code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # AzureML Pipeline, AutoML, AKS Deployment for Sentence Similarity
# 
# This notebook builds off of the [AutoML Local Deployment ACI](automl_local_deployment_aci.ipynb) notebook and demonstrates how to use [Azure Machine Learning](https://azure.microsoft.com/en-us/services/machine-learning-service/
# ) pipelines and Automated Machine Learning ([AutoML](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-automated-ml
# )) to streamline the creation of a machine learning workflow for predicting sentence similarity. The pipeline contains two steps:
# 1. PythonScriptStep: embeds sentences using a popular sentence embedding model, Google Universal Sentence Encoder
# 2. AutoMLStep: demonstrates how to use Automated Machine Learning (AutoML) to automate model selection for predicting sentence similarity (regression)
#
# After creating the pipeline, the notebook demonstrates the deployment of our sentence similarity model using Azure Kubernetes Service ([AKS](https://docs.microsoft.com/en-us/azure/aks/intro-kubernetes
# )).
#
# This notebook showcases how to use the following AzureML features:
# - AzureML Pipelines (PythonScriptStep and AutoMLStep)
# - Automated Machine Learning
# - AmlCompute
# - Datastore
# - Logging
# ## Table of Contents
# 1. [Introduction](#1.-Introduction)
# * 1.1 [What are AzureML Pipelines?](#1.1-What-are-AzureML-Pipelines?)
# * 1.2 [What is Azure AutoML?](#1.2-What-is-Azure-AutoML?)
# * 1.3 [Modeling Problem](#1.3-Modeling-Problem)
# 2. [Data Preparation](#2.-Data-Preparation)
# 3. [AzureML Setup](#3.-AzureML-Setup)
# * 3.1 [Link to or create a `Workspace`](#3.1-Link-to-or-create-a-Workspace)
# * 3.2 [Set up an `Experiment` and Logging](#3.2-Set-up-an-Experiment-and-Logging)
# * 3.3 [Link `AmlCompute` compute target](#3.3-Link-AmlCompute-compute-target)
# * 3.4 [Upload data to `Datastore`](#3.4-Upload-data-to-Datastore)
# 4. [Create AzureML Pipeline](#4.-Create-AzureML-Pipeline)
# * 4.1 [Set up run configuration file](#4.1-Set-up-run-configuration-file)
# * 4.2 [PythonScriptStep](#4.2-PythonScriptStep)
# * 4.2.1 [Define python script to run](#4.2.1-Define-python-script-to-run)
# * 4.2.2 [Create PipelineData object](#4.2.2-Create-PipelineData-object)
# * 4.2.3 [Create PythonScriptStep](#4.2.3-Create-PythonScriptStep)
# * 4.3 [AutoMLStep](#4.3-AutoMLStep)
# * 4.3.1 [Define get_data script to load data](#4.3.1-Define-get_data-script-to-load-data)
# * 4.3.2 [Create AutoMLConfig object](#4.3.2-Create-AutoMLConfig-object)
# * 4.3.3 [Create AutoMLStep](#4.3.3-Create-AutoMLStep)
# 5. [Run Pipeline](#5.-Run-Pipeline)
# 6. [Deploy Sentence Similarity Model](#6.-Deploy-Sentence-Similarity-Model)
# * 6.1 [Register/Retrieve AutoML and Google Universal Sentence Encoder Models for Deployment](#6.1-Register/Retrieve-AutoML-and-Google-Universal-Sentence-Encoder-Models-for-Deployment)
# * 6.2 [Create Scoring Script](#6.2-Create-Scoring-Script)
# * 6.3 [Create a YAML File for the Environment](#6.3-Create-a-YAML-File-for-the-Environment)
# * 6.4 [Image Creation](#6.4-Image-Creation)
# * 6.5 [Provision the AKS Cluster](#6.5-Provision-the-AKS-Cluster)
# * 6.6 [Deploy the image as a Web Service to Azure Kubernetes Service](#6.6-Deploy-the-image-as-a-Web-Service-to-Azure-Kubernetes-Service)
# * 6.7 [Test Deployed Model](#6.7-Test-Deployed-Webservice)
#
#
# ## 1. Introduction
# ### 1.1 What are AzureML Pipelines?
#
# [AzureML Pipelines](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines) define reusable machine learning workflows that can be used as a template for your machine learning scenarios. Pipelines allow you to optimize your workflow and spend time on machine learning rather than infrastructure. A Pipeline is defined by a series of steps; the following steps are available: AdlaStep, AutoMLStep, AzureBatchStep, DataTransferStep, DatabricksStep, EstimatorStep, HyperDriveStep, ModuleStep, MpiStep, and PythonScriptStep (see [here](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/?view=azure-ml-py) for details of each step). When the pipeline is run, cached results are used for all steps that have not changed, optimizing the run time. Data sources and intermediate data can be used across multiple steps in a pipeline, saving time and resources. Below we see an example of an AzureML pipeline.
# 
# ### 1.2 What is Azure AutoML?
#
# Automated machine learning ([AutoML](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-automated-ml)) is a capability of Microsoft's [Azure Machine Learning service](https://azure.microsoft.com/en-us/services/machine-learning-service/
# ). The goal of AutoML is to improve the productivity of data scientists and democratize AI by allowing for the rapid development and deployment of machine learning models. To acheive this goal, AutoML automates the process of selecting a ML model and tuning the model. All the user is required to provide is a dataset (suitable for a classification, regression, or time-series forecasting problem) and a metric to optimize in choosing the model and hyperparameters. The user is also given the ability to set time and cost constraints for the model selection and tuning.
# 
# The AutoML model selection and tuning process can be easily tracked through the Azure portal or directly in python notebooks through the use of widgets. AutoML quickly selects a high quality machine learning model tailored for your prediction problem. In this notebook, we walk through the steps of preparing data, setting up an AutoML experiment, and evaluating the results of our best model. More information about running AutoML experiments in Python can be found [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train).
# ### 1.3 Modeling Problem
#
# The regression problem we will demonstrate is predicting sentence similarity scores on the STS Benchmark dataset. The [STS Benchmark dataset](http://ixa2.si.ehu.es/stswiki/index.php/STSbenchmark#STS_benchmark_dataset_and_companion_dataset) contains a selection of English datasets that were used in Semantic Textual Similarity (STS) tasks 2012-2017. The dataset contains 8,628 sentence pairs with a human-labeled integer representing the sentences' similarity (ranging from 0, for no meaning overlap, to 5, meaning equivalence).
#
# For each sentence in the sentence pair, we will use Google's pretrained Universal Sentence Encoder (details provided below) to generate a $512$-dimensional embedding. Both embeddings in the sentence pair will be concatenated and the resulting $1024$-dimensional vector will be used as features in our regression problem. Our target variable is the sentence similarity score.
# +
# Set the environment path to find NLP
import sys
sys.path.append("../../")
import time
import logging
import csv
import os
import pandas as pd
import shutil
import numpy as np
import sys
from scipy.stats import pearsonr
from scipy.spatial import distance
from sklearn.externals import joblib
import json
# Import utils
from utils_nlp.azureml import azureml_utils
from utils_nlp.dataset import stsbenchmark
from utils_nlp.dataset.preprocess import (
to_lowercase,
to_spacy_tokens,
rm_spacy_stopwords,
)
from utils_nlp.common.timer import Timer
# Google Universal Sentence Encoder loader
import tensorflow_hub as hub
# AzureML packages
import azureml as aml
import logging
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics=True)
from azureml.core import Datastore, Experiment, Workspace
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core.webservice import AksWebservice, Webservice
from azureml.core.compute import AksCompute, ComputeTarget
from azureml.core.image import ContainerImage
from azureml.core.model import Model
from azureml.train.automl import AutoMLStep, AutoMLStepRun, AutoMLConfig
from azureml.pipeline.core import Pipeline, PipelineData, TrainingOutput
from azureml.pipeline.steps import PythonScriptStep
from azureml.data.data_reference import DataReference
from azureml.widgets import RunDetails
print("System version: {}".format(sys.version))
print("Azure ML SDK Version:", aml.core.VERSION)
print("Pandas version: {}".format(pd.__version__))
# -
BASE_DATA_PATH = "../../data"
# + tags=["parameters"]
automl_settings = {
"task": 'regression', # type of task: classification, regression or forecasting
"iteration_timeout_minutes": 15, # How long each iteration can take before moving on
"iterations": 50, # Number of algorithm options to try
"primary_metric": "spearman_correlation", # Metric to optimize
"preprocess": True, # Whether dataset preprocessing should be applied
"verbosity": logging.INFO,
"blacklist_models": ['XGBoostRegressor'] #this model is blacklisted due to installation issues
}
config_path = (
"./.azureml"
) # Path to the directory containing config.json with azureml credentials
# Azure resources
subscription_id = "YOUR_SUBSCRIPTION_ID"
resource_group = "YOUR_RESOURCE_GROUP_NAME"
workspace_name = "YOUR_WORKSPACE_NAME"
workspace_region = "YOUR_WORKSPACE_REGION" #Possible values eastus, eastus2 and so on.
# -
# # 2. Data Preparation
# **STS Benchmark Dataset**
#
# As described above, the STS Benchmark dataset contains 8.6K sentence pairs along with a human-annotated score for how similar the two sentences are. We will load the training, development (validation), and test sets provided by STS Benchmark and preprocess the data (lowercase the text, drop irrelevant columns, and rename the remaining columns) using the utils contained in this repo. Each dataset will ultimately have three columns: _sentence1_ and _sentence2_ which contain the text of the sentences in the sentence pair, and _score_ which contains the human-annotated similarity score of the sentence pair.
# Load in the raw datasets as pandas dataframes
train_raw = stsbenchmark.load_pandas_df(BASE_DATA_PATH, file_split="train")
dev_raw = stsbenchmark.load_pandas_df(BASE_DATA_PATH, file_split="dev")
test_raw = stsbenchmark.load_pandas_df(BASE_DATA_PATH, file_split="test")
# Clean each dataset by lowercasing text, removing irrelevant columns,
# and renaming the remaining columns
train_clean = stsbenchmark.clean_sts(train_raw)
dev_clean = stsbenchmark.clean_sts(dev_raw)
test_clean = stsbenchmark.clean_sts(test_raw)
# Convert all text to lowercase
train = to_lowercase(train_clean)
dev = to_lowercase(dev_clean)
test = to_lowercase(test_clean)
print("Training set has {} sentences".format(len(train)))
print("Development set has {} sentences".format(len(dev)))
print("Testing set has {} sentences".format(len(test)))
train.head()
# +
# Save the cleaned data
if not os.path.isdir("data"):
os.mkdir("data")
train.to_csv("data/train.csv", index=False)
test.to_csv("data/test.csv", index=False)
dev.to_csv("data/dev.csv", index=False)
# -
# # 3. AzureML Setup
# Now, we set up the necessary components for running this as an AzureML experiment
# 1. Create or link to an existing `Workspace`
# 2. Set up an `Experiment` with `logging`
# 3. Create or attach existing `AmlCompute`
# 4. Upload our data to a `Datastore`
# ## 3.1 Link to or create a Workspace
# The following cell looks to set up the connection to your [Azure Machine Learning service Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace). You can choose to connect to an existing workspace or create a new one.
#
# **To access an existing workspace:**
# 1. If you have a `config.json` file, you do not need to provide the workspace information; you will only need to update the `config_path` variable that is defined above which contains the file.
# 2. Otherwise, you will need to supply the following:
# * The name of your workspace
# * Your subscription id
# * The resource group name
#
# **To create a new workspace:**
#
# Set the following information:
# * A name for your workspace
# * Your subscription id
# * The resource group name
# * [Azure region](https://azure.microsoft.com/en-us/global-infrastructure/regions/) to create the workspace in, such as `eastus2`.
#
# This will automatically create a new resource group for you in the region provided if a resource group with the name given does not already exist.
ws = azureml_utils.get_or_create_workspace(
config_path=config_path,
subscription_id=subscription_id,
resource_group=resource_group,
workspace_name=workspace_name,
workspace_region=workspace_region,
)
print(
"Workspace name: " + ws.name,
"Azure region: " + ws.location,
"Subscription id: " + ws.subscription_id,
"Resource group: " + ws.resource_group,
sep="\n",
)
# ## 3.2 Set up an Experiment and Logging
# +
# Make a folder for the project
project_folder = "./automl-sentence-similarity"
os.makedirs(project_folder, exist_ok=True)
# Set up an experiment
experiment_name = "NLP-SS-googleUSE"
experiment = Experiment(ws, experiment_name)
# Add logging to our experiment
run = experiment.start_logging()
# -
# ## 3.3 Link AmlCompute Compute Target
# To use AzureML Pipelines we need to link a compute target as they can not be run locally. The different options include AmlCompute, Azure Databricks, Remote VMs, etc. All [compute options](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#supported-compute-targets) can be found in this table with details about whether the given options work with automated ML, pipelines, and GPU. For the following example, we will use an AmlCompute target because it supports Azure Pipelines and GPU.
# +
# choose your cluster
cluster_name = "gpu-test"
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print("Found existing compute target.")
except ComputeTargetException:
print("Creating a new compute target...")
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_NC6", max_nodes=4
)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# use get_status() to get a detailed status for the current AmlCompute.
print(compute_target.get_status().serialize())
# -
# ## 3.4 Upload data to Datastore
# This step uploads our local data to a `Datastore` so that the data is accessible from the remote compute target and creates a `DataReference` to point to the location of the data on the Datastore. A DataStore is backed either by a Azure File Storage (default option) or Azure Blob Storage ([how to decide between these options](https://docs.microsoft.com/en-us/azure/storage/common/storage-decide-blobs-files-disks)) and data is made accessible by mounting or copying data to the compute target. `ws.datastores` lists all options for datastores and `ds.account_name` gets the name of the datastore that can be used to find it in the Azure portal.
# +
# Select a specific datastore or you can call ws.get_default_datastore()
datastore_name = "workspacefilestore"
ds = ws.datastores[datastore_name]
# Upload files in data folder to the datastore
ds.upload(
src_dir="./data",
target_path="stsbenchmark_data",
overwrite=True,
show_progress=True,
)
# -
# We also set up a `DataReference` object that points to the data we just uploaded into the stsbenchmark_data folder. DataReference objects point to data that is accessible from a datastore and will be used an an input into our pipeline.
input_data = DataReference(
datastore=ds,
data_reference_name="stsbenchmark",
path_on_datastore="stsbenchmark_data/",
overwrite=False,
)
# # 4. Create AzureML Pipeline
# Now we set up our pipeline which is made of two steps:
# 1. `PythonScriptStep`: takes each sentence pair from the data in the `Datastore` and concatenates the Google USE embeddings for each sentence into one vector. This step saves the embedding feature matrix back to our `Datastore` and uses a `PipelineData` object to represent this intermediate data.
# 2. `AutoMLStep`: takes the intermediate data produced by the previous step and passes it to an `AutoMLConfig` which performs the automatic model selection
# ## 4.1 Set up run configuration file
# First we set up a `RunConfiguration` object which configures the execution environment for an experiment (sets up the conda dependencies, etc.)
# + format="row"
# create a new RunConfig object
conda_run_config = RunConfiguration(framework="python")
# Set compute target to AmlCompute
conda_run_config.target = compute_target
conda_run_config.environment.docker.enabled = True
conda_run_config.environment.docker.base_image = aml.core.runconfig.DEFAULT_CPU_IMAGE
# Specify our own conda dependencies for the execution environment
conda_run_config.environment.python.user_managed_dependencies = False
conda_run_config.environment.python.conda_dependencies = CondaDependencies.create(
pip_packages=[
"azureml-sdk[automl]==1.0.48",
"azureml-dataprep==1.1.8",
"azureml-train-automl==1.0.48",
],
conda_packages=[
"numpy",
"py-xgboost<=0.80",
"pandas",
"tensorflow",
"tensorflow-hub",
"scikit-learn",
],
pin_sdk_version=False,
)
print("run config is ready")
# -
# ## 4.2 PythonScriptStep
# `PythonScriptStep` is a step which runs a user-defined Python script ([documentation](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.python_script_step.pythonscriptstep?view=azure-ml-py) here). In this `PythonScriptStep`, we will convert our sentences into a numerical representation in order to use them in our machine learning model. We will embed both sentences using the Google Universal Sentence Encoder (provided by tensorflow-hub) and concatenate their representations into a $1024$-dimensional vector to use as features for AutoML.
#
# **Google Universal Sentence Encoder:**
# We'll use a popular sentence encoder called Google Universal Sentence Encoder (see [original paper](https://static.googleusercontent.com/media/research.google.com/en//pubs/archive/46808.pdf)). Google provides two pretrained models based on different design goals: a Transformer model (targets high accuracy even if this reduces model complexity) and a Deep Averaging Network model (DAN; targets efficient inference). Both models are trained on a variety of web sources (Wikipedia, news, question-answers pages, and discussion forums) and produced 512-dimensional embeddings. This notebook utilizes the Transformer-based encoding model which can be downloaded [here](https://tfhub.dev/google/universal-sentence-encoder-large/3) because of its better performance relative to the DAN model on the STS Benchmark dataset (see Table 2 in Google Research's [paper](https://static.googleusercontent.com/media/research.google.com/en//pubs/archive/46808.pdf)). The Transformer model produces sentence embeddings using the "encoding sub-graph of the transformer architecture" (original architecture introduced [here](https://arxiv.org/abs/1706.03762)). "This sub-graph uses attention to compute context aware representations of words in a sentence that take into account both the ordering and identity of all the other workds. The context aware word representations are converted to a fixed length sentence encoding vector by computing the element-wise sum of the representations at each word position." The input to the model is lowercase PTB-tokenized strings and the model is designed to be useful for multiple different tasks by using multi-task learning. More details about the model can be found in the [paper](https://static.googleusercontent.com/media/research.google.com/en//pubs/archive/46808.pdf) by Google Research.
# ### 4.2.1 Define python script to run
#
# Define the script (called embed.py) that the `PythonScriptStep` will execute:
# +
# %%writefile $project_folder/embed.py
import argparse
import os
import azureml.core
import pandas as pd
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
tf.logging.set_verbosity(tf.logging.ERROR) # reduce logging output
def google_encoder(dataset):
""" Function that embeds sentences using the Google Universal
Sentence Encoder pretrained model
Parameters:
----------
dataset: pandas dataframe with sentences and scores
Returns:
-------
emb1: 512-dimensional representation of sentence1
emb2: 512-dimensional representation of sentence2
"""
sts_input1 = tf.placeholder(tf.string, shape=(None))
sts_input2 = tf.placeholder(tf.string, shape=(None))
# Apply embedding model and normalize the input
sts_encode1 = tf.nn.l2_normalize(embedding_model(sts_input1), axis=1)
sts_encode2 = tf.nn.l2_normalize(embedding_model(sts_input2), axis=1)
with tf.Session() as session:
session.run(tf.global_variables_initializer())
session.run(tf.tables_initializer())
emb1, emb2 = session.run(
[sts_encode1, sts_encode2],
feed_dict={
sts_input1: dataset["sentence1"],
sts_input2: dataset["sentence2"],
},
)
return emb1, emb2
def feature_engineering(dataset):
"""Extracts embedding features from the dataset and returns
features and target in a dataframe
Parameters:
----------
dataset: pandas dataframe with sentences and scores
Returns:
-------
df: pandas dataframe with embedding features
scores: list of target variables
"""
google_USE_emb1, google_USE_emb2 = google_encoder(dataset)
n_google = google_USE_emb1.shape[1] # length of the embeddings
df = np.concatenate((google_USE_emb1, google_USE_emb2), axis=1)
names = ["USEEmb1_" + str(i) for i in range(n_google)] + [
"USEEmb2_" + str(i) for i in range(n_google)
]
df = pd.DataFrame(df, columns=names)
return df, dataset["score"]
def write_output(df, path, name):
"""Write dataframes to correct path"""
os.makedirs(path, exist_ok=True)
print("%s created" % path)
df.to_csv(path + "/" + name, index=False)
# Parse arguments
parser = argparse.ArgumentParser()
parser.add_argument("--sentence_data", type=str)
parser.add_argument("--embedded_data", type=str)
args = parser.parse_args()
# Import the Universal Sentence Encoder's TF Hub module
module_url = "https://tfhub.dev/google/universal-sentence-encoder-large/3"
embedding_model = hub.Module(module_url)
# Read data
train = pd.read_csv(args.sentence_data + "/train.csv")
dev = pd.read_csv(args.sentence_data + "/dev.csv")
# Get Google USE features
training_data, training_scores = feature_engineering(train)
validation_data, validation_scores = feature_engineering(dev)
# Write out training data to Datastore
write_output(training_data, args.embedded_data, "X_train.csv")
write_output(
pd.DataFrame(training_scores, columns=["score"]), args.embedded_data, "y_train.csv"
)
# Write out validation data to Datastore
write_output(validation_data, args.embedded_data, "X_dev.csv")
write_output(
pd.DataFrame(validation_scores, columns=["score"]), args.embedded_data, "y_dev.csv"
)
# -
# ### 4.2.2 Create PipelineData object
# `PipelineData` objects represent a piece of intermediate data in a pipeline. Generally they are produced by one step (as an output) and then consumed by the next step (as an input), introducing an implicit order between steps in a pipeline. We create a PipelineData object that can represent the data produced by our first pipeline step that will be consumed by our second pipeline step.
embedded_data = PipelineData("embedded_data", datastore=ds)
# ### 4.2.3 Create PythonScriptStep
# This step defines the `PythonScriptStep`. We give the step a name, tell the step which python script to run (embed.py) and what directory that script is located in (source_directory).
#
# We also link the compute target and run configuration that we made previously. Our input is the `DataReference` object (input_data) where our raw sentence data was uploaded and our ouput is the `PipelineData` object (embedded_data) where the embedded data produced by this step will be stored. These are also passed in as arguments so that we have access to the correct data paths.
embed_step = PythonScriptStep(
name="Embed",
script_name="embed.py",
arguments=["--embedded_data", embedded_data, "--sentence_data", input_data],
inputs=[input_data],
outputs=[embedded_data],
compute_target=compute_target,
runconfig=conda_run_config,
source_directory=project_folder,
allow_reuse=True,
)
# ## 4.3 AutoMLStep
# `AutoMLStep` creates an AutoML step in a pipeline (see [documentation](https://docs.microsoft.com/en-us/python/api/azureml-train-automl/azureml.train.automl.automlstep?view=azure-ml-py) and [basic example](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-with-automated-machine-learning-step.ipynb)). When using AutoML on remote compute, rather than passing our data directly into the `AutoMLConfig` object as we did in the local example, we must define a get_data.py script with a get_data() function to pass as the data_script argument. This workflow can be used for both local and remote executions (see [details](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-auto-train-remote)).
#
#
# ### 4.3.1 Define get_data script to load data
# Define the get_data.py file and get_data() function that the `AutoMLStep` will execute to collect data. When AutoML is used with a remote compute, the data can not be passed directly as parameters. Rather, a get_data function must be defined to access the data (see [this resource](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-auto-train-remote) for further details). Note that we can directly access the path of the intermediate data (called embedded_data) through `os.environ['AZUREML_DATAREFERENCE_embedded_data']`. This is necessary because the AutoMLStep does not accept additional parameters like the PythonScriptStep does with `arguments`.
# +
# %%writefile $project_folder/get_data.py
import os
import pandas as pd
# get location of the embedded_data for future use
EMBEDDED_DATA_REF = os.environ["AZUREML_DATAREFERENCE_embedded_data"]
def get_data():
"""Function needed to load data for use on remote AutoML experiments"""
X_train = pd.read_csv(EMBEDDED_DATA_REF + "/X_train.csv")
y_train = pd.read_csv(EMBEDDED_DATA_REF + "/y_train.csv")
X_dev = pd.read_csv(EMBEDDED_DATA_REF + "/X_dev.csv")
y_dev = pd.read_csv(EMBEDDED_DATA_REF + "/y_dev.csv")
return {"X": X_train.values, "y": y_train.values.flatten(), "X_valid": X_dev.values, "y_valid": y_dev.values.flatten()}
# -
# ### 4.3.2 Create AutoMLConfig object
# Now, we specify the parameters for the `AutoMLConfig` class:
# **task**
# AutoML supports the following base learners for the regression task: Elastic Net, Light GBM, Gradient Boosting, Decision Tree, K-nearest Neighbors, LARS Lasso, Stochastic Gradient Descent, Random Forest, Extremely Randomized Trees, XGBoost, DNN Regressor, Linear Regression. In addition, AutoML also supports two kinds of ensemble methods: voting (weighted average of the output of multiple base learners) and stacking (training a second "metalearner" which uses the base algorithms' predictions to predict the target variable). Specific base learners can be included or excluded in the parameters for the AutoMLConfig class (whitelist_models and blacklist_models) and the voting/stacking ensemble options can be specified as well (enable_voting_ensemble and enable_stack_ensemble)
# **preprocess**
# AutoML also has advanced preprocessing methods, eliminating the need for users to perform this manually. Data is automatically scaled and normalized but an additional parameter in the AutoMLConfig class enables the use of more advanced techniques including imputation, generating additional features, transformations, word embeddings, etc. (full list found [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-create-portal-experiments#preprocess)). Note that algorithm-specific preprocessing will be applied even if preprocess=False.
# **primary_metric**
# The regression metrics available are the following: Spearman Correlation (spearman_correlation), Normalized RMSE (normalized_root_mean_squared_error), Normalized MAE (normalized_mean_absolute_error), and R2 score (r2_score)
# **Constraints:**
# There is a cost_mode parameter to set cost prediction modes (see options [here](https://docs.microsoft.com/en-us/python/api/azureml-train-automl/azureml.train.automl.automlconfig?view=azure-ml-py)). To set constraints on time there are multiple parameters including experiment_exit_score (target score to exit the experiment after achieving), experiment_timeout_minutes (maximum amount of time for all combined iterations), and iterations (total number of different algorithm and parameter combinations to try).
automl_config = AutoMLConfig(
debug_log="automl_errors.log",
path=project_folder,
compute_target=compute_target,
run_configuration=conda_run_config,
data_script=project_folder
+ "/get_data.py", # local path to script with get_data() function
**automl_settings #where the AutoML main settings are defined
)
# ### 4.3.3 Create AutoMLStep
# Finally, we create `PipelineData` objects for the model data (our outputs) and then create the `AutoMLStep`. The `AutoMLStep` requires a `AutoMLConfig` object and we pass our intermediate data (embedded_data) in as the inputs.
# +
# Create PipelineData objects for tracking AutoML metrics
metrics_data = PipelineData(
name="metrics_data",
datastore=ds,
pipeline_output_name="metrics_output",
training_output=TrainingOutput(type="Metrics"),
)
model_data = PipelineData(
name="model_data",
datastore=ds,
pipeline_output_name="best_model_output",
training_output=TrainingOutput(type="Model"),
)
# -
automl_step = AutoMLStep(
name="AutoML",
automl_config=automl_config, # the AutoMLConfig object created previously
inputs=[
embedded_data
], # inputs is the PipelineData that was the output of the previous pipeline step
outputs=[
metrics_data,
model_data,
], # PipelineData objects to reference metric and model information
allow_reuse=True,
)
# # 5. Run Pipeline
# Now we set up our pipeline which requires specifying our `Workspace` and the ordering of the steps that we created (steps parameter). We submit the pipeline and inspect the run details using a RunDetails widget. For remote runs, the execution of iterations is asynchronous.
pipeline = Pipeline(
description="pipeline_embed_automl", # give a name for the pipeline
workspace=ws,
steps=[embed_step, automl_step],
)
pipeline_run = experiment.submit(pipeline)
# Inspect the run details using the provided widget
RunDetails(pipeline_run).show()
# 
# Alternatively, block until the run has completed.
pipeline_run.wait_for_completion(
show_output=True
) # show console output while run is in progress
# **Cancel the Run**
#
# Interrupting/Restarting the jupyter kernel will not properly cancel the run, which can lead to wasted compute resources. To avoid this, we recommend explicitly canceling a run with the following code:
#
# `pipeline_run.cancel()`
# # 6. Deploy Sentence Similarity Model
#
# Deploying an Azure Machine Learning model as a web service creates a REST API. You can send data to this API and receive the prediction returned by the model.
# In general, you create a webservice by deploying a model as an image to a Compute Target.
#
# Some of the Compute Targets are:
# 1. Azure Container Instance
# 2. Azure Kubernetes Service
# 3. Local web service
#
# The general workflow for deploying a model is as follows:
# 1. Register a model
# 2. Prepare to deploy
# 3. Deploy the model to the compute target
# 4. Test the deployed model (webservice)
#
# In this notebook we walk you through the process of creating a webservice running on Azure Kubernetes Service ([AKS](https://docs.microsoft.com/en-us/azure/aks/intro-kubernetes
# )) by deploying the model as an image. AKS is good for high-scale production deployments. It provides fast response time and autoscaling of the deployed service. Cluster autoscaling is not supported through the Azure Machine Learning SDK.
#
# You can find more information on deploying and serving models [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-deploy-and-where)
#
# ## 6.1 Register/Retrieve AutoML and Google Universal Sentence Encoder Models for Deployment
#
# Registering a model means registering one or more files that make up a model. The Machine Learning models are registered in your current Aure Machine Learning Workspace. The model can either come from Azure Machine Learning or another location, such as your local machine.
#
# See other ways to register a model [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-deploy-and-where)
#
# Below we show how to register a new model and also how to retrieve and register an existing model.
#
# ### Register a new automl model
# Register the best AutoML model based on the pipeline results or load the saved model
# +
automl_step_run = AutoMLStepRun(step_run=pipeline_run.find_step_run("AutoML")[0])
# to register the fitted_mode
description = "Pipeline AutoML Model"
tags = {"area": "nlp", "type": "sentencesimilarity pipelines"}
model = automl_step_run.register_model(description=description, tags=tags)
automl_model_name = automl_step_run.model_id
print(
automl_step_run.model_id
) # Use this id to deploy the model as a web service in Azure.
# -
# ### Retrieve existing model from Azure
# If you already have a best model then you can skip registering the model by just retrieving the latest version of model by providing its name
automl_model_name = "711e9373160c4a8best" # best fit model registered in the workspace
model = Model(ws, name=automl_model_name)
print("Found model with name", automl_model_name)
# ### Register Google Universal Sentence Encoder Model
# Register the Google Universal Sentence Encoder model if not already registered in your workspace
# set location for where to download google tensorflow model
os.environ["TFHUB_CACHE_DIR"] = "./googleUSE"
# download model
hub.Module("https://tfhub.dev/google/universal-sentence-encoder-large/3")
# register model
embedding_model = Model.register(
model_path="googleUSE",
model_name="googleUSEmodel",
tags={"Model": "GoogleUSE"},
description="Google Universal Sentence Embedding pretrained model",
workspace=ws,
)
print("Registered googleUSEembeddings model")
# ### Retrieve existing Google USE model from Azure
embedding_model = Model(ws, name="googleUSEmodel")
print("Found model with name googleUSEembeddings")
# ## 6.2 Create Scoring Script
#
# In this section we show an example of an entry script, which is called from the deployed webservice. `score.py` is our entry script. The script must contain:
# 1. init() - This function loads the model in a global object.
# 2. run() - This function is used for model prediction. The inputs and outputs to `run()` typically use JSON for serialization and deserilization.
# +
# %%writefile score.py
import pickle
import json
import numpy as np
import azureml.train.automl
from sklearn.externals import joblib
from azureml.core.model import Model
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
import os
tf.logging.set_verbosity(tf.logging.ERROR) # reduce logging output
def google_encoder(dataset):
""" Function that embeds sentences using the Google Universal
Sentence Encoder pretrained model
Parameters:
----------
dataset: pandas dataframe with sentences and scores
Returns:
-------
emb1: 512-dimensional representation of sentence1
emb2: 512-dimensional representation of sentence2
"""
global embedding_model, sess
sts_input1 = tf.placeholder(tf.string, shape=(None))
sts_input2 = tf.placeholder(tf.string, shape=(None))
# Apply embedding model and normalize the input
sts_encode1 = tf.nn.l2_normalize(embedding_model(sts_input1), axis=1)
sts_encode2 = tf.nn.l2_normalize(embedding_model(sts_input2), axis=1)
sess.run(tf.global_variables_initializer())
sess.run(tf.tables_initializer())
emb1, emb2 = sess.run(
[sts_encode1, sts_encode2],
feed_dict={sts_input1: dataset["sentence1"], sts_input2: dataset["sentence2"]},
)
return emb1, emb2
def feature_engineering(dataset):
"""Extracts embedding features from the dataset and returns
features and target in a dataframe
Parameters:
----------
dataset: pandas dataframe with sentences and scores
Returns:
-------
df: pandas dataframe with embedding features
scores: list of target variables
"""
google_USE_emb1, google_USE_emb2 = google_encoder(dataset)
return np.concatenate((google_USE_emb1, google_USE_emb2), axis=1)
def init():
global model, googleUSE_dir_path
model_path = Model.get_model_path(
model_name="<<modelid>>"
) # this name is model.id of model that we want to deploy
# deserialize the model file back into a sklearn model
model = joblib.load(model_path)
# load the path for google USE embedding model
googleUSE_dir_path = Model.get_model_path(model_name="googleUSEmodel")
os.environ["TFHUB_CACHE_DIR"] = googleUSE_dir_path
def run(rawdata):
global embedding_model, sess, googleUSE_dir_path, model
try:
# load data and convert to dataframe
data = json.loads(rawdata)["data"]
data_df = pd.DataFrame(data, columns=["sentence1", "sentence2"])
# begin a tensorflow session and load tensorhub module
sess = tf.Session()
embedding_model = hub.Module(
googleUSE_dir_path + "/96e8f1d3d4d90ce86b2db128249eb8143a91db73"
)
# Embed sentences using Google USE model
embedded_data = feature_engineering(data_df)
# Predict using AutoML saved model
result = model.predict(embedded_data)
except Exception as e:
result = str(e)
sess.close()
return json.dumps({"error": result})
sess.close()
return json.dumps({"result": result.tolist()})
# +
# Substitute the actual model id in the script file.
script_file_name = "score.py"
with open(script_file_name, "r") as cefr:
content = cefr.read()
with open(script_file_name, "w") as cefw:
cefw.write(content.replace("<<modelid>>", automl_model_name))
# -
# ## 6.3 Create a YAML File for the Environment
#
# To ensure the fit results are consistent with the training results, the SDK dependency versions need to be the same as the environment that trains the model. The following cells create a file, pipeline_env.yml, which specifies the dependencies from the run.
# +
myenv = CondaDependencies.create(
conda_packages=[
"numpy",
"scikit-learn",
"py-xgboost<=0.80",
"pandas",
"tensorflow",
"tensorflow-hub",
],
pip_packages=["azureml-sdk[automl]==1.0.48.*"],
python_version="3.6.8",
)
conda_env_file_name = "pipeline_env.yml"
myenv.save_to_file(".", conda_env_file_name)
# -
# ## 6.4 Image Creation
#
# In this step we create a container image which is wrapper containing the entry script, yaml file with package dependencies and the model. The created image is then deployed as a webservice in the next step. This step can take up to 10 minutes and even longer if the model is large.
# +
# trying to add dependencies
image_config = ContainerImage.image_configuration(
execution_script=script_file_name,
runtime="python",
conda_file=conda_env_file_name,
description="Image with aml pipeline model",
tags={"area": "nlp", "type": "sentencesimilarity pipeline"},
)
image = ContainerImage.create(
name="pipeline-automl-image",
# this is the model object
models=[model, embedding_model], # add both embedding and autoML models
image_config=image_config,
workspace=ws,
)
image.wait_for_creation(show_output=True)
# -
# If the above step fails, then use below command to see logs.
# +
# image.get_logs()
# -
# ## 6.5 Provision the AKS Cluster
#
# **Time estimate:** Approximately 20 minutes.
#
# Creating or attaching an AKS cluster is a one time process for your workspace. You can reuse this cluster for multiple deployments. If you delete the cluster or the resource group that contains it, you must create a new cluster the next time you need to deploy. You can have multiple AKS clusters attached to your workspace.
#
# **Note:** Check the Azure Portal to make sure that the AKS Cluster has been provisioned properly before moving forward with this notebook
# +
# create aks cluser
# Use the default configuration (can also provide parameters to customize)
prov_config = AksCompute.provisioning_configuration()
# Create the cluster
aks_target = ComputeTarget.create(
workspace=ws, name="nlp-aks-cluster", provisioning_configuration=prov_config
)
# -
#
# ## 6.6 Deploy the Image as a Web Service on Azure Kubernetes Service
#
# In the case of deployment on AKS, in addition to the Docker image, we need to define computational resources. This is typically a cluster of CPUs or a cluster of GPUs. If we already have a Kubernetes-managed cluster in our workspace, we can use it, otherwise, we can create a new one.
#
# In this notebook we will use the cluster in the above cell.
# Set the web service configuration
aks_config = AksWebservice.deploy_configuration()
# We are now ready to deploy our web service. We will deploy from the Docker image. It contains our AutoML model as well as the Google Universal Sentence Encoder model and the conda environment needed for the scoring script to work properly. The parameters to pass to the Webservice.deploy_from_image() command are similar to those used for deployment on Azure Container Instance ([ACI](https://azure.microsoft.com/en-us/services/container-instances/
# )). The only major difference is the compute target (aks_target), i.e. the CPU cluster we just spun up.
#
# **Note:** This deployment takes a few minutes to complete.
# +
# deploy image as web service
aks_service_name = "aks-pipelines-service"
aks_service = Webservice.deploy_from_image(
workspace=ws,
name=aks_service_name,
image=image,
deployment_config=aks_config,
deployment_target=aks_target,
)
aks_service.wait_for_deployment(show_output=True)
print(aks_service.state)
# -
# If the above step fails then use below command to see logs
# +
# aks_service.get_logs()
# -
# ## 6.7 Test Deployed Webservice
#
# Testing the deployed model means running the created webservice. <br>
# The deployed model can be tested by passing a list of sentence pairs. The output will be a score between 0 and 5, with 0 indicating no meaning overlap between the sentences and 5 meaning equivalence.
#
# The run method expects input in json format. The Run() method retrieves API keys behind the scenes to make sure that the call is authenticated. The service has a timeout (default of ~30 seconds) which does not allow passing the large test dataset. To overcome this, you can batch data and send it to the service.
sentences = [
["This is sentence1", "This is sentence1"],
["A hungry cat.", "A sleeping cat"],
["Its summer time ", "Winter is coming"],
]
data = {"data": sentences}
data = json.dumps(data)
# +
# Set up a Timer to see how long the model takes to predict
t = Timer()
t.start()
score = aks_service.run(input_data=data)
t.stop()
print("Time elapsed: {}".format(t))
result = json.loads(score)
try:
output = result["result"]
print("Number of samples predicted: {}".format(len(output)))
print(output)
except:
print(result["error"])
# -
# Finally, we'll calculate the Pearson Correlation on the test set.
#
# **What is Pearson Correlation?**
#
# Our evaluation metric is Pearson correlation ($\rho$) which is a measure of the linear correlation between two variables. The formula for calculating Pearson correlation is as follows:
#
# $$\rho_{X,Y} = \frac{E[(X-\mu_X)(Y-\mu_Y)]}{\sigma_X \sigma_Y}$$
#
# This metric takes a value in [-1,1] where -1 represents a perfect negative correlation, 1 represents a perfect positive correlation, and 0 represents no correlation. We utilize the Pearson correlation metric as this is the main metric that [SentEval](http://nlpprogress.com/english/semantic_textual_similarity.html), a widely-used evaluation toolkit for evaluation sentence representations, uses for the STS Benchmark dataset.
# load test set sentences
data = pd.read_csv("data/test.csv")
train_y = data["score"].values.flatten()
train_x = data.drop("score", axis=1).values.tolist()
data = {"data": train_x[:500]}
data = json.dumps(data)
# +
# Set up a Timer to see how long the model takes to predict
t = Timer()
t.start()
score = aks_service.run(input_data=data)
t.stop()
print("Time elapsed: {}".format(t))
result = json.loads(score)
try:
output = result["result"]
print("Number of sample predicted : {}".format(len(output)))
except:
print(result["error"])
# -
# get Pearson Correlation
print(pearsonr(output, train_y[:500])[0])
# ## Conclusion
#
# This notebook demonstrated how to use AzureML Pipelines and AutoML to streamline the creation of a machine learning workflow for predicting sentence similarity. After creating the pipeline, the notebook demonstrated the deployment of our sentence similarity model using AKS. The model results reported in this notebook (using Google USE embeddings) are much stronger than the results from using AutoML with its built-in embedding capabilities (as in [AutoML Local Deployment ACI](automl_local_deployment_aci.ipynb)).
| examples/sentence_similarity/automl_with_pipelines_deployment_aks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aayushkumar20/Advance-Python-projects/blob/main/Chat%20backup/WhatsApp_chats_backup_to_a_excel_file.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Gjud5PfJh5EQ"
# !pip install pandas
# !pip install openpyxl
# !pip install pushbullet.py == 0.9.1
# + id="BLtn46uIicpJ"
#Install all this modules for proper functioning.
#Setup Pushbullet on both smartphone and computer by creating a login account
#Generate and download the API i.e. access token.
# + id="V7V0afc-iagQ"
# Import following modules
import urllib.request
import pandas as pd
from pushbullet import PushBullet
# Get Access Token from pushbullet.com
Access_token = "Your Access Token" # Download / copy the API fron the website.
# Authentication
pb = PushBullet(Access_token)
# All pushes created by you
all_pushes = pb.get_pushes()
# Get the latest push
latest_one = all_pushes[0]
# Fetch the latest file URL link
url = latest_one['file_url']
# Create a new text file for storing
# all the chats
Text_file = "All_Chats.txt"
# Retrieve all the data store into
# Text file
urllib.request.urlretrieve(url, Text_file)
# Create an empty chat list
chat_list = []
# Open the Text file in read mode and
# read all the data
with open(Text_file, mode='r', encoding='utf8') as f:
# Read all the data line-by-line
data = f.readlines()
# Excluded the first item of the list
# first items contains some garbage
# data
final_data_set = data[1:]
# Run a loop and read all the data
# line-by-line
for line in final_data_set:
# Extract the date, time, name,
# message
date = line.split(",")[0]
tim = line.split("-")[0].split(",")[1]
name = line.split(":")[1].split("-")[1]
message = line.split(":")[2][:-1]
# Append all the data in a List
chat_list.append([date, tim, name, message])
# Create a dataframe, for storing
# all the data in a excel file
df = pd.DataFrame(chat_list,
columns = ['Date', 'Time',
'Name', 'Message'])
df.to_excel("BackUp.xlsx", index = False)
| Chat backup/WhatsApp_chats_backup_to_a_excel_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sentiment Classification & How To "Frame Problems" for a Neural Network
#
# by <NAME>
#
# - **Twitter**: @iamtrask
# - **Blog**: http://iamtrask.github.io
# ### What You Should Already Know
#
# - neural networks, forward and back-propagation
# - stochastic gradient descent
# - mean squared error
# - and train/test splits
#
# ### Where to Get Help if You Need it
# - Re-watch previous Udacity Lectures
# - Leverage the recommended Course Reading Material - [Grokking Deep Learning](https://www.manning.com/books/grokking-deep-learning) (40% Off: **traskud17**)
# - Shoot me a tweet @iamtrask
#
#
# ### Tutorial Outline:
#
# - Intro: The Importance of "Framing a Problem"
#
#
# - Curate a Dataset
# - Developing a "Predictive Theory"
# - **PROJECT 1**: Quick Theory Validation
#
#
# - Transforming Text to Numbers
# - **PROJECT 2**: Creating the Input/Output Data
#
#
# - Putting it all together in a Neural Network
# - **PROJECT 3**: Building our Neural Network
#
#
# - Understanding Neural Noise
# - **PROJECT 4**: Making Learning Faster by Reducing Noise
#
#
# - Analyzing Inefficiencies in our Network
# - **PROJECT 5**: Making our Network Train and Run Faster
#
#
# - Further Noise Reduction
# - **PROJECT 6**: Reducing Noise by Strategically Reducing the Vocabulary
#
#
# - Analysis: What's going on in the weights?
# + [markdown] nbpresent={"id": "56bb3cba-260c-4ebe-9ed6-b995b4c72aa3"}
# # Lesson: Curate a Dataset
# + nbpresent={"id": "eba2b193-0419-431e-8db9-60f34dd3fe83"}
def pretty_print_review_and_label(i):
print(labels[i] + "\t:\t" + reviews[i][:80] + "...")
g = open('reviews.txt','r') # What we know!
reviews = list(map(lambda x:x[:-1],g.readlines()))
g.close()
g = open('labels.txt','r') # What we WANT to know!
labels = list(map(lambda x:x[:-1].upper(),g.readlines()))
g.close()
# -
len(reviews)
# + nbpresent={"id": "bb95574b-21a0-4213-ae50-34363cf4f87f"}
reviews[0]
# + nbpresent={"id": "e0408810-c424-4ed4-afb9-1735e9ddbd0a"}
labels[0]
# -
# # Lesson: Develop a Predictive Theory
# + nbpresent={"id": "e67a709f-234f-4493-bae6-4fb192141ee0"}
print("labels.txt \t : \t reviews.txt\n")
pretty_print_review_and_label(2137)
pretty_print_review_and_label(12816)
pretty_print_review_and_label(6267)
pretty_print_review_and_label(21934)
pretty_print_review_and_label(5297)
pretty_print_review_and_label(4998)
# -
# # Project 1: Quick Theory Validation
from collections import Counter
import numpy as np
positive_counts = Counter()
negative_counts = Counter()
total_counts = Counter()
for i in range(len(reviews)):
if(labels[i] == 'POSITIVE'):
for word in reviews[i].split(" "):
positive_counts[word] += 1
total_counts[word] += 1
else:
for word in reviews[i].split(" "):
negative_counts[word] += 1
total_counts[word] += 1
positive_counts.most_common()
# +
pos_neg_ratios = Counter()
for term,cnt in list(total_counts.most_common()):
if(cnt > 100):
pos_neg_ratio = positive_counts[term] / float(negative_counts[term]+1)
pos_neg_ratios[term] = pos_neg_ratio
for word,ratio in pos_neg_ratios.most_common():
if(ratio > 1):
pos_neg_ratios[word] = np.log(ratio)
else:
pos_neg_ratios[word] = -np.log((1 / (ratio+0.01)))
# -
# words most frequently seen in a review with a "POSITIVE" label
pos_neg_ratios.most_common()
# words most frequently seen in a review with a "NEGATIVE" label
list(reversed(pos_neg_ratios.most_common()))[0:30]
# # Transforming Text into Numbers
# +
from IPython.display import Image
review = "This was a horrible, terrible movie."
Image(filename='sentiment_network.png')
# +
review = "The movie was excellent"
Image(filename='sentiment_network_pos.png')
# -
# # Project 2: Creating the Input/Output Data
vocab = set(total_counts.keys())
vocab_size = len(vocab)
print(vocab_size)
list(vocab)
# +
import numpy as np
layer_0 = np.zeros((1,vocab_size))
layer_0
# -
from IPython.display import Image
Image(filename='sentiment_network.png')
# +
word2index = {}
for i,word in enumerate(vocab):
word2index[word] = i
word2index
# +
def update_input_layer(review):
global layer_0
# clear out previous state, reset the layer to be all 0s
layer_0 *= 0
for word in review.split(" "):
layer_0[0][word2index[word]] += 1
update_input_layer(reviews[0])
# -
layer_0
def get_target_for_label(label):
if(label == 'POSITIVE'):
return 1
else:
return 0
labels[0]
get_target_for_label(labels[0])
labels[1]
get_target_for_label(labels[1])
# # Project 3: Building a Neural Network
# - Start with your neural network from the last chapter
# - 3 layer neural network
# - no non-linearity in hidden layer
# - use our functions to create the training data
# - create a "pre_process_data" function to create vocabulary for our training data generating functions
# - modify "train" to train over the entire corpus
# ### Where to Get Help if You Need it
# - Re-watch previous week's Udacity Lectures
# - Chapters 3-5 - [Grokking Deep Learning](https://www.manning.com/books/grokking-deep-learning) - (40% Off: **traskud17**)
| sentiment_network/Sentiment Classification - Mini Project 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Planet Tasking API Order Creation
#
# ---
# ## Introduction
#
# ---
#
# This tutorial is an introduction on how to create tasking orders using [Planet](https://www.planet.com)'s Tasking API. It provides code samples on how to write simple Python code to do this.
#
# The API reference documentation can be found at https://developers.planet.com/docs/tasking
# ### Requirements
#
# ---
#
# #### Software & Modules
#
# This tutorial assumes familiarity with the [Python](https://python.org) programming language throughout. Familiarity with basic REST API concepts and usage is also assumed.
#
# We'll be using a **"Jupyter Notebook"** (aka Python Notebook) to run through the examples.
# To learn more about and get started with using Jupyter, visit: [Jupyter](https://jupyter.org/) and [IPython](https://ipython.org/).
#
# For the best experience, download this notebook and run it on your system, and make sure to install the modules listed below first. You can also copy the examples' code to a separate Python files an run them directly with Python on your system if you prefer.
#
# #### Planet API Key
#
# You should have an account on the Planet Platform to access the Tasking API. You may retrieve your API key from your [account page](https://www.planet.com/account/), or from the "API Tab" in [Planet Explorer](https://www.planet.com/explorer).
# ## Overview
#
# ---
#
# ### The basic workflow
#
# 1. Create a tasking order
# 1. Check the status of the tasking order
# 1. Download images captured by the tasking order
#
# Examples on how to edit or cancel existing tasking orders can be found in the notebook [planet_tasking_api_order_edit_and_cancel.ipynb](planet_tasking_api_order_edit_and_cancel.ipynb)
# ### API Endpoints
#
# This tutorial will cover the following API ***endpoint***:
#
# * [`/order`](https://api.planet.com/tasking/v2/order/)
# * [`https://api.planet.com/data/v1/quick-search`](https://api.planet.com/data/v1/quick-search)
#
# ## Basic Setup
#
# ---
#
# Before interacting with the Planet Tasking API using Python, we will set up our environment with some useful modules and helper functions.
#
# * We'll configure *authentication* to the Planet Tasking API
# * We'll use the `requests` Python module to make HTTP communication easier.
# * We'll use the `json` Python module to help us work with JSON responses from the API.
# * We'll use the `pytz` Python module to define the time frame for the order that we will be creating.
# * We'll create a function called `p` that will print Python dictionaries nicely.
#
# Then we'll be ready to make our first call to the Planet Tasking API by hitting the base endpoint at `https://api.planet.com/tasking/v2`.
#
# Let's start by configuring authentication:
# ### Authentication
#
# Authentication with the Planet Tasking API can be achieved using a valid Planet **API key**.
# You can *export* your API Key as an environment variable on your system:
#
# `export PL_API_KEY="YOUR API KEY HERE"`
#
# Or add the variable to your path, etc.
#
# To start our Python code, we'll setup an API Key variable from an environment variable to use with our requests:
# +
# Import the os module in order to access environment variables
import os
#If you are running this notebook outside of the docker environment that comes with the repo, you can uncomment the next line to provide your API key
#os.environ['PL_API_KEY']=input('Please provide your API Key')
# Setup the API Key from the `PL_API_KEY` environment variable
PLANET_API_KEY = os.getenv('PL_API_KEY')
# -
# ### Helper Modules and Functions
# Import helper modules
import json
import requests
import pytz
from time import sleep
from datetime import datetime, timedelta
# Helper function to printformatted JSON using the json module
def p(data):
print(json.dumps(data, indent=2))
# +
# Setup Planet Tasking PLANET_API_HOST
TASKING_API_URL = "https://api.planet.com/tasking/v2"
# Setup the session
session = requests.Session()
# Authenticate
session.headers.update({
'Authorization': f'api-key {PLANET_API_KEY}',
'Content-Type': 'application/json'
})
# -
# ### 1 | Creating a tasking order
# +
## Compose the tasking order
We want to create a tasking order that can return an image to us. To keep things simple we are going to create a Point order, which takes a single latitude/longitude coordinate pair. Since this is your tasking order, you need to provide the details of what the tasing order is called and the coordinates for the tasking order.
To make things easier we will default the start and end time to start tomorrow and end 7 days from now. Of course feel free to change this to suit your needs but if you do take note that all times should be in UTC format. The start and end times are optional,but we include them in this tutorial to provide a better picture of what can be done.
# +
# Define the name and coordinates for the order
name=input("Give the order a name")
latitude=float(input("Provide the latitude"))
longitude=float(input("Provide the longitude"))
# Because the geometry is GeoJSON, the coordinates must be longitude,latitude
order = {
'name': name,
'geometry': {
'type': 'Point',
'coordinates': [
longitude,
latitude
]
}
}
# Set a start and end time, giving the order a week to complete
tomorrow = datetime.now(pytz.utc) + timedelta(days=1)
one_week_later = tomorrow + timedelta(days=7)
datetime_parameters = {
'start_time': tomorrow.isoformat(),
'end_time': one_week_later.isoformat()
}
# Add the datetime parameters
order.update(datetime_parameters)
# -
#View the payload before posting
p(order)
# +
# The creation of an order is a POST request to the /orders endpoint
res = session.request('POST', TASKING_API_URL + '/orders/', json=order)
if res.status_code == 403:
print('Your PLANET_API_KEY is valid, but you are not authorized.')
elif res.status_code == 401:
print('Your PLANET_API_KEY is incorrect')
elif res.status_code == 201:
print('Your order was created successfully')
else:
print(f'Received status code {res.status_code} from the API. Please contact support.')
# View the response
p(res.json())
# -
# **Congratulations!** You just created your first tasking order to the Planet Tasking API. Depending on the start and end time that you provided, a satellite will be attempting to take an image over your given coordinates in the near future.
# ### 2 | Check the status of the tasking order
#
# To see the status an existing tasking order, the tasking order id is required. Depending on the tasking order, it can take some time for the status of the tasking order to change, and so you may need to come back to this section once some time has elapsed before changes to the tasking order can be seen. It is recommended to run the next part of this notebook to extract the ID of the newly created order and save that for later use.
# Get the response JSON and extract the ID of the order
response = res.json()
new_order_id = response["id"]
p(new_order_id)
def check_order_status(order_id):
# Make a GET request with the order_id concatenated to the end of the /orders url; e.g. https://api.planet.com/tasking/v2/orders/<ORDER_ID>
res = session.request('GET', TASKING_API_URL + '/orders/' + order_id)
if res.status_code == 403:
print('Your PLANET_API_KEYPLANET_API_KEY is valid, but you are not authorized to view this order.')
elif res.status_code == 401:
print('Your PLANET_API_KEYPLANET_API_KEY is incorrect')
elif res.status_code == 404:
print(f'Your order ({order_id}) does not exist')
elif res.status_code != 200:
print(f'Received status code {res.status_code} from the API. Please contact support.')
else:
order = res.json()
p(res.json())
print(f'Your order is {order["status"]} with {order["capture_status_published_count"]} published captures '
f'and {order["capture_assessment_success_count"]} successful captures')
check_order_status(new_order_id)
# ### 3 | Download successfully captured images
#
# Once the status of the tasking order has reached "FULFILLED" you can be certain that there are images associated with the tasking order that can be downloaded. To do this we need to use another api, the Planet Data API, to retreive the images. If you want to know more about the Planet Data API,there is Jupyter Notebok 'jupyter-notebooks/data-api-tutorials/planet_data_api_introduction.ipynb' which can provide a more complete tutorial.
#
# As with monitoring the tasking order, the tasking order id is required.
def download_successful_captures(order_id):
# Make a GET request to the captures endpoint
res = session.request('GET', TASKING_API_URL + '/captures/?order_id' + order_id + '&fulfilling=true')
if res.status_code == 403:
print('Your API KEY is valid, but you are not authorized to view this order.')
elif res.status_code == 401:
print('Your API KEY is incorrect')
elif res.status_code != 200:
print(f'Received status code {res.status_code} from the API. Please contact support.')
else:
p(res.json())
# Retrieve the captures from the response
captures = res.json()['results']
# For each capture, take the strip ID and create a payload that will be sent to the Data API
strip_ids = [capture['strip_id'] for capture in captures]
search_data = {
"filter": {
"config": strip_ids,
"field_name": "strip_id",
"type": "StringInFilter"
},
"item_types": ["SkySatCollect"]
}
# Make a POST requst to the Data API
data_api_response = session.request('POST', 'https://api.planet.com/data/v1/quick-search',search_data)
asset_urls = [feature['_links']['assets'] for feature in data_api_response.json()['features']]
# Activate the ortho_visual asset(s)
ortho_visual_urls = []
for asset_url in asset_urls:
assets = requests.get(asset_url, headers=headers).json()
activation_url = assets['ortho_visual']['_links']['activate']
requests.get(activation_url, headers=headers)
ortho_visual_urls.append(assets['ortho_visual']['_links']['_self'])
# Wait for activation and print
for ortho_visual_url in ortho_visual_urls:
ortho_visual = requests.get(ortho_visual_url, headers=headers).json()
while 'location' not in ortho_visual:
sleep(10)
print('Waiting 10 seconds for asset to unlock...')
ortho_visual = requests.get(ortho_visual_url, headers=headers).json()
print(f'Open the following link in a browser or download it to a file:\n{ortho_visual["location"]}')
download_successful_captures(new_order_id)
| jupyter-notebooks/tasking-api/planet_tasking_api_order_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 
# ___
# # Chapter 8 - Basic Algorithmic Learning
# ## Segment 2 - Logistic Regression
# +
import numpy as np
import pandas as pd
from pandas import Series, DataFrame
import scipy
from scipy.stats import spearmanr
import matplotlib.pyplot as plt
from pylab import rcParams
import seaborn as sb
import sklearn
from sklearn.preprocessing import scale
from sklearn.linear_model import LogisticRegression
from sklearn.cross_validation import train_test_split
from sklearn import metrics
from sklearn import preprocessing
# -
# %matplotlib inline
rcParams['figure.figsize'] = 5, 4
sb.set_style('whitegrid')
# ### Logistic regression on mtcars
address = 'C:/Users/<NAME>/Desktop/Exercise Files/Ch08/08_02/mtcars.csv'
cars = pd.read_csv(address)
cars.columns = ['car_names','mpg','cyl','disp', 'hp', 'drat', 'wt', 'qsec', 'vs', 'am', 'gear', 'carb']
cars.head()
# +
cars_data = cars.ix[:,(5,11)].values
cars_data_names = ['drat','carb']
y = cars.ix[:,9].values
# -
# #### Checking for independence between features
sb.regplot(x='drat', y='carb', data=cars, scatter=True)
# +
drat = cars['drat']
carb = cars['carb']
spearmanr_coefficient, p_value = spearmanr(drat, carb)
print 'Spearman Rank Correlation Coefficient %0.3f' % (spearmanr_coefficient)
# -
# #### Checking for missing values
cars.isnull().sum()
# #### Checking that your target is binary or ordinal
sb.countplot(x='am', data=cars, palette='hls')
# #### Checking that your dataset size is sufficient
cars.info()
# #### Deploying and evaluating your model
X = scale(cars_data)
# +
LogReg = LogisticRegression()
LogReg.fit(X,y)
print LogReg.score(X,y)
# -
y_pred = LogReg.predict(X)
from sklearn.metrics import classification_report
print(classification_report(y, y_pred))
| Ch08/08_02/08_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
mlbyte
from sklearn import *
train = pd.read_hdf('train', 'Sam')
test = pd.read_hdf('test', 'Sam')
def RMSLE(y, pred):
return metrics.mean_squared_error(y, pred)**0.5
train.drop('day_of_week', axis= 1, inplace=True)
test.drop('day_of_week', axis= 1, inplace=True)
col = [c for c in train if c not in ['id', 'air_store_id', 'visit_date','visitors']]
test.head()
from xgboost import XGBRegressor
model2 = neighbors.KNeighborsRegressor(n_jobs=-1, n_neighbors=4)
model3 = XGBRegressor(learning_rate=0.2, seed=3, n_estimators=200, subsample=0.8,
colsample_bytree=0.8, max_depth =10)
# +
#model1.fit(train[col], np.log1p(train['visitors'].values))
model2.fit(train[col], np.log1p(train['visitors'].values))
model3.fit(train[col], np.log1p(train['visitors'].values))
#preds1 = model1.predict(train[col])
preds2 = model2.predict(train[col])
preds3 = model3.predict(train[col])
#print('RMSE GradientBoostingRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds1))
print('RMSE KNeighborsRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds2))
print('RMSE XGBRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds3))
# -
from lightgbm import LGBMRegressor
model1 = LGBMRegressor(learning_rate=0.3, num_leaves=1400, max_depth=15, max_bin=300, min_child_weight=5)
model1.fit(train[col], np.log1p(train['visitors'].values))
preds1 = model1.predict(train[col])
print('RMSE LightGBM: ', RMSLE(np.log1p(train['visitors'].values), preds1))
preds1 = model1.predict(test[col])
preds2 = model2.predict(test[col])
preds3 = model3.predict(test[col])
pred = (preds1+preds2+preds3) / 3.0
test['visitors'] = pred
test['visitors'] = np.expm1(test['visitors']).clip(lower=0.)
sub1 = test[['id','visitors']].copy()
| Recruit Restaurant Visitor Forecasting/Notebooks/.ipynb_checkpoints/DAE-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="l-23gBrt4x2B"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="HMUDt0CiUJk9"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="77z2OchJTk0l"
# # Migrating feature_columns to TF2's Keras Preprocessing Layers
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/migrate/migrating_feature_columns">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/migrate/migrating_feature_columns.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/migrate/migrating_feature_columns.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/migrate/migrating_feature_columns.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + id="g3Zltl6_3Gy2"
# Temporarily install tf-nightly as the notebook depends on symbols in 2.6.
# !pip uninstall -q -y tensorflow keras
# !pip install -q tf-nightly
# + [markdown] id="-5jGPDA2PDPI"
# Training a model will usually come with some amount of feature preprocessing, particularly when dealing with structured data. When training a `tf.estimator.Estimator` in TF1, this feature preprocessing is usually done with the `tf.feature_column` API. In TF2, this preprocessing can be done directly with Keras layers, called _preprocessing layers_.
#
# In this migration guide, you will perform some common feature transformations using both feature columns and preprocessing layers, followed by training a complete model with both APIs.
#
# First, start with a couple of necessary imports,
# + id="iE0vSfMXumKI"
import tensorflow as tf
import tensorflow.compat.v1 as tf1
import math
# + [markdown] id="NVPYTQAWtDwH"
# and add a utility for calling a feature column for demonstration:
# + id="LAaifuuytJjM"
def call_feature_columns(feature_columns, inputs):
# This is a convenient way to call a `feature_column` outside of an estimator
# to display its output.
feature_layer = tf1.keras.layers.DenseFeatures(feature_columns)
return feature_layer(inputs)
# + [markdown] id="ZJnw07hYDGYt"
# ## Input handling
#
# To use feature columns with an estimator, model inputs are always expected to be a dictionary of tensors:
# + id="y0WUpQxsKEzf"
input_dict = {
'foo': tf.constant([1]),
'bar': tf.constant([0]),
'baz': tf.constant([-1])
}
# + [markdown] id="xYsC6H_BJ8l3"
# Each feature column needs to be created with a key to index into the source data. The output of all feature columns is concatenated and used by the estimator model.
# + id="3fvIe3V8Ffjt"
columns = [
tf1.feature_column.numeric_column('foo'),
tf1.feature_column.numeric_column('bar'),
tf1.feature_column.numeric_column('baz'),
]
call_feature_columns(columns, input_dict)
# + [markdown] id="hvPfCK2XGTyl"
# In Keras, model input is much more flexible. A `tf.keras.Model` can handle a single tensor input, a list of tensor features, or a dictionary of tensor features. You can handle dictionary input by passing a dictionary of `tf.keras.Input` on model creation. Inputs will not be concatenated automatically, which allows them to be used in much more flexible ways. They can be concatenated with `tf.keras.layers.Concatenate`.
# + id="5sYWENkgLWJ2"
inputs = {
'foo': tf.keras.Input(shape=()),
'bar': tf.keras.Input(shape=()),
'baz': tf.keras.Input(shape=()),
}
# Inputs are typically transformed by preprocessing layers before concatenation.
outputs = tf.keras.layers.Concatenate()(inputs.values())
model = tf.keras.Model(inputs=inputs, outputs=outputs)
model(input_dict)
# + [markdown] id="GXkmiuwXTS-B"
# ## One-hot encoding integer IDs
#
# A common feature transformation is one-hot encoding integer inputs of a known range. Here is an example using feature columns:
# + id="XasXzOgatgRF"
categorical_col = tf1.feature_column.categorical_column_with_identity(
'type', num_buckets=3)
indicator_col = tf1.feature_column.indicator_column(categorical_col)
call_feature_columns(indicator_col, {'type': [0, 1, 2]})
# + [markdown] id="iSCkJEQ6U-ru"
# Using Keras preprocessing layers, these columns can be replaced by a single `tf.keras.layers.CategoryEncoding` layer with `output_mode` set to `'one_hot'`:
# + id="799lbMNNuAVz"
one_hot_layer = tf.keras.layers.CategoryEncoding(
num_tokens=3, output_mode='one_hot')
one_hot_layer([0, 1, 2])
# + [markdown] id="kNzRtESU7tga"
# Note: For large one-hot encodings, it is much more efficient to use a sparse representation of the output. If you pass `sparse=True` to the `CategoryEncoding` layer, the output of the layer will be a `tf.sparse.SparseTensor`, which can be efficiently handled as input to a `tf.keras.layers.Dense` layer.
# + [markdown] id="Zf7kjhTiAErK"
# ## Normalizing numeric features
#
# When handling continuous, floating-point features with feature columns, you need to use a `tf.feature_column.numeric_column`. In the case where the input is already normalized, converting this to Keras is trivial. You can simply use a `tf.keras.Input` directly into your model, as shown above.
#
# A `numeric_column` can also be used to normalize input:
# + id="HbTMGB9XctGx"
def normalize(x):
mean, variance = (2.0, 1.0)
return (x - mean) / math.sqrt(variance)
numeric_col = tf1.feature_column.numeric_column('col', normalizer_fn=normalize)
call_feature_columns(numeric_col, {'col': tf.constant([[0.], [1.], [2.]])})
# + [markdown] id="M9cyhPR_drOz"
# In contrast, with Keras, this normalization can be done with `tf.keras.layers.Normalization`.
# + id="8bcgG-yOdqUH"
normalization_layer = tf.keras.layers.Normalization(mean=2.0, variance=1.0)
normalization_layer(tf.constant([[0.], [1.], [2.]]))
# + [markdown] id="d1InD_4QLKU-"
# ## Bucketizing and one-hot encoding numeric features
# + [markdown] id="k5e0b8iOLRzd"
# Another common transformation of continuous, floating point inputs is to bucketize then to integers of a fixed range.
#
# In feature columns, this can be achieved with a `tf.feature_column.bucketized_column`:
# + id="_rbx6qQ-LQx7"
numeric_col = tf1.feature_column.numeric_column('col')
bucketized_col = tf1.feature_column.bucketized_column(numeric_col, [1, 4, 5])
call_feature_columns(bucketized_col, {'col': tf.constant([1., 2., 3., 4., 5.])})
# + [markdown] id="PCYu-XtwXahx"
# In Keras, this can be replaced by `tf.keras.layers.Discretization`:
# + id="QK1WOG2uVVsL"
discretization_layer = tf.keras.layers.Discretization(bin_boundaries=[1, 4, 5])
one_hot_layer = tf.keras.layers.CategoryEncoding(
num_tokens=4, output_mode='one_hot')
one_hot_layer(discretization_layer([1., 2., 3., 4., 5.]))
# + [markdown] id="5bm9tJZAgpt4"
# ## One-hot encoding string data with a vocabulary
#
# Handling string features often requires a vocabulary lookup to translate strings into indices. Here is an example using feature columns to lookup strings and then one-hot encode the indices:
# + id="3fG_igjhukCO"
vocab_col = tf1.feature_column.categorical_column_with_vocabulary_list(
'sizes',
vocabulary_list=['small', 'medium', 'large'],
num_oov_buckets=0)
indicator_col = tf1.feature_column.indicator_column(vocab_col)
call_feature_columns(indicator_col, {'sizes': ['small', 'medium', 'large']})
# + [markdown] id="8rBgllRtY738"
# Using Keras preprocessing layers, use the `tf.keras.layers.StringLookup` layer with `output_mode` set to `'one_hot'`:
# + id="arnPlSrWvDMe"
string_lookup_layer = tf.keras.layers.StringLookup(
vocabulary=['small', 'medium', 'large'],
num_oov_indices=0,
output_mode='one_hot')
string_lookup_layer(['small', 'medium', 'large'])
# + [markdown] id="f76MVVYO8LB5"
# Note: For large one-hot encodings, it is much more efficient to use a sparse representation of the output. If you pass `sparse=True` to the `StringLookup` layer, the output of the layer will be a `tf.sparse.SparseTensor`, which can be efficiently handled as input to a `tf.keras.layers.Dense` layer.
# + [markdown] id="c1CmfSXQZHE5"
# ## Embedding string data with a vocabulary
#
# For larger vocabularies, an embedding is often needed for good performance. Here is an example embedding a string feature using feature columns:
# + id="C3RK4HFazxlU"
vocab_col = tf1.feature_column.categorical_column_with_vocabulary_list(
'col',
vocabulary_list=['small', 'medium', 'large'],
num_oov_buckets=0)
embedding_col = tf1.feature_column.embedding_column(vocab_col, 4)
call_feature_columns(embedding_col, {'col': ['small', 'medium', 'large']})
# + [markdown] id="3aTRVJ6qZZH0"
# Using Keras preprocessing layers, this can be achieved by combining a `tf.keras.layers.StringLookup` layer and an `tf.keras.layers.Embedding` layer. The default output for the `StringLookup` will be integer indices which can be fed directly into an embedding.
#
# Note: The `Embedding` layer contains trainable parameters. While the `StringLookup` layer can be applied to data inside or outside of a model, the `Embedding` must always be part of a trainable Keras model to train correctly.
# + id="8resGZPo0Fho"
string_lookup_layer = tf.keras.layers.StringLookup(
vocabulary=['small', 'medium', 'large'], num_oov_indices=0)
embedding = tf.keras.layers.Embedding(3, 4)
embedding(string_lookup_layer(['small', 'medium', 'large']))
# + [markdown] id="3I5loEx80MVm"
# ## Complete training example
#
# To show a complete training workflow, first prepare some data with three features of different types:
# + id="D_7nyBee0ZBV"
features = {
'type': [0, 1, 1],
'size': ['small', 'small', 'medium'],
'weight': [2.7, 1.8, 1.6],
}
labels = [1, 1, 0]
predict_features = {'type': [0], 'size': ['foo'], 'weight': [-0.7]}
# + [markdown] id="e_4Xx2c37lqD"
# Define some common constants for both TF1 and TF2 workflows:
# + id="3cyfQZ7z8jZh"
vocab = ['small', 'medium', 'large']
one_hot_dims = 3
embedding_dims = 4
weight_mean = 2.0
weight_variance = 1.0
# + [markdown] id="ywCgU7CMIfTH"
# ### With feature columns
#
# Feature columns must be passed as a list to the estimator on creation, and will be called implicitly during training.
# + id="Wsdhlm-uipr1"
categorical_col = tf1.feature_column.categorical_column_with_identity(
'type', num_buckets=one_hot_dims)
# Convert index to one-hot; e.g. [2] -> [0,0,1].
indicator_col = tf1.feature_column.indicator_column(categorical_col)
# Convert strings to indices; e.g. ['small'] -> [1].
vocab_col = tf1.feature_column.categorical_column_with_vocabulary_list(
'size', vocabulary_list=vocab, num_oov_buckets=1)
# Embed the indices.
embedding_col = tf1.feature_column.embedding_column(vocab_col, embedding_dims)
normalizer_fn = lambda x: (x - weight_mean) / math.sqrt(weight_variance)
# Normalize the numeric inputs; e.g. [2.0] -> [0.0].
numeric_col = tf1.feature_column.numeric_column(
'weight', normalizer_fn=normalizer_fn)
estimator = tf1.estimator.DNNClassifier(
feature_columns=[indicator_col, embedding_col, numeric_col],
hidden_units=[1])
def _input_fn():
return tf1.data.Dataset.from_tensor_slices((features, labels)).batch(1)
estimator.train(_input_fn)
# + [markdown] id="qPIeG_YtfNV1"
# The feature columns will also be used to transform input data when running inference on the model.
# + id="K-AIIB8CfSqt"
def _predict_fn():
return tf1.data.Dataset.from_tensor_slices(predict_features).batch(1)
next(estimator.predict(_predict_fn))
# + [markdown] id="baMA01cBIivo"
# ### With Keras preprocessing layers
#
# Keras preprocessing layers are more flexible in where they can be called. A layer can be applied directly to tensors, used inside a `tf.data` input pipeline, or built directly into a trainable Keras model.
#
# In this example, you will apply preprocessing layers inside a `tf.data` input pipeline. To do this, you can define a separate `tf.keras.Model` to preprocess your input features. This model is not trainable, but is a convenient way to group preprocessing layers.
# + id="NMz8RfMQdCZf"
inputs = {
'type': tf.keras.Input(shape=(), dtype='int64'),
'size': tf.keras.Input(shape=(), dtype='string'),
'weight': tf.keras.Input(shape=(), dtype='float32'),
}
# Convert index to one-hot; e.g. [2] -> [0,0,1].
type_output = tf.keras.layers.CategoryEncoding(
one_hot_dims, output_mode='one_hot')(inputs['type'])
# Convert size strings to indices; e.g. ['small'] -> [1].
size_output = tf.keras.layers.StringLookup(vocabulary=vocab)(inputs['size'])
# Normalize the numeric inputs; e.g. [2.0] -> [0.0].
weight_output = tf.keras.layers.Normalization(
axis=None, mean=weight_mean, variance=weight_variance)(inputs['weight'])
outputs = {
'type': type_output,
'size': size_output,
'weight': weight_output,
}
preprocessing_model = tf.keras.Model(inputs, outputs)
# + [markdown] id="NRfISnj3NGlW"
# Note: As an alternative to supplying a vocabulary and normalization statistics on layer creation, many preprocessing layers provide an `adapt()` method for learning layer state directly from the input data. See the [preprocessing guide](https://www.tensorflow.org/guide/keras/preprocessing_layers#the_adapt_method) for more details.
#
# You can now apply this model inside a call to `tf.data.Dataset.map`. Please note that the function passed to `map` will automatically be converted into
# a `tf.function`, and usual caveats for writing `tf.function` code apply (no side effects).
# + id="c_6xAUnbNREh"
# Apply the preprocessing in tf.data.Dataset.map.
dataset = tf.data.Dataset.from_tensor_slices((features, labels)).batch(1)
dataset = dataset.map(lambda x, y: (preprocessing_model(x), y),
num_parallel_calls=tf.data.AUTOTUNE)
# Display a preprocessed input sample.
next(dataset.take(1).as_numpy_iterator())
# + [markdown] id="8_4u3J4NdJ8R"
# Next, you can define a separate `Model` containing the trainable layers. Note how the inputs to this model now reflect the preprocessed feature types and shapes.
# + id="kC9OZO5ldmP-"
inputs = {
'type': tf.keras.Input(shape=(one_hot_dims,), dtype='float32'),
'size': tf.keras.Input(shape=(), dtype='int64'),
'weight': tf.keras.Input(shape=(), dtype='float32'),
}
# Since the embedding is trainable, it needs to be part of the training model.
embedding = tf.keras.layers.Embedding(len(vocab), embedding_dims)
outputs = tf.keras.layers.Concatenate()([
inputs['type'],
embedding(inputs['size']),
tf.expand_dims(inputs['weight'], -1),
])
outputs = tf.keras.layers.Dense(1)(outputs)
training_model = tf.keras.Model(inputs, outputs)
# + [markdown] id="ir-cn2H_d5R7"
# You can now train the `training_model` with `tf.keras.Model.fit`.
# + id="6TS3YJ2vnvlW"
# Train on the preprocessed data.
training_model.compile(
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True))
training_model.fit(dataset)
# + [markdown] id="pSaEbOE4ecsy"
# Finally, at inference time, it can be useful to combine these separate stages into a single model that handles raw feature inputs.
# + id="QHjbIZYneboO"
inputs = preprocessing_model.input
outpus = training_model(preprocessing_model(inputs))
inference_model = tf.keras.Model(inputs, outpus)
predict_dataset = tf.data.Dataset.from_tensor_slices(predict_features).batch(1)
inference_model.predict(predict_dataset)
# + [markdown] id="O01VQIxCWBxU"
# This composed model can be saved as a [SavedModel](https://www.tensorflow.org/guide/saved_model) for later use.
# + id="6tsyVZgh7Pve"
inference_model.save('model')
restored_model = tf.keras.models.load_model('model')
restored_model.predict(predict_dataset)
# + [markdown] id="IXMBwzggwUjI"
# Note: Preprocessing layers are not trainable, which allows you to apply them *asynchronously* using `tf.data`. This has performence benefits, as you can both [prefetch](https://www.tensorflow.org/guide/data_performance#prefetching) preprocessed batches, and free up any accelerators to focus on the differentiable parts of a model. As this guide shows, seperating preprocessing during training and composing it during inference is a flexible way to leverage these performance gains. However, if your model is small or preprocessing time is negligable, it may be simpler to build preprocessing into a complete model from the start. To do this you can build a single model starting with `tf.keras.Input`, followed by preprocessing layers, followed by trainable layers.
# + [markdown] id="2pjp7Z18gRCQ"
# ## Feature column equivalence table
#
# For reference, here is an approximate correspondence between feature columns and
# preprocessing layers:<table>
# <tr>
# <th>Feature Column</th>
# <th>Keras Layer</th>
# </tr>
# <tr>
# <td>`feature_column.bucketized_column`</td>
# <td>`layers.Discretization`</td>
# </tr>
# <tr>
# <td>`feature_column.categorical_column_with_hash_bucket`</td>
# <td>`layers.Hashing`</td>
# </tr>
# <tr>
# <td>`feature_column.categorical_column_with_identity`</td>
# <td>`layers.CategoryEncoding`</td>
# </tr>
# <tr>
# <td>`feature_column.categorical_column_with_vocabulary_file`</td>
# <td>`layers.StringLookup` or `layers.IntegerLookup`</td>
# </tr>
# <tr>
# <td>`feature_column.categorical_column_with_vocabulary_list`</td>
# <td>`layers.StringLookup` or `layers.IntegerLookup`</td>
# </tr>
# <tr>
# <td>`feature_column.crossed_column`</td>
# <td>Not implemented.</td>
# </tr>
# <tr>
# <td>`feature_column.embedding_column`</td>
# <td>`layers.Embedding`</td>
# </tr>
# <tr>
# <td>`feature_column.indicator_column`</td>
# <td>`output_mode='one_hot'` or `output_mode='multi_hot'`*</td>
# </tr>
# <tr>
# <td>`feature_column.numeric_column`</td>
# <td>`layers.Normalization`</td>
# </tr>
# <tr>
# <td>`feature_column.sequence_categorical_column_with_hash_bucket`</td>
# <td>`layers.Hashing`</td>
# </tr>
# <tr>
# <td>`feature_column.sequence_categorical_column_with_identity`</td>
# <td>`layers.CategoryEncoding`</td>
# </tr>
# <tr>
# <td>`feature_column.sequence_categorical_column_with_vocabulary_file`</td>
# <td>`layers.StringLookup`, `layers.IntegerLookup`, or `layer.TextVectorization`†</td>
# </tr>
# <tr>
# <td>`feature_column.sequence_categorical_column_with_vocabulary_list`</td>
# <td>`layers.StringLookup`, `layers.IntegerLookup`, or `layer.TextVectorization`†</td>
# </tr>
# <tr>
# <td>`feature_column.sequence_numeric_column`</td>
# <td>`layers.Normalization`</td>
# </tr>
# <tr>
# <td>`feature_column.weighted_categorical_column`</td>
# <td>`layers.CategoryEncoding`</td>
# </tr>
# </table>
#
# \* `output_mode` can be passed to `layers.CategoryEncoding`, `layers.StringLookup`, `layers.IntegerLookup`, and `layers.TextVectorization`.
#
# † `layers.TextVectorization` can handle freeform text input directly (e.g. entire sentences or paragraphs). This is not one-to-one replacement for categorical sequence handling in TF1, but may offer a convinient replacement for ad-hoc text preprocessing.
# + [markdown] id="AQCJ6lM3YDq_"
# ## Next Steps
#
# - For more information on keras preprocessing layers, see [the guide to preprocessing layers](https://www.tensorflow.org/guide/keras/preprocessing_layers).
# - For a more in-depth example of applying preprocessing layers to structured data, see [the structured data tutorial](https://www.tensorflow.org/tutorials/structured_data/preprocessing_layers).
| site/en-snapshot/guide/migrate/migrating_feature_columns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.1 64-bit
# language: python
# name: python3
# ---
### Importando bibliotecas
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
import warnings
warnings.filterwarnings('ignore')
# #### Importando e analisando dados
df = pd.read_csv('https://assets.datacamp.com/production/repositories/628/datasets/444cdbf175d5fbf564b564bd36ac21740627a834/diabetes.csv', sep=',')
X = df.drop(['diabetes'], axis=1)
y = df['diabetes']
y = pd.DataFrame(y)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state=42)
logreg = LogisticRegression()
logreg.fit(X_train, y_train)
### Previsão
y_predic = logreg.predict(X_test)
### Analisando acurácia
acur = logreg.score(X_test, y_test)
acur
### Analisando acurácia
print(confusion_matrix(y_test, y_predic))
print('\n')
print(classification_report(y_test, y_predic))
# #### Curva ROC
from sklearn.metrics import roc_curve
y_prob = logreg.predict_log_proba(X_test)[:,1]
fpr, tpr, thresholds = roc_curve(y_test, y_prob)
### Plotando gráfico
plt.style.use('ggplot')
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr, tpr)
plt.xlabel('Falso Positivo')
plt.ylabel('Verdadeiro Positivo')
plt.title('Curva ROC')
plt.show()
# #### AUC
### Importando bibliotecas
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import cross_val_score
roc_auc_score(y_test, y_prob)
cross_val_score(logreg, X, y, cv=4, scoring='roc_auc')
| Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from tensorflow.keras.utils import to_categorical
import tensorflow.keras.datasets as datasets
from tensorflow.keras.layers import Conv2D, MaxPool2D, Dense, Dropout, Flatten, Reshape, Activation, BatchNormalization
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import SGD, Adam
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# -
MAX_WORDS = 10000
# +
np_load_old = np.load
# modify the default parameters of np.load
np.load = lambda *a,**k: np_load_old(*a, allow_pickle=True, **k)
(train_words, train_labels), (test_words, test_labels) = datasets.reuters.load_data(num_words=MAX_WORDS)
print(type(train_words), train_words.shape)
print(type(train_labels), train_labels.shape)
# restore np.load for future normal usage
np.load = np_load_old
max_val = 0
for i in range(len(train_words)):
if max_val < max(train_words[i]):
max_val = max(train_words[i])
print('Maximum word value: ', max_val)
# -
word_index = datasets.reuters.get_word_index()
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
decoded_newswire = ' '.join([reverse_word_index.get(i-3, '?') for i in train_words[0]])
print(decoded_newswire)
def vectorize_sequences(sequences, dimensions=MAX_WORDS):
output = np.zeros((len(sequences), dimensions))
for i, sequence in enumerate(sequences):
output[i, sequence] = 1
return output
# +
x_train = vectorize_sequences(train_words)
x_test = vectorize_sequences(test_words)
y_train = to_categorical(train_labels)
y_test = to_categorical(test_labels)
print(x_train.shape, x_test.shape)
print(y_train.shape, y_test.shape)
# -
print(x_train[0], sum(x_train[0]))
model = Sequential([Dense(64, activation='relu', input_shape=(MAX_WORDS,)),
Dropout(0.5),
Dense(64, activation='relu'),
# Dense(128, activation='relu'),
# Dropout(0.5),
# Dense(128, activation='relu'),
Dense(46, activation='softmax')])
model.summary()
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(x_train, y_train, epochs=40, batch_size=512, validation_data=(x_test, y_test))
train_loss = history.history['loss']
test_loss = history.history['val_loss']
train_acc = history.history['acc']
test_acc = history.history['val_acc']
plt.plot(list(range(len(train_loss))), train_loss, label='Training loss')
plt.plot(list(range(len(test_loss))), test_loss, label='Test loss')
plt.legend()
plt.plot(list(range(len(train_loss))), train_acc, label='Training accuracy')
plt.plot(list(range(len(test_loss))), test_acc, label='Test accuracy')
plt.legend()
| mlp/keras/keras_reuters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IPython extension for drawing circuit diagrams with LaTeX/Circuitikz
# Created by <NAME>
# This fork by <NAME>
#
# https://github.com/mopfeil/tikzmagic
# ## Requirements
# This IPython magic command uses the following external dependencies: `pdflatex`, `pdfcrop`, the [Circuitikz](http://www.ctan.org/pkg/circuitikz) package and
#
# * for PNG output: `convert` from ImageMagick and `Ghostscript`
# * for SVG output: `pdf2svg`
# ## Installation
# Clone the tikzmagic repository into a directory of your own. Assume that it resides in 'C:/Users/mopfe/Dropbox/HRW/ipython_circuitikz/
import os
import sys
try:
import ipython_circuitikz.circuitikz
except ImportError:
# Assuming CWD is C:/user/mopfeil/ipython_circuitikz/
assert os.path.isfile('C:/Users/mopfe/Dropbox/HRW/ipython_circuitikz/circuitikz.py')
sys.path.append('C:/Users/mopfe/Dropbox/HRW/')
import ipython_circuitikz.circuitikz
# ## Load the extension
# %reload_ext circuitikz
# ## Example: SQUID
# +
# %%circuitikz filename=squid dpi=125
\begin{circuitikz}[scale=1]
\draw ( 0, 0) [short, *-] node[anchor=south] {$\Phi_J$} to (0, -1);
% right
\draw ( 0, -1) to (2, -1) to node[anchor=west] {$\Phi_{J}^2$} (2, -2) to (3, -2)
to [barrier, l=$E_J^2$] (3, -4) to (2, -4)to (2, -5) to (0, -5) node[ground] {};
\draw ( 2, -2) to (1, -2) to [capacitor, l=$C_J^2$] (1, -4) to (1, -4) to (2, -4);
% left
\draw ( 0, -1) to (-2, -1) to node[anchor=west] {$\Phi_{J}^1$} (-2, -2) to (-3, -2)
to [capacitor, l=$C_J^1$] (-3, -4) to (-2, -4) to (-2, -5) to (0, -5);
\draw (-2, -2) to (-1, -2) to [barrier, l=$E_J^1$] (-1, -4) to (-1, -4) to (-2, -4);
\end{circuitikz}
# -
# ## Example: Transmission line
# +
# %%circuitikz filename=tm dpi=150
\begin{circuitikz}[scale=1.25]
\draw (-1,0) node[anchor=east] {} to [short, *-*] (1,0);
\draw (-1,2) node[anchor=east] {} to [inductor, *-*, l=$\Delta x L$] (1,2);
\draw (-1,0) to [open, l=$\cdots$] (-1,2);
\draw (3, 0) to (1, 0) to [capacitor, l=$\Delta x C$, *-*] (1, 2) to [inductor, *-*, l=$\Delta x L$] (3, 2);
\draw (5, 0) to (3, 0) to [capacitor, l=$\Delta x C$, *-*] (3, 2) to [inductor, *-*, l=$\Delta x L$] (5, 2);
\draw (7, 0) to (5, 0) to [capacitor, l=$\Delta x C$, *-*] (5, 2) to [inductor, *-*, l=$\Delta x L$] (7, 2);
\draw (9, 0) to (7, 0) to [capacitor, l=$\Delta x C$, *-*] (7, 2) to [inductor, *-*, l=$\Delta x L$] (9, 2);
\draw (9,0) node[anchor=east] {} to [short, *-*] (9,0);
\draw (10,0) to [open, l=$\cdots$] (10,2);
\end{circuitikz}
| Circuitikz-examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysing the final propositions generated from VisDial dataset and used in the experiments
# +
import copy
import itertools
import json
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import pandas as pd
import seaborn as sns
import random
from collections import Counter
# %matplotlib inline
sns.set_style("darkgrid")
plt.rcParams['font.family'] = 'DejaVu Sans'
# +
PATH_VISDIAL = 'data/visual_dialog/'
DIR = 'propositions/'
SPLITS = ('train', 'val', 'test')
# -
# Select either original to analyse all or downsampled/balanced to analyse the final versions used in the paper.
which_probes = {'train': 'downsampled-balanced-', 'val': 'downsampled-', 'test': 'downsampled-'}
# Load datasets:
# +
datasets = {}
visdial = {}
propositions = {}
for split in SPLITS:
with open(f'{PATH_VISDIAL}visdial_1.0_{split}.json', 'r') as data:
visdial[split] = json.load(data)
datasets[split] = visdial[split]['data']['dialogs']
for split in SPLITS:
with open(f'{DIR}{which_probes[split]}propositions_{split}.json', 'r') as data:
props = json.load(data)
propositions[split] = props['dialogues']
# -
# Checking how many of the original dialogues ended up in the samples:
print('Train: ', len([k for k,v in propositions['train'].items() if v]))
print('Val: ', len([k for k,v in propositions['val'].items() if v]))
print('Test: ', len([k for k,v in propositions['test'].items() if v]))
# We'll remove the empty, unused dialogues from the analysis:
propositions['train'] = {k:v for k,v in propositions['train'].items() if v !={}}
propositions['val'] = {k:v for k,v in propositions['val'].items() if v !={}}
propositions['test'] = {k:v for k,v in propositions['test'].items() if v !={}}
# ### Examining an example of generated probes for a dialogue:
def get_vd_dialogue(vd_dialog, split, add_punct=False):
caption = vd_dialog['caption']
if add_punct:
caption += '.'
turns = []
for qa in vd_dialog['dialog']:
q = visdial[split]['data']['questions'][qa['question']]
if add_punct:
q += '?'
try:
a = visdial[split]['data']['answers'][qa['answer']]
if add_punct:
a += '.'
except KeyError:
a = ''
turns.append((q, a))
return caption, turns
# Pick a random dialogue:
split = 'val'
ID = random.randint(0, len(datasets[split])) # 46907
print(ID)
print(visdial[split]['data']['dialogs'][ID]['image_id'])
# +
caption, turns = get_vd_dialogue(datasets[split][ID], split, add_punct=True)
dialogue = [('', caption)] + turns
probes = [[] for n in range(11)]
for p in propositions[split][str(ID)].values():
turn = int(p['turn_shared'])
probes[turn].append(p['proposition'])
for n in range(11):
print(f'{n} {dialogue[n][0]} {dialogue[n][1]}')
if probes[n]:
for i in range(0, len(probes[n]), 2):
print(f'\t{probes[n][i]}\n\t{probes[n][i+1]}')
else:
print(' --')
# -
# # Analysis of the resulting propositions
# ## Manipulated turns per dialogue
# How many turns in a dialogue have been turned into propositions?
# +
manipulated_turns = {}
mean_manip_turns = {}
df_manip_turns = {}
keys = list(range(1, 12))
for split in SPLITS:
manipulated_turns[split] = Counter([len(set([p['turn_shared'] for p in dialogue.values()]))
for dialogue in propositions[split].values()])
mean_manip_turns[split] = sum([turns*freq for turns, freq in manipulated_turns[split].items()]) / sum(manipulated_turns[split].values())
df_manip_turns[split] = pd.DataFrame([manipulated_turns[split][i] for i in keys], index=keys, columns=['frequency']).transpose()
print(f'On average, the {split} set has {mean_manip_turns[split]} manipulated turns per dialogue.')
# +
fig, axes = plt.subplots(1, 3, figsize=(13,3))
#fig.suptitle('How many turns in a dialogue were turned into propositions')
for i, (ax, split) in enumerate(zip(axes, SPLITS)):
sns.barplot(data=df_manip_turns[split], ax=ax, palette='rocket_r')
ax.set_title(split, fontsize=16)
ax.tick_params(labelsize=14)
if i != 1:
ax.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: '{:,.0f}'.format(x/1000) + 'K' if x != 0 else 0))
if i == 0:
ax.set_ylabel('n dialogues', fontsize=16)
if i == 1:
ax.set_xlabel('n manipulated turns', fontsize=16)
split = 'train'
plt.savefig(f'plots/howmany-turns-{which_probes[split]}{split}.pdf', bbox_inches="tight")
plt.show()
# -
# The test set has a different distribution because it has incomplete dialogues. The balanced train set has a different distribution because of the sampling.
# ## At which turn propositions shift from private to shared
shift_turn = {}
df_shift_turn = {}
keys = list(range(11))
for split in SPLITS:
shift_turn[split]= Counter([p['turn_shared'] for dialogue in propositions[split].values()
for p in dialogue.values()])
df_shift_turn[split] = pd.DataFrame([shift_turn[split][i] for i in keys], index=keys, columns=['frequency']).transpose()
# The point where private/shared is balanced is between turn 5-6 on the balanced training set and at turn 3 on the original set (due to the many caption propositions).
# +
split = 'train'
total = sum([value for key, value in shift_turn[split].items()])
print(total)
cum = 0
for i in range(11):
cum += shift_turn[split][i]
print(i, cum / total)
# +
fig, axes = plt.subplots(1, 3, figsize=(13,3))
#fig.suptitle('Which QA pair was manipulated?')
for i, (ax, split) in enumerate(zip(axes, SPLITS)):
sns.barplot(data=df_shift_turn[split], ax=ax, palette='dark:salmon_r')
ax.set_title(split, fontsize=16)
ax.tick_params(labelsize=14)
if i == 0:
ax.set_ylabel('frequency', fontsize=16)
ax.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: '{:,.0f}'.format(x/1000) + 'K' if x != 0 else 0))
if i == 1:
ax.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: '{:,.1f}'.format(x/1000) + 'K' if x != 0 else 0))
ax.set_xlabel('turn', fontsize=16)
if i == 2:
ax.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: '{:,.0f}'.format(x/1000) + 'K' if x != 0 else 0))
split = 'train'
plt.savefig(f'plots/which-turn-{which_probes[split]}{split}.pdf', bbox_inches="tight")
#plt.savefig('propositions/original-propositions_turn-distribution.png', bbox_inches="tight")
plt.show()
# -
# Constructing the balanced train set causes the distribution so be slightly skewed towards later turns... this will induce some bias on the private dimension, but mid point is still between turn 5 and 6.
# ## Propositions per dialogue
# How many propositions were generated for each dialogue?
# +
n_propositions = {}
mean_n_props = {}
df_n_props = {}
N_MAX = 50
keys = list(range(N_MAX))
for split in SPLITS:
n_propositions[split] = Counter([len(dialogue.values()) for dialogue in propositions[split].values()])
df_n_props[split] = pd.DataFrame([n_propositions[split][i] for i in keys], index=keys, columns=['frequency']).transpose()
mean_n_props[split] = sum([turns*freq for turns, freq in n_propositions[split].items()]) / len(propositions[split])
print(f'On average, the {split} set has {mean_n_props[split]} propositions per dialogue.')
min_p = min(n_propositions[split].keys())
max_p = max(n_propositions[split].keys())
print(f'The {split} set has dialogues with minimum {min_p} and maximum {max_p} propositions. \n')
# +
fig, axes = plt.subplots(1, 3, figsize=(15,5))
fig.suptitle('Propositions per dialogue')
for i, (ax, split) in enumerate(zip(axes, SPLITS)):
g = sns.barplot(data=df_n_props[split], ax=ax, palette='rocket_r')
g.set_xticks([0, 10, 20, 30, 40])
g.set_xticklabels(['0', '10', '20', '30', '40'])
if i == 0:
ax.set(ylabel='frequency')
if i == 1:
ax.set(xlabel='n propositions')
plt.show()
# -
# n_propositions is always an even number by construction. The balanced train set differs because of the sampling that does not guarantee the occurrence of both entailment+contradiction cases for all probes (which is good to reduce bias).
# ## Polarity of original answers
# - Positive: propositions deriving from questions whose answers in the dialogue were yes.
# - Negative: propositions deriving from questions whose answers in the dialogue were no.
# - Nan: propositions deriving from captions or questions that were not polar questions (e.g. what color is the dog? black); can be also considered positive facts about the image.
# +
prop_polarity = {}
for split in SPLITS:
prop_polarity[split] = Counter([p['qa_fact'] for dialogue in propositions[split].values()
for p in dialogue.values()])
polarities = prop_polarity['train'].keys()
d_polar = {split: [prop_polarity[split][p]*100 / sum(prop_polarity[split].values()) for p in polarities] for split in SPLITS}
# -
df_polarity = pd.DataFrame(data=d_polar, index=polarities)
df_polarity.style.set_caption("Percentage of QA polarity for the generated propositions")
df_polarity
# The balanced train has has fewer probes coming from rules that have no polarity, because it's harder to find a match for them in the balanced set. But this is good because it keeps the training set balanced with respect to positive and negative facts about the images.
# ## Types of rules
# +
prop_rule = {}
for split in SPLITS:
prop_rule[split] = Counter([p['rule'] for dialogue in propositions[split].values() for p in dialogue.values()])
print(f'There are {len(prop_rule[split])} rules on the {split} set.')
rules = set([x for split in SPLITS for x in prop_rule[split].keys()])
d_rules = {split: [prop_rule[split][r]*100 / sum(prop_rule[split].values()) for r in rules] for split in SPLITS}
# -
df_rules = pd.DataFrame(data=d_rules, index=rules)
df_rules.style.set_caption("Percentage of rules used to generate propositions")
df_rules
# ## Turns
# Proportion of probes deriving from each of the 0+10 dialogue turns.
# +
prop_turns = {}
for split in SPLITS:
prop_turns[split] = Counter([p['turn_shared'] for dialogue in propositions[split].values() for p in dialogue.values()])
turns = list(range(0, 11))
d_turns = {split: [prop_turns[split][t]*100 / sum(prop_turns[split].values()) for t in turns] for split in SPLITS}
df_turns = pd.DataFrame(data=d_turns, index=turns)
df_turns.style.set_caption("Proportion of turns from which a proposition derives")
df_turns
# -
print(df_turns.to_latex())
# ## Rules vs types
# On train set:
# +
cross_dic = {'train': {}, 'val':{}, 'test':{}}
for split in SPLITS:
for dialogue in propositions[split].values():
for p in dialogue.values():
rule = p['rule']
if rule not in cross_dic[split]:
cross_dic[split][rule] = []
cross_dic[split][rule].append(p['qa_fact'])
split = 'train'
d_cross = {
'positive_polarity': [100*Counter(cross_dic[split][rule])['positive'] / len(cross_dic[split][rule]) for rule in rules],
'negative_polarity': [100*Counter(cross_dic[split][rule])['negative'] / len(cross_dic[split][rule]) for rule in rules],
'neutral': [100*Counter(cross_dic[split][rule])[None] / len(cross_dic[split][rule]) for rule in rules]
}
# -
df = pd.DataFrame(data=d_cross, index=rules)
df.style.set_caption("Distribution of polarity per rule")
df
# ### A thinks it's true or false?
# This is balanced by constrution. Each probe has an entailment and a contradiction. The balanced training set also ensures that for each probe that is an entailment of A's perpective with respect an image, another case is included where the same probe contradict A's perspective with respect to another image.
# +
prop_a_thinks_TF = {}
for split in SPLITS:
prop_a_thinks_TF[split] = Counter([p['a_thinks_true'] for dialogue in propositions[split].values() for p in dialogue.values()])
TF = [0, 1]
d_a_thinks_TF = {split: [prop_a_thinks_TF[split][t]*100 / sum(prop_a_thinks_TF[split].values()) for t in TF] for split in SPLITS}
df_a_thinks_TF = pd.DataFrame(data=d_a_thinks_TF, index=TF)
df_a_thinks_TF.style.set_caption("Proportion of propositions that the answerer considers true or false")
df_a_thinks_TF
# -
# ## Diversity
uniq_props = {}
for split in SPLITS:
uniq_props[split] = Counter(p['proposition'] for dialogue in propositions[split].values() for p in dialogue.values())
# Proportion of most common propositions on train set:
for split in SPLITS:
print(split.upper())
total = sum(uniq_props[split].values())
for p, freq in uniq_props[split].most_common(20):
print(round(100*freq/total, 2), '\t', p)
for split in ['val', 'test']:
proportion = 100*len(set(uniq_props[split].keys()).intersection(uniq_props['train'].keys())) / len(uniq_props[split])
print(f'{proportion}% of the {split} propositions appear on the train propositions.')
# ## Sizes
vocab = Counter()
separate_vocab = {'train': Counter(), 'val': Counter(), 'test': Counter()}
for split in ['train', 'val', 'test']:
for d, dialogue in propositions[split].items():
for p, prop in dialogue.items():
vocab.update(prop['proposition'].strip('.').split())
separate_vocab[split].update(prop['proposition'].strip('.').split())
# +
size = {}
vocabs = {}
number_of_props = {}
for split in SPLITS:
size[split] = len(propositions[split])
vocabs[split] = len(separate_vocab[split])
number_of_props[split] = sum([len(dialogue.values()) for dialogue in propositions[split].values()])
# -
datapoints = {}
datapoints['train'] = number_of_props['train'] * 11
datapoints['val'] = number_of_props['val'] * 11
# +
total_datapoints_test = 0
with open('propositions/visdial_1.0_test_dialogueLens.txt', 'r') as file:
lines = file.readlines()
for line in lines:
idx, dialogue_len = line.strip('\n').split('\t')
if idx in propositions['test']:
n_props = len(propositions['test'][idx])
total_datapoints_test += n_props * int(dialogue_len)
datapoints['test'] = total_datapoints_test
# +
types = ['dialogues', 'propositions', 'proposition types', 'datapoints', 'vocab size']
data_table = {split: [size[split], number_of_props[split], len(uniq_props[split]), datapoints[split], vocabs[split]]
for split in SPLITS}
# -
df = pd.DataFrame(data=data_table, index=types)
df.style.set_caption("Dataset sizes")
df
print(df.to_latex())
# ## Classes distribution
# Verifying the distribution of classes on the splits:
# +
# Functions from the main_task dataloader class
labels = {(1, 1): 0, # 'a thinks true and shared'
(0, 1): 1, # 'a thinks false and shared'
(1, 0): 2, # 'a thinks true and private'
(0, 0): 3, # 'a thinks false and private'
}
id2labels = {0: 'a_thinks_true and shared', 1: 'a_thinks_false and shared',
2: 'a_thinks_true and private', 3: 'a_thinks_false and private'}
def create_labels(a_thinks_true, turn, n_turns):
lbs = [labels[(a_thinks_true, 0)] if x < turn else labels[(a_thinks_true, 1)]
for x in range(n_turns)]
return lbs
def create_new_items(n_items, n_turns, labels, idx_d, idx_p):
"""Return a list with new datapoints."""
