d-98415ca0bed0"
# + id="6ND0fHTVxRtB"
# randomly choose one buzzword, action, and outcome from each list
verb = random.choice (verbs)
adjective = random.choice(adjectives)
noun = random.choice(nouns)
# + id="95c40fea-40d0-49e6-afe5-c181da731368"
# build the phrase by "adding" the words together
phrase = verb + '' + adjective + '' + noun
# + id="3e36c71d-ef4b-40d5-ac7f-a7717460e28f"
# output the phrase
# + colab={"base_uri": "https://localhost:8080/"} id="71838d24-05d5-44cb-ae5a-1195fd805622" outputId="58e7a381-7f88-4d82-b7b9-e210b5985d8b"
print(phrase)
| Business_Buzzword_Generator_STUDENTS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="fQhSr_LqSIg4" colab_type="code" colab={}
# !wget -q https://l1nna.com/372/Assignment/A2-3/train.csv
# !wget -q https://l1nna.com/372/Assignment/A2-3/test.csv
# + id="uaABvQpluwjs" colab_type="code" colab={}
import pandas as pd
import csv
xy_train_df = pd.read_csv('train.csv')
x_test_df = pd.read_csv('test.csv', index_col='id')
xy_train_df['length'] = xy_train_df.apply(lambda x: len(x.review), axis=1)
xy_train_df = xy_train_df.sort_values('length')
xy_train_df
# + id="dHl0DGCyvA7l" colab_type="code" colab={}
from tensorflow.keras.preprocessing.text import Tokenizer
from sklearn.model_selection import train_test_split
vocab_size = 10000
max_len = 256
xy_train, xy_validation = train_test_split(xy_train_df, test_size=0.2)
# build vocabulary from training set
tokenizer = Tokenizer(num_words=vocab_size)
tokenizer.fit_on_texts(xy_train.review)
# padding is done inside:
x_train = tokenizer.texts_to_matrix(xy_train.review, mode='binary')[:, :max_len]
y_train = xy_train.rating
x_valid = tokenizer.texts_to_matrix(xy_validation.review, mode='binary')[:, :max_len]
y_valid = xy_validation.rating
x_test = tokenizer.texts_to_matrix(x_test_df.review, mode='binary')[:, :max_len]
print(x_train.shape)
print(x_valid.shape)
print(x_test.shape)
# + id="_EtEv2RivFKP" colab_type="code" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import collections
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow import keras
import tensorflow as tf
from tensorflow.keras.optimizers import Adam
model = keras.Sequential()
model.add(keras.layers.Embedding(vocab_size, 20))
model.add(keras.layers.CuDNNGRU(100))
model.add(keras.layers.Dense(1, activation=tf.nn.sigmoid))
model.compile(
optimizer=Adam(clipnorm=4.),
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(x_train,
y_train,
epochs=15,
batch_size=64,
validation_data=(x_valid, y_valid),
verbose=1)
# + id="z_naN0rn3OlN" colab_type="code" colab={}
model.evaluate(x_valid, y_valid)
# + id="Byj-weFZvaaS" colab_type="code" colab={}
y_predict = np.squeeze(model.predict_classes(x_valid))
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix
print(f1_score(y_valid, y_predict, average='micro'))
# + id="8FDSaQaxvG0q" colab_type="code" colab={}
# run on testing set:
y_predict = np.squeeze(model.predict_classes(x_test))
pd.DataFrame(
{'id': x_test_df.index,
'rating':y_predict}).to_csv('sample_submission.csv', index=False)
| A3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to the Interstellar Medium
# ### <NAME>
# ### Figure 6.15: simple model of heating and cooling in an HII region
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker
# %matplotlib inline
# +
# scale wavelengths to energy via Hydrogen
E_IP = 13.6 # eV
lambda_IP = 91.2 # nm
#------------- OII --------------
E21_O2 = E_IP * lambda_IP / 372.9
A21_O2 = 3.6e-5
Omega21_O2 = 0.80
E31_O2 = E_IP * lambda_IP / 372.6
A31_O2 = 1.8e-4
Omega31_O2 = 0.54
A32_O2 = 1.3e-7
#--------------------------------
#------------- OIII -------------
E31_O3 = E_IP * lambda_IP / 232.1
A31_O3 = 0.22
Omega31_O3 = 0.28
E32_O3 = E_IP * lambda_IP / 436.3
A32_O3 = 1.8
Omega32_O3 = 0.62
E21_O3 = E_IP * lambda_IP / 500.7
A21_O3 = 2.0e-2 + 6.7e-3
Omega21_O3 = 2.17
# fine structure
E31_O3fine = E_IP * lambda_IP / (32.7 * 1000)
A31_O3fine = 3.0e-11
Omega31_O3fine = 0.27
E32_O3fine = E_IP * lambda_IP / (51.8 * 1000)
A32_O3fine = 9.8e-5
Omega32_O3fine = 1.29
E21_O3fine = E_IP * lambda_IP / (88.4 * 1000)
A21_O3fine = 2.6e-5
Omega21_O3fine = 0.54
#--------------------------------
k = 8.617e-5 # eV/K
X_O = 5.4e-4 # Oxygen abundance relative to Hydrogen (Sun)
X_O = 3e-4 # Oxygen abundance relative to Hydrogen (ISM)
X_O *= 0.25 # assume 25% is O2, 25% is O3
# 50% unaccounted (kludge to make heating = cooling at ~8000 K)
nsteps = 30
Te0 = 2e3
Te1 = 15e3
Te = Te0 + (Te1-Te0)*np.arange(nsteps)/(nsteps-1)
heating = np.zeros(nsteps)
cooling_O2 = np.zeros(nsteps)
cooling_O3 = np.zeros(nsteps)
cooling_O3fine = np.zeros(nsteps)
for j,Te1 in enumerate(Te):
# OII
gamma12_O2 = 8.63e-12 * Omega21_O2 * np.exp(-E21_O2/(k*Te1)) / Te1**0.5
gamma13_O2 = 8.63e-12 * Omega31_O2 * np.exp(-E31_O2/(k*Te1)) / Te1**0.5
n1 = X_O
n3 = n1 * gamma13_O2 / (A31_O2 + A32_O2)
n2 = n1 * gamma12_O2 / A21_O2 + n3 * A32_O2 / A21_O2
I31 = n3 * A31_O2 * E31_O2
I21 = n2 * A21_O2 * E21_O2
cooling_O2[j] = I31 + I21
# OIII
gamma12_O3 = 8.63e-12 * Omega21_O3 * np.exp(-E21_O3/(k*Te1)) / Te1**0.5
gamma13_O3 = 8.63e-12 * Omega31_O3 * np.exp(-E31_O3/(k*Te1)) / Te1**0.5
n1 = X_O
n3 = n1 * gamma13_O3 / (A31_O3 + A32_O3)
n2 = n1 * gamma12_O3 / A21_O3 + n3 * A32_O3 / A21_O3
I31 = n3 * A31_O3 * E31_O3
I32 = n3 * A32_O3 * E32_O3
I21 = n2 * A21_O3 * E21_O3
cooling_O3[j] = I31 + I32 + I21
# OIII fine structure
gamma12_O3fine = 8.63e-12 * Omega21_O3fine * np.exp(-E21_O3fine/(k*Te1)) / Te1**0.5
gamma13_O3fine = 8.63e-12 * Omega31_O3fine * np.exp(-E31_O3fine/(k*Te1)) / Te1**0.5
n1 = X_O
n3 = n1 * gamma13_O3fine / (A31_O3fine + A32_O3fine)
n2 = n1 * gamma12_O3fine / A21_O3fine + n3 * A32_O3fine / A21_O3fine
I31 = n3 * A31_O3fine * E31_O3fine
I32 = n3 * A32_O3fine * E32_O3fine
I21 = n2 * A21_O3fine * E21_O3fine
cooling_O3fine[j] = I31 + I32 + I21
# heating per unit vol (divided by ne**2)
alpha2 = 2.6e-19 * (1e4/Te1)**0.85
E_ave = 5.0 # eV per ionization for 40000 K star (from planck_average.py)
heating[j] = alpha2 * E_ave
#print("{0:6g} {1} {2} {3}".format(Te, cooling_O2, cooling_O3, heating))
fig = plt.figure(figsize=(6,4))
ax = fig.add_subplot(111)
ax.set_xlim(4000,12000.0)
ax.set_ylim(5e-20,1e-17)
ax.set_yscale("log", nonposy='clip')
ax.set_xlabel(r'$T_e\ {\rm (K)}$', fontsize=14)
ax.set_ylabel(r'$\Gamma, \Lambda\ \ \rm{(eV\ m^3\ s^{-1})}$', fontsize=14)
x_labels = ['4000','6000','8000','10000','12000']
x_loc = np.array([float(x) for x in x_labels])
ax.set_xticks(x_loc)
ax.set_xticklabels(x_labels)
ax.plot(Te, heating, 'k-', lw=2)
ax.plot(Te, cooling_O2, 'k--', lw=1)
ax.plot(Te, cooling_O3, 'k-.', lw=1)
ax.plot(Te, cooling_O3fine, 'k-.', lw=1)
ax.plot(Te, cooling_O2 + cooling_O3 + cooling_O3fine, 'k-', lw=2)
#ax.plot([8000,8000], [1e-20,1e-16], 'k-', lw=0.5, alpha=0.5)
ax.text(4500, 2.45e-18, 'Heating', fontsize=10, rotation=-9)
ax.text(10600, 3.6e-18, 'Cooling', fontsize=10, rotation=10)
ax.text(8500, 2.6e-19, 'OII', fontsize=10, rotation=20)
ax.text(6300, 3.7e-19, 'OIII (optical)', fontsize=10, rotation=27)
ax.text(8500, 0.85e-19, 'OIII (infrared)', fontsize=10, rotation=-2)
fig.tight_layout()
plt.savefig('heating_cooling.pdf')
# -
| ionized/.ipynb_checkpoints/heating_cooling-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import numpy as np
import pandas as pd
import math
# +
pathToFile = '/mnt/home/jbielecki1/NEMA/'
dataParts = []
fileName = 'NEMA_IQ_384str_N0_1000_COINCIDENCES_PREPARED_part'
for i in range(8):
dataParts = dataParts.append(pickle.load(open(pathToFile + fileName + '0' + str(i+1), 'rb')))
for i in range(8):
dataParts = dataParts.append(pickle.load(open(pathToFile + fileName + '1' + str(i), 'rb')))
data = pd.concat(dataParts)
# -
dataPositive = data[data['class'] == 1]
dataRec = dataPositive.iloc[:,:16]
dataRec.to_csv(pathToFile + 'posReconstruction', sep = "\t", header = False, index = False)
| Notebooks/Reconstrucion/.ipynb_checkpoints/posReconstruction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pymongo import MongoClient
client=MongoClient("mongodb+srv://luharukas:<EMAIL>/myFirstDatabase?retryWrites=true&w=majority")
db=client.get_database('Bloguer')
record=db.Login
record
record.count_documents({})
record.insert_one(new_user)
new_user={
'username':'1rf19cs052',
'password':'<PASSWORD>'
}
list(record.find())
record.find_one({'username':'1rf19cs052'})
update_user={
'password':'<PASSWORD>'
}
record.update_one({'username':'1rf19cs052'},{'$set':update_user})
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Importing Library
from blmarkdown import Learning, LearningData
# ### Variables Instantiation
learning_subject = 'PHP Basic'
author = '<NAME>'
# ### Implementing blmarkdown
learning = Learning(learning_subject, author)
# +
# Data1
data1 = LearningData('Operator')
data1.add('Addition')
data1.add('Subtraction')
data1.add('Multiplication')
data1.add('Division')
# Adding Data1 to Learning
learning.add(data1)
# +
# Data2
data2 = LearningData('Control Structure')
data2.add('Loop')
data2.add('Branching')
# Adding Data2 to Learning
learning.add(data2)
# -
# This function will open a new tab in your default browser for viewing the generated markdown
learning.preview()
# ### Get Learning Markdown
learning.markdown()
# ### Save Learning Markdown
learning.save_markdown()
| old_version/v2.0/examples/example_simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (reco_base)
# language: python
# name: reco_base
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Data transformation (collaborative filtering)
# It is usually observed in the real-world datasets that users may have different types of interactions with items. In addition, same types of interactions (e.g., click an item on the website, view a movie, etc.) may also appear more than once in the history. Given that this is a typical problem in practical recommendation system design, the notebook shares data transformation techniques that can be used for different scenarios.
#
# Specifically, the discussion in this notebook is only applicable to collaborative filtering algorithms.
# ## 0 Global settings
# +
# set the environment path to find Recommenders
import sys
import pandas as pd
import numpy as np
import datetime
import math
print("System version: {}".format(sys.version))
# -
# ## 1 Data creation
# Two dummy datasets are created to illustrate the ideas in the notebook.
# ### 1.1 Explicit feedback
#
# In the "explicit feedback" scenario, interactions between users and items are numerical / ordinal **ratings** or binary preferences such as **like** or **dislike**. These types of interactions are termed as *explicit feedback*.
#
# The following shows a dummy data for the explicit rating type of feedback. In the data,
# * There are 3 users whose IDs are 1, 2, 3.
# * There are 3 items whose IDs are 1, 2, 3.
# * Items are rated by users only once. So even when users interact with items at different timestamps, the ratings are kept the same. This is seen in some use cases such as movie recommendations, where users' ratings do not change dramatically over a short period of time.
# * Timestamps of when the ratings are given are also recorded.
data1 = pd.DataFrame({
"UserId": [1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3],
"ItemId": [1, 1, 2, 2, 2, 1, 2, 1, 2, 3, 3, 3, 3, 3, 1],
"Rating": [4, 4, 3, 3, 3, 4, 5, 4, 5, 5, 5, 5, 5, 5, 4],
"Timestamp": [
'2000-01-01', '2000-01-01', '2000-01-02', '2000-01-02', '2000-01-02',
'2000-01-01', '2000-01-01', '2000-01-03', '2000-01-03', '2000-01-03',
'2000-01-01', '2000-01-03', '2000-01-03', '2000-01-03', '2000-01-04'
]
})
data1
# ### 1.2 Implicit feedback
#
# Many times there are no explicit ratings or preferences given by users, that is, the interactions are usually implicit. For example, a user may puchase something on a website, click an item on a mobile app, or order food from a restaurant. This information may reflect users' preference towards the items in an **implicit** manner.
#
# As follows, a data set is created to illustrate the implicit feedback scenario.
#
# In the data,
# * There are 3 users whose IDs are 1, 2, 3.
# * There are 3 items whose IDs are 1, 2, 3.
# * There are no ratings or explicit feedback given by the users. Sometimes there are the types of events. In this dummy dataset, for illustration purposes, there are three types for the interactions between users and items, that is, **click**, **add** and **purchase**, meaning "click on the item", "add the item into cart" and "purchase the item", respectively.
# * Sometimes there is other contextual or associative information available for the types of interactions. E.g., "time-spent on visiting a site before clicking" etc. For simplicity, only the type of interactions is considered in this notebook.
# * The timestamp of each interaction is also given.
data2 = pd.DataFrame({
"UserId": [1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3],
"ItemId": [1, 1, 2, 2, 2, 1, 2, 1, 2, 3, 3, 3, 3, 3, 1],
"Type": [
'click', 'click', 'click', 'click', 'purchase',
'click', 'purchase', 'add', 'purchase', 'purchase',
'click', 'click', 'add', 'purchase', 'click'
],
"Timestamp": [
'2000-01-01', '2000-01-01', '2000-01-02', '2000-01-02', '2000-01-02',
'2000-01-01', '2000-01-01', '2000-01-03', '2000-01-03', '2000-01-03',
'2000-01-01', '2000-01-03', '2000-01-03', '2000-01-03', '2000-01-04'
]
})
data2
# ## 2 Data transformation
# Many collaborative filtering algorithms are built on a user-item sparse matrix. This requires that the input data for building the recommender should contain unique user-item pairs.
#
# For explicit feedback datasets, this can simply be done by deduplicating the repeated user-item-rating tuples.
data1 = data1.drop_duplicates()
data1
# In the implicit feedback use cases, there are several methods to perform the deduplication, depending on the requirements of the actual business user cases.
# ### 2.1 Data aggregation
#
# Usually, data is aggregated by user to generate some scores that represent preferences (in some algorithms like SAR, the score is called *affinity score*, for simplicity reason, hereafter the scores are termed as *affinity*).
#
# It is worth mentioning that in such case, the affinity scores are different from the ratings in the explicit data set, in terms of value distribution. This is usually termed as an [ordinal regression](https://en.wikipedia.org/wiki/Ordinal_regression) problem, which has been studied in [Koren's paper](https://pdfs.semanticscholar.org/934a/729409d6fbd9894a94d4af66bd82222b5515.pdf). In this case, the algorithm used for training a recommender should be carefully chosen to consider the distribution of the affinity scores rather than discrete integer values.
# #### 2.2.1 Count
#
# The most simple technique is to count times of interactions between user and item for producing affinity scores. The following shows the aggregation of counts of user-item interactions in `data2` regardless the interaction type.
data2_count = data2.groupby(['UserId', 'ItemId']).agg({'Timestamp': 'count'}).reset_index()
data2_count.columns = ['UserId', 'ItemId', 'Affinity']
data2_count
# #### 2.2.1 Weighted count
# It is useful to consider the types of different interactions as weights in the count aggregation. For example, assuming weights of the three differen types, "click", "add", and "purchase", are 1, 2, and 3, respectively. A weighted-count can be done as the following
# +
# Add column of weights
data2_w = data2.copy()
conditions = [
data2_w['Type'] == 'click',
data2_w['Type'] == 'add',
data2_w['Type'] == 'purchase'
]
choices = [1, 2, 3]
data2_w['Weight'] = np.select(conditions, choices, default='black')
# Convert to numeric type.
data2_w['Weight'] = pd.to_numeric(data2_w['Weight'])
# -
# Do count with weight.
data2_wcount = data2_w.groupby(['UserId', 'ItemId'])['Weight'].sum().reset_index()
data2_wcount.columns = ['UserId', 'ItemId', 'Affinity']
data2_wcount
# #### 2.2.2 Time dependent count
# In many scenarios, time dependency plays a critical role in preparing dataset for building a collaborative filtering model that captures user interests drift over time. One of the common techniques for achieving time dependent count is to add a time decay factor in the counting. This technique is used in [SAR](https://github.com/Microsoft/Recommenders/blob/master/notebooks/02_model/sar_deep_dive.ipynb). Formula for getting affinity score for each user-item pair is
#
# $$a_{ij}=\sum_k w_k \left(\frac{1}{2}\right)^{\frac{t_0-t_k}{T}} $$
#
# where $a_{ij}$ is the affinity score, $w_k$ is the interaction weight, $t_0$ is a reference time, $t_k$ is the timestamp for the $k$-th interaction, and $T$ is a hyperparameter that controls the speed of decay.
#
# The following shows how SAR applies time decay in aggregating counts for the implicit feedback scenario.
# In this case, we use 5 days as the half-life parameter, and use the latest time in the dataset as the time reference.
# +
T = 5
t_ref = pd.to_datetime(data2_w['Timestamp']).max()
# +
# Calculate the weighted count with time decay.
data2_w['Timedecay'] = data2_w.apply(
lambda x: x['Weight'] * np.power(0.5, (t_ref - pd.to_datetime(x['Timestamp'])).days / T),
axis=1
)
# -
data2_w
# Affinity scores of user-item pairs can be calculated then by summing the 'Timedecay' column values.
data2_wt = data2_w.groupby(['UserId', 'ItemId'])['Timedecay'].sum().reset_index()
data2_wt.columns = ['UserId', 'ItemId', 'Affinity']
data2_wt
# ### 2.2 Negative sampling
# The above aggregation is based on assumptions that user-item interactions can be interpreted as preferences by taking the factors like "number of interation times", "weights", "time decay", etc. Sometimes these assumptions are biased, and only the interactions themselves matter. That is, the original dataset with implicit interaction records can be binarized into one that has only 1 or 0, indicating if a user has interacted with an item, respectively.
#
# For example, the following generates data that contains existing interactions between users and items.
data2_b = data2[['UserId', 'ItemId']].copy()
data2_b['Feedback'] = 1
data2_b = data2_b.drop_duplicates()
data2_b
# "Negative sampling" is a technique that samples negative feedback. Similar to the aggregation techniques, negative feedback cna be defined differently in different scenarios. In this case, for example, we can regard the items that a user has not interacted as those that the user does not like. This may be a strong assumption in many user cases, but it is reasonable to build a model when the interaction times between user and item are not that many.
#
# The following shows that, on top of `data2_b`, there are another 2 negative samples are generated which are tagged with "0" in the "Feedback" column.
users = data2['UserId'].unique()
items = data2['ItemId'].unique()
# +
interaction_lst = []
for user in users:
for item in items:
interaction_lst.append([user, item, 0])
data_all = pd.DataFrame(data=interaction_lst, columns=["UserId", "ItemId", "FeedbackAll"])
# -
data_all
data2_ns = pd.merge(data_all, data2_b, on=['UserId', 'ItemId'], how='outer').fillna(0).drop('FeedbackAll', axis=1)
data2_ns
# Also note that sometimes the negative sampling may also impact the count-based aggregation scheme. That is, the count may start from 0 instead of 1, and 0 means there is no interaction between the user and item.
# # References
#
# 1. <NAME> *et al*, Neural Collaborative Filtering, WWW 2017.
# 2. <NAME> *et al*, Collaborative filtering for implicit feedback datasets, ICDM 2008.
# 3. Simple Algorithm for Recommendation (SAR). See notebook [sar_deep_dive.ipynb](../02_model_collaborative_filtering/sar_deep_dive.ipynb).
# 4. <NAME> and <NAME>, OrdRec: an ordinal model for predicting personalized item rating distributions, RecSys 2011.
| examples/01_prepare_data/data_transform.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#export
from fastai2.torch_basics import *
from fastai2.data.all import *
from fastai2.text.core import *
from nbdev.showdoc import *
# +
#default_exp text.data
#default_cls_lvl 3
# -
# # Text data
#
# > Functions and transforms to help gather text data in a `Datasets`
# ## Numericalizing
#export
def make_vocab(count, min_freq=3, max_vocab=60000):
"Create a vocab of `max_vocab` size from `Counter` `count` with items present more than `min_freq`"
vocab = [o for o,c in count.most_common(max_vocab) if c >= min_freq]
for o in reversed(defaults.text_spec_tok): #Make sure all special tokens are in the vocab
if o in vocab: vocab.remove(o)
vocab.insert(0, o)
vocab = vocab[:max_vocab]
return vocab + [f'xxfake' for i in range(0, 8-len(vocab)%8)]
count = Counter(['a', 'a', 'a', 'a', 'b', 'b', 'c', 'c', 'd'])
test_eq(set([x for x in make_vocab(count) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a'.split()))
test_eq(len(make_vocab(count))%8, 0)
test_eq(set([x for x in make_vocab(count, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c d'.split()))
test_eq(set([x for x in make_vocab(count,max_vocab=12, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c'.split()))
#export
class TensorText(TensorBase): pass
class LMTensorText(TensorText): pass
# export
class Numericalize(Transform):
"Reversible transform of tokenized texts to numericalized ids"
def __init__(self, vocab=None, min_freq=3, max_vocab=60000, sep=' '):
self.vocab,self.min_freq,self.max_vocab,self.sep = vocab,min_freq,max_vocab,sep
self.o2i = None if vocab is None else defaultdict(int, {v:k for k,v in enumerate(vocab)})
def setups(self, dsets):
if dsets is None: return
if self.vocab is None:
count = dsets.counter if hasattr(dsets, 'counter') else Counter(p for o in dsets for p in o)
self.vocab = make_vocab(count, min_freq=self.min_freq, max_vocab=self.max_vocab)
self.o2i = defaultdict(int, {v:k for k,v in enumerate(self.vocab) if v != 'xxfake'})
def encodes(self, o): return TensorText(tensor([self.o2i [o_] for o_ in o]))
def decodes(self, o): return TitledStr(self.sep.join([self.vocab[o_] for o_ in o if self.vocab[o_] != PAD]))
num = Numericalize(min_freq=1, sep=' ')
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'This is an example of text this another'.split()))
test_eq(len(num.vocab)%8, 0)
start = 'This is an example of text'
t = num(start.split())
test_eq(t, tensor([11, 9, 12, 13, 14, 10]))
test_eq(num.decode(t), start)
num = Numericalize(min_freq=2, sep=' ')
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'is text'.split()))
test_eq(len(num.vocab)%8, 0)
t = num(start.split())
test_eq(t, tensor([0, 9, 0, 0, 0, 10]))
test_eq(num.decode(t), f'{UNK} is {UNK} {UNK} {UNK} text')
#hide
df = pd.DataFrame({'texts': ['This is an example of text', 'this is another text']})
tl = TfmdLists(df, [attrgetter('text'), Tokenizer.from_df('texts'), Numericalize(min_freq=2, sep=' ')])
test_eq(tl, [tensor([2, 8, 9, 10, 0, 0, 0, 11]), tensor([2, 9, 10, 0, 11])])
# ## LM_DataLoader -
#export
def _maybe_first(o): return o[0] if isinstance(o, tuple) else o
#export
def _get_tokenizer(ds):
tok = getattr(ds, 'tokenizer', None)
if isinstance(tok, Tokenizer): return tok
if isinstance(tok, (list,L)):
for t in tok:
if isinstance(t, Tokenizer): return t
#export
def _get_lengths(ds):
tok = _get_tokenizer(ds)
if tok is None: return
return tok.get_lengths(ds.items)
ReindexCollection
#export
#TODO: add backward
@delegates()
class LMDataLoader(TfmdDL):
def __init__(self, dataset, lens=None, cache=2, bs=64, seq_len=72, num_workers=0, **kwargs):
self.items = ReindexCollection(dataset, cache=cache, tfm=_maybe_first)
self.seq_len = seq_len
if lens is None: lens = _get_lengths(dataset)
if lens is None: lens = [len(o) for o in self.items]
self.lens = ReindexCollection(lens, idxs=self.items.idxs)
# The "-1" is to allow for final label, we throw away the end that's less than bs
corpus = round_multiple(sum(lens)-1, bs, round_down=True)
self.bl = corpus//bs #bl stands for batch length
self.n_batches = self.bl//(seq_len) + int(self.bl%seq_len!=0)
self.last_len = self.bl - (self.n_batches-1)*seq_len
self.make_chunks()
super().__init__(dataset=dataset, bs=bs, num_workers=num_workers, **kwargs)
self.n = self.n_batches*bs
def make_chunks(self): self.chunks = Chunks(self.items, self.lens)
def shuffle_fn(self,idxs):
self.items.shuffle()
self.make_chunks()
return idxs
def create_item(self, seq):
if seq>=self.n: raise IndexError
sl = self.last_len if seq//self.bs==self.n_batches-1 else self.seq_len
st = (seq%self.bs)*self.bl + (seq//self.bs)*self.seq_len
txt = self.chunks[st : st+sl+1]
return LMTensorText(txt[:-1]),txt[1:]
@delegates(TfmdDL.new)
def new(self, dataset=None, seq_len=72, **kwargs):
lens = self.lens.coll if dataset is None else None
return super().new(dataset=dataset, lens=lens, seq_len=seq_len, **kwargs)
#hide
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
list(dl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [5, 6, 7], [10, 11, 12], [15, 16, 17]]),
tensor([[1, 2, 3], [6, 7, 8], [11, 12, 13], [16, 17, 18]])],
[tensor([[3, 4], [8, 9], [13, 14], [18, 19]]),
tensor([[4, 5], [9, 10], [14, 15], [19, 20]])]])
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22,23],[24]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
#hide
#Check lens work
dl = LMDataLoader(ints, lens=ints.map(len), bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
dl = LMDataLoader(ints, bs=bs, seq_len=sl, shuffle=True)
for x,y in dl: test_eq(x[:,1:], y[:,:-1])
((x0,y0), (x1,y1)) = tuple(dl)
#Second batch begins where first batch ended
test_eq(y0[:,-1], x1[:,0])
test_eq(type(x0), LMTensorText)
# ### Showing
#export
@patch
def truncate(self:TitledStr, n):
words = self.split(' ')[:n]
return TitledStr(' '.join(words))
#export
@typedispatch
def show_batch(x: TensorText, y, samples, ctxs=None, max_n=10, trunc_at=150, **kwargs):
if ctxs is None: ctxs = get_empty_df(min(len(samples), max_n))
samples = L((s[0].truncate(trunc_at),*s[1:]) for s in samples)
ctxs = show_batch[object](x, y, samples, max_n=max_n, ctxs=ctxs, **kwargs)
display_df(pd.DataFrame(ctxs))
return ctxs
#export
@typedispatch
def show_batch(x: LMTensorText, y, samples, ctxs=None, max_n=10, **kwargs):
return show_batch[TensorText](x, None, samples, ctxs=ctxs, max_n=max_n, **kwargs)
# ## Integration example
path = untar_data(URLs.IMDB_SAMPLE)
df = pd.read_csv(path/'texts.csv')
df.head(2)
splits = ColSplitter()(df)
tfms = [attrgetter('text'), Tokenizer.from_df('text'), Numericalize()]
dsets = Datasets(df, [tfms], splits=splits, dl_type=LMDataLoader)
dls = dsets.dataloaders(bs=16, seq_len=72)
dls.show_batch(max_n=6)
b = dls.one_batch()
test_eq(type(x), LMTensorText)
test_eq(len(dls.valid_ds[0][0]), dls.valid.lens[0])
# ## Classification
#export
def pad_input(samples, pad_idx=1, pad_fields=0, pad_first=False, backwards=False):
"Function that collect samples and adds padding. Flips token order if needed"
pad_fields = L(pad_fields)
max_len_l = pad_fields.map(lambda f: max([len(s[f]) for s in samples]))
if backwards: pad_first = not pad_first
def _f(field_idx, x):
if field_idx not in pad_fields: return x
idx = pad_fields.items.index(field_idx) #TODO: remove items if L.index is fixed
sl = slice(-len(x), sys.maxsize) if pad_first else slice(0, len(x))
pad = x.new_zeros(max_len_l[idx]-x.shape[0])+pad_idx
x1 = torch.cat([pad, x] if pad_first else [x, pad])
if backwards: x1 = x1.flip(0)
return retain_type(x1, x)
return [tuple(map(lambda idxx: _f(*idxx), enumerate(s))) for s in samples]
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0),
[(tensor([1,2,3]),1), (tensor([4,5,0]),2), (tensor([6,0,0]), 3)])
test_eq(pad_input([(tensor([1,2,3]), (tensor([6]))), (tensor([4,5]), tensor([4,5])), (tensor([6]), (tensor([1,2,3])))], pad_idx=0, pad_fields=1),
[(tensor([1,2,3]),(tensor([6,0,0]))), (tensor([4,5]),tensor([4,5,0])), ((tensor([6]),tensor([1, 2, 3])))])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, pad_first=True),
[(tensor([1,2,3]),1), (tensor([0,4,5]),2), (tensor([0,0,6]), 3)])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, backwards=True),
[(tensor([3,2,1]),1), (tensor([5,4,0]),2), (tensor([6,0,0]), 3)])
x = test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, backwards=True),
[(tensor([3,2,1]),1), (tensor([5,4,0]),2), (tensor([6,0,0]), 3)])
#hide
#Check retain type
x = [(TensorText([1,2,3]),1), (TensorText([4,5]), 2), (TensorText([6]), 3)]
y = pad_input(x, pad_idx=0)
for s in y: test_eq(type(s[0]), TensorText)
#export
def pad_input_chunk(samples, pad_idx=1, pad_first=True, seq_len=72):
max_len = max([len(s[0]) for s in samples])
def _f(x):
l = max_len - x.shape[0]
pad_chunk = x.new_zeros((l//seq_len) * seq_len) + pad_idx
pad_res = x.new_zeros(l % seq_len) + pad_idx
x1 = torch.cat([pad_chunk, x, pad_res]) if pad_first else torch.cat([x, pad_res, pad_chunk])
return retain_type(x1, x)
return [(_f(s[0]), *s[1:]) for s in samples]
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),1), (tensor([1,2,3]), 2), (tensor([1,2]), 3)], pad_idx=0, seq_len=2),
[(tensor([1,2,3,4,5,6]),1), (tensor([0,0,1,2,3,0]),2), (tensor([0,0,0,0,1,2]), 3)])
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),), (tensor([1,2,3]),), (tensor([1,2]),)], pad_idx=0, seq_len=2),
[(tensor([1,2,3,4,5,6]),), (tensor([0,0,1,2,3,0]),), (tensor([0,0,0,0,1,2]),)])
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),), (tensor([1,2,3]),), (tensor([1,2]),)], pad_idx=0, seq_len=2, pad_first=False),
[(tensor([1,2,3,4,5,6]),), (tensor([1,2,3,0,0,0]),), (tensor([1,2,0,0,0,0]),)])
# +
#export
def _default_sort(x): return len(x[0])
@delegates(TfmdDL)
class SortedDL(TfmdDL):
def __init__(self, dataset, sort_func=None, res=None, **kwargs):
super().__init__(dataset, **kwargs)
self.sort_func = _default_sort if sort_func is None else sort_func
if res is None and self.sort_func == _default_sort: res = _get_lengths(dataset)
self.res = [self.sort_func(self.do_item(i)) for i in range_of(self.dataset)] if res is None else res
if len(self.res) > 0: self.idx_max = np.argmax(self.res)
def get_idxs(self):
idxs = super().get_idxs()
if self.shuffle: return idxs
return sorted(idxs, key=lambda i: self.res[i], reverse=True)
def shuffle_fn(self,idxs):
idxs = np.random.permutation(len(self.dataset))
idx_max = np.where(idxs==self.idx_max)[0][0]
idxs[0],idxs[idx_max] = idxs[idx_max],idxs[0]
sz = self.bs*50
chunks = [idxs[i:i+sz] for i in range(0, len(idxs), sz)]
chunks = [sorted(s, key=lambda i: self.res[i], reverse=True) for s in chunks]
sort_idx = np.concatenate(chunks)
sz = self.bs
batches = [sort_idx[i:i+sz] for i in range(0, len(sort_idx), sz)]
sort_idx = np.concatenate(np.random.permutation(batches[1:-1])) if len(batches) > 2 else np.array([],dtype=np.int)
sort_idx = np.concatenate((batches[0], sort_idx) if len(batches)==1 else (batches[0], sort_idx, batches[-1]))
return iter(sort_idx)
@delegates(TfmdDL.new)
def new(self, dataset=None, **kwargs):
res = self.res if dataset is None else None
return super().new(dataset=dataset, res=res, **kwargs)
# -
ds = [(tensor([1,2]),1), (tensor([3,4,5,6]),2), (tensor([7]),3), (tensor([8,9,10]),4)]
dl = SortedDL(ds, bs=2, before_batch=partial(pad_input, pad_idx=0))
test_eq(list(dl), [(tensor([[ 3, 4, 5, 6], [ 8, 9, 10, 0]]), tensor([2, 4])),
(tensor([[1, 2], [7, 0]]), tensor([1, 3]))])
ds = [(tensor(range(random.randint(1,10))),i) for i in range(101)]
dl = SortedDL(ds, bs=2, create_batch=partial(pad_input, pad_idx=-1), shuffle=True, num_workers=0)
batches = list(dl)
max_len = len(batches[0][0])
for b in batches:
assert(len(b[0])) <= max_len
test_ne(b[0][-1], -1)
splits = RandomSplitter()(range_of(df))
dsets = Datasets(df, splits=splits, tfms=[tfms, [attrgetter("label"), Categorize()]], dl_type=SortedDL)
dls = dsets.dataloaders(before_batch=pad_input)
dls.show_batch(max_n=2)
# ## TransformBlock for text
#export
class TextBlock(TransformBlock):
@delegates(Numericalize.__init__)
def __init__(self, tok_tfm, vocab=None, is_lm=False, seq_len=72, **kwargs):
return super().__init__(type_tfms=[tok_tfm, Numericalize(vocab, **kwargs)],
dl_type=LMDataLoader if is_lm else SortedDL,
dls_kwargs={} if is_lm else {'before_batch': partial(pad_input_chunk, seq_len=seq_len)})
@classmethod
@delegates(Tokenizer.from_df, keep=True)
def from_df(cls, text_cols, vocab=None, is_lm=False, seq_len=72, **kwargs):
return cls(Tokenizer.from_df(text_cols, **kwargs), vocab=vocab, is_lm=is_lm, seq_len=seq_len)
@classmethod
@delegates(Tokenizer.from_folder, keep=True)
def from_folder(cls, path, vocab=None, is_lm=False, seq_len=72, **kwargs):
return cls(Tokenizer.from_folder(path, **kwargs), vocab=vocab, is_lm=is_lm, seq_len=seq_len)
# ## TextDataLoaders -
# +
#export
class TextDataLoaders(DataLoaders):
@classmethod
@delegates(DataLoaders.from_dblock)
def from_folder(cls, path, train='train', valid='valid', valid_pct=None, seed=None, vocab=None, text_vocab=None, is_lm=False,
tok_tfm=None, seq_len=72, **kwargs):
"Create from imagenet style dataset in `path` with `train`,`valid`,`test` subfolders (or provide `valid_pct`)."
splitter = GrandparentSplitter(train_name=train, valid_name=valid) if valid_pct is None else RandomSplitter(valid_pct, seed=seed)
blocks = [TextBlock.from_folder(path, text_vocab, is_lm, seq_len) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm, seq_len)]
if not is_lm: blocks.append(CategoryBlock(vocab=vocab))
get_items = partial(get_text_files, folders=[train,valid]) if valid_pct is None else get_text_files
dblock = DataBlock(blocks=blocks,
get_items=get_items,
splitter=splitter,
get_y=None if is_lm else parent_label)
return cls.from_dblock(dblock, path, path=path, seq_len=seq_len, **kwargs)
@classmethod
@delegates(DataLoaders.from_dblock)
def from_df(cls, df, path='.', valid_pct=0.2, seed=None, text_col=0, label_col=1, label_delim=None, y_block=None,
text_vocab=None, is_lm=False, valid_col=None, tok_tfm=None, seq_len=72, **kwargs):
blocks = [TextBlock.from_df(text_col, text_vocab, is_lm, seq_len) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm, seq_len)]
if y_block is None and not is_lm:
blocks.append(MultiCategoryBlock if is_listy(label_col) and len(label_col) > 1 else CategoryBlock)
if y_block is not None and not is_lm: blocks += (y_block if is_listy(y_block) else [y_block])
splitter = RandomSplitter(valid_pct, seed=seed) if valid_col is None else ColSplitter(valid_col)
dblock = DataBlock(blocks=blocks,
get_x=ColReader(text_col),
get_y=None if is_lm else ColReader(label_col, label_delim=label_delim),
splitter=splitter)
return cls.from_dblock(dblock, df, path=path, seq_len=seq_len, **kwargs)
@classmethod
def from_csv(cls, path, csv_fname='labels.csv', header='infer', delimiter=None, **kwargs):
df = pd.read_csv(Path(path)/csv_fname, header=header, delimiter=delimiter)
return cls.from_df(df, path=path, **kwargs)
TextDataLoaders.from_csv = delegates(to=TextDataLoaders.from_df)(TextDataLoaders.from_csv)
# -
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/31_text.data.ipynb |
# +
# Example of a Gaussian Process Regression with multiple local minima
# in the marginal log-likelihood as a function of the hyperparameters
# Based on: https://github.com/probml/pmtk3/blob/master/demos/gprDemoMarglik.m
# Authors: <NAME> & <NAME>
import numpy as np
import matplotlib.pyplot as plt
try:
import probml_utils as pml
except ModuleNotFoundError:
# %pip install -qq git+https://github.com/probml/probml-utils.git
import probml_utils as pml
from numpy.linalg import inv, slogdet
from scipy.optimize import minimize
def k(u, v, sigma_f, l=1):
return sigma_f**2 * np.exp(-((u - v) ** 2) / (2 * l**2))
def gp_predictive_post(xstar, x, y, k, sigma_y, *args, **kwargs):
"""
Compute predictive distribution of a 1D-Gaussian Process for
regression
Parameters
----------
xstar: array(nt, 1)
Values to perform inference on
x: array(n, 1)
Training independent variables
y: array(n, 1)
Training dependent variables
k: function
Kernel function to evaluate the GP
sigma_y: float
data-noise term
*args: additional arguments of k
**kwargs: additional keyword-arguments of k
Returns
-------
* array(nt, 1):
Array of predicted (mean) values
* array(nt, nt):
Posterior covariance matrix
"""
n, _ = x.shape
kstar = k(x, xstar.T, *args, **kwargs)
Kxx = k(x, x.T, *args) + sigma_y**2 * np.eye(n)
kxx_star = k(xstar, xstar.T, *args, **kwargs)
Kxx_inv = inv(Kxx)
ystar = kstar.T @ Kxx_inv @ y
Sigma_post = kxx_star - kstar.T @ Kxx_inv @ kstar
return ystar, Sigma_post
def log_likelihood(x, y, sigma_f, l, sigma_y):
"""
Compute marginal log-likelihood of a regression GP
with rbf kernel
Parameters
----------
x: array(n, 1)
Training independent variables
y: array(n, 1)
Training dependent variables
sigma_f: float
Vertical-scale parameter
l: float
Horizontal-scale parameter
sigma_y: float
data noise
Returns
-------
* float:
Marginal log-likelihood as the specified hyperparameters
"""
n, _ = x.shape
x = x / np.exp(l)
Kxx = k(x, x.T, sigma_f) + np.exp(2 * sigma_y) * np.eye(n)
_, DKxx = slogdet(Kxx)
l = -1 / 2 * (y.T @ inv(Kxx) @ y + DKxx + n * np.log(2 * np.pi))
return l.item()
def plot_gp_pred(x, y, xstar, k, sigma_f, l, sigma_y, ax):
ystar, Sigma_post = gp_predictive_post(xstar, x, y, k, sigma_y, sigma_f, l)
upper_bound = ystar.ravel() + 2 * np.sqrt(np.diag(Sigma_post))
lower_bound = ystar.ravel() - 2 * np.sqrt(np.diag(Sigma_post))
ax.scatter(x, y, marker="+", s=100, c="black")
ax.plot(xstar, ystar, c="black")
ax.fill_between(xstar.ravel(), lower_bound, upper_bound, color="tab:gray", alpha=0.3, edgecolor="none")
ax.set_xlim(-7.5, 7.5)
ax.set_ylim(-2, 2.5)
def plot_marginal_likelihood_surface(x, y, sigma_f, l_space, sigma_y_space, ax, levels=None):
P = np.stack(np.meshgrid(l_space, sigma_y_space), axis=0)
Z = np.apply_along_axis(lambda p: log_likelihood(x, y, sigma_f, *p), 0, P)
ax.contour(*np.exp(P), Z, levels=levels)
ax.set_xlabel("characteristic length scale")
ax.set_ylabel("noise standard deviation")
ax.set_xscale("log")
ax.set_yscale("log")
if __name__ == "__main__":
plt.rcParams["axes.spines.right"] = False
plt.rcParams["axes.spines.top"] = False
sigma_f = 1.0
x = np.array([-1.3089, 6.7612, 1.0553, -1.1734, -2.9339, 7.2530, -6.5843])[:, None]
y = np.array([1.6218, 1.8558, 0.4102, 1.2526, -0.0133, 1.6380, 0.2189])[:, None]
xstar = np.linspace(-7.5, 7.5, 201)
ngrid = 41
l_space = np.linspace(np.log(0.5), np.log(80), ngrid)
sigma_y_space = np.linspace(np.log(0.03), np.log(3), ngrid)
P = np.stack(np.meshgrid(l_space, sigma_y_space), axis=0)
configs = [(1.0, 0.2), (10, 0.8)]
fig, ax = plt.subplots()
plot_gp_pred(x, y, xstar, k, sigma_f, *configs[0], ax)
pml.savefig("gpr_config0.pdf")
fig, ax = plt.subplots()
plot_gp_pred(x, y, xstar, k, sigma_f, *configs[1], ax)
pml.savefig("gpr_config1.pdf")
ngrid = 41
w01 = np.array([np.log(1), np.log(0.1)])
w02 = np.array([np.log(10), np.log(0.8)])
s0 = minimize(lambda p: -log_likelihood(x, y, sigma_f, *p), w01)
s1 = minimize(lambda p: -log_likelihood(x, y, sigma_f, *p), w02)
levels = -np.array([8.3, 8.5, 8.9, 9.3, 9.8, 11.5, 15])[::-1]
l_space = np.linspace(np.log(0.5), np.log(80), ngrid)
sigma_y_space = np.linspace(np.log(0.03), np.log(3), ngrid)
fig, ax = plt.subplots()
plot_marginal_likelihood_surface(x, y, sigma_f, l_space, sigma_y_space, ax, levels=levels)
plt.scatter(*np.exp(s0.x), marker="+", s=100, c="tab:blue")
plt.scatter(*np.exp(s1.x), marker="+", s=100, c="tab:blue")
pml.savefig("gpr_marginal_likelihood.pdf")
plt.show()
| notebooks/book1/17/gpr_demo_marglik.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importance of known positives versus known negatives
#
# In this notebook we will show how to compute performance curves (ROC and PR curves to be specific) based on a data set with known positives, known negatives and unlabeled data. We will show how to use our approach starting from known negatives instead of known positives, which is useful of the number of known negatives is much larger.
#
# This notebook is intended to enable you to easily change the configuration of experiments, so feel free to make modifications where you see fit to see their effect.
import random
import operator as op
import optunity.metrics
import semisup_metrics as ss
import numpy as np
from matplotlib import pyplot as plt
import pickle
import csv
import util
# %matplotlib inline
# # Create or load data set
# Generate simulated data. Feel free to configure the code below as you please.
# +
# fraction of positives/negatives that are known
# known_neg_frac == 0 implies PU learning
known_pos_frac = 0.05
known_neg_frac = 0.5
# if you simulate a smaller data set you can see some effects better
# but estimated bounds will be much wider
# because the ECDF confidence interval becomes large
num_pos = 10000
num_neg = 10000
distid = 2 # can be 1, 2 or 3, these correspond to certain curves in ROC space
# generate rankings and label vectors and compute corresponding beta
# beta is based on known_pos_frac, known_neg_frac and the number of pos and negs
# labels is a list of {True, False, None}, where None indicates unlabeled
# true_labels is a list of {True, False}
labels, true_labels, decision_values, beta = util.simulate_data(distid, num_pos, num_neg,
known_pos_frac,
known_neg_frac)
# -
# ## Data set characteristics
# Known parameters:
print('total number of instances: \t %d' % len(labels))
print('number of known positives: \t %d' % len(list(filter(lambda x: x == True, labels))))
print('number of known negatives: \t %d' % len(list(filter(lambda x: x == False, labels))))
print('number of unlabeled instances: \t %d' % len(list(filter(lambda x: x == None, labels))))
# Unknown parameters:
print('number of latent positives: \t %d' % len(list(filter(lambda x: x[0] == None and x[1] == True, zip(labels, true_labels)))))
print('number of latent negatives: \t %d' % len(list(filter(lambda x: x[0] == None and x[1] == False, zip(labels, true_labels)))))
print('beta: \t %1.4f' % beta)
# # Estimate beta
#
# As this notebook is intended to show the effect of known positives vs known negatives, we will continue with the true value of beta. Change this parameter to see its effect in the figures later on.
# +
betahat = beta
print('true value of beta\t%1.4f' % beta)
print('point estimate of beta\t%1.4f' % betahat)
# -
# # 1. Compute cumulative rank distribution of known positives
#
# We start by computing the rank CDF of known positives, and then determine a confidence interval for it. We will use a standard bootstrap approach.
# +
# sort the labels in descending order of corresponding decision values
sort_labels, sort_dv, sort_true_labels = zip(*sorted(zip(labels, decision_values, true_labels),
key=op.itemgetter(1), reverse=True))
# ranks of the known positives
known_pos_ranks = [idx for idx, lab in enumerate(sort_labels) if lab]
# compute rank ECDF of known positives
known_pos_ecdf = ss.compute_ecdf_curve(known_pos_ranks)
# -
# Next, we determine a confidence interval on the rank CDF of known positives. We can do this in several ways, our code provides a bootstrap approach and a method based on the Dvoretzky–Kiefer–Wolfowitz (DKW) inequality. Feel free to experiment.
# +
ci_width = 0.95 # width of the confidence band on ECDF to be used
use_bootstrap = True # use bootstrap to compute confidence band
nboot = 2000 # number of bootstrap iterations to use, not used if use_bootstrap = False
if use_bootstrap:
pos_cdf_bounds = ss.bootstrap_ecdf_bounds(labels, decision_values, nboot=nboot, ci_width=ci_width)
else:
pos_cdf_bounds = ss.dkw_bounds(labels, decision_values, ci_width=ci_width)
# -
# For reference, we will also compute the rank ECDF of latent positives. In practical applications this is impossible, as the latent positives are by definition not known.
latent_positives = map(lambda x, y: x == True and y == None, true_labels, labels)
sort_lps, _ = zip(*sorted(zip(latent_positives, decision_values),
key=op.itemgetter(1), reverse=True))
latent_pos_ranks = [idx for idx, lab in enumerate(sort_lps) if lab]
latent_pos_ecdf = ss.compute_ecdf_curve(latent_pos_ranks)
# Plot the rank CDFs of known and latent positives. It may occur that the rank CDF of latent positives is not within the confidence interval of known positives, in which case the corresponding bounds on performance will not be strict.
# +
# convenience plot functions
def plot_proxy(color, alpha):
p = plt.Rectangle((0, 0), 0, 0, color=color, alpha=alpha)
ax = plt.gca()
ax.add_patch(p)
return p
def fix_plot_shape(fig):
ax = fig.add_subplot(111, aspect='equal')
axes = fig.gca()
axes.set_xlim([0,1])
axes.set_ylim([0,1])
xs = list(range(len(labels)))
plt.figure(1)
plt.fill_between(xs, list(map(pos_cdf_bounds.lower, xs)), list(map(pos_cdf_bounds.upper, xs)),
color='blue', alpha=0.4)
plt.plot(*zip(*known_pos_ecdf), color='black', linestyle='dashed', linewidth=2)
plt.plot(*zip(*latent_pos_ecdf), color='black', linewidth=2)
plot_proxy('blue', 0.4)
plt.xlabel('rank')
plt.ylabel('TPR')
plt.legend(['known positives', 'latent positives', 'expected region'], loc=4)
plt.title('Rank CDF')
plt.show()
# -
# # 2. Compute cumulative rank distribution of known negatives
#
# The code below is analogous to what we did earlier for known positives, but now for known negatives. We will first flip known class labels, as described in the text.
# +
# ranks of the known positives
known_neg_ranks = [idx for idx, lab in enumerate(sort_labels) if lab == False]
# compute rank ECDF of known positives
known_neg_ecdf = ss.compute_ecdf_curve(sorted(known_neg_ranks))
# flip class labels
labels_flipped = [not label if label is not None else None for label in labels]
sort_labels_flipped = [not label if label is not None else None for label in sort_labels]
# compute confidence interval in the same way as we did for known positives
if use_bootstrap:
neg_cdf_bounds = ss.bootstrap_ecdf_bounds(labels_flipped, decision_values, nboot=nboot, ci_width=ci_width)
else:
neg_cdf_bounds = ss.dkw_bounds(labels_flipped, decision_values, ci_width=ci_width)
# for reference, rank CDF of latent negatives
latent_negatives = map(lambda x, y: x == False and y == None, true_labels, labels)
sort_lns, _ = zip(*sorted(zip(latent_negatives, decision_values),
key=op.itemgetter(1), reverse=True))
latent_neg_ranks = [idx for idx, lab in enumerate(sort_lns) if lab]
latent_neg_ecdf = ss.compute_ecdf_curve(latent_neg_ranks)
# -
# Plot rank CDFs of known positives and known negatives along with their CIs.
# +
xs = list(range(len(labels)))
plt.figure(1)
plt.fill_between(xs, list(map(pos_cdf_bounds.lower, xs)), list(map(pos_cdf_bounds.upper, xs)),
color='blue', alpha=0.4)
plt.plot(*zip(*known_pos_ecdf), color='blue', linestyle='dashed', linewidth=2)
plt.plot(*zip(*latent_pos_ecdf), color='blue', linewidth=2)
plot_proxy('blue', 0.4)
plt.fill_between(xs, list(map(neg_cdf_bounds.lower, xs)), list(map(neg_cdf_bounds.upper, xs)),
color='red', alpha=0.4)
plt.plot(*zip(*known_neg_ecdf), color='red', linestyle='dashed', linewidth=2)
plt.plot(*zip(*latent_neg_ecdf), color='red', linewidth=2)
plot_proxy('red', 0.4)
plt.xlabel('rank')
plt.ylabel('TPR')
plt.legend(['known positives', 'latent positives', 'known negatives', 'latent negatives', 'CI for pos', 'CI for neg'],
loc="upper left", bbox_to_anchor=(1,1))
plt.title('Rank CDF')
plt.show()
# -
# # 3. Compute contingency tables for each rank
# We will compute contingency tables based on the rank distribution of known positives and based on the rank distribution of known negatives.
#
# Applying our approach to known negatives requires a few modifications:
# 1. Flip known class labels, so negatives become positives
# 2. Modify beta (new beta = 1 - beta)
# 3. Run our approach
# 4. Adjust resulting contingency tables (i.e., flip class labels back). Note also that the optimistic contingency table becomes the pessimistic table when flipping labels and vice versa.
# +
# compute contingency tables based on CI of rank CDF of known positives
tables_pos = ss.compute_contingency_tables(labels=sort_labels, decision_values=sort_dv,
reference_lb=pos_cdf_bounds.lower,
reference_ub=pos_cdf_bounds.upper,
beta=betahat, presorted=True)
# compute contingency tables based on CI of rank CDF of known negatives
# this requires flipping labels, changing beta and post-processing the resulting contingency tables
betahat_flipped = 1.0 - betahat
tables_neg = ss.compute_contingency_tables(labels=sort_labels_flipped, decision_values=sort_dv,
reference_lb=neg_cdf_bounds.lower,
reference_ub=neg_cdf_bounds.upper,
beta=betahat_flipped, presorted=True)
postprocess_ct = lambda ct: ss.ContingencyTable(TP=ct.FP,
FP=ct.TP,
TN=ct.FN,
FN=ct.TN)
tables_neg_post = ss._lb_ub(lower=list(map(postprocess_ct, tables_neg.upper)),
upper=list(map(postprocess_ct, tables_neg.lower)))
# -
# # 4. Compute and plot performance estimates based on the contingency tables
# ## 4.1 ROC curves
#
# Start off with the easy stuff: the true ROC curve (unknown in practice), and the curve we obtain by assuming beta=0.
# compute the true ROC curve (we use Optunity's ROC function)
_, roc_true = optunity.metrics.roc_auc(true_labels, decision_values, return_curve=True)
# Compute bounds based on the proposed method.
# +
# we can directly use the contingency tables we already computed anyways
roc_bounds = lambda tables: ss._lb_ub(lower=ss.roc_from_cts(tables.lower),
upper=ss.roc_from_cts(tables.upper))
# bounds starting from known positives
roc_bounds_pos = roc_bounds(tables_pos)
# bounds starting from known negatives
roc_bounds_neg = roc_bounds(tables_neg_post)
# -
# Plot the resulting curves
xs = [float(x) / 100 for x in range(101)]
roc_pos_up = ss.zoh(*zip(*roc_bounds_pos.upper))
roc_pos_lo = ss.zoh(*zip(*roc_bounds_pos.lower))
roc_neg_up = ss.zoh(*zip(*roc_bounds_neg.upper))
roc_neg_lo = ss.zoh(*zip(*roc_bounds_neg.lower))
fig = plt.figure(2)
fix_plot_shape(fig)
plt.plot(*zip(*roc_true), color='black', linewidth=2)
plt.fill_between(xs, list(map(roc_pos_lo, xs)), list(map(roc_pos_up, xs)), color='blue', alpha=0.3)
plt.fill_between(xs, list(map(roc_neg_lo, xs)), list(map(roc_neg_up, xs)), color='red', alpha=0.7)
plot_proxy('blue', 0.3)
plot_proxy('red', 0.7)
plt.xlabel('FPR')
plt.ylabel('TPR')
plt.legend(['true curve', 'expected region via known pos', 'expected region via known neg'],
loc="upper left", bbox_to_anchor=(1,1))
plt.title('Receiver Operating Characteristic curve')
plt.show()
# The resulting curves show clearly that in this case computing bounds based on known negatives is better, because we have far more known negatives.