# global id, unique identifier for each element
ids = list(range(n_items, n_items + n_turns))
# original dialogue id
d_ids = [int(idx_d)] * n_turns
# original proposition/probe id
p_ids = [int(idx_p)] * n_turns
# dialogue turns
turns = list(range(n_turns))
return zip(ids, d_ids, p_ids, turns, labels)
# +
datapoint_classes = {'train': {}, 'val':{}, 'test':{}}
n_datapoints_test = 0
for split in ('train', 'val', 'test'):
for idx_d, dialogue in propositions[split].items():
n_turns = len(datasets[split][int(idx_d)]['dialog']) + 1 # add one because of caption
if split == 'test':
n_turns -= 1 # because last question has no answer
for idx_p, prop in dialogue.items():
a_thinks_true = prop['a_thinks_true']
turn_shared = prop['turn_shared']
classes = create_labels(a_thinks_true, turn_shared, n_turns)
n_items = len(datapoint_classes[split])
new_items = create_new_items(n_items, n_turns, classes, idx_d, idx_p)
datapoint_classes[split].update({x[0]: x[1:] for x in new_items})
if split == 'test':
n_datapoints_test += (n_turns * len(dialogue))
assert len(datapoint_classes['val']) == np.sum(number_of_props['val']) * 11
assert len(datapoint_classes['train']) == np.sum(number_of_props['train']) * 11
assert len(datapoint_classes['test']) == n_datapoints_test
# -
# Checking why valid labels are not more balanced. It's because of the captions.
c_val = Counter([p['turn_shared'] for props in propositions['val'].values() for p in props.values()])
# Shared datapoints:
[c_val[i]*(11-i) for i in range(11)]
# Private datapoints:
[c_val[i]*(i) for i in range(11)]
# Captions are always shared, so the first element is 0 in the private datapoints.
# +
idx_of_class = 3 # in tuples (d_ids, p_ids, turns, labels)
classes_train = Counter([c[idx_of_class] for c in datapoint_classes['train'].values()])
classes_val = Counter([c[idx_of_class] for c in datapoint_classes['val'].values()])
classes_test = Counter([c[idx_of_class] for c in datapoint_classes['test'].values()])
# +
types = [key for key in classes_train.keys()]
d = {'train': [100*classes_train[c]/len(datapoint_classes['train']) for c in types],
'val': [100*classes_val[c]/len(datapoint_classes['val']) for c in types],
'test': [100*classes_test[c]/len(datapoint_classes['test']) for c in types]
}
types = [id2labels[key] for key in types]
# -
df = pd.DataFrame(data=d, index=types)
df.style.set_caption("Distribution of classes")
df
print(df.to_latex())
print('There are {:,} items on the train set, {:,} on the val set and {:,} on the test set.'.format(
len(datapoint_classes['train']), len(datapoint_classes['val']), len(datapoint_classes['test'])))
# +
classes_per_turn = {'train': {x:Counter() for x in range(11)},
'val':{x:Counter() for x in range(11)},
'test':{x:Counter() for x in range(11)}}
for split in ('train', 'val', 'test'):
for global_idx, (d_idx, p_idx, turn, label) in datapoint_classes[split].items():
classes_per_turn[split][turn].update([label])
# -
# On turn 10, everything is shared. On turn 0, everything is private except for captions.
# +
indexes = [0, 1, 2, 3]
split = 'train'
df = pd.DataFrame(columns=[id2labels[x] for x in indexes])
for turn in range(11):
total = sum(classes_per_turn[split][turn].values())
values = [100*classes_per_turn[split][turn][idx] / total for idx in indexes]
df.loc[turn] = values
df
# -
# There is a roughly balanced division between private and shared around turns 5-6 on the training set.
# +
indexes = [0, 1, 2, 3]
split = 'val'
df = pd.DataFrame(columns=[id2labels[x] for x in indexes])
for turn in range(11):
total = sum(classes_per_turn[split][turn].values())
values = [100*classes_per_turn[split][turn][idx] / total for idx in indexes]
df.loc[turn] = values
df
# -
# There is a roughly balanced division between private and shared around turns 4-5 on the training set.
# +
indexes = [0, 1, 2, 3]
split = 'test'
df = pd.DataFrame(columns=[id2labels[x] for x in indexes])
for turn in range(10):
total = sum(classes_per_turn[split][turn].values())
values = [100*classes_per_turn[split][turn][idx] / total for idx in indexes]
df.loc[turn] = values
df
# -
# The test set has no turn 10, because it is a question that has no published answer on VisDial competition.
# ## In how many dialogues each proposition appears
# +
sentences = {'train': {}, 'val': {}, 'test': {}}
sentences_status = {'train': {}, 'val': {}, 'test': {}}
sent_counters = {}
for split in SPLITS:
for dialogue in propositions[split].values():
for p in dialogue.values():
if p['proposition'] not in sentences[split]:
sentences[split][p['proposition']] = 0
sentences_status[split][p['proposition']] = []
sentences[split][p['proposition']] += 1
sentences_status[split][p['proposition']].append(p['a_thinks_true'])
sent_counters[split] = Counter(list(sentences[split].values()))
# -
singletons = {}
for split in SPLITS:
singletons[split] = len([x for x in sentences[split].values() if x == 1])
x = 100 * singletons[split] / len(sentences[split])
print(f'{x}% of the {split} propositions occur in only one dialogue.')
sentences_with_fixed_truefalse_status = set()
for split in SPLITS:
for sentence, status_list in sentences_status[split].items():
if len(set(status_list)) == 1:
sentences_with_fixed_truefalse_status.add(sentence)
immutable_sentences = len(sentences_with_fixed_truefalse_status)
all_sentences = len(sentences['train']) + len(sentences['val']) + len(sentences['test'])
print('{:.2f}% of the sentences are either always considered to be true or always considered to be false by A accross dialogues.'.format(100*immutable_sentences / all_sentences))
# ignore singletons whose status cannot be more than one anyway
frequent_sentences_with_fixed_truefalse_status = set()
for split in SPLITS:
for sentence, status_list in sentences_status[split].items():
if len(set(status_list)) == 1 and len(status_list) != 1:
frequent_sentences_with_fixed_truefalse_status.add(sentence)
frequent_immutable_sentences = len(frequent_sentences_with_fixed_truefalse_status)
print('{:.2f}% of the sentences that appear more than once are either always considered to be true or always considered to be false by A accross dialogues.'.format(100*frequent_immutable_sentences / all_sentences))
# So, in general, propositions occur only in one dialogue. Among those that occur across dialogues, a small portion has a fixed true/false status that could be learnt by heart by the model. But this should not be an issue on the balanced train set.
print('On average, a proposition appears in {} dialogues in the training set'.format(np.mean(list(sentences['train'].values()))))
print('On average, a proposition appears in {} dialogues in the valid set'.format(np.mean(list(sentences['val'].values()))))
print('On average, a proposition appears in {} dialogues in the test set'.format(np.mean(list(sentences['test'].values()))))
# ## Coreference resolution
with open('data/coref/stats.json', 'r') as data:
corefs = json.load(data)
with open('data/coref/fails.json', 'r') as data:
failed_corefs = json.load(data)
print('On average, {:.3f} pronouns were solved per dialogue on the training set, {:.3f} on the val set and {:.3f} on the test set.'.format(
np.mean(list(corefs['train'].values())),
np.mean(list(corefs['val'].values())),
np.mean(list(corefs['test'].values()))
))
# ## Vocabulary
len(vocab)
vocab.most_common(100)
# Common vocabulary among each set:
print('{}% of val words in train words'.format(100*len(set(separate_vocab['train'].keys()) & set(separate_vocab['val'].keys())) / len(separate_vocab['val'].keys())))
print('{}% of val words in train words'.format(100*len(set(separate_vocab['train'].keys()) & set(separate_vocab['test'].keys())) / len(separate_vocab['test'].keys())))
max_len = 0
for split in ['train', 'val', 'test']:
for d, dialogue in propositions[split].items():
for p, prop in dialogue.items():
sentence = prop['proposition'].split()
if len(sentence) > max_len:
max_len = len(sentence)
# Maximum length of 15 tokens was enforced:
assert max_len == 15
# Checking longish sentences:
for split in ['train', 'val', 'test']:
for d, dialogue in propositions[split].items():
for p, prop in dialogue.items():
sentence = prop['proposition']
if len(sentence.split()) > 10:
print(sentence)
# # What causes the bias in the private dimension for balanced train set
# Checking the balanced probes:
# +
split = 'train'
a_thinks_true = []
sents = []
turns = []
for idx_d, dialogue in propositions[split].items():
for idx_p, prop in dialogue.items():
a_thinks_true.append(prop['a_thinks_true'])
sents.append(prop['proposition'])
turns.append(prop['turn_shared'])
turns_shared = list(zip(sents, turns))
# -
# Select a probe:
# +
s = 'there are no buildings.'
example = Counter()
for x in turns_shared:
if x[0] == s:
example.update([x[1]])
plt.bar(example.keys(), example.values())
plt.xlabel('manipulated turn')
plt.ylabel('# probes in training set')
plt.title(s)
plt.show()
# -
# Plot a few examples for the paper:
# +
sents = ['there are no buildings.', 'the sky is blue.', 'there are no people.', 'there are windows.',
'the image is not in color.', 'there are trees.']
fig, axes = plt.subplots(2, 3, figsize=(15, 10))
# https://stackoverflow.com/questions/6541123/improve-subplot-size-spacing-with-many-subplots-in-matplotlib
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=None, hspace=0.3)
for i, ax in enumerate(axes.flatten()):
s = sents[i]
example = Counter()
for x in turns_shared:
if x[0] == s:
example.update([x[1]])
ax = sns.barplot(x=list(example.keys()), y=list(example.values()), palette="Oranges_d", ax=ax)
if i == 4:
ax.set_xlabel('manipulated turn', fontsize=14)
if i == 0 or i == 3:
ax.set_ylabel('# probes in training set', fontsize=14)
ax.set_title(s, fontsize=14)
# split = 'train'
#plt.savefig(f'plots/props-skewedPS_{which_probes[split]}.pdf')
plt.show()
# +
sents = ['the image is in color.', 'there are trees.']
fig, axes = plt.subplots(1, 2, figsize=(9, 3))
# https://stackoverflow.com/questions/6541123/improve-subplot-size-spacing-with-many-subplots-in-matplotlib
plt.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=None, hspace=0.3)
for i, ax in enumerate(axes.flatten()):
s = sents[i]
example = Counter()
for x in turns_shared:
if x[0] == s:
example.update([x[1]])
ax = sns.barplot(x=list(example.keys()), y=list(example.values()), palette="Oranges_d", ax=ax)
ax.set_xlabel('turn', fontsize=14)
ax.tick_params(labelsize=14)
if i == 0 or i == 3:
ax.set_ylabel('# propositions (train)', fontsize=14)
ax.set_title(s, fontsize=18)
split = 'train'
plt.savefig(f'plots/props-skewedPS_{which_probes[split]}{split}.pdf', bbox_inches="tight")
plt.show()
# -
# Some probes have a distribution over manipulated turns that is skewed towards private (or shared). This may be a shorcut for the classifier.
# # Examples for the paper
# The image used in the paper is from the training data, index 8778.
ID = 8778
split = 'train'
get_vd_dialogue(visdial[split]['data']['dialogs'][ID], split)
# +
caption, turns = get_vd_dialogue(datasets[split][ID], split, add_punct=True)
dialogue = [('', caption)] + turns
probes = [[] for n in range(11)]
for p in propositions[split][str(ID)].values():
turn = int(p['turn_shared'])
probes[turn].append(p['proposition'])
for n in range(11):
print(f'{n} {dialogue[n][0]} {dialogue[n][1]}')
if probes[n]:
for i in range(0, len(probes[n]), 2):
print(f'\t{probes[n][i]}\n\t{probes[n][i+1]}')
else:
print(' --')
# +
labels = {(1, 1): 0, # 'a thinks true and shared'
(0, 1): 1, # 'a thinks false and shared'
(1, 0): 2, # 'a thinks true and private'
(0, 0): 3, # 'a thinks false and private'
}
label_names = {0: 'a thinks true \n shared', 1: 'a thinks false \n shared',
2: 'a thinks true \n private', 3: 'a thinks false \n private'}
# -
def get_vd_scoreboard(ID, split):
scorekeeping = np.zeros([11, len(propositions[split][str(ID)])])
caption = datasets[split][ID]['caption'] + '.'
turns = []
for qa in datasets[split][ID]['dialog']:
q = visdial[split]['data']['questions'][qa['question']] + '?'
a = visdial[split]['data']['answers'][qa['answer']] + '.'
turns.append((q, a))
turns = [('', caption)] + turns
probes = [[] for n in range(11)]
for p, prop in enumerate(propositions[split][str(ID)].values()):
turn = int(prop['turn_shared'])
tf = prop['a_thinks_true']
probes[turn].append(prop['proposition'])
scorekeeping[:, p] = np.array([[labels[(tf, 0)] if x < turn else labels[(tf, 1)]
for x in range(11)]])
return turns, probes, scorekeeping
# ### Examples of generated propositions on full dialogues for the appendix
# For this we'll use the original-downsampled training set.
split = 'train'
with open(f'{DIR}downsampled-propositions_{split}.json', 'r') as data:
props = json.load(data)
downsampled_train = props['dialogues']
for d, dialogue in enumerate(visdial[split]['data']['dialogs']):
if dialogue['image_id'] == 318405: #121622, 16677, 364032, 141086, 84859, 283921, 469898, 391229
ID = d
break
# +
caption, turns = get_vd_dialogue(visdial[split]['data']['dialogs'][ID], split)
dialogue = [('', caption)] + turns
probes = [[] for n in range(11)]
for p in downsampled_train[str(ID)].values():
turn = int(p['turn_shared'])
probes[turn].append(p['proposition'])
for n in range(11):
print(f'{dialogue[n][0]}? {dialogue[n][1]}.')
if probes[n]:
for i in range(0, len(probes[n]), 2):
print(f' {probes[n][i]}\n {probes[n][i+1]}')
else:
print(' none')
# -
# ### Example for the scorekeeping matrix:
# For this we use the original dataset.
with open(f'{DIR}original/propositions_{split}.json', 'r') as data:
props = json.load(data)
original_train = props['dialogues']
# rewrite from above, original_train is hardcoded!
def get_vd_original_scoreboard(ID, split):
scorekeeping = np.zeros([11, len(original_train[str(ID)])])
caption = datasets[split][ID]['caption'] + '.'
turns = []
for qa in datasets[split][ID]['dialog']:
q = visdial[split]['data']['questions'][qa['question']] + '?'
a = visdial[split]['data']['answers'][qa['answer']] + '.'
turns.append((q, a))
turns = [('', caption)] + turns
probes = [[] for n in range(11)]
for p, prop in enumerate(original_train[str(ID)].values()):
turn = int(prop['turn_shared'])
tf = prop['a_thinks_true']
probes[turn].append(prop['proposition'])
scorekeeping[:, p] = np.array([[labels[(tf, 0)] if x < turn else labels[(tf, 1)]
for x in range(11)]])
return turns, probes, scorekeeping
for d, dialogue in enumerate(visdial[split]['data']['dialogs']):
if dialogue['image_id'] == 176904: # 208589, 478148, 209419, 407235, 3926
ID = d
break
# +
turns, probes, scoreboard = get_vd_original_scoreboard(ID, split)
for n in range(11):
print(f'{turns[n][0]} {turns[n][1]}')
if probes[n]:
for i in range(0, len(probes[n]), 2):
print(f' {probes[n][i]}\n {probes[n][i+1]}')
else:
print(' none')
# -
columns = [q + ' ' + a for (q, a) in turns]
index = [x for p in probes for x in p]
df = pd.DataFrame(data=scoreboard.T, columns=columns, index=index)
# +
fig, ax1 = plt.subplots(1, 1, figsize=(7.5, 4))
cbar_ax = fig.add_axes([.2, .03, .6, .05])
custom_palette = sns.color_palette("Paired")[6:6+n]
vmap = {k: " and\n".join(label_names[k].split(' and ')) for k in range(4)}
n = len(vmap)
ax1.xaxis.set_ticks_position('top')
ax1.xaxis.set_label_position('top')
ax1.set_yticklabels(labels=ax1.get_yticklabels(), va='center', size=16)
matrix = sns.heatmap(df.transpose(), annot=False, cmap=custom_palette, ax=ax1, linewidths=0.1, cbar_ax=cbar_ax, cbar_kws={'orientation': 'horizontal'})
matrix.set_xticklabels(matrix.get_xticklabels(), rotation=55, ha='left', size=16)
# https://stackoverflow.com/questions/38836154/discrete-legend-in-seaborn-heatmap-plot
colorbar = ax1.collections[0].colorbar
r = colorbar.vmax - colorbar.vmin
colorbar.set_ticks([colorbar.vmin + 0.5 * r / (n) + r * i / (n) for i in range(n)])
colorbar.set_ticklabels(list(vmap.values()))
colorbar.ax.tick_params(labelsize=12)
plt.savefig('plots/orig-scoreboard-example.pdf', bbox_inches="tight")
plt.show()
| generating_propositions/analysis_final_probes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Molecular Modeling Software: OpenForceField"
# > Putting together open-source molecular modeling software.
#
# - toc: false
# - badges: true
# - comments: false
# - categories: [molecular modeling, scientific computing, grad school]
#
# The open-source molecular modelling community is a small (but growing) trend within academics.
# A lot of academics (professors, lab scientists, grad students) are putting together libraries and API that help fulfill a small task or purpose, and 21st century software engineering standards make them usable by others.
# Even in these early stages of open-source molecular modelling, these libraries are striving for interoperability, where two independently-developed API have gotten to the point where now they want to interact and communicate with each other.
#
# This is a very interesting point in time, where molecular modellers are now tasked with the effort of making many different libraries and API work together to successfully run simulations and complete research projects.
# Usually, scientists work within a singular software package that was designed by some core developers, and those scientists didn't need to venture outside that single software package, the license they paid for it, and the manual.
#
# With the release of [OpenForceField 1.0](https://openforcefield.org/news/introducing-openforcefield-1.0/), I was curious to use their SMRINOFF force field.
# To my understanding (don't quote me on this), the idea behind SMIRNOFF is to simplify molecular mechanics force fields, cut down on redundant atom types/parameters, and parametrize molecules based on "chemical perception" (chemical context and local bonding environment).
# Armed with these simplified, context-based force field methodologies, the frustrating, in-the-weeds obstacles associated with force field development might be ameliorated - the kinds of obstacles of which molecular modellers are painfully aware.
#
# Beyond the SMIRNOFF force field, I was curious how parameters might compare to the older OPLS all-atom force fields. As a personal challenge, I wanted to see how much "modern computational science" I could use, specifically trying to exercise the interoperability between different open-source molecular modelling packages.
# ## Building the our molecular system and model
# We begin with some imports. We can already see a variety of packages being used: mBuild, Foyer, ParmEd, OpenForceField, Simtk, OpenMM, MDTraj, and NGLView.
#
# Take note of all the different data structure interconversions happening. There are *a lot*. This is good that we can get these API working together this often, but maybe not-so-good that we have to do these interconversions so often
#
# Note: OpenForceField also utilizes RDKit and OpenEyeToolkit. mBuild also utilizes OpenBabel
# +
# MoSDeF tools for initializing and parametrizing systems
import mbuild
from mbuild.examples import Ethane
import foyer
# ParmEd for interconverting data structures
import parmed
# Omnia suite of molecular modelling tools
from openforcefield.topology import Topology, Molecule
from openforcefield.typing.engines.smirnoff import ForceField
from simtk import openmm, unit
# For post-simulation analysis and visualization
import mdtraj # Also Omnia
import nglview
# -
# We will use mBuild to create a generic Ethane ($C_2H_6$) molecule.
# While this is imported from the examples, mBuild functionality allows users to construct chemical systems in a lego-like fashion by declaring particles and bonding them.
# Under the hood, rigid transformations are performed to orient particles-to-be-bonded
mbuild_compound = Ethane()
mbuild_compound.visualize(backend='nglview')
# Another operation we can do within mBuild is to take this compound, convert it to an `openbabel.Molecule` object,
# and obtain the SMILES string for it.
ethane_obmol = mbuild_compound.to_pybel()
ethane_obmol.write("smi", 'out.smi', overwrite=True)
smiles_string = open('out.smi', 'r').readlines()
print(smiles_string)
# Using foyer, we can convert an `mbuild.Compound` object to an `openmm.Topology` object.
# `openmm.Topology` objects don't actually know positions, they just know certain atomic and bonding information, but no coordinates/velocities/force field information.
# This foyer function helps recover the positions in a simple array of `simtk.Quantity`
omm_topology, xyz = foyer.forcefield.generate_topology(mbuild_compound, residues='Ethane')
print(omm_topology)
print(xyz)
# To translate these objects into `openforcefield.Topology` objects, we need to identify the unique molecules, which helps identify the isolated subgraphs - individual molecules that don't bond to anything outside its molecular network.
#
# Using the SMILES string, we can generate an `openforcefield.Molecule` object, which is this self-enclosed bonding entity (chemically speaking, this is a molecule)
ethane_molecule = Molecule.from_smiles(smiles_string[0].split()[0])
ethane_molecule
# Now that we have isolated the unique molecules, we can construct our `openforcefield.Topology` object from our `openmm.Topology` and `openmm.Molecule` objects.
off_topology = Topology.from_openmm(omm_topology, unique_molecules=[ethane_molecule])
off_topology
# ## Adding in a force field, evaluating energy
# Next, we need to create our `openforcefield.Forcefield` object. These are created from `offxml` files, and the OpenForceField group publishes new ones fairly regularly.
#
# In the comments is an example (but out-of-date) force field within the [main openforcefield package](https://github.com/openforcefield/openforcefield).
#
# The one we are using is the most-recent SMIRNOFF force field (I think this one is Parsley, or maybe just 1.0.0).
# The SMIRNOFF force fields are being housed in [a separate repo](https://github.com/openforcefield/smirnoff99Frosst), but utilize pythonic `entry_points` to help one repo into another.
#off_forcefield = ForceField('test_forcefields/smirnoff99Frosst.offxml')
off_forcefield = ForceField('smirnoff99Frosst-1.1.0.offxml')
off_forcefield
# With the `openforcefield.Topology` and `openforcefield.Forcefield` objects, we can create an `openmm.System`.
# Note the discrepancy/interplay between the objects - `openforcefield` for the molecular mechanics building blocks, but `openmm` is ultimately the workhorse for simulating and representing these systems (although you could opt to simulate with other engines via `parmed`).
#
# Note the use of AM1-BCC methods to identify partial charges.
smirnoff_omm_system = off_forcefield.create_openmm_system(off_topology)
smirnoff_omm_system
# This is a utility function we will use to evaluate the energy of a molecular system.
# Given an `openmm.system` (force field, parameters, topological, atomic information) and atomic coordinates, we can get a potential energy associated with that set of coordinatess
def get_energy(system, positions):
"""
Return the potential energy.
Parameters
----------
system : simtk.openmm.System
The system to check
positions : simtk.unit.Quantity of dimension (natoms,3) with units of length
The positions to use
Returns
---------
energy
Notes
-----
Taken from an openforcefield notebook
"""
integrator = openmm.VerletIntegrator(1.0 * unit.femtoseconds)
context = openmm.Context(system, integrator)
context.setPositions(positions)
state = context.getState(getEnergy=True)
energy = state.getPotentialEnergy().in_units_of(unit.kilocalories_per_mole)
return energy
# Next, we will try to calculate the potential energy of our ethane system under the SMIRNOFF force field.
# As a (small) obstacle to doing this, we need to change the dimensions of our simulation box because some force fields and simulations use cutoffs, and cutoffs cannot be larger than the simulation box itself.
#
# Okay 45 kcal/mol, cool. Potential energies of single configurations are usually not helpful for any real physical analysis, but can be helpful in comparing force fields.
new_vectors = [[10*unit.nanometer, 0*unit.nanometer, 0*unit.nanometer],
[0*unit.nanometer, 10*unit.nanometer, 0*unit.nanometer],
[0*unit.nanometer, 0* unit.nanometer, 10*unit.nanometer]]
smirnoff_omm_system.setDefaultPeriodicBoxVectors(*new_vectors)
get_energy(smirnoff_omm_system, xyz)
# ## Tangent: Interfacing with other simulation engines
# We can use `parmed` to convert the `openmm.Topology`, `openmm.System`, and coordinates into a `parmed.Structure`.
# From a `parmed.Structure`, we can spit out files appropriate for different simulation packages.
# Word of caution, while the developers of `parmed` did an excellent job building the conversion tools, please do your due diligence to make sure the output is as you expect
pmd_structure = parmed.openmm.load_topology(omm_topology, system=smirnoff_omm_system, xyz=xyz)
pmd_structure
# ## Comparing to the OPLS-AA force field
# Let's use a different force field.
# `foyer` ships with an XML of the OPLS-AA force field.
# We will use `foyer` (which utilizes some `parmed` and `openmm` api) to build our molecular model of ethane with OPLS
#
# * Create the `foyer.Forcefield` object
# * Apply it to our `mbuild.Compound`, get a `parmed.Structure`
# * Convert the `parmed.Structure` to an `openmm.System`
# * Reset the box vectors to be consistent with the SMIRNOFF example
# * Evaluate the energy
foyer_ff = foyer.Forcefield(name='oplsaa')
opls_pmd_structure = foyer_ff.apply(mbuild_compound)
opls_omm_system = opls_pmd_structure.createSystem()
opls_omm_system.setDefaultPeriodicBoxVectors(*new_vectors)
get_energy(opls_omm_system, opls_pmd_structure.positions)
# 37.5 kcal/mol versus 45.0 kcal/mol, for a *single* ethane. This is a little alarming because this energetic difference stems from how the interactions are quantified and parametrized.
#
# However, this isn't a deal-breaker since most physically-interesting phenomena depend on changes in (free) energies. So a singular energy isn't important - how it varies when you change configurations or sample configurational space is generally more important
# ## Cracking open the models and looking at parameters
#
# In this whole process, we've been dealing with data structures and API that are fairly transparent.
# What's great now is that we can look in-depth at these data structures.
# Specifically, we could look at the force field parameters, either within the force field files or molecular models themselves.
#
# We're going to crack open these `openmm.System` objects, looking at how some of these forces are parametrized
# Within the SMIRNOFF force field applied to ethane, we have some harmonic bonds, harmonic angles, periodic torsions, and nonbonded forces
smirnoff_omm_system.getForces()
# Within the OPLS force field applied to ethane, we have some harmonic bonds, harmonic angles, Ryckaert-Belleman torsions, and nonbonded forces.
# Don't worry about the center of mass motion remover - that's more for running a simulation.
opls_omm_system.getForces()
# We are going to compare the *nonbonded parameters* between these `openmm.System` objects.
# For every particle in our system, we're going to look at their charges, LJ sigmas, and LJ epsilsons (both of these systems utilize Coulombic electrostatics and Lennard-Jones potentials)
# Based on the charges and frequency-of-appearance, we can see which ones are carbons and which ones are hydrogens.
#
# The OPLS-system is more-charged, carbons are more negative and hydrogens are more positive.
# The SMIRNOFF-system actually isn't electro-neutral, and that might be consequence of having used AM1-BCC for such a small system.
#
# The sigmas are pretty similar between FF implementations. The hydrogen epsilsons in SMIRNOFF are about half of those in OPLS. The carbon epsilons in SMIRNOFF are almost double those in OPLS. This is kind of interesting, while SMIRNOFF-ethane has weaker electrostatics (weaker charges), the LJ might compensate with the greater carbon-epsilon.
opls_omm_nonbond_force = opls_omm_system.getForce(3)
smirnoff_omm_nonbond_force = smirnoff_omm_system.getForce(0)
for i in range(opls_omm_nonbond_force.getNumParticles()):
opls_params = opls_omm_nonbond_force.getParticleParameters(i)
smirnoff_params = smirnoff_omm_nonbond_force.getParticleParameters(i)
print(opls_params)
print(smirnoff_params)
print('---')
# ## Running some molecular dynamics simulations
# We've come this far in building our model with different force fields, we might as well build up the rest of the simulation.
# `openmm` will be used to run our simulation, since we already have an `openmm.System` object.
# We need an integrator that describes our equations of motion, timestep, and temperature behavior.
#
# As a side note, we forcibly made our simulation box really big to address cutoffs, but we can probably go with a smaller box that still fits the bill. The smaller box helps speed up the computation.
integrator = openmm.LangevinIntegrator(323 * unit.kelvin, 1.0/unit.picoseconds, 0.001 * unit.picoseconds)
smallbox_vectors = [[2*unit.nanometer, 0*unit.nanometer, 0*unit.nanometer],
[0*unit.nanometer, 2*unit.nanometer, 0*unit.nanometer],
[0*unit.nanometer, 0* unit.nanometer, 2*unit.nanometer]]
smirnoff_omm_system.setDefaultPeriodicBoxVectors(*smallbox_vectors)
# We combine our `openmm.Topology`, `openmm.System`, and `openmm.Integrator` to make our `openmm.Simulation`, then set the positions
smirnoff_simulation = openmm.app.Simulation(omm_topology, smirnoff_omm_system, integrator)
smirnoff_simulation.context.setPositions(xyz)
# Before running the simulation, we need to report some information.
# Otherwise, the simulation's going to run and we won't have anything to show for it.
# This is handled in `openmm` by creating `openmmm.reporters` and attaching them to your `openmm.Simulation`
# We will write out the timeseries of coordinates (trajectory) in a `dcd` format,
# but also a `pdb` format to show a singular configuration.
# In this case, we're printing a `pdb` file that corresponds to the first configuration, before any simulation was run.
smirnoff_simulation.reporters.append(openmm.app.DCDReporter('trajectory.dcd', 10))
pdbreporter = openmm.app.PDBReporter('first_frame.pdb', 5000)
pdbreporter.report(smirnoff_simulation, smirnoff_simulation.context.getState(-1))
# Now we can run our simulation!
smirnoff_simulation.step(1000)
# After it's finished, we can load the trajectory files into an `mdtraj.Trajectory` object, and visualize in a jupyter notebook with `nglview`. From this `mdtraj.Trajectory` object, you have pythonic-access to all the coordinates over time, and also access to various analysis libraries within `mdtraj`.
#
# The ethane is jumping around the boundaries of the periodic box, but you can see it wiggling.
# Unfortunately, markdown doesn't show NGL widgets, so I advise people to look at the notebook.
# Not super interesting, but simulations from open-source software are doable.
# If I had a more powerful computer, maybe I'd try a larger system, but I'll leave it to others to build off of my notebook (you can find this in my website's [git repo](https://github.com/ahy3nz/ahy3nz.github.io/tree/master/files/notebooks))
traj = mdtraj.load('trajectory.dcd', top='first_frame.pdb')
nglview.show_mdtraj(traj)
| _notebooks/2019-10-15-openforcefield1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulating Language Lab 1, Intro to Python (walkthrough)
# **A function which calculates and returns the product of two numbers.**
#
# I'll go through this in excruciating detail!
#
# ```python
# def product(x, y):
# ```
#
# Every function definition starts something like this. We use `def` to tell Python we are about to provide it with a definition of a function. We then give the function name: in this case I chose product, since that seems like a sensible name for a function that is going to return the product of two numbers, but I could have called it anything. Then, between brackets (...), I give the arguments that this function will take: in this case, we are going to pass two numbers to the function (which it will then multiply), so it has two arguments. Here, I decided to call them `x` and `y`, but again I could have called them anything I wanted: `n1` `n2`,`jack` and `jill`, whatever. Finally, we have a colon :. The body of the function - the stuff that actually happens when we use this function - follows this colon.
#
# ```python
# return x * y
# ```
#
# This is the body of the function - since this function is so simple, it’s just one line. Notice that, if you type this into the notebook, it automatically indents for you after the colon at the end of the previous line: the body of the function is a block of code, and blocks of code are indicated by indentation.
#
# We use return to indicate that we want this function to return some value - we are going to pass two values in to this function, we want it to pass us a single value back (namely, the product of the two numbers we passed in), and return is how we do this. Then we have an expression which tells it what to return: in this case, `x * y`, i.e. the product of the two numbers we passed in to the function as arguments. So when we actually use this function it will multiply whatever two numbers we give it, and return (pass back) the product of those numbers.
# **A function which returns the first item in a list.**
#
# ```python
# def first_in_list(alist):
# ```
#
# Again, we start with our function definition: in this case, I have decided to call the function `first_in_list`, and I have called the single argument that this function takes `alist`: we pass a list to the function, what we want the function to do is pass back the first element from that list.
#
# ```python
# return alist[0]
# ```
#
# Again, the notebook automatically indents the next line for us, because it knows that every def line is followed by a block of code (the body of the function, the bit of code that does the stuff the function is supposed to do). Once again, all we want to do is return something, so we have a return statement followed by the thing to be returned. In this case, the thing to be returned is `alist[0]`. This is the 0th element from the list called `alist`, in other words the “first” item from the list we passed in to the function.
# **A function which returns the last item in a list.**
#
# ```python
# def last_in_list(alist):
# ```
#
# Once again, we start of with our `def` line: just like `first_in_list`, `last_in_list` takes a single argument, which should be the list we want to know the last member of.
#
# ```python
# return alist[len(alist) - 1]
# ```
#
# Just like `first_in_list`, we are going to return some item from the list we are calling `alist`, so we have a return statement followed by the thing we want to return. Just like in `first_in_list`, we are returning some item from `alist`, so we have `alist[...]`. The interesting stuff with this function happens inside the square brackets. We want the last item from `alist`, and to get it we are using the `len` function.
#
# `len` is one of the built-in functions that is automatically defined when you start Python. If this was a programming course, we’d ask you to define your own `len` function, for fun, but basic functions like this are almost always provided for you, so we won’t bother with that. `len(alist)` will return the length of alist: so if `alist` has 1 element, `len(alist)` will return 1, if it has 4 elements, `len(alist)` will return 4, and so on.
#
# We subtract 1 from the number provided by `len(alist)`: that’s what `len(alist) - 1` means. This is the position in the list we want: if `alist` only has one thing in it, it has length 1 and the last element is the 0th element in that list (which is `len(alist) - 1`); if `alist` has 4 things in it, it has length 4 and the last element in that list has index 3 (which is `len(alist) - 1`). If this seems a bit confusing, remember that the “first” item in a list has index 0.
#
# So to recap: this line of code returns the item with index `len(alist) - 1` from `alist`, which should be the last item in that list.
#
# **A function which takes a list and prints out the square of each value in the list in turn.**
#
# This function uses a for loop to work through the list.
#
# ```python
# def square_list(alist):
# ```
#
# A fairly standard opening line: again, this function takes a single argument.
#
# ```python
# for x in alist:
# ```
#
# Now we have the body of the function, which is a new code block and therefore indented. This is a for loop: it will work through `alist`, taking each element in turn from that list (starting with the 0th, then the 1st, etc), temporarily calling that element `x` (although we could have used some other variable - we could have called each element `i`, or `n`, or `darling`), and does something with that `x`. The thing that it actually does with each `x` from the list is given in the next line.
#
# ```python
# print(x * x)
# ```
#
#
# Notice first that the notebook has automatically indented this line even further: the previous line setting up the for loop is followed by a new code block (the code block we are going to execute for each `x` in `alist`), and we identify the start and end of that code block using indenting. In this case, all we are doing with `x` is printing `x` out times itself: this is what `print(x * x)` achieves. Note also that we're not using `return` anywhere. That's because what the function is doing is printing stuff on the screen, rather than giving us an answer back.
# **A function which returns the largest number in a list.**
#
# This function is actually quite complex. Before I go through the code, I’ll explain the basic procedure that my code uses. Often, if you want to write a function that does something interesting, you should start by working out a step-by-step procedure for how this thing is to be achieved - once you know what you are doing, the code should be easy(ish!) to write, but if you try to write the code without really knowing what you are trying to do you’ll just end up in a mess.
#
# In this case, my general idea is that I am going to work through the list of numbers, left to right, and keep a note of the highest value I have encountered so far. I’ll call this something like “current max”. So as I go through the list I know what current max is. For each new element in the list, I compare it to current max. If it’s greater than current max, then it becomes my new current max: I forget what the old current max was, and replace it with this new value I have just encountered. On the other hand, if this new number is not greater than current max, I don’t have to do anything: I just forget about this new value, and just move on down the list, keeping current max as it was.
#
# Finally, when I have worked through this entire list, I have to remember to return my current max: this is what the function is supposed to do, and by the time I have gone through the list I will definitely have encountered and remembered the highest value in that list, which I am calling current max, so I just return that.
#
# I think that’ll work. The only other thing I have to decide is what value of current max I should start with. I guess I could choose some extremely low value that’s probably going to be lower than anything in the list, but that’s a bit risky (what if I guess wrong?), so instead the sensible thing to do is take the very first item from the list as my initial highest value, then work down the rest of the list, checking to see if I can find anything higher.
#
# OK, now I have a plan I can work through the code, explaining how it executes the procedure I have just outlined.
#
# ```python
# def max_in_list(alist):
# ```
#
# As usual, I start by naming my function and its argument.
#
# ```python
# current_max = alist[0]
# ```
#
# I introduce a new variable, which I call `current_max`. This is where I am storing the current maximum value that I have encountered. Initially I assign this variable the value `alist[0]`, i.e. the 0th element in the list I am working with.
#
# ```python
# for x in alist[1:]:
# ```
#
# Now we have a for loop, which is going to help me work through the list. Compare this line with the equivalent line in my `square_list` function: it basically looks the same, but instead of working through all of `alist`, I am going to work through `alist[1:]`. `alist[1:]` is the list of elements in alist from index 1 to the end - in other words everything in alist apart from element 0. Check back over the notes in the worksheet where this “splice” notation is explained to refresh your memory. It wouldn't actually matter if I just worked through all of alist (i.e. replaced this line of code with `for x in alist:`), but since I’ve already looked at element 0 in the list it seems a bit of a waste to look at it again.
#
# So, to recap, we’re going to take every element from the rest of the list in turn, call it `x`, and do something with it.
#
# ```python
# if x > current_max:
# ```
#
# The notebook has indented us a bit more: we are now inside the code block that is executed for every `x` in `alist[1:]`. And what we have is a conditional statement: so we compare the element `x` to `current_max`, and if `x` is greater than `current_max` we do whatever it says in the next code block.
#
# ```python
# current_max = x
# ```
#
# And what we do, if our condition is met, is overwrite `current_max` with `x`: so if the value we are currently considering (which we are calling `x`) is greater than the highest value we have encountered so far (which we are calling `current_max`), then we store `x` as our new `current_max` (and forget whatever the old value of `current_max` was). Notice we're indented again one more time because this is the code block for the result of the conditional statement.
#
# ```python
# return current_max
# ```
#
# The indenting at this point gets interesting: we have actually come out two levels, which means we are out of the body of the `if` conditional, and also out of the body of the `for` loop: so in other words, this line of code is executed once the for loop has completely finished. This line of code simply states that we return the value of `current_max` that we have ended up with after working through the list using the for loop. Remember, this was the plan: work through the list, keep a note of the maximum value encountered, and then when we have gone all the way through the list we return that value.
#
# **The geeky solution...**
#
# See if you can figure out why this other solution to the max in list problem also works:
#
# ```python
# def geeky_max_in_list(alist):
# if len(alist) > 1:
# max_of_rest = geeky_max_in_list(alist[1:])
# if alist[0] < max_of_rest:
# return max_of_rest
# return alist[0]
# ```
#
# Don't worry too much if you can't figure it out. But think through it line by line. Notice that in this function, we're actually calling the function from within itself on the third line! This is called "recursion" and is actually very similar to the kind of recursion that linguists talk about. When you're writing a function, you can sort of pretend that the function is already finished and working and then use it when you're writing it. What's happening here, is that we're thinking of the problem of finding the maximum in a list as being the same as comparing the *first* number of a list with the maximum of the *rest* of the list. If the first number is bigger than the all the other numbers in the list, then the maximum is the first number, otherwise the maximum is the biggest number in the rest of the list... Of course, if the list has only one element in it (i.e., it's length is 1) then that element must be the maximum by default.
| lab1_walkthrough.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7GYCbf4MxTS9"
# # Power method
# + id="DRjmKc4HxRX5"
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="M8Als-QFxXg8" outputId="28b7ac0f-2265-4c74-c8e2-f95830c378c9"
A = np.array([
[2.0, 1.0, 5.0],
[5.0, 7.0, 9.0],
[4.0, 6.0, 1.0],
])
print('A= ')
print(A)
# + colab={"base_uri": "https://localhost:8080/"} id="NcqD4py6T72X" outputId="1bdb8047-5889-4047-97b5-55411f630900"
w, v = np.linalg.eig(A)
print('eigenvalues = ', w)
# + [markdown] id="w-jp3s8mQBgP"
# ## Power method to find the largest (in absolute value) eigenvalue
# + [markdown] id="kejmXaobU1Gn"
# ### Algorithm 0 - test
#
# > $10$ iterations on power method
# + colab={"base_uri": "https://localhost:8080/"} id="RDUILF86xwJS" outputId="b19a28b0-1205-44ec-c527-f075a61157ae"
# initial guess
u = np.random.random((3,1))
# itmax: max. iteration number
itmx = 10
# initial iteration
k=0
while (k<itmx):
v = A.dot(u)
lamb = np.linalg.norm(v)
u = v/lamb
k = k+1
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[0]))
# + [markdown] id="7xyDiqAIRXYU"
# ### Algorithm 1
#
# $$
# \lambda^{(k+1)} = \|\hat{x}^{(k+1)}\| \to |\lambda_1|
# $$
# + colab={"base_uri": "https://localhost:8080/"} id="AMVFE2PURYQn" outputId="39b69383-73fd-4f59-9476-88ea06e2d282"
# initial guess
u = np.random.random((3,1))
u = u/np.linalg.norm(u)
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
lamb0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
while ( (k<itmx) and (rel_diff>Tol) ):
v = A.dot(u)
lamb1 = np.linalg.norm(v)
u = v/lamb1
rel_diff = abs((lamb1-lamb0)/lamb0)
k = k+1
lamb0 = lamb1
print('k= ', k, ' lamb= ', lamb1, ' error = ', abs(lamb1-w[0]))
# + [markdown] id="ci868OfVU3Qp"
# ### Algorithm 2
#
# $$
# \lambda^{(k+1)} = \ell(\hat{x}^{(k+1)}) \to \lambda_1
# $$
# + colab={"base_uri": "https://localhost:8080/"} id="xq87gFyVUz57" outputId="1fe1640a-c5ea-4cc1-c591-fbfc636f48f6"
# initial guess
u = np.random.random((3,1))
u = u/u[1,0]
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
lamb0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
while ( (k<itmx) and (rel_diff>Tol) ):
v = A.dot(u)
lamb1 = v[1,0]
u = v/lamb1
rel_diff = abs((lamb1-lamb0)/lamb0)
k = k+1
lamb0 = lamb1
print('k= ', k, ' lamb= ', lamb1, ' error = ', abs(lamb1-w[0]))
# + [markdown] id="R6WhxV8tVTIe"
# ## Inverse power method to find the smallest (in absolute value) eigenvalue
# + [markdown] id="hOrk3ulsSRoE"
# ### Algorithm 2
#
# $$
# \mu^{(k+1)} = \ell(\hat{x}^{(k+1)}) \to \frac{1}{\lambda_1}
# $$
# + id="-khCPxm9yT-F" colab={"base_uri": "https://localhost:8080/"} outputId="8b9f9f32-ae60-4086-d70b-f5e029190b3c"
# Initial guess
u = np.random.random((3,1))
u = u/u[1,0]
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
mu0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
while ( (k<itmx) and (rel_diff>Tol) ):
v = np.linalg.solve(A, u)
mu1 = v[1,0]
u = v/mu1
rel_diff = abs((mu1-mu0)/mu0)
k = k+1
mu0 = mu1
# eigenvalue = 1/mu
lamb = 1.0/mu1
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[1]))
# + [markdown] id="2kkYWS7KV46B"
# ## Shift-inverse power method to find the eigenvalue that is closest to a given one
# + [markdown] id="FXyuXgmLTWgC"
# ### Algorithm 2
#
# $$
# \mu^{(k+1)} = \ell(\hat{x}^{(k+1)}) \to \frac{1}{\lambda_1-\sigma}
# $$
# + id="CuUeMh0DFTK3" colab={"base_uri": "https://localhost:8080/"} outputId="cac2a176-8f97-4091-a1b2-98b233a2c6ad"
# Initial guess
u = np.random.random((3,1))
u = u/u[1,0]
# shift
sigma = 10.0
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
mu0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
# As: shifted matrix
# As = A - sigma*I
As = A - sigma*np.identity(3)
while ( (k<itmx) and (rel_diff>Tol) ):
v = np.linalg.solve(As, u)
mu1 = v[1,0]
u = v/mu1
rel_diff = abs((mu1-mu0)/mu0)
k = k+1
mu0 = mu1
# eigenvalue = sigma+1/mu
lamb = sigma+1.0/mu1
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[0]))
# + [markdown] id="gYBhZWrxWkKg"
# ## Inverse power method with variant shift to find one of the eigenvalue
# + [markdown] id="yd8HTW-zgTMg"
# ### Algorithm 2
#
# $$
# \sigma^{(k+1)} = \sigma^{(k)} + \frac{1}{\ell(\hat{x}^{(k+1)})} \to \lambda
# $$
# + id="G3BiOtbOWUC8" colab={"base_uri": "https://localhost:8080/"} outputId="2a3dfb3a-32ad-4a95-bc5f-06659d15b204"
# Initial guess
u = np.random.random((3,1))
u = u/u[1,0]
# shift
sigma = 10.0
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
mu0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
while ( (k<itmx) and (rel_diff>Tol) ):
sigma0 = sigma
# As: shifted matrix
# As = A - sigma*I
As = A - sigma*np.identity(3)
v = np.linalg.solve(As, u)
mu1 = v[1,0]
u = v/mu1
sigma = sigma + 1.0/mu1
rel_diff = abs((sigma0-sigma)/sigma0)
k = k+1
# eigenvalue = sigma
lamb = sigma
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[0]))
# + [markdown] id="pb0prd6KgVXq"
# ### Algorithm 3: With Rayleigh quotient
#
# $$
# \sigma^{(k+1)} =(x^{(k+1)})^TAx^{(k+1)} \to \lambda
# $$
# + id="DOYUPi7uhdZX" colab={"base_uri": "https://localhost:8080/"} outputId="f2c4d802-5e1b-40af-a0a7-c3f45c346f9c"
# Initial guess
u = np.random.random((3,1))
u = u/np.linalg.norm(u)
# shift
sigma = 10.0
# itmax: max. iteration number
itmx = 100
# initial iteration
k=0
# initial guess of largest eigenvalue
mu0 = 1.0
# tolerance
Tol = 1e-10
# initial relative difference
rel_diff = 1.0
while ( (k<itmx) and (rel_diff>Tol) ):
sigma0 = sigma
# As: shifted matrix
# As = A - sigma*I
As = A - sigma*np.identity(3)
v = np.linalg.solve(As, u)
mu1 = np.linalg.norm(v)
u = v/mu1
sigma = (u.T).dot(A.dot(u))
rel_diff = abs((sigma0-sigma)/sigma0)
k = k+1
# eigenvalue = sigma
lamb = sigma
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[0]))
# + [markdown] id="QyQ4gpNNg4Qe"
# ## For symmetric matrix
# + [markdown] id="s9dYLd69hAfb"
# ### Power iteration with Rayleigh Quotient to find the largest (in magnitude) eigenvalue
# + id="g1eCye3EgtZp"
Asym = A+A.T
# + colab={"base_uri": "https://localhost:8080/"} id="pi8JgPzagy49" outputId="fc0081fd-f066-45d9-fd9d-098c296d2b6e"
w, v = np.linalg.eig(Asym)
print('eigenvalues = ', w)
# + id="UbmMDWhJqxyb" colab={"base_uri": "https://localhost:8080/"} outputId="f1de205f-66de-4877-fd9d-739a36cf1f73"
# initial guess
u = np.random.random((3,1))
u = u/np.linalg.norm(u)
# itmax: max. iteration number
itmx = 10
# initial iteration
k=0
while (k<itmx):
v = Asym.dot(u)
lamb = (u.T).dot(v)
u = v/np.linalg.norm(v)
k = k+1
print('k= ', k, ' lamb= ', lamb, ' error = ', abs(lamb-w[0]))
# + id="wDhJpCQ3gvP2"
| power_method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Loss Functions
#
# This python script illustrates the different loss functions for regression and classification.
#
# We start by loading the ncessary libraries and resetting the computational graph.
# + deletable=true editable=true
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.python.framework import ops
ops.reset_default_graph()
# + [markdown] deletable=true editable=true
# ### Create a Graph Session
# + deletable=true editable=true
sess = tf.Session()
# + [markdown] deletable=true editable=true
# ## Numerical Predictions
#
# ---------------------------------
#
# To start with our investigation of loss functions, we begin by looking at numerical loss functions. To do so, we must create a sequence of predictions around a target. For this exercise, we consider the target to be zero.
# + deletable=true editable=true
# Various Predicted X-values
x_vals = tf.linspace(-1., 1., 500)
# Create our target of zero
target = tf.constant(0.)
# + [markdown] deletable=true editable=true
# ### L2 Loss
#
# The L2 loss is one of the most common regression loss functions. Here we show how to create it in TensorFlow and we evaluate it for plotting later.
# + deletable=true editable=true
# L2 loss
# L = (pred - actual)^2
l2_y_vals = tf.square(target - x_vals)
l2_y_out = sess.run(l2_y_vals)
# + [markdown] deletable=true editable=true
# ### L1 Loss
#
# An alternative loss function to consider is the L1 loss. This is very similar to L2 except that we take the `absolute value` of the difference instead of squaring it.
# + deletable=true editable=true
# L1 loss
# L = abs(pred - actual)
l1_y_vals = tf.abs(target - x_vals)
l1_y_out = sess.run(l1_y_vals)
# + [markdown] deletable=true editable=true
# ### Pseudo-Huber Loss
#
# The psuedo-huber loss function is a smooth approximation to the L1 loss as the (predicted - target) values get larger. When the predicted values are close to the target, the pseudo-huber loss behaves similar to the L2 loss.
# + deletable=true editable=true
# L = delta^2 * (sqrt(1 + ((pred - actual)/delta)^2) - 1)
# Pseudo-Huber with delta = 0.25
delta1 = tf.constant(0.25)
phuber1_y_vals = tf.multiply(tf.square(delta1), tf.sqrt(1. + tf.square((target - x_vals)/delta1)) - 1.)
phuber1_y_out = sess.run(phuber1_y_vals)
# Pseudo-Huber with delta = 5
delta2 = tf.constant(5.)
phuber2_y_vals = tf.multiply(tf.square(delta2), tf.sqrt(1. + tf.square((target - x_vals)/delta2)) - 1.)
phuber2_y_out = sess.run(phuber2_y_vals)
# + [markdown] deletable=true editable=true
# ### Plot the Regression Losses
#
# Here we use Matplotlib to plot the L1, L2, and Pseudo-Huber Losses.
# + deletable=true editable=true
x_array = sess.run(x_vals)
plt.plot(x_array, l2_y_out, 'b-', label='L2 Loss')
plt.plot(x_array, l1_y_out, 'r--', label='L1 Loss')
plt.plot(x_array, phuber1_y_out, 'k-.', label='P-Huber Loss (0.25)')
plt.plot(x_array, phuber2_y_out, 'g:', label='P-Huber Loss (5.0)')
plt.ylim(-0.2, 0.4)
plt.legend(loc='lower right', prop={'size': 11})
plt.show()
# + [markdown] deletable=true editable=true
# ## Categorical Predictions
#
# -------------------------------
#
# We now consider categorical loss functions. Here, the predictions will be around the target of 1.
# + deletable=true editable=true
# Various predicted X values
x_vals = tf.linspace(-3., 5., 500)
# Target of 1.0
target = tf.constant(1.)
targets = tf.fill([500,], 1.)
# + [markdown] deletable=true editable=true
# ### Hinge Loss
#
# The hinge loss is useful for categorical predictions. Here is is the `max(0, 1-(pred*actual))`.
# + deletable=true editable=true
# Hinge loss
# Use for predicting binary (-1, 1) classes
# L = max(0, 1 - (pred * actual))
hinge_y_vals = tf.maximum(0., 1. - tf.multiply(target, x_vals))
hinge_y_out = sess.run(hinge_y_vals)
# + [markdown] deletable=true editable=true
# ### Cross Entropy Loss
#
# The cross entropy loss is a very popular way to measure the loss between categorical targets and output model logits. You can read about the details more here: https://en.wikipedia.org/wiki/Cross_entropy
# + deletable=true editable=true
# Cross entropy loss
# L = -actual * (log(pred)) - (1-actual)(log(1-pred))
xentropy_y_vals = - tf.multiply(target, tf.log(x_vals)) - tf.multiply((1. - target), tf.log(1. - x_vals))
xentropy_y_out = sess.run(xentropy_y_vals)
# + [markdown] deletable=true editable=true
# ### Sigmoid Entropy Loss
#
# TensorFlow also has a sigmoid-entropy loss function. This is very similar to the above cross-entropy function except that we take the sigmoid of the predictions in the function.
# + deletable=true editable=true
# L = -actual * (log(sigmoid(pred))) - (1-actual)(log(1-sigmoid(pred)))
# or
# L = max(actual, 0) - actual * pred + log(1 + exp(-abs(actual)))
x_val_input = tf.expand_dims(x_vals, 1)
target_input = tf.expand_dims(targets, 1)
xentropy_sigmoid_y_vals = tf.nn.softmax_cross_entropy_with_logits(logits=x_val_input, labels=target_input)
xentropy_sigmoid_y_out = sess.run(xentropy_sigmoid_y_vals)
# + [markdown] deletable=true editable=true
# ### Weighted (Softmax) Cross Entropy Loss
#
# Tensorflow also has a similar function to the `sigmoid cross entropy` loss function above, but we take the softmax of the actuals and weight the predicted output instead.
# + deletable=true editable=true
# Weighted (softmax) cross entropy loss
# L = -actual * (log(pred)) * weights - (1-actual)(log(1-pred))
# or
# L = (1 - pred) * actual + (1 + (weights - 1) * pred) * log(1 + exp(-actual))
weight = tf.constant(0.5)
xentropy_weighted_y_vals = tf.nn.weighted_cross_entropy_with_logits(x_vals, targets, weight)
xentropy_weighted_y_out = sess.run(xentropy_weighted_y_vals)
# + [markdown] deletable=true editable=true
# ### Plot the Categorical Losses
# + deletable=true editable=true
# Plot the output
x_array = sess.run(x_vals)
plt.plot(x_array, hinge_y_out, 'b-', label='Hinge Loss')
plt.plot(x_array, xentropy_y_out, 'r--', label='Cross Entropy Loss')
plt.plot(x_array, xentropy_sigmoid_y_out, 'k-.', label='Cross Entropy Sigmoid Loss')
plt.plot(x_array, xentropy_weighted_y_out, 'g:', label='Weighted Cross Entropy Loss (x0.5)')
plt.ylim(-1.5, 3)
#plt.xlim(-1, 3)
plt.legend(loc='lower right', prop={'size': 11})
plt.show()
# + [markdown] deletable=true editable=true
# ### Softmax entropy and Sparse Entropy
#
# Since it is hard to graph mutliclass loss functions, we will show how to get the output instead
# + deletable=true editable=true
# Softmax entropy loss
# L = -actual * (log(softmax(pred))) - (1-actual)(log(1-softmax(pred)))
unscaled_logits = tf.constant([[1., -3., 10.]])
target_dist = tf.constant([[0.1, 0.02, 0.88]])
softmax_xentropy = tf.nn.softmax_cross_entropy_with_logits(logits=unscaled_logits,
labels=target_dist)
print(sess.run(softmax_xentropy))
# Sparse entropy loss
# Use when classes and targets have to be mutually exclusive
# L = sum( -actual * log(pred) )
unscaled_logits = tf.constant([[1., -3., 10.]])
sparse_target_dist = tf.constant([2])
sparse_xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(logits=unscaled_logits,
labels=sparse_target_dist)
print(sess.run(sparse_xentropy))
| 02_TensorFlow_Way/04_Implementing_Loss_Functions/04_loss_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import nltk
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import RegexpTokenizer
import ast
import numpy as np
import os
import ast
import urllib.request
from urllib.request import urlopen
from bs4 import BeautifulSoup
import os.path
from datetime import datetime
from collections import Counter
nltk.download('stopwords')
nltk.download('words')
nltk.download('wordnet')
# -
month = "january"
char_blacklist = list(chr(i) for i in range(32, 127) if i <= 64 or i >= 91 and i <= 96 or i >= 123)
stopwords = nltk.corpus.stopwords.words('english')
stopwords.extend(char_blacklist)
english_vocab = set(w.lower() for w in nltk.corpus.words.words())
english_tolerance = 50
english_confidence = []
words_threshold = 10
top = 2500
toker = RegexpTokenizer(r'((?<=[^\w\s])\w(?=[^\w\s])|(\W))+', gaps=True)
words_frequency = {}
# +
# Read new generated data set file
df = pd.read_csv("../Datasets/full_data_{}.csv".format(month))
# Generate most frequent words list for each category
words_frequency = {}
for category in set(df['main_category'].values):
print(category)
all_words = []
for row in df[df['main_category'] == category]['tokenized_words'].tolist():
for word in ast.literal_eval(row):
all_words.append(word)
most_common = nltk.FreqDist(w for w in all_words).most_common(top)
words_frequency[category] = most_common
# Extract only words
for category in set(df['main_category'].values):
words_frequency[category] = [word for word, number in words_frequency[category]]
# Save words_frequency model
import pickle
words_filename = "../Models/{}/word_frequency_{}_test.picle".format(month.title(), month)
if not os.path.isfile(words_filename):
pickle_out = open(words_filename,"wb")
pickle.dump(words_frequency, pickle_out)
pickle_out.close()
# Create labels and features set for ML
features = np.zeros(df.shape[0] * top).reshape(df.shape[0], top)
labels = np.zeros(df.shape[0])
counter = 0
for i, row in df.iterrows():
c = [word for word, word_count in Counter(row['tokenized_words']).most_common(top)]
labels[counter] = list(set(df['main_category'].values)).index(row['main_category'])
for word in c:
if word in words_frequency[row['main_category']]:
features[counter][words_frequency[row['main_category']].index(word)] = 1
counter += 1
# Features and labels splitting to training and testing data
from sklearn.metrics import accuracy_score
from scipy.sparse import coo_matrix
X_sparse = coo_matrix(features)
from sklearn.utils import shuffle
X, X_sparse, y = shuffle(features, X_sparse, labels, random_state=0)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# Train and validate data using ML algorithms
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(X_train, y_train)
predictions = lr.predict(X_test)
lr_score = lr.score(X_test, y_test)
print('LogisticRegression')
print('Score: ', lr_score)
print('Top: ', top)
print('Tolerance: ', english_tolerance)
print('Dataset length: ', df.shape[0])
print()
from sklearn.svm import LinearSVC
lsvm = LinearSVC()
lsvm.fit(X_train, y_train)
predictions = lsvm.predict(X_test)
lsvm_score = lsvm.score(X_test, y_test)
print('LSVM')
print('Score: ', lsvm_score)
print('Top: ', top)
print('Tolerance: ', english_tolerance)
print('Dataset length: ', df.shape[0])
# -
import pandas as pd
import nltk
import ast
import numpy as np
import os
import ast
import urllib.request
from urllib.request import urlopen
from bs4 import BeautifulSoup
import os.path
nltk.download('stopwords')
nltk.download('words')
nltk.download('punkt')
# # Dataset creation if it is not existing.
# __Dataset is filtered by these set of rules:__
# 1. Main category != Not_working (Exclude non working URL's)
# 2. Main category:confidence > 0.5 (Leave url's with likely know categories)
# 3. Non responding URL's are excluded
# 4. Non english language URL's are excluded.
#
# ### Caution, the full data set creation may take ~15 hours.
# +
def no_filter_data():
file = 'Datasets/URL-categorization-DFE.csv'
df = pd.read_csv(file)[['main_category', 'main_category:confidence', 'url']]
df = df[(df['main_category'] != 'Not_working') & (df['main_category:confidence'] > 0.5)]
df['tokenized_words'] = ''
counter = 0
for i, row in df.iterrows():
counter += 1
print("{}, {}/{}".format(row['url'], counter, len(df)))
try:
hdr = {'User-Agent': 'Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.11 (KHTML, like Gecko) Chrome/23.0.1271.64 Safari/537.11',
'Accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8',
'Accept-Charset': 'ISO-8859-1,utf-8;q=0.7,*;q=0.3',
'Accept-Encoding': 'none',
'Accept-Language': 'en-US,en;q=0.8',
'Connection': 'keep-alive'}
req = urllib.request.Request(url, headers=hdr)
html = urlopen(req).read()
# html = urlopen('http://' + row['url'], timeout=15).read()
except:
continue
soup = BeautifulSoup(html, "html.parser")
[tag.decompose() for tag in soup("script")]
[tag.decompose() for tag in soup("style")]
text = soup.get_text()
lines = (line.strip() for line in text.splitlines())
chunks = (phrase.strip() for line in lines for phrase in line.split(" "))
text = '\n'.join(chunk.lower() for chunk in chunks if chunk)
tokens = nltk.word_tokenize(text)
df.at[i, 'tokenized_words'] = tokens if len(tokens) > 0 else ''
df = df[~df['tokenized_words'].isnull()]
df.to_csv("Datasets/full_data_v3.csv")
if not os.path.isfile("Datasets/full_data_v3.csv"):
no_filter_data()
# -
# ### Reading data set and creating list of stopwords and english vocabulary for further investigation
df = pd.read_csv("Datasets/full_data_december.csv")
df = df[~df['tokenized_words'].isnull()]
char_blacklist = list(chr(i) for i in range(32, 127) if i <= 64 or i >= 91 and i <= 96 or i >= 123)
stopwords = nltk.corpus.stopwords.words('english')
stopwords.extend(char_blacklist)
english_vocab = set(w.lower() for w in nltk.corpus.words.words())
# # Filter webpages with english language
# If the webpage contains at least 20 % english words of total words, then the webpage is considered as english
english_confidence = []
english_tolerance = 50
for i, row in df.iterrows():
english_words = 0
words = ast.literal_eval(row['tokenized_words'])
for word in words:
if word.lower() in english_vocab:
english_words += 1
english_confidence.append(english_words / len(words) * 100)
df['english:confidence'] = english_confidence
df = df[df['english:confidence'] > english_tolerance]
# # Make the most popular word list for each catgegory
# +
top = 2500
words_frequency = {}
for category in set(df['main_category'].values):
all_words = []
for row in df[df['main_category'] == category]['tokenized_words'].tolist():
for word in ast.literal_eval(row):
all_words.append(word)
allWordExceptStopDist = nltk.FreqDist(
w.lower() for w in all_words if w not in stopwords and len(w) >= 3 and w[0] not in char_blacklist)
most_common = allWordExceptStopDist.most_common(top)
words_frequency[category] = most_common
for category in set(df['main_category'].values):
words_frequency[category] = [word for word, number in words_frequency[category]]
# -
# ### Remove most frequent words in all categories
from collections import Counter
words = []
for category in words_frequency.keys():
words.extend(words_frequency[category][0:15])
words_counter = Counter(words)
words_filter = {x : words_counter[x] for x in words_counter if words_counter[x] >= 7}
words_stop = list(words_filter.keys())
for category in words_frequency.keys():
words_frequency[category] = [word for word in words_frequency[category] if word not in words_stop]
words_filter
# # Create features and labels for Machine learning training
# +
from collections import Counter
features = np.zeros(df.shape[0] * top).reshape(df.shape[0], top)
labels = np.zeros(df.shape[0])
counter = 0
for i, row in df.iterrows():
c = [word for word, word_count in Counter(ast.literal_eval(row['tokenized_words'])).most_common(top)]
labels[counter] = list(set(df['main_category'].values)).index(row['main_category'])
for word in c:
if word in words_frequency[row['main_category']]:
features[counter][words_frequency[row['main_category']].index(word)] = 1
counter += 1
# -
# # Create seperate training/testing datasets and shuffle them
# +
from sklearn.metrics import accuracy_score
from scipy.sparse import coo_matrix
X_sparse = coo_matrix(features)
from sklearn.utils import shuffle
X, X_sparse, y = shuffle(features, X_sparse, labels, random_state=0)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# -
# # Predictions
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(X_train, y_train)
predictions = lr.predict(X_test)
score = lr.score(X_test, y_test)
print('LogisticRegression')
print('Score: ', score)
print('Top: ', top)
print('Tolerance: ', english_tolerance)
print('Dataset length: ', df.shape[0])
print()
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(X_train, y_train)
predictions = dtc.predict(X_test)
score = dtc.score(X_test, y_test)
print('DecisionTreeClassifier')
print('Score: ', score)
print('Top: ', top)
print('Tolerance: ', english_tolerance)
print('Dataset length: ', df.shape[0])
print()
from sklearn.svm import LinearSVC
clf = LinearSVC()
clf.fit(X_train, y_train)
predictions = clf.predict(X_test)
score = clf.score(X_test, y_test)
print('SVM')
print('Score: ', score)
print('Top: ', top)
print('Tolerance: ', english_tolerance)
print('Dataset length: ', df.shape[0])
# ### Save ML model
# +
month = 'December'
from sklearn.externals import joblib
filename = "Models/{}/LR_model_v3_stop_{}.joblib".format(month, month)
if not os.path.isfile(filename):
joblib.dump(lr, filename)
import pickle
words_filename = "Models/{}/word_frequency_v3_stop_{}.picle".format(month, month)
if not os.path.isfile(words_filename):
pickle_out = open(words_filename,"wb")
pickle.dump(words_frequency, pickle_out)
pickle_out.close()
filename = "Models/{}/LR_maxtrain_v3.joblib_stop_{}".format(month, month)
if not os.path.isfile(filename):
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(X, y)
joblib.dump(lr, filename)
# +
# import matplotlib.pyplot as plt; plt.rcdefaults()
# import numpy as np
# import matplotlib.pyplot as plt
# objects = ('English', 'Italic', 'Russian', 'Japan', 'China', 'Belgium')
# y_pos = np.arange(len(objects))
# performance = [8143,260,646,338,125,100]
# plt.bar(y_pos, performance, align='center', alpha=0.5)
# plt.xticks(y_pos, objects)
# plt.ylabel('URLs')
# plt.title('Languages diversity in the data set')
# plt.show()
# plt.savefig("language_diversity.png")
# df[df['main_category'] == 'Business_and_Industry']['url']
# +
# import matplotlib.pyplot as plt; plt.rcdefaults()
# import numpy as np
# import matplotlib.pyplot as plt
# from collections import Counter
# words = []
# for category in words_frequency.keys():
# words.extend(words_frequency[category][0:15])
# words_counter = Counter(words)
# words_filter = {x : words_counter[x] for x in words_counter if words_counter[x] >= 7}
# objects = tuple(words_filter.keys())
# y_pos = np.arange(len(objects))
# performance = list(words_filter.values())
# plt.barh(y_pos, performance, align='center', alpha=1)
# plt.xticks(range(1, max(performance) + 1))
# plt.yticks(y_pos, objects)
# plt.xlabel('Word diversity in categories (TOP 15 words)')
# plt.title('Words diversity in each category TOP 15 most frequent words')
# plt.show()
# plt.savefig("words_diversity.png")
| Jupyter-notebook/.ipynb_checkpoints/FullDataset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import sys
sys.executable
# +
sys.path.append(r'B:GitRepos\Python Projects\pySB')
import numpy as np
from .Utils.COGridGen import COGridGen
from .Utils.mapper import mapper
from shapely.geometry import shape
import matplotlib.pyplot as plt
# +
print("The excutable is {}".format(sys.executable))
grid = COGridGen(r"..\data\GreenRiver\centerline.shp", 0.5, nx=25, ny=11, width=350)
print(grid.description())
# +
x, y = grid.cl.getpoints()
# print(x)
xo, yo = grid.cl.getinterppts(grid.nx, 100)
# print(xo)
xgrid, ygrid = grid.getXYGrid()
# print(xgrid)
plt.scatter(xgrid,ygrid)
plt.plot(x,y)
plt.plot(xo, yo)
plt.show()
# -
type(x.tolist())
x.tolist()
# +
import numpy as np
import holoviews as hv
hv.extension('bokeh', 'matplotlib')
# +
# %%opts Curve [aspect='equal']
# %%opts Scatter [aspect='equal']
# Note aspect does not seem to work at this point, search suggests that it will
# eventually
# # %%opts Curve [height=400, width=400, aspect='equal']
Easting = hv.Dimension('easting', label='Easting', unit='m')
Northing = hv.Dimension('northing', label='Northing', unit='m')
cl_curve = hv.Curve((x.tolist(), y.tolist()), Easting, Northing, label='centerline')
sp_curve = hv.Curve((xo.tolist(), yo.tolist()), label = 'spline')
gridpts = hv.Scatter((xgrid.tolist(), ygrid.tolist()), label = 'grid')
cl_curve * sp_curve * gridpts
# +
# from holoviews.operation import Operation
# class build_grid(Operation):
# def _process(self, grid, key=None):
# xo, yo = grid.cl.getinterppts(grid.nx)
# sp_curve = hv.Curve((xo.tolist(), yo.tolist()), label = 'spline')
# return sp_curve
# +
# cl_curve * build_grid(grid)
# -
def clcurve(tension):
x, y = grid.cl.getpoints()
xo, yo = grid.cl.getinterppts(grid.nx, tension)
print(tension)
c1 = hv.Curve((x.tolist(), y.tolist()), Easting, Northing, label='centerline')
c2 = hv.Curve((xo.tolist(), yo.tolist()), label = 'spline')
return c1*c2
dmap = hv.DynamicMap(clcurve, kdims=['tension'])
dmap.redim.range(tension=(0.1, 100))
map = mapper(grid, 'Elevation', 60, 5, 1, r"..\data\GreenRiver\channeltopoz.shp", shp_has_geo=True)
xd, yd, zd = map.getData()
plt.scatter(xd, yd, s = 0.25, c=zd)
plt.scatter(xgrid, ygrid, s=1, c='black')
plt.show()
map.MapwCLTemplate()
tmp = 1000
print(tmp)
map.plotmapgrid()
grid.plotmapgrid('Elevation')
| notebooks/test-hv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This script consolidates *.csv formatted output from Mindboggle software (https://mindboggle.info).
# As written, it pulls data on area, cortical thickness (mean), travel depth (mean), and mean curvature (mean) for 62 cortical brain regions.
# The script consolidates volumetric estimates from *.csv outputs. However, as this list is not comprehensive, we advice using the freesurfer_wmparc_labels_in_hybrid_graywhite.nii.gz output instead (a script to extract volumetric estimates from these files, may be found at https://github.com/TeddyTuresky/BrainMorphometry_DiminishedGrowth_BEANstudy_2021)
side = 'left' # Need to also change to pull right side values
tag = 'label' # Set to consolidate estimates from label_shapes.csv, but can also pull from sulcal_shapes.csv by renaming to 'sulcal'
prog = 'freesurfer'# Set to examine volumes according to FreeSurfer labels, but can use ANTs labels with 'ants'
import os
import csv
import pandas as pd
import glob
g = sorted(glob.glob('mindboggle-tables/*_{}_{}_shapes.csv'.format(side,tag)))
v = sorted(glob.glob('mindboggle-tables/*_volume_per_{}_label.csv'.format(prog)))
i = 0
for file in g:
i += 1
file1 = file.split('/')[1]
sub = file1.split('_')[0]
print(sub)
df = pd.read_csv(file)
df.columns = df.columns.str.strip().str.lower().str.replace(' ', '_').str.replace(':', '')
area1 = df.area # This line and the three below are where you specify which out of 100 surface-based measures you wish to consolidate
depth1 = df.travel_depth_mean
curve1 = df.mean_curvature_mean
thick1 = df.freesurfer_thickness_mean
if i == 1:
areas = area1
depths = depth1
curves = curve1
thicks = thick1
else:
areas = pd.concat([areas, area1], axis=1)
depths = pd.concat([depths, depth1], axis=1)
curves = pd.concat([curves, curve1], axis=1)
thicks = pd.concat([thicks, thick1], axis=1)
i = 0
for vfile in v:
i += 1
vfile1 = vfile.split('/')[1]
vsub = vfile1.split('_')[0]
dfv = pd.read_csv(vfile)
dfv.columns = dfv.columns.str.strip().str.lower().str.replace(' ', '_').str.replace(':', '')
vol1 = dfv.volume
if i == 1:
vols = vol1
else:
vols = pd.concat([vols, vol1], axis=1)
#print(areas)
#print(depths)
#print(curves)
#print(thicks)
areas.to_csv('{}_{}_shapes_areas.csv'.format(side,tag))
depths.to_csv('{}_{}_shapes_depths.csv'.format(side,tag))
curves.to_csv('{}_{}_shapes_curves.csv'.format(side,tag))
thicks.to_csv('{}_{}_shapes_thicks.csv'.format(side,tag))
vols.to_csv('{}_vols.csv'.format(prog))
# -
| consolidateShapes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data passing tutorial
# Data passing is the most important aspect of Pipelines.
#
# In Kubeflow Pipelines, the pipeline authors compose pipelines by creating component instances (tasks) and connecting them together.
#
# Component have inputs and outputs. They can consume and produce arbitrary data.
#
# Pipeline authors establish connections between component tasks by connecting their data inputs and outputs - by passing the output of one task as an argument to another task's input.
#
# The system takes care of storing the data produced by components and later passing that data to other components for consumption as instructed by the pipeline.
#
# This tutorial shows how to create python components that produce, consume and transform data.
# It shows how to create data passing pipelines by instantiating components and connecting them together.
# Put your KFP cluster endpoint URL here if working from GCP notebooks (or local notebooks). ('https://xxxxx.notebooks.googleusercontent.com/')
kfp_endpoint='https://XXXXX.{pipelines|notebooks}.googleusercontent.com/'
# Install Kubeflow Pipelines SDK. Add the --user argument if you get permission errors.
# !PIP_DISABLE_PIP_VERSION_CHECK=1 python3 -m pip install 'kfp>=1.4.0' --quiet --user
# +
from typing import NamedTuple
import kfp
from kfp.components import InputPath, InputTextFile, OutputPath, OutputTextFile
from kfp.components import func_to_container_op
# -
# ## Small data
#
# Small data is the data that you'll be comfortable passing as program's command-line argument. Small data size should not exceed few kilobytes.
#
# Some examples of typical types of small data are: number, URL, small string (e.g. column name).
#
# Small lists, dictionaries and JSON structures are fine, but keep an eye on the size and consider switching to file-based data passing methods taht are more suitable for bigger data (more than several kilobytes) or binary data.
#
# All small data outputs will be at some point serialized to strings and all small data input values will be at some point deserialized from strings (passed as command-line argumants). There are built-in serializers and deserializers for several common types (e.g. `str`, `int`, `float`, `bool`, `list`, `dict`). All other types of data need to be serialized manually before returning the data. Make sure to properly specify type annotations, otherwize there would be no automatic deserialization and the component function will receive strings instead of deserialized objects.
# ### Consuming small data
# +
@func_to_container_op
def print_small_text(text: str):
'''Print small text'''
print(text)
def constant_to_consumer_pipeline():
'''Pipeline that passes small constant string to to consumer'''
consume_task = print_small_text('Hello world') # Passing constant as argument to consumer
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(constant_to_consumer_pipeline, arguments={})
# +
def pipeline_parameter_to_consumer_pipeline(text: str):
'''Pipeline that passes small pipeline parameter string to to consumer'''
consume_task = print_small_text(text) # Passing pipeline parameter as argument to consumer
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(
pipeline_parameter_to_consumer_pipeline,
arguments={'text': 'Hello world'}
)
# -
# ### Producing small data
# +
@func_to_container_op
def produce_one_small_output() -> str:
return 'Hello world'
def task_output_to_consumer_pipeline():
'''Pipeline that passes small data from producer to consumer'''
produce_task = produce_one_small_output()
# Passing producer task output as argument to consumer
consume_task1 = print_small_text(produce_task.output) # task.output only works for single-output components
consume_task2 = print_small_text(produce_task.outputs['output']) # task.outputs[...] always works
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(task_output_to_consumer_pipeline, arguments={})
# -
# ### Producing and consuming multiple arguments
# +
@func_to_container_op
def produce_two_small_outputs() -> NamedTuple('Outputs', [('text', str), ('number', int)]):
return ("data 1", 42)
@func_to_container_op
def consume_two_arguments(text: str, number: int):
print('Text={}'.format(text))
print('Number={}'.format(str(number)))
def producers_to_consumers_pipeline(text: str = "Hello world"):
'''Pipeline that passes data from producer to consumer'''
produce1_task = produce_one_small_output()
produce2_task = produce_two_small_outputs()
consume_task1 = consume_two_arguments(produce1_task.output, 42)
consume_task2 = consume_two_arguments(text, produce2_task.outputs['number'])
consume_task3 = consume_two_arguments(produce2_task.outputs['text'], produce2_task.outputs['number'])
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(producers_to_consumers_pipeline, arguments={})
# -
# ### Consuming and producing data at the same time
# +
@func_to_container_op
def get_item_from_list(list_of_strings: list, index: int) -> str:
return list_of_strings[index]
@func_to_container_op
def truncate_text(text: str, max_length: int) -> str:
return text[0:max_length]
def processing_pipeline(text: str = "Hello world"):
truncate_task = truncate_text(text, max_length=5)
get_item_task = get_item_from_list(list_of_strings=[3, 1, truncate_task.output, 1, 5, 9, 2, 6, 7], index=2)
print_small_text(get_item_task.output)
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(processing_pipeline, arguments={})
# -
# ## Bigger data (files)
#
# Bigger data should be read from files and written to files.
#
# The paths for the input and output files are chosen by the system and are passed into the function (as strings).
#
# Use the `InputPath` parameter annotation to tell the system that the function wants to consume the corresponding input data as a file. The system will download the data, write it to a local file and then pass the **path** of that file to the function.
#
# Use the `OutputPath` parameter annotation to tell the system that the function wants to produce the corresponding output data as a file. The system will prepare and pass the **path** of a file where the function should write the output data. After the function exits, the system will upload the data to the storage system so that it can be passed to downstream components.
#
# You can specify the type of the consumed/produced data by specifying the type argument to `InputPath` and `OutputPath`. The type can be a python type or an arbitrary type name string. `OutputPath('TFModel')` means that the function states that the data it has written to a file has type 'TFModel'. `InputPath('TFModel')` means that the function states that it expect the data it reads from a file to have type 'TFModel'. When the pipeline author connects inputs to outputs the system checks whether the types match.
#
# Note on input/output names: When the function is converted to component, the input and output names generally follow the parameter names, but the "\_path" and "\_file" suffixes are stripped from file/path inputs and outputs. E.g. the `number_file_path: InputPath(int)` parameter becomes the `number: int` input. This makes the argument passing look more natural: `number=42` instead of `number_file_path=42`.
#
# ### Writing and reading bigger data
# +
# Writing bigger data
@func_to_container_op
def repeat_line(line: str, output_text_path: OutputPath(str), count: int = 10):
'''Repeat the line specified number of times'''
with open(output_text_path, 'w') as writer:
for i in range(count):
writer.write(line + '\n')
# Reading bigger data
@func_to_container_op
def print_text(text_path: InputPath()): # The "text" input is untyped so that any data can be printed
'''Print text'''
with open(text_path, 'r') as reader:
for line in reader:
print(line, end = '')
def print_repeating_lines_pipeline():
repeat_lines_task = repeat_line(line='Hello', count=5000)
print_text(repeat_lines_task.output) # Don't forget .output !
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(print_repeating_lines_pipeline, arguments={})
# -
# ### Processing bigger data
# +
@func_to_container_op
def split_text_lines(source_path: InputPath(str), odd_lines_path: OutputPath(str), even_lines_path: OutputPath(str)):
with open(source_path, 'r') as reader:
with open(odd_lines_path, 'w') as odd_writer:
with open(even_lines_path, 'w') as even_writer:
while True:
line = reader.readline()
if line == "":
break
odd_writer.write(line)
line = reader.readline()
if line == "":
break
even_writer.write(line)
def text_splitting_pipeline():
text = '\n'.join(['one', 'two', 'three', 'four', 'five', 'six', 'seven', 'eight', 'nine', 'ten'])
split_text_task = split_text_lines(text)
print_text(split_text_task.outputs['odd_lines'])
print_text(split_text_task.outputs['even_lines'])
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(text_splitting_pipeline, arguments={})
# -
# ### Processing bigger data with pre-opened files
# +
@func_to_container_op
def split_text_lines2(source_file: InputTextFile(str), odd_lines_file: OutputTextFile(str), even_lines_file: OutputTextFile(str)):
while True:
line = source_file.readline()
if line == "":
break
odd_lines_file.write(line)
line = source_file.readline()
if line == "":
break
even_lines_file.write(line)
def text_splitting_pipeline2():
text = '\n'.join(['one', 'two', 'three', 'four', 'five', 'six', 'seven', 'eight', 'nine', 'ten'])
split_text_task = split_text_lines2(text)
print_text(split_text_task.outputs['odd_lines']).set_display_name('Odd lines')
print_text(split_text_task.outputs['even_lines']).set_display_name('Even lines')
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(text_splitting_pipeline2, arguments={})
# -
# ### Example: Pipeline that generates then sums many numbers
# +
# Writing many numbers
@func_to_container_op
def write_numbers(numbers_path: OutputPath(str), start: int = 0, count: int = 10):
with open(numbers_path, 'w') as writer:
for i in range(start, count):
writer.write(str(i) + '\n')
# Reading and summing many numbers
@func_to_container_op
def sum_numbers(numbers_path: InputPath(str)) -> int:
sum = 0
with open(numbers_path, 'r') as reader:
for line in reader:
sum = sum + int(line)
return sum
# Pipeline to sum 100000 numbers
def sum_pipeline(count: 'Integer' = 100000):
numbers_task = write_numbers(count=count)
print_text(numbers_task.output)
sum_task = sum_numbers(numbers_task.outputs['numbers'])
print_text(sum_task.output)
# Running the pipeline
kfp.Client(host=kfp_endpoint).create_run_from_pipeline_func(sum_pipeline, arguments={})
| courses/machine_learning/deepdive2/production_ml/labs/samples/tutorials/Data passing in python components.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Heroes Of Pymoli Data Analysis
# * Of the 1163 active players, the vast majority are male (84%). There also exists, a smaller, but notable proportion of female players (14%).
#
# * Our peak age demographic falls between 20-24 (44.8%) with secondary groups falling between 15-19 (18.60%) and 25-29 (13.4%).
# -----
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
import numpy as np
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data_df = pd.read_csv(file_to_load, encoding="utf-8")
# -
# ## Player Count
# * Display the total number of players
#
#Display the total number of players
player_count = len(purchase_data_df["SN"].unique())
#Place all the data into player count dataframe
player_count_table = pd.DataFrame({"Total Players":[player_count]})
player_count_table
# ## Purchasing Analysis (Total)
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
# +
#Obtain number of unique items
unique_items = len(purchase_data_df["Item ID"].unique())
number_of_purchases = purchase_data_df["Purchase ID"].count()
#Obtain average price
average_price = purchase_data_df["Price"].sum()/number_of_purchases
#Obtain total revenue
total_revenue = purchase_data_df["Price"].sum()
#Create DataFrame using Dictionary of Arrays
purchasing_analysis = pd.DataFrame({
"Number of Unique Items":unique_items,
"Average Price": average_price,
"Number of Purchases":number_of_purchases,
"Total Revenue":[total_revenue]
})
#Formating the Price
purchasing_analysis["Average Price"] = purchasing_analysis["Average Price"].map("${:.2f}".format)
purchasing_analysis["Total Revenue"] = purchasing_analysis["Total Revenue"].map("${:.2f}".format)
purchasing_analysis
# -
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
#Creating dataframe for SN , Gender column and remove duplicate
gender_name_df = pd.DataFrame(purchase_data_df, columns=["SN","Gender"]).drop_duplicates()
#Count the player with gender after removing duplicates
gender_count = gender_name_df["Gender"].value_counts()
gender_percentage = round((gender_name_df["Gender"].value_counts()/len(gender_name_df["Gender"]))*100,2)
data_gender = {"Total Count":gender_count,"Percentage of Players":gender_percentage}
#Create new dataframe for Gender Demographics results
gender_demographics = pd.DataFrame(data_gender)
gender_demographics
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
#Grouping of purchase count by gender
purchase_count = purchase_data_df.groupby(["Gender"])["Purchase ID"].count()
#Calculate average purchase count
avg_purchase_price = round(purchase_data_df.groupby(["Gender"])["Price"].mean(),2)
total_purchase_value = purchase_data_df.groupby(["Gender"])["Price"].sum()
player_count_by_gender = purchase_data_df.groupby(["Gender"])["SN"].count()
avg_purchase_total_per_person = round(total_purchase_value/gender_count,2)
#Get new dataframe for Purchasing Analysis
purchasing_analysis = pd.DataFrame({
"Purchase Count": purchase_count,
"Average Purchase Price": avg_purchase_price,
"Total Purchase Value": total_purchase_value.map("${:.2f}".format),
"Avg Total Purchase per Person": avg_purchase_total_per_person.map("${:.2f}".format)
})
#Formating of price results
purchasing_analysis["Average Purchase Price"] = purchasing_analysis["Average Purchase Price"].map("${:.2f}".format)
purchasing_analysis
# -
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
# +
#This is to define the bins
bins = [1, 9, 14, 19, 24, 29, 34, 39, 90]
#Define the groups of the bins
group_names = ["<10","10-14","15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
#Categorize the existing players using the age bins.
purchase_data_df["age_group"] = pd.cut(purchase_data_df["Age"], bins, labels=group_names)
#Create new dataframe and remove the duplicate rows
age_group_df = pd.DataFrame(purchase_data_df, columns = ["SN","age_group"]).drop_duplicates()
#Calculate the numbers by age group
numbers_by_age_group = age_group_df["age_group"].value_counts()
#Calculate the percentages by age group
percentages_by_age_group = round(((numbers_by_age_group/player_count)*100),2)
#Create new datframe for the Age Demographics results
age_demographics = pd.DataFrame({
"Total Count": numbers_by_age_group,
"Percentage of Players":percentages_by_age_group
})
age_demographics = age_demographics.sort_index(0, ascending= True)
age_demographics
# -
# ## Purchasing Analysis (Age)
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
#This is to define the bins
bins = [1, 9, 14, 19, 24, 29, 34, 39, 90]
#Define the groups of the bins
group_names = ["<10","10-14","15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
#Categorize the existing players using the age bins.
purchase_data_df["age_group"] = pd.cut(purchase_data_df["Age"], bins, labels=group_names)
#Create new dataframe
purchase_data_age_group_df = pd.DataFrame(purchase_data_df, columns = ["SN","age_group", "Price"])#.drop_duplicates()
#Get Series for Purchase count by age
purchase_count = purchase_data_age_group_df.groupby(["age_group"])["Price"].count()
#purchase_count = purchase_data_age_group_df["age_group"].value_counts()
# Get Series for Average Price in age group
purchase_price_by_age_group = purchase_data_age_group_df.groupby(["age_group"])["Price"].sum()
avg_purchase_price = round((purchase_price_by_age_group/purchase_count),2)
player_count_by_age_group = purchase_data_age_group_df.groupby(["age_group"])["SN"].count()
#Format the price data and add $ sign
avg_total_purchase_per_person = purchase_price_by_age_group/numbers_by_age_group
#Create Datafranme for Ourchasung Analysis
purchasing_analysis = pd.DataFrame({
"Purchase Count": purchase_count,
"Average Purchase Price": avg_purchase_price.map("${:.2f}".format),
"Total Purchase Value": purchase_price_by_age_group.map("${:.2f}".format),
"Avg Total Purchase per Person": avg_total_purchase_per_person.map("${:.2f}".format)
})
purchasing_analysis
# -
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
# +
purchase_count = purchase_data_df.groupby(["SN"])["Item ID"].count()
total_purchase_value = purchase_data_df.groupby(["SN"])["Price"].sum()
avg_purchase_price = round((total_purchase_value/purchase_count),2)
#Create a summary data frame to hold the results
top_spenders = pd.DataFrame({
"Purchase Count": purchase_count,
"Average Purchase Price": avg_purchase_price,
"Total Purchase Value": total_purchase_value
})
# Sort the total purchase value column in descending order
top_spenders = top_spenders.sort_values("Total Purchase Value", ascending= False)
top_spenders["Average Purchase Price"] = top_spenders["Average Purchase Price"].map("${:.2f}".format)
top_spenders["Total Purchase Value"] = top_spenders["Total Purchase Value"].map("${:.2f}".format)
# Display a preview of the summary data frame
top_spenders.head()
# -
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
# Retrieve the Item ID, Item Name, and Item Price columns
popular_items_df = pd.DataFrame(purchase_data_df, columns =["Item ID","Item Name","Price"])
#Get Series for Purchase count by Item ID and Item Name
purchase_count = popular_items_df.groupby(["Item ID", "Item Name"])["Price"].count() ##get series for items price
total_purchase_value = round(purchase_data_df.groupby(["Item ID", "Item Name"])["Price"].sum(),2)
item_price = round(purchase_data_df.groupby(["Item ID","Item Name"])["Price"].mean(),2)
most_popular_items = pd.DataFrame({
"Purchase Count": purchase_count,
"Item Price": item_price.map("${:.2f}".format),
"Total Purchase Value":total_purchase_value
})
# Sort the purchase count column in descending order
most_popular_items_sortby_pc = most_popular_items.sort_values("Purchase Count", ascending= False)
most_popular_items_sortby_pc ["Total Purchase Value"] = most_popular_items_sortby_pc ["Total Purchase Value"].map("${:.2f}".format)
# Display a preview of the summary data frame
most_popular_items_sortby_pc .head()
# -
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
# +
#Sort the table by total purchase value in descending order
most_profitable_items = most_popular_items.sort_values("Total Purchase Value", ascending= False)
most_profitable_items["Total Purchase Value"] = most_profitable_items["Total Purchase Value"].map("${:.2f}".format)
# Display a preview of the data frame
most_profitable_items.head()
# -
| HeroesOfPymoli/HeroesOfPymoli_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DataKind Red Cross Project NFIRS-SVI Quickstart
#
#
# +
from pathlib import Path
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
pd.set_option('display.max_columns',500)
sns.set()
# +
# Data Import and Cleaning
# +
p = Path.cwd()
data_path = p.parent / 'data' / 'Master Project Data'
nfirs_path = data_path / 'NFIRS Fire Incident Data.csv'
svi2016_path = data_path / 'SVI Tract Data.csv'
svi2016_top = pd.read_csv(svi2016_path,nrows=1000)
svi_col_dtypes = {'ST':str,'STCNTY':str,'FIPS':str}
svi2016 = pd.read_csv(svi2016_path,
index_col=0,
dtype = svi_col_dtypes)
p = Path.cwd()
data_path = p.parent / 'data' / 'Master Project Data'
nfirs_path = data_path / 'NFIRS Fire Incident Data.csv'
cols_to_use = ['State','FDID','City','Zip','inc_date','oth_inj','oth_death','prop_loss',
'cont_loss','tot_loss','GEOID']
col_dtypes = {'GEOID':str}
nfirs = pd.read_csv(nfirs_path,
dtype = col_dtypes,
usecols = cols_to_use,
encoding='latin-1')
nfirs['inc_date'] = pd.to_datetime(nfirs['inc_date'], infer_datetime_format=True)
# -
# Add the severe fire column to the dataset
sev_fire_mask = (nfirs['oth_death'] > 0) | (nfirs['oth_inj'] > 0) | (nfirs['tot_loss'] >= 10000)
nfirs['severe_fire'] = 'not_sev_fire'
nfirs.loc[sev_fire_mask,'severe_fire'] = 'sev_fire'
nfirs['had_inj'] = np.where(nfirs['oth_inj']>0,'had_inj','no_inj')
nfirs['had_death'] = np.where(nfirs['oth_death']>0,'had_death','no_death')
nfirs['10k_loss'] = np.where(nfirs['tot_loss']>=10000,'had_10k_loss','no_10k_loss')
# ## Fix GEOIDs (add leading zeros to correct columns)
nfirs['GEOID'] = (nfirs['GEOID'].str[:-2]
.str.zfill(11))
# Add a year column to be used to groupby in addition to GEOID
nfirs['year'] = nfirs['inc_date'].dt.year
| notebooks/SVI_Starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysing Hillslope Morphology at the Landscape Scale
#
# Hillslope morphology reflects the competition between surface uplift and river erosion, which sets the baselevel conditions for hillslopes. In lab 2 you ran a simple numerical model in order to explore how changes in uplift and erosion influence hillslope development. In this lab, you will look at some hillslope morphology data from a real landscape in order to try and interpret the distribution of erosion in order to infer rates of tectonic processes. The data has already been generated for you by running software developed by Hurst et al. (2012) and Grieve et al (2016) to extract information about hillslope morphology from digitial elevation models (DEMs). This software is slow and was run on a very large dataset so the results have been provided in the data download from Moodle. The aims of this lab are:
# * Follow and execute python code to analyse hillslope morphology extracted from a DEM.
# * Further understand the concept of steady-state and transience in hillslope morphology
# * Quantify metrics of dimensionless hillslope morphology
# * Analyse hillslope morphology at a landscape scale to evaluate the distribution of erosion/uplift.
# * Create and explain figures that show variation in hillslope form in a tectonically active landscape
#
# ## Bolinas
# The landscape you will work on is the Bolinas Ridgeline, which runs adjacent to the San Andreas Fault in California, just north of San Francisco, over the Golden Gate Bridge. You will analyse hillslope morphology in 40 catchments along the fault that drain towards the fault. These catchments are of similar size, but relatively small, no more than ~4 km in length. The elevation of the landscape generally increases from NW to SE.
#
# 
#
# The bedrock beneath these catchments is uniform, consisting of poorly-consolidated metasedimentary rocks (sandstones) of the Franciscan Fm. The key point here is that there should be little variation in channel or hillslope form due to differences in bedrock type. The hillslopes here are all soil-mantled, and thus you can assume transport-limited conditions and that the nonlinear sediment flux law is appropriate to govern downslope soil flux on the hillslopes.
#
# Previous work by Kirby et al (2007) has examined the morphology of the river profiles and found that rivers get progressively steeper, with increasing $M_{\chi}$ from NW to SE. The parameter $M_{\chi}$ is an estimate of channel slope normalised to drainage area, and is considered a good proxy for erosion rate.
#
# 
#
# Kirby et al (2007) suggested that channel erosion rates increased from NW to SE and thus inferred that there was an increase in tectonic uplift rate along this landform. This is important because the high uplift rates make this site a potential nucleus for future earthquakes and San Francisco has a prominent history of being hit by large earthquakes. Millions of people live in the Bay Area, which also has one of the USA's biggest ports, and there is a great deal of industry near by from hi-tech manufacturing to financial services. We will explore whether hillslope morphology reflects the patterns of erosion inferred from the channel network.
#
# ## Python
# The programming language we are using in this lab is called Python. No prior knowledge of programming is required for this lab. Learning how to be a programmer is not the aim! However, this sort of scientific computing is becoming more common place in research and consultancy, so it won't do you any harm to see it in action. Python is multifunctional, for example it can interface with ArcGIS (the software used in the previous lab) to automate workflows. In the last lab we saw how it can be used to construct numerical models and visualise results. In this lab we'll use it to visualise the results of analyses performed on topographic data.
#
# **To run a code block, click in a cell, hold down shift, and press enter.** An asterisk in square brackets `In [*]:` will appear while the code is being executed, and this will change to a number `In [1]:` when the code is finished. *The order in which you execute the code blocks matters, they must be run in sequence.*
#
# Inside blocks of python code there are comments indicated by lines that start with `#`. These lines are not computer code but rather comments providing information about what the code is doing to help you follow along. Before we get started we need to tell python which tools we want to use (these are called modules):
#
# +
# import modules for numerical calculations, for plotting, and for data manipulation
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import pandas as pd
# import operating system functions
import os
# tell python to allow plotting to occur within the page
# %matplotlib inline
# Customise figure style, changing the font to arial and the size to 16pt
from matplotlib import rc
rc('font',**{'family':'sans-serif','sans-serif':['Arial']})
rc('font',size=12)
# -
# Before we get started we need to define the workspace we are working in so that python knows where to go to look for files and data.
# +
# Get the current directory
# DataDirectory = os.getcwd()+"\\data\\" # use this line for windows
DataDirectory = os.getcwd()+"/data/" # use this line if on a mac or on binder
# Set the project name
ProjectName = "Bolinas"
# -
# ## Load the hillslope data
# The file named **Bolinas_HilltopData.csv** contains the results of all the hillslope traces collected along the Bolinas ridge. Each trace starts at a hilltop and follows the steepest path down to an adjacent channel. Open up this file in excel and take a look at the data table. Information about the location of the hilltop ($X$,$Y$,$i$,$j$) are included, as are the location where the traces stopped in the channel network ($a$,$b$,*StreamID*). For our purposes, the hilltop curvature $C_{HT}$, hillslope length $L_H$, hillslope relief $R$ and average hillslope gradient $S$ are all recorded, and this is the data we wil be using mostly. The code below reads this data into Python so that we can explore it, and then checks the data for errors.
# +
# load the hillslopes data
Filename = ProjectName+'_HilltopData.csv'
HillslopeData = pd.read_csv(DataDirectory+Filename)
# drop any rows with no data (hillslope traces to outside the study domain)
# or with value of -9999 for Basin ID
HillslopeData = HillslopeData.dropna()
HillslopeData = HillslopeData[HillslopeData.BasinID != -9999]
# -
# ## Determine *S<sub>C</sub>*
# The critical slope gradient *S<sub>C</sub>* is an important parameter to constrain. This can be done based on topographic data following the work of Grieve et al. (2016). Recall from equation X that *S<sub>C</sub>* is the gradient toward which hillslope sediment transport accelerates rapidly toward infinity. Therefore it should not be possible for a hillslope to be steeper than *S<sub>C</sub>*.
#
# We can determine *S<sub>C</sub>* by looking at the percentage of hillslopes that are steeper than a range of *S<sub>C</sub>* values. We will arbritarily assume that *S<sub>C</sub>* is the value at which >99.5% of hillslopes in the entire Bolinas landscape have a lower slope.
# +
# set up range of S_c values to test from 0.5 to 1.
Sc_test = np.arange(0.5,1.,0.01)
# set up empty array for results
Percent_Less_Than = np.zeros(len(Sc_test))
NoHillslopes = len(HillslopeData)
#loop across the
for i, Sc in enumerate(Sc_test):
# get max hillslope relief
R_test = HillslopeData.Lh*Sc
#compare to actual hillslope relief and count how many fall below max line
#first for segments
BelowMask = HillslopeData.R < R_test
NumberBelow = np.sum(BelowMask)
Percent_Less_Than[i] = 100.*float(NumberBelow)/float(NoHillslopes)
# find the location of the 99.5th percentile
ind = np.argmin(np.abs(99-Percent_Less_Than))
# SET S_C BASED ON THE RESULTS
Sc = Sc_test[ind]
# plot the resulting curve
plt.plot(Sc_test,Percent_Less_Than,'k-',color=[0.5,0.5,0.5])
# plot the percentile
plt.plot([0.5,Sc_test[ind],Sc_test[ind]],[99,99,0.],'r--',lw=0.5)
#plot the value of Sc
plt.text(Sc_test[ind],85,"$S_C = $"+str(Sc_test[ind]),rotation=-90)
#label the axes
plt.xlabel("$S_C$")
plt.ylabel("Percent with lower Relief")
plt.ylim(70,100)
plt.xlim(0.5,1.0)
plt.tight_layout()
plt.savefig(ProjectName+"Sc_Percent.pdf")
# -
# ## Hillslope morphology within an individual catchment
# During the first part of this lab, you will explore hillslope morphology within an individual catchment along the Bolinas ridge. There are 40 catchments to choose from so you can explore the catchments to decide which you think might be the most interesting. The catchments are numbered from NW to SE.
#
# 
# ### Hillslope Traces
#
# The data you will be working with consists of individual hillslope traces from every hilltop down the path of steepest descent to a neigbouring channel pixel. Only the hilltops falling within or on the boundary of the 41 catchments pictured above have been extracted. There are over 150,000 individual hillslope traces, so this is alot of data to analyse and will require us to group the data spatially and calculate summary statistics. The image below shows you some examples of individual hillslope traces across a DEM. The hilltops are marked in black and the channels in blue, the hillslope traces are depicted with red lines.
#
# 
# ### Plot Histograms of Hillslope Morphology
#
# You will start by looking at the distribution of hillslope morphology within your catchments by plotting histograms of the important hillslope metrics: hillslope length, slope, relief and hilltop curvature:
# +
# SET THE BASIN NUMBER YOU WISH TO WORK WITH HERE
Basin = 24
# get the hillslope data only for the basin you are working on
Basins = np.sort(HillslopeData.BasinID.unique())
BasinHillslopeData = HillslopeData[HillslopeData.BasinID == Basins[Basin]]
# setup the figure
# you can modify the figure size here if you wish
fig_width_inches = 6.6
fig_height_inches = 6.6
plt.figure(figsize=(fig_width_inches,fig_height_inches))
# this figure is going to have four parts, called subplots
# we create a subplot by providing a 3 digit number,
# the first two digits refer to the dimensions of the plot (2x2 subplots)
# the third number refers to which of these we want to populate
# HILLSLOPE LENGTH
plt.subplot(221)
# generate the frequency data for a histogram for hillslope length
freq, bin_edges = np.histogram(BasinHillslopeData.Lh,bins=np.arange(0,400,20.))
# normalise the frequency data to the largest value in the dataset
freq_norm = freq.astype(np.float)/float(np.max(freq))
# plot the results as a bar chart
plt.bar(bin_edges[:-1],freq_norm,width=20,align='edge',edgecolor='k',linewidth=0.5,color='steelblue')
plt.xlabel("Hillslope Length (m)")
plt.ylabel("Normalised Frequency")
plt.xlim(0,400)
# HILLSLOPE GRADIENT
plt.subplot(222)
freq, bin_edges = np.histogram(BasinHillslopeData.S,bins=np.arange(0,0.8,0.04))
freq_norm = freq.astype(np.float)/float(np.max(freq))
plt.bar(bin_edges[:-1],freq_norm,width=0.04,align='edge',edgecolor='k',linewidth=0.5,color='thistle')
plt.xlabel("Mean Slope (m/m)")
plt.ylabel("Normalised Frequency")
plt.xlim(0,0.8)
# HILLTOP CURVATURE
plt.subplot(223)
freq, bin_edges = np.histogram(BasinHillslopeData.R,bins=np.arange(0,200.,10.))
freq_norm = freq.astype(np.float)/float(np.max(freq))
plt.bar(bin_edges[:-1],freq_norm,width=10.,align='edge',edgecolor='k',linewidth=0.5,color='sandybrown')
plt.xlabel("Hillslope Relief (m)")
plt.ylabel("Normalised Frequency")
plt.xlim(0,200)
# HILLSLOPE RELIEF
plt.subplot(224)
freq, bin_edges = np.histogram(BasinHillslopeData.Cht,bins=np.arange(-0.1,0,0.005))
freq_norm = freq.astype(np.float)/float(np.max(freq))
plt.bar(bin_edges[:-1],freq_norm,width=0.005,align='edge',edgecolor='k',linewidth=0.5,color='salmon')
plt.xlabel("Hilltop Curvature (m$^{-1}$)")
plt.ylabel("Normalised Frequency")
plt.xlim(0,-0.1)
# FINALISE THE FIGURE AND SAVE
# add a figure title
plt.suptitle("Basin "+str(Basin)+" Hillslopes")
# check the layout, leave some white space around the edges
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
# save the figure
plt.savefig(ProjectName + "_Basin" + "%02d" % Basin + "_HillslopeHists.png", dpi=300)
# -
# ### Calculate the dimensionless hillslope morphology within the basin
#
# Recall from the lecture that hillslopes with different lengths will have different relief. Therefore it is useful to calculate the dimensionless properties of hillslopes so that we can compare between different landscapes.
#
# The nonlinear sediment transport equation predicts a relationship between dimensionless erosion rate and dimensionless relief **for steady state hillslopes** of the form:
# $$
# \begin{equation}
# R^* = {{1}\over{E^*}}\left(\sqrt{1+(E^*)^2} - \ln \left[ {{1}\over{2}} \left( 1+\sqrt{1+(E^*)^2}\right) \right] -1 \right)
# \end{equation}
# $$
# +
# Calculate Theoretical Steady-State E* vs R*
# First declare a range of E* values from 10^-1 (0.1) to 10^3 (1000)
EStarSS = np.logspace(-1,3,1000)
# Calculate RStar as a function of EStar for steady-state hillslopes from equation X
RStarSS = (1./EStarSS)*(np.sqrt(1.+(EStarSS**2.)) - np.log(0.5*(1.+np.sqrt(1+EStarSS**2.))) - 1.)
# -
# When $E^*$ < 1, the model predictions are similar to those of a linear sediment flux law, but for $E^*$ > 1 hillslope relief is limited and hillslope gradients approach the critical value $S_C$ and the hillslopes become steep and planar. Conveniently, the relationship between $E^*$ and $R^*$ can also be cast in terms of properties of hillslopes that are measureable from topography. This means that by measuring these properties we can test whether steady-state hillslopes conform with this flux law, or if we are confident the flux law is appropriate we can test whether hillslopes are at steady state:
#
# Now we will calculate $E^*$ and $R^*$ based on the observed topography from our hillslope traces.
#
# **This next cell of code has been intentionally left blank so that you can complete it yourself.**
#
# You can access the data for hillslope length with `BasinHillslopeData.Lh`, hilltop curvature with `BasinHillslopeData.Cht` and hillslope gradient with `BasinHillslopeData.S`. The equations for these are:
#
# $$
# \begin{equation}
# E^* = {{- 2 \:C_{HT} \:L_H}\over{S_C}} %\tag{}
# \end{equation}
# $$
#
# $$
# \begin{equation}
# R^* = {{S}\over{S_C}} %\tag{}
# \end{equation}
# $$
#
# **Complete the cells below to calculate $E^*$ and $R^*$ for your basin**
# You may need to look back at the histogram code to figure out how to access the specific data you need within the BasinHillslopeData object.
# +
#### YOU NEED TO COMPLETE THIS CELL
# Calculate E* vs R* for our hillslope data
EStar =
RStar =
# -
# Now we will plot the results so we can compare the observed hillslope morphology to the steady-state predictions:
# +
# setup the figure
# you can modify the figure size here if you wish
fig_width_inches = 6.6
fig_height_inches = 5
fig = plt.figure(figsize=(fig_width_inches,fig_height_inches))
ax = plt.subplot()
# Set up the plot to have a lograthmic scale on both axes
plt.loglog()
# Plot analytical steady-state relationship as a dashed line
ax.plot(EStarSS,RStarSS,'k--',zorder=10)
# Plot the observed hillslope data as a grey scatter plot
ax.plot(EStar,RStar,'.',color=[0.5,0.5,0.5])
# Finalise the figure
plt.xlabel('$E^*={{-2\:C_{HT}\:L_H}/{S_C}}$')
plt.ylabel('$R^*=S/S_C$')
plt.xlim(0.1,1000.)
plt.ylim(0.01,1.)
plt.suptitle("Basin "+str(Basin)+" Dimensionless Hillslope Morphology")
plt.tight_layout(rect=[0, 0.03, 1, 0.95])
# -
# The results look quite messy. Some of the data sit near the line, and some of the data sit quite far away. **Remember** these are thousands of individual hillslope traces! To try and make a bit more sense of this data we will do some grouping. The hillslope traces record where in the river network the hillslope terminates, and the channel network has been split into 100 m long segments. We will collect all the hillslope data belonging to each channel segment and conduct some statistics. Because we do not know whether the data is normally distributed, you will use the median as our central tendency indicator (instead of mean). To get an idea of the spread of the data, you will use the 16th and 84th percentiles (instead of standard deviation). These concepts should be familiar to you by now but please ask if you're not sure why this is important.
# +
# get a list of the unique channel segments in the hillslope data
Segments = BasinHillslopeData.StreamID.unique()
# we will create a new dataset to store the results in
# this sets up the new dataset and give names to the columns
SegmentsData = pd.DataFrame(columns=['SegmentNo','EStar','EStarLower','EStarUpper','RStar','RStarLower','RStarUpper','NTraces'])
# loop across all segments in the dataset
for i, Segment in enumerate(Segments):
# Get segment hillslope data as a new data object
SegmentHillslopeData = BasinHillslopeData[BasinHillslopeData.StreamID == float(Segment)]
# Calculate EStar and RStar for the current segment
TempEs = (-2.*SegmentHillslopeData.Cht*SegmentHillslopeData.Lh)/Sc
TempRs = SegmentHillslopeData.S/Sc
# Get the median and quartiles for EStar and RStar
EStarSeg = TempEs.quantile(0.5)
EStarSegUpper = TempEs.quantile(0.84)
EStarSegLower = TempEs.quantile(0.16)
RStarSeg = TempRs.quantile(0.5)
RStarSegUpper = TempRs.quantile(0.84)
RStarSegLower = TempRs.quantile(0.16)
NTraces = SegmentHillslopeData.size
#add to the new data frame
SegmentsData.loc[i] = [Segment,EStarSeg,EStarSegLower,EStarSegUpper,RStarSeg,RStarSegLower,RStarSegUpper,NTraces]
# only keep segments with more than 100 hillslope traces so that we have a good size dataset
SegmentsData = SegmentsData[SegmentsData.NTraces > 100]
# -
# Now that you have segmented the hillslope data, let's plot this instead of/on top of the raw hillslope data and see if it provides a clearer picture of what is going on:
# +
# Plot symbols with error bars with colours but faded (increase alpha)
for i, row in SegmentsData.iterrows():
EStarErr = np.array([[row.EStarLower],[row.EStarUpper]])
RStarErr = np.array([[row.RStarLower],[row.RStarUpper]])
ax.plot([row.EStar,row.EStar],RStarErr,'k-', lw=1,alpha=0.5,zorder=9)
ax.plot(EStarErr,[row.RStar,row.RStar],'k-', lw=1,alpha=0.5,zorder=9)
ax.plot(row.EStar,row.RStar,'ko',ms=4,zorder=32)
# save the figure
plt.savefig(ProjectName + "_Basin" + "%02d" % Basin + "_EStarRStar.png", dpi=300)
# display the updated plot
fig
# -
# ## Hillslope morphology along the Bolinas Ridge
# You have perhaps observed that hillslope morphology data is quite noisy and quite varied. In order to see through the variability within a basin you will now make a series of plots designed to show how hillslope morphology varies from basin to basin. To do this you will calculate the median, 16th and 84th percentiles of the various hillslope metrics and also the channel steepness data derived by Kirby et al. (2007):
# +
# we have already loaded the hillslope data, now we need the channel data
Filename = ProjectName+'_MChiSegmented.csv'
ChannelData = pd.read_csv(DataDirectory+Filename)
# Get rid of sections where there is no data
Segments2Remove = ChannelData[ChannelData.chi == -9999].segment_number.unique()
ChannelData = ChannelData[~ChannelData.segment_number.isin(Segments2Remove)]
# get list of basins
Basins = HillslopeData.BasinID.unique()
Basins.sort()
# create a new data object
BasinData = pd.DataFrame(columns=['Basin','CHT','CHT16','CHT84','LH','LH16','LH84','S','S16','S84','MChi','MChi16','MChi84'])
# calculate catchment summary data for each basin
for i, Basin in enumerate(Basins):
# isolate hillslope data for this basin
BasinHillslopeData = HillslopeData[HillslopeData.BasinID == Basin]
# isolate channel data for this basin
BasinChannelData = ChannelData[ChannelData.basin_key == i]
# curvature stats
CHT = BasinHillslopeData.Cht.quantile(0.5)
CHT16 = BasinHillslopeData.Cht.quantile(0.16)
CHT84 = BasinHillslopeData.Cht.quantile(0.84)
# hillslope length stats
LH = BasinHillslopeData.Lh.quantile(0.5)
LH16 = BasinHillslopeData.Lh.quantile(0.16)
LH84 = BasinHillslopeData.Lh.quantile(0.84)
# slope stats
S = BasinHillslopeData.S.quantile(0.5)
S16 = BasinHillslopeData.S.quantile(0.16)
S84 = BasinHillslopeData.S.quantile(0.84)
# Channel Steepness stats
MChi = BasinChannelData.m_chi.quantile(0.5)
MChi16 = BasinChannelData.m_chi.quantile(0.16)
MChi84 = BasinChannelData.m_chi.quantile(0.84)
# add stats to the new data frame
BasinData.loc[i] = [i,CHT,CHT16,CHT84,LH,LH16,LH84,S,S16,S84,MChi,MChi16,MChi84]
# -
# ### Compare hillslope morphology along the length of the Bolinas Ridge
#
# Now you will make some plots to show how the hillslope and channel morphology vary along the length of the Bolinas Ridge. In the lecture, we identified that hilltop curvature should be a proxy for erosion rates if the hillslopes are adjusted to their baselevel conditions. Hillslope gradient is also expected to increase with erosion rates, but become limited ***IF*** a process transition to more frequent landsliding has occurred. Think about what sort of relationship you might expect between hilltop curvature and hillslope gradient if this is the case. I suggest sketching out the expected relationship before you make the next plot.
#
# Now go ahead and create a plot of hilltop curvature against hillslope gradient:
# +
# setup the figure
# you can modify the figure size here if you wish
fig_width_inches = 6.6
fig_height_inches = 5
fig = plt.figure(figsize=(fig_width_inches,fig_height_inches))
ax = plt.subplot()
#set up a colour map for showing basin number
ColourMap = cm.viridis
# Plot symbols with error bars with colours but faded (increase alpha)
for i, row in BasinData.iterrows():
CHTErr = np.array([[row.CHT16],[row.CHT84]])
SErr = np.array([[row.S16],[row.S84]])
Colour = float(i)/float(len(BasinData))
ax.plot([row.CHT,row.CHT],SErr,'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(CHTErr,[row.S,row.S],'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(row.CHT,row.S,'o',color=ColourMap(Colour),ms=5,zorder=32)
# label the axes
plt.xlabel("Hilltop Curvature (m$^{-1}$)")
plt.ylabel("Hillslope Gradient (m/m)")
# add the colour bar
m = cm.ScalarMappable(cmap=ColourMap)
m.set_array(BasinData.Basin)
cbar = plt.colorbar(m)
cbar.set_label("Basin No.")
# save the figure
plt.savefig(ProjectName + "_Cht_S.png", dpi=300)
# -
# The results are interesting, and have been colour-coded by basin number from NW to SE to help you to interpret the results. But these results don't allow us to compare the hillslope morphology to the theoretical predictions of the nonlinear sediment flux equation. To do that, we again need to calculate $E^*$ and $R^*$.
# +
# Calculate E* vs R* for our hillslope data
EStar = (-2*BasinData.CHT*BasinData.LH)/(Sc)
RStar = BasinData.S/Sc
# Calculate E* vs R* range
EStar16 = (-2*BasinData.CHT16*BasinData.LH16)/(Sc)
RStar16 = BasinData.S16/Sc
EStar84 = (-2*BasinData.CHT84*BasinData.LH84)/(Sc)
RStar84 = BasinData.S84/Sc
# setup the figure
# you can modify the figure size here if you wish
fig_width_inches = 6.6
fig_height_inches = 4.4
fig = plt.figure(figsize=(fig_width_inches,fig_height_inches))
ax = plt.subplot()
#make a logarithmic plot
plt.loglog()
#plot the theoretical relationship between EStar and RStar
ax.plot(EStarSS,RStarSS,'k--',zorder=10)
#set up a colour map for showing basin number
ColourMap = cm.viridis
# Plot symbols with error bars with colours but faded (increase alpha)
for i in range(0,len(Basins)):
EStarErr = np.array([[EStar16[i]],[EStar84[i]]])
RStarErr = np.array([[RStar16[i]],[RStar84[i]]])
Colour = float(i)/float(len(BasinData))
ax.plot([EStar[i],EStar[i]],RStarErr,'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(EStarErr,[RStar[i],RStar[i]],'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(EStar[i],RStar[i],'o',color=ColourMap(Colour),ms=5,zorder=32)
# label the axes and limit
plt.xlabel('$E^*={{-2\:C_{HT}\:L_H}/{S_C}}$')
plt.ylabel('$R^*=S/S_C$')
plt.xlim(0.1,100.)
plt.ylim(0.02,1.)
# add the colour bar
m = cm.ScalarMappable(cmap=ColourMap)
m.set_array(BasinData.Basin)
cbar = plt.colorbar(m)
cbar.set_label("Basin No.")
# save the figure
plt.savefig(ProjectName + "_Estar_Rstar.png", dpi=300)
# -
# It would seem that hillslope morphology is varying systematically along the length of the Bolinas Ridge. The boundary conditions for the hillslopes are set erosion in the adjacent streams. Kirby et al. (2007) suggested the steepness of the channels increasing along the Bolinas Ridge was reflecting increased erosion in response to a gradient in tectonic uplift rates. Let's use our hillslope data to investigate whether hillslope and channel metrics for erosion rates are comparable by plotting channel steepness ($M_\chi$) against $E^*$:
# +
# setup the figure
# you can modify the figure size here if you wish
fig_width_inches = 6.6
fig_height_inches = 4.4
fig = plt.figure(figsize=(fig_width_inches,fig_height_inches))
ax = plt.subplot()
#set up a colour map for showing basin number
ColourMap = cm.viridis
# Get MChi data
MChi = BasinData.MChi
MChi16 = BasinData.MChi16
MChi84 = BasinData.MChi84
# Plot symbols with error bars with colours but faded (increase alpha)
for i in range(0,len(Basins)):
EStarErr = np.array([[EStar16[i]],[EStar84[i]]])
MChiErr = np.array([[MChi16[i]],[MChi84[i]]])
Colour = float(i)/float(len(BasinData))
ax.plot([MChi[i],MChi[i]],EStarErr,'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(MChiErr,[EStar[i],EStar[i]],'-',color=ColourMap(Colour),lw=1,alpha=0.5,zorder=9)
ax.plot(MChi[i],EStar[i],'o',color=ColourMap(Colour),ms=5,zorder=32)
# label the axes and limit
plt.xlabel('Channel Steepness Index $M_{Chi}$')
plt.ylabel('$E^*={{-2\:C_{HT}\:L_H}/{S_C}}$')
# add the colour bar
m = cm.ScalarMappable(cmap=ColourMap)
m.set_array(BasinData.Basin)
cbar = plt.colorbar(m)
cbar.set_label("Basin No.")
# save the figure
plt.savefig(ProjectName + "_MChi_EStar.png", dpi=300)
# -
# <div class="alert alert-block alert-info">
# <font color="black">
# <h3>ASSESSMENT TASK</h3>
# <p>
# Describe the distribution of hillslope morphology within an individual catchment and along the length of the Bolinas Ridge. Evaluate how hillslope morphology compares to the steady-state predictions of the non-linear sediment transport equation and to the boundary conditions set by the channel steepness. You may wish to refer to existing academic literature on this site, and on other locations where hillslope morphology has been analysed in this way to help you. You can choose which figures to include, and remember that your figures will need to have a figure caption. You should approach this task as a short report/essay.
# </p>
# <p></p>
# </font>
# </div>
| Hillslope_Morphology_Tectonics_Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
# +
from VisionSystem import VisionSystem, VisualObject, VideoStream
from VisionSystem.DetectionModel import ThreshBlob
livestream = VideoStream()
# +
my_face = VisualObject(real_size=(0.15, 0.1, 0.3), detection_model=ThreshBlob(), result_limit=1)
vision = VisionSystem(resolution=livestream.resolution, objects_to_track={'me': my_face})
# +
from DisplayPane import DisplayPane
from DisplayPane.Interactor.VisionSystemTuner import VisionSystemTuner
tuner = VisionSystemTuner(vision_system=vision)
DisplayPane(video_stream=livestream, vision_system=vision, interactors=[tuner])
# -
| notebooks/VisionSystemTutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## <NAME> - 12017318 ##
# ### Tic Tac Toe Project ###
# +
# Importing all the required modules
import pygame as pg
import sys
from pygame.locals import *
import numpy as np
pg.init()
# Pygame Screen size
WIDTH = 400
HEIGHT = 400
# Colors
WHITE = (255, 255, 255)
BLACK = (0, 0, 0)
GRAY = (200, 200, 200)
ORANGE = (255, 100, 10)
# Tic Tac Toe Board
MarkedBOX = (np.array([
[0, 0, 0],
[0, 0, 0],
[0, 0, 0]
]))
DRAW = False
WINNER = None
XO = 'o'
mainscreen = pg.display.set_mode((WIDTH, HEIGHT + 100))
pg.display.set_caption("Tic Tac Toe ")
mainscreen.fill(BLACK)
# Window and box(Grid)
def start():
font = pg.font.Font(None, 82)
mainscreen.fill(GRAY)
mainscreen.blit(pg.image.load("pasted image 0.png"), (0, 0))
pg.display.update()
pg.time.wait(1000)
mainscreen.fill(WHITE)
# Two Vertical lines
pg.draw.line(mainscreen, BLACK, (WIDTH / 3, 0), (WIDTH / 3, HEIGHT), 5)
pg.draw.line(mainscreen, BLACK, (WIDTH / 3 * 2, 0), (WIDTH / 3 * 2, HEIGHT), 5)
# Two Horizontal lines
pg.draw.line(mainscreen, BLACK, (0, 0), (WIDTH, 0), 5)
pg.draw.line(mainscreen, BLACK, (0, HEIGHT / 3), (WIDTH, HEIGHT / 3), 5)
pg.draw.line(mainscreen, BLACK, (0, HEIGHT / 3 * 2), (WIDTH, HEIGHT / 3 * 2), 5)
pg.draw.line(mainscreen, BLACK, (0, HEIGHT), (WIDTH, HEIGHT), 5)
# Getting Mouse x and y coordinates
def mousepoint():
global row, col
x, y = pg.mouse.get_pos()
# getting width of the box
if (x < WIDTH / 3) and (y < HEIGHT / 3):
row = 0
col = 0
elif (x > WIDTH / 3 and x < WIDTH / 3 * 2) and (y < HEIGHT / 3):
row = 0
col = 1
elif (x > WIDTH / 3 * 2) and (y < HEIGHT / 3):
row = 0
col = 2
elif (x < WIDTH / 3) and (y > HEIGHT / 3 and y < HEIGHT / 3 * 2):
row = 1
col = 0
elif (x > WIDTH / 3 and x < WIDTH / 3 * 2) and (y > HEIGHT / 3 and y < HEIGHT / 3 * 2):
row = 1
col = 1
elif (x > WIDTH / 3 * 2) and (y > HEIGHT / 3 and y < HEIGHT / 3 * 2):
row = 1
col = 2
elif (x < WIDTH / 3) and (y > HEIGHT / 3 * 2):
row = 2
col = 0
elif (x > WIDTH / 3 and x < WIDTH / 3 * 2) and (y > HEIGHT / 3 * 2):
row = 2
col = 1
elif (x > WIDTH / 3 * 2) and (y > HEIGHT / 3 * 2):
row = 2
col = 2
else:
row = None
col = None
fig(row, col)
# Drawing X and O on window
def fig(row, col):
global XO
if (MarkedBOX[row, col] == 0):
global DRAW
if row == 0:
posx = 65
if row == 1:
posx = WIDTH / 3 + 65
if row == 2:
posx = WIDTH / 3 * 2 + 65
if col == 0:
posy = 65
if col == 1:
posy = HEIGHT / 3 + 65
if col == 2:
posy = HEIGHT / 3 * 2 + 65
# Drawing X and O on mainscreen
if (XO == 'o'):
pg.draw.circle(mainscreen, BLACK, (posy, posx), 40, 8)
MarkedBOX[row][col] = 1
XO = 'x'
else:
pg.draw.line(mainscreen, BLACK, (posy - 30, posx - 30),
(posy + 30, posx + 30), 8)
pg.draw.line(mainscreen, BLACK, (posy + 30, posx - 30),
(posy - 30, posx + 30), 8)
MarkedBOX[row][col] = 2
XO = 'o'
pg.display.update()
Whowins()
else:
pass
# Show Player Message turns
message = XO.upper() + "'s Turn"
font = pg.font.Font(None, 70)
text = font.render(message, True, ORANGE, (WHITE))
textRect = text.get_rect()
textRect.center = (200, 450)
mainscreen.blit(text, textRect)
pg.display.update()
# This Function check winner and draw
def Whowins():
global WINNER
for row in range(0, 3):
if ((MarkedBOX[row][0] == MarkedBOX[row][1] == MarkedBOX[row][2]) and (MarkedBOX[row][0] != 0)):
# this row won
WINNER = MarkedBOX[row][0]
pg.draw.line(mainscreen, BLACK, (0, (row + 1) * HEIGHT / 3 - HEIGHT / 6), \
\
(WIDTH, (row + 1) * HEIGHT / 3 - HEIGHT / 6), 4)
wowinnmess(WINNER)
break
# check for winning columns
for col in range(0, 3):
if (MarkedBOX[0][col] == MarkedBOX[1][col] == MarkedBOX[2][col]) and (MarkedBOX[0][col] != 0):
# this column won
WINNER = MarkedBOX[0][col]
# draw winning line
pg.draw.line(mainscreen, BLACK, ((col + 1) * WIDTH / 3 - WIDTH / 6, 0), \
\
((col + 1) * WIDTH / 3 - WIDTH / 6, HEIGHT), 4)
wowinnmess(WINNER)
break
# check for diagonal WINNERs
if (MarkedBOX[0][0] == MarkedBOX[1][1] == MarkedBOX[2][2]) and (MarkedBOX[0][0] != 0):
# checking if player win diagonally left to right
WINNER = MarkedBOX[0][0]
pg.draw.line(mainscreen, BLACK, (50, 50), (350, 350), 4)
wowinnmess(WINNER)
if (MarkedBOX[0][2] == MarkedBOX[1][1] == MarkedBOX[2][0]) and (MarkedBOX[0][2] != 0):
# checking if player win diagonally right to left
WINNER = MarkedBOX[0][2]
pg.draw.line(mainscreen, BLACK, (350, 50), (50, 350), 4)
wowinnmess(WINNER)
if (all([all(row) for row in MarkedBOX]) and WINNER is None):
# checking if match is draw
DRAW = True
wowinnmess('Match is Draw')
def wowinnmess(winner):
font = pg.font.Font(None, 70)
if winner == 1:
winner = " <NAME> "
elif winner == 2:
winner = " <NAME> "
else:
pass
text = font.render(winner, True, ORANGE, (WHITE))
textRect = text.get_rect()
textRect.center = (200, 450)
mainscreen.blit(text, textRect)
pg.display.update()
pg.time.wait(2000)
reset()
def reset():
global MarkedBOX, DRAW, WINNER
MarkedBOX = (np.array([
[0, 0, 0],
[0, 0, 0],
[0, 0, 0]
]))
XO = 'x'
DRAW = False
WINNER = None
start()
start()
while (True):
for event in pg.event.get():
if event.type == QUIT:
pg.quit()
sys.exit()
elif event.type == MOUSEBUTTONDOWN:
mousepoint()
pg.display.update()
pg.display.flip()
# -
| Tic Tac Toe/Tic Tac Toe .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hyyN-2qyK_T2"
# # RL-Scope - Getting Started
#
# This tutorial will show you the basics of using RL-Scope to collect traces from your training script and visualize their results. We will demonstrate this by applying RL-Scope to the "evaluation loop" of an RL model in the PyTorch based `stable-baselines3` framework.
#
# This tutorial is based off a tutorial for using `stable-baselines3` (from [here](https://github.com/araffin/rl-tutorial-jnrr19/blob/sb3/1_getting_started.ipynb)). So, we will begin by showing how to use `stable-baselines3`, then show how to annotate and profile our code with RL-Scope.
#
# ### Concepts
#
# In this notebook, we will cover:
# - How to install RL-Scope
# - Basic RL-Scope API usage
# - How to collect traces and generate plots using RL-Scope
# - How to use the PyTorch based `stable-baselines3` RL framework to train a model
# - How to write an evaluation inference loop in `stable-baselines3`
#
# + [markdown] id="rcri-PzG1bi-"
# # 1. Using `stable-baselines3` to train and evaluate a model
#
# We will begin by learning the basics for using stable baselines library: how to create a RL model, train it and evaluate it. Because all algorithms share the same interface, we will see how simple it is to switch from one algorithm to another.
#
# <!-- ## Install Dependencies and Stable Baselines3 Using Pip
#
# List of full dependencies can be found in the [README](https://github.com/DLR-RM/stable-baselines3).
#
#
# ```
# pip install stable-baselines3[extra]
# ``` -->
# + [markdown] id="NoUnVxrS7GIm"
# ## Install `stable-baselines3` and PyTorch
# + [markdown] id="JkAf3O61AmtM"
# First, install `stable-baselines3` apt dependencies:
# + colab={"base_uri": "https://localhost:8080/"} id="CtJ3ewjf211W" outputId="801b9a7c-1041-41ac-e856-3097c0b66ae3"
# stable-baselines3 dependencies for visualizing video of trained agents.
# !sudo apt-get install -y ffmpeg freeglut3-dev xvfb
# + [markdown] id="SZ5u_X3YAiPS"
# Next, install `stable-baselines3` `pip` package:
# + colab={"base_uri": "https://localhost:8080/"} id="gWskDE2c9WoN" outputId="9678c128-1e24-4be1-ea9d-91089d5457b5"
# Install stable-baselines3 RL framework.
# !pip install stable-baselines3[extra]==v0.10.0
# + [markdown] id="_HFjowCP8tvX"
# Next, we will figure out which version of CUDA is installed in our host environment. We'll assume its the one at `/usr/local/cuda`:
# > **ASIDE:** depending on your host configuration, you could may have multiple CUDA version installed. The important thing is that the CUDA version of PyTorch and RL-Scope match.
# + colab={"base_uri": "https://localhost:8080/"} id="SvjXPUvBgSXs" outputId="f22e7d51-2aa3-4ea3-c087-0014e2726e7e"
# Determine CUDA version at /usr/local/cuda
# !ls -ld /usr/local/cuda*
import os
import re
m = re.search(r'^cuda-(?P<cuda_version>.*)', os.path.basename(os.path.realpath('/usr/local/cuda')))
cuda_version = m.group('cuda_version')
cu_version = re.sub('\.', '', cuda_version)
cu_suffix = "+cu{ver}".format(ver=cu_version)
# For historical record, this collab instance running Ubuntu 18.04,
# but it should work on newer Ubuntu versions (e.g., 20.04).
# !echo && lsb_release -a
print()
print("> Host CUDA version: {cuda_version}".format(cuda_version=cuda_version))
# + [markdown] id="2xGUGu038lYq"
# Next, we will install the PyTorch package whose CUDA version matches the CUDA version of our host environment.
# + colab={"base_uri": "https://localhost:8080/"} id="B7wIi8A4211Y" outputId="dcbb40db-85f3-4812-d228-28fdde1d93d9"
# 1. Get version of torch installed with stable-baselines3
# 2. Re-install torch with CUDA version that matches /usr/local/cuda
# torch_version = !python -c 'import torch; import re; torch_version = re.sub(r"\+cu.*$", "", torch.__version__); print(torch_version);'
torch_version = torch_version[0]
# !pip uninstall -y torch || true
pip_torch_version = "torch=={ver}{cu}".format(
ver=torch_version,
cu=cu_suffix)
# !pip install $pip_torch_version -f https://download.pytorch.org/whl/torch_stable.html
# torch_version = !python -c 'import torch; import re; torch_version = re.sub(r"\+cu.*$", "", torch.__version__); print(torch_version);'
torch_version = torch_version[0]
# torch_cuda_version = !python -c 'import torch; print(torch.version.cuda);'
torch_cuda_version = torch_cuda_version[0]
torch_cuda_version
print()
print("> PyTorch version : {torch_version}".format(torch_version=torch_version))
print(" PyTorch CUDA version: {torch_cuda_version}".format(torch_cuda_version=torch_cuda_version))
# + [markdown] id="FtY8FhliLsGm"
# ## Imports
# + [markdown] id="gcX8hEcaUpR0"
# Stable-Baselines3 works on environments that follow the [gym interface](https://stable-baselines3.readthedocs.io/en/master/guide/custom_env.html).
# You can find a list of available environment [here](https://gym.openai.com/envs/#classic_control).
#
# It is also recommended to check the [source code](https://github.com/openai/gym) to learn more about the observation and action space of each env, as gym does not have a proper documentation.
# Not all algorithms can work with all action spaces, you can find more in this [recap table](https://stable-baselines3.readthedocs.io/en/master/guide/algos.html)
# + id="BIedd7Pz9sOs"
import gym
import numpy as np
# + [markdown] id="Ae32CtgzTG3R"
# The first thing you need to import is the RL model, check the documentation to know what you can use on which problem
# + id="R7tKaBFrTR0a"
from stable_baselines3 import PPO
# + [markdown] id="-0_8OQbOTTNT"
# The next thing you need to import is the policy class that will be used to create the networks (for the policy/value functions).
# This step is optional as you can directly use strings in the constructor:
#
# ```PPO('MlpPolicy', env)``` instead of ```PPO(MlpPolicy, env)```
#
# Note that some algorithms like `SAC` have their own `MlpPolicy`, that's why using string for the policy is the recommened option.
# + id="ROUJr675TT01"
from stable_baselines3.ppo.policies import MlpPolicy
# + [markdown] id="RapkYvTXL7Cd"
# ## Create the Gym env and instantiate the agent
#
# For this example, we will use CartPole environment, a classic control problem.
#
# "A pole is attached by an un-actuated joint to a cart, which moves along a frictionless track. The system is controlled by applying a force of +1 or -1 to the cart. The pendulum starts upright, and the goal is to prevent it from falling over. A reward of +1 is provided for every timestep that the pole remains upright. "
#
# Cartpole environment: [https://gym.openai.com/envs/CartPole-v1/](https://gym.openai.com/envs/CartPole-v1/)
#
# 
#
#
# We chose the MlpPolicy because the observation of the CartPole task is a feature vector, not images.
#
# The type of action to use (discrete/continuous) will be automatically deduced from the environment action space
#
# Here we are using the [Proximal Policy Optimization](https://stable-baselines3.readthedocs.io/en/master/modules/ppo2.html) algorithm, which is an Actor-Critic method: it uses a value function to improve the policy gradient descent (by reducing the variance).
#
# It combines ideas from [A2C](https://stable-baselines3.readthedocs.io/en/master/modules/a2c.html) (having multiple workers and using an entropy bonus for exploration) and [TRPO](https://stable-baselines.readthedocs.io/en/master/modules/trpo.html) (it uses a trust region to improve stability and avoid catastrophic drops in performance).
#
# PPO is an on-policy algorithm, which means that the trajectories used to update the networks must be collected using the latest policy.
# It is usually less sample efficient than off-policy alorithms like [DQN](https://stable-baselines.readthedocs.io/en/master/modules/dqn.html), [SAC](https://stable-baselines3.readthedocs.io/en/master/modules/sac.html) or [TD3](https://stable-baselines3.readthedocs.io/en/master/modules/td3.html), but is much faster regarding wall-clock time.
#
# + id="pUWGZp3i9wyf"
env = gym.make('CartPole-v1')
model = PPO(MlpPolicy, env, verbose=0)
# + [markdown] id="4efFdrQ7MBvl"
# We create a helper function to evaluate the agent:
# + id="63M8mSKR-6Zt"
def evaluate(model, num_episodes=100):
"""
Evaluate a RL agent
:param model: (BaseRLModel object) the RL Agent
:param num_episodes: (int) number of episodes to evaluate it
:return: (float) Mean reward for the last num_episodes
"""
# This function will only work for a single Environment
env = model.get_env()
all_episode_rewards = []
for i in range(num_episodes):
episode_rewards = []
done = False
obs = env.reset()
while not done:
# _states are only useful when using LSTM policies
action, _states = model.predict(obs)
# here, action, rewards and dones are arrays
# because we are using vectorized env
obs, reward, done, info = env.step(action)
episode_rewards.append(reward)
all_episode_rewards.append(sum(episode_rewards))
mean_episode_reward = np.mean(all_episode_rewards)
print("Mean reward:", mean_episode_reward, "Num episodes:", num_episodes)
return mean_episode_reward
# + [markdown] id="zjEVOIY8NVeK"
# Let's evaluate the un-trained agent, this should be a random agent.
# + colab={"base_uri": "https://localhost:8080/"} id="xDHLMA6NFk95" outputId="2aa72bf3-bcd2-4373-8c82-0175ef9a6f7a"
# Random Agent, before training
mean_reward_before_train = evaluate(model, num_episodes=100)
# + [markdown] id="QjjPxrwkYJ2i"
# Stable-Baselines already provides you with that helper:
# + id="8z6K9YImYJEx"
from stable_baselines3.common.evaluation import evaluate_policy
# + colab={"base_uri": "https://localhost:8080/"} id="4oPTHjxyZSOL" outputId="edaaf565-8fa1-42bf-e87c-08984462ef8d"
mean_reward, std_reward = evaluate_policy(model, env, n_eval_episodes=100)
print(f"mean_reward:{mean_reward:.2f} +/- {std_reward:.2f}")
# + [markdown] id="r5UoXTZPNdFE"
# ## Train the agent and evaluate it
# + colab={"base_uri": "https://localhost:8080/"} id="e4cfSXIB-pTF" outputId="391ff6d7-6adb-4ef5-f76e-725bac9913a1"
# Train the agent for 10000 steps
model.learn(total_timesteps=10000)
# + colab={"base_uri": "https://localhost:8080/"} id="ygl_gVmV_QP7" outputId="c9b7c5ff-8892-4b27-8579-9e405bd3be67"
# Evaluate the trained agent
mean_reward, std_reward = evaluate_policy(model, env, n_eval_episodes=100)
print(f"mean_reward:{mean_reward:.2f} +/- {std_reward:.2f}")
# + [markdown] id="A00W6yY3NkHG"
# Apparently the training went well, the mean reward increased a lot !
# + [markdown] id="_NmTPtsOhzHZ"
# #2. Using RL-Scope to profile the `stable-baselines3` evaluation inference loop
# Lets annotate the evaluation inference loop with RL-Scope annotations to understand where time is spent.
# + [markdown] id="VT3i_nd74Eso"
# ##Install RL-Scope
#
# First, we'll install some required external `apt` dependencies:
# + colab={"base_uri": "https://localhost:8080/"} id="nNC2S0W_4f7L" outputId="31a8b489-adb9-4690-a56c-88c41e182219"
# REQUIRED:
# RL-Scope external dependencies.
# Needed for generating plots.
# !sudo apt install -y texlive-extra-utils # installs 'pdfcrop' command
# !sudo apt install -y poppler-utils # installs 'pdftoppm' command (PDF -> PNG)
# OPTIONAL:
# Command-line tool useful in this tutorial for inspecting trace file directories.
# !sudo apt install -y tree
# + [markdown] id="d9n-YHML5Ei7"
# Next, we will install the RL-Scope `pip` package whose CUDA version matches the CUDA version of our host environment.
#
# + colab={"base_uri": "https://localhost:8080/"} id="ojGXqViF5FAo" outputId="3502dbee-d814-4323-d78b-e9ef03cc76eb"
# RL-Scope version; this corresponds to a tag (with release files uploaded)
# on our github repo:
# https://github.com/UofT-EcoSystem/rlscope
rlscope_version = '0.0.1'
# RL-Scope pre-compiles C++ components for various CUDA versions (e.g., 10.1, 11.0),
# and adds a "+cu101" suffix to the version number (e.g., for 10.1).
# We must use a CUDA version that matches our DL framework (PyTorch) and host environment
# (whatever /usr/local/cuda points to).
# NOTE: this is the same approach taken by PyTorch.
rlscope_pip_version = "rlscope=={ver}{cu}".format(
ver=rlscope_version,
cu=cu_suffix)
# !pip uninstall -y rlscope || true
# !pip install $rlscope_pip_version -f https://uoft-ecosystem.github.io/rlscope/whl
# installed_rlscope_version = !python -c 'import rlscope; rlscope_version = rlscope.__version__; print(rlscope_version);'
installed_rlscope_version = installed_rlscope_version[0]
# rlscope_git_version = !python -c 'import rlscope; print(rlscope.version.git_version);'
rlscope_git_version = rlscope_git_version[0]
print()
print("> RL-Scope version : {rlscope_version}".format(rlscope_version=installed_rlscope_version))
print(" RL-Scope git version: {rlscope_git_version}".format(rlscope_git_version=rlscope_git_version))
# + [markdown] id="a4oij9fC4MAl"
# ##TLDR: How to use RL-Scope
#
# Below, you will find the same code as before, but with additional lines for RL-Scope annotations. The main steps to using RL-Scope are:
#
# 1. Import RL-Scope:
# - ```python
# import rlscope.api as rlscope
# ```
# 2. Add RL-Scope command-line arguments:
# - ```python
# parser = argparse.ArgumentParser(...)
# rlscope.add_rlscope_arguments(parser)
# ```
# 3. Determine where you want profiling to start:
# - ```python
# with rlscope.prof.profile(...):
# # code to profile
# ```
# 4. Add meaningful operation annotations to your code:
# - ```python
# with rlscope.prof.operation('training_loop'):
# for i in range(num_episodes):
# with rlscope.prof.operation('inference'):
# action = model.predict(...)
# with rlscope.prof.operation('step'):
# obs = env.step(action)
# ```
# + [markdown] id="KUqMvP3T4OB8"
# ##Running RL-Scope: `rls-prof`
#
# In order to collect traces, we need to run our training script from the command line using `rls-prof`, so instead of evaluating code piece-by-piece in this notebook, we will write it to `rlscope_tutorial.py` then invoke it using the jupyter's shell syntax:
# <!-- (`!rls-prof python rlscope_tutorial.py ...`). -->
# + colab={"base_uri": "https://localhost:8080/"} id="DgUupt6AiwZj" outputId="11d97e23-cf6e-4936-d17a-3718cbac30b8"
# %%writefile rlscope_tutorial.py
# Executing this will write to ./rlscope_tutorial.py
import gym
import numpy as np
from stable_baselines3 import PPO
from stable_baselines3.ppo.policies import MlpPolicy
import argparse
# Import rlscope.api to interact with rlscope.
import rlscope.api as rlscope
def main():
parser = argparse.ArgumentParser(description="Evaluate an RL policy")
# rlscope will add custom arguments to the argparse argument parser
# that allow you to customize (e.g., "--rlscope-directory <dir>"
# for where to store results).
rlscope.add_rlscope_arguments(parser)
args = parser.parse_args()
# Using the parsed arguments, rlscope will instantiate a singleton
# profiler instance (rlscope.prof).
rlscope.handle_rlscope_args(
parser=parser,
args=args,
)
# Provide a name for the algorithm and simulator (env) used so we can
# generate meaningful plot labels.
# The "process_name" and "phase_name" are useful identifiers for
# multi-process workloads.
rlscope.prof.set_metadata({
'algo': 'PPO',
'env': 'CartPole-v1',
})
process_name = 'PPO_CartPole'
phase_name = process_name
env = gym.make('CartPole-v1')
# Random Agent, before training
model = PPO(MlpPolicy, env, verbose=0)
# "with rlscope.prof.profile(...)" encapsulates the code you wish to profile.
# You can put "setup code" you don't wish to measure before this block.
with rlscope.prof.profile(process_name=process_name, phase_name=phase_name):
mean_reward_before_train = evaluate_rlscope(model,
# num_episodes=10000,
# num_episodes=1000,
num_episodes=100,
)
def evaluate_rlscope(model, num_episodes=100):
"""
Evaluate a RL agent
:param model: (BaseRLModel object) the RL Agent
:param num_episodes: (int) number of episodes to evaluate it
:return: (float) Mean reward for the last num_episodes
"""
# This function will only work for a single Environment
env = model.get_env()
all_episode_rewards = []
# 'training_loop' will capture any time spent in the training loop below
# that isn't captured by a nested annotation (e.g., 'inference')
with rlscope.prof.operation('training_loop'):
for i in range(num_episodes):
# rlscope.prof.report_progress(
# percent_complete=i/float(num_episodes),
# num_timesteps=i,
# total_timesteps=num_episodes)
episode_rewards = []
done = False
obs = env.reset()
while not done:
# 'inference' is the time spent determining the next action to take.
with rlscope.prof.operation('inference'):
# _states are only useful when using LSTM policies
action, _states = model.predict(obs)
# 'step' is the time spent running the simulator on the given action.
with rlscope.prof.operation('step'):
# here, action, rewards and dones are arrays
# because we are using vectorized env
obs, reward, done, info = env.step(action)
episode_rewards.append(reward)
all_episode_rewards.append(sum(episode_rewards))
mean_episode_reward = np.mean(all_episode_rewards)
print("Mean reward:", mean_episode_reward, "Num episodes:", num_episodes)
return mean_episode_reward
if __name__ == '__main__':
main()
# + [markdown] id="L_FAb_xbcWQk"
# Next, we delete any trace files from a previous run of this notebook:
# + id="uGtIXusm51PC"
# !rm -rf ./rlscope_tutorial
# + [markdown] id="oIdhNlDUcceN"
# Next, we use `rls-prof` to run our training script with profiling enabled.
# `--rlscope-directory ./rlscope_tutorial` tells rlscope to all of its profiling traces and graphs in `./rlscope_tutorial`.
#
# > **NOTE:** if you run your training script without `rls-prof`, then no profiling will be performed. This makes it easy to switch between profiling and debugging.
#
# Your training script will be run multiple times to calibrate for overhead correction, so trace collection takes longer than just running your code as usual.
# If you have multiple GPUs, experiments will be run across available GPUs.
#
# After all the configurations are run, `rls-prof` will analyze the collected trace files and generate time breakdown graphs; so lets run the command and check them out:
# + colab={"base_uri": "https://localhost:8080/"} id="dnph6enooRgu" outputId="74b417cc-e8d8-4feb-a0fe-91ab1cbf7b42"
# Run RL-Scope: collect traces, analyze results, and generate plots in ./rlscope_tutorial
# !rls-prof python rlscope_tutorial.py --rlscope-directory ./rlscope_tutorial
# + [markdown] id="AuVDBIgreGDN"
# If `rls-prof` completes without errors, it will generate time breakdown plots. RL-Scope will output `pdf` and `png` versions of the plots for convenience.
# + colab={"base_uri": "https://localhost:8080/"} id="LY-JSYIR211g" outputId="7dc31836-66ff-4f9f-84d3-68e114df3325"
# Look at the plot files generated by RL-Scope
# !ls -l ./rlscope_tutorial/*.png
# + [markdown] id="XGXqZuA8e7az"
# ##RL-Scope plots
#
# Lets look at two of the plots RL-Scope generated, namely the time breakdown plot, and the transition plot.
#
#
#
# + [markdown] id="GarRqNMZfUxb"
# First, lets define some helper functions for displaying our plots.
# + id="MuJFQQ-P211g"
# Define helper functions for plotting PNG files.
from IPython.core.display import display
from IPython.display import IFrame
from IPython.display import Image
from glob import glob
def display_pdfs(glob_expr):
"""
Display all PDFs found in file glob expansion.
e.g.
"""
paths = glob(glob_expr)
pdfs = []
for path in paths:
# pdfs.append(WImage(path))
pdfs.append(IFrame(path, width="100%", height="500"))
for path, pdf in zip(paths, pdfs):
display(path)
display(pdf)
# NOTE: using width="100%" doesn't work in collab...instead hardcode width.
IMAGE_WIDTH_PIXELS = 600
def display_imgs(glob_expr):
"""
Display all PDFs found in file glob expansion.
e.g.
"""
paths = glob(glob_expr)
imgs = []
for path in paths:
imgs.append(Image(path, width=IMAGE_WIDTH_PIXELS))
for path, img in zip(paths, imgs):
display(path)
display(img)
# + [markdown] id="C175BkHOlBvW"
# 1. **Time breakdown plot:**
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 399} id="uJ0KJZNfk8wT" outputId="96f546d9-4a5a-4d73-efbb-3f251ddae18d"
display_imgs('./rlscope_tutorial/*.operation_training_time.png')
# + [markdown] id="-fSEarN1lF7e"
# This plot shows us the total training time broken down by operation, resource type (`CPU`, `GPU`), and fine-grained category (e.g., `CUDA`, `Backend`). The vertical bars show us the operation names we added to our program (`training_loop`, `inference`, `step`). Within each operation, stacked bars further break down the time into resource type, which are colour coded as
# $\color{lightgreen}{\text{GPU}}$,
# $\color{lightblue}{\text{CPU + GPU}}$,
# $\color{red}{\text{CPU}}$.
#
# Within each resource type (colour), time is further divided into fine-grained categories (hatch pattern):
# - **Backend** is CPU time spent in the C++ backend of the DL framework (PyTorch here).
# - **CUDA** is CPU time spent executing CUDA API calls (e.g., `cudaLaunchKernel`, `cudaMemcpy`).
# - **Python** is CPU time spent executing python code in the python interpretter.
# + [markdown] id="MzN4SWfVlvVk"
# 2. **Transition plot:**
# + colab={"base_uri": "https://localhost:8080/", "height": 380} id="5L--9gnelA6C" outputId="4385c7b8-1f73-43c4-9b56-2fbdf59ad256"
display_imgs('./rlscope_tutorial/CategoryTransitionPlot.combined.png')
# + [markdown] id="9b6TUI9xlHOX"
# This plot show the number of language transitions.
#
# - *Backend*: Python$\rightarrow$*Backend*
#
# Calls from Python into the C++ backend of the DL framework.
#
# - *CUDA*: Backend$\rightarrow$*CUDA*
#
# CUDA API calls made by the framework.
#
# A large number of language transitions can result in large times spent CPU-bound. Eager execution suffers more from this compared to more optimized execution models (e.g., AutoGraph in TensorFlow, TorchScript in PyTorch, Graph in TensorFlow v1).
# + [markdown] id="FrI6f5fWnzp-"
# # Conclusion and next steps
#
# In this notebook, we covered:
# - How to install RL-Scope
# - Basic RL-Scope API usage
# - How to collect traces and generate plots using RL-Scope
# - How to use the PyTorch based `stable-baselines3` RL framework to train a model
# - How to write an evaluation inference loop in `stable-baselines3`
#
# ### Next steps
#
# For simplicity, we only profiled the evaluation inference loop. However, to profile RL algorithm training time, we will need to add RL-Scope annotations to the underlying RL framework. This typically requires modifying the underlying RL framework so can profile portions of the training loop.
#
# In a future tutorial (TODO) we will show how to modify the `stable-baselines3` framework to profile an RL training time.
# + [markdown] id="LbVUKuqf3MyA"
# # Resources / Links
#
# For more information on the RL-Scope profiling tool:
# - Installing RL-Scope: https://rl-scope.readthedocs.io/en/latest/installation.html
# - Reproducing RL-Scope paper figures: https://rl-scope.readthedocs.io/en/latest/artifacts.html
# - Developing/buildilng RL-Scope in a docker development environment: https://rl-scope.readthedocs.io/en/latest/host_config.html
#
# For more information on the PyTorch based RL framework `stable-baselines3`:
# - Jupyter notebook github: https://github.com/araffin/rl-tutorial-jnrr19/tree/sb3/
# - `stable-baselines3` github: https://github.com/DLR-RM/stable-baselines3
# - `stable-baselines3` documentation: https://stable-baselines3.readthedocs.io/en/master/
# - RL Baselines3 zoo: https://github.com/DLR-RM/rl-baselines3-zoo
#
# RL Baselines3 Zoo is a collection of pre-tuned and pre-trained Reinforcement Learning agents using `stable-Baselines3`.
| jupyter/01_rlscope_getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''.venv'': venv)'
# name: python3
# ---
# # Maximum Mean Discrepancy drift detector on CIFAR-10
#
# ### Method
#
# The [Maximum Mean Discrepancy (MMD)](http://jmlr.csail.mit.edu/papers/v13/gretton12a.html) detector is a kernel-based method for multivariate 2 sample testing. The MMD is a distance-based measure between 2 distributions *p* and *q* based on the mean embeddings $\mu_{p}$ and $\mu_{q}$ in a reproducing kernel Hilbert space $F$:
#
# \begin{align}
# MMD(F, p, q) & = || \mu_{p} - \mu_{q} ||^2_{F} \\
# \end{align}
#
# We can compute unbiased estimates of $MMD^2$ from the samples of the 2 distributions after applying the kernel trick. We use by default a [radial basis function kernel](https://en.wikipedia.org/wiki/Radial_basis_function_kernel), but users are free to pass their own kernel of preference to the detector. We obtain a $p$-value via a [permutation test](https://en.wikipedia.org/wiki/Resampling_(statistics)) on the values of $MMD^2$. This method is also described in [Failing Loudly: An Empirical Study of Methods for Detecting Dataset Shift](https://arxiv.org/abs/1810.11953).
#
# ### Backend
#
# The method is implemented in both the *PyTorch* and *TensorFlow* frameworks with support for CPU and GPU. Various preprocessing steps are also supported out-of-the box in Alibi Detect for both frameworks and illustrated throughout the notebook. Alibi Detect does however not install PyTorch for you. Check the [PyTorch docs](https://pytorch.org/) how to do this.
#
# ### Dataset
#
# [CIFAR10](https://www.cs.toronto.edu/~kriz/cifar.html) consists of 60,000 32 by 32 RGB images equally distributed over 10 classes. We evaluate the drift detector on the CIFAR-10-C dataset ([Hendrycks & Dietterich, 2019](https://arxiv.org/abs/1903.12261)). The instances in
# CIFAR-10-C have been corrupted and perturbed by various types of noise, blur, brightness etc. at different levels of severity, leading to a gradual decline in the classification model performance. We also check for drift against the original test set with class imbalances.
# +
from functools import partial
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from alibi_detect.cd import MMDDrift
from alibi_detect.models.tensorflow.resnet import scale_by_instance
from alibi_detect.utils.fetching import fetch_tf_model
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.datasets import fetch_cifar10c, corruption_types_cifar10c
# -
# ### Load data
#
# Original CIFAR-10 data:
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.cifar10.load_data()
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
y_train = y_train.astype('int64').reshape(-1,)
y_test = y_test.astype('int64').reshape(-1,)
# For CIFAR-10-C, we can select from the following corruption types at 5 severity levels:
corruptions = corruption_types_cifar10c()
print(corruptions)
# Let's pick a subset of the corruptions at corruption level 5. Each corruption type consists of perturbations on all of the original test set images.
corruption = ['gaussian_noise', 'motion_blur', 'brightness', 'pixelate']
X_corr, y_corr = fetch_cifar10c(corruption=corruption, severity=5, return_X_y=True)
X_corr = X_corr.astype('float32') / 255
# We split the original test set in a reference dataset and a dataset which should not be rejected under the *H<sub>0</sub>* of the MMD test. We also split the corrupted data by corruption type:
np.random.seed(0)
n_test = X_test.shape[0]
idx = np.random.choice(n_test, size=n_test // 2, replace=False)
idx_h0 = np.delete(np.arange(n_test), idx, axis=0)
X_ref,y_ref = X_test[idx], y_test[idx]
X_h0, y_h0 = X_test[idx_h0], y_test[idx_h0]
print(X_ref.shape, X_h0.shape)
# check that the classes are more or less balanced
classes, counts_ref = np.unique(y_ref, return_counts=True)
counts_h0 = np.unique(y_h0, return_counts=True)[1]
print('Class Ref H0')
for cl, cref, ch0 in zip(classes, counts_ref, counts_h0):
assert cref + ch0 == n_test // 10
print('{} {} {}'.format(cl, cref, ch0))
n_corr = len(corruption)
X_c = [X_corr[i * n_test:(i + 1) * n_test] for i in range(n_corr)]
# We can visualise the same instance for each corruption type:
# + tags=["hide_input"]
i = 4
n_test = X_test.shape[0]
plt.title('Original')
plt.axis('off')
plt.imshow(X_test[i])
plt.show()
for _ in range(len(corruption)):
plt.title(corruption[_])
plt.axis('off')
plt.imshow(X_corr[n_test * _+ i])
plt.show()
# -
# We can also verify that the performance of a classification model on CIFAR-10 drops significantly on this perturbed dataset:
dataset = 'cifar10'
model = 'resnet32'
clf = fetch_tf_model(dataset, model)
acc = clf.evaluate(scale_by_instance(X_test), y_test, batch_size=128, verbose=0)[1]
print('Test set accuracy:')
print('Original {:.4f}'.format(acc))
clf_accuracy = {'original': acc}
for _ in range(len(corruption)):
acc = clf.evaluate(scale_by_instance(X_c[_]), y_test, batch_size=128, verbose=0)[1]
clf_accuracy[corruption[_]] = acc
print('{} {:.4f}'.format(corruption[_], acc))
# Given the drop in performance, it is important that we detect the harmful data drift!
#
# ### Detect drift with TensorFlow backend
#
# First we try a drift detector using the *TensorFlow* framework for both the preprocessing and the *MMD* computation steps.
#
# We are trying to detect data drift on high-dimensional (*32x32x3*) data using a multivariate MMD permutation test. It therefore makes sense to apply dimensionality reduction first. Some dimensionality reduction methods also used in [Failing Loudly: An Empirical Study of Methods for Detecting Dataset Shift](https://arxiv.org/pdf/1810.11953.pdf) are readily available: a randomly initialized encoder (**UAE** or Untrained AutoEncoder in the paper), **BBSDs** (black-box shift detection using the classifier's softmax outputs) and **PCA** (using `scikit-learn`).
#
# #### Random encoder
#
# First we try the randomly initialized encoder:
# +
from tensorflow.keras.layers import Conv2D, Dense, Flatten, InputLayer, Reshape
from alibi_detect.cd.tensorflow import preprocess_drift
tf.random.set_seed(0)
# define encoder
encoding_dim = 32
encoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(32, 32, 3)),
Conv2D(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(128, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(512, 4, strides=2, padding='same', activation=tf.nn.relu),
Flatten(),
Dense(encoding_dim,)
]
)
# define preprocessing function
preprocess_fn = partial(preprocess_drift, model=encoder_net, batch_size=512)
# initialise drift detector
cd = MMDDrift(X_ref, backend='tensorflow', p_val=.05,
preprocess_fn=preprocess_fn, n_permutations=100)
# we can also save/load an initialised detector
filepath = 'my_path' # change to directory where detector is saved
save_detector(cd, filepath)
cd = load_detector(filepath)
# -
# Let's check whether the detector thinks drift occurred on the different test sets and time the prediction calls:
# +
from timeit import default_timer as timer
labels = ['No!', 'Yes!']
def make_predictions(cd, x_h0, x_corr, corruption):
t = timer()
preds = cd.predict(x_h0)
dt = timer() - t
print('No corruption')
print('Drift? {}'.format(labels[preds['data']['is_drift']]))
print(f'p-value: {preds["data"]["p_val"]:.3f}')
print(f'Time (s) {dt:.3f}')
if isinstance(x_corr, list):
for x, c in zip(x_corr, corruption):
t = timer()
preds = cd.predict(x)
dt = timer() - t
print('')
print(f'Corruption type: {c}')
print('Drift? {}'.format(labels[preds['data']['is_drift']]))
print(f'p-value: {preds["data"]["p_val"]:.3f}')
print(f'Time (s) {dt:.3f}')
# -
make_predictions(cd, X_h0, X_c, corruption)
# As expected, drift was only detected on the corrupted datasets.
# #### BBSDs
#
# For **BBSDs**, we use the classifier's softmax outputs for black-box shift detection. This method is based on [Detecting and Correcting for Label Shift with Black Box Predictors](https://arxiv.org/abs/1802.03916). The ResNet classifier is trained on data standardised by instance so we need to rescale the data.
X_ref_bbsds = scale_by_instance(X_ref)
X_h0_bbsds = scale_by_instance(X_h0)
X_c_bbsds = [scale_by_instance(X_c[i]) for i in range(n_corr)]
# Initialisation of the drift detector. Here we use the output of the softmax layer to detect the drift, but other hidden layers can be extracted as well by setting *'layer'* to the index of the desired hidden layer in the model:
# +
from alibi_detect.cd.tensorflow import HiddenOutput
# define preprocessing function
preprocess_fn = partial(preprocess_drift, model=HiddenOutput(clf, layer=-1), batch_size=128)
# initialise drift detector
cd = MMDDrift(X_ref_bbsds, backend='tensorflow', p_val=.05,
preprocess_fn=preprocess_fn, n_permutations=100)
# -
make_predictions(cd, X_h0_bbsds, X_c_bbsds, corruption)
# Again drift is only flagged on the perturbed data.
#
# ### Detect drift with PyTorch backend
#
# We can do the same thing using the *PyTorch* backend. We illustrate this using the randomly initialized encoder as preprocessing step:
# +
import torch
import torch.nn as nn
# set random seed and device
seed = 0
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
# -
# Since our *PyTorch* encoder expects the images in a *(batch size, channels, height, width)* format, we transpose the data:
# +
def permute_c(x):
return np.transpose(x.astype(np.float32), (0, 3, 1, 2))
X_ref_pt = permute_c(X_ref)
X_h0_pt = permute_c(X_h0)
X_c_pt = [permute_c(xc) for xc in X_c]
print(X_ref_pt.shape, X_h0_pt.shape, X_c_pt[0].shape)
# +
from alibi_detect.cd.pytorch import preprocess_drift
# define encoder
encoder_net = nn.Sequential(
nn.Conv2d(3, 64, 4, stride=2, padding=0),
nn.ReLU(),
nn.Conv2d(64, 128, 4, stride=2, padding=0),
nn.ReLU(),
nn.Conv2d(128, 512, 4, stride=2, padding=0),
nn.ReLU(),
nn.Flatten(),
nn.Linear(2048, encoding_dim)
).to(device).eval()
# define preprocessing function
preprocess_fn = partial(preprocess_drift, model=encoder_net, device=device, batch_size=512)
# initialise drift detector
cd = MMDDrift(X_ref_pt, backend='pytorch', p_val=.05,
preprocess_fn=preprocess_fn, n_permutations=100)
# -
make_predictions(cd, X_h0_pt, X_c_pt, corruption)
# The drift detector will attempt to use the GPU if available and otherwise falls back on the CPU. We can also explicitly specify the device. Let's compare the GPU speed up with the CPU implementation:
# +
device = torch.device('cpu')
preprocess_fn = partial(preprocess_drift, model=encoder_net.to(device),
device=device, batch_size=512)
cd = MMDDrift(X_ref_pt, backend='pytorch', preprocess_fn=preprocess_fn, device='cpu')
# -
make_predictions(cd, X_h0_pt, X_c_pt, corruption)
# Notice the over **30x acceleration** provided by the GPU.
#
# Similar to the *TensorFlow* implementation, *PyTorch* can also use the hidden layer output from a pretrained model for the preprocessing step via:
#
# ```python
# from alibi_detect.cd.pytorch import HiddenOutput
# ```
| examples/cd_mmd_cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''base'': conda)'
# name: python37464bitbaseconda09becdb38b044ff7a798497c9762d813
# ---
# +
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# +
import pandas as pd
from sklearn.model_selection import train_test_split
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['status_group'], random_state=42)
train.shape, test.shape, val.shape
# +
import numpy as np
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# return the wrangled dataframe
return X
# -
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# +
target = 'status_group'
train_features = train.drop(columns=[target])
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
cardinality = train_features.select_dtypes(exclude='number').nunique()
categorical_features = cardinality[cardinality <= 50].index.tolist()
features = numeric_features + categorical_features
# -
X_train
| module2-random-forests/torch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
import os
for dirname, _, filenames in os.walk('/kaggle/input/cord-19-eda-parse-json-and-generate-clean-csv'):
for filename in filenames:
print(os.path.join(dirname, filename))
# -
from fastai.text import *
from sklearn.model_selection import train_test_split
from langdetect import detect
# !pip install googletrans
from googletrans import Translator
# [[](http://)](http://)
#
# * https://www.kaggle.com/xhlulu/cord-19-eda-parse-json-and-generate-clean-csv
# * https://www.kaggle.com/danielwolffram/topic-modeling-finding-related-articles
#
#
# First we load the data inside each of the csv files.
#
path = '/kaggle/input/cord-19-eda-parse-json-and-generate-clean-csv/'
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
pmc_custom_license_df = pd.read_csv(path + "clean_pmc.csv")
noncomm_use_df = pd.read_csv(path + "clean_noncomm_use.csv")
comm_use_df = pd.read_csv(path + "clean_comm_use.csv")
biorxiv = pd.read_csv(path + "biorxiv_clean.csv")
# -
data = pd.concat([pmc_custom_license_df, noncomm_use_df, comm_use_df, biorxiv], axis =0)
# We subset the dataset to have the paper id and text of the document only
covid_papers = data[data.text.str.contains('COVID-19|SARS-CoV-2|2019-nCov|SARS Coronavirus 2|2019 Novel Coronavirus')][['paper_id', 'text']]
data = covid_papers
# To detect the language of the papers we use the function detect from the langdetect library
data['language'] = data['text'].map(detect)
data.head()
data.shape
data['language'].value_counts()
# Four papers are written in Spanish.
indices_to_translate = data[data['language']=='es']['text'].index
translator = Translator()
for i in indices_to_translate:
if (len(data.loc[i, 'text']) > 3900):
data.loc[i, 'text'] = data.loc[i, 'text'][:3900]
paper = translator.translate(data.loc[i, 'text'])
data.loc[i, 'text'] = paper.text
data.loc[indices_to_translate, 'text']
# The papers are now translated
data.to_csv('data_covid19_papers.csv')
# # Language model data
#
#
# * Training on 90% of the data and validating on 10%
# * The function TextLMDataBunch creates a TextDataBunch suitable for training a language model.
#
train, validation = train_test_split(data, test_size=0.1, random_state=42, shuffle= True)
data_lm = TextLMDataBunch.from_df('.', train, validation,
label_cols='text')
# Saving the results to save time in the future.
data_lm.save('data_lm_export.pkl')
data_lm = load_data('.', 'data_lm_export.pkl')
# Now making the language model learn. It has already learnt some features of basic English using Wikipedia and now I am training it to learn the peculiarities of Covid-19 papers and articles.
learn = language_model_learner(data_lm, AWD_LSTM, drop_mult=0.5)
learn.lr_find()
learn.recorder.plot()
# Training for 4 epochs
learn.fit_one_cycle(4, 2e-2)
learn.lr_find()
learn.recorder.plot()
learn.unfreeze()
learn.fit_one_cycle(10, slice(1e-4, 1e-3))
learn.save_encoder('ft_enc')
#
# # Inference
#
np.random.seed(42)
learn.predict("risks factors", n_words=100, temperature=0.1)
learn.predict("smoking", n_words=100, no_unk=True, temperature=0.1)
learn.predict("pre-existing pulmonary disease", n_words=50, no_unk=True, temperature=0.05)
learn.predict("Co-infections", n_words=50, no_unk=True, temperature=0.1)
learn.predict("comorbidities", n_words=50, no_unk=True, temperature=0.1)
learn.predict("Neonates", n_words=50, no_unk=True, temperature=0.1)
learn.predict("pregnant women", n_words=50, no_unk=True, temperature=0.1)
learn.predict("Socio-economic factors", n_words=100, no_unk=True, temperature=0.1)
learn.predict("behavioral factors", n_words=80, no_unk=True, temperature=0.1)
learn.predict("Transmission dynamics of the virus", n_words=50, no_unk=True, temperature=0.1)
learn.predict("basic reproductive number", n_words=80, no_unk=True, temperature=0.1)
learn.predict("incubation period", n_words=50, no_unk=True, temperature=0.1)
learn.predict("serial interval", n_words=50, no_unk=True, temperature=0.1)
learn.predict("environmental factors", n_words=50, no_unk=True, temperature=0.1)
learn.predict("modes of transmission", n_words=50, no_unk=True, temperature=0.1)
learn.predict("Severity of disease", n_words=50, no_unk=True, temperature=0.1)
learn.predict("risk of fatality among symptomatic hospitalized patients", n_words=50, no_unk=True, temperature=0.1)
learn.predict("high-risk patient", n_words=50, no_unk=True, temperature=0.1)
learn.predict("Susceptibility of populations", n_words=50, no_unk=True, temperature=0.1)
learn.predict("Public health mitigation measures", n_words=50, no_unk=True, temperature=0.1)
learn.predict("What do we know about COVID-19 risk factors?", n_words=100, no_unk=True, temperature=0.1)
data.head()
| Covid-19_LitLM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi-Task Learning Example
# This is a simple example to show how to use mxnet for multi-task learning.
#
# The network is jointly going to learn whether a number is odd or even and to actually recognize the digit.
#
#
# For example
#
# - 1 : 1 and odd
# - 2 : 2 and even
# - 3 : 3 and odd
#
# etc
#
# In this example we don't expect the tasks to contribute to each other much, but for example multi-task learning has been successfully applied to the domain of image captioning. In [A Multi-task Learning Approach for Image Captioning](https://www.ijcai.org/proceedings/2018/0168.pdf) by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, they train a network to jointly classify images and generate text captions
# +
import logging
import random
import time
import matplotlib.pyplot as plt
import mxnet as mx
from mxnet import gluon, np, npx, autograd
import numpy as onp
# -
# ### Parameters
batch_size = 128
epochs = 5
ctx = mx.gpu() if mx.context.num_gpus() > 0 else mx.cpu()
lr = 0.01
# ## Data
#
# We get the traditionnal MNIST dataset and add a new label to the existing one. For each digit we return a new label that stands for Odd or Even
# 
train_dataset = gluon.data.vision.MNIST(train=True)
test_dataset = gluon.data.vision.MNIST(train=False)
def transform(x,y):
x = x.transpose((2,0,1)).astype('float32')/255.
y1 = y
y2 = y % 2 #odd or even
return x, onp.float32(y1), onp.float32(y2)
# We assign the transform to the original dataset
train_dataset_t = train_dataset.transform(transform)
test_dataset_t = test_dataset.transform(transform)
# We load the datasets DataLoaders
train_data = gluon.data.DataLoader(train_dataset_t, shuffle=True, last_batch='rollover', batch_size=batch_size, num_workers=5)
test_data = gluon.data.DataLoader(test_dataset_t, shuffle=False, last_batch='rollover', batch_size=batch_size, num_workers=5)
print("Input shape: {}, Target Labels: {}".format(train_dataset[0][0].shape, train_dataset_t[0][1:]))
# ## Multi-task Network
#
# The output of the featurization is passed to two different outputs layers
class MultiTaskNetwork(gluon.HybridBlock):
def __init__(self):
super(MultiTaskNetwork, self).__init__()
self.shared = gluon.nn.HybridSequential()
self.shared.add(
gluon.nn.Dense(128, activation='relu'),
gluon.nn.Dense(64, activation='relu'),
gluon.nn.Dense(10, activation='relu')
)
self.output1 = gluon.nn.Dense(10) # Digist recognition
self.output2 = gluon.nn.Dense(1) # odd or even
def forward(self, x):
y = self.shared(x)
output1 = self.output1(y)
output2 = self.output2(y)
return output1, output2
# We can use two different losses, one for each output
loss_digits = gluon.loss.SoftmaxCELoss()
loss_odd_even = gluon.loss.SigmoidBCELoss()
# We create and initialize the network
mx.np.random.seed(42)
random.seed(42)
net = MultiTaskNetwork()
net.initialize(mx.init.Xavier(), ctx=ctx)
net.hybridize() # hybridize for speed
trainer = gluon.Trainer(net.collect_params(), 'adam', {'learning_rate':lr})
# ## Evaluate Accuracy
# We need to evaluate the accuracy of each task separately
def evaluate_accuracy(net, data_iterator):
acc_digits = mx.gluon.metric.Accuracy(name='digits')
acc_odd_even = mx.gluon.metric.Accuracy(name='odd_even')
for i, (data, label_digit, label_odd_even) in enumerate(data_iterator):
data = data.as_in_ctx(ctx)
label_digit = label_digit.as_in_ctx(ctx)
label_odd_even = label_odd_even.as_in_ctx(ctx).reshape(-1,1)
output_digit, output_odd_even = net(data)
acc_digits.update(label_digit, npx.softmax(output_digit))
acc_odd_even.update(label_odd_even, npx.sigmoid(output_odd_even) > 0.5)
return acc_digits.get(), acc_odd_even.get()
# ## Training Loop
# We need to balance the contribution of each loss to the overall training and do so by tuning this alpha parameter within [0,1].
alpha = 0.5 # Combine losses factor
for e in range(epochs):
# Accuracies for each task
acc_digits = mx.gluon.metric.Accuracy(name='digits')
acc_odd_even = mx.gluon.metric.Accuracy(name='odd_even')
# Accumulative losses
l_digits_ = 0.
l_odd_even_ = 0.
for i, (data, label_digit, label_odd_even) in enumerate(train_data):
data = data.as_in_ctx(ctx)
label_digit = label_digit.as_in_ctx(ctx)
label_odd_even = label_odd_even.as_in_ctx(ctx).reshape(-1,1)
with autograd.record():
output_digit, output_odd_even = net(data)
l_digits = loss_digits(output_digit, label_digit)
l_odd_even = loss_odd_even(output_odd_even, label_odd_even)
# Combine the loss of each task
l_combined = (1-alpha)*l_digits + alpha*l_odd_even
l_combined.backward()
trainer.step(data.shape[0])
l_digits_ += l_digits.mean()
l_odd_even_ += l_odd_even.mean()
acc_digits.update(label_digit, npx.softmax(output_digit))
acc_odd_even.update(label_odd_even, npx.sigmoid(output_odd_even) > 0.5)
print("Epoch [{}], Acc Digits {:.4f} Loss Digits {:.4f}".format(
e, acc_digits.get()[1], l_digits_.item()/(i+1)))
print("Epoch [{}], Acc Odd/Even {:.4f} Loss Odd/Even {:.4f}".format(
e, acc_odd_even.get()[1], l_odd_even_.item()/(i+1)))
print("Epoch [{}], Testing Accuracies {}".format(e, evaluate_accuracy(net, test_data)))
# ## Testing
def get_random_data():
idx = random.randint(0, len(test_dataset))
img = test_dataset[idx][0]
data, _, _ = test_dataset_t[idx]
data = np.expand_dims(data.as_in_ctx(ctx), axis=0)
plt.imshow(img.squeeze().asnumpy(), cmap='gray')
return data
# +
data = get_random_data()
digit, odd_even = net(data)
digit = digit.argmax(axis=1)[0].asnumpy()
odd_even = (npx.sigmoid(odd_even)[0] > 0.5).asnumpy()
print("Predicted digit: {}, odd: {}".format(digit, odd_even))
| example/multi-task/multi-task-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="X6WDvajSqIDs"
# # Lab 3: Gesture Recognition using Convolutional Neural Networks
#
# **Deadlines**: Feb 8, 5:00PM
#
# **Late Penalty**: There is a penalty-free grace period of one hour past the deadline. Any work that is submitted between 1 hour and 24 hours past the deadline will receive a 20% grade deduction. No other late work is accepted. Quercus submission time will be used, not your local computer time. You can submit your labs as many times as you want before the deadline, so please submit often and early.
#
# **Grading TAs**:
# <NAME>
#
# This lab is based on an assignment developed by Prof. <NAME>.
#
# This lab will be completed in two parts. In Part A you will you will gain experience gathering your own data set (specifically images of hand gestures), and understand the challenges involved in the data cleaning process. In Part B you will train a convolutional neural network to make classifications on different hand gestures. By the end of the lab, you should be able to:
#
# 1. Generate and preprocess your own data
# 2. Load and split data for training, validation and testing
# 3. Train a Convolutional Neural Network
# 4. Apply transfer learning to improve your model
#
# Note that for this lab we will not be providing you with any starter code. You should be able to take the code used in previous labs, tutorials and lectures and modify it accordingly to complete the tasks outlined below.
#
# ### What to submit
#
# **Submission for Part A:**
# Submit a zip file containing your images. Three images each of American Sign Language gestures for letters A - I (total of 27 images). You will be required to clean the images before submitting them. Details are provided under Part A of the handout.
#
# Individual image file names should follow the convention of student-number_Alphabet_file-number.jpg
# (e.g. 100343434_A_1.jpg).
#
#
# **Submission for Part B:**
# Submit a PDF file containing all your code, outputs, and write-up
# from parts 1-5. You can produce a PDF of your Google Colab file by
# going to **File > Print** and then save as PDF. The Colab instructions
# has more information. Make sure to review the PDF submission to ensure that your answers are easy to read. Make sure that your text is not cut off at the margins.
#
# **Do not submit any other files produced by your code.**
#
# Include a link to your colab file in your submission.
#
# Please use Google Colab to complete this assignment. If you want to use Jupyter Notebook, please complete the assignment and upload your Jupyter Notebook file to Google Colab for submission.
# + [markdown] id="LfiFE_WOqIDu"
# ## Colab Link
#
# Include a link to your colab file here
#
# Colab Link: https://colab.research.google.com/drive/1nA0F7VYFxMIpm07UY6Xx2CD7qwOynFLP?usp=sharing
# + [markdown] id="kvTXpH_kqIDy"
# ## Part A. Data Collection [10 pt]
#
# So far, we have worked with data sets that have been collected, cleaned, and curated by machine learning
# researchers and practitioners. Datasets like MNIST and CIFAR are often used as toy examples, both by
# students and by researchers testing new machine learning models.
#
# In the real world, getting a clean data set is never that easy. More than half the work in applying machine
# learning is finding, gathering, cleaning, and formatting your data set.
#
# The purpose of this lab is to help you gain experience gathering your own data set, and understand the
# challenges involved in the data cleaning process.
#
# ### American Sign Language
#
# American Sign Language (ASL) is a complete, complex language that employs signs made by moving the
# hands combined with facial expressions and postures of the body. It is the primary language of many
# North Americans who are deaf and is one of several communication options used by people who are deaf or
# hard-of-hearing.
#
# The hand gestures representing English alphabet are shown below. This lab focuses on classifying a subset
# of these hand gesture images using convolutional neural networks. Specifically, given an image of a hand
# showing one of the letters A-I, we want to detect which letter is being represented.
#
# 
#
#
# ### Generating Data
# We will produce the images required for this lab by ourselves. Each student will collect, clean and submit
# three images each of Americal Sign Language gestures for letters A - I (total of 27 images)
# Steps involved in data collection
#
# 1. Familiarize yourself with American Sign Language gestures for letters from A - I (9 letters).
# 2. Take three pictures at slightly different orientation for each letter gesture using your
# mobile phone.
# - Ensure adequate lighting while you are capturing the images.
# - Use a white wall as your background.
# - Use your right hand to create gestures (for consistency).
# - Keep your right hand fairly apart from your body and any other obstructions.
# - Avoid having shadows on parts of your hand.
# 3. Transfer the images to your laptop for cleaning.
#
# ### Cleaning Data
# To simplify the machine learning the task, we will standardize the training images. We will make sure that
# all our images are of the same size (224 x 224 pixels RGB), and have the hand in the center of the cropped
# regions.
#
# You may use the following applications to crop and resize your images:
#
# **Mac**
# - Use Preview:
# – Holding down CMD + Shift will keep a square aspect ratio while selecting the hand area.
# – Resize to 224x224 pixels.
#
# **Windows 10**
# - Use Photos app to edit and crop the image and keep the aspect ratio a square.
# - Use Paint to resize the image to the final image size of 224x224 pixels.
#
# **Linux**
# - You can use GIMP, imagemagick, or other tools of your choosing.
# You may also use online tools such as http://picresize.com
# All the above steps are illustrative only. You need not follow these steps but following these will ensure that
# you produce a good quality dataset. You will be judged based on the quality of the images alone.
# Please do not edit your photos in any other way. You should not need to change the aspect ratio of your
# image. You also should not digitally remove the background or shadows—instead, take photos with a white
# background and minimal shadows.
#
# ### Accepted Images
# Images will be accepted and graded based on the criteria below
# 1. The final image should be size 224x224 pixels (RGB).
# 2. The file format should be a .jpg file.
# 3. The hand should be approximately centered on the frame.
# 4. The hand should not be obscured or cut off.
# 5. The photos follows the ASL gestures posted earlier.
# 6. The photos were not edited in any other way (e.g. no electronic removal of shadows or background).
#
# ### Submission
# Submit a zip file containing your images. There should be a total of 27 images (3 for each category)
# 1. Individual image file names should follow the convention of student-number_Alphabet_file-number.jpg
# (e.g. 100343434_A_1.jpg)
# 2. Zip all the images together and name it with the following convention: last-name_student-number.zip
# (e.g. last-name_100343434.zip).
# 3. Submit the zipped folder.
# We will be anonymizing and combining the images that everyone submits. We will announce when the
# combined data set will be available for download.
#
# 
# + [markdown] id="bJxMgWGNqID2"
# ## Part B. Building a CNN [50 pt]
#
# For this lab, we are not going to give you any starter code. You will be writing a convolutional neural network
# from scratch. You are welcome to use any code from previous labs, lectures and tutorials. You should also
# write your own code.
#
# You may use the PyTorch documentation freely. You might also find online tutorials helpful. However, all
# code that you submit must be your own.
#
# Make sure that your code is vectorized, and does not contain obvious inefficiencies (for example, unecessary
# for loops, or unnecessary calls to unsqueeze()). Ensure enough comments are included in the code so that
# your TA can understand what you are doing. It is your responsibility to show that you understand what you
# write.
#
# **This is much more challenging and time-consuming than the previous labs.** Make sure that you
# give yourself plenty of time by starting early.
# + [markdown] id="MiDuQaAh56sT"
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# ### 1. Data Loading and Splitting [5 pt]
#
# Download the anonymized data provided on Quercus. To allow you to get a heads start on this project we will provide you with sample data from previous years. Split the data into training, validation, and test sets.
#
# Note: Data splitting is not as trivial in this lab. We want our test set to closely resemble the setting in which
# our model will be used. In particular, our test set should contain hands that are never seen in training!
#
# Explain how you split the data, either by describing what you did, or by showing the code that you used.
# Justify your choice of splitting strategy. How many training, validation, and test images do you have?
#
# For loading the data, you can use plt.imread as in Lab 1, or any other method that you choose. You may find
# torchvision.datasets.ImageFolder helpful. (see https://pytorch.org/docs/stable/torchvision/datasets.html?highlight=image%20folder#torchvision.datasets.ImageFolder
# )
# + id="3FHOjQJCm-qT"
# First we must import all the libraries that we require
import numpy as np
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
from torch.utils.data.sampler import SubsetRandomSampler
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
import os
# + colab={"base_uri": "https://localhost:8080/"} id="WBrH5kBqRLa6" outputId="97e2cf3d-20d0-42b4-cf80-11057bfcdd86"
from google.colab import drive
drive.mount('/content/gdrive')
main_dir = '/content/gdrive/My Drive/Colab Notebooks/lab3'
# + id="_3ELzBLYrXoP"
def load_data(batch_size=64, num_workers=1, small=False):
# Compose allows us to have multiple transformations to occur, where we first ensure our images are 224x224, and then
# convert the images to a tensor in the form (torch.Tensor([3, 224, 224])
transform_it = transforms.Compose([transforms.CenterCrop(224), transforms.ToTensor()])
# Check to see if we want to load the small dataset for over fitting
if small:
small_path = main_dir + '/Lab3a'
small_data = torchvision.datasets.ImageFolder(small_path, transform=transform_it)
return small_data
# Save the paths of each of the different types of data that are located in my drive
train_path = main_dir + '/Gesture_Dataset_Sorted/train_data'
val_path = main_dir + '/Gesture_Dataset_Sorted/val_data'
test_path = main_dir + '/Gesture_Dataset_Sorted/test_data'
# Load all of the data from my google drive
train_data = torchvision.datasets.ImageFolder(train_path, transform=transform_it)
val_data = torchvision.datasets.ImageFolder(val_path, transform=transform_it)
test_data = torchvision.datasets.ImageFolder(test_path, transform=transform_it)
return train_data, val_data, test_data
# + id="nZ_saxXyHTwA" colab={"base_uri": "https://localhost:8080/"} outputId="1c94f98a-0358-4826-ac2d-576d4091ec33"
train_data, val_data, test_data = load_data()
small_data = load_data(small=True)
# TRAINING EXAMPLES
print("Training Examples:", len(train_data))
#VALIDATION EXMAPLES
print("Validation Examples:", len(val_data))
#TEST EXAMPLES
print("Test Examples:", len(test_data))
# SMALL EXAMPLES
print("Small Examples:", len(small_data))
# + id="9MGNM5l9Cid0"
# For the model checkpoints
def get_model_name(name, batch_size, learning_rate, epoch):
""" Generate a name for the model consisting of all the hyperparameter values
Args:
config: Configuration object containing the hyperparameters
Returns:
path: A string with the hyperparameter name and value concatenated
"""
path = "model_{0}_bs{1}_lr{2}_epoch{3}".format(name,
batch_size,
learning_rate,
epoch)
return path
# + [markdown] id="Ek980ebCJrZ7"
# **EXPLAINATION FOR PART 1**
#
# The load_data function essentially loads all of my sample data from the gesture dataset, and returns them as a usable tensor.
#
# The way I have split up my data, was that ~70% of the total data is used for training, which is approximatly 1686 images out of 2431. Then, the rest of the 30% was split evenly into validation and testing data. Where 372 images were used for validation, and 373 images were used for testing. The dataset was slightly uneven, because from the given gesture dataset the letter I had 249 images instead of 272 from A-H.
#
# The reason for the split being 70% training, 15% validation and 15% test data was that in order to ensure that a neural net is being trained with an adequetely, a significant portion of the data must be used for training purposes. Then, we require a similar sized datasets for validation and testing because a smaller validation dataset prevents the net from performing well. Then, in order to ensure we are testing the net properly to simulate real life data, we must also have data that is a proper generalization of the real time data.
#
# + [markdown] id="5VWX4DGY5gQE"
# ### 2. Model Building and Sanity Checking [15 pt]
#
# ### Part (a) Convolutional Network - 5 pt
#
# Build a convolutional neural network model that takes the (224x224 RGB) image as input, and predicts the gesture
# letter. Your model should be a subclass of nn.Module. Explain your choice of neural network architecture: how
# many layers did you choose? What types of layers did you use? Were they fully-connected or convolutional?
# What about other decisions like pooling layers, activation functions, number of channels / hidden units?
# + id="2dtx1z5951fS"
#Convolutional Neural Network Architecture
class GestureClassifier(nn.Module):
def __init__(self):
self.name = "GestureClassifier"
super(GestureClassifier, self).__init__()
self.conv1 = nn.Conv2d(3, 5, 5) #in_channels, out_chanels, kernel_size
self.pool = nn.MaxPool2d(2, 2) #kernel_size, stride
self.conv2 = nn.Conv2d(5, 10, 5) #in_channels, out_chanels, kernel_size
self.fc1 = nn.Linear(10*53*53, 30)
self.fc2 = nn.Linear(30, 9)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 10*53*53)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
# + [markdown] id="uDQkBksL01U7"
# **EXPLAINATION FOR PART 2a**
#
# I chose to build a simple CNN architecture, where there are two convolution layers, two pooling layers, two fully connected layers, and one output layer.
#
# The follwing will explain the steps in how each layer of my CNN works.
#
# * Conv2d layer 1 - it takes in 1 channel, and outputs 5 channels using a 5x5 size kernel
#
# **Our image here becomes (5x220x220)**
#
# * max pooling layer 1 - it filters out the image by only taking the max value in every 2x2 within the image, reducing image size.
#
# **Our image here becomes (5x110x10)**
#
# * Conv2d layer 2 - which takes in 5 channels, since output of first convolution layer was 5, and outputs 10 channels, with the same kernel size of 5
#
# **Our image here becomes (10x106x106)**
#
# * max pooling layer 2 - it once again filters out the images, by taking max value.
#
# **Our image here becomes (10x53x53)**
#
#
# * Fully Connected Layer 1 - image goes into the fully connected layer
# * Fully Connected Layer 2 - which then goes through another layer and outputs 9
# * Output layer - Then, the image is output
#
#
#
#
#
#
#
#
# + [markdown] id="XeGvelvb515e"
# ### Part (b) Training Code - 5 pt
#
# Write code that trains your neural network given some training data. Your training code should make it easy
# to tweak the usual hyperparameters, like batch size, learning rate, and the model object itself. Make sure
# that you are checkpointing your models from time to time (the frequency is up to you). Explain your choice
# of loss function and optimizer.
# + id="hWUUKPCGjpF-"
# Obtain the accuracy of the model on the dataset
def get_accuracy(model, loader):
data = loader
correct = 0
total = 0
for imgs, labels in data:
#############################################
#To Enable GPU Usage
if use_cuda and torch.cuda.is_available():
imgs = imgs.cuda()
labels = labels.cuda()
#############################################
output = model(imgs)
#select index with maximum prediction score
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(labels.view_as(pred)).sum().item()
# Debug
#print(labels)
#print(pred)
total += imgs.shape[0]
return correct / total
# + id="sgcUiUv88Q2A"
def train(model, batch_size=64, learning_rate=0.001, num_epochs=20):
torch.manual_seed(1000)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=learning_rate)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, shuffle=True)
val_loader = torch.utils.data.DataLoader(val_data, batch_size=batch_size, shuffle=True)
iters, losses, train_acc, val_acc = [], [], [], []
# training
n = 0 # the number of iterations
for epoch in range(num_epochs):
for imgs, labels in iter(train_loader):
#############################################
#To Enable GPU Usage
if use_cuda and torch.cuda.is_available():
#print("GPU is Available")
imgs = imgs.cuda()
labels = labels.cuda()
#############################################
out = model(imgs) # forward pass
loss = criterion(out, labels) # compute the total loss
loss.backward() # backward pass (compute parameter updates)
optimizer.step() # make the updates for each parameter
optimizer.zero_grad() # a clean up step for PyTorch
# save the current training information
iters.append(n)
losses.append(float(loss)/batch_size) # compute *average* loss
train_acc.append(get_accuracy(model, train_loader)) # compute training accuracy
val_acc.append(get_accuracy(model, val_loader)) # compute validation accuracy
n += 1
# Print the accuracies of validation and training for each epoch to observe how it changes over time
#print("epoch = {}, Training Accuracy = {}, Validation Accuracy = {}".format(epoch, train_acc[epoch], val_acc[epoch]))
print("epoch number: ", epoch+1, "Training accuracy: ",train_acc[epoch], "Validation accuracy: ", val_acc[epoch])
# Save the current model (checkpoint) to a file
model_path = get_model_name(model.name, batch_size, learning_rate, epoch)
torch.save(model.state_dict(), model_path)
# plotting
plt.title("Training Curve")
plt.plot(iters, losses, label="Train")
plt.xlabel("Iterations")
plt.ylabel("Loss")
plt.show()
plt.title("Training Curve")
plt.plot(iters, train_acc, label="Train")
plt.plot(iters, val_acc, label="Validation")
plt.xlabel("Iterations")
plt.ylabel("Training Accuracy")
plt.legend(loc='best')
plt.show()
print("Final Training Accuracy: {}".format(train_acc[-1]))
print("Final Validation Accuracy: {}".format(val_acc[-1]))
# + [markdown] id="QlhYGJPDOp22"
# **EXPLAINATION FOR PART 2b**
#
# I opted to use the Cross Entropy Loss for my loss function, because it tends to much better for multi classification models. I also opted for the Adam optimizer because it is much faster than the SDG. Additionally, the Adam optimizer has an adaptive learning rate (starts with bigger steps, which gradually becomes smaller), this enables it be a better optimizer on average than others.
# + [markdown] id="bk1RNgAj54rZ"
# ### Part (c) “Overfit” to a Small Dataset - 5 pt
#
# One way to sanity check our neural network model and training code is to check whether the model is capable
# of “overfitting” or “memorizing” a small dataset. A properly constructed CNN with correct training code
# should be able to memorize the answers to a small number of images quickly.
#
# Construct a small dataset (e.g. just the images that you have collected). Then show that your model and
# training code is capable of memorizing the labels of this small data set.
#
# With a large batch size (e.g. the entire small dataset) and learning rate that is not too high, You should be
# able to obtain a 100% training accuracy on that small dataset relatively quickly (within 200 iterations).
# + id="alD8YRgTleYJ"
def small_net(model, batch_size=27, learning_rate=0.001, num_epochs=1):
torch.manual_seed(1000)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=learning_rate)
iters, losses, train_acc, val_acc = [], [], [], []
small_loader = torch.utils.data.DataLoader(small_data, batch_size=batch_size, shuffle=True)
# training
n = 0 # the number of iterations
for epoch in range(num_epochs):
for imgs, labels in iter(small_loader):
#############################################
#To Enable GPU Usage
if use_cuda and torch.cuda.is_available():
#print("GPU is Available")
imgs = imgs.cuda()
labels = labels.cuda()
#############################################
out = model(imgs) # forward pass
loss = criterion(out, labels) # compute the total loss
loss.backward() # backward pass (compute parameter updates)
optimizer.step() # make the updates for each parameter
optimizer.zero_grad() # a clean up step for PyTorch
# save the current training information
iters.append(n)
losses.append(float(loss)/batch_size) # compute *average* loss
train_acc.append(get_accuracy(model, small_loader)) # compute training accuracy
n += 1
# Print the accuracies of validation and training for each epoch to observe how it changes over time
#print("epoch = {}, Training Accuracy = {}, Validation Accuracy = {}".format(epoch, train_acc[epoch], val_acc[epoch]))
print("epoch number: ", epoch+1, "Training accuracy: ",train_acc[epoch])
# Save the current model (checkpoint) to a file
model_path = get_model_name(model.name, batch_size, learning_rate, epoch)
torch.save(model.state_dict(), model_path)
# plotting
plt.title("Training Curve")
plt.plot(iters, losses, label="Train")
plt.xlabel("Iterations")
plt.ylabel("Loss")
plt.show()
plt.title("Training Curve")
plt.plot(iters, train_acc, label="Train")
#plt.plot(iters, val_acc, label="Validation")
plt.xlabel("Iterations")
plt.ylabel("Training Accuracy")
plt.legend(loc='best')
plt.show()
print("Final Training Accuracy: {}".format(train_acc[-1]))
#print("Final Validation Accuracy: {}".format(val_acc[-1]))
# + colab={"base_uri": "https://localhost:8080/"} id="lXYRBhQO6d3u" outputId="5f4526e3-133b-4626-d2e3-8e60cf268561"
# Use GPU
use_cuda = True
small_model= GestureClassifier()
if use_cuda and torch.cuda.is_available():
small_model.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="HHKE7F7-aR5e" outputId="c5046bf3-0c4f-4d67-acfc-e61457ed496a"
small_net(small_model, batch_size=27, learning_rate=0.001, num_epochs=200)
# + [markdown] id="nvDLw-Vz6eVS"
# ### 3. Hyperparameter Search [10 pt]
#
# ### Part (a) - 1 pt
#
# List 3 hyperparameters that you think are most worth tuning. Choose at least one hyperparameter related to
# the model architecture.
# + [markdown] id="tQ3A1854CPYF"
# **EXPLAINATION FOR 3a**
#
# The hyperparameters that are worth tuning are the *batch size*, *learning rate*, and *number of convolution layers*
# + [markdown] id="zeD6EzPB6kSW"
# ### Part (b) - 5 pt
#
# Tune the hyperparameters you listed in Part (a), trying as many values as you need to until you feel satisfied
# that you are getting a good model. Plot the training curve of at least 4 different hyperparameter settings.
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="PolG-_OeEBmp" outputId="f097131d-2e6b-4297-c4b6-b2134fd34766"
# Use GPU
use_cuda = True
ges_model1= GestureClassifier()
if use_cuda and torch.cuda.is_available():
ges_model1.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
# Setting 1 - Using Batch Size 64, learning rate = 0.001, 2 CNN Layers
train(ges_model1, batch_size=64, learning_rate=0.001,num_epochs=15)
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="YtC-zAfvXATP" outputId="eae0d412-2f92-4b97-b296-0d8e8d875596"
# Use GPU
use_cuda = True
ges_model2= GestureClassifier()
if use_cuda and torch.cuda.is_available():
ges_model2.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
# Setting 2 - Using Batch Size 128, lr =0.001 , 2 CNN Layers
train(ges_model2, batch_size=128, learning_rate=0.001, num_epochs=15)
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="Wth9eKYPRRiE" outputId="e4b26379-b040-4b5d-ee27-9a85a9e29513"
# Setting 3 - Using Batch Size 64, lr =0.003 , 2 CNN Layers
# Use GPU
use_cuda = True
ges_model3= GestureClassifier()
if use_cuda and torch.cuda.is_available():
ges_model3.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
train(ges_model3, batch_size=64, learning_rate=0.003, num_epochs=15)
# + id="Jvi2C3IZ2VqQ"
#Convolutional Neural Network Architecture for a 3 layer CNN
class GestureClassifierThree(nn.Module):
def __init__(self):
self.name = "GestureClassifierThree"
super(GestureClassifierThree, self).__init__()
self.conv1 = nn.Conv2d(3, 5, 5) #in_channels, out_chanels, kernel_size
self.pool = nn.MaxPool2d(2, 2) #kernel_size, stride
self.conv2 = nn.Conv2d(5, 10, 5) #in_channels, out_chanels, kernel_size
self.conv3 = nn.Conv2d(10, 20, 4) #in_channels, out_chanels, kernel_size
self.fc1 = nn.Linear(20*25*25, 30)
self.fc2 = nn.Linear(30, 9)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
x = x.view(-1, 20*25*25)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="jD5ZbDXuippW" outputId="239af5e0-3ebf-4f48-baa1-55ef420c1540"
# Setting 4 - Using Batch Size 64, lr =0.003 , 3 CNN Layers
# Use GPU
use_cuda = True
ges_model4= GestureClassifierThree()
if use_cuda and torch.cuda.is_available():
ges_model4.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
train(ges_model4, batch_size=64, learning_rate=0.003, num_epochs=15)
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="H7K93OeBLeD-" outputId="93183e37-467e-4382-9ba4-037f10681a36"
# Setting 5 - Using Batch Size 64, lr =0.001 , 3 CNN Layers
# Use GPU
use_cuda = True
ges_model5= GestureClassifierThree()
if use_cuda and torch.cuda.is_available():
ges_model5.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
train(ges_model5, batch_size=64, learning_rate=0.001, num_epochs=15)
# + [markdown] id="H93iN5_l60BO"
# ### Part (c) - 2 pt
# Choose the best model out of all the ones that you have trained. Justify your choice.
# + [markdown] id="cwdWKqjlC1DE"
# **EXPLAINATION FOR PART 3c**
#
# The best model (model 2) that I have trained was where the hyperparameters were:
#
# * Batch Size = 128
# * Learning Rate = 0.001
# * CNN Architecture Layers = 2 layers
#
# I chose model 2 to be my best model. However, there was another model that stood out to me as well, model 5. The difference between the two models were the number of CNN layers that were used and the batch size. The one with three CNN layers (model 5) gave me a higher training accuracy at 92%. However both models gave me the validation accuracy at 72%, whereas the model that I finally opted with (model 2) with 2 CNN layers had a slightly lower training accuracy at 82%. However, it can be seen that the loss function for both models tends to decrease significantly with less fluculations relative to the most other models (model 3 was just overtrained, so loss was lowest). It is better to choose the model with a higher validation accuracy, because this is an indicator of the model being able to generalize for real time data, however this was not what was my deciding factor was. I made a decision based on the loss curve, and the curve that was closer to the training accuracy, which means that the difference between the training accuracy and validation accuracy should be smaller (smaller error). It is much better to pick a model with lower training accuracy than validation accuracy, to prevent overfitting the data. This allows for the model to become more confident in its predictions, which can also be observed with the loss curve decreasing over more epochs, and the numbers became smallest relative to other models. Therefore, the model I chose to be my best model was model 2, with less CNN layers since it gave me the highest validation accuracy with an exponentially decreasing loss curve, indicating that a smaller learning rate and a slightly larger batch size improved the model.
#
#
# + [markdown] id="QzNA5oup67JO"
# ### Part (d) - 2 pt
# Report the test accuracy of your best model. You should only do this step once and prior to this step you should have only used the training and validation data.
# + id="2eJ7AbVl6-ax" colab={"base_uri": "https://localhost:8080/"} outputId="8879a4bb-6986-4406-a4af-4c8124b54618"
# Test accuracy of my model
# Get the test loader first
test_loader = torch.utils.data.DataLoader(test_data, batch_size=128, shuffle=True)
test_accuracy = get_accuracy(ges_model2, test_loader)
print("Test Accuracy for Model: ", test_accuracy)
# + [markdown] id="Wrem-iXV6_Bz"
# ### 4. Transfer Learning [15 pt]
# For many image classification tasks, it is generally not a good idea to train a very large deep neural network
# model from scratch due to the enormous compute requirements and lack of sufficient amounts of training
# data.
#
# One of the better options is to try using an existing model that performs a similar task to the one you need
# to solve. This method of utilizing a pre-trained network for other similar tasks is broadly termed **Transfer
# Learning**. In this assignment, we will use Transfer Learning to extract features from the hand gesture
# images. Then, train a smaller network to use these features as input and classify the hand gestures.
#
# As you have learned from the CNN lecture, convolution layers extract various features from the images which
# get utilized by the fully connected layers for correct classification. AlexNet architecture played a pivotal
# role in establishing Deep Neural Nets as a go-to tool for image classification problems and we will use an
# ImageNet pre-trained AlexNet model to extract features in this assignment.
# + [markdown] id="rWdQJz4Q7O2F"
# ### Part (a) - 5 pt
# Here is the code to load the AlexNet network, with pretrained weights. When you first run the code, PyTorch
# will download the pretrained weights from the internet.
# + id="BJKcTW9C7TZk"
import torchvision.models
alexnet = torchvision.models.alexnet(pretrained=True)
# + [markdown] id="NQ0GZYaP7VAR"
# The alexnet model is split up into two components: *alexnet.features* and *alexnet.classifier*. The
# first neural network component, *alexnet.features*, is used to compute convolutional features, which are
# taken as input in *alexnet.classifier*.
#
# The neural network alexnet.features expects an image tensor of shape Nx3x224x224 as input and it will
# output a tensor of shape Nx256x6x6 . (N = batch size).
#
# Compute the AlexNet features for each of your training, validation, and test data. Here is an example code
# snippet showing how you can compute the AlexNet features for some images (your actual code might be
# different):
# + id="oX7SjVdB7XAE"
# img = ... a PyTorch tensor with shape [N,3,224,224] containing hand images ...
#features = alexnet.features(img)
# + [markdown] id="DYcjHg_A7cCM"
# **Save the computed features**. You will be using these features as input to your neural network in Part
# (b), and you do not want to re-compute the features every time. Instead, run *alexnet.features* once for
# each image, and save the result.
# + id="TBo1BpL373LX"
# Created a new folder where it is called AlexNet_Features
new_main_dir = main_dir + '/Alex_Features'
# Load all of the data such as training, validation and test
train_loader = torch.utils.data.DataLoader(train_data, batch_size=1, num_workers=1, shuffle=True)
val_loader = torch.utils.data.DataLoader(val_data, batch_size=1, num_workers=1, shuffle=True)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=1, num_workers=1, shuffle=True)
small_loader = torch.utils.data.DataLoader(small_data, batch_size=1, num_workers=1, shuffle=True)
# Create a function to save all the features into the folders
def load_features(loader, new_main_dir, folder):
classes = ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I']
n = 0
for img, label in loader:
# Get the features of each image and transform to a tensor
feature = alexnet.features(img)
feature_tensor = torch.from_numpy(feature.detach().numpy())
# Save those features into folder
feature_dir = new_main_dir + '/' + str(folder) + '/' + str(classes[label])
if not os.path.isdir(feature_dir):
os.mkdir(feature_dir)
torch.save(feature_tensor.squeeze(0), feature_dir + '/' + str(n) + '.tensor')
n+=1
print("Features done for ", folder)
# + colab={"base_uri": "https://localhost:8080/"} id="575szVuQgBGe" outputId="f292597d-eca9-45a3-d787-aed0243a7d42"
load_features(train_loader, new_main_dir, 'Train')
load_features(val_loader,new_main_dir,'Validation')
load_features(test_loader,new_main_dir,'Test')
load_features(small_loader,new_main_dir,'Small')
# + [markdown] id="OFWvvhFN73qY"
# ### Part (b) - 3 pt
# Build a convolutional neural network model that takes as input these AlexNet features, and makes a
# prediction. Your model should be a subclass of nn.Module.
#
# Explain your choice of neural network architecture: how many layers did you choose? What types of layers
# did you use: fully-connected or convolutional? What about other decisions like pooling layers, activation
# functions, number of channels / hidden units in each layer?
#
# Here is an example of how your model may be called:
# + id="oVTuHUeV78-U"
# features = ... load precomputed alexnet.features(img) ...
#output = model(features)
#prob = F.softmax(output)
# + id="OEavy7zhnd5S"
#Convolutional Neural Network Architecture for Alex
class AlexClassifier(nn.Module):
def __init__(self):
self.name = "AlexClassifier"
super(AlexClassifier, self).__init__()
self.conv1 = nn.Conv2d(256, 100, 2) #in_channels, out_chanels, kernel_size
self.conv2 = nn.Conv2d(100, 10, 2) #in_channels, out_chanels, kernel_size
self.fc1 = nn.Linear(10*4*4, 30)
self.fc2 = nn.Linear(30, 9)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = x.view(-1, 10*4*4)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
# + [markdown] id="3uW-AS3pqIKC"
# **EXPLAINATION FOR 4b**
#
# I chose to have a simple CNN where I disregarded max pooling since the AlexNet features reduced the tensor to 256x6x6, so there was no need to make the image smaller. I kept two convolution layers because I found from part 3 that it generally gave best results, but with AlexNet features there was no need to have more than two conv2d filters to extract the features. I had two fully connected layers as well, with 30 hidden units. I chose to have the output channels to be at 100 for conv2d layer 1 and 10 for conv2d layer 2, with kernel size to be 2 since the image tensor was 256x6x6. AlexNet should be able to enhance the results for a simple CNN, since it was working relatively well in part 3.