# ## 4.2 PR curves
# +
# we can directly use the contingency tables we already computed anyways
pr_bounds = lambda tables: ss._lb_ub(lower=ss.pr_from_cts(tables.lower),
upper=ss.pr_from_cts(tables.upper))
pr_bounds_pos = pr_bounds(tables_pos)
pr_bounds_neg = pr_bounds(tables_neg_post)
# an alternative without all these intermediate steps would be:
# roc_bounds_point = ss.roc_bounds(labels, decision_values, beta=betahat)
# compute the true ROC curve (we use Optunity's ROC function)
_, pr_true = optunity.metrics.pr_auc(true_labels, decision_values, return_curve=True)
# -
pr_pos_up = ss.zoh(*zip(*pr_bounds_pos.upper))
pr_pos_lo = ss.zoh(*zip(*pr_bounds_pos.lower))
pr_neg_up = ss.zoh(*zip(*pr_bounds_neg.upper))
pr_neg_lo = ss.zoh(*zip(*pr_bounds_neg.lower))
fig = plt.figure(3)
fix_plot_shape(fig)
plt.plot(*zip(*pr_true), color='black', linewidth=2)
plt.fill_between(xs, list(map(pr_pos_lo, xs)), list(map(pr_pos_up, xs)), color='blue', alpha=0.3)
plt.fill_between(xs, list(map(pr_neg_lo, xs)), list(map(pr_neg_up, xs)), color='red', alpha=0.7)
plot_proxy('blue', 0.3)
plot_proxy('red', 0.7)
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.legend(['true curve', 'expected region via known pos', 'expected region via known neg'],
loc="upper left", bbox_to_anchor=(1,1))
plt.title('Precision-Recall curve')
plt.show()
| known-pos-vs-known-neg-python3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BFS: Shortest Reach in a Graph
#
# <br>
#
# 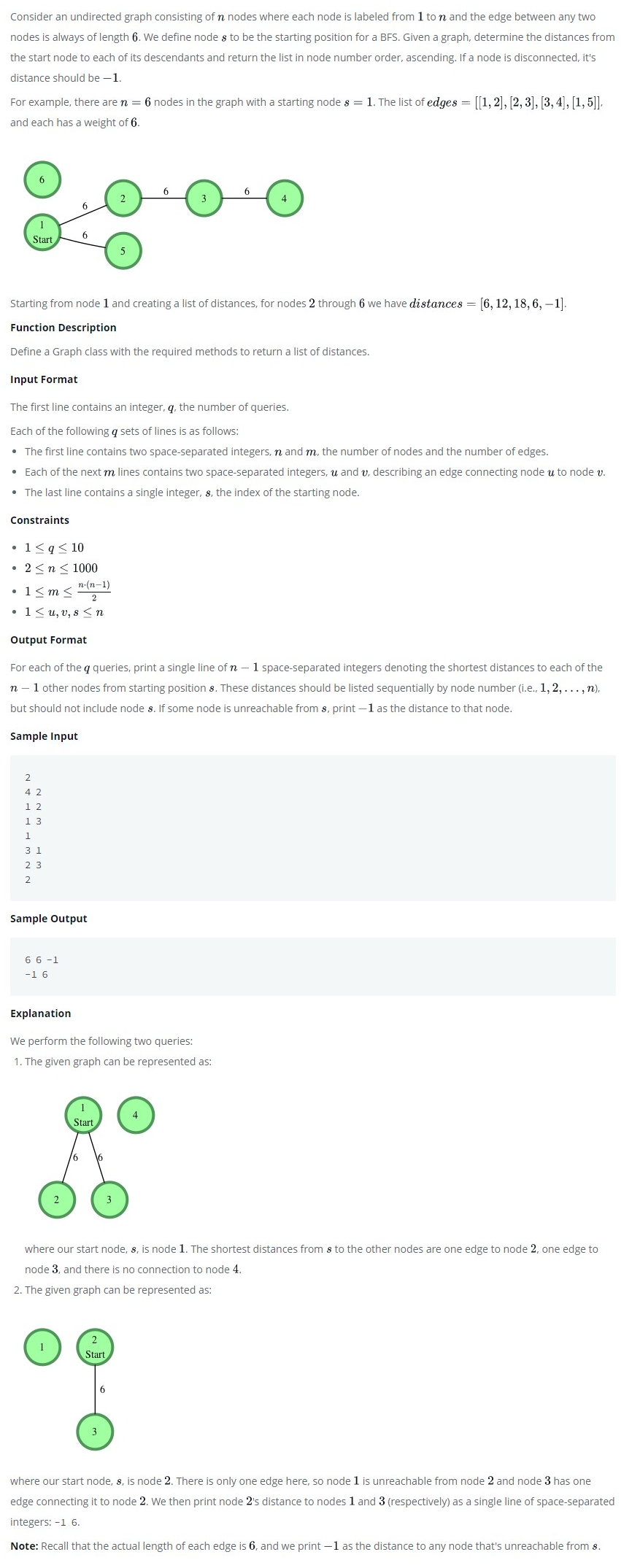
# +
from collections import deque
class Graph:
num = 0
def __init__(self, num):
self.num = num
self.lst = [[] for _ in range(num)]
def connect(self, x, y):
self.lst[x] += [y]
self.lst[y] += [x]
def find_all_distances(self, start):
q = deque()
q.append(start)
visit = [-1] * self.num
visit[start] = 0
while q:
curr = q.popleft()
for i in self.lst[curr]:
if visit[i] == -1:
visit[i] = visit[curr] + 6
q.append(i)
print(*[val for (idx, val) in enumerate(visit) if idx != start])
for i in range(int(input())):
n, m = map(int, input().split())
graph = Graph(n)
for i in range(m):
x, y = [int(x) for x in input().split()]
graph.connect(x - 1, y - 1)
s = int(input())
graph.find_all_distances(s - 1)
| Interview Preparation Kit/10. Graphs/BFS_Shortest Reach in a Graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing Road Crash Data
#
# ##### 1.1 Data Preparation and Loading
# **1. Write the code to create a SparkContext object using SparkSession, which tells Spark how to access a cluster. To create a SparkSession you first need to build a SparkConf object that contains information about your application. Give an appropriate name for your application and run.**
# +
# Code to create a SparkContext object using SparkSession
# Import SparkConf class into program
from pyspark import SparkConf
# local[*]: run Spark in local mode with as many working processors as logical cores on your machine
# If we want Spark to run locally with 'k' worker threads, we can specify as "local[k]".
master = "local[*]"
# The `appName` field is a name to be shown on the Spark cluster UI page
# app_name = "Assignment 1"
# Setup configuration parameters for Spark
spark_conf = SparkConf().setMaster(master).setAppName(app_name)
# Import SparkContext and SparkSession classes
from pyspark import SparkContext # Spark
from pyspark.sql import SparkSession # Spark SQL
# Using SparkSession
spark = SparkSession.builder.config(conf=spark_conf).getOrCreate()
sc = spark.sparkContext
sc.setLogLevel('ERROR')
# -
# **2. Import all the “Units” csv files from 2015-2019 into a single RDD.**
# creating a Units csv rdd
units_rdd = sc.textFile('*_DATA_SA_Units.csv')
# **3. Import all the “Crashes” csv files from 2015-2019 into a single RDD.**
# creating a Crashes csv rdd
crashes_rdd = sc.textFile('*_DATA_SA_Crash.csv')
# **4. For each Units and Crashes RDDs, remove the header rows and display the total
# count and first 10 records.**
# +
# to remove the next line character
units_rdd = units_rdd.map(lambda line: line.split(','))
# filter out header
header_unit = units_rdd.first()
filt_units_rdd = units_rdd.filter(lambda row: row != header_unit)
print("#### For Units ####")
print(f"Number of lines: {filt_units_rdd.count()}") # to display the total count
filt_units_rdd.take(10) # to display first 10 records
# +
# to remove the next line character
crashes_rdd = crashes_rdd.map(lambda line: line.split(','))
# to remove the header
header_crash = crashes_rdd.first()
filt_crashes_rdd = crashes_rdd.filter(lambda row: row != header_crash)
print("\n\n#### For Crashes ####")
print(f"Number of lines: {filt_crashes_rdd.count()}") # to display the total count
filt_crashes_rdd.take(10) # to display first 10 records
# -
# ##### 1.2 Data Partitioning in RDD
# +
# function to print number of records in each partition
from pyspark.rdd import RDD
def print_partitions(data):
if isinstance(data, RDD):
numPartitions = data.getNumPartitions()
partitions = data.glom().collect()
else:
numPartitions = data.rdd.getNumPartitions()
partitions = data.rdd.glom().collect()
print(f"####### NUMBER OF PARTITIONS: {numPartitions}")
for index, partition in enumerate(partitions):
# show partition if it is not empty
if len(partition) > 0:
print(f"Partition {index}: {len(partition)} records")
# -
print("Number of partitions:{}".format(filt_units_rdd.getNumPartitions())) # to display number of partitions
print("Partitioner:{}".format(filt_units_rdd.partitioner))
print_partitions(filt_units_rdd)
print("Number of partitions:{}".format(filt_crashes_rdd.getNumPartitions())) # to display number of partitions
print("Partitioner:{}".format(filt_crashes_rdd.partitioner))
print_partitions(filt_crashes_rdd)
# By default Spark is partitioning data according to **Random equal partitioning** method. As we can see the number of records in each partition has less variation and both the rdds have 5 partitions each.
# **a. Create a Key Value Pair RDD with Lic State as the key and rest of the other columns as value.**
# +
# to create Key Value Pair RDD from units rdd
result_pair = filt_units_rdd.map(lambda x: (x[9], (x[0], x[1], x[2], x[3], x[4], x[5], x[6],\
x[7], x[8], x[10], x[11], x[12], x[13], x[14], x[15], x[16], x[17])))
result_pair.take(2)
# -
# **b. Write the code to implement this partitioning in RDD using appropriate partitioning functions.**
# Using hash partitioning function to partition rdd
def hash_function(key):
val = 0
if key == '"SA"':
val = 2
else:
val = 3
return val
# performing hash partitioning with our function
num_partitions = 2
hash_partition_rdd = result_pair.partitionBy(num_partitions, hash_function)
# **c. Write the code to print the number of records in each partition. What does it tell about the data skewness?**
# using the above created print_partitions function to get records in each partition
print_partitions(hash_partition_rdd)
# By looking at the nnumber of records in each partition we can identify that **Partition 0** that denotes records belonging to **SA** are highly greater than that for other states. This implies that our data is highly skewed.
# ##### Average age of male and female drivers separately.
# +
# filtering done to remove empty and non-essential records from the AGE column
clean_rdd = filt_units_rdd.filter(lambda x: '"XXX"' not in x[8]).filter(lambda x: x[8] != '')
# to get the desired columns of GENDER and AGE
male_female_rdd = clean_rdd.map(lambda x: (x[7].replace('"',''), int(x[8].replace('"',''))))
# -
# to aggregate only the male records in gender column
male_rdd = male_female_rdd.filter(lambda x: x[0] == 'Male')
grouped_male = male_rdd.groupByKey().map(lambda x: (x[0], sum(x[1])/len(x[1])))
grouped_male.collect()
# to aggregate only the female records in gender column
female_rdd = male_female_rdd.filter(lambda x: x[0] == 'Female')
grouped_female = female_rdd.groupByKey().map(lambda x: (x[0], sum(x[1])/len(x[1])))
grouped_female.collect()
# **Oldest and the newest vehicle year involved in the accident? Display the Registration State, Year and Unit type of the vehicle.**
# +
# to get the required columns from the rdd
vehicle_reqq = filt_units_rdd.filter(lambda x: (x[5]!='' and 'XXXX' not in x[5]))\
.map(lambda x: (x[3],x[4],int(x[5].replace('"',''))))
# to find the details of newest vehicle
vehicle_reqq.max(key=lambda x: x[2])
# -
# to find the details of oldest vehicle
vehicle_reqq.min(key=lambda x: x[2])
# ##### 2.1 Data Preparation and Loading
# **1. Load all units and crash data into two separate dataframes**
# to load data in two dataframes
units_df = spark.read.csv("*_DATA_SA_Units.csv",header=True)
crashes_df = spark.read.csv("*_DATA_SA_Crash.csv",header=True)
# #### 2. Display the schema of the final two dataframes
# to display the schema
units_df.printSchema()
crashes_df.printSchema()
# to import necessary functionality
from pyspark.sql.functions import *
from pyspark.sql.types import IntegerType
# +
# to convert 'Total Cas' column to integer type
crashes_df = crashes_df.withColumn("Total Cas", crashes_df["Total Cas"].cast(IntegerType()))
# to show all crashes in Adelaide with casualities greater than 3
crashes_df.where((col("Suburb") == 'ADELAIDE') & (col("Total Cas") > 3)).show()
# -
# **2. Display 10 crash events with highest casualties.**
# to show top ten crashes
crashes_df.orderBy("Total Cas", ascending=False).show(10)
# **3. Find the total number of fatalities for each crash type**
# to convert 'Total Fats' column to integer type
crashes_df = crashes_df.withColumn("Total Fats", crashes_df["Total Fats"].cast(IntegerType()))
# to group the different crash types and display their total fatalities
crash_group_df = crashes_df.groupBy('Crash Type')
crash_group_df.agg(sum('Total Fats').alias('Number_of_Fatalities')).orderBy("Number_of_Fatalities", ascending=False).show()
# **4. Find the total number of casualties for each suburb when the vehicle was driven by an unlicensed driver. You are required to display the name of the suburb and the total number of casualties.**
#
# +
# join the two dataframes
joined_df = units_df.join(crashes_df, units_df.REPORT_ID==crashes_df.REPORT_ID,how='inner')
# to display the casualities when the driver is unlicenced
joined_df.select("Suburb", "Total Cas")\
.where((col("Licence Type") == 'Unlicenced'))\
.groupBy('Suburb').agg(sum('Total Cas').alias('Number_of_Casualities'))\
.orderBy("Number_of_Casualities", ascending=False)\
.show()
# -
# **1. Find the total number of crash events for each severity level. Which severity level is the most common?**
#
# to find the total number of crash events for each severity level
crashes_df.groupBy("CSEF Severity").agg(count("*").alias('Number_of_events'))\
.orderBy('Number_of_events',ascending=False).show()
# **2. Compute the total number of crash events for each severity level and the percentage for the four different scenarios.**
# **a. When the driver is tested positive on drugs**
# to get the required conditions in dataframe
crash_count_df = crashes_df.select('CSEF Severity')\
.where(col("Drugs Involved") == 'Y')\
.groupBy("CSEF Severity").agg(count("*").alias('Count'))
# used the below links to get strings in required format
#https://stackoverflow.com/a/43992110
#https://sparkbyexamples.com/spark/usage-of-spark-sql-string-functions/
total = crash_count_df.select("Count").agg({"Count": "sum"}).collect().pop()['sum(Count)']
crash_count_df.withColumn('Percentage', format_string("%2.3f%%", round((crash_count_df['Count']/total * 100),2))).show()
# **b. When the driver is tested positive for blood alcohol concentration.**
# to get the required conditions in dataframe
crash_alcohol_df = crashes_df.select('CSEF Severity')\
.where(col('DUI Involved') != '')\
.groupBy("CSEF Severity").agg(count("*").alias('Count'))
#to get the output in required format
total = crash_alcohol_df.select("Count").agg({"Count": "sum"}).collect().pop()['sum(Count)']
crash_alcohol_df.withColumn('Percentage', format_string("%2.3f%%", round((crash_alcohol_df['Count']/total * 100),2))).show()
# **c. When the driver is tested positive for both drugs and blood alcohol**
# to get the required conditions in dataframe
crash_both_df = crashes_df.select('CSEF Severity')\
.where((col('DUI Involved') != '') & (col("Drugs Involved") == 'Y'))\
.groupBy("CSEF Severity").agg(count("*").alias('Count'))
# to get the output in required format
total = crash_both_df.select("Count").agg({"Count": "sum"}).collect().pop()['sum(Count)']
crash_both_df.withColumn('Percentage', format_string("%2.3f%%", round((crash_both_df['Count']/total * 100),2))).show()
# **d. When the driver is tested negative for both (no alcohol and no drugs).**
# to get the required conditions in dataframe
negative_crash_df = crashes_df.select('CSEF Severity')\
.where((col('DUI Involved').isNull()) & (col("Drugs Involved").isNull()))\
.groupBy("CSEF Severity").agg(count("*").alias('Count'))
# to get the output in required format
total = negative_crash_df.select("Count").agg({"Count": "sum"}).collect().pop()['sum(Count)']
negative_crash_df.withColumn('Percentage', format_string("%2.3f%%", round((negative_crash_df['Count']/total * 100),2))).show()
# **1. Find the Date and Time of Crash, Number of Casualties in each unit and the Gender, Age, License Type of the unit driver for the suburb "Adelaide".**
# +
# %%time
# Spark RDD
req_crashes_rdd = filt_crashes_rdd.map(lambda x: (x[0].replace('"',''),(x[2].replace('"',''),x[10].replace('"',''),\
x[11].replace('"',''),x[12].replace('"',''),x[13].replace('"',''),x[6].replace('"',''))))
req_unit_rdd = filt_units_rdd.map(lambda x: (x[0].replace('"',''),(x[7].replace('"',''), x[8].replace('"',''),\
x[11].replace('"',''))))
# join the rdd on key
join_rdd = req_crashes_rdd.join(req_unit_rdd).filter(lambda x: x[1][0][0] == 'ADELAIDE')
join_rdd.map(lambda x: (x[1][0][1] +"-"+ x[1][0][2] +"-"+ x[1][0][3], x[1][0][4], x[1][0][5],\
x[1][1][0], x[1][1][1],x[1][1][2])).collect()
# +
# %%time
# Spark DataFrame
# join the two dataframes
joined_df = units_df.join(crashes_df, units_df.REPORT_ID==crashes_df.REPORT_ID,how='inner')
# used the below link to find out concat function
#https://www.edureka.co/community/2280/concatenate-columns-in-apache-spark-dataframe
joined_df.select(concat("Year", lit("-"), "Month", lit("-"),"Day").alias("Date"), "Time", "Total Cas", "Sex", "Age", "Licence Type")\
.where(col("Suburb")=='ADELAIDE').show()
# +
# %%time
# Spark SQL
# Create Views from Dataframes
crashes_df.createOrReplaceTempView("sql_crashes")
units_df.createOrReplaceTempView("sql_units")
# perform the sql operations
spark.sql('''
SELECT Year || '-' || Month||'-'|| Day AS Date, Time, `Total Cas`, Sex, Age, `Licence Type`
FROM sql_crashes d JOIN sql_units w ON d.REPORT_ID=w.REPORT_ID
WHERE d.Suburb == 'ADELAIDE'
''').show()
# -
# **2. Find the total number of casualties for each suburb when the vehicle was driven by an unlicensed driver. You are required to display the name of the suburb and the total number of casualties.**
# +
# %%time
# Spark DataFrame
# join the two dataframes
joined_df = units_df.join(crashes_df, units_df.REPORT_ID==crashes_df.REPORT_ID,how='inner')
joined_df.select("Suburb", "Total Cas")\
.where((col("Licence Type") == 'Unlicenced'))\
.groupBy('Suburb').agg(sum('Total Cas').alias('Number_of_Casualities'))\
.orderBy("Number_of_Casualities", ascending=False)\
.show()
# +
# %%time
# Spark SQL
# Create Views from Dataframes
crashes_df.createOrReplaceTempView("sql_crashes")
units_df.createOrReplaceTempView("sql_units")
spark.sql('''
SELECT Suburb, sum(`Total Cas`) AS Number_of_Casualities
FROM sql_crashes d JOIN sql_units w ON d.REPORT_ID=w.REPORT_ID
WHERE w.`Licence Type` == 'Unlicenced'
GROUP BY Suburb
ORDER BY Number_of_Casualities desc
''').show()
| Pyspark - Road Crash Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SciViz!
# ## 2. Using yt
import yt
# We'll use a dataset originally from the yt hub: http://yt-project.org/data/
#
# Specifically, we'll use the IsolatedGalaxy dataset: http://yt-project.org/data/IsolatedGalaxy.tar.gz
#
# Now, lets grab a dataset & upload it. Here's where mine is stored (in data):
ds = yt.load("/Users/jillnaiman/Downloads/IsolatedGalaxy/galaxy0030/galaxy0030")
# Print out various stats of this dataset:
ds.print_stats()
# This is basically telling us something about the number of data points in the dataset. Don't worry if you don't know what levels, grids or cells are at this point we'll get to it later.
# Same thing with field list, its cool if some of these look less familiar then others:
ds.field_list
ds.derived_field_list
# This is a 3D simululation of a galaxy, lets check out some stats about the box:
ds.domain_right_edge, ds.domain_left_edge
# What this is saying is the box goes from (0,0,0) to (1,1,1) in "code_length" units. Basically, this is just a normalized box.
# You can also do fun things like print out max & min densities:
ds.r[:].max("density"), ds.r[:].min("density")
# The above is for the whole box.
#
# We can also ask where the maximum density is in this simulation box:
ds.r[:].argmax("density")
# So this gives us x/y/z positions for where the maximum density is.
#
# Ok, lets make a quick plot 1/2 down the z-direction.
# +
# if the plot is too big for class try:
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [3, 3]
p = ds.r[:, :, 0.5].plot("density")
# -
# Let's zoom:
p.zoom(10)
# So, unless you're an astronomer you might be a little confused about these "kpc" units. But yt allows us to change them! Behold cool yt units things:
(yt.units.kpc).in_units("cm")
# So we have now changed these weird kpc units.
#
# yt also can do cool things with units like, `yt.units` figures out some math stuff like, making things into cubed cm:
(yt.units.kpc**3).in_units("cm**3")
# So let's set some units of our plot! Let's change the units of density from $g/cm^3$ to $kg/m^3$
p.set_unit("density","kg/m**3")
# We can also include annotations on this plot:
p.annotate_velocity()
# This shows how material is moving in this simulation this is shown with velocity vectors.
#
# We can combine some of our coding around finding max values of density and combine with some region plots.
#
# Let's project the maximum density along the z axis i.e. lets make a plot of the maximum density along the z-axis of our plot:
p2 = ds.r[:].max("density", axis="z").plot()
# We can zoom this as well:
p2.zoom(10)
# If we scroll back up we can see that there is indeed a different between this and our slice plot. Here, we are much more "smeared" since we're picking only the max density $\rightarrow$ everything looks brighter.
#
# We can also do plots based on region selection but over specific values of z (and x & y). If we recall our box goes from 0$\rightarrow$1 in each x/y/z direction, we can plot a zoom in like so:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].max("density", axis="z").plot()
# So, this shows the maximum density but only in a thin slice of the z-axis which is offset from the center.
#
# Since the galaxy lives at the center, and is the highest density gas region, it makes sense that our densities are lower and our features look different -- more "fuzzy ball" outside of the galaxy then gas flowing onto a galaxy disk.
#
# Let's redo the same plot but for the temperature of the gas:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].mean("temperature", axis="z").plot()
# We might want to highlight the temperature of the most dense regions. Why? Well maybe we want to, instead of depicting the straight temperature, we want to depict the temperature of the *majority of the gas*. We can do this by specifying a "weight" in our projection:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].mean("temperature", weight="density", axis="z").plot()
# So why is there this blocky structure? In space, we don't see cubes around galaxies... yet anyway...
#
# This is becuase this is a simulation of a galaxy, not an actual galaxy. We can show why this might be by plotting the "grids" of this simulation over this thing:
p.annotate_grids()
# From this we can see that our grids sort of align where the temperature looks funny. This is a good indicator that we have some numerical artifacts in our simulation.
#
# Ok! Let's try some more analysis-like plots some of the helpful yt included plots is:
ds.r[:].profile("density", "temperature").plot()
# So this is plotting the temperature of the gas in our simulation, in each binned density.
#
# In our actual simulation, we have temperaturates at a variety of densities, and this is usually the case, so by default what is plotted is the temperature (our 2nd param) plotted at each density bin, but weighted by the mass of material (gas) in each cell.
#
# We can weight by other things, like in this case density:
ds.r[:].profile("density", "temperature", weight_field="density").plot()
# So, similar shape (since mass and density are related) but a little different.
# # Activity #2: Brain data with yt
#
# We can also use yt to play with other sorts of data:
import h5py # might have to pip install
# Let's read our datafile into something called "scan_data":
with h5py.File("/Users/jillnaiman/Downloads/single_dicom.h5", "r") as f:
scan_data = f["/scan"][:]
# If we recall, we had a weird shape of this data:
scan_data.shape
# So to import this data into yt to have yt make images for us, we need to do some formatting with numpy:
import numpy as np
dsd = yt.load_uniform_grid({'scan': scan_data},
[36, 512, 512],
length_unit = yt.units.cm, # specify the units of this dataset
bbox = np.array([[0., 10], [0, 10], [0, 10]]), # give a "size" to this dataset
)
dsd.r[:].mean("scan", axis="y").plot(); # this takes the mean along the specified axis "y" and plots
# Can also do .max or .min
#
# Note here that the number of fields available is much less:
dsd.field_list
# We can also look at different potions of the z-y axis by specifying the x-axis:
p = dsd.r[0.75,:,:].plot('scan')
# # Activity #3: Output images and objects (3D) with yt
# Note: we'll do more with 3D objects next week/the last week, but this is a good first view of some cool ways we can output objects with yt.
#
# Let's go back to to our galaxy object and make a surface.
#
# First, we'll cut down to a sphere and check that out:
sphere = ds.sphere("max", (500.0, "kpc"))
sphere.mean("density", axis="y").plot(); # this takes the mean along the specified axis "y" and plots
# Let's generate a surface of constant density i.e. we'll connect points on a surface where the density has a single value:
surface = ds.surface(sphere, "density", 1e-27)
surface.export_obj('/Users/jillnaiman/Downloads/myGalFiles',color_field='temperature')
# the above might take a while
# At this point you can upload this to SketchFab, or use PyGEL3D if you were able to install this.
#
# #### If you have PyGEL3D installed:
# +
# for checking out our surfaces right here
#http://www2.compute.dtu.dk/projects/GEL/PyGEL/
# #!pip install PyGEL3D
# you might have to link where pip installs things
# you can find this in your activated DataViz environment with `pip show PyGEL3D`
from sys import path
path.append('/Users/jillnaiman/opt/anaconda3/lib/python3.7/site-packages/')
# +
from PyGEL3D import gel
from PyGEL3D import js
# for navigating
js.set_export_mode()
m = gel.obj_load("/Users/jillnaiman/Downloads/myGalFiles.obj")
viewer = gel.GLManifoldViewer()
viewer.display(m)
# press ESC to quit? Yes, but then it takes a while so
# -
# to get rid of the window
del viewer
# Now, lets try with an inline viewer -- also display in the notebook:
# +
import numpy as np # if you haven't yet
#js.display(m,wireframe=False)
# comment out after you've run since we'll re-run below
# -
# Now let's try with an inline viewer & data colors:
surf_temp = surface['temperature']
surf_temp.shape
# We see that this is infact a long list of values temperatures on each surface *face*.
#
# If we look at the shape of the object:
m.positions().shape, surf_temp.shape[0]*3
# We see we have (surf_temp.shape)X3 times the number of points in x/y/z. This is because these are *vertex* values. So, if we want to color by something, we should use 3X the number of faces.
js.display(m, data=np.repeat(np.log10(surf_temp),3),wireframe=False)
# We can also process for 3D printing:
surface.export_obj('/Users/jillnaiman/Downloads/myGalFiles_print',dist_fac=0.001)
# ## Outputing images for things like clothing
p = ds.r[:, :, 0.5].plot("density")
p.zoom(20)
myImage = p.frb # fixed resoltuion binary
# We can then grab a simple image array:
plt.imshow(np.array(myImage['density']))
# ... or we can turn off labels and grab a lovely image:
p = ds.r[:, :, 0.5].plot("density")
p.zoom(10)
p.hide_colorbar(); p.hide_axes();
p
# Save the image:
p.save('/Users/jillnaiman/Downloads/myImage.png')
# Now you have a lovely image that you can upload and put on things like sweaters or whatnot.
| week15/prep_notebook_week15_part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/intro/colab_intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="24_OboyL7tqe"
# # Introduction to colab
#
# This notebook illustrates how to install the PML code and various other libraries. More details in the [official documentation](https://colab.research.google.com/notebooks/intro.ipynb) and the [official introduction](https://colab.research.google.com/notebooks/basic_features_overview.ipynb).
#
# + colab={"base_uri": "https://localhost:8080/"} id="ZjFsGQJ41k32" outputId="fe9e76a3-ee97-41ca-ad44-144cdff8d06e"
IS_COLAB = ('google.colab' in str(get_ipython()))
print(IS_COLAB)
# + [markdown] id="8lAbDqny-vDq"
# # How to import standard libraries
# + [markdown] id="XHO2_uKXMbD4"
# Colab comes with most of the packages we need pre-installed.
#
#
#
#
# + id="B4KQOCig_xf1"
# Standard Python libraries
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import time
import glob
from typing import Any, Callable, Dict, Iterator, Mapping, Optional, Sequence, Tuple
# + [markdown] id="U9PghW_NT1HY"
# To install new packages, use the following (see [this page](https://colab.research.google.com/notebooks/snippets/importing_libraries.ipynb) for details):
#
# ```
# # # !pip install foo
# ```
#
#
# + [markdown] id="fOBdg02-_Jws"
# ## Numpy
# + id="AzP2LAtN_L1m" colab={"base_uri": "https://localhost:8080/"} outputId="775ab507-3a70-4816-d76e-e2d463aa571d"
import numpy as np
np.set_printoptions(precision=3)
A = np.random.randn(2,3)
print(A)
# + [markdown] id="76jPgsuk_1IP"
# ## Pandas
# + id="GimloDqo_4No" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="1ddca722-d652-4b44-c4e0-982c722dabeb"
import pandas as pd
pd.set_option('precision', 2) # 2 decimal places
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', 30)
pd.set_option('display.width', 100) # wide windows
url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data'
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Year', 'Origin', 'Name']
df = pd.read_csv(url, names=column_names, sep='\s+', na_values="?")
df.head()
# + [markdown] id="hUCC261x_7zZ"
# ## Sklearn
# + id="RCSwx_lE_7Jn" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="4036c988-bb39-4ad4-d9af-f45bc6330231"
import sklearn
from sklearn.datasets import load_iris
iris = load_iris()
# Extract numpy arrays
X = iris.data
y = iris.target
import matplotlib.pyplot as plt
plt.scatter(X[:,0], X[:,1])
# + [markdown] id="PJXF4csdBhsN"
# ## JAX
# + id="8JiSxcJJ79Bv" colab={"base_uri": "https://localhost:8080/"} outputId="97f3ebb7-93b8-4255-bccb-781d01ccb4cc"
# JAX (https://github.com/google/jax)
import jax
import jax.numpy as jnp
A = jnp.zeros((3,3))
# Check if JAX is using GPU
print("jax backend {}".format(jax.lib.xla_bridge.get_backend().platform))
# + [markdown] id="l99YLyorBdYE"
# ## Tensorflow
# + colab={"base_uri": "https://localhost:8080/"} id="StpReaSICLUm" outputId="5dfac104-8e5a-49cd-8400-eadb56650acd"
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
print("tf version {}".format(tf.__version__))
print([d for d in tf.config.list_physical_devices()])
if not tf.config.list_physical_devices('GPU'):
print("No GPU was detected. DNNs can be very slow without a GPU.")
if IS_COLAB:
print("Go to Runtime > Change runtime and select a GPU hardware accelerator.")
# + [markdown] id="grUUK1GrBfIY"
# ## PyTorch
# + id="Oi4Zmzla73A_" colab={"base_uri": "https://localhost:8080/"} outputId="558ed73a-5dfe-491a-be1c-64c949283e48"
import torch
import torchvision
print("torch version {}".format(torch.__version__))
if torch.cuda.is_available():
print(torch.cuda.get_device_name(0))
else:
print("Torch cannot find GPU")
# + [markdown] id="M4ayVFuc0FD9"
# # Plotting
#
# Colab lets you make static plots using matplotlib, as shown below.
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="j_k4tv4D1VaC" outputId="dda6cc16-8351-4aff-9c09-ab7ad5dc968f"
import matplotlib.pyplot as plt
import PIL
import imageio
import seaborn as sns;
sns.set(style="ticks", color_codes=True)
from IPython import display
plt.figure()
plt.plot(range(10))
plt.savefig('myplot.png')
# + [markdown] id="qLh3fxl63IHW"
# Colab also lets you create interactive plots using various javascript libraries - see [here](https://colab.research.google.com/notebooks/charts.ipynb#scrollTo=QSMmdrrVLZ-N) for details.
# + [markdown] id="7aCgO-moU2WA"
# # Accessing local files
#
# Clicking on the file folder icon on the left hand side of colab lets you browse local files. Right clicking on a filename lets you download it to your local machine. Double clicking on a file will open it in the file viewer/ editor, which appears on the right hand side.
#
# The result should look something like this:
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-image-viewer.png?raw=true">
#
#
# You can also use standard unix commands to manipulate files, as we show below.
# + colab={"base_uri": "https://localhost:8080/"} id="dRh4BOIxHpEX" outputId="104c8f4f-9eff-4b9b-fc81-c084740e475b"
# !pwd
# + colab={"base_uri": "https://localhost:8080/"} id="T7i8bvaghwy7" outputId="bb0012d2-bc82-4d5d-dfe3-acf2ba3e77a7"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="HDNijfMPjPsE" outputId="d2bb4d90-86fc-44b2-9900-6050507e082b"
# !echo 'foo bar' > foo.txt
# !cat foo.txt
# + [markdown] id="Ell60Ff4UE-P"
# You can open text files in the editor by clicking on their filename in the file browser, or programmatically as shown below.
# + colab={"base_uri": "https://localhost:8080/", "height": 16} id="CcxDCpyYjhfU" outputId="d521b728-16e6-43d7-f8e9-2c2d3c0b8eef"
from google.colab import files
files.view('foo.txt')
# + [markdown] id="PMet3XdcVF9O"
# If you make changes to a file containing code, the new version of the file will not be noticed unless you use the magic below.
# + colab={"base_uri": "https://localhost:8080/"} id="0ufY8AO1VEUh" outputId="13aeca9f-992b-4881-98d4-89508b0ad23a"
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="6a6nkLsKWQpu"
# # Syncing with Google drive
#
# Files that you generate in, or upload to, colab are ephemeral, since colab is a temporary environment with an idle timeout of 90 minutes and an absolute timeout of 12 hours (24 hours for Colab pro). To save any files permanently, you need to mount your google drive folder as we show below. (Executing this command will open a new window in your browser - you need cut and paste the password that is shown into the prompt box.)
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="cYZpcMiQkl15" outputId="b9e259a1-c252-48c0-cdd8-72f97531c2f0"
from google.colab import drive
drive.mount('/content/gdrive')
# !pwd
with open('/content/gdrive/MyDrive/foo.txt', 'w') as f:
f.write('Hello Google Drive!')
# !cat /content/gdrive/MyDrive/foo.txt
# + [markdown] id="CeGz5P_RUyzG"
# To ensure that local changes are detected by colab, use this piece of magic.
# + id="K6RR9dfoUyMG"
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="buZsxpmUS37n"
# # Working with github
#
# It is possible to download code (or data) from githib into a local directory on this virtual machine. It is also possible to upload local files back to github, although that is more complex. See details below.
# + [markdown] id="rVvGT6GUBg2Q"
# ## Cloning a repo from github
#
# Below we show how to clone the repo for this book, so you can access its code and data.
# + id="uVZWqzdW7_ZG" colab={"base_uri": "https://localhost:8080/"} outputId="fa8feb47-d11b-49c1-c92c-6a96ef874985"
# !rm -rf pyprobml # Remove any old local directory to ensure fresh install
# !git clone https://github.com/probml/pyprobml
# + colab={"base_uri": "https://localhost:8080/"} id="XdC34HzKT8L8" outputId="86394ce4-76e0-4802-8917-f3e142f3cba9"
# !ls
# + [markdown] id="HuplkSqtkDuR"
# We can access data as shown below.
# + colab={"base_uri": "https://localhost:8080/"} id="_8m9VQ-3kFcq" outputId="4f078099-482d-4f09-c1b1-a950cd703d33"
datadir = 'pyprobml/data'
import re
fname = os.path.join(datadir, 'timemachine.txt')
with open(fname, 'r') as f:
lines = f.readlines()
sentences = [re.sub('[^A-Za-z]+', ' ', st).lower().split()
for st in lines]
for i in range(5):
words = sentences[i]
print(words)
# + [markdown] id="MNWWINngc5rn"
# We can run any script as shown below.
# (Note we first have to define the environment variable for where the figures will be stored.)
# + colab={"base_uri": "https://localhost:8080/", "height": 856} id="aYXkQP-DdApw" outputId="76b33a43-44a0-4539-c1a4-91151c38f803"
import os
os.environ['PYPROBML']='pyprobml'
# %run pyprobml/scripts/activation_fun_plot.py
# + [markdown] id="YTT5eJ_qUDFe"
# We can also import code, as we show below.
# + colab={"base_uri": "https://localhost:8080/"} id="6rTAz3nECmZr" outputId="e63dde3d-9c4c-4928-f6f7-4a359230cb6c"
#os.chdir('pyprobml/scripts')
import pyprobml.scripts.pyprobml_utils as pml
pml.pyprobml_test()
# + [markdown] id="yaISmcnNmnS7"
# ## Pushing local files back to github
#
# You can easily save your entire colab notebook to github by choosing 'Save a copy in github' under the File menu in the top left. But if you want to save individual files (eg code that you edited in the colab file editor, or a bunch of images or data files you created), the process is more complex.
#
# You first need to do some setup to create SSH keys on your current colab VM (virtual machine), manually add the keys to your github account, and then copy the keys to your mounted google drive so you can reuse the same keys in the future. This only has to be done once.
#
# After setup, you can use the `git_command` function we define below to securely execute git commands. This works by copying your SSH keys from your google drive to the current colab VM, executing the git command, and then deleting the keys from the VM for safety.
#
# + [markdown] id="0gOzFmcKoUuO"
# ### Setup
#
# Follow these steps. (These instructions are text, not code, since they require user interaction.)
#
# ```
# # # !ssh-keygen -t rsa -b 4096
# # # !ssh-keyscan -t rsa github.com >> ~/.ssh/known_hosts
# # # !cat /root/.ssh/id_rsa.pub
# ```
# The cat command will display your public key in the colab window.
# Cut and paste this and manually add to your github account following [these instructions](https://github.com/settings/keys).
#
# Test it worked
# ```
# # # !ssh -T <EMAIL>@github.com
# ```
#
# Finally, save the generated keys to your Google drive
#
# ```
# from google.colab import drive
# drive.mount('/content/drive')
# # # !mkdir /content/drive/MyDrive/ssh/
# # # !cp -r ~/.ssh/* /content/drive/MyDrive/ssh/
# # # !ls /content/drive/MyDrive/ssh/
# ```
#
# + [markdown] id="oSiVyBG1xm44"
# ### Test previous setup
#
# Let us check that we can see our SSH keys in our mounted google drive.
# + colab={"base_uri": "https://localhost:8080/"} id="cCUxHiHAxcY2" outputId="a60a0325-da4d-41d5-8f57-43ec0d119497"
from google.colab import drive
drive.mount('/content/drive')
# !ls /content/drive/MyDrive/ssh/
# + [markdown] id="C-lPchgDpD7t"
# ### Executing git commands from colab via SSH
#
# The following function lets you securely doing a git command via SSH.
# It copies the keys from your google drive to the local VM, excecutes the command, then removes the keys.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="8Vsx7hE0SCpA" outputId="7c7ed95f-2c3c-4866-f147-cb40a078acb7"
# !rm -rf git_colab.py # remove any old copies of this file
# !wget https://raw.githubusercontent.com/probml/pyprobml/master/scripts/git_colab.py
# + [markdown] id="umvOwzMfvpmU"
# Below we show how to use this. We first clone the repo to this colab VM.
# + colab={"base_uri": "https://localhost:8080/"} id="B-o1fImxucGz" outputId="06372b1d-e1c6-4a68-c5f7-e250cfb1c89c"
from google.colab import drive
drive.mount('/content/drive') # must do this before running git_colab
import git_colab as gc
# !rm -rf pyprobml # remove any old copies of this directory
# #!git clone https://github.com/probml/pyprobml.git # clones using wrong credentials
gc.git_ssh("git clone https://github.com/probml/pyprobml.git") # clone using your credentials
# + [markdown] id="ft_pJJ4ZTLdl"
# Next we add a file and push it to github.
#
# + colab={"base_uri": "https://localhost:8080/"} id="3hfluZVYTSNd" outputId="0a64118d-d176-4101-db77-2029f573aef8"
# !pwd
# !ls
# To add stuff to github, you must be inside the git directory
# %cd /content/pyprobml
# !echo 'this is a test' > scripts/foo.txt
gc.git_ssh("git add scripts; git commit -m 'push from colab'; git push")
# %cd /content
# + [markdown] id="DAqqAhzzTS7F"
# [Here](https://github.com/probml/pyprobml/blob/master/scripts/foo.txt) is a link to the file we just pushed.
#
# Finally we clean up our mess.
# + colab={"base_uri": "https://localhost:8080/"} id="r16zYiNTu_Mz" outputId="0fc340b5-4550-4466-b554-b746914ec706"
# %cd /content/pyprobml
gc.git_ssh("git rm scripts/foo*.txt; git commit -m 'colab cleanup'; git push")
# %cd /content
# + [markdown] id="q-kRtmdm5d7X"
# # Software engineering tools
#
# [<NAME> has argued](https://docs.google.com/presentation/d/1n2RlMdmv1p25Xy5thJUhkKGvjtV-dkAIsUXP-AL4ffI/edit) that notebooks are bad for developing complex software, because they encourage creating monolithic notebooks instead of factoring out code into separate, well-tested files.
#
# [<NAME> has responded to Joel's critiques here](https://www.youtube.com/watch?v=9Q6sLbz37gk&feature=youtu.be). In particular, the FastAI organization has created [nbdev](https://github.com/fastai/nbdev) which has various tools that make notebooks more useful.
#
#
# + [markdown] id="PuSsmj_fZ106"
# ## Avoiding problems with global state
#
# One of the main drawbacks of colab is that all variables are globally visible, so you may accidently write a function that depends on the current state of the notebook, but which is not passed in as an argument. Such a function may fail if used in a different context.
#
# One solution to this is to put most of your code in files, and then have the notebook simply import the code and run it, like you would from the command line. Then you can always run the notebook from scratch, to ensure consistency.
#
# Another solution is to use the [localscope](https://localscope.readthedocs.io/en/latest/README.html) package can catch some of these errors.
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Q0FmEeIgc0YI" outputId="0d71c4a8-a4fb-45bd-e137-c391e408c83f"
# !pip install localscope
# + id="9zfUiUB8d-jh"
from localscope import localscope
# + colab={"base_uri": "https://localhost:8080/"} id="wI5wXzUPdlOS" outputId="500dfc1f-1cbc-4fa6-a30e-02f32258fa31"
a = 'hello world'
def myfun():
print(a) # silently accesses global variable
myfun()
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="T3V1iV32czq8" outputId="b346cb7a-f252-41bc-992b-dd72e5aec825"
a = 'hello world'
@localscope
def myfun():
print(a)
myfun()
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="5t48_AMbAN8V" outputId="71fdaacf-4650-4a52-9fb5-da5873c33e73"
def myfun2():
return 42
@localscope
def myfun3():
return myfun2()
# + colab={"base_uri": "https://localhost:8080/"} id="DAqLZ8PdAquy" outputId="021e8282-a283-414c-ef1d-46521da89e49"
@localscope.mfc # allow for global methods, functions, classes
def myfun4():
return myfun2()
myfun4()
# + [markdown] id="iZaeVouoAhXP"
# ## Factoring out functionality into files stored on github
#
# The recommended workflow is to develop your code in the colab in the usual way, and when it is working, to factor out the core code into separate files. You can edit these files locally in the colab editor, and then push the code to github when ready (see details above). To run functions defined in a local file, just import them. For example, suppose we have created the file /content/pyprobml/scripts/fit_flax.py; we can use this idiom to run its test suite:
# ```
# import pyprobml.scripts.fit_flax as ff
# ff.test()
# ```
# If you make local edits, you want to be sure
# that you always import the latest version of the file (not a cached version). So you need to use this piece of colab magic first:
# ```
# # # %load_ext autoreload
# # # %autoreload 2
# ```
#
# + [markdown] id="DdXlYCe1AlJa"
#
# ## File editors
#
# Colab has a simple file editor, illustrated below for an example file.
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-file-editor.png?raw=true">
#
# + [markdown] id="4qi2xKMbAnlj"
#
# ## VScode
# The default colab file editor is very primitive.
# See [this article](https://amitness.com/vscode-on-colab/) for how to run VScode
# from inside your Colab browser. Unfortunately this is a bit slow. It is also possible to run VScode locally on your laptop, and have it connect to colab via SSH, but this is more complex (see [this medium post](https://medium.com/@robertbracco1/configuring-google-colab-like-a-pro-d61c253f7573#4cf4) for details).
#
# + [markdown] id="v0G2d13kIEz5"
# # GPUs
#
# If you select the 'Runtime' menu at top left, and then select 'Change runtime type' and then select 'GPU', you can get free access to a GPU.
#
#
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-change-runtime.png?raw=true" height=300>
# <img src="https://github.com/probml/pyprobml/blob/
# master/book1/intro/figures/colab-select-gpu.png?raw=true" height=200>
#
#
# + [markdown] id="MGVZB0esI0QG"
#
#
# To get access to more powerful machines (with faster GPU, TPU, more memory, and/or to use colab for up to 24h instead of 12h (and with longer idle timeouts), you can subscript to [Colab Pro](https://colab.research.google.com/signup). At the time of writing (Jan 2021), the cost is $10/month (USD). This is a good deal if you use GPUs a lot. Using my pro account, when I select large memory, I get the spec below. (The free version has roughtly half the memory and half the speed for GPUs.)
#
#
# + id="FikkXWQqBU9O" colab={"base_uri": "https://localhost:8080/"} outputId="655ec017-3552-4390-bd7e-21e48ebc4ea9"
# gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
print(gpu_info)
# + colab={"base_uri": "https://localhost:8080/"} id="GU6nII1F5S2S" outputId="d220a1a6-cc83-4718-c77e-2df38f13bf9c"
# !grep Model: /proc/driver/nvidia/gpus/*/information | awk '{$1="";print$0}'
# + colab={"base_uri": "https://localhost:8080/"} id="DXvETYYWImlV" outputId="d400b03c-52e7-4f21-b30d-b38251c4b18d"
from psutil import virtual_memory, cpu_count
ram_gb = virtual_memory().total / 1e9
print('RAM (GB)', ram_gb)
print('num cores', cpu_count())
# + id="TiHgJVbyJ2Br" colab={"base_uri": "https://localhost:8080/"} outputId="cdcd6ac6-c3bf-4fc4-b8bf-b9bc307a7930"
# !cat /proc/version
# + colab={"base_uri": "https://localhost:8080/"} id="FAO3U-1DZZm8" outputId="22503da3-d169-450f-bd89-e47bab67488e"
# !cat /proc/cpuinfo
# + colab={"base_uri": "https://localhost:8080/"} id="Ijp8wNZgZfAY" outputId="5ee7242b-2f1a-42ac-a99b-504987c598c7"
# !cat /proc/meminfo
# + id="hzi1OpMaZlAc"
| book1/intro/colab_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib widget
import sys
sys.path.append("..")
import os
import numpy as np
import spotlob
from spotlob.register import PROCESS_REGISTER as prg
filename = os.path.abspath("testdata5.JPG")
gui = spotlob.make_gui(filename)
@spotlob.use_in(gui)
@prg.binarization_plugin([("lower_threshold",(0,255,100)),
("upper_threshold",(0,255,200)),
("invert", True)])
def my_threshold(image, lower_threshold, upper_threshold, invert):
out = np.logical_and( image > lower_threshold,
image < upper_threshold)
out = out.astype(np.uint8)*255
if invert:
out = ~out
return out
spotlob.show_gui(gui)
gui.results().describe()
| notebooks/gui_extension_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import numpy as np
import matplotlib.pyplot as plt
from sklearn import cluster, datasets
from sklearn.neighbors import kneighbors_graph
from sklearn.preprocessing import StandardScaler
np.random.seed(0)
# Generate datasets. We choose the size big enough to see the scalability
# of the algorithms, but not too big to avoid too long running times
n_samples = 800
noisy_circles = datasets.make_circles(n_samples=n_samples, factor=.5,
noise=.1)
noisy_moons = datasets.make_moons(n_samples=n_samples, noise=.05)
blobs = datasets.make_blobs(n_samples=n_samples, random_state=8)
no_structure = np.random.rand(n_samples, 2), None
colors = np.array([x for x in 'bgrcmykbgrcmykbgrcmykbgrcmyk'])
colors = np.hstack([colors] * 20)
# clustering_names = [
# 'MiniBatchKMeans', 'AffinityPropagation', 'MeanShift',
# 'SpectralClustering', 'Ward', 'AgglomerativeClustering',
# 'DBSCAN', 'Birch']
clustering_names = [
# 'MiniBatchKMeans', 'AffinityPropagation', 'MeanShift',
'Ward', 'Average', 'single', 'complete'
]
plt.figure(figsize=(len(clustering_names) * 2 + 3, 9.5))
plt.subplots_adjust(left=.02, right=.98, bottom=.001, top=.96, wspace=.05,
hspace=.01)
plot_num = 1
# datasets = [noisy_circles, noisy_moons, blobs, no_structure]
datasets = [noisy_moons]
for i_dataset, dataset in enumerate(datasets):
X, y = dataset
# normalize dataset for easier parameter selection
X = StandardScaler().fit_transform(X)
# estimate bandwidth for mean shift
bandwidth = cluster.estimate_bandwidth(X, quantile=0.3)
# connectivity matrix for structured Ward
connectivity = kneighbors_graph(X, n_neighbors=4, include_self=False)
# make connectivity symmetric
connectivity = 0.5 * (connectivity + connectivity.T)
# create clustering estimators
ms = cluster.MeanShift(bandwidth=bandwidth, bin_seeding=True)
two_means = cluster.MiniBatchKMeans(n_clusters=2)
ward = cluster.AgglomerativeClustering(n_clusters=2, linkage='ward',
connectivity=connectivity)
spectral = cluster.SpectralClustering(n_clusters=2,
eigen_solver='arpack',
affinity="nearest_neighbors")
dbscan = cluster.DBSCAN(eps=.2)
affinity_propagation = cluster.AffinityPropagation(damping=.9,
preference=-200)
average_linkage = cluster.AgglomerativeClustering(
linkage="average", affinity="euclidean", n_clusters=2,
connectivity=connectivity)
single_linkage = cluster.AgglomerativeClustering(
linkage="single", affinity="euclidean", n_clusters=2,
connectivity=connectivity)
complete_linkage = cluster.AgglomerativeClustering(
linkage="complete", affinity="euclidean", n_clusters=2,
connectivity=connectivity)
birch = cluster.Birch(n_clusters=2)
# clustering_algorithms = [
# two_means, affinity_propagation, ms, spectral, ward, average_linkage,
# dbscan, birch]
clustering_algorithms = [ward, average_linkage, single_linkage, complete_linkage]
for name, algorithm in zip(clustering_names, clustering_algorithms):
# predict cluster memberships
t0 = time.time()
algorithm.fit(X)
t1 = time.time()
if hasattr(algorithm, 'labels_'):
y_pred = algorithm.labels_.astype(np.int)
else:
y_pred = algorithm.predict(X)
# plot
plt.subplot(4, len(clustering_algorithms), plot_num)
if i_dataset == 0:
plt.title(name, size=18)
plt.scatter(X[:, 0], X[:, 1], color=colors[y_pred].tolist(), s=10)
if hasattr(algorithm, 'cluster_centers_'):
centers = algorithm.cluster_centers_
center_colors = colors[:len(centers)]
plt.scatter(centers[:, 0], centers[:, 1], s=100, c=center_colors)
plt.xlim(-2, 2)
plt.ylim(-2, 2)
plt.xticks(())
plt.yticks(())
plt.text(.99, .01, ('%.2fs' % (t1 - t0)).lstrip('0'),
transform=plt.gca().transAxes, size=15,
horizontalalignment='right')
plot_num += 1
plt.show()
# -
plt.show()
| twomoon_simulation/moon_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This is a small tutorial on data cleaning, preprocessing and the idea of a pipeline for changing raw data to clean data. The primary purpose of this tutorial is to showcase benefits of my package 'preprocessor' in the above mentioned scenarios.
from preprocessor.misc import read_csv
import numpy as np
import pandas as pd
# ### Using our read_csv is a wrapper over pandas function of same name which is better at reading datetime columns
data = read_csv("example.csv", verbose =True, encoding = 'latin')
data.head()
# ### Lets look at values in each column
data['mixed'].value_counts()
# #### On first glance we can tell that the first column 'mixed' has very small and large numbers as well as text in some of its values, now lets look at second column
data['cat'].value_counts()
# #### The second column look like a regular categorical column, however, we can see there are some values which have just one occurance
data['date'].value_counts()[:10]
# #### The third column look like a regular datetime column, but lets say we want to use this in a machine learning model, in that we cant use dates in their raw form
data['num'].value_counts()
# #### Finally a regular numerical column, with nans ofcourse
# ### Now lets use our preprocessor package to create a piple line for this data
verbose = True
# +
# First lets try to deal with the mixed column(s)
from preprocessor.feature_extractor import extract_numericals_forall
# extract_numericals_forall goes through all categorical columns and tries to see if it is a mixed column
# if it is, it creates a new column with the column name + '_numerical' as suffix and puts all numerical
# values in the new column
df1 = extract_numericals_forall(data,verbose =verbose)
df1.head()
# -
# #### We see we were able to generate two new numerical columns, lets analyze them
print(df1['mixed_numerical'].value_counts(), df1['cat_numerical'].value_counts().nlargest(10))
# #### We see that mixed_numerical looks like a sensible column while on first glance cat_numerical seems to be constructed of values in cat column which just happen to be interpretable as numericals. Therefore we decide to just extract numericals from one column instead of all categorical columns in our pipleline
# +
from preprocessor.feature_extractor import extract_numericals
df1 = extract_numericals(data,col = 'mixed',verbose =verbose)
df1.head()
# -
# #### Now that we have extracted the numbers from 'mixed', we want to remove numericals from the actual 'mixed' column in order to make it purely categorical
# +
from preprocessor.imputer import remove_numericals_from_categories
# Delete all numericals from 'mixed' column
df2 = remove_numericals_from_categories(df1,include=['mixed'],verbose =verbose)
df2['mixed'].value_counts()
# -
# #### At this point we can assume that 'mixed' is not a useful column anymore so we can delete it, since our df2 object is a regular pandas data structure we can use all pandas functions without any problem
df3 = df2.drop(['mixed'], axis=1)
df3.head()
# #### Awsome, now that we have dealt with 'mixed', lets focus on date column now
# +
# Lets try to extract some features from the date column
from preprocessor.feature_extractor import extract_datetime_features_forall
from preprocessor.imputer import remove_datetimes
# extract_datetime_features_forall goes through all datetime columns and tries to extract 15 predefined features from them
df4 = extract_datetime_features_forall(df3,verbose =verbose)
# since we have the features extracted, no need for keeping datetime columns anymore
df4 = remove_datetimes(df4, verbose = verbose)
df4.head()
# -
# #### Great, so we have extracted 15 features from one datetime column, and we could have extracted n*15 where n is number of date time columns in the data
# #### Next, Lets try to see if we have any infs or -infs in the data, lets say for now I want to make a new feature column for whenever we find infs in a column
# +
# Lets try to extract some features from the date column
from preprocessor.feature_extractor import extract_is_inf_forall
# extract_is_inf_forall goes through all numerical columns and create a new column for infs if any
# if seperate_ninf is true then the new column has 3 unique values (1 for inf, 0 for no inf & -1 for ninf)
# otherwise the new column is a boolean which is true if any kind of inf was encountered
df5 = extract_is_inf_forall(df4,verbose =verbose, seperate_ninf = True)
df5.head()
# -
# #### We can see that since no infs were found in any of the data, we might conclude that we might never encounter infs in our data and hence never include this step in our final piple line
# #### Now if we remember correctly some of our numerical columns had numbers of varying range, may be outliers which might affect rest of our statistics, So lets identify outliers and remove if we find any
# +
from preprocessor.feature_extractor import extract_is_outlier_forall
# extract_is_outlier_forall goes through all numerical columns and against each column creates a new boolean column which has true if the value is marked outlier
# replace_with if None then leaves outliers intact in the actual column
df6 = extract_is_outlier_forall(df4,verbose =verbose, replace_with = np.nan)
df6.head()
# -
# #### Looks like we could only find outliers for some datetime features, depending on if we find it a valid thing in the context of our problem we can ignore or keep this step. For this specific example, in the final pipeline, I'll move this step before extracting date time features in order to don't search outliers in datetime feature columns.