# + [markdown] id="wVAGuURu7-9q"
# ### Part (c) - 5 pt
# Train your new network, including any hyperparameter tuning. Plot and submit the training curve of your
# best model only.
#
# Note: Depending on how you are caching (saving) your AlexNet features, PyTorch might still be tracking
# updates to the **AlexNet weights**, which we are not tuning. One workaround is to convert your AlexNet
# feature tensor into a numpy array, and then back into a PyTorch tensor.
# + id="JCmiH11x7-q1"
tensor = torch.from_numpy(tensor.detach().numpy())
# + id="J6xVdpX1vN0a"
# Get the path of the alexnet features
train_alex_path = main_dir + '/Alex_Features/Train'
val_alex_path = main_dir + '/Alex_Features/Validation'
test_alex_path = main_dir + '/Alex_Features/Test'
small_alex_path = main_dir + '/Alex_Features/Small'
train_alex_data = torchvision.datasets.DatasetFolder(train_alex_path, loader=torch.load, extensions=('.tensor'))
val_alex_data = torchvision.datasets.DatasetFolder(val_alex_path, loader=torch.load, extensions=('.tensor'))
test_alex_data = torchvision.datasets.DatasetFolder(test_alex_path, loader=torch.load, extensions=('.tensor'))
small_alex_data = torchvision.datasets.DatasetFolder(small_alex_path, loader=torch.load, extensions=('.tensor'))
# + id="zzH3yA1stCzQ"
def trainAlex(model, batch_size=64, learning_rate=0.001, num_epochs=20):
torch.manual_seed(1000)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=learning_rate)
train_loader = torch.utils.data.DataLoader(train_alex_data, batch_size=batch_size, shuffle=True)
val_loader = torch.utils.data.DataLoader(val_alex_data, batch_size=batch_size, shuffle=True)
iters, losses, train_acc, val_acc = [], [], [], []
# training
n = 0 # the number of iterations
for epoch in range(num_epochs):
for imgs, labels in iter(train_loader):
#############################################
#To Enable GPU Usage
if use_cuda and torch.cuda.is_available():
#print("GPU is Available")
imgs = imgs.cuda()
labels = labels.cuda()
#############################################
out = model(imgs) # forward pass
loss = criterion(out, labels) # compute the total loss
loss.backward() # backward pass (compute parameter updates)
optimizer.step() # make the updates for each parameter
optimizer.zero_grad() # a clean up step for PyTorch
# save the current training information
iters.append(n)
losses.append(float(loss)/batch_size) # compute *average* loss
train_acc.append(get_accuracy(model, train_loader)) # compute training accuracy
val_acc.append(get_accuracy(model, val_loader)) # compute validation accuracy
print("epoch number: ", epoch+1, "Training accuracy: ",train_acc[epoch], "Validation accuracy: ", val_acc[epoch])
n += 1
# Print the accuracies of validation and training for each epoch to observe how it changes over time
#print("epoch = {}, Training Accuracy = {}, Validation Accuracy = {}".format(epoch, train_acc[epoch], val_acc[epoch]))
# Save to a checkpoint
model_path = "model_{0}_bs{1}_lr{2}_epoch{3}".format(model.name,batch_size,learning_rate,epoch)
torch.save(model.state_dict(), model_path)
# plotting
plt.title("Training Curve")
plt.plot(iters, losses, label="Train")
plt.xlabel("Iterations")
plt.ylabel("Loss")
plt.show()
plt.title("Training Curve")
plt.plot(iters, train_acc, label="Train")
plt.plot(iters, val_acc, label="Validation")
plt.xlabel("Iterations")
plt.ylabel("Training Accuracy")
plt.legend(loc='best')
plt.show()
print("Final Training Accuracy: {}".format(train_acc[-1]))
print("Final Validation Accuracy: {}".format(val_acc[-1]))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="OwlmW6FfwmcH" outputId="917626f4-d6aa-458a-83db-92bd3a240b8a"
# Use GPU
use_cuda = True
alex_model= AlexClassifier()
if use_cuda and torch.cuda.is_available():
alex_model.cuda()
print('CUDA is available! Training on GPU ...')
else:
print('CUDA is not available. Training on CPU ...')
# Train using the features
trainAlex(alex_model, batch_size=64, learning_rate=0.001, num_epochs=15)
# + [markdown] id="hQ2tvqJ68Mqb"
# ### Part (d) - 2 pt
# Report the test accuracy of your best model. How does the test accuracy compare to Part 3(d) without transfer learning?
# + [markdown] id="gWgpEiAXPYAX"
# **EXPLAINATION OF PART 4d**
#
# The test accuracy is much higher with transfer learning, where it is at 91%. Transfer learning is generally used for when we ourselves lack a significant amount of data. When we applied transfer learning, the model did not tend to overfit the training data. It also allowed the CNN to be much more enhanced with it already pretrained, with a high training and validation accuracy where they are relatively similar. This means, the data is able to generalize much better for real time data compared to my model in part 3d, where test accuracy was only 72%. This is due to the lack of training compared to the AlexNet model, so my model in part 3 was unable to generalize that well, with a greater error margin in its predictions. This is evident in the loss curve of the training as well, as the loss curve gets significantly smaller with less fluctuations over the number of iterations, unlike my loss curves in part 3d.
# + id="yCp_kFSg8Q2T" colab={"base_uri": "https://localhost:8080/"} outputId="0f426f47-61bf-41e7-9bcd-bd4f8f440541"
# Test accuracy of my model
# Get the test loader first
test_loader = torch.utils.data.DataLoader(test_alex_data, batch_size=64, shuffle=True)
test_accuracy = get_accuracy(alex_model, test_loader)
print("Test Accuracy for Model: ", test_accuracy)
# + [markdown] id="vPBcaDuNcsA8"
# ### 5. Additional Testing [5 pt]
# As a final step in testing we will be revisiting the sample images that you had collected and submitted at the start of this lab. These sample images should be untouched and will be used to demonstrate how well your model works at identifying your hand guestures.
#
# Using the best transfer learning model developed in Part 4. Report the test accuracy on your sample images and how it compares to the test accuracy obtained in Part 4(d)? How well did your model do for the different hand guestures? Provide an explanation for why you think your model performed the way it did?
# + [markdown] id="68QRBn7DRz69"
# **EXPLAINATION FOR PART 5**
#
# The test accuracy for my own submission dataset is similar to part 4d, where it was 91.6% and for my submission, it was 92.5%. There was a slight improvement when it came to my own dataset because of several reasons.
#
# When comparing my submission vs the test dataset, all of the images in my submission was by just me, which means there was less variety when it came to the images. There was a not as much of data augmentation as the testing dataset, because the lighting was similar, and it was just my hand which means the model was able to predict my own set of images slightly better due to a dataset images being similar to one another.
#
# Apart from the lack of variation, my dataset was also smaller compared to the test dataset, which is not the best representation of real time data as it may have just coincidentally predicted my submission data well, but may not predict it well for other data, where there is more variety within the images.
#
#
# + id="mquOqqZ7csA8" colab={"base_uri": "https://localhost:8080/"} outputId="55a67b0b-27ac-40c3-9baf-1f23c0607ed5"
# Get the small loader first
small_loader = torch.utils.data.DataLoader(small_alex_data, batch_size=64, shuffle=True)
small_accuracy = get_accuracy(alex_model, small_loader)
print("Test Accuracy for Model: ", small_accuracy)
| Primary Models/transfer_learning/lab3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dataset size and model complexity
#
# So far we've looked at the relationship between training dataset size and model performance. Now we'll add another dimension to the picture, and try to get a qualitative sense of how our training and validation curves vary as a function of model complexity—which in this case we'll operationalize as the number of predictors our linear regression estimator gets to use.
#
# To save ourselves a lot of code, we'll use a `plot_learning_curves` helper function I've written that wraps the `learning_curve` utility we used to produce the curves in the previous plot. It'll enale us to easily generate plots with multiple panels, where each panel shows the learning curve when plotted for a different dataset and/or estimator.
#
# Let's start by asking how well we can predict age given our three different predictor sets—the domains, facets, and items, respectively. These sets contain 5, 30, and 300 features, respectively. We'll once again evaluate performance for sample sizes ranging from 100 to 50,000. For the time being, we'll stick with ordinary least-squares regression as our estimator.
# +
# the aforementioned plotting helper
from support import plot_learning_curves
# we'll compare performance for three different sets of predictors:
# the 5 domains, 30 facets, and 300 items. note the *X_sets
# notation; this is an advanced iterable unpacking trick that
# only works in Python 3. it lets the variable preceded by the
# star "swallow" any variables that aren't explicitly assigned.
*X_sets, age = get_features(data, 'domains', 'facets', 'items', 'AGE')
# training sizes remain the same in our previous examples
train_sizes = [100, 200, 500, 1000, 2000, 5000, 10000, 20000, 50000]
# titles for each of the plot panels
labels = ['5 domains', '30 facets', '300 items']
# we pass plot_learning_curves an estimator, the list of feature
# #sets, the outcome variable, and the train_sizes and labels
plot_learning_curves(LinearRegression(), X_sets, age, train_sizes, labels=labels)
# -
# There are a couple of new points to note here:
#
# 1. Our ability to predict age varies dramatically depending on which features we use. Looking at the terminal point of each test curve (representing the K-fold cross-validated performance estimate at the largest sample size), we see that we do much better with 300 items than with 30 facets, and much better with the 30 facets than with just the 5 domains.
#
# 2. When sample size is small, we actually do a better job predictively using a smaller set of features. For example, at n = 500, we can explain about 20% of the variance in the test set using 30 facets, whereas our 300-item model is basically useless. By contrast, if we paid attention solely to the (overfitted) training set estimates, we would be misled into thinking that the 300-item model performs much better than the 30-facet model (~80% of the variance vs. 30%).
#
# 3. The point at which the training and test curves converge shifts systematically to the right with increasing model complexity.
#
# Points (2) and (3) illustrate a general relationship between model complexity and dataset size: the more complex a model, the greater its capacity to learn from the data, but the more data we need in order to avoid overfitting. This tradeoff is unavoidable, and means that we need to think carefully (and conduct validation analyses like the one above!) when constructing our model.
| content/courses/ml_intro/12_validation/04_validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1XtnSll862HF"
# # Malaria Cell Classification using CNN
#
# The dataset comprises of segmented cells from the thin blood smear slide images from the Malaria Screener research activity conducted by Lister Hill National Center for Biomedical Communications (LHNCBC), part of National Library of Medicine (NLM). The dataset has been manually annotated by experts into two categories:
# - Parasitic
# - Uninfected
#
# The dataset can be found in the URL: https://lhncbc.nlm.nih.gov/LHC-publications/pubs/MalariaDatasets.html
#
# We will try to use a Dense Convolution Neural Network algorithm to classify the images into parasitized or uninfected. \
# The original dataset contains 27,558 images evenly balanced in two categories. However, due to high computation resource unavailability we will randomly sample some 2000 images from each class for training and test. Avid researchers with high computational resources are encouraged to perform training on the complete dataset.
# + id="gYgQt0rHCpdu" executionInfo={"status": "ok", "timestamp": 1635977528362, "user_tz": 240, "elapsed": 2536, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Import the required libararies
import numpy as np
np.random.seed(365) # set a random seed for replication
import matplotlib.pyplot as plt
import os
import shutil
import glob
import random
import cv2
from PIL import Image
from tensorflow import keras
os.environ['KERAS_BACKEND'] = 'tensorflow'
from sklearn.model_selection import train_test_split
from tensorflow.keras.utils import to_categorical
# + colab={"base_uri": "https://localhost:8080/"} id="VxH1W8hpL5EX" executionInfo={"status": "ok", "timestamp": 1635977528365, "user_tz": 240, "elapsed": 33, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X<KEY>mCUg=s64", "userId": "04140676807795382520"}} outputId="cad3b983-55b5-45fa-8f17-0bc4242e647a"
# cd /content/drive/MyDrive/Colab Notebooks/Malaria
# + [markdown] id="zaGRetQ99B-F"
# ### Random Sampling of Images
# + id="BhvR6iQ5LGGR" executionInfo={"status": "ok", "timestamp": 1635978358523, "user_tz": 240, "elapsed": 830181, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Sample some records for training
num_img = 2000
sampled_parasitized = random.sample(glob.glob("./cell_images/Parasitized/*.png"), num_img)
for f in enumerate(sampled_parasitized, 1):
dest = os.path.join(os.getcwd() + "/data/Parasitized/")
if not os.path.exists(dest):
os.makedirs(dest)
shutil.copy(f[1], dest)
sampled_parasitized = random.sample(glob.glob("./cell_images/Uninfected/*.png"), num_img)
for f in enumerate(sampled_parasitized, 1):
dest = os.path.join(os.getcwd() + "/data/Uninfected/")
if not os.path.exists(dest):
os.makedirs(dest)
shutil.copy(f[1], dest)
# + colab={"base_uri": "https://localhost:8080/"} id="uoa6wrpYulQW" executionInfo={"status": "ok", "timestamp": 1635978358524, "user_tz": 240, "elapsed": 25, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}} outputId="30c87b3f-4648-49e4-f96a-0155630cac05"
print("Parasitized Data: ", len(glob.glob("./data/Parasitized/*.png")))
print("Uninfected Data: ", len(glob.glob("./data/Uninfected/*.png")))
# + id="d8qOIj2wL8EH" executionInfo={"status": "ok", "timestamp": 1635978358526, "user_tz": 240, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Iterate through all images in Parasitized folder, resize to 64 x 64
# Then save as numpy array with name 'dataset'
# Place holders to define add labels. We will add 0 to all parasitized images and 1 to uninfected.
image_directory = 'data/'
SIZE = 64
dataset = []
label = []
# + id="nxL-LjWKu1hF" executionInfo={"status": "ok", "timestamp": 1635978366944, "user_tz": 240, "elapsed": 8429, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Iterate through all images in the sampled Parasitized folder
# Since the images are a various sized, resize them all to 64 x 64
# Then save into the same numpy array 'dataset' but with label 0
parasitized_images = os.listdir(image_directory + 'Parasitized/')
for i, image_name in enumerate(parasitized_images):
if (image_name.split('.')[1] == 'png'):
image = cv2.imread(image_directory + 'Parasitized/' + image_name)
image = Image.fromarray(image, 'RGB')
image = image.resize((SIZE, SIZE))
dataset.append(np.array(image))
label.append(0)
# + id="F94GTSiFu4mk" executionInfo={"status": "ok", "timestamp": 1635978375114, "user_tz": 240, "elapsed": 8203, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Iterate through all images in the sampled Uninfected folder
# Since the images are a various sized, resize them all to 64 x 64
# Then save into the same numpy array 'dataset' but with label 1
uninfected_images = os.listdir(image_directory + 'Uninfected/')
for i, image_name in enumerate(uninfected_images):
if (image_name.split('.')[1] == 'png'):
image = cv2.imread(image_directory + 'Uninfected/' + image_name)
image = Image.fromarray(image, 'RGB')
image = image.resize((SIZE, SIZE))
dataset.append(np.array(image))
label.append(1)
# + [markdown] id="XxKftlxpvkIE"
# ## Model
# + id="a8wG-d2wvgk9" executionInfo={"status": "ok", "timestamp": 1635978375114, "user_tz": 240, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Input Images change to (SIZE, SIZE, 3)
INPUT_SHAPE = (SIZE, SIZE, 3)
inp = keras.layers.Input(shape=INPUT_SHAPE)
# + colab={"base_uri": "https://localhost:8080/"} id="mnXMtd8BvtjZ" executionInfo={"status": "ok", "timestamp": 1635978375822, "user_tz": 240, "elapsed": 725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}} outputId="3caf590b-5ddb-46a1-f904-446f0988f44f"
# Create Model
# Convolutional Neural Network
conv1 = keras.layers.Conv2D(32, kernel_size=(3, 3),
activation='relu', padding='same')(inp)
pool1 = keras.layers.MaxPooling2D(pool_size=(2, 2))(conv1)
norm1 = keras.layers.BatchNormalization(axis = -1)(pool1)
drop1 = keras.layers.Dropout(rate=0.2)(norm1)
conv2 = keras.layers.Conv2D(32, kernel_size=(3, 3),
activation='relu', padding='same')(drop1)
pool2 = keras.layers.MaxPooling2D(pool_size=(2, 2))(conv2)
norm2 = keras.layers.BatchNormalization(axis = -1)(pool2)
drop2 = keras.layers.Dropout(rate=0.2)(norm2)
# Flatten the matrix to get it ready for dense.
flat = keras.layers.Flatten()(drop2)
# Dense Fully Connected Network
hidden1 = keras.layers.Dense(512, activation='relu')(flat)
norm3 = keras.layers.BatchNormalization(axis = -1)(hidden1)
drop3 = keras.layers.Dropout(rate=0.2)(norm3)
hidden2 = keras.layers.Dense(256, activation='relu')(drop3)
norm4 = keras.layers.BatchNormalization(axis = -1)(hidden2)
drop4 = keras.layers.Dropout(rate=0.2)(norm4)
out = keras.layers.Dense(2, activation='sigmoid')(drop4) #units=1 gives error
model = keras.Model(inputs=inp, outputs=out)
model.summary()
# + id="Psn8TjWWv0o1" executionInfo={"status": "ok", "timestamp": 1635978375822, "user_tz": 240, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Compile the model with Adam optimizer and categorical crossentropy loss
# https://www.machinecurve.com/index.php/2019/10/22/how-to-use-binary-categorical-crossentropy-with-keras/
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# + id="QlHtuHKxv9hv" executionInfo={"status": "ok", "timestamp": 1635978375823, "user_tz": 240, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Split the data into training and test with 80-20 ratio
X_train, X_test, y_train, y_test = train_test_split(dataset, to_categorical(np.array(label)), test_size = 0.20, random_state = 0)
# + id="kMVMLwtSxT4c" executionInfo={"status": "ok", "timestamp": 1635978375823, "user_tz": 240, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
# Define Callbacks to save the best model and not just the last one
if not os.path.exists('./models'):
os.makedirs('models/')
model_path = './models/malaria.h5'
callbacks = keras.callbacks.ModelCheckpoint(filepath=model_path, mode='max', monitor='val_accuracy', verbose=2, save_best_only=True)
# + [markdown] id="bglSrSKt_Ha2"
# ### Training
# + colab={"base_uri": "https://localhost:8080/"} id="9tlkB7j9wED6" executionInfo={"status": "ok", "timestamp": 1635978483270, "user_tz": 240, "elapsed": 107451, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}} outputId="46859f7f-7722-4a29-f2fa-21230eb556f7"
#Fit the model
history = model.fit(np.array(X_train),
y_train,
batch_size = 64,
verbose = 1,
epochs = 100, #Changed to 3 from 50 for testing purposes.
validation_split = 0.1,
shuffle = False,
callbacks=[callbacks]
)
# + [markdown] id="TL5BU2fi_SQG"
# ### Accuracy and Loss
# + id="sH1SosCVwseD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1635978483857, "user_tz": 240, "elapsed": 622, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}} outputId="24f3fc3d-ded0-472a-cf5c-f19dbb9de7cc"
print("Test_Accuracy: {:.2f}%".format(model.evaluate(np.array(X_test), np.array(y_test))[1]*100))
# + colab={"base_uri": "https://localhost:8080/", "height": 308} id="rT59jJGk1dIU" executionInfo={"status": "ok", "timestamp": 1635978484831, "user_tz": 240, "elapsed": 982, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}} outputId="e142a164-3dfc-42e3-da95-99e5a65dc596"
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
t = f.suptitle('CNN Performance', fontsize=12)
f.subplots_adjust(top=0.85, wspace=0.3)
max_epoch = len(history.history['accuracy'])+1
epoch_list = list(range(1,max_epoch))
ax1.plot(epoch_list, history.history['accuracy'], label='Train Accuracy')
ax1.plot(epoch_list, history.history['val_accuracy'], label='Validation Accuracy')
ax1.set_xticks(np.arange(1, max_epoch, 10))
ax1.set_ylabel('Accuracy Value')
ax1.set_xlabel('Epoch')
ax1.set_title('Accuracy')
l1 = ax1.legend(loc="best")
ax2.plot(epoch_list, history.history['loss'], label='Train Loss')
ax2.plot(epoch_list, history.history['val_loss'], label='Validation Loss')
ax2.set_xticks(np.arange(1, max_epoch, 10))
ax2.set_ylabel('Loss Value')
ax2.set_xlabel('Epoch')
ax2.set_title('Loss')
l2 = ax2.legend(loc="best")
# + [markdown] id="rHH6XU2H_WHC"
# ### Save Model
# + id="--OKQ1-d1ksP" executionInfo={"status": "ok", "timestamp": 1635978485234, "user_tz": 240, "elapsed": 21, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhlSJDlZ0s-MgWUH_Pwct_zj3X1Lfh7O8awMGmCUg=s64", "userId": "04140676807795382520"}}
#Save the model
model.save('models/malaria_cnn.h5')
| DL/Malaria Cell Classification/FCNN/MalariaClassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn as sk
import numpy as np
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from tqdm import tqdm
# +
train = pd.read_csv("../input/train.csv", index_col=0)
test = pd.read_csv("../input/test.csv", index_col=0)
submission = pd.read_csv("../input/sample_submission.csv", index_col=0)
category = "wheezy-copper-turtle-magic"
# +
################
# QDA
################
# Config
features = [column for column in train.columns if column not in ["id", "target", category]]
probabilities = pd.Series(np.zeros(len(train)), index=train.index)
test_predictions = pd.Series(np.zeros(len(test)), index=test.index)
# Loop through wheezy-copper-turtle-magic
for i in tqdm(range(512)):
# Subset train and test
# where wheezy == i and features only
train_ = train.loc[train[category] == i, :]
test_ = test.loc[test[category] == i, :]
# VarianceThreshold
from sklearn.feature_selection import VarianceThreshold
feature_selector = VarianceThreshold(threshold = 1.5).fit(train_.loc[:, features])
train_2 = feature_selector.transform(train_.loc[:, features])
test_2 = feature_selector.transform(test_.loc[:, features])
# At this moment train_ and test_ contain all columns and only samples from wheezy == i
# and train_2 and test_2 contain only selected columns and samples from wheezy == i
# Stratified k-fold
skf = sk.model_selection.StratifiedKFold(n_splits=10, random_state=26, shuffle=True)
for split_train_index, split_test_index in skf.split(train_2, train_["target"]):
# QDA
qda = QuadraticDiscriminantAnalysis()
qda.fit(train_2[split_train_index, :], train_["target"][split_train_index])
# Getting probabilities of the test part of the split
split_probabilities = qda.predict_proba(train_2[split_test_index, :])[:, 1]
# Saving predictions
probabilities[train_.index[split_test_index]] += split_probabilities
test_predictions[test_.index] += qda.predict_proba(test_2)[:, 1] / skf.n_splits
print(sk.metrics.roc_auc_score(train["target"], probabilities))
# +
###################
# Data augmentation
###################
# Psuedo-labeling
mask = test_predictions[np.logical_or(test_predictions <= test_predictions.quantile(0.25),
test_predictions >= test_predictions.quantile(0.75))].index
train_only_psuedo = test.loc[mask, :]
train_only_psuedo = train_only_psuedo.assign(target=[1 if el > 0.5 else 0 for el in test_predictions[mask]])
# Check if indices match and columns match
assert all(test_predictions[mask].index == train_only_psuedo.index)
assert all(train.columns == train_only_psuedo.columns)
# Concatenate psudeo train with train
train_with_psuedo = pd.concat([train, train_only_psuedo])
# +
# The idea here is motivated by others and is inspired
# by the nature of the dataset
# Since we know make_classification with y_flip introduces flips
# Let's reverse them
# -
print(train_with_psuedo.columns)
| kernels/QDA-baseline-plus-data-augmentation.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Lesson 1 - Background and Terminology
//
// Author: <a href='mailto:<EMAIL>'><EMAIL></a>
// + [markdown] toc=true
// <h1>Table of Contents<span class="tocSkip"></span></h1>
// <div class="toc"><ul class="toc-item"><li><span><a href="#Terms,-concepts-and-background" data-toc-modified-id="Terms,-concepts-and-background-1"><span class="toc-item-num">1 </span>Terms, concepts and background</a></span><ul class="toc-item"><li><span><a href="#What-is-an-IDE,-and-why-it-doesn't-matter?" data-toc-modified-id="What-is-an-IDE,-and-why-it-doesn't-matter?-1.1"><span class="toc-item-num">1.1 </span>What is an IDE, and why it doesn't matter?</a></span></li><li><span><a href="#Why-you-will-and-should-learn-multiple-languages-and-why-the-language-doesn't-matter" data-toc-modified-id="Why-you-will-and-should-learn-multiple-languages-and-why-the-language-doesn't-matter-1.2"><span class="toc-item-num">1.2 </span>Why you will and should learn multiple languages and why the language doesn't matter</a></span><ul class="toc-item"><li><span><a href="#Python,-Java,-C,-C++,-JS" data-toc-modified-id="Python,-Java,-C,-C++,-JS-1.2.1"><span class="toc-item-num">1.2.1 </span>Python, Java, C, C++, JS</a></span></li><li><span><a href="#Syntax-is-not-programming" data-toc-modified-id="Syntax-is-not-programming-1.2.2"><span class="toc-item-num">1.2.2 </span>Syntax is not programming</a></span></li></ul></li><li><span><a href="#OOP---what-is-it-and-why-do-we-use-it" data-toc-modified-id="OOP---what-is-it-and-why-do-we-use-it-1.3"><span class="toc-item-num">1.3 </span>OOP - what is it and why do we use it</a></span></li><li><span><a href="#Git-and-github-and-standing-on-the-shoulders-of-giants" data-toc-modified-id="Git-and-github-and-standing-on-the-shoulders-of-giants-1.4"><span class="toc-item-num">1.4 </span>Git and github and standing on the shoulders of giants</a></span></li><li><span><a href="#The-Oracle-JVM,-JDK-and-JRE" data-toc-modified-id="The-Oracle-JVM,-JDK-and-JRE-1.5"><span class="toc-item-num">1.5 </span>The Oracle JVM, JDK and JRE</a></span></li><li><span><a href="#Why-Java-and-why-not-always-Java" data-toc-modified-id="Why-Java-and-why-not-always-Java-1.6"><span class="toc-item-num">1.6 </span>Why Java and why not always Java</a></span></li><li><span><a href="#Staying-Up-to-Date" data-toc-modified-id="Staying-Up-to-Date-1.7"><span class="toc-item-num">1.7 </span>Staying Up to Date</a></span></li></ul></li></ul></div>
// -
// <img src='https://images.unsplash.com/photo-1489875347897-49f64b51c1f8?ixlib=rb-1.2.1&ixid=eyJhcHBfaWQiOjEyMDd9&auto=format&fit=crop&w=1950&q=80'>
// ## Terms, concepts and background
//
// ### What is an IDE, and why it doesn't matter?
// IDE, or Integrated Development Environment, is the GUI in which you program. This can have many flavours and usually programmers have their favorite that they go to. Most people use <a href='https://www.jetbrains.com/idea/'>Jetbrains' IntelliJ</a> or <a href='https://www.eclipse.org/downloads/packages/release/kepler/sr1/eclipse-ide-java-developers'>Eclips</a>.
//
// Personally I prefer <a href='https://code.visualstudio.com/'>VS Code</a> as it is a one stop shop for all your programming language, rather than being geared to one spesific language. It has plently of add-ons that make it as feature rish, if not more, than most IDEs. However, sometimes the choice isn't yours and your team, company, or university will govern which IDE use.
//
// However, for these lessons, I'll actually be using <a href='https://jupyter.org/'>Jupyter Notebooks</a> as it makes it easy to illustrate code and comment accordingly. You can set up a Java kernel for Juypter by going to <a href='https://github.com/SpencerPark/IJava'>IJava</a> git repo.
//
// At the end of the day it doens't matter what IDE you use. Sometime you'll prefer one of the other, but as long as you are writing good, clean, working code your programming and any additional tools are just there to speed up the process.
// ### Why you will and should learn multiple languages and why the language doesn't matter
// Through your coding career you'll undoubtible learn more than just _one_ language. And you should! Learning different languages highlights strenghts and weaknesses of the various langauges and gives you a good understanding as to why people prefer one language over another for a specif task.
//
// At the end of the day programming is not the language program in, it is a mindset that you'll learn and refine.
// #### Python, Java, C, C++, JS
// Some of the common languages you'll encounter are: C/C++, Java, .NET/C#, Ruby, Python, PHP, Javascript. A small explanation is shown below taken from <a href='https://www.quora.com/What-are-the-pros-cons-and-uses-of-the-major-programming-languages-What-programming-language-should-I-learn-I-wanted-a-general-overview-of-what-certain-languages-are-used-for-what-is-good-or-bad-easy-hard-high-maintenance-or-low-maintenance'>this</a> blog post.
//
// ###### C/C++:
// Best/generally used for lower-level system stuff. Operating systems, device drivers, etc. Also very popular in game development as well as in the Linux community. Potential cons: very (relatively speaking) low-level, not ideal for developing ordinary applications and seldom used for web development. (Disclaimer: These are two different languages, but they are often grouped together like this and the above description does apply to both.)
//
// ###### Java:
// Probably most popular language overall, widely used for enterprise applications, web applications (though I wouldn't personally recommend it for that—read on), a good deal of open source stuff, and Android. Potential cons: not very "sexy", fairly boring in terms of language features, somewhat old-fashioned in many ways.
//
// ###### .NET (primarily C#):
// Similar to Java but not as widely used. I actually really like C# but wouldn't recommend it unless you have a specific need to learn .NET. Potential cons: perceived by many as a Java clone, useful in the same scenarios as Java but not as widely used.
// Objective-C: primarily popular again because of iOS. Related to C/C++ but with some very different syntax mixed in. Potential cons: very narrow applicability (though make no mistake: iOS is a pretty big reason to learn it).
//
// ###### Ruby:
// Very popular and "cool" esp. in the startup community. Great for writing web applications using Rails, hugely popular framework, or other lightweight frameworks that exist (e.g., Sinatra). Better than Java for creating a web application quickly in my opinion. Potential cons: fast moving language, lots of out-of-date online documentation, not really useful for developing GUI applications—pretty much web-only, realistically speaking.
//
// ###### Python:
// Python is easy to use, powerful, and versatile, making it a great choice for beginners and experts alike. Python’s readability makes it a great first programming language — it allows you to think like a programmer and not waste time with confusing syntax. A slogan for Python is: "The second best language for everything" - in other words, there is usually a better, faster, more optimised language for an application, however, Python will also get you there in half the time. This makes it a great language to have in your arsinal.
//
// ###### PHP:
// A language people love to hate. Wildly ubiquitous but considered by many to be a terrible (i.e., poorly designed) language. That said, used on a huge number of successful websites, including Facebook and Wordpress. To my knowledge, used exclusively for web applications. Potential cons: perception of low quality within the industry, lots of bad code examples online.
//
// ###### JavaScript:
// Amazing language that seems to be in the middle of a renaissance. Formerly used exclusively for client-side functionality that runs in users' web browsers, but now also used as a server-side language in some cases (using Node.js). Potential cons: considered a "toy" language by some (wrongly, in my opinion), still mostly used only for browser-side functionality in practice.
// #### Syntax is not programming
// To reiterate: the language you are programming in, is not programming, rather it is just the syntax used to execute concepts. Below I show how to declare a variable and print out "Hallo World" in each of the languages above.
//
// ###### C/C++:
//
// ```C
// #include <stdio.h>
// int main() {
// // initialise string variable (which is an array of chars) with its size of chars: 200.
// char message[200] = "Hallo World!";
// printf(message);
// return 0;
// }
// ```
//
// ###### Java:
//
// ```java
// public class HelloWorld {
// public static void main(String[] args) {
// // Prints "Hello, World" to the terminal window.
// String message = "Hello, World";
// System.out.println(message);
// }
// }
// ```
//
// ###### .NET (primarily C#):
//
// ```C#
// using System;
// using System.Collections.Generic;
// using System.Linq;
// using System.Text;
// using System.Threading.Tasks;
//
// namespace ConsoleApp1
// {
// class Program
// {
// static void Main(string[] args){
// string message = "Hello World!!";
// Console.WriteLine(message);
// Console.ReadLine();}
// }
// }
// ```
//
// ###### Ruby:
//
// ```ruby
// message = 'Hello World!'
// puts message
// ```
//
// ###### Python:
//
// ```python
// message = 'Hello World!'
// print(message)
// ```
//
// ###### PHP:
//
// ```html
// <!DOCTYPE html>
// <html>
// <body>
//
// <?php
// $message = "Hallo World!";
// echo "" . $color . "<br>";
// ?>
//
// </body>
// </html>
// ```
//
// ###### JavaScript:
//
// ```javascript
// var message = "Hallo World!"
// console.log(message
// ```
//
// As you can see, you can do the same thing in every language, the syntax and how you do it is all that differs. So don't get hampered by syntax, just google it, rather think about the problem you are trying to solve.
// ### OOP - what is it and why do we use it
// OOP - short for Object Orientated Programming, is a concept that you'll hear everyday the deeper you go into programming. It is a mindset of abstracting small logical blocs in a modular way to make reusing them easy.
//
// For example, we might have a class full of student and each of them has a lenght and weight. We could declare a list of string containing the details of students, like:
//
// ```javascript
// students [{"name":"John", "length":180, "weight": 80},
// {"name":"Jane", "length":175, "weight": 70},
// ...]
// ```
//
// or we can create a Student **Class**, also called an **Object**, that holds the information of each student. For example:
//
// ```java
// public class Student {
// String name;
// int length;
// int weight;
//
// public Student(String n, int l, int w) {
// name = n;
// length = l;
// weight = w;
// }
//
// public static void main(String[] args) {
// Student John = new Student("John", 180, 80);
// Student Jane = new Student("Jane", 175, 70);
// System.out.println(John.name + " " + John.length + " " + John.weight);
// System.out.println(Jane.name + " " + Jane.length + " " + Jane.weight);
// }
// }
//
// // Outputs
// // John 180 80
// // Jane 175 70
// ```
//
// This might seem like extra effort now, but it makes modifying, maintaining and integrating code so much easier. To the point where you can't live without it. More on this later in the later.
// + [markdown] heading_collapsed=true
// ### Git and github and standing on the shoulders of giants
// + [markdown] hidden=true
// You'll probably not learn about git just yet, but it is of paramount importance if you want to program. For now we'll just say it is a way to keep track of versions of your software. It enables you to release versions of your code and fallback to previous changes if something doens't work or breaks do to a change.
//
// To get you use to how git (and github) works, these notebooks are hosted on github on <a href='https://github.com/louwjlabuschagne/com-sci-101'>this</a> repository. For now, just look at the lessons on github - we'll unpack git and github as we go along, but rest assured. It is quite a complicated concept that takes a few years to understand, so best to start getting your feet wet now.
//
// The advantage of using git and github is that you can reuse a lot of code that many people have previously writen - and you should! Rule 1 in programming is don't re-invent the wheel - if somebody has done the work before, why should you do it again? Just make sure you give credit to any code you copy and use - especially at Universities they can be a bit heavy on copying code, but once you get into the real world and have to get something working, nobody cares where you got the code, as long as it works.
// + [markdown] heading_collapsed=true
// ### The Oracle JVM, JDK and JRE
// + [markdown] hidden=true
// Java is a language that is used widely because code that you write in Java is highly portable. The same code you wrote on you Mac will work on Windows, Android, an ATM or anything else running the JVM.
//
// The **JVM** is a the **Java Virtual Machine** and you can think of it as a layer between Java and the host you are running your Java code on. It basically deals with translating your Java code to whatever is needed on the host.
//
// In order ot develop Java you'll need the JDK, or **Java Development Kit** installed. This has all the internals needed to create, compile, debug and test your Java programs.
//
// However, if you want to deploy an application you don't always need the JDK, but _just_ the **JRE** or **Java Runtime Environment**. The JRE has a much smaller footprint compared to the JDK and allows your java programs to execute.
// + [markdown] heading_collapsed=true
// ### Why Java and why not always Java
// + [markdown] hidden=true
// Java is great for many things. It is a great mid-point language regarding complexcity as well as ease of use (although it might not feel that way right now). This is why it is one of the go to languages for teaching people to code. It elludes to low level concepts like memory allocation, but Java also has some user friendly helpers like the Java garbage collector.
//
// However, be open to learning new languages whenever you can as this will aid you in learning Java even more.
// -
// ### Staying Up to Date
// Programming is a highly dynamic field. Every week there is something new that gets released. Sometimes it's a new version of a language, or an entirely new language, or an update to a core package of a language that changes everything.
//
// To be a succesful programmer you have to stay up to date with the trends and what new features come out. I recommend searching for a YouTube channel that speaks to you, a twitter handle or a sub-reddit that keeps you up to date with new features. As with learning new languages, being aware of new features gives you an understanding of what and why something is currenlty lacking and broadens your knowledge about a language and programming in general.
| classes/Lesson_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/teomotun/Restaurant-Plug/blob/main/Yelp_restaurant_classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5MbTZBygBR0N"
# # Configuration
# + id="fdJL9mi2wHTx" outputId="dedb3830-8676-47ed-d8c4-0e84cf43928c" colab={"base_uri": "https://localhost:8080/", "height": 357}
#@title GPU INFO
# gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
if gpu_info.find('failed') >= 0:
print('Select the Runtime > "Change runtime type" menu to enable a GPU accelerator, ')
print('and then re-execute this cell.')
else:
print(gpu_info)
# + id="vgxq4u6lw3c3" outputId="1127d811-94a7-4db1-c16c-6a5570de3af5" colab={"base_uri": "https://localhost:8080/", "height": 68}
#@title More memory
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print('Your runtime has {:.1f} gigabytes of available RAM\n'.format(ram_gb))
if ram_gb < 20:
print('To enable a high-RAM runtime, select the Runtime > "Change runtime type"')
print('menu, and then select High-RAM in the Runtime shape dropdown. Then, ')
print('re-execute this cell.')
else:
print('You are using a high-RAM runtime!')
# + id="KVgpRq6BxOyw" outputId="19716857-cb68-48bd-9e0b-fe7faa54f07e" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="Du-vOUJrxXNt"
# !apt install -y caffe-cuda
# !pip install --upgrade kaggle
# !kaggle -v
# + id="ryMELqxozsx8"
# !mkdir -p ~/.kaggle
# !cp /content/drive/My\ Drive/Yelp-Restaurant-Classification/kaggle.json ~/.kaggle/
# Change the permission
# !chmod 600 ~/.kaggle/kaggle.json
# Change dir to where data folder
# %cd drive/
# %cd My\ Drive/
# %cd Yelp-Restaurant-Classification/Model/
# %cd data
# !pwd
# Download the kaggle dataset
# !kaggle competitions download -c yelp-restaurant-photo-classification
# Unzip and remove zip files
# !unzip \*.zip && rm *.zip
# + id="MV1OdXC10gyH" outputId="3be6e4cb-69aa-43df-bf51-d35e8e9db244" colab={"base_uri": "https://localhost:8080/", "height": 119}
import tarfile
import os
def extract_tgz(filename):
print("Working on: " + filename)
tar = tarfile.open(filename, "r:gz")
tar.extractall()
tar.close()
os.remove(filename)
print("-----------")
return
tgzs = [
"sample_submission.csv.tgz", "test_photo_to_biz.csv.tgz",
"test_photos.tgz", "train.csv.tgz",
"train_photo_to_biz_ids.csv.tgz", "train_photos.tgz"
]
for tgz in tgzs:
try:
extract_tgz(tgz)
except:
pass
# + id="5QqkxlNeinHG" outputId="165d927a-7ced-4e83-c244-43c0d8ecd279" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !apt install -y caffe-cuda
# + [markdown] id="s0P-PGoaHABQ"
# # First Stage
#
# + [markdown] id="bzGYpJMNpNi8"
# Total training set contains 234842 photos of 2000 restaurants
#
# + id="NSEoXEwF3emt" outputId="03997fc3-7469-48cb-af28-bb37820d249d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# #%%writefile training_image_features.py
import numpy as np
import pandas as pd
import tarfile
import skimage
import io
import h5py
import os
import caffe
import time
# Paths
CAFFE_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/caffe/"
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
DATA_ = "/content/"
# Model creation
# Using bvlc_reference_caffenet model for training
import os
if os.path.isfile(CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'):
print('CaffeNet found.')
else:
print('Downloading pre-trained CaffeNet model...')
#os.system('/caffe/scripts/download_model_binary.py /caffe/models/bvlc_reference_caffenet')
# !python /content/drive/My\ Drive/Yelp-Restaurant-Classification/Model/caffe/scripts/download_model_binary.py /content/drive/My\ Drive/Yelp-Restaurant-Classification/Model/caffe//models/bvlc_reference_caffenet
model_def = CAFFE_HOME + 'models/bvlc_reference_caffenet/deploy.prototxt'
model_weights = CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'
# Create a net object
model = caffe.Net(model_def, # defines the structure of the model
model_weights, # contains the trained weights
caffe.TEST) # use test mode (e.g., don't perform dropout)
# set up transformer - creates transformer object
transformer = caffe.io.Transformer({'data': model.blobs['data'].data.shape})
# transpose image from HxWxC to CxHxW
transformer.set_transpose('data', (2, 0, 1))
transformer.set_mean('data', np.load(CAFFE_HOME + 'python/caffe/imagenet/ilsvrc_2012_mean.npy').mean(1).mean(1))
# set raw_scale = 255 to multiply with the values loaded with caffe.io.load_image
transformer.set_raw_scale('data', 255)
# swap image channels from RGB to BGR
transformer.set_channel_swap('data', (2, 1, 0))
def extract_features(image_paths):
"""
This function is used to extract feature from the current batch of photos.
Features are extracted using the pretrained bvlc_reference_caffenet
Instead of returning 1000-dim vector from SoftMax layer, using fc7 as the final layer to get 4096-dim vector
"""
test_size = len(image_paths)
model.blobs['data'].reshape(test_size, 3, 227, 227)
model.blobs['data'].data[...] = list(map(lambda x: transformer.preprocess('data', skimage.img_as_float(skimage.io.imread(x)).astype(np.float32) ), image_paths))
out = model.forward()
return model.blobs['fc7'].data
if not os.path.isfile(FEATURES_HOME + 'train_features.h5'):
"""
If this file doesn't exist, create a new one and set up two columns: photoId, feature
"""
file = h5py.File(FEATURES_HOME + 'train_features.h5', 'w')
photoId = file.create_dataset('photoId', (0,), maxshape=(None,), dtype='|S54')
feature = file.create_dataset('feature', (0, 4096), maxshape=(None, 4096), dtype=np.dtype('int16'))
file.close()
# If this file exists, then track how many of the images are already done.
file = h5py.File(FEATURES_HOME + 'train_features.h5', 'r+')
already_extracted_images = len(file['photoId'])
file.close()
# Get training images and their business ids
train_data = pd.read_csv(DATA_ + 'train_photo_to_biz_ids.csv')
train_photo_paths = [os.path.join(DATA_ + 'train_photos/', str(photo_id) + '.jpg') for photo_id in
train_data['photo_id']]
# Each batch will have 500 images for feature extraction
train_size = len(train_photo_paths)
batch_size = 500
batch_number = round(already_extracted_images / batch_size + 1,3)
hours_elapsed = 0
print("Total images:", train_size)
print("already_done_images: ", already_extracted_images)
# Feature extraction of the train dataset
for image_count in range(already_extracted_images, train_size, batch_size):
start_time = round(time.time(),3)
# Get the paths for images in the current batch
image_paths = train_photo_paths[image_count: min(image_count + batch_size, train_size)]
# Feature extraction for the current batch
features = extract_features(image_paths)
# Update the total count of images done so far
total_done_images = image_count + features.shape[0]
# Storing the features in h5 file
file = h5py.File(FEATURES_HOME + 'train_features.h5', 'r+')
file['photoId'].resize((total_done_images,))
file['photoId'][image_count: total_done_images] = np.array(image_paths,dtype='|S54')
file['feature'].resize((total_done_images, features.shape[1]))
file['feature'][image_count: total_done_images, :] = features
file.close()
print("Batch No:", batch_number, "\tStart:", image_count, "\tEnd:", image_count + batch_size, "\tTime elapsed:", hours_elapsed, "hrs", "\tCompleted:", round(float(
image_count + batch_size) / float(train_size) * 100,3), "%")
batch_number += 1
hours_elapsed += round(((time.time() - start_time)/60)/60,3)
# + [markdown] id="azRPPhVypchy"
# Test set contains 1190225 of 10000 restaurants but I could only load 395500 due to memory constraints
# + id="PIZL1bIjoi7y" outputId="d9f22cde-2499-42c3-98ae-cd962544d152" colab={"base_uri": "https://localhost:8080/", "height": 34}
# #%%writefile test_image_features.py
import numpy as np
import pandas as pd
import tarfile
import skimage
import io
import h5py
import os
import caffe
import time
# Paths
CAFFE_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/caffe/"
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
DATA_ = "/content/"
# Model creation
# Using bvlc_reference_caffenet model for training
import os
if os.path.isfile(CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'):
print('CaffeNet found.')
else:
print('Downloading pre-trained CaffeNet model...')
#os.system('/caffe/scripts/download_model_binary.py /caffe/models/bvlc_reference_caffenet')
# !python /content/drive/My\ Drive/Yelp-Restaurant-Classification/Model/caffe/scripts/download_model_binary.py /content/drive/My\ Drive/Yelp-Restaurant-Classification/Model/caffe//models/bvlc_reference_caffenet
model_def = CAFFE_HOME + 'models/bvlc_reference_caffenet/deploy.prototxt'
model_weights = CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'
# Create a net object
model = caffe.Net(model_def, # defines the structure of the model
model_weights, # contains the trained weights
caffe.TEST) # use test mode (e.g., don't perform dropout)
# set up transformer - creates transformer object
transformer = caffe.io.Transformer({'data': model.blobs['data'].data.shape})
# transpose image from HxWxC to CxHxW
transformer.set_transpose('data', (2, 0, 1))
transformer.set_mean('data', np.load(CAFFE_HOME + 'python/caffe/imagenet/ilsvrc_2012_mean.npy').mean(1).mean(1))
# set raw_scale = 255 to multiply with the values loaded with caffe.io.load_image
transformer.set_raw_scale('data', 255)
# swap image channels from RGB to BGR
transformer.set_channel_swap('data', (2, 1, 0))
def extract_features(image_paths):
"""
This function is used to extract feature from the current batch of photos.
Features are extracted using the pretrained bvlc_reference_caffenet
Instead of returning 1000-dim vector from SoftMax layer, using fc7 as the final layer to get 4096-dim vector
"""
test_size = len(image_paths)
model.blobs['data'].reshape(test_size, 3, 227, 227)
model.blobs['data'].data[...] = list(map(lambda x: transformer.preprocess('data', skimage.img_as_float(skimage.io.imread(x)).astype(np.float32) ), image_paths))
out = model.forward()
return model.blobs['fc7'].data
if not os.path.isfile(FEATURES_HOME + 'test_features.h5'):
"""
If this file doesn't exist, create a new one and set up two columns: photoId, feature
"""
file = h5py.File(FEATURES_HOME + 'test_features.h5', 'w')
photoId = file.create_dataset('photoId', (0,), maxshape=(None,), dtype='|S54')
feature = file.create_dataset('feature', (0, 4096), maxshape=(None, 4096), dtype=np.dtype('int16'))
file.close()
# If this file exists, then track how many of the images are already done.
file = h5py.File(FEATURES_HOME + 'test_features.h5', 'r+')
already_extracted_images = len(file['photoId'])
file.close()
# Get testing images and their business ids
test_data = pd.read_csv(DATA_HOME + 'test_photo_to_biz.csv')
test_photo_paths = [os.path.join(DATA_ + 'test_photos/', str(photo_id) + '.jpg') for photo_id in
test_data['photo_id']]
# Each batch will have 500 images for feature extraction
test_size = 395500#len(test_photo_paths)
batch_size = 500
batch_number = round(already_extracted_images / batch_size + 1,3)
hours_elapsed = 0
print("Total images:", test_size)
print("already_done_images: ", already_extracted_images-500)
# Feature extraction of the test dataset
for image_count in range(already_extracted_images, test_size, batch_size):
start_time = round(time.time(),3)
# Get the paths for images in the current batch
image_paths = test_photo_paths[image_count: min(image_count + batch_size, test_size)]
# Feature extraction for the current batch
features = extract_features(image_paths)
# Update the total count of images done so far
total_done_images = image_count + features.shape[0]
# Storing the features in h5 file
file = h5py.File(FEATURES_HOME + 'test_features.h5', 'r+')
try:
file['photoId'].resize((total_done_images,))
file['photoId'][image_count: total_done_images] = np.array(image_paths,dtype='|S54')
file['feature'].resize((total_done_images, features.shape[1]))
file['feature'][image_count: total_done_images, :] = features
file.close()
except Exception as e:
print(e)
file.close()
print("Batch No:", batch_number, "\tStart:", image_count, "\tEnd:", image_count + batch_size, "\tTime elapsed:", hours_elapsed, "hrs", "\tCompleted:", round(float(
image_count + batch_size) / float(test_size) * 100,3), "%")
batch_number += 1
hours_elapsed += round(((time.time() - start_time)/60)/60,3)
# + [markdown] id="Ww61ekEpS5vj"
# # Second Stage
# + [markdown] id="xJ9lmPd_p3kS"
# Restaurant aggregation of training features
# + id="8CHbv3hXn0yv" outputId="799a10a3-2bbc-46ac-c9c4-3ffd325170bd" colab={"base_uri": "https://localhost:8080/", "height": 34}
# #%%writefile training_restaurant_features.py
import pandas as pd
import h5py
# Paths
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
# Get photo->business mapping from the file provided
train_photo_to_biz_ids = pd.read_csv(DATA_HOME + 'train_photo_to_biz_ids.csv')
# Get labels for businesses in the training data
train_data_business = pd.read_csv(DATA_HOME + 'train.csv').dropna()
# Sort these labels in the ascending order for simplicity e.g. (0, 6, 4, 2, 5) -> (0, 2, 4, 5, 6)
train_data_business['labels'] = train_data_business['labels'].apply(
lambda feature_vector: tuple(sorted(int(feature) for feature in feature_vector.split())))
train_data_business.set_index('business_id', inplace=True)
# Get business ids
business_ids = train_data_business.index.unique()
print("Total train business:", len(business_ids))
# Reading stored features from h5 file
train_features_file = h5py.File(FEATURES_HOME + 'train_features.h5', 'r')
train_features = np.copy(train_features_file['feature'])
train_features_file.close()
# Create a pandas dataframe to make the data ready for training the SVM classifier in the following format
train_df = pd.DataFrame(columns=['business_id', 'label', 'feature'])
for business_id in business_ids:
"""
For each business, write the values for the above triplet in the file viz. ['business_id', 'label', 'feature']
"""
business_id = int(business_id)
# Get the labels for the current business
label = train_data_business.loc[business_id]['labels']
# Get all the images which represent the current business with business_id
images_for_business_id = train_photo_to_biz_ids[train_photo_to_biz_ids['business_id'] == business_id].index.tolist()
# As a feature for current business, take the average over all the images
feature = list(np.mean(train_features[images_for_business_id], axis=0))
# Put the triplet into the data frame
train_df.loc[business_id] = [business_id, label, feature]
print("Train business feature extraction is completed.")
# Write the above data frame into a csv file
with open(FEATURES_HOME + 'train_aggregate_features.csv', 'w') as business_features_file:
train_df.to_csv(business_features_file, index=False)
# + id="zeQFt9V8wI4t" outputId="c0affa26-12f7-4cc0-905e-6e3906f8f41c" colab={"base_uri": "https://localhost:8080/", "height": 204}
train_df.head()
# + id="osIEJ2mZwKJk" outputId="0f69a5bd-324e-49d7-acb1-e64412d63f92" colab={"base_uri": "https://localhost:8080/", "height": 34}
train_df.shape
# + [markdown] id="QICHq4tDqIxA"
# Restaurant aggregation of test features
# + id="rr5DQ56DwJqV" outputId="8220fbb5-e896-4ab3-9d78-<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 34}
# #%%writefile testing_restaurant_features.py
import numpy as np
import pandas as pd
import h5py
# Paths
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
# Get photo->business mapping from the file provided
test_photo_to_biz_ids = pd.read_csv(DATA_HOME + 'test_photo_to_biz.csv')[:395500]
# Get business ids
business_ids = test_photo_to_biz_ids['business_id'].unique()
print("Total test business:", len(business_ids))
# Reading stored features from h5 file
test_features_file = h5py.File(FEATURES_HOME + 'test_features.h5', 'r')
# test_features = test_features_file['feature']
# test_features_file.close()
print(test_features_file['feature'][0])
# Create a pandas dataframe to make the data ready for training the SVM classifier in the following format
# Note that there will not be 'label' column as this is the actual testing data provided by Yelp
test_df = pd.DataFrame(columns=['business_id', 'feature'])
id = 0
for business_id in business_ids:
"""
For each business, write the values for the above tuple in the file viz. ['business_id', 'feature']
"""
# Get all the images which represent the current business with business_id
images_for_business_id = test_photo_to_biz_ids[test_photo_to_biz_ids['business_id'] == business_id].index.tolist()
# images_for_business_id[0]:(images_for_business_id[-1]+1)
# As a feature for current business, take the average over all the images
feature = list(
np.mean(np.asarray(test_features_file['feature'][:395500,:][images_for_business_id[0]:(images_for_business_id[-1] + 1)]),
axis=0))
# Put the tuple into the data frame
test_df.loc[business_id] = [business_id, feature]
id += 1
if id % 100 == 0:
print("ID:", id)
print("Test business feature extraction is completed.")
test_features_file.close()
# Write the above data frame into a csv file
with open(FEATURES_HOME + 'test_aggregated_features.csv', 'w') as business_features_file:
test_df.to_csv(business_features_file, index=False)
# + id="5ollIysVih4A" outputId="8d0ccb9d-4697-4ea7-ef0b-e773207a1065" colab={"base_uri": "https://localhost:8080/", "height": 204}
test_df.head()
# + id="MEGpPGo0jAZ0" outputId="64d567c1-a2f7-4bf6-e6bc-404bb6f5d7eb" colab={"base_uri": "https://localhost:8080/", "height": 34}
test_df.shape
# + [markdown] id="Co3DQEb6Uu7k"
# # 3rd Stage
# + [markdown] id="keEtp7gqqbzU"
# Training Restaurant Label Classifier on Training and Validation Set
# + [markdown] id="F3OU52hUnfju"
# SVC, KNN, RF, Extratrees Model, Stacking Model of 4 of them and a Stack of 3 KNN AND 1 RF
# + id="Y73MWTXv2k-7" outputId="e6e41166-483b-41b9-c90f-8a9d8d82b41d" colab={"base_uri": "https://localhost:8080/", "height": 598}
# #%%writefile training_classification_model.py
import numpy as np
import statistics
import pandas as pd
import time
import os
from sklearn.metrics import f1_score, accuracy_score
from sklearn.multiclass import OneVsRestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.model_selection import cross_val_predict
from sklearn.externals import joblib
def get_labels(label_string):
"""
This function converts label from string to array of labels
Input: "(1, 2, 3, 4, 5)"
Output: [1, 2, 3, 4, 5]
"""
label_array = label_string[1:-1]
label_array = label_array.split(',')
label_array = [int(label) for label in label_array if len(label) > 0]
return label_array
def get_features(feature_string):
"""
This function converts feature vector from string to array of features
Input: "(1.2, 3.4, ..., 9.10)"
Output: [1.2, 3.4, ..., 9.10]
"""
feature_array = feature_string[1:-1]
feature_array = feature_array.split(',')
feature_array = [float(label) for label in feature_array]
return feature_array
# Set home paths for data and features
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
MODELS_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/model/'
# Read training data and test data
train_data = pd.read_csv(FEATURES_HOME + 'train_aggregate_features.csv')
# Separate the labels from features in the training data
trainX = np.array([get_features(feature) for feature in train_data['feature']])
trainY = np.array([get_labels(label) for label in train_data['label']])
# Use validation data for calculating the training accuracy, random_state ensures reproducible results without overfitting
trainX, validationX, trainY, validationY = train_test_split(trainX, trainY, test_size=0.3, random_state=42)
# Binary representation (just like one-hot vector) (1, 3, 5, 9) -> (1, 0, 1, 0, 1, 0, 0, 0, 1)
mlb = MultiLabelBinarizer()
trainY = mlb.fit_transform(trainY)
# Do the same for validation labels
actual_labels = validationY
mlb = MultiLabelBinarizer()
validationY = mlb.fit_transform(validationY)
svc_clf = OneVsRestClassifier(SVC(kernel='linear', probability=True, verbose=True))
rf_clf = RandomForestClassifier(n_estimators=200, oob_score=True, n_jobs=-1, random_state=42)
knn_clf = KNeighborsClassifier()
extra_tree_clf = ExtraTreesClassifier(n_estimators=195, max_leaf_nodes=16, n_jobs=-1, random_state=42)
for clf in [svc_clf, rf_clf, knn_clf, extra_tree_clf]:
if not os.path.isfile(MODELS_HOME + f'{clf.__class__.__name__}.pkl'):
# Start time
start_time = time.time()
# Fit the classifier on the training data and labels
clf.fit(trainX, trainY)
cross_val = cross_val_predict(clf, validationX, validationY, cv=3)
print(f"{clf.__class__.__name__} trained.")
joblib.dump((mlb,clf), MODELS_HOME + f'{clf.__class__.__name__}.pkl')
print("Model saved.")
# End time
end_time = time.time()
print(f"Overall F1 Score for {clf.__class__.__name__}:", f1_score(cross_val, validationY, average='micro'))
print(f"Individual F1 Score for {clf.__class__.__name__}:", f1_score(cross_val, validationY, average=None))
print(f"Variance of {clf.__class__.__name__} is:", statistics.variance(f1_score(cross_val, validationY, average=None)))
print(f"Time taken for training the {clf.__class__.__name__}", end_time - start_time, "sec")
print("======================================================")
print("\n")
mlb,clf = joblib.load(MODELS_HOME + f'{clf.__class__.__name__}'+".pkl")
print(f"{clf.__class__.__name__} Model loaded.")
# Predict the labels for the validation data
preds_binary = clf.predict(validationX)
# Predicted labels are converted back
# (1, 0, 1, 0, 1, 0, 0, 0, 1) -> (1, 3, 5, 9)
predicted_labels = mlb.inverse_transform(preds_binary)
print("Validation Set Results:")
print(f"Overall F1 Score for {clf.__class__.__name__}:", f1_score(preds_binary, validationY, average='micro'))
print(f"Individual F1 Score for {clf.__class__.__name__}:", f1_score(preds_binary, validationY, average=None))
print(f"Variance of {clf.__class__.__name__} is:", statistics.variance(f1_score(preds_binary, validationY, average=None)))
print("======================================================")
X_train_1, X_train_2, y_train_1, y_train_2 = train_test_split(trainX, trainY, random_state=42)
svc_clf = OneVsRestClassifier(SVC(kernel='linear', probability=True, verbose=True))
rf_clf = RandomForestClassifier(n_estimators=200, oob_score=True, n_jobs=-1, random_state=42)
knn_clf = KNeighborsClassifier()
extra_tree_clf = ExtraTreesClassifier(n_estimators=195, max_leaf_nodes=16, n_jobs=-1, random_state=42)
start_time = time.time()
rnd_clf_2 = RandomForestClassifier(random_state=42)
for p in [svc_clf, rf_clf, knn_clf, extra_tree_clf]:
p.fit(X_train_1, y_train_1)
svc_clf_p = svc_clf.predict(X_train_2)
rf_clf_p = rf_clf.predict(X_train_2)
knn_clf_p = knn_clf.predict(X_train_2)
held_out = np.column_stack((svc_clf_p, rf_clf_p, knn_clf_p))
rnd_clf_2.fit(held_out, y_train_2)
result_1 = []
for p in [svc_clf, rf_clf, knn_clf]:
result_1.append(p.predict(validationX))
y_pred_s = rnd_clf_2.predict(np.column_stack(tuple(result_1)))
# End time
end_time = time.time()
print(f"Time taken for training the Stacked Model:", end_time - start_time, "sec")
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average='micro'))
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average=None))
print(f"Variance of Stacked Model is:", statistics.variance(f1_score(y_pred_s, validationY, average=None)))
# + [markdown] id="4kk_AyVCar1B"
# [link text](https://)Stack of 3 KNN AND 1 RF
# + id="pPixqDgGZTRo" outputId="21e7a9bd-1c68-4b4e-bcb8-1192c6d578ac" colab={"base_uri": "https://localhost:8080/", "height": 102}
# #%%writefile training_classification_model.py
import numpy as np
import statistics
import pandas as pd
import time
import os
from sklearn.metrics import f1_score, accuracy_score
from sklearn.multiclass import OneVsRestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import confusion_matrix
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.model_selection import cross_val_predict
from sklearn.externals import joblib
def get_labels(label_string):
"""
This function converts label from string to array of labels
Input: "(1, 2, 3, 4, 5)"
Output: [1, 2, 3, 4, 5]
"""
label_array = label_string[1:-1]
label_array = label_array.split(',')
label_array = [int(label) for label in label_array if len(label) > 0]
return label_array
def get_features(feature_string):
"""
This function converts feature vector from string to array of features
Input: "(1.2, 3.4, ..., 9.10)"
Output: [1.2, 3.4, ..., 9.10]
"""
feature_array = feature_string[1:-1]
feature_array = feature_array.split(',')
feature_array = [float(label) for label in feature_array]
return feature_array
# Set home paths for data and features
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
FEATURES_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/features/'
MODELS_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/model/'
# Read training data and test data
train_data = pd.read_csv(FEATURES_HOME + 'train_aggregate_features.csv')
# Separate the labels from features in the training data
trainX = np.array([get_features(feature) for feature in train_data['feature']])
trainY = np.array([get_labels(label) for label in train_data['label']])
# Use validation data for calculating the training accuracy, random_state ensures reproducible results without overfitting
trainX, validationX, trainY, validationY = train_test_split(trainX, trainY, test_size=0.3, random_state=42)
# Binary representation (just like one-hot vector) (1, 3, 5, 9) -> (1, 0, 1, 0, 1, 0, 0, 0, 1)
mlb = MultiLabelBinarizer()
trainY = mlb.fit_transform(trainY)
# Do the same for validation labels
actual_labels = validationY
mlb = MultiLabelBinarizer()
validationY = mlb.fit_transform(validationY)
svc_clf = OneVsRestClassifier(SVC(kernel='linear', probability=True, verbose=True))
rf_clf = RandomForestClassifier(n_estimators=200, oob_score=True, n_jobs=-1, random_state=42)
knn_clf = KNeighborsClassifier()
extra_tree_clf = ExtraTreesClassifier(n_estimators=195, max_leaf_nodes=16, n_jobs=-1, random_state=42)
for clf in [svc_clf, rf_clf, knn_clf, extra_tree_clf]:
if not os.path.isfile(MODELS_HOME + f'{clf.__class__.__name__}.pkl'):
# Start time
start_time = time.time()
# Fit the classifier on the training data and labels
clf.fit(trainX, trainY)
cross_val = cross_val_predict(clf, validationX, validationY, cv=3)
print(f"{clf.__class__.__name__} trained.")
joblib.dump((mlb,clf), MODELS_HOME + f'{clf.__class__.__name__}.pkl')
print("Model saved.")
# End time
end_time = time.time()
print(f"Overall F1 Score for {clf.__class__.__name__}:", f1_score(cross_val, validationY, average='micro'))
print(f"Individual F1 Score for {clf.__class__.__name__}:", f1_score(cross_val, validationY, average=None))
print(f"Variance of {clf.__class__.__name__} is:", statistics.variance(f1_score(cross_val, validationY, average=None)))
print(f"Time taken for training the {clf.__class__.__name__}", end_time - start_time, "sec")
print("======================================================")
print("\n")
mlb,clf = joblib.load(MODELS_HOME + f'{clf.__class__.__name__}'+".pkl")
print(f"{clf.__class__.__name__} Model loaded.")
# Predict the labels for the validation data
preds_binary = clf.predict(validationX)
# Predicted labels are converted back
# (1, 0, 1, 0, 1, 0, 0, 0, 1) -> (1, 3, 5, 9)
predicted_labels = mlb.inverse_transform(preds_binary)
print("Validation Set Results:")
print(f"Overall F1 Score for {clf.__class__.__name__}:", f1_score(preds_binary, validationY, average='micro'))
print(f"Individual F1 Score for {clf.__class__.__name__}:", f1_score(preds_binary, validationY, average=None))
print(f"Variance of {clf.__class__.__name__} is:", statistics.variance(f1_score(preds_binary, validationY, average=None)))
print("======================================================")
X_train_1, X_train_2, y_train_1, y_train_2 = train_test_split(trainX, trainY, random_state=42)
svc_clf = OneVsRestClassifier(SVC(kernel='linear', probability=True, verbose=True))
rf_clf = RandomForestClassifier(n_estimators=200, oob_score=True, n_jobs=-1, random_state=42)
knn_clf = KNeighborsClassifier()
extra_tree_clf = ExtraTreesClassifier(n_estimators=195, max_leaf_nodes=16, n_jobs=-1, random_state=42)
start_time = time.time()
rnd_clf_2 = RandomForestClassifier(random_state=42)
for p in [svc_clf, rf_clf, knn_clf, extra_tree_clf]:
p.fit(X_train_1, y_train_1)
svc_clf_p = svc_clf.predict(X_train_2)
rf_clf_p = rf_clf.predict(X_train_2)
knn_clf_p = knn_clf.predict(X_train_2)
held_out = np.column_stack((svc_clf_p, rf_clf_p, knn_clf_p))
rnd_clf_2.fit(held_out, y_train_2)
result_1 = []
for p in [svc_clf, rf_clf, knn_clf]:
result_1.append(p.predict(validationX))
y_pred_s = rnd_clf_2.predict(np.column_stack(tuple(result_1)))
# End time
end_time = time.time()
print(f"Time taken for training the Stacked Model:", end_time - start_time, "sec")
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average='micro'))
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average=None))
print(f"Variance of Stacked Model is:", statistics.variance(f1_score(y_pred_s, validationY, average=None)))
X_train_1, X_train_2, y_train_1, y_train_2 = train_test_split(trainX, trainY, random_state=42)
start_time = time.time()
rnd_clf1 = RandomForestClassifier(random_state=42)
knn_clf1 = KNeighborsClassifier()
knn_clf2 = KNeighborsClassifier()
knn_clf3 = KNeighborsClassifier()
knn_clf4 = KNeighborsClassifier()
rnd_clf2 = RandomForestClassifier(random_state=42)
for p in [knn_clf1, knn_clf2, knn_clf3, rnd_clf1]:
p.fit(X_train_1, y_train_1)
knn_clf1_p = knn_clf1.predict(X_train_2)
knn_clf2_p = knn_clf2.predict(X_train_2)
knn_clf3_p = knn_clf3.predict(X_train_2)
rnd_clf1_p = rnd_clf1.predict(X_train_2)
held_out = np.column_stack((knn_clf1_p, knn_clf2_p, knn_clf3_p, rnd_clf1_p))
rnd_clf_2.fit(held_out, y_train_2)
result_1 = []
for p in [knn_clf1, knn_clf1, knn_clf3, rnd_clf1]:
result_1.append(p.predict(validationX))
y_pred_s = rnd_clf_2.predict(np.column_stack(tuple(result_1)))
# End time
end_time = time.time()
print(f"Time taken for training the Stacked Model:", end_time - start_time, "sec")
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average='micro'))
print(f"Overall Stacked F1 Score for:", f1_score(y_pred_s, validationY, average=None))
print(f"Variance of Stacked Model is:", statistics.variance(f1_score(y_pred_s, validationY, average=None)))
# + [markdown] id="AuCgsjTnjkkl"
# Random Forest algorithm:
# Average F1 => 0.80516;
# Time taken to train => 17.7761 sec;
# Variance => 0.01443
#
# SVC algorithm:
# Average F1 => 0.79910;
# Time taken to train => 295.4199 sec;
# Variance => 0.01100
#
# KNN algorithm:
# Average F1 => 0.80656;
# Time taken to train => 2.7335 sec;
# Variance => 0.01132
#
# Extratrees algorithm:
# Average F1 => 0.76833;
# Time taken to train => 3.6577 sec;
# Variance => 0.03619
#
# Stacked algorithm (RF, SVC, KNN, Extratrees):
# Average F1 => 0.79852;
# Time taken to train => 165.1282 sec;
# Variance => 0.01412
#
# Stacked algorithm (RF, KNN, KNN, KNN):
# Average F1 => 0.77767;
# Time taken to train => 34.3231 sec;
# Variance => 0.01467
#
#
# + [markdown] id="C-78noS1tqSB"
# KNN and Random Forest appears to outperform the other models in terms of time, variance and F1 score with both over 80% across all classes. Stacking the best two predictors seem to have an adverse effect on the model.
#
# I'll stick with KNN for the rest of my analysis and testing
# + [markdown] id="1Fc6oQXmURok"
# # KNN Model Analysis
# + [markdown] id="cZqAJpYc0OqG"
# Compute ROC curve and ROC area for each class and Confusion Matrix of each class
# + id="mc2zMjNMubs3" outputId="698a194d-<PASSWORD>8-4<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# #%%writefile model_analysis.py
import numpy as np
import statistics
import pandas as pd
import time
import os
print(__doc__)
from itertools import cycle
from sklearn.metrics import f1_score, accuracy_score
from sklearn.metrics import roc_curve
from sklearn.metrics import auc
from sklearn.metrics import multilabel_confusion_matrix, confusion_matrix
import matplotlib.pyplot as plt
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.externals import joblib
MODELS_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/model/'
mlb,clf = joblib.load(MODELS_HOME + "KNeighborsClassifier.pkl")
# Predict the labels for the validation data
preds_binary = clf.predict(validationX)
# Predicted labels are converted back
# (1, 0, 1, 0, 1, 0, 0, 0, 1) -> (1, 3, 5, 9)
predicted_labels = mlb.inverse_transform(preds_binary)
conf_mx = multilabel_confusion_matrix(preds_binary,validationY)
i = 0
for conf in conf_mx:
print(i)
print(pd.DataFrame(conf))
print("======\n")
i += 1
n_classes = preds_binary.shape[1]
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(validationY[:, i], preds_binary[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
n_classes = preds_binary.shape[1]
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(validationY[:, i], preds_binary[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(preds_binary.ravel(), validationY.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
##############################################################################
# Plot ROC curves for the multiclass problem
# Compute macro-average ROC curve and ROC area
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += np.interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
plt.figure()
plt.plot(fpr["micro"], tpr["micro"],
label='micro-avg (area = {0:0.2f})'
''.format(roc_auc["micro"]),
linewidth=2)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-avg (area = {0:0.2f})'
''.format(roc_auc["macro"]),
linewidth=2)
for i in range(n_classes):
plt.plot(fpr[i], tpr[i], label='Class {0} (area = {1:0.2f})'
''.format(i, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Some extension of Receiver operating characteristic to multi-class')
plt.legend(loc="lower right")
plt.show()
# + [markdown] id="iNQb4-OVUC_D"
# # Testing 3 stage model on new data
#
#
# + id="E_A8oHvTkTz-" outputId="28b4f0ff-cea3-45e2-df2e-0f77b20bc834" colab={"base_uri": "https://localhost:8080/", "height": 71}
# #%%writefile get_prediction.py
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.ensemble import RandomForestClassifier
from sklearn.externals import joblib
import numpy as np
import pandas as pd
import tarfile
import skimage
import io
import h5py
import os
import caffe
import time
def get_predictions(RESTAURANT_HOME, CAFFE_HOME, DATA_HOME, MODELS_HOME):
"""
This function is used to make restaurant class prediction of photos from several directory paths.
Features are extracted using the pretrained bvlc_reference_caffenet
Instead of returning 1000-dim vector from SoftMax layer, using fc7 as the final layer to get 4096-dim vector.
The features are the passed to a KNN multi label classifier
"""
# Model creation
# Using bvlc_reference_caffenet model for training
import os
if os.path.isfile(CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'):
print('CaffeNet found.')
model_def = CAFFE_HOME + 'models/bvlc_reference_caffenet/deploy.prototxt'
model_weights = CAFFE_HOME + 'models/bvlc_reference_caffenet/bvlc_reference_caffenet.caffemodel'
# Create a net object
model = caffe.Net(model_def, # defines the structure of the model
model_weights, # contains the trained weights
caffe.TEST) # use test mode (e.g., don't perform dropout)
# set up transformer - creates transformer object
transformer = caffe.io.Transformer({'data': model.blobs['data'].data.shape})
# transpose image from HxWxC to CxHxW
transformer.set_transpose('data', (2, 0, 1))
transformer.set_mean('data', np.load(CAFFE_HOME + 'python/caffe/imagenet/ilsvrc_2012_mean.npy').mean(1).mean(1))
# set raw_scale = 255 to multiply with the values loaded with caffe.io.load_image
transformer.set_raw_scale('data', 255)
# swap image channels from RGB to BGR
transformer.set_channel_swap('data', (2, 1, 0))
def extract_features(image_paths):
"""
This function is used to extract feature from the current batch of photos.
Features are extracted using the pretrained bvlc_reference_caffenet
Instead of returning 1000-dim vector from SoftMax layer, using fc7 as the final layer to get 4096-dim vector
"""
test_size = len(image_paths)
model.blobs['data'].reshape(test_size, 3, 227, 227)
model.blobs['data'].data[...] = list(map(lambda x: transformer.preprocess('data', skimage.img_as_float(skimage.io.imread(x)).astype(np.float32) ), image_paths))
out = model.forward()
return model.blobs['fc7'].data
features = extract_features(image_paths)
mlb,clf = joblib.load(MODELS_HOME + "KNeighborsClassifier.pkl")
# Predict the labels for the validation data
preds_binary = clf.predict(features)
# Predicted labels are converted back
# (1, 0, 1, 0, 1, 0, 0, 0, 1) -> (1, 3, 5, 9)
predicted_labels = mlb.inverse_transform(preds_binary)
return predicted_labels
# + id="S8vAHo1Dl-V9" outputId="a14e7843-bd41-4869-eea2-8031c5539f69" colab={"base_uri": "https://localhost:8080/", "height": 850}
# Paths
CAFFE_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/caffe/"
DATA_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/data/"
MODELS_HOME = '/content/drive/My Drive/Yelp-Restaurant-Classification/Model/model/'
RESTAURANT_HOME = "/content/drive/My Drive/Yelp-Restaurant-Classification/Model/restaurant-images/"
image_paths = [RESTAURANT_HOME+f.strip() for f in os.listdir(RESTAURANT_HOME) if os.path.isfile(RESTAURANT_HOME + f)]
get_predictions(image_paths, CAFFE_HOME, DATA_HOME, MODELS_HOME)
# + id="RuIdDS7f2Srm" outputId="74721dfc-0651-4b4b-9a92-c7913ee3e60f" colab={"base_uri": "https://localhost:8080/", "height": 37}
RESTAURANT_HOME
# + id="5BLEjV4R2cil"
| Yelp_restaurant_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Protocols
# Python is a protocol based language.
# If you're coming from Java, you can think of protocols the same way you think of interfaces.
# Except Python does not have this very strict idea of an interface.
# You simply add some functions to your class using a specific name, and if Python finds it there, it will use it. The onus is on you to get the naming right, and correctly implementing any specific set of these functions that loosely make up the protocol yourself.
# #### The `str` and `repr` Protocols
# Let's take a look at a very simple example.
# When we have an object, we can as for it's string representation in two ways:
a = 10
str(a)
repr(a)
# These look identical, but that is not always the case. In general `str` is used for end-user display, and `repr` is used for development (or debugging) display.
# For example:
from fractions import Fraction
f = Fraction(1, 2)
str(f)
repr(f)
# Each class may implement it's own mechanism for returning a value for either `str` or `repr`.
# This is done by implementing the correct protocol.
# Let's create our own class and implement both the `str` and the `repr` protocols:
class Person:
def __init__(self, name):
self.name = name
def __str__(self):
return self.name.strip()
def __repr__(self):
return f"Person(name='{self.name}')"
# As you can see we simply implemented to specially name instance methods: `__str__` and `__repr__`.
#
# Let's use them:
p = Person('<NAME>ton')
# Now these are just instance methods, and can be called that way:
p.__str__()
p.__repr__()
# But, because of the special names we used, when we use the `str()` and `repr()` functions, Python will find and use our custom `__str__` and `__repr__` methods instead:
str(p)
repr(p)
# In Python, every class directly or indirectly, inherits from the `object` class. This class provides standard implementations for a lot of protocols.
class Point:
def __init__(self, x, y):
self.x = x
self.y = y
p = Point(1, 2)
str(p)
repr(p)
# As you can see the default string and representations simply document the class that was used to create that object, and the memory address of the instance. As you saw, we can override this default behavior by implementing our own special functions.
# #### The `addition` Protocol
# When we write something like this in Python:
1 + 2
# What is actually happening, is that integres implement the addition protocol, and when Python sees
#
# ```
# 1 + 2
# ```
#
# it actually uses the addition protocol defined by integers to evaluate that statement.
# We can implement this protocol in our custom classes too.
# Let's start by creating a basic vector class:
class Vector:
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return f"Vector({self.x}, {self.y})"
def __str__(self):
return f"({self.x}, {self.y})"
v1 = Vector(1, 2)
v2 = Vector(10, 20)
# We implemented the str and repr protocols, so we can do this:
print(str(v1))
print(repr(v1))
# But we cannot add those two vectors:
v1 + v2
# As you can see Python is telling us it does not know how to add two `Vector` instances together.
# We can tell Python how to do that, by simply implementin the `add` protocol:
class Vector:
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return f"Vector({self.x}, {self.y})"
def __str__(self):
return f"({self.x}, {self.y})"
def __add__(self, other):
new_x = self.x + other.x
new_y = self.y + other.y
return Vector(new_x, new_y)
# Note: technically it would be better to check that `other` is also a `Vector` instance, but let's ignore that for now.
v1 = Vector(1, 2)
v2 = Vector(10, 20)
# And now we can add those two vectors together:
v1 + v2
# Ok, let's just go back and fix the `__add__` method, to at least make sure we are adding two vectors, because here's what happens right now:
v1 + 10
# In fact, the weird things is that if we have another object with those `x` and `y` attributes, the addition may actually work!
class NotAVector:
def __init__(self, x, y, z):
self.x = x
self.y = y
self.x = z
nav = NotAVector(10, 20, 30)
v1 + nav
# So, we may want to restrict our addition to only two vectors:
class Vector:
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return f"Vector({self.x}, {self.y})"
def __str__(self):
return f"({self.x}, {self.y})"
def __add__(self, other):
if not isinstance(other, Vector):
raise TypeError('Addition is only supported between two Vector instances.')
new_x = self.x + other.x
new_y = self.y + other.y
return Vector(new_x, new_y)
v1 = Vector(1, 2)
v2 = Vector(10, 20)
nav = NotAVector(10, 20, 30)
v1 + v2
v1 + nav
v1 + 10
# but what if we wanted to support something like this:
v1 + (10, 20)
# or
v1 + [10, 20]
# We can enhance our `__add__` method to allow this:
class Vector:
def __init__(self, x, y):
self.x = x
self.y = y
def __repr__(self):
return f"Vector({self.x}, {self.y})"
def __str__(self):
return f"({self.x}, {self.y})"
def __add__(self, other):
if isinstance(other, (list, tuple)) and len(other) >= 2:
new_x = self.x + other[0]
new_y = self.y + other[1]
elif isinstance(other, Vector):
new_x = self.x + other.x
new_y = self.y + other.y
else:
raise TypeError(f"Unsupported type for Vector addition: {type(other)}")
return Vector(new_x, new_y)
v1 = Vector(1, 2)
v2 = Vector(10, 20)
nav = NotAVector(10, 20, 30)
v1 + v2
v1 + (100, 200)
v1 + [100, 200]
v1 + nav
# #### Other Protocols
# Most of the operators in Python, as well as various behavior traits of objects, are controlled in custom classes using these protocols, which you can find documented here:
#
# https://docs.python.org/3/reference/datamodel.html
| 16 - Protocols.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Campaign management
# Predict Interaction rate from ad texts
from fastai import *
from fastai.text import *
import os
import pathlib as path
import pandas as pd
import re
# ### Data loading and cleaning
HOME = path.Path('/home/jovyan/work')
DATA = HOME/'data'
MODELS = DATA/'corpus/models'
bs = 256 # Batch size
df = pd.read_csv(DATA/'podaci.csv', header=2, error_bad_lines=True, low_memory=False)
print(df.shape)
# for col_name in df.columns:
# print(col_name)
# Last 350 rows are totals
df[-351:-340]
# Bye totals
df.drop(df.tail(350).index, inplace=True)
df[-3:]
# Clean and preprocess text, lowercase all
def clean_text(dirty_text):
text = re.sub(r'http\S+','', str(dirty_text))
punctuation = '.!"#$%&()*+-/:;<=>?@[\\]^_`{|}~'
text = ''.join(ch for ch in text if ch not in set(punctuation))
text = text.lower()
# remove numbers
# text = text.replace("[0-9]", "")
text = re.sub("[0-9]", "", text)
text = re.sub("®", "", text)
# remove ekupi
text = re.sub('ekupi','', text, flags=re.IGNORECASE)
# Remove Keywords, KeyWords
text = re.sub('keyword','', text, flags=re.IGNORECASE)
# Remove whitespaces
text = ' '.join(text.split())
return text
# #### Prepare data set for training of LM
# Take all texts, doesn't matter are metrics OK
# Drop all non text ads
df = df.drop(df[(df["Ad type"] == 'Image ad') | (df["Ad type"] == 'Display ad')].index)
df_clean_lm = df.Headline.map(clean_text) + ' ' + df['Headline 1'].map(clean_text) + ' ' + df['Headline 2'].map(clean_text) + ' ' + df['Headline 3'].map(clean_text) + ' ' + df['Headline 4'].map(clean_text)+ ' '+ df['Headline 5'].map(clean_text) + ' '+ df.Description.map(clean_text) + ' '+ df['Description 1'].map(clean_text) + ' '+ df['Description 2'].map(clean_text)+ ' '+ df['Description 3'].map(clean_text)+ df['Description 4'].map(clean_text)+ ' '+ df['Description 5'].map(clean_text)
df_train_lm = pd.concat([df_clean_lm,df['Impr.']], axis= 1) # We need second column though is not used in training
df_train_lm.columns = ['text', 'impr']
df_train_lm[:20] # Let's check first 20 rows
print(df_train_lm.shape[0])
# Now let's drop duplicates
df_train_lm.drop_duplicates(subset ="text", keep = 'first', inplace = True)
df_train_lm.shape[0]
# Let's see some random texts
df[1200:1225]
# #### Language model train
# Previously trained ULMFIT for Croatian on Wiki data
campaign_lm = (TextList.from_df(df_train_lm, DATA)
.split_by_rand_pct(0.1)
.label_for_lm()
.databunch(bs=bs))
campaign_lm.save('campaign_lm.pkl')
campaign_lm.vocab.itos[:10]
# Save new Language model
campaign_lm.vocab.save(MODELS/'campaign_vocab.pkl') # pickle file
data_lm = load_data(DATA, 'campaign_lm.pkl', bs=bs)
data_lm.show_batch()
FILE_LM_ENCODER = MODELS/'pretrained_model'
FILE_ITOS = MODELS/'pretrained_itos'
learn = language_model_learner(data_lm, AWD_LSTM, pretrained=True, pretrained_fnames=[FILE_LM_ENCODER, FILE_ITOS], drop_mult=0.3)
learn.lr_find()
learn.recorder.plot(skip_end=12)
learn.fit_one_cycle(1, 1e-3, moms=(0.8,0.7))
learn.unfreeze()
learn.fit_one_cycle(10, 7e-3, moms=(0.8,0.7))
learn.predict('Kupite najbolji uređaj', temperature=1.1, min_p=0.001, n_words=10)
learn.save_encoder(MODELS/'campaign_encoder')
# #### Prepare data set for training
# Interaction rate (former CTR) and Impressions to numbers, drop all rows with few impr. or too high IR
df['Impr.'] = df['Impr.'].apply(lambda x: int(x.replace(',',''))) # Change Impressions to sum
df['Impr.'].sum() # Nr. of impressions, litle less then 27B
# Take percentage texts as sloats
df['Interaction rate'] = df['Interaction rate'].apply(lambda x: float(x.replace('%','').replace('--','0')))
df[df['Interaction rate'] > 25].shape[0] # How much is over 25%, we're gonna drop that
# Now let's delete all rows with no Impressions and Image ads
df = df.drop(df[(df['Impr.'] < 100) | (df['Interaction rate'] > 25)].index)
print(f'Number of rows for training: {df.shape[0]}')
df[:20]
# #### Dataframes for training
# Prepare and save dataframes with headlines and dascriptions for interaction regression
# +
df_clean_headline = df.Headline.map(clean_text) + ' ' + df['Headline 1'].map(clean_text) + ' ' + df['Headline 2'].map(clean_text) + ' ' + df['Headline 3'].map(clean_text) + ' ' + df['Headline 4'].map(clean_text)+ ' '+ df['Headline 5'].map(clean_text)
df_clean_description = df.Description.map(clean_text) + ' '+ df['Description 1'].map(clean_text) + ' '+ df['Description 2'].map(clean_text)+ ' '+ df['Description 3'].map(clean_text)+ df['Description 4'].map(clean_text)+ ' '+ df['Description 5'].map(clean_text)
df_train_headline = pd.concat([df_clean_headline, df['Interaction rate']], axis= 1)
df_train_description = pd.concat([df_clean_description, df['Interaction rate']], axis= 1)
# Rename columns
df_train_headline.columns = ['text', 'rate']
df_train_description.columns = ['text', 'rate']
# Drop duplicates
df_train_headline.drop_duplicates(subset ="text", keep = 'first', inplace = True)
print(f'Number of different headline texts: {df_train_headline.shape[0]}')
df_train_description.drop_duplicates(subset ="text", keep = 'first', inplace = True)
print(f'Number of different description texts: {df_train_description.shape[0]}')
df_train_headline.to_csv(DATA/'headlines.csv', index=False)
df_train_headline.to_csv(DATA/'descriptions.csv', index=False)
# -
# ### Training regression
# +
df_train = pd.read_csv(DATA/'headlines.csv')
df_train.head()
# -
df_train.shape # Lost lot of rows
# See possible approaches https://forums.fast.ai/t/regression-using-fine-tuned-language-model/29091/56
data_regr = (TextList.from_df(df=df_train, path=DATA, cols='text', vocab=data_lm.vocab)
.split_by_rand_pct(0.2)
.label_from_df('rate', label_cls=FloatList)
.databunch(bs=bs))
# Loss function - mse, optimizer - Adam ()-default, last layer - linear
learn = text_classifier_learner(data_regr,
arch = AWD_LSTM,
drop_mult=0.3,
metrics = rmse)
learn.loss_func = MSELossFlat()
learn.load_encoder(MODELS/'campaign_encoder')
learn.freeze()
learn.lr_find()
learn.recorder.plot()
learn.fit_one_cycle(1, 6e-2, moms=(0.8,0.7))
learn.save(MODELS/'headline-first-run')
learn.load(MODELS/'headline-first-run')
learn.freeze_to(-2)
learn.fit_one_cycle(1, slice(6e-2/(2.6**4),6e-1), moms=(0.8,0.7))
# Save to possibly retrain again
learn.save(MODELS/'headline-second-run')
# Load again
learn.load(MODELS/'headline-second-run')
pass # don't show model definition again
learn.freeze_to(-3)
learn.fit_one_cycle(2, slice(1e-2/(2.6**4),1e-2), moms=(0.8,0.7))
# Save to possibly retrain again
learn.save(MODELS/'headline-third-run')
# Load again
learn.load(MODELS/'headline-third-run')
pass
learn.unfreeze()
learn.fit_one_cycle(3, slice(1e-2/(2.6**4),1e-2), moms=(0.8,0.7))
learn.save(MODELS/'headline-final')
learn.predict("Kupi mobitel, najbolji popust")
learn.export(MODELS/'headline-learner.pkl')
| Campaign-Headline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#linearregreesion
import numpy as np
import pandas as pd
import pandas_profiling
import matplotlib.pyplot as plt
from scipy import stats
import seaborn as sns
from sklearn.linear_model import LinearRegression
a=np.array([[1,2,3,4,5,6,7,8]])
b=np.array([-1,2,5,8,11,13,16,19])
lr=LinearRegression()
a.shape
b.shape
a=a.reshape(-1,1)
lr.fit(a,b)
lr.score(a,b)
lr.predict([[10]])#
lr.intercept_
lr.coef_
#training and testing and validation
Train = pd.read_csv('/home/manikanta/Documents/dataset/csvfile/sir_train.csv')
Test = pd.read_csv('/home/manikanta/Documents/dataset/csvfile/sir_test.csv')
Train.head()
Test.head()
Train = Train.dropna()
Test = Test.dropna()
sns.pairplot(Train)
sns.pairplot(Test)
lr=LinearRegression()
Train.columns
Test.columns
x_train=Train[['x']]
y_train=Train['y']
x_test = Test[['x']]
y_test = Test['y']
lr=LinearRegression(normalize=True)
lr.fit(x_train,y_train)
Train_Score =lr.score(x_train,y_train)
Train_Score
Test_Score= lr.score(x_test,y_test)
Test_Score
new_prediction=lr.predict(x_test)
new_prediction
d=pd.DataFrame({'new_pre':new_prediction,'actual_data':y_test})
from sklearn.metrics import mean_absolute_error,mean_squared_error,r2_score
mean_squared_error(new_prediction,y_test)
mean_absolute_error(new_prediction,y_test)
##model score
r2_score(new_prediction,y_test)
from sklearn.model_selection import cross_val_score
cv = cross_val_score(lr,x_train,y_train,cv = 20)
cv
np.mean(cv)
np.max(cv)
np.min(cv)
sns.lmplot(x='actual_data',y='new_pre',data=d)
| Linear_Regrssion/linearRegression_trining_and_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <div style="align:center; font-size:large">
# <img src='img/logo-igm.png' width=50%>
# </div>
#
# **L1 Mathématiques - L1 Informatique — Semestre 1**
#
# # Projet 1 - Cours du 6/12/2021
# + [markdown] slideshow={"slide_type": "slide"}
# ## Contenu du cours (indicatif)
#
# Cours 1
# - Présentation de l'UE
# - Présentation du sujet
# - Conseils de travail
# - Complément de cours : itérables
# - Questions / réponses
# + [markdown] slideshow={"slide_type": "subslide"}
# Cours 2
# - Complément de cours : types mutables et immutables
# - Complément de cours : `fltk`
# - Compléments et exercices selon demande
# - Questions / réponses
# + [markdown] slideshow={"slide_type": "subslide"}
# Cours 3
# - Conseils de rédaction du rapport
# - Conseils pour la soutenance
# - Compléments et exercices selon demande
# - Questions / réponses
# + [markdown] slideshow={"slide_type": "slide"}
# ## Présentation de l'UE
#
# - Organisation
# - Consignes
# - Évaluation
# - Réponse aux questions
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Organisation
#
# Calendrier :
# - début lundi 6 décembre
# - fin : vendredi 7 janvier
# - soutenances : semaine du 10 janvier (sauf changement)
# + [markdown] slideshow={"slide_type": "subslide"}
# Rythme :
# - cours en amphi les lundis 6/12, 13/12 et 3/1
# - TP les mêmes semaines, en salles 102, 104 et 106
# - 2h de TP encadré ; 2h de TP en autonomie
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Consignes
#
# Réalisation d'un projet de programmation
# - par équipes de 2 ou 3 du **même TP**
# - pas d'équipes de 1 ni de 4 !
# - en utilisant `fltk` pour la partie graphique
# + [markdown] slideshow={"slide_type": "subslide"}
# Constitution des équipes
# - renseigner son groupe de TP sur elearning
# - former une équipe de 2 ou 3 personnes
# - choisir un numéro d'équipe et y inscrire chaque participant
# + [markdown] slideshow={"slide_type": "subslide"}
# Rédaction d'un rapport
# - 5 pages environ, hors page de garde, hors annexes (images, code...)
# - contenu précisé dans un prochain amphi
# + [markdown] slideshow={"slide_type": "subslide"}
# Soutenance
# - 15 mn environ
# - présentation + questions du jury
# - tout le monde doit savoir répondre à toutes les questions
# + [markdown] slideshow={"slide_type": "slide"}
# ## Présentation du sujet
#
# [<NAME>](https://igm.univ-mlv.fr/~ameyer/pr1/)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Conseils de travail
#
# 1. Démarrez le projet tout de suite (vous avez seulement un mois, vacances comprises !)
# 2. Répartissez les responsabilités
# - Qui est **responsable** de quelle tâche (1 à 4 + rapport) ?
# - Qui garde la version la plus à jour du code ?
# - Qui surveille l'avancement du projet et les délais ?
# 3. Décidez d'une méthode de travail
# - Comment communique-t-on entre équipiers (Discord, Zoom, Signal, téléphone...) ?
# - Comment avance-t-on sur le code (ensemble ou chacun.e de son côté) ?
# + [markdown] slideshow={"slide_type": "subslide"}
# Pour pouvoir programmer en même temps sur **différentes** parties du code :
# - Se mettre d'accord sur la représentation des objets
# - Utiliser des fonctions avec une spécification claire (notion d'**interface**)
# - Mettre le code en commun **régulièrement** et **TESTER !!!**
# + [markdown] slideshow={"slide_type": "subslide"}
# Exemple :
# - Tu écris les fonctions qui créent et modifient l'état du jeu
# - J'écris une fonction qui reçoit l'état du jeu et le dessine avec `fltk`
#
# Chacun peut avancer de son côté, puis on met en commun
# + [markdown] slideshow={"slide_type": "subslide"}
# Pour pouvoir programmer en même temps sur les **mêmes** parties du code :
# - Être ensemble tout le temps pour travailler (en Ader, à la bibliothèque, chez soi...)
# - OU partager son écran en ligne (avec Discord, Zoom ou autre)
# - OU utiliser un éditeur collaboratif en ligne comme https://replit.com/
# + [markdown] slideshow={"slide_type": "slide"}
# ## Complément de cours : Itérables
# + [markdown] slideshow={"slide_type": "-"}
# Le [glossaire Python](https://docs.python.org/fr/3/glossary.html#term-iterable) définit **itérable** comme:
#
# > Objet capable de renvoyer ses éléments un à un.
#
# Par exemple, les `list` et les `str` sont des *itérables*, ainsi que les `tuple` et les objets renvoyés par la fonction `range`.
# + [markdown] slideshow={"slide_type": "subslide"}
# Le principal intérêt d'un *itérable* est qu'on peut l'utiliser dans une boucle `for` pour parcourir ses éléments un par un. Par exemple pour une liste :
# + slideshow={"slide_type": "-"}
lst = ['a', 'b', 'c']
for element in lst:
print(element)
# + [markdown] slideshow={"slide_type": "subslide"}
# Vous avez déjà rencontré des boucles de cette forme utilisant la fonction `range` :
# + slideshow={"slide_type": "-"}
for i in range(3):
print(i)
# + [markdown] slideshow={"slide_type": "subslide"}
# Cela fonctionne parce que `range` renvoie un objet **itérable** (de type... `range`)
# + slideshow={"slide_type": "-"}
type(range(10))
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :** Écrire une fonction `affiche` qui reçoit un itérable `elems` et affiche ses éléments (un par ligne) à l'aide d'une boucle `for`, et la tester avec divers itérables (de type `list`, `tuple`, `str` et `range`)
# + slideshow={"slide_type": "-"}
def affiche(elems):
for elem in elems:
print(elem)
affiche((1, 2, 3, 4))
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :** Écrire une fonction `appartient` qui reçoit un itérable `elems` et une valeur `x` et renvoie `True` si `x` est un élément de `elems` et `False` sinon, et la tester avec divers itérables (de type `list`, `tuple`, `str` et `range`).
#
# *Remarque : cette fonctionnalité existe en Python (opérateur `in`)*
#
# *Contraintes :* boucle `for`, pas de `range`, pas de test `x in elems`
# + slideshow={"slide_type": "-"}
def appartient(elems, x):
for elem in elems:
if elem == x:
return True
return False # SURTOUT PAS dans la boucle !!!!!
appartient("bonjour", "jour")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Que peut-on faire d'autre avec un itérable ?
#
# N'importe quel itérable permet aussi, entre autres :
# - de créer une liste avec `list(elems)` ou un tuple avec `tuple(elems)`
# - de tester l'appartenance d'un élément avec `x in elems` et `x not in elems`
# - *hors programme:* d'utiliser toute fonction recevant en paramètre un ou plusieurs itérables (comme `sum`, `min`, `max`, `any`, `all`, `zip`, `map`, `filter`, `sorted`, etc.)
# - *hors programme:* d'écrire une *mutation de liste*
# + slideshow={"slide_type": "subslide"}
list("bonjour")
# + slideshow={"slide_type": "subslide"}
tuple('abc')
# + slideshow={"slide_type": "subslide"}
7 in range(10)
# + slideshow={"slide_type": "subslide"}
phrase = 'We are the knights who say "Ni!"'
mots = phrase.split()
[len(mot) for mot in mots]
# + slideshow={"slide_type": "subslide"}
list(zip(range(1, 4),
['un', 'deux', 'trois'],
['one', 'two', 'three'],
['eins', 'tzwei', 'drei']))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Intervalles d'entiers : `range`
#
# https://docs.python.org/fr/3/library/stdtypes.html#ranges
#
# La fonction `range` fabrique des intervalles d'entiers:
#
# * `range(i)` : entiers de `0` à `i-1`
# * `range(i, j)` : entiers de `i` à `j-1`
# * `range(i, j, k)` : entiers de `i` à `j-1` par
# pas de `k`
# + [markdown] slideshow={"slide_type": "subslide"}
# Len opérations autorisées sur un `range` `r` sont :
#
# * `x in r, x not in r`
# * `r[i]`
# * `len(r), min(r), max(r), r.index(x), r.count(x)`
# * `list(r)` (conversion en liste)
# + [markdown] slideshow={"slide_type": "subslide"}
# Toutes les autres opérations sont interdites sur un range (car un range est non mutable, contrairement à une liste). Plus précisément, sont interdits :
#
# - Concaténation, répétition, affectation d'élément
# - Autres méthodes de listes (append, etc.)
# + [markdown] slideshow={"slide_type": "subslide"}
# Caractéristique importante :
#
# Un `range` n'est pas construit entièrement en mémoire. Au lieu de cela, ses éléments sont fabriqués « à la demande ». Donc, par exemple, un appel à `range(10000)` ne prend pas plus de place en mémoire (ni de temps à construire) que `range(2)`.
# + [markdown] slideshow={"slide_type": "subslide"}
# Utilisation fréquente des `range` : parcours des indices d'une liste
# + slideshow={"slide_type": "-"}
lst = ['a', 'b', 'c']
for i in range(len(lst)):
print(i, '->', lst[i])
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :** Écrire une fonction `premier_indice(elems, x)` qui renvoie la première position de `x` dans la liste `elems` (ou `None` si `x` n'est pas dans `elems`) en utilisant une boucle `for` et la fonction `range`
#
# *Remarque : cette fonctionnalité existe en Python (méthode `index`)*
# + slideshow={"slide_type": "-"}
def premier_indice(elems, x):
for i in range(len(elems)):
if elems[i] == x:
return i # ATTENTION ICI !
return None
premier_indice((1, 2, 1, 2, 1), 2)
# + [markdown] slideshow={"slide_type": "-"}
# Pourquoi est-il nécessaire d'utiliser `range` ici ?
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :** Écrire une fonction `dernier_indice(elems, x)` qui renvoie la dernière position de `x` dans la liste `elems` (ou `None` si `x` n'est pas dans `elems`) en utilisant une boucle `for` et la fonction `range`
# + slideshow={"slide_type": "-"}
# + [markdown] slideshow={"slide_type": "slide"}
# ### <img src='img/non-exigible.png' width='35px' style='display:inline'> Un dernier type de parcours d'itérable : `enumerate`
#
# *La connaissance de cette notion n'est pas exigible à l'examen.*
#
# La fonction `enumerate(iterable)` renvoie un nouvel itérable constitué des couples `(i, elem)` contenant un indice et un élément.
# + slideshow={"slide_type": "-"}
list(enumerate(['a', 'b', 'c']))
# + [markdown] slideshow={"slide_type": "subslide"}
# Reprenons un exemple précédent :
# + slideshow={"slide_type": "-"}
lst = ['a', 'b', 'c']
for i, elem in enumerate(lst):
print(i, '->', elem)
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercice :** Réécrire la fonction `premier_indice(elems, x)` en utilisant une boucle `for` et la fonction `enumerate`
# + slideshow={"slide_type": "-"}
# + [markdown] slideshow={"slide_type": "slide"}
# ## Questions-réponses
#
# À vous...
# -
| pr1-amphi1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nlp
# language: python
# name: nlp
# ---
import sys
FOLDER_PATH = '/home/taindp/Jupyter/custom_dqn'
sys.path.append(FOLDER_PATH)
# %load_ext autoreload
# %autoreload 2
from user_simulator import UserSimulator
from error_model_controller import ErrorModelController
from dqn_agent import DQNAgent
from db_query import DBQuery
# from dqn_agent import get_action
from state_tracker import StateTracker
import pickle, argparse, json, math
from utils import remove_empty_slots
from user import User
import time
import json
from tqdm import tqdm
# %cd $FOLDER_PATH
from db_query import DBQuery
# from tqdm import tqdm
# +
CONSTANTS_FILE_PATH = f'{FOLDER_PATH}/constants.json'
constants_file = CONSTANTS_FILE_PATH
with open(constants_file) as f:
constants = json.load(f)
# -
# Load file path constants
file_path_dict = constants['db_file_paths']
DATABASE_FILE_PATH = file_path_dict['database']
DICT_FILE_PATH = file_path_dict['dict']
USER_GOALS_FILE_PATH = file_path_dict['user_goals']
database= json.load(open(DATABASE_FILE_PATH,encoding='utf-8'))
db_dict = json.load(open(DICT_FILE_PATH,encoding='utf-8'))[0]
user_goals = json.load(open(USER_GOALS_FILE_PATH,encoding='utf-8'))
test_goal = json.load(open('/home/taindp/Jupyter/reference/dqn_ref/data/activity_user_goals_full_slots_multi_req_slots_800.json'))
list_rule= []
for item in test_goal:
rule = list(item.values())[0]
if rule:
list_rule.append(list(rule.keys())[0])
set(list_rule)
rule_requests=['name_activity', 'type_activity', 'holder', 'time', 'city', 'district', 'ward', 'name_place', 'street',
'reward', 'contact', 'register', 'works', 'joiner']
set(rule_requests) == set(list_rule)
list_rule= []
for item in test_goal:
rule = list(item.values())[0]
if rule:
list_rule.append(list(rule.keys())[0])
set(list_rule)
| check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="gzCv2vbt1HnJ"
# # Nikkei High Dividend Yield 50 Analysis with Portfolio Optimizer
#
#
# + [markdown] id="4icbKzYm1HnL"
# ## Initialization
# + [markdown] id="yxrvVg1O1HnM"
# If you are using CoLab, first install the dependency PyPortfolioOpt
# + id="v8m3TiH91HnM" outputId="1a9474ed-9750-4dbe-84d8-a0284303ef3e" colab={"base_uri": "https://localhost:8080/"}
# !pip install git+https://github.com/robertmartin8/PyPortfolioOpt.git
# + [markdown] id="xZ5S5Vao1HnO"
# Then get our PortfolioOptimizer library and the necessary datasets from our repository
# + id="ikhEw5mi1HnO" outputId="a40760d6-7792-4953-8146-21616b7f33ab" colab={"base_uri": "https://localhost:8080/"}
# !wget https://raw.githubusercontent.com/cartasuzuki/phynance/master/PortfolioOptimizer.py
# + [markdown] id="9ndVE59o1HnO"
# Import libraries
# + id="do5vma0F1HnP"
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from PortfolioOptimizer import PortfolioOptimizer
# + id="0dXct09W1HnP"
stock_symbols = pd.read_csv('https://raw.githubusercontent.com/cartasuzuki/phynance/master/datasets/nikkei_high_dividend_yield_50_weight_en.csv')
# + id="-CEegZu01HnP"
# + [markdown] id="w4Bqx_Qp1HnQ"
# Once we have loaded our stock symbols dataset we can either use the prices in the csv file or download prices from alphadvantage to get updated data.
# + [markdown] id="2J4qcM4w1HnQ"
# ### Filter out some stocks from the index
# + [markdown] id="6ffhJmYY1HnQ"
# Chose a minimum div/yield. Set to 0 if you want to use all stocks in the index
# + id="d7en_0Hw1HnR"
min_yield = 3.5
# + id="O5uAu8RU1HnR"
selected_stocks = stock_symbols[stock_symbols['Dividend']>min_yield]
# + [markdown] id="Z2U22dpi1HnR"
# Create a filter string (highDivString) to be used later to filter stocks with lower yield than min_yield
# + id="r4doUNhH1HnR"
highDivString = selected_stocks['Code'].values.astype(int)
highDivString = highDivString.astype(str)
string = '.TOK'
#highDivString = [x + string for x in highDivString]
# + [markdown] id="cw30vyhi1HnS"
# ### Method 1: use csv file
# + [markdown] id="vdDAWM8Q1HnS"
# Read the stock prices from the csv provided in our repository
# + id="fSFd0yoM1HnS"
stocks = pd.read_csv('https://raw.githubusercontent.com/cartasuzuki/phynance/master/datasets/nikkei50.csv', index_col= ['timestamp'], parse_dates= ['timestamp'])
#stocks = pd.read_csv('https://raw.githubusercontent.com/cartasuzuki/phynance/master/datasets/nikkei_high_dividend_yield_50_prices.csv', index_col= ['timestamp'], parse_dates= ['timestamp'])
# + id="2mWeZhgE21vd" outputId="b5bdc085-7305-438d-ef3d-06181a67db67" colab={"base_uri": "https://localhost:8080/"}
highDivString
# + [markdown] id="OE1m75wU1HnT"
# If you want higher yield filter out using the filter previously created
# + id="0vx4HQQJ1HnT"
stocks = stocks[highDivString]
# + id="qunr5O1Z64T0"
stocks = stocks.replace(',','', regex=True)
cols=[i for i in stocks.columns if i not in ["timestamp"]]
for col in cols:
stocks[col]=pd.to_numeric(stocks[col])
# + id="BN_0kZZR68fn" outputId="af025598-062f-4636-d74d-2e7b3ddd00e9" colab={"base_uri": "https://localhost:8080/", "height": 302}
stocks.head()
# + id="3WZ6fsy67QJI"
# + id="g3VmBvGv1HnU"
# + [markdown] id="qj7NMn6W1HnV"
# ## Portfolio Optimization
# + id="RCkbiX_p1HnV"
weights, sharpe, ret = PortfolioOptimizer.optimize_portfolio(stocks,0,0.2)
# + id="DuLd3SwB1HnV" outputId="248970aa-0c93-4345-e8d0-69dc08d5a09b" colab={"base_uri": "https://localhost:8080/", "height": 384}
PortfolioOptimizer.print_portfolio_result(weights, sharpe, ret)
PortfolioOptimizer.portfolioAsPieChart(weights)
# + [markdown] id="goSiHsF41HnW"
# Average yield
# + id="4KbedtcX1HnW" outputId="1346436f-1375-471a-9e66-385c8d8f9153" colab={"base_uri": "https://localhost:8080/"}
selected_stocks['Dividend'].mean()
# + [markdown] id="GanME8Xr1HnW"
# Remove 0s and TOK string.
# + id="g6kFQ7IK1HnW" outputId="26c8ce45-4d09-4fce-d797-583e85ea17c9" colab={"base_uri": "https://localhost:8080/"}
www ={x:y for x,y in weights.items() if y>0.001}
portfolio =list(www.keys())
portfoliovalues = list(www.values())
portfolio
# + [markdown] id="7l-lnC161HnW"
# ### Resulting Portfolio
# + id="zX5wm4m_1HnW"
selected_stocks = selected_stocks[selected_stocks['Code'].isin(portfolio)]
# + id="-9AcJwNZ1HnX" outputId="c115a33e-ea86-44f1-a612-fdbc9925257a" colab={"base_uri": "https://localhost:8080/", "height": 206}
selected_stocks
# + id="KxGMp9H91HnX" outputId="e030623a-c2b5-4748-e5c6-93a8cb8b7b84" colab={"base_uri": "https://localhost:8080/"}
Div_Yield = np.average(selected_stocks['Dividend'], weights=selected_stocks['Weight'])
round(Div_Yield, 2)
# + id="pl7CwvrY1HnX"
| NikkeiHighDiv50_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
# +
import pandas as pd
import numpy as np
import scipy
import scipy.sparse
import scipy.stats
import os
import scipy.io as sio
import regex as re
from collections import Counter, defaultdict
import sys
import gzip
def distance(astring, bstring) :
distance = 0
limit = len(astring)
diff = len(bstring) - len(astring)
if len(bstring) < len(astring) :
limit = len(bstring)
diff = len(astring) - len(bstring)
for i in range(limit) :
if astring[i] != bstring[i] :
distance += 1
return distance + diff
# +
r1_rna = 'Undetermined_S0_R1_001.fastq.gz'
#r2_rna = 'Undetermined_S0_R2_001.fastq.gz'
r_indx = 'Undetermined_S0_I1_001.fastq.gz'
proximal_regex = re.compile(r"(AAAAAAAAAAAAAAAAAAAA){s<=3}")
proximal_regex_prefix = re.compile(r"(AAA)(AAAAAAAAAAAAAAAAA){s<=3}")
wildtype_downstream_regex = re.compile(r"(GATGTCTCGTGATCTGGTGT){s<=2}")
upstream_regex = re.compile(r"(CAATTCTGCT){s<=2}[ACGTN]{40}(CTAAAATATA){s<=2}")
downstream_regex = re.compile(r"(AGTATGAAAC){s<=2}[ACGTN]{20}(ACCCTTATCC){s<=2}")
seq_regex = re.compile(r"(CAATTCTGCT){s<=2}[ACGTN]{40}(CTAAAATATA){s<=2}.*(AGTATGAAAC){s<=2}[ACGTN]{20}(ACCCTTATCC){s<=2}")
# +
f1 = gzip.open(r1_rna,'rt')
i1 = gzip.open(r_indx, 'rt')
#f2 = open(r2_rna,'r')
head, seq, pr, q, head2, seq2, pr2, q2, headi, seqi, pri, qi = ({} for i in range(12))
count = 0
total_proximal_rna_count = 0
num_upstream_region_extractions = 0
num_downstream_region_extractions = 0
print('Processing RNA reads.')
out = open('tomm5_rna_polyatail_3errors_test1.csv','w')
out.write('upstream_seq,downstream_seq,seq,umi,polya,polya_prefixed,is_proximal\n')
while True:
head = f1.readline()[:-1]
seq = f1.readline()[:-1]
pr = f1.readline()[:-1]
q = f1.readline()[:-1]
headi = i1.readline()[:-1]
seqi = i1.readline()[:-1]
pri = i1.readline()[:-1]
qi = i1.readline()[:-1]
if len(q) == 0:
break # End of File
upstream_flank = re.search(upstream_regex, seq)
downstream_flank = re.search(downstream_regex, seq[70:220])
both_flank = re.search(seq_regex, seq)
if upstream_flank is not None:
num_upstream_region_extractions += 1
upstream_flank_seq = upstream_flank.group()
proximal_test_outcome = re.search(proximal_regex, seq)
umi = seqi
polya_pos = -1
polya_pos_prefixed = -1
downstream_flank_seq = ''
is_prox = 0
if downstream_flank is not None :
num_downstream_region_extractions += 1
downstream_flank_seq = downstream_flank.group()
elif proximal_test_outcome is not None :
total_proximal_rna_count += 1
polya_pos = proximal_test_outcome.start()
is_prox = 1
prefixed_test_outcome = re.search(proximal_regex_prefix, seq)
if prefixed_test_outcome is not None :
polya_pos_prefixed = prefixed_test_outcome.start()
both_flank_seq = ''
if both_flank is not None :
both_flank_seq = both_flank.group()
out.write(upstream_flank_seq)
out.write(',' + downstream_flank_seq)
out.write(',' + both_flank_seq)
out.write(',' + umi)
out.write(',' + str(polya_pos))
out.write(',' + str(polya_pos_prefixed))
out.write(',' + str(is_prox))
out.write('\n')
if count % 1000000 == 0:
print('Count: ' + str(count))
print('Number of upstream regions extracted: ' + str(num_upstream_region_extractions))
print('Number of downstream regions extracted: ' + str(num_downstream_region_extractions))
print(str(total_proximal_rna_count) + ' proximal RNA reads')
count += 1
print('COMPLETE')
print('Number of upstream regions extracted: ' + str(num_upstream_region_extractions))
print('Number of downstream regions extracted: ' + str(num_downstream_region_extractions))
print(str(total_proximal_rna_count) + ' proximal RNA reads')
out.close()
f1.close()
#f2.close()
i1.close()
# -
# +
proximal_regex = re.compile(r"(AAAAAAAAAAAAAAAAAAAA){s<=3}")
test_re = re.search(proximal_regex, 'TTTAAGTTTTTTTGATAGTAAGGCCCATTACCTGAGGCCGCAATTCTGCTTGTTAAGAACAATCCCAGTTCTGGTAACTGACCTTCAAAGCTAAAATATAAAACTATTTGGGAAGTATGAAAAAAAAAAAAAAAAAAAAACCGGTTTCCGGATGGGGAGGGCGCCCGGGGGGGGGGCGGGCCCGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG')
print(test_re.start())
# +
proximal_regex = re.compile(r"(AAA)(AAAAAAAAAAAAAAAAA){s<=3}")
test_re = re.search(proximal_regex, 'TTTAAGTTTTTTTGATAGTAAGGCCCATTACCTGAGGCCGCAATTCTGCTTGTTAAGAACAATCCCAGTTCTGGTAACTGACCTTCAAAGCTAAAATATAAAACTATTTGGGAAGTATGAAAAAAAAAAAAAAAAAAAAACCGGTTTCCGGATGGGGAGGGCGCCCGGGGGGGGGGCGGGCCCGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGG')
print(test_re.start())
| data/random_mpra/individual_library/tomm5/unprocessed_data/tomm5_rna_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Segmentation of images from the UDIAT dataset
#
# UDIAT is a publically available dataset of breast mass ultrasound images. For the details, we suggest to read the original paper by [Yap et al., IEEE JBHI paper](https://doi.org/10.1109/JBHI.2017.2731873). The dataset can be downloaded at the project's [website](http://goo.gl/SJmoti). After downloading, extract the files to 'data/UDIAT/'.
# +
import numpy as np
import matplotlib.pyplot as plt
import cv2
from os import listdir
from seg_lib import dice_coef_np, selective_unet
# +
images_udiat, rois_udiat = [], []
dsize = (224, 224)
path = 'data/udiat/'
img_file = listdir(path+'original')
for i, file in enumerate(img_file):
img = cv2.imread(path+'original/'+file, 0)
roi = cv2.imread(path+'GT/'+file, 0)/255
img = cv2.resize(img, dsize, interpolation=cv2.INTER_CUBIC)
roi = cv2.resize(roi, dsize, interpolation=cv2.INTER_NEAREST)
images_udiat.append(img)
rois_udiat.append(roi)
images_udiat = np.array(images_udiat, dtype=np.float32)
images_udiat = np.expand_dims(images_udiat, 3)
rois_udiat = np.array(rois_udiat, dtype=np.int16)
rois_udiat = np.expand_dims(rois_udiat, 3)
# -
# [SK-U-Net weights](https://drive.google.com/file/d/1cVEAcoyA5wLHxoCtOAIX2bKusxJsBvYM/view?usp=sharing) (via Google Drive)
model = selective_unet()
model.load_weights('models/skunet_weights.h5')
rois_predicted = model.predict(images_udiat).squeeze().round()
# +
dices = np.zeros(rois_predicted.shape[0])
for i in range(rois_predicted .shape[0]):
dices[i] = dice_coef_np(rois_predicted[i], rois_udiat[i])
# -
# Results are slightly different compared to our paper. Originally, we additionally preprocessed the images in Matlab.
print('Dice scores | mean:', np.mean(dices).round(3), 'median:', np.median(dices).round(3), 'mean Dice>0.5:', np.mean(dices[dices>0.5]).round(3))
| UDIAT example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Belief Propagation from Geo-Located Imagery
# +
# If new server on Descartes Labs, need to install rioxarray
try: import rioxarray
except:
# %pip install rioxarray
import demo_functions as df
# -
# __________
# Let's begin with the input parameters. These include the label file, confidence in the labels and the data types we will use. Once we confirm the data types we will be asked for paths to the files containing the imagery. Post-event must be provided but pre-event is optional. If a pre-event image is provided the data used will be the difference between the images which contains more information than the post event image alone.
inputs = df.parameter_input()
# ______
# Now let's load up the map of our ground labels and define an area for the model. Then below the map we'll pick the model parameter to run on the data from the selected area. If we wish to group classes together we will also be offered some clustering options.
parameters = df.model_parameters(inputs)
# ________
# Now we have all the parameters for the model, let's import and classify the data according to our selections. If you have already imported the data and just adjusted the model parameters then just re-classify rather than re-importing.
# + jupyter={"source_hidden": true}
import helper_functions as hf
import imports as ip
import demo_functions as demo
def import_data(v):
# Retrieve file locations from inputs
for j in range(len(v['dataTypes'])):
try: v['preFile'+str(j)], v['postFile'+str(j)] = [i.value for i in v['bxfile'+str(j)].trait_values()['children'][1::2]]
except KeyError: raise KeyError('Please make sure you have confirmed the data types.')
for i in v.keys(): globals()[i] = v[i] # Retrieve variables to use
# Reproject Data if necessary
v = demo.reproject_data(v)
# Import Files
print("------Importing Data Files---------")
# Import first data type
df, crop = ip.img_to_df(postFile0, testPoly, crs=crs)
if preFile0:
preDf, _ = ip.img_to_df(preFile0, testPoly, crs=crs)
df -= preDf
# Import other data types
if len(dataTypes) > 1:
crop.rio.to_raster("croptemp.tif")
for i in range(1, len(dataTypes)):
ip.resample_tif(globals()['postFile'+str(i)], testPoly, 'posttemp'+str(i)+'.tif')
globals()['dataArray'+str(i)] = ip.tif_to_array('posttemp'+str(i)+'.tif', 'resample')
if globals()['preFile'+str(i)]:
ip.resample_tif(globals()['preFile'+str(i)], testPoly, 'pretemp'+str(i)+'.tif')
globals()['dataArray'+str(i)] -= ip.tif_to_array('pretemp'+str(i)+'.tif', 'resample')
ip.del_file_endings(".", "temp*.tif")
# Concatenate data types
data = df.copy()
for j in range(1, len(dataTypes)):
da = globals()['dataArray'+str(1)]
for k in range(min(da.shape)):
data[str(dataTypes[j])+str(k)]=da.reshape(min(da.shape),-1)[k]
data.dropna(inplace=True)
print("------Finished Data Import---------")
typesUsed = [list(df.columns.values)]
for j in range(1,len(dataTypes)): typesUsed.append(list(data.columns[[dataTypes[j] in str(i) for i in data.columns]]))
v.update({'data':data, 'typesUsed':typesUsed})
return v
import os
import random
import importlib
import ground_truth
import numpy as np
import pandas as pd
import rasterio as ro
import rioxarray as rxr
import geopandas as gpd
import helper_functions as hf
import shapely.geometry as sg
from rasterio.io import MemoryFile
from rasterio.enums import Resampling
from rasterio.windows import from_bounds
from rasterio.warp import calculate_default_transform, reproject, Resampling
def img_to_df(file, poly=False, crs=False, label='img', columns=False, crsPoly='epsg:4326', verbose=True):
# Import raster
img = rxr.open_rasterio(file, masked=True).squeeze()
# Crop image if polygon supplied
if poly:
_, extent = hf.get_extent(poly, crsPoly=crsPoly, crs=crs)
img = img.rio.clip(extent.geometry.apply(sg.mapping))
named = img.rename('img')
# Convert to dataframe
xm, ym = np.meshgrid(np.array(named.coords['x']), np.array(named.coords['y']))
mi = pd.MultiIndex.from_arrays([ym.flatten(),xm.flatten()],names=('y','x'))
size = min(named.shape) if len(named.shape) > 2 else 1
df = pd.DataFrame(named.data.reshape(size,-1).transpose(), index=mi)
if verbose: print(file+" read completed.")
return df, named
# -
imports = import_data(parameters)
classified = classify_data(imports)
# ____________
# OK, the data is formatted the model parameters are all checked. Let's build the graph of nodes & edges and run the belief propagation!
# +
for i in classified.keys(): globals()[i] = classified[i]
initial = classified['initial']
trainSplit = bxNodes.trait_values()['children'][3].value
confidence = list(bxConf.trait_values()['children'][1].value)
neighbours = [i.value for i in bxEdges.trait_values()['children'][1].trait_values()['children']]
adjacent, geoNeighbours = [i.value for i in bxAdjacent.trait_values()['children'][1::2]]
# Split pixels in to train and test sets
X_train, X_test, y_train, y_test = hf.train_test_split(labelsUsed, cn, hf.get_polygon(testPoly, conv=True), testSplit=(1-(trainSplit/100)))
# Create nodes
nodes = hf.create_nodes(initial, X_train)
import numpy as np
summary = nodes.groupby(cn).size()
equivUse = True
if equivUse:
equiv = gpd.GeoDataFrame()
for i in summary.index.values:
equiv = equiv.append(nodes[nodes[cn] == i][0:min(summary)])
equiv = equiv.append(nodes[[np.isnan(x) for x in nodes[cn]]])
nodes=equiv.copy()
initial = initial.loc[nodes.index.values].reset_index()
# Assign prior beliefs from assessments
priors = hf.prior_beliefs(nodes, beliefColumns = initial.columns[-nClasses:], beliefs=confidence, classNames=classNames, column = cn)
classes = classNames
d = dict(enumerate(classes))
gdf = gpd.sjoin(initial, X_test, how='left', op='within').dropna(subset=[cn])
summary = gdf.groupby(cn).size()
equivTest = True
if equivTest:
equiv = gpd.GeoDataFrame()
for i in summary.index.values:
equiv = equiv.append(gdf[gdf[cn] == i][0:min(summary)])
equiv = equiv.append(gdf[[np.isnan(x) for x in gdf[cn]]])
y_true = equiv[cn]
y_true_l = list(equiv[cn])
else: y_true = gdf[cn]
# + jupyter={"outputs_hidden": true}
import sklearn as skl
import numpy as np
# Edge creation
measures = [0,0,0,0,0,0,0]
from tqdm import tqdm
scores = []
num = 11
for geoNeighbours in tqdm(range(num)):
for ed1 in tqdm(range(num)):
for ed2 in tqdm(range(num)):
neighbours = [ed1,ed2]
if all(values is 0 for values in neighbours) and (geoNeighbours is 0):
edges, beliefs = [], priors
else:
edges = hf.create_edges(nodes, adjacent=adjacent, geo_neighbours=geoNeighbours, values=typesUsed, neighbours=neighbours)
beliefs, _ = nc.netconf(edges,priors,verbose=False,limit=1e-3)
# Get y_true vs y_pred for test set
y_pred = skl.preprocessing.normalize(beliefs[y_true.index], norm='l1')
yp_clf = np.argmax(y_pred, axis=1)
pred_clf = [i for i in yp_clf]
f1 = skl.metrics.f1_score(y_true_l, pred_clf,average='weighted',zero_division=0)
a = skl.metrics.accuracy_score(y_true_l, pred_clf)
r = skl.metrics.recall_score(y_true_l, pred_clf,average='weighted',zero_division=0)
log_loss = skl.metrics.log_loss(y_true_l, y_pred, labels=[0,1])
measures = np.vstack((measures, [geoNeighbours, ed1, ed2, f1,a,r,log_loss]))
# -
np.savetxt('results/beirutedges.csv', measures, delimiter=',')
output = run_bp(classified,limit=1e-4)
# _____
# Now let's use the test set to evaluate the effectiveness of the model.
plots = evaluate_output(output)
# Want to save the plot? Run the cell below. If you want to specify a location replace the False boolean with the filepath.
df.save_plot(plots, location=False)
import geopandas as gpd
import helper_functions as hf
import netconf as nc
def run_bp(v, limit=1e-5):
# Retrieve data from inputs
for i in v.keys(): globals()[i] = v[i]
initial = v['initial']
trainSplit = bxNodes.trait_values()['children'][3].value
confidence = list(bxConf.trait_values()['children'][1].value)
neighbours = [i.value for i in bxEdges.trait_values()['children'][1].trait_values()['children']]
adjacent, geoNeighbours = [i.value for i in bxAdjacent.trait_values()['children'][1::2]]
# Split pixels in to train and test sets
X_train, X_test, y_train, y_test = hf.train_test_split(labelsUsed, cn, hf.get_polygon(testPoly, conv=True), testSplit=(1-(trainSplit/100)))
# Create nodes
nodes = hf.create_nodes(initial, X_train)
import numpy as np
summary = nodes.groupby(cn).size()
equivUse = True
if equivUse:
equiv = gpd.GeoDataFrame()
for i in summary.index.values:
equiv = equiv.append(nodes[nodes[cn] == i][0:min(summary)])
equiv = equiv.append(nodes[[np.isnan(x) for x in nodes[cn]]])
nodes=equiv.copy()
initial = initial.loc[nodes.index.values].reset_index()
# Assign prior beliefs from assessments
priors = hf.prior_beliefs(nodes, beliefColumns = initial.columns[-nClasses:], beliefs=confidence, classNames=classNames, column = cn)
if all(values is 0 for values in neighbours) and (geoNeighbours is 0):
edges, beliefs = [], priors
else:
# Create edges
edges = hf.create_edges(nodes, adjacent=adjacent, geo_neighbours=geoNeighbours, values=typesUsed, neighbours=neighbours)
# Run belief propagation
beliefs, _ = nc.netconf(edges,priors,verbose=True,limit=limit)
v.update({'trainSplit':trainSplit, 'confidence':confidence, 'neighbours':neighbours, 'adjacent':adjacent, 'geoNeighbours':geoNeighbours, 'X_train':X_train, 'X_test':X_test, 'nodes':nodes, 'priors':priors, 'edges':edges,'beliefs':beliefs,'initial':initial})
return v
# +
import sklearn as skl
import plotting as pl
def evaluate_output(v):
for i in v.keys(): globals()[i] = v[i]
# Get y_true vs y_pred for test set
y_true, y_pred = get_labels(initial, X_test, beliefs, column=cn)
# Classification metrics
true_clf, pred_clf = hf.class_metrics(y_true, y_pred, classes=usedNames, orig=unique)
fig, axs = pl.create_subplots(1,2, figsize=[12,5])
# Confusion matrix
axs = pl.confusion_matrix(axs, true_clf, pred_clf, usedNames)
# Cross entropy / Confidence metrics
if nClasses == 2: axs = cross_entropy_metrics(axs, y_true, y_pred[:,1].reshape(-1,1), usedNames)
else: axs[1] = pl.cross_entropy_multiclass(axs[1], true_clf, y_pred, usedNames)
pl.show_plot()
v.update({'y_true':y_true, 'y_pred':y_pred, 'true_clf':true_clf, 'pred_clf':pred_clf, 'fig':fig})
return v
def cross_entropy_metrics(axs, y_true, y_pred, classes, dmgThresh=0.5, initBelief=0.5):
try: ax = axs[1]
except: ax = axs
try:
int(classes[0]), int(classes[1])
label1, label2 = 'True class '+str(classes[0]), 'True class '+str(classes[1])
except: label1, label2 = 'True '+str(classes[0]), 'True '+str(classes[1])
p1 = ax.hist(y_pred[(np.array(1-y_true)*y_pred).nonzero()[0]], range = [0,1], bins = 100, label = label1, color = 'g', alpha = 0.5)
if len(classes) > 1:
p2 = ax.hist(y_pred[(np.array(y_true)*y_pred).nonzero()[0]], range = [0,1], bins = 100, label = label2, color = 'r', alpha = 0.5)
# ax.axvline(x=dmgThresh, color='k',linestyle='--', linewidth=1, label='Classification Threshold')
ax.axvline(x=initBelief, color='b',linestyle='--', linewidth=1, label='Initial probability')
log_loss = skl.metrics.log_loss(y_true, y_pred, labels=[0,1])
ax.set_title('Belief Propagation\nCross-entropy loss: {:.3f}'.format(log_loss),size=14)
ax.legend(loc='best',fontsize=12),
try:
int(classes[0]), int(classes[1])
ax.set_xlabel('Class '+str(classes[1])+' Probability',fontsize=12)
ax.text(dmgThresh/2, 0.6, 'Class '+str(classes[0])+'\n Prediction', ha='center', va='center', transform=ax.transAxes,fontsize=12)
ax.text(dmgThresh+(1-dmgThresh)/2, 0.6, 'Class '+str(classes[1])+'\n Prediction', ha='center', va='center', transform=ax.transAxes,fontsize=12)
except:
ax.set_xlabel(str(classes[1])+' Probability',fontsize=12)
ax.text(dmgThresh/2, 0.6, str(classes[0])+'\n Prediction', ha='center', va='center', transform=ax.transAxes)
ax.text(dmgThresh+(1-dmgThresh)/2, 0.6, str(classes[1])+'\n Prediction', ha='center', va='center', transform=ax.transAxes)
ax.set_ylabel('Number of predictions',fontsize=12)
return axs, log_loss
# Get y_true and y_pred for test set oof nodes
def get_labels(init, X_test, beliefs, column, values = False):
gdf = gpd.sjoin(init, X_test, how='left', op='within').dropna(subset=[column])
summary = gdf.groupby(cn).size()
equivTest = True
if equivTest:
equiv = gpd.GeoDataFrame()
for i in summary.index.values:
equiv = equiv.append(gdf[gdf[cn] == i][0:min(summary)])
equiv = equiv.append(gdf[[np.isnan(x) for x in gdf[cn]]])
y_true = equiv[column]
else: y_true = gdf[column]
if values: y_true = y_true.map(values)
y_pred = skl.preprocessing.normalize(beliefs[y_true.index], norm='l1')
return np.array(y_true).reshape(-1,1).astype(type(y_true.values[0])), y_pred
# -
import demo_functions as demo
import imports as ip
def import_data(v):
# Retrieve file locations from inputs
v['dataTypes'] = [i.value.split(' ')[0] for i in v['bxDataTypes'].trait_values()['children'][1:] if len(i.value) > 0]
for j in range(len(v['dataTypes'])):
try: v['preFile'+str(j)], v['postFile'+str(j)] = [i.value for i in v['bxfile'+str(j)].trait_values()['children'][1::2]]
except KeyError: raise KeyError('Please make sure you have confirmed the data types.')
for i in v.keys(): globals()[i] = v[i] # Retrieve variables to use
# Reproject Data if necessary
v = demo.reproject_data(v)
# Import Files
print("------Importing Data Files---------")
# Import first data type
df, crop = ip.img_to_df(postFile0, testPoly, crs=crs)
if preFile0:
preDf, _ = ip.img_to_df(preFile0, testPoly, crs=crs)
df -= preDf
# Import other data types
if len(dataTypes) > 1:
crop.rio.to_raster("croptemp.tif")
for i in range(1, len(dataTypes)):
ip.resample_tif(globals()['postFile'+str(i)], testPoly, 'posttemp'+str(i)+'.tif')
globals()['dataArray'+str(i)] = ip.tif_to_array('posttemp'+str(i)+'.tif', 'resample')
if globals()['preFile'+str(i)]:
ip.resample_tif(globals()['preFile'+str(i)], testPoly, 'pretemp'+str(i)+'.tif')
globals()['dataArray'+str(i)] -= ip.tif_to_array('pretemp'+str(i)+'.tif', 'resample')
ip.del_file_endings(".", "temp*.tif")
# Concatenate data types
data = df.copy()
for j in range(1, len(dataTypes)): data[dataTypes[j]]=globals()['dataArray'+str(j)].flatten()
data.dropna(inplace=True)
print("------Finished Data Import---------")
typesUsed = [list(df.columns.values)]
for j in range(1,len(dataTypes)): typesUsed.append(list(data.columns[[dataTypes[j] in str(i) for i in data.columns]]))
v.update({'data':data, 'typesUsed':typesUsed})
return v
output['initial'].loc[output['nodes'].index.values].reset_index()
# +
import ipyleaflet as ipl
import sys
# Converting gdf columns to GeoData for plotting
def to_geodata(gdf, color, name='Data', fill=0.7):
plotGdf = ipl.GeoData(geo_dataframe = gdf,
style={'color': color, 'radius':2, 'fillColor': color, 'opacity':fill+0.1, 'weight':1.9, 'dashArray':'2', 'fillOpacity':fill},
hover_style={'fillColor': 'white' , 'fillOpacity': 0.2},
point_style={'radius': 3, 'color': color, 'fillOpacity': 0.8, 'fillColor': color, 'weight': 3},
name = name)
return plotGdf
# Plotting for building footprints with attached assessments
def plot_assessments(gdf, mapName, cn='decision', classes=['GREEN','YELLOW','RED','TOTAL','LAND'], colors=['green','yellow','red','maroon','cyan'],
layer_name='Data', layer_only=False, no_leg=False, fill=0.7, legName=False):
classes = inputs['labels']['decision'].unique() if classes is False else classes
leg = {}
globals()['layer'+layer_name] = ipl.LayerGroup(name = layer_name)
for i, cl in enumerate(classes):
try: globals()['layer'+layer_name].add_layer(to_geodata(gdf.loc[gdf[cn].str.contains(cl)],colors[i],layer_name,fill))
except: globals()['layer'+layer_name].add_layer(to_geodata(gdf.loc[gdf[cn] == cl],colors[i],layer_name,fill))
leg.update({cl:colors[i]})
if not layer_only:
mapName.add_layer(globals()['layer'+layer_name])
if not 'l1' in globals() and no_leg is False: # Add legend if forming map for first time
l1 = ipl.LegendControl(leg, name=cn if legName is False else legName, position="bottomleft")
mapName.add_control(l1)
return mapName
else: return globals()['layer'+layer_name]
# + jupyter={"outputs_hidden": true}
# Visualise spatial results
import plotting as pl
for i in plots.keys(): globals()[i] = plots[i] # Retrieve variables to use
from ipyleaflet import LayersControl
import ipywidgets as ipw
from branca.colormap import linear
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.colors as clrs
ngrid=100
mf = pl.create_map(lat, lon, zoom, basemap=ipl.basemaps.OpenStreetMap.BlackAndWhite)
plot_assessments(labels, mf, cn=cn, layer_name='Ground truth', fill=0.4, legName='Ground Truth')
plot_assessments(nodes.to_crs({'init':crs}).dropna(), mf, layer_name='Train Locations', no_leg=True, classes=sorted([x for x in nodes.decision.unique() if str(x) != 'nan']), colors = ['green', 'red'] if nClasses==2 else None)
import geopandas as gpd
a = gpd.sjoin(initial, X_test, how='left', op='within').dropna(subset=[cn])
a['prediction']=pred_clf
plot_assessments(a.to_crs({'init':crs}).dropna(), mf, cn='prediction', layer_name='Test Predictions', no_leg=True, classes=[x for x in a.prediction.unique() if str(x) != 'nan'], colors = ['green', 'red'] if nClasses==2 else None)
from scipy.interpolate import griddata
xi, yi = np.linspace(nodes.geometry.x.min(), nodes.geometry.x.max(), ngrid), np.linspace(nodes.geometry.y.min(), nodes.geometry.y.max(), ngrid)
zi = griddata((nodes.geometry.x, nodes.geometry.y), (beliefs[:,0]-beliefs[:,1]+0.5), (xi[None, :], yi[:, None]), method='nearest')
#cs = plt.contourf(xi, yi, zi, norm=matplotlib.colors.Normalize(vmin=zi.min(), vmax=zi.max()),levels=20)
levs = math.floor((zi.max()-zi.min())/0.1)
print(levs)
cs = plt.contourf(xi, yi, zi, levels=levs-1, extend='both')
plt.close()
# add contours as polygons
# hardwired colors for now: these correspons to the 8-level default of matplotlib with an added orange color
# add contours as polygons
# hardwired colors for now: these correspons to the 8-level default of matplotlib with an added orange color
# colors10 = ["#ff0000", "#ff3232", "#ff6666", "#ff9999", "#ffcccc", "#ccf5cc", "#99eb99", "#66e166", "#32d732","#00cd00"]
# colors20 = ["#ff0000","#ff0000", "#ff1919", "#ff3232","#ff4c4c", "#ff6666", "#ff7f7f", "#ff9999","#ffb2b2", "#ffcccc","#ccf5cc","#ccf5cc","#ccf5cc","#ccf5cc","#ccf5cc","#b2f0b2", "#99eb99","#7fe67f","#66e166","#4cdc4c","#32d732","#19d219","#00cd00","#00cd00"]
#"#ffe5e5","#ffffff","#e5fae5"
colorsRed = ['#e50000','#ff0000','#ff3232','#ff6666','#ff9999']
colorsGreen = ['#b2f0b2','#99eb99','#66e166','#32d732','#00b800']
colors=[]
print(len(cs.allsegs))
for i in range(math.floor(len(cs.allsegs)/2-5)-math.floor(((zi.max()-1-(0-zi.min()))/0.1)/2)): colors.append('#ff0000')
colors += colorsRed
colors += colorsGreen
for i in range(math.ceil(len(cs.allsegs)/2-5)+math.floor(((zi.max()-1-(0-zi.min()))/0.1)/2)): colors.append('#32d732')
allsegs, allkinds = cs.allsegs, cs.allkinds
print(colors)
contourLayer = ipl.LayerGroup(name = 'Assessment Contours')
for clev in range(len(cs.allsegs)):
print(clev)
kinds = None if allkinds is None else allkinds[clev]
segs = split_contours(allsegs[clev], kinds)
polygons = ipl.Polygon(
locations=[p.tolist() for p in segs],
# locations=segs[14].tolist(),
color=colors[clev],
weight=1,
opacity=0.5,
fill_color=colors[clev],
fill_opacity=0.4,
name='layer_name'
)
contourLayer.add_layer(polygons)
mf.add_layer(contourLayer)
control = ipl.LayersControl(position='topright')
leg = dict(zip([str(round(x-0.1,1))+'-'+str(round(x,1)) for x in np.linspace(1,0.1,10).tolist()],colorsRed+colorsGreen))
l2 = ipl.LegendControl(leg, name='Damage Prob', position="topleft")
mf.add_control(l2)
mf.add_control(control)
zoom_slider = ipw.IntSlider(description='Zoom level:', min=7, max=18, value=14)
ipw.jslink((zoom_slider, 'value'), (mf, 'zoom'))
widget_control1 = ipl.WidgetControl(widget=zoom_slider, position='topright')
mf.add_control(widget_control1)
mf.add_control(ipl.FullScreenControl(position='topright'))
mf.zoom_control = False
mf
# -
def split_contours(segs, kinds=None):
"""takes a list of polygons and vertex kinds and separates disconnected vertices into separate lists.
The input arrays can be derived from the allsegs and allkinds atributes of the result of a matplotlib
contour or contourf call. They correspond to the contours of one contour level.
Example:
cs = plt.contourf(x, y, z)
allsegs = cs.allsegs
allkinds = cs.allkinds
for i, segs in enumerate(allsegs):
kinds = None if allkinds is None else allkinds[i]
new_segs = split_contours(segs, kinds)
# do something with new_segs
More information:
https://matplotlib.org/3.3.3/_modules/matplotlib/contour.html#ClabelText
https://matplotlib.org/3.1.0/api/path_api.html#matplotlib.path.Path
"""
if kinds is None:
return segs # nothing to be done
# search for kind=79 as this marks the end of one polygon segment
# Notes:
# 1. we ignore the different polygon styles of matplotlib Path here and only
# look for polygon segments.
# 2. the Path documentation recommends to use iter_segments instead of direct
# access to vertices and node types. However, since the ipyleaflet Polygon expects
# a complete polygon and not individual segments, this cannot be used here
# (it may be helpful to clean polygons before passing them into ipyleaflet's Polygon,
# but so far I don't see a necessity to do so)
new_segs = []
for i, seg in enumerate(segs):
segkinds = kinds[i]
boundaries = [0] + list(np.nonzero(segkinds == 79)[0])
for b in range(len(boundaries)-1):
new_segs.append(seg[boundaries[b]+(1 if b>0 else 0):boundaries[b+1]])
return new_segs
#contour = plt.contourf(xi, yi, zi, levels=14, cmap='RdYlGn')
import math
cs = plt.contourf(xi, yi, zi, levels=28, extend='both')
plt.colorbar()
def classify_data(v,seed=1):
# Retrieve data from inputs
for i in v.keys(): globals()[i] = v[i]
max_nodes = bxNodes.trait_values()['children'][1].value
nClasses = bxNClasses.trait_values()['children'][1].value
classAssign = False if ('bxAssign' not in v) or (bxCluster.trait_values()['children'][1].value is True) else [list(i.value) for i in bxAssign.trait_values()['children']]
classNames = False if 'bxClNames' not in v else [i.value for i in bxClNames.trait_values()['children']]
# Sample data and create geodataframe
print("------Data Sampling---------")
if max_nodes < 2: raise ValueError("Insufficient Nodes for belief propagation")
gdf = ip.get_sample_gdf(data, max_nodes, crs,seed=1)
print("------Data Classification---------")
defClasses, labelsUsed, dataUsed = len(labels[cn].unique()), labels.to_crs(crs).copy(), gdf.copy() # Default classes from labels
usedNames = labels[cn].unique() if nClasses==defClasses or nClasses is False else classNames
initial = hf.init_beliefs(dataUsed, classes=nClasses, columns=usedNames, crs=crs) # Initial class value for each data pixel
if not nClasses or nClasses == defClasses:
nClasses = defClasses # If default classes used
classesUsed = usedNames.copy()
elif nClasses > defClasses: raise NameError('Cannot assign more classes than in original data') # If invalid input
elif nClasses < defClasses: # Perform class grouping
items = [item for sublist in classAssign for item in sublist] if classAssign is not False else False
if (classAssign is False) or not any(classAssign) or (len(items) is not (len(set(items)))): # Perform clustering
if classAssign is not False: print('Incorrect class assignment - Proceeding with clustering. Please assign a single class for each value.')
# Assign labels to each pixel
allPixels = hf.create_nodes(initial, labelsUsed[['geometry',cn]][labelsUsed.within(hf.get_polygon(testPoly, conv=True))])
# Run PCA if set to True
#X = hf.run_PCA(dataUsed[typesUsed[0]].values.transpose(), pcaComps).components_.transpose() if pca else dataUsed[typesUsed[0]]
types = [item for sublist in typesUsed for item in sublist]
X = dataUsed[types]
# Run clustering
meanCluster = True
kmeans, clusterClasses, initLabels = hf.run_cluster(X.iloc[allPixels[cn].dropna().index].values.reshape(-1,len(types)), allPixels[cn].dropna(), meanCluster, nClasses)
print('Clustered classes:{} , original classes:{}'.format(clusterClasses, initLabels))
# Create groups of classes
classesUsed = []
for j in range(nClasses): classesUsed.append([initLabels[i] for i, x in enumerate(list(clusterClasses)) if x==j])
else:
if len(set(items)) is not defClasses:
print('Not all labels have been assigned to class. Sampling data to include only labels selected.')
labelsUsed = labelsUsed.loc[labelsUsed[cn].isin(items)]
classesUsed = classAssign
#used = [i in flatten_list(classesUsed) for i in labelsUsed[cn]]
initial = hf.init_beliefs(dataUsed, classes=nClasses, columns=usedNames, crs=crs)
# Assign labels for each pixel after clustering
labelsUsed[cn] = hf.group_classes(labelsUsed[cn], classesUsed)
print("------Finished Data Classification---------")
# Update variables
v.update({'max_nodes':max_nodes, 'nClasses':nClasses, 'classAssign':classAssign,'classNames':classNames, 'labelsUsed':labelsUsed,'initial':initial, 'usedNames':usedNames, 'classesUsed':classesUsed, 'dataUsed':dataUsed})
return v
# + jupyter={"source_hidden": true}
# Visualise spatial results
import plotting as pl
for i in plots.keys(): globals()[i] = plots[i] # Retrieve variables to use
from ipyleaflet import LayersControl
import ipywidgets as ipw
mf = pl.create_map(lat, lon, zoom, basemap=ipl.basemaps.OpenStreetMap.BlackAndWhite)
plot_assessments(labels, mf, cn=cn, layer_name='Ground truth', fill=0.1)
plot_assessments(nodes.to_crs({'init':crs}).dropna(), mf, layer_name='Train Locations', no_leg=True)
import geopandas as gpd
a = gpd.sjoin(initial, X_test, how='left', op='within').dropna(subset=[cn])
a['prediction']=pred_clf
plot_assessments(a.to_crs({'init':crs}).dropna(), mf, cn='prediction', layer_name='Test Predictions', no_leg=True)
control = ipl.LayersControl(position='topright')
mf.add_control(control)
zoom_slider = ipw.IntSlider(description='Zoom level:', min=7, max=18, value=14)
ipw.jslink((zoom_slider, 'value'), (mf, 'zoom'))
widget_control1 = ipl.WidgetControl(widget=zoom_slider, position='topright')
mf.add_control(widget_control1)
mf.add_control(ipl.FullScreenControl())
mf
| archive/run_netconf_widgets-edges.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.4 64-bit
# metadata:
# interpreter:
# hash: 5c4d2f1fdcd3716c7a5eea90ad07be30193490dd4e63617705244f5fd89ea793
# name: python3
# ---
# # Data Science Bootcamp - The Bridge
# ## Precurso
# En este notebook vamos a ver, uno a uno, los conceptos básicos de Python. Constarán de ejercicios prácticos acompañados de una explicación teórica dada por el profesor.
#
# Los siguientes enlaces están recomendados para el alumno para profundizar y reforzar conceptos a partir de ejercicios y ejemplos:
#
# - https://facundoq.github.io/courses/aa2018/res/02_python.html
#
# - https://www.w3resource.com/python-exercises/
#
# - https://www.practicepython.org/
#
# - https://es.slideshare.net/egutierrezru/python-paraprincipiantes
#
# - https://www.sololearn.com/Play/Python#
#
# - https://github.com/mhkmcp/Python-Bootcamp-from-Basic-to-Advanced
#
# Ejercicios avanzados:
#
# - https://github.com/darkprinx/100-plus-Python-programming-exercises-extended/tree/master/Status (++)
#
# - https://github.com/mahtab04/Python-Programming-Practice (++)
#
# - https://github.com/whojayantkumar/Python_Programs (+++)
#
# - https://www.w3resource.com/python-exercises/ (++++)
#
# - https://github.com/fupus/notebooks-ejercicios (+++++)
#
# Tutor de ayuda PythonTutor:
#
# - http://pythontutor.com/
#
# ## 1. Variables y tipos
#
# ### Cadenas
# +
# Entero - Integer - int
x = 7
# Cadena - String - Lista de caracteres
x = "lorena"
print(x)
x = 7
print(x)
# -
# built-in
type
# +
x = 5
y = 7
z = x + y
print(z)
# +
x = "'lorena'\" "
l = 'silvia----'
g = x + l
# Las cadenas se concatenan
print(g)
# -
print(g)
# type muestra el tipo de la variable
type(g)
type(3)
# +
# print es una función que recibe varios argumentos y cada argumento está diferenciado por la coma. Después de cada coma, la función 'print' añade un espacio.
# mala praxis
print( g,z , 6, "cadena")
# buena praxis - PEP8
print(g, z, 6, "cadena")
# +
u = "g"
silvia = "<NAME> "
anos = " años"
suma = silvia + u + anos
print(suma)
# +
n = 2
m = "3"
print(n + m)
# +
# Cambiar de int a str
j = 2
print(j)
print(type(j))
j = str(j)
print(j)
print(type(j))
# +
# Cambiar de int a str
j = 2
print(j)
print(type(j))
j = str(j) + " - " + silvia
print(j)
print(type(j))
# -
k = 22
k = str(k)
print(k)
# +
# Cambiar de str a int
lor = "98"
lor = int(lor)
print(lor)
# +
# Para ver la longitud de una lista de caracteres (lista)
mn = "lista de caracteres$%·$% "
#lenght
print(len(mn))
# +
h = len(mn)
print(h + 7)
# -
h = 8
print(h)
# +
x = 2
print(x)
gabriel_vazquez = "<NAME>"
# +
print("Hello Python world!")
print("Nombre de compañero")
companero_clase = "Compañero123123"
print(companero_clase)
# -
print(compañero)
x = (2 + 4) + 7
print(x)
# +
# String, Integer, Float, List, None (NaN)
# str, int, float, list,
string_ = "23"
numero = 23
print(type(string_))
# -
print(string_)
numero2 = 10
suma = numero + numero2
print(suma)
string2 = "33"
suma2 = string_ + string2
print(suma2)
m = (numero2 + int(string2))
print(m)
m = ((((65 + int("22")) * 2)))
m
print(type(int(string2)))
y = 22
y = str(y)
print(type(y))
string2 = int(string2)
print(type(string2))
# +
string3 = "10"
numero_a_partir_de_string = int(string3)
print(numero_a_partir_de_string)
print(string3)
print(type(numero_a_partir_de_string))
print(type(string3))
# -
h = "2"
int(h)
# +
# los decimales son float. Python permite las operaciones entre int y float
x = 4
y = 4.2
print(x + y)
# -
# La división normal (/) es siempre float
# La división absoluta (//) puede ser:
# - float si uno de los dos números (o los dos) son float
# - int si los dos son int
j = 15
k = 4
division = j // k
print(division)
print(type(division))
# +
num1 = 12
num2 = 3
suma = num1 + num2
resta = num1 - num2
multiplicacion = num1 * num2
division = num1 / num2
division_absoluta = num1 // num2
gabriel_vazquez = "<NAME>"
print("suma:", suma)
print("resta:", resta)
print("multiplicacion:", multiplicacion)
print("division:", division)
print("division_absoluta:", division_absoluta)
print(type(division))
print(type(division_absoluta))
# -
print(x)
j = "2"
j
print(j)
# +
x = 2
j = 6
g = 4
h = "popeye"
# Jupyter notebook permite que la última línea se imprima por pantalla ( la variable )
print(g)
print(j)
print(h)
x
# -
int(5.6/2)
float(2)
g = int(5.6/2)
print(g)
5
# +
x = int(5.6//2)
x
# -
# Soy un comentario
# print("Hello Python world!")
# Estoy creando una variable que vale 2
"""
Esto es otro comentario
"""
print(x)
x = 25
x = 76
x = "1"
message2 = "One of Python's strengths is its diverse community."
print(message2)
# ## Ejercicio:
# ### Crear una nueva celda.
# ### Declarar tres variables:
# - Una con el nombre "edad" con valor vuestra edad
# - Otra "edad_compañero_der" que contengan la edad de tipo entero de vuestro compañero de la derecha
# - Otra "suma_anterior" que contenga la suma de las dos variables anteriormente declaradas
#
# Mostrar por pantalla la variable "suma_anterior"
#
#
edad = 99
edad_companero_der = 30
suma_anterior = edad_companero_der + edad
print(suma_anterior)
h = 89 + suma_anterior
h
edad = 18
edad_companero_der = 29
suma_anterior = edad + edad_companero_der
suma_anterior
i = "hola"
o = i.upper()
o
o.lower()
name = "<NAME>"
x = 2
print(name.upper())
print(name.lower())
print(name.upper)
print(name.upper())
x = 2
x = x + 1
x
x += 1
x
# int
x = 1
# float
y = 2.
# str
s = "string"
# type --> muestra el tipo de la variable o valor
print(type(x))
type(y)
type(s)
5 + 2
x = 2
x = x + 1
x += 1
# +
x = 2
y = 4
print(x, y, "Pepito", "Hola")
# -
s = "<NAME> Soraya:"
s + "789"
print(s, 98, 29, sep="")
print(s, 98, 29)
type( x )
2 + 6
# ## 2. Números y operadores
# +
### Enteros ###
x = 3
print("- Tipo de x:")
print(type(x)) # Imprime el tipo (o `clase`) de x
print("- Valor de x:")
print(x) # Imprimir un valor
print("- x+1:")
print(x + 1) # Suma: imprime "4"
print("- x-1:")
print(x - 1) # Resta; imprime "2"
print("- x*2:")
print(x * 2) # Multiplicación; imprime "6"
print("- x^2:")
print(x ** 2) # Exponenciación; imprime "9"
# Modificación de x
x += 1
print("- x modificado:")
print(x) # Imprime "4"
x *= 2
print("- x modificado:")
print(x) # Imprime "8"
print("- El módulo de x con 40")
print(40 % x)
print("- Varias cosas en una línea:")
print(1, 2, x, 5*2) # imprime varias cosas a la vez
# +
# El módulo muestra el resto de la división entre dos números
2 % 2
# -
3 % 2
4 % 5
# +
numero = 99
numero % 2 # Si el resto es 0, el número es par. Sino, impar.
# -
numero % 2
99 % 100
y = 2.5
print("- Tipo de y:")
print(type(y)) # Imprime el tipo de y
print("- Varios valores en punto flotante:")
print(y, y + 1, y * 2.5, y ** 2) # Imprime varios números en punto flotante
# ## Título
#
# Escribir lo que sea
#
# 1. uno
# 2. dos
# # INPUT
# +
edad = input("Introduce tu edad")
print("Diego tiene", edad, "años")
# +
# Input recoge una entrada de texto de tipo String
num1 = int(input("Introduce el primer número"))
num2 = int(input("Introduce el segundo número"))
print(num1 + num2)
# -
# ## 3. Tipo None
x = None
n = 5
s = "Cadena"
print(x + s)
# ## 4. Listas y colecciones
# +
# Lista de elementos:
# Las posiciones se empiezan a contar desde 0
s = "Cadena"
primer_elemento = s[0]
#ultimo_elemento = s[5]
ultimo_elemento = s[-1]
print(primer_elemento)
print(ultimo_elemento)
# -
bicycles = ['trek', 'cannondale', 'redline', 'specialized']
bicycles[0]
tamano_lista = len(bicycles)
tamano_lista
ultimo_elemento_por_posicion = tamano_lista - 1
bicycles[ultimo_elemento_por_posicion]
# +
bicycles = ['trek', 'cannondale', 'redline', 'specialized']
message = "My first bicycle was a " + bicycles[0]
print(bicycles)
print(message)
# -
print(type(bicycles))
s = "String"
s.lower()
print(s.lower())
print(s)
s = s.lower()
s
# +
# Existen dos tipos de funciones:
# 1. Los que modifican los valores sin que tengamos que especificar una reasignación a la variable
# 2. Los que solo devuelven la operación y no modifican el valor de la variable. Tenemos que forzar la reasignación si queremos modificar la variable.
cars = ['bmw', 'audi', 'toyota', 'subaru']
print(cars)
cars.reverse()
print(cars)
# +
cars = ['bmw']
print(cars)
cars.reverse()
print(cars)
# +
s = "Hola soy Clara"
print(s[::-1])
# -
s
l = "hola"
len(l)
l[3]
# +
# POP
lista = [2, 4, 6, "a", 6, 8]
lista.pop(4)
lista
# +
lista = [2, 4, 6, "a", 6]
# Remove borra el primer elemento con el valor dado
lista.remove(6)
lista
# +
# pop retorna el valor borrado
lista = [2, 4, 6, "a", 6, 8]
x = lista.pop(4)
print(x)
lista
# +
lista = [2, 4, 6, "a", 6, 8]
x = lista.remove(6)
print(x)
# +
def remove2(a):
x = a + 2
y = remove2(6)
print(y)
# +
lista = [2, 4, 6, "a", 6, 8]
def f(s):
print(s)
x = f(s=2)
print(x)
# +
# Para acceder a varios elementos, se especifica con la nomenclatura "[N:M]". N es el primer elemento a obtener, M es el último elemento a obtener pero no incluido. Ejemplo:
# Queremos mostrar desde las posiciones 3 a la 7. Debemos especificar: [3:8]
# Si M no tiene ningún valor, se obtiene desde N hasta el final.
# Si N no tiene ningún valor, es desde el principio de la colección hasta M
s[3:len(s)]
# -
s[:3]
s[3:10]
# +
motorcycles = ['honda', 'yamaha', 'suzuki', 'ducati']
print(motorcycles)
too_expensive = 'ducati'
motorcycles.remove(too_expensive)
print(motorcycles)
print(too_expensive + " is too expensive for me.")
# -
# Agrega un valor a la última posición de la lista
motorcycles.append("ducati")
motorcycles
lista = ['honda', 2, 8.9, [2, 3], 'yamaha', 'suzuki', 'ducati']
lista[3]
lista.remove(8.9)
lista
lista
l = lista[1]
l
lista.remove(l)
lista
lista.remove(lista[2])
lista = ['honda', 2, 8.9, [2, 3], 'yamaha', 'suzuki', 'honda', 'ducati']
lista
# remove elimina el primer elemento que se encuentra que coincide con el valor del argumento
lista = ['honda', 2, 8.9, [2, 3], 'yamaha', 'suzuki', 'honda', 'ducati']
lista.remove("honda")
lista
# Accedemos a la posición 1 del elemento que está en la posición 2 de lista
lista[2][1]
lista[3][2]
p = lista.remove("honda")
print(p)
l = [2, 4, 6, 8]
l.reverse()
l
# ### Colecciones
#
# 1. Listas
# 2. String (colección de caracteres)
# 3. Tuplas
# 4. Conjuntos (Set)
# Listas --> Mutables
lista = [2, 5, "caract", [9, "g", ["j"]]]
print(lista[-1][-1][-1])
lista[3][2][0]
lista.append("ultimo")
lista
# +
# Tuplas --> Inmutables
tupla = (2, 5, "caract", [9, "g", ["j"]])
tupla
# -
s = "String"
s[2]
tupla[3].remove(9)
tupla
tupla[3][1].remove("j")
tupla
tupla2 = (2, 5, 'caract', ['g', ['j']])
tupla2[-1].remove(["j"])
tupla2
tupla2 = (2, 5, 'caract', ['g', ['j']])
tupla2[-1].remove("g")
tupla2
tupla2[-1].remove(["j"])
tupla2
if False == 0:
print(0)
print(type(lista))
print(type(tupla))
# Update listas
lista = [2, "6", ["k", "m"]]
lista[1] = 1
lista
lista = [2, "6", ["k", "m"]]
lista[2] = 0
lista
tupla = (2, "6", ["k", "m"])
tupla[2] = 0
tupla
tupla = (2, "6", ["k", "m"])
tupla[2][1] = 0
tupla
# +
# Conjuntos
conjunto = [2, 4, 6, "a", "z", "h", 2]
conjunto = set(conjunto)
conjunto
# -
conjunto = ["a", "z", "h", 2, 2, 4, 6, True, True, False]
conjunto = set(conjunto)
conjunto
conjunto = ["a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False]
conjunto = set(conjunto)
conjunto
conjunto_tupla = ("a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False)
conjunto = set(conjunto_tupla)
conjunto
conjunto = {"a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False}
conjunto
s = "String"
lista_s = list(s)
lista_s
s = "String"
conj = {s}
conj
tupla = (2, 5, "h")
tupla = list(tupla)
tupla.remove(2)
tupla = tuple(tupla)
tupla
tupla = (2, 5, "h")
tupla = list(tupla)
tupla.remove(2)
tupla = (((((tupla)))))
tupla
tupla = (2, 5, "h")
tupla = list(tupla)
tupla.remove(2)
tupla = tuple(tupla)
tupla
conjunto = {2, 5, "h"}
lista_con_conjunto = [conjunto]
lista_con_conjunto[0]
# No podemos acceder a elementos de un conjunto
lista_con_conjunto[0][0]
lista = [1, 5, 6, True, 6]
set(lista)
lista = [True, 5, 6, 1, 6]
set(lista)
1 == True
# ## 5. Condiciones: if, elif, else
# ### Boolean
True
False
# Operadores comparativos
x = (1 == 1)
x
1 == 2
"a" == "a"
# Diferente
"a" != "a"
2 > 4
4 > 2
4 >= 4
4 > 4
4 <= 5
4 < 3
input()
1 == 1
"""
== -> Igualdad
!= -> Diferecia
< -> Menor que
> -> Mayor que
<= -> Menor o igual
>= -> Mayor o igual
"""
# and solo devolverá True si TODOS son True
True and False
# +
# or devolverá True si UNO es True
(1 == 1) or (1 == 2)
# -
(1 == 1) or (1 == 2) and (1 == 1)
(1 == 1) or ((1 == 2) and (1 == 1))
(1 == 1) and (1 == 2) and (1 == 1)
(1 == 1) and (1 == 2) and ((1 == 1) or (0 == 0))
# True and False and True
print("Yo soy\n Gabriel")
if 1 != 1:
print("Son iguales")
else:
print("No entra en if")
if 1 == 1:
print("Son iguales")
else:
print("No entra en if")
if 1>3:
print("Es mayor")
elif 2==2:
print("Es igual")
elif 3==3:
print("Es igual2")
else:
print("Ninguna de las anteriores")
if 2>3:
print(1)
else:
print("Primer else")
if 2==2:
print(2)
else:
print("Segundo else")
if 2>3:
print(1)
else:
print("Primer else")
if 3==3:
print(" 3 es 3 ")
if 2==2:
print(2)
else:
print("Segundo else")
if 2>3:
print(1)
else:
print("Primer else")
# -------
if 3==4:
print(" 3 es 3 ")
# --------
if 2==2:
print(2)
print(5)
x = 6
print(x)
else:
print("Segundo else")
if 2>3:
print(1)
else:
print("Primer else")
# -------
if 3==4:
print(" 3 es 3 ")
# --------
if 2==2:
print(2)
print(5)
x = 6
print(x)
# ------
if x == 7:
print("X es igual a 6")
# ------
y = 7
print(y)
else:
print("Segundo else")
if (not (1==1)):
print("Hola")
if not None:
print(1)
if "a":
print(2)
if 0:
print(0)
# +
# Sinónimos de False para las condiciones:
# None
# False
# 0 (int or float)
# Cualquier colección vacía --> [], "", (), {}
# El None no actúa como un número al compararlo con otro número
"""
El True lo toma como un numérico 1
"""
# +
lista = []
if lista:
print(4)
# +
lista = ["1"]
if lista:
print(4)
# -
if 0.0:
print(2)
if [] or False or 0 or None:
print(4)
if [] and False or 0 or None:
print(4)
if not ([] or False or 0 or None):
print(4)
if [] or False or not 0 or None:
print(True)
else:
print(False)
# +
x = True
y = False
x + y
# +
x = True
y = False
str(x) + str(y)
# -
t = True
f = False
2 == 2
# +
l1 = [1, 2]
l2 = [1, 2]
l1 == l2
# -
id(l1)
id(l2)
l1 == l2
2 is 2
id(2)
l1 is l2
id("a")
"a" is "a"
h = "a"
g = "a"
h == g
id(h)
id(g)
h is g
id(2)
b = 875757
id(b)
f = 875757
id(f)
b is f
# +
d = 2
s = 2
id(d)
# -
id(s)
# +
def funcion_condicion():
if y > 4:
print("Es mayor a 4")
else:
print("No es mayor a 4")
funcion_condicion(y=4)
# +
def funcion_primera(x):
if x == 4:
print("Es igual a 4")
else:
funcion_condicion(y=x)
funcion_primera(x=5)
# +
def funcion_final(apellido):
if len(apellido) > 5:
print("Cumple condición")
else:
print("No cumpole condición")
funcion_final(apellido="Vazquez")
# -
#
# # Bucles For, While
# +
lista = [1, "dos", 3, "Pepito"]
print(lista[0])
print(lista[1])
print(lista[2])
print(lista[3])
# -
if 2 != 2:
print(True)
print(4)
lista = [1, "dos", 3, "Pepito"]
for elnombrequequiera in lista:
print(elnombrequequiera)
print("-----")
# De esta forma NO muestra el 6
for g in [2, 4, 6, 8]:
if g == 6:
break
print(g)
print(8)
print(9)
print("ESto está fuera del bucle")
# De esta forma muestra el 6 -- versión 1
for g in [2, 4, 6, 8]:
if g == 8:
break
print(g)
print(8)
print(9)
print("ESto está fuera del bucle")
# De esta forma muestra el 6 -- versión 2
for g in [2, 4, 6, 8]:
print(g)
if g == 6:
break
print(8)
print(9)
print("ESto está fuera del bucle")
# De esta forma muestra el 6 -- versión 3
for g in [2, 4, 6, 8]:
if g > 6:
break
print(g)
print(8)
print(9)
print("ESto está fuera del bucle")
# De esta forma solo muestra el 6 -- versión 4
for g in [2, 4, 6, 8]:
if g == 6:
print(g)
break
print(8)
print(9)
print("ESto está fuera del bucle")
==
!=
>
<
>=
<=
and
or
not
# De esta forma solo muestra el 6 -- versión 4
for g in [2, 4, 6, 8]:
if g == 6:
print(g)
break
print(8)
print(9)
print("ESto está fuera del bucle")
k = 1, 2
k
file1 = "Primera Fila", ["Maria", "Angeles", "Juan"]
file1
file1[1][1]
# Intento <NAME>
if type(x) == str:
for x in lista:
print(x)
# Intento <NAME> solved
for x in lista:
if type(x) == str:
print(x)
file1 = "Primera Fila", ["Maria", "Angeles", "Juan"]
file2 = "Segunda Fila", ["Marta", "Daniel", "Leo", "Miguel1", "Estela"]
file3 = "<NAME>", ["Kapil", "Roberto", "Alfonso", "Miguel2"]
fileR = "Remoto", ["Mar", "Alex", "Anais", "Antonio", "Ariadna", "Javi", "JaviOlcoz"]
# +
altura1 = [1.78, 1.63, 1.75, 1.68]
altura2 = [2.00, 1.82]
altura3 = [1.65, 1.73, 1.75]
altura4 = [1.72, 1.71, 1.71, 1.62]
lista_alturas = [altura1, altura2, altura3, altura4]
print(lista_alturas[0][1])
for x in lista_alturas:
print(x[3])
# +
lista_con_lista = [2, "4", "c", [0, "a"]]
print(lista_con_lista[3][1])
# -
lista_con_lista[3][1] = "modificado"
lista_con_lista
# +
lista_con_lista = [2, "4", "c", (0, "a")]
tupla_lista = list(lista_con_lista[3])
lista_con_lista[3] = tupla_lista
lista_con_lista
# +
lista_con_lista = [2, "4", "c", [7, "a", 7]]
pos_7 = lista_con_lista[3].index(7)
pos_7
# +
#for x in lista_con_lista: # Valor
# for pos in range(len(lista_con_lista)): # range funciona por posición
# for pos, val in enumerate(lista_con_lista): # enumerate tanto por posición como valor
lista_con_lista = [2, "4", "c", [7, "a", 7]]
i = 0
for x in lista_con_lista:
if i == 3:
print("-------")
print("pos de 7:", x.index(7))
z += 1
print(i, ":", x)
i += 1
# +
altura1 = [1.78, 1.63, 1.75, 1.68]
altura2 = [2.00, 1.82]
altura3 = [1.65, 1.73, 1.75]
altura4 = [1.72, 1.71, 1.71, 1.62]
lista_alturas = [altura1, altura2, altura3, altura4]
print(lista_alturas[0][1])
for x in lista_alturas:
if len(x) > 3: # La lista ha de tener 4 elementos
print(x[3])
# -
print(range(6))
type(range(6))
print(list(range(6)))
r = list(range(6))
for i in r:
print(r)
for i in range(6): # Ejecuta 6 iteraciones
print(r)
for i in range(6):
print(i)
list(range(4))
for i in range(4): # bucle con 4 iteraciones
s = input("Introduce letra:")
print(s)
# +
lista = ("juan", "pepito", "silvia", "6", 7)
tamano = len(lista)
for i in range(tamano):
print(lista[i])
# +
# Con range, creamos una colección que empieza en 0 hasta un número N. En ese caso, trabajamos con posiciones.
# +
# enumerate
lista = ("juan", "pepito", "silvia", "6", 7)
for posicion, valor in enumerate(lista):
print("position:", posicion)
print("valor:", valor)
print("-------")
# -
k = 2, 7
k
tupla = ("juan", "pepito", "silvia", "6", 7)
for k in enumerate(tupla):
print(k)
tupla = ("juan", "pepito", "silvia", "6", 7)
for pos, val in enumerate(tupla):
print(pos)
print(val)
# continue
tupla = ("juan", "pepito", "silvia", "6", 7)
for x in tupla: # O(n)
if x == "silvia":
continue # continue salta a la siguiente iteración
print(x)
list(range(4))
for i in range(4): # Lo que haya debajo se ejecuta 4 veces
print("Iteración ", i+1," del primer bucle")
print("-------------------")
# +
# bucle ejecutando todo N veces
# acumuladores
# Queremos que se muestren todos los elementos de tupla 4 veces excepto "silvia" la última iteración
tupla = ("juan", "pepito", "silvia", "6", 7)
N = 4
for i in range(N): # Lo que haya debajo se ejecuta 4 veces
print("Iteración", i+1,"del primer bucle")
print("-------------------")
for x in tupla: # Recorre cada valor de tupla y x tiene cada valor del elemento que se está recorriendo
if x == "silvia" and i == (N - 1):
continue # continue salta a la siguiente iteración
print(x)
print(2)
# +
# bucle ejecutando todo N veces
# acumuladores
# Queremos que se muestren todos los elementos de tupla 4 veces excepto "silvia" la última iteración
tupla = ("juan", "pepito", "silvia", "6", 7)
N = 4
for i in range(N): # Lo que haya debajo se ejecuta 4 veces
print("-------------------")
print("Iteración", i+1,"del primer bucle")
print("-------------------")
for pos, x in enumerate(tupla): # Recorre cada valor de tupla y x tiene cada valor del elemento que se está recorriendo
if x == "silvia" and i == (N - 1):
continue # continue salta a la siguiente iteración
print("~~~~~~~~~~")
print("It", pos+1, "del segundo bucle")
print("~~~~~~~~~~")
print(x)
print(2)
# +
lista = ["a", "b", "c", "d"]
for i in range(len(lista)):
print(i, lista[i])
# -
lista = ["a", "b", "c", "d"]
for i, x in enumerate(lista):
print(i, x)
lista = ["a", "b", "c", "d"]
acum = 0
for x in lista: # el valor del elemento
print(acum, x)
acum += 1
lista = ["a", "b", "c", "d"]
acum = -1
for x in lista: # el valor del elemento
acum += 1
print(acum, x)
list(range(len(lista)))
# ## Funciones
# +
l = ["primer", 2, "tercer", 4]
if "tercer" in l:
l.pop(0)
l
# -
l.remove(4)
l
l2 = [2, 3, 3, 3, 3, 3, 4]
for x in l2:
if x == 3:
l2.remove(x)
print(l2)
l2
list(range(7))
l2 = [2, 3, 3, 3, 3, 3, 4]
for i in range(len(l2)): # hará tantas iteraciones como elementos tenga l2
if 3 in l2:
l2.remove(3)
print(l2)
l2 = [2, 3, 3, 3, 3, 3, 4]
tamano = len(l2)
for i in range(tamano):
print("tamaño:", tamano)
if 3 in l2:
l2.remove(3)
print(l2)
# +
# La función retorna None por defecto
def nombre_funcion():
print("Hola")
x = 2
print(x)
return None
r = nombre_funcion()
print(r)
# +
def nombre_funcion():
print("Hola")
x = 2
print(x)
return 7
r = nombre_funcion()
print(r)
# +
def suma_2_argumentos(a, b):
return int(a) + int(b)
lo_que_retorna = suma_2_argumentos(a="76", b=40)
print("lo_que_retorna:", lo_que_retorna)
# +
l1 = [2, 6, "10"]
l2 = ["l", "aa"]
l3 = l1 + l2
l3
# +
def suma_2_argumentos(a, b):
a = int(a)
b = int(b)
print(a)
return a + b
lo_que_retorna = suma_2_argumentos(a="76", b=40)
print("lo_que_retorna:", lo_que_retorna)
r = suma_2_argumentos(a="8", b=40)
# -
l = [2, 4]
l = int(l)
l
# +
def suma_2_argumentos(a, b):
a = int(a)
b = int(b)
print(a)
return a + b
lo_que_retorna = suma_2_argumentos(a=[2,4], b=[3, 5, 7])
# +
def suma_2_argumentos(a, b):
a = int(a)
b = int(b)
print(a)
return a + b
lo_que_retorna = suma_2_argumentos(2, 6)
# +
def suma_2_argumentos(a, b):
""" Esto es la descripción de la función """
a = int(a)
b = int(b)
return a + b
suma_2_argumentos(2,1)
# +
def f():
x = int(input())
r = int(input())
return x / r
f()
# +
def add_element(lista, to_add):
lista.append(to_add)
l = [2, 4, 6]
to_add = 9
add_element(lista=l, to_add=9)
print(l)
# +
# Todas las variables que están dentro de una función son borradas de memoria al terminar la ejecución de la función.
def k():
x = 2
y = 3
m = 8
k()
print(m)
# +
def k():
x = 2
y = 3
m = 8
return 7
lo_que_devuelve_k = k()
print(lo_que_devuelve_k)
# +
def k():
x = 2
y = 3
m = 8
return 7
print(k())
# +
def g(l):
""" Devuelve True si la lista 'l' contiene un número mayor a 5
Args:
l (list): Lista de elementos
"""
for i in l:
if i > 5:
return True
lista = [2, 4, 5, 9, 999]
z = g(l=lista)
print(z)
# +
acum += 1
acum
# +
def s(l):
""" Devuelve el número de veces que hay número mayor a 5 en la lista 'l'
"""
acum = 0
for i in l:
if i > 5:
acum = acum + 1
return acum
lista = [2, 4, 5, 9, 999]
z = s(l=lista)
print(z)
# +
def p1(l):
acum = 0
for pos, val in enumerate(l):
# if pos == 0: # Esto se cumple en la primera iteración
if pos == 2: # Esto se cumple en la tercera iteración
break
lista = [2, 4, 5, 9, 999]
z = s(l=lista)
print(z)
# +
lista3 = [2, 4, 5, 9, 999]
def p1(l):
for pos, val in enumerate(l):
if pos == len(l) - 1: # Se cumple cuando sea la última iteración
return 2
else:
print("True")
z = p1(l=lista3)
print(z)
# +
def df(l):
acum = 0
return acum
print(acum)
for i in l:
if i > 5:
acum = acum + 1
lista = [2, 4, 5, 9, 999]
z = df(l=lista)
z
# +
def df(l):
acum = 0
print(acum)
for i in l:
if i > 5:
acum = acum + 1
lista = [2, 4, 5, 9, 999]
z = df(l=lista)
print(z)
# +
# Función que retorne cuantos números hay mayores a 5 en los elementos de 'l'
l = ["Ana", "Dorado", ["m", 2], 7]
def r_n_m_5(l):
acum = 0
for elem in l:
if type(elem) == int and elem > 5:
print(elem)
acum += 1
return acum
r_n_m_5(l=l)
# +
def r_n_m_5(l):
acum = 0
for elem in l:
if type(elem) == int:
if elem > 5:
print(elem)
acum += 1
return acum
r_n_m_5(l=l)
# +
# Función que retorne cuantos números hay mayores a 5 en los elementos de 'l'
l = ["Ana", "Dorado", ["m", 2], 7]
def r_n_m_5(l):
acum = 0
for elem in l:
if isinstance(elem, int) and elem > 5:
print(elem)
acum += 1
return acum
r_n_m_5(l=l)
# -
i = 2.2
print(type(i))
i = int(i)
print(type(i))
print(i)
# +
def mostrar_cada_elemento_de_lista(lista):
for x in lista:
print(x)
mostrar_cada_elemento_de_lista(lista=lista_alturas)
# -
mostrar_cada_elemento_de_lista(lista=lista)
for x in lista_alturas:
if len(x) > 2:
print(x[2])
else:
print(x[1])
# # Diccionarios
# +
hola = "saludo internacional"
#clave/key = valor
diccionario = {"key":"valor"}
diccionario
# -
print(diccionario["key"])
diccionario2 = {2:"este es el valor", 2:"este es el valor"}
diccionario2
# +
diccionario3 = {"<EMAIL>":["password", "<PASSWORD>", 606122333, "C/Pepito", "Profesor"]}
diccionario3["<EMAIL>"]
# -
diccionario4 = {"juan": 8, "silvia":10, "juan2":9}
diccionario5 = {9.8:"S"}
diccionario5
diccionario6 = {"<EMAIL>":["password", "<PASSWORD>", 606122333, "C/Pepito", "Profesor"],
"<EMAIL>":["password", "<PASSWORD>", 606142333, "C/Pepito", "Profesor"],
"<EMAIL>":["password", "<PASSWORD>", 602122333, "C/Pepito", "Profesor"]}
diccionario6["<EMAIL>"]
diccionario6["password"]
del diccionario6["<EMAIL>"]
diccionario6
diccionario7 = {"k":"v", 8:[7,"y"], 6:{1.1:[5,"b"]}}
type(diccionario7[6][1.1][1])
list(diccionario7.keys())
diccionario7.keys()
diccionario7.values()
list(diccionario7.values())
diccionario7
for key in diccionario7.keys():
print(key)
for value in diccionario7.values():
print(value)
diccionario7
for key, value in diccionario7.items():
print(key, "--->", value)
for key, value in diccionario7.items():
if (type(key) == int or type(key) == float) and key > 5:
print(key, "--->", value)
for pos, (key, value) in enumerate(diccionario7.items()):
print("Iteración número:", pos+1)
print(key, "--->", value)
print("#####################")
lista = [2, 4, 6, "k"]
for pos, value in enumerate(lista):
print("Iteración número:", pos)
# +
lista = [2, 4, 6, "k"]
pos = 0
for value in lista:
print("Iteración número:", pos)
pos += 1
print("Real valor de pos:", pos)
print("#################")
print("Último valor de pos:", pos)
# +
# Creación de conjunto (set)
d = {3, 5, 1, "z", "a"}
d
# +
# Creación de diccionario
d = {"k": 2, "k2": [8, "p"]}
d
# -
# Acceder a un valor a través de una key
key_a_buscar = "k2"
d[key_a_buscar]
# Buscar/Mostrar todas las keys
list(d.keys())
# Buscar/Mostrar todos los values
list(d.values())
# Buscar/Mostrar todos los keys/values
list(d.items())
k = 2, "g"
k
# +
# Buscar en bucle todos los elementos
for key, value in d.items():
if key == "k2":
print(key, "-->", value)
# -
d
# Recorrer la lista value de la clave "k2"
for key, value in d.items():
if key == "k2":
for x in value:
print(x)
d
# Borrar clave de diccionario
del d["k"]
d
# Actualizar/Modificar el valor asociado a una clave
d["k2"] = [8, "a"]
d
# +
# Añadir un diccionario a otro
d1 = {8: [8, 9]}
d2 = {"a":"hola"}
d1.update(d2)
print(d1)
# +
def suma_dos_valores(a, b):
x = a + b
return x
k = suma_dos_valores(a=9, b=1)
print(k)
k + 7
# -
l = [2, 7]
l = l.append("a")
print(l)
# +
def add_to_list(lista, to_add):
lista.append(to_add)
return lista
l = [2, 7]
l = add_to_list(lista=l, to_add=6)
print(l)
# +
def add_to_list(lista, to_add):
lista.append(to_add)
l = [2, 7]
add_to_list(lista=l, to_add=6)
print(l)
# -
d["k2"].append(3)
d
l8 = [1,2]
d = {2: "h", "j":l8}
l = [5, "m", d]
l[-1]
# +
# Retornar más de un elemento
k = 2, 7
k
# -
k, j = 2, 7
print(k)
print(j)
k, j = 2, 7, 3
print(k)
print(j)
k, j = [2, 0], (4, 7, "m")
print(k)
print(j)
# +
def f():
return 3, 6, 8
def cualquiercosa():
return 5, 2, 3
def t():
return 6, 2, 3
g, h = f(), cualquiercosa()
print(g)
print(h)
# -
# ### Funciones con valores por defecto
#
# +
# función con un parámetro
def nombre_funcion(param):
return param
x = nombre_funcion(1)
x = nombre_funcion(param=1)
x
# +
# Función con parámetro/valor por defecto
def nombre_funcion_por_defecto(param=2):
return param
x = nombre_funcion_por_defecto()
x
# +
def nombre_funcion_por_defecto(param=2):
return param
x = nombre_funcion_por_defecto(param=7)
x
# +
# Si un argumento de la función no tiene un valor por defecto, es obligatorio darle un valor al llamar a la función
# Si le damos un valor a un argumento de la función al ser llamada, ese valor tiene prioridad al valor por defecto
def nombre_funcion_por_defecto(y, param=5):
return param + y
x = nombre_funcion_por_defecto(4, 7)
x
# +
def nombre_funcion_por_defecto(y=4, param=5):
return param - y
x = nombre_funcion_por_defecto(param=7, y=8)
x
# +
def nombre_funcion_por_defecto(y=4, param=5):
return param - y
x = nombre_funcion_por_defecto(param=2, y=4)
x
# +
def nombre_funcion_por_defecto(y, param=2, h=1, p=0):
return param - y
x = nombre_funcion_por_defecto(3,6,h=1)
x
# +
def nombre_funcion_por_defecto(y, param=2, h=1, p=0):
return param - y
x = nombre_funcion_por_defecto(y=4, h=2)
x
# +
def nombre_funcion_por_defecto(y, param=2, h=1, p=0):
if y == 2:
return param - y
else:
return True
x = nombre_funcion_por_defecto(4, 2,)
x
# -
lista = [2, 4, 6, 8,]
lista
def gh(c=6):
return c
# +
def get_last_value_from_list(lista, position=-1):
if len(lista) > 0: # Si hay algún elemento
return lista[position]
else:
return "No hay elementos"
lista = ["m", 2]
x = get_last_value_from_list(lista=lista)
print(x)
# +
lista = [2, "8", [6, "joven"], "casa", "coche", "pepino", [9, 8]]
def muestra_todos_excepto_joven_casa_9(lista):
for x in lista:
if x == lista[2]:
print(x[0])
elif x == lista[3]:
continue
elif x == lista[-1]:
print(x[1])
else:
print(x)
muestra_todos_excepto_joven_casa_9(lista=lista)
# -
# # While
# +
lista = [7, 5, "m", "Z"]
for elem in lista:
print(elem)
# +
lista = [2, 4, 6, 8, 10]
contador = 0
for elem in lista:
if elem > 5:
#contador = contador + 1
contador += 1
print(contador)
# -
while True:
print(5)
break
print("Fuera del while")
s = "Start"
while s == "Start":
print("Entra en el while")
s = input()
print("Fuera del while")
# +
password = "TB"
while True:
s = input("Introduce la contraseña:")
if s == password:
print("Contraseña correcta")
break
else:
print("Contraseña incorrecta")
# -
password = "TB"
s = ""
while s != password:
s = input("Introduce la contraseña:")
if s == password:
print("Contraseña correcta")
else:
print("Contraseña incorrecta")
print("Fuera del while")
# +
contador = 0
while contador != 5:
print(contador)
contador += 1
print("Fuera del while")
# +
lista = ["a", "b", "c", "d"]
print(lista[0])
print(lista[1])
print(lista[2])
print(lista[3])
# +
lista = ["a", "b", "c", "d"]
acum = 0
while acum < len(lista):
if acum == 2:
break
print(lista[acum])
acum += 1
# +
lista = ["a", "b", "c", "d"]
acum = 0
while True:
if acum == 2:
continue
print(lista[acum])
acum += 1
# +
# No ejecutar porque no para
lista = ["a", "b", "c", "d"]
acum = 0
while True:
if acum == 0:
continue
print(8)
break
# +
import time
while True:
print("Hola")
time.sleep(3)
print("Adios")
break
# +
lista = ["a", "b", "c", "d", "e"]
acum = 1
while 1 <= acum and acum <= 3:
print(lista[acum])
acum += 1
# +
lista = ["a", "b", "c", "d", "e"]
acum = 1
while 1 <= acum and acum <= 3:
if acum == 2:
acum += 1
continue
print(lista[acum])
acum += 1
# +
lista = ["a", "b", "c", "d", "e"]
acum = 1
while 1 <= acum and acum <= 3:
print(lista[acum])
acum += 2
# +
lista = ["a", "b", "c", "d", "e", "f"]
acum = 0
while acum < len(lista):
print(lista[acum])
acum += 2
# +
lista = ["a", "b", "c", "d", "e", "f"]
acum = 1
while acum < len(lista):
print(lista[acum])
acum += 2
# +
lista = ["a", "b", "c", "d", "e", "f"]
acum = 1
while acum >= 1 and acum <= len(lista) - 1:
print(lista[acum])
acum += 2
# -
# # Try Except
# +
def suma(a, b="3"):
resultado = a + b
return resultado
s = suma(a=2)
s
# +
def suma(a, b="3"):
try:
resultado = a + b
except:
print("Ha ocurrido un error al sumar")
a = int(a)
b = int(b)
resultado = a + b
return resultado
s = suma(a=2)
s
# +
def suma_positiva(a, b):
""" Suma dos valores numéricos de tipo entero. Si algún parámetro incorrecto, la función devolverá (0x086080808)"""
resultado = -1
try:
resultado = a + b
except:
print("Ha ocurrido un error al sumar. Los argumentos 'a' y 'b' deben ser de tipo Entero. El código del error es: '0x86080808'")
return resultado
s = suma_positiva(a=2, b="4")
s
# +
a = 2
b = "g"
resultado = a + b
print("hola")
# +
a = 2
b = "g"
try:
resultado = a + b
except Exception as error:
print("Ha ocurrido un error:", error)
resultado = -1
print("hola")
# +
def minicalculadora(a, b, operador, DEBUG=0):
try:
a = int(a)
b = int(b)
if operador == "+":
return a + b
elif operador == "/":
return a / b
else:
print("Esta calculadora solo permite suma y división")
except Exception as error:
if DEBUG == 1:
print(error)
print("Ha ocurrido un error, solo se admiten números")
s = input("Escribe un operador")
a = input("Escribe un número")
b = input("Escribe otro número")
minicalculadora(a=a, b=b, operador=s, DEBUG=1)
# -
# ### Assert
# +
def mostrar_dinero_banco(password):
assert (password==<PASSWORD>), "No has introducido la contraseña correcta"
return <PASSWORD>
mostrar_dinero_banco(password=<PASSWORD>)
# -
try:
assert (1==2), "Error, 1 no es 2"
except Exception as error:
print("error:", error)
print("Ha ocurrido un error")
# ### Zip List/Dict
# +
l1 = ["a", 3, 7]
l2 = ["b", 2, 0]
l3 = list(zip(l1, l2))
l3
# +
l3 = ["a", "b", "c"]
l4 = [2, 4, 6]
for i in range(len(l3)):
print(l3[i])
print(l4[i])
print("-----")
# -
l5 = list(zip(l3, l4))
l5
k = 2, 4
k
j, h = 2, 4
l5
for elem in l5:
print(elem)
for e1, e2 in l5:
print(e1)
print(e2)
print("-----")
# +
l1 = [["a"], [3, 2], (7, "m")]
l2 = [(5, 7), ["x"], ["y"]]
l3 = list(zip(l1,l2))
print(l3)
# -
for e1, e2 in l3:
print(e1)
# +
l1 = [["a","b"], [3, 2], (7, "m")]
l2 = [(5, 7), ["x",0], ["y", -1]]
l3 = list(zip(l1,l2))
print(l3)
# -
l3
for elem in l3:
print(elem)
for e1, e2 in l3:
print(e1)
print("--------")
for t1 in e1:
print(t1)
# +
# Este for da el error en la línea del segundo for ya que se le está pidiendo t1, t2 cuando solo se está recorriendo un elemento que tiene un elemento.
l1 = [([['a'], ['b']], (5, 7)), ([3, 2], ['x', 0]), ((7, 'm'), ['y', -1])]
for e1, e2 in l1:
print(e1)
print("--------")
for t1, t2 in e1:
print(t1)
# -
# ### Zip para generar diccionarios
d1 = {"clave":3, "k":"j", 2:["8", "x"]}
d1
# +
l1 = ["a", 3, 7]
l2 = ["b", 2, 0]
d3 = dict(zip(l1, l2))
d3
# +
# Añadir un elemento (clave, valor) a un diccionario
d = {}
d["key"] = 2
d
# -
d["key"] = 4
d
d[2] = [8, 4]
d
d[2] = [8, 4, 6]
d
d[2].append("x")
d
k = 2, 4
k
d[4] = [2, 5, 9], (8,"a", "b")
d
d[4] = list(d[4])
d[4].append(4)
d
d[4][1] = list(d[4][1])
d
d[4] = tuple(d[4])
d
# +
# Zip trata de crear tantos elementos como el número de elementos que tenga la colección con menos elementos contando con todas las colecciones.
l1 = ["x", "y"]
l2 = [0, 1]
l3 = (True, False)
l4 = [300, 5000, 999999]
s = "abdf"
l5 = list(zip(l1, l2, l3, l4, s))
l5
# -
e1, e2, e3, e4, e5 = 1, 2, 3, 4, 5
e5
for e1, e2, e3, e4, e5 in l5:
print(e5)
for pos, elem in enumerate(l5):
print(pos)
print(elem)
for pos, (e1, e2, e3, e4, e5) in enumerate(l5):
print(pos)
print(e3)
print("------")
l4 = {6, 5000, 999999}
l4
# ### List/Dict comprehesion
list(range(21))
l = [3]
for i in range(21):
l.append(i)
l
p = 0
l = [p for elem in [2, 5, 9] ]
l
p = 0
l = [elem for elem in [2, 5, 9] if elem > 4]
l
p = 0
l = [elem for elem in [2, 5., 9] if isinstance(elem, int)]
l
p = 0
l = [elem for elem in [2, 5., 9] if isinstance(elem, int)]
l
p = 0
l = [elem/2 for elem in [2, 5., 9, 17] if isinstance(elem, int)]
l
type("x") == int
p = 1
l = [(elem/2) + p for elem in [2, 5., 9, 17, "x"] if isinstance(elem, int)]
l
# +
o = [2, 5., 9, 17, "x"]
l = [(elem/2) + p for elem in o if isinstance(elem, int)]
k = []
for elem in o:
if isinstance(elem, int):
k.append((elem/2) + p)
k
# +
o = [2, 5., 9, 17, "x"]
l = [(elem/2) + p for elem in o if isinstance(elem, int)]
k = [2.0]
p = 1
def is_int(elem):
return type(elem) == int
for elem in o:
if is_int(elem=elem):
k.append((elem/2) + p)
# +
dict1 = {'a': 1, 'b': 2, 'c': 3, 'd': 4, 'e': 5}
for (k,v) in dict1.items():
print("Clave:", k)
print("Valor:", v)
print("------")
# +
dict1 = {'a': 1, 'b': 2, 'c': 3, 'd': 4, 'e': 5}
double_dict1 = {(k*2):(v*2) for (k,v) in dict1.items()}
print(double_dict1)
# +
dict1 = {'a': 1, 'b': 2, 'c': 3, 'd': 4, 'e': 5}
double_dict1 = {(k*2):(v*2) for (k,v) in dict1.items() if v > 2}
print(double_dict1)
# -
# ### Recursividad
# +
def f1(cont):
return cont + 1
r = f1(cont=2)
r
# +
import time
def p():
print("Hola")
time.sleep(2)
p()
p()
# +
def f1(cont):
if cont == 0:
print("CASO BASE")
print("Valor de cont la última vez:", cont)
return cont
else:
print("Valor de cont:", cont)
f1(cont=cont-1)
r = f1(cont=2)
print(r)
# +
def f1(cont):
if cont == 0:
print("CASO BASE")
print("Valor de cont la última vez:", cont)
return cont
else:
print("Valor de cont:", cont)
return f1(cont=cont-1)
r = f1(cont=2)
r
# +
l = []
for i in range(6):
l.append(i)
print(l)
# -
lc = [i for i in range(6)]
lc
# +
def fn(k=None):
if k: # True, 1 == 1, [1], 4
return []
else: # None, False, conjunto vacío, 0
return [i for i in range(6)]
p = fn()
print(p)
# +
def ct(val):
if val > 5:
return True
else:
return False
lc = [x for x in range(10) if ct(val=x)]
lc
# -
for i in range(6):
if ct(val=i):
print("Mayor que 5")
else:
print("Menor que 5")
# +
def ct(val, limite):
a_retornar = False
if val > limite:
a_retornar = True
return a_retornar
def main(lista, limite):
for x in lista:
if ct(val=x, limite=limite):
return [i for i in range(x) if i < 5]
l = list(range(10))
limite = 8
print(main(lista=l, limite=limite))
# -
ct(val=1000, limite=100)
def ct(val, limite):
if val > limite:
return True
# +
l = ["st4", 3, [0.1, 1], (8, 6), {2:"valor"}, {9, "s"}]
lista_numerica = []
for x in l:
if isinstance(x, (int, float)):
lista_numerica.append(x)
else:
if isinstance(x, dict):
for k, v in x.items():
if isinstance(k, (int, float)):
lista_numerica.append(k)
if isinstance(v, (int, float)):
lista_numerica.append(v)
else: # x es una colección
if isinstance(x, str):
continue
else: # colección no str
for elem in x:
if isinstance(elem, (int, float)):
lista_numerica.append(elem)
lista_numerica
# +
l = ["st4", 3, [0.1, 1], (8, 6), {2:"valor"}, {9, "s"}]
lista_numerica = []
def check_and_add(elem, types, lista):
if isinstance(elem, types):
lista.append(elem)
for x in l:
if isinstance(x, (int, float)):
lista_numerica.append(x)
else:
if isinstance(x, dict):
for k, v in x.items():
if isinstance(k, (int, float)):
lista_numerica.append(k)
if isinstance(v, (int, float)):
lista_numerica.append(v)
else: # x es una colección
if isinstance(x, str):
continue
else: # colección no str
for elem in x:
if isinstance(elem, (int, float)):
lista_numerica.append(elem)
# +
l = ["st4", 3, [0.1, 1], (8, 6), {2:"valor"}, {9, "s"}]
lista_numerica = []
def check_and_add(elem, types, lista):
if isinstance(elem, types):
lista.append(elem)
for x in l:
if isinstance(x, (int, float)):
lista_numerica.append(x)
else:
if isinstance(x, dict):
for k, v in x.items():
check_and_add(k, (int, float), lista_numerica)
check_and_add(v, (int, float), lista_numerica)
else: # x es una colección
if isinstance(x, str):
continue
else: # colección no str
for elem in x:
check_and_add(elem, (int, float), lista_numerica)
# -
d = {2:"valor"}
list(d.items())
d = {2:"valor"}
for k, v in d.items():
print(k)
print(v)
# ### Lambda
# +
def nombre_f(d, a):
return d + 2
nombre_f(d=2, a=None)
# +
nombre_g = lambda d, a: (d + 2)/a
nombre_g(d=2, a=5)
# +
def nombre_f(k):
return k
nombre_f = lambda k: k
nombre_f(k=2)
# +
def nombre_f(k):
if k > 0:
return k
else:
return 2
# Si escribimos una condición durante la creación de una función lambda, necesitamos tanto el if como else obligatoriamente
nombre_f = lambda k: k if k > 0 else 2
r = nombre_f(k=4)
print(r)
# +
def nombre_f(k):
if k > 0:
return k
else:
return 2
# Si escribimos una condición durante la creación de una función lambda, necesitamos tanto el if como else obligatoriamente
lambda_f = lambda d: print(d+2)
def f1(d):
print(d+2)
return d
"""
def nombre_f(k):
if k > 0:
return f1(d=k)
else:
return 2
"""
nombre_f = lambda k: f1(d=k) if k > 0 else 2
r = nombre_f(k=4)
print(r)
# +
def is_higher_5(elem):
if elem > 5:
return elem
l = [i for i in range(10)]
l
# -
l = ["st4", 3, [0.1, 1], (8, 6), {2:"valor"}, {9, "s"}]
lista_numerica = [x for x in l if isinstance(x, (int, float)) for y in l if isinstance(y, (list, tuple, set)) for x in y if isinstance(x, (int, float))]
lista_numerica
# ## Else list comprehesion
l = [x if x > 0 else 2 for x in range(5)]
l
| week3/day1/Python_IX_Precurse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import pandas as pd
from codex import CodexKg
#Load csv data
df = pd.read_csv("sample_data/tech_companies.csv")
# -
df
# +
#Make new codex object
codexkg = CodexKg()
#Create new keyspace
codexkg.create_db("tech_example")
# -
#Load data into Grakn
codexkg.create_entity(df, "Company", entity_key="name")
# +
# Find Company that has a name equal to Google
ans = codexkg.find(
concept="Company",
concept_attrs=["name"],
concept_conds=["equals"],
concept_values=["Google"],
)
#Display data as a DataFrame
logging.info(ans)
# {'Company': name budget
# 0 Google 999.99}
# +
ans = codexkg.find(
concept="Company",
concept_attrs=["name"],
concept_conds=["equals"],
concept_values=["Apple"],
)
#Display data as a DataFrame
logging.info(ans)
# {'Company': name budget
# 0 Google 999.99}
# -
ans['Company']
| codex_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import math
from scratch.probability import normal_cdf, binomial, normal_pdf, inverse_normal_cdf
import matplotlib.pyplot as plt
import seaborn as sns
import random
# + [markdown] heading_collapsed=true
# # Coin flipping Example
# + [markdown] hidden=true
# Null hypothesis is default assumption i.e. coin is fair P(Head) = p = 0.5
#
# Alternative hypothesis is p!=5
#
# To find fairness of coin, we will do experiment where we flip coin n=1000 times and assumed p = 0.5.
#
# Counting the number of heads X is a random variable of Binomial(n,p)
#
# **Note:** We can never accept null hypothesis. We either reject them or "fail to reject" them.
# + [markdown] hidden=true
# ### Binomial $\approx$ Normal for large n
# + hidden=true
n = 1000
p = 0.5
# Output of 20 such experiments
n_exp = 20
#expected number of heads
E_binomial = sum([binomial(n,p) for i in range(n_exp)])/n_exp
print(E_binomial)
# + [markdown] hidden=true
# Binomial(n,p) can be approximated using Normal of mu = p*n and sigma = sqrt(p*(p-1)*n)
# + hidden=true
def normal_approximation_to_binomial(n, p):
"""finds mu and sigma corresponding to a Binomial(n, p)"""
mu = p * n
sigma = math.sqrt(p * (1 - p) * n)
return mu, sigma
mu, sigma = normal_approximation_to_binomial(n,p)
mu,sigma
# + hidden=true
# expected number of heads
E_normal = inverse_normal_cdf(p,mu,sigma)
E_normal
# + hidden=true
percent_diff = abs(E_binomial - E_normal)*100.0/E_normal
percent_diff
# + [markdown] hidden=true
# As you can see above E_normal $\approx$ E_binomial when n is large
# + hidden=true
import pandas
n_list = []
percent_diff_list = []
for n in [10,100,1000,10000,100000]:
p = 0.5
# Output of 20 such experiments
n_exp = 20
#expected number of heads
E_binomial = sum([binomial(n,p) for i in range(n_exp)])/n_exp
mu, sigma = normal_approximation_to_binomial(n,p)
E_normal = inverse_normal_cdf(p,mu,sigma)
percent_diff = abs(E_binomial - E_normal)*100.0/E_normal
n_list.append(n)
percent_diff_list.append(percent_diff)
plot_df = pandas.DataFrame({'n':n_list,'percent_diff':percent_diff_list})
plot_df
# + hidden=true
sns.scatterplot(x='n',y='percent_diff',data=plot_df)
# + [markdown] hidden=true
# ## Decision making
#
# * The first step is to state the relevant null and alternative hypotheses.
# * Null hypothesis is p=0.5
# * alternate hypothesis is p$\neq$0.5
# * Select a significance level (α), a probability threshold below which the null hypothesis will be rejected. Common values are 5% and 1%.
# * Lets take 5% significance level i.e. 5% type 1 error (false positive error) when we incorrectly reject null hypothesis
# * i.e. We "fail to reject" null hypothesis with 95% confidence
#
# **Note:** A "negative" is a decision in favor of the null hypothesis and a "positive" is a decision in favor of the alternative hypothesis.
#
#
# Hence, type 1 error = incorrectly reject null hypothesis = False positive
#
# where, Reject null hypothesis = "positive" decision in favor of the alternative hypothesis.
#
# Similarly:
#
# type 2 error = incorrectly "fail to reject" null hypothesis = False negative
#
# where, "fail to reject" null hypothesis = decision in favor of the null hypothesis
#
#
# + [markdown] hidden=true
# 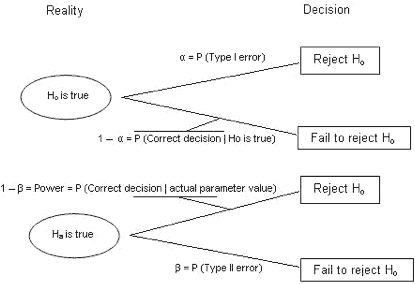
# + [markdown] hidden=true
# ## Significance level of test
# + hidden=true
# the normal cdf _is_ the probability the variable is below a threshold
normal_probability_below = normal_cdf
# it's above the threshold if it's not below the threshold
def normal_probability_above(lo, mu=0, sigma=1):
return 1 - normal_cdf(lo, mu, sigma)
# it's between if it's less than hi, but not less than lo
def normal_probability_between(lo, hi, mu=0, sigma=1):
return normal_cdf(hi, mu, sigma) - normal_cdf(lo, mu, sigma)
# it's outside if it's not between
def normal_probability_outside(lo, hi, mu=0, sigma=1):
return 1 - normal_probability_between(lo, hi, mu, sigma)
# + hidden=true
def normal_upper_bound(probability, mu=0, sigma=1):
"""returns the z for which P(Z <= z) = probability"""
return inverse_normal_cdf(probability, mu, sigma)
def normal_lower_bound(probability, mu=0, sigma=1):
"""returns the z for which P(Z >= z) = probability"""
return inverse_normal_cdf(1 - probability, mu, sigma)
def normal_two_sided_bounds(probability, mu=0, sigma=1):
"""returns the symmetric (about the mean) bounds
that contain the specified probability"""
tail_probability = (1 - probability) / 2
# upper bound should have tail_probability above it
upper_bound = normal_lower_bound(tail_probability, mu, sigma)
# lower bound should have tail_probability below it
lower_bound = normal_upper_bound(tail_probability, mu, sigma)
return lower_bound, upper_bound
# + hidden=true
n = 1000
p = 0.5
mu_0, sigma_0 = normal_approximation_to_binomial(1000, 0.5)
mu_0, sigma_0
# + hidden=true
# 5% error that coin is fair, hence probability is 0.95
lo, hi= normal_two_sided_bounds(0.95, mu_0, sigma_0)
lo,hi
# + [markdown] hidden=true
# if number of heads is in range [lo, hi] then we "fail to reject" H0/"null hypothesis" with 95% confidence or 5% error.
#
# Confidence interval of [lo,hi] for significance of 5%.
# + [markdown] hidden=true
# Assuming p really equals 0.5 (i.e., H0 is true), there is just a 5% chance we observe an
# X that lies outside this interval, which is the exact significance we wanted. Said differently,
# if H0 is true, then, approximately 19 times out of 20, this test will give the correct
# result.
# + [markdown] hidden=true
# ## Power of test
# + [markdown] hidden=true
# **Power** of a test, which is the probability of not making a type 2 error, in which we fail to reject H0 even though it’s false.
# + [markdown] hidden=true
# Power = 1 - $\beta$, where $\beta$ = P(type 2 error)
#
# type 2 error = false negative = "fail to reject H0" when H0 is False
# + [markdown] hidden=true
# Since H0 is False let actual p=0.55. Note we had assumed H0 of p=0.5
# + hidden=true
mu_0, sigma_0 = normal_approximation_to_binomial(n=1000,p=0.5) #since we assumed p=0.5
lo, hi = normal_two_sided_bounds(0.95, mu_0, sigma_0)
lo,hi
# + hidden=true
# a type 2 error means we fail to reject the null hypothesis
# which will happen when X is still in our original interval
"""Probability that number of heads is in range [lo,hi] calculated by assuming p=0.5
but actually p!=0.5, p=0.55 = type_2_probability"""
# actual mu and sigma based on p = 0.55
mu_1, sigma_1 = normal_approximation_to_binomial(1000, 0.55)
type_2_probability = normal_probability_between(lo, hi, mu_1, sigma_1)
type_2_probability
# + hidden=true
power = 1 - type_2_probability
power
# + [markdown] hidden=true
# ## P-value
#
# **P-value:** P-value is the probability of obtaining results as extreme as the observed results of a statistical hypothesis test, assuming that the null hypothesis is correct. A smaller p-value means that there is stronger evidence in favor of the alternative hypothesis.
#
# For our example of H0: p=0.5
# Lets assume H0 is true than number of heads should be in confidence interval of [lo,hi] = [469.01, 530.98]
# Let observed value is 530
#
# For our two-sided test:
# + hidden=true
def two_sided_p_value(x, mu=0, sigma=1):
if x >= mu:
# if x is greater than the mean, the tail is what's greater than x
return 2 * normal_probability_above(x, mu, sigma)
else:
# if x is less than the mean, the tail is what's less than x
return 2 * normal_probability_below(x, mu, sigma)
"""Why did we use 529.5 instead of 530? This is what’s called a continuity correction.
It reflects the fact that normal_probability_between(529.5, 530.5, mu_0, sigma_0)"""
two_sided_p_value(529.5, mu_0,sigma_0)
# + [markdown] hidden=true
# One way to convince yourself that this is a sensible estimate is with a simulation, as shown below
# + hidden=true
extreme_value_count = 0
for _ in range(100000):
"""
count # of heads in 1000 flips
and count how often the # is 'extreme'
i.e.