# #### Now lets deal with nans in our numerical columns, we will first create _isnull column against all numerical columns to preserve the nan information
# +
from preprocessor.feature_extractor import extract_is_nan_forall
# extract_is_nan_forall goes through all numerical columns and against each column creates a new boolean column which has
# true if the value was nan, this perserves the information of nans for when we finally substitute a valid numerical
# value against all nulls
df7 = extract_is_nan_forall(df6,verbose =verbose)
df7.head()
# -
# #### Once we have all the required information preserved, lets replace nans with median for each column, means are usually susceptible to outliers (One can't be too careful!)
# +
from preprocessor.imputer import fillnans
# fillnans goes through all numerical columns and fills nans with either Mean, Median or any other provided value
df8 = fillnans(df7,verbose =verbose, by = 'median')
df8.head()
# -
# #### We have come to far, now all that remains is our categorical column, we will simply on hot encode it, however, we will use a cutoff of 100 (an educated guess, which might differ for your data). The idea is to not create new columns for a value that only occured insignificant time in the data.
# +
from preprocessor.feature_extractor import onehot_encode_all
# onehot_encode_all goes through all categorical columns and one hot encodes them
# allow_na if True means treat None or Null as a class
# onehot_encode_all drops the actual column after one hot encoding it
df9 = onehot_encode_all(df8,verbose =verbose, cutoff = 100, cutoff_class = 'other', allow_na = True)
df9.head()
# -
# #### finally looks like our data is all numbers, with no nans, no nulls and almost ready for any algorthim that operates on numbers, but wait!
# #### Some algorthims are really sensitive if features are in a very different scale of magnitude, therefore we normalize
# +
from sklearn.preprocessing import RobustScaler, StandardScaler
from preprocessor.imputer import normalize
# normalize, normalizes each column of the dataframe using provided scaler
df_scaled_standard = normalize(df9, verbose = verbose) #StandardScaler is dfault scaler
# You can read benefits of robust scaler at
# https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html
df_scaled_robust = normalize(df9, verbose = verbose, scaler = RobustScaler())
# -
# #### Lets visualize at the changes in the scales of a feature after scaling
# +
from preprocessor.plotter import plot_line
# %matplotlib inline
col = 'num'
plot_line(data,col,title = 'raw')
plot_line(df9,col,title = 'unscaled_processed')
plot_line(df_scaled_standard,col,title = 'standard scaled')
plot_line(df_scaled_robust,col,title = 'robust scaled')
# -
# #### Looking at the results we can choose which ever scale of features suits us and work with it.
# ### Final pipeline
# +
# imports
from preprocessor.feature_extractor import extract_is_nan_forall, extract_is_outlier_forall,extract_datetime_features_forall,\
extract_is_inf_forall, onehot_encode_all, extract_numericals_forall, extract_numericals
from preprocessor.imputer import fillnans, remove_datetimes, fillinfs, normalize, remove_numericals_from_categories,\
remove_single_value_features
verbose = False
df10 = data
df10 = extract_numericals(df10,col = 'mixed',verbose =verbose)
df10 = remove_numericals_from_categories(df10,include=['mixed'],verbose =verbose)
df10 = df10.drop(['mixed'], axis=1)
df10 = extract_is_outlier_forall(df10,verbose =verbose, replace_with = np.nan)
df10 = extract_datetime_features_forall(df10,verbose =verbose)
df10 = remove_datetimes(df10, verbose = verbose)
df10 = extract_is_nan_forall(df10,verbose =verbose)
df10 = fillnans(df10,verbose =verbose, by = 'median')
df10 = onehot_encode_all(df10,verbose =verbose, cutoff = 100, cutoff_class = 'other', allow_na = True)
df10 = normalize(df10, verbose = verbose)
df10.head()
# -
# #### Final Comments: We were able to make a pipeline which is able to make our 4 raw columns into 202 columns what any data analysis algorthim can use
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from IPython.display import Markdown
Markdown(filename='README.md')
# + urth={"dashboard": {}}
# %matplotlib inline
# + urth={"dashboard": {}}
import datetime
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import requests
# + urth={"dashboard": {}}
mpl.style.use('ggplot')
figsize = (14,7)
# -
now = datetime.datetime.utcnow()
print(f'This notebook was last rendered at {now} UTC')
# + [markdown] urth={"dashboard": {"hidden": true}}
# First, let's load the historical data into a DataFrame indexed by date.
# + urth={"dashboard": {}}
hits_df = pd.read_csv('ipynb_counts.csv', index_col=0, header=0, parse_dates=True)
hits_df.reset_index(inplace=True)
hits_df.drop_duplicates(subset='date', inplace=True)
hits_df.set_index('date', inplace=True)
hits_df.sort_index(ascending=True, inplace=True)
# + urth={"dashboard": {"hidden": true}}
hits_df.tail(3)
# + [markdown] urth={"dashboard": {"hidden": true}}
# There might be missing counts for days that we failed to sample. We build up the expected date range and insert NaNs for dates we missed.
# + urth={"dashboard": {}}
til_today = pd.date_range(hits_df.index[0], hits_df.index[-1])
# + urth={"dashboard": {}}
hits_df = hits_df.reindex(til_today)
# + [markdown] urth={"dashboard": {"hidden": true}}
# Now we plot the known notebook counts.
# -
fig, ax = plt.subplots(figsize=figsize)
ax.set_title(f'GitHub search hits for {len(hits_df)} days')
ax.plot(hits_df.hits, 'ko', markersize=1, label='hits')
ax.legend(loc='upper left')
ax.set_xlabel('Date')
ax.set_ylabel('# of ipynb files');
# Growth appears exponential until December 2020, at which point the count dropped suddenly and resumed growth from a new origin.
# The total change in the number of `*.ipynb` hits between the first day we have data and today is:
# + urth={"dashboard": {"hidden": true}}
total_delta_nbs = hits_df.iloc[-1] - hits_df.iloc[0]
total_delta_nbs
# + [markdown] urth={"dashboard": {"hidden": true}}
# The mean daily change for the entire duration is:
# + urth={"dashboard": {"hidden": true}}
avg_delta_nbs = total_delta_nbs / len(hits_df)
avg_delta_nbs
# + [markdown] urth={"dashboard": {"hidden": true}}
# The change in hit count between any two consecutive days for which we have data looks like the following:
# + urth={"dashboard": {}}
daily_deltas = (hits_df.hits - hits_df.hits.shift())
# -
fig, ax = plt.subplots(figsize=figsize)
ax.plot(daily_deltas, 'ko', markersize=2)
ax.set_xlabel('Date')
ax.set_ylabel('$\Delta$ # of ipynb files')
ax.set_title('Day-to-Day Change');
# The large jumps in the data are from GitHub reporting drastically different counts from one day to the next.
#
# Let's drop outliers defined as values more than two standard deviations away from a centered 90 day rolling mean.
daily_delta_rolling = daily_deltas.rolling(window=90, min_periods=0, center=True)
outliers = abs(daily_deltas - daily_delta_rolling.mean()) > 2*daily_delta_rolling.std()
outliers.value_counts()
cleaned_hits_df = hits_df.copy()
cleaned_hits_df[outliers] = np.NaN
cleaned_daily_deltas = (cleaned_hits_df.hits - cleaned_hits_df.hits.shift())
fig, ax = plt.subplots(figsize=figsize)
ax.plot(cleaned_daily_deltas, 'ko', markersize=2)
ax.set_xlabel('Date')
ax.set_ylabel('$\Delta$ # of ipynb files')
ax.set_title('Day-to-Day Change Sans Outliers');
| estimate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
#array slicing
x=np.arange(10)
x
x[2:8:2]
x[:3]
x[3:]
x[3:9]#creating a sub-array of a single dimension array
x1=np.random.randint(6, size=(2,3,4))
x1[:2,:3]
#reshaping arrays
np.arange(1,17).reshape((4,4))
| notes/notes_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Dictionaries (Data Structure)
# ===
# + [markdown] slideshow={"slide_type": "fragment"}
# Dictionaries allow us to store connected bits of information. For example, you might store a person's name and age together.
# + [markdown] slideshow={"slide_type": "skip"}
# <a name="top"></a>Contents
# ===
# - [What are dictionaries?](#what)
# - [General Syntax](#general_syntax)
# - [Example](#example)
# - [Exercises](#exercises_what)
# - [Common operations with dictionaries](#common_operations)
# - [Adding new key-value pairs](#adding_pairs)
# - [Modifying values in a dictionary](#modifying_values)
# - [Removing key-value pairs](#removing_pairs)
# - [Modifying keys in a dictionary](#modifying_keys)
# - [Exercises](#exercises_common_operations)
# - [Looping through a dictionary](#looping)
# - [Looping through all key-value pairs](#loop_all_keys_values)
# - [Looping through all keys in a dictionary](#loop_all_keys)
# - [Looping through all values in a dictionary](#loop_all_values)
# - [Looping through a dictionary in order](#looping_in_order)
# - [Exercises](#exercises_looping)
# - [Nesting](#nesting)
# - [Lists in a dictionary](#lists_in_dictionary)
# - [Dictionaries in a dictionary](#dictionaries_in_dictionary)
# - [An important note about nesting](#important_note)
# - [Exercises](#exercises_nesting)
# - [Overall Challenges](#overall_challenges)
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='what'></a>What are dictionaries?
# ===
# Dictionaries are a way to store information that is connected in some way. Dictionaries store information in *key-value* pairs, so that any one piece of information in a dictionary is connected to at least one other piece of information.
#
# Dictionaries do not store their information in any particular order, so you may not get your information back in the same order you entered it.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='general_syntax'></a>General Syntax
# ---
# A general dictionary in Python looks something like this:
# + slideshow={"slide_type": "fragment"}
dictionary_name = {key_1: value_1, key_2: value_2, key_3: value_3}
# + [markdown] slideshow={"slide_type": "fragment"}
# Since the keys and values in dictionaries can be long, we often write just one key-value pair on a line. You might see dictionaries that look more like this:
# + slideshow={"slide_type": "fragment"}
dictionary_name = {key_1: value_1,
key_2: value_2,
key_3: value_3,
}
# + [markdown] slideshow={"slide_type": "fragment"}
# This is a bit easier to read, especially if the values are long.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='example'></a>Example
# ---
# A simple example involves modeling an actual dictionary.
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# + [markdown] slideshow={"slide_type": "subslide"}
# We can get individual items out of the dictionary, by giving the dictionary's name, and the key in square brackets:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
print("\nWord: %s" % 'list')
print("Meaning: %s" % python_words['list'])
print("\nWord: %s" % 'dictionary')
print("Meaning: %s" % python_words['dictionary'])
print("\nWord: %s" % 'function')
print("Meaning: %s" % python_words['function'])
# + [markdown] slideshow={"slide_type": "subslide"}
# This code looks pretty repetitive, and it is. Dictionaries have their own for-loop syntax, but since there are two kinds of information in dictionaries, the structure is a bit more complicated than it is for lists. Here is how to use a for loop with a dictionary:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Print out the items in the dictionary.
for word, meaning in python_words.items():
print("\nWord: %s" % word)
print("Meaning: %s" % meaning)
# + [markdown] slideshow={"slide_type": "subslide"}
# The output is identical, but we did it in 3 lines instead of 6. If we had 100 terms in our dictionary, we would still be able to print them out with just 3 lines.
#
# The only tricky part about using for loops with dictionaries is figuring out what to call those first two variables. The general syntax for this for loop is:
# + slideshow={"slide_type": "fragment"}
for key_name, value_name in dictionary_name.items():
print(key_name) # The key is stored in whatever you called the first variable.
print(value_name) # The value associated with that key is stored in your second variable.
# + [markdown] slideshow={"slide_type": "skip"}
# <a name='exercises_what'></a>Exercises
# ---
# #### Ex: 6.1: Pet Names
# - Create a dictionary to hold information about pets. Each key is an animal's name, and each value is the kind of animal.
# - For example, 'ziggy': 'canary'
# - Put at least 3 key-value pairs in your dictionary.
# - Use a for loop to print out a series of statements such as "Willie is a dog."
#
# #### Ex 6.2: Polling Friends
# - Think of a question you could ask your friends. Create a dictionary where each key is a person's name, and each value is that person's response to your question.
# - Store at least three responses in your dictionary.
# - Use a for loop to print out a series of statements listing each person's name, and their response.
# + slideshow={"slide_type": "skip"}
# Ex 6.1 : Pet Names
# put your code here
# + slideshow={"slide_type": "skip"}
# Ex 6.2 : Polling Friends
# put your code here
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='common_operations'></a>Common operations with dictionaries
# ===
# There are a few common things you will want to do with dictionaries. These include adding new key-value pairs, modifying information in the dictionary, and removing items from dictionaries.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='adding_pairs'></a>Adding new key-value pairs
# ---
# To add a new key-value pair, you give the dictionary name followed by the new key in square brackets, and set that equal to the new value. We will show this by starting with an empty dictionary, and re-creating the dictionary from the example above.
# + slideshow={"slide_type": "fragment"}
# Create an empty dictionary.
python_words = {}
# Fill the dictionary, pair by pair.
python_words['list'] ='A collection of values that are not connected, but have an order.'
python_words['dictionary'] = 'A collection of key-value pairs.'
python_words['function'] = 'A named set of instructions that defines a set of actions in Python.'
# Print out the items in the dictionary.
for word, meaning in python_words.items():
print("\nWord: %s" % word)
print("Meaning: %s" % meaning)
# +
# Dealing with Hashing Functions
hashings = {}
hashings['function'] = hash('function')
hashings['list'] = hash('list')
hashings['dictionary'] = hash('dictionary')
for k, v in hashings.items():
print(k,': ',v)
from collections import OrderedDict
hashings = OrderedDict()
hashings['function'] = hash('function')
hashings['list'] = hash('list')
hashings['dictionary'] = hash('dictionary')
for k, v in hashings.items():
print(k,': ',v)
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='modifying_values'></a>Modifying values in a dictionary
# ---
# At some point you may want to modify one of the values in your dictionary. Modifying a value in a dictionary is pretty similar to modifying an element in a list. You give the name of the dictionary and then the key in square brackets, and set that equal to the new value.
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
print('dictionary: ' + python_words['dictionary'])
# Clarify one of the meanings.
python_words['dictionary'] = 'A collection of key-value pairs. \
Each key can be used to access its corresponding value.'
print('\ndictionary: ' + python_words['dictionary'])
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='removing_pairs'></a>Removing key-value pairs
# ---
# You may want to remove some key-value pairs from one of your dictionaries at some point. You can do this using the same `del` command you learned to use with lists. To remove a key-value pair, you give the `del` command, followed by the name of the dictionary, with the key that you want to delete. This removes the key and the value as a pair.
# + slideshow={"slide_type": "subslide"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Remove the word 'list' and its meaning.
_ = python_words.pop('list')
# Show the current set of words and meanings.
print("\n\nThese are the Python words I know:")
for word, meaning in python_words.items():
print("\nWord: %s" % word)
print("Meaning: %s" % meaning)
# + [markdown] slideshow={"slide_type": "subslide"}
# If you were going to work with this code, you would certainly want to put the code for displaying the dictionary into a function.
#
# def means define function which is like a mini-program that we can use again later, instead of re-writing that code. In this case the function is called 'show_words_meanings'.
#
# Let's see what this looks like:
# + slideshow={"slide_type": "fragment"}
def show_words_meanings(python_words):
# This function takes in a dictionary of python words and meanings,
# and prints out each word with its meaning.
print("\n\nThese are the Python words I know:")
for word, meaning in python_words.items():
print("\nWord: %s" % word)
print("Meaning: %s" % meaning)
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
show_words_meanings(python_words)
# Remove the word 'list' and its meaning.
del python_words['list']
show_words_meanings(python_words)
# + [markdown] slideshow={"slide_type": "subslide"}
# As long as we have a nice clean function to work with, let's clean up our output a little:
# + slideshow={"slide_type": "fragment"}
def show_words_meanings(python_words):
# This function takes in a dictionary of python words and meanings,
# and prints out each word with its meaning.
print("\n\nThese are the Python words I know:")
for word, meaning in python_words.items():
print("\n%s: %s" % (word, meaning))
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
show_words_meanings(python_words)
# Remove the word 'list' and its meaning.
del python_words['list']
show_words_meanings(python_words)
# + [markdown] slideshow={"slide_type": "fragment"}
# This is much more realistic code.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='modifying_keys'></a>Modifying keys in a dictionary
# ---
# Modifying a value in a dictionary was straightforward, because nothing else depends on the value. Modifying a key is a little harder, because each key is used to unlock a value. We can change a key in two steps:
#
# - Make a new key, and copy the value to the new key.
# - Delete the old key, which also deletes the old value.
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's what this looks like. We will use a dictionary with just one key-value pair, to keep things simple.
# +
# We have a spelling mistake!
python_words = {'list': 'A collection of values that are not connected, but have an order.'}
# Create a new, correct key, and connect it to the old value.
# Then delete the old key.
python_words['list'] = python_words['lisst']
del python_words['lisst']
# Print the dictionary, to show that the key has changed.
print(python_words)
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "skip"}
# <a name='exercises_common_operations'></a>Exercises
# ---
# #### Ex 6.3: Pet Names 2
# - Make a copy of your program from [Pet Names](#exercises_what).
# - Use a for loop to print out a series of statements such as "Willie is a dog."
# - Modify one of the values in your dictionary. You could clarify to name a breed, or you could change an animal from a cat to a dog.
# - Use a for loop to print out a series of statements such as "Willie is a dog."
# - Add a new key-value pair to your dictionary.
# - Use a for loop to print out a series of statements such as "Willie is a dog."
# - Remove one of the key-value pairs from your dictionary.
# - Use a for loop to print out a series of statements such as "Willie is a dog."
# - Bonus: Use a function to do all of the looping and printing in this problem.
#
# #### Ex 6.4: Weight Lifting
# - Make a dictionary where the keys are the names of weight lifting exercises, and the values are the number of times you did that exercise.
# - Use a for loop to print out a series of statements such as "I did 10 bench presses".
# - Modify one of the values in your dictionary, to represent doing more of that exercise.
# - Use a for loop to print out a series of statements such as "I did 10 bench presses".
# - Add a new key-value pair to your dictionary.
# - - Use a for loop to print out a series of statements such as "I did 10 bench presses".
# - Remove one of the key-value pairs from your dictionary.
# - - Use a for loop to print out a series of statements such as "I did 10 bench presses".
# - Bonus: Use a function to do all of the looping and printing in this problem.
# + slideshow={"slide_type": "skip"}
# Ex 6.3 : Pet Names 2
# put your code here
# + slideshow={"slide_type": "skip"}
# Ex 6.4 : Weight Lifting
# put your code here
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='looping'></a>Looping through a dictionary
# ===
# + [markdown] slideshow={"slide_type": "subslide"}
# Since dictionaries are really about connecting bits of information, you will often use them in the ways described above, where you add key-value pairs whenever you receive some new information, and then you retrieve the key-value pairs that you care about. Sometimes, however, you will want to loop through the entire dictionary. There are several ways to do this:
#
# - You can loop through all key-value pairs;
# - You can loop through the keys, and pull out the values for any keys that you care about;
# - You can loop through the values.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='loop_all_keys_values'></a>Looping through all key-value pairs
# ---
# This is the kind of loop that was shown in the first example. Here's what this loop looks like, in a general format:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
for key, value in my_dict.items():
print('\nKey: %s' % key)
print('Value: %s' % value)
# + [markdown] slideshow={"slide_type": "subslide"}
# This works because the method `.items()` pulls all key-value pairs from a dictionary into a list of tuples:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
print(my_dict.items())
# + [markdown] slideshow={"slide_type": "subslide"}
# The syntax `for key, value in my_dict.items():` does the work of looping through this list of tuples, and pulling the first and second item from each tuple for us.
#
# There is nothing special about any of these variable names, so Python code that uses this syntax becomes really readable. Rather than create a new example of this loop, let's just look at the original example again to see this in a meaningful context:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
for word, meaning in python_words.items():
print("\nWord: %s" % word)
print("Meaning: %s" % meaning)
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='loop_all_keys'></a>Looping through all keys in a dictionary
# ---
# Python provides a clear syntax for looping through just the keys in a dictionary:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
for key in my_dict.keys():
print('Key: %s' % key)
# + [markdown] slideshow={"slide_type": "subslide"}
# This is actually the default behavior of looping through the dictionary itself. So you can leave out the `.keys()` part, and get the exact same behavior:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
for key in my_dict:
print('Key: %s' % key)
# + [markdown] slideshow={"slide_type": "subslide"}
# The only advantage of using the `.keys()` in the code is a little bit of clarity. But anyone who knows Python reasonably well is going to recognize what the second version does. In the rest of our code, we will leave out the `.keys()` when we want this behavior.
#
# You can pull out the value of any key that you are interested in within your loop, using the standard notation for accessing a dictionary value from a key:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
for key in my_dict:
print('Key: %s' % key)
if key == 'key_2':
print(" The value for key_2 is %s." % my_dict[key])
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's show how we might use this in our Python words program. This kind of loop provides a straightforward way to show only the words in the dictionary:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Show the words that are currently in the dictionary.
print("The following Python words have been defined:")
for word in python_words:
print("- %s" % word)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can extend this slightly to make a program that lets you look up words. We first let the user choose a word. When the user has chosen a word, we get the meaning for that word, and display it:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Show the words that are currently in the dictionary.
print("The following Python words have been defined:")
for word in python_words:
print("- %s" % word)
# Allow the user to choose a word, and then display the meaning for that word.
requested_word = raw_input("\nWhat word would you like to learn about? ")
print("\n%s: %s" % (requested_word, python_words[requested_word]))
# + [markdown] slideshow={"slide_type": "subslide"}
# This allows the user to select one word that has been defined. If we enclose the input part of the program in a while loop, the user can see as many definitions as they'd like:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Show the words that are currently in the dictionary.
print("The following Python words have been defined:")
for word in python_words:
print("- %s" % word)
requested_word = ''
while requested_word != 'quit':
# Allow the user to choose a word, and then display the meaning for that word.
requested_word = raw_input("\nWhat word would you like to learn about? (or 'quit') ")
if requested_word in python_words.keys():
print("\n %s: %s" % (requested_word, python_words[requested_word]))
else:
# Handle misspellings, and words not yet stored.
print("\n Sorry, I don't know that word.")
# + [markdown] slideshow={"slide_type": "subslide"}
# This allows the user to ask for as many meanings as they want, but it takes the word "quit" as a requested word. Let's add an `elif` clause to clean up this behavior:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Show the words that are currently in the dictionary.
print("The following Python words have been defined:")
for word in python_words:
print("- %s" % word)
requested_word = ''
while requested_word != 'quit':
# Allow the user to choose a word, and then display the meaning for that word.
requested_word = raw_input("\nWhat word would you like to learn about? (or 'quit') ")
if requested_word in python_words.keys():
# This is a word we know, so show the meaning.
print("\n %s: %s" % (requested_word, python_words[requested_word]))
elif requested_word != 'quit':
# This is not in python_words, and it's not 'quit'.
print("\n Sorry, I don't know that word.")
else:
# The word is quit.
print "\n Bye!"
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='loop_all_values'></a>Looping through all values in a dictionary
# ---
# Python provides a straightforward syntax for looping through all the values in a dictionary, as well:
# + slideshow={"slide_type": "fragment"}
my_dict = {'key_1': 'value_1',
'key_2': 'value_2',
'key_3': 'value_3',
}
for value in my_dict.values():
print('Value: %s' % value)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can use this loop syntax to have a little fun with the dictionary example, by making a little quiz program. The program will display a meaning, and ask the user to guess the word that matches that meaning. Let's start out by showing all the meanings in the dictionary:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
for meaning in python_words.values():
print("Meaning: %s" % meaning)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now we can add a prompt after each meaning, asking the user to guess the word:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Print each meaning, one at a time, and ask the user
# what word they think it is.
for meaning in python_words.values():
print("\nMeaning: %s" % meaning)
guessed_word = raw_input("What word do you think this is? ")
# The guess is correct if the guessed word's meaning matches the current meaning.
if python_words[guessed_word] == meaning:
print("You got it!")
else:
print("Sorry, that's just not the right word.")
# + [markdown] slideshow={"slide_type": "subslide"}
# This is starting to work, but we can see from the output that the user does not get the chance to take a second guess if they guess wrong for any meaning. We can use a while loop around the guessing code, to let the user guess until they get it right:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
# Print each meaning, one at a time, and ask the user
# what word they think it is.
for meaning in python_words.values():
print("\nMeaning: %s" % meaning)
# Assume the guess is not correct; keep guessing until correct.
correct = False
while not correct:
guessed_word = input("\nWhat word do you think this is? ")
# The guess is correct if the guessed word's meaning matches the current meaning.
if python_words[guessed_word] == meaning:
print("You got it!")
correct = True
else:
print("Sorry, that's just not the right word.")
# + [markdown] slideshow={"slide_type": "subslide"}
# This is better. Now, if the guess is incorrect, the user is caught in a loop that they can only exit by guessing correctly. The final revision to this code is to show the user a list of words to choose from when they are asked to guess:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
def show_words(python_words):
# A simple function to show the words in the dictionary.
display_message = ""
for word in python_words.keys():
display_message += word + ' '
print display_message
# Print each meaning, one at a time, and ask the user
# what word they think it is.
for meaning in python_words.values():
print("\n%s" % meaning)
# Assume the guess is not correct; keep guessing until correct.
correct = False
while not correct:
print("\nWhat word do you think this is?")
show_words(python_words)
guessed_word = raw_input("- ")
# The guess is correct if the guessed word's meaning matches the current meaning.
if python_words[guessed_word] == meaning:
print("You got it!")
correct = True
else:
print("Sorry, that's just not the right word.")
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name="looping_in_order"></a>Looping through a dictionary in order
# ===
# Dictionaries are quite useful because they allow bits of information to be connected. One of the problems with dictionaries, however, is that they are not stored in any particular order. When you retrieve all of the keys or values in your dictionary, you can't be sure what order you will get them back. There is a quick and easy way to do this, however, when you want them in a particular order.
#
# Let's take a look at the order that results from a simple call to *dictionary.keys()*:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
for word in python_words.keys():
print(word)
# + [markdown] slideshow={"slide_type": "subslide"}
# The resulting list is not in order. The list of keys can be put in order by passing the list into the *sorted()* function, in the line that initiates the for loop:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
for word in sorted(python_words.keys()):
print(word)
# + [markdown] slideshow={"slide_type": "subslide"}
# This approach can be used to work with the keys and values in order. For example, the words and meanings can be printed in alphabetical order by word:
# + slideshow={"slide_type": "fragment"}
python_words = {'list': 'A collection of values that are not connected, but have an order.',
'dictionary': 'A collection of key-value pairs.',
'function': 'A named set of instructions that defines a set of actions in Python.',
}
for word in sorted(python_words.keys()):
print("%s: %s" % (word.title(), python_words[word]))
# + [markdown] slideshow={"slide_type": "fragment"}
# In this example, the keys have been put into alphabetical order in the for loop only; Python has not changed the way the dictionary is stored at all. So the next time the dictionary is accessed, the keys could be returned in any order. There is no way to permanently specify an order for the items in an ordinary dictionary, but if you want to do this you can use the [OrderedDict](http://docs.python.org/3.3/library/collections.html#ordereddict-objects) structure.
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "skip"}
# <a name='exercises_looping'></a>Exercises
# ---
# #### <a name='exercise_mountain_heights'></a>Ex 6.5: Mountain Heights
# - Wikipedia has a list of the [tallest mountains in the world](http://en.wikipedia.org/wiki/List_of_mountains_by_elevation), with each mountain's elevation. Pick five mountains from this list.
# - Create a dictionary with the mountain names as keys, and the elevations as values.
# - Print out just the mountains' names, by looping through the keys of your dictionary.
# - Print out just the mountains' elevations, by looping through the values of your dictionary.
# - Print out a series of statements telling how tall each mountain is: "Everest is 8848 meters tall."
# - Revise your output, if necessary.
# - Make sure there is an introductory sentence describing the output for each loop you write.
# - Make sure there is a blank line between each group of statements.
#
# #### Ex 6.6: Mountain Heights 2
# - Revise your final output from Mountain Heights, so that the information is listed in alphabetical order by each mountain's name.
# - That is, print out a series of statements telling how tall each mountain is: "Everest is 8848 meters tall."
# - Make sure your output is in alphabetical order.
# + slideshow={"slide_type": "skip"}
# Ex 6.5 : Mountain Heights
# put your code here
# + slideshow={"slide_type": "skip"}
# Ex 6.6 : Mountain Heights 2
# put your code here
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='nesting'></a>Nesting
# ===
# Nesting is one of the most powerful concepts we have come to so far. Nesting involves putting a list or dictionary inside another list or dictionary. We will look at two examples here, lists inside of a dictionary and dictionaries inside of a dictionary. With nesting, the kind of information we can model in our programs is expanded greatly.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='lists_in_dictionary'></a>Lists in a dictionary
# ---
# A dictionary connects two pieces of information. Those two pieces of information can be any kind of data structure in Python. Let's keep using strings for our keys, but let's try giving a list as a value.
#
# The first example will involve storing a number of people's favorite numbers. The keys consist of people's names, and the values are lists of each person's favorite numbers. In this first example, we will access each person's list one at a time.
# + slideshow={"slide_type": "subslide"}
# This program stores people's favorite numbers, and displays them.
favorite_numbers = {'eric': [3, 11, 19, 23, 42],
'ever': [2, 4, 5],
'willie': [5, 35, 120],
}
# Display each person's favorite numbers.
print("Eric's favorite numbers are:")
print(favorite_numbers['eric'])
print("\nEver's favorite numbers are:")
print(favorite_numbers['ever'])
print("\nWillie's favorite numbers are:")
print(favorite_numbers['willie'])
# + [markdown] slideshow={"slide_type": "subslide"}
# We are really just working our way through each key in the dictionary, so let's use a for loop to go through the keys in the dictionary:
# +
# Example for a Sparse Matrix Implementation
sparse_matrix = {}
sparse_matrix[0] = {1: 12.3, 23: 25.5}
sparse_matrix[1] = {3: 12.0, 15: 25.5}
sparse_matrix[1][15]
full_matrix = [[1, 3, 4],
[2, 5, 3]]
full_matrix[1][2]
# + slideshow={"slide_type": "fragment"}
# This program stores people's favorite numbers, and displays them.
favorite_numbers = {'eric': [3, 11, 19, 23, 42],
'ever': [2, 4, 5],
'willie': [5, 35, 120],
}
# Display each person's favorite numbers.
for name in favorite_numbers:
print("\n%s's favorite numbers are:" % name.title())
print(favorite_numbers[name])
# + [markdown] slideshow={"slide_type": "subslide"}
# This structure is fairly complex, so don't worry if it takes a while for things to sink in. The dictionary itself probably makes sense; each person is connected to a list of their favorite numbers.
#
# This works, but we'd rather not print raw Python in our output. Let's use a for loop to print the favorite numbers individually, rather than in a Python list.
# + slideshow={"slide_type": "fragment"}
# This program stores people's favorite numbers, and displays them.
favorite_numbers = {'eric': [3, 11, 19, 23, 42],
'ever': [2, 4, 5],
'willie': [5, 35, 120],
}
# Display each person's favorite numbers.
for name in favorite_numbers:
print("\n%s's favorite numbers are:" % name.title())
# Each value is itself a list, so we need another for loop
# to work with the list.
for favorite_number in favorite_numbers[name]:
print(favorite_number)
# + [markdown] slideshow={"slide_type": "subslide"}
# Things get a little more complicated inside the for loop. The value is a list of favorite numbers, so the for loop pulls each *favorite\_number* out of the list one at a time. If it makes more sense to you, you are free to store the list in a new variable, and use that to define your for loop:
#
# + slideshow={"slide_type": "fragment"}
# This program stores people's favorite numbers, and displays them.
favorite_numbers = {'eric': [3, 11, 19, 23, 42],
'ever': [2, 4, 5],
'willie': [5, 35, 120],
}
# Display each person's favorite numbers.
for name in favorite_numbers:
print("\n%s's favorite numbers are:" % name.title())
# Each value is itself a list, so let's put that list in a variable.
current_favorite_numbers = favorite_numbers[name]
for favorite_number in current_favorite_numbers:
print(favorite_number)
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='dictionaries_in_dictionary'></a>Dictionaries in a dictionary
# ---
# The most powerful nesting concept we will cover right now is nesting a dictionary inside of a dictionary.
#
# To demonstrate this, let's make a dictionary of pets, with some information about each pet. The keys for this dictionary will consist of the pet's name. The values will include information such as the kind of animal, the owner, and whether the pet has been vaccinated.
# + slideshow={"slide_type": "subslide"}
# This program stores information about pets. For each pet,
# we store the kind of animal, the owner's name, and
# the breed.
pets = {'willie': {'kind': 'dog', 'owner': 'eric', 'vaccinated': True},
'walter': {'kind': 'cockroach', 'owner': 'eric', 'vaccinated': False},
'peso': {'kind': 'dog', 'owner': 'chloe', 'vaccinated': True},
}
# Let's show all the information for each pet.
print("Here is what I know about Willie:")
print("kind: " + pets['willie']['kind'])
print("owner: " + pets['willie']['owner'])
print("vaccinated: " + str(pets['willie']['vaccinated']))
print("\nHere is what I know about Walter:")
print("kind: " + pets['walter']['kind'])
print("owner: " + pets['walter']['owner'])
print("vaccinated: " + str(pets['walter']['vaccinated']))
print("\nHere is what I know about Peso:")
print("kind: " + pets['peso']['kind'])
print("owner: " + pets['peso']['owner'])
print("vaccinated: " + str(pets['peso']['vaccinated']))
# + [markdown] slideshow={"slide_type": "subslide"}
# Clearly this is some repetitive code, but it shows exactly how we access information in a nested dictionary. In the first set of `print` statements, we use the name 'willie' to unlock the 'kind' of animal he is, the 'owner' he has, and whether or not he is 'vaccinated'. We have to wrap the vaccination value in the `str` function so that Python knows we want the words 'True' and 'False', not the values `True` and `False`. We then do the same thing for each animal.
#
# Let's rewrite this program, using a for loop to go through the dictionary's keys:
# + slideshow={"slide_type": "fragment"}
# This program stores information about pets. For each pet,
# we store the kind of animal, the owner's name, and
# the breed.
pets = {'willie': {'kind': 'dog', 'owner': 'eric', 'vaccinated': True},
'walter': {'kind': 'cockroach', 'owner': 'eric', 'vaccinated': False},
'peso': {'kind': 'dog', 'owner': 'chloe', 'vaccinated': True},
}
# Let's show all the information for each pet.
for pet_name, pet_information in pets.items():
print("\nHere is what I know about %s:" % pet_name.title())
print("kind: " + pet_information['kind'])
print("owner: " + pet_information['owner'])
print("vaccinated: " + str(pet_information['vaccinated']))
# + [markdown] slideshow={"slide_type": "subslide"}
# This code is much shorter and easier to maintain. But even this code will not keep up with our dictionary. If we add more information to the dictionary later, we will have to update our print statements. Let's put a second for loop inside the first loop in order to run through all the information about each pet:
# + slideshow={"slide_type": "fragment"}
# This program stores information about pets. For each pet,
# we store the kind of animal, the owner's name, and
# the breed.
pets = {'willie': {'kind': 'dog', 'owner': 'eric', 'vaccinated': True},
'walter': {'kind': 'cockroach', 'owner': 'eric', 'vaccinated': False},
'peso': {'kind': 'dog', 'owner': 'chloe', 'vaccinated': True},
}
# Let's show all the information for each pet.
for pet_name, pet_information in pets.items():
print("\nHere is what I know about %s:" % pet_name.title())
# Each animal's dictionary is in 'information'
for key in pet_information:
print(key + ": " + str(pet_information[key]))
# + [markdown] slideshow={"slide_type": "subslide"}
# This nested loop can look pretty complicated, so again, don't worry if it doesn't make sense for a while.
#
# - The first loop gives us all the keys in the main dictionary, which consist of the name of each pet.
# - Each of these names can be used to 'unlock' the dictionary of each pet.
# - The inner loop goes through the dictionary for that individual pet, and pulls out all of the keys in that individual pet's dictionary.
# - We print the key, which tells us the kind of information we are about to see, and the value for that key.
# - You can see that we could improve the formatting in the output.
# - We could capitalize the owner's name.
# - We could print 'yes' or 'no', instead of True and False.
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's show one last version that uses some if statements to clean up our data for printing:
# + slideshow={"slide_type": "subslide"}
# This program stores information about pets. For each pet,
# we store the kind of animal, the owner's name, and
# the breed.
pets = {'willie': {'kind': 'dog', 'owner': 'eric', 'vaccinated': True},
'walter': {'kind': 'cockroach', 'owner': 'eric', 'vaccinated': False},
'peso': {'kind': 'dog', 'owner': 'chloe', 'vaccinated': True},
}
# Let's show all the information for each pet.
for pet_name, pet_information in pets.items():
print("\nHere is what I know about %s:" % pet_name.title())
# Each animal's dictionary is in pet_information
for key in pet_information:
if key == 'owner':
# Capitalize the owner's name.
print(key + ": " + pet_information[key].title())
elif key == 'vaccinated':
# Print 'yes' for True, and 'no' for False.
vaccinated = pet_information['vaccinated']
if vaccinated:
print('vaccinated: yes')
else:
print('vaccinated: no')
else:
# No special formatting needed for this key.
print(key + ": " + pet_information[key])
# + [markdown] slideshow={"slide_type": "fragment"}
# This code is a lot longer, and now we have nested if statements as well as nested for loops. But keep in mind, this structure would work if there were 1000 pets in our dictionary, and it would work if we were storing 1000 pieces of information about each pet. One level of nesting lets us model an incredible array of information.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='important_note'></a>An important note about nesting
# ---
# While one level of nesting is really useful, nesting much deeper than that gets really complicated, really quickly. There are other structures such as classes which can be even more useful for modeling information. In addition to this, we can use Python to store information in a database, which is the proper tool for storing deeply nested information.
#
# Often times when you are storing information in a database you will pull a small set of that information out and put it into a dictionary, or a slightly nested structure, and then work with it. But you will rarely, if ever, work with Python data structures nested more than one level deep.
# + [markdown] slideshow={"slide_type": "skip"}
# <a name='exercises_nesting'></a>Exercises
# ---
# #### Ex 6.7: Mountain Heights 3
# - This is an extension of [Mountain Heights](#exercise_mountain_heights). Make sure you save this program under a different filename, such as *mountain\_heights_3.py*, so that you can go back to your original program if you need to.
# - The list of [tallest mountains in the world](http://en.wikipedia.org/wiki/List_of_mountains_by_elevation) provided all elevations in meters. Convert each of these elevations to feet, given that a meter is approximately 3.28 feet. You can do these calculations by hand at this point.
# - Create a new dictionary, where the keys of the dictionary are still the mountains' names. This time however, the values of the dictionary should be a list of each mountain's elevation in meters, and then in feet: {'everest': [8848, 29029]}
# - Print out just the mountains' names, by looping through the keys of your dictionary.
# - Print out just the mountains' elevations in meters, by looping through the values of your dictionary and pulling out the first number from each list.
# - Print out just the mountains' elevations in feet, by looping through the values of your dictionary and pulling out the second number from each list.
# - Print out a series of statements telling how tall each mountain is: "Everest is 8848 meters tall, or 29029 feet."
# - Bonus:
# - Start with your original program from [Mountain Heights](#exercise_mountain_heights). Write a function that reads through the elevations in meters, and returns a list of elevations in feet. Use this list to create the nested dictionary described above.
#
# #### Ex 6.8: Mountain Heights 4
# - This is one more extension of Mountain Heights.
# - Create a new dictionary, where the keys of the dictionary are once again the mountains' names. This time, the values of the dictionary are another dictionary. This dictionary should contain the elevation in either meters or feet, and the range that contains the mountain. For example: {'everest': {'elevation': 8848, 'range': 'himalaya'}}.
# - Print out just the mountains' names.
# - Print out just the mountains' elevations.
# - Print out just the range for each mountain.
# - Print out a series of statements that say everything you know about each mountain: "Everest is an 8848-meter tall mountain in the Himalaya range."
# + slideshow={"slide_type": "skip"}
# Ex 6.7 : Mountain Heights 3
# put your code here
# + slideshow={"slide_type": "skip"}
# Ex 6.8 : Mountain Heights 4
# put your code here
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "skip"}
# <a name='overall_challenges'></a>Overall Challenges
# ===
# #### Word Wall
# - A word wall is a place on your wall where you keep track of the new words and meanings you are learning. Write a terminal app that lets you enter new words, and a meaning for each word.
# - Your app should have a title bar that says the name of your program.
# - Your program should give users the option to see all words and meanings that have been entered so far.
# - Your program should give users the option to enter a new word and meaning.
# - Your program must not allow duplicate entries.
# - Your program should store existing words and meanings, even after the program closes.
# - Your program should give users the option to modify an existing meaning.
# - Bonus Features
# - Allow users to modify the spelling of words.
# - Allow users to categorize words.
# - Turn the program into a game that quizzes users on words and meanings.
# - (later on) Turn your program into a website that only you can use.
# - (later on) Turn your program into a website that anyone can register for, and use.
# - Add a visualization feature that reports on some statistics about the words and meanings that have been entered.
#
# #### Periodic Table App
# - The [Periodic Table](http://www.ptable.com/) of the Elements was developed to organize information about the elements that make up the Universe. Write a terminal app that lets you enter information about each element in the periodic table.
# - Make sure you include the following information:
# - symbol, name, atomic number, row, and column
# - Choose at least one other piece of information to include in your app.
# - Provide a menu of options for users to:
# - See all the information that is stored about any element, by entering that element's symbol.
# - Choose a property, and see that property for each element in the table.
# - Bonus Features
# - Provide an option to view the symbols arranged like the periodic table. ([hint](#hints_periodic_table))
# + slideshow={"slide_type": "skip"}
# Challenge: Word Wall
# put your code here
# + slideshow={"slide_type": "skip"}
# Challenge: Periodic Table App
# put your code here
# + [markdown] slideshow={"slide_type": "skip"}
# [top](#top)
# + [markdown] slideshow={"slide_type": "skip"}
# Hints
# ===
# #### <a name='hints_periodic_table'></a>Periodic Table App
# - You can use a for loop to loop through each element. Pick out the elements' row numbers and column numbers.
# - Use two nested for loops to print either an element's symbol or a series of spaces, depending on how full that row is.
| 06_dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# # Módulos y bibliotecas en Python
#
# Hasta ahora hemos desarrollado programas no muy complicados que cabían en una celda de un Notebook, pero poco a poco se va complicando más la cosa. ¿Qué ocurrirá cuando tengamos varias funciones definidas, datos declarados, Clases (lo veremos el próximo día), llamadas a páginas web...? ¿Lo vamos a tener todo en una celda/Notebook?
#
# Sería poco manejable, ¿no? Para ayudarnos con esto utilizaremos la sentencia `import`, pudiendo dividir nuestro programa en varios scripts de Python, y no solo eso, sino que tendremos también la opción de usar librerías de terceros en nuestro código, lo que va a ayudar mucho en el desarrollo.
#
# Para este Notebook se recomienda tener otro IDE como VSCode, Spyder o Pycharm.
#
# 1. [import](#1.-import)
# 2. [Bibliotecas](#2.-Bibliotecas)
# 3. [Resumen](#3.-Resumen)
#
# *NOTA: Cuando veamos las clases, volveremos para crearnos nuestro módulo con clases y así repasar de nuevo todo esto
# ## 1. import
# Recuerda que hay que **evitar en la medida de lo posible tener código repetido o duplicado**, por lo que funcionalidades que uses en diferentes partes del código no tendrás que copiar y pegarlas en todos lados, sino que tienes la posibilidad de definirlas una sola vez, e importarlas después, cuando consideres.
#
# Por tanto, según vamos complicando nuestros programas, surge la necesidad de modularizarlos, es decir, poder dividirlos y "paquetizarlos". Como habrás podido imaginar, programas productivos, como una página web o un juego, no van en un Notebook, sino en varios scripts de Python. Según las diferentes funcionalidades del código, lo iremos dividiendo en varias partes. Por ejemplo, en un script pueden ir funciones auxiliares, en otro constantes, en otro tus Clases... Veamos algunos conceptos
#
# **Script**
#
# No es más que un archivo con código Python. Estos archivos tienen la extensión `.py`, por lo que un ejemplo podría ser `primer_script.py`.
#
# **Módulo/Biblioteca/Librería/Paquete**
#
# Son términos parecidos que se suelen utilizar para referirse a lo mismo. A efectos prácticos, los consideraremos igual. Se trata de paquetes de código que tienen una funcionalidad bien definida y podremos importar en nuestros programas de Python, utilizando lo que haya dentro (funciones, variables, clases...).
#
# **import**
#
# Palabra reservada que se usa en Python para importar cualquier objeto en nuestro código. Se usa para importar variables, funciones o módulos.
#
# Veamos cómo podemos importar objetos de otros scripts.
#
# Para ello, utilizaremos un comando nuevo para nosotros que no es Python (no lo volveremos a usar, es solo para el desarrollo de esta explicación) pero que nos permitirá crearnos un fichero desde el notebook, que será esa primera línea: "%%file nombre_archivo"
#
# Lo que hace es crear un fichero llamado nombre_archivo, y dentro de él mete todo lo que escribamos en las líneas posteriores. Compruébalo:
# +
# %%file primer_script.py
"""
Example of a python module. Contains a variable called my_variable,
and a function called my_function.
"""
my_variable = 0
my_variable2 = (20,)
my_variable3 = "var"
def my_function1():
"""
Example function
"""
return my_variable
def my_function2(var1, var2):
"""
Example 2 function. Returns a list that contains a concatenation of my_variable2 and my_variable3 and var1 and var2
"""
return list(my_variable2) + [my_variable3] + [var1] + [var2]
# -
# Tenemos en otro script una variable y una función definidas. Ahora, lo importamos:
import primer_script
help(primer_script)
# Como siempre en cualquier lenguaje de programación **tenemos que fijarnos siempre dónde está apuntando el Notebook**, es decir, en qué directorio de trabajo va a leer el script. Al poner `import primer_script`, va a buscar ese script en el **mismo directorio donde esté el Notebook**. Si lo tenemos en otro lado del ordenador, no lo detectará. Cuando realizas la importación, Python no recorre todas las carpetas del ordenador, lo busca en el mismo sitio donde está el Notebook. Más delante veremos como leer scripts de otros directorios.
#
# Si queremos acceder a cualquier objeto del módulo, simplemente utilizamos la sintaxis `nombre_modulo.objeto`
print(primer_script.my_variable)
print(primer_script.my_variable2)
print(primer_script.my_variable3)
print(primer_script.my_function1)
print(primer_script.my_function2)
primer_script.my_function1()
# [Aquí](https://docs.python.org/3/reference/import.html) tienes el enlace a la documentación del `import` por si tienes dudas, o necesitas realizar operaciones más complejas con el importado de módulos.
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/error.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>ERRORES Importado de módulos</h3>
#
# </td></tr>
# </table>
# Típico error cuando intentamos acceder a un módulo que pensamos que está instalado en el intérprete de Anaconda, o queremos acceder a un *.py* que en realidad no está en la ruta deseada.
import mi_modulo
# O si intentamos acceder una función/atributo que en realidad no existe en el módulo.
primer_script.my_function3
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Crear un script</h3>
#
# Crea un nuevo script de Python (es un archivo .py) y ponle de nombre "ejer_prueba.py". Lo puedes crear desde el propio Jupyter Lab desde File -> New -> Text File. O también desde VSCode.
#
# Declara dos variables en el script
# a = 1
# b = 2
#
# Importa ambas variables e imprímelas por pantalla.
#
# </td></tr>
# </table>
import ejer_prueba
print(ejer_prueba.a)
print(ejer_prueba.b)
# Tienes otras maneras de importar datos, ya que en ocasiones desconocemos lo que hay dentro del módulo, o simplemente no queremos utilizar todo lo que hay.
# +
import primer_script
primer_script.my_variable
# print(my_variable) ERROR. Solo estamos referenciando el modulo, no cargando lo de dentro
# +
from primer_script import my_variable, my_function1
print(my_variable)
print(my_function1)
# -
# En este caso ya no es necesario usar la sintaxis `modulo.objeto`, sino que el objeto ya está cargado en memoria. Es como si lo hubiésemos ejecuctado en una celda.
#
# Otra opción es importar todo. **NO es lo recomendable**. Porque el módulo podría tener muchas variables o funciones que no necesitemos, y ocupan espacio en memoria.
# +
from primer_script import *
# Equivalente a:
# from primer_script import my_variable, my_variable2, my_variable3, my_function1, my_function2
# -
# **¿Y si tenemos el archivo en otra carpeta?** La sintaxis es la misma. Si dentro de la carpeta donde está el Notebook, hay otra carpeta llamada `direc_segundo`, y dentro de esa carpeta hay otro script llamado `segundo_script.py`, podrás acceder a los objetos de ese escript mediante la sintaxis `import direc_segundo.segundo_script`
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Crear un script dentro de una carpeta</h3>
#
# En el mismo sitio donde se encuentra este Notebook, crea la carpeta "direc_segundo", y dentro de la carpeta crea otro script llamado "segundo_script.py".
#
# Declara dos variables en el script
# c = 3
# d = 4
#
# Importa ambas variables e imprimelas por pantalla.
#
# </td></tr>
# </table>
# +
import direc_segundo.segundo_script
print(direc_segundo.segundo_script.c)
print(direc_segundo.segundo_script.d)
# -
# En estos casos resulta muy útil ponerle un alias al `import`. Esto nos ayudará a acortar el nombre y a poner uno más intuitivo
# +
import direc_segundo.segundo_script as variables
print(variables.c)
print(variables.d)
# -
# Por útimo, **¿y si nuestros módulos no están en una carpeta dentro del proyecto donde estamos trabajando, sino en otra carpeta del ordenador, o en directorios anteriores?** Esto lo podemos solucionar mediante `sys.path`. `sys` es el módulo de Python que se usa para manejar las variables del sistema y el intérprete. Y `path` es una lista de strings con una serie de rutas donde acude el intérprete de Python cuando tiene que buscar un módulo. [Tienes aqui la documentación de `sys`](https://docs.python.org/3/library/sys.html)
import sys
sys.path
# Si queremos que el intérprete de Python entienda de otros módulos que **no están en la carpeta de Notebook**, tendremos que añadir esa ruta a la lista de paths mediante la sintaxis `sys.path.append(la_nueva_ruta)`. Recuerda que `sys.path` es una lista, por lo que podrás aplicarle lo que ya sabes sobre listas.
# <table align="left">
# <tr><td width="80"><img src="../../imagenes/ejercicio.png" style="width:auto;height:auto"></td>
# <td style="text-align:left">
# <h3>Añadir un nuevo path</h3>
#
# Crea un nuevo directorio en otra ruta del ordenador, por ejemplo en el escritorio. Llámalo "direc_tercero" e introduce dentro un nuevo script que se llame "tercer_script.py". Tendrás que añadir el path de ese directorio a los paths de sys.
#
# Declara dos variables en el script
# e = 5
# f = 6
#
#
# Importa ambas variables e imprimelas por pantalla.
#
# </td></tr>
# </table>
# Una vez creado el nuevo directorio, se recomienda copiar su ruta y pegarla en el `append`.
sys.path.append('C:/Users/TheBridge/Desktop/Bloque_0_-_Ramp_Up/semana_2/dia_2/direc_tercero')
sys.path
import tercer_script as ts
print(ts.e)
print(ts.f)
# Con lo visto en este Notebook tienes conocimiento de sobra para manejar tu programa en varios scripts, pero si quieres aprender más sobre el concepto `import`, te recomiendo que te leas [este artículo](https://chrisyeh96.github.io/2017/08/08/definitive-guide-python-imports.html).