"""
num_heads = sum(1 if random.random() < 0.5 else 0 for _ in range(1000))
if num_heads >= 530 or num_heads <= (mu_0-(530-mu_0)):
extreme_value_count += 1
print(extreme_value_count / 100000) # 0.062
# + [markdown] hidden=true
# Since the p-value is greater than our 5% significance, we don’t reject the null.
#
# If we instead saw 532 heads, the p-value would be
# + hidden=true
two_sided_p_value(531.5, mu_0, sigma_0)
# + [markdown] hidden=true
# Which is smaller than the 5% significance, which means we would reject the null. It’s
# the exact same test as before. It’s just a different way of approaching the statistics.
# + [markdown] heading_collapsed=true
# # Coin flipping example 2
#
# null hypothesis is coin is not biased toward heads, or that p ≤ 0 . 5. Let significance level is 5%
# + hidden=true
hi = normal_upper_bound(0.95, mu_0, sigma_0) # 0.95 since significance level is 5%
hi
# + [markdown] hidden=true
# If number of heads if less equal to than 526 we decide in favor of null hypothesis with confidence of 95%.
# + hidden=true
#Actually p = 0.55
mu_1, sigma_1 = normal_approximation_to_binomial(1000, 0.55)
# type_2_probability = incorrectly decide in favour of null hypothesis i.e. Number of heads is less than hi = 526
type_2_probability = normal_probability_below(hi,mu_1, sigma_1)
power = 1 - type_2_probability
power
# + [markdown] hidden=true
# Coin flipping example 2 is more powerful test
# + [markdown] hidden=true
# ## P-value
# + [markdown] hidden=true
# This is one sided test hence if observed value is 525 than
# + hidden=true
upper_p_value = normal_probability_above
lower_p_value = normal_probability_below
p_value = upper_p_value(524.5, mu_0, sigma_0) # 0.061
p_value
# -
# # Confidence Intervals
# If we have seen 540 heads
p_hat = 540 / 1000
mu = p_hat
sigma = math.sqrt(p_hat * (1 - p_hat) / 1000) # 0.0158
normal_two_sided_bounds(0.95, mu, sigma) # [0.5091, 0.5709]
# # P-hacking
def run_experiment():
"""flip a fair coin 1000 times, True = heads, False = tails"""
return [random.random() < 0.5 for _ in range(1000)]
def reject_fairness(experiment):
"""using the 5% significance levels"""
num_heads = len([flip for flip in experiment if flip])
return num_heads < 469 or num_heads > 531
random.seed(0)
experiments = [run_experiment() for _ in range(1000)]
num_rejections = len([experiment
for experiment in experiments
if reject_fairness(experiment)])
num_rejections
num_rejections*100.0/1000
# # Example: Running an A/B Test
def estimated_parameters(N, n):
p = n / N
sigma = math.sqrt(p * (1 - p) / N)
return p, sigma
def a_b_test_statistic(N_A, n_A, N_B, n_B):
p_A, sigma_A = estimated_parameters(N_A, n_A)
p_B, sigma_B = estimated_parameters(N_B, n_B)
return (p_B - p_A) / math.sqrt(sigma_A ** 2 + sigma_B ** 2)
z = a_b_test_statistic(1000, 200, 1000, 180)
z
two_sided_p_value(z)
z = a_b_test_statistic(1000, 200, 1000, 150) # -2.94
two_sided_p_value(z)
| DS_scratch/notebooks/07_Hypothesis_and_Inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pose Estimation
#
# _You can view [IPython Nootebook](README.ipynb) report._
#
# ----
#
# ## Contents
#
# - [GOAL](#GOAL)
# - [Basics](#Basics)
# - [Render the Axes](#Render-the-Axes)
# - [Render a Cube](#Render-a-Cube)
#
# ## GOAL
#
# In this section:
#
# - We will learn to exploit calib3d module to create some 3D effects in images.
#
# ## Basics
#
# ### Render the Axes
#
# This is going to be a small section. During the last session on camera calibration, you have found the camera matrix, distortion coefficients etc. Given a pattern image, we can utilize the above information to calculate its pose, or how the object is situated in space, like how it is rotated, how it is displaced etc. For a planar object, we can assume Z=0, such that, the problem now becomes how camera is placed in space to see our pattern image. So, if we know how the object lies in the space, we can draw some 2D diagrams in it to simulate the 3D effect. Let's see how to do it.
#
# Our problem is, we want to draw our 3D coordinate axis (X, Y, Z axes) on our chessboard's first corner. X axis in blue color, Y axis in green color and Z axis in red color. So in-effect, Z axis should feel like it is perpendicular to our chessboard plane.
#
# First, let's load the camera matrix and distortion coefficients from the previous calibration result.
#
# ```python
# import numpy as np
# import cv2 as cv
# import glob
#
# # Load previously saved data
# with np.load('B.npz') as X:
# mtx, dist, _, _ = [X[i] for i in ('mtx', 'dist', 'rvecs', 'tvecs')]
# ```
#
# Now let's create a function, draw which takes the corners in the chessboard (obtained using [cv.findChessboardCorners()](https://docs.opencv.org/3.4.1/d9/d0c/group__calib3d.html#ga93efa9b0aa890de240ca32b11253dd4a)) and axis points to draw a 3D axis.
#
# ```python
# def draw(img, corners, imgpts):
# corner = tuple(corners[0].ravel())
# img = cv.line(img, corner, tuple(imgpts[0].ravel()), (255,0,0), 5)
# img = cv.line(img, corner, tuple(imgpts[1].ravel()), (0,255,0), 5)
# img = cv.line(img, corner, tuple(imgpts[2].ravel()), (0,0,255), 5)
# return img
# ```
#
# Then as in previous case, we create termination criteria, object points (3D points of corners in chessboard) and axis points. Axis points are points in 3D space for drawing the axis. We draw axis of length 3 (units will be in terms of chess square size since we calibrated based on that size). So our X axis is drawn from (0,0,0) to (3,0,0), so for Y axis. For Z axis, it is drawn from (0,0,0) to (0,0,-3). Negative denotes it is drawn towards the camera.
#
# ```python
# criteria = (cv.TERM_CRITERIA_EPS + cv.TERM_CRITERIA_MAX_ITER, 30, 0.001)
# objp = np.zeros((6*7,3), np.float32)
# objp[:, :2] = np.mgrid[0:7, 0:6].T.reshape(-1, 2)
# axis = np.float32([[3, 0, 0], [0, 3, 0], [0, 0, -3]]).reshape(-1, 3)
# ```
#
# Now, as usual, we load each image. Search for 7x6 grid. If found, we refine it with subcorner pixels. Then to calculate the rotation and translation, we use the function, [cv.solvePnPRansac()](https://docs.opencv.org/3.4.1/d9/d0c/group__calib3d.html#ga50620f0e26e02caa2e9adc07b5fbf24e). Once we those transformation matrices, we use them to project our axis points to the image plane. In simple words, we find the points on image plane corresponding to each of (3,0,0), (0,3,0), (0,0,3) in 3D space. Once we get them, we draw lines from the first corner to each of these points using our draw() function. Done !!!
#
# ```python
# for fname in glob.glob("../../data/left*.jpg"):
# print(fname)
# img = cv.imread(fname)
# gray = cv.cvtColor(img,cv.COLOR_BGR2GRAY)
# ret, corners = cv.findChessboardCorners(gray, (7, 6), None)
# if ret is True:
# corners2 = cv.cornerSubPix(gray, corners, (11, 11), (-1, -1), criteria)
#
# # Find the rotation and translation vectors.
# ret, rvecs, tvecs = cv.solvePnP(objp, corners2, mtx, dist)
#
# # Project 3D points to image plane
# imgpts, jac = cv.projectPoints(axis, rvecs, tvecs, mtx, dist)
# img = draw(img, corners2, imgpts)
# cv.imshow("Image - {}".format(str(fname[11:])), img)
# k = cv.waitKey(5000) & 0xFF
# if k == 27: # Press "Esc" to exit
# break
# elif k == ord('n'): # Press 'n' to open next image
# cv.destroyWindow("Image - {}".format(str(fname[11:])))
# elif k == ord('s'): # Press 's' to save current image
# cv.destroyWindow("Image - {}".format(str(fname[11:])))
# cv.imwrite("output-files/" + "axis-" + str(fname[11:17]) +
# ".png", img)
# cv.destroyAllWindows()
# ```
#
# See some results below. Notice that each axis is 3 squares long:
#
# 
#
# ### Render a Cube
#
# If you want to draw a cube, modify the draw() function and axis points as follows.
#
# Modified draw() function:
#
# ```python
# def draw(img, corners, imgpts):
# imgpts = np.int32(imgpts).reshape(-1, 2)
#
# # Draw ground floor in green
# img = cv.drawContours(img, [imgpts[:4]], -1, (0, 255, 0), -3)
#
# # Draw pillars in blue color
# for i, j in zip(range(4), range(4, 8)):
# img = cv.line(img, tuple(imgpts[i]), tuple(imgpts[j]), (255), 3)
#
# # Draw top layer in red color
# img = cv.drawContours(img, [imgpts[4:]], -1, (0, 0, 255), 3)
# return img
# ```
#
# Modified axis points. They are the 8 corners of a cube in 3D space:
#
# ```python
# axis = np.float32([[0, 0, 0], [0, 3, 0], [3, 3, 0], [3, 0, 0],
# [0, 0, -3], [0, 3, -3], [3, 3, -3], [3, 0, -3]])
# ```
#
# And look at the result below:
#
# 
#
# If you are interested in graphics, augmented reality etc, you can use OpenGL to render more complicated figures.
| calibration-reconstruction/pose-estimation/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 4교시 조인 연산
#
# ### 목차
# * [1. 조인 유형](#1.-조인-유형)
# * [2. Inner Join](#2.-Inner-Join)
# * [3. Outer Join](#3.-Outer-Join)
# * [4. Left Semi Join](#3.-Outer-Join)
# * [5. Left Anti Join](#3.-Outer-Join)
# * [6. Natural Join](#3.-Outer-Join)
# * [7. Cross Join - Cartesian Join](#7.-Cross-Join---Cartesian-Join)
# * [8. 조인시 유의할 점](#4.-조인시-유의할-점)
# * [참고자료](#참고자료)
# +
from pyspark.sql import *
from pyspark.sql.functions import *
from pyspark.sql.types import *
from IPython.display import display, display_pretty, clear_output, JSON
spark = (
SparkSession
.builder
.config("spark.sql.session.timeZone", "Asia/Seoul")
.getOrCreate()
)
# 노트북에서 테이블 형태로 데이터 프레임 출력을 위한 설정을 합니다
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # display enabled
spark.conf.set("spark.sql.repl.eagerEval.truncate", 100) # display output columns size
# 공통 데이터 위치
home_jovyan = "/home/jovyan"
work_data = f"{home_jovyan}/work/data"
# work_dir=!pwd
work_dir = work_dir[0]
# 로컬 환경 최적화
spark.conf.set("spark.sql.shuffle.partitions", 5) # the number of partitions to use when shuffling data for joins or aggregations.
spark.conf.set("spark.sql.streaming.forceDeleteTempCheckpointLocation", "true")
spark
# -
# ## 1. 조인 유형
#
# + 스파크의 조인 타입
# + inner join, outer join, left outer join, right outer join
# + left semi join: 왼쪽 데이터셋의 키가 오른쪽 데이터셋에 있는 경우에는 키가 일치하는 왼쪽 데이터셋만 유지
# + left anti join: 왼쪽 데이터셋의 키가 오른쪽 데이터셋에 없는 경우에는 키가 일치하지 않는 왼쪽 데이터셋만 유지
# + natural join: 두 데이터셋에서 동일한 이름을 가진 컬럼을 암시적으로 결합하는 조인을 수행
# + cross join: 왼쪽 데이터셋의 모든 로우와 오른쪽 데이터 셋의 모든 로우를 조합
#
# 
# ### JOIN 학습을 위해 상품은 단 하나만 구매할 수 있다고 가정하여 아래와 같은 테이블이 존재합니다
# #### 정보 1. 고객은 4명이지만, 1명은 탈퇴하여 존재하지 않습니다
# | 고객 아이디 (u_id) | 고객 이름 (u_name) | 고객 성별 (u_gender) |
# | - | - | - |
# | 1 | 정휘센 | 남 |
# | 2 | 김싸이언 | 남 |
# | 3 | 박트롬 | 여 |
#
# #### 정보 2. 구매 상품은 3개이며, 탈퇴한 고객의 상품정보가 남아있습니다
# | 구매 고객 아이디 (u_id) | 구매 상품 이름 (p_name) | 구매 상품 가격 (p_amount) |
# | - | - | - |
# | 2 | LG DIOS | 2,000,000 |
# | 3 | LG Cyon | 1,800,000 |
# | 4 | LG Computer | 4,500,000 |
#
# +
user = spark.createDataFrame([
(1, "정휘센", "남"),
(2, "김싸이언", "남"),
(3, "박트롬", "여")
]).toDF("u_id", "u_name", "u_gender")
user.printSchema()
display(user)
purchase = spark.createDataFrame([
(2, "LG DIOS", 2000000),
(3, "LG Cyon", 1800000),
(4, "LG Computer", 4500000)
]).toDF("p_uid", "p_name", "p_amont")
purchase.printSchema()
display(purchase)
# -
# ## 2. Inner Join
#
# > join type 을 명시하지 않았을 때 기본적으로 내부조인을 수행함
#
# * 왼쪽 테이블과 오른쪽 테이블에서 동일한 칼럼을 가져야함.
# * 참으로 평가되는 로우만 결합
# * 세 번째 파라미터로 조인 타입을 명확하게 지정할 수 있음
# * 두 테이블의 키가 중복되거나 여러 복사본으로 있다면 성능이 저하됨. 조인이 여러 키를 최소화하기 위해 일종의 카테시안 조인으로 변환되어 버림
#
# ### 2.1 구매 정보와 일치하는 고객 정보를 조인 (inner)
display(user.join(purchase, user.u_id == purchase.p_uid))
user.join(purchase, user.u_id == purchase.p_uid, "inner").count()
# ### <font color=green>1. [기본]</font> 고객 정보 f"{work_data}/tbl_user", 제품 정보 f"{work_data}/tbl_purchase" CSV 파일을 읽고
# #### 1. 각각 스키마를 출력하세요
# #### 2. 각각 데이터를 출력하세요
# #### 3. 고객(tbl_user) 테이블의 u_id 와 제품(tbl_purchase) 테이블의 p_uid 는 고객아이디 입니다
# #### 4. 고객 테이블을 기준으로 어떤 제품을 구매하였는지 inner join 을 통해 조인해 주세요
# #### 5. 조인된 최종 테이블의 스키마와 데이터를 출력해 주세요
#
# <details><summary>[실습1] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_user.csv")
# )
# left.printSchema()
# left.show()
#
# right = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_purchase.csv")
# )
# right.printSchema()
# right.show()
#
# join_codition = left.u_id == right.p_uid
# answer = left.join(right, join_codition, "inner")
# answer.printSchema()
# display(answer)
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ## 3. Outer Join
#
# > 왼쪽과 오른쪽의 모든 로우를 제공.
#
# * 왼쪽이나 오른쪽 DataFrame에 일치하는 로우가 없다면 해당 위치에 null을 삽입
# * 공통 로우가 거의 없는 테이블에서 사용하면 결과값이 매우 커지고 성능 저하
#
# ### 3.1 모든 고객의 정보 구매 정보를 조인 (left_outer)
user.join(purchase, user.u_id == purchase.p_uid, "left_outer").orderBy(purchase.p_uid.asc())
# ### 3-2. 모든 상품에 대한 고객 정보를 조인 (right_outer)
user.join(purchase, user.u_id == purchase.p_uid, "right_outer").orderBy(purchase.p_uid.asc())
# ### 3-3. 모든 고객과 상품에 대한 정보를 조인 (full_outer)
user.join(purchase, user.u_id == purchase.p_uid, "full_outer").orderBy(purchase.p_uid.asc())
# ## 4. Left Semi Join
#
# > 오른쪽에 존재하는 것을 기반으로 왼쪽 로우만 제공
#
# * 값이 존재하는 지 확인 용도, 값이 있다면 왼쪽 DataFrame에 중복 키가 있더라도 해당 로우는 결과에 포함
# * 기존 조인 기능과는 달리 DataFrame의 필터 기능과 유사
# * 하나의 테이블만 확실히 고려되고, 다른 테이블은 조인 조건만 확인하기 때문에 성능이 매우 좋음
#
user.join(purchase, user.u_id == purchase.p_uid, "left_semi").orderBy(user.u_id.asc())
# ## 5. Left Anti Join
# + 왼쪽 세미 조인의 반대 개념, 즉 오른쪽 DataFrame의 어떤 값도 포함하지 않음
# + SQL의 NOT IN과 같은 스타일의 필터
user.join(purchase, user.u_id == purchase.p_uid, "left_anti").orderBy(user.u_id.asc())
# ## 6. Natural Join
# > 조인하려는 컬럼을 암시적으로 추정
#
# * 암시적인 처리는 언제나 위험하므로 비추천
# * Python join 함수는 이 기능을 지원하지 않음
#
user.createOrReplaceTempView("user")
purchase.createOrReplaceTempView("purchase")
spark.sql("show tables")
user.printSchema()
purchase.printSchema()
# 지정된 필드의 값이 일치하는 경우 해당 필드를 기준으로 조인
spark.sql("SELECT * FROM user NATURAL JOIN purchase")
# ## 7. Cross Join - Cartesian Join
#
# > 교차 조인은 조건절을 기술하지 않은 내부 조인을 의미
# * 왼쪽의 모든 로우를 오른쪽의 모든 로우와 결합함(결과의 로우 수 = 왼쪽 로우 수 * 오른쪽 로우 수)
# * 큰 데이터에서 사용할 경우 out-of-memory exception 발생. 가장 좋지 않은 성능을 가진 조인. 주의해서 사용하며 특정 사례에서만 사용해야함.
""" 크로스 조인이지만 조건을 설정해야 하며, 조건에 부합된 결과를 출력하여 inner조인과 동일."""
joinExpr = user.u_id == purchase.p_uid
joinType = "cross"
user.join(purchase, on=joinExpr, how=joinType)
user.crossJoin(purchase)
# ### <font color=green>2. [기본]</font> 고객 정보 f"{work_data}/tbl_user", 제품 정보 f"{work_data}/tbl_purchase" CSV 파일을 읽고
# #### 1. 각각 스키마를 출력하세요
# #### 2. 각각 데이터를 출력하세요
# #### 3. 고객(tbl_user) 테이블의 u_id 와 제품(tbl_purchase) 테이블의 p_uid 는 고객아이디 입니다
# #### 4. 모든 상품을 기준으로 구매하 고객정보를 조인해 주세요 (left: purchase, right: user, join: left_outer)
# #### 5. 조인된 최종 테이블의 스키마와 데이터를 출력해 주세요
# #### 6. 출력시에 상품 가격의 내림차순으로 정렬해 주세요
#
# <details><summary>[실습2] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_purchase.csv")
# )
# left.printSchema()
# # left.show()
#
# right = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_user.csv")
# )
# right.printSchema()
# # right.show()
#
# join_codition = left.p_uid == right.u_id
# answer = left.join(right, join_codition, "left_outer")
# answer.printSchema()
# display(answer.orderBy(desc("p_amount")))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=blue>3. [중급]</font> 고객 정보 f"{work_data}/tbl_user", 제품 정보 f"{work_data}/tbl_purchase" CSV 파일을 읽고
# #### 1. 모든 고객을 기준으로 구매한 상품 정보를 조인해 주세요 (left: user, right: purchase, join: left_outer)
# * 고객(tbl_user) 테이블의 u_id 와 제품(tbl_purchase) 테이블의 p_uid 는 고객아이디 입니다
# * 조인된 최종 테이블의 스키마와 데이터를 출력해 주세요
# * 출력시에 상품 가격(tbl_purchase.p_amount)의 내림차순으로 정렬해 주세요
# * 상품가격이 없는 경우에는 등록일자(tbl_user.u_signup)가 최신으로 정렬해 주세요
# * 가능한 Structured API 를 사용하여 작성하되 최대한 간결하게 작성해 보세요
#
# <details><summary>[실습3] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = spark.read.csv(f"{work_data}/tbl_user.csv", inferSchema=True, header=True)
# right = spark.read.csv(f"{work_data}/tbl_purchase.csv", inferSchema=True, header=True)
# answer = left.join(right, left.u_id == right.p_uid, "left_outer")
# answer.printSchema()
# display(answer.orderBy(desc("p_amount"), desc("u_signup")))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=blue>4. [중급]</font> 고객 정보 f"{work_data}/tbl_user", 제품 정보 f"{work_data}/tbl_purchase" CSV 파일을 읽고
# #### 1. 모든 고객과 모든 상품 정보가 출력될 수 있도록 조인해 주세요
# * 고객(tbl_user) 테이블의 u_id 와 제품(tbl_purchase) 테이블의 p_uid 는 고객아이디 입니다
# * 조인된 최종 테이블의 스키마와 데이터를 출력해 주세요
# * 출력시에 상품 가격(tbl_purchase.p_amount)의 내림차순으로 정렬해 주세요 (단, null 이 먼저 출력 되도록)
# * 상품가격이 없는 경우에는 등록일자(tbl_user.u_signup)가 최신으로 정렬해 주세요 (단, null 이 마지막에 출력 되도록)
# * 가능한 Structured API 를 사용하여 작성하되 최대한 간결하게 작성해 주세요
#
# <details><summary>[실습4] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = spark.read.csv(f"{work_data}/tbl_user.csv", inferSchema=True, header=True)
# right = spark.read.csv(f"{work_data}/tbl_purchase.csv", inferSchema=True, header=True)
# answer = left.join(right, left.u_id == right.p_uid, "full_outer")
# answer.printSchema()
# display(answer.orderBy(desc_nulls_first("p_amount"), desc_nulls_last("u_signup")))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=green>5. [기본]</font> 아래의 조인 연산 결과에서 null 값에 대한 치환을 해주세요
# #### 1. 고객아이디(u_id)와 고객이름(u_name), 성별(u_gender), 가입일자(u_signup) 기본값을 채워주세요
# ##### u_id = 0, u_name = '미확인', u_gender = '미확인', u_signup = '19700101'
#
# <details><summary>[실습5] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_purchase.csv")
# )
# left.printSchema()
# # left.show()
#
# right = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load(f"{work_data}/tbl_user.csv")
# )
# right.printSchema()
# # right.show()
#
# join_codition = left.p_uid == right.u_id
# user_fill = { "u_id":0, "u_name":"미확인", "u_gender":"미확인", "u_signup":"19700101" }
# answer = left.join(right, join_codition, "left_outer").na.fill(user_fill)
# answer.printSchema()
# display(answer.orderBy(asc("u_signup")))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ## 8. 조인시 유의할 점
# +
u = spark.createDataFrame([
(1, "정휘센", "남"),
(2, "김싸이언", "남"),
(3, "박트롬", "여")
]).toDF("id", "name", "gender")
u.printSchema()
display(u)
p = spark.createDataFrame([
(2, "LG DIOS", 2000000),
(3, "LG Cyon", 1800000),
(4, "LG Computer", 4500000)
]).toDF("id", "name", "amount")
p.printSchema()
display(p)
# -
# ### 8.1 중복 컬럼명 처리가 되지 않은 경우
# > #### AnalysisException: "Reference 'id' is ambiguous, could be: id, id.;"
up = u.join(p, u.id == p.id)
up.show()
# up.select("id")
# ### 3.2 중복 컬럼명 해결방안 - 데이터 프레임의 컬럼 명을 다르게 만든다
# +
u1 = u.withColumnRenamed("id", "u_uid")
p1 = p.withColumnRenamed("id", "p_uid")
u1.printSchema()
p1.printSchema()
up = u1.join(p1, u1.u_uid == p1.p_uid)
display(up)
up.select("u_uid")
# -
# ### 3.3 중복 컬럼명 해결방안 - 조인 직후 중복 컬럼을 제거합니다
up = u.join(p, u.id == p.id).drop(p.id)
display(up)
display(up.select("id"))
# ### <font color=red>6. [고급]</font> 고객 정보 f"{work_data}/tbl_user_id", 제품 정보 f"{work_data}/tbl_purchase_id" CSV 파일을 읽고
# #### 1. 가장 비싼 `제품`을 구매한 고객의 고객정보와 제품정보를 출력해 주세요
# * **고객(tbl_user) 테이블의 아이디도 id 이고 제품(tbl_purchase) 테이블의 아이디도 id** 인 점을 주의 하세요
# * 최종 출력되는 컬럼은 고객아이디(u_id), 고객이름(u_name), 상품이름(p_name), 상품가격(p_amount) 이렇게 4개 컬럼입니다
# * 상품가격 (p_amount) 내림차순으로 정렬해 주세요
# * 가능한 Structured API 를 사용하여 작성하되 최대한 간결하게 작성해 보세요
#
# <details><summary>[실습6] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# left = spark.read.csv(f"{work_data}/tbl_purchase_id.csv", inferSchema=True, header=True)
# right = spark.read.csv(f"{work_data}/tbl_user_id.csv", inferSchema=True, header=True)
# u_left = left.withColumnRenamed("id", "u_id")
# u_right = right.withColumnRenamed("id", "p_uid")
# answer = u_left.join(u_right, u_left.u_id == u_right.p_uid, "inner").where("p_amount > 0").select("u_id", "u_name", "p_name", "p_amount")
# answer.printSchema()
# display(answer.orderBy(desc("p_amount")))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ## 9. 스파크의 조인 수행 방식
# ### 9.1 Broadcast Hash Join : **SmallSize** join **AnySize**
# - 큰 데이터셋과 이보다 작은 데이터 셋 간의 조인은 큰 데이터셋의 파티션이 있는 모든 익스큐터에 작은 데이터셋이 브로드캐스트되어 수행
# - 기본값: spark.sql.autoBroadcastJoinThreshold = 10mb
# - spark.sql.autoBroadcastJoinThreshold 는 조인을 수행 할 때 모든 작업자 노드에 브로드캐스트되는 테이블의 최대 크기를 구성
# - 참고
# - [Spark Doc - performance Tuning](https://spark.apache.org/docs/latest/sql-performance-tuning.html)
# - [Does spark.sql.autoBroadcastJoinThreshold work for joins using Dataset's join operator?](https://stackoverflow.com/questions/43984068/does-spark-sql-autobroadcastjointhreshold-work-for-joins-using-datasets-join-op)
# - 수행
# - 작은 테이블은 드라이버에 모였다가 다시 모든 노드에 복사
# - 작은 테이블은 메모리에 올라가고
# - 큰 테이블은 스트림을 통해서 조인 수행
# - 브로드캐스트 힌트 `person.join(broadcast(grudateProgram),Seq(“id”))`
# - `autoBroadcastJoinThreshold`에 관계없이 힌트가 있는 조인 측이 브로드캐스트됨.
# - 조인의 양쪽에 브로드캐스트 힌트가 있으면 실제 크기가 더 작은 쪽이 브로드 캐스트됨.
# - 힌트가 없고 테이블의 실제 물리적 추정값이 autoBroadcastJoinThreshold 보다 작으면, 해당 테이블은 모든 실행기 노드로 브로드캐스트됨.
# - 성능
# - 한 쪽의 데이터가 하나의 machine 에 fit-in-meory 될 정도로 작으면 성능 좋음.
# - table broadcast는 네트워크를 많이 사용하므로, 브로드캐스트 된 테이블이 크면 때때로 out-of-memory나 성능저하가 발생할 수 있음
# - 셔플링이 없기 때문에, 브로드캐스트 되는 쪽이 데이터가 작으면 다른 알고리즘보다 빠름.
# - 브로드캐스트 지원
# - Full outer join 은 지원하지 않음.
# - letf-outer join 에서는 오른쪽 테이블만 브로드캐스트, right-outer join 에서는 왼쪽 테이블만 브로드캐스트 가능.
# ### 9.2 Shuffle hash join : **MiddleSize** join **LargeSize**
# - 두 테이블 모두 shuffle 을 통해 노드에 분산되고
# - 비교적 작은 테이블이 메모리 버퍼에 올라가고
# - 큰 테이블은 스트림을 통해서 조인 수행
# - 파티션이 전체 익스큐터로 분배
# - 셔플은 비용이 많이 듦. 파티션과 셔플 배포가 최적으로 수행되는지 확인하기 위해 로직을 분석하는 것이 중요.
# - 큰 데이터는 join에 필요한 부분만 filtering 을 하거나
# - repartioning 을 고려해야함
# - Map Reduce Fundamentals 유사
# - Map - 두 개의 서로 다른 Data frames/table
# - Output key를 join 조건에서 필드로 사용
# - Shuflle - output key로 두 데이터 세트를 섞음
# - Reduce - join 결과
# 
# #### 성능을 최적화하려면
# - join할 키가 균등하게 dirstribute되어있거나,
# - parallelism위한 적절한 수의 키가 있을 때
#
# #### 성능이 나쁜 경우 - 고르지 않은 sharding 및 제한된 parallelism
# - data skewness 처럼 하나의 단일 파티션이 다른 파티션에 비해 너무 많은 데이터를 가지고 있을 때
# - 각 스테이트에서 50개 키만 셔플할 수 있음 -> 스파크 클러스터가 크면 고른 sharding과 parallelism 으로 해결 못함
# 
# ### 9.3 Sort merge join : **LargeSize** join **LargeSize**
# - 일치하는 조인키를 sort할 수 있고, 브로드캐스트 조인, 셔플 해시 조인에 적합하지 않은 경우 사용
# - shuffle hash join 과 비교했을 때, 클러스터에서 데이터 이동(shuffling)을 최소화함
# - 수행
# - 두 테이블 모두 셔플 및 정렬이 발생하고
# - 그나마 작은 쪽이 버퍼를 하고 큰 쪽이 스트리밍으로 조인을 수행한다
# - partition 은 join 작업 전에 조인키 정렬
# - 참고. [SortMergeJoinExec Binary Physical Operator for Sort Merge Join](https://jaceklaskowski.gitbooks.io/mastering-spark-sql/spark-sql-SparkPlan-SortMergeJoinExec.html)
# ### 9.4 BroadcastNestedLoopJoin
# - 적용 : 조인 키가 지정되어 있지 않고 브로드 캐스트 힌트가 있거나 조인의 한쪽이 브로드캐스트 될 수 있고, spark.sql.autoBroadcastJoinThreshold보다 작은 경우
# - 브로드캐스트 된 데이터 세트가 크면 매우 느릴 수 있으며 OutOfMemoryExceptions을 일으킬 수 있음
# ---
# ### 9.5 테이블 크기에 따른 조인 동작방식
# #### 큰 테이블과 큰 테이블 조인
# + 전체 노드 간 통신이 발생하는 셔플 조인이 발생됨
# #### 큰 테이블과 작은 테이블 조인
# + 작은 DataFrame을 클러스터 전체 워커에 복제한 후 통신없이 진행
# + 모든 단일 노드에서 개별적으로 조인이 수행되므로 CPU가 가장 큰 병목 구간이 됨
# + broadcast 함수(힌트)를 통해 브로드캐스트 조인을 설정할 수 있으나 강제할 수는 없음(옵티마이저가 무시 가능)
from pyspark.sql.functions import broadcast
user.join(broadcast(purchase), user.u_id == purchase.p_uid).select("*")
user.join(broadcast(purchase), user.u_id == purchase.p_uid).select("*").explain()
# #### 작은 테이블과 작은 테이블 조인
# + 스파크가 결정하도록 내버려두는 것이 제일 좋은 선택
# ### 9.6 고찰
# - 브로드캐스트조인을 가능한 한 사용하고 조인 전에 관련없는 행을 조인 키로 필터링하여 불필요한 데이터 셔플링을 피하자
# - 필요한 경우 spark.sql.autoBroadcastJoinThreshold를 적절히 조정
# - sort-merge join 이 default이고, 대부분의 시나리오에서 잘 수행됨.
# - Shuffle Hash 조인이 Sort-Merge 조인보다 낫다고 확신이 있으면,Sort-Merge join을 비활성화해서 shuffle hash join이 수행되도록 함.
# - builde size 가 stream size보다 작으면 Shuffle Hash 조인이 나음
# - unique한 조인키가 없거나 조인키가 없는 조인은 수행비용이 비싸므로 최대한 피해야함
# ---
# ## 10. 설명 이해 위한 배경 지식
# ### 10.1 파티셔닝 partioning
# - 어떤 데이터를 어디에 저장할 것인지 제어할 수 있는 기능
# - RDD는 데이터 파티션으로 구성되고 모든 연산은 RDD의 데이터 파티션에서 수행됨.
# - 파티션 개수는 RDD 트랜스포메이션 실행할 태스크 수에 직접적인 영향을 줌
# - 파티션 개수 너무 적으면 -> 많은 데이터에서 아주 일부의 CPU/코어만 사용 -> 성능 저하, 클러스터 제대로 활용 못함
# - 파티션 개수 너무 많으면 -> 실제 필요한 것보다 많은 자원을 사용 -> 멀티테넌트 환경에서는 자원 부족 현상 발생
# - Partioner 에 의해 RDD 파티셔닝이 실행된. 파티셔너는 파티션 인덱스를 RDD 엘리먼트에 할당.
# ### 10.2 Shuffling
# - 파티셔너가 어떤 파티션을 사용하든 많은 연산이 RDD의 파티션 전체에 걸쳐 데이터 리파티셔닝Repartioning 이 발생함
# - 새로운 파티션이 생성되거나 파티션이 축소, 병합될 수 있음.
# - 리파티셔닝에 필요한 모든 데이터 이동을 **셔플링 Shuffling**이라고 함.
# - **Shuffling 을 할 때, Disk I/O + Network I/O 과도하게 발생**
# - 셔플링은 계산을 동일 익스큐터의 메모리에서 더 이상 진행하지 않고 익스큐터 간에 데이터를 교환함 -> 많은 성능 지연을 초래할 수 있음
# - 스파크 잡의 실행 프로세스를 결정, 잡이 스테이지로 분할되는 부분에 영향을 미침
# - 셔플링이 많을 수록 스파크 잡이 실행될 때 더 많은 스테이지가 발생하기 때문에 성능에 영향을 미침
# - 리파티셔닝을 유발하는 연산은 **조인**, 리듀스, 그룹핑, 집계 연산이 있음
# ### 10.3 버켓팅 Bucketing
# - Bucketing = pre-(shffle + sort) inputs on join Keys
# - 각 파일에 저장된 데이터를 제어할 수 있는 또 다른 파일 조직화 기법
# - 동일한 버킷 ID를 가진 데이터가 하나의 물리적 파티션에 모두 모여 있기 때문에 데이터를 읽을 때 셔플을 피할 수 있음
# - **데이터가 이후 사용 방식에 맞춰 사전에 파티셔닝되므로 조인이나 집계할 때 발생하는 고비용의 셔플을 피할 수 있음.**
# - 같은 키로 계속 조인이 발생하는 경우, 일별 누적으로 쌓여가는 테이블 을 버킷팅을 하면 효과를 볼 수 있음
# ### 10.4 Broadcast variable
# - 브로드캐스트 변수는 모든 익스큐터에서 사용할 수 있는 공유 변수 shared variable
# - 드라이버에서 한 번 생성되면 익스큐터에서만 읽을 수 있음.
# - 전체 데이터셋이 스파크 클러스터에서 전파될 수 있어서 익스큐터에서는 브로드캐스트 변수의 데이터에 접근할 수 있음
# - **익스큐터 내부에서 실행되는 모든 태스크는 모두 브로드캐스트 변수에 접근할 수 있음**
# ---
# ## 참고자료
#
# * [Spark Programming Guide](https://spark.apache.org/docs/latest/sql-programming-guide.html)
# * [PySpark SQL Modules Documentation](https://spark.apache.org/docs/latest/api/python/pyspark.sql.html)
# * <a href="https://spark.apache.org/docs/3.0.1/api/sql/" target="_blank">PySpark 3.0.1 Builtin Functions</a>
# * [PySpark Search](https://spark.apache.org/docs/latest/api/python/search.html)
# * [Pyspark Functions](https://spark.apache.org/docs/latest/api/python/pyspark.sql.html?#module-pyspark.sql.functions)
# - [Spark SQL, DataFrames and Datasets Guide](https://spark.apache.org/docs/latest/sql-programming-guide.html)
# - [Spark summit 2017 - Hive Bucketing in Apache Spark with Tejas Patil](https://youtu.be/6BD-Vv-ViBw?t=30) / [slide](https://www.slideshare.net/databricks/hive-bucketing-in-apache-spark-with-tejas-patil) / [한글 요약본](https://www.notion.so/Hive-Bucketing-in-Apache-Spark-Tejas-Patil-9374879e0ca744cc8e7047e82cf5fdfa)
# - [Spark summit 2017 - Optimizing Apache Spark SQL Joins: Spark Summit East talk by <NAME>](https://www.youtube.com/watch?v=fp53QhSfQcI) / [slide](https://www.slideshare.net/databricks/optimizing-apache-spark-sql-joins)
# - [Everyday I'm Shuffling - Tips for Writing Better Apache Spark Programs](https://www.youtube.com/watch?v=Wg2boMqLjCg)
# - [Spark Memory Management by 0x0fff](https://0x0fff.com/spark-memory-management/)
# - [Apache Spark에서 컬럼 기반 저장 포맷 Parquet(파케이) 제대로 활용하기](http://engineering.vcnc.co.kr/2018/05/parquet-and-spark/)
# - [Understanding Database Sharding](https://www.digitalocean.com/community/tutorials/understanding-database-sharding)
| day3/notebooks/lgde-spark-core/lgde-spark-core-4-join.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Align
# %load_ext autoreload
# %autoreload 2
import molsysmt as msm
molsys_1 = msm.convert('pdb_id:181l', to_form='molsysmt.MolSys',
selection='molecule_type=="protein"')
molsys_2 = msm.convert('pdb_id:1l17', to_form='molsysmt.MolSys',
selection='molecule_type=="protein"')
molsys_2on1 = msm.structure.align(molsys_2, reference_molecular_system=molsys_1)
msm.view([molsys_1, molsys_2on1])
| docs/contents/structure/align.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Name: con_example.ipynb
Authors: <NAME>, <NAME>
Example of the realistic simulations that can be done
"""
# %load_ext autoreload
# %autoreload 2
# +
# General imports
import numpy as np
import matplotlib.pyplot as plt
import sys
import pandas as pd
import yaml
from copy import deepcopy
from cProfile import Profile
import networkx as nx
# picture path
PICS = '../pics/'
# Module imports
from contagion import Contagion, config
from contagion.config import _baseconfig
from distributed import client
from itertools import product
from distributed import Client, performance_report, progress
# -
my_config = yaml.safe_load(open("param_scan_config.yaml"))
# +
class ParScan(object):
def __init__(self):
self._par_range = None
def par_update(self, val):
pass
def make_generator(self, configs):
for conf in configs:
for par in self._par_range:
yield self.par_update(par, conf)
class ScanMu(ParScan):
def __init__(self):
super().__init__()
self._par_range = [0.01, 0.05, 0.1, 0.2, 0.5, 0.7]
self.par_name = "Mu"
self.title = "Fraction of external connections"
def par_update(self, par, conf):
conf = deepcopy(conf)
conf["population"]["nx"]["kwargs"]["mu"] = par
return conf
class ScanScenario(ParScan):
def __init__(self):
super().__init__()
self.t_dur_initial = 28
t_starts = [7, 14, 28, 35]
t_durs_soft = [30, 3*30, 6*30, 9*30, 12*30, 14*30]
hard_scalings = [1/1.78, 2/7.8]
soft_scalings = [2/7.8, 3/7.8, 5/7.8]
self._par_range = product(
t_starts,
t_durs_soft,
hard_scalings,
soft_scalings)
self.par_name = "TStart"
self.title = "Lockdown Start"
def par_update(self, par, conf):
t_start, t_dur_soft, hard_scaling, soft_scaling = par
conf = deepcopy(conf)
conf["population"]["nx"]["kwargs"]["mu"] = par
conf["scenario"]["class"] = "SocialDistancing"
conf["scenario"]["t_steps"] = [
t_start,
t_start + self.t_dur_initial,
t_start + self.t_dur_initial + t_dur_soft]
conf["scenario"]["contact_rate_scalings"] = [hard_scaling, soft_scaling, 1]
return conf
# +
scan = ScanScenario()
gen = scan.make_generator([my_config])
scan2 = ScanMu()
gen2 = scan2.make_generator(gen)
# -
def submit_func(conf):
contagion = Contagion(conf)
if hasattr(contagion.pop, "_graph"):
g = deepcopy(contagion.pop._graph)
else:
g = None
contagion.sim()
stats = pd.DataFrame(contagion.statistics)
inf_per_day = []
for day in contagion.trace_contacts:
inf_per_day.append(np.unique(day))
return (stats, inf_per_day)
client = Client(scheduler_file="scheduler.json")
#client.restart()
futures = []
for conf in gen2:
futures.append(client.submit(submit_func, conf, pure=False))
progress(futures, notebook=True)
with performance_report(filename="dask-report.html"):
results = client.gather(futures)
futures
traceback.print_tb(res.traceback(), )
client
| examples/cluster_param_scan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
from keras.models import Model
from keras.layers import Input
from keras.layers.recurrent import GRU
from keras import backend as K
import json
from collections import OrderedDict
def format_decimal(arr, places=6):
return [round(x * 10**places) / 10**places for x in arr]
DATA = OrderedDict()
# ### GRU
# **[recurrent.GRU.0] units=4, activation='tanh', recurrent_activation='hard_sigmoid'**
#
# Note dropout_W and dropout_U are only applied during training phase
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid')
layer_0 = Input(shape=data_in_shape)
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3200 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.0'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
[w.shape for w in model.get_weights()]
# **[recurrent.GRU.1] units=5, activation='sigmoid', recurrent_activation='sigmoid'**
#
# Note dropout_W and dropout_U are only applied during training phase
# +
data_in_shape = (8, 5)
rnn = GRU(5, activation='sigmoid', recurrent_activation='sigmoid')
layer_0 = Input(shape=data_in_shape)
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3300 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.1'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.2] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=True**
#
# Note dropout_W and dropout_U are only applied during training phase
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=True)
layer_0 = Input(shape=data_in_shape)
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3400 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.2'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.3] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=False, go_backwards=True**
#
# Note dropout_W and dropout_U are only applied during training phase
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=False, go_backwards=True)
layer_0 = Input(shape=data_in_shape)
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3410 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.3'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.4] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=True, go_backwards=True**
#
# Note dropout_W and dropout_U are only applied during training phase
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=True, go_backwards=True)
layer_0 = Input(shape=data_in_shape)
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3420 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.4'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.5] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=False, go_backwards=False, stateful=True**
#
# Note dropout_W and dropout_U are only applied during training phase
#
# **To test statefulness, model.predict is run twice**
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=False, go_backwards=False, stateful=True)
layer_0 = Input(batch_shape=(1, *data_in_shape))
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3430 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.5'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.6] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=True, go_backwards=False, stateful=True**
#
# Note dropout_W and dropout_U are only applied during training phase
#
# **To test statefulness, model.predict is run twice**
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=True, go_backwards=False, stateful=True)
layer_0 = Input(batch_shape=(1, *data_in_shape))
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3440 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.6'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.7] units=4, activation='tanh', recurrent_activation='hard_sigmoid', return_sequences=False, go_backwards=True, stateful=True**
#
# Note dropout_W and dropout_U are only applied during training phase
#
# **To test statefulness, model.predict is run twice**
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid',
return_sequences=False, go_backwards=True, stateful=True)
layer_0 = Input(batch_shape=(1, *data_in_shape))
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3450 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U', 'b']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.7'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# **[recurrent.GRU.8] units=4, activation='tanh', recurrent_activation='hard_sigmoid', use_bias=False, return_sequences=True, go_backwards=True, stateful=True**
#
# Note dropout_W and dropout_U are only applied during training phase
#
# **To test statefulness, model.predict is run twice**
# +
data_in_shape = (3, 6)
rnn = GRU(4, activation='tanh', recurrent_activation='hard_sigmoid', use_bias=False,
return_sequences=True, go_backwards=True, stateful=True)
layer_0 = Input(batch_shape=(1, *data_in_shape))
layer_1 = rnn(layer_0)
model = Model(inputs=layer_0, outputs=layer_1)
# set weights to random (use seed for reproducibility)
weights = []
for i, w in enumerate(model.get_weights()):
np.random.seed(3460 + i)
weights.append(2 * np.random.random(w.shape) - 1)
model.set_weights(weights)
weight_names = ['W', 'U']
for w_i, w_name in enumerate(weight_names):
print('{} shape:'.format(w_name), weights[w_i].shape)
print('{}:'.format(w_name), format_decimal(weights[w_i].ravel().tolist()))
data_in = 2 * np.random.random(data_in_shape) - 1
result = model.predict(np.array([data_in]))
result = model.predict(np.array([data_in]))
data_out_shape = result[0].shape
data_in_formatted = format_decimal(data_in.ravel().tolist())
data_out_formatted = format_decimal(result[0].ravel().tolist())
print('')
print('in shape:', data_in_shape)
print('in:', data_in_formatted)
print('out shape:', data_out_shape)
print('out:', data_out_formatted)
DATA['recurrent.GRU.8'] = {
'input': {'data': data_in_formatted, 'shape': data_in_shape},
'weights': [{'data': format_decimal(w.ravel().tolist()), 'shape': w.shape} for w in weights],
'expected': {'data': data_out_formatted, 'shape': data_out_shape}
}
# -
# ### export for Keras.js tests
print(json.dumps(DATA))
| notebooks/layers/recurrent/GRU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="9d315388"
# #### Transfer Learning Modell 1
# - Backbone VGG-16
# - Images cropped and resized
# - Colorscheme original
# - Preprocess input will scale pixels between -1 and 1, sample-wise.
# + [markdown] id="556ee7ab"
# Sources for use and issue resolving:
# - https://machinelearningmastery.com/use-pre-trained-vgg-model-classify-objects-photographs/
# - https://arxiv.org/abs/1409.1556
# - https://github.com/keras-team/keras-applications/blob/master/keras_applications/imagenet_utils.py#L157
# + colab={"base_uri": "https://localhost:8080/"} id="ibx3brHsJ8Qb" executionInfo={"status": "ok", "timestamp": 1628229699797, "user_tz": -120, "elapsed": 20940, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="9af04371-05d2-477d-c987-9e78fd1f79a1"
from google.colab import drive
drive.mount('/content/drive')
# + id="bff767d5" executionInfo={"status": "ok", "timestamp": 1628229762102, "user_tz": -120, "elapsed": 204, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import Image, display
# + id="5411cbe9" executionInfo={"status": "ok", "timestamp": 1628229767125, "user_tz": -120, "elapsed": 3920, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation, Flatten, Dropout
from keras_preprocessing.image import ImageDataGenerator
# + colab={"base_uri": "https://localhost:8080/"} id="9a37e09d" executionInfo={"status": "ok", "timestamp": 1628229767126, "user_tz": -120, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="4a87b8dd-de65-4ee3-b4fc-7e8bae277a5d"
import tensorflow as tf
print(tf.__version__)
# + id="fd5d00d5" executionInfo={"status": "ok", "timestamp": 1628229770351, "user_tz": -120, "elapsed": 533, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
df = pd.read_csv('/content/drive/MyDrive/Datensätze/train4_small.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="BIY67kkVKrwM" executionInfo={"status": "ok", "timestamp": 1628229771586, "user_tz": -120, "elapsed": 217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="ee0b8c03-0db4-415c-c6c1-0fed29bc00e6"
df.head()
# + id="4c4bbe44" executionInfo={"status": "ok", "timestamp": 1628229775183, "user_tz": -120, "elapsed": 212, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
# Original Images
in_dir = '/content/drive/MyDrive/Datensätze/four'
# + [markdown] id="b390b24c"
# ### Image Generator
# + id="f8e97a88" executionInfo={"status": "ok", "timestamp": 1628229784112, "user_tz": -120, "elapsed": 197, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
datagen = ImageDataGenerator(rescale=1./255., validation_split = 0.25)
# + colab={"base_uri": "https://localhost:8080/"} id="6bae5961" executionInfo={"status": "ok", "timestamp": 1628229786255, "user_tz": -120, "elapsed": 723, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="32d10362-e29f-4449-bede-1377efa58d0d"
train_gen = datagen.flow_from_dataframe(dataframe = df,
directory = in_dir,
x_col = "filename",
y_col = 'labels',
batch_size = 10,
seed = 2,
shuffle = True,
class_mode = "categorical",
classes = ['opacity', 'glaucoma','md', 'normal'],
subset='training',
target_size = (224,224))
# + colab={"base_uri": "https://localhost:8080/"} id="a2b3e2ac" executionInfo={"status": "ok", "timestamp": 1628229788660, "user_tz": -120, "elapsed": 518, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="a979c2aa-dcd5-48a1-9246-c6871ce6de2a"
val_gen = datagen.flow_from_dataframe(dataframe = df,
directory = in_dir,
x_col = "filename",
y_col = 'labels',
batch_size = 10,
seed = 2,
shuffle = False,#Labels nicht shuffeln für CM
class_mode = "categorical",
classes = ['opacity', 'glaucoma','md', 'normal'],
subset='validation',
target_size = (224,224))
# + id="3b168db2" executionInfo={"status": "ok", "timestamp": 1628229794106, "user_tz": -120, "elapsed": 3020, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
imgs, labels = next(train_gen)
# + id="1b87f1e3" executionInfo={"status": "ok", "timestamp": 1628229796823, "user_tz": -120, "elapsed": 195, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
def plotImages(images_arr):
fig, axes = plt.subplots(1, 5, figsize=(20,20))
axes = axes.flatten()
for img, ax in zip( images_arr, axes):
ax.imshow(img)
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="4fb73401" executionInfo={"status": "ok", "timestamp": 1628229800556, "user_tz": -120, "elapsed": 1843, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="c779f4c0-82d7-4ad0-a395-881be04ce8c2"
plotImages(imgs)
print(labels)
# + [markdown] id="e2799732"
# ### Get VGG16 Pretrained Model
# + id="16220992" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628229810007, "user_tz": -120, "elapsed": 6333, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="1d4d13d1-f521-4183-d39b-ee916e13173a"
model_vgg16 = tf.keras.applications.vgg16.VGG16()
# + colab={"base_uri": "https://localhost:8080/"} id="ad21d283" executionInfo={"status": "ok", "timestamp": 1628229812501, "user_tz": -120, "elapsed": 215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="55ed9bcd-381e-4809-9268-dd7c476c9dc0"
model_vgg16.summary()
# + id="a0ddb39a" executionInfo={"status": "ok", "timestamp": 1628229816938, "user_tz": -120, "elapsed": 201, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
# Build sequentail model
model = Sequential()
for layer in model_vgg16.layers[:-1]:
model.add(layer)
# + id="bb9d945d" executionInfo={"status": "ok", "timestamp": 1628229818526, "user_tz": -120, "elapsed": 190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
# freeze trainable parameters for all layers in the model
for layer in model.layers:
layer.trainable = False
# + id="0262d606" executionInfo={"status": "ok", "timestamp": 1628229821114, "user_tz": -120, "elapsed": 192, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
# my output layer for four output classes, activation for multi-class classification
model.add(Dense(units=4, activation='softmax'))
# + colab={"base_uri": "https://localhost:8080/"} id="24707c9e" executionInfo={"status": "ok", "timestamp": 1628240147740, "user_tz": -120, "elapsed": 38, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="eced6db5-9c82-4352-ca37-fe71113f96c1"
model.summary()
# + id="58c9cc74" executionInfo={"status": "ok", "timestamp": 1628229831113, "user_tz": -120, "elapsed": 200, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
step_size_train = train_gen.n//train_gen.batch_size
step_size_val = val_gen.n//val_gen.batch_size
# + id="4857f041" executionInfo={"status": "ok", "timestamp": 1628229832720, "user_tz": -120, "elapsed": 194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}}
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="3835d8c0" executionInfo={"status": "ok", "timestamp": 1628240147237, "user_tz": -120, "elapsed": 10311471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="1c0acca4-c059-4c28-f5ea-397ed9e12110"
history = model.fit(x=train_gen, validation_data=val_gen, steps_per_epoch=step_size_train,
validation_steps=step_size_val, epochs=10,verbose=2)
# + colab={"base_uri": "https://localhost:8080/", "height": 499} id="d8776457" executionInfo={"status": "ok", "timestamp": 1628240147739, "user_tz": -120, "elapsed": 516, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="06994303-ed03-4864-cd92-e0ece42005e7"
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(10)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
# + id="b6d57603" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628240311700, "user_tz": -120, "elapsed": 7439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhQN7oZcOXhMM0AgecritM7JP6IWKvkL4HS8bJW=s64", "userId": "13385611080783470639"}} outputId="bef49489-5766-48b8-ab32-03bb8eaeb1a1"
model.save('/content/drive/MyDrive/Transfer Learning/Savemodel_2.1')
# + id="6c1f5355"
# + id="69d9b571"
| 03_Multi-Class-Classification/Transfer_2.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Сегодня мы обойдемся без DeepPavlov
# +
import pandas as pd
import numpy as np
import dask.dataframe as dd
import matplotlib.pyplot as plt
# -
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer, TfidfTransformer
df = pd.read_csv("Log.dms", sep='\t')
df.request = df.request.replace({"https://yandex\.ru/search/\?text=":""}, regex=True)
df["text"] = df["request"].map(str) + df["urls"].map(str)
df.head()
len(df)
# Интернет нужен для порно :)
#
# А поскольку оно не идет по ТВ, значит все, что попадает сюда — точно не относится к ТВ.
pron_keywords = ['porn', 'порн', 'xxx']
pron = df.text.str.contains("|".join(pron_keywords), na=False)
pron = pron.replace(True, -1)
pron = pron.replace(False, 0)
len(pron[pron == -1])
df["pron"] = pron
# Без обучающей выборки будем искать по ключевым словам
other_keywords = ["скачать", "mp3", "facebook.com", "vk.com", "погода", "читать", "maps", "купить"]
tv_keywords = [".tv", "tv.ru", "сериал"]
other = df.text.str.contains("|".join(other_keywords), na=False)
other = other.replace(True, -1)
other = other.replace(False, 0)
#other = other.astype('bool')
print(other.head())
print(len(other))
other.hist()
pd.crosstab(other, other)
# +
tv = df.text.str.contains("|".join(tv_keywords), na=False)
tv = tv.astype('int')
print(tv.head())
print(len(tv))
tv.hist()
pd.crosstab(tv, tv)
# -
tv_column = tv + other + pron
tv_column = tv_column.replace(-2,-1)
#tv_column = tv_column.replace(0,np.NaN)
df["tv"] = tv_column
pd.crosstab(df["tv"], df["tv"])
train = df[df["tv"].notnull()]
# +
tfidf_vectorizer = TfidfVectorizer(max_df=0.95, min_df=2,
max_features=1000,
stop_words=['english','russian']
)
tfidf = tfidf_vectorizer.fit_transform(train["request"])
# -
tfidf.shape
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB().fit(tfidf, train["tv"])
# +
test_new = ['tv.is', 'Random flood?', "mp3"]
test_new_tfidf = tfidf_vectorizer.transform(test_new)
predicted = clf.predict(test_new_tfidf)
predicted
for doc, category in zip(test_new, predicted):
print('%r => %s' % (doc, category))
# +
from sklearn.linear_model import SGDClassifier
from sklearn.calibration import CalibratedClassifierCV
from sklearn.pipeline import Pipeline
# text_clf = Pipeline([('tfidf_std', tfidf_vectorizer),
# ('clf', SGDClassifier(random_state=42, max_iter=5)),
# ])
text_clf = Pipeline([('tfidf_std', tfidf_vectorizer),
('clf', CalibratedClassifierCV()),
])
text_clf.fit(train["text"], train["tv"])
# -
predicted = text_clf.predict(train["text"])
np.mean(predicted == train["tv"])
from sklearn import metrics
print(metrics.classification_report(train["tv"], predicted))
predicted = text_clf.predict(df["text"])
predict_proba
df["tv_final"] = predicted + df["tv"]
pd.crosstab(df["tv_final"], df["tv_final"])
df["tv_final"].value_counts(normalize = True)
# Всего связано с TV как минимум 12% запросов. Но, возможно, из оставшихся 67% есть еще...
df[df["tv_final"] >= 2].request.head()
df[df["tv_final"] <= -2].request.head()
# +
tfidf_results = text_clf.steps[0][1].transform(df[df["tv_final"] == 2]["request"])
# -
def tfidf_scores(vectorizer, tfidf_result):
# http://stackoverflow.com/questions/16078015/
results = []
scores = zip(vectorizer.get_feature_names(),
np.asarray(tfidf_result.sum(axis=0)).ravel())
sorted_scores = sorted(scores, key=lambda x: x[1], reverse=True)
for item in sorted_scores:
results.append("{0:50} Score: {1}".format(item[0], item[1]))
return results
tfidf_scores(text_clf.steps[0][1], tfidf_results)[:50]
| tv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wlrma0108/Pytorch/blob/main/Bidirectional%20Recurrent%20Neural%20Network.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="LQORfLNUm_Tu" outputId="db32a45e-f292-48e6-91d9-80a56df60555"
# !pip3 install torch
# + id="1GvcN2bdnDg7"
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
import numpy as np
from torch.nn.utils import clip_grad_norm
# + id="I18psUoKnDn7"
device=torch.device=('cuda' if torch.cuda.is_available() else 'cpu')
sequence_length=28
input_size=28
hidden_size=128
num_layers=2
num_classes=10
batch_size=100
num_epochs=2
learning_rate=0.003
# + id="OxmktRfvnDvC"
train_dataset=torchvision.datasets.MNIST(root='../../data/',train=True,transform=transforms.ToTensor(),download=True)
test_dataset=torchvision.datasets.MNIST(root='../../data/',train=False,transform=transforms.ToTensor())
train_loader=torch.utils.data.DataLoader(dataset=train_dataset,batch_size=batch_size,shuffle=True)
test_loader=torch.utils.data.DataLoader(dataset=test_dataset,batch_size=batch_size,shuffle=False)
# + id="e3mcRF09nD2S"
class BiRNN(nn.Module):
def __init__(self,input_size,hidden_size, num_layers,num_classes):
super(BiRNN,self).__init__()
self.hidden_size=hidden_size
self.num_layers=num_layers
self.lstm=nn.LSTM(input_size,hidden_size,num_layers,batch_first=True,bidirectional=True)
self.fc=nn.Linear(hidden_size*2,num_classes)
def forward(self,x):
h0=torch.zeros(self.num_layers*2,x.size(0),self.hidden_size).to(device)
c0=torch.zeros(self.num_layers*2,x.size(0),self.hidden_size).to(device)
out,_=self.lstm(x,(h0,c0))
out=self.fc(out[:,-1,:])
return out
model=BiRNN(input_size,hidden_size,num_layers,num_classes).to(device)
# + colab={"base_uri": "https://localhost:8080/"} id="6xfzxYiKnD9S" outputId="facf4f91-484b-4ab8-a2fd-a2ac60a89c1c"
criterion=nn.CrossEntropyLoss()
optimizer=torch.optim.Adam(model.parameters(), lr=learning_rate)
total_step=len(train_loader)
for epoch in range(num_epochs):
for i, (images,labels)in enumerate(train_loader):
images=images.reshape(-1,sequence_length,input_size).to(device)
labels=labels.to(device)
outputs=model(images)
loss=criterion(outputs,labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
if(i+1)%100==0:
print('Epoch[{}/{}], step []{}/{}],Loss:{:.4f}'.format(epoch+1,num_epochs,i+1,total_step,loss.item()))
with torch.no_grad():
correct=0
total=0
for images,labels in test_loader:
images=images.reshape(-1,sequence_length,input_size).to(device)
labels=labels.to(device)
outputs=model(images)
_,predicted=torch.max(outputs.data,1)
total+=labels.size(0)
correct+=(predicted==labels).sum().item()
print('Test Accuracy og the model on the 10000 test images: {}%'.format(100*correct/total))
torch.save(model.state_dict(),'codel.ckpt')
| Bidirectional Recurrent Neural Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## A detour – to DL machinery with pytorch
#
# ## Goal:
# * Get familiar with a basic pytorch neural network
#
# ## Steps:
# 1. Define Network, loss & optimizer
# 2. Get Dataset - MNIST
# 3. Construct minibatch dataloader
# 4. Forward pass
# 5. Calculate error/loss
# 6. Do backprop
# 7. Update weights using Gradient Descent
# 8. Train & Test
# ### Reference :
# * pytorch Udacity git https://github.com/udacity/deep-learning-v2-pytorch
# ### 1. Install the required packages
#
# * No esoteric requirements
# * You can run them without docker
# * pip install -r requirements.txt
# * Requirements
# * python 3.6, pytorch, openAI gym, numpy, matplotlib
# * anaconda is easier but not needed
# * Miniconda works fine
# ### 2. Define imports
#
# python 3, numpy, matplotlib, torch, gym
# +
# General imports
import gym
import PIL # for in-line display of certain environments
import sys
import numpy as np
import random
from collections import namedtuple, deque, defaultdict
import matplotlib.pyplot as plt
# %matplotlib inline
# torch imports
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
#
print("== Version Numbers ==")
print("Python : %s.%s.%s" % sys.version_info[:3])
print(" Torch : ",torch.__version__)
# -
# * == Version Numbers ==
# * Python : 3.7.1
# * Torch : 1.0.0
# ### 2.1. Global Constants and other variables
# Constants Definitions
BUFFER_SIZE = 512 # int(1e5) # replay buffer size
BATCH_SIZE = 64 # minibatch size
GAMMA = 0.99 # discount factor
TAU = 1e-3 # for soft update of target parameters
LR = 5e-4 # learning rate
UPDATE_EVERY = 4 # how often to update the network
# Number of neurons in the layers of the Network
FC1_UNITS = 128
FC2_UNITS = 64
FC3_UNITS = 32
# Store models flag. Store during calibration runs and do not store during hyperparameter search
STORE_MODELS = False
# ### 3. pytorch DL machinery
# ### 3.1. Define Network
class ANetwork(nn.Module):
def __init__(self, input_size, output_size, seed=42, fc1_units = FC1_UNITS, fc2_units = FC2_UNITS,
fc3_units = FC3_UNITS):
"""Initialize parameters and build model."""
super(ANetwork, self).__init__()
self.seed = torch.manual_seed(seed)
self.fc1 = nn.Linear(input_size,fc1_units)
self.fc2 = nn.Linear(fc1_units,fc2_units)
# self.fc3 = nn.Linear(fc2_units,fc3_units)
# self.fc4 = nn.Linear(fc3_units,action_size)
self.fc4 = nn.Linear(fc2_units,output_size)
def forward(self, state):
"""Build a network that maps state -> action values."""
x = F.relu(self.fc1(state))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
x = F.softmax(self.fc4(x),dim=-1)
return x
# ### 3.2 Get MNIST Dataset
# +
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
# transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
transforms.Normalize([0.5], [0.5]),
])
# Download and load the training data
trainset = datasets.MNIST('MNIST_data/', download=True, train=True, transform=transform)
# -
# ### 3.3 Construct minibatch dataloader
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# ### 3.4 Create network instance & loss
# ### And a quick functional test
model = ANetwork(784,10)
# Define the loss
criterion = nn.CrossEntropyLoss()
model
# ### 3.4 Forward pass
# +
# Get our data
images, labels = next(iter(trainloader))
# Flatten images
images = images.view(images.shape[0], -1)
# Forward pass, get our logits
logits = model(images)
# -
# ### 3.5 Calculate error/loss
# +
# Calculate the loss with the logits and the labels
loss = criterion(logits, labels)
print(loss)
# -
# ### 3.6 Do backprop
# +
print('Before backward pass: \n', model.fc1.weight.grad)
loss.backward()
print('After backward pass: \n', model.fc1.weight.grad)
# -
# ### Create Optimizer
# +
from torch import optim
# Optimizers require the parameters to optimize and a learning rate
optimizer = optim.SGD(model.parameters(), lr=0.01)
# -
# ### 3.7 Update weights using Gradient Descent
# Take an update step and few the new weights
optimizer.step()
print('Updated weights - ', model.fc1.weight)
# ## Train
# ### Now let us do full training
# +
import time
from datetime import datetime, timedelta
start_time = time.time()
# This is another way (probably easier) of building a network in pytorch
# Instead of softmax, it uses log softmax
model = nn.Sequential(nn.Linear(784, 128),
nn.ReLU(),
nn.Linear(128, 64),
nn.ReLU(),
nn.Linear(64, 10),
nn.LogSoftmax(dim=1))
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003)
epochs = 5
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Flatten MNIST images into a 784 long vector
images = images.view(images.shape[0], -1)
# Training pass
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
print("Training loss : ", running_loss/len(trainloader))
print('Elapsed : {}'.format(timedelta(seconds=time.time() - start_time)))
# -
# ### 4. Test
# #### I like the Udacity helper class - very good to test the MNIST results
# +
# %matplotlib inline
import helper
images, labels = next(iter(trainloader))
img = images[0].view(1, 784)
# Turn off gradients to speed up this part
with torch.no_grad():
logps = model(img)
# Output of the network are log-probabilities, need to take exponential for probabilities
ps = torch.exp(logps)
helper.view_classify(img.view(1, 28, 28), ps)
# -
# ### Calculate Accuracy
# #### For this we need to load the test dataset, run the forward pass and then calculate the accuracy
# Download and load the test data
testset = datasets.MNIST('MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# +
images, labels = next(iter(testloader))
print(images.shape,labels.shape)
img = images = images.view(images.shape[0], -1)
# Get the class probabilities
ps = torch.exp(model(img))
# Make sure the shape is appropriate, we should get 10 class probabilities for 64 examples
print(ps.shape)
# -
top_p, top_class = ps.topk(1, dim=1)
print(top_class.shape)
print(top_p.shape)
equals = top_class == labels.view(*top_class.shape) # pay attention to shapes and sizes
accuracy = torch.mean(equals.type(torch.FloatTensor))
print('Accuracy : %.2f %%' % (accuracy.item()*100))
# ## _That's all Folks !_
| pytorch-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: lidar
# language: python
# name: lidar
# ---
# +
import os
from os import listdir
import glob
import multiprocessing as mp
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from plyfile import PlyData, PlyElement
import open3d as o3d
# -
# # **CREATE CSVs FOR EACH OBJECT**
# + tags=[]
ply0 = PlyData.read('../../data/paris_lille/paris_lille/Paris.ply')
ply1 = PlyData.read('../../data/paris_lille/paris_lille/Lille1.ply')
ply2 = PlyData.read('../../data/paris_lille/paris_lille/Lille2.ply')
def plyToData(ply_file):
data = ply_file.elements[0].data
data_pd = pd.DataFrame(data)
data_np = np.zeros(data_pd.shape, dtype=np.float)
property_names = data[0].dtype.names
for i, name in enumerate(property_names):
data_np[:,i] = data_pd[name]
return data_np, data_pd
data_np0, data_pd0 = plyToData(ply0)
data_np1, data_pd1 = plyToData(ply1)
data_np2, data_pd2 = plyToData(ply2)
# -
def createCSVObjectFile(df):
for i in df.label.unique():
df_ = df[df.label == i]
if df_['class'].iloc[0] == 0 or df_['class'].iloc[0] == 100000000:
continue
else:
file_name = "../../data/paris_lille/csv_objects/" + str(int(df_.iloc[0]['class'])) + "_" + str(int(df_.iloc[0].label)) + ".csv"
df_.to_csv(file_name, index=False)
# +
num_cores = mp.cpu_count()
pool = mp.Pool(num_cores)
start = time.time()
csvs = pool.map(createCSVObjectFile, [data_pd0, data_pd1, data_pd2])
end = time.time()
print(end - start)
pool.close()
# -
# # **VOXELIZE EACH CSV**
def voxelize(filename):
df = pd.read_csv(filename)
df = df[['x', 'y', 'z']]
points = df.to_numpy()
points = points[np.logical_not(np.isnan(points).any(axis=1))]
origin = (np.min(points[:, 0]), np.min(points[:, 1]), np.min(points[:, 2]))
points[:, 0] -= origin[0]
points[:, 1] -= origin[1]
points[:, 2] -= origin[2]
voxel_size=(24, 24, 24)
padding_size=(32, 32, 32)
resolution=0.1
OCCUPIED = 1
FREE = 0
x_logical = np.logical_and((points[:, 0] < voxel_size[0] * resolution), (points[:, 0] >= 0))
y_logical = np.logical_and((points[:, 1] < voxel_size[1] * resolution), (points[:, 1] >= 0))
z_logical = np.logical_and((points[:, 2] < voxel_size[2] * resolution), (points[:, 2] >= 0))
xyz_logical = np.logical_and(x_logical, np.logical_and(y_logical, z_logical))
inside_box_points = points[xyz_logical]
voxels = np.zeros(padding_size)
center_points = inside_box_points + (padding_size[0] - voxel_size[0]) * resolution / 2
x_idx = (center_points[:, 0] / resolution).astype(int)
y_idx = (center_points[:, 1] / resolution).astype(int)
z_idx = (center_points[:, 2] / resolution).astype(int)
voxels[x_idx, y_idx, z_idx] = OCCUPIED
npy_filename = '../../data/paris_lille/npy_objects_method1/' + filename.split('.')[0].split('/')[-1] + '.npy'
np.save(npy_filename, voxels)
# + tags=[]
csv_dir = '/home/jupyter-seanandrewchen/shared/cusp-capstone/data/paris_lille/csv_objects/'
input_path = os.path.join(csv_dir, '*.csv')
num_cores = mp.cpu_count()
pool = mp.Pool(num_cores)
start = time.time()
voxelizations = pool.map(voxelize, glob.glob(input_path))
end = time.time()
print(end - start)
pool.close()
# +
#csv_dir = '/home/jupyter-seanandrewchen/shared/cusp-capstone/data/paris_lille/csv_objects/'
#input_path = os.path.join(csv_dir, '*.csv')
#for csv in glob.iglob(input_path):
# voxelize(csv)
# -
# # **INSPECT FILES**
def plot3DVoxel(voxels):
fig = plt.figure('Point Cloud 3D Voxelization')
plt3d = fig.gca(projection='3d')
occupied = (voxels == 1)
free = (voxels == 0)
# # set the colors of each object
colors = np.zeros(voxels.shape + (4,))
colors[free] = [0.1, 0.1, 0.1, 0.1]
colors[occupied] = [0.8, 0.8, 0.8, 1.0]
# setting camera angle
plt3d.set_xlabel('X', fontsize=9)
plt3d.set_ylabel('Y', fontsize=9)
plt3d.set_zlabel('Z', fontsize=9)
plt3d.set_xlim3d(0, voxels.shape[0])
plt3d.set_ylim3d(0, voxels.shape[1])
plt3d.set_zlim3d(0, voxels.shape[2])
# and plot everything
# plt3d.voxels(occupied | free, facecolors=colors, edgecolor='k', linewidth=0.8)
plt3d.voxels(occupied, facecolors=colors, edgecolor='k', linewidth=0.8)
dx, dy, dz = voxels.shape
x = y= z = 0
xx = [x, x, x+dx, x+dx, x]
yy = [y, y+dy, y+dy, y, y]
kwargs = {'alpha': 1, 'color': 'red'}
plt3d.plot3D(xx, yy, [z]*5, **kwargs)
plt3d.plot3D(xx, yy, [z+dz]*5, **kwargs)
plt3d.plot3D([x, x], [y, y], [z, z+dz], **kwargs)
plt3d.plot3D([x, x], [y+dy, y+dy], [z, z+dz], **kwargs)
plt3d.plot3D([x+dx, x+dx], [y+dy, y+dy], [z, z+dz], **kwargs)
plt3d.plot3D([x+dx, x+dx], [y, y], [z, z+dz], **kwargs)
plt.show()
voxels = np.load('../../data/paris_lille/npy_objects_method1/302020400_159.npy').astype(np.float32)
plot3DVoxel(voxels)
# # **LOAD ALL NPY FILES**
# +
path = os.path.join('../../data/paris_lille/npy_objects/', '*.npy')
voxels_list = []
label_list = []
for npy_file in glob.iglob(path):
file_name = npy_file.split('/')[-1].split('_')[0]
label_list.append(file_name)
voxels = np.load(npy_file).astype(np.float32)
voxels_list.append(voxels)
| lidar_data_processing_voxelization_method1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as pl
from expose.instruments import MAVIS
from expose.telescopes import VLT
from expose.sources import galaxy_source
from expose.sky import sky_source
# -
# ## First initilize the source and sky model
skyModel = sky_source(offline=True)
sourceModel = galaxy_source()
# ## Set source and sky parameters.
#
# In this case we're assuming a relatively dark sky with FLI=0.3 and a bright-ish source at z=0.01.
skyModel.set_params(fli=0.3, airmass=1.2, pwv=10)
sourceModel.set_params(template='Sc', redshift=1, obs_mag=22, obs_band='sdss_r')
# ## initialize the telescope and instrument.
# In principle there is nothing stopping mixing and matching instruments on different
# telescopes, modulo changes to the FoV etc.
tel = VLT()
spec = MAVIS(mode='LR-red', pix_scale=0.025, jitter=40)
# ## Run the observations and plot the output. Returns predicted S/N per pixel.
wave_mavis, sn_mavis = spec.observe(sourceModel, tel, sky=skyModel, seeing=0.8, dit=900, ndit=4, binning=1)
# ## Just to check, pull out ensquared energy profile
# +
fig = pl.figure(figsize=(8,3))
ax = fig.add_axes([0.15,0.15,0.8,0.8])
ax.plot(wave_mavis, spec.obs_ee*100, '-', color='0.6')
ax.axis([0.37,1.007,0,spec.obs_ee.max()*1.2*100])
ax.set_ylabel('Ensquared energy [%]', size=12)
ax.set_xlabel('Wavelength [$\mu$m]', size=12)
pl.show()
# -
# ## Plot S/N
# +
fig = pl.figure(figsize=(8,3))
ax = fig.add_axes([0.15,0.15,0.8,0.8])
ax.plot(wave_mavis, sn_mavis, '-', color='0.6')
ax.set_ylabel('S/N [pixel$^{-1}$]', size=12)
ax.set_xlabel('Wavelength [$\mu$m]', size=12)
pl.show()
# -
# ## Plot throughput curve
# +
fig = pl.figure(figsize=(8,3))
ax = fig.add_axes([0.15,0.15,0.8,0.8])
ax.plot(wave_mavis, spec.tpt*100, '-', color='0.6')
ax.set_ylabel('Throughput [%]', size=12)
ax.set_xlabel('Wavelength [$\mu$m]', size=12)
ax.axis([0.37,1.007,0,spec.tpt.max()*100*1.2])
pl.show()
# -
| examples/MAVIS_ETC_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multiclass Support Vector Machines
# In this exercise you will:
# - implement a fully-vectorized loss function for the multi-class SVM
# - implement the fully-vectorized expression for its analytic gradient
# - check your implementation using numerical gradient
# - use a validation set to tune the learning rate and regularization strength
# - optimize the loss function with SGD
# - visualize the final learned parameters
# +
# Run some setup code for this notebook.
import random
import numpy as np
from data_utils import load_CIFAR10
import matplotlib.pyplot as plt
# This is a bit of magic to make matplotlib figures appear inline in the
# notebook rather than in a new window.
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# -
# ## CIFAR-10 Data Loading and Preprocessing
# Open up a terminal window and navigate to the datasets folder inside the hw3 folder. Run the get_datasets.sh script. On my Mac, I just type in ./get_datasets.sh at the shell prompt. A new folder called cifar_10_batches_py will be created and it will contain 50000 labeled images for training and 10000 labeled images for testing. The function further partitions the 50000 training images into a train set and a validation set for selection of hyperparameters. We have provided a function to read this data in **data_utils.py**. Each image is a 32×32
# array of RGB triples. It is preprocessed by subtracting the mean image from all images. We flatten each image into a 1-dimensional array of size 3072 (i.e., 32×32×3). Then a 1 is appended to the front of that vector to handle the intercept term. So the training set is a numpy matrix of size 49000×3073, the validation set is a matrix of size 1000×3073 and the set-aside test set is of size 10000×3073. We also have a random sample of 500 images from the training data to serve as a development set or dev set to test our gradient and loss function implementations.
# +
import data_utils
# Get the CIFAR-10 data broken up into train, validation and test sets
X_train, y_train, X_val, y_val, X_dev, y_dev, X_test, y_test = data_utils.get_CIFAR10_data()
# -
# ## SVM Classifier
# Your code for this section will all be written inside **linear_svm.py**.
# You will need to write the function **naive_loss_svm** which uses for loops to evaluate the multiclass SVM loss function.
# +
# Evaluate the naive implementation of the loss we provided for you:
from linear_svm import svm_loss_naive
import time
# generate a random SVM coefficient matrix of small numbers
theta = np.random.randn(3073, 10) * 0.0001
loss, grad = svm_loss_naive(theta, X_train, y_train, 0.00001)
print('loss: %f' % (loss, ))
print('grad: %s', grad)
# -
# The grad returned from the function above is right now all zero. Derive and implement the gradient for the SVM cost function and implement it inline inside the function **svm_loss_naive**. You will find it helpful to interleave your new code inside the existing function.
# To check that you have correctly implemented the gradient, you can numerically estimate the gradient of the loss function and compare the numeric estimate to the gradient that you computed. We have provided code that does this for you.
# +
# Once you've implemented the gradient, recompute it with the code below
# and gradient check it with the function we provided for you
# Compute the loss and its gradient at theta.
loss, grad = svm_loss_naive(theta, X_dev, y_dev, 0.0)
# Numerically compute the gradient along several randomly chosen dimensions, and
# compare them with your analytically computed gradient. The numbers should match
# almost exactly along all dimensions.
from gradient_check import grad_check_sparse
f = lambda th: svm_loss_naive(th, X_dev, y_dev, 0.0)[0]
grad_numerical = grad_check_sparse(f, theta, grad)
# do the gradient check once again with regularization turned on
# you didn't forget the regularization gradient did you?
loss, grad = svm_loss_naive(theta, X_dev, y_dev, 1e2)
f = lambda w: svm_loss_naive(theta, X_dev, y_dev, 1e2)[0]
grad_numerical = grad_check_sparse(f, theta, grad)
# +
# Next implement the function svm_loss_vectorized; for now only compute the loss;
# we will implement the gradient in a moment.
tic = time.time()
loss_naive, grad_naive = svm_loss_naive(theta, X_dev, y_dev, 0.00001)
toc = time.time()
print('Naive loss: %e computed in %fs' % (loss_naive, toc - tic))
from linear_svm import svm_loss_vectorized
tic = time.time()
loss_vectorized, _ = svm_loss_vectorized(theta, X_dev, y_dev, 0.00001)
toc = time.time()
print('Vectorized loss: %e computed in %fs' % (loss_vectorized, toc - tic))
# The losses should match but your vectorized implementation should be much faster.
print('Difference: %f' % (loss_naive - loss_vectorized))
# -
# ## Vectorized version of the gradient computation
# Complete the implementation of svm_loss_vectorized, and compute the gradient
# of the loss function in a vectorized way.
#
#
# +
# The naive implementation and the vectorized implementation should match, but
# the vectorized version should still be much faster.
tic = time.time()
_, grad_naive = svm_loss_naive(theta, X_dev, y_dev, 0.00001)
toc = time.time()
print('Naive loss and gradient: computed in %fs' % (toc - tic))
tic = time.time()
_, grad_vectorized = svm_loss_vectorized(theta, X_dev, y_dev, 0.00001)
toc = time.time()
print('Vectorized loss and gradient: computed in %fs' % (toc - tic))
# The loss is a single number, so it is easy to compare the values computed
# by the two implementations. The gradient on the other hand is a matrix, so
# we use the Frobenius norm to compare them.
difference = np.linalg.norm(grad_naive - grad_vectorized, ord='fro')
print('Difference: %f' % difference)
# -
# In the file linear_classifier.py, we have implemented SGD in the function
# LinearClassifier.train() and you can run it with the code below.
from linear_classifier import LinearSVM
svm = LinearSVM()
tic = time.time()
loss_hist = svm.train(X_train, y_train, learning_rate=1e-7, reg=5e4,
num_iters=1500, verbose=True)
toc = time.time()
print('That took %fs' % (toc - tic))
# A useful debugging strategy is to plot the loss as a function of
# iteration number:
plt.plot(loss_hist)
plt.xlabel('Iteration number')
plt.ylabel('Loss value')
plt.show()
# ## Prediction with an SVM
# Compute $\theta^T x$ for a new example $x$ and pick the class with the highest score.
# Write the LinearSVM.predict function and evaluate the performance on both the
# training and validation set
y_train_pred = svm.predict(X_train)
print('training accuracy: %f' % (np.mean(y_train == y_train_pred), ))
y_val_pred = svm.predict(X_val)
print('validation accuracy: %f' % (np.mean(y_val == y_val_pred), ))
# +
# Use the validation set to tune hyperparameters (regularization strength and
# learning rate). You should experiment with different ranges for the learning
# rates and regularization strengths; if you are careful you should be able to
# get a classification accuracy of about 0.38 or higher on the validation set.
learning_rates = [1e-8, 5e-8, 1e-7, 5e-7, 1e-6]
regularization_strengths = [1e4, 5e4, 1e5, 5e5]
# results is dictionary mapping tuples of the form
# (learning_rate, regularization_strength) to tuples of the form
# (training_accuracy, validation_accuracy). The accuracy is simply the fraction
# of data points that are correctly classified.
results = {}
best_val = -1 # The highest validation accuracy that we have seen so far.
best_svm = None # The LinearSVM object that achieved the highest validation rate.
################################################################################
# TODO: #
# Write code that chooses the best hyperparameters by tuning on the validation #
# set. For each combination of hyperparameters, train a linear SVM on the #
# training set, compute its accuracy on the training and validation sets, and #
# store these numbers in the results dictionary. In addition, store the best #
# validation accuracy in best_val and the LinearSVM object that achieves this #
# accuracy in best_svm. #
# #
# Hint: You should use a small value for num_iters as you develop your #
# validation code so that the SVMs don't take much time to train; once you are #
# confident that your validation code works, you should rerun the validation #
# code with a larger value for num_iters. #
################################################################################
################################################################################
# END OF YOUR CODE #
################################################################################
# Print out results.
for lr, reg in sorted(results):
train_accuracy, val_accuracy = results[(lr, reg)]
print('lr %e reg %e train accuracy: %f val accuracy: %f' % (
lr, reg, train_accuracy, val_accuracy))
print('best validation accuracy achieved during cross-validation: %f' % best_val)
# +
# Visualize the cross-validation results
import math
x_scatter = [math.log10(x[0]) for x in results]
y_scatter = [math.log10(x[1]) for x in results]
# plot training accuracy
marker_size = 100
colors = [results[x][0] for x in results]
plt.subplot(2, 1, 1)
plt.scatter(x_scatter, y_scatter, marker_size, c=colors)
plt.colorbar()
plt.xlabel('log learning rate')
plt.ylabel('log regularization strength')
plt.title('CIFAR-10 training accuracy')
# plot validation accuracy
colors = [results[x][1] for x in results] # default size of markers is 20
plt.subplot(2, 1, 2)
plt.scatter(x_scatter, y_scatter, marker_size, c=colors)
plt.colorbar()
plt.xlabel('log learning rate')
plt.ylabel('log regularization strength')
plt.title('CIFAR-10 validation accuracy')
plt.tight_layout()
plt.show()
# -
# Evaluate the best svm on test set
y_test_pred = best_svm.predict(X_test)
test_accuracy = np.mean(y_test == y_test_pred)
print('linear SVM on raw pixels final test set accuracy: %f' % test_accuracy)
# +
# Visualize the learned weights for each class.
# Depending on your choice of learning rate and regularization strength, these may
# or may not be nice to look at.
theta = best_svm.theta[:-1,:] # strip out the bias
theta = theta.reshape(32, 32, 3, 10)
theta_min, theta_max = np.min(theta), np.max(theta)
classes = ['plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
for i in range(10):
plt.subplot(2, 5, i + 1)
# Rescale the weights to be between 0 and 255
thetaimg = 255.0 * (theta[:, :, :, i].squeeze() - theta_min) / (theta_max - theta_min)
plt.imshow(thetaimg.astype('uint8'))
plt.axis('off')
plt.title(classes[i])
# -
| HW4/hw4/multiclass_svm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''AITraining'': virtualenvwrapper)'
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JoshuaShunk/NSDropout/blob/main/mnist_implementation_of_New_Dropout.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dtYgI3SFHqm4"
# # MNIST Numbers Implementation of My New Layer
# + id="s2GytIidUnpd"
import matplotlib.pyplot as plt
import numpy as np
import random
import keras
from keras.datasets import mnist
import tensorflow as tf
import pandas as pd
from sklearn.model_selection import train_test_split
import statistics
# + id="0aLxFoLMU2jC"
np.set_printoptions(threshold=np.inf)
# + id="06HD9nTuEVHD"
np.random.seed(seed=22) #Random seed used for comparison between old dropout
# + colab={"base_uri": "https://localhost:8080/"} id="cag8ZraxEZbF" outputId="8ce6c916-47cb-48f9-e00f-26f054b3abca"
print(np.random.random(size=3)) #Check that seeds line up
# + colab={"base_uri": "https://localhost:8080/"} id="X4kKr-GRMeU2" outputId="d93eb3ec-f156-4328-b77f-3d69ad34c06b"
a = np.array([1,0,0,1,1,1])
b = np.array([1,0,1,1,0,1])
count = 0
for i, j in zip(a,b):
if i != j:
count += 1
print(count)
print(f'Difference: {np.count_nonzero(a==b)}')
# + id="-NkY3EiBU4tR" cellView="form"
#@title Load Layers (Credit to <NAME> & <NAME> for raw python implementation)
# Dense layer
class Layer_Dense:
# Layer initialization
def __init__(self, n_inputs, n_neurons,
weight_regularizer_l1=0, weight_regularizer_l2=0,
bias_regularizer_l1=0, bias_regularizer_l2=0):
# Initialize weights and biases
self.weights = 0.01 * np.random.randn(n_inputs, n_neurons)
self.biases = np.zeros((1, n_neurons))
# Set regularization strength
self.weight_regularizer_l1 = weight_regularizer_l1
self.weight_regularizer_l2 = weight_regularizer_l2
self.bias_regularizer_l1 = bias_regularizer_l1
self.bias_regularizer_l2 = bias_regularizer_l2
# Forward pass
def forward(self, inputs):
# Remember input values
self.inputs = inputs
# Calculate output values from inputs, weights and biases
self.output = np.dot(inputs, self.weights) + self.biases
# Backward pass
def backward(self, dvalues):
# Gradients on parameters
self.dweights = np.dot(self.inputs.T, dvalues)
self.dbiases = np.sum(dvalues, axis=0, keepdims=True)
# Gradients on regularization
# L1 on weights
if self.weight_regularizer_l1 > 0:
dL1 = np.ones_like(self.weights)
dL1[self.weights < 0] = -1
self.dweights += self.weight_regularizer_l1 * dL1
# L2 on weights
if self.weight_regularizer_l2 > 0:
self.dweights += 2 * self.weight_regularizer_l2 * \
self.weights
# L1 on biases
if self.bias_regularizer_l1 > 0:
dL1 = np.ones_like(self.biases)
dL1[self.biases < 0] = -1
self.dbiases += self.bias_regularizer_l1 * dL1
# L2 on biases
if self.bias_regularizer_l2 > 0:
self.dbiases += 2 * self.bias_regularizer_l2 * \
self.biases
# Gradient on values
self.dinputs = np.dot(dvalues, self.weights.T)
# ReLU activation
class Activation_ReLU:
# Forward pass
def forward(self, inputs):
# Remember input values
self.inputs = inputs
# Calculate output values from inputs
self.output = np.maximum(0, inputs)
# Backward pass
def backward(self, dvalues):
# Since we need to modify original variable,
# let's make a copy of values first
self.dinputs = dvalues.copy()
# Zero gradient where input values were negative
self.dinputs[self.inputs <= 0] = 0
# Softmax activation
class Activation_Softmax:
# Forward pass
def forward(self, inputs):
# Remember input values
self.inputs = inputs
# Get unnormalized probabilities
exp_values = np.exp(inputs - np.max(inputs, axis=1,
keepdims=True))
# Normalize them for each sample
probabilities = exp_values / np.sum(exp_values, axis=1,
keepdims=True)
self.output = probabilities
# Backward pass
def backward(self, dvalues):
# Create uninitialized array
self.dinputs = np.empty_like(dvalues)
# Enumerate outputs and gradients
for index, (single_output, single_dvalues) in \
enumerate(zip(self.output, dvalues)):
# Flatten output array
single_output = single_output.reshape(-1, 1)
# Calculate Jacobian matrix of the output
jacobian_matrix = np.diagflat(single_output) - \
np.dot(single_output, single_output.T)
# Calculate sample-wise gradient
# and add it to the array of sample gradients
self.dinputs[index] = np.dot(jacobian_matrix,
single_dvalues)
def predictions(self, outputs):
return np.argmax(outputs, axis=1)
# Sigmoid activation
class Activation_Sigmoid:
# Forward pass
def forward(self, inputs):
# Save input and calculate/save output
# of the sigmoid function
self.inputs = inputs
self.output = 1 / (1 + np.exp(-inputs))
# Backward pass
def backward(self, dvalues):
# Derivative - calculates from output of the sigmoid function
self.dinputs = dvalues * (1 - self.output) * self.output
# SGD optimizer
class Optimizer_SGD:
# Initialize optimizer - set settings,
# learning rate of 1. is default for this optimizer
def __init__(self, learning_rate=1., decay=0., momentum=0.):
self.learning_rate = learning_rate
self.current_learning_rate = learning_rate
self.decay = decay
self.iterations = 0
self.momentum = momentum
# Call once before any parameter updates
def pre_update_params(self):
if self.decay:
self.current_learning_rate = self.learning_rate * \
(1. / (1. + self.decay * self.iterations))
# Update parameters
def update_params(self, layer):
# If we use momentum
if self.momentum:
# If layer does not contain momentum arrays, create them
# filled with zeros
if not hasattr(layer, 'weight_momentums'):
layer.weight_momentums = np.zeros_like(layer.weights)
# If there is no momentum array for weights
# The array doesn't exist for biases yet either.
layer.bias_momentums = np.zeros_like(layer.biases)
# Build weight updates with momentum - take previous
# updates multiplied by retain factor and update with
# current gradients
weight_updates = \
self.momentum * layer.weight_momentums - \
self.current_learning_rate * layer.dweights
layer.weight_momentums = weight_updates
# Build bias updates
bias_updates = \
self.momentum * layer.bias_momentums - \
self.current_learning_rate * layer.dbiases
layer.bias_momentums = bias_updates
# Vanilla SGD updates (as before momentum update)
else:
weight_updates = -self.current_learning_rate * \
layer.dweights
bias_updates = -self.current_learning_rate * \
layer.dbiases
# Update weights and biases using either
# vanilla or momentum updates
layer.weights += weight_updates
layer.biases += bias_updates
# Call once after any parameter updates
def post_update_params(self):
self.iterations += 1
# Adagrad optimizer
class Optimizer_Adagrad:
# Initialize optimizer - set settings
def __init__(self, learning_rate=1., decay=0., epsilon=1e-7):
self.learning_rate = learning_rate
self.current_learning_rate = learning_rate
self.decay = decay
self.iterations = 0
self.epsilon = epsilon
# Call once before any parameter updates
def pre_update_params(self):
if self.decay:
self.current_learning_rate = self.learning_rate * \
(1. / (1. + self.decay * self.iterations))
# Update parameters
def update_params(self, layer):
# If layer does not contain cache arrays,
# create them filled with zeros
if not hasattr(layer, 'weight_cache'):
layer.weight_cache = np.zeros_like(layer.weights)
layer.bias_cache = np.zeros_like(layer.biases)
# Update cache with squared current gradients
layer.weight_cache += layer.dweights ** 2
layer.bias_cache += layer.dbiases ** 2
# Vanilla SGD parameter update + normalization
# with square rooted cache
layer.weights += -self.current_learning_rate * \
layer.dweights / \
(np.sqrt(layer.weight_cache) + self.epsilon)
layer.biases += -self.current_learning_rate * \
layer.dbiases / \
(np.sqrt(layer.bias_cache) + self.epsilon)
# Call once after any parameter updates
def post_update_params(self):
self.iterations += 1
# RMSprop optimizer
class Optimizer_RMSprop:
# Initialize optimizer - set settings
def __init__(self, learning_rate=0.001, decay=0., epsilon=1e-7,
rho=0.9):
self.learning_rate = learning_rate
self.current_learning_rate = learning_rate
self.decay = decay
self.iterations = 0
self.epsilon = epsilon
self.rho = rho
# Call once before any parameter updates
def pre_update_params(self):
if self.decay:
self.current_learning_rate = self.learning_rate * \
(1. / (1. + self.decay * self.iterations))
# Update parameters
def update_params(self, layer):
# If layer does not contain cache arrays,
# create them filled with zeros
if not hasattr(layer, 'weight_cache'):
layer.weight_cache = np.zeros_like(layer.weights)
layer.bias_cache = np.zeros_like(layer.biases)
# Update cache with squared current gradients
layer.weight_cache = self.rho * layer.weight_cache + \
(1 - self.rho) * layer.dweights ** 2
layer.bias_cache = self.rho * layer.bias_cache + \
(1 - self.rho) * layer.dbiases ** 2
# Vanilla SGD parameter update + normalization
# with square rooted cache
layer.weights += -self.current_learning_rate * \
layer.dweights / \
(np.sqrt(layer.weight_cache) + self.epsilon)
layer.biases += -self.current_learning_rate * \
layer.dbiases / \
(np.sqrt(layer.bias_cache) + self.epsilon)
# Call once after any parameter updates
def post_update_params(self):
self.iterations += 1
# Adam optimizer
class Optimizer_Adam:
# Initialize optimizer - set settings
def __init__(self, learning_rate=0.02, decay=0., epsilon=1e-7,
beta_1=0.9, beta_2=0.999):
self.learning_rate = learning_rate
self.current_learning_rate = learning_rate
self.decay = decay
self.iterations = 0
self.epsilon = epsilon
self.beta_1 = beta_1
self.beta_2 = beta_2
# Call once before any parameter updates
def pre_update_params(self):
if self.decay:
self.current_learning_rate = self.learning_rate * \
(1. / (1. + self.decay * self.iterations))
# Update parameters
def update_params(self, layer):
# If layer does not contain cache arrays,
# create them filled with zeros
if not hasattr(layer, 'weight_cache'):
layer.weight_momentums = np.zeros_like(layer.weights)
layer.weight_cache = np.zeros_like(layer.weights)
layer.bias_momentums = np.zeros_like(layer.biases)
layer.bias_cache = np.zeros_like(layer.biases)
# Update momentum with current gradients
layer.weight_momentums = self.beta_1 * \
layer.weight_momentums + \
(1 - self.beta_1) * layer.dweights
layer.bias_momentums = self.beta_1 * \
layer.bias_momentums + \
(1 - self.beta_1) * layer.dbiases
# Get corrected momentum
# self.iteration is 0 at first pass
# and we need to start with 1 here
weight_momentums_corrected = layer.weight_momentums / \
(1 - self.beta_1 ** (self.iterations + 1))
bias_momentums_corrected = layer.bias_momentums / \
(1 - self.beta_1 ** (self.iterations + 1))
# Update cache with squared current gradients
layer.weight_cache = self.beta_2 * layer.weight_cache + \
(1 - self.beta_2) * layer.dweights ** 2
layer.bias_cache = self.beta_2 * layer.bias_cache + \
(1 - self.beta_2) * layer.dbiases ** 2
# Get corrected cache
weight_cache_corrected = layer.weight_cache / \
(1 - self.beta_2 ** (self.iterations + 1))
bias_cache_corrected = layer.bias_cache / \
(1 - self.beta_2 ** (self.iterations + 1))
# Vanilla SGD parameter update + normalization
# with square rooted cache
layer.weights += -self.current_learning_rate * \
weight_momentums_corrected / \
(np.sqrt(weight_cache_corrected) +
self.epsilon)
layer.biases += -self.current_learning_rate * \
bias_momentums_corrected / \
(np.sqrt(bias_cache_corrected) +
self.epsilon)
# Call once after any parameter updates
def post_update_params(self):
self.iterations += 1
# Common loss class
class Loss:
# Regularization loss calculation
def regularization_loss(self, layer):
# 0 by default
regularization_loss = 0
# L1 regularization - weights
# calculate only when factor greater than 0
if layer.weight_regularizer_l1 > 0:
regularization_loss += layer.weight_regularizer_l1 * \
np.sum(np.abs(layer.weights))
# L2 regularization - weights
if layer.weight_regularizer_l2 > 0:
regularization_loss += layer.weight_regularizer_l2 * \
np.sum(layer.weights *
layer.weights)
# L1 regularization - biases
# calculate only when factor greater than 0
if layer.bias_regularizer_l1 > 0:
regularization_loss += layer.bias_regularizer_l1 * \
np.sum(np.abs(layer.biases))
# L2 regularization - biases
if layer.bias_regularizer_l2 > 0:
regularization_loss += layer.bias_regularizer_l2 * \
np.sum(layer.biases *
layer.biases)
return regularization_loss
# Set/remember trainable layers
def remember_trainable_layers(self, trainable_layers):
self.trainable_layers = trainable_layers
# Calculates the data and regularization losses
# given model output and ground truth values
def calculate(self, output, y, *, include_regularization=False):
# Calculate sample losses
sample_losses = self.forward(output, y)
# Calculate mean loss
data_loss = np.mean(sample_losses)
# Return loss
return data_loss
# Calculates accumulated loss
def calculate_accumulated(self, *, include_regularization=False):
# Calculate mean loss
data_loss = self.accumulated_sum / self.accumulated_count
# If just data loss - return it
if not include_regularization:
return data_loss
# Return the data and regularization losses
return data_loss, self.regularization_loss()
# Reset variables for accumulated loss
def new_pass(self):
self.accumulated_sum = 0
self.accumulated_count = 0
# Cross-entropy loss
class Loss_CategoricalCrossentropy(Loss):
# Forward pass
def forward(self, y_pred, y_true):
# Number of samples in a batch
samples = len(y_pred)
# Clip data to prevent division by 0
# Clip both sides to not drag mean towards any value
y_pred_clipped = np.clip(y_pred, 1e-7, 1 - 1e-7)
# Probabilities for target values -
# only if categorical labels
if len(y_true.shape) == 1:
correct_confidences = y_pred_clipped[
range(samples),
y_true
]
# Mask values - only for one-hot encoded labels
elif len(y_true.shape) == 2:
correct_confidences = np.sum(
y_pred_clipped * y_true,
axis=1
)
# Losses
negative_log_likelihoods = -np.log(correct_confidences)
return negative_log_likelihoods
# Backward pass
def backward(self, dvalues, y_true):
# Number of samples
samples = len(dvalues)
# Number of labels in every sample
# We'll use the first sample to count them
labels = len(dvalues[0])
# If labels are sparse, turn them into one-hot vector
if len(y_true.shape) == 1:
y_true = np.eye(labels)[y_true]
# Calculate gradient
self.dinputs = -y_true / dvalues
# Normalize gradient
self.dinputs = self.dinputs / samples
# Softmax classifier - combined Softmax activation
# and cross-entropy loss for faster backward step
class Activation_Softmax_Loss_CategoricalCrossentropy():
# Creates activation and loss function objects
def __init__(self):
self.activation = Activation_Softmax()
self.loss = Loss_CategoricalCrossentropy()
# Forward pass
def forward(self, inputs, y_true):
# Output layer's activation function
self.activation.forward(inputs)
# Set the output
self.output = self.activation.output
# Calculate and return loss value
return self.loss.calculate(self.output, y_true)
# Backward pass
def backward(self, dvalues, y_true):
# Number of samples
samples = len(dvalues)
# If labels are one-hot encoded,
# turn them into discrete values
if len(y_true.shape) == 2:
y_true = np.argmax(y_true, axis=1)
# Copy so we can safely modify
self.dinputs = dvalues.copy()
# Calculate gradient
self.dinputs[range(samples), y_true] -= 1
# Normalize gradient
self.dinputs = self.dinputs / samples
# Binary cross-entropy loss
class Loss_BinaryCrossentropy(Loss):
# Forward pass
def forward(self, y_pred, y_true):
# Clip data to prevent division by 0
# Clip both sides to not drag mean towards any value
y_pred_clipped = np.clip(y_pred, 1e-7, 1 - 1e-7)
# Calculate sample-wise loss
sample_losses = -(y_true * np.log(y_pred_clipped) +
(1 - y_true) * np.log(1 - y_pred_clipped))
sample_losses = np.mean(sample_losses, axis=-1)
# Return losses
return sample_losses
# Backward pass
def backward(self, dvalues, y_true):
# Number of samples
samples = len(dvalues)
# Number of outputs in every sample
# We'll use the first sample to count them
outputs = len(dvalues[0])
# Clip data to prevent division by 0
# Clip both sides to not drag mean towards any value
clipped_dvalues = np.clip(dvalues, 1e-7, 1 - 1e-7)
# Calculate gradient
self.dinputs = -(y_true / clipped_dvalues -
(1 - y_true) / (1 - clipped_dvalues)) / outputs
# Normalize gradient
self.dinputs = self.dinputs / samples
# Common accuracy class
class Accuracy:
# Calculates an accuracy
# given predictions and ground truth values
def calculate(self, predictions, y):
# Get comparison results
comparisons = self.compare(predictions, y)
# Calculate an accuracy
accuracy = np.mean(comparisons)
# Add accumulated sum of matching values and sample count
# Return accuracy
return accuracy
# Calculates accumulated accuracy
def calculate_accumulated(self):
# Calculate an accuracy
accuracy = self.accumulated_sum / self.accumulated_count
# Return the data and regularization losses
return accuracy
# Reset variables for accumulated accuracy
def new_pass(self):
self.accumulated_sum = 0
self.accumulated_count = 0
# Accuracy calculation for classification model
class Accuracy_Categorical(Accuracy):
def __init__(self, *, binary=False):
# Binary mode?
self.binary = binary
# No initialization is needed
def init(self, y):
pass
# Compares predictions to the ground truth values
def compare(self, predictions, y):
if not self.binary and len(y.shape) == 2:
y = np.argmax(y, axis=1)
return predictions == y
# Accuracy calculation for regression model
class Accuracy_Regression(Accuracy):
def __init__(self):
# Create precision property
self.precision = None
# Calculates precision value
# based on passed-in ground truth values
def init(self, y, reinit=False):
if self.precision is None or reinit:
self.precision = np.std(y) / 250
# Compares predictions to the ground truth values
def compare(self, predictions, y):
return np.absolute(predictions - y) < self.precision
class model:
def __init__(self):
pass
def predict(self, classes, samples):
self.classes = classes
self.samples = samples
self.X, self.y = spiral_data(samples=self.samples, classes=self.classes)
dense1.forward(self.X)
activation1.forward(dense1.output)
dense2.forward(activation1.output)
activation2.forward(dense2.output)
# Calculate the data loss
self.loss = loss_function.calculate(activation2.output, self.y)
self.predictions = (activation2.output > 0.5) * 1
self.accuracy = np.mean(self.predictions == self.y)
print(f'Accuracy: {self.accuracy}')
# + [markdown] id="UA4GFMbIPUkI"
# # Old Dropout Layer
# + id="ZxoiO43tPbTa"
class Layer_Dropout:
# Init
def __init__(self, rate):
# Store rate, we invert it as for example for dropout
# of 0.1 we need success rate of 0.9
self.rate = 1 - rate
# Forward pass
def forward(self, inputs):
# Save input values
self.inputs = inputs
# Generate and save scaled mask
self.binary_mask = np.random.binomial(1, self.rate,
size=inputs.shape) / self.rate
# Apply mask to output values
self.output = inputs * self.binary_mask
# Backward pass
def backward(self, dvalues):
# Gradient on values
self.dinputs = dvalues * self.binary_mask
#print(self.dinputs.shape)
# + [markdown] id="OV_Og9ZrbKtV"
# # New Dropout Layers
# + id="PXBWsHDIUSfh"
class Layer_BinaryNSDropout:
# Init
def __init__(self, rate):
self.rate = 1 - rate
self.iterations = 0
def forward(self, inputs, val_inputs):
self.inputs = inputs
self.val_inputs = val_inputs
nummask = round(len(self.inputs[0]) * self.rate)
#Averaging Values
self.meanarray1 = np.mean(inputs, axis=0)
self.meanarray2 = np.mean(val_inputs, axis=0)
if self.iterations != 0:
# Calculating value
self.difference = self.meanarray1 - self.meanarray2
ind = np.argpartition(self.difference, -nummask)[-nummask:]
mask = np.ones(self.meanarray1.shape, dtype=bool)
mask[ind] = False
self.difference[~mask] = 1
self.difference[mask] = 0.