# ## 2. Bibliotecas
# Hasta ahora, todas las funcionalidades que necesitabas en Python las has ido desarrollando con tu propio código. Esto está bien, pero no siempre es necesario. **Existen muchísimas librerías que pueden hacer el trabajo por ti**, por lo que antes de implementar algo, ***no reinventes la rueda***. Se listo y busca si ya hay algo hecho en la web. Esto es lo bueno de trabajar con un lenguaje open source, que existe una comunidad con muchísimos desarrolladores y hay una gran cantidad de código publicado de manera gratuita. Por tanto, ¿de dónde podemos sacar estos códigos mágicos?
#
# * **[Biblioteca estándar de Python](https://docs.python.org/3/library/index.html#the-python-standard-library)**: aqui tienes muchas de las cosas que hemos visto ya (tipos de datos, el móduglo math...) y mucho más (lectura de archivos, compresión, acceso a BBDD...).
# * **Bibliotecas de terceros**: librerías que no vienen en el paquete por defecto de Python. Estas librerías o bien las implementa un particular o una empresa, y son de código abierto. Anaconda está muy orientado a la ciencia de datos, por lo que tiene librerías muy interesantes que utilizaremos durante el curso (pandas, numpy o sklearn). Además de estás librerías, podrás añadir al intérprete de Python todas las que necesites
#
# Veamos un ejemplo de importación de una librería
import math
math.sqrt(36)
# Importando `math`, podremos acceder a todas sus funciones mediante la sintaxis `math.funcion()`. [En la documentación](https://docs.python.org/3/library/math.html#math.log) verás todo lo que puedes hacer con `math`.
#
# Veamos un ejemplo en el que nos ayude a no escribir código de más.
help(math)
help(math.sqrt)
print(dir(math))
help(dir)
# Otro ejemplo del uso de librerías de terceros. En él, vemos cómo podemos leer ficheros desde Pythonn de 2 formas distintas, donde el uso de pandas (librería que veremos en detalle en unas semanas) nos facilita mucho el trabajo. De momento, no intentes entender el código, quédate solo con que se pueden usar librerías de terceros en tus scripts:
# + jupyter={"outputs_hidden": true}
import csv
with open('datos_prueba.csv', 'r') as file:
my_reader = csv.reader(file, delimiter=',')
for row in my_reader:
print(row)
# + jupyter={"outputs_hidden": true}
import pandas as pd
mi_data = pd.read_csv("datos_prueba.csv", sep=",")
#data.to_excel("miexcel.xlsx")
mi_data
# +
import seaborn as sns
graph = sns.barplot(x = "origin", y = "distance", data = mi_data)
# -
# ### Bibliotecas de terceros
# **¿Y si encontramos una librería interesante, pero no viene en el paquete de Anaconda?** La podremos instalar. Anaconda viene con muchas bibliotecas ya instaladas, la mayoría orientadas a trabajar con datos, pero por supuesto, no contiene todas las bibliotecas. Si quieres ver los paquetes de Anaconda, abre un *Prompt de Anaconda* y escribe `pip freeze` (o añade un ```!``` a ese comando y ejecútalo desde una celda ;) ).
#
# + jupyter={"outputs_hidden": true}
# !pip freeze
# -
# Entonces, **¿cómo se instala un paquete nuevo?** La mayoría de paquetes los vas a encontrar en [PyPI (Python Package Index)](https://pypi.org), que es el repositorio oficial de paquetes de Python. Existen paquetes de terceros que no pertenecen a PyPI. Eso no quiere decir que no los puedas instalar, sino que probablemente estén menos testados que los "oficiales".
#
# Veamos un ejemplo de cómo instalar una nueva librería. En este caso, instalaremos **wget**, que nos sirve para obtener archivos de la web.
# ! where python
import wget
# Al hace el `import` no existe, por lo que tendremos que instalarla. Para instalarla, abre un prompr de Anaconda y ejecuta `pip install wget`
#
# !pip install wget
# +
import wget
url = 'http://www.futurecrew.com/skaven/song_files/mp3/razorback.mp3'
filename = wget.download(url)
filename
# -
# ¡Perfecto! has podido instalar tu primer paquete en el intérprete de Python 3.7
# ## 3. Resumen
# Como ves, **gran parte de la potencia de Python reside en sus librerías**. Combinando los conocimientos básicos ya aprendidos con las librerías adecuadas, podremos crear dashboards interactivos, modelos de machine learning, páginas webs, automatizar procesos... ¡Los límites los pones tú! :)
# +
# Sintaxis básica para importar un modulo
import primer_script
# Para acceder a su documentación
help(primer_script)
# PAra acceder a los objetos del modulo
print(primer_script.my_variable)
print(primer_script.my_function1)
# Una manera de acceder a todo
from primer_script import *
# Renombrar el modulo también es util
import direc_segundo.segundo_script as variables
print(variables.c)
# Si queremos acceder rutas en otras partes del ordenador, hay que añadirlas a sys.path
sys.path.append('C:/Users/TheBridge/Desktop/Bloque_0_-_Ramp_Up/semana_2/dia_2/direc_tercero')
sys.path
# Por otro lado, tenemos las librerías del estandar de Python
import math
math.sqrt(36)
# O librerías de terceros que podrás instalar mediante un pip install libreria, en un Prompt de Anaconda.
import wget
| semana_3/dia_1/RESU_Modulos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Topic Modeling
# There are two popular choices for models: Latent Semantic Indexing (LSI) and Latent Dirichlet Allocation (LDA). LDA is a more complex process, and thus takes more resources and longer to run, but has higher accuracy. LSI is a much simpler process and can be run quite quickly.
# - LSI looks at words in a documents and its relationships to other words, with the important assumption that every word can only mean one thing. (cf. https://en.wikipedia.org/wiki/Latent_semantic_indexing)
# - LDA seeks to remedy this fault by allowing words to exist in multiple topics, first grouping them by topic, and each document is compared across each topic to determine the best fit. (cf. https://en.wikipedia.org/wiki/Latent_Dirichlet_allocation)
# +
from nltk.stem.snowball import SnowballStemmer
from nltk.corpus import stopwords
from string import punctuation
from gensim import corpora, models, similarities
stemmer = SnowballStemmer("english")
docs_tm = [tokenize_only(x) for x in docs]
docs_tm = [[x for x in i if x not in stopwords.words("english") and x not in punctuation] for i in docs_tm]
# -
# We first create a dict of word IDs and their respective word frequency for all documents.
# create a Gensim dictionary from the texts
dictionary = corpora.Dictionary(docs_tm)
# remove extremes (similar to the min/max df step used when creating the tf-idf matrix)
# no_below is absolute # of docs, no_above is fraction of corpus
dictionary.filter_extremes(no_below=40, no_above=.70)
# The corpus we now create with doc2bow is a vector of all words (IDs from the dict), and frequency for each document.
# convert the dictionary to a bag of words corpus for reference
corpus = [dictionary.doc2bow(i) for i in docs_tm]
# We'll make a tfidf, *term freqency inverse document frequency*, matrix. A tfidf takes into account the frequency of a word in the entire corpus, and offsets it based on its frequency among documents (more here: https://en.wikipedia.org/wiki/Tf–idf):
tfidf = models.TfidfModel(corpus)
corpus_tfidf = tfidf[corpus]
# ## LSI
lsi = models.LsiModel(corpus_tfidf, id2word=dictionary, num_topics=6)
corpus_lsi = lsi[corpus_tfidf]
lsi.print_topics(6)
# ## LDA
# +
# we run chunks of 15 books, and update after every 2 chunks, and make 10 passes
lda = models.LdaModel(corpus, num_topics=6,
update_every=2,
id2word=dictionary,
chunksize=15,
passes=10)
lda.show_topics()
# -
corpus_lda = lda[corpus_tfidf]
for i,doc in enumerate(corpus_lda): # both bow->tfidf and tfidf->lsi transformations are actually executed here, on the fly
print(titles[i],doc)
print ()
# For more with gensim, see the tutorials here: https://radimrehurek.com/gensim/tutorial.html
| text-analysis/templates/topic-modeling-general.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import cv2
def process_image(img):
dim = (224,224)
img = plt.imread(img)
img = cv2.resize(img, dim, interpolation=cv2.INTER_LINEAR)
return img
images_dir = 'images/'
# +
from os import listdir
test_images = []
for file in listdir(images_dir):
test_images.append(process_image(images_dir+file))
# -
fig, axes = plt.subplots(3, 3, figsize=(10,10))
ax = axes.flatten()
for i in range(len(test_images)):
ax[i].imshow(test_images[i])
ax[i].set_xlabel(test_images[i].shape)
# +
import keras
# from keras.models import load_model
from keras.utils import CustomObjectScope
from keras.initializers import glorot_uniform
from keras.models import model_from_json
with CustomObjectScope({'GlorotUniform': glorot_uniform()}):
json_file = open('kangaroo3.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("kangaroo3.h5")
# -
def draw_mask(box):
masks = np.zeros([224, 224], dtype='uint8')
row_s, row_e = box[1], box[3]
col_s, col_e = box[0], box[2]
masks[row_s:row_e, col_s:col_e] = 1
plt.imshow(masks[:, :], cmap='gray', alpha=0.4)
i=9
pred = np.round(loaded_model.predict(np.array([test_images[i]]))[0]).astype(int)
plt.imshow(test_images[i])
draw_mask(pred)
pred
| kangaroo-detection-localization/kangaroo localisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# libraries
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy.stats.stats import pearsonr
import matplotlib.pyplot as plt
# %matplotlib inline
#set ggplot style
# plt.style.use('ggplot')
duplicatejoins = pd.read_csv('rawdata/duplicate-joins.tsv', sep='\t')
kval = '01'
duplicatejoins = duplicatejoins[duplicatejoins['size']== str(kval) + 'k_rows']
duplicatejoins.head()
duplicatejoins = duplicatejoins[['tool', 'size', 'duplicates', '%involved', 'totaltime']]
duplicatejoins
# ### Config 1
c1 = duplicatejoins[duplicatejoins['%involved'].isin([5, 10, 20])]
c1.head()
c1.shape
c1 = c1.sort_values(by=['size', '%involved', 'duplicates'])
c1 = c1.reset_index(drop=True)
c1
# ### Config 2
c2 = duplicatejoins[duplicatejoins['%involved'].isin([30, 40, 50])]
c2.head()
c2.shape
c2 = c2.sort_values(by=['size', '%involved', 'duplicates'])
c2 = c2.reset_index(drop=True)
c2.head()
import itertools
comb = list(itertools.permutations(list(range(1, 5)), 2))
comb.append((1,1))
comb.append((2,2))
comb.append((3,3))
comb.append((4,4))
comb = sorted(comb)
xydf = pd.DataFrame(comb, columns=['Configurations', 'Configuration'])
tools = ['rdfizer', 'rmlmapper']
configs = [c1, c2]
confignames = ['Low_Dup', 'High_Dup']
z = {'z': [], 'xtick': [], 'ytick': []}
colors = {'c': []}
for row in xydf.iterrows():
ix = row[0]
x = row[1][0]
y = row[1][1]
# print(ix, x, y)
xix = 0 if 4/x >= 2 else 1
xtool = tools[xix]
yix = 0 if 4/y >= 2 else 1
ytool = tools[yix]
xconfigix = 1 if x%2 == 0 else (x % 2 - 1)
xconfig = configs[xconfigix]
yconfigix = 1 if y%2 == 0 else (y % 2 - 1)
yconfig = configs[yconfigix]
z['xtick'].append(xtool+'-'+ confignames[xconfigix])
z['ytick'].append(ytool+'-'+ confignames[yconfigix])
v = np.corrcoef( xconfig[((xconfig['tool']== xtool))]['totaltime'],
yconfig[((yconfig['tool']==ytool))]['totaltime']
)[0, 1]
k = v
if k > 0.999999999:
c = "#e7eff6"
elif k < 0:
c = '#fe4a49'
else:
c = '#03396c'
colors['c'].append(c)
z['z'].append(np.abs(v))
# +
xydf['z'] = z['z']
xydf
# -
xt = xydf.xtick.unique()
xt
# +
colors = pd.DataFrame(colors)
xt = xydf.xtick.unique()
ploto = xydf.plot.scatter('Configurations', 'Configuration', s=xydf.z * 2500, color=colors.c, alpha=0.8, edgecolors="black", linewidth=.1)
ploto.set_title('Join Duplicates')
ploto.set_xlabel('Configurations')
ploto.set_ylabel('Configurations')
ploto.set_xticks([0,1,2,3,4,5])
ploto.set_yticks([0,1,2,3,4,5])
# ploto.set_xticklabels(['rdfizer-Low_Dup',
# 'rdfizer-High_Dup',
# 'rmlmapper-Low_Dup',
# 'rmlmapper-High_Dup'],
# rotation=90)
# ploto.set_yticklabels(['rdfizer-Low_Dup',
# 'rdfizer-High_Dup',
# 'rmlmapper-Low_Dup',
# 'rmlmapper-High_Dup'])
# ploto.set_yticklabels(xydf.ytick)
ploto.spines['top'].set_visible(False)
ploto.spines['right'].set_visible(False)
#adds major gridlines
ploto.grid(color='grey', linestyle='-', linewidth=0.15, alpha=0.5)
#adds legend
#ploto.legend(categories.unique())
plt.figure(figsize=(35, 4))
# -
# # !mkdir figures/dexa
fig = ploto.get_figure()
fig.savefig("duplicate_joins/duplicate_joins_"+ str(kval) + "k_bubble.png", dpi=300, bbox_inches='tight')
| results/Plot Duplicate Joins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
import pandas as pd
import utils
import random
# +
# hyperparameters
batch_iterations = 100
batch_size = 32
full_iterations = 25
learning_rate = 0.01
reg_eta = 0.01
# dimensionalities
dim_lstm = 300
dim_word = 300
dim_aspect = 5
dim_sentence = 80
dim_polarity = 3
# setup utils object
isSample = True
u = utils.UTILS(batch_size, dim_sentence, dim_polarity, isSample)
# -
# define tf placeholders
X = tf.placeholder(tf.int32, [None, dim_sentence])
y = tf.placeholder(tf.float32, [None, dim_polarity])
seqlen = tf.placeholder(tf.int32, [None])
lr = tf.placeholder(tf.float32, [])
# define tf variables
with tf.variable_scope('bilstm_vars'):
with tf.variable_scope('weights', reuse = tf.AUTO_REUSE):
lstm_w = tf.get_variable(
name = 'softmax_w',
shape = [dim_lstm * 2, dim_polarity],
initializer = tf.random_uniform_initializer(-0.003, 0.003),
regularizer = tf.contrib.layers.l2_regularizer(reg_eta)
)
with tf.variable_scope('biases', reuse = tf.AUTO_REUSE):
lstm_b = tf.get_variable(
name = 'softmax_b',
shape = [dim_polarity],
initializer = tf.random_uniform_initializer(-0.003, 0.003),
regularizer = tf.contrib.layers.l2_regularizer(reg_eta)
)
# define lstm model
def dynamic_lstm(inputs, seqlen):
inputs = tf.nn.dropout(inputs, keep_prob=1.0)
with tf.name_scope('bilstm_model'):
forward_lstm_cell = tf.contrib.rnn.LSTMCell(dim_lstm)
backward_lstm_cell = tf.contrib.rnn.LSTMCell(dim_lstm)
outputs, states = tf.nn.bidirectional_dynamic_rnn(
forward_lstm_cell,
backward_lstm_cell,
inputs = inputs,
sequence_length = seqlen,
dtype = tf.float32,
scope = 'bilstm'
)
forward_outputs, backward_outputs = outputs
backward_outputs = tf.reverse_sequence(backward_outputs, tf.cast(seqlen, tf.int64), seq_dim=1)
outputs = tf.concat([forward_outputs, backward_outputs], 2)
size = tf.shape(outputs)[0]
index = tf.range(0, size) * dim_sentence + seqlen - 1
output = tf.gather(tf.reshape(outputs, [-1, dim_lstm * 2]), index) # batch_size * n_hidden * 2
predict = tf.matmul(output, lstm_w) + lstm_b
return predict
# define operations
# tf.reset_default_graph()
pred = dynamic_lstm(tf.nn.embedding_lookup(u.gloveDict, X), seqlen)
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(logits = pred, labels = y))
optimizer = tf.train.AdamOptimizer(lr).minimize(loss)
correct = tf.equal(tf.argmax(pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
# batch training
test_X, test_y, test_seqlen, _ = u.getData('test')
lrval = 0.01
with tf.Session() as sess:
sess.run(init)
for i in range(batch_iterations):
batch_X, batch_y, batch_seqlen, _ = u.nextBatch(batch_size)
lrval *= (1. / (1. + 0.2 * i))
sess.run(optimizer, feed_dict = {X: batch_X, y: batch_y, seqlen: batch_seqlen, lr: lrval})
if i > 0 and i % 4 == 0:
loss_train, accuracy_train = sess.run([loss, accuracy], feed_dict = {X: batch_X, y: batch_y, seqlen: batch_seqlen})
print('step: %s, train loss: %s, train accuracy: %s' % (i, loss_train, accuracy_train))
loss_test, accuracy_test = sess.run([loss, accuracy], feed_dict = {X: test_X, y: test_y, seqlen: test_seqlen})
print('step: %s, test loss: %s, test accuracy: %s' % (i, loss_test, accuracy_test))
# full dataset training
test_X, test_y, test_seqlen, _ = u.getData('test')
train_X, train_y, train_seqlen, _ = u.getData('train')
lrval = 0.01
with tf.Session() as sess:
sess.run(init)
for i in range(full_iterations):
lrval *= (1. / (1. + 0.2 * i))
sess.run(optimizer, feed_dict = {X: train_X, y: train_y, seqlen: train_seqlen, lr: lrval})
if i > 0 and i % 4 == 0:
loss_train, accuracy_train = sess.run([loss, accuracy], feed_dict = {X: train_X, y: train_y, seqlen: train_seqlen, lr: 0.01})
print('step: %s, train loss: %s, train accuracy: %s' % (i, loss_train, accuracy_train))
loss_test, accuracy_test = sess.run([loss, accuracy], feed_dict = {X: test_X, y: test_y, seqlen: test_seqlen, lr: 0.01})
print('step: %s, test loss: %s, test accuracy: %s' % (i, loss_test, accuracy_test))
| data_analysis/yelp_preprocess/.ipynb_checkpoints/aspect-bilstm-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, execute, Aer, IBMQ, QuantumRegister
from qiskit.compiler import transpile, assemble
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
import numpy as np
import qiskit as qk
from fractions import Fraction
from qiskit.providers.ibmq import least_busy
from qiskit.tools.monitor import job_monitor
# Loading your IBM Q account(s)
provider = IBMQ.load_account()
# -
simulator = qk.BasicAer.get_backend('qasm_simulator')
real = least_busy(provider.backends(filters=lambda x: x.configuration().n_qubits > 4,
operational=True, simulator=False))
print(real)
# ### A Bernstein-Vazirani Probléma
#
# Adott egy feketedoboz függvény ami egy $\underline{x} = (x_1, x_2, ..., x_n)$ bitsorhoz rendel hozzá 0-t vagy 1-et:<br><br>\begin{equation}
# f(x_1, x_2, ..., x_n) = 0 \text{ or } 1.
# \end{equation}
# (Minden $i$-re $x_i$ 0 vagy 1.) Tudjuk, hogy a függvény minden $\underline{x}$ bitsor esetén egy $\underline{s}$ bitsorral vett bitenkénti szorzatának 2 szerinti maradékát adja vissza. A bitenkénti szorzat alatt a következő műveletet értjük: $\underline{s}\cdot\underline{x} = x_1\cdot s_1 +x_2\cdot s_2 +...+x_n\cdot s_n$. Tehát a függvényt a következő alakban írható fel: $f(\underline{x}) = \underline{s}\cdot\underline{x} \text{ mod } 2$. Az a feladat, hogy találjuk ki, hogy mi az $\underline{s}$ bitsor.<br>
# **1. feladat**
#
# Gondoljuk végig, hogy hogyan oldanánk meg A Berstein-Vazarini problémát egy klasszikus számítógéppel. Hányszor kéne ehhez elvégezni az $f$ függvényt megvalósító műveletet?
# +
n = 4
s = np.random.randint(0, 2, n)
def f(x, s): # x egy és s azonos hosszúságú numpy array-ok
if len(x) != n:
raise ValueError("x and s have to be of the same length")
return np.dot(x, s)%2
# -
# **2. feladat**
# Határozzuk meg a véletlenszerűen generált $s$ bitsort, anélkül, hogy kiírnánk az értékét. Használjuk az f(x, s) függvényt, ami az $\underline{s}\cdot\underline{x} \text{ mod } 2$ értéket adja vissza.
# +
# megoldás helye
# -
# **Állítás**: Az alábbi ábrán látható kvantumáramkör elvégzése után pontosan az $s$ bitsort mérjük. (Az utolsó qubitet nem kell mérni, annak értéke nem érdekel minket.) Tehát elég egyszer elvégeznünk az $f$ függvényt megvalósító műveletet.
#
# 
# A $|-\rangle$ állapotot úgy állíthatjuk elő, hogy az $|{1}\rangle$ állapotra hatunk egy $H$ kapuval.
def black_box(s): # s egy bitsor
n = len(s)
qc = QuantumCircuit(n+1)
for i in range(len(s)):
if s[n-i-1] == 1:
qc.cx(i, n)
qc.name = "f"
return qc
"""az i egész szám bináris alakját írja be a függvény n darab qubitbe"""
def encode(i, n):
if 2**n <= i:
raise ValueError("'i' is too big to be stored on n qubits")
bits = np.array(list(format(i, "b")), dtype=int)
while len(bits) < n:
bits = np.insert(bits, 0, 0)
qc = QuantumCircuit(n)
for j in range(len(bits)):
if bits[j] == 1:
qc.x(n-j-1)
qc.name = "%i" %i
return qc
# **3. feladat (szorgalmi)**
#
# Ellenőrizzük, hogy a black_box(s) kvantumkapu úgy működik-e ahogy azt elvárjuk tőle az $\underline{s}=(1, 0, 1, 1)$ bitsor esetén:
# - Hozzunk létre egy 5 qubites kvantumáramkört.
# - Írjunk bele egy $\underline{x}$ bitsort az első 4 qubitbe. Ehhez használhatjuk az encode($i$, $n$) függvényt, ami az $i$ egész szám bináris alakját írja bele $n$ darab qubitbe, de a függvény nélkül is könnyen megoldható a feladat.
# - Hattassuk a black_box(s) kaput az 5 qubitre, majd mérjük meg az 5. qubitet.
# - Ha a black_box(s) kvantumkapu jól működik az $\underline{x}_0 = (0, 0, 0, 1)$, $\underline{x}_1 = (0, 0, 1, 0)$, $\underline{x}_2 = (0, 1, 0, 0)$, $\underline{x}_3 = (1, 0, 0, 0)$ bemeneti bitsorokra, akkor minden bemeneti bitsorra jól működik.
#
# +
s = np.array([1, 0, 1, 1])
# megoldás helye
# -
# **4.feladat**
# Rakjuk össze a fenti ábrán látható áramkört. Az $f$-el jelölt kapu helyére rakjuk a black_box($\underline{s}$) kaput. Legyen $\underline{s} = (1, 0, 1, 1)$.
# Ellenőrizzük, hogy a kvantumáramkör mérésekor tényleg visszakapjuk-e az $s$ bitsort. (Az áramkört futtathatjuk $\underline{s}$ más értékeire is.) Próbáljuk ki a kvantumáramkört szimulátoron is és igazi kvantumszámítógépen is.
# +
# megoldás helye
| szakkor_files/Berstein-Vazirani.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Masuyama 氏らの手法に基づく位相復元 (ADMM法とする)
# <NAME>, <NAME> and <NAME>, "Griffin-Lim like phase recovery via alternating directionmethod of multipliers," IEEE Signal Processing Letters, vol.26, no.1, pp.184--188, Jan. 2019. https://ieeexplore.ieee.org/document/8552369
import numpy as np
from scipy.io import wavfile
import librosa
from pypesq import pesq
from IPython.display import Audio
IN_WAVE_FILE = "in.wav" # モノラル音声
OUT_WAVE_FILE = "out_admm_gla.wav" # 復元音声
FRAME_LENGTH = 1024 # フレーム長 (FFTサイズ)
HOP_LENGTH = 80 # フレームのシフト長
ITERATION = 100 # 位相推定の最大繰り返し数
MULTIPLIER = 0.01 # ADMM法の強さを制御; 0.0のときはGriffin-Lim法に一致
# 音声のロード
fs, data = wavfile.read(IN_WAVE_FILE)
data = data.astype(np.float64)
# ## 振幅スペクトル(位相復元なので手に入るのはこれのみ)
# 振幅スペクトル(位相復元なので手に入るのはこれのみ)
amp_spec = np.abs(
librosa.core.stft(
data, n_fft=FRAME_LENGTH, hop_length=HOP_LENGTH, win_length=FRAME_LENGTH
)
)
# 乱数の種を指定して再現性を保証
np.random.seed(seed=0)
# ## ADMM法に基づく位相スペクトルの推定
for i in range(ITERATION):
if i == 0:
# 初回は乱数で初期化
phase_spec = np.random.rand(*amp_spec.shape)
control_spec = np.zeros(amp_spec.shape)
else:
# 振幅スペクトルと推定された位相スペクトルから複素スペクトログラムを復元
recovered_spec = amp_spec * np.exp(1j * phase_spec)
# 短時間フーリエ逆変換で音声を復元
combined = recovered_spec + control_spec
recovered = librosa.core.istft(
combined, hop_length=HOP_LENGTH, win_length=FRAME_LENGTH
)
# 復元音声から複素スペクトログラムを再計算
complex_spec = librosa.core.stft(
recovered,
n_fft=FRAME_LENGTH,
hop_length=HOP_LENGTH,
win_length=FRAME_LENGTH,
)
complex_spec = MULTIPLIER * combined + complex_spec
complex_spec /= 1.0 + MULTIPLIER
# 初回以降は計算済みの複素スペクトログラムから位相スペクトルを推定
control_spec = control_spec + recovered_spec - complex_spec
phase_spec = np.angle(complex_spec - control_spec)
# 音声を復元
recovered_spec = amp_spec * np.exp(1j * phase_spec)
recovered_admm = librosa.core.istft(
recovered_spec, hop_length=HOP_LENGTH, win_length=FRAME_LENGTH
)
# pesqで音質を評価
print("PESQ = ", pesq(recovered_admm, data[: len(recovered)], fs))
# ## 比較のため、Griffin-Lim法による位相復元
# <NAME> and <NAME>, “Signal estimation from modified short-time Fourier transform,”
# IEEE Trans. ASSP, vol.32, no.2, pp.236–243, Apr. 1984.
# https://ieeexplore.ieee.org/document/1164317
# Griffin-Lim法に基づく位相スペクトルの推定
for i in range(ITERATION):
if i == 0:
# 初回は乱数で初期化
phase_spec = np.random.rand(*amp_spec.shape)
else:
# 振幅スペクトルと推定された位相スペクトルから複素スペクトログラムを復元
recovered_spec = amp_spec * np.exp(1j * phase_spec)
# 短時間フーリエ逆変換で音声を復元
recovered = librosa.core.istft(recovered_spec, hop_length=HOP_LENGTH,
win_length=FRAME_LENGTH)
# 復元音声から複素スペクトログラムを再計算
complex_spec = librosa.core.stft(recovered, n_fft=FRAME_LENGTH,
hop_length=HOP_LENGTH,
win_length=FRAME_LENGTH)
# 初回以降は計算済みの複素スペクトログラムから位相スペクトルを推定
phase_spec = np.angle(complex_spec)
# + code_folding=[0]
# 音声を復元
recovered_spec = amp_spec * np.exp(1j * phase_spec)
recovered_gla = librosa.core.istft(recovered_spec, hop_length=HOP_LENGTH,
win_length=FRAME_LENGTH)
# pesqで音質を評価
print("PESQ = ", pesq(recovered_gla, data[: len(recovered_gla)], fs))
# -
# ## 比較のため、Fast Griffin-Lim法による位相復元
# <NAME>., <NAME>., & <NAME>. “A fast Griffin-Lim algorithm,”
# IEEE Workshop on Applications of Signal Processing to Audio and Acoustics (pp. 1-4), Oct. 2013.
# https://ieeexplore.ieee.org/document/6701851
# 比較のため、Griffin-Lim法により位相復元
recovered_fgla = librosa.griffinlim(
amp_spec, n_iter=ITERATION, hop_length=HOP_LENGTH, random_state=0
)
# pesqで音質を評価
print("PESQ = ", pesq(recovered_fgla, data[: len(recovered_fgla)], fs))
# ## 音声の聴き比べ
# ### 元音声
Audio(data, rate=fs)
# ### ADMM法による復元
Audio(recovered_admm, rate=fs)
# ### 従来のGriffin-Lim法による復元
Audio(recovered_gla, rate=fs)
# ### Fast Griffin-Lim法による復元
Audio(recovered_fgla, rate=fs)
| SpeechAnalysis/feat_gla_admm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # Vocabulary and Matching
# So far we've seen how a body of text is divided into tokens, and how individual tokens are parsed and tagged with parts of speech, dependencies and lemmas.
#
# In this section we will identify and label specific phrases that match patterns we can define ourselves.
# ## Rule-based Matching
# spaCy offers a rule-matching tool called `Matcher` that allows you to build a library of token patterns, then match those patterns against a Doc object to return a list of found matches. You can match on any part of the token including text and annotations, and you can add multiple patterns to the same matcher.
# Perform standard imports
import spacy
nlp = spacy.load('en_core_web_sm')
# Import the Matcher library
from spacy.matcher import Matcher
matcher = Matcher(nlp.vocab)
# <font color=green>Here `matcher` is an object that pairs to the current `Vocab` object. We can add and remove specific named matchers to `matcher` as needed.</font>
# ### Creating patterns
# In literature, the phrase 'solar power' might appear as one word or two, with or without a hyphen. In this section we'll develop a matcher named 'SolarPower' that finds all three:
# +
pattern1 = [{'LOWER': 'solarpower'}]
pattern2 = [{'LOWER': 'solar'}, {'LOWER': 'power'}]
pattern3 = [{'LOWER': 'solar'}, {'IS_PUNCT': True}, {'LOWER': 'power'}]
matcher.add('SolarPower',
[pattern1, pattern2, pattern3],
on_match=None)
# -
# Let's break this down:
# * `pattern1` looks for a single token whose lowercase text reads 'solarpower'
# * `pattern2` looks for two adjacent tokens that read 'solar' and 'power' in that order
# * `pattern3` looks for three adjacent tokens, with a middle token that can be any punctuation.<font color=green>*</font>
#
# <font color=green>\* Remember that single spaces are not tokenized, so they don't count as punctuation.</font>
# <br>Once we define our patterns, we pass them into `matcher` with the name 'SolarPower', and set *callbacks* to `None` (more on callbacks later).
# ### Applying the matcher to a Doc object
doc = nlp(u'The Solar Power industry continues to grow as demand \
for solarpower increases. Solar-power cars are gaining popularity.')
found_matches = matcher(doc)
print(found_matches)
# `matcher` returns a list of tuples. Each tuple contains an ID for the match, with start & end tokens that map to the span `doc[start:end]`
for match_id, start, end in found_matches:
string_id = nlp.vocab.strings[match_id] # get string representation
span = doc[start:end] # get the matched span
print(match_id, string_id, start, end, span.text)
# The `match_id` is simply the hash value of the `string_ID` 'SolarPower'
# ### Setting pattern options and quantifiers
# You can make token rules optional by passing an `'OP':'*'` argument. This lets us streamline our patterns list:
# +
# Redefine the patterns:
pattern1 = [{'LOWER': 'solarpower'}]
pattern2 = [{'LOWER': 'solar'}, {'IS_PUNCT': True, 'OP':'*'}, {'LOWER': 'power'}]
# Remove the old patterns to avoid duplication:
#matcher.remove('SolarPower')
# Add the new set of patterns to the 'SolarPower' matcher:
matcher.add('SolarPower', [pattern1, pattern2], on_match=None)
# -
found_matches = matcher(doc)
print(found_matches)
# This found both two-word patterns, with and without the hyphen!
#
# The following quantifiers can be passed to the `'OP'` key:
# <table><tr><th>OP</th><th>Description</th></tr>
#
# <tr ><td><span >\!</span></td><td>Negate the pattern, by requiring it to match exactly 0 times</td></tr>
# <tr ><td><span >?</span></td><td>Make the pattern optional, by allowing it to match 0 or 1 times</td></tr>
# <tr ><td><span >\+</span></td><td>Require the pattern to match 1 or more times</td></tr>
# <tr ><td><span >\*</span></td><td>Allow the pattern to match zero or more times</td></tr>
# </table>
#
# ### Be careful with lemmas!
# If we wanted to match on both 'solar power' and 'solar powered', it might be tempting to look for the *lemma* of 'powered' and expect it to be 'power'. This is not always the case! The lemma of the *adjective* 'powered' is still 'powered':
# +
pattern1 = [{'LOWER': 'solarpower'}]
pattern2 = [{'LOWER': 'solar'}, {'IS_PUNCT': True, 'OP':'*'}, {'LEMMA': 'power'}] # CHANGE THIS PATTERN
# Remove the old patterns to avoid duplication:
#matcher.remove('SolarPower')
# Add the new set of patterns to the 'SolarPower' matcher:
matcher.add('SolarPower',[pattern1, pattern2])
# -
doc2 = nlp(u'Solar-powered energy runs solar-powered cars.')
found_matches = matcher(doc2)
print(found_matches)
# <font color=green>The matcher found the first occurrence because the lemmatizer treated 'Solar-powered' as a verb, but not the second as it considered it an adjective.<br>For this case it may be better to set explicit token patterns.</font>
# +
pattern1 = [{'LOWER': 'solarpower'}]
pattern2 = [{'LOWER': 'solar'}, {'IS_PUNCT': True, 'OP':'*'}, {'LOWER': 'power'}]
pattern3 = [{'LOWER': 'solarpowered'}]
pattern4 = [{'LOWER': 'solar'}, {'IS_PUNCT': True, 'OP':'*'}, {'LOWER': 'powered'}]
# Remove the old patterns to avoid duplication:
#matcher.remove('SolarPower')
# Add the new set of patterns to the 'SolarPower' matcher:
matcher.add('SolarPower',[pattern1, pattern2, pattern3, pattern4])
# -
found_matches = matcher(doc2)
print(found_matches)
# ## Other token attributes
# Besides lemmas, there are a variety of token attributes we can use to determine matching rules:
# <table><tr><th>Attribute</th><th>Description</th></tr>
#
# <tr ><td><span >`ORTH`</span></td><td>The exact verbatim text of a token</td></tr>
# <tr ><td><span >`LOWER`</span></td><td>The lowercase form of the token text</td></tr>
# <tr ><td><span >`LENGTH`</span></td><td>The length of the token text</td></tr>
# <tr ><td><span >`IS_ALPHA`, `IS_ASCII`, `IS_DIGIT`</span></td><td>Token text consists of alphanumeric characters, ASCII characters, digits</td></tr>
# <tr ><td><span >`IS_LOWER`, `IS_UPPER`, `IS_TITLE`</span></td><td>Token text is in lowercase, uppercase, titlecase</td></tr>
# <tr ><td><span >`IS_PUNCT`, `IS_SPACE`, `IS_STOP`</span></td><td>Token is punctuation, whitespace, stop word</td></tr>
# <tr ><td><span >`LIKE_NUM`, `LIKE_URL`, `LIKE_EMAIL`</span></td><td>Token text resembles a number, URL, email</td></tr>
# <tr ><td><span >`POS`, `TAG`, `DEP`, `LEMMA`, `SHAPE`</span></td><td>The token's simple and extended part-of-speech tag, dependency label, lemma, shape</td></tr>
# <tr ><td><span >`ENT_TYPE`</span></td><td>The token's entity label</td></tr>
#
# </table>
# ### Token wildcard
# You can pass an empty dictionary `{}` as a wildcard to represent **any token**. For example, you might want to retrieve hashtags without knowing what might follow the `#` character:
# >`[{'ORTH': '#'}, {}]`
# ___
# ## PhraseMatcher
# In the above section we used token patterns to perform rule-based matching. An alternative - and often more efficient - method is to match on terminology lists. In this case we use PhraseMatcher to create a Doc object from a list of phrases, and pass that into `matcher` instead.
# Perform standard imports, reset nlp
import spacy
nlp = spacy.load('en_core_web_sm')
# Import the PhraseMatcher library
from spacy.matcher import PhraseMatcher
matcher = PhraseMatcher(nlp.vocab)
# For this exercise we're going to import a Wikipedia article on *Reaganomics*<br>
# Source: https://en.wikipedia.org/wiki/Reaganomics
with open('../TextFiles/reaganomics.txt', encoding='unicode_escape') as f:
doc3 = nlp(f.read())
# +
# First, create a list of match phrases:
phrase_list = ['voodoo economics', 'supply-side economics', 'trickle-down economics', 'free-market economics']
# Next, convert each phrase to a Doc object:
phrase_patterns = [nlp(text) for text in phrase_list]
# Pass each Doc object into matcher (note the use of the asterisk!):
matcher.add('VoodooEconomics', None, *phrase_patterns)
# Build a list of matches:
matches = matcher(doc3)
# -
# (match_id, start, end)
matches
for match_id, start, end in found_matches:
string_id = nlp.vocab.strings[match_id] # get string representation
span = doc3[start-5:end+5] # get the matched span
print(match_id, string_id, start, end, span.text)
# <font color=green>The first four matches are where these terms are used in the definition of Reaganomics:</font>
doc3[:70]
# ## Viewing Matches
# There are a few ways to fetch the text surrounding a match. The simplest is to grab a slice of tokens from the doc that is wider than the match:
doc3[665:685] # Note that the fifth match starts at doc3[673]
doc3[2975:2995] # The sixth match starts at doc3[2985]
# Another way is to first apply the `sentencizer` to the Doc, then iterate through the sentences to the match point:
# +
# Build a list of sentences
sents = [sent for sent in doc3.sents]
# In the next section we'll see that sentences contain start and end token values:
print(sents[0].start, sents[0].end)
# -
# Iterate over the sentence list until the sentence end value exceeds a match start value:
for sent in sents:
if matches[4][1] < sent.end: # this is the fifth match, that starts at doc3[673]
print(sent)
break
# For additional information visit https://spacy.io/usage/linguistic-features#section-rule-based-matching
# ## Next Up: NLP Basics Assessment
| 05-Vocabulary-and-Matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !python main.py --model baseline --epochs 30
# !python main.py --mode test --model baseline
# !python main.py --model baseline_attetion --epochs 30
# !python main.py --model baseline_attetion --mode test
# !python main.py --model baseline_attetion_fc --epochs 30
# !python main.py --model baseline_attetion_fc --mode test
# !python main.py --model dense_attention --epochs 30
# !python main.py --model dense_attention --mode test
# !python main.py --model baseline_preconv_attention --epochs 30
# !python main.py --model baseline_preconv_attention --mode test
| bin/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Riddler Battle Royale
#
#
#
# > [538's *The Riddler* Asks](http://fivethirtyeight.com/features/the-battle-for-riddler-nation-round-2/): *In a distant, war-torn land, there are 10 castles. There are two warlords: you and your archenemy, with whom you’re competing to collect the most victory points. Each castle has its own strategic value for a would-be conqueror. Specifically, the castles are worth 1, 2, 3, …, 9, and 10 victory points. You and your enemy each have 100 soldiers to distribute, any way you like, to fight at any of the 10 castles. Whoever sends more soldiers to a given castle conquers that castle and wins its victory points. If you each send the same number of troops, you split the points. You don’t know what distribution of forces your enemy has chosen until the battles begin. Whoever wins the most points wins the war. Submit a plan distributing your 100 soldiers among the 10 castles.*
#
#
# Load some useful modules
# %matplotlib inline
import matplotlib.pyplot as plt
import csv
import random
from collections import Counter
from statistics import mean
# Let's play with this and see if we can find a good solution. Some implementation choices:
# * A `Plan` will be a tuple of 10 soldier counts (one for each castle).
# * `castles` will hold the indexes of the castles. Note that index 0 is castle 1 (worth 1 point) and index 9 is castle 10 (worth 10 points).
# * `half` is half the total number of points; if you get more than this you win.
# * `plans` will hold a set of plans that were submitted in the previous contest.
# * `play(A, B)` gives the single game reward for Plan A against Plan B: 1 if A wins, 0 if A loses, and 1/2 for a tie.
# * `reward(a, b, payoff)` returns payoff, payoff/2, or 0, depending on whether `a` is bigger than `b`.
# +
Plan = tuple
castles = range(10)
half = 55/2
plans = {Plan(map(int, row[:10]))
for row in csv.reader(open('battle_royale.csv'))}
def play(A, B):
"Play Plan A against Plan B and return a reward (0, 1/2, or 1)."
A_points = sum(reward(A[c], B[c], c + 1) for c in castles)
return reward(A_points, half)
def reward(a, b, payoff=1): return (payoff if a > b else payoff / 2 if a == b else 0)
# -
# Some tests:
# +
assert reward(6, 5, 9) == 9 # 6 soldiers defeat 5, winning all 9 of the castle's points
assert reward(6, 6, 8) == 4 # A tie on an 8-point castle is worth 4 points
assert reward(6, 7, 7) == 0 # No points for a loss
assert reward(30, 25) == 1 # 30 victory points beats 25
assert len(plans) == 1202
assert play((26, 5, 5, 5, 6, 7, 26, 0, 0, 0),
(25, 0, 0, 0, 0, 0, 0, 25, 25, 25)) == 1 # A wins game
assert play((26, 5, 5, 5, 6, 7, 26, 0, 0, 0),
(0, 25, 0, 0, 0, 0, 0, 25, 25, 25)) == 0 # B wins game
assert play((25, 5, 5, 5, 6, 7, 26, 0, 0, 0),
(25, 0, 0, 0, 0, 0, 0, 25, 25, 25)) == 1/2 # Tie game
# -
# Let's run a tournament, playing each plan against every other, and returning a list of `[(plan, mean_game_points),...]`. I will also define `show` to pretty-print these results and display a histogram:
# +
def tournament(plans):
"Play each plan against each other; return a sorted list of [(plan: mean_points)]"
rankdict = {A: mean_points(A, plans) for A in plans}
return Counter(rankdict).most_common()
def mean_points(A, opponents):
"Mean points for A playing against all opponents (but not against itself)."
return mean(play(A, B) for B in opponents if B is not A)
def show(rankings, n=10):
"Pretty-print the n best plans, and display a histogram of all plans."
print('Top', n, 'of', len(rankings), 'plans:')
for (plan, points) in rankings[:n]:
print(pplan(plan), pct(points))
plt.hist([s for (p, s) in rankings], bins=20)
def pct(x): return '{:6.1%}'.format(x)
def pplan(plan): return '(' + ', '.join('{:2}'.format(c) for c in plan) + ')'
# -
# This is what the result of a tournament looks like:
tournament({(26, 5, 5, 5, 6, 7, 26, 0, 0, 0),
(25, 0, 0, 0, 0, 0, 0, 25, 25, 25),
(0, 25, 0, 0, 0, 0, 0, 25, 25, 25)})
# A tournament with all 1202 plans:
rankings = tournament(plans)
show(rankings)
# It looks like there are a few really bad plans in there. Let's just keep the top 1000 plans (out of 1202), and re-run the rankings:
plans = {A for (A, _) in rankings[:1000]}
rankings = tournament(plans)
show(rankings)
# The top 10 plans are still winning over 80%, and the top plan remains `(0, 3, 4, 7, 16, 24, 4, 34, 4, 4)`. This is an interesting plan: it places most of the soldiers on castles 4+5+6+8, which totals only 23 points, so it needs to pick up 5 more points from the other castles (that have mostly 4 soldiers attacking each one). Is this a good strategy? Where should we optiomally allocate soldiers?
#
# To gain some insight, I'll create a plot with 10 curves, one for each castle. Each curve maps the number of soldiers sent to the castle (on the x-axis) to the expected points won (against the 1000 plans) on the y-axis:
#
# +
def plotter(plans, X=range(41)):
X = list(X)
def mean_reward(c, s): return mean(reward(s, p[c], c+1) for p in plans)
for c in range(10):
plt.plot(X, [mean_reward(c, s) for s in X], '.-')
plt.xlabel('Number of soldiers (on each of the ten castles)')
plt.ylabel('Expected points won')
plt.grid()
plotter(plans)
# -
# For example, this says that for castle 10 (the orange line at top), there is a big gain in expected return as we increase from 0 to 4 soldiers, and after that the gains are relatively less steep. This plot is interesting, but I can't see how to directly read off a best plan from it.
#
# ## Hillclimbing
#
# Instead I'll see if I can improve the existing plans, using a simple *hillclimbing* strategy: Take a Plan A, and change it by randomly moving some soldiers from one castle to another. If that yields more `mean_points`, then keep the updated plan, otherwise discard it. Repeat.
# +
def hillclimb(A, plans=plans, steps=1000):
"Try to improve Plan A, repeat `steps` times; return new plan and total."
m = mean_points(A, plans)
for _ in range(steps):
B = mutate(A)
m, A = max((m, A),
(mean_points(B, plans), B))
return A, m
def mutate(plan):
"Return a new plan that is a slight mutation."
plan = list(plan) # So we can modify it.
i, j = random.sample(castles, 2)
plan[i], plan[j] = random_split(plan[i] + plan[j])
return Plan(plan)
def random_split(n):
"Split the integer n into two integers that sum to n."
r = random.randint(0, n)
return r, n - r
# -
# Let's see how well this works. Remember, the best plan so far had a score of `87.4%`. Can we improve on that?
hillclimb((0, 3, 4, 7, 16, 24, 4, 34, 4, 4))
# We got an improvement. Let's see what happens if we start with other plans:
hillclimb((10, 10, 10, 10, 10, 10, 10, 10, 10, 10))
hillclimb((0, 1, 2, 3, 4, 18, 18, 18, 18, 18))
hillclimb((2, 3, 5, 5, 5, 20, 20, 20, 10, 10))
hillclimb((0, 0, 5, 5, 25, 3, 25, 3, 31, 3))
# What if we hillclimb 20 times longer?
hillclimb((0, 3, 4, 7, 16, 24, 4, 34, 4, 4), steps=20000)
# ## Opponent modeling
#
# To have a chance of winning the second round of this contest, we have to predict what the other entries will be like. Nobody knows for sure, but I can hypothesize that the entries will be slightly better than the first round, and try to approximate that by hillclimbing from each of the first-round plans for a small number of steps:
def hillclimbers(plans, steps=100):
"Return a sorted list of [(improved_plan, mean_points), ...]"
pairs = {hillclimb(plan, plans, steps) for plan in plans}
return sorted(pairs, key=lambda pair: pair[1], reverse=True)
# For example:
hillclimbers({(26, 5, 5, 5, 6, 7, 26, 0, 0, 0),
(25, 0, 0, 0, 0, 0, 0, 25, 25, 25),
(0, 25, 0, 0, 0, 0, 0, 25, 25, 25)})
# I will define `plans2` (and `rankings2`) to be my estimate of the entries for round 2:
# %time rankings2 = hillclimbers(plans)
plans2 = {A for (A, _) in rankings2}
show(rankings2)
# Even though we only took 100 steps, the `plans2` plans are greatly improved: Almost all of them defeat 75% or more of the first-round `plans`. The top 10 plans are all very similar, targeting castles 4+6+8+10 (for 28 points), but reserving 20 or soldiers to spread among the other castles. Let's look more carefully at every 40th plan, plus the last one:
for (p, m) in rankings2[::40] + [rankings2[-1]]:
print(pplan(p), pct(m))
# We see a wider variety in plans as we go farther down the rankings. Now for the plot:
plotter(plans2)
# We see that many castles (e.g. 9 (green), 8 (blue), 7 (black), 6 (yellowish)) have two plateaus. Castle 7 (black) has a plateau at 3.5 points for 6 to 20 soldiers (suggesting that 6 soldiers is a good investment and 20 soldiers a bad investment), and then another plateau at 7 points for everything above 30 soldiers.
#
# Now that we have an estimate of the opponents, we can use `hillclimbers` to try to find a plan that does well against all the others:
# %time rankings3 = hillclimbers(plans2)
show(rankings3)
# We can try even harder to improve the champ:
champ, _ = rankings3[0]
hillclimb(champ, plans2, 10000)
# Here are some champion plans from previous runs of this notebook:
champs = {
(0, 1, 3, 16, 20, 3, 4, 5, 32, 16),
(0, 1, 9, 16, 15, 24, 5, 5, 8, 17),
(0, 1, 9, 16, 16, 24, 5, 5, 7, 17),
(0, 2, 9, 16, 15, 24, 5, 5, 8, 16),
(0, 2, 9, 16, 15, 25, 5, 4, 7, 17),
(0, 3, 4, 7, 16, 24, 4, 34, 4, 4),
(0, 3, 5, 6, 20, 4, 4, 33, 8, 17),
(0, 4, 5, 7, 20, 4, 4, 33, 7, 16),
(0, 4, 6, 7, 19, 4, 4, 31, 8, 17),
(0, 4, 12, 18, 21, 7, 6, 4, 8, 20),
(0, 4, 12, 19, 25, 4, 5, 6, 8, 17),
(0, 5, 6, 7, 18, 4, 5, 32, 7, 16),
(0, 5, 7, 3, 18, 4, 4, 34, 8, 17),
(1, 2, 9, 16, 15, 24, 5, 4, 7, 17),
(1, 2, 9, 16, 15, 24, 5, 4, 8, 16),
(1, 2, 11, 16, 15, 24, 5, 4, 7, 15),
(1, 3, 14, 18, 24, 4, 5, 6, 8, 17),
(1, 6, 3, 16, 16, 24, 5, 5, 7, 17),
(2, 3, 7, 16, 16, 25, 5, 5, 8, 13),
(2, 3, 8, 16, 12, 25, 5, 4, 8, 17),
(2, 3, 8, 16, 15, 24, 5, 4, 7, 16),
(2, 3, 8, 16, 15, 25, 4, 5, 8, 14),
(2, 3, 8, 16, 16, 24, 5, 5, 8, 13),
(2, 3, 9, 15, 12, 25, 4, 5, 8, 17),
(2, 3, 9, 16, 12, 24, 5, 5, 8, 16),
(2, 4, 12, 18, 24, 4, 6, 5, 8, 17),
(3, 3, 7, 16, 16, 24, 5, 5, 8, 13),
(3, 3, 8, 16, 12, 25, 4, 4, 8, 17),
(3, 3, 8, 16, 15, 25, 5, 4, 7, 14),
(3, 4, 12, 18, 23, 4, 6, 5, 8, 17),
(3, 4, 15, 18, 23, 4, 5, 6, 8, 14),
(3, 5, 7, 16, 5, 4, 5, 34, 7, 14),
(3, 6, 13, 17, 23, 4, 6, 5, 8, 15),
(4, 3, 12, 18, 23, 4, 5, 6, 8, 17),
(4, 5, 3, 15, 11, 23, 5, 5, 10, 19),
(4, 6, 3, 16, 14, 25, 5, 5, 8, 14),
(4, 6, 3, 16, 16, 24, 5, 5, 7, 14),
(4, 6, 3, 16, 16, 24, 5, 5, 8, 13),
(5, 3, 12, 17, 23, 4, 5, 6, 8, 17),
(5, 5, 3, 16, 12, 25, 4, 5, 8, 17),
(5, 6, 3, 16, 16, 24, 5, 5, 7, 13),
(5, 6, 7, 3, 21, 4, 27, 5, 8, 14),
(5, 6, 8, 3, 18, 4, 27, 5, 8, 16),
(5, 6, 8, 3, 20, 4, 27, 5, 8, 14),
(5, 6, 8, 3, 21, 4, 27, 5, 8, 13)}
# We can evaluate each of them against the original `plans`, against the improved `plans2`, against their fellow champs, and against all of those put together:
# +
def μ(plan, plans): return pct(mean_points(plan,plans))
all = plans | plans2 | champs
print('Plan plans plans2 champs all')
for p in sorted(champs, key=lambda p: -mean_points(p, all)):
print(pplan(p), μ(p, plans), μ(p, plans2), μ(p, champs), μ(p, all))
# -
# Which plan is best? In the end, we don't know, because we don't know the pool we will be competing against.
| ipynb/Riddler Battle Royale.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import os
import joblib
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
from scipy.signal import gaussian
from scipy.ndimage import filters
from sklearn.utils import resample
save = False
cm = 1/2.54 # centimeters in inches
sns.set_context('paper', rc={'font.size': 10, 'xtick.labelsize': 8, 'ytick.labelsize': 8,
'figure.titleweight': 'bold', 'axes.labelsize':10, 'axes.titlesize':12})
dir_path = os.path.abspath('')
output_path = dir_path + '/Figures/'
# -
def smooth_response(response, filterWidth=8):
"""
:param response: Trials x Times
:param filterWidth: SD of Gaussian
:return: Smoothed response
"""
if len(response.shape) == 1:
response = response[np.newaxis, :]
gauss = gaussian(10 * filterWidth, filterWidth)
return filters.convolve1d(response, gauss / gauss.sum(), axis=1)
# +
tracker = joblib.load(dir_path + '/ModelAnalysis/FiringRates_readout.pkl')
colors = sns.color_palette(sns.xkcd_palette([ "orange yellow","windows blue", "greyish", "faded green", "dusty purple","orange", "grey"]))
# +
offset = 0
onset = 100
features = 256 # 512
time_steps = np.arange(-onset, tracker['neutral']['MUA'].shape[1] - onset) #np.arange(offset, tracker[25]['neutral']['MUA_centre'].shape[1])
conds = ['precision','inputGain','outputGain']
n = tracker['neutral']['MUA'].shape[0]
num_draws = 500
np.random.seed(3)
idx = np.arange(n)
FR_avgs = pd.DataFrame(np.zeros((len(conds) * 3 * n,4)), columns=['Mechanism', 'Attention', 'Image','FR'])
FR_avgs['Attention'] = np.repeat(np.arange(3)[np.newaxis,:],n *len(conds), axis=0).flatten()
FR_avgs['Mechanism'] = np.repeat(np.arange(len(conds))[np.newaxis,:],n *3, axis=1).flatten()
FR_avgs['Image'] = np.repeat(np.repeat(np.arange(n)[:,np.newaxis],3).flatten()[np.newaxis,:], len(conds), axis=0).flatten()
replace_map = {'Attention': {0: 'valid', 1:'neutral', 2:'invalid'},
'Mechanism': {0: conds[0], 1: conds[1], 2: conds[2]}}
FR_avgs.replace(replace_map, inplace=True)
resp = tracker['neutral']['MUA'][:n, offset:]
resp_gauss = smooth_response(resp)
mean_draws_neutral = list()
for d in range(num_draws):
# Resampling, with replacement, taking the class imbalance into account
draw = resample(idx, n_samples=n, random_state=d)
mean_draws_neutral.append(np.mean(resp_gauss[draw], axis=0))
# +
fig, ax = plt.subplots(1, 3, figsize=(13.5*cm, 5*cm), sharey=True, sharex=True)
ax = ax.flatten()
for c, cond in enumerate(conds):
for a, att in enumerate(['valid', 'invalid', 'neutral']):
if att == 'neutral':
resp = tracker[att]['MUA'][:n, offset:]
resp_gauss = smooth_response(resp)
mean_draws = mean_draws_neutral
else:
resp = tracker[cond][att]['MUA'][:n, offset:]
resp_gauss = smooth_response(resp)
mean_draws = list()
for d in range(num_draws):
# Resampling, with replacement, taking the class imbalance into account
draw = resample(idx, n_samples=n, random_state=d)
mean_draws.append(np.mean(resp_gauss[draw], axis=0))
ax[c].plot(time_steps,np.mean(resp_gauss, axis=0), color=colors[a])
ax[c].fill_between(time_steps, np.percentile(np.array(mean_draws), 2.5, axis=0),np.percentile(mean_draws, 97.5, axis=0),
color=colors[a], alpha=0.4)
ax[c].set_ylim([0,1])
ax[0].set_ylabel('Spike count')
ax[0].set_xlabel('Time (ms)')
ax[0].set_title('Precision')
ax[0].set_yticks([0, 0.5, 1])
ax[1].set_title('Input gain')
ax[1].set_xlabel('Time (ms)')
ax[2].set_title('Connection gain')
ax[2].set_xlabel('Time (ms)')
sns.despine()
print(cond)
FR_avgs.loc[(FR_avgs['Mechanism'] == cond) & (FR_avgs['Attention'] == 'valid'), 'FR' ] =((np.sum(
tracker[cond]['valid']['MUA'][:n,250:], axis=1))/features) * (1000 / 450)
FR_avgs.loc[(FR_avgs['Mechanism'] == cond) & (FR_avgs['Attention'] == 'invalid'), 'FR'] = ((np.sum(
tracker[cond]['invalid']['MUA'][:n, 250:], axis=1)) / features) * (1000 / 450)
FR_avgs.loc[(FR_avgs['Mechanism'] == cond) & (FR_avgs['Attention'] == 'neutral'), 'FR'] = ((np.sum(
tracker['neutral']['MUA'][:n, 250:], axis=1)) / features) * (1000 / 450)
plt.subplots_adjust(wspace=0.01, hspace=0.01)
plt.tight_layout()
if save == True:
fig.savefig( output_path + 'FiringRates_allConditions.pdf', dpi=300, transparent=True)
else:
plt.show()
# +
dodge = True
capsize = 0.1
aspect = 1.7
errwidth = 1.3
g = sns.catplot(y="Mechanism", x="FR", hue="Attention", data=FR_avgs[FR_avgs['Attention'] != 'neutral'],
height=4 * cm, aspect=aspect, kind="point",
palette={'valid': colors[0], 'invalid': colors[1], 'neutral': colors[2]},
legend=False, zorder=6, join=False, dodge=dodge,
capsize=capsize, errwidth=errwidth, orient='h')
g.set_yticklabels(['Precision', 'Input gain', 'Connection\ngain'])
means = FR_avgs.groupby(['Attention']).mean()
means = means.reset_index()
mean = means.loc[means['Attention'] == 'neutral', 'FR'].values
limits = np.array([1, 3.4])
plt.xlim(limits)
g.ax.set_xticks([1, 2, 3])
g.ax.set_xlabel('Firing rate (Hz)')
g.ax.set_ylabel(' ')
g.ax.axvline(mean, c=colors[2], ls='--', zorder=0)
plt.tight_layout()
plt.axvspan(mean * 1.05, mean * 1.3, color=colors[2], alpha=0.3, zorder=0)
plt.axvspan(mean * 0.7, mean * 0.95, color=colors[2], alpha=0.3, zorder=0)
sns.despine(top=True, right=True)
if save == True:
g.savefig(figure_path + 'FiringRates_' + mode + '.pdf', dpi=300, transparent=True)
else:
plt.show()
# +
import random
comps = ['valid vs invalid', 'valid vs neutral', 'invalid vs neutral']
results = {}
results_df = pd.DataFrame([], columns=['Mechanism', 'Comparison', 'Difference', 'p-values', 'significant'])
# Define p (number of permutations):
p = 10000
results['Permutations'] = p
results['alpha-level'] = 0.05 / len(comps)
for cond in conds:
if cond not in results:
results[cond] = {}
print(cond)
validPreds = FR_avgs.loc[(FR_avgs['Mechanism']==cond) & (FR_avgs['Attention']=='valid'), 'FR'].values
invalidPreds = FR_avgs.loc[(FR_avgs['Mechanism']==cond) & (FR_avgs['Attention']=='invalid'), 'FR'].values
neutralPreds = FR_avgs.loc[(FR_avgs['Mechanism'] == cond) & (FR_avgs['Attention'] == 'neutral'), 'FR'].values
idx = np.arange(len(validPreds) * 2)
for comp in comps:
if comp not in results[cond]:
print(comp)
if comp == 'valid vs invalid':
diff = np.abs(np.mean(validPreds) - np.mean(invalidPreds))
samplePreds = np.concatenate([validPreds, invalidPreds], axis=0)
elif comp == 'valid vs neutral':
diff = np.abs(np.mean(validPreds) - np.mean(neutralPreds))
samplePreds = np.concatenate([validPreds, neutralPreds], axis=0)
elif comp == 'invalid vs neutral':
diff = np.abs(np.mean(invalidPreds) - np.mean(neutralPreds))
samplePreds = np.concatenate([invalidPreds, neutralPreds], axis=0)
else:
raise ValueError('Comparison is not specified.')
# Initialize permutation:
pD = []
# Permutation loop:
for i in range(0, p):
random.shuffle(idx)
pD.append(np.abs(np.mean(samplePreds[idx[:int(len(idx)//2)]]) - np.mean(samplePreds[idx[int(len(idx) // 2):]])))
p_val = len(np.where(pD>=diff)[0])/p
print(cond + ': ' + str(p_val))
results[cond][comp] = {}
results[cond][comp]['Observed difference'] = diff
results[cond][comp]['Difference distribution'] = pD
results[cond][comp]['p-value']= p_val
results_df = results_df.append({'Mechanism':cond, 'Comparison':comp, 'Difference':diff, 'p-values':p_val, 'significant':p_val<results['alpha-level']},ignore_index=True )
joblib.dump(results,
dir_path + '/ModelEvaluation/FRPermutationTestsComparisons.pkl',
compress=True)
results_df.to_csv(dir_path +'/ModelEvaluation/FRPermutationTestsComparisons_df.csv')
# -
| Figures/Figure5AD_FiringRates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Many to One Classification
#
# Simple example for Many to one (word sentiment classification) by Recurrent Neural Networks
#
#
# - Creating the **data pipeline** with `tf.data`
# - Preprocessing word sequences (variable input sequence length) using `padding technique` by `user function (pad_seq)`
# - Using `tf.nn.embedding_lookup` for getting vector of tokens (eg. word, character)
# - Creating the model as **Class**
# - Reference
# - https://github.com/golbin/TensorFlow-Tutorials/blob/master/10%20-%20RNN/02%20-%20Autocomplete.py
# - https://github.com/aisolab/TF_code_examples_for_Deep_learning/blob/master/Tutorial%20of%20implementing%20Sequence%20classification%20with%20RNN%20series.ipynb
#
# ### Setup
# +
import os, sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import string
# %matplotlib inline
slim = tf.contrib.slim
print(tf.__version__)
# -
# ### Prepare example data
words = ['good', 'bad', 'amazing', 'so good', 'bull shit', 'awesome']
y = [[1.,0.], [0.,1.], [1.,0.], [1., 0.],[0.,1.], [1.,0.]]
# Character quantization
char_space = string.ascii_lowercase
char_space = char_space + ' ' + '*'
char_space
char_dic = {char : idx for idx, char in enumerate(char_space)}
print(char_dic)
# ### Create pad_seq function
def pad_seq(sequences, max_len, dic):
seq_len, seq_indices = [], []
for seq in sequences:
seq_len.append(len(seq))
seq_idx = [dic.get(char) for char in seq]
seq_idx += (max_len - len(seq_idx)) * [dic.get('*')] # 27 is idx of meaningless token "*"
seq_indices.append(seq_idx)
return seq_len, seq_indices
# ### Apply pad_seq function to data
max_length = 10
X_length, X_indices = pad_seq(sequences = words, max_len = max_length, dic = char_dic)
print(X_length)
print(np.shape(X_indices))
# ### Define CharRNN class
class CharRNN:
def __init__(self, X_length, X_indices, y, n_of_classes, hidden_dim, dic):
# data pipeline
with tf.variable_scope('input_layer'):
self._X_length = X_length
self._X_indices = X_indices
self._y = y
one_hot = tf.eye(len(dic), dtype = tf.float32)
self._one_hot = tf.get_variable(name='one_hot_embedding', initializer = one_hot,
trainable = False) # embedding vector training 안할 것이기 때문
self._X_batch = tf.nn.embedding_lookup(params = self._one_hot, ids = self._X_indices)
# RNN cell
with tf.variable_scope('rnn_cell'):
rnn_cell = tf.contrib.rnn.BasicRNNCell(num_units = hidden_dim, activation = tf.nn.tanh)
_, state = tf.nn.dynamic_rnn(cell = rnn_cell, inputs = self._X_batch,
sequence_length = self._X_length, dtype = tf.float32)
with tf.variable_scope('output_layer'):
self._score = slim.fully_connected(inputs = state, num_outputs = n_of_classes, activation_fn = None)
with tf.variable_scope('loss'):
self.ce_loss = tf.losses.softmax_cross_entropy(onehot_labels = self._y, logits = self._score)
with tf.variable_scope('prediction'):
self._prediction = tf.argmax(input = self._score, axis = -1, output_type = tf.int32)
def predict(self, sess, X_length, X_indices):
feed_prediction = {self._X_length : X_length, self._X_indices : X_indices}
return sess.run(self._prediction, feed_dict = feed_prediction)
# ### Create a model of CharRNN
# hyper-parameter#
lr = .003
epochs = 10
batch_size = 2
total_step = int(np.shape(X_indices)[0] / batch_size)
print(total_step)
## create data pipeline with tf.data
tr_dataset = tf.data.Dataset.from_tensor_slices((X_length, X_indices, y))
tr_dataset = tr_dataset.shuffle(buffer_size = 20)
tr_dataset = tr_dataset.batch(batch_size = batch_size)
tr_iterator = tr_dataset.make_initializable_iterator()
print(tr_dataset)
X_length_mb, X_indices_mb, y_mb = tr_iterator.get_next()
char_rnn = CharRNN(X_length = X_length_mb, X_indices = X_indices_mb, y = y_mb, n_of_classes = 2,
hidden_dim = 16, dic = char_dic)
# ### Creat training op and train model
## create training op
opt = tf.train.AdamOptimizer(learning_rate = lr)
training_op = opt.minimize(loss = char_rnn.ce_loss)
# +
sess = tf.Session()
sess.run(tf.global_variables_initializer())
tr_loss_hist = []
for epoch in range(epochs):
avg_tr_loss = 0
tr_step = 0
sess.run(tr_iterator.initializer)
try:
while True:
_, tr_loss = sess.run(fetches = [training_op, char_rnn.ce_loss])
avg_tr_loss += tr_loss
tr_step += 1
except tf.errors.OutOfRangeError:
pass
avg_tr_loss /= tr_step
tr_loss_hist.append(avg_tr_loss)
print('epoch : {:3}, tr_loss : {:.3f}'.format(epoch + 1, avg_tr_loss))
# -
plt.plot(tr_loss_hist, label = 'train')
yhat = char_rnn.predict(sess = sess, X_length = X_length, X_indices = X_indices)
print('training acc: {:.2%}'.format(np.mean(yhat == np.argmax(y, axis = -1))))
| week07/01_many_to_one_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get all Links
# [](https://colab.research.google.com/github/educatorsRlearners/book-maturity/blob/master/00_get_search_pages.ipynb)
import os
import string
import requests
import json
import pandas as pd
from bs4 import BeautifulSoup
from csv import writer
# ## Create links to every page
base_url = 'https://www.commonsensemedia.org/book-reviews'
page = '?page='
all_pages = range(1,291)
all_pages_list = [base_url+page+str(p) for p in all_pages]
# ## Create a folder to hold the scraped pages
# !mkdir lexile/raw/lists
# ## Make the Soup for the first page
page = requests.get(base_url)
soup = BeautifulSoup(page.text, 'html.parser')
# ### Write the Soup to a file for future analysis
with open('lexile/raw/lists/page=0.html', 'wb') as f:
f.write(soup.encode('utf-8'))
# ## Make the Soup for every page and save it for future analysis
for p in all_pages_list:
page = requests.get(p)
soup = BeautifulSoup(page.text, 'html.parser')
with open("lexile/raw/lists/" + p.split("?")[1] + ".html", 'wb') as f:
f.write(soup.encode('utf-8'))
| 00_get_search_pages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2: Naive Bayes
# Welcome to week two of this specialization. You will learn about Naive Bayes. Concretely, you will be using Naive Bayes for sentiment analysis on tweets. Given a tweet, you will decide if it has a positive sentiment or a negative one. Specifically you will:
#
# * Train a naive bayes model on a sentiment analysis task
# * Test using your model
# * Compute ratios of positive words to negative words
# * Do some error analysis
# * Predict on your own tweet
#
# You may already be familiar with Naive Bayes and its justification in terms of conditional probabilities and independence.
# * In this week's lectures and assignments we used the ratio of probabilities between positive and negative sentiments.
# * This approach gives us simpler formulas for these 2-way classification tasks.
#
# Load the cell below to import some packages.
# You may want to browse the documentation of unfamiliar libraries and functions.
from utils import process_tweet, lookup
import pdb
from nltk.corpus import stopwords, twitter_samples
import numpy as np
import pandas as pd
import nltk
import string
from nltk.tokenize import TweetTokenizer
from os import getcwd
# If you are running this notebook in your local computer,
# don't forget to download the twitter samples and stopwords from nltk.
#
# ```
# nltk.download('stopwords')
# nltk.download('twitter_samples')
# ```
# add folder, tmp2, from our local workspace containing pre-downloaded corpora files to nltk's data path
filePath = f"{getcwd()}/../tmp2/"
nltk.data.path.append(filePath)
# +
# get the sets of positive and negative tweets
all_positive_tweets = twitter_samples.strings('positive_tweets.json')
all_negative_tweets = twitter_samples.strings('negative_tweets.json')
# split the data into two pieces, one for training and one for testing (validation set)
test_pos = all_positive_tweets[4000:]
train_pos = all_positive_tweets[:4000]
test_neg = all_negative_tweets[4000:]
train_neg = all_negative_tweets[:4000]
train_x = train_pos + train_neg
test_x = test_pos + test_neg
# avoid assumptions about the length of all_positive_tweets
train_y = np.append(np.ones(len(train_pos)), np.zeros(len(train_neg)))
test_y = np.append(np.ones(len(test_pos)), np.zeros(len(test_neg)))
# -
# # Part 1: Process the Data
#
# For any machine learning project, once you've gathered the data, the first step is to process it to make useful inputs to your model.
# - **Remove noise**: You will first want to remove noise from your data -- that is, remove words that don't tell you much about the content. These include all common words like 'I, you, are, is, etc...' that would not give us enough information on the sentiment.
# - We'll also remove stock market tickers, retweet symbols, hyperlinks, and hashtags because they can not tell you a lot of information on the sentiment.
# - You also want to remove all the punctuation from a tweet. The reason for doing this is because we want to treat words with or without the punctuation as the same word, instead of treating "happy", "happy?", "happy!", "happy," and "happy." as different words.
# - Finally you want to use stemming to only keep track of one variation of each word. In other words, we'll treat "motivation", "motivated", and "motivate" similarly by grouping them within the same stem of "motiv-".
#
# We have given you the function `process_tweet()` that does this for you.
# +
custom_tweet = "RT @Twitter @chapagain Hello There! Have a great day. :) #good #morning http://chapagain.com.np"
# print cleaned tweet
print(process_tweet(custom_tweet))
# -
# ## Part 1.1 Implementing your helper functions
#
# To help train your naive bayes model, you will need to build a dictionary where the keys are a (word, label) tuple and the values are the corresponding frequency. Note that the labels we'll use here are 1 for positive and 0 for negative.
#
# You will also implement a `lookup()` helper function that takes in the `freqs` dictionary, a word, and a label (1 or 0) and returns the number of times that word and label tuple appears in the collection of tweets.
#
# For example: given a list of tweets `["i am rather excited", "you are rather happy"]` and the label 1, the function will return a dictionary that contains the following key-value pairs:
#
# {
# ("rather", 1): 2
# ("happi", 1) : 1
# ("excit", 1) : 1
# }
#
# - Notice how for each word in the given string, the same label 1 is assigned to each word.
# - Notice how the words "i" and "am" are not saved, since it was removed by process_tweet because it is a stopword.
# - Notice how the word "rather" appears twice in the list of tweets, and so its count value is 2.
#
# #### Instructions
# Create a function `count_tweets()` that takes a list of tweets as input, cleans all of them, and returns a dictionary.
# - The key in the dictionary is a tuple containing the stemmed word and its class label, e.g. ("happi",1).
# - The value the number of times this word appears in the given collection of tweets (an integer).
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>Please use the `process_tweet` function that was imported above, and then store the words in their respective dictionaries and sets.</li>
# <li>You may find it useful to use the `zip` function to match each element in `tweets` with each element in `ys`.</li>
# <li>Remember to check if the key in the dictionary exists before adding that key to the dictionary, or incrementing its value.</li>
# <li>Assume that the `result` dictionary that is input will contain clean key-value pairs (you can assume that the values will be integers that can be incremented). It is good practice to check the datatype before incrementing the value, but it's not required here.</li>
# </ul>
# </p>
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def count_tweets(result, tweets, ys):
'''
Input:
result: a dictionary that will be used to map each pair to its frequency
tweets: a list of tweets
ys: a list corresponding to the sentiment of each tweet (either 0 or 1)
Output:
result: a dictionary mapping each pair to its frequency
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
for y, tweet in zip(ys, tweets):
for word in process_tweet(tweet):
# define the key, which is the word and label tuple
pair = (word, y)
# if the key exists in the dictionary, increment the count
if pair in result:
result[pair] += 1
# else, if the key is new, add it to the dictionary and set the count to 1
else:
result[pair] = 1
### END CODE HERE ###
return result
# +
# Testing your function
result = {}
tweets = ['i am happy', 'i am tricked', 'i am sad', 'i am tired', 'i am tired']
ys = [1, 0, 0, 0, 0]
count_tweets(result, tweets, ys)
# -
# **Expected Output**: {('happi', 1): 1, ('trick', 0): 1, ('sad', 0): 1, ('tire', 0): 2}
# # Part 2: Train your model using Naive Bayes
#
# Naive bayes is an algorithm that could be used for sentiment analysis. It takes a short time to train and also has a short prediction time.
#
# #### So how do you train a Naive Bayes classifier?
# - The first part of training a naive bayes classifier is to identify the number of classes that you have.
# - You will create a probability for each class.
# $P(D_{pos})$ is the probability that the document is positive.
# $P(D_{neg})$ is the probability that the document is negative.
# Use the formulas as follows and store the values in a dictionary:
#
# $$P(D_{pos}) = \frac{D_{pos}}{D}\tag{1}$$
#
# $$P(D_{neg}) = \frac{D_{neg}}{D}\tag{2}$$
#
# Where $D$ is the total number of documents, or tweets in this case, $D_{pos}$ is the total number of positive tweets and $D_{neg}$ is the total number of negative tweets.
# #### Prior and Logprior
#
# The prior probability represents the underlying probability in the target population that a tweet is positive versus negative. In other words, if we had no specific information and blindly picked a tweet out of the population set, what is the probability that it will be positive versus that it will be negative? That is the "prior".
#
# The prior is the ratio of the probabilities $\frac{P(D_{pos})}{P(D_{neg})}$.
# We can take the log of the prior to rescale it, and we'll call this the logprior
#
# $$\text{logprior} = log \left( \frac{P(D_{pos})}{P(D_{neg})} \right) = log \left( \frac{D_{pos}}{D_{neg}} \right)$$.
#
# Note that $log(\frac{A}{B})$ is the same as $log(A) - log(B)$. So the logprior can also be calculated as the difference between two logs:
#
# $$\text{logprior} = \log (P(D_{pos})) - \log (P(D_{neg})) = \log (D_{pos}) - \log (D_{neg})\tag{3}$$
# #### Positive and Negative Probability of a Word
# To compute the positive probability and the negative probability for a specific word in the vocabulary, we'll use the following inputs:
#
# - $freq_{pos}$ and $freq_{neg}$ are the frequencies of that specific word in the positive or negative class. In other words, the positive frequency of a word is the number of times the word is counted with the label of 1.
# - $N_{pos}$ and $N_{neg}$ are the total number of positive and negative words for all documents (for all tweets), respectively.
# - $V$ is the number of unique words in the entire set of documents, for all classes, whether positive or negative.
#
# We'll use these to compute the positive and negative probability for a specific word using this formula:
#
# $$ P(W_{pos}) = \frac{freq_{pos} + 1}{N_{pos} + V}\tag{4} $$
# $$ P(W_{neg}) = \frac{freq_{neg} + 1}{N_{neg} + V}\tag{5} $$
#
# Notice that we add the "+1" in the numerator for additive smoothing. This [wiki article](https://en.wikipedia.org/wiki/Additive_smoothing) explains more about additive smoothing.
# #### Log likelihood
# To compute the loglikelihood of that very same word, we can implement the following equations:
#
# $$\text{loglikelihood} = \log \left(\frac{P(W_{pos})}{P(W_{neg})} \right)\tag{6}$$
# ##### Create `freqs` dictionary
# - Given your `count_tweets()` function, you can compute a dictionary called `freqs` that contains all the frequencies.
# - In this `freqs` dictionary, the key is the tuple (word, label)
# - The value is the number of times it has appeared.
#
# We will use this dictionary in several parts of this assignment.
# +
# Build the freqs dictionary for later uses
freqs = count_tweets({}, train_x, train_y)
# -
# #### Instructions
# Given a freqs dictionary, `train_x` (a list of tweets) and a `train_y` (a list of labels for each tweet), implement a naive bayes classifier.
#
# ##### Calculate $V$
# - You can then compute the number of unique words that appear in the `freqs` dictionary to get your $V$ (you can use the `set` function).
#
# ##### Calculate $freq_{pos}$ and $freq_{neg}$
# - Using your `freqs` dictionary, you can compute the positive and negative frequency of each word $freq_{pos}$ and $freq_{neg}$.
#
# ##### Calculate $N_{pos}$, $N_{neg}$, $V_{pos}$, and $V_{neg}$
# - Using `freqs` dictionary, you can also compute the total number of positive words and total number of negative words $N_{pos}$ and $N_{neg}$.
# - Similarly, use `freqs` dictionary to compute the total number of **unique** positive words, $V_{pos}$, and total **unique** negative words $V_{neg}$.
#
# ##### Calculate $D$, $D_{pos}$, $D_{neg}$
# - Using the `train_y` input list of labels, calculate the number of documents (tweets) $D$, as well as the number of positive documents (tweets) $D_{pos}$ and number of negative documents (tweets) $D_{neg}$.
# - Calculate the probability that a document (tweet) is positive $P(D_{pos})$, and the probability that a document (tweet) is negative $P(D_{neg})$
#
# ##### Calculate the logprior
# - the logprior is $log(D_{pos}) - log(D_{neg})$
#
# ##### Calculate log likelihood
# - Finally, you can iterate over each word in the vocabulary, use your `lookup` function to get the positive frequencies, $freq_{pos}$, and the negative frequencies, $freq_{neg}$, for that specific word.
# - Compute the positive probability of each word $P(W_{pos})$, negative probability of each word $P(W_{neg})$ using equations 4 & 5.
#
# $$ P(W_{pos}) = \frac{freq_{pos} + 1}{N_{pos} + V}\tag{4} $$
# $$ P(W_{neg}) = \frac{freq_{neg} + 1}{N_{neg} + V}\tag{5} $$
#
# **Note:** We'll use a dictionary to store the log likelihoods for each word. The key is the word, the value is the log likelihood of that word).
#
# - You can then compute the loglikelihood: $log \left( \frac{P(W_{pos})}{P(W_{neg})} \right)$.
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def train_naive_bayes(freqs, train_x, train_y):
'''
Input:
freqs: dictionary from (word, label) to how often the word appears
train_x: a list of tweets
train_y: a list of labels correponding to the tweets (0,1)
Output:
logprior: the log prior. (equation 3 above)
loglikelihood: the log likelihood of you Naive bayes equation. (equation 6 above)
'''
loglikelihood = {}
logprior = 0
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# calculate V, the number of unique words in the vocabulary
vocab = set([pair[0] for pair in freqs.keys()])
V = len(vocab)
# calculate N_pos, N_neg, V_pos, V_neg
N_pos = N_neg = V_pos = V_neg = 0
for pair in freqs.keys():
# if the label is positive (greater than zero)
if pair[1] > 0:
# increment the count of unique positive words by 1
V_pos += 1
# Increment the number of positive words by the count for this (word, label) pair
N_pos += freqs[pair]
# else, the label is negative
else:
# increment the count of unique negative words by 1
V_neg += 1
# increment the number of negative words by the count for this (word,label) pair
N_neg += freqs[pair]
# Calculate D, the number of documents
D = len(train_x)
# Calculate D_pos, the number of positive documents
D_pos = sum(train_y==1) / D
# Calculate D_neg, the number of negative documents
D_neg = sum(train_y==0) / D
# Calculate logprior
logprior = np.log(D_pos)-np.log(D_neg)
# For each word in the vocabulary...
for word in vocab:
freq_pos = lookup(freqs,word,1)
freq_neg = lookup(freqs,word,0)
# calculate the probability that each word is positive, and negative
p_w_pos = (freq_pos+1)/(N_pos+V)
p_w_neg = (freq_neg+1)/(N_neg+V)
# calculate the log likelihood of the word
loglikelihood[word] = np.log(p_w_pos/p_w_neg)
### END CODE HERE ###
print(D_neg)
print(D)
return logprior, loglikelihood
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# You do not have to input any code in this cell, but it is relevant to grading, so please do not change anything
logprior, loglikelihood = train_naive_bayes(freqs, train_x, train_y)
print(logprior)
print(len(loglikelihood))
# **Expected Output**:
#
# 0.0
#
# 9089
# # Part 3: Test your naive bayes
#
# Now that we have the `logprior` and `loglikelihood`, we can test the naive bayes function by making predicting on some tweets!
#
# #### Implement `naive_bayes_predict`
# **Instructions**:
# Implement the `naive_bayes_predict` function to make predictions on tweets.
# * The function takes in the `tweet`, `logprior`, `loglikelihood`.
# * It returns the probability that the tweet belongs to the positive or negative class.
# * For each tweet, sum up loglikelihoods of each word in the tweet.
# * Also add the logprior to this sum to get the predicted sentiment of that tweet.
#
# $$ p = logprior + \sum_i^N (loglikelihood_i)$$
#
# #### Note
# Note we calculate the prior from the training data, and that the training data is evenly split between positive and negative labels (4000 positive and 4000 negative tweets). This means that the ratio of positive to negative 1, and the logprior is 0.
#
# The value of 0.0 means that when we add the logprior to the log likelihood, we're just adding zero to the log likelihood. However, please remember to include the logprior, because whenever the data is not perfectly balanced, the logprior will be a non-zero value.
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def naive_bayes_predict(tweet, logprior, loglikelihood):
'''
Input:
tweet: a string
logprior: a number
loglikelihood: a dictionary of words mapping to numbers
Output:
p: the sum of all the logliklihoods of each word in the tweet (if found in the dictionary) + logprior (a number)
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# process the tweet to get a list of words
word_l = process_tweet(tweet)
# initialize probability to zero
p = 0
# add the logprior
p += logprior
for word in word_l:
# check if the word exists in the loglikelihood dictionary
if word in loglikelihood:
# add the log likelihood of that word to the probability
p += loglikelihood[word]
### END CODE HERE ###
return p
# +
# UNQ_C5 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# You do not have to input any code in this cell, but it is relevant to grading, so please do not change anything
# Experiment with your own tweet.
my_tweet = '<NAME>.'
p = naive_bayes_predict(my_tweet, logprior, loglikelihood)
print('The expected output is', p)
# -
# **Expected Output**:
# - The expected output is around 1.57
# - The sentiment is positive.
# #### Implement test_naive_bayes
# **Instructions**:
# * Implement `test_naive_bayes` to check the accuracy of your predictions.
# * The function takes in your `test_x`, `test_y`, log_prior, and loglikelihood
# * It returns the accuracy of your model.
# * First, use `naive_bayes_predict` function to make predictions for each tweet in text_x.
# UNQ_C6 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def test_naive_bayes(test_x, test_y, logprior, loglikelihood):
"""
Input:
test_x: A list of tweets
test_y: the corresponding labels for the list of tweets
logprior: the logprior
loglikelihood: a dictionary with the loglikelihoods for each word
Output:
accuracy: (# of tweets classified correctly)/(total # of tweets)
"""
accuracy = 0 # return this properly
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
y_hats = []
for tweet in test_x:
# if the prediction is > 0
if naive_bayes_predict(tweet, logprior, loglikelihood) > 0:
# the predicted class is 1
y_hat_i = 1
else:
# otherwise the predicted class is 0
y_hat_i = 0
# append the predicted class to the list y_hats
y_hats.append(y_hat_i)
# error is the average of the absolute values of the differences between y_hats and test_y
error = np.mean(np.abs(y_hats-test_y))
# Accuracy is 1 minus the error
accuracy = 1 - error
### END CODE HERE ###
return accuracy
print("Naive Bayes accuracy = %0.4f" %
(test_naive_bayes(test_x, test_y, logprior, loglikelihood)))
# **Expected Accuracy**:
#
# 0.9940
# +
# UNQ_C7 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# You do not have to input any code in this cell, but it is relevant to grading, so please do not change anything
# Run this cell to test your function
for tweet in ['I am happy', 'I am bad', 'this movie should have been great.', 'great', 'great great', 'great great great', 'great great great great']:
# print( '%s -> %f' % (tweet, naive_bayes_predict(tweet, logprior, loglikelihood)))
p = naive_bayes_predict(tweet, logprior, loglikelihood)
# print(f'{tweet} -> {p:.2f} ({p_category})')
print(f'{tweet} -> {p:.2f}')
# -
# **Expected Output**:
# - I am happy -> 2.15
# - I am bad -> -1.29
# - this movie should have been great. -> 2.14
# - great -> 2.14
# - great great -> 4.28
# - great great great -> 6.41
# - great great great great -> 8.55
# Feel free to check the sentiment of your own tweet below
my_tweet = 'Finally at Part 4. Hurray!'
naive_bayes_predict(my_tweet, logprior, loglikelihood)
# # Part 4: Filter words by Ratio of positive to negative counts
#
# - Some words have more positive counts than others, and can be considered "more positive". Likewise, some words can be considered more negative than others.
# - One way for us to define the level of positiveness or negativeness, without calculating the log likelihood, is to compare the positive to negative frequency of the word.
# - Note that we can also use the log likelihood calculations to compare relative positivity or negativity of words.
# - We can calculate the ratio of positive to negative frequencies of a word.
# - Once we're able to calculate these ratios, we can also filter a subset of words that have a minimum ratio of positivity / negativity or higher.
# - Similarly, we can also filter a subset of words that have a maximum ratio of positivity / negativity or lower (words that are at least as negative, or even more negative than a given threshold).
#
# #### Implement `get_ratio()`
# - Given the `freqs` dictionary of words and a particular word, use `lookup(freqs,word,1)` to get the positive count of the word.
# - Similarly, use the `lookup()` function to get the negative count of that word.
# - Calculate the ratio of positive divided by negative counts
#
# $$ ratio = \frac{\text{pos_words} + 1}{\text{neg_words} + 1} $$
#
# Where pos_words and neg_words correspond to the frequency of the words in their respective classes.
# <table>
# <tr>
# <td>
# <b>Words</b>
# </td>
# <td>
# Positive word count
# </td>
# <td>
# Negative Word Count
# </td>
# </tr>
# <tr>
# <td>
# glad
# </td>
# <td>
# 41
# </td>
# <td>
# 2
# </td>
# </tr>
# <tr>
# <td>
# arriv
# </td>
# <td>
# 57
# </td>
# <td>
# 4
# </td>
# </tr>
# <tr>
# <td>
# :(
# </td>
# <td>
# 1
# </td>
# <td>
# 3663
# </td>
# </tr>
# <tr>
# <td>
# :-(
# </td>
# <td>
# 0
# </td>
# <td>
# 378
# </td>
# </tr>
# </table>
# UNQ_C8 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_ratio(freqs, word):
'''
Input:
freqs: dictionary containing the words
word: string to lookup
Output: a dictionary with keys 'positive', 'negative', and 'ratio'.
Example: {'positive': 10, 'negative': 20, 'ratio': 0.5}
'''
pos_neg_ratio = {'positive': 0, 'negative': 0, 'ratio': 0.0}
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
# use lookup() to find positive counts for the word (denoted by the integer 1)
pos_neg_ratio['positive'] = lookup(freqs,word,1)
# use lookup() to find negative counts for the word (denoted by integer 0)
pos_neg_ratio['negative'] = lookup(freqs,word,0)
# calculate the ratio of positive to negative counts for the word
pos_neg_ratio['ratio'] = (pos_neg_ratio['positive'] + 1) / (pos_neg_ratio['negative'] + 1)
### END CODE HERE ###
return pos_neg_ratio
get_ratio(freqs, 'happi')
# #### Implement `get_words_by_threshold(freqs,label,threshold)`
#
# * If we set the label to 1, then we'll look for all words whose threshold of positive/negative is at least as high as that threshold, or higher.
# * If we set the label to 0, then we'll look for all words whose threshold of positive/negative is at most as low as the given threshold, or lower.
# * Use the `get_ratio()` function to get a dictionary containing the positive count, negative count, and the ratio of positive to negative counts.
# * Append a dictionary to a list, where the key is the word, and the dictionary is the dictionary `pos_neg_ratio` that is returned by the `get_ratio()` function.
# An example key-value pair would have this structure:
# ```
# {'happi':
# {'positive': 10, 'negative': 20, 'ratio': 0.5}
# }
# ```
# +
# UNQ_C9 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def get_words_by_threshold(freqs, label, threshold):
'''
Input:
freqs: dictionary of words
label: 1 for positive, 0 for negative
threshold: ratio that will be used as the cutoff for including a word in the returned dictionary
Output:
word_set: dictionary containing the word and information on its positive count, negative count, and ratio of positive to negative counts.
example of a key value pair:
{'happi':
{'positive': 10, 'negative': 20, 'ratio': 0.5}
}
'''
word_list = {}
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
for key in freqs.keys():
word, _ = key
# get the positive/negative ratio for a word
pos_neg_ratio = get_ratio(freqs,word)
# if the label is 1 and the ratio is greater than or equal to the threshold...
if label == 1 and pos_neg_ratio['ratio'] >= threshold:
# Add the pos_neg_ratio to the dictionary
word_list[word] = pos_neg_ratio
# If the label is 0 and the pos_neg_ratio is less than or equal to the threshold...
elif label == 0 and pos_neg_ratio['ratio'] <= threshold:
# Add the pos_neg_ratio to the dictionary
word_list[word] = pos_neg_ratio
# otherwise, do not include this word in the list (do nothing)
else:
pass
### END CODE HERE ###
return word_list
# print(key)
# print(pos_neg_ratio)
# -
# Test your function: find negative words at or below a threshold
get_words_by_threshold(freqs, label=0, threshold=0.05)
# Test your function; find positive words at or above a threshold
get_words_by_threshold(freqs, label=1, threshold=10)
# pos_neg_ratio['ratio']
p = get_ratio(freqs, 'hahaha')
p['ratio']
# Notice the difference between the positive and negative ratios. Emojis like :( and words like 'me' tend to have a negative connotation. Other words like 'glad', 'community', and 'arrives' tend to be found in the positive tweets.
# # Part 5: Error Analysis
#
# In this part you will see some tweets that your model missclassified. Why do you think the misclassifications happened? Were there any assumptions made by the naive bayes model?
# Some error analysis done for you
print('Truth Predicted Tweet')
for x, y in zip(test_x, test_y):
y_hat = naive_bayes_predict(x, logprior, loglikelihood)
if y != (np.sign(y_hat) > 0):
print('%d\t%0.2f\t%s' % (y, np.sign(y_hat) > 0, ' '.join(
process_tweet(x)).encode('ascii', 'ignore')))
# # Part 6: Predict with your own tweet
#
# In this part you can predict the sentiment of your own tweet.
# +
# Test with your own tweet - feel free to modify `my_tweet`
my_tweet = 'I am happy because I am learning :)'
p = naive_bayes_predict(my_tweet, logprior, loglikelihood)
print(p)
# -
# Congratulations on completing this assignment. See you next week!
| NLP/Natural Language Processing with Classification and Vector Spaces/2/bayes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="gjwDhSG_89hb"
# # Text Extraction with BERT
# + [markdown] id="x18sQgfwNpsm"
# Resource: [Text Extraction with Bert](https://keras.io/examples/nlp/text_extraction_with_bert/)
# + colab={"base_uri": "https://localhost:8080/"} id="3J74DSyH9VzL" executionInfo={"status": "ok", "timestamp": 1622729839951, "user_tz": -180, "elapsed": 19357, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="b8eed164-932d-477e-99bd-96df8ea902df"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="ROpJgf1l9Eav" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1622729855106, "user_tz": -180, "elapsed": 11803, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="25e2b142-1465-4a11-919e-f19237d0117a"
# ! pip install transformers
# ! pip install tokenizers
# + id="Lnl0Y7z989hk" executionInfo={"status": "ok", "timestamp": 1622729860039, "user_tz": -180, "elapsed": 4937, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
import json
import os
import random
import numpy as np
import collections
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from transformers import BertTokenizer, TFBertModel, BertConfig
from tokenizers import BertWordPieceTokenizer
# + id="VHOU9WH289hm" executionInfo={"status": "ok", "timestamp": 1622729860041, "user_tz": -180, "elapsed": 16, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
class WikiElement:
def __init__(self, question, context, answer, answer_start, answer_end):
self.question = question
self.context = context
self.answer = answer
self.answer_start = answer_start
self.answer_end = answer_end
def preprocess(self):
# create context vector with answers marked
context_vector = [0] * len(self.context)
for index in range(self.answer_start, self.answer_end):
context_vector[index] = 1
# tokenize context
tokenized_context = tokenizer.encode(self.context)
# find answer token indices
answer_token_index = []
for index, (start, end) in enumerate(tokenized_context.offsets):
if sum(context_vector[start:end]) > 0: # if token is answer
answer_token_index.append(index)
if len(answer_token_index) == 0:
return 0
# start and end token index
start_token_index = answer_token_index[0]
end_token_index = answer_token_index[-1]
# tokenize question
tokenized_question = tokenizer.encode(self.question)
# create inputs
input_ids = tokenized_context.ids + tokenized_question.ids[1:]
token_type_ids = [0] * len(tokenized_context.ids) + [1] * len(tokenized_question.ids[1:])
attention_mask = [1] * len(input_ids)
# padding for equal lenght sequence
padding_length = max_len - len(input_ids)
if padding_length > 0: # pad
input_ids = input_ids + ([0] * padding_length)
attention_mask = attention_mask + ([0] * padding_length) # len(input) [1] + padding [0]
token_type_ids = token_type_ids + ([0] * padding_length) # context [0] + question [1] + padding [0]
elif padding_length < 0:
return 0
self.input_ids = input_ids
self.token_type_ids = token_type_ids
self.attention_mask = attention_mask
self.start_token_index = start_token_index
self.end_token_index = end_token_index
self.context_token_to_char = tokenized_context.offsets
return 1
def class_print(self):
print("Question: {}\nAnswer: {}\nAnswer Start: {}\nAnswer End: {}\nContext: {}".format(self.question,
self.answer,
self.answer_start,
self.answer_end,
self.context))
# + id="-MpTho0t89hn" executionInfo={"status": "ok", "timestamp": 1622729891530, "user_tz": -180, "elapsed": 189, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
def read_json(file_name):
with open(file_name, "r", encoding="utf-8") as json_file:
data = json.load(json_file)
return data
def json_to_list(json_dataset):
dataset = []
for paragraph_element in json_dataset["data"]:
for question_element in paragraph_element["qas"]:
dataset.append(WikiElement(question_element["question"],
paragraph_element["text"],
question_element["answer"],
question_element["answer_start"],
question_element["answer_end"]))
print("Number of questions: ", len(dataset))
return dataset
def create_input_targets(dataset):
dataset_dict = {
"input_ids": [],
"token_type_ids": [],
"attention_mask": [],
"start_token_index": [],
"end_token_index": [],
}
i=0
for item in dataset:
# print(i)
i = i + 1
# print(item.class_print())
for key in dataset_dict:
dataset_dict[key].append(getattr(item, key))
for key in dataset_dict:
dataset_dict[key] = np.array(dataset_dict[key])
x = [
dataset_dict["input_ids"],
dataset_dict["token_type_ids"],
dataset_dict["attention_mask"],
]
y = [dataset_dict["start_token_index"], dataset_dict["end_token_index"]]
return x, y
def find_max_length(dataset):
max_ = 0
index = 0
i = 0
for element in dataset:
tokenized_question = tokenizer.encode(element.question)
tokenized_context = tokenizer.encode(element.context)
input_ids = tokenized_context.ids + tokenized_question.ids[1:]
if len(input_ids) > max_:
max_ = len(input_ids)
index = i
i += 1
print("Max length: {}, Index: {}".format(max_, index))
return max_
def train_test_split(dataset):
random.shuffle(dataset)
cut = int(len(dataset)*0.1)
train, test = dataset[:-cut], dataset[-cut:]
return train, test
def create_model():
## BERT encoder
encoder = TFBertModel.from_pretrained(MODEL_NAME)
# QA model
input_ids = layers.Input(shape=(max_len,), dtype=tf.int32)
token_type_ids = layers.Input(shape=(max_len,), dtype=tf.int32)
attention_mask = layers.Input(shape=(max_len,), dtype=tf.int32)
embedding = encoder.bert(input_ids,
token_type_ids=token_type_ids,
attention_mask=attention_mask)[0]
start_logits = layers.Dense(1, name="start_logit", use_bias=False)(embedding)
start_logits = layers.Flatten()(start_logits)
end_logits = layers.Dense(1, name="end_logit", use_bias=False)(embedding)
end_logits = layers.Flatten()(end_logits)
start_probs = layers.Activation(keras.activations.softmax)(start_logits)
end_probs = layers.Activation(keras.activations.softmax)(end_logits)
model = keras.Model(
inputs=[input_ids, token_type_ids, attention_mask],
outputs=[start_probs, end_probs],
)
loss = keras.losses.SparseCategoricalCrossentropy(from_logits=False)
optimizer = keras.optimizers.Adam(lr=5e-5)
model.compile(optimizer=optimizer, loss=[loss, loss])
return model
# + id="AhJivsbAC1i7" executionInfo={"status": "ok", "timestamp": 1622729895037, "user_tz": -180, "elapsed": 217, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
import pickle
def save_data_as_file(data, file_name):
with open(path + file_name + ".dat", "wb") as f:
pickle.dump(data, f)
def read_saved_data(file_name):
with open(path + file_name + ".dat", "rb") as f:
data = pickle.load(f)
return data
# + [markdown] id="87wT60ih89hp"
#
# ## Load Tokenizer
# + id="nl9YM7Mb9x9L" executionInfo={"status": "ok", "timestamp": 1622729898535, "user_tz": -180, "elapsed": 192, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
path = "/content/gdrive/MyDrive/Q&A projesi/"
models_path = path + "models/"
MODEL_NAME = "dbmdz/bert-base-turkish-cased"
# + id="7JXP38Km89hp" colab={"base_uri": "https://localhost:8080/", "height": 114, "referenced_widgets": ["4e2f0f8926944d9bbf15e14e30c24ffc", "a91e513f935644d7af33f0a6c689c9a1", "<KEY>", "15fd2fface0c4a46ba776cce5066594f", "<KEY>", "c93da2d9ce184bd68557ceea9a45e3e7", "<KEY>", "<KEY>", "677dd16227b248b7a092d2ea75ff4abc", "edd5f4051bf64e7a893b50ba37401393", "560ed9f8e2d64a588bbdb366627cf7fe", "6e733372fb254e12a9f28285fd4c2a9e", "6a0e3b2e90a34a5597a393a22f975cb7", "<KEY>", "db0b4276b92a4c04af6a570d237fb6e9", "69d34ac12fdb45a28d074946b9e25375"]} executionInfo={"status": "ok", "timestamp": 1622729902829, "user_tz": -180, "elapsed": 2886, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="4e5fd59a-7c5d-4871-95d0-6710bb9b6e29"
slow_tokenizer = BertTokenizer.from_pretrained(MODEL_NAME, do_lower_case=False)
splitted_model = MODEL_NAME.split("/")
save_path = models_path + splitted_model[0] + "-" + splitted_model[1] + "/"
if not os.path.exists(save_path):
os.makedirs(save_path)
slow_tokenizer.save_pretrained(save_path)
tokenizer = BertWordPieceTokenizer(save_path + "vocab.txt", lowercase=False)
# + [markdown] id="atU_OyxE89hq"
# ## Load Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="e8KUmNG989hr" executionInfo={"status": "ok", "timestamp": 1622663238397, "user_tz": -180, "elapsed": 191, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="6d588d50-c9bd-4e4a-b1cb-0d5d1c1ebd97"
file_path = path + "json_dataset/Wiki_Dataset_Final.json"
json_dataset = read_json(file_path)
json_dataset["data"][144]["qas"][8]
# + colab={"base_uri": "https://localhost:8080/"} id="vSj-Qe9589hs" executionInfo={"status": "ok", "timestamp": 1622663239880, "user_tz": -180, "elapsed": 211, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="d9396f68-ee8f-41e6-cb8b-9ea3b157df24"
raw_dataset = json_to_list(json_dataset)
raw_dataset[0].class_print()
# + colab={"base_uri": "https://localhost:8080/"} id="Dc8ETKx-89ht" executionInfo={"status": "ok", "timestamp": 1622663251030, "user_tz": -180, "elapsed": 4508, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="5a938c16-b541-4971-cb62-09ba1f6ad94e"
# max_len = find_max_length(raw_dataset)
max_len = 384
dataset = []
for data in raw_dataset:
result = data.preprocess()
if result != 0:
dataset.append(data)
print("Dataset len: ", len(dataset))
# + id="rLxDSHcnD83G"
train, test = train_test_split(dataset)
# + id="Ex1nrjOLDFlq"
save_data_as_file(test, "test_384")
save_data_as_file(train, "train_384")
# + [markdown] id="DpM2dGf2PZat"
# ## Training
#
#
# + id="LD57dcMNPhDp" executionInfo={"status": "ok", "timestamp": 1622732108364, "user_tz": -180, "elapsed": 1971, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
max_len = 384
train = read_saved_data("train_" + str(max_len) + "_bert")
test = read_saved_data("test_" + str(max_len) + "_bert")
# + colab={"base_uri": "https://localhost:8080/"} id="RFqXr-R389hu" executionInfo={"status": "ok", "timestamp": 1622732110657, "user_tz": -180, "elapsed": 720, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="5ac2d19d-c651-499c-de59-ff7e1ac86de6"
x_train, y_train = create_input_targets(train)
x_test, y_test = create_input_targets(test)
print(len(x_train[0]), len(x_test[0]))
# + id="uti4Gpd989hv" executionInfo={"status": "ok", "timestamp": 1622732112914, "user_tz": -180, "elapsed": 238, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
configuration = BertConfig() # default parameters and configuration for BERT
# + colab={"base_uri": "https://localhost:8080/"} id="v4hArJC8-_u0" executionInfo={"status": "ok", "timestamp": 1622732640157, "user_tz": -180, "elapsed": 48430, "user": {"displayName": "<NAME>ehra \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="3663b6b3-aefa-42d9-8bd6-12ca6baace76"
use_tpu = True
if use_tpu:
# Create distribution strategy
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
# Create model
with strategy.scope():
model = create_model()
else:
model = create_model()
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="37Sl7nE989hw" executionInfo={"status": "ok", "timestamp": 1622733058076, "user_tz": -180, "elapsed": 55414, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="46c11e45-d91f-4ca0-aeb3-a9ed1bb92577"
# Load Weights from Drive
# model.load_weights(path + "models/bertV1_weights.h5")
model.fit(
x_train,
y_train,
epochs=5, # For demonstration, 3 epochs are recommended
verbose=2,
batch_size=64,
)
# + id="2kL_E5cv89hw" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1622733077004, "user_tz": -180, "elapsed": 1241, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="bf498a3d-e61a-41b4-9a41-5f38bafb6ac9"
pred_start, pred_end = model.predict(x_test)
count = 0
results = []
total_f1 = 0
for idx, (start, end) in enumerate(zip(pred_start, pred_end)):
element = test[idx]
offsets = element.context_token_to_char
start = np.argmax(start)
end = np.argmax(end)
if start >= len(offsets):
continue
pred_char_start = offsets[start][0]
if end < len(offsets):
pred_char_end = offsets[end][1]
pred_ans = element.context[pred_char_start:pred_char_end]
else:
pred_ans = element.context[pred_char_start:]
pred_tokens = pred_ans.split()
true_tokens = element.answer.split()
common = collections.Counter(true_tokens) & collections.Counter(pred_tokens)
num_same = sum(common.values())
if len(true_tokens) == 0 or len(pred_tokens) == 0:
# If either is no-answer, then F1 is 1 if they agree, 0 otherwise
f1 = int(true_tokens == pred_tokens)
elif num_same == 0:
f1 = 0
else:
precision = 1.0 * num_same / len(pred_tokens)
recall = 1.0 * num_same / len(true_tokens)
f1 = (2 * precision * recall) / (precision + recall)
total_f1 += f1
results.append({
"question": element.question,
"true answer": element.answer,
"predicted answer": pred_ans,
"context": element.context,
"f1 score": f1,
})
# print(f"Question: {element.question}")
# print(f"Prediction: {pred_ans}\nTrue Answer: {element.answer}")
# print(f"Context: {element.context}")
# print("\n")
if pred_ans == element.answer:
count += 1
acc = count / len(y_test[0])
F1 = total_f1 / len(y_test[0])
print(f"exact match:={acc:.2f} f1:={F1:.2f}")
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="LZTn_LuP_CfF" executionInfo={"status": "ok", "timestamp": 1618665582591, "user_tz": -180, "elapsed": 463, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}} outputId="32811ed8-b8a4-4311-c692-161e37f90671"
results[0]["question"]
# + id="xgNdtluN-Dg5" executionInfo={"status": "ok", "timestamp": 1622730500195, "user_tz": -180, "elapsed": 560, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
save_model_name = splitted_model[0] + "-" + splitted_model[1]
file_name = save_path + "test-results/" + save_model_name + "_10epochs_results.txt"
with open(file_name, "w") as f:
for result in results:
f.write('%s\n' %result)
# + [markdown] id="nxgo-YipwZ6f"
# ### Save Weights to Google Drive
# + id="U8xpx5lV2iYh" executionInfo={"status": "ok", "timestamp": 1622732085100, "user_tz": -180, "elapsed": 3652, "user": {"displayName": "<NAME> \u00c7etin", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjzWxwkSm6SQwLvKuI7eKiYtfg5Yy0_bJzt0tQigQ=s64", "userId": "14276538161093709229"}}
model.save_weights(save_path + "weights/" + save_model_name + "_seqlen512_epochs12"".h5")
| Text Extraction with BERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from requests_html import HTMLSession
import re
def get_single_record(records, photoId):
session = HTMLSession()
url = 'http://ucr.emuseum.com/view/objects/asitem/3631/' + str(photoId)
response = session.get(url, timeout = 20)
sel = '#singlemedia > div:nth-child(1) > a > img[src^="/internal/media/dispatcher/"]'
singlemedia = response.html.find(sel, first=True)
# src = singlemedia.('/internal/media/dispatcher/{}/resize:format=preview')
src = singlemedia.attrs['src']
index = re.search(r'\d+', src).group()
# indexToHtml[index] = url
# print(singlemedia)
print(index)
selData = '#singledata > div'
# sel2 = '#singledata > div'
# selLabels = '#singledata > div > span'
# selDatas = '#singledata > div'
data = response.html.find(selData)
# singledata = response.html.find(sel2)
# labels = response.html.find(selLabels)
# datas = response.html.find(selDatas)
info = {}
info['Webpage'] = url
for j in range(len(data)):
# print(data[j].text)
# print(data[j])
line = data[j].text.strip()
if (line != ''):
l = line.split(':', 1)[0].strip()
r = line.split(':', 1)[1].strip()
if (l == 'Subjects'):
r = r.split('\n')
# print(r)
# print(l + ":\n" + r + "\n")
info[l] = r
records[index] = info
# print(records[index])
records = {} # index -> data
# indexToHtml = {}
for i in range(0, 45197):
# for i in range(3):
print("---start %s ---" % i)
get_single_record(records, i)
print("Done!")
# get_single_record(records, 2)
# print(records)
# +
# import pandas as pd
# df = pd.DataFrame(records)
# # df = pd.DataFrame.from_dict(records, orient='index', columns=records['26348'].keys())
# df.to_csv('photo_infos_1370.csv', encoding='utf-8', index=True)
# df
# print(records['14400'])
# print(indexToHtml['86636'])
# +
from requests_html import HTMLSession
import re
from multiprocessing import Pool, Manager, Process
import pandas as pd
from functools import partial
import json
# def scrape(url):
# try:
# print requests.get(url)
# except ConnectionError:
# print 'Error Occured ', url
# finally:
# print 'URL ', url, ' Scraped'
# records = {} # index -> data
# count = 0
def get_single_record(records, photoId):
# print("---start %s ---" % photoId)
session = HTMLSession()
url = 'http://ucr.emuseum.com/view/objects/asitem/3631/' + str(photoId)
response = session.get(url, timeout=20)
sel = '#singlemedia > div:nth-child(1) > a > img[src^="/internal/media/dispatcher/"]'
singlemedia = response.html.find(sel, first=True)
# src = singlemedia.('/internal/media/dispatcher/{}/resize:format=preview')
src = singlemedia.attrs['src']
index = re.search(r'\d+', src).group()
# print(singlemedia)
print(index)
selData = '#singledata > div'
# sel2 = '#singledata > div'
# selLabels = '#singledata > div > span'
# selDatas = '#singledata > div'
data = response.html.find(selData)
# singledata = response.html.find(sel2)
# labels = response.html.find(selLabels)
# datas = response.html.find(selDatas)
info = {}
info['Webpage'] = url
for j in range(len(data)):
# print(data[j].text)
line = data[j].text.strip()
if (line != ''):
l = line.split(':', 1)[0].strip()
r = line.split(':', 1)[1].strip()
if (l == 'Subjects'):
r = r.split('\n')
# print(r)
# print(l + ":\n" + r + "\n")
info[l] = r
records[index] = info
# print(records[index])
# ff = open('photo-infos/'+ str(index) + '.txt','w+',encoding='utf-8')
# ff.write(str(records))
print(str(photoId) + "(" + str(index) + ")" + "Done!")