self.binary_mask = self.difference / self.rate
else:
self.binary_mask = np.random.binomial(1, self.rate,
size=inputs.shape) / self.rate
self.output = inputs * self.binary_mask
def backward(self, dvalues):
# Gradient on values
self.dinputs = dvalues * self.binary_mask
def post_update_params(self):
self.iterations += 1
# + id="WiuCzwWxbRl0"
class Layer_CatagoricalNSDropout:
# Init
def __init__(self, rate):
self.rate = rate
self.iterations = 0
def forward(self, X_test, y_test, X, y):
if self.iterations != 0:
#Adding sorted data into dictionaries
sorted_x = {}
sorted_y = {}
for classes in range(len(set(y))):
sorted_x["class_{0}".format(classes)] = X[y == classes]
sorted_y["label_{0}".format(classes)] = y[y == classes]
sorted_x_test = {}
sorted_y_test = {}
for classes in range(len(set(y))):
sorted_x_test["class_{0}".format(classes)] = X_test[y_test == classes]
sorted_y_test["label_{0}".format(classes)] = y_test[y_test == classes]
#Averaging sorted data from each class then finding the difference between the averaged train and test inputs
differnce_classes = {}
for i, classes, test_classes in zip(range(len(set(y))), sorted_x, sorted_x_test):
differnce_classes["diff_{0}".format(i)] = np.mean(sorted_x[classes], axis=0) - np.mean(sorted_x_test[classes], axis=0)
#Masking the data taking the high values(greatest difference between train and test) and setting their values to 0
self.diff_mask = {}
for i, classes, test_classes, diff in zip(range(len(set(y))), sorted_x, sorted_x_test, differnce_classes):
ind = np.argpartition(differnce_classes[diff], -round(len(X[0]) * self.rate))[-round(len(X[0]) * self.rate):]
mask = np.ones(np.mean(sorted_x[classes],axis=0).shape, dtype=bool)
mask[ind] = False
differnce_classes[diff][~mask] = 0.
differnce_classes[diff][mask] = 1
self.diff_mask["mask_{0}".format(i)] = differnce_classes[diff]
#Goes through each input values and applies the apprioprite mask based on what the true output should be.
binary_mask = np.empty(shape=X.shape)
for i, (input, label) in enumerate(zip(X,y)):
for true, diff in enumerate(self.diff_mask):
if label == true:
self.binary_mask[i] = self.diff_mask[diff]
else:
self.binary_mask = np.random.binomial(1, (1-self.rate), size=X.shape)
self.cached_binary_mask = self.binary_mask
self.output = (self.binary_mask/(1-self.rate)) * X
def backward(self, dvalues):
# Gradient on values
self.dinputs = dvalues * self.binary_mask
def infrence(self, input, label):
self.input = input
self.label = label
idx = np.argsort(self.label)
input_sorted = input[idx]
label_sorted = label[idx]
self.infrence_binary_mask = np.empty(shape=self.input.shape)
for i, (input, label) in enumerate(zip(self.input, self.label)):
#for true, diff in zip(range(len(set(self.label))),self.diff_mask):
for true, diff in enumerate(self.diff_mask):
if label == true:
self.infrence_binary_mask[i] = self.diff_mask[diff]
self.output = self.infrence_binary_mask * self.input
def post_update_params(self):
self.iterations += 1
# + [markdown] id="XRB57nFublm3"
# Initializing Caches
# + id="_kyAX0txV-cF"
loss_cache = []
val_loss_cache = []
acc_cache = []
val_acc_cache = []
lr_cache = []
epoch_cache = []
test_acc_cache = []
test_loss_cache = []
binary_mask_cache = []
max_val_accuracyint = 0
# + [markdown] id="F_7VWnIlF8yx"
# Initializing Summary List
# + id="Xtnu5VToGAq0"
summary = []
# + [markdown] id="W1Eu0pm-WjKI"
# # Loading Data
# + [markdown] id="x-24YuBKre0f"
# Vizulizing Data
# + colab={"base_uri": "https://localhost:8080/"} id="4kUOJ9avrho8" outputId="0b4ead6e-2745-4846-afb4-7522d10fd1fa"
(X, y), (X_val, y_val) = tf.keras.datasets.fashion_mnist.load_data()
# Label index to label name relation
fashion_mnist_labels = {
0: 'T-shirt/top',
1: 'Trouser',
2: 'Pullover',
3: 'Dress',
4: 'Coat',
5: 'Sandal',
6: 'Shirt',
7: 'Sneaker',
8: 'Bag',
9: 'Ankle boot'
}
# Shuffle the training dataset
keys = np.array(range(X.shape[0]))
np.random.shuffle(keys)
X = X[keys]
y = y[keys]
input = X
label = y
X = X[:10000,:,:]
#X_test = X_test[:1600,:,:]
y = y[:10000]
#y_test = y_test[:1600]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=(.2))
# Scale and reshape samples
X = (X.reshape(X.shape[0], -1).astype(np.float32) - 127.5) / 127.5
X_train = (X_train.reshape(X_train.shape[0], -1).astype(np.float32) - 127.5) / 127.5
X_test = (X_test.reshape(X_test.shape[0], -1).astype(np.float32) - 127.5) / 127.5
X_val = (X_val.reshape(X_val.shape[0], -1).astype(np.float32) - 127.5) / 127.5
input = (input.reshape(input.shape[0], -1).astype(np.float32) - 127.5) / 127.5
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# + [markdown] id="i_nW7mqGTnem"
# Sorting Training Data
# + colab={"base_uri": "https://localhost:8080/"} id="AtvA9y81TpGf" outputId="4e5fb0de-a3da-4062-f0c2-40cf9ddb3a65"
idx = np.argsort(y_train)
X_sorted = X_train[idx]
y_sorted = y_train[idx]
sorted_x = {}
sorted_y = {}
for classes in range(len(set(y))):
sorted_x["X_{0}".format(classes)] = X_train[y_train == classes]
sorted_y["y_{0}".format(classes)] = y_train[y_train == classes]
for sorted_lists in sorted_x:
print(f'Number of Samples for {sorted_lists}: {sorted_x[sorted_lists].shape[0]}')
# + [markdown] id="vpaNaUO3kP2G"
# Sorting Testing Data
# + colab={"base_uri": "https://localhost:8080/"} id="TBLFeGAUkSOs" outputId="ecda42ff-1556-40a2-88c4-f04c4fd560e9"
idx = np.argsort(y_test)
X_test_sorted = X_test[idx]
y_test_sorted = y_test[idx]
class_list = []
sorted_x_test = {}
sorted_y_test = {}
for classes in range(len(set(y))):
sorted_x_test["X_test_{0}".format(classes)] = X_test[y_test == classes]
sorted_y_test["y_test_{0}".format(classes)] = y_test[y_test == classes]
for sorted_lists in sorted_x_test:
print(f'Number of Samples for {sorted_lists}: {sorted_x_test[sorted_lists].shape[0]}')
class_list.append(sorted_x_test[sorted_lists].shape[0])
# + colab={"base_uri": "https://localhost:8080/"} id="SSOlE-g40VbR" outputId="018bb086-f88a-4222-eedd-87c11279b7cc"
idx = np.argsort(y_val)
X_val_sorted = X_val[idx]
y_val_sorted = y_val[idx]
class_list = []
sorted_x_val = {}
sorted_y_val = {}
for classes in range(len(set(y))):
sorted_x_val["X_val_{0}".format(classes)] = X_val[y_val == classes]
sorted_y_val["y_val_{0}".format(classes)] = y_val[y_val == classes]
for sorted_lists in sorted_x_val:
print(f'Number of Samples for {sorted_lists}: {sorted_x_val[sorted_lists].shape[0]}')
class_list.append(sorted_x_val[sorted_lists].shape[0])
# + colab={"base_uri": "https://localhost:8080/"} id="6hbnq4TJp1cl" outputId="1ef84a44-6519-42c4-b19d-29f61ec632fe"
print(f'Found {X.shape[0]} images belonging to {len(set(y))} unique classes')
# + [markdown] id="Fd_dSHDNW1Rn"
# # Initializing Layers
# + id="aly5fwUCW_4l"
# Create Dense layer with 2 input features and 64 output values
dense1 = Layer_Dense(X.shape[1], 128, weight_regularizer_l2=5e-4,
bias_regularizer_l2=5e-4)
activation1 = Activation_ReLU()
dropout1 = Layer_CatagoricalNSDropout(0.2)
dense2 = Layer_Dense(128, 128)
activation2 = Activation_ReLU()
dense3 = Layer_Dense(128,128)
activation3 = Activation_ReLU()
dense4 = Layer_Dense(128,len(set(y)))
activation4 = Activation_Softmax()
loss_function = Loss_CategoricalCrossentropy()
softmax_classifier_output = \
Activation_Softmax_Loss_CategoricalCrossentropy()
# Create optimizer
optimizer = Optimizer_Adam(decay=5e-7,learning_rate=0.005)
#optimizer = Optimizer_SGD(learning_rate=0.01)
accuracy = Accuracy_Categorical()
accuracy.init(y)
# + [markdown] id="-xmbxDuwXIBk"
# # Training Loop
# + colab={"base_uri": "https://localhost:8080/"} id="14yHOjq9XLee" outputId="fc58b2ee-d21c-45c0-afa8-3801676bb0e5"
epochs = 223
bmc = []
full_bmc = []
for epoch in range(epochs + 1):
dense1.forward(X_train)
activation1.forward(dense1.output)
if epoch != 0:
cached_val_inputs = cached_val_inputs
cached_train_inputs = activation1.output
else:
cached_val_inputs = np.random.random(size=128) #Never used just needed to pass to dropout
cached_train_inputs = activation1.output
dropout1.forward(X=activation1.output, y=y_train, X_test=cached_val_inputs, y_test=y_test)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
data_loss = loss_function.calculate(activation4.output, y_train)
regularization_loss = \
loss_function.regularization_loss(dense1) + \
loss_function.regularization_loss(dense2) + \
loss_function.regularization_loss(dense3) + \
loss_function.regularization_loss(dense4)
loss = data_loss + regularization_loss
#Accuracy
predictions = activation4.predictions(activation4.output)
train_accuracy = accuracy.calculate(predictions, y_train)
# Backward pass
softmax_classifier_output.backward(activation4.output, y_train)
activation4.backward(softmax_classifier_output.dinputs)
dense4.backward(activation4.dinputs)
activation3.backward(dense4.dinputs)
dense3.backward(activation3.dinputs)
activation2.backward(dense3.dinputs)
dense2.backward(activation2.dinputs)
dropout1.backward(dense2.dinputs)
activation1.backward(dropout1.dinputs)
dense1.backward(activation1.dinputs)
# Update weights and biases
optimizer.pre_update_params()
optimizer.update_params(dense1)
optimizer.update_params(dense2)
optimizer.update_params(dense3)
optimizer.update_params(dense4)
optimizer.post_update_params()
dropout1.post_update_params()
#print(dropout1.binary_mask.shape)
#print(dropout1.binary_mask[0])
bmc.append(dropout1.binary_mask[0].tolist())
full_bmc.append(dropout1.binary_mask.tolist())
#print(bmc[epoch-1])
# Validation
dense1.forward(X_test)
activation1.forward(dense1.output)
if epoch == 0:
dense2.forward(activation1.output)
else:
dropout1.infrence(activation1.output,y_test)
dense2.forward(dropout1.output)
dense1_outputs = dense1.output
meanarray = np.mean(dense1.output, axis=0)
cached_val_inputs = activation1.output
trainout = meanarray
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
valloss = loss_function.calculate(activation4.output, y_test)
predictions = activation4.predictions(activation4.output)
valaccuracy = accuracy.calculate(predictions, y_test)
#Unseen Validaiton Accuracy
dense1.forward(X_val)
activation1.forward(dense1.output)
if epoch == 0:
dense2.forward(activation1.output)
else:
dropout1.infrence(activation1.output,y_val)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
testloss = loss_function.calculate(activation4.output, y_val)
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, y_val)
#Updating List
loss_cache.append(loss)
val_loss_cache.append(valloss)
acc_cache.append(train_accuracy)
val_acc_cache.append(valaccuracy)
lr_cache.append(optimizer.current_learning_rate)
epoch_cache.append(epoch)
test_acc_cache.append(testaccuracy)
test_loss_cache.append(testloss)
#Summary Items
if valaccuracy >= .8 and len(summary) == 0:
nintypercent = f'Model hit 80% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if valaccuracy >= .85 and len(summary) == 1:
nintypercent = f'Model hit 85% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if valaccuracy >= .9 and len(summary) == 2:
nintypercent = f'Model hit 90% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if valaccuracy >= .95 and len(summary) == 3:
nintypercent = f'Model hit 95% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if valaccuracy >= .975 and len(summary) == 4:
nintypercent = f'Model hit 97.5% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if valaccuracy >= 1 and len(summary) == 5:
nintypercent = f'Model hit 100% validation accuracy in {epoch} epochs'
summary.append(nintypercent)
if epoch == epochs:
if valaccuracy > max_val_accuracyint:
max_val_accuracyint = valaccuracy
max_val_accuracy = f'Max accuracy was {valaccuracy * 100}% at epoch {epoch}.'
summary.append(max_val_accuracy)
else:
summary.append(max_val_accuracy)
else:
if valaccuracy > max_val_accuracyint:
max_val_accuracyint = valaccuracy
max_val_accuracy = f'Max accuracy was {valaccuracy * 100}% at epoch {epoch}.'
if not epoch % 1:
print(f'epoch: {epoch}, ' +
f'acc: {train_accuracy:.3f}, ' +
f'loss: {loss:.3f} (' +
f'data_loss: {data_loss:.3f}, ' +
f'reg_loss: {regularization_loss:.3f}), ' +
f'lr: {optimizer.current_learning_rate:.9f} ' +
f'validation, acc: {valaccuracy:.3f}, loss: {valloss:.3f} ' +
f'Unseen, acc: {testaccuracy:.3f}, loss: {testloss:.3f} ')
# + [markdown] id="GR0u0Jm7QCrw"
# # Summary
# + colab={"base_uri": "https://localhost:8080/"} id="ay7eT5DBihZ0" outputId="cd52bec5-3fc7-4e2f-b03f-92da0ccb9cdd"
range(len(bmc))
# + colab={"base_uri": "https://localhost:8080/"} id="pibv5C5bX6Ad" outputId="66bd2229-bc7f-483a-89e6-fa51188f89c4"
count = 0
count_list = []
for i in bmc:
i = np.array(i)
for i in range(len(bmc)):
count = 0
for j in range(len(bmc[i])):
if i != len(bmc) - 1:
if bmc[i][j] != bmc[i+1][j]:
count += 1
count_list.append(count)
print(count_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="NOfygjgHb5Sw" outputId="37800118-189f-4292-9009-3f62129220e1"
plt.plot(count_list)
plt.title('Values Changed')
plt.xlabel('Epoch')
plt.ylabel('Values Changed')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="yTkECLnyeSbI" outputId="b4534c58-88ef-4dd0-ba45-13bea255b9fb"
print(len(full_bmc[1]))
# + colab={"base_uri": "https://localhost:8080/"} id="n0CK2ke1eOnH" outputId="22fa8692-d427-437a-add9-07e4e2d04806"
count = 0
average_count = 0
count_list = []
average_list = []
for q in range(len(full_bmc[1])):
count_list = []
for i in range(len(full_bmc)):
count = 0
for j in range(len(full_bmc[i][q])):
if i != len(full_bmc) - 1:
if full_bmc[i][q][j] != full_bmc[i+1][q][j]:
count += 1
count_list.append(count)
average_list.append(count_list)
final_list = []
average_values = []
for i in range(len(average_list[1])):
for j in range(len(average_list)):
average_values.append(average_list[j][i])
final_list.append(statistics.mean(average_values))
print(final_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 294} id="uxPM6iRcgrCa" outputId="b2cb7fc7-6ee4-44df-b60f-98284e22bbf7"
plt.plot(final_list)
plt.title('Values Changed / Epoch')
plt.xlabel('Epoch')
plt.ylabel('Values Changed')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="nrQbkCEMR4Z4" outputId="e508cbe0-3f18-4670-89ee-860111b5746b"
print(len(binary_mask_cache))
# + id="1KNnDUP_U8Xn" colab={"base_uri": "https://localhost:8080/"} outputId="0eb19004-d2d9-4175-8d32-386abde10416"
print(np.mean(acc_cache))
# + id="2NbXMisqQKqF" colab={"base_uri": "https://localhost:8080/"} outputId="420c8b52-12b6-4be4-a4ce-f9dc230497f1"
for milestone in summary:
print(milestone)
# + [markdown] id="_rVqT3yaXS5k"
# # Testing
# + id="smwSXsZVU8Xo" colab={"base_uri": "https://localhost:8080/"} outputId="503adac4-b3fa-4173-d9ee-ffda1c8b4607"
accuracy = Accuracy_Categorical()
accuracy.init(y_test)
dense1.forward(X_test)
activation1.forward(dense1.output)
dropout1.infrence(activation1.output,y_test)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
index = 27
print(f'{(activation4.output[index][np.where(activation4.output[index] == np.amax(activation4.output[index]))][0]*100):.3f}% Confident True is {fashion_mnist_labels[np.where(activation4.output[index] == np.amax(activation4.output[index]))[0][0]]}. True is actually {fashion_mnist_labels[y_test[index]]}')
# Calculate the data loss
loss = loss_function.calculate(activation4.output, y_test)
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, y_test)
print(f'Accuracy: {testaccuracy:.3f}, loss: {loss:.3f}')
# + id="1k0Ve2M0bPG3"
training_diff = []
testing_diff = []
combined_diff = []
# + [markdown] id="MByL_RwvlIx3"
# Individual Training Classes
# + id="YTOnqnDXa0ME" colab={"base_uri": "https://localhost:8080/"} outputId="17732184-c669-42cb-d78b-f6bc7106b853"
accuracy = Accuracy_Categorical()
for classes, (X_sorted_lists, y_sorted_lists) in enumerate(zip(sorted_x, sorted_y)):
accuracy = Accuracy_Categorical()
y = sorted_y[y_sorted_lists]
X = sorted_x[X_sorted_lists]
accuracy.init(y)
dense1.forward(X)
activation1.forward(dense1.output)
train_train_mean = activation1.output
dropout1.infrence(activation1.output,y)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
loss = loss_function.calculate(activation4.output, y)
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, y)
print(f'{fashion_mnist_labels[classes]} Train Accuracy: {testaccuracy:.3f}, loss: {loss:.3f}')
# + id="scjb7Wh_sn6b" colab={"base_uri": "https://localhost:8080/"} outputId="406ccc17-2a5a-4256-9d09-d3d86e60b330"
accuracy = Accuracy_Categorical()
for classes, (X_sorted_lists, y_sorted_lists) in enumerate(zip(sorted_x_val, sorted_y_val)):
accuracy.init(sorted_y_val[y_sorted_lists])
#print(sorted_y[y_sorted_lists].shape)
#print(sorted_x[X_sorted_lists].shape)
dense1.forward(sorted_x_val[X_sorted_lists])
activation1.forward(dense1.output)
testmean = np.mean(activation1.output, axis=0)
testing_diff.append(testmean)
dropout1.infrence(activation1.output,sorted_y_val[y_sorted_lists])
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
loss = loss_function.calculate(activation4.output, sorted_y_val[y_sorted_lists])
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, sorted_y_val[y_sorted_lists])
print(f'{fashion_mnist_labels[classes]} Test Accuracy: {testaccuracy:.3f}, loss: {loss:.3f}')
# + [markdown] id="R2u-O8oNZ0qA"
# # Full mnist test
# + [markdown] id="UbD4KrLMnTcR"
# Training data
# + id="TMfBGUHeZ4L5" colab={"base_uri": "https://localhost:8080/"} outputId="d60544f7-fc73-4e50-a9fc-847c5152688a"
accuracy = Accuracy_Categorical()
accuracy.init(label)
dense1.forward(input)
activation1.forward(dense1.output)
train_train_mean = activation1.output
dropout1.infrence(activation1.output,label)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
loss = loss_function.calculate(activation4.output, label)
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, label)
print(f'Found {input.shape[0]} images belonging to {len(set(label))} unique classes')
print(f'Full Training Accuracy: {testaccuracy:.5f}, loss: {loss:.3f}')
# + [markdown] id="Aat-uVF6nYu7"
# Testing data
# + id="4hyU1tBDna8x" colab={"base_uri": "https://localhost:8080/"} outputId="29d4ae4d-21e6-4abf-9823-13083a56d19b"
(X, y), (X_val, y_val) = tf.keras.datasets.fashion_mnist.load_data()
X_val = (X_val.reshape(X_val.shape[0], -1).astype(np.float32) - 127.5) / 127.5 # Reshape X_val if cell below was already ran
accuracy = Accuracy_Categorical()
accuracy.init(y_val)
dense1.forward(X_val)
activation1.forward(dense1.output)
dropout1.infrence(activation1.output,y_val)
dense2.forward(dropout1.output)
activation2.forward(dense2.output)
dense3.forward(activation2.output)
activation3.forward(dense3.output)
dense4.forward(activation3.output)
activation4.forward(dense4.output)
# Calculate the data loss
loss = loss_function.calculate(activation4.output, y_val)
predictions = activation4.predictions(activation4.output)
testaccuracy = accuracy.calculate(predictions, y_val)
print(f'Found {X_val.shape[0]} images belonging to {len(set(y_val))} unique classes')
print(f'Full Testing Accuracy: {testaccuracy:.5f}, loss: {loss:.3f}')
# + id="dHjYxBoURAgv" colab={"base_uri": "https://localhost:8080/", "height": 736} outputId="c023466c-ca22-49f4-ba55-c144b1f81b88"
predicted_list = []
true_list = []
for sample in range(len(X_val)):
predicted_list.append(fashion_mnist_labels[np.where(activation4.output[sample] == np.amax(activation4.output[sample]))[0][0]])
true_list.append(fashion_mnist_labels[y_val[sample]])
from sklearn import metrics
import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
array = metrics.confusion_matrix(true_list, predicted_list, labels=['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot'])
df_cm = pd.DataFrame(array, range(len(set(true_list))), range(len(set(true_list))))
df_cm.round(9)
plt.figure(figsize=(10,7))
sn.set(font_scale=1.2) # for label size
sn.heatmap(df_cm, annot=True, annot_kws={"size": 12}, fmt='g') # font size
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
# Printing the precision and recall, among other metrics
print(metrics.classification_report(true_list, predicted_list, labels=['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']))
# + id="l3JzV--WNNT7" colab={"base_uri": "https://localhost:8080/"} outputId="431cc989-6b22-4f22-8ab0-48a33d2bf5bb"
for index in range(10000):
if y_val[index] != np.where(activation4.output[index] == np.amax(activation4.output[index]))[0][0]:
print(index)
# + [markdown] id="AbIMZ7Pk_Tnp"
# Change idex to get confidence of different samples of testing data. Index values 0-1600 were refrenced in training. Anything past was never seen during training. Lowest confidence is at index 2732 when trained with 488 epochs and numpy seed set to 22.
# + id="JaxWcRIr_BCV" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="9c0bd661-0855-424f-ac6d-0e2fc6660b51"
index = 5674
print(f'{(activation4.output[index][np.where(activation4.output[index] == np.amax(activation4.output[index]))][0]*100):.3f}% Confident True is {fashion_mnist_labels[np.where(activation4.output[index] == np.amax(activation4.output[index]))[0][0]]}. True is actually {fashion_mnist_labels[y_val[index]]}')
X_val.resize(X_val.shape[0],28,28)
image = X_val[index]
plt.rcParams['axes.grid'] = False
fig = plt.figure
plt.title(f'{fashion_mnist_labels[y_val[index]]}')
plt.imshow(image, cmap='gray')
plt.show()
# + id="p6KeoLkg7k0u" colab={"base_uri": "https://localhost:8080/"} outputId="69d7c493-ecdf-499f-cd67-94a89e69d0a6"
confidence_list = []
for index in range(10000):
confidence_list.append(activation4.output[index][np.where(activation4.output[index] == np.amax(activation4.output[index]))][0])
print(confidence_list.index(min(confidence_list)))
a = confidence_list[:]
a.sort()
print(confidence_list.index(a[1]))
# + [markdown] id="MRXGM4hyXmr7"
# Plotting Graphs
# + id="h5c5xUTNXk2v" colab={"base_uri": "https://localhost:8080/", "height": 881} outputId="08e9ef04-3df2-4d77-9e55-5dd535358330"
plt.plot(epoch_cache, val_loss_cache, label='Validation Loss')
plt.plot(epoch_cache, loss_cache, label='Training Loss')
plt.title('Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend(loc = "upper right")
plt.show()
plt.plot(epoch_cache, val_acc_cache, label='Validation Accuracy')
plt.plot(epoch_cache, acc_cache, label='Training Accuracy')
plt.title('Accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.legend(loc = "upper right")
plt.show()
plt.plot(epoch_cache, lr_cache, label='LR')
plt.title('Learning Rate')
plt.xlabel('Epoch')
plt.ylabel('Learning Rate')
plt.show()
| mnist_implementation_of_New_Dropout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
a = np.arange(12).reshape(3, 4)
print(a)
b = np.arange(-5, 7).reshape(3, 4)
print(b)
a_copysign = np.copysign(a, b)
print(a_copysign)
print(a_copysign.dtype)
print(np.copysign(10, -5))
print(type(np.copysign(10, -5)))
print(a)
b_small = np.array([-100, -100, 100, 100])
print(b_small)
print(a + b_small)
print(np.copysign(a, b_small))
b_mismatch = np.array([-100, -100, 100])
print(b_mismatch)
# +
# print(np.copysign(a, b_mismatch))
# ValueError: operands could not be broadcast together with shapes (3,4) (3,)
# -
print(np.copysign(b, -10))
print(np.abs(b) * -1)
print(np.abs(b) * -1.0)
a_special = np.array([0.0, -0.0, np.inf, -np.inf, np.nan])
print(a_special)
print(np.copysign(a_special, 1))
print(np.copysign(a_special, -1))
print(np.copysign([10, 10, 10, 10, 10], a_special))
print(np.copysign([-10, -10, -10, -10, -10], a_special))
print(np.copysign(10, 0))
print(np.copysign(0, 10))
print(np.copysign(0, -10))
a_complex = np.array([10 + 10j, -10 + 10j])
print(a_complex)
# +
# print(np.copysign(a_complex, 1))
# TypeError: ufunc 'copysign' not supported for the input types, and the inputs could not be safely coerced to any supported types according to the casting rule ''safe''
# +
# print(np.copysign([1, 1], a_complex))
# TypeError: ufunc 'copysign' not supported for the input types, and the inputs could not be safely coerced to any supported types according to the casting rule ''safe''
| notebook/numpy_copysign.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import pandas as pd
import yfinance as yf
from ta.others import DailyReturnIndicator,CumulativeReturnIndicator
# + pycharm={"name": "#%%\n"}
gspc= yf.Ticker("^GSPC")
df = gspc.history(start= '2007-01-01', end= '2020-12-31')
# + pycharm={"name": "#%%\n"}
df.head()
# + pycharm={"name": "#%%\n"}
daily_returns = DailyReturnIndicator(close=df["Close"])
df["daily_returns"] = daily_returns.daily_return()
# + pycharm={"name": "#%%\n"}
df["daily_returns"]
# + pycharm={"name": "#%%\n"}
df["daily_returns"].to_csv('sp500_returns.csv')
| SB3/SP500_Returns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Quick concept of Machine Learning
#
# Most machine learning algorithms share the same pipeline structure as the one shown below:
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_0.png" width="600"/>
#
#
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_1.png" width="600"/>
#
#
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_2.png" width="600"/>
#
#
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_3.png" width="600"/>
#
#
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_4.png" width="600"/>
#
#
# <img style="border:1px solid black;" src="https://raw.githubusercontent.com/qingkaikong/20171206_ML_basics_THW/master/images/image_5.png" width="600"/>
#
# # Scikit-learn Basics
#
# This session will cover the basics of Scikit-Learn, a popular package containing a collection of tools for machine learning written in Python. See more at http://scikit-learn.org.
# ## Loading an example dataset
#
# Let's start by loading some [pre-existing datasets](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.datasets) in the scikit-learn, which comes with a few standard datasets. For example, the [iris](https://en.wikipedia.org/wiki/Iris_flower_data_set) and [digits](http://archive.ics.uci.edu/ml/datasets/Pen-Based+Recognition+of+Handwritten+Digits) datasets for classification and the [boston house prices](http://archive.ics.uci.edu/ml/datasets/Housing) dataset for regression. Using these existing datasets, we can easily test the algorithms that we are interested in.
#
# A dataset is a dictionary-like object that holds all the data and some metadata about the data. This data is stored in the .data member, which is a n_samples, n_features array. In the case of supervised problem, one or more response variables are stored in the .target member. More details on the different datasets can be found in the dedicated section.
# ### Load iris
#
# The iris dataset consists of 50 samples from each of three species of Iris (Iris setosa, Iris virginica and Iris versicolor). Four features were measured from each sample: the length and the width of the sepals and petals, in centimetres.
#
#
# | [](https://en.wikipedia.org/wiki/Iris_setosa) | [](https://en.wikipedia.org/wiki/Iris_virginica) | [](https://en.wikipedia.org/wiki/Iris_versicolor) |
# |:---:|:---:|:---:|
# | <NAME>| Iris Virginica| Iris Versicolor|
# +
import warnings
warnings.filterwarnings("ignore")
from sklearn import datasets
import matplotlib.pyplot as plt
import numpy as np
plt.style.use('seaborn-poster')
# %matplotlib inline
# -
iris = datasets.load_iris()
print(iris.feature_names)
# only print the first 10 samples
print(iris.data[:10])
print('We have %d data samples with %d features'%(iris.data.shape[0], iris.data.shape[1]))
# The data is always a 2D array, shape (n_samples, n_features), although the original data may have had a different shape. The following prints out the target names and the representatoin of the target using 0, 1, 2. Each of them represent a class.
print(iris.target_names)
print(set(iris.target))
# ### Load Digits
#
# This dataset is made up of 1797 8x8 images. Each image, like the ones shown below, is of a hand-written digit. In order to utilize an 8x8 figure like this, we’d have to first transform it into a feature vector with length 64.
digits = datasets.load_digits()
print('We have %d samples'%len(digits.target))
print(digits.data)
print('The targets are:')
print(digits.target_names)
# In the digits, each original sample is an image of shape (8, 8) and it is flattened into a 64 dimension vector here.
print(digits.data.shape)
## plot the first 64 samples, and get a sense of the data
fig = plt.figure(figsize = (8,8))
fig.subplots_adjust(left=0, right=1, bottom=0, top=1, hspace=0.05, wspace=0.05)
for i in range(64):
ax = fig.add_subplot(8, 8, i+1, xticks=[], yticks=[])
ax.imshow(digits.images[i],cmap=plt.cm.binary,interpolation='nearest')
ax.text(0, 7, str(digits.target[i]))
# ### Load Boston Housing Data
#
# The Boston housing dataset reports the median value of owner-occupied homes in various places in the Boston area, together with several variables which might help to explain the variation in median value, such as Crime (CRIM), areas of non-retail business in the town (INDUS), the age of people who own the house (AGE), and there are many other attributes that you can find the details [here](https://archive.ics.uci.edu/ml/datasets/housing).
boston = datasets.load_boston()
print(boston.DESCR)
boston.feature_names
# let's just plot the average number of rooms per dwelling with the price
plt.figure(figsize = (10,8))
plt.plot(boston.data[:,5], boston.target, 'o')
plt.xlabel('Number of rooms')
plt.ylabel('Price (thousands)')
# ## The Scikit-learn Estimator Object
#
# Every algorithm is exposed in scikit-learn via an ''Estimator'' object. For instance a linear regression is implemented as so:
from sklearn.linear_model import LinearRegression
# **Estimator parameters**: All the parameters of an estimator can be set when it is instantiated, and have suitable default values:
# +
# you can check the parameters as
# LinearRegression?
# -
# let's change one parameter
model = LinearRegression(normalize=True)
print(model.normalize)
print(model)
# **Estimated Model parameters**: When data is *fit* with an estimator, parameters are estimated from the data at hand. All the estimated parameters are attributes of the estimator object ending by an underscore:
# ### Simple regression problem
#
# Let's fit a simple linear regression model to see what is the sklearn API looks like. We use a very simple datasets with 10 samples with added in noise.
x = np.arange(10)
y = 2 * x + 1
plt.figure(figsize = (10,8))
plt.plot(x,y,'o')
# +
# generate noise between -1 to 1
# this seed is just to make sure your results are the same as mine
np.random.seed(42)
noise = 2 * np.random.rand(10) - 1
# add noise to the data
y_noise = y + noise
# -
plt.figure(figsize = (10,8))
plt.plot(x,y_noise,'o')
# The input data for sklearn is 2D: (samples == 10 x features == 1)
X = x[:, np.newaxis]
print(X)
print(y_noise)
# model fitting is via the fit function
model.fit(X, y_noise)
# underscore at the end indicates a fit parameter
print(model.coef_)
print(model.intercept_)
# then we can use the fitted model to predict new data
predicted = model.predict(X)
plt.figure(figsize = (10,8))
plt.plot(x,y_noise,'o')
plt.plot(x,predicted, label = 'Prediction')
plt.legend()
# ## Exercise
#
# In the next section, we will use support vector machine (SVM) to do a classification problem, have a look of the [sklearn API](http://scikit-learn.org/stable/modules/classes.html), and find out which class we will use for a classification problem using SVM. (hint: we will use the one with C support).
# %load ../solutions/solution_01.py
| code_examples/python_regression_classification/notebooks/01_Scikit-learn_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D2_DynamicNetworks/student/W3D2_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Oh1wn4afauo0"
# # Neuromatch Academy: Week 3, Day 2, Tutorial 2
#
# # Neuronal Network Dynamics: Wilson-Cowan Model
#
# + [markdown] colab_type="text" id="A9pNslIYayZt"
# ## Objectives
# In the previous tutorial, you became familiar with a neuronal network consisting of only an excitatory population. Here, we extend the approach we used to include both excitatory and inhibitory neuronal populations in our network. A simple, yet powerful, model to study the dynamics of two interacting populations of excitatory and inhibitory neurons is the so-called **Wilson-Cowan** rate model, which will be the subject of this tutorial.
#
# The objectives of this tutorial are to:
#
# - Write the **Wilson-Cowan** equations for the firing rate dynamics of a 2D system composed of an excitatory (E) and an inhibitory (I) population of neurons
# - Simulate the dynamics of the system, i.e., Wilson-Cowan model.
# - Plot the frequency-current (F-I) curves for both populations (i.e., E and I).
# - Visualize and inspect the behavior of the system using **phase plane analysis**, **vector fields**, and **nullclines**.
#
# Bonus steps:
#
# - Find and plot the **fixed points** of the Wilson-Cowan model.
# - Investigate the stability of the Wilson-Cowan model by linearizing its dynamics and examining the **Jacobian matrix**.
# - Learn how the Wilson-Cowan model can reach an oscillatory state.
#
# Bonus steps (applications):
# - Visualize the behavior of an Inhibition-stabilized network.
# - Simulate working memory using the Wilson-Cowan model.
#
# \\
# Reference paper:
#
# _[<NAME> and <NAME> (1972) Excitatory and inhibitory interactions in localized populations of model neurons. Biophysical Journal 12](https://doi.org/10.1016/S0006-3495(72)86068-5)_
# + [markdown] colab_type="text" id="ACCJbmDqtLfS"
# ## Setup
# + [markdown] colab_type="text" id="BptrqpMma6c3"
# Please execute the cell below to initialize the notebook environment.
# + cellView="both" colab={} colab_type="code" id="QG7r5GjEadue"
# Imports
import matplotlib.pyplot as plt # import matplotlib
import numpy as np # import numpy
import scipy.optimize as opt # import root-finding algorithm
import ipywidgets as widgets # interactive display
# + cellView="form" colab={} colab_type="code" id="JteKm2l1tYrV"
#@title Figure Settings
# %matplotlib inline
fig_w, fig_h = 8, 4.5
my_fontsize = 16
my_params = {'axes.labelsize': my_fontsize,
'axes.titlesize': my_fontsize,
'figure.figsize': [fig_w, fig_h],
'font.size': my_fontsize,
'legend.fontsize': my_fontsize-4,
'lines.markersize': 8.,
'lines.linewidth': 2.,
'xtick.labelsize': my_fontsize-2,
'ytick.labelsize': my_fontsize-2}
plt.rcParams.update(my_params)
# + cellView="form" colab={} colab_type="code" id="T0GZb4qxbJCj"
#@title Helper functions
def default_pars( **kwargs):
pars = {}
### Excitatory parameters ###
pars['tau_E'] = 1. # Timescale of the E population [ms]
pars['a_E'] = 1.2 # Gain of the E population
pars['theta_E'] = 2.8 # Threshold of the E population
### Inhibitory parameters ###
pars['tau_I'] = 2.0 # Timescale of the I population [ms]
pars['a_I'] = 1.0 # Gain of the I population
pars['theta_I'] = 4.0 # Threshold of the I population
### Connection strength ###
pars['wEE'] = 9. # E to E
pars['wEI'] = 4. # I to E
pars['wIE'] = 13. # E to I
pars['wII'] = 11. # I to I
### External input ###
pars['I_ext_E'] = 0.
pars['I_ext_I'] = 0.
### simulation parameters ###
pars['T'] = 50. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
pars['E_init'] = 0.2 # Initial value of E
pars['I_init'] = 0.2 # Initial value of I
### External parameters if any ###
for k in kwargs:
pars[k] = kwargs[k]
pars['range_t'] = np.arange(0, pars['T'], pars['dt']) # Vector of discretized time points [ms]
return pars
def F(x,a,theta):
"""
Population activation function.
Args:
x : the population input
a : the gain of the function
theta : the threshold of the function
Returns:
f : the population activation response f(x) for input x
"""
# add the expression of f = F(x)
f = (1+np.exp(-a*(x-theta)))**-1 - (1+np.exp(a*theta))**-1
return f
def my_test_plot(t, E1, I1, E2, I2):
ax = plt.subplot(2,1,1)
ax.plot(pars['range_t'], E1, 'b', label='E population')
ax.plot(pars['range_t'], I1, 'r', label='I population')
ax.set_xlabel('t (ms)')
ax.set_ylabel('Activity')
ax.legend(loc='best')
ax = plt.subplot(2,1,2)
ax.plot(pars['range_t'], E2, 'b', label='E population')
ax.plot(pars['range_t'], I2, 'r', label='I population')
ax.set_xlabel('t (ms)')
ax.set_ylabel('Activity')
ax.legend(loc='best')
plt.tight_layout()
def my_plot_nullcline(pars):
E_grid = np.linspace(-0.01,0.96,100)
E_nullcline = get_E_nullcline(pars, E_grid)# calculate E nullclines
I_grid = np.linspace(-.01,0.8,100)
I_nullcline = get_I_nullcline(pars, I_grid)# calculate I nullclines
plt.plot(E_grid, E_nullcline, 'b', label='E nullcline')
plt.plot(I_nullcline, I_grid, 'r', label='I nullcline')
plt.xlabel('E')
plt.ylabel('I')
plt.legend(loc='best',fontsize=12)
def my_plot_vector(pars, n_skip=2., scale=5):
EI_grid = np.linspace(0., 1., 20)
E_meshgrid, I_meshgrid = np.meshgrid(EI_grid,EI_grid)
dEdt, dIdt = EIderivs(E_meshgrid, I_meshgrid, pars)
n_skip = 2
plt.quiver(E_meshgrid[::n_skip,::n_skip], I_meshgrid[::n_skip,::n_skip],
dEdt[::n_skip,::n_skip], dIdt[::n_skip,::n_skip],
angles='xy', scale_units='xy', scale=5,facecolor='c')
plt.xlabel('E')
plt.ylabel('I')
def my_plot_trajectory(pars, mycolor, x_init, mylabel):
pars['E_init'], pars['I_init'] = x_init[0], x_init[1]
E_tj, I_tj= simulate_wc(pars)
plt.plot(E_tj, I_tj, color=mycolor, label=mylabel)
plt.plot(x_init[0], x_init[1], 'o', color=mycolor, ms=8)
plt.xlabel('E')
plt.ylabel('I')
def my_plot_trajectories(pars, dx, n, mylabel):
"""
Solve for I along the E_grid from dE/dt = 0.
Expects:
pars : Parameter dictionary
dx : increment of initial values
n : n*n trjectories
mylabel : label for legend
Returns:
figure of trajectory
"""
for ie in range(n):
for ii in range(n):
pars['E_init'], pars['I_init'] = dx*ie, dx*ii
E_tj, I_tj= simulate_wc(pars)
if (ie==n-1)&(ii==n-1):
plt.plot(E_tj, I_tj, 'k', alpha=0.3, label=mylabel)
else:
plt.plot(E_tj, I_tj, 'k', alpha=0.3)
plt.xlabel('E')
plt.ylabel('I')
def check_fp(x_fp):
dEdt, dIdt = EIderivs(x_fp[0], x_fp[1], pars)
return dEdt**2 + dIdt**2<1e-6
def plot_fp(x_fp, mycolor):
plt.plot(x_fp_2[0], x_fp_2[1], 'o', color=mycolor, ms=8)
def dF(x,a,theta):
"""
Population activation function.
Args:
x : the population input
a : the gain of the function
theta : the threshold of the function
Returns:
the population activation response F(x) for input x
"""
dFdx = a*np.exp(-a*(x-theta))*(1+np.exp(-a*(x-theta)))**-2
return dFdx
# + [markdown] colab_type="text" id="8YerINCmfH6O"
# # Wilson-Cowan model of excitatory and inhibitory populations
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 519} colab_type="code" id="o6sG8Enm0xBG" outputId="31d990b9-11cf-479f-d3d5-6330e27a091a"
#@title Video: Phase analysis of the Wilson-Cowan E-I model
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="EgEad5Me_Ro", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="kHzQkZQ4QzdD"
# Many of the rich dynamics recorded in the brain are generated by the interaction of excitatory and inhibitory subtype neurons. Here, similar to what we did in the previous tutorial, we will model two coupled populations of E and I neurons (**Wilson-Cowan** model). We can, in fact, write two rate-based equations, one for each population, with interacting terms:
#
# \begin{align}
# \tau_E \frac{dE}{dt} &= -E + F_E(w_{EE}E -w_{EI}I + I^{\text{ext}}_E;a_E,\theta_E)\\
# \tau_I \frac{dI}{dt} &= -I + F_I(w_{IE}E -w_{II}I + I^{\text{ext}}_I;a_I,\theta_I) \qquad (1)
# \end{align}
#
# $E(t)$ represents the average activation (or firing rate) of the excitatory population at time $t$, and $I(t)$ the activation (or firing rate) of the inhibitory population. The parameters $\tau_E$ and $\tau_I$ control the timescales of the dynamics of each population. Connection strengths are given by: $w_{EE}$ (E $\rightarrow$ E), $w_{EI}$ (I $\rightarrow$ E), $w_{IE}$ (E $\rightarrow$ I), and $w_{II}$ (I $\rightarrow$ I). The terms $w_{EI}$ and $w_{IE}$ are the interacting terms and, respectively, represent connections from inhibitory to excitatory population and vice versa. The transfer functions (or F-I curves) $F_E(x;a_E,\theta_E)$ and $F_I(x;a_I,\theta_I)$ can be different for the excitatory and the inhibitory populations.
#
# \\
#
# Now execute the cell below to initialize simulation parameters and define the handler functions we will use throughout the tutorial.
# + [markdown] colab_type="text" id="nTVNcF9ebXhm"
# ## Exercise 1: plot out the f-I curves for the E and I populations
#
# Let's first plot out the f-I curves for the E and I populations by the function the defined above with the default values
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="SKmZ16knbboa" outputId="2e1a2362-2741-40c2-c9b6-5874bcd4f474"
# Exercise 1
pars = default_pars() # get the default value
x = np.arange(0,10,.1) # set the input
print(pars['a_E'], pars['theta_E'])
print(pars['a_I'], pars['theta_I'])
###################################################################
## TODO for students: compute and plot the F-I curve here #
## Note: aE, thetaE, aI and theta_I are in the dictionray 'pars' #
###################################################################
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 369} colab_type="text" id="Ek7KFT1fMDv0" outputId="0db4603e-1f44-4b43-d661-c313215e318e"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_187031a9.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=513 height=357 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_187031a9_3.png>
#
#
# + [markdown] colab_type="text" id="lQSpCEuTbjpj"
# ## Simulation scheme for the Wilson-Cowan model
#
# Using the Euler method, the E-I dynamical system can be simulated on a time-grid of stepsize $\Delta t$. The updates for the activity of the excitatory and the inhibitory populations can be written as:
#
# \begin{align}
# E[k+1] &= E[k] + \frac{\Delta t}{\tau_E}[-E[k] + F_E(w_{EE}E[k] -w_{EI}I[k] + I^{\text{ext}}_E[k];a_E,\theta_E)]\\
# I[k+1] &= I[k] + \frac{\Delta t}{\tau_I}[-I[k] + F_I(w_{IE}E[k] -w_{II}I[k] + I^{\text{ext}}_I[k];a_I,\theta_I)]
# \end{align}
# + [markdown] colab_type="text" id="UoDULUySMoPj"
# ### Exercise 2: Numerically integrate the Wilson-Cowan equations
# + colab={} colab_type="code" id="jwbliV-0Mpsx"
# Exercise 2
def simulate_wc(pars):
"""
Simulate the Wilson-Cowan equations
Args:
pars : Parameter dictionary
Returns:
E : Activity of excitatory population (array)
I : Activity of inhibitory population (array)
"""
# Set parameters
tau_E, a_E, theta_E = pars['tau_E'], pars['a_E'], pars['theta_E']
tau_I, a_I, theta_I = pars['tau_I'], pars['a_I'], pars['theta_I']
wEE, wEI = pars['wEE'], pars['wEI']
wIE, wII = pars['wIE'], pars['wII']
I_ext_E, I_ext_I = pars['I_ext_E'], pars['I_ext_I']
E_init, I_init = pars['E_init'], pars['I_init']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize activity
E = np.zeros(Lt)
I = np.zeros(Lt)
E[0] = E_init
I[0] = I_init
I_ext_E = I_ext_E * np.ones(Lt)
I_ext_I = I_ext_I * np.ones(Lt) #ensure the external input an array
# simulate the Wilson-Cowan equations
for k in range(Lt-1):
######################################################################
## TODO for students: compute dE and dI, remove NotImplementedError #
######################################################################
# dE = ...
# dI = ...
raise NotImplementedError("Student excercise: compute the change in E/I")
E[k+1] = E[k] + dE
I[k+1] = I[k] + dI
return E, I
# Uncomment the below lines after completing the simulate_wc function
# Here are tow trjectories with close intial values
#pars = default_pars()
#pars['E_init'], pars['I_init'] = 0.32, 0.15
#E1,I1 = simulate_wc(pars)
#pars['E_init'], pars['I_init'] = 0.33, 0.15
#E2,I2 = simulate_wc(pars)
#my_test_plot(pars['range_t'], E1, I1, E2, I2)
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 397} colab_type="text" id="RkLzEO8EyuT9" outputId="dba7fce8-4c01-4770-d46e-26b1ceb12f40"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_88ca5a4b.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=560 height=380 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_88ca5a4b_0.png>
#
#
# + [markdown] colab_type="text" id="WXBTwdWbznem"
# ### Interactive Demo: population trajectories with different intial values
# In this interactive demo we will simulate the Wilson-Cowan model and plot the trajectories of each population for different initial conditions.
#
#
# **Remember to enable the demo by running the cell.**
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 401, "referenced_widgets": ["d63cf6caae6f4205be1f79e4703282f7", "122569ee0f8b42548ec7dbd7ede90c20", "9b998de360c8495490376ecb82cad7f3", "dfe575aad96a4fd3adc24e0f52cc02c7", "f529c05e88294648964cace55f091554", "37b9014d388c4d628585436413d8a2d4", "c88b507eb7c14c51a7c6b72fce2b87d5"]} colab_type="code" id="FcwiqGyNbuLD" outputId="e3ef1926-3bd3-427b-bb07-d15d3c98e688"
#@title System trajectories with different initial conditions
def plot_EI_diffinitial(E_init = 0.0):
pars = default_pars()
pars['E_init'], pars['I_init'] = E_init, 0.15
E, I = simulate_wc(pars)
plt.figure(figsize=(8, 5.5))
plt.plot(pars['range_t'], E, 'b', label='E population')
plt.plot(pars['range_t'], I, 'r', label='I population')
plt.xlabel('t (ms)', fontsize=14)
plt.ylabel('Activity', fontsize=14)
plt.legend(loc='best', fontsize=12)
plt.show()
_ = widgets.interact(plot_EI_diffinitial, E_init = (0.30, 0.35, .01))
# + [markdown] colab_type="text" id="UfOMo8Nkbx6z"
# Question: It is evident that the steady states of the neuronal response can be different when the initial states are chosen to be different. Why is that? If this phenomenon confuses you, we will give the answer right below.
# + [markdown] colab_type="text" id="b4yv1eyhby70"
# ## Phase plane analysis
#
# We will next introduce the phase plane analysis to understand the behavior of the E and I populations in the Wilson-Cowan model. So far, we have plotted the activities of the two populations as a function of time, i.e. in the `Activity-t` plane, either the $(t, E(t))$ plane or the and $(t, I(t))$ one. Instead, we can plot the two activities $E(t)$ and $I(t)$ against each other at any time point $t$. This characterization in the `I-E` plane $(I(t), E(t))$ is called the **phase plane**. Each line in the phase plane indicates how both $E$ and $I$ evolve with time.
# + [markdown] colab_type="text" id="YoxJ32W9tEHz"
# ### Interactive Demo: From the `Activity-t` plane to the `I-E` phase plane
#
# In this demo widget, we will visualize the system dynamics using both the `Activity-time` and the `(E, I)` phase plane. The circles indicate the activities at a given time $t$, while the lines represent the evolution of the system for the entire duration of the simulation.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 423, "referenced_widgets": ["0ec265da2a054acbbb4e1dc47c85424e", "fa53197403334efeb8cfc531ee7fc0b6", "5c6cbfe03aaa42ec81ae7a95ffe1f909", "ba01b9bb19454367a941cb5af608f69d", "cf2f4d9691cb4cadbaf7eb50bc63e415", "05628c37fc6947deb2a5bf1af39e5637", "a678271849d248218439cc6697d67bb8"]} colab_type="code" id="V3f1OK_vsvqM" outputId="0ebd3060-0969-48b8-8263-33801ab41ed7"
#@title `Activity-t` plane vs `I-E` phase plane
pars = default_pars(T=10)
pars['E_init'], pars['I_init'] = 0.6, 0.8
E,I = simulate_wc(pars)
def plot_activity_phase(n_t):
plt.figure(figsize=(8, 5.5))
plt.subplot(2,1,1)
plt.plot(pars['range_t'], E, 'b', label='E')
plt.plot(pars['range_t'], I, 'r', label='I')
plt.plot(pars['range_t'][n_t], E[n_t], 'bo')
plt.plot(pars['range_t'][n_t], I[n_t], 'ro')
plt.axvline(pars['range_t'][n_t], 0, 1, color='k', ls='--')
plt.xlabel('t (ms)', fontsize=14)
plt.ylabel('Activity', fontsize=14)
plt.legend(loc='best', fontsize=14)
plt.subplot(2,1,2)
plt.plot(E, I, 'k')
plt.plot(E[n_t], I[n_t], 'ko')
plt.xlabel('E', fontsize=18, color='b')
plt.ylabel('I', fontsize=18, color='r')
plt.tight_layout()
plt.show()
_ = widgets.interact(plot_activity_phase, n_t = (0, len(pars['range_t']-1), 1))
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 519} colab_type="code" id="O4Ci9Lmp7HvG" outputId="6f091ea0-2d59-4c68-abfc-9edf490abc09"
#@title Video: Nullclines and Vector Fields
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="BnwMK9dxCnk", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="KAoqPzL6ooIN"
# ### Nullclines of the Wilson-Cowan Equations
#
# An important concept in the phase plane analysis is the "nullcline", which is defined as the set of points in the phase plane where the activity of one population (but not necessarily the other) does not change.
#
# In other words, the $E$ and $I$ nullclines of Equation $(1)$ are defined as the points where $\displaystyle{\frac{dE}{dt}}=0$, for the excitatory nullcline, or $\displaystyle\frac{dI}{dt}=0$ for the inhibitory nullcline. That is:
#
# \begin{align}
# -E + F_E(w_{EE}E -w_{EI}I + I^{\text{ext}}_E;a_E,\theta_E) &= 0 \qquad (2)\\[1mm]
# -I + F_I(w_{IE}E -w_{II}I + I^{\text{ext}}_I;a_I,\theta_I) &= 0 \qquad (3)
# \end{align}
#
# Formally, Equations $2$ and $3$ can be written in the following form:
#
# \begin{align}
# I = \frac{1}{w_{EI}}\big{[}w_{EE}E - F_E^{-1}(E; a_E,\theta_E) + I^{\text{ext}}_E \big{]} \qquad (4)
# \end{align}
#
# Where $F_E^{-1}(E; a_E,\theta_E)$ is the inverse of the excitatory transfer function. Equation $4$ defines the $E$ nullcline.
#
#
# Similarly, the $I$ nullcline is found as:
# \begin{align}
# E = \frac{1}{w_{IE}} \big{[} w_{II}I + F_I^{-1}(I;a_I,\theta_I) - I^{\text{ext}}_I \big{]}. \qquad (5)
# \end{align}
#
# Where $F_I^{-1}(x; a_I,\theta_I)$ is the inverse of the inhibitory transfer function. Equation $5$ defines the $I$ nullcline.
#
# \\
# + [markdown] colab_type="text" id="uJTqbUyCQD1T"
# #### Exercise 3: Compute the nullclines of the Wilson-Cowan model
#
# In the next exercise, we will compute and plot the $E$ and the $I$ nullclines using Equations $4$ - $5$.
# Note that, when computing the nullclines with Equations $4$-$5$, we also need to calculate the inverse of the transfer functions.
# + colab={} colab_type="code" id="Msd1elDBbvpI"
# Exercise 3: Nullclines of Wilson-Cowan model
# Define the inverse of F
def F_inv(x,a,theta):
"""
Args:
x : the population input
a : the gain of the function
theta : the threshold of the function
Returns:
F_inverse : value of the inverse function
"""
#####################################################################
## TODO for students: compute F_inverse, remove NotImplementedError #
#####################################################################
# F_inverse = ...
raise NotImplementedError("Student excercise: compute the inverse of F(x)")
return F_inverse
# get the nullcline for E, solve Equation. (4) along the E-grid
def get_E_nullcline(pars, E_grid):
"""
Solve for I along the E_grid from dE/dt = 0.
Args:
pars : Parameter dictionary
E_grid : a single value or an array
Returns:
I : values of inhibitory population along the nullcline on the E-grid
"""
a_E, theta_E = pars['a_E'], pars['theta_E']
wEE, wEI = pars['wEE'], pars['wEI']
I_ext_E = pars['I_ext_E']
###########################################
## TODO for students: compute E nullcline #
###########################################
# I = ...
raise NotImplementedError("Student excercise: compute the E nullcline")
return I
# get the nullcline for I, solve Equation. (5) along the I-grid
def get_I_nullcline(pars, I_grid):
"""
Solve for E along the I_grid from dI/dt = 0.
Args:
pars : Parameter dictionary
I_grid : a single value or an array
Returns:
E : values of the excitatory population along the nullcline on the I-grid
"""
a_I, theta_I = pars['a_I'], pars['theta_I']
wIE, wII = pars['wIE'], pars['wII']
I_ext_I = pars['I_ext_I']
###########################################
## TODO for students: compute I nullcline #
###########################################
# E = ...
raise NotImplementedError("Student excercise: compute the I nullcline")
return E
# Uncomment the below lines after completing the all above functions
# pars = default_pars() # get parameters
# my_plot_nullcline(pars)
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="text" id="5_nTmGL51k6B" outputId="8902ada6-8267-474e-8c26-6617f4c122b6"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_a1723906.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=513 height=357 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_a1723906_0.png>
#
#
# + [markdown] colab_type="text" id="AV8XDjhMb-jk"
# ### Vector field
#
# How can the phase plane and the nullcline curves help us understand the behavior of the Wilson-Cowan model?
#
# The activities of the $E$ and $I$ populations $E(t)$ and $I(t)$ at each time point $t$ correspond to a single point in the phase plane, with coordinates $(E(t),I(t))$. Therefore, the time-dependent trajectory of the system can be described as a continuous curve in the phase plane, and the tangent vector to the trajectory, which is defined as the vector $\bigg{(}\displaystyle{\frac{dE(t)}{dt},\frac{dI(t)}{dt}}\bigg{)}$, indicates the direction towards which the activity is evolving and how fast is the activity changing along each axis. In fact, for each point $(E,I)$ in the phase plane, we can compute the tangent vector $\bigg{(}\displaystyle{\frac{dE}{dt},\frac{dI}{dt}}\bigg{)}$, which will indicate the behavior of the system when it traverses that point. The map of tangent vectors in the phase plane is called **vector field**. The behavior of any trajectory in the phase plane is determined by i) the initial conditions $(E(0),I(0))$, and ii) the vector field $(\displaystyle{\frac{dE(t)}{dt},\frac{dI(t)}{dt}})$.
#
# In general, the value of the vector field at a particular point in the phase plane is represented by an arrow. The orientation and the size of the arrow reflect the direction and the norm of the vector, respectively.
# + [markdown] colab_type="text" id="4r-JYJ9USlRY"
# #### Exercise 4: Compute and plot the vector field $\displaystyle{\Big{(}\frac{dE}{dt}, \frac{dI}{dt} \Big{)}}$
# + colab={} colab_type="code" id="cq2654EzTzZ2"
# Exercise 4
# Define the value of the derivatives according to Equation. (1)
def EIderivs(E_grid, I_grid, pars):
"""
Time derivatives for E/I variables (dE/dt, dI/dt).
"""
tau_E, a_E, theta_E = pars['tau_E'], pars['a_E'], pars['theta_E']
tau_I, a_I, theta_I = pars['tau_I'], pars['a_I'], pars['theta_I']
wEE, wEI = pars['wEE'], pars['wEI']
wIE, wII = pars['wIE'], pars['wII']
I_ext_E, I_ext_I = pars['I_ext_E'], pars['I_ext_I']
##########################################################################
## TODO for students: compute dEdt and dI/dt, remove NotImplementedError #
##########################################################################
# dEdt = ...
# dIdt = ...
raise NotImplementedError("Student excercise: compute the vector field")
return dEdt, dIdt
# Uncomment these lines after completing the EIderivs function
# pars = default_pars()
# my_plot_trajectories(pars, 0.2, 6, 'Sample trajectories \nof different initials')
# my_plot_trajectory(pars, 'orange', [0.6, 0.8], 'Sample trajectory to \nlow activity')
# my_plot_trajectory(pars, 'm', [0.6, 0.6], 'Sample trajectory to \nhigh activity')
# my_plot_vector(pars)
# my_plot_nullcline(pars)
# plt.legend(loc=[1.02, 0.6], fontsize=12, handlelength=1)
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 377} colab_type="text" id="4bJm2sqg3KMd" outputId="901b2ab3-4c6b-4b7a-9263-526eceff6d26"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_a20da002.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=617 height=387 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_a20da002_0.png>
#
#
# + [markdown] colab_type="text" id="ZM_pPKMScDh5"
# ### Think!
#
# The last phase plane plot showed us that:
# - trajectories seem to follow the direction of the vector field
# - different trajectories eventually always reach one of two points depending on the initial conditions. Note that the two points are the intersection of the two nullcline curves.
#
# There are, in total, three intersection points, but one of them is never the final state of a trajectory. Why is that?
# + [markdown] colab_type="text" id="bx7aiRLtpFAw"
# ## Summary
#
# Congratulations! You have finished the second day of the last week of neuromatch academy! Here, you learned how to simulate a rate based model consisting of excitatory and inhibitory population of neurons.
#
# In the last tutorial on dynamical neuronal networks you learned to:
# - Implement and simulate a 2D system composed of an E and an I population of neurons using the **Wilson-Cowan** model
# - Plot the frequency-current (F-I) curves for both populations
# - Examine the behavior of the system using phase **plane analysis**, **vector fields**, and **nullclines**.
#
# Do you have more time? Have you finished early? We have more fun material for you!
#
# Below are some, more advanced concepts on dynamical systems:
#
# - You will learn how to find the fixed points on such a system, and to investigate its stability by linearizing its dynamics and examining the **Jacobian matrix**.
# - You will see how the Wilson-Cowan model can reach an oscillatory state.
#
# If you need even more, there are two applications of the Wilson-Cowan model:
#
# - Visualization of an Inhibition-stabilized network
# - Simulation of working memory
# + [markdown] colab_type="text" id="8v2qsjq2OSOo"
# ## Bonus 1: Fixed points, stability analysis, and limit cycles in the Wilson-Cowan model
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 519} colab_type="code" id="XbeIet5NOPjo" outputId="062d67c6-ecd8-4ecb-cec5-2b67f7320e74"
#@title Video: Fixed points and their stability
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="RgysOunhhwM", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="kueE2dLycGbA"
# ### Fixed Point of the E/I system
#
# Clearly, the intersection points of the two nullcline curves are the fixed points of the Wilson-Cowan model in Equation $(1)$.
#
# In the next exercise, we will find the coordinate of all fixed points for a given set of parameters.
#
# Let's start by inspecting and then executing the cell below.
# + cellView="both" colab={} colab_type="code" id="Dni0QzyfcB66"
def my_fp(pars, E_init, I_init):
"""
use opt.root function to solve Equations. (4)-(5) from an initial [E_init, I_init]
"""
tau_E, a_E, theta_E = pars['tau_E'], pars['a_E'], pars['theta_E']
tau_I, a_I, theta_I = pars['tau_I'], pars['a_I'], pars['theta_I']
wEE, wEI = pars['wEE'], pars['wEI']
wIE, wII = pars['wIE'], pars['wII']
I_ext_E, I_ext_I = pars['I_ext_E'], pars['I_ext_I']
# define the right hand of wilson-cowan equations
def my_WCr(x):
E = x[0]
I = x[1]
dEdt=(-E + F(wEE*E-wEI*I+I_ext_E,a_E,theta_E))/tau_E
dIdt=(-I + F(wIE*E-wII*I+I_ext_I,a_I,theta_I))/tau_I
y = np.array([dEdt, dIdt])
return y
x0 = np.array([E_init, I_init])
x_fp = opt.root(my_WCr, x0).x
return x_fp
# + [markdown] colab_type="text" id="6grnXKjq4pQf"
# #### Exercise 5: Find the fixed points of the Wilson-Cowan model
#
# From the above nullclines, we notice that the system features three fixed points with the parameters we used. To find their coordinates, we need to choose the proper initial value to give to the `opt.root` function inside of the function `my_fp` we just defined, since the algorithm can only find fixed points in the vicinity of the initial value.
#
# In this exercise, you will use the function `my_fp` to find each of the fixed points by varying the initial values.
# + colab={"base_uri": "https://localhost:8080/", "height": 370} colab_type="code" id="TXka86oM4plz" outputId="761f0a92-97d8-4cd3-f537-1be57045625f"
# Exercise 5
pars = default_pars()
plt.figure(figsize=(8, 5.5))
my_plot_nullcline(pars)
#####################################################
## TODO for students: #
# Calculate the fixed point with your initial value #
# verify your fixed point and plot the corret ones #
#####################################################
#x_fp = my_fp(pars, )
#check if x_fp is the intersection of the lines with the give function check_fp(x_fp)
#vary different initial values to find the correct fixed point
#if check_fp(x_fp):
# plot_fp(x_fp) #you can plot fixedpoit directly by plt.plot(...)
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="text" id="Unwet1TfcM_h" outputId="70569cd0-99c3-461d-b30a-8e1fe68841d2"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_954c437d.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=513 height=357 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_954c437d_0.png>
#
#
# + [markdown] colab_type="text" id="qastrWoycQjn"
# ### Stability of a fixed point and eigenvalues of the Jacobian Matrix
#
# First, let's first rewrite the system $1$ as:
#
# \begin{align}
# &\frac{dE}{dt} = G_E(E,I)\\[0.5mm]
# &\frac{dI}{dt} = G_I(E,I) \qquad (11)
# \end{align}
# where
#
# \begin{align}
# &G_E(E,I) = \frac{1}{\tau_E} [-E + F_E(w_{EE}E -w_{EI}I + I^{\text{ext}}_E;a,\theta)]\\[1mm]
# &G_I(E,I) = \frac{1}{\tau_I} [-I + F_I(w_{IE}E -w_{II}I + I^{\text{ext}}_I;a,\theta)]
# \end{align}
#
# By definition, $\displaystyle\frac{dE}{dt}=0$ and $\displaystyle\frac{dI}{dt}=0$ at each fixed point. Therefore, if the initial state is exactly at the fixed point, the state of the system will not change as time evolves. However, if the initial state deviates slightly from the fixed point, two possibilities will happen (1) the trajectory will be attracted back to the fixed point (2) the trajectory will diverge from the fixed point. These two possibilities define the type of fixed point, i.e., stable or unstable. Similar to the 1d system studied in the previous tutorial, the stability of a fixed point $(E, I)$ can be determined by linearizing the dynamics of the system (can you figure out how?). The linearization will yield a matrix of first-order derivatives called the Jacobian matrix:
#
# \begin{equation}
# J=
# \left[ {\begin{array}{cc}
# \displaystyle{\frac{\partial G_E}{\partial E}} & \displaystyle{\frac{\partial G_E}{\partial I}}\\[1mm]
# \displaystyle\frac{\partial G_I}{\partial E} & \displaystyle\frac{\partial G_I}{\partial I} \\
# \end{array} } \right]. \quad (12)
# \end{equation}
#
# The eigenvalues of the Jacobian matrix calculated at the fixed point will determine whether it is a stable or unstable fixed point.
#
# \\
#
# We can now compute the derivatives needed to build the Jacobian matrix. Using the chain and product rules, the derivatives for the excitatory population are given by:
# \begin{align}
# &\frac{\partial G_E}{\partial E} = \frac{1}{\tau_E} [-1 + w_{EE} F_E'(w_{EE}E -w_{EI}I + I^{\text{ext}}_E)] \\[1mm]
# &\frac{\partial G_E}{\partial I} = \frac{1}{\tau_E} [-w_{EI} F_I'(w_{EE}E -w_{EI}I + I^{\text{ext}}_E)]
# \end{align}
#
# The same applies to the inhibitory population.
# + [markdown] colab_type="text" id="3uy5zBscbI0c"
# #### Exercise 6: Compute the Jacobian Matrix for the Wilson-Cowan model
# + colab={} colab_type="code" id="HbzRXGAwcUet"
# Exercise 6
def get_eig_Jacobian(pars, fp):
"""
Simulate the Wilson-Cowan equations
Args:
pars : Parameter dictionary
fp : fixed point (E, I), array
Returns:
evals : 2x1 vector of eigenvalues of the Jacobian matrix
"""
#get the parameters
tau_E, a_E, theta_E = pars['tau_E'], pars['a_E'], pars['theta_E']
tau_I, a_I, theta_I = pars['tau_I'], pars['a_I'], pars['theta_I']
wEE, wEI = pars['wEE'], pars['wEI']
wIE, wII = pars['wIE'], pars['wII']
I_ext_E, I_ext_I = pars['I_ext_E'], pars['I_ext_I']
#initialization
E = fp[0]
I = fp[1]
J = np.zeros((2,2))
######################################################################
## TODO for students: compute J, then remove the NotImplementedError #
######################################################################
# J[i, j] = ...
raise NotImplementedError("Student excercise: compute the Jacobian matrix")
# Eigenvalues
evals = np.linalg.eig(J)[0]
return evals
# Uncomment these lines after completing the get_eig_Jacobian function
# only when you get the correct fixed point above you print their eigenvalues below
#eig_1 = get_eig_Jacobian(pars, x_fp_1)
#eig_2 = get_eig_Jacobian(pars, x_fp_2)
#eig_3 = get_eig_Jacobian(pars, x_fp_3)
#print(eig_1, 'Stable point')
#print(eig_2, 'Unstable point')
#print(eig_3, 'Stable point')
# + [markdown] cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="text" id="F9uswAXA5FRo" outputId="9a748b91-49a7-47ad-d59f-8fc746cbc75b"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_0349af2c.py)
#
#
# + [markdown] colab_type="text" id="CvpweBaIcaew"
# As is evident, the stable point corresponds to the negative eigenvalues, while unstable point corresponds to at least one positive eigenvalue.
# + [markdown] colab_type="text" id="tn7kTA9ZcWTE"
# Below we investigate the effect of $w_{EE}$ on the nullclines and the eigenvalues of the dynamical system. <font color='black'> _Critical change is referred as **pitchfork bifurcation**._</font>
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 371} colab_type="code" id="GPXlFbGU3Ydc" outputId="013ab82d-e8f0-4b26-c0f0-d5a421ca3c77"
#@title Effect of `wEE` on the nullclines and the eigenvalues
eig_1_M = []
eig_2_M = []
eig_3_M = []
pars = default_pars()
wEE_grid = np.linspace(6,10,40)
my_thre = 7.9
for wEE in wEE_grid:
x_fp_1 = [0., 0.]
x_fp_2 = [.4, .1]
x_fp_3 = [.8, .1]
pars['wEE'] = wEE
if wEE < my_thre:
x_fp_1 = my_fp(pars, x_fp_1[0], x_fp_1[1])
eig_1 = get_eig_Jacobian(pars, x_fp_1)
eig_1_M.append(np.max(np.real(eig_1)))
else:
x_fp_1 = my_fp(pars, x_fp_1[0], x_fp_1[1])
eig_1 = get_eig_Jacobian(pars, x_fp_1)
eig_1_M.append(np.max(np.real(eig_1)))
x_fp_2 = my_fp(pars, x_fp_2[0], x_fp_2[1])
eig_2 = get_eig_Jacobian(pars, x_fp_2)
eig_2_M.append(np.max(np.real(eig_2)))
x_fp_3 = my_fp(pars, x_fp_3[0], x_fp_3[1])
eig_3 = get_eig_Jacobian(pars, x_fp_3)
eig_3_M.append(np.max(np.real(eig_3)))
eig_1_M = np.array(eig_1_M)
eig_2_M = np.array(eig_2_M)
eig_3_M = np.array(eig_3_M)
plt.figure(figsize=(8, 5.5))
plt.plot(wEE_grid, eig_1_M, 'ko', alpha=0.5)
plt.plot(wEE_grid[wEE_grid>=my_thre], eig_2_M, 'bo', alpha=0.5)
plt.plot(wEE_grid[wEE_grid>=my_thre], eig_3_M, 'ro', alpha=0.5)
plt.xlabel(r'$w_{\mathrm{EE}}$')
plt.ylabel('maximum real part of eigenvalue')
plt.show()
# + [markdown] colab_type="text" id="ZgK-hVuCcM89"
# #### Interactive Demo: Nullclines position in the phase plane changes with parameter values.
#
# In this interactive widget, we will explore how the nullclines move for different values of the parameter $w_{EE}$.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 423, "referenced_widgets": ["54a4af4754754e5681e26006de5a6e56", "d6c78211049447b987efa0c6dcab3083", "7661f3c67e014e79b44a1462bfa0a69a", "08e33a5f30e34932b7824759e2e779d6", "d42075db409241af8f97d00c80b9e73f", "d852e82d2cfe46ae8733c7f6ea6eca9d", "7031b4e5a1244c80b0a900ab8e0fe208"]} colab_type="code" id="hm4tdQXcclC2" outputId="624f1060-38de-4282-dbc9-ad13c37cccd4"
#@title Nullcline Explorer
def plot_nullcline_diffwEE(wEE):
'''
plot nullclines for different values of wEE
'''
pars = default_pars()
pars['wEE'] = wEE
# plot the E, I nullclines
E_grid = np.linspace(-0.01,.96,100)
E_nullcline = get_E_nullcline(pars, E_grid)
I_grid = np.linspace(-.01,.8,100)
I_nullcline = get_I_nullcline(pars, I_grid)
plt.figure(figsize=(12, 5.5))
plt.subplot(1, 2, 1)
plt.plot(E_grid, E_nullcline, 'r', label='E nullcline')
plt.plot(I_nullcline, I_grid, 'b', label='I nullcline')
#plt.xlim(0.6, 1.0)
#plt.ylim(0.3, 0.6)
plt.xlabel('E')
plt.ylabel('I')
plt.legend(loc='best')
plt.subplot(2, 2, 2)
pars['E_init'], pars['I_init'] = 0.2, 0.2
E, I = simulate_wc(pars)
plt.plot(pars['range_t'], E, 'r', label='E population', clip_on=False)
plt.plot(pars['range_t'], I, 'b', label='I population', clip_on=False)
plt.xlabel('t (ms)')
plt.ylabel('Activity')
plt.legend(loc='best')
plt.ylim(-0.05, 1.05)
plt.title('E/I activity', fontsize=10, fontweight='bold')
plt.subplot(2, 2, 4)
pars['E_init'], pars['I_init'] = 0.4, 0.1
E, I = simulate_wc(pars)
plt.plot(pars['range_t'], E, 'r', label='E population', clip_on=False)
plt.plot(pars['range_t'], I, 'b', label='I population', clip_on=False)
plt.xlabel('t (ms)')
plt.ylabel('Activity')
plt.legend(loc='best')
plt.ylim(-0.05, 1.05)
plt.tight_layout()
plt.show()
_ = widgets.interact(plot_nullcline_diffwEE, wEE = (6., 10., .01))
# + [markdown] colab_type="text" id="n8ZRNtt2couO"
# ##### Task: effect of other parameters
# We can also investigate the effect of different $w_{EI}$, $w_{IE}$, $w_{II}$, $\tau_{E}$, $\tau_{I}$, and $I_{E}^{\text{ext}}$ on the stability of fixed points. In addition, we can also consider the perturbation on the parameters of the gain curve $F(\cdot)$
# + [markdown] colab_type="text" id="d913ob7Xcqw3"
# ## Limit cycle
#
# If we use a different set of parameters, $w_{EE}=6.4$, $w_{EI}=4.8$, $w_{IE}=6.$, $w_{II}=1.2$, and $I_{E}^{\text{ext}}=0.8$, then we shall observe that the E and I population activity start to oscillate! Please execute the cell below to check the oscillatory behavior.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 370} colab_type="code" id="DCw-lgqxcm0Q" outputId="a5051e4c-74ee-479c-c89d-b86fc25286a3"
#@title Oscillations
pars = default_pars(T=100.)
pars['wEE'], pars['wEI'] = 6.4, 4.8
pars['wIE'], pars['wII'] = 6.0, 1.2
pars['I_ext_E'] = 0.8
pars['E_init'], pars['I_init'] = 0.25, 0.25
E,I = simulate_wc(pars)
plt.figure(figsize=(8, 5.5))
plt.plot(pars['range_t'], E, 'r')
plt.plot(pars['range_t'], I, 'b')
plt.xlabel('t (ms)')
plt.ylabel('E(t), I(t)')
plt.show()
# + [markdown] colab_type="text" id="tX0xMzVscy5a"
# ##### Exercise 7: Plot the phase plane
#
# We can also understand the oscillations of the population behavior using the phase plane. By plotting a set of trajectories with different initial states, we can see that these trajectories will move in a circle instead of converging to a fixed point. This circle is called "limit cycle", and shows the periodic oscillations of the $E$ and $I$ population behavior under some conditions.
#
# Try to plot the phase plane using the previously defined functions.
# + colab={} colab_type="code" id="lPJTsupA08CA"
pars = default_pars(T=100.)
pars['wEE'], pars['wEI'] = 6.4, 4.8
pars['wIE'], pars['wII'] = 6.0, 1.2
pars['I_ext_E'] = 0.8
###############################################################################
## TODO for students: plot phase plane: nullclines, trajectories, fixed point #
###############################################################################
## please make sure you find the corret fixed point
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="text" id="0a_nAz8OdStx" outputId="f8198fb0-499a-433a-d08c-f8ec22e0915a"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_fedbbbea.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=513 height=357 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D2_DynamicNetworks/static/W3D2_Tutorial2_Solution_fedbbbea_1.png>
#
#
# + [markdown] colab_type="text" id="5IPxPCWfcwQa"
# #### Interactive Demo: Limit cycle and oscillations.
#
# From the above examples, the change of model parameters changes the shape of the nullclines and accordingly, the behavior of the $E$ and $I$ populations from steady fixed points to oscillations. However, the shape of the nullclines is unable to fully determine the behavior of the network. The vector field also matters. To demonstrate this, here we will investigate the effect of time constants on the population behavior. By changing the inhibitory time constant $\tau_I$, the nullclines do not change, but the network behavior changes substantially from steady state to oscillations with different frequencies.
#
# Such a dramatic change in the system behavior is referred to as a **bifurcation**.
#
# \\
# Please execute the code below to check this out.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 446, "referenced_widgets": ["dcc7c0a2811c4fc3a80a8e9ae959bbc2", "6f16291ab2884ec3871c83147727998e", "5753eacc6711433b98b6b6675df96991", "a467da6202354753a878328a9f3be074", "b3afb82870b04d80a35ed5e8d96346dd", "670a4be1d38d451bb00d684c1db1e22b", "31cdf6f3c2144e439cd2d3beadc07eec"]} colab_type="code" id="L2QzZmryc70S" outputId="d70cc40c-9180-41ec-89e3-a2572bdf4529"
#@title Limit Cycle Explorer
def time_constant_effect(tau_s=0.5):
pars = default_pars(T=100.)
pars['wEE'], pars['wEI'] = 6.4, 4.8
pars['wIE'], pars['wII'] = 6.0, 1.2
pars['I_ext_E'] = 0.8
pars['tau_I'] = tau_s
E_grid = np.linspace(0.0,.9,100)
I_grid = np.linspace(0.0,.6,100)
E_nullcline = get_E_nullcline(pars, E_grid)
I_nullcline = get_I_nullcline(pars, I_grid)
with plt.xkcd():
plt.figure(figsize=(12.5, 5.5))
plt.subplot(1,2,1) #nullclines
plt.plot(E_grid, E_nullcline, 'r', label='E nullcline')
plt.plot(I_nullcline, I_grid, 'b', label='I nullcline')
plt.xlabel('E')
plt.ylabel('I')
#fixed point
x_fp_1 = my_fp(pars, 0.5, 0.5)
plt.plot(x_fp_1[0], x_fp_1[1], 'ko')
eig_1 = get_eig_Jacobian(pars, x_fp_1)
print('tau_I=%.1f ms,' %tau_s, 'eigenvalue of J matrix ', eig_1)
#trajectories
E_tj = np.zeros((pars['range_t'].size, 5, 5))
I_tj = np.zeros((pars['range_t'].size, 5, 5))
for ie in range(5):
for ii in range(5):
pars['E_init'], pars['I_init'] = 0.1*ie, 0.1*ii
E_tj[:, ie, ii], I_tj[:, ie, ii] = simulate_wc(pars)
plt.plot(E_tj[:, ie, ii], I_tj[:, ie, ii],'k',alpha=0.3)
#vector field
EI_grid_E = np.linspace(0., 1.0, 20)
EI_grid_I = np.linspace(0., 0.6, 20)
E_meshgrid, I_meshgrid = np.meshgrid(EI_grid_E, EI_grid_I)
dEdt, dIdt = EIderivs(E_meshgrid, I_meshgrid, pars)
n_skip = 2
plt.quiver(E_meshgrid[::n_skip,::n_skip], I_meshgrid[::n_skip,::n_skip],
dEdt[::n_skip,::n_skip], dIdt[::n_skip,::n_skip],
angles='xy', scale_units='xy', scale=10,facecolor='c')
plt.title(r'$\tau_I=$'+'%.1f ms' % tau_s)
plt.subplot(1,2,2) # sample E/I trajectories
pars['E_init'], pars['I_init'] = 0.25, 0.25
E,I = simulate_wc(pars)
plt.plot(pars['range_t'], E, 'r')
plt.plot(pars['range_t'], I, 'b')
plt.xlabel('t (ms)')
plt.ylabel('E(t), I(t)')
plt.title(r'$\tau_I=$'+'%.1f ms' % tau_s)
plt.tight_layout()
plt.show()
_ = widgets.interact(time_constant_effect,tau_s = (0.1, 3, .1))
# + [markdown] colab_type="text" id="F6Q2LcERqXI7"
# ## Bonus 2: Inhibition-stabilized network (ISN)
#
# As described above, one can obtain the linear approximation around the fixed point as
#
# \begin{equation}
# \frac{d}{dr} \vec{X}=
# \left[ {\begin{array}{cc}
# \displaystyle{\frac{\partial G_E}{\partial E}} & \displaystyle{\frac{\partial G_E}{\partial I}}\\[1mm]
# \displaystyle\frac{\partial G_I}{\partial E} & \displaystyle\frac{\partial G_I}{\partial I} \\
# \end{array} } \right] \vec{X},
# \end{equation}
# where $\vec{X} = [E, I]^{\rm T}$ is the vector of the E/I activity.
#
# Let's direct our attention to the excitatory subpopulation which follows:
#
# \begin{equation}
# \frac{dE}{dt} = \frac{\partial G_E}{\partial E}\cdot E + \frac{\partial G_E}{\partial I} \cdot I
# \end{equation}
#
# Recall that:
# \begin{align}
# &\frac{\partial G_E}{\partial E} = \frac{1}{\tau_E} [-1 + w_{EE} F'(w_{EE}E -w_{EI}I + I^{\text{ext}}_E)] \qquad (13)\\[1mm]
# &\frac{\partial G_E}{\partial I} = \frac{1}{\tau_E} [-w_{EI} F'(w_{EE}E -w_{EI}I + I^{\text{ext}}_E)] \qquad (14)
# \end{align} \\
#
#
# From Equation. (8), it is clear that $\displaystyle{\frac{\partial G_E}{\partial I}}$ is negative, since the $\displaystyle{\frac{dF}{dx}}$ is always positive. It can be understood by that the recurrent inhibition from the inhibitory activity I can reduce the E activity. However, as described above, $\displaystyle{\frac{\partial G_E}{\partial E}}$ has negative terms related to the "leak" effect, and positive term related to the recurrent excitation. Therefore, it leads to two different regimes:
#
# - $\displaystyle{\frac{\partial G_E}{\partial E}}<0$, **noninhibition-stabilized
# network (non-ISN) regime**
#
# - $\displaystyle{\frac{\partial G_E}{\partial E}}>0$, **inhibition-stabilized
# network (ISN) regime**
#
#
# + [markdown] colab_type="text" id="m1XpwD0k7U6S"
# #### Exercise 8: Compute $\displaystyle{\frac{\partial G_E}{\partial E}}$
# Implemet the function to calculate the $\displaystyle{\frac{\partial G_E}{\partial E}}$ for the default parameters, and the parameters of the limit cycle case.
# + colab={} colab_type="code" id="dzbXN5m43Aw_"
# Exercise 8
def get_dGdE(pars, fp):
"""
Simulate the Wilson-Cowan equations
Args:
pars : Parameter dictionary
fp : fixed point (E, I), array
Returns:
J : the 2x2 Jacobian matrix
"""
# get the parameters
tau_E, a_E, theta_E = pars['tau_E'], pars['a_E'], pars['theta_E']
wEE, wEI = pars['wEE'], pars['wEI']
I_ext_E = pars['I_ext_E']
# initialization
E = fp[0]
I = fp[1]
####################################################################
## TODO for students: compute dGdE, remove the NotImplementedError #
####################################################################
#dGdE =...
raise NotImplementedError("Student excercise: compute the dG/dE, Equation. (13)")
return dGdE
# Uncomment these lines to make the output once you've completed the function
# pars = default_pars()
# x_fp_1 = my_fp(pars, 0.1, 0.1)
# x_fp_2 = my_fp(pars, 0.3, 0.3)
# x_fp_3 = my_fp(pars, 0.8, 0.6)
# dGdE1 = get_dGdE(pars, x_fp_1)
# dGdE2 = get_dGdE(pars, x_fp_2)
# dGdE3 = get_dGdE(pars, x_fp_3)
# print ('For the default case:')
# print ('dG/dE(fp1) = %.3f' %(dGdE1))
# print ('dG/dE(fp2) = %.3f' %(dGdE2))
# print ('dG/dE(fp3) = %.3f' %(dGdE3))
# print ('\n')
# pars = default_pars()
# pars['wEE'], pars['wEI'] = 6.4, 4.8
# pars['wIE'], pars['wII'] = 6.0, 1.2
# pars['I_ext_E'] = 0.8
# x_fp_lc = my_fp(pars, 0.8, 0.8)
# dGdE_lc = get_dGdE(pars, x_fp_lc)
# print ('For the limit cycle case:')
# print ('dG/dE(fp_lc) = %.3f' %(dGdE_lc))
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 156} colab_type="text" id="ZJzQuEDJKENe" outputId="a5c0f0ea-9430-4746-8a5d-452394ca54b5"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D2_DynamicNetworks/solutions/W3D2_Tutorial2_Solution_74c52d9b.py)
#
#
# + [markdown] colab_type="text" id="7i9ZTBMp73Nd"
# ### Interactive Demo: Paradoxical effect in ISN
#
# In this interactive widget, we inject excitatory ($I^{\text{ext}}_I>0$) or inhibitory ($I^{\text{ext}}_I<0$) drive into the inhibitory population when the system is at its equilibrium (with parameters $w_{EE}=6.4$, $w_{EI}=4.8$, $w_{IE}=6.$, $w_{II}=1.2$, $I_{E}^{\text{ext}}=0.8$, $\tau_I = 0.8$, and $I^{\text{ext}}_I=0$). Then check how the firing rate of the $I$ population changes.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 488, "referenced_widgets": ["9a3db3a11b79470e831e57592d7d65aa", "83942da47c4b4d808b6e5c0a4c2832c5", "281c73b3f48c4f97a8724fee238e4cf9", "b9e2e1f2d7d44049897b52fff4b6650d", "4e24dace7c784fa6b9e2520861266eed", "f46a7f32ea80435bbd26b80b33373074", "296bd299b2b94f218c5ee227a171f4f4"]} colab_type="code" id="pv5S7g0N7A1s" outputId="369f6c5e-3098-46cd-cb76-9e16807a28c2"
#@title ISN Explorer
pars = default_pars(T=50., dt=0.1)
pars['wEE'], pars['wEI'] = 6.4, 4.8
pars['wIE'], pars['wII'] = 6.0, 1.2
pars['I_ext_E'] = 0.8
pars['tau_I'] = 0.8
def ISN_I_perturb(dI=0.1):
Lt = len(pars['range_t'])
pars['I_ext_I'] = np.zeros(Lt)
pars['I_ext_I'][int(Lt/2):] = dI
pars['E_init'], pars['I_init'] = 0.6, 0.26
E,I = simulate_wc(pars)
plt.figure(figsize=(8, 1.5))
plt.plot(pars['range_t'], pars['I_ext_I'], 'k')
plt.xlabel('t (ms)')
plt.ylabel(r'$I_I^{\mathrm{ext}}$')
plt.ylim(pars['I_ext_I'].min()-0.01, pars['I_ext_I'].max()+0.01)
plt.show()
plt.figure(figsize=(8, 4.5))
plt.plot(pars['range_t'], E, 'r')
plt.plot(pars['range_t'], E[int(Lt/2)-1]*np.ones(Lt), 'r--')
plt.plot(pars['range_t'], I, 'b')
plt.plot(pars['range_t'], I[int(Lt/2)-1]*np.ones(Lt), 'b--')
plt.ylim(0, 0.8)
plt.xlabel('t (ms)')
plt.ylabel('E(t), I(t)')
plt.show()
_ = widgets.interact(ISN_I_perturb, dI = (-0.2, 0.21, .05))
# + [markdown] colab_type="text" id="I1KCkJWhABCd"
# ## Bonus 3: Fixed point and working memory
# + [markdown] colab_type="text" id="O7EPxbBPD9iA"
# The input into the neurons measured in the experiment is often very noisy ([links](http://www.scholarpedia.org/article/Stochastic_dynamical_systems)) . Here, the noisy synaptic input current is modeled as an Ornstein-Uhlenbeck (OU)process, which has been discussed several times in the previous tutorials.
#
# please execute the following cell of function `my_OU(pars, sig, myseed=False)`
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 370} colab_type="code" id="keZx8QBxAuBz" outputId="1cbe668a-2871-4866-c4b4-be5508db08c5"
#@title `my_OU(pars, sig, myseed=False)`
def my_OU(pars, sig, myseed=False):
'''
Expects:
pars : parameter dictionary
sig : noise amplitute
myseed : random seed. int or boolean
Returns:
I : Ornstein-Uhlenbeck input current
'''
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tau_ou = pars['tau_ou'] # [ms]
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# Initialize
noise = np.random.randn(Lt)
I = np.zeros(Lt)
I[0] = noise[0] * sig
#generate OU
for it in range(Lt-1):
I[it+1] = I[it] + dt/tau_ou*(0.-I[it]) + np.sqrt(2.*dt/tau_ou) * sig * noise[it+1]
return I
pars = default_pars(T=50)
pars['tau_ou'] = 1. #[ms]
sig_ou = 0.1
I_ou = my_OU(pars, sig=sig_ou, myseed=1998)
plt.figure(figsize=(8, 5.5))
plt.plot(pars['range_t'], I_ou, 'b')
plt.xlabel('Time (ms)')
plt.ylabel(r'$I_{\mathrm{OU}}$')
plt.show()
# + [markdown] colab_type="text" id="DQmkMAjfGMDL"
# With the default parameters, the system fluctuates around a resting state with the noisy input.
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 370} colab_type="code" id="aa3l3pgkAGKG" outputId="36bb97be-7b06-46c7-a413-50f383d30ed9"
#@title WC with OU
pars = default_pars(T=100)
pars['tau_ou'] = 1. #[ms]
sig_ou = 0.1
pars['I_ext_E'] = my_OU(pars, sig=sig_ou, myseed=20201)
pars['I_ext_I'] = my_OU(pars, sig=sig_ou, myseed=20202)
pars['E_init'], pars['I_init'] = 0.1, 0.1
E,I = simulate_wc(pars)
plt.figure(figsize=(8, 5.5))
ax = plt.subplot(111)
ax.plot(pars['range_t'], E, 'r', label='E population')
ax.plot(pars['range_t'], I, 'b', label='I population')
ax.set_xlabel('t (ms)')
ax.set_ylabel('Activity')
ax.legend(loc='best')
plt.show()
# + [markdown] colab_type="text" id="TbxX1ha6GpTW"
# ### Short pulse induced persistent activity
# Then, let's use a brief 10-ms positive current to the E population when the system is at its equilibrium. When this amplitude is sufficiently large, a persistent activity is produced that outlasts the transient input. What is the firing rate of the persistent activity, and what is the critical input strength? Try to understand the phenomena from the above phase-plane analysis.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 407, "referenced_widgets": ["59ee5c4c5353418db60f903da05038b6", "4a64d3b07fd04d139362a62f18b40a7e", "27bd5b2c0b62443e937a6e195fa94cab", "0eec5812c6cf4498bea39b4fc5bf6eff", "f57ee9b002b04bfa8201f103b9ea7d58", "b12d8700b098449c8a7ce163048d2f4e", "5eeab31ded6d445db9f971362c5c44d1"]} colab_type="code" id="Z1O10EF5InFG" outputId="87e8b76c-93a8-4633-b8f4-415b35565db7"
#@title Pulse Explorer
def my_inject(pars, t_start, t_lag=10.):
'''
Expects:
pars : parameter dictionary
t_start : pulse starts [ms]
t_lag : pulse lasts [ms]
Returns:
I : extra pulse time
'''
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize
I = np.zeros(Lt)
#pulse timing
N_start = int(t_start/dt)
N_lag = int(t_lag/dt)
I[N_start:N_start+N_lag] = 1.
return I
pars = default_pars(T=100)
pars['tau_ou'] = 1. #[ms]
sig_ou = 0.1
pars['I_ext_I'] = my_OU(pars, sig=sig_ou, myseed=20202)
pars['E_init'], pars['I_init'] = 0.1, 0.1
#pulse
I_pulse = my_inject(pars, t_start=20., t_lag=10.)
L_pulse = sum(I_pulse>0.)
def WC_with_pulse(SE=0.):
pars['I_ext_E'] = my_OU(pars, sig=sig_ou, myseed=20201)
pars['I_ext_E'] += SE*I_pulse
E,I = simulate_wc(pars)
plt.figure(figsize=(8, 5.5))
ax = plt.subplot(111)
ax.plot(pars['range_t'], E, 'r', label='E population')
ax.plot(pars['range_t'], I, 'b', label='I population')
ax.plot(pars['range_t'][I_pulse>0.], 1.0*np.ones(L_pulse), 'r', lw=2.)
ax.text(25, 1.05, 'stimulus on', horizontalalignment='center',
verticalalignment='bottom')
ax.set_ylim(-0.03, 1.2)
ax.set_xlabel('t (ms)')
ax.set_ylabel('Activity')
ax.legend(loc='best')
plt.show()
_ = widgets.interact(WC_with_pulse, SE = (0.45, 0.5, .01))
# + [markdown] colab_type="text" id="ThodUE_9OaE8"
# Explore what happened when a second, brief current is applied onto the inhibitory population.
| tutorials/W3D2_DynamicNetworks/student/W3D2_Tutorial2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3D Transformation Matrices
# ---
# - Author: <NAME>
# - GitHub: [github.com/diegoinacio](https://github.com/diegoinacio)
# - Notebook: [3DTransformation_Matrix.ipynb](https://github.com/diegoinacio/computer-vision-notebooks/blob/master/Computer-Graphics/3DTransformation_Matrix.ipynb)
# ---
# Overview and application of tri-dimensional transformation matrices.
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
plt.rcParams['figure.figsize'] = (16, 4)
X, Y, Z = np.mgrid[0:1:5j, 0:1:5j, 0:1:5j]
x, y, z = X.ravel(), Y.ravel(), Z.ravel()
# ## 1. Translation
# ---
# $$
# \large x'=x + t_x \\
# \large y'=y + t_y \\
# \large z'=z + t_z
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} = \begin{bmatrix} 1 & 0 & 0 & t_x \\ 0 & 1 & 0 & t_y \\ 0 & 0 & 1 & t_z \\ 0 & 0 & 0 & 1 \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
# +
def trans_translate(x, y, z, tx, ty, tz):
T = [[1, 0, 0, tx],
[0, 1, 0, ty],
[0, 0, 1, tz],
[0, 0, 0, 1 ]]
T = np.array(T)
P = np.array([x, y, z, [1]*x.size])
return np.dot(T, P)
fig, ax = plt.subplots(1, 4, subplot_kw={'projection': '3d'})
T_ = [[2.3, 0, 0], [0, 1.7, 0], [0, 0, 2.5], [2, 2, 2]]
for i in range(4):
tx, ty, tz = T_[i]
x_, y_, z_, _ = trans_translate(x, y, z, tx, ty, tz)
ax[i].view_init(20, -30)
ax[i].scatter(x_, y_, z_)
ax[i].set_title(r'$t_x={0:.2f}$ , $t_y={1:.2f}$ , $t_z={2:.2f}$'.format(tx, ty, tz))
ax[i].set_xlim([-0.5, 4])
ax[i].set_ylim([-0.5, 4])
ax[i].set_zlim([-0.5, 4])
plt.show()
# -
# 
# ## 2. Scaling
# ---
# Relative to the point $(p_x, p_y, p_z)$
#
# $$
# \large x'=s_x(x - p_x) + p_x = s_x x + p_x(1 - s_x) \\
# \large y'=s_y(y - p_y) + p_y = s_y y + p_y(1 - s_y) \\
# \large z'=s_z(z - p_z) + p_z = s_z z + p_z(1 - s_z)
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} = \begin{bmatrix} s_x & 0 & 0 & p_x(1 - s_x) \\ 0 & s_y & 0 & p_y(1 - s_y) \\ 0 & 0 & s_z & p_z(1 - s_z) \\ 0 & 0 & 0 & 1 \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
# +
def trans_scale(x, y, z,
px, py, pz,
sx, sy, sz):
T = [[sx, 0 , 0 , px*(1 - sx)],
[0 , sy, 0 , py*(1 - sy)],
[0 , 0 , sz, pz*(1 - sz)],
[0 , 0 , 0 , 1 ]]
T = np.array(T)
P = np.array([x, y, z, [1]*x.size])
return np.dot(T, P)
fig, ax = plt.subplots(1, 4, subplot_kw={'projection': '3d'})
S_ = [[1.8, 1, 1], [1, 1.7, 1], [1, 1, 1.9], [2, 2, 2]]
P_ = [[0, 0, 0], [0, 0, 0], [0.45, 0.45, 0.45], [1.1, 1.1, 1.1]]
for i in range(4):
sx, sy, sz = S_[i]; px, py, pz = P_[i]
x_, y_, z_, _ = trans_scale(x, y, z, px, py, pz, sx, sy, sz)
ax[i].view_init(20, -30)
ax[i].scatter(x_, y_, z_)
ax[i].scatter(px, py, pz, s=50)
ax[i].set_title(
r'$p_x={0:.2f}$ , $p_y={1:.2f}$ , $p_z={2:.2f}$'.format(px, py, pz) + '\n'
r'$s_x={0:.2f}$ , $s_y={1:.2f}$ , $s_z={2:.2f}$'.format(sx, sy, sz)
)
ax[i].set_xlim([-2, 2])
ax[i].set_ylim([-2, 2])
ax[i].set_zlim([-2, 2])
plt.show()
# -
# 
# ## 3. Rotation
# ---
# Relative to the point $(p_x, p_y, p_z)$
# $$ \large
# R=R_x(\alpha)R_y(\beta)R_z(\gamma)
# $$
#
# ### 3.1. Rotation around the x-axis
# ---
#
# $$
# \large y'=(y - p_y)\cos\alpha-(z - p_z)\sin \alpha + p_y = y \cos \alpha - z \sin \alpha + p_y(1 - \cos \alpha) + p_z \sin \alpha \\
# \large z'=(y - p_y)\sin\alpha+(z - p_z)\cos \alpha + p_z = y \sin \alpha + z \cos \alpha + p_z(1 - \cos \alpha) - p_y \sin \alpha
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} = \begin{bmatrix}
# 1 & 0 & 0 & 0 \\
# 0 & \cos\alpha & -\sin\alpha & p_y(1 - \cos \alpha) + p_z \sin \alpha \\
# 0 & \sin\alpha & \cos\alpha & p_z(1 - \cos \alpha) - p_y \sin \alpha \\
# 0 & 0 & 0 & 1
# \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
#
# ### 3.2. Rotation around the y-axis
# ---
#
# $$
# \large x'=(x - p_x)\cos\beta+(z - p_z)\sin \beta + p_x = x \cos \beta + z \sin \beta + p_x(1 - \cos \beta) - p_z \sin \beta \\
# \large z'=-(y - p_y)\sin\beta+(z - p_z)\cos \beta + p_z = -x \sin \beta + z \cos \beta + p_z(1 - \cos \beta) + p_x \sin \beta
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} = \begin{bmatrix}
# \cos\beta & 0 & \sin\beta & p_x(1 - \cos \beta) - p_z \sin \beta \\
# 0 & 1 & 0 & 0 \\
# -\sin\beta & 0 & \cos\beta & p_z(1 - \cos \beta) + p_x \sin \beta \\
# 0 & 0 & 0 & 1
# \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
#
# ### 3.3. Rotation around the z-axis
# ---
#
# $$
# \large x'=(x - p_x)\cos\gamma-(y - p_y)\sin \gamma + p_x = x \cos \gamma - y \sin \gamma + p_x(1 - \cos \gamma) + p_y \sin \gamma \\
# \large y'=(x - p_x)\sin\gamma+(y - p_y)\cos \gamma + p_y = x \sin \gamma + y \cos \gamma + p_y(1 - \cos \gamma) - p_x \sin \gamma
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} = \begin{bmatrix}
# \cos\gamma & -\sin\gamma & 0 & p_x(1 - \cos \gamma) + p_y \sin \gamma \\
# \sin\gamma & \cos\gamma & 0 & p_y(1 - \cos \gamma) - p_x \sin \gamma \\
# 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 1
# \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
# +
def trans_rotate(x, y, z, px, py, pz, alpha, beta, gamma):
alpha, beta, gamma = np.deg2rad(alpha), np.deg2rad(beta), np.deg2rad(gamma)
Rx = [[1, 0 , 0 , 0 ],
[0, np.cos(alpha), -np.sin(alpha), py*(1 - np.cos(alpha)) + pz*np.sin(alpha)],
[0, np.sin(alpha), np.cos(alpha), pz*(1 - np.cos(alpha)) - py*np.sin(alpha)],
[0, 0 , 0 , 1 ]]
Ry = [[ np.cos(beta), 0, np.sin(beta), px*(1 - np.cos(beta)) - pz*np.sin(beta)],
[ 0 , 1, 0 , 0 ],
[-np.sin(beta), 0, np.cos(beta), pz*(1 - np.cos(beta)) + px*np.sin(beta)],
[ 0 , 0, 0 , 1 ]]
Rz = [[np.cos(gamma), -np.sin(gamma), 0, px*(1 - np.cos(gamma)) + py*np.sin(gamma)],
[np.sin(gamma), np.cos(gamma), 0, py*(1 - np.cos(gamma)) - px*np.sin(gamma)],
[0 , 0 , 1, 0 ],
[0 , 0 , 0, 1 ]]
Rx = np.array(Rx); Ry = np.array(Ry); Rz = np.array(Rz)
P = np.array([x, y, z, [1]*x.size])
return np.dot(np.dot(np.dot(Rx, Ry), Rz), P)
fig, ax = plt.subplots(1, 4, subplot_kw={'projection': '3d'})
R_ = [[45, 0, 0],
[0, 45, 0],
[0, 0, 45],
[0, 0, 0]]
P_ = [[0, 0, 0],
[0, 0, 0],
[0.5, -0.5, 0.5],
[1.1, 1.1, 1.1]]
for i in range(4):
alpha, beta, gamma = R_[i]; px, py, pz = P_[i]
x_, y_, z_, _ = trans_rotate(x, y, z, px, py, pz, alpha, beta, gamma)
ax[i].view_init(20, -30)
ax[i].scatter(x_, y_, z_)
ax[i].scatter(px, py, pz)
ax[i].set_title(
r'$p_x={0:.2f}$ , $p_y={1:.2f}$ , $p_z={2:.2f}$'.format(px, py, pz) + '\n'
r'$\alpha={0:03d}^o$ , $\beta={1:03d}^o$ , $\gamma={2:03d}^o$'.format(alpha, beta, gamma)
)
ax[i].set_xlim([-2, 2])
ax[i].set_ylim([-2, 2])
ax[i].set_zlim([-2, 2])
plt.show()
# -
# 
# ## 4. Shearing
# ---
# Relative to the point $(p_x, p_y, p_z)$
#
# $$
# \large x' = x + \lambda_x^y(y - p_x) + \lambda_x^z(z - p_x) = x + \lambda_x^y y + \lambda_x^z z - (\lambda_x^y + \lambda_x^z) p_x\\
# \large y' = y + \lambda_y^x(x - p_y) + \lambda_y^z(z - p_y) = y + \lambda_y^x x + \lambda_y^z z - (\lambda_y^x + \lambda_y^z) p_y\\
# \large z' = z + \lambda_z^x(x - p_z) + \lambda_z^y(y - p_z) = z + \lambda_z^x x + \lambda_z^y y - (\lambda_z^x + \lambda_z^y) p_z
# $$
#
# using homogeneous matrix
#
# $$ \large
# \begin{bmatrix} x' \\ y' \\ z' \\ 1 \end{bmatrix} =\begin{bmatrix}
# 1 & \lambda_x^y & \lambda_x^z & -(\lambda_x^y + \lambda_x^z) p_x \\
# \lambda_y^x & 1 & \lambda_y^z & -(\lambda_y^x + \lambda_y^z) p_y \\
# \lambda_z^x & \lambda_z^y & 1 & -(\lambda_z^x + \lambda_z^y) p_z \\
# 0 & 0 & 0 & 1 \end{bmatrix}\begin{bmatrix} x \\ y \\ z \\ 1 \end{bmatrix}
# $$
# +
def trans_shear(x, y, z, px, py, pz,
lambdaxy, lambdaxz,
lambdayx, lambdayz,
lambdazx, lambdazy):
T = [[1 , lambdaxy, lambdaxz, -(lambdaxy+lambdaxz)*px],
[lambdayx, 1 , lambdayz, -(lambdayx+lambdayz)*py],
[lambdazx, lambdazy, 1 , -(lambdazx+lambdazy)*py],
[0 , 0 , 0 , 1 ]]
T = np.array(T)
P = np.array([x, y, z, [1]*x.size])
return np.dot(T, P)
fig, ax = plt.subplots(1, 4, subplot_kw={'projection': '3d'})
L_ = [[[2, 0], [0, 0], [0, 0]],
[[0, 0], [2, 0], [1, 0]],
[[0, 1], [0, 0], [0, 2]],
[[2, 0], [0, 2], [2, 0]]]
P_ = [[0, 0, 0], [0, 0, 0], [0, 1.5, 0], [1.1, 1.1, 1.1]]
for i in range(4):
lambdax, lambday, lambdaz = L_[i]; px, py, pz = P_[i]
x_, y_, z_, _ = trans_shear(x, y, z, px, py, pz,
*lambdax, *lambday, *lambdaz)
ax[i].view_init(20, -30)
ax[i].scatter(x_, y_, z_)
ax[i].scatter(px, py)
ax[i].set_title(
r'$p_x={0:.2f}$ , $p_y={1:.2f}$ , $p_z={2:.2f}$'.format(px, py, pz) + '\n'
r'$\lambda_x^y={0:.2f}$ , $\lambda_y^x={1:.2f}$ , $\lambda_z^x={2:.2f}$'.format(lambdax[0], lambday[0], lambdaz[0]) + '\n'
r'$\lambda_x^z={0:.2f}$ , $\lambda_y^z={1:.2f}$ , $\lambda_z^y={2:.2f}$'.format(lambdax[1], lambday[1], lambdaz[1])
)
ax[i].set_xlim([-3, 3])
ax[i].set_ylim([-3, 3])
ax[i].set_zlim([-3, 3])
plt.show()
# -
# 
| Computer-Graphics/3DTransformation_Matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ALCxPdSdX3NU"
# # Exporting an MNIST Classifier in SavedModel Format
#
# In this exercise, we will learn on how to create models for TensorFlow Hub. You will be tasked with performing the following tasks:
#
# * Creating a simple MNIST classifier and evaluating its accuracy.
# * Exporting it into SavedModel.
# * Hosting the model as TF Hub Module.
# * Importing this TF Hub Module to be used with Keras Layers.
# + colab={} colab_type="code" id="swaA66rjiRTd"
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
from os import getcwd
# + [markdown] colab_type="text" id="UMZdLgyN7gby"
# ## Create an MNIST Classifier
#
# We will start by creating a class called `MNIST`. This class will load the MNIST dataset, preprocess the images from the dataset, and build a CNN based classifier. This class will also have some methods to train, test, and save our model.