# print(str(++count) + "Done!")
# if __name__ == '__main__':
manager = Manager()
records = manager.dict()
# l = manager.list(range(10))
pool = Pool(processes=4)
pool.starmap(get_single_record, [(records, i) for i in range(45197)])
pool.close()
pool.join()
# for i in range(4):
# p1 = Process(target=get_single_record, args=(records, l))
# p2 = Process(target=get_single_record, args=(records, l))
# p1.start()
# p2.start()
# p1.join
# p2.join
# print(records)
# df = pd.DataFrame(records)
# df.to_csv('photo_infos_1000.csv', encoding='utf-8', index=False)
# print("photo_infos_1000.csv created!")
with open('my_dict_script_10.json', 'w+') as f:
json.dump(records.copy(), f)
# records = {} # index -> data
# for i in range(45197):
# # for i in range(3):
# print("---start %s ---" % i)
# get_single_record(records, i)
# print("Done!")
# get_single_record(records, 2)
# print(records)
# +
# from multiprocessing import Pool
# def show_video_stats(options):
# pool = Pool(8)
# video_page_urls = get_video_page_urls()
# results = pool.map(get_video_data, video_page_urls)
import json
with open('my_dict_10.json', 'w+') as f:
json.dump(records, f)
# elsewhere...
# with open('my_dict.json') as f:
# my_dict = json.load(f)
| web-crawler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle as pkl
import numpy as np
import pandas as pd
from zipfile import ZipFile
from sklearn.metrics import accuracy_score, matthews_corrcoef, confusion_matrix
from sklearn.model_selection import train_test_split
from tensorflow.keras.backend import clear_session
from twitter_nlp_toolkit.tweet_sentiment_classifier import tweet_sentiment_classifier
from twitter_nlp_toolkit.file_fetcher import file_fetcher
# -
# Install Spacy library
# !python -m spacy download en_core_web_sm
clear_session()
chunk = 1 # Fraction of data to train on - you can reduce for debugging for speed
model_path = '.models'
redownload=True
# Here we download pre-trained models and a validation dataset. The models have been pre-trained on the Sentiment140 dataset, taken form here: https://www.kaggle.com/kazanova/sentiment140
#
# The validation data is hand-labeled airline customer feedback taken from https://www.figure-eight.com/data-for-everyone/
# + jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
if redownload:
# Validation data
file_fetcher.download_file('https://www.dropbox.com/s/muqov03s9vtbe92/tweets_airline.zip?dl=1',"tweets_airline.zip")
# +
# Load the validation data
test_data = pd.read_csv('tweets_airline.zip', header=0,
names=['Index', 'Sentiment', 'Sentiment_confidence',
'Negative_reason', 'Negative_reason_confidence',
'Airline', 'Airline_sentiment_gold', 'Handle',
'Negative_reason_gold', 'Retweet_count', 'Text',
'Tweet_coord', 'Time', 'Location', 'Timezone'])
# -
test_data.head(5)
# We drop the tweets with "neural" sentiment:
# +
# Remove the unlabeled test data
test_data['Labels'] = (test_data['Sentiment'] == 'positive') * 2
test_data['Labels'] = test_data['Labels'] + (test_data['Sentiment'] == 'neutral') * 1
test_data['Labels'] = test_data['Labels'] / 2
test_data.set_index('Labels')
test_data = test_data[test_data.Labels != 0.5]
# +
# Download the small ensemble
# Executing this cell starts a 500MB download
Classifier = tweet_sentiment_classifier.SentimentAnalyzer()
Classifier.load_small_ensemble()
# -
# We sanity check the models:
Classifier.predict(['I am happy', 'I am sad', 'I am cheerful', 'I am mad'])
# We test the model on an airline customer feedback dataset.
# +
# Executing this cell takes several minuites on a laptop
predictions = Classifier.predict(test_data['Text'])
# -
print('Test Accuracy: {:.3f}'.format(accuracy_score(test_data['Labels'], predictions)))
print('Test MCC: {:.3f}'.format(matthews_corrcoef(test_data['Labels'], predictions)))
confusion_matrix(test_data['Labels'], predictions)
# We have accuracy of just over 80%.
#
# We split our evaluation dataset into validation and testing and check for poor-performing models:
valX, testX, valY, testY = train_test_split(test_data['Text'], test_data['Labels'], test_size=0.5, stratify=test_data['Labels'])
# +
# Executing this cell takes several minuites on a laptop
Classifier.trim_models(valX, valY, threshold=0.7)
# -
# All three models perform well, so none have been pruned
# +
predictions = Classifier.predict(testX)
print('Test Accuracy: {:.3f}'.format(accuracy_score(testY, predictions)))
print('Test MCC: {:.3f}'.format(matthews_corrcoef(testY, predictions)))
confusion_matrix(testY, predictions)
# -
# To improve our accuracy, we can refine the models on our airline data. The early stopping procedure (enabled by default to use 20% of the training data for validation) should minimize overfitting.
Classifier.refine(valX, valY)
# +
test_predictions = Classifier.predict(testX)
print('Test Accuracy: {:.3f}'.format(accuracy_score(testY, test_predictions)))
print('Test MCC: {:.3f}'.format(matthews_corrcoef(testY, test_predictions)))
confusion_matrix(testY, predictions)
# -
# Now we have accuracies of over 90%!
Classifier.evaluate(testX, testY)
| demo_sentiment_classifier_trainer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # The Discrete-Time Fourier Transform
#
# *This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the bachelors module Signals and Systems, Comunications Engineering, Universität Rostock. Please direct questions and suggestions to [<EMAIL>](mailto:<EMAIL>).*
# -
# ## Theorems
#
# The theorems of the discrete-time Fourier transform (DTFT) relate basic operations applied to discrete signals to their equivalents in the DTFT domain. They are of use to transform signals composed from modified [standard signals](../discrete_signals/standard_signals.ipynb), for the computation of the response of a linear time-invariant (LTI) system and to predict the consequences of modifying a signal or system by certain operations.
# ### Convolution Theorem
#
# The [convolution theorem](https://en.wikipedia.org/wiki/Convolution_theorem) states that the DTFT of the linear convolution of two discrete signals $x[k]$ and $y[k]$ is equal to the scalar multiplication of their DTFTs $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$ and $Y(e^{j \Omega}) = \mathcal{F}_* \{ y[k] \}$
#
# \begin{equation}
# \mathcal{F}_* \{ x[k] * y[k] \} = X(e^{j \Omega}) \cdot Y(e^{j \Omega})
# \end{equation}
#
# The theorem can be proven by introducing the [definition of the linear convolution](../discrete_systems/linear_convolution.ipynb) into the [definition of the DTFT](definition.ipynb) and changing the order of summation
#
# \begin{align}
# \mathcal{F} \{ x[k] * y[k] \} &= \sum_{k = -\infty}^{\infty} \left( \sum_{\kappa = -\infty}^{\infty} x[\kappa] \cdot y[k - \kappa] \right) e^{-j \Omega k} \\
# &= \sum_{\kappa = -\infty}^{\infty} \left( \sum_{k = -\infty}^{\infty} y[k - \kappa] \, e^{-j \Omega k} \right) x[\kappa] \\
# &= Y(e^{j \Omega}) \cdot \sum_{\kappa = -\infty}^{\infty} x[\kappa] \, e^{-j \Omega \kappa} \\
# &= Y(e^{j \Omega}) \cdot X(e^{j \Omega})
# \end{align}
#
# The convolution theorem is very useful in the context of LTI systems. The output signal $y[k]$ of an LTI system is given as the convolution of the input signal $x[k]$ with its impulse response $h[k]$. Hence, the signals and the system can be represented equivalently in the time and frequency domain
#
# 
#
# Calculation of the system response by transforming the problem into the DTFT domain can be beneficial since this replaces the computation of the linear convolution by a scalar multiplication. The (inverse) DTFT is known for many signals or can be derived by applying the properties and theorems to standard signals and their transforms. In many cases this procedure simplifies the calculation of the system response significantly.
#
# The convolution theorem can also be useful to derive the DTFT of a signal. The key is here to express the signal as convolution of two other signals for which the transforms are known. This is illustrated in the following example.
# #### Transformation of the trapezoidal and triangular signal
#
# The linear convolution of two [rectangular signals](../discrete_signals/standard_signals.ipynb#Rectangular-Signal) of lengths $N$ and $M$ defines a [signal of trapezoidal shape](../discrete_systems/linear_convolution.ipynb#Finite-Length-Signals)
#
# \begin{equation}
# x[k] = \text{rect}_N[k] * \text{rect}_M[k]
# \end{equation}
#
# Application of the convolution theorem together with the [DTFT of the rectangular signal](definition.ipynb#Transformation-of-the-Rectangular-Signal) yields its DTFT as
#
# \begin{equation}
# X(e^{j \Omega}) = \mathcal{F}_* \{ \text{rect}_N[k] \} \cdot \mathcal{F}_* \{ \text{rect}_M[k] \} =
# e^{-j \Omega \frac{N+M-2}{2}} \cdot \frac{\sin(\frac{N \Omega}{2}) \sin(\frac{M \Omega}{2})}{\sin^2 ( \frac{\Omega}{2} )}
# \end{equation}
#
# The transform of the triangular signal can be derived from this result. The convolution of two rectangular signals of equal length $N=M$ yields the triangular signal $\Lambda[k]$ of length $2N - 1$
#
# \begin{equation}
# \Lambda_{2N - 1}[k] = \begin{cases} k + 1 & \text{for } 0 \leq k < N \\
# 2N - 1 - k & \text{for } N \leq k < 2N - 1 \\
# 0 & \text{otherwise}
# \end{cases}
# \end{equation}
#
# From above result the DTFT of the triangular signal is derived by substitution of $N$ by $M$
#
# \begin{equation}
# \mathcal{F}_* \{ \Lambda_{2N - 1}[k] \} =
# e^{-j \Omega (N-1)} \cdot \frac{\sin^2(\frac{N \Omega}{2}) }{\sin^2 ( \frac{\Omega}{2} )}
# \end{equation}
#
# Both the triangular signal and the magnitude of its DTFT are plotted for illustration
# +
import sympy as sym
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
N = 7
x = np.convolve(np.ones(N), np.ones(N), mode='full')
plt.stem(x)
plt.xlabel('$k$')
plt.ylabel('$x[k]$')
W = sym.symbols('Omega')
X = sym.exp(-sym.I*W * (N-1)) * sym.sin(N*W/2)**2 / sym.sin(W/2)**2
sym.plot(sym.Abs(X), (W, -5, 5), xlabel='$\Omega$',
ylabel='$|X(e^{j \Omega})|$')
# -
# **Exercise**
#
# * Change the length of the triangular signal in above example. How does its DTFT change?
# * The triangular signal introduced above is of odd length $2N - 1$
# * Define a triangular signal of even length by convolving two rectangular signals
# * Derive its DTFT
# * Compare the DTFTs of a triangular signal of odd/even length
# ### Shift Theorem
#
# The [shift of a signal](../discrete_signals/operations.ipynb#Shift) $x[k]$ can be expressed by a convolution with a shifted Dirac impulse
#
# \begin{equation}
# x[k - \kappa] = x[k] * \delta[k - \kappa]
# \end{equation}
#
# for $\kappa \in \mathbb{Z}$. This follows from the sifting property of the Dirac impulse. Applying the DTFT to the left- and right-hand side and exploiting the convolution theorem yields
#
# \begin{equation}
# \mathcal{F}_* \{ x[k - \kappa] \} = X(e^{j \Omega}) \cdot e^{- j \Omega \kappa}
# \end{equation}
#
# where $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$. Note that $\mathcal{F}_* \{ \delta(k - \kappa) \} = e^{- j \Omega \kappa}$ can be derived from the definition of the DTFT together with the sifting property of the Dirac impulse. Above relation is known as shift theorem of the DTFT.
#
# Expressing the DTFT $X(e^{j \Omega}) = |X(e^{j \Omega})| \cdot e^{j \varphi(e^{j \Omega})}$ by its absolute value $|X(e^{j \Omega})|$ and phase $\varphi(e^{j \Omega})$ results in
#
# \begin{equation}
# \mathcal{F}_* \{ x[k - \kappa] \} = | X(e^{j \Omega}) | \cdot e^{j (\varphi(e^{j \Omega}) - \Omega \kappa)}
# \end{equation}
#
# Shifting of a signal does not change the absolute value of its spectrum but it subtracts the linear contribution $\Omega \kappa$ from its phase.
# ### Multiplication Theorem
#
# The transform of a multiplication of two signals $x[k] \cdot y[k]$ is derived by introducing the signals into the definition of the DTFT, expressing the signal $x[k]$ by its spectrum $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$ and rearranging terms
#
# \begin{align}
# \mathcal{F}_* \{ x[k] \cdot y[k] \} &= \sum_{k=-\infty}^{\infty} x[k] \cdot y[k] \, e^{-j \Omega k} \\
# &= \sum_{k=-\infty}^{\infty} \left( \frac{1}{2 \pi} \int_{-\pi}^{\pi} X(e^{j \nu}) \, e^{j \nu k} \; d \nu \right) y[k] \, e^{-j \Omega k} \\
# &= \frac{1}{2 \pi} \int_{-\pi}^{\pi} X(e^{j \nu}) \sum_{k=-\infty}^{\infty} y[k] \, e^{-j (\Omega - \nu) k} \; d\nu \\
# &= \frac{1}{2 \pi} \int_{-\pi}^{\pi} X(e^{j \nu}) \cdot Y(e^{j (\Omega - \nu)}) d\nu
# \end{align}
#
# where $Y(e^{j \Omega}) = \mathcal{F}_* \{ y[k] \}$.
#
# The [periodic (cyclic/circular) convolution](https://en.wikipedia.org/wiki/Circular_convolution) of two aperiodic signals $h(t)$ and $g(t)$ is defined as
#
# \begin{equation}
# h(t) \circledast_{T} g(t) = \int_{-\infty}^{\infty} h(\tau) \cdot g_\text{p}(t - \tau) \; d\tau
# \end{equation}
#
# where $T$ denotes the period of the convolution, $g_\text{p}(t) = \sum_{n=-\infty}^{\infty} g(t + n T)$ the periodic summation of $g(t)$ and $\tau \in \mathbb{R}$ an arbitrary constant. The periodic convolution is commonly abbreviated by $\circledast_{T}$. With $h_\text{p}(t)$ denoting the periodic summation of $h(t)$ the periodic convolution can be rewritten as
#
# \begin{equation}
# h(t) \circledast_{T} g(t) = \int_{\tau_0}^{\tau_0 + T} h_\text{p}(\tau) \cdot g_\text{p}(t - \tau) \; d\tau
# \end{equation}
#
# where $\tau_0 \in \mathbb{R}$ denotes an arbitrary constant. The latter definition holds also for two [periodic signals](../periodic_signals/spectrum.ipynb) $h(t)$ and $g(t)$ with period $T$.
#
# Comparison of the DTFT of two multiplied signals with the definition of the periodic convolution reveals that the preliminary result above can be expressed as
#
# \begin{equation}
# \mathcal{F}_* \{ x[k] \cdot y[k] \} = \frac{1}{2\pi} \, X(e^{j \Omega}) \circledast_{2 \pi} Y(e^{j \Omega})
# \end{equation}
#
# The DTFT of a multiplication of two signals $x[k] \cdot y[k]$ is given by the periodic convolution of their transforms $X(e^{j \Omega})$ and $Y(e^{j \Omega})$ weighted with $\frac{1}{2 \pi}$. The periodic convolution has a period of $T = 2 \pi$. Note, the convolution is performed with respect to the normalized angular frequency $\Omega$.
#
# Applications of the multiplication theorem include the modulation and windowing of signals. The former leads to the modulation theorem introduced later, the latter is illustrated by the following example.
# **Example**
#
# Windowing of signals is used to derive signals of finite duration from signals of infinite duration or to truncate signals to a shorter length. The signal $x[k]$ is multiplied by a weighting function $w[k]$ in order to derive the finite length signal
#
# \begin{equation}
# y[k] = w[k] \cdot x[k]
# \end{equation}
#
# Application of the multiplication theorem yields the spectrum $Y(e^{j \Omega}) = \mathcal{F}_* \{ y[k] \}$ of the windowed signal as
#
# \begin{equation}
# Y(e^{j \Omega}) = \frac{1}{2 \pi} W(e^{j \Omega}) \circledast X(e^{j \Omega})
# \end{equation}
#
# where $W(e^{j \Omega}) = \mathcal{F}_* \{ w[k] \}$ and $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$. In order to illustrate the consequence of windowing, a cosine signal $x[k] = \cos(\Omega_0 k)$ is truncated to a finite length using a rectangular signal
#
# \begin{equation}
# y[k] = \text{rect}_N[k] \cdot \cos(\Omega_0 k)
# \end{equation}
#
# where $N$ denotes the length of the truncated signal and $\Omega_0$ its normalized angular frequency. Using the DTFT of the [rectangular signal](definition.ipynb#Transformation-of-the-Rectangular-Signal) and the [cosine signal](properties.ipynb#Transformation-of-the-cosine-and-sine-signal) yields
#
# \begin{align}
# Y(e^{j \Omega}) &= \frac{1}{2 \pi} e^{-j \Omega \frac{N-1}{2}} \cdot \frac{\sin \left(\frac{N \Omega}{2} \right)}{\sin \left( \frac{\Omega}{2} \right)} \circledast \frac{1}{2} \left[ {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega + \Omega_0}{2 \pi} \right) + {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega - \Omega_0}{2 \pi} \right) \right] \\
# &= \frac{1}{2} \left[ e^{-j (\Omega+\Omega_0) \frac{N-1}{2}} \cdot \frac{\sin \left(\frac{N (\Omega+\Omega_0)}{2} \right)}{\sin \left( \frac{\Omega+\Omega_0}{2} \right)} + e^{-j (\Omega-\Omega_0) \frac{N-1}{2}} \cdot \frac{\sin \left(\frac{N (\Omega-\Omega_0)}{2} \right)}{\sin \left( \frac{\Omega-\Omega_0}{2} \right)} \right]
# \end{align}
#
# The latter identity results from the sifting property of the Dirac impulse and the periodicity of both spectra. The signal $y[k]$ and its magnitude spectrum $|Y(e^{j \Omega})|$ are plotted for specific values of $N$ and $\Omega_0$.
# +
N = 20
W0 = 2*np.pi/10
k = np.arange(N)
x = np.cos(W0 * k)
plt.stem(k, x)
plt.xlabel('$k$')
plt.ylabel('$y[k]$')
# +
W = sym.symbols('Omega')
Y = 1/2 * ((sym.exp(-sym.I*(W+W0)*(N-1)/2) * sym.sin(N*(W+W0)/2) / sym.sin((W+W0)/2)) +
(sym.exp(-sym.I*(W-W0)*(N-1)/2) * sym.sin(N*(W-W0)/2) / sym.sin((W-W0)/2)))
sym.plot(sym.Abs(Y), (W, -sym.pi, sym.pi),
xlabel='$\Omega$', ylabel='$|Y(e^{j \Omega})|$')
# -
# **Exercise**
#
# * Change the length $N$ of the signal by modifying the example. How does the spectrum change if you decrease or increase the length?
#
# * What happens if you change the normalized angular frequency $\Omega_0$ of the signal?
#
# * Assume a signal that is composed from a superposition of two finite length cosine signals with different frequencies. What qualitative condition has to hold that you can derive these frequencies from inspection of the spectrum?
# ### Modulation Theorem
#
# The complex modulation of a signal $x[k]$ is defined as $e^{j \Omega_0 k} \cdot x[k]$ with $\Omega_0 \in \mathbb{R}$. The DTFT of the modulated signal is derived by applying the multiplication theorem
#
# \begin{equation}
# \mathcal{F}_* \left\{ e^{j \Omega_0 k} \cdot x[k] \right\} = \frac{1}{2 \pi} \cdot {\bot \!\! \bot \!\! \bot}\left( \frac{\Omega - \Omega_0}{2 \pi} \right) \circledast X(e^{j \Omega})
# = X \big( e^{j \, (\Omega - \Omega_0)} \big)
# \end{equation}
#
# where $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$. Above result states that the complex modulation of a signal leads to a shift of its spectrum. This result is known as modulation theorem.
# **Example**
#
# An example for the application of the modulation theorem is the
# [downsampling/decimation](https://en.wikipedia.org/wiki/Decimation_(signal_processing)) of a discrete signal $x[k]$. Downsampling refers to lowering the sampling rate of a signal. The example focuses on the special case of removing every second sample, hence halving the sampling rate. The downsampling is modeled by defining a signal $x_\frac{1}{2}[k]$ where every second sample is set to zero
#
# \begin{equation}
# x_\frac{1}{2}[k] = \begin{cases}
# x[k] & \text{for even } k \\
# 0 & \text{for odd } k
# \end{cases}
# \end{equation}
#
# In order to derive the spectrum $X_\frac{1}{2}(e^{j \Omega}) = \mathcal{F}_* \{ x_\frac{1}{2}[k] \}$, the signal $u[k]$ is introduced where every second sample is zero
#
# \begin{equation}
# u[k] = \frac{1}{2} ( 1 + e^{j \pi k} ) = \begin{cases} 1 & \text{for even } k \\
# 0 & \text{for odd } k \end{cases}
# \end{equation}
#
# Using $u[k]$, the process of setting every second sample of $x[k]$ to zero can be expressed as
#
# \begin{equation}
# x_\frac{1}{2}[k] = u[k] \cdot x[k]
# \end{equation}
#
# Now the spectrum $X_\frac{1}{2}(e^{j \Omega})$ is derived by applying the multiplication theorem and introducing the [DTFT of the exponential signal](definition.ipynb#Transformation-of-the-Exponential-Signal). This results in
#
# \begin{equation}
# X_\frac{1}{2}(e^{j \Omega}) = \frac{1}{4 \pi} \left( {\bot \!\! \bot \!\! \bot}\left( \frac{\Omega}{2 \pi} \right) +
# {\bot \!\! \bot \!\! \bot}\left( \frac{\Omega - \pi}{2 \pi} \right) \right) \circledast X(e^{j \Omega}) =
# \frac{1}{2} X(e^{j \Omega}) + \frac{1}{2} X(e^{j (\Omega- \pi)})
# \end{equation}
#
# where $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$. The spectrum $X_\frac{1}{2}(e^{j \Omega})$ consists of the spectrum of the original signal $X(e^{j \Omega})$ superimposed by the shifted spectrum $X(e^{j (\Omega- \pi)})$ of the original signal. This may lead to overlaps that constitute aliasing. In order to avoid aliasing, the spectrum of the signal $x[k]$ has to be band-limited to $-\frac{\pi}{2} < \Omega < \frac{\pi}{2}$ before downsampling.
# ### Parseval's Theorem
#
# [Parseval's theorem](https://en.wikipedia.org/wiki/Parseval's_theorem) relates the energy of a discrete signal to its spectrum. The squared absolute value of a signal $x[k]$ represents its instantaneous power. It can be expressed as
#
# \begin{equation}
# | x[k] |^2 = x[k] \cdot x^*[k]
# \end{equation}
#
# where $x^*[k]$ denotes the complex conjugate of $x[k]$. Transformation of the right-hand side and application of the multiplication theorem results in
#
# \begin{equation}
# \mathcal{F}_* \{ x[k] \cdot x^*[k] \} = \frac{1}{2 \pi} \cdot X(e^{j \Omega}) \circledast_{2 \pi} X^*(e^{-j \Omega})
# \end{equation}
#
# Introducing the definition of the DTFT and the periodic convolution
#
# \begin{equation}
# \sum_{k = -\infty}^{\infty} x[k] \cdot x^*[k] \, e^{-j \Omega k} =
# \frac{1}{2 \pi} \int_{-\pi}^{\pi} X(e^{j \nu}) \cdot X^*(e^{j (\Omega - \nu)}) \; d\nu
# \end{equation}
#
# Setting $\Omega = 0$ followed by the substitution $\nu = \Omega$ yields Parseval's theorem
#
# \begin{equation}
# \sum_{k = -\infty}^{\infty} | x[k] |^2 = \frac{1}{2 \pi} \int_{-\pi}^{\pi} | X(e^{j \Omega}) |^2 \; d\Omega
# \end{equation}
#
# The sum over the samples of the squared absolute signal is equal to the integral over its squared absolute spectrum divided by $2 \pi$. Since the left-hand side represents the energy $E$ of the signal $x[k]$, Parseval's theorem states that the energy can be computed alternatively in the spectral domain by integrating over the squared absolute value of the spectrum.
# + [markdown] nbsphinx="hidden"
# **Copyright**
#
# This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Sascha Spors, Continuous- and Discrete-Time Signals and Systems - Theory and Computational Examples*.
| discrete_time_fourier_transform/theorems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Crawler For Non-MNIST Data Set<br/><br/>
#
# ### https://github.com/hellock/icrawler<br/>
#
# ### Try it with [pip install icrawler] or [conda install -c hellock icrawler]<br/><br/>
#
# This is a GitHub Library for Image Crawling.<br/>
# I downloaded 1000 images with 20 different keywords.
# +
from icrawler.builtin import GoogleImageCrawler
google_crawler = GoogleImageCrawler(storage={'root_dir': 'non-mnist'})
oogle_crawler = GoogleImageCrawler(
feeder_threads=1,
parser_threads=1,
downloader_threads=4,
storage={'root_dir': 'non-mnist'})
filters = dict(
type='photo',
size='medium',
color='color')
# +
keywords=['human','cat','dog','colorful','happy'\
,'kakao','unity','nodejs','macbook','peace'\
,'smile','raniy','sunny','banana','apple'\
,'setting','blender','memo','crap','king']
for i in range(20):
google_crawler.crawl(keyword=keywords[i], max_num=50, filters=filters, min_size=(100,100)\
,file_idx_offset=i*50)
# -
| Tensorflow/ImageCrawler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from plot_service import *
from numpy import *
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from sklearn.model_selection import learning_curve
from keras.wrappers.scikit_learn import KerasRegressor
# +
train_file = 'datasets/redshifts.csv'
X = loadtxt(train_file, usecols=(1, 2, 3, 4, 5, 6, 7, 8, 9, 10), unpack=True, delimiter=',').T
Y = loadtxt(train_file, unpack=True, usecols=(11), delimiter=',')
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.2, random_state=42)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.2, random_state=42)
print('Data loaded!')
# -
plot_simple_table(X_train.T[:, :30])
def build_learning_data(model, X, y):
train_sizes, train_scores, test_scores = learning_curve(model, X, y, cv=3, scoring='neg_mean_squared_error', train_sizes=linspace(0.01, 1.0, 50))
# Create means and standard deviations of training set scores
train_mean = mean(train_scores, axis=1)
train_std = std(train_scores, axis=1)
# Create means and standard deviations of test set scores
test_mean = mean(test_scores, axis=1)
test_std = std(test_scores, axis=1)
return train_sizes, train_mean, train_std, test_mean, test_std
def create_baseline_model():
model = Sequential()
model.add(Dense(300, input_dim=10, activation='relu'))
model.add(Dropout(0.1))
model.add(Dense(150, activation='relu'))
model.add(Dropout(0.05))
model.add(Dense(1, activation='linear'))
model.compile(loss='mse', optimizer='adam', metrics=['mse'])
return model
model = KerasRegressor(build_fn=create_baseline_model)
hist = model.fit(X_train, y_train, validation_data=(X_val, y_val), epochs=30)
preds = model.predict(X_test)
# +
pred = preds.reshape(len(preds))
real = y_test
plot_table(real, pred)
# -
score = model.score(X_val, y_val)
print('Cross-Val Score:', score)
plot_scatter(X_train, y_train, X_val, y_val, X_test, y_test, preds, show_only=True)
plot(hist.history, 'mean_squared_error', show_only=True)
plot_hm(real, pred, show_only=True)
train_sizes, train_mean, train_std, test_mean, test_std = build_learning_data(model, X_train, y_train)
plot_curves(train_sizes, train_mean, train_std, test_mean, test_std, show_only=True)
| jupyter/notebooks/ann_full_conn_redshift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Envelope Selection
#
# In this notebook, we will explore how to select the envelope from information layers.
# ## 0. Initialization
#
# ### 0.1. Load required libraries
import os
import topogenesis as tg
import pyvista as pv
import numpy as np
import pandas as pd
# extra import function
def lattice_from_csv(file_path):
# read metadata
meta_df = pd.read_csv(file_path, nrows=3)
shape = np.array(meta_df['shape'])
unit = np.array(meta_df['unit'])
minbound = np.array(meta_df['minbound'])
# read lattice
lattice_df = pd.read_csv(file_path, skiprows=5)
# create the buffer
buffer = np.array(lattice_df['value']).reshape(shape)
# create the lattice
l = tg.to_lattice(buffer, minbound=minbound, unit=unit)
return l
# ### 0.2. Load Sun Access Lattice
# loading the lattice from csv
sun_acc_path = os.path.relpath('../data/sun_access.csv')
sun_acc_lattice = lattice_from_csv(sun_acc_path)
# ## 1. Envelope Selection
#
# ### 1.1. Visualizing the selection
# +
p = pv.Plotter(notebook=True)
base_lattice = sun_acc_lattice
# Set the grid dimensions: shape + 1 because we want to inject our values on the CELL data
grid = pv.UniformGrid()
grid.dimensions = np.array(base_lattice.shape) + 1
# The bottom left corner of the data set
grid.origin = base_lattice.minbound - base_lattice.unit * 0.5
# These are the cell sizes along each axis
grid.spacing = base_lattice.unit
# adding the boundingbox wireframe
p.add_mesh(grid.outline(), color="grey", label="Domain")
# adding the avilability lattice
# init_avail_lattice.fast_vis(p)
# adding axes
p.add_axes()
p.show_bounds(grid="back", location="back", color="#aaaaaa")
def create_mesh(value):
lattice = np.copy(sun_acc_lattice)
lattice[sun_acc_lattice < value] *= 0.0
# Add the data values to the cell data
grid.cell_arrays["Agents"] = lattice.flatten(order="F") # Flatten the array!
# filtering the voxels
threshed = grid.threshold([0.001, 1.0])
# adding the voxels
p.add_mesh(threshed, name='sphere', show_edges=True, opacity=1.0, show_scalar_bar=False, clim=[0.0, 1.0])
return
p.add_slider_widget(create_mesh, [0, 1], title='Time', value=0, event_type="always", style="classic", pointa=(0.1, 0.1), pointb=(0.9, 0.1))
p.show(use_ipyvtk=True)
# -
# ### 1.2. Generating an envelope based on the selection
threshold = 0.675
new_avail_lattice = sun_acc_lattice > threshold
# ### 1.3. Visualize the new available lattice
# +
p = pv.Plotter(notebook=True)
# adding the avilability lattice
new_avail_lattice.fast_vis(p)
p.show(use_ipyvtk=True)
# -
# ### 1.4. Save new envelope to CSV
csv_path = os.path.relpath('../data/new_envelope_lattice.csv')
new_avail_lattice.to_csv(csv_path)
__author__ = "<NAME>"
__license__ = "MIT"
__version__ = "1.0"
__url__ = "https://github.com/shervinazadi/spatial_computing_workshops"
__summary__ = "Spatial Computing Design Studio Workshop [+]"
| notebooks/w+1_envelope_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
import pandas as pd
import numpy
import numpy as np
import random as rn
import functools
import re
'''
Task 1:
Check Pandas Version
'''
print('Task 1:')
print(pd.__version__)
# +
'''
Task 2:
Numpy Array
Create three columns with Zero values
'''
print('Task 2:')
dtype = [('Col1','int32'), ('Col2','float32'), ('Col3','float32')]
values = numpy.zeros(20, dtype=dtype)
index = ['Row'+str(i) for i in range(1, len(values)+1)]
df = pandas.DataFrame(values, index=index)
print(df)
df = pandas.DataFrame(values)
print(df)
# -
'''
Task 3:
iLoc in Pandas
Print first five rows
'''
print('Task 3:')
df = pandas.read_csv('data1.csv', sep=';', header=None)
print(df.iloc[:4]) # 0 - 4 = 5 values
'''
Task 4:
Create Random integer between 2 to 10 with 4 items
'''
print('Task 4:')
values = np.random.randint(2, 10, size=4)
print(values)
'''
Task 5:
Create Random integer between 2 to 10 with 4 items
'''
print('Task 5:')
df = pd.DataFrame(np.random.randint(0, 100, size=(3, 2)), columns=list('xy'))
print(df)
'''
Task 6:
Create Random integer between 2 to 10 with 4 items
'''
print('Task 6:')
df = pd.DataFrame(np.random.randint(0, 100, size=(2, 4)), columns=['A', 'B', 'C', 'D'])
print(df)
'''
Task 7:
'''
print('Task 7:')
values = np.random.randint(5, size=(2, 4))
print(values)
'''
Task 8:
Numpy Random Seed 0
'''
print('Task 8:')
# +
'''
Task 9:
3 rows, 2 columns in pandas
1st column = random between 10 to 20
2nd column = random between 80 and 90
3rd column = random between 40 and 50
'''
print('Task 9:')
dtype = [('One','int32'), ('Two','int32')]
values = np.zeros(3, dtype=dtype)
index = ['Row'+str(i) for i in range(1, 4)]
df = pandas.DataFrame(values, index=index)
print(df)
# +
'''
Task 10:
Fill Random Science and Math Marks
'''
print('Task 10:')
dtype = [('Science','int32'), ('Maths','int32')]
values = np.zeros(3, dtype=dtype)
#values = np.random.randint(10, 100, size = (3, 5))
#print(type(dtype))
#values = np.random.randint(5, size=(2, 4))
print(values)
index = ['Row'+str(i) for i in range(1, 4)]
df = pandas.DataFrame(values, index=index)
print(df)
# +
'''
Task 11:
CSV to Dataframe (from_csv)
Note: from_csv is Deprecated since version 0.21.0: Use pandas.read_csv() instead.
'''
print('Task 11:')
csv = pd.DataFrame.from_csv('uk-500.csv')
print(csv)
# -
'''
Task 12:
CSV to Dataframe (from_csv)
'''
print('Task 12:')
#df = df.from_csv(path, header, sep, index_col, parse_dates, encoding, tupleize_cols, infer_datetime_format)
df = pd.DataFrame.from_csv('uk-500.csv')
print(df)
'''
Task 13:
first 4 rows of CSV
'''
print('Task 13:')
df = pandas.read_csv('data1.csv', sep=',')
print(df.shape)
#print(df[2:14])
print(df.iloc[0:4,0:2])
#print(df[df.columns[0]])
'''
Task 14:
show even rows and first three columns
'''
print('Task 14:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
#print(df)
#print(df[2:14])
print(df.iloc[::2, 0:3])
#print(df.iloc[::1, 0:3])
#print(df[df.columns[0]])
# +
'''
Task 15:
New columns as sum of all
'''
print('Task 15:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
print(df)
df['total'] = df.sum(axis=1)
print(df)
#print(df[2:14])
#print(df.iloc[:,0:2])
#print(df[df.columns[0]])
# +
'''
Task 16:
Delete Rows of one column where the value is less than 50
'''
print('Task 16:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
print(df)
df = df[df.science > 50]
print(df)
# +
'''
Task 17:
Delete with Query
Note: Query doesn't work if your column has space in it
'''
print('Task 17:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
print(df)
df = df.query('science > 45')
print(df)
# -
'''
Task 18:
Skip single row
'''
print('Task 18:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8", skiprows=[5])
print(df.shape)
print(df)
# +
'''
Task 19:
Skip multiple rows
'''
print('Task 19:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8", skiprows=[1, 5, 7])
print(df.shape)
#print(df)
#df = df[df[[1]] > 45]
print(df)
# +
'''
Task 20:
Select Column by Index
Note:
df[[1]] doesn't work in later Pandas
'''
print('Task 20:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
print(df)
#df = df[int(df.columns[2]) > 45]
print(df)
print(type(df.columns[2]))
# +
'''
Task 21:
Skip rows
Note:
df[[1]] doesn't work in later Pandas
'''
print('Task 21:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8", skiprows=[0])
print(df.shape)
print(df)
#df = df[int(df.columns[2]) > 45]
#print(df)
print(df.columns[2])
# +
'''
Task 22:
String to Dataframe
Note:
df[[1]] doesn't work in later Pandas
'''
print('Task 22:')
from io import StringIO
s = """
1, 2
3, 4
5, 6
"""
df = pd.read_csv(StringIO(s), header=None)
print(df.shape)
print(df)
#df = df[int(df.columns[2]) > 45]
#print(df)
# -
'''
Task 23:
New columns as max of other columns
float to int used
'''
print('Task 23:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
df['sum'] = df.sum(axis=1)
df['max'] = df.max(axis=1)
df['min'] = df.min(axis=1)
df['average'] = df.mean(axis=1).astype(int)
print(df)
# +
'''
Task 24:
New columns as max of other columns
float to int used
Math is considered more, so double the marks for maths
'''
def apply_math_special(row):
return (row.maths * 2 + row.language / 2 + row.history / 3 + row.science) / 4
print('Task 24:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
df['sum'] = df.sum(axis=1)
df['max'] = df.max(axis=1)
df['min'] = df.min(axis=1)
df['average'] = df.mean(axis=1).astype(int)
df['math_special'] = df.apply(apply_math_special, axis=1).astype(int)
print(df)
# +
'''
Task 25:
New columns as max of other columns
35 marks considered as pass
If the student fails in math, consider fail
If the student passes in language and science, consider as pass
'''
def pass_one_subject(row):
if(row.maths > 34):
return 'Pass'
if(row.language > 34 and row.science > 34):
return 'Pass'
return 'Fail'
print('Task 25:')
df = pandas.read_csv('abc.csv', sep=',', encoding = "utf-8")
print(df.shape)
df['pass_one'] = df.apply(pass_one_subject, axis=1)
print(df)
# +
'''
Task 26:
fill with average
df.fillna(df.mean, inplace=True)
this will cause
Fatal Python error: Cannot recover from stack overflow.
Current thread 0x00008324 (most recent call first):
'''
print('Task 26:')
df = pandas.read_csv('abc2.csv', sep=',', encoding = "utf-8")
print(df.shape)
print(df)
df.fillna(df.mean(), inplace=True)
#df['pass_one'] = df.apply(pass_one_subject, axis=1)
print(df)
# -
'''
Task 28:
New columns as sum of all
'''
print('Task 28:')
df = pd.DataFrame(np.random.rand(10, 5))
df.iloc[0:3, 0:4] = np.nan # throw in some na values
print(df)
df.loc[:, 'test'] = df.iloc[:, 2:].sum(axis=1)
print(df)
'''
Task 29:
Unicode issue and fix
'''
print('Task 29:')
df = pandas.read_csv('score.csv', sep=',', encoding = "ISO-8859-1")
print(df.shape)
#print(df[2:14])
#print(df.iloc[:,0:2])
#print(df[df.columns[0]])
# +
'''
Task 27:
fill with average
df.fillna(df.mean, inplace=True)
this will cause
Fatal Python error: Cannot recover from stack overflow.
Current thread 0x00008324 (most recent call first):
'''
print('Task 27:')
df = pd.DataFrame(np.random.rand(3,4), columns=list("ABCD"))
print(df.shape)
print(df)
df.fillna(df.mean(), inplace=True)
#df['pass_one'] = df.apply(pass_one_subject, axis=1)
print(df)
# -
'''
Task 30:
Last 4 rows
'''
print('Task 30:')
df = pandas.read_csv('data1.csv', sep=';')
print(df[-4:])
'''
Task 31:
Simple Data analysis
'''
print('Task 31:')
'''
Task 32:
get 3 and 4th row
'''
print('Task 32:')
df = pandas.read_csv('data1.csv', sep=';')
print(df[2:4])
'''
Task 33:
Last 4th to 1st
'''
print('Task 33:')
df = pandas.read_csv('data1.csv', sep=';')
print(df[-4:-1])
'''
Task 34:
iloc position slice
'''
print('Task 34:')
df = pandas.read_csv('data1.csv', sep=';')
print(df.iloc[1:9])
'''
Task 35:
Loc - iloc - ix - at - iat
'''
print('Task 35:')
df = pandas.read_csv('data1.csv', sep=';')
# +
'''
Task 36:
Random data
'''
print('Task 36:')
def xrange(x):
return iter(range(x))
rnd_1 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
rnd_2 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
rnd_3 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
date = pd . date_range ( '2012-4-10' , '2015-1-4' )
print(len(date))
data = pd . DataFrame ({ 'date' : date , 'rnd_1' : rnd_1 , 'rnd_2' : rnd_2 , 'rnd_3' : rnd_3 })
# +
'''
Task 37:
filter with the value comparison
'''
print('Task 37:')
below_20 = data[data['rnd_1'] < 20]
print(below_20)
# -
'''
Task 38:
Filter between 5 and 10 on col 1
'''
print('Task 38:')
def xrange(x):
return iter(range(x))
rnd_1 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
rnd_2 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
rnd_3 = [ rn.randrange ( 1 , 20 ) for x in xrange ( 1000 )]
date = pd . date_range ( '2012-4-10' , '2015-1-4' )
print(len(date))
data = pd . DataFrame ({ 'date' : date , 'rnd_1' : rnd_1 , 'rnd_2' : rnd_2 , 'rnd_3' : rnd_3 })
below_20 = data[data['rnd_1'] < 20]
ten_to_20 = data[(data['rnd_1'] >= 5) & (data['rnd_1'] < 10)]
#print(ten_to_20)
# +
'''
Task 39:
15 to 20
'''
print('Task 39:')
date = pd . date_range ( '2018-08-01' , '2018-08-15' )
date_count = len(date)
def fill_rand(start, end, count):
return [rn.randrange(1, 20 ) for x in xrange( count )]
rnd_1 = fill_rand(1, 20, date_count)
rnd_2 = fill_rand(1, 20, date_count)
rnd_3 = fill_rand(1, 20, date_count)
#print(len(date))
data = pd . DataFrame ({ 'date' : date , 'rnd_1' : rnd_1 , 'rnd_2' : rnd_2 , 'rnd_3' : rnd_3 })
#print(len(date))
ten_to_20 = data[(data['rnd_1'] >= 15) & (data['rnd_1'] < 20)]
print(ten_to_20)
# +
'''
Task 40:
15 to 33
'''
print('Task 40:')
date = pd . date_range ( '2018-08-01' , '2018-08-15' )
date_count = len(date)
def fill_rand(start, end, count):
return [rn.randrange(1, 20 ) for x in xrange( count )]
rnd_1 = fill_rand(1, 20, date_count)
rnd_2 = fill_rand(1, 20, date_count)
rnd_3 = fill_rand(1, 20, date_count)
#print(len(date))
data = pd . DataFrame ({ 'date' : date , 'rnd_1' : rnd_1 , 'rnd_2' : rnd_2 , 'rnd_3' : rnd_3 })
#print(len(date))
ten_to_20 = data[(data['rnd_1'] >= 15) & (data['rnd_1'] < 20)]
print(ten_to_20)
# +
'''
Task 41:
set index
'''
print('Task 41:')
date = pd . date_range ( '2018-08-01' , '2018-08-15' )
date_count = len(date)
def xrange(x):
return iter(range(x))
def fill_rand(start, end, count):
return [rn.randrange(1, 20 ) for x in xrange( count )]
rnd_1 = fill_rand(1, 20, date_count)
rnd_2 = fill_rand(1, 20, date_count)
rnd_3 = fill_rand(1, 20, date_count)
#print(len(date))
data = pd . DataFrame ({ 'date' : date , 'rnd_1' : rnd_1 , 'rnd_2' : rnd_2 , 'rnd_3' : rnd_3 })
filter_loc = data.loc[ 2 : 4 , [ 'rnd_2' , 'date' ]]
print(filter_loc)
# -
'''
Task 42:
'''
print('Task 42:')
date_date = data.set_index( 'date' )
print(date_date.head())
# +
'''
Task 43:
Change columns based on other columns
'''
print('Task 43:')
df = pd.DataFrame({
'a' : [1,2,3,4],
'b' : [9,8,7,6],
'c' : [11,12,13,14]
});
print(df)
print('changing on one column')
# Change columns
df.loc[df.a >= 2,'b'] = 9
print(df)
# -
'''
Task 44:
Change multiple columns based on one column values
'''
print('Task 44:')
print('changing on multipe columns')
df.loc[df.a > 2,['b', 'c']] = 45
print(df)
'''
Task 45:
Pandas Mask
'''
print('Task 45:')
print(df)
df_mask = pd.DataFrame({
'a' : [True] * 4,
'b' : [False] * 4,
'c' : [True, False] * 2
})
print(df.where(df_mask,-1000))
'''
Task 46:
'''
print('Task 46:')
print(df)
df['logic'] = np.where(df['a'] > 5, 'high', 'low')
print(df)
'''
Task 47:
Student Marks (Pass or Fail)
'''
print('Task 47:')
marks_df = pd.DataFrame({
'Language' : [60, 45, 78, 4],
'Math' : [90, 80, 23, 60],
'Science' : [45, 90, 95, 20]
});
print(marks_df)
marks_df['language_grade'] = np.where(marks_df['Language'] >= 50, 'Pass', 'Fail')
marks_df['math_grade'] = np.where(marks_df['Math'] >= 50, 'Pass', 'Fail')
marks_df['science_grade'] = np.where(marks_df['Science'] >= 50, 'Pass', 'Fail')
print(marks_df)
# +
'''
Actors by age and movies acted
'''
'''
Task 48:
Get passed grades
'''
print('Task 48:')
marks_df = pd.DataFrame({
'Language' : [60, 45, 78, 4],
'Math' : [90, 80, 23, 60],
'Science' : [45, 90, 95, 20]
});
print(marks_df)
marks_df_passed_in_language = marks_df[marks_df.Language >=50 ]
print(marks_df_passed_in_language)
# -
'''
Task 49:
Students passed in Language and Math
'''
print('Task 49:')
marks_df_passed_in_lang_math = marks_df[(marks_df.Language >=50) & (marks_df.Math >= 50)]
print(marks_df_passed_in_lang_math)
# +
'''
Task 50:
Students passed in Language and Science
'''
print('Task 50:')
marks_df_passed_in_lang_and_sc = marks_df.loc[(marks_df.Language >=50) & (marks_df.Science >= 50)]
print(marks_df_passed_in_lang_and_sc)
# -
'''
Task 51:
Loc with Label oriented slicing
possible error:
pandas.errors.UnsortedIndexError
'''
print('Task 51:')
stars = {
'age' : [31, 23, 65, 50],
'movies' : [51, 23, 87, 200],
'awards' : [42, 12, 4, 78]
}
star_names = ['dhanush', 'simbu', 'kamal', 'vikram']
stars_df = pd.DataFrame(data=stars, index=[star_names])
print(stars_df)
'''
Task 52:
iloc with positional slicing
'''
print('Task 52:')
print(stars_df.iloc[1:3])
# +
'''
Task 53:
stars with names
'''
print('Task 40:')
numbers = pd.DataFrame({
'one' : [10, 50, 80, 40],
'two' : [2, 6, 56, 45]
},
index = [12, 14, 16, 18])
print(numbers)
print('label between 12 and 16')
print(numbers.loc[12:16])
print('index between 1 and 3')
print(numbers.iloc[1:3])
# -
'''
Task 54:
stars with names
'''
print('Task 54:')
stars = {
'age' : [31, 23, 65, 50],
'movies' : [51, 23, 87, 200],
'awards' : [42, 12, 4, 78]
}
star_names = ['dhanush', 'simbu', 'kamal', 'vikram']
stars_df = pd.DataFrame(data=stars, index=[star_names])
numbers = pd.DataFrame({
'one' : [10, 50, 80, 40],
'two' : [2, 6, 56, 45]
},
index = [12, 14, 16, 18])
print(numbers)
# +
'''
Task 55:
Row label selection
Age is above 25 and movies above 25
'''
print('Task 55:')
age_movies_25 = stars_df[(stars_df.movies > 25 ) & (stars_df.age > 25)]
print(age_movies_25)
# -
'''
Task 56:
stars in in certain ages
'''
print('Task 56:')
custom_stars = stars_df[stars_df.age.isin([31, 65])]
print(custom_stars)
'''
Task 57:
inverse opeartor
!( above one.45 and below two.50 )
'''
print('Task 57:')
print(numbers)
print(numbers[~( (numbers.one > 45) & (numbers.two < 50) )])
'''
Task 58:
Extend a panal frame by transposing
'''
print('Task 58:')
'''
Task 59:
adding a new dimension
'''
print('Task 59:')
'''
Task 60:
Creating new Columns using Applymap
Sides & applymap
'''
print('Task 60:')
sides_df = pd.DataFrame({
'a' : [1, 1, 2, 4],
'b' : [2, 1, 3, 4]
})
print(sides_df)
source_cols = sides_df.columns
print(source_cols)
new_cols = [str(x)+"_side" for x in source_cols]
side_category = {
1 : 'North',
2 : 'East',
3 : 'South',
4 : 'West'
}
sides_df[new_cols] = sides_df[source_cols].applymap(side_category.get)
print(sides_df)
# +
'''
Task 61:
Replacing some values with mean of the rest of a group
'''
print('Task 61:')
df = pd.DataFrame({'A' : [1, 1, 2, 2], 'B' : [1, -1, 1, 2]})
print(df)
gb = df.groupby('A')
def replace(g):
mask = g < 0
g.loc[mask] = g[~mask].mean()
return g
gbt = gb.transform(replace)
print(gbt)
# -
'''
Task 62:
Students passed in Language or Science (any one subject)
'''
print('Task 62:')
marks_df = pd.DataFrame({
'Language' : [60, 45, 78, 4],
'Math' : [90, 80, 23, 60],
'Science' : [45, 90, 95, 20]
});
print(marks_df)
marks_df_passed_in_lang_or_sc = marks_df.loc[(marks_df.Language >=50) | (marks_df.Science >= 50)]
print(marks_df_passed_in_lang_or_sc)
'''
Task 63:
possible errors:
TypeError: 'Series' objects are mutable, thus they cannot be hashed
'''
print('Task 63:')
marks_df['passed_one_subject'] = 'Fail'
marks_df.loc[(marks_df.Language >=50) , 'passed_one_subject'] = 'Pass'
print(marks_df)
# +
'''
Task 64:
argsort
Select rows with data closest to certain value using argsort
'''
print('Task 64:')
df = pd.DataFrame({
"a": np.random.randint(0, 100, size=(5,)),
"b": np.random.randint(0, 70, size=(5,))
})
print(df)
par = 65
print('with argsort')
df1 = df.loc[(df.a-par).abs().argsort()]
print(df1)
print(df.loc[(df.b-2).abs().argsort()])
# -
'''
Task 65:
argsort with stars
old stars (near by 50 age) argsort
'''
print('Task 65:')
stars = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"movies": [2, 3, 90, 45, 34, 2]
})
print(stars.loc[(stars.age - 50).abs().argsort()])
'''
Task 66:
Argsort with actors
young stars (near by 17)
'''
print('Task 66:')
print(stars.loc[(stars.age - 17).abs().argsort()])
'''
Task 67:
Binary operators
Stars with
younger than 19 - very young
more movies acted
'''
print('Task 67:')
stars = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"movies": [22, 33, 90, 75, 34, 2]
})
print(stars)
print('Young and more movies acted')
young = stars.age < 30
more_movies = stars.movies > 30
young_more = [young, more_movies]
young_more_Criteria = functools.reduce(lambda x, y : x & y, young_more)
print(stars[young_more_Criteria])
'''
Task 68:
Young, Higher Salary, and Higher Position
'''
print('Task 68:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8]
})
print(employees)
print('Young, Higher Salary, and Higher Position')
young = employees.age < 30
high_salary = employees.salary > 60
high_position = employees.grade > 6
young_salary_position = [young, high_salary, high_position]
young_salary_position_Criteria = functools.reduce(lambda x, y : x & y, young_salary_position)
print(employees[young_salary_position_Criteria])
'''
Task 69:
Rename columns
'''
print('Task 69:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8]
})
print(employees)
employees.rename(columns={'age': 'User Age', 'salary': 'Salary 2018'}, inplace=True)
print(employees)
'''
Task 70:
Add a new column
'''
print('Task 70:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8]
})
print(employees)
employees['group'] = pd.Series(np.random.randn(len(employees)))
print(employees)
'''
Task 71:
Drop a column
'''
print('Task 71:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8]
})
print(employees)
employees['group'] = pd.Series(np.random.randn(len(employees)))
print(employees)
employees.drop(employees.columns[[0]], axis=1, inplace = True)
print(employees)
'''
Task 72:
Drop multiple columns
'''
print('Task 72:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8]
})
print(employees)
employees['group'] = pd.Series(np.random.randn(len(employees)))
print(employees)
employees.drop(employees.columns[[1, 2]], axis=1, inplace = True)
print(employees)
# +
'''
Task 73:
Drop first and last column
'''
print('Task 73:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees.drop(employees.columns[[0, len(employees.columns)-1]], axis=1, inplace = True)
print(employees)
# +
'''
Task 74:
Delete by pop function
'''
print('Task 74:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
group = employees.pop('group')
print(employees)
print(group)
# -
'''
Task 75:
DataFrame.from_items
'''
print('Task 75:')
df = pd.DataFrame.from_items([('A', [1, 2, 3]), ('B', [4, 5, 6]), ('C', [7,8, 9])], orient='index', columns=['one', 'two', 'three'])
print(df)
# +
'''
Task 76:
Pandas to list
'''
print('Task 76:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees_list1 = list(employees.columns.values)
employees_list2 = employees.values.tolist()
#employees_list = list(employees)
print(employees_list1)
print(employees_list2)
# +
'''
Task 77:
Pandas rows to list
'''
print('Task 77:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees_list2 = employees.values.tolist()
print(employees_list2)
print(type(employees_list2))
print(len(employees_list2))
# +
'''
Task 78:
Pandas rows to array
Note: as_matrix is deprecated
'''
print('Task 78:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees_list2 = employees.values
print(employees_list2)
print(type(employees_list2))
print(employees_list2.shape)
# +
'''
Task 79:
Pandas rows to map
'''
print('Task 79:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees_list2 = map(list, employees.values)
print(employees_list2)
print(type(employees_list2))
# +
'''
Task 80:
Pandas rows to map
'''
print('Task 80:')
employees = pd.DataFrame({
"age": [17, 50, 24, 45, 65, 18],
"salary": [75, 33, 90, 175, 134, 78],
"grade" : [7, 8, 9, 2, 7, 8],
"group" : [1, 1, 2, 2, 2, 1]
})
print(employees)
employees_list2 = list(map(list, employees.values))
print(employees_list2)
print(type(employees_list2))
# +
'''
Task 81:
Drop duplicates
'''
print('Task 81:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users.drop_duplicates('id', inplace=True, keep='last')
print(users)
# +
'''
Task 82:
Selecting multiple columns
'''
print('Task 82:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users1 = users[['id', 'city']]
print(users1)
# +
'''
Task 83:
Selecting multiple columns
'''
print('Task 83:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
columns = ['id', 'count']
users1 = pd.DataFrame(users, columns=columns)
print(users1)
# +
'''
Task 84:
Row and Column Slicing
'''
print('Task 84:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users1 = users.iloc[0:2, 1:3]
print(users1)
# +
'''
Task 85:
Iterating rows
'''
print('Task 85:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
for index, row in users.iterrows():
print(row['city'], "==>", row['count'])
# +
'''
Task 86:
Iterating tuples
'''
print('Task 86:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
for row in users.itertuples(index=True, name='Pandas'):
print(getattr(row, 'city'))
for row in users.itertuples(index=True, name='Pandas'):
print(row.count)
# +
'''
Task 87:
Iterating rows and columns
'''
print('Task 87:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
for i, row in users.iterrows():
for j, col in row.iteritems():
print(col)
# +
'''
Task 88:
List of Dictionary to Dataframe
'''
print('Task 88:')
pointlist = [
{'points': 50, 'time': '5:00', 'year': 2010},
{'points': 25, 'time': '6:00', 'month': "february"},
{'points':90, 'time': '9:00', 'month': 'january'},
{'points_h1':20, 'month': 'june'}
]
print(pointlist)
pointDf = pd.DataFrame(pointlist)
print(pointDf)
pointDf1 = pd.DataFrame.from_dict(pointlist)
print(pointDf1)
# -
'''
Task 89:
'''
print('Task 89:')
df = pd.DataFrame(np.random.randn(10,6))
# Make a few areas have NaN values
df.iloc[1:3,1] = np.nan
df.iloc[5,3] = np.nan
df.iloc[7:9,5] = np.nan
print(df)
df1 = df.isnull()
print(df1)
'''
Task 90:
Sum of all nan
'''
print('Task 90:')
df = pd.DataFrame(np.random.randn(10,6))
# Make a few areas have NaN values
df.iloc[1:3,1] = np.nan
df.iloc[5,3] = np.nan
df.iloc[7:9,5] = np.nan
print(df)
print(df.isnull().sum())
print(df.isnull().sum(axis=1))
print(df.isnull().sum().tolist())
'''
Task 91:
Sum of all nan rowwise
'''
print('Task 91:')
df = pd.DataFrame(np.random.randn(10,6))
# Make a few areas have NaN values
df.iloc[1:3,1] = np.nan
df.iloc[5,3] = np.nan
df.iloc[7:9,5] = np.nan
print(df)
print(df.isnull().sum(axis=1))
'''
Task 92:
Sum of all nan as list
'''
print('Task 92:')
df = pd.DataFrame(np.random.randn(10,6))
# Make a few areas have NaN values
df.iloc[1:3,1] = np.nan
df.iloc[5,3] = np.nan
df.iloc[7:9,5] = np.nan
print(df)
print(df.isnull().sum().tolist())
# +
'''
Task 93:
Change the order of columns
Note:
FutureWarning: '.reindex_axis' is deprecated and will be removed in a future version
'''
print('Task 93:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users1 = users.reindex_axis(['city', 'count', 'id'], axis=1)
print(users1)
users2 = users.reindex(columns=['city', 'id', 'count'])
print(users2)
# +
'''
Task 94:
Drop multiple rows
'''
print('Task 94:')
numbers = pd.DataFrame({
"id": [1, 2, 3, 4, 5, 6],
"number": [10, 20, 30, 30, 23, 12]
})
print(numbers)
numbers.drop(numbers.index[[0, 3, 5]], inplace=True)
print(numbers)
# +
'''
Task 95:
Drop multiple rows by row name
'''
print('Task 95:')
numbers = pd.DataFrame({
"id": [1, 2, 3, 4, 5, 6],
"number": [10, 20, 30, 30, 23, 12]
}, index=['one', 'two', 'three', 'four', 'five', 'six'])
print(numbers)
numbers1 = numbers.drop(['two','six'])
print(numbers1)
numbers2 = numbers.drop('two')
print(numbers2)
# -
'''
Task 96:
Drop a single row
'''
print('Task 96:')
'''
Task 97:
Get the the odd row
'''
print('Task 97:')
x = numpy.array([
[ 1, 2, 3, 4, 5],
[ 6, 7, 8, 9, 10],
[11, 12, 13, 14, 15],
[16, 17, 18, 19, 20]
]
)
print(x)
print(x[::2])
'''
Task 98:
Get the even columns
'''
print('Task 98:')
x = numpy.array([
[ 1, 2, 3, 4, 5],
[ 6, 7, 8, 9, 10],
[11, 12, 13, 14, 15],
[16, 17, 18, 19, 20]
]
)
print(x)
print(x[:, 1::2])
# +
'''
Task 99:
Odd rows and even columns
'''
print('Task 99:')
x = numpy.array([
[ 1, 2, 3, 4, 5],
[ 6, 7, 8, 9, 10],
[11, 12, 13, 14, 15],
[16, 17, 18, 19, 20]
]
)
print(x)
print(x[::2, 1::2])
# +
'''
Task 100:
Drop duplicates
'''
print('Task 100:')
users = pd.DataFrame({
"id": [1, 1, 2, 2, 3, 3],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users.drop_duplicates('id', inplace=True)
print(users)
# +
'''
Task 101:
Drop all duplicates
'''
print('Task 101:')
users = pd.DataFrame({
"name": ['kevin', 'james', 'kumar', 'kevin', 'kevin', 'james'],
"city": ['Toronto', 'Montreal', 'Calgary', 'Montreal', 'Montreal', 'Ottawa'],
"count" : [7, 8, 9, 2, 7, 8]
})
print(users)
users.drop_duplicates('name', inplace=True, keep='last')
print(users)
users1 = users.drop_duplicates('name', keep=False)
print(users1)
# -
'''
Task 102:
Multi Indexing:
'''
print('Task 102:')
'''
Task 103:
Missing Data:
Make A'th 3rd coulmn Nan
'''
print('Task 103:')
df = pd.DataFrame(np.random.randn(6,1), index=pd.date_range('2013-08-01', periods=6, freq='B'), columns=list('A'))
print(df)
df.loc[df.index[3], 'A'] = np.nan
print(df)
'''
Task 104:
'''
print('Task 104:')
df1 = df.reindex(df.index[::-1]).ffill()
print(df1)
'''
Task 105:
Cumsum reset Nan
'''
print('Task 105:')
'''
Task 106:
'''
print('Task 106:')
df = pd.DataFrame([['http://wap.blah.com/xxx/id/11/someproduct_step2;jsessionid=....']],columns=['A'])
df1 = df['A'].str.findall("\\d\\d\\/(.*?)(;|\\?)",flags=re.IGNORECASE).apply(lambda x: pd.Series(x[0][0],index=['first']))
print(df1)
'''
Task 107:
Grouping:
'''
print('Task 107:')
animals_df = pd.DataFrame({'animal': 'cat dog cat fish dog cat cat'.split(),
'size': list('SSMMMLL'),
'weight': [8, 10, 11, 1, 20, 12, 12],
'adult' : [False] * 5 + [True] * 2})
print(animals_df)
'''
Task 108:
Basic Group by
'''
print('Task 108:')
animals_df1 = animals_df.groupby('animal').apply(lambda x: x['size'][x['weight'].idxmax()])
print(animals_df1)
'''
Task 109:
Get Group
'''
print('Task 109:')
cats = animals_df.groupby(['animal']).get_group('cat')
print(cats)
'''
Task 110:
'''
print('Task 110:')
weights = animals_df.groupby(['weight']).get_group(20)
print(weights)
# +
'''
Task 111:
'''
print('Task 111:')
def GrowUp(x):
avg_weight = sum(x[x['size'] == 'series1'].weight * 1.5)
avg_weight += sum(x[x['size'] == 'M'].weight * 1.25)
avg_weight += sum(x[x['size'] == 'L'].weight)
avg_weight /= len(x)
return pd.Series(['L',avg_weight,True], index=['size', 'weight', 'adult'])
animals_df = pd.DataFrame({'animal': 'cat dog cat fish dog cat cat'.split(),
'size': list('SSMMMLL'),
'weight': [8, 10, 11, 1, 20, 12, 12],
'adult' : [False] * 5 + [True] * 2})
gb = animals_df.groupby(['animal'])
expected_df = gb.apply(GrowUp)
print(expected_df)
# -
'''
Task 112:
Expanding Apply:
'''
print('Task 112:')
series1 = pd.Series([i / 100.0 for i in range(1,6)])
print(series1)
def CumRet(x,y):
return x * (1 + y)
def Red(x):
return functools.reduce(CumRet,x,1.0)
s2 = series1.expanding().apply(Red)
# s2 = series1.expanding().apply(Red, raw=True) # is not working
print(s2)
'''
Task 113:
Create Random integer between 0 to 100 with 10 itmes (2 rows, 5 columns)
'''
print('Task 113:')
df = pd.DataFrame(np.random.randint(0, 100, size=(3, 5)), columns=['Toronto', 'Ottawa', 'Calgary', 'Montreal', 'Quebec'])
print(df)
| pandas_150.ipynb |
# + cell_id="f2b872cafc114de78f994e0277570619" editable=false
@file:DependsOn(".")
# + cell_id="0b1e78afb57648ff9846db7db77ef10f" editable=false
import backend.*
# + [markdown] cell_id="08d37d33b32449d7889e7693f77b52bc" editable=false lambdacheck={"layout": {"column_offset": 0, "column_span": 4, "row_span": 10}}
# This is the interpreter's main function.
# + cell_id="4a91cd8f89e548a7a7aaa6fe08ae5aa1" editable=false lambdacheck={"layout": {"column_offset": 4, "column_span": 8}}
fun run(program: Expr) {
try {
val data = program.eval(Context())
println("> ${data}")
} catch(e: Exception) {
println("[error] ${e}")
}
}
# + cell_id="1083ec4271464b2994d55c9b976046ca" editable=false lambdacheck={"check": {"_check_id": "1083ec4271464b2994d55c9b976046ca", "_immediate": true, "_name": "", "display.text/plain": true, "output.stderr": true, "output.stdout": true, "result.text/plain": true}, "outputs": [{"name": "stdout", "output_type": "stream", "text": "> Int(10)\n"}]}
run(IntLiteral(10))
# + cell_id="cb311c8bd9de428cb192f6a892469cb9" editable=false lambdacheck={"check": {"_check_id": "cb311c8bd9de428cb192f6a892469cb9", "_immediate": true, "_name": "", "display.text/plain": true, "output.stderr": true, "output.stdout": true, "result.text/plain": true}, "outputs": [{"name": "stdout", "output_type": "stream", "text": "[error] java.lang.Exception: function does not exist: f\n"}]}
run(
Invoke("f", listOf())
)
# + [markdown] cell_id="1a5de7466e4e483aac20b4b8f2f37b06" lambdacheck={"layout": {"column_offset": 0, "column_span": 12}, "workunit": {"id": "1a5de7466e4e483aac20b4b8f2f37b06", "type": "title"}}
# ## Work Unit 1
# + [markdown] cell_id="aafd05155daf4b5c88b858e23f51fdc6" editable=false lambdacheck={"layout": {"column_offset": 0, "column_span": 4, "row_span": 2}, "workunit": {"id": "1a5de7466e4e483aac20b4b8f2f37b06", "type": "text"}}
# Implement a program which is equivalent to the following:
#
# ```
# function factorial(i) {
# if(i < 2)
# 1
# else
# i * factorial(i-1)
# }
#
# factorial(10)
# ```
# + cell_id="ce499345e4a148058cca324923d456de" lambdacheck={"layout": {"column_offset": 4, "column_span": 8}, "workunit": {"id": "1a5de7466e4e483aac20b4b8f2f37b06", "type": "student-solution"}}
val program1 = Block(listOf(
Declare("factorial",
listOf("i"),
IfElse(
Compare(Comparator.LT, Deref("i"), IntLiteral(2)),
IntLiteral(1),
Arithmetic(Op.Mul,
Deref("i"),
Invoke("factorial", listOf(Arithmetic(Op.Sub,
Deref("i"),
IntLiteral(1)))))
)),
Invoke("factorial", listOf(IntLiteral(10)))
))
# + cell_id="eee391513d4d42e9a6b07da31b22ef66" editable=false lambdacheck={"check": {"_check_id": "eee391513d4d42e9a6b07da31b22ef66", "_immediate": true, "_name": "", "display.text/plain": true, "output.stderr": true, "output.stdout": true, "result.text/plain": true}, "outputs": [{"name": "stdout", "output_type": "stream", "text": "> Int(3628800)\n"}]}
run(program1)
# + [markdown] cell_id="cbd8c3f0134340869c42ba830bbb6f58" lambdacheck={"layout": {"column_offset": 0, "column_span": 12}, "workunit": {"id": "cbd8c3f0134340869c42ba830bbb6f58", "type": "title"}}
# ## Work Unit 2
# + [markdown] cell_id="aac814431e4a4a178a67b4c41141042f" editable=false lambdacheck={"layout": {"column_offset": 0, "column_span": 4, "row_span": 2}, "workunit": {"id": "cbd8c3f0134340869c42ba830bbb6f58", "type": "text"}}
# Implement a program equivalent to the following:
#
# ```
# function fac(i) {
# prod = 1
# counter = 1
# loop(counter <= i) {
# prod = prod * counter
# counter = counter + 1
# }
# }
#
# fac(10)
# ```
# + cell_id="03895701b66045e481596a71b155cc0d" lambdacheck={"layout": {"column_offset": 4, "column_span": 8}, "workunit": {"id": "cbd8c3f0134340869c42ba830bbb6f58", "type": "student-solution"}}
val program2 = Block(listOf(
Declare(
"fac",
listOf("i"),
Block(
listOf(
Bind("prod", IntLiteral(1)),
Bind("counter", IntLiteral(1)),
Loop(
Compare(Comparator.LE, Deref("counter"), Deref("i")),
Block(
listOf(
Bind("prod", Arithmetic(Op.Mul,
Deref("prod"),
Deref("counter"))),
Bind("counter", Arithmetic(Op.Add,
Deref("counter"),
IntLiteral(1))),
)
)
),
Deref("prod")
)
)),
Invoke("fac", listOf(IntLiteral(10)))
))
# + cell_id="19ecbfc190764cff831bb65c5cb6c1dd" editable=false lambdacheck={"check": {"_check_id": "19ecbfc190764cff831bb65c5cb6c1dd", "_immediate": true, "_name": "", "display.text/plain": true, "output.stderr": true, "output.stdout": true, "result.text/plain": true}, "outputs": [{"name": "stdout", "output_type": "stream", "text": "> Int(3628800)\n"}]}
run(program2)
| 2020_notebooks/Labs/lab_6/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Remove input cells at runtime (nbsphinx)
import IPython.core.display as d
d.display_html('<script>jQuery(function() {if (jQuery("body.notebook_app").length == 0) { jQuery(".input_area").toggle(); jQuery(".prompt").toggle();}});</script>', raw=True)
# # Direction Look-Up-Tables (LUTs)
# **Datasample:** gamma-1 (goes into energy training)
#
# **Data level:** DL1b (telescope-wise image parameters)
#
# **Scope:**
#
# To obtain an estimate for an image, given its intensity, width and length, how reliable its axis is as a measure of the shower axis' orientation.
# The values from the LUTs can be used to set relative weights for the different telescopes in the stereoscopic reconstruction of events with three or more valid images.
#
# **Approach:**
#
# - calculate for each image the miss parameter, aka the distance from the image axis to the point on the camera which corresponds to the true gamma-ray direction
#
# - build a LUT per telescope type, containing in bins of image intensity and width/length, the square of \<miss>.
# ## Table of contents
# - [Counts](#Counts-LUTs)
# - [Counts ratio between protopipe and CTAMARS](#Count-LUTs-ratio-between-protopipe-and-CTAMARS)
# - [Direction LUT](#Direction-LUT)
# - [Direction LUT comparisons between protopipe and CTAMARS](#Direction-LUT-ratio-between-protopipe-and-CTAMARS)
# - [Profile along Y-axis (width/length)](#Profile-along-Y-axis-(width/length))
# - [Ratio between the LUTs](#Ratio-between-the-LUTs)
# + [markdown] nbsphinx="hidden"
# ## Imports
# +
from pathlib import Path
import numpy as np
from scipy.stats import binned_statistic_2d
import pandas
import tables
import uproot
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
from ctapipe.image import camera_to_shower_coordinates
# + [markdown] nbsphinx="hidden"
# ## Functions
# -
def get_camera_names(inputPath = None, fileName = None):
"""Read the names of the cameras.
Parameters
==========
infile : str
Full path of the input DL1 file.
fileName : str
Name of the input DL1 file.
Returns
=======
camera_names : list(str)
Table names as a list.
"""
if (inputPath is None) or (fileName is None):
print("ERROR: check input")
h5file = tables.open_file(inputPath / fileName, mode='r')
group = h5file.get_node("/")
camera_names = [x.name for x in group._f_list_nodes()]
h5file.close()
return camera_names
def load_reset_infile_protopipe(inputPath = None, fileName = None, camera_names=None, cols=None):
"""(Re)load the file containing DL1(a) data and extract the data per telescope type.
Parameters
==========
infile : str
Full path of the input DL1 file.
fileName : str
Name of the input DL1 file.
Returns
=======
dataFrames : dict(pandas.DataFrame)
Dictionary of tables per camera.
"""
if (inputPath is None) or (fileName is None):
print("ERROR: check input")
if camera_names is None:
print("ERROR: no cameras specified")
# load DL1 images
dataFrames = {camera : pandas.read_hdf(inputPath / fileName, f"/{camera}") for camera in camera_names}
return dataFrames
# + [markdown] nbsphinx="hidden"
# ## Input data
# -
# First we check if a _plots_ folder exists already.
# If not, we create it.
Path("./plots").mkdir(parents=True, exist_ok=True)
# + [markdown] nbsphinx="hidden"
# ### CTAMARS
# +
indir_CTAMARS = Path("/Volumes/DataCEA_PERESANO/Data/CTA/ASWG/Prod3b/Release_2019/CTAMARS_reference_data/TRAINING/DL1")
filename_CTAMARS = "DirLUT.root"
filepath_CTAMARS = Path(indir_CTAMARS, filename_CTAMARS)
CTAMARS_cameras = ["LSTCam", "NectarCam"]
CTAMARS_histograms = ["DirLookupTable", "DirLookupTable_degrees", "DirEventStatistics"]
CTAMARS = dict.fromkeys(CTAMARS_cameras)
with uproot.open(filepath_CTAMARS) as infile_CTAMARS:
for camera_index in range(len(CTAMARS_cameras)):
CTAMARS[CTAMARS_cameras[camera_index]] = dict.fromkeys(CTAMARS_histograms)
CTAMARS[CTAMARS_cameras[camera_index]][f"DirLookupTable"] = infile_CTAMARS[f"DirLookupTable_type{camera_index}"]
CTAMARS[CTAMARS_cameras[camera_index]][f"DirLookupTable_degrees"] = infile_CTAMARS[f"DirLookupTable_degrees_type{camera_index}"]
CTAMARS[CTAMARS_cameras[camera_index]][f"DirEventStatistics"] = infile_CTAMARS[f"DirEventStatistics_type{camera_index}"]
CTAMARS_X_edges = CTAMARS["LSTCam"]["DirLookupTable"].axes[0].edges()
CTAMARS_Y_edges = CTAMARS["LSTCam"]["DirLookupTable"].axes[1].edges()
# + [markdown] nbsphinx="hidden"
# ### protopipe
# -
# EDIT ONLY THIS CELL
indir_protopipe = Path("/Users/michele/Applications/ctasoft/dirac/shared_folder/analyses/v0.4.0_dev1/data/TRAINING/for_energy_estimation")
filename_protopipe = "TRAINING_energy_tail_gamma_merged.h5"
# +
cameras = get_camera_names(inputPath = indir_protopipe,
fileName = filename_protopipe)
PROTOPIPE = load_reset_infile_protopipe(inputPath = indir_protopipe,
fileName = filename_protopipe,
camera_names=cameras)
# -
# - ``miss`` is here defined as the absolute value of the component transverse to the main shower axis of the distance between the true source position (0,0 in case of on-axis simulation) and the COG of the cleaned image,
# - it is calculated for ALL images of the gamma1 sample and added to the tables for each camera,
# - then we select only images for which miss < 1.0 deg in each camera
# +
PROTOPIPE_selected = {}
for camera in cameras:
hillas_x = PROTOPIPE[camera]["hillas_x_reco"]
hillas_y = PROTOPIPE[camera]["hillas_y_reco"]
hillas_psi = PROTOPIPE[camera]["hillas_psi_reco"]
# Components of the distance between center of the camera (for on-axis simulations) and reconstructed position of the image
longitudinal, transverse = camera_to_shower_coordinates(x = 0.,
y = 0.,
cog_x = hillas_x,
cog_y = hillas_y,
psi = np.deg2rad(hillas_psi))
# Take the absolute value of the transverse component
# Add miss to the dataframe
PROTOPIPE[camera]["miss"] = np.abs(transverse)
# miss < 1 deg
mask = PROTOPIPE[camera]["miss"] < 1.0
# Make a smaller dataframe with just what we actually need and select for miss < 1 deg
PROTOPIPE_selected[camera] = PROTOPIPE[camera][['hillas_intensity_reco', 'hillas_width_reco', 'hillas_length_reco', 'miss']].copy()
PROTOPIPE_selected[camera] = PROTOPIPE_selected[camera][mask]
# -
# ## Counts
# [back to top](#Table-of-contents)
# This is just the 2D grid that will host the LUT, showing how many events fall in each bin.
#
# In CTAMARS an additional image quality cut for direction reconstruction selects for images that fall in a bin which contains >10 images
# +
fig = plt.figure(figsize=(12, 5))
plt.subplots_adjust(wspace = 0.25)
PROTOPIPE_COUNTS = {}
for i, camera in enumerate(cameras):
plt.subplot(1, 2, i+1)
intensity = PROTOPIPE_selected[camera]["hillas_intensity_reco"]
width = PROTOPIPE_selected[camera]["hillas_width_reco"]
length = PROTOPIPE_selected[camera]["hillas_length_reco"]
PROTOPIPE_COUNTS[camera], _, _, _ = plt.hist2d(x = np.log10(intensity),
y = width / length,
bins = [CTAMARS_X_edges, CTAMARS_Y_edges],
norm = LogNorm(),
cmap = "rainbow")
plt.title(camera)
cb = plt.colorbar()
cb.set_label("Number of images")
plt.xlabel("log10(intensity) [phe]")
plt.ylabel("width / length")
# -
# ## Counts ratio between protopipe and CTAMARS
# [back to top](#Table-of-contents)
# +
fig = plt.figure(figsize=(15, 7))
plt.subplots_adjust(wspace = 0.4)
font_size = 20
for i, camera in enumerate(cameras):
RATIO = PROTOPIPE_COUNTS[camera]/CTAMARS[camera]["DirEventStatistics"].values()
plt.subplot(1, 2, i+1)
plt.pcolormesh(CTAMARS_X_edges,
CTAMARS_Y_edges,
np.transpose(PROTOPIPE_COUNTS[camera]/CTAMARS[camera]["DirEventStatistics"].values()),
norm = LogNorm()
)
# add value labels for better visualization
for i, x in enumerate(CTAMARS[camera]["DirLookupTable_degrees"].axes[0].centers()):
for j, y in enumerate(CTAMARS[camera]["DirLookupTable_degrees"].axes[1].centers()):
plt.text(x,
y,
np.round(RATIO[i][j]),
ha='center',va='center',
size=10,color='b')
plt.title(camera, fontsize=font_size)
ax = plt.gca()
cb = plt.colorbar()
cb.set_label("Counts ratio protopipe/CTAMARS", fontsize=font_size)
ax.tick_params(axis='both', which='major', labelsize=font_size)
ax.tick_params(axis='both', which='minor', labelsize=font_size)
plt.xlabel("log10(intensity) [phe]", fontsize=font_size)
plt.ylabel("width / length", fontsize=font_size)
# -
# ## Direction LUT
# [back to top](#Table-of-contents)
# +
# Build the LUT by using,
# - ``np.log10(intensity)`` as ``x`` axis,
# - ``width/length`` as ``y``axis,
# For each 2D bin we calculate the ``mean of miss`` for the images which fall into that bin.
mean_miss = {}
for camera in cameras:
intensity = PROTOPIPE_selected[camera]["hillas_intensity_reco"]
width = PROTOPIPE_selected[camera]["hillas_width_reco"]
length = PROTOPIPE_selected[camera]["hillas_length_reco"]
miss = PROTOPIPE_selected[camera]["miss"]
mean_miss[camera], _, _, _ = binned_statistic_2d(x = np.log10(intensity),
y = width/length,
values = miss,
statistic='mean',
bins=[CTAMARS_X_edges, CTAMARS_Y_edges]
)
# +
# After obtaining such a 2D binned statistic we square the value of each bin.
# That is the final LUT
LUT = {}
for camera in cameras:
LUT[camera] = np.square(mean_miss[camera])
# +
fig = plt.figure(figsize=(12, 5))
plt.subplots_adjust(wspace = 0.4)
for i, camera in enumerate(cameras):
plt.subplot(1, 2, i+1)
plt.pcolormesh(CTAMARS_X_edges,
CTAMARS_Y_edges,
np.transpose( LUT[camera] ),
norm = LogNorm(vmin = 1.e-4, vmax = 2.e-1),
cmap = "rainbow"
)
plt.title(camera)
cb = plt.colorbar()
cb.set_label("<miss>**2")
plt.xlabel("log10(intensity [phe])")
plt.ylabel("width / length")
plt.xlim(CTAMARS_X_edges[1], CTAMARS_X_edges[-2])
# -
# ## Direction LUT comparisons between protopipe and CTAMARS
# [back to top](#Table-of-contents)
# ### Profile along Y-axis (width/length)
# [back to top](#Table-of-contents)
# Here we select as an example the bin #9, containing images with 0.45 < width / length < 0.55
# +
plt.figure(figsize=(15,10))
plt.subplots_adjust(hspace=0.2, wspace=0.2)
for i, camera in enumerate(cameras):
plt.subplot(2, 2, i*2+1)
H = np.transpose(CTAMARS[camera]["DirLookupTable_degrees"].values())
plt.errorbar(x = CTAMARS[camera]["DirLookupTable_degrees"].axes[0].centers(),
y = H[9],
xerr = np.diff(CTAMARS_X_edges)/2,
yerr = None,
fmt="o",
label="CTAMARS")
plt.errorbar(x = CTAMARS[camera]["DirLookupTable_degrees"].axes[0].centers(),
y = np.transpose(LUT[camera])[9],
xerr = np.diff(CTAMARS_X_edges)/2,
yerr = None,
fmt="o",
label="protopipe")
plt.xlabel("log10(intensity) [phe]")
plt.ylabel("<miss>**2 [deg**2]")
plt.grid()
plt.yscale("log")
plt.title(camera)
plt.legend()
plt.xlim(CTAMARS_X_edges[1], CTAMARS_X_edges[-1])
plt.ylim(1.e-4, 2.e-1)
plt.subplot(2, 2, i*2+2)
ratio = np.transpose(LUT[camera])[9] / H[9]
plt.errorbar(x = CTAMARS[camera]["DirLookupTable_degrees"].axes[0].centers()[1:-1],
y = np.log10(ratio[1:-1]),
xerr = np.diff(CTAMARS_X_edges[1:-1])/2,
yerr = None,
ls = "-",
fmt="o",)
plt.hlines(0., plt.gca().get_xlim()[0], plt.gca().get_xlim()[1], colors="red", linestyles='solid')
plt.xlabel("log10(intensity) [phe]")
plt.ylabel("log10(protopipe / CTAMARS)")
plt.grid()
plt.title(camera)
plt.xlim(CTAMARS_X_edges[1], CTAMARS_X_edges[-1])
plt.ylim(-2,2.)
# -
# ### Ratio between the LUTs
# [back to top](#Table-of-contents)
# +
# we use the same bin edges of CTAMARS reference data
fig = plt.figure(figsize=(12, 5))
plt.subplots_adjust(wspace = 0.25)
for i, camera in enumerate(cameras):
plt.subplot(1, 2, i+1)
plt.pcolormesh(CTAMARS_X_edges,
CTAMARS_Y_edges,
np.transpose( LUT[camera] / CTAMARS[camera]["DirLookupTable_degrees"].values()),
norm=LogNorm(),
cmap = "rainbow"
)
plt.title(camera)
cb = plt.colorbar()
cb.set_label("<miss>**2 ratio protopipe/CTAMARS")
plt.xlabel("log10(intensity) [phe]")
plt.ylabel("width / length")
plt.xlim(CTAMARS_X_edges[1], CTAMARS_X_edges[-2])
# -
# Same, but zomming in the regime of current image quality cuts
#
# - 0.1 < width/length < 0.6
# - intensity > 50 phe
# +
fig = plt.figure(figsize=(12, 5))
plt.subplots_adjust(wspace = 0.25)
for i, camera in enumerate(cameras):
plt.subplot(1, 2, i+1)
plt.pcolormesh(CTAMARS_X_edges[2:-2],
CTAMARS_Y_edges[2:13],
np.transpose( LUT[camera] / CTAMARS[camera]["DirLookupTable_degrees"].values())[2:12,2:-2],
cmap = "rainbow"
)
plt.title(camera)
cb = plt.colorbar()
cb.set_label("<miss>**2 ratio protopipe/CTAMARS")
plt.xlabel("log10(intensity) [phe]")
plt.ylabel("width / length")
# -
| docs/contribute/benchmarks/TRAINING/benchmarks_DL1_DirectionLUT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# Click this button to open the notebook in SageMaker Studio Lab. [](https://studiolab.sagemaker.aws/import/github/neo4j-partners/hands-on-lab-neo4j-and-sagemaker/blob/main/Lab%205%20-%20Graph%20Data%20Science/embedding.ipynb)
# + [markdown] id="JtcD7PrPzSqE"
# # Install Prerequisites
# First off, you'll also need to install a few packages.
# + id="qwKogqD_He_e" vscode={"languageId": "python"}
# %pip install --quiet --upgrade neo4j
# %pip install pandas
# + [markdown] id="IdMFRbqGzSqF"
# # Working with Neo4j
# You'll need to enter the credentials from your Neo4j instance below.
#
# The default DB_NAME is always neo4j.
# + id="P41l_P4zzSqF" vscode={"languageId": "python"}
# Edit this variable!
DB_URL = 'neo4j://172.16.58.3:7687'
# You can leave these defaults
DB_USER = 'neo4j'
DB_PASS = '<PASSWORD>'
DB_NAME = 'neo4j'
# + id="8lUkSvmozSqF" vscode={"languageId": "python"}
import pandas as pd
from neo4j import GraphDatabase
driver = GraphDatabase.driver(DB_URL, auth=(DB_USER, DB_PASS))
# + [markdown] id="ZtJy4eO_zSqF"
# First we're going to create an in memory graph represtation of the data in Neo4j Graph Data Science (GDS).
# + id="URRShWv0zSqG" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.read_transaction(
lambda tx: tx.run(
"""
CALL gds.graph.create(
'mygraph',
['Company', 'Manager', 'Holding'],
{
OWNS: {orientation: 'UNDIRECTED'},
PARTOF: {orientation: 'UNDIRECTED'}
}
)
YIELD
graphName AS graph,
relationshipProjection AS readProjection,
nodeCount AS nodes,
relationshipCount AS rels
"""
).data()
)
df = pd.DataFrame(result)
display(df)
# + [markdown] id="hFwZUeZY0jd7"
# Note, if you get an error saying the graph already exists, that's probably because you ran this code before. You can destroy it using this command:
# + id="_bxCuwC_0gWZ" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.read_transaction(
lambda tx: tx.run(
"""
CALL gds.graph.drop('mygraph')
"""
).data()
)
# + [markdown] id="ZnnO932bBexn"
# Now, let's list the details of the graph to make sure the projection was created as we want.
# + id="7ZBPLijbrSCt" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.read_transaction(
lambda tx: tx.run(
"""
CALL gds.graph.list()
"""
).data()
)
print(result)
# + [markdown] id="4pFaTeUSzSqG"
# Now we can generate an embedding from that graph. This is a new feature we can use in our predictions. We're using FastRP, which is a more full featured and higher performance of Node2Vec. You can learn more about that [here](https://neo4j.com/docs/graph-data-science/current/algorithms/fastrp/).
# + id="VVxhVtTq4Kfc" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.read_transaction(
lambda tx: tx.run(
"""
CALL gds.fastRP.mutate('mygraph',{
embeddingDimension: 16,
randomSeed: 1,
mutateProperty:'embedding'
})
"""
).data()
)
df = pd.DataFrame(result)
display(df)
# + [markdown] id="_GR9GCbFitFy"
# That creates an embedding for each node type. However, we only want the embedding on the nodes of type holding.
#
# We're going to take the embedding from our projection and write it to the holding nodes in the underlying database.
# + id="C-EKqaR-inUe" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.run(
"""
CALL gds.graph.writeNodeProperties('mygraph', ['embedding'], ['Holding'])
YIELD writeMillis
"""
)
print(result)
# + id="RLTjaLlMzSqH" vscode={"languageId": "python"}
with driver.session(database=DB_NAME) as session:
result = session.read_transaction(
lambda tx: tx.run(
"""
MATCH (n:Holding) RETURN n
"""
).data()
)
# + [markdown] id="IGeVDYfrCEZx"
# Note that this query will take 2-3 minutes to run as it's grabbing nearly half a million nodes along with all their properties and our new embedding.
# + id="WUpaQ69smfJ9" vscode={"languageId": "python"}
df = pd.DataFrame([dict(record.get('n')) for record in result])
df
# + [markdown] id="ND0BKcgNDPgH"
# Note that the embedding row is an array. To make this dataset more consumable, we should flatten that out into multiple individual features: embedding_0, embedding_1, ... embedding_n.
#
# + id="W11q18eTC6-N" vscode={"languageId": "python"}
embeddings = pd.DataFrame(df['embedding'].values.tolist()).add_prefix("embedding_")
merged = df.drop(columns=['embedding']).merge(embeddings, left_index=True, right_index=True)
merged
# + [markdown] id="_ooXlcpRbFd0"
# Now that we have the data formatted properly, let's split it into a training and a testing set and write those to disk.
# + id="UdPgEiDdDu-B" vscode={"languageId": "python"}
df = merged
df['split']=df['reportCalendarOrQuarter']
df['split']=df['split'].replace(['03-31-2021', '06-30-2021', '09-30-2021'], ['TRAIN', 'VALIDATE', 'TEST'])
df = df.drop(columns=['reportCalendarOrQuarter'])
df.to_csv('embedding.csv', index=False)
# + [markdown] id="Azb1inEVAC7f"
# # Upload to Amazom S3
# Now let's create a bucket and upload our data set to it. Then we'll be able to access the data in SageMaker in the next lab.
# + id="mRKBGEi3AC7f" vscode={"languageId": "python"}
filename='embedding.csv'
#to do
| Lab 5 - Graph Data Science/embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Using TensorFlow Scripts in SageMaker - Quickstart
#
# Starting with TensorFlow version 1.11, you can use SageMaker's TensorFlow containers to train TensorFlow scripts the same way you would train outside SageMaker. This feature is named **Script Mode**.
#
# This example uses
# [Multi-layer Recurrent Neural Networks (LSTM, RNN) for character-level language models in Python using Tensorflow](https://github.com/sherjilozair/char-rnn-tensorflow).
# You can use the same technique for other scripts or repositories, including
# [TensorFlow Model Zoo](https://github.com/tensorflow/models) and
# [TensorFlow benchmark scripts](https://github.com/tensorflow/benchmarks/tree/master/scripts/tf_cnn_benchmarks).
# ## Test locally using SageMaker Python SDK TensorFlow Estimator
# You can use the SageMaker Python SDK [`TensorFlow`](https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/tensorflow/README.rst#training-with-tensorflow) estimator to easily train locally and in SageMaker.
#
# Let's start by setting the training script arguments `--num_epochs` and `--data_dir` as hyperparameters. Remember that we don't need to provide `--model_dir`:
# +
from sagemaker import get_execution_role
role = get_execution_role()
# -
hyperparameters = {'train_steps': 10, 'model_name': 'DeepFM'}
# This notebook shows how to use the SageMaker Python SDK to run your code in a local container before deploying to SageMaker's managed training or hosting environments. Just change your estimator's train_instance_type to local or local_gpu. For more information, see: https://github.com/aws/sagemaker-python-sdk#local-mode.
#
# In order to use this feature you'll need to install docker-compose (and nvidia-docker if training with a GPU). Running following script will install docker-compose or nvidia-docker-compose and configure the notebook environment for you.
#
# Note, you can only run a single local notebook at a time.
# !/bin/bash ./utils/setup.sh
# To train locally, you set `train_instance_type` to [local](https://github.com/aws/sagemaker-python-sdk#local-mode):
# +
import subprocess
train_instance_type='local'
if subprocess.call('nvidia-smi') == 0:
## Set type to GPU if one is present
train_instance_type = 'local_gpu'
print("Train instance type = " + train_instance_type)
# -
# We create the `TensorFlow` Estimator, passing the `git_config` argument and the flag `script_mode=True`. Note that we are using Git integration here, so `source_dir` should be a relative path inside the Git repo; otherwise it should be a relative or absolute local path. the `Tensorflow` Estimator is created as following:
#
# +
import os
import sagemaker
from sagemaker.tensorflow import TensorFlow
estimator = TensorFlow(entry_point='train_estimator.py',
source_dir='.',
train_instance_type=train_instance_type,
train_instance_count=1,
hyperparameters=hyperparameters,
role=role,
framework_version='2.2.0',
py_version='py37',
script_mode=True,
model_dir='/opt/ml/model')
# -
# To start a training job, we call `estimator.fit(inputs)`, where inputs is a dictionary where the keys, named **channels**,
# have values pointing to the data location. `estimator.fit(inputs)` downloads the TensorFlow container with TensorFlow Python 3, CPU version, locally and simulates a SageMaker training job.
# When training starts, the TensorFlow container executes **train.py**, passing `hyperparameters` and `model_dir` as script arguments, executing the example as follows:
# ```bash
# python -m train --num-epochs 1 --data_dir /opt/ml/input/data/training --model_dir /opt/ml/model
# ```
#
# +
inputs = {'training': f'file:///home/ec2-user/SageMaker/deepctr_sagemaker/data/'}
estimator.fit(inputs)
# -
# Let's explain the values of `--data_dir` and `--model_dir` with more details:
#
# - **/opt/ml/input/data/training** is the directory inside the container where the training data is downloaded. The data is downloaded to this folder because `training` is the channel name defined in ```estimator.fit({'training': inputs})```. See [training data](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html#your-algorithms-training-algo-running-container-trainingdata) for more information.
#
# - **/opt/ml/model** use this directory to save models, checkpoints, or any other data. Any data saved in this folder is saved in the S3 bucket defined for training. See [model data](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html#your-algorithms-training-algo-envvariables) for more information.
#
# ### Reading additional information from the container
#
# Often, a user script needs additional information from the container that is not available in ```hyperparameters```.
# SageMaker containers write this information as **environment variables** that are available inside the script.
#
# For example, the example above can read information about the `training` channel provided in the training job request by adding the environment variable `SM_CHANNEL_TRAINING` as the default value for the `--data_dir` argument:
#
# ```python
# if __name__ == '__main__':
# parser = argparse.ArgumentParser()
# # reads input channels training and testing from the environment variables
# parser.add_argument('--data_dir', type=str, default=os.environ['SM_CHANNEL_TRAINING'])
# ```
#
# Script mode displays the list of available environment variables in the training logs. You can find the [entire list here](https://github.com/aws/sagemaker-containers/blob/master/README.rst#list-of-provided-environment-variables-by-sagemaker-containers).
# # Training in SageMaker
# After you test the training job locally, upload the dataset to an S3 bucket so SageMaker can access the data during training:
# +
import sagemaker
inputs = sagemaker.Session().upload_data(path='/home/ec2-user/SageMaker/deepctr_sagemaker/data', key_prefix='DEMO-tensorflow-deepctr')
print(inputs)
# -
# The returned variable inputs above is a string with a S3 location which SageMaker Tranining has permissions
# to read data from.
# To train in SageMaker:
# - change the estimator argument `train_instance_type` to any SageMaker ml instance available for training.
# - set the `training` channel to a S3 location.
# +
estimator = TensorFlow(entry_point='train_estimator.py',
source_dir='.',
train_instance_type='ml.p3.2xlarge', # Executes training in a ml.p2.xlarge/ml.p3.2xlarge/ml.p3.8xlarge instance
train_instance_count=1,
hyperparameters=hyperparameters,
role=role,
framework_version='2.2.0',
py_version='py37',
script_mode=True,
model_dir='/opt/ml/model')
estimator.fit({'training': inputs})
# -
# ## Git Support
# +
git_config = {'repo': 'https://github.com/whn09/deepctr_sagemaker.git', 'branch': 'main'}
estimator = TensorFlow(entry_point='train.py',
source_dir='.',
git_config=git_config,
train_instance_type='ml.p3.2xlarge', # Executes training in a ml.p2.xlarge instance
train_instance_count=1,
hyperparameters=hyperparameters,
role=role,
framework_version='2.2.0',
py_version='py37',
script_mode=True,
model_dir='/opt/ml/model')
estimator.fit({'training': inputs})
# -
# ## Deploy the trained model to an endpoint
#
# The deploy() method creates a SageMaker model, which is then deployed to an endpoint to serve prediction requests in real time. We will use the TensorFlow Serving container for the endpoint, because we trained with script mode. This serving container runs an implementation of a web server that is compatible with SageMaker hosting protocol. The Using your own inference code document explains how SageMaker runs inference containers.
predictor = estimator.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
# ## Invoke the endpoint
#
# Let's download the training data and use that as input for inference.
# +
import json
def test_REST_serving():
'''
test rest api
'''
fea_dict1 = {'I1':[0.0],'I2':[0.001332],'I3':[0.092362],'I4':[0.0],'I5':[0.034825],'I6':[0.0],'I7':[0.0],'I8':[0.673468],'I9':[0.0],'I10':[0.0],'I11':[0.0],'I12':[0.0],'I13':[0.0],'C1':[0],'C2':[4],'C3':[96],'C4':[146],'C5':[1],'C6':[4],'C7':[163],'C8':[1],'C9':[1],'C10':[72],'C11':[117],'C12':[127],'C13':[157],'C14':[7],'C15':[127],'C16':[126],'C17':[8],'C18':[66],'C19':[0],'C20':[0],'C21':[3],'C22':[0],'C23':[1],'C24':[96],'C25':[0],'C26':[0]}
fea_dict2 = {'I1':[0.0],'I2':[0.0],'I3':[0.00675],'I4':[0.402298],'I5':[0.059628],'I6':[0.117284],'I7':[0.003322],'I8':[0.714284],'I9':[0.154739],'I10':[0.0],'I11':[0.03125],'I12':[0.0],'I13':[0.343137],'C1':[11],'C2':[1],'C3':[98],'C4':[98],'C5':[1],'C6':[6],'C7':[179],'C8':[0],'C9':[1],'C10':[89],'C11':[58],'C12':[97],'C13':[79],'C14':[7],'C15':[72],'C16':[26],'C17':[7],'C18':[52],'C19':[0],'C20':[0],'C21':[47],'C22':[0],'C23':[7],'C24':[112],'C25':[0],'C26':[0]}
fea_dict3 = {'I1':[0.0],'I2':[0.000333],'I3':[0.00071],'I4':[0.137931],'I5':[0.003968],'I6':[0.077873],'I7':[0.019934],'I8':[0.714284],'I9':[0.505803],'I10':[0.0],'I11':[0.09375],'I12':[0.0],'I13':[0.17647],'C1':[0],'C2':[18],'C3':[39],'C4':[52],'C5':[3],'C6':[4],'C7':[140],'C8':[2],'C9':[1],'C10':[93],'C11':[31],'C12':[122],'C13':[16],'C14':[7],'C15':[129],'C16':[97],'C17':[8],'C18':[49],'C19':[0],'C20':[0],'C21':[25],'C22':[0],'C23':[6],'C24':[53],'C25':[0],'C26':[0]}
fea_dict4 = {'I1':[0.0],'I2':[0.004664],'I3':[0.000355],'I4':[0.045977],'I5':[0.033185],'I6':[0.094967],'I7':[0.016611],'I8':[0.081632],'I9':[0.028046],'I10':[0.0],'I11':[0.0625],'I12':[0.0],'I13':[0.039216],'C1':[0],'C2':[45],'C3':[7],'C4':[117],'C5':[1],'C6':[0],'C7':[164],'C8':[1],'C9':[0],'C10':[20],'C11':[61],'C12':[104],'C13':[36],'C14':[1],'C15':[43],'C16':[43],'C17':[8],'C18':[37],'C19':[0],'C20':[0],'C21':[156],'C22':[0],'C23':[0],'C24':[32],'C25':[0],'C26':[0]}
fea_dict5 = {'I1':[0.0],'I2':[0.000333],'I3':[0.036945],'I4':[0.310344],'I5':[0.003922],'I6':[0.067426],'I7':[0.013289],'I8':[0.65306],'I9':[0.035783],'I10':[0.0],'I11':[0.03125],'I12':[0.0],'I13':[0.264706],'C1':[0],'C2':[11],'C3':[59],'C4':[77],'C5':[1],'C6':[5],'C7':[18],'C8':[1],'C9':[1],'C10':[45],'C11':[171],'C12':[162],'C13':[96],'C14':[4],'C15':[36],'C16':[121],'C17':[8],'C18':[14],'C19':[5],'C20':[3],'C21':[9],'C22':[0],'C23':[0],'C24':[5],'C25':[1],'C26':[47]}
data = {"instances": [fea_dict1,fea_dict2,fea_dict3,fea_dict4,fea_dict5]}
# print(data)
json_response = predictor.predict(data)
predictions = json_response['predictions']
# print(predictions)
return predictions
# -
test_REST_serving()
# ## Delete the endpoint
#
# Let's delete the endpoint we just created to prevent incurring any extra costs.
sagemaker.Session().delete_endpoint(predictor.endpoint)
| byos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit ('3.8.5')
# name: python385jvsc74a57bd038b4776f74ef7846326f679ebb3c9e296e2ab09fe7ffa3e73b643f9a7922e0a6
# ---
# +
import pandas as pd
url = "https://gist.githubusercontent.com/guilhermesilveira/2d2efa37d66b6c84a722ea627a897ced/raw/10968b997d885cbded1c92938c7a9912ba41c615/tracking.csv"
data = pd.read_csv(url)
x = data[['home', 'how_it_works', 'contact']]
y = data[['bought']]
# +
# Mannualy separate test/train data
import numpy as np
from sklearn.svm import LinearSVC
from sklearn.metrics import accuracy_score
x_train = x[:75]
y_train = y[:75]
x_test = x[75:]
y_test = y[75:]
print('Training with %d items & testing with %d items' % (len(x_train), len(x_test)))
model = LinearSVC()
model.fit(x_train, np.ravel(y_train))
predictions = model.predict(x_test)
accuracy = accuracy_score(y_test, predictions) * 100
print('Accuracy is %.2f%%' % accuracy)
# +
# Using SKLearn to separate test/train data
from sklearn.model_selection import train_test_split
SEED = 20
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.25, stratify=y, random_state=SEED)
print('Training with %d items & testing with %d items' % (len(x_train), len(x_test)))
model = LinearSVC()
model.fit(x_train, np.ravel(y_train))
predictions = model.predict(x_test)
accuracy = accuracy_score(y_test, predictions) * 100
print('Accuracy is %.2f%%' % accuracy)
# -
| website_tracking.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # ArrayList
// ## Specifications
// - All elements are of the same type
// - All elements are stored in adjacent memory locations
// - The element at index `i` is at memory address `x + bi` => we can access specific element we want very quickly: in O(1) time. (This is called **random access**)
//
// ```
// You create an array of integers (assume each integer is exactly 4 bytes) in memory, and the beginning of the array (i.e., the start of the very first cell of the array) happens to be at memory address 1000 (in decimal, not binary). What is the memory address of the start of cell 6 of the array, assuming 0-based indexing (i.e., cell 0 is the first cell of the array)?
// ```
//
// 0: 1000
//
// 1: 1004
//
// 2: 1008
//
// 3: 1012
//
// 4: 1016
//
// 5: 1020
//
// 6: `1024`
//
// ## Assumptions
// - we will assume that a user can only add elements to indices between 0 and n (inclusive), `n = num of total elements that exist in the list prior to the new insertion`
// - user can only add elements to the front or back of the array (push/pop)
//
// ## Dynamic ArrayList
// 1. allocate some default "large" amount of memory initially
// 2. insert elements into this initial array
// 3. once the array is full, they create a new larger array (typically twice as large as the old array),
// 4. copy all elements from the old array into the new array,
// 5. replace any references to the old array with references to the new array.
//
// In C++, this is `vector` equivalent.
//
// ## Thinking challenge
// ```
// Array structures (e.g. the array, or the Java ArrayList, or the C++ vector, etc.) require that all elements be the same size. However, array structures can contain strings, which can be different lengths (and thus different sizes in memory). How is this possible?
// ```
//
// Answer: store pointers to strings in an array. Pointers are of the same size anyways.
//
// ## Insertion
// - Worst case: `O(n)` = Inserting into the first index (You have to move up all other elements by one index)
// - Best case: `O(1)` = Inserting at the end
//
// ## Binary search in ArrayList
// **Everything is under the assumption that elements are sorted in advance**
//
// 1. Have the array sorted in advance
// 2. Compare the element in search against the middle element
// 3. If less than the middle element, repeat search on the left half of the array / If more, do it on the right half
//
// ## Removal
// - Best case: `O(1)` = remove the end
// - Worst case: `O(n)` = remove at the beginning (comes from our restriction that all elements in our array must be contiguous)
//
// ## Thinking challenge
// ```
// When we remove from the very beginning of the backing array of an Array List, even before we move the remaining elements to the left, the remaining elements are all still contiguous, so our restriction is satisfied. Can we do something clever with our implementation to avoid having to perform this move operation when we remove from the very beginning of our list?
// ```
//
// Answer: if we were to remove `arr[0]`, set index 1 as index 0, 2 as 1, 3 as 2... and n as n-1 and so on. Then it could be done in `O(1)`. In short: adjust indices.
// # LinkedList
// - Improved version of ArrayList for better time & space complexities
// - Uses nodes
// - Head & tail pointer
// - Singly-linked list: each node has 1 pointer towards the tail
// - Doubly-linked list: each node has 2 pointers back and forth
//
// ## `find`
// - Takes `O(n)`. Start from the head/tail.
//
// ```c++
// bool find(Node* node, int element) {
// while(true){
// if (node->value == element){
// return true;
// }
// else if(node->next == NULL){
// return false;
// }
// else{
// node = node->next;
// }
// }
// }
//
// ```
//
// ## `insert`
// - find the insertion site,
// - rearrange pointers to fit the new node in its rightful spot.
//
// ```c++
// void insert(Node* head, Node* newnode, int index) {
// Node* crnt = head;
// for(int i = 0; i < index - 1; i++){
// crnt = crnt->next;
// }
// Node* tmp = crnt->next;
// crnt->next = newnode;
// newnode->next = tmp;
// }
// ```
//
// ## `remove`
//
// ```c++
// void remove(Node* head, int index){
// if (--index == 0){
// // reached the target index
// head->next = head->next->next;
// }
// else{
// // not yet there
// remove(head->next, index);
// }
// }
// ```
//
// ## summary
// - add/remove: O(1)
// - find: O(n) always (Theta). Cannot do something like binary search.
// - no wasted memory, compared to ArrayList
//
// # Skip lists
//
// ## Review
//
// ### ArrayList
// - Worst case for `find`: `O(logn)` (for a sorted one)
// - Always, for `insert`/`delete`: `O(n)`.
//
// ### LinkedList
// - Worst case for `insert`/`remove` to front/back of the structure : `O(1)`
// - `find`: `O(n)`, because LinkedList does not have **random access** property.
//
// ## Then skip list?
// - Comprised of nodes, each of them containing 1 key and multiple pointers
// - Multiple layers. Each layer is a node with a forward pointer
// 
// 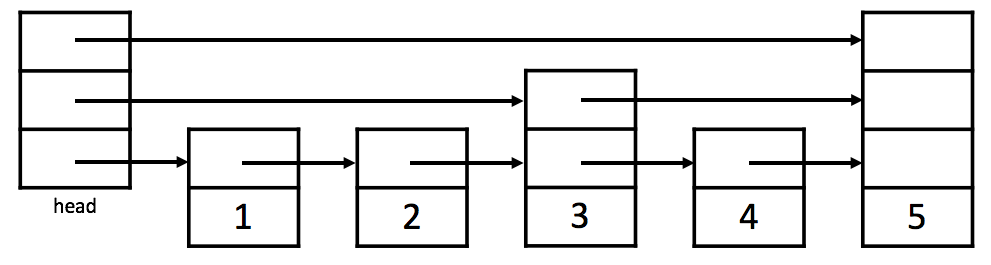
// - For each layer `i`, the `i`-th pointer in head points to the first node that has a height of `i`.
// - The head also has multiple layers.
//
// ## `find`
// 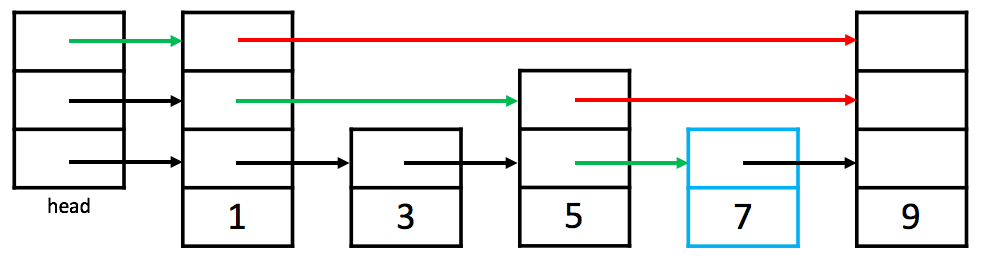
// - start at head
// - traverse the forward pointer
//
// ## `remove`
// 
// 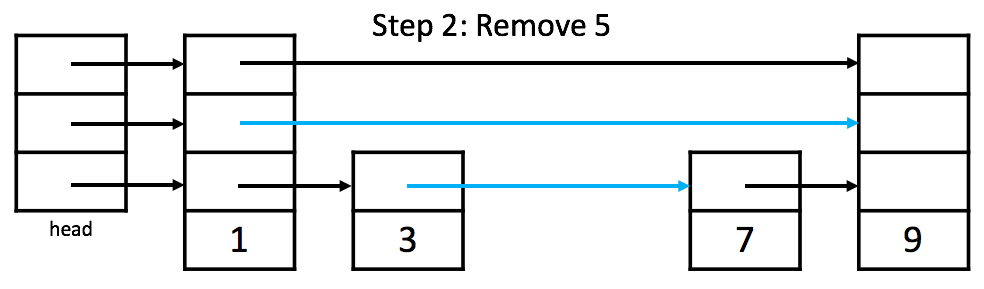
// - do `find`
// - rearrange pointers
//
// ## Time complexity for the worst-case
// 
// - worst-case: if the Skip List's node heights are all equal or descending
// - you can do no more than `n` times of efficient traversal.
// - worst-case time complexity for `find`/`remove`: O(n).
//
// ## Time complexity for the opitmally-distributed SkipList
//
// 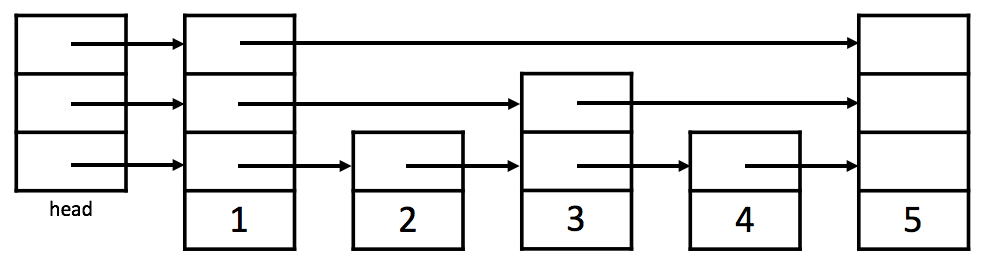
// - O(logn)
// - each "jump" allows you to traverse half of the remainder of the list
// - The size of the search space starts at n, but then you jump to the middle and cut the search space in half (n/2), then cut in half again (n/4), until eventually the search space becomes 1 (the element you want). The series n, n/2, n/4, ..., 2, 1 has O(log n) elements
// - **The distribution of heights really matters. With optimal distribution, it can perform O(logn), which is effectively the TC of binary search.**
//
// ## How to design an optiamlly distributed SkipList
//
// 1. Find where to put first
// 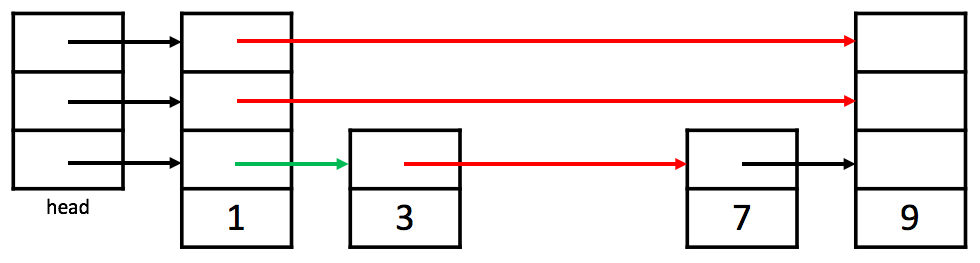
// 2. Determine the height of the node
// - start at height = 0
// - coin's probability of heads is p.
// - If we flip heads, we increase our height by 1.
// - If we flip tails, we stop playing the game and keep our current height.
//
// ## Why do we use the probability `p`?
// - we designate the new node's height from Bernoulli distribution, which means that we "randomly" sample from the distribution.
// - There is absolutely no guarantee of optimality in a randomized structure: the hope is that, on average, the results will be good
//
// ## No multiple coin-flips
// - we do not need to flip a coin every single time to increase/stop increasing the height.
// - coin-flip is a **Bernoulli distribution (or a binary distribution), which is just the formal name for a probability distribution in which we only have two possible outcomes: success and failure**.
// - you know how many times you will flip the coin. say, `k`.
// - then use **the formula for P(X = k) for geometric distribution, which is Bernoulli distribution done multiple times (n = k in the picture)**.
//
//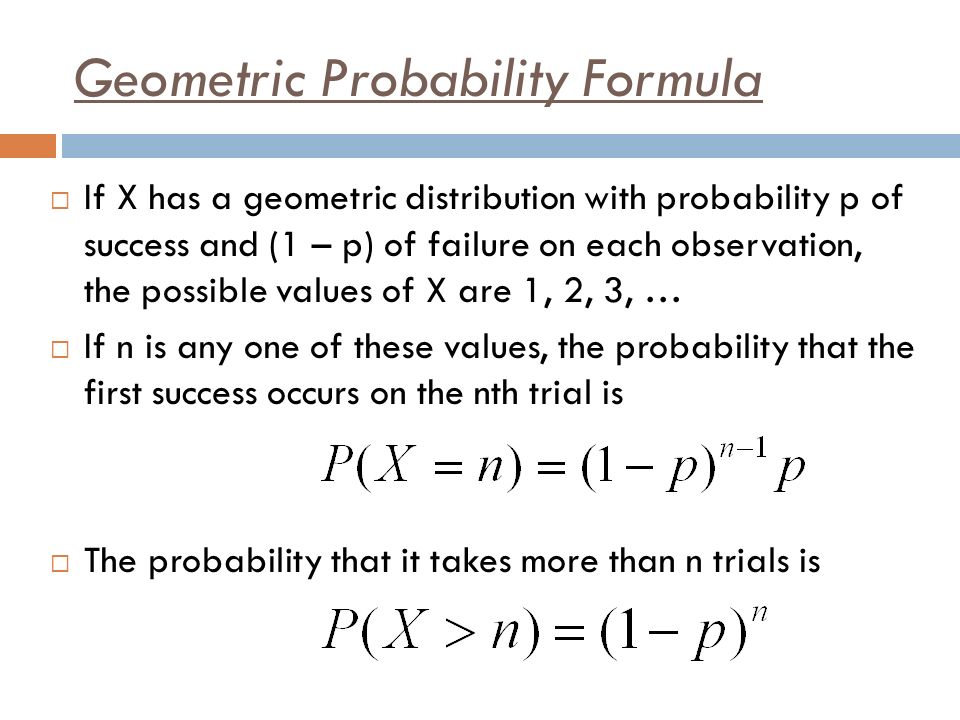 
//
// But, we want the number of flips until the first failure. So reverse it:
// $$P(X = k) = p^{k}(1-p)$$
// Note that **k = the number of coin flips just before the fisrt failure in this formula.**
//
// ## Problem
// ```
// To determine the heights of new nodes, you use a coin with a probability of success of p = 0.3. What is the probability that a new node will have a height of 0? (Enter your answer as a decimal rounded to the nearest thousandth)
// ```
// Answer: 1 - 0.3 = 0.7
//
// ## Problem 2
// ```
// To determine the heights of new nodes, you use a coin with a probability of success of p = 0.3. What is the probability that a new node will have a height of 2? (Enter your answer as a decimal rounded to the nearest thousandth)
// ```
//
// Answer: 0.3 * 0.3 * 0.7 (two "successes" and one "fail")
// Height 0: 0.3
// Height 1: 0.3 * 0.3
// Height 2: 0.3 * 0.3 * 0.7
//
// ## TC
// - Worst-case TC to find/insert/remove is O(n) (poorly distributed)
// - Average-case TC is O(logn) (optimally distributed). => Has a proof
// - Expected number of comparisons must be done in $1 + (1/p)log_{1/p}(n)+1/(1-p)$
//
// ## Finding a 'good' `p`
// 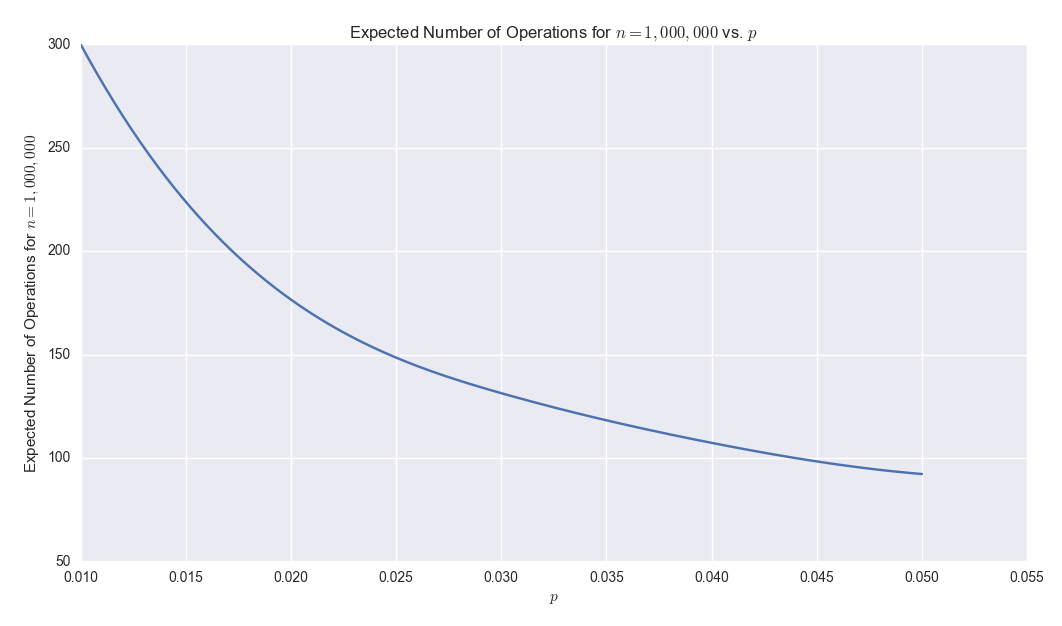
// - You already know how big `n` is roughly going to be
// - The curve begins to flatten at around p = 0.025, so that might be a good value to pick for p.
// - **Why should you do this? Because as p increases, the amount of space we need to use to store pointers also increases.**
//
// ## Finding a max height
// Once we've chosen a value for p, it can be formally proven (via a proof that is a bit out-of-scope) that, for good performance, the maximum height of the Skip List should be no smaller than **$log_{1/𝑝}n$**.
//
// ```
// Imagine you are implementing a Skip List, and you chose a value for p = 0.1. If you are expecting that you will be inserting roughly n = 1,000 elements, what should you pick as your Skip List's maximum height?
// ```
//
// $log_{1/0.1}1000 = log_{10}10^{3} = 3$
//
// ## Summary
// - SkipList works efficiently only with optimizations:
// - Choose a good `p` for a geometric probability distribution
// - Choose a good `height`
// - Otherwise, it would work just like LinkedList.
// # Circular array
// You want to only get the good parts from Array Lists and LinkedLists:
// - random access
// - inserting at the beginning and at the end in O(1) time
//
// A circular array is a:
// - Array list mimicking the behavior of a Linked List.
// - Array list that has head (first) and tail (last) indices
//
// ## More explained
// 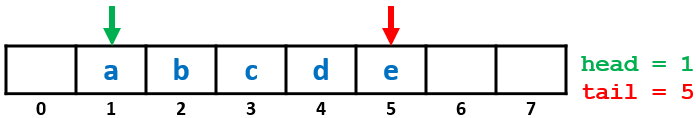
// - You only care about the head (1) & tail (5) indices
// - So you can represent the array like a circle as well:
// 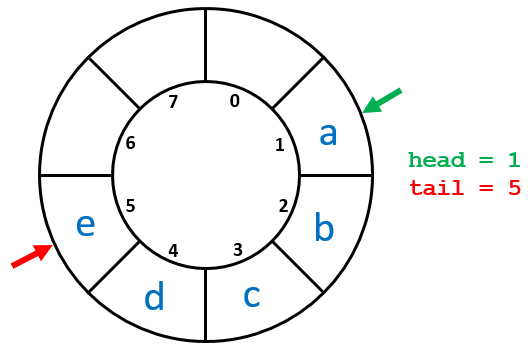
//
// Note: head index should be 'before' the tail index
// - i.e. (0,2) or (5, 7) or (6, 0), or (7, 1) in a circular array of capacity of 8 that contains 3 elements.
// - i.e. NOT (1,7) or (0,6)
//
// ## When the array becomes full
// - create a new backing array (typically of twice the size)
// - simply copy all elements from the old backing array into the new backing array.
// - To ensure the same order, let the elements in indices 0 through n-1 stay in the same indices in the new array.
//
// 
//
// ## Worst-case insertion at the front / back of a circular array
// - the backing array can be full
// - need to allocate a new backing array
// - copy all n elements from the old array to the new one
// - O(n)
//
// ## Worst-case insertion at the front / back of a circular array that's not full
// - trivial. O(1). Just insert at the empty index.
//
// ## Accessing elements in the middle
// - `(head + i) % array.length` would give you the correct index in the real array, where head = 0 and i = index given that head = 0.
// - For example, the element at i = 2 of our list is at index (7 + 2) % 8 = 9 % 8 = 1 of the backing array
// 
// # Abstract data types
// - we don't necessarily care about how the data structure executes these tasks: we just care that it gets the job done.
// - a model for data types where the **data type is defined by its behavior from the point of view of a user of the data** (i.e., by what functions the user claims it needs to have) is an Abstract Data Type (ADT).
//
// ## ADT vs Data structures
// - ADT: it cares about **what functions it should be able to perform, but it does not at all depend on how it actually goes about doing** those functions (i.e., it is not implementation-specific)
// - Data structure:
//
// ## Again explained
// - An Abstract Data Type does NOT contain details on how it should be implemented
// - An Abstract Data Type is designed from the perspective of a user, not an implementer
// - Any implementations of an Abstract Data Type have a strict set of functions they must support
// # Deque(Dequeue/Double-ended queue)
// - works pretty similar to browser history functions
//
// ## Deque ADT
//
// ```
// addFront(element): Add element to the front of the Deque
// addBack(element): Add element to the back of the Deque
// peekFront(): Look at the element at the front of the Deque
// peekBack(): Look at the element at the back of the Deque
// removeFront(): Remove the element at the front of the Deque
// removeBack(): Remove the element at the back of the Deque
// ```
//
// ## Which data structures to choose?
// - Doubly Linked List:
// - would do O(1) for all those ADTs listed
// - but accessing in the middle would require O(n)
// - Circular array:
// - find/remove: O(1)
// - insert: O(1) ~ O(n) (when the backing array is full)
// # Queues
| 2-Data-Structures-An-Active-Learning-Approach/4-Introductory-data-structures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### **Time-series Examples**
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
from src.helpers import plot_ts_and_save
data_path = os.path.join(os.getcwd(), 'data')
plot_path = os.path.join(os.getcwd(), 'plots')
data_files = ['friends_episodes_votes_ts.csv', 'retail_revenue_ts.csv',
'shampoo_ts.csv', 'temperature_ts.csv']
def load_ts_data(file_list, d_path):
"""loads csv datatsets from a list of filenames and a path.
args:
file_list: list of file names
path: data path string
"""
dfs = []
for f in file_list:
data_file_path = os.path.join(d_path, f)
print('Reading {}'.format(data_file_path))
df = pd.read_csv(data_file_path)
df.name = f[:-4]
dfs.append(df)
return dfs
# load data
friends_df, retail_df, shampoo_df, temp_df = load_ts_data(data_files, data_path)
# #### Shampoo Sales Time-Series
plot_ts_and_save(shampoo_df, 'Month', 'Sales', 'Shampoo Sales Time-Series', plot_path)
# #### Retail Revenue Time-Series
plot_ts_and_save(retail_df, 'Period', 'Revenue', 'Retail Revenue Time-Series', plot_path)
# #### Friends TV Time-Series Votes
friends_df['year_row_idx'] = friends_df.groupby(['Year_of_prod','Season']).cumcount()+1
friends_df['year_season_episode'] = friends_df['Year_of_prod'].astype(str) + friends_df['Season'].astype(str) + friends_df['year_row_idx'].astype(str)
plot_ts_and_save(friends_df, 'year_season_episode', 'Votes', 'Friends TV Series Votes Time-Series', plot_path)
# #### Temperature Time Series
plot_ts_and_save(temp_df, 'Date', 'Temp', 'Temperature Time-Series', plot_path)
| sample_time_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Pymaceuticals Observations and Insights
# Using python and its libraries, Scipy.stats, Mumpy, Pandas and Matplotlib, to analyze data for Pymaceuticals's Capomulin animal study.
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
from scipy.stats import linregress
# Study data files
mouse_metadata_path = "./Resources/Mouse_metadata.csv"
study_results_path = "./Resources/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# -
# Viewing data
mouse_metadata
# Viewing data
study_results
# Combine the data into a single dataset
mouse_study = pd.merge(mouse_metadata, study_results, how = "outer", on = "Mouse ID")
# Display the data table for preview
mouse_study
# Checking the number of mice.
len(mouse_study["Mouse ID"].unique())
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
# Source: https://stackoverflow.com/questions/14657241/how-do-i-get-a-list-of-all-the-duplicate-items-using-pandas-in-python
mouse_study[mouse_study[['Mouse ID', 'Timepoint']].duplicated()]
# +
# Optional: Get all the data for the duplicate mouse ID.
# -
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
mouse_study.drop((mouse_study[mouse_study['Mouse ID'] == 'g989'].index), inplace = True)
# Renaming dataframe
study_clean_data = mouse_study
study_clean_data
# Checking the number of mice in the clean DataFrame.
len(study_clean_data["Mouse ID"].unique())
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation,
# and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Assemble the resulting series into a single summary dataframe.
drug_groupby = study_clean_data.groupby('Drug Regimen')
study_mean_drug = drug_groupby['Tumor Volume (mm3)'].mean()
study_median_drug = drug_groupby['Tumor Volume (mm3)'].median()
study_variance_drug = drug_groupby['Tumor Volume (mm3)'].var()
study_standard_dev_drug = drug_groupby['Tumor Volume (mm3)'].std()
study_sem_drug = drug_groupby['Tumor Volume (mm3)'].sem()
# -
study_median_drug
# +
# Generate a summary statistics table of mean, median, variance, standard deviation,
# and SEM of the tumor volume for each regimen
summary_table = {
'Mean': study_mean_drug,
"Median": study_median_drug,
'Variance': study_variance_drug,
'Standard Deviation': study_standard_dev_drug,
'Standard Error of Mean(SEM)': study_sem_drug
}
summary_table
# Using the aggregation method, produce the same summary statistics in a single line
# -
# Making summary table a data frame
pd.DataFrame(summary_table)
# ## Bar and Pie Charts
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
drug_groupby = study_clean_data.groupby('Drug Regimen') # Grouping by Drug Regimen
study_mice_drug = drug_groupby['Mouse ID'].count() #Number of mice in each regimen
drug_mice_ct = study_mice_drug.to_frame() # Converting to dataframe
drug_mice_ct = drug_mice_ct.rename(columns = {"Mouse ID": 'Count of Mice'}) # Renaming columns
drug_mice_ct = drug_mice_ct.sort_values(by=['Count of Mice']) # Sorting by count
# Bar Chart
drug_mice_ct.plot(kind ='bar', ylabel = 'Number of Mice Tested', legend = False)
# Saving Bar Chart
plt.savefig("./Charts/Number of Mice Tested.png")
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
x_axis = np.arange(len(drug_mice_ct)) # Determine number of bars needed
tick_locations = [value for value in x_axis] # to tell the plot where to place tick marks
# Bar Chart
plt.bar(x_axis, drug_mice_ct["Count of Mice"], color='r', alpha=0.5, align="center")
plt.xticks(tick_locations, drug_mice_ct.index.values, rotation="vertical")
# +
# Generate a pie plot showing the distribution of female versus male mice using pandas
drug_groupby_gender = study_clean_data.groupby('Sex') # Grouping by sex
study_mice_gender = drug_groupby_gender['Mouse ID'].count() #Determine number in each group
gender_mice_ct = study_mice_gender.to_frame() # Converting to dataframe
gender_mice_ct = gender_mice_ct.rename(columns = {"Mouse ID": 'Count of Mice'}) #Renaming columns
gender_mice_ct = gender_mice_ct.sort_values(by=['Count of Mice']) # Sorting Values
# Creating pie chart using pandas
gender_mice_ct.plot(y='Count of Mice', kind ='pie', autopct="%1.1f%%", colors = ('coral', 'seagreen'),
ylabel = 'Sex', title = 'Mice Gender Distribution', legend = False)
# Saving distribution
plt.savefig("./Charts/Mice Gender Distribution.png")
# -
# Generate a pie plot showing the distribution of female versus male mice using pyplot
plt.pie(gender_mice_ct['Count of Mice'], labels = ['Female', 'Male'], autopct="%1.1f%%", colors = ('coral', 'seagreen'))
plt.title("Mice Gender Distribution")
plt.ylabel("Sex")
# Saving distribution
plt.savefig("./Charts/Mice Gender Distribution(2).png")
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# Start by getting the last (greatest) timepoint for each mouse
groupby_drug_id = study_clean_data.groupby(['Drug Regimen', 'Mouse ID'])
study_last_timepoint = groupby_drug_id['Timepoint'].max()
last_timepoint = pd.DataFrame(study_last_timepoint)
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
last_timepoint = pd.merge(last_timepoint, study_clean_data, how = 'left', on = ['Drug Regimen', 'Mouse ID', 'Timepoint'])
# -
# Merging to create a dataframe with last timepoint and final tumor volume
study_data_last_time = pd.merge(study_clean_data, last_timepoint, how = 'outer', on = ['Drug Regimen', 'Mouse ID'], suffixes = ('_original', '_final'))
study_data_last_time
# Removing duplicate/unneeded columns from merge
study_data_last_time = study_data_last_time.drop(columns = ['Sex_final', 'Age_months_final', 'Weight (g)_final'])
# +
# Put treatments into a list for for loop (and later for plot labels)
treatments = study_data_last_time['Drug Regimen'].unique()
treatments
# Create empty list to fill with tumor vol data (for plotting)
tumor_vol = []
# Calculate the IQR and quantitatively determine if there are any potential outliers.
outliers = []
for regimen in treatments:
# Locate the rows which contain mice on each drug and get the tumor volumes
regimen_df = last_timepoint.loc[last_timepoint['Drug Regimen'] == regimen]
# add subset
quartiles = regimen_df['Tumor Volume (mm3)'].quantile([.25,.5,.75])
# Determine outliers using upper and lower bounds
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
outlier_occupancy = regimen_df.loc[(regimen_df['Tumor Volume (mm3)'] < lower_bound) | (regimen_df['Tumor Volume (mm3)'] > upper_bound)]
outliers.append({regimen: outlier_occupancy['Mouse ID'].count()})
# Print list of drug regimen with potential outliers
print("List of drug regimen's and the number of potential outliers:")
# zip source: https://stackoverflow.com/questions/1663807/how-to-iterate-through-two-lists-in-parallel
for (regimen, row) in zip(treatments, outliers):
if row[regimen] > 0:
print(f"{regimen}'s has {row[regimen]} potential outlier(s).")
# -
# To see list of treatments
treatments
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
treatment_choices = ['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin'] # Drug choices
tumor_vol = [] # For tumor volume in for loop
data = [] # To save data for each regimen
#Setting up plot
fig1, ax1 = plt.subplots()
ax1.set_title('Final Tumor Volume (Capomulin, Ramicane, Infubinol, Ceftamin)')
ax1.set_ylabel('Tumor Volume (mm3)')
for regimen in treatment_choices:
regimen_df = last_timepoint.loc[last_timepoint['Drug Regimen'] == regimen]
tumor_vol.append(regimen_df['Tumor Volume (mm3)'])
tum = tumor_vol
data.append(tum)
tumor_vol = []
# Extracting from list fro boxplot
x1 = data[0]
x2 = data[1]
x3 = data[2]
x4 = data[3]
# Convert to array source: https://www.educative.io/edpresso/how-to-convert-a-list-to-an-array-in-python
ax1.boxplot([np.array(x1[0]), np.array(x2[0]), np.array(x3[0]), np.array(x4[0])], labels = treatment_choices)
# Saving figure
plt.savefig("./Charts/Final Tumor Volume.png")
# Showing image
plt.show()
# -
# ## Line and Scatter Plots
# Capomulin mice ID, this give a list of options for the next cell
capomulin = study_clean_data.loc[study_clean_data["Drug Regimen"]== "Capomulin"]
capomulin["Mouse ID"].unique()
# +
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
#Choose mouse from list above
mouse_choice = input("What Capomulin mouse are you looking for? (List above) " )
tumor_vol = study_clean_data.loc[study_clean_data["Mouse ID"]== mouse_choice, ["Tumor Volume (mm3)", "Timepoint"]]
# +
# Plotting line with above mouse
y_values = tumor_vol["Tumor Volume (mm3)"]
timepoints = tumor_vol["Timepoint"]
plt.plot(timepoints, y_values, color="green")
plt.title("Capomulin treatment of mouse " + mouse_choice)
plt.xlabel("Timepoints")
plt.ylabel("Tumor Volume (mm3)")
# saving image
plt.savefig("./Charts/Tumor Volume over time for mouse of choice.png")
# +
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
mouse_group = capomulin.groupby('Mouse ID')
x_values = mouse_group['Weight (g)'].mean()
y_values = mouse_group['Tumor Volume (mm3)'].mean()
plt.scatter(x_values,y_values)
plt.xlabel('Weight (g)')
plt.ylabel('Tumor Volume (mm3)')
# Saving image
plt.savefig("./Charts/Scatter Plot_Avg Tumor Volume vs. Mouse Weight.png")
#Showing plot
plt.show()
# -
# ## Correlation and Regression
# +
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
mouse_group = capomulin.groupby('Mouse ID')
x_values = mouse_group['Weight (g)'].mean()
y_values = mouse_group['Tumor Volume (mm3)'].mean()
# linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
# creating equation
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plotting
plt.scatter(x_values,y_values) #Scatterplot
plt.plot(x_values,regress_values,"r-") #regression
plt.annotate(line_eq,(20,35),fontsize=15,color="red") #printing equation
plt.xlabel('Weight (g)')
plt.ylabel('Tumor Volume (mm3)')
#Correlation
print(f"The correlation between mouse weight and the average tumor volume is {'{:,.2f}'.format(rvalue)}")
# Saving image
plt.savefig("./Charts/Correlation Linear Reg.png")
#Showing plot
plt.show()
# -
| pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="JSjG64ra4aFu" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="eba951e4-6056-4433-81d3-dd26de83caea"
from google.colab import drive
drive.mount('/content/drive')
# + colab_type="code" id="V8-7SARDZErK" colab={}
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.optim as optim
from matplotlib import pyplot as plt
import copy
import pickle
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
# + id="dNZo88NV5ZUO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="98ab3307-0c6a-435e-ef3a-efd5915b09e2"
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
# + id="tZSGIGxk5ZUX" colab_type="code" colab={}
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
# + id="jtdsP4Lq5ZUc" colab_type="code" colab={}
dataiter = iter(trainloader)
background_data_train=[]
background_label_train=[]
foreground_data_train=[]
foreground_label_train=[]
batch_size=10
for i in range(5000): #5000*batch_size = 50000 data points
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data_train.append(img)
background_label_train.append(labels[j])
else:
img = images[j].tolist()
foreground_data_train.append(img)
foreground_label_train.append(labels[j])
foreground_data_train = torch.tensor(foreground_data_train)
foreground_label_train = torch.tensor(foreground_label_train)
background_data_train = torch.tensor(background_data_train)
background_label_train = torch.tensor(background_label_train)
# + id="y7eYWwWUTnS8" colab_type="code" colab={}
dataiter = iter(testloader)
background_data_test=[]
background_label_test=[]
foreground_data_test=[]
foreground_label_test=[]
batch_size=10
for i in range(1000): #1000*batch_size = 10000 data points
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data_test.append(img)
background_label_test.append(labels[j])
else:
img = images[j].tolist()
foreground_data_test.append(img)
foreground_label_test.append(labels[j])
foreground_data_test = torch.tensor(foreground_data_test)
foreground_label_test = torch.tensor(foreground_label_test)
background_data_test = torch.tensor(background_data_test)
background_label_test = torch.tensor(background_label_test)
# + id="4sasFFybUPOS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="f68a1f6e-fde6-453a-d356-36660dfd0e8b"
print(foreground_data_train.size())
print(foreground_data_test.size())
# + id="SLR6EvK5DqGC" colab_type="code" colab={}
foreground_data_combined = torch.cat([foreground_data_train , foreground_data_test], dim=0)
foreground_label_combined = torch.cat([foreground_label_train , foreground_label_test], dim=0)
background_data_combined = torch.cat([background_data_train , background_data_test], dim=0)
background_label_combined = torch.cat([background_label_train , background_label_test], dim=0)
# + id="XX8KV_9g_SY-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="fc40e5c1-b6b6-4146-c2c1-9e4068478f85"
print(foreground_data_train.size() , foreground_data_test.size() , foreground_data_combined.size())
print(foreground_label_train.size() , foreground_label_test.size() , foreground_label_combined.size())
print(background_data_train.size() , background_data_test.size() , background_data_combined.size())
print(background_label_train.size() , background_label_test.size() , background_label_combined.size())
# + id="oZa2mMli5ZUt" colab_type="code" colab={}
def create_mosaic_img(background_data, foreground_data, foreground_label, bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx] #-7 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
# + id="ZMSidmeagYPV" colab_type="code" colab={}
def init_mosaic_creation(bg_size, fg_size, desired_num, background_data, foreground_data, foreground_label):
# bg_size = 35000
# fg_size = 15000
# desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,bg_size,8)
fg_idx = np.random.randint(0,fg_size)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(background_data, foreground_data, foreground_label ,bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
return mosaic_list_of_images, mosaic_label, fore_idx
# + id="KBymhsvlhstt" colab_type="code" colab={}
mosaic_list_of_images, mosaic_label, fore_idx = init_mosaic_creation(bg_size = 42000,
fg_size = 15000,
desired_num = 30000,
background_data = background_data_combined,
foreground_data = foreground_data_train,
foreground_label = foreground_label_train)
# + id="_W4WPsAzBcgM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="11f89b3d-28d6-423a-c897-2a735da9e68d"
print(len(mosaic_list_of_images),len(mosaic_label))
# + colab_type="code" id="nSO9SFE25Lrk" colab={}
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
# + id="F71gWrhugFUO" colab_type="code" colab={}
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
# + id="7KSYzkDitY9H" colab_type="code" colab={}
# model initialisation
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0)
self.conv4 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv6 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32)
self.batch_norm2 = nn.BatchNorm2d(128)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(128,64)
self.fc2 = nn.Linear(64, 32)
self.fc3 = nn.Linear(32, 10)
self.fc4 = nn.Linear(10, 1)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv6(x)))
x = self.pool(x)
x = x.view(x.size(0), -1)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.dropout2(x)
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
# + id="aexGRZ2svm-3" colab_type="code" colab={}
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.module1 = Focus().double()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0)
self.conv4 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv6 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32)
self.batch_norm2 = nn.BatchNorm2d(128)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(128,64)
self.fc2 = nn.Linear(64, 32)
self.fc3 = nn.Linear(32, 10)
self.fc4 = nn.Linear(10, 3)
def forward(self,z): #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
x = x.to("cuda")
y = y.to("cuda")
for i in range(9):
x[:,i] = self.module1.forward(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
y1 = self.conv1(y)
y1 = F.relu(self.batch_norm1(y1))
y1 = (F.relu(self.conv2(y1)))
y1 = self.pool(y1)
y1 = self.conv3(y1)
y1 = F.relu(self.batch_norm2(y1))
y1 = (F.relu(self.conv4(y1)))
y1 = self.pool(y1)
y1 = self.dropout1(y1)
y1 = self.conv5(y1)
y1 = F.relu(self.batch_norm2(y1))
y1 = (F.relu(self.conv6(y1)))
y1 = self.pool(y1)
y1 = y1.view(y1.size(0), -1)
y1 = self.dropout2(y1)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = self.dropout2(y1)
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1 , x, y
# + id="bCXAyn9NvqV4" colab_type="code" colab={}
classify = Classification().double()
classify = classify.to("cuda")
# + id="bpaPbDFsvsZe" colab_type="code" colab={}
import torch.optim as optim
criterion_classify = nn.CrossEntropyLoss()
optimizer_classify = optim.SGD(classify.parameters(), lr=0.01, momentum=0.9)
# + id="YqZqomaQtUU_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6ff35dbd-d17a-4797-91ac-d5d93b825d6e"
# train
nos_epochs = 300
loss_list = []
for epoch in range(nos_epochs): # loop over the dataset multiple times
running_loss = 0.0
epoch_loss = []
cnt=0
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_classify.zero_grad()
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion_classify(outputs, labels)
loss.backward()
optimizer_classify.step()
running_loss += loss.item()
mini = 40
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
loss_list.append(np.mean(epoch_loss))
if(np.mean(epoch_loss) <= 0.03):
break;
print('Finished Training')
# + id="6fQIyZlgyvqA" colab_type="code" colab={}
name = "train_ds3"
# + id="uzDugMRftUyS" colab_type="code" colab={}
# save model
torch.save(classify.state_dict(),"/content/drive/My Drive/Research/genralisation_on_unseen_CIFAR/"+name+".pt")
# + id="qVcTZ1RQzTqy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="2c98ca47-03a1-4386-d734-bafe80f78b4f"
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="lSntUxwSgHNs" colab_type="code" colab={}
mosaic_list_of_images_1, mosaic_label_1, fore_idx_1 = init_mosaic_creation(bg_size = 35000,
fg_size = 15000,
desired_num = 10000,
background_data = background_data_train,
foreground_data = foreground_data_train,
foreground_label = foreground_label_train)
msd1 = MosaicDataset(mosaic_list_of_images_1, mosaic_label_1 , fore_idx_1)
test_loader_1 = DataLoader( msd1, batch_size= batch ,shuffle = False)
# + id="gsRII_ZuzjvJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="d7d26450-3730-495f-a632-5a5fab39fbf3"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_1:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="pmBGlzdRugSn" colab_type="code" colab={}
del test_loader_1
del msd1
del mosaic_list_of_images_1
del mosaic_label_1
del fore_idx_1
# + id="1YsXLEKBjyLf" colab_type="code" colab={}
mosaic_list_of_images_2, mosaic_label_2, fore_idx_2 = init_mosaic_creation(bg_size = 7000,
fg_size = 15000,
desired_num = 10000,
background_data = background_data_test,
foreground_data = foreground_data_train,
foreground_label = foreground_label_train)
msd2 = MosaicDataset(mosaic_list_of_images_2, mosaic_label_2 , fore_idx_2)
test_loader_2 = DataLoader( msd2, batch_size= batch ,shuffle = False)
# + id="L8Z7CSKM0VBB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="d0ff2313-cd58-422f-8d1f-ebc7a06ecd09"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_2:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="AJFJhvNR0tT6" colab_type="code" colab={}
del test_loader_2
del msd2
del mosaic_list_of_images_2
del mosaic_label_2
del fore_idx_2
# + id="jb3FD-4Sohhq" colab_type="code" colab={}
mosaic_list_of_images_3, mosaic_label_3, fore_idx_3 = init_mosaic_creation(bg_size = 42000,
fg_size = 15000,
desired_num = 10000,
background_data = background_data_combined,
foreground_data = foreground_data_train,
foreground_label = foreground_label_train)
msd3 = MosaicDataset(mosaic_list_of_images_3, mosaic_label_3 , fore_idx_3)
test_loader_3 = DataLoader( msd3, batch_size= batch ,shuffle = False)
# + id="e7CPi2wT0Xd7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="24f2f936-1595-4170-bce6-b79958ab8d74"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_3:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="zPfZgbTR00w4" colab_type="code" colab={}
del test_loader_3
del msd3
del mosaic_list_of_images_3
del mosaic_label_3
del fore_idx_3
# + colab_type="code" id="ywQU0yG2o3T_" colab={}
mosaic_list_of_images_4, mosaic_label_4, fore_idx_4 = init_mosaic_creation(bg_size = 35000,
fg_size = 3000,
desired_num = 10000,
background_data = background_data_train,
foreground_data = foreground_data_test,
foreground_label = foreground_label_test)
msd4 = MosaicDataset(mosaic_list_of_images_4, mosaic_label_4 , fore_idx_4)
test_loader_4 = DataLoader( msd4, batch_size= batch ,shuffle = False)
# + id="DLeW4vdv0Y7T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="0b2e3ecb-7592-46b7-8b83-d35cececfbe1"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_4:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="qVFYNQc104dS" colab_type="code" colab={}
del test_loader_4
del msd4
del mosaic_list_of_images_4
del mosaic_label_4
del fore_idx_4
# + colab_type="code" id="aK2aYdLdo3UK" colab={}
mosaic_list_of_images_5, mosaic_label_5, fore_idx_5 = init_mosaic_creation(bg_size = 7000,
fg_size = 3000,
desired_num = 10000,
background_data = background_data_test,
foreground_data = foreground_data_test,
foreground_label = foreground_label_test)
msd5 = MosaicDataset(mosaic_list_of_images_5, mosaic_label_5 , fore_idx_5)
test_loader_5 = DataLoader( msd5, batch_size= batch ,shuffle = False)
# + id="5gcb6dgr0bzq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="d89a58a2-69fb-4146-f074-dc51cb3a1f7b"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_5:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="4oHtjoe808QI" colab_type="code" colab={}
del test_loader_5
del msd5
del mosaic_list_of_images_5
del mosaic_label_5
del fore_idx_5
# + colab_type="code" id="ACIcvFpso3UT" colab={}
mosaic_list_of_images_6, mosaic_label_6, fore_idx_6 = init_mosaic_creation(bg_size = 42000,
fg_size = 3000,
desired_num = 10000,
background_data = background_data_combined,
foreground_data = foreground_data_test,
foreground_label = foreground_label_test)
msd6 = MosaicDataset(mosaic_list_of_images_6, mosaic_label_6 , fore_idx_6)
test_loader_6 = DataLoader( msd6, batch_size= batch ,shuffle = False)
# + id="H4rozEI40dbI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="3d2981a2-f98a-44b8-c89a-12f9ec475bc2"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_6:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="AX4uG3Sn1A5p" colab_type="code" colab={}
del test_loader_6
del msd6
del mosaic_list_of_images_6
del mosaic_label_6
del fore_idx_6
# + colab_type="code" id="gLt4xNT_q_Y2" colab={}
mosaic_list_of_images_7, mosaic_label_7, fore_idx_7 = init_mosaic_creation(bg_size = 35000,
fg_size = 18000,
desired_num = 10000,
background_data = background_data_train,
foreground_data = foreground_data_combined,
foreground_label = foreground_label_combined)
msd7 = MosaicDataset(mosaic_list_of_images_7, mosaic_label_7 , fore_idx_7)
test_loader_7 = DataLoader( msd7, batch_size= batch ,shuffle = False)
# + id="brH70rlP0hWi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="e9c04a20-ea42-453b-e921-384307f49372"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_7:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="A6owhcwU1HVq" colab_type="code" colab={}
del test_loader_7
del msd7
del mosaic_list_of_images_7
del mosaic_label_7
del fore_idx_7
# + colab_type="code" id="OfnNbZqfq_ZT" colab={}
mosaic_list_of_images_8, mosaic_label_8, fore_idx_8 = init_mosaic_creation(bg_size = 7000,
fg_size = 18000,
desired_num = 10000,
background_data = background_data_test,
foreground_data = foreground_data_combined,
foreground_label = foreground_label_combined)
msd8 = MosaicDataset(mosaic_list_of_images_8, mosaic_label_8 , fore_idx_8)
test_loader_8 = DataLoader( msd8, batch_size= batch ,shuffle = False)
# + id="nzFo1vH30k_1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="d31f3c11-603e-465d-cfe9-1ac3342bc12c"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_8:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="CEneUJNd1LQy" colab_type="code" colab={}
del test_loader_8
del msd8
del mosaic_list_of_images_8
del mosaic_label_8
del fore_idx_8
# + colab_type="code" id="CBLBjzAJq_Zc" colab={}
mosaic_list_of_images_9, mosaic_label_9, fore_idx_9 = init_mosaic_creation(bg_size = 42000,
fg_size = 18000,
desired_num = 10000,
background_data = background_data_combined,
foreground_data = foreground_data_combined,
foreground_label = foreground_label_combined)
msd9 = MosaicDataset(mosaic_list_of_images_9, mosaic_label_9 , fore_idx_9)
test_loader_9 = DataLoader( msd9, batch_size= batch ,shuffle = False)
# + id="626PTbsW0moI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="c863af78-f3fd-4528-91b0-765b7edf3066"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader_9:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="HHlKwKma1PsS" colab_type="code" colab={}
del test_loader_9
del msd9
del mosaic_list_of_images_9
del mosaic_label_9
del fore_idx_9
# + id="hzcMFWrB5ZWT" colab_type="code" colab={}
# %matplotlib inline
# + id="nivyY0qX5ZWZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="3d5e89ef-cf22-4549-eaf1-75b6eb0efab1"
plt.plot(loss_list)
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="Pex6d6CO5ZWf" colab_type="code" colab={}
| 1_mosaic_data_attention_experiments/9_generalisation_on_unseen_CIFAR10/train_datasets_codes/train_ds3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Code Synonyms
#
# This is useful for when you want to highlight two languages such as `python` and `ipython`
# syntax but want each to be executable within the same document.
#
# This can be done in the `conf.py` file through the `jupyter_lang_synonyms = ["ipython"]` option
import numpy as np
a = np.random.rand(10000)
# and now an ipython block
%%file us_cities.txt
new york: 8244910
los angeles: 3819702
chicago: 2707120
houston: 2145146
philadelphia: 1536471
phoenix: 1469471
san antonio: 1359758
san diego: 1326179
dallas: 1223229
# this will generate a file `us_cities.txt` in the local directory using an ipython
# cellmagic
| tests/base/ipynb/code_synonyms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pre-Processing Flow
# How to run scripts in "Plotly-Dash\stable_scripts\From_SD_to_CSV"
#
# 1. 1_SD_pandas_parse_.py
# - concats ".csv" files in JSON format from SD cards into one CSV
# -
# 2. 2_cbas_post_SD_resample.py
# - Filters outliers using "filterCriteria_0th"
# - fill NAs
# - Resamples by sample rate
# 3. 3_extradata.py
# - merges observational notes from Gsheet
# - ffill/bbfil
#
# 4. comfort\generateComfortMetrics.py
# -
#
# ##### Notes
# Timestamps remain in GMT for all scripts, only converted if plotting
#
# ## Usage
# ### 1_SD_pandas_parse_.py
# Location: stable_scripts\From_SD_to_CSV\1_SD_pandas_parse_.py
# ##### usage
# Run this in terminal **"python 1_SD_pandas_parse_.py [Board]**
# EX: "1_SD_pandas_parse_.py protoCBAS-A"
# > Currently available board names
# protoCBAS-A
# protoCBAS-B
# protoCBAS-C
# protoCBAS-D
# protoCBAS-G
#
# Input dir: (base_path, "CSV", "3Raw", "From_SDcard",board)
#
# Output dir: (base_path, "CSV", "2Interim", "1_SD_pandas_parse_")
#
# output columns
#
# ### 2_cbas_post_SD_resample.py
# Location: stable_scripts\From_SD_to_CSV\2_cbas_post_SD_resample.py
# ##### usage
# Run this in terminal **"python 2_cbas_post_SD_resample.py [sample rate]**
#
# EX: "2_cbas_post_SD_resample.py 5T" Resample at 5 min Interval
#
#
# ###### sample rates
# |Alias |Description|
# |------|------|
# |B |business day frequency|
# |C |custom business day frequency|
# |D |calendar day frequency|
# |W |weekly frequency|
# |M |month end frequency|
# |SM |semi-month end frequency| (15th and end of month)
# |BM |business month end frequency|
# |CBM |custom business month end frequency|
# |MS |month start frequency|
# |SMS |semi-month start frequency| (1st and 15th)
# |BMS |business month start frequency|
# |CBMS |custom business month start frequency|
# |Q |quarter end frequency|
# |BQ |business quarter end frequency|
# |QS |quarter start frequency|
# |BQS |business quarter start frequency|
# |A, Y |year end frequency|
# |BA, BY |business year end frequency|
# |AS, YS |year start frequency|
# |BAS, BYS |business year start frequency|
# |BH |business hour frequency|
# |H |hourly frequency|
# |T, min |minutely frequency|
# |S |secondly frequency|
# |L, ms |milliseconds
# |U, us |microseconds
# |N |nanoseconds|
#
# Input dir: (base_path, "CSV", "2Interim", "1_SD_pandas_parse_")
#
# samprt_path = 'resampled('+samprt+')'
# Output dir: (base_path,"CSV","2Interim","2_cbas_post_SD_resample",samprt_path)
#
# ### 3_extradata.py
# Location: stable_scripts\From_SD_to_CSV\3_extradata.py
# ##### usage
# Run this in terminal **"python 3_extradata.py [sample rate]**
# No arguments needed
#
#
# Input dir: (base_path, "CSV", "2Interim", "2_cbas_post_SD_resample","resampled(" + samprt + ")")
#
# Output dir: (base_path, "CSV", "2Interim", "3_extradata")
#
# Data Directory Structure
# -----------------
# ```
# `
# ├── CSV
# │ ├── 1processed <- Final data sets.
# ├── 1234 <- Folder labled by processes run on data.
# │ ├── 2interim <- Intermediate data transformed by scrpits.
# │ ├── 1_SD_pandas_parse_
# │ ├── 2_cbas_post_SD_resample
# │ ├── 3_extradata
# │ ├── 4_generateComfortMetrics
# │ ├── 3raw <- SDcard dumps and ingestion script output.
# │ ├── From_SDcard
# │ ├── ingestions_local <- from local machines
# │ ├── ingestions_rasp <- from RasperryPi (replaced w VM)
# │ ├── ingestions_VM <- from Gcloud VM
# `
# ```
#
# %run C:/Users/samgt/Documents/GitHub/Plotly-Dash/stable_scripts/From_SD_to_CSV/1_SD_pandas_parse_.py protoCBAS-A
# %run C:/Users/samgt/Documents/GitHub/Plotly-Dash/stable_scripts/From_SD_to_CSV/1_SD_pandas_parse_.py protoCBAS-B
# %run C:/Users/samgt/Documents/GitHub/Plotly-Dash/stable_scripts/From_SD_to_CSV/1_SD_pandas_parse_.py protoCBAS-C
# %run C:/Users/samgt/Documents/GitHub/Plotly-Dash/stable_scripts/From_SD_to_CSV/1_SD_pandas_parse_.py protoCBAS-D
# %run C:/Users/samgt/Documents/GitHub/Plotly-Dash/stable_scripts/From_SD_to_CSV/1_SD_pandas_parse_.py protoCBAS-G
| Plotly_dash/stable_scripts/from_SD_to_CSV/.ipynb_checkpoints/Pre-processing Instructions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Importing Numpy package
import numpy as np
#Check if a value is scalar
np.isscalar(23)
np.isscalar(False) # Boolean value is also scalar
np.isscalar([23]) # as it is a list it is not considered as scalar value
np.isscalar(45.6)
# +
# Scalar Computations
s1 = 6
s2 = 4
print(s1+s2)
print(s1-s2)
print(s1*s2)
print(s1/s2)
print(s1//s2)
# -
#Vectors
a = [1,2,3]
a * 2
#Multiplying scalar to a vector
np.multiply(a,2)
np.array(a) * 2
#Adding scalar to a vector
np.add(a,2)
#Multiple vectors
a = [10,15,18]
b = [11,23,42]
a + b
#Vector Addition
np.add(a,b)
#Vector Subtraction
np.subtract(a,b)
#Vector Addition
a = np.array([10,15,18])
b = np.array([11,23,42])
a + b
#Multiply a scalar with a vector
np.multiply(a,3)
# Dot product of two vectors
np.dot(a,b)
# Cross product of two vectors
np.cross(a,b)
#Finding out the magnitude of a vector
np.linalg.norm(a)
| NumPy - Scalars and Vectors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + slideshow={"slide_type": "skip"}
"""
IPython Notebook v4.0 para python 2.7
Librerías adicionales: numpy, matplotlib
Contenido bajo licencia CC-BY 4.0. Código bajo licencia MIT. (c) <NAME>.
"""
# Configuracion para recargar módulos y librerías
# %reload_ext autoreload
# %autoreload 2
from IPython.core.display import HTML
HTML(open("style/mat281.css", "r").read())
# + [markdown] slideshow={"slide_type": "slide"}
# <header class="w3-container w3-teal">
# <img src="images/utfsm.png" alt="" height="100px" align="left"/>
# <img src="images/mat.png" alt="" height="100px" align="right"/>
# </header>
# <br/><br/><br/><br/><br/>
# # MAT281
# ## Aplicaciones de la Matemática en la Ingeniería
#
# ### <NAME>
#
# https://www.github.com/sebastiandres/mat281
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Clase anterior
#
# * Uno
# * Dos
# * Tres
# + [markdown] slideshow={"slide_type": "slide"}
# ## ¿Qué contenido aprenderemos?
#
# * Uno
# * Dos
# * Tres
# + [markdown] slideshow={"slide_type": "slide"}
# ## ¿Porqué aprenderemos ese contenido?
#
# * Uno
# * Dos
# * Tres
# + [markdown] slideshow={"slide_type": "slide"}
# # Título
# Texto
#
# ## Subtítulo
# Texto
#
# ### Sección
# Texto
#
# #### Subsección
# Texto
#
# ##### Subsubsección
# Texto
# + [markdown] slideshow={"slide_type": "slide"}
# Imagen:
# <img src="images/utfsm.png" alt="" height="50px" align="middle"/>
#
# Lista:
# * Texto <span class="good"> bueno </span>, <span class="warning"> masomeno </span>
# * y <span class="bad"> malo </span> o ***muy malo***
#
# Y código
# -
# Codigo
1+2
| clases/ipynb-base/base.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # step_function_sample_sat
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/sat/step_function_sample_sat.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/ortools/sat/samples/step_function_sample_sat.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# #!/usr/bin/env python3
# Copyright 2010-2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Implements a step function."""
from ortools.sat.python import cp_model
class VarArraySolutionPrinter(cp_model.CpSolverSolutionCallback):
"""Print intermediate solutions."""
def __init__(self, variables):
cp_model.CpSolverSolutionCallback.__init__(self)
self.__variables = variables
self.__solution_count = 0
def on_solution_callback(self):
self.__solution_count += 1
for v in self.__variables:
print('%s=%i' % (v, self.Value(v)), end=' ')
print()
def solution_count(self):
return self.__solution_count
def step_function_sample_sat():
"""Encode the step function."""
# Model.
model = cp_model.CpModel()
# Declare our primary variable.
x = model.NewIntVar(0, 20, 'x')
# Create the expression variable and implement the step function
# Note it is not defined for x == 2.
#
# - 3
# -- -- --------- 2
# 1
# -- --- 0
# 0 ================ 20
#
expr = model.NewIntVar(0, 3, 'expr')
# expr == 0 on [5, 6] U [8, 10]
b0 = model.NewBoolVar('b0')
model.AddLinearExpressionInDomain(
x, cp_model.Domain.FromIntervals([(5, 6), (8, 10)])).OnlyEnforceIf(b0)
model.Add(expr == 0).OnlyEnforceIf(b0)
# expr == 2 on [0, 1] U [3, 4] U [11, 20]
b2 = model.NewBoolVar('b2')
model.AddLinearExpressionInDomain(
x, cp_model.Domain.FromIntervals([(0, 1), (3, 4),
(11, 20)])).OnlyEnforceIf(b2)
model.Add(expr == 2).OnlyEnforceIf(b2)
# expr == 3 when x == 7
b3 = model.NewBoolVar('b3')
model.Add(x == 7).OnlyEnforceIf(b3)
model.Add(expr == 3).OnlyEnforceIf(b3)
# At least one bi is true. (we could use a sum == 1).
model.AddBoolOr([b0, b2, b3])
# Search for x values in increasing order.
model.AddDecisionStrategy([x], cp_model.CHOOSE_FIRST,
cp_model.SELECT_MIN_VALUE)
# Create a solver and solve with a fixed search.
solver = cp_model.CpSolver()
# Force the solver to follow the decision strategy exactly.
solver.parameters.search_branching = cp_model.FIXED_SEARCH
# Enumerate all solutions.
solver.parameters.enumerate_all_solutions = True
# Search and print out all solutions.
solution_printer = VarArraySolutionPrinter([x, expr])
solver.Solve(model, solution_printer)
step_function_sample_sat()
| examples/notebook/sat/step_function_sample_sat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/stevenrdz/IR-SOES/blob/master/CS.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Wl1i7nz4iwZb" outputId="169f9876-3257-4b4e-de84-ea4632446ec3" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import files
from google.colab import drive
drive.mount('/content/gdrive')
# + id="t2vC2XqZivNj" outputId="9a4e7259-4c4d-40fc-d99f-48b51ed5c0de" colab={"base_uri": "https://localhost:8080/", "height": 102}
import nltk
nltk.download('stopwords')
nltk.download('punkt')
# + id="qbRym5Z0isTs" outputId="f6dce551-dc74-4939-b5d8-8ec37eb2f579" colab={"base_uri": "https://localhost:8080/", "height": 34}
from time import time
start_nb = time()
start = time()
import pandas as pd
from nltk.corpus import stopwords
import re
from nltk import word_tokenize
es_stops = set(stopwords.words('spanish'))
print('Importación de Librerias. %.2f segundos' % (time() - start))
# + id="cErokgQliSxH" outputId="82f8e81d-128d-40bc-ab61-58b75dfbf7a9" colab={"base_uri": "https://localhost:8080/", "height": 306}
start = time()
data = pd.read_csv('//content/gdrive/My Drive/ultimate.csv',
sep=';', encoding='utf-8-sig')
print('Dataset generado. %.2f segundos' % (time() - start))
data.head()
# + id="FqBc45cmil_S"
def preprocess(doc):
code = re.compile('[¿?""(),:'']') # Eliminación de signos de puntuacion.
doc = re.sub(code, '', doc)
punt = re.compile('[./@_]') # Eliminación de signos de puntuacion.
doc = re.sub(punt, ' ', doc)
code = re.compile('<pre><code>[\s\S]*</code></pre>') # Eliminación de codigo.
doc = re.sub(code, '', doc)
tag = re.compile('<.*?>') # Eliminación de etiquetas.
doc = re.sub(tag, '', doc)
doc = doc.lower() # Minuscula todo el texto.
doc = word_tokenize(doc) # Dividir en palabras.
doc = [w for w in doc if not w in es_stops] # Eliminar stopwords.
# doc = [w for w in doc if w.isalpha()] # Eliminar numbers and punctuation.
return doc
# + id="TcoMFH-PiX4M"
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import pandas as pd
import numpy as np
import string
titles_cosim = data['Processed_Title'].to_list()
id_index = data['Id'].to_list()
# Create the Document Term Matrix
vectorizer = CountVectorizer()
# + id="9Gu0tjGiiZDj"
def print_res(data, id):
with pd.option_context('display.max_colwidth', 500):
print(data.iloc[[id], [0,1]])
# + id="IKVIWU9uiaQa"
# Create the Document Term Matrix
count_vectorizer = CountVectorizer()
sparse_matrix = count_vectorizer.fit_transform(titles_cosim)
doc_term_matrix = sparse_matrix.todense()
dtm = pd.DataFrame(doc_term_matrix,
columns=count_vectorizer.get_feature_names())
# + id="T_5dL03ribrb" outputId="9608ce6b-236c-4f0e-aa87-f16d0eb2d659" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Create the Query Matrix
query = preprocess("Problemas con OSRM con R y Ubuntu 18.04")
print(query)
query_matrix = count_vectorizer.transform(query)
query_term_matrix = query_matrix.todense()
qtm = pd.DataFrame(query_term_matrix,
columns=count_vectorizer.get_feature_names())
# + id="G7ki9zRcidKD" outputId="0cc11484-cf95-48b2-8a1b-468d0dd093e9" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Results and format
k = 5
cossim_matrix = cosine_similarity(qtm, dtm)
last = len(cossim_matrix)-1
exa = []
arg_sort = cossim_matrix[last].argsort()[-k:][::-1]
print(arg_sort)
value_sort = np.sort(cossim_matrix[last])[-k:][::-1]
print(value_sort)
# + id="BgjlQOBuiekT"
for i in range(k):
exa.append((arg_sort[i], value_sort[i]))
# + id="Gc_8Ql_oifw6" outputId="1eb3cf4c-6659-4d8c-9c54-ba359eae5761" colab={"base_uri": "https://localhost:8080/", "height": 357}
for i in range(len(exa)):
print_res(data, exa[i][0])
print("sim: ",exa[i])
print("-------------------------------------------------------")
| CS.ipynb |