#
# In the cell below, fill in the missing code and create the following Keras `Sequential` model:
#
# ```
# Model: "sequential"
# _________________________________________________________________
# Layer (type) Output Shape Param #
# =================================================================
# lambda (Lambda) (None, 28, 28, 1) 0
# _________________________________________________________________
# conv2d (Conv2D) (None, 28, 28, 8) 80
# _________________________________________________________________
# max_pooling2d (MaxPooling2D) (None, 14, 14, 8) 0
# _________________________________________________________________
# conv2d_1 (Conv2D) (None, 14, 14, 16) 1168
# _________________________________________________________________
# max_pooling2d_1 (MaxPooling2 (None, 7, 7, 16) 0
# _________________________________________________________________
# conv2d_2 (Conv2D) (None, 7, 7, 32) 4640
# _________________________________________________________________
# flatten (Flatten) (None, 1568) 0
# _________________________________________________________________
# dense (Dense) (None, 128) 200832
# _________________________________________________________________
# dense_1 (Dense) (None, 10) 1290
# =================================================================
#
# ```
#
# Notice that we are using a ` tf.keras.layers.Lambda` layer at the beginning of our model. `Lambda` layers are used to wrap arbitrary expressions as a `Layer` object:
#
# ```python
# tf.keras.layers.Lambda(expression)
# ```
#
# The `Lambda` layer exists so that arbitrary TensorFlow functions can be used when constructing `Sequential` and Functional API models. `Lambda` layers are best suited for simple operations.
# -
class MNIST:
def __init__(self, export_path, buffer_size=1000, batch_size=32,
learning_rate=1e-3, epochs=10):
self._export_path = export_path
self._buffer_size = buffer_size
self._batch_size = batch_size
self._learning_rate = learning_rate
self._epochs = epochs
self._build_model()
self.train_dataset, self.test_dataset = self._prepare_dataset()
# Function to preprocess the images.
def preprocess_fn(self, x):
# EXERCISE: Cast x to tf.float32 using the tf.cast() function.
# You should also normalize the values of x to be in the range [0, 1].
x = tf.cast(x,dtype=tf.float32) / 255.0 # YOUR CODE HERE
return x
def _build_model(self):
# EXERCISE: Build the model according to the model summary shown above.
self._model = tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(28, 28, 1), dtype=tf.uint8),
# Use a Lambda layer to use the self.preprocess_fn function
# defined above to preprocess the images.
# YOUR CODE HERE
tf.keras.layers.Lambda(self.preprocess_fn),
# Create a Conv2D layer with 8 filters, a kernel size of 3
# and padding='same'.
# YOUR CODE HERE
tf.keras.layers.Conv2D(8, (3,3), activation='relu',padding='same'),
# Create a MaxPool2D() layer. Use default values.
# YOUR CODE HERE
tf.keras.layers.MaxPooling2D(),
# Create a Conv2D layer with 16 filters, a kernel size of 3
# and padding='same'.
# YOUR CODE HERE
tf.keras.layers.Conv2D(16, (3,3), activation='relu',padding='same'),
# Create a MaxPool2D() layer. Use default values.
# YOUR CODE HERE
tf.keras.layers.MaxPooling2D(),
# Create a Conv2D layer with 32 filters, a kernel size of 3
# and padding='same'.
# YOUR CODE HERE
tf.keras.layers.Conv2D(32, (3,3), activation='relu',padding='same'),
# Create the Flatten and Dense layers as described in the
# model summary shown above.
# YOUR CODE HERE
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10, activation='sigmoid')
])
# EXERCISE: Define the optimizer, loss function and metrics.
# Use the tf.keras.optimizers.Adam optimizer and set the
# learning rate to self._learning_rate.
optimizer_fn = tf.keras.optimizers.Adam(learning_rate=self._learning_rate) # YOUR CODE HERE
# Use sparse_categorical_crossentropy as your loss function.
loss_fn = "sparse_categorical_crossentropy"# YOUR CODE HERE
# Set the metrics to accuracy.
metrics_list = ['accuracy'] # YOUR CODE HERE
# Compile the model.
self._model.compile(optimizer_fn, loss=loss_fn, metrics=metrics_list)
def _prepare_dataset(self):
filePath = f"{getcwd()}/../tmp2"
# EXERCISE: Load the MNIST dataset using tfds.load(). Make sure to use
# the argument data_dir=filePath. You should load the images as well
# as their corresponding labels and load both the test and train splits.
dataset = tfds.load(name = 'mnist', data_dir=filePath,as_supervised=True)# YOUR CODE HERE
# EXERCISE: Extract the 'train' and 'test' splits from the dataset above.
# print(dataset)
train_dataset, test_dataset = dataset['train'], dataset['test']# YOUR CODE HERE
#print(len(train_dataset),len(test_dataset))
# print(dataset)
return train_dataset, test_dataset
def train(self):
# EXERCISE: Shuffle and batch the self.train_dataset. Use self._buffer_size
# as the shuffling buffer and self._batch_size as the batch size for batching.
dataset_tr = self.train_dataset.shuffle(self._buffer_size).batch(self._batch_size)
#self._model.summary()
# Train the model for specified number of epochs.
self._model.fit(dataset_tr, epochs=self._epochs)
def test(self):
# EXERCISE: Batch the self.test_dataset. Use a batch size of 32.
dataset_te = self.test_dataset.batch(32)# YOUR CODE HERE
# Evaluate the dataset
results = self._model.evaluate(dataset_te)
# Print the metric values on which the model is being evaluated on.
for name, value in zip(self._model.metrics_names, results):
print("%s: %.3f" % (name, value))
def export_model(self):
# Save the model.
tf.saved_model.save(self._model, self._export_path)
# + [markdown] colab_type="text" id="-dDAjgDe7lp4"
# ## Train, Evaluate, and Save the Model
#
# We will now use the `MNIST` class we created above to create an `mnist` object. When creating our `mnist` object we will use a dictionary to pass our training parameters. We will then call the `train` and `export_model` methods to train and save our model, respectively. Finally, we call the `test` method to evaluate our model after training.
#
# **NOTE:** It will take about 12 minutes to train the model for 5 epochs.
# + colab={"base_uri": "https://localhost:8080/", "height": 287} colab_type="code" id="w6Ba6ileois3" outputId="a280b504-6619-4a1e-c020-6c787b5b76b4"
# Define the training parameters.
args = {'export_path': './saved_model',
'buffer_size': 1000,
'batch_size': 32,
'learning_rate': 1e-3,
'epochs': 5
}
# Create the mnist object.
mnist = MNIST(**args)
# Train the model.
mnist.train()
# Save the model.
mnist.export_model()
# Evaluate the trained MNIST model.
mnist.test()
# + [markdown] colab_type="text" id="sotJ7pQm7umV"
# ## Create a Tarball
#
# The `export_model` method saved our model in the TensorFlow SavedModel format in the `./saved_model` directory. The SavedModel format saves our model and its weights in various files and directories. This makes it difficult to distribute our model. Therefore, it is convenient to create a single compressed file that contains all the files and folders of our model. To do this, we will use the `tar` archiving program to create a tarball (similar to a Zip file) that contains our SavedModel.
# -
# Create a tarball from the SavedModel.
# !tar -cz -f module.tar.gz -C ./saved_model .
# ## Inspect the Tarball
#
# We can uncompress our tarball to make sure it has all the files and folders from our SavedModel.
# + colab={"base_uri": "https://localhost:8080/", "height": 143} colab_type="code" id="NknIrjE1ovkF" outputId="ca2a4d3b-b448-45af-cc7a-44e1096b7974"
# Inspect the tarball.
# !tar -tf module.tar.gz
# + [markdown] colab_type="text" id="n8LjCeO474N4"
# ## Simulate Server Conditions
#
# Once we have verified our tarball, we can now simulate server conditions. In a normal scenario, we will fetch our TF Hub module from a remote server using the module's handle. However, since this notebook cannot host the server, we will instead point the module handle to the directory where our SavedModel is stored.
# + colab={"base_uri": "https://localhost:8080/", "height": 143} colab_type="code" id="C-8vmmtVxJVF" outputId="05176438-367d-4914-d38e-db61d6e69978"
# !rm -rf ./module
# !mkdir -p module
# !tar xvzf module.tar.gz -C ./module
# + colab={} colab_type="code" id="TSmU1oZgxJZS"
# Define the module handle.
MODULE_HANDLE = './module'
# -
# ## Load the TF Hub Module
# + colab={} colab_type="code" id="b2lOfoKab5Rv"
# EXERCISE: Load the TF Hub module using the hub.load API.
model = hub.load(MODULE_HANDLE) # YOUR CODE HERE
# -
# ## Test the TF Hub Module
#
# We will now test our TF Hub module with images from the `test` split of the MNIST dataset.
# + colab={} colab_type="code" id="dCmeWVj_ovno"
filePath = f"{getcwd()}/../tmp2"
# EXERCISE: Load the MNIST 'test' split using tfds.load().
# Make sure to use the argument data_dir=filePath. You
dataset = tfds.load(name = 'mnist',split=tfds.Split.TEST, data_dir=filePath,as_supervised=True)# YOUR CODE HERE
# EXERCISE: Batch the dataset using a batch size of 32.
test_dataset = dataset.batch(32)# YOUR CODE HERE
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="wY9bVLTayn3H" outputId="72dd5ad9-359c-4f71-a054-55a2cb04d6a8"
# Test the TF Hub module for a single batch of data
for batch_data in test_dataset.take(1):
outputs = model(batch_data[0])
outputs = np.argmax(outputs, axis=-1)
print('Predicted Labels:', outputs)
print('True Labels: ', batch_data[1].numpy())
# -
# We can see that the model correctly predicts the labels for most images in the batch.
# + [markdown] colab_type="text" id="ciRPFhPg8FWH"
# ## Evaluate the Model Using Keras
#
# In the cell below, you will integrate the TensorFlow Hub module into the high level Keras API.
# + colab={} colab_type="code" id="YMjnPFOjxmus"
# EXERCISE: Integrate the TensorFlow Hub module into a Keras
# sequential model. You should use a hub.KerasLayer and you
# should make sure to use the correct values for the output_shape,
# and input_shape parameters. You should also use tf.uint8 for
# the dtype parameter.
model = tf.keras.Sequential([
hub.KerasLayer(MODULE_HANDLE,
input_shape=(28,28,1),
dtype=tf.uint8,
output_shape=(10)
)
])
# Compile the model.
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="ShGHxh0Wx7lW" outputId="cce8cbc9-8f95-4965-d0cb-a8ce95e68156"
# Evaluate the model on the test_dataset.
results = model.evaluate(test_dataset)
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="wZ6jUqbDx7s4" outputId="b408ecf5-e91f-4f5c-e147-30047b267131"
# Print the metric values on which the model is being evaluated on.
for name, value in zip(model.metrics_names, results):
print("%s: %.3f" % (name, value))
# -
# # Submission Instructions
# +
# Now click the 'Submit Assignment' button above.
# -
# # When you're done or would like to take a break, please run the two cells below to save your work and close the Notebook. This frees up resources for your fellow learners.
# + language="javascript"
# <!-- Save the notebook -->
# IPython.notebook.save_checkpoint();
# + language="javascript"
# <!-- Shutdown and close the notebook -->
# window.onbeforeunload = null
# window.close();
# IPython.notebook.session.delete();
| Advanced Deployment Scenarios with TensorFlow/week 2/utf-8''TF_Serving_Week_2_Exercise_Question.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python3]
# language: python
# name: conda-env-python3-py
# ---
import numpy as np
import matplotlib.pyplot as plt
plt.pcolormesh(np.random.rand(100,100), cmap='RdBu_r')
clb = plt.colorbar(); plt.clim([0, 1])
clb.set_ticks([0, 0.5, 1])
plt.savefig("colbar.pdf")
| scripts/jupyter/make_video_colorbar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This cell is added by sphinx-gallery
# !pip install mrsimulator --quiet
# %matplotlib inline
import mrsimulator
print(f'You are using mrsimulator v{mrsimulator.__version__}')
# -
#
# # Coesite, ¹⁷O (I=5/2) DAS
#
# ¹⁷O (I=5/2) Dynamic-angle spinning (DAS) simulation.
#
# The following is a Dynamic Angle Spinning (DAS) simulation of Coesite. Coesite has
# five crystallographic $^{17}\text{O}$ sites. In the following, we use the
# $^{17}\text{O}$ EFG tensor information from Grandinetti `et al.` [#f1]_
#
#
# +
import matplotlib.pyplot as plt
from mrsimulator import Simulator
from mrsimulator.methods import Method2D
from mrsimulator import signal_processing as sp
# -
# Create the Simulator object and load the spin systems database or url address.
#
#
# +
sim = Simulator()
# load the spin systems from url.
filename = "https://sandbox.zenodo.org/record/687656/files/coesite.mrsys"
sim.load_spin_systems(filename)
# -
# Use the generic 2D method, `Method2D`, to simulate a DAS spectrum by customizing the
# method parameters, as shown below. Note, the Method2D method simulates an infinite
# spinning speed spectrum.
#
#
# +
das = Method2D(
name="Dynamic Angle Spinning",
channels=["17O"],
magnetic_flux_density=11.74, # in T
spectral_dimensions=[
{
"count": 256,
"spectral_width": 5e3, # in Hz
"reference_offset": 0, # in Hz
"label": "DAS isotropic dimension",
"events": [
{
"fraction": 0.5,
"rotor_angle": 37.38 * 3.14159 / 180,
"transition_query": [{"P": [-1], "D": [0]}],
},
{
"fraction": 0.5,
"rotor_angle": 79.19 * 3.14159 / 180,
"transition_query": [{"P": [-1], "D": [0]}],
},
],
},
# The last spectral dimension block is the direct-dimension
{
"count": 256,
"spectral_width": 2e4, # in Hz
"reference_offset": 0, # in Hz
"label": "MAS dimension",
"events": [
{
"rotor_angle": 54.735 * 3.14159 / 180,
"transition_query": [{"P": [-1], "D": [0]}],
}
],
},
],
)
sim.methods = [das] # add the method
# A graphical representation of the method object.
plt.figure(figsize=(5, 3.5))
das.plot()
plt.show()
# -
# Run the simulation
#
#
sim.run()
# The plot of the simulation.
#
#
# +
data = sim.methods[0].simulation
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(data / data.max(), aspect="auto", cmap="gist_ncar_r")
plt.colorbar(cb)
ax.invert_xaxis()
ax.invert_yaxis()
plt.tight_layout()
plt.show()
# -
# Add post-simulation signal processing.
#
#
processor = sp.SignalProcessor(
operations=[
# Gaussian convolution along both dimensions.
sp.IFFT(dim_index=(0, 1)),
sp.apodization.Gaussian(FWHM="0.3 kHz", dim_index=0),
sp.apodization.Gaussian(FWHM="0.15 kHz", dim_index=1),
sp.FFT(dim_index=(0, 1)),
]
)
processed_data = processor.apply_operations(data=data)
processed_data /= processed_data.max()
# The plot of the simulation after signal processing.
#
#
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(processed_data.real, cmap="gist_ncar_r", aspect="auto")
plt.colorbar(cb)
ax.invert_xaxis()
ax.invert_yaxis()
plt.tight_layout()
plt.show()
# .. [#f1] <NAME>., <NAME>., <NAME>., <NAME>.,
# <NAME>. and <NAME>.
# Solid-State $^{17}\text{O}$ Magic-Angle and Dynamic-Angle Spinning NMR
# Study of the $\text{SiO}_2$ Polymorph Coesite, J. Phys. Chem. 1995,
# **99**, *32*, 12341-12348.
# `DOI: 10.1021/j100032a045 <https://doi.org/10.1021/j100032a045>`_
#
#
| docs/notebooks/examples/2D_simulation(crystalline)/plot_4_DAS_Coesite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Fraction:
def __init__(self, num = 0, den = 1):
if den == 0:
den = 1
self.num = num
self.den = den
def print(self):
if self.num == 0:
print(0)
elif self.den == 1:
print(self.num)
else:
print(f'{self.num} / {self.den}')
def simplify(self):
current = min(self.num, self.den)
while current > 1:
if self.num % current == 0 and self.den % current == 0:
break
current -= 1
self.num = self.num // current
self.den = self.den // current
def add(self, second_object):
newNUM = second_object.den*self.num + second_object.num*self.den
newDEN = second_object.den*self.den
self.num = newNUM
self.den = newDEN
def multiplicatio(self, second_object):
numer = second_object.num*self.num
deno = second_object.den*self.den
print(f'{numer} / {deno}')
f = Fraction(10,5)
f.print()
f.simplify()
f.print()
# + pycharm={"name": "#%%\n"}
f1 = Fraction(2,3)
f2 = Fraction(3,2)
f1.print()
f2.print()
f1.add(f2)
f1.print()
# + pycharm={"name": "#%%\n"}
f3 = Fraction(2,2)
f4 = Fraction(2,2)
f3.print()
f4.print()
f3.multiplicatio(f4)
| 05. OOPS Part-1/6.Fraction-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: research
# language: python
# name: research
# ---
# + [markdown] heading_collapsed=true
# # Setup
# + hidden=true
"""
Add parent directorys to current path
"""
import os.path
import sys
for p in ['..', '../..', '../../..', '../../../..']:
d = os.path.abspath(p)
if d not in sys.path:
sys.path.insert(0,d)
"""
Add tiger-env directory to current path
Still not sure why this is needed.
"""
d = [os.path.abspath('../../../../../custom_envs/gym-tiger'),
os.path.abspath('../../../../../custom_envs/gym-dummy/')]
for _d in d:
if _d not in sys.path:
sys.path.insert(0, _d)
"""
Enable hot-reloading
"""
from notebook_utils import import_module_by_name, reload_module_by_name
def reload():
"""Helper function for hot-reloading QLearnerObsSingle class from source"""
reload_module_by_name(
'experiments.qlearning.basic.qlearner_obs_single.qlearner_obs_single',
'QLearnerObsSingle')
global QLearnerObsSingle
from experiments.qlearning.basic.qlearner_obs_single.qlearner_obs_single \
import QLearnerObsSingle
import gym
import gym_tiger
import gym_dummy
import matplotlib.pyplot as plt
from experiments.qlearning.basic.utils import play_one, plot_running_avg
from experiments.qlearning.basic.qlearner_obs_single.qlearner_obs_single \
import QLearnerObsSingle
# -
# # Tiger-v0
# +
def exp_reward_open_right_growl_left(obs_acc, r_gold, r_tiger,
p_a_listen, p_a_oleft, p_a_oright):
if (P_A_LISTEN + P_A_OLEFT + P_A_ORIGHT) != 1:
raise Exception('Probabilities of previous action must sum to 1.')
p_tiger_left = (obs_acc)*p_a_listen + .5*p_a_oleft + .5*p_a_oright
p_tiger_right = (1-obs_acc)*p_a_listen + .5*p_a_oleft + .5*p_a_oright
print('p_tiger_left', p_tiger_left)
print('p_tiger_right', p_tiger_right)
return r_gold*p_tiger_left + r_tiger*p_tiger_right
OBS_ACC = .85
R_GOLD = 49
R_TIGER = -51
P_A_LISTEN = 1/2
P_A_OLEFT = 1/2
P_A_ORIGHT = 0
exp_reward_open_right_growl_left(OBS_ACC, R_GOLD, R_TIGER,
P_A_LISTEN, P_A_OLEFT, P_A_ORIGHT)
# + [markdown] heading_collapsed=true
# ## Setup ENV and Model
# + hidden=true
import gym
import gym_tiger
import matplotlib.pyplot as plt
from experiments.qlearning.basic.utils import play_one, plot_running_avg
from experiments.qlearning.basic.qlearner_obs_single.qlearner_obs_single \
import QLearnerObsSingle
env = gym.make('Tiger-v0')
env.__init__(reward_tiger=-100, reward_gold=10, reward_listen=-1,
max_steps_per_episode=500)
model = QLearnerObsSingle(env, initial_alpha=.5, gamma=.9, alpha_decay=.4)
eps = 1
n = 0
ot = env.reset()
# + [markdown] heading_collapsed=true
# ## Take one action and update Q
# + [markdown] hidden=true
# $$
# Q(s_{t-1}, a_{t-1}) = Q(s_{t-1}, a_{t-1}) + \alpha \big[ r_t + \gamma \cdot Q(s_t, a_t) - Q(s_{t-1}, a_{t-1}) \big]
# $$
# + hidden=true
print ('Q values at n=0')
print(model)
otm1 = ot
atm1 = model.sample_action(otm1, eps)
ot, r, done, info = env.step(atm1)
at = model.best_action(ot)
model.update(otm1, atm1, r, ot, at)
_otm1 = env.translate_obs(otm1)
_atm1 = env.translate_action(atm1)
print(_otm1, ',', _atm1, ',', r)
print(model)
# + [markdown] heading_collapsed=true
# ## Play 1 Episode (500 steps)
# + hidden=true
play_one(env, model, eps, verbose=True)
# + hidden=true
print(model)
# -
# ## Play 100 Episodes
# +
env = gym.make('Tiger-v0')
env.__init__(reward_tiger=-85, reward_gold=15, reward_listen=-1,
max_steps_per_episode=500)
model = QLearnerObsSingle(env, initial_alpha=.01, gamma=.9, alpha_decay=0)
eps = .1
n = 0
ot = env.reset()
N = 100
totalrewards = np.empty(N)
print()
for n in range(N):
if n % (N/5) == 0:
print('Q values at n={}'.format(n))
print(model)
if n > 25:
eps = 0
else:
eps = 1.0/np.sqrt(n+1)
totalreward = play_one(env, model, eps)
totalrewards[n] = totalreward
print('Q values at n={}'.format(N))
print(model)
print()
print("avg reward for last {} episodes:".format(N/5),
totalrewards[int(-1*(N/5)):].mean())
fig, ax = plt.subplots(1, 1, figsize=(20, 5))
ax.plot(totalrewards)
ax.set_title("Rewards")
plot_running_avg(totalrewards[25:], window=N//25)
# + [markdown] heading_collapsed=true
# # GreaterThanZero-v0
# + hidden=true
class GreaterThanZeroFeatureTransformer:
def __init__(self):
pass
def transform(self, o):
if o[0] < 0:
return 0
else:
return 1
env = gym.make('GreaterThanZero-v0')
env.__init__()
model = QLearnerObsSingle(env, initial_alpha=.5, gamma=.8,
feature_transformer=DummyFeatureTransformer(),
alpha_decay=.4)
eps = 1
n = 0
ot = env.reset()
N = 500
totalrewards = np.empty(N)
print(model)
for n in range(N):
if n >= N - (N/5):
eps = 0
else:
eps = 1.0/np.sqrt(n+1)
totalreward = play_one(env, model, eps)
totalrewards[n] = totalreward
if n % (N/5) == 0:
print(model)
print("avg reward for last {} episodes:".format(N/5),
totalrewards[int(-1*(N/5)):].mean())
fig, ax = plt.subplots(1, 1, figsize=(20, 5))
ax.plot(totalrewards)
ax.set_title("Rewards")
plot_running_avg(totalrewards, window=5)
# + hidden=true
| experiments/qlearning/basic/qlearner_obs_single/qlearner-obs-single.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Knuth-Morris-Pratt Algorithm
# Problem: find all occurences of a pattern P of length n in text T of length n
T = "xyxxyxyxyyxyxyxyyxyxyxxy"
P = "xyxyyxyxyxx"
# KMP = O(m+n) solution to the above problem
# Reference: http://www.cs.ubc.ca/labs/beta/Courses/CPSC445-08/Handouts/kmp.pdf
# Overview:
# If we have matched P[0,...,q] with T[i-q,...,i], we check if P[q] == T[i+1]
# If true, we increment i and q | note the occurence of the pattern if q = m and slide the pattern to the right | end the algorithm if we have reached the end of the text
# If false, we slide the pattern to the right | increment i if we can no longer slide it to the right (S[q] = -1)
# Sliding:
# We precompute a table S so that S[q] gives us the longest proper prefix of substring P[0...q] that is also a suffix of the substring. Proper prefix = any prefix that isn't the whole substring
# Computing array S:
# S[0] = -1
# At step i, we want to compute S[i] and we have S[i-1]
# q <- S[i-1]
# While(True):
# If P[i] = P[q+1] => S[i] = q+1 + break
# Else q <- S[q] + if q = -1 break (we go to the previous longest proper prefix we found)
#
# Extra explanation for the last line: Say we have the substring xyxyp.....xyxyt
# p != t => we consider the longest common prefix of the substring 'xyxy' (the previous longest common prefix that is also a suffix) ('xy') and check if x == t and so on.
# Why not consider 'xyx'? Because we already know that 'xyx' is not a suffix of 'xyxy', we already computed what to try out next.
def compute_prefix_table(P):
S = [-1] * len(P)
for ii in range(1,len(P)):
#print(ii)
q = S[ii-1]
while(True):
if(P[ii] == P[q+1]):
S[ii] = q+1
break
else:
if(q == -1):
break
q = S[q]
if(q == -1):
break
return S
S = compute_prefix_table(P)
S
# Sanity check for non-zero values of S:
for ii in range(len(S)):
if(S[ii] == -1):
continue
print(str(S[ii]) + ': ' + P[:ii+1] + ' ' + P[:S[ii]+1])
# KMP Algorithm:
# Returns an array of indices where pattern P occurs in T
def kmp(T,P):
S = compute_prefix_table(P)
occ = []
if(len(P) > len(T)):
return []
if(len(P) == len(T)):
return [0] if P == T else []
jj = -1 # index in P such that P[0...jj-1] is matched with T[ii-jj+1,...,ii]
ii = -1
while(ii < len(T)-1):
if(P[jj+1] == T[ii+1]):
if(jj+1 == len(P) - 1):
occ.append(ii-len(P)+2)
jj = S[len(P)-1]
else:
jj += 1
ii += 1
else:
if(jj == -1):
ii += 1
else:
jj = S[jj]
return occ
occ = kmp(T,P)
P
T[occ[0]:]
occ
# +
T = "aababcabcdabcdeabcdef"
P = "abcdef"
occs = kmp(T,P)
for occ in occs:
print(T[occ:occ+len(P)] + ' @pos: ' + str(occ))
# -
| ads_simple_implementations/Knuth-Morris-Pratt Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulation exercises
import numpy as np
# ## Penalty shootout technique
#
# Jen is the penalty taker for her football team.
#
# She's been doing this for a long time. She thinks she normally has a 80%
# chance of scoring.
#
# In the last 15 penalties she has taken, she has been trying a new technique.
# She scored on all 15 penalties.
#
# How certain can she be that this would not have happened, using the old
# technique?
# ## Aim big
#
# John is playing Monopoly. His piece, the top hat, is sitting in a really bad
# spot, just in front of some expensive hotels. He is about to roll the two
# 6-sided die. He needs a score of 10 or more to skip over the hotels. What are
# John's chances?
#
# Solve by simulation.
#
# Hint: consider `np.random.randint`. Read the help with `np.random.randint?`.
# ## Blackjack
#
# Given any three random playing cards, what is the chance that the ranks of the
# three cards add up to 21?
#
# 10, jack, queen and king all count as 10. For example, one way of getting 21
# is a seven, a four and a king.
# ### Simple version
#
# Assume the three cards are each dealt from the top of a full shuffled deck.
# Therefore, the procedure is:
#
# * you shuffle, look at the top card, record the rank, you put it back.
# * repeat twice more.
#
# Assume that the ace counts as 1. What are the chances of getting
# a total rank of 21?
#
# Hint: start with this array:
ranks = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 10, 10, 10])
# Investigate `np.random.choice` to use this array for the problem.
# ## Less simple version
#
# Assume the cards are drawn as they were for the problem above.
#
# Now the ace can count as 1 or as 11, whichever gives a total of 21. Now what are the chances of a total rank of 21?
#
# Hint: you can change values of 1 to 11 like this:
# Make an example array
some_cards = np.array([1, 3, 5, 1, 2, 10, 1, 4])
some_cards
# Make a Boolean array that has True for positions where some_cards == 1
card_eq_1 = some_cards == 1
card_eq_1
# In the found positions, change the value to 11
some_cards[card_eq_1] = 11
some_cards
# You might want to use this kind of trick more than once in your solution.
| exercises/simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from itertools import combinations
from hashlib import sha256
from nltk import ngrams
from struct import pack
# # PPRL with field-level Bloom filters
# For a Bloom filter we define:
#
# - a bit vector $v$ of length $l$ with all values initially set to $0$;
# - a set $H$ with $k$ independent hash functions with range $0$ to $l - 1$.
#
# To convert a set $S$ to a Bloom filter:
#
# - each element $x_i \in S$ is hashed using the $k$ hash functions;
# - all bits in $v$ with indices $h_j(x_i)$, for $1 \leq j \leq k$, are set to $1$;
#
# First, all fields are converted to a set of n-grams. This n-gram set is used to create field-level Bloom filters (FBFs).
# +
l = 512
k = 2
H = [sha256(sha256(bytes(i)).digest()) for i in range(k)]
def n_grams(field):
"""Converts a field to a set of n-grams."""
return [''.join(ng) for ng in ngrams(' {} '.format(field), 2)]
def bit_vector(size):
"""Returns a bit vector with all values set to zero."""
return [0 for _ in range(size)]
def hash_indices(x):
"""Returns the indices generated by h(x)."""
if type(x) is str:
x = x.encode('UTF-8')
for h in H:
g = h.copy()
g.update(x)
digest = int(g.hexdigest(), 16)
yield digest % l
# +
p1 = ('John', 'Smith', 'Amsterdam')
p2 = ('John', 'Smyth', 'Amsterdam')
p3 = ('Johhny', 'Smith', '<NAME>')
p1_S = [n_grams(f) for f in p1]
p2_S = [n_grams(f) for f in p2]
p3_S = [n_grams(f) for f in p3]
p1_v = [bit_vector(l) for _ in p1]
p2_v = [bit_vector(l) for _ in p2]
p3_v = [bit_vector(l) for _ in p3]
all_p = [p1] + [p2] + [p3]
all_S = p1_S + p2_S + p3_S
all_v = p1_v + p2_v + p3_v
# Construct FBFs.
for (S, v) in zip(all_S, all_v):
for x in S:
for i in hash_indices(x):
v[i] = 1
# -
# Similarity between FBFs is calculated using the Dice-coefficient:
#
# $$Dice\_sim(v_1, v_2) = \frac{2c}{x_1 + x_2}$$
#
# - $c$ is the number of common bit positions in $v_1$ and $v_2$ that are set to $1$;
# - $x_1$ is the number of bit positions in $v_1$ set to $1$;
# - $x_2$ is the number of bit positions in $v_2$ set to $1$.
#
# The similarity of two records is the mean of all pairwise FBF similarities:
#
# $$record\_sim(V_a, V_b) = \frac{1}{n}\sum_{i=1}^{n}{Dice\_sim(v_{i}^{(a)}, v_{i}^{(b)})}$$
#
# - $V_a$ is the ordered list of $n$ FBFs constructed from record $a$;
# - $V_b$ is the ordered list of $n$ FBFs constructed from record $b$;
# - $v_{i}^{(a)}$ is the $i$-th FBF from $V_a$;
# - $v_{i}^{(b)}$ is the $i$-th FBF from $V_b$;
# +
def dice_sim(a, b):
c = sum([1 for (i, j) in zip(a, b) if i + j == 2])
x1 = sum(a)
x2 = sum(b)
return (2*c) / (x1 + x2)
def record_sim(Va, Vb):
s = 0
for (va, vb) in zip(Va, Vb):
s += dice_sim(va, vb)
return s / len(Va)
for (a, b) in combinations([(p1, p1_v), (p2, p2_v), (p3, p3_v)], 2):
record_a = ' '.join(a[0])
record_b = ' '.join(b[0])
similarity = record_sim(a[1], b[1])
print('Similarity between \'{0}\' and \'{1}\': {2:.3}'.format(record_a, record_b, similarity))
# -
# ## References
# - [Privacy-preserving record linkage using Bloom filters](https://www.semanticscholar.org/paper/Privacy-preserving-record-linkage-using-Bloom-filt-Schnell-Bachteler/8392fa13e5013073b617e947b0229bf1734990ac)
# - [Composite Bloom filters for secure record linkage](https://www.semanticscholar.org/paper/Composite-Bloom-Filters-for-Secure-Record-Linkage-Durham-Kantarcioglu/bb1dbac86d922b91daabf61ec75f56dde47997be)
| record-linkage/PPRL with field-level Bloom filters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Hyperalignment Tutorial
#
# This jupyter notebook is an example to use searchlight hyperalignment (Guntupalli et al., 2016) on fMRI movie data and benchmark its performance.
#
# In this example, we will use some minimal data from the Guntupalli et al. (2016) paper to save computation time. This minimal dataset contains 3 subjects, 2 movie runs per subject, and left hemisphere data only. The data have been preprocessed with motion correction, surface-based alignment, and denoising.
# ## 0. Preparations
#
# We will use the docker image from https://github.com/Summer-MIND/mind-tools
#
# Reopen the container by typing
# ```
# docker start MIND && docker attach MIND
# ```
# in the command line. (Or
# ```
# docker run -it -p 9999:9999 --name MIND -v ~/Desktop:/mnt ejolly/mind-tools
# ```
# if you haven't used it before).
#
# Then, within the docker container, let's create the directory and download the tutorial data.
# ```
# # mkdir /mnt/hyperalignment
# # cd /mnt/hyperalignment
# wget http://discovery.dartmouth.edu/~fma/hyper_data.tar.gz
# wget http://discovery.dartmouth.edu/~fma/hyperalignment_tutorial.ipynb
# tar xzvf hyper_data.tar.gz
# ```
#
# Finally, prepare the python packages we will use. Here we will use python2 because PyMVPA dependency h5py is not compatible with python3.
# ```
# source activate py27
# pip install h5py nibabel pprocess pymvpa2
# ```
#
# After all these, you can start a jupyter notebook using
# ```
# jupyter notebook --port=9999 --no-browser --ip=0.0.0.0 --allow-root
# ```
# And copy the url from the terminal to your web browser.
# ## 1. Import python functions and classes
# %matplotlib inline
import numpy as np
from scipy.spatial.distance import pdist, cdist
from mvpa2.datasets.base import Dataset
from mvpa2.mappers.zscore import zscore
from mvpa2.misc.surfing.queryengine import SurfaceQueryEngine
from mvpa2.algorithms.searchlight_hyperalignment import SearchlightHyperalignment
from mvpa2.base.hdf5 import h5save, h5load
# Alternatively, all those above can be imported using
# from mvpa2.suite import *
import matplotlib.pyplot as plt
from mvpa2.support.nibabel.surf import read as read_surface
# ## 2. Read data
# The data are read from numpy npy files and wrapped as Datasets. Features (vertices) are normalized to have unit variance.
# +
dss_train = []
dss_test = []
subjects = ['rid000005', 'rid000011', 'rid000014']
for subj in subjects:
ds = Dataset(np.load('raiders/{subj}_run00_lh.npy'.format(subj=subj)))
ds.fa['node_indices'] = np.arange(ds.shape[1], dtype=int)
zscore(ds, chunks_attr=None)
dss_train.append(ds)
ds = Dataset(np.load('raiders/{subj}_run01_lh.npy'.format(subj=subj)))
ds.fa['node_indices'] = np.arange(ds.shape[1], dtype=int)
zscore(ds, chunks_attr=None)
dss_test.append(ds)
# -
# Each run has 336 time points and 10242 features per subject.
print(dss_train[0].shape)
print(dss_test[0].shape)
# ## 3. Create SearchlightHyperalignment instance
# The QueryEngine is used to find voxel/vertices within a searchlight. This SurfaceQueryEngine use a searchlight radius of 5 mm based on the fsaverage surface.
sl_radius = 5.0
qe = SurfaceQueryEngine(read_surface('fsaverage.lh.surf.gii'), radius=sl_radius)
hyper = SearchlightHyperalignment(
queryengine=qe,
compute_recon=False, # We don't need to project back from common space to subject space
nproc=1, # Number of processes to use. Change "Docker - Preferences - Advanced - CPUs" accordingly.
)
# ## 4. Create common template space with training data
# This step may take a long time. In my case it's 10 minutes with `nproc=1`.
# +
# mappers = hyper(dss_train)
# h5save('mappers.hdf5.gz', mappers, compression=9)
mappers = h5load('mappers.hdf5.gz') # load pre-computed mappers
# -
# ## 5. Project testing data to the common space
dss_aligned = [mapper.forward(ds) for ds, mapper in zip(dss_test, mappers)]
_ = [zscore(ds, chunks_attr=None) for ds in dss_aligned]
# ## 6. Benchmark inter-subject correlations
def compute_average_similarity(dss, metric='correlation'):
"""
Returns
=======
sim : ndarray
A 1-D array with n_features elements, each element is the average
pairwise correlation similarity on the corresponding feature.
"""
n_features = dss[0].shape[1]
sim = np.zeros((n_features, ))
for i in range(n_features):
data = np.array([ds.samples[:, i] for ds in dss])
dist = pdist(data, metric)
sim[i] = 1 - dist.mean()
return sim
sim_test = compute_average_similarity(dss_test)
sim_aligned = compute_average_similarity(dss_aligned)
plt.figure(figsize=(6, 6))
plt.scatter(sim_test, sim_aligned)
plt.xlim([-.2, .5])
plt.ylim([-.2, .5])
plt.xlabel('Surface alignment', size='xx-large')
plt.ylabel('SL Hyperalignment', size='xx-large')
plt.title('Average pairwise correlation', size='xx-large')
plt.plot([-1, 1], [-1, 1], 'k--')
plt.show()
# ## 7. Benchmark movie segment classifications
def movie_segment_classification_no_overlap(dss, window_size=6, dist_metric='correlation'):
"""
Parameters
==========
dss : list of ndarray or Datasets
window_size : int, optional
dist_metric : str, optional
Returns
=======
cv_results : ndarray
An n_subjects x n_segments boolean array, 1 means correct classification.
"""
dss = [ds.samples if hasattr(ds, 'samples') else ds for ds in dss]
def flattern_movie_segment(ds, window_size=6):
n_seg = ds.shape[0] // window_size
ds = ds[:n_seg*window_size, :].reshape((n_seg, window_size, -1))
ds = ds.reshape((n_seg, -1))
return ds
dss = [flattern_movie_segment(ds, window_size=window_size) for ds in dss]
n_subj, n_seg = len(dss), dss[0].shape[0]
ds_sum = np.sum(dss, axis=0)
cv_results = np.zeros((n_subj, n_seg), dtype=bool)
for i, ds in enumerate(dss):
dist = cdist(ds, (ds_sum - ds) / float(n_subj - 1), dist_metric)
predicted = np.argmin(dist, axis=1)
acc = (predicted == np.arange(n_seg))
cv_results[i, :] = acc
return cv_results
acc_test = movie_segment_classification_no_overlap(dss_test)
acc_aligned = movie_segment_classification_no_overlap(dss_aligned)
print('Classification accuracy with surface alignment: %.1f%%' % (acc_test.mean()*100, ))
print('Classification accuracy with SL hyperalignment: %.1f%%' % (acc_aligned.mean()*100, ))
print('Classification accuracy with surface alignment per subject:', acc_test.mean(axis=1))
print('Classification accuracy with SL hyperalignment per subject:', acc_aligned.mean(axis=1))
# ## Extras
#
# If you have completed all the practices above and want to try more, here are some possible options:
#
# ### 1
#
# Try to apply this method to your own surface data. For example, you can create a common template space with movie data and project retinotopic data to the common space. Gifti files can be loaded using `mvpa2.datasets.gifti.gifti_dataset`.
#
# ### 2
#
# Try to use ROI hyperalignment (`mvpa2.algorithms.hyperalignment.Hyperalignment`) instead of searchlight hyperalignment, and compare computation time and results.
#
# ### 3
#
# Read (and practice) more with the more content-rich hyperalignment tutorial http://nbviewer.jupyter.org/url/www.pymvpa.org/notebooks/hyperalignment.ipynb
#
# Data can be downloaded from http://data.pymvpa.org/datasets/hyperalignment_tutorial_data/hyperalignment_tutorial_data.hdf5.gz
#
| Tutorials/hyperalignment/hyperalignment_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Introduction
# This notebook illustrates the motion of an electron into a rectangular waveguide.
#
# # References
# * Semenov et al., Multipactor in Rectangular Waveguides, Physics of Plasmas 14, 033501 (2007).
# Python modules import
# %pylab
# %matplotlib inline
import numpy as np # numpy
from scipy.constants import c, pi, m_e, e, mu_0 # some physical constants
# We consider here the fundamental mode TE10 in a rectangular waveguide of width $a$ ($-a/2<x<a/2$) and height $b$ ($0<y<b$):
# $$
# \begin{array}{ccl}
# E_y &=& -E_0 \cos(k_\perp x) \sin(\omega t - k z) \\
# H_x &=& \frac{c k}{\omega} E_0 \cos(k_\perp x) \sin(\omega t - k z) \\
# H_z &=& \frac{c k_\perp}{\omega} E_0 \sin(k_\perp x) \cos(\omega t - k z)
# \end{array}
# $$
def EH_field_rect_wg(x, z, t, f, E0, a):
"""
Returns the E and H field component of the TE10 mode
in a rectangular waveguide (a x b).
x being defined as [-a/2,a/2] and y in [0,b]
Inputs:
- x, z, t : spatial and time coordinates
- f: frequency in [Hz]
- E0: Electric field amplitude in [V/m]
- a: waveguide width [m]
Returns:
- Ey, Hx, Hz
"""
k_perp = pi/a
omega = 2*pi*f
k = np.sqrt((omega/c)**2 - k_perp**2)
Ey = -E0*np.cos(k_perp*x)*np.sin(omega*t - k*z)
Hx = c*k/omega*E0*np.cos(k_perp*x)*np.sin(omega*t - k*z)
Hz = c*k_perp/omega*E0*np.sin(k_perp*x)*np.cos(omega*t - k*z)
return Ey, Hx, Hz
# The motion of the electron inside the waveguide is given by $m\ddot{\mathbf{x}}=q(\mathbf{E} + \dot{\mathbf{x}}\times\mathbf{B})$, i.e.:
# $$
# \begin{array}{ccl}
# m \ddot{y} &=& -e E_y - e H_x \dot{z}/c + e H_z \dot{x}/c \\
# m \ddot{x} &=& -e H_z \dot{y}/c \\
# m \ddot{z} &=& e H_x \dot{y}/c
# \end{array}
# $$
# The previous system is rewritten with only first time derivatives system $\frac{d \mathbf{u}}{dt}=\mathbf{f}(\mathbf{u},t)$, with:
# $$
# \mathbf{u} =
# \left(
# \begin{array}{c}
# x \\ y \\ z \\ \dot{x} \\ \dot{y} \\ \dot{z}
# \end{array}
# \right)
# $$
# and
# $$
# \mathbf{f} =
# \left(
# \begin{array}{c}
# \dot{x} \\ \dot{y} \\ \dot{z} \\
# -\frac{e}{m c} H_z \dot{y} \\
# -\frac{e}{m} E_y - \frac{e}{m c} H_x \dot{z} + \frac{e}{m c} H_z \dot{x} \\
# \frac{e}{m c} H_x \dot{y}
# \end{array}
# \right)
# $$
def fun(u, t, f, E0, a):
"""
Computes the derivatives f(u,t).
Inputs:
- u: (x,y,z,xdot,ydot,zdot) (6x1 array)
- t: time [s]
- E0: Electric field amplitude [V/m]
- a: rectangular waveguide width [m]
Returns:
- f(u,t) (6x1 array)
"""
x, y, z, xdot, ydot, zdot = u # unpacking
Ey, Hx, Hz = EH_field_rect_wg(x, z, t, f, E0, a)
# Additional DC field along y
H0 = 0/mu_0
f = [xdot,
ydot,
zdot,
-e/(m_e*c)*Hz*ydot + e/(m_e*c)*H0*zdot,
-e/m_e*Ey - e/(m_e*c)*Hx*zdot + e/(m_e*c)*Hz*xdot,
e/(m_e*c)*Hx*ydot - e/(m_e*c)*H0*xdot]
return f
# Constants
a = 72e-3
b = 34e-3
f = 3.7e9
E0 = 1e5
t = linspace(0e-9, 200/f, 501) # time range to solve : 200 RF periods
# +
from scipy.integrate import odeint
# electron initial location
x0, y0, z0 = [-a/8, 0, 0]
# electron initial velocity
vx0, vy0, vz0 = [0, 0, 0]
# initial condition
u0 = [x0, y0, z0, vx0, vy0, vz0]
# solve
u_num = odeint(fun, u0, t, args=(f, E0, a))
# -
# plot the (x(t), y(t)) motion of the electron in the waveguide cross section
plot(u_num[:,0], u_num[:,1], color='r', lw=2)
# superpose the Efield
x = linspace(-a/2, a/2, 101)
fill_between(x, b*cos(pi/a*x), alpha=0.1)
# shade the waveguide walls for illustration
axis('equal')
axis([-a/2-5e-3, +a/2+5e-3, 0-5e-3, b+5e-3])
axhspan(ymin=0-10e-3, ymax=0, color='#555555')
axhspan(ymin=b, ymax=b+10e-3, color='#555555')
axvspan(xmin=-a/2-10e-3, xmax=-a/2, color='#555555')
axvspan(xmin=+a/2, xmax=a/2+10e-3, color='#555555')
# (x,z) plot
plot(u_num[:,0], u_num[:,2])
axis([-a/2-2e-3, a/2+2e-3, -10e-3, +10e-3], 'equal')
axvline(-a/2, color='k')
axvline(+a/2, color='k')
xlabel('x [m]')
ylabel('z [m]')
# In order to illustrate the motion for different starting point, we solve for many starting points and plot the result.
# +
x0_vec = linspace(-a/2, a/2, 11)
u_num_vec = []
for x0 in x0_vec:
# initial condition
u0 = [x0, y0, z0, vx0, vy0, vz0]
# solve
u_num_vec.append(odeint(fun, u0, t, args=(f, E0, a)))
# +
for u_num in u_num_vec:
plot(u_num[:,0], u_num[:,1], color='r')
# shade the waveguide walls for illustration
axis('equal')
axis([-a/2-5e-3, +a/2+5e-3, 0-5e-3, b+5e-3])
axhspan(ymin=0-10e-3, ymax=0, color='#555555')
axhspan(ymin=b, ymax=b+10e-3, color='#555555')
axvspan(xmin=-a/2-10e-3, xmax=-a/2, color='#555555')
axvspan(xmin=+a/2, xmax=a/2+10e-3, color='#555555')
# -
# In order to be resonant with the wall height $b$, a particle of velocity $V$ must travel a distance $\approx b$ (if one neglects the RF magnetic field in this case) during a half RF-period $T/2=1/(2f)=\pi/\omega$. More generally, it can be a odd number of (half-period) : 1, 3, 5...
# Thus, the resonance coundition is expressed as :
# $$
# b \approx V j / (2 f)
# $$
# thus
# $$
# b \omega \approx V j \pi
# $$
# The particle velocity is the addition of its initial emission velocity $V_{y0}$ (if emitted only in the normal direction) and the RF kick, given by :
# $$
# V_{RF} \approx \frac{e E_y}{m \omega}
# $$
# again in the parallel plate approximation.
| notebooks/Rectangular Waveguide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
import time
import pickle
# # cells defined in earlier notebooks
def R_nonbinding_3eq(y,t):
"""
system of ODEs from Zaytsev 2016, simplified using two mass balances
with the following components:
- a: inactive Aurora B kinase
- A: active Aurora B kinase
- AA: enzyme-substrate complex of inactive + active Aurora B kinase
- Ph: phosphatase
- PhA: enzyme-substrate complex of phosphatase + active Aurora B kinase
- a0: total Aurora B kinase
- p0: total phosphatase
"""
# set variable space
A, AA, Ph = y
# mass balances
PhA = p0 - Ph
a = a0 - A - 2*AA - PhA
# reaction equations
dAdt = (kcis - kfa*A)*a + (kra+2*kca)*AA - kfp*A*Ph + krp*PhA
dAAdt = kfa*A*a - (kra+kca)*AA
dPhdt = -kfp*A*Ph + (krp + kcp)*PhA
return dAdt, dAAdt, dPhdt
# +
"""
parameters from Zaytsev 2016
"""
kcis = 7.29*10**-6 # 1/s # rate constant for 'in cis' Aurora B activation
kfa = 0.1 # 1/(uM*s) # rate constant for AA complex formation
kca = 2.7*10**-2 # 1/s # rate constant for AA catalysis
Kma = 51 # uM # Michaelis constant for AA 'in trans' activation
kra = kfa*Kma-kca # 1/ # rate constant for AA complex dissociation
kfp = 0.6 # 1/(uM*s) # rate constant for PhA complex formation
kcp = 2.4*10**-2 # 1/s # rate constant for PhA catalysis
Kmp = 1.95 # uM # Michaelis constant for PhA 'in trans' activation
krp = kfp*Kmp-kcp # 1/s # rate constant for PhA complex dissociation
# +
"""
algorithm to find high + low steady states
for different phosphatase concentrations + 10 uM total kinase
uses zero for the low state in phosphatase concentrations with monostable high states
to be used as initial active kinase concentrations for spatial simulations
"""
t = np.linspace(0,2000*60,2000*60)
a0 = 10
# build phosphatase concentration list
p0_range = [0,.2]
for n in np.arange(.4,.61,.01):
p0_range.append(round(n,2))
# temporal evolution to find steady state concentrations with low initial kinase activity
lo_ss_nbns = []
for p0 in p0_range:
y = odeint(R_nonbinding_3eq,[0,0,p0],t)
A, aA, Ph = y[-1,:]
# use zero initial active kinase for points with monostable high states
if A > 1:
lo_ss_nbns.append((str(p0),0,a0,0,p0,0))
else:
PhA = p0 - Ph
a = a0-A-2*aA-PhA
lo_ss_nbns.append((str(p0),A, a, aA, Ph, PhA))
# temporal evolution to find steady state concentrations with high initial kinase activity
hi_ss_nbns = []
for p0 in p0_range:
y = odeint(R_nonbinding_3eq,[a0,0,p0],t)
A, aA, Ph = y[-1,:]
PhA = p0 - Ph
a = a0-A-2*aA-PhA
hi_ss_nbns.append((str(p0),A, a, aA, Ph, PhA))
# -
def mesh_fourier(zmin,zmax,nz):
"""
discrete approximation of the spatial derivative operator (Laplacian)
uses spectral symmetry to simplify operations, forces periodic boundary conditions
"""
dz = np.zeros((nz,nz))
for i in range(nz):
for j in range(nz):
if i == j:
dz[i,i] = 0
else:
dz[i,j] = np.pi*(-1)**((i-1)+(j-1))/(zmax-zmin)/np.tan(((i-1)-(j-1))*np.pi/(nz))
return dz
def R_nonbinding_5eq(y,t):
"""
system of ODEs from Zaytsev 2016, without simplifications
"""
# unpack species profiles
A, a, aA, Ph, PhA = y
# calculate reaction equations
dadt = -(kcis + kfa*A)*a + kra*aA + kcp*PhA
dAdt = (kcis - kfa*A)*a + (kra + 2*kca)*aA - kfp*A*Ph + krp*PhA
daAdt = kfa*A*a - (kra + kca)*aA
dPhdt = -kfp*A*Ph + (krp + kcp)*PhA
dPhAdt = -dPhdt
# output concentration changes
return dAdt, dadt, daAdt, dPhdt, dPhAdt
def spatial_simulation_nonbind_ss_perturb(lo_ss,hi_ss,t_end,dt,t_save,L,N,perturb_width):
"""
reaction-diffusion algorithm with a perturbed center width as initial conditions
combines the above kinase-phosphatase reaction network + simple diffusion algorithm
"""
# extract the information from the initial condition array function inputs
lostr, A0_lo, a0_lo, aA0_lo, Ph0_lo, PhA0_lo = lo_ss
histr, A0_hi, a0_hi, aA0_hi, Ph0_hi, PhA0_hi = hi_ss
# initilize perturbed conditions for each reacting species
A = np.ones(N)*A0_lo
A[round(N/2-perturb_width/2):round(N/2+perturb_width/2)] = np.ones(perturb_width)*A0_hi
a = np.ones(N)*a0_lo
a[round(N/2-perturb_width/2):round(N/2+perturb_width/2)] = np.ones(perturb_width)*a0_hi
aA = np.ones(N)*aA0_lo
aA[round(N/2-perturb_width/2):round(N/2+perturb_width/2)] = np.ones(perturb_width)*aA0_hi
Ph = np.ones(N)*Ph0_lo
Ph[round(N/2-perturb_width/2):round(N/2+perturb_width/2)] = np.ones(perturb_width)*Ph0_hi
PhA = np.ones(N)*PhA0_lo
PhA[round(N/2-perturb_width/2):round(N/2+perturb_width/2)] = np.ones(perturb_width)*PhA0_hi
# combine species profiles into a single variable
y = A, a, aA, Ph, PhA
A_arr = np.zeros((N,round(t_end/t_save)+1))
t_vec = np.zeros(round(t_end/t_save)+1)
A_arr[:,0] = A
dz = mesh_fourier(0,L,N)
dz2 = np.dot(dz,dz)
counter = 0
counter_save = 0
t = 0
for i in range(round(t_end/dt)+1):
counter += 1
# solve reaction equations
dy = R_nonbinding_5eq(y,t)
# evolve species profiles according to reaction + diffusion
A += dt*( dy[0] + D*np.dot(dz2,A) ) # dA/dt = R(A,a,aA,Ph,PhA) + D * dA^2/dz^2
a += dt*( dy[1] + D*np.dot(dz2,a) )
aA += dt*( dy[2] + D*np.dot(dz2,aA) )
Ph += dt*( dy[3] + D*np.dot(dz2,Ph) )
PhA += dt*( dy[4] + D*np.dot(dz2,PhA) )
y = A, a, aA, Ph, PhA
t += dt
if counter == round(t_save/dt):
counter = 0
counter_save += 1
A_arr[:,counter_save] = A
t_vec[counter_save] = t
arrays = A_arr,t_vec
y = A, a, aA, Ph, PhA
# output saved data arrays + last concentration profile variable in case extension is desired
return arrays, y
# # effect of diffusion speed on traveling front behavior
# +
"""
parameters used for the set of simulations below
"""
Ds = [
10**-2,
10**-3,
10**-4,
10**-5,
10**-6,
10**-7,
10**-8
]
t_ends = [
60*60,
160*60,
260*60,
600*60,
1000*60,
1000*60,
1000*60
]
dts = [
0.025,
0.1,
0.25,
0.25,
0.25,
0.25,
0.25
]
t_save = 60
N = 480
L = 20
x_span = np.linspace(-L/2,L/2,N)
# +
"""
simulates set of ~1.7 um perturbations with 0.55 uM phosphatase + varying diffusion coefficients
- perturbation width chosen to be identical to the chromosomal binding profile width
"""
start = time.time()
## algorithm takes ~16 min
# actual simulated perturbation width = (pw+1) * L / (N-1) = 9*500/399 ~ 11.3 um
pw = 40
print(f'actual pw: {round(x_span[int(N/2+pw/2)]*2,1)} um')
# phosphatase concentration: 0.55 uM
idx = 17
print(f'P = {lo_ss_nbns[idx][0]} uM')
perturbs_P055_varyDs = []
for D,t_end,dt in zip(Ds,t_ends,dts):
print(f'D: {D:.0E} um^2/sec')
arrays, y = spatial_simulation_nonbind_ss_perturb(lo_ss_nbns[idx],hi_ss_nbns[idx],t_end,dt,t_save,L,N,pw)
perturbs_P055_varyDs.append(arrays)
pickle.dump(perturbs_P055_varyDs,open('perturbs_P055_varyDs','wb'))
end = time.time()
print(f'~ {round( (end - start)/60, 1 )} min')
# +
"""
Figure 14 + Supplemental 16
plots simulation results with a 20 min separation between each spatial profile
- D: 1E-02 um^2/s : no traveling front develops
- D: 1E-03 um^2/s : wide traveling front develops
- traveling front narrows/sharpens/slows as diffusion speed increases
- D: 1E-08 um^2/s : front is effectively stationary
"""
perturbs_P055_varyDs = pickle.load(open('perturbs_P055_varyDs','rb'))
for arrays,D,t_end in zip(perturbs_P055_varyDs,Ds,t_ends):
A_arr, t_vec = arrays
lin_range = range(0,int(t_end/60+1),20)
colors = sns.color_palette('viridis', n_colors=len(lin_range))
for n,i in enumerate(lin_range):
# send first + last spatial profiles to the legend
if i == 0 or i == lin_range[-1]:
plt.plot(x_span,A_arr[:,i],color=colors[n], label=f'{i} min')
else:
plt.plot(x_span,A_arr[:,i],color=colors[n])
plt.xlabel("Distance (\u03BCm)")
plt.ylabel("[ABKp] (\u03BCM)")
plt.title(f'D = {D:.0E} \u03BCm^2/s')
plt.ylim(0,6)
plt.xlim(-3,3)
plt.legend()
plt.show()
# -
# # traveling front + localization behavior with reduced diffusion speed
def R_binding(y):
"""
system of ODEs from Zaytsev 2016 where kinase binding to sites along a chromosome is included,
previously defined a/A/aA/PhA components involve a diffusible/unbound Aurora B kinase,
the following components are introduced in this model:
- b: inactive/bound Aurora B kinase
- B: active/bound Aurora B kinase
- aB/bA/bB: enzyme-substrate complexes of inactive/active/bound/diffusible Aurora B kinase
- PhB: enzyme-substrate complex of phosphatase + active/bound Aurora B kinase
- BS: free binding sites
"""
A, B, a, b, aA, aB, bA, bB, Ph, PhA, PhB, BS = y
dAdt = (kcis - kfa*A)*a + (kra + 2*kca)*aA - kfp*A*Ph + krp*PhA +\
kca*aB - kfa*A*b + (kra + kca)*bA + koff*B - kon*BS*A
dBdt = (kcis - kfb*B)*b + (krb + 2*kca)*bB - kfp*B*Ph + krp*PhB +\
kca*bA - kfa*B*a + (kra + kca)*aB - koff*B + kon*BS*A
dadt = -(kcis + kfa*A + kfa*B)*a + kra*(aA + aB) + kcp*PhA + koff*b - kon*BS*a
dbdt = -(kcis + kfb*B + kfa*A)*b + krb*bB + kra*bA + kcp*PhB - koff*b + kon*BS*a
daAdt = kfa*a*A - (kra + kca)*aA
daBdt = kfa*a*B - (kra + kca)*aB
dbAdt = kfa*b*A - (kra + kca)*bA
dbBdt = kfb*b*B - (krb + kca)*bB
dPhdt = -kfp*Ph*(A + B) + (krp + kcp)*(PhA + PhB)
dPhAdt = kfp*Ph*A - (krp + kcp)*PhA
dPhBdt = kfp*Ph*B - (krp + kcp)*PhB
return dAdt, dBdt, dadt, dbdt, daAdt, daBdt, dbAdt, dbBdt, dPhdt, dPhAdt, dPhBdt
# +
"""
additional parameters for the binding model
previously defined parameters for the nonbinding model still apply
"""
# sterically limited b*B in trans autoactivation
kfb = kfa * 0.01 # 1/(uM*s)
Kmb = Kma * 100 # uM
kra = kfa*Kma-kca # 1/s
krb = kfb*Kma-kca # 1/s
# rate constants for kinase binding + dissociating from binding sites
kon = 2.9 # 1/(uM*s)
koff = 0.014 # 1/s
# -
def spatial_simulation_binding(A0,Atot,P0,t_end,dt,t_save,L,N,BSmin,BSw):
"""
reaction-diffusion algorithm with added binding/localization reactions
available chromosomal binding sites are situated in a Gaussian-like distribution
initial kinase/phosphatase conditions are constant across domain
"""
# initilize conditions for each reacting species
# flat initial kinase + phosphatase profiles
A = np.zeros(N) + A0
B = np.zeros(N)
a = np.zeros(N) + Atot - A
b = np.zeros(N)
aA = np.zeros(N)
aB = np.zeros(N)
bA = np.zeros(N)
bB = np.zeros(N)
Ph = np.zeros(N) + P0
PhA = np.zeros(N)
PhB = np.zeros(N)
x = np.linspace(-L/2, L/2, N)
# initialize binding site profile as a Gaussian distribution
BS0 = (Atot - BSmin) * np.exp(- Atot / BSw * x**2) + BSmin
y = A, B, a, b, aA, aB, bA, bB, Ph, PhA, PhB, BS0
A_arr = np.zeros((N,round(t_end/t_save)+1))
B_arr = np.zeros((N,round(t_end/t_save)+1))
a_arr = np.zeros((N,round(t_end/t_save)+1))
b_arr = np.zeros((N,round(t_end/t_save)+1))
aA_arr = np.zeros((N,round(t_end/t_save)+1))
aB_arr = np.zeros((N,round(t_end/t_save)+1))
bA_arr = np.zeros((N,round(t_end/t_save)+1))
bB_arr = np.zeros((N,round(t_end/t_save)+1))
Ph_arr = np.zeros((N,round(t_end/t_save)+1))
PhA_arr = np.zeros((N,round(t_end/t_save)+1))
PhB_arr = np.zeros((N,round(t_end/t_save)+1))
BS_arr = np.zeros((N,round(t_end/t_save)+1))
t_vec = np.zeros(round(t_end/t_save)+1)
# pulls non-zero concentration profiles to the saved data arrays
A_arr[:,0] = A
a_arr[:,0] = a
Ph_arr[:,0] = Ph
BS_arr[:,0] = BS0
dz = mesh_fourier(0,L,N)
dz2 = np.dot(dz,dz)
counter = -1
counter_save = 0
t = 0
for i in range(round(t_end/dt)+1):
counter += 1
dy = R_binding(y)
A += dt*(dy[0] + D*np.dot(dz2,A))
B += dt*dy[1]
a += dt*(dy[2] + D*np.dot(dz2,a))
b += dt*dy[3]
aA += dt*(dy[4] + D*np.dot(dz2,aA))
aB += dt*dy[5]
bA += dt*dy[6]
bB += dt*dy[7]
Ph += dt*(dy[8] + D*np.dot(dz2,Ph))
PhA += dt*(dy[9] + D*np.dot(dz2,PhA))
PhB += dt*dy[10]
# calculate binding site profile via mass balance
BS = BS0 - B - b - aB - bA - 2*bB - PhB
y = A, B, a, b, aA, aB, bA, bB, Ph, PhA, PhB, BS
t += dt
if counter == round(t_save/dt):
counter = 0
counter_save += 1
A_arr[:,counter_save] = A
B_arr[:,counter_save] = B
a_arr[:,counter_save] = a
b_arr[:,counter_save] = b
aA_arr[:,counter_save] = aA
aB_arr[:,counter_save] = aB
bA_arr[:,counter_save] = bA
bB_arr[:,counter_save] = bB
Ph_arr[:,counter_save] = Ph
PhA_arr[:,counter_save] = PhA
PhB_arr[:,counter_save] = PhB
BS_arr[:,counter_save] = BS
t_vec[counter_save] = t
arrays = A_arr, B_arr, a_arr, b_arr, aA_arr, aB_arr, bA_arr, bB_arr, Ph_arr, PhA_arr, PhB_arr, BS_arr, t_vec
y = A, B, a, b, aA, aB, bA, bB, Ph, PhA, PhB, BS
return arrays, y
# +
"""
constructs/plots kinase chromosomal binding site profile to be used for simulations below
"""
N = 500
L = 10
x_span = np.linspace(-L/2,L/2,N)
# scales initial binding site profile to approximate Figure 6 - Supplement 1C, Zaytsev, 2016
Atot = 10
BSmin = 1.5
BSw = 1.5
BS0 = (Atot - BSmin) * np.exp(- Atot / BSw * x_span**2) + BSmin
plt.plot(x_span,BS0)
plt.ylim(0,13)
plt.xlim(-3,3)
plt.ylabel('[Binding Sites] (\u03BCM)')
plt.xlabel('Distance (\u03BCm)');
# +
"""
parameters for the simulation below
"""
Ds = [
10**-4,
10**-5,
10**-6
]
t_end = 2000*60
dt = 0.035
t_save = 60
A0 = 0
P0 = 0.55
# +
"""
simulates reaction/diffusion/localization with 0.55 uM phosphatase + varying diffusion coefficients
total kinase kept at 10 uM across domain length, initial active kinase at zero
binding site profile used is plotted above
"""
start = time.time()
## algorithm takes >2 hrs
bindingsims_P055_varyDs = []
for D in Ds:
print(f'D: {D:.0E} um^2/sec')
arrays, y = spatial_simulation_binding(A0,Atot,P0,t_end,dt,t_save,L,N,BSmin,BSw)
bindingsims_P055_varyDs.append(arrays)
pickle.dump(bindingsims_P055_varyDs,open('bindingsims_P055_varyDs','wb'))
end = time.time()
print(f'~ {round( (end - start)/60, 1 )} min')
# +
"""
Figure 15 + Supplemental Figure 17
plots simulation results in two ways:
- time evolution of spatial profiles with a 50 min separation between profiles
- spatiotemporal heatmap
- D: 1E-04 um^2/s
- localization force towards the Gaussian peak in the center of the chromosome pulls kinase,
concentrating active kinase until autoactivation
- linearly progressing traveling front develops outward along chromosomes
- D: 1E-05 um^2/s
- stalling behavior emerges between autoactivation + traveling front progression
- shown by the increased density in spatial profiles at the centromere's boundaries
- shown also in the heatmap by the shallow slope around 250 - 1250 minutes
- represents a transient minimum in net force behind traveling front
- D: 1E-06 um^2/s
- autoactivation but no traveling front develops, pinned to the chromosomal binding sites
"""
bindingsims_P055_varyDs = pickle.load(open('bindingsims_P055_varyDs','rb'))
# first simulation limited to first 500 min
t_ends = [
500,
2000,
2000
]
for arrays,D,t_end in zip(bindingsim_P055_varyDs,Ds,t_ends):
print(f'D: {D:.0E} um^2/sec')
A_arr, B_arr, a_arr, b_arr, aA_arr, aB_arr, bA_arr, bB_arr, Ph_arr, PhA_arr, PhB_arr, BS_arr, t_vec = arrays
lin_range = range(0,t_end+1,50)
colors = sns.color_palette('viridis', n_colors=len(lin_range))
for n,i in enumerate(lin_range):
if i == 0 or i == t_end:
plt.plot(x_span, A_arr[:,i]+B_arr[:,i], color=colors[n], label=f'{i} min')
else:
plt.plot(x_span, A_arr[:,i]+B_arr[:,i], color=colors[n])
plt.ylim(0,13)
plt.xlim(-3,3)
plt.ylabel('[ABKp] (\u03BCm)')
plt.xlabel('Distance (\u03BCm)')
plt.legend()
plt.show()
# uses pcolormesh() to plot spatial profiles along the y-axis, evolving through time on the x-axis
heatmap = plt.pcolormesh(A_arr[:,:t_end]+B_arr[:,:t_end])
cbar = plt.colorbar(heatmap)
cbar.ax.set_title('[ABKp] (\u03BCM)')
# changes tich marks from spatial discretization points to distance in micrometers
plt.yticks(np.linspace(0,500,7), np.arange(-3, 4, 1))
plt.ylabel('Distance (\u03BCm)')
plt.xlabel('Time (min)')
plt.show()
# -
| 6_4_traveling_fronts_mass_action_model_reduced_diffusion_speed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Stats Quality for 2016 D-I College Nationals
#
# As one of the biggest tournaments hosted by USAU, the D-I College Nationals is one of the few tournaments where player statistics are relatively reliably tracked. For each tournament game, each player's aggregate scores, assists, Ds, and turns are counted, although its quite possible the definition of a "D" or a "Turn" could differ across stat-keepers.
#
# Data below was scraped from the [USAU website](http://play.usaultimate.org/events/USA-Ultimate-D-I-College-Championships/). First we'll set up some imports to be able to load this data.
# + run_control={"frozen": false, "read_only": false}
import usau.reports
import usau.fantasy
# + run_control={"frozen": false, "read_only": false}
from IPython.display import display, HTML
import pandas as pd
pd.options.display.width = 200
pd.options.display.max_colwidth = 200
pd.options.display.max_columns = 200
# + run_control={"frozen": false, "read_only": false}
def display_url_column(df):
"""Helper for formatting url links"""
df.url = df.url.apply(lambda url: "<a href='{base}{url}'>Match Report Link</a>"
.format(base=usau.reports.USAUResults.BASE_URL, url=url))
display(HTML(df.to_html(escape=False)))
# -
# Since we should already have the data downloaded as csv files in this repository, we will not need to re-scrape the data. Omit this cell to directly download from the USAU website (may be slow).
# + run_control={"frozen": false, "read_only": false}
# Read data from csv files
usau.reports.d1_college_nats_men_2016.load_from_csvs()
usau.reports.d1_college_nats_women_2016.load_from_csvs()
# -
# Let's take a look at the games for which the sum of the player goals/assists is less than the final score of the game:
# + run_control={"frozen": false, "read_only": false}
display_url_column(pd.concat([usau.reports.d1_college_nats_men_2016.missing_tallies,
usau.reports.d1_college_nats_women_2016.missing_tallies])
[["Score", "Gs", "As", "Ds", "Ts", "Team", "Opponent", "url"]])
# -
# All in all, not too bad! A few of the women's consolation games are missing player statistics, and there are several other games for which a couple of goals or assists were missed. For missing assists, it is technically possible that there were one or more callahans scored in those game, but obviously that's not the case with all ~14 missing assists. Surprisingly, there were 10 more assists recorded by the statkeepers than goals; I would have guessed that assists would be harder to keep track.
# Turns and Ds are the other stats available. In past tournaments these haven't been tracked very well, but actually there was only one game where no Turns or Ds were recorded:
# + run_control={"frozen": false, "read_only": false}
men_matches = usau.reports.d1_college_nats_men_2016.match_results
women_matches = usau.reports.d1_college_nats_women_2016.match_results
display_url_column(pd.concat([men_matches[(men_matches.Ts == 0) & (men_matches.Gs > 0)],
women_matches[(women_matches.Ts == 0) & (women_matches.Gs > 0)]])
[["Score", "Gs", "As", "Ds", "Ts", "Team", "Opponent", "url"]])
# -
# This implies that there was a pretty good effort made to keep up with counting turns and Ds. By contrast, see how many teams did not keep track of Ds and turns last year (2015)!
# + run_control={"frozen": false, "read_only": false}
# Read last year's data from csv files
usau.reports.d1_college_nats_men_2015.load_from_csvs()
usau.reports.d1_college_nats_women_2015.load_from_csvs()
display_url_column(pd.concat([usau.reports.d1_college_nats_men_2015.missing_tallies,
usau.reports.d1_college_nats_women_2015.missing_tallies])
[["Score", "Gs", "As", "Ds", "Ts", "Team", "Opponent", "url"]])
# + run_control={"frozen": false, "read_only": false}
| notebooks/2016-D-I_College_Nationals_Data_Quality.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hn9rVSpNNJcq"
# <a href="https://colab.research.google.com/github/https-deeplearning-ai/tensorflow-1-public/blob/master/C2/W3/ungraded_lab/C2_W3_Lab_1_transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bT0to3TL2q7H"
# # Ungraded Lab: Transfer Learning
#
# In this lab, you will see how you can use a pre-trained model to achieve good results even with a small training dataset. This is called _transfer learning_ and you do this by leveraging the trained layers of an existing model and adding your own layers to fit your application. For example, you can:
#
# 1. just get the convolution layers of one model
# 2. attach some dense layers onto it
# 3. train just the dense network
# 4. evaluate the results
#
# Doing this will allow you to save time building your application because you will essentially skip weeks of training time of very deep networks. You will just use the features it has learned and tweak it for your dataset. Let's see how these are done in the next sections.
# + [markdown] id="Qvrr8pLRzJMV"
# **IMPORTANT NOTE:** This notebook is designed to run as a Colab. Running the notebook on your local machine might result in some of the code blocks throwing errors.
# + [markdown] id="-12slkPL6_JH"
# ## Setup the pretrained model
#
# You will need to prepare pretrained model and configure the layers that you need. For this exercise, you will use the convolution layers of the [InceptionV3](https://arxiv.org/abs/1512.00567) architecture as your base model. To do that, you need to:
#
# 1. Set the input shape to fit your application. In this case. set it to `150x150x3` as you've been doing in the last few labs.
#
# 2. Pick and freeze the convolution layers to take advantage of the features it has learned already.
#
# 3. Add dense layers which you will train.
#
# Let's see how to do these in the next cells.
# + [markdown] id="3VqhFEK2Y-PK"
# First, in preparing the input to the model, you want to fetch the pretrained weights of the `InceptionV3` model and remove the fully connected layer at the end because you will be replacing it later. You will also specify the input shape that your model will accept. Lastly, you want to freeze the weights of these layers because they have been trained already.
# + id="1xJZ5glPPCRz" colab={"base_uri": "https://localhost:8080/"} outputId="5eb7cbed-a5af-4442-b72e-4eb405549865"
# Download the pre-trained weights. No top means it excludes the fully connected layer it uses for classification.
# !wget --no-check-certificate \
# https://storage.googleapis.com/mledu-datasets/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5 \
# -O /tmp/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5
# + id="KsiBCpQ1VvPp"
from tensorflow.keras.applications.inception_v3 import InceptionV3
from tensorflow.keras import layers
# Set the weights file you downloaded into a variable
local_weights_file = '/tmp/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5'
# Initialize the base model.
# Set the input shape and remove the dense layers.
pre_trained_model = InceptionV3(input_shape = (150, 150, 3),
include_top = False,
weights = None)
# Load the pre-trained weights you downloaded.
pre_trained_model.load_weights(local_weights_file)
# Freeze the weights of the layers.
for layer in pre_trained_model.layers:
layer.trainable = False
# + [markdown] id="1y2rEnqFaa9k"
# You can see the summary of the model below. You can see that it is a very deep network. You can then select up to which point of the network you want to use. As Laurence showed in the exercise, you will use up to `mixed_7` as your base model and add to that. This is because the original last layer might be too specialized in what it has learned so it might not translate well into your application. `mixed_7` on the other hand will be more generalized and you can start with that for your application. After the exercise, feel free to modify and use other layers to see what the results you get.
# + id="qeGP0Ust5kCR" colab={"base_uri": "https://localhost:8080/"} outputId="9d9fb6a5-09c4-46bc-830b-55d59292416e"
pre_trained_model.summary()
# + id="jDmGO9tg5iPc" colab={"base_uri": "https://localhost:8080/"} outputId="fc2ee2e8-95ff-4548-a9f4-deb263b47e4c"
# Choose `mixed_7` as the last layer of your base model
last_layer = pre_trained_model.get_layer('mixed7')
print('last layer output shape: ', last_layer.output_shape)
last_output = last_layer.output
# + [markdown] id="UXT9SDMK7Ioa"
# ## Add dense layers for your classifier
#
# Next, you will add dense layers to your model. These will be the layers that you will train and is tasked with recognizing cats and dogs. You will add a [Dropout](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dropout) layer as well to regularize the output and avoid overfitting.
# + id="BMXb913pbvFg" colab={"base_uri": "https://localhost:8080/"} outputId="b15351fd-33b5-4004-9572-50fcc80e2a76"
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras import Model
# Flatten the output layer to 1 dimension
x = layers.Flatten()(last_output)
# Add a fully connected layer with 1,024 hidden units and ReLU activation
x = layers.Dense(1024, activation='relu')(x)
# Add a dropout rate of 0.2
x = layers.Dropout(0.2)(x)
# Add a final sigmoid layer for classification
x = layers.Dense (1, activation='sigmoid')(x)
# Append the dense network to the base model
model = Model(pre_trained_model.input, x)
# Print the model summary. See your dense network connected at the end.
model.summary()
# + id="SAwTTkWr56uC"
# Set the training parameters
model.compile(optimizer = RMSprop(learning_rate=0.0001),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
# + [markdown] id="aYLGw_RO7Z_X"
# ## Prepare the dataset
#
# Now you will prepare the dataset. This is basically the same code as the one you used in the data augmentation lab.
# + id="O4s8HckqGlnb" colab={"base_uri": "https://localhost:8080/"} outputId="4e2eafea-338d-443f-dbbe-b9de6516354a"
# Download the dataset
# !wget https://storage.googleapis.com/tensorflow-1-public/course2/cats_and_dogs_filtered.zip
# + id="WOV8jON3c3Jv" colab={"base_uri": "https://localhost:8080/"} outputId="744a51d4-c27c-49f5-b48d-fc8d2e9277ff"
import os
import zipfile
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# Extract the archive
zip_ref = zipfile.ZipFile("./cats_and_dogs_filtered.zip", 'r')
zip_ref.extractall("tmp/")
zip_ref.close()
# Define our example directories and files
base_dir = 'tmp/cats_and_dogs_filtered'
train_dir = os.path.join( base_dir, 'train')
validation_dir = os.path.join( base_dir, 'validation')
# Directory with training cat pictures
train_cats_dir = os.path.join(train_dir, 'cats')
# Directory with training dog pictures
train_dogs_dir = os.path.join(train_dir, 'dogs')
# Directory with validation cat pictures
validation_cats_dir = os.path.join(validation_dir, 'cats')
# Directory with validation dog pictures
validation_dogs_dir = os.path.join(validation_dir, 'dogs')
# Add our data-augmentation parameters to ImageDataGenerator
train_datagen = ImageDataGenerator(rescale = 1./255.,
rotation_range = 40,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
# Note that the validation data should not be augmented!
test_datagen = ImageDataGenerator( rescale = 1.0/255. )
# Flow training images in batches of 20 using train_datagen generator
train_generator = train_datagen.flow_from_directory(train_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (150, 150))
# Flow validation images in batches of 20 using test_datagen generator
validation_generator = test_datagen.flow_from_directory( validation_dir,
batch_size = 20,
class_mode = 'binary',
target_size = (150, 150))
# + [markdown] id="3m3S6AZb7h-B"
# ## Train the model
#
# With that, you can now train the model. You will do 20 epochs and plot the results afterwards.
# + id="Blhq2MAUeyGA" colab={"base_uri": "https://localhost:8080/"} outputId="16319b72-e579-40df-a0db-d3714576c03c"
# Train the model.
history = model.fit(
train_generator,
validation_data = validation_generator,
steps_per_epoch = 100,
epochs = 20,
validation_steps = 50,
verbose = 1)
# + [markdown] id="RwcB2bPj7lIx"
# ## Evaluate the results
#
# You will use the same code to plot the results. As you can see, the validation accuracy is also trending upwards as your training accuracy improves. This is a good sign that your model is no longer overfitting!
# + id="C2Fp6Se9rKuL" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="142d6d10-8c76-4ec4-ae46-2678e93840e3"
import matplotlib.pyplot as plt
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend(loc=0)
plt.figure()
plt.show()
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 543} id="_sDF7xtPXtH2" outputId="ec366bbe-558e-4525-e0a0-0b61b7a195ae"
import numpy as np
from google.colab import files
from keras.preprocessing import image
uploaded = files.upload()
for fn in uploaded.keys():
# predicting images
path = '/content/' + fn
img = image.load_img(path, target_size=(150, 150))
x = image.img_to_array(img)
x /= 255
x = np.expand_dims(x, axis=0)
images = np.vstack([x])
classes = model.predict(images, batch_size=10)
print(classes[0])
if classes[0]>0.5:
print(fn + " is a dogs")
else:
print(fn + " is a cats")
# + id="KMuWmRyPYU79"
| Convolutional Neural Networks in TensorFlow/C2_W3_Lab_1_transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Winery classification with the bivariate Gaussian
#
# Our first generative model for Winery classification used just one feature. Now we use two features, modeling each class by a **bivariate Gaussian**.
# ## 1. Load in the data set
# As in the univariate case, we start by loading in the Wine data set. Make sure the file `wine.data.txt` is in the same directory as this notebook.
#
# Recall that there are 178 data points, each with 13 features and a label (1,2,3). As before, we will divide this into a training set of 130 points and a test set of 48 points.
# Standard includes
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Useful module for dealing with the Gaussian density
from scipy.stats import norm, multivariate_normal
# installing packages for interactive graphs
import ipywidgets as widgets
from IPython.display import display
from ipywidgets import interact, interactive, fixed, interact_manual, IntSlider
# Load data set.
data = np.loadtxt('wine.data.txt', delimiter=',')
# Names of features
featurenames = ['Alcohol', 'Malic acid', 'Ash', 'Alcalinity of ash','Magnesium', 'Total phenols',
'Flavanoids', 'Nonflavanoid phenols', 'Proanthocyanins', 'Color intensity', 'Hue',
'OD280/OD315 of diluted wines', 'Proline']
# Split 178 instances into training set (trainx, trainy) of size 130 and test set (testx, testy) of size 48
np.random.seed(0)
perm = np.random.permutation(178)
trainx = data[perm[0:130],1:14]
trainy = data[perm[0:130],0]
testx = data[perm[130:178], 1:14]
testy = data[perm[130:178],0]
# ## 2. Look at the distribution of two features from one of the wineries
# Our goal is to plot the distribution of two features from a particular winery. We will use several helper functions for this. It is worth understanding each of these.
# The first helper function fits a Gaussian to a data set, restricting attention to specified features.
# It returns the mean and covariance matrix of the Gaussian.
# Fit a Gaussian to a data set using the selected features
def fit_gaussian(x, features):
mu = np.mean(x[:,features], axis=0)
covar = np.cov(x[:,features], rowvar=0, bias=1)
return mu, covar
# For example, let's look at the Gaussian we get for winery 1, using features 0 ('alcohol') and 6 ('flavanoids').
f1 = 0
f2 = 6
label = 1
mu, covar = fit_gaussian(trainx[trainy==label,:], [f1,f2])
print("Mean:\n" + str(mu))
print("Covariance matrix:\n" + str(covar))
# Next, we will construct a routine for displaying points sampled from a two-dimensional Gaussian, as well as a few contour lines. Part of doing this involves deciding what range to use for each axis. We begin with a little helper function that takes as input an array of numbers (values along a single feature) and returns the range in which these numbers lie.
# Find the range within which an array of numbers lie, with a little buffer
def find_range(x):
lower = min(x)
upper = max(x)
width = upper - lower
lower = lower - 0.2 * width
upper = upper + 0.2 * width
return lower, upper
# Next we define a routine that plots a few contour lines of a given two-dimensional Gaussian.
# It takes as input:
# * `mu`, `cov`: the parameters of the Gaussian
# * `x1g`, `x2g`: the grid (along the two axes) at which the density is to be computed
# * `col`: the color of the contour lines
def plot_contours(mu, cov, x1g, x2g, col):
rv = multivariate_normal(mean=mu, cov=cov)
z = np.zeros((len(x1g),len(x2g)))
for i in range(0,len(x1g)):
for j in range(0,len(x2g)):
z[j,i] = rv.logpdf([x1g[i], x2g[j]])
sign, logdet = np.linalg.slogdet(cov)
normalizer = -0.5 * (2 * np.log(6.28) + sign * logdet)
for offset in range(1,4):
plt.contour(x1g,x2g,z, levels=[normalizer - offset], colors=col, linewidths=2.0, linestyles='solid')
# The function **two_features_plot** takes an input two features and a label, and displays the distribution for the specified winery and pair of features.
#
# The first line allows you to specify the parameters interactively using sliders.
@interact_manual( f1=IntSlider(0,0,12,1), f2=IntSlider(6,0,12,1), label=IntSlider(1,1,3,1) )
def two_features_plot(f1,f2,label):
if f1 == f2: # we need f1 != f2
print("Please choose different features for f1 and f2.")
return
# Set up plot
x1_lower, x1_upper = find_range(trainx[trainy==label,f1])
x2_lower, x2_upper = find_range(trainx[trainy==label,f2])
plt.xlim(x1_lower, x1_upper) # limit along x1-axis
plt.ylim(x2_lower, x2_upper) # limit along x2-axis
# Plot the training points along the two selected features
plt.plot(trainx[trainy==label, f1], trainx[trainy==label, f2], 'ro')
# Define a grid along each axis; the density will be computed at each grid point
res = 200 # resolution
x1g = np.linspace(x1_lower, x1_upper, res)
x2g = np.linspace(x2_lower, x2_upper, res)
# Now plot a few contour lines of the density
mu, cov = fit_gaussian(trainx[trainy==label,:], [f1,f2])
plot_contours(mu, cov, x1g, x2g, 'k')
# Finally, display
plt.xlabel(featurenames[f1], fontsize=14, color='red')
plt.ylabel(featurenames[f2], fontsize=14, color='red')
plt.title('Class ' + str(label), fontsize=14, color='blue')
plt.show()
# ## 3. Fit a Gaussian to each class
# We now define a function that will fit a Gaussian generative model to the three classes, restricted to a given list of features. The function returns:
# * `mu`: the means of the Gaussians, one per row
# * `covar`: covariance matrices of each of the Gaussians
# * `pi`: list of three class weights summing to 1
# Assumes y takes on values 1,2,3
def fit_generative_model(x, y, features):
k = 3 # number of classes
d = len(features) # number of features
mu = np.zeros((k+1,d)) # list of means
covar = np.zeros((k+1,d,d)) # list of covariance matrices
pi = np.zeros(k+1) # list of class weights
for label in range(1,k+1):
indices = (y==label)
mu[label,:], covar[label,:,:] = fit_gaussian(x[indices,:], features)
pi[label] = float(sum(indices))/float(len(y))
return mu, covar, pi
# Now we will plot the three Gaussians.
@interact_manual( f1=IntSlider(0,0,12,1), f2=IntSlider(6,0,12,1) )
def three_class_plot(f1,f2):
if f1 == f2: # we need f1 != f2
print("Please choose different features for f1 and f2.")
return
# Set up plot
x1_lower, x1_upper = find_range(trainx[:,f1])
x2_lower, x2_upper = find_range(trainx[:,f2])
plt.xlim(x1_lower, x1_upper) # limit along x1-axis
plt.ylim(x2_lower, x2_upper) # limit along x2-axis
# Plot the training points along the two selected features
colors = ['r', 'k', 'g']
for label in range(1,4):
plt.plot(trainx[trainy==label,f1], trainx[trainy==label,f2], marker='o', ls='None', c=colors[label-1])
# Define a grid along each axis; the density will be computed at each grid point
res = 200 # resolution
x1g = np.linspace(x1_lower, x1_upper, res)
x2g = np.linspace(x2_lower, x2_upper, res)
# Show the Gaussian fit to each class, using features f1,f2
mu, covar, pi = fit_generative_model(trainx, trainy, [f1,f2])
for label in range(1,4):
gmean = mu[label,:]
gcov = covar[label,:,:]
plot_contours(gmean, gcov, x1g, x2g, colors[label-1])
# Finally, display
plt.xlabel(featurenames[f1], fontsize=14, color='red')
plt.ylabel(featurenames[f2], fontsize=14, color='red')
plt.title('Wine data', fontsize=14, color='blue')
plt.show()
# ## 4. Predict labels for the test points
# How well we can predict the class (1,2,3) based just on these two features?
#
# We start with a testing procedure that is analogous to what we developed in the 1-d case.
# Now test the performance of a predictor based on a subset of features
@interact( f1=IntSlider(0,0,12,1), f2=IntSlider(6,0,12,1) )
def test_model(f1, f2):
if f1 == f2: # need f1 != f2
print("Please choose different features for f1 and f2.")
return
features= [f1,f2]
mu, covar, pi = fit_generative_model(trainx, trainy, features)
k = 3 # Labels 1,2,...,k
nt = len(testy) # Number of test points
score = np.zeros((nt,k+1))
for i in range(0,nt):
for label in range(1,k+1):
score[i,label] = np.log(pi[label]) + \
multivariate_normal.logpdf(testx[i,features], mean=mu[label,:], cov=covar[label,:,:])
predictions = np.argmax(score[:,1:4], axis=1) + 1
# Finally, tally up score
errors = np.sum(predictions != testy)
print("Test error using feature(s): ")
for f in features:
print("'" + featurenames[f] + "'" + " ")
print("Errors: " + str(errors) + "/" + str(nt))
# Now test the performance of a predictor based on a subset of features
# ### <font color="magenta">Fast exercise 1</font>
# Different pairs of features yield different test errors.
# * What is the smallest achievable test error?--> 3
# * Which pair of features achieves this minimum test error? --> (9,6)
#
# *Make a note of your answers to these questions, as you will need to enter them as part of this week's assignment.*
# ## 5. The decision boundary
# The function **show_decision_boundary** takes as input two features, builds a classifier based only on these two features, and shows a plot that contains both the training data and the decision boundary.
#
# To compute the decision boundary, a dense grid is defined on the two-dimensional input space and the classifier is applied to every grid point. The built-in `pyplot.contour` function can then be invoked to depict the boundary.
@interact( f1=IntSlider(0,0,12,1), f2=IntSlider(6,0,12,1) )
def show_decision_boundary(f1,f2):
# Fit Gaussian to each class
mu, covar, pi = fit_generative_model(trainx, trainy, [f1,f2])
# Set up dimensions of plot
x1_lower, x1_upper = find_range(trainx[:,f1])
x2_lower, x2_upper = find_range(trainx[:,f2])
plt.xlim([x1_lower,x1_upper])
plt.ylim([x2_lower,x2_upper])
# Plot points in training set
colors = ['r', 'k', 'g']
for label in range(1,4):
plt.plot(trainx[trainy==label,f1], trainx[trainy==label,f2], marker='o', ls='None', c=colors[label-1])
# Define a dense grid; every point in the grid will be classified according to the generative model
res = 200
x1g = np.linspace(x1_lower, x1_upper, res)
x2g = np.linspace(x2_lower, x2_upper, res)
# Declare random variables corresponding to each class density
random_vars = {}
for label in range(1,4):
random_vars[label] = multivariate_normal(mean=mu[label,:],cov=covar[label,:,:])
# Classify every point in the grid; these are stored in an array Z[]
Z = np.zeros((len(x1g), len(x2g)))
for i in range(0,len(x1g)):
for j in range(0,len(x2g)):
scores = []
for label in range(1,4):
scores.append(np.log(pi[label]) + random_vars[label].logpdf([x1g[i],x2g[j]]))
Z[i,j] = np.argmax(scores) + 1
# Plot the contour lines
plt.contour(x1g,x2g,Z.T,3,cmap='seismic')
# Finally, show the image
plt.xlabel(featurenames[f1], fontsize=14, color='red')
plt.ylabel(featurenames[f2], fontsize=14, color='red')
plt.show()
# Let's use the function above to draw the decision boundary using features 0 ('alcohol') and 6 ('flavanoids').
show_decision_boundary(0,6)
# ### <font color="magenta">Fast exercise 2</font>
# Can you add interactive sliders to function **show_decision_boundary**? Done
# ### <font color="magenta">Fast exercise 3</font>
# Produce a plot similar to that of **show_decision_boundary**, but in which just the **test** data is shown.
# Look back at your answer to *Fast exercise 1*. Is it corroborated by your plot? Are the errors clearly visible?
@interact( f1=IntSlider(0,0,12,1), f2=IntSlider(6,0,12,1) )
def show_decision_boundary_test(f1,f2):
# Fit Gaussian to each class
mu, covar, pi = fit_generative_model(testx, testy, [f1,f2])
# Set up dimensions of plot
x1_lower, x1_upper = find_range(testx[:,f1])
x2_lower, x2_upper = find_range(testx[:,f2])
plt.xlim([x1_lower,x1_upper])
plt.ylim([x2_lower,x2_upper])
# Plot points in test set
colors = ['r', 'k', 'g']
for label in range(1,4):
plt.plot(testx[testy==label,f1], testx[testy==label,f2], marker='o', ls='None', c=colors[label-1])
# Define a dense grid; every point in the grid will be classified according to the generative model
res = 200
x1g = np.linspace(x1_lower, x1_upper, res)
x2g = np.linspace(x2_lower, x2_upper, res)
# Declare random variables corresponding to each class density
random_vars = {}
for label in range(1,4):
random_vars[label] = multivariate_normal(mean=mu[label,:],cov=covar[label,:,:])
# Classify every point in the grid; these are stored in an array Z[]
Z = np.zeros((len(x1g), len(x2g)))
for i in range(0,len(x1g)):
for j in range(0,len(x2g)):
scores = []
for label in range(1,4):
scores.append(np.log(pi[label]) + random_vars[label].logpdf([x1g[i],x2g[j]]))
Z[i,j] = np.argmax(scores) + 1
# Plot the contour lines
plt.contour(x1g,x2g,Z.T,3,cmap='seismic')
# Finally, show the image
plt.xlabel(featurenames[f1], fontsize=14, color='red')
plt.ylabel(featurenames[f2], fontsize=14, color='red')
plt.show()
| Assignment 2/winery-bivariate/winery-classification-bivariate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### merge emr outputs in sorted order and dump in txt file
# +
import sys
from itertools import groupby
from operator import itemgetter
def read_mapper_output(file, separator='\t'):
for line in file:
yield line.rstrip().split(separator, 1)
def main(separator='\t'):
output_file=open("Twitter_Rugby/word_count_rugby.txt","a");
file =open("Twitter_Rugby/emr_rugby");
hashtable={}; # maintain dictionary <word,count> to store count of word
# input comes from STDIN (standard input)
data = read_mapper_output(file, separator=separator);
for current_word, group in groupby(data, itemgetter(0)):
try:
total_count = sum(int(count) for current_word, count in group)
if current_word not in hashtable:
hashtable[current_word] = total_count;
else:
hashtable[current_word] += total_count;
except ValueError:
# count was not a number, so silently discard this item
pass
sortedTuple = sorted(hashtable.items(), key=itemgetter(1),reverse=True); # sort hashtable by word occurence count
for tuples in sortedTuple:
line=tuples[0]+separator+str(tuples[1]);
output_file.write(line);
output_file.write("\n");
# print("%s%s%d" % (tuples[0], separator, tuples[1]));
output_file.close();
if __name__ == "__main__":
main()
# -
| part3/Twitter/Data/Twitter_MR_Output/.ipynb_checkpoints/merge-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-1.2 Intro Python
# ## Sequence: String
# - Accessing String Character with index
# - **Accessing sub-strings with index slicing**
# - Iterating through Characters of a String
# - More String Methods
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - Work with String Characters
# - **Slice strings into substrings**
# - Iterate through String Characters
# - Use String Methods
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Accessing sub-strings
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/251ad8c1-588b-47de-8638-a5bcd0f29800/Unit2_Section1.2a-Index_Slicing-Substrings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/251ad8c1-588b-47de-8638-a5bcd0f29800/Unit2_Section1.2a-Index_Slicing-Substrings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Index Slicing [start:stop]
# String slicing returns a string section by addressing the start and stop indexes
#
# ```python
# # assign string to student_name
# student_name = "Colette"
# # addressing the 3rd, 4th and 5th characters
# student_name[2:5]
# ```
# The slice starts at index 2 and ends at index 5 (but does not include index 5)
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# +
# [ ] review and run example
# assign string to student_name
student_name = "Colette"
# addressing the 3rd, 4th and 5th characters using a slice
print("slice student_name[2:5]:",student_name[2:5])
# +
# [ ] review and run example
# assign string to student_name
student_name = "Colette"
# addressing the 3rd, 4th and 5th characters individually
print("index 2, 3 & 4 of student_name:", student_name[2] + student_name[3] + student_name[4])
# -
# [ ] review and run example
long_word = 'Acknowledgement'
print(long_word[2:11])
print(long_word[2:11], "is the 3rd char through the 11th char")
print(long_word[2:11], "is the index 2, \"" + long_word[2] + "\",", "through index 10, \"" + long_word[10] + "\"")
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
#
# ## slice a string
# ### start & stop index
# [ ] slice long_word to print "act" and to print "tic"
long_word = "characteristics"
print(long_word[4:7])
print(long_word[11:14])
# [ ] slice long_word to print "sequence"
long_word = "Consequences"
print(long_word[3:11])
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Accessing beginning of sub-strings
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/368b352f-6061-488c-80a4-d75e455f4416/Unit2_Section1.2b-Index_Slicing_Beginnings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/368b352f-6061-488c-80a4-d75e455f4416/Unit2_Section1.2b-Index_Slicing_Beginnings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Index Slicing [:stop]
# String slicing returns a string section from index 0 by addressing only the stop index
#
# ```python
# student_name = "Colette"
# # addressing the 1st, 2nd & 3rd characters
# student_name[:3]
# ```
# **default start for a slice is index 0**
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
# [ ] review and run example
student_name = "Colette"
# addressing the 1st, 2nd & 3rd characters
print(student_name[:3])
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
#
# [ ] print the first half of the long_word
long_word = "Consequences"
print(long_word[:6])
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## Accessing ending of sub-strings
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/29beb75a-aee7-43df-9569-e9ad22cffac4/Unit2_Section1.2c-Index_Slicing_Endings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/29beb75a-aee7-43df-9569-e9ad22cffac4/Unit2_Section1.2c-Index_Slicing_Endings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Index Slicing [start:]
# String slicing returns a string section including by addressing only the start index
#
# ```python
# student_name = "Colette"
# # addressing the 4th, 5th and 6th characters
# student_name[3:]
# ```
# **default end index returns up to and including the last string character**
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
# [ ] review and run example
student_name = "Colette"
# 4th, 5th, 6th and 7th characters
student_name[3:]
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
#
# [ ] print the second half of the long_word
long_word = "Consequences"
print(long_word[6:])
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## accessing sub-strings by step size
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/62c65917-4979-4d26-9a05-09e1ed02cc51/Unit2_Section1.2d-Index_Slicing-Step_Sizes.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/62c65917-4979-4d26-9a05-09e1ed02cc51/Unit2_Section1.2d-Index_Slicing-Step_Sizes.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Index Slicing [:], [::2]
# - **[:]** returns the entire string
# - **[::2]** returns the first char and then steps to every other char in the string
# - **[1::3]** returns the second char and then steps to every third char in the string
#
# the number **2**, in the print statement below, represents the **step**
#
# ```python
# print(long_word[::2])
# ```
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# [ ] review and run example
student_name = "Colette"
# return all
print(student_name[:])
# [ ] review and run example
student_name = "Colette"
# return every other
print(student_name[::2])
# [ ] review and run example
student_name = "Colette"
# return every third, starting at 2nd character
print(student_name[1::2])
# [ ] review and run example
long_word = "Consequences"
# starting at 2nd char (index 1) to 9th character, return every other character
print(long_word[1:9:2])
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
#
# [ ] print the 1st and every 3rd letter of long_word
long_word = "Acknowledgement"
print(long_word[0::2])
# [ ] print every other character of long_word starting at the 3rd character
long_word = "Acknowledgement"
print(long_word[2::2])
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
#
# ## Accessing sub-strings continued
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/2e59f526-fadb-434e-822e-afe3732f75df/Unit2_Section1.2e-Index_Slicing-Reverse.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/2e59f526-fadb-434e-822e-afe3732f75df/Unit2_Section1.2e-Index_Slicing-Reverse.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### stepping backwards
#
# ```python
# print(long_word[::-1])
# ```
#
# use **[::-1]** to reverse a string
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
# [ ] review and run example of stepping backwards using [::-1]
long_word = "characteristics"
# make the step increment -1 to step backwards
print(long_word[::-1])
# [ ] review and run example of stepping backwards using [6::-1]
long_word = "characteristics"
# start at the 7th letter backwards to start
print(long_word[6::-1])
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 5</B></font>
# use slicing
# [ ] reverse long_word
long_word = "stressed"
print(long_word[::-1])
# [ ] print the first 5 letters of long_word in reverse
long_word = "characteristics"
print(long_word[4::-1])
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 6</B></font>
# use slicing
# +
# [ ] print the first 4 letters of long_word
# [ ] print the first 4 letters of long_word in reverse
# [ ] print the last 4 letters of long_word in reverse
# [ ] print the letters spanning indexes 3 to 6 of long_word in Reverse
long_word = "timeline"
print(long_word[:4])
print(long_word[3::-1])
print(long_word[:3:-1])
print(long_word[6:2:-1])
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Fundamentals/Module_1_2_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using `jit`
#
# We know how to find hotspots now, how do we improve their performance?
#
# We `jit` them!
#
# We'll start with a trivial example but get to some more realistic applications shortly.
# ### Array sum
#
# The function below is a naive `sum` function that sums all the elements of a given array.
def sum_array(inp):
J, I = inp.shape
#this is a bad idea
mysum = 0
for j in range(J):
for i in range(I):
mysum += inp[j, i]
return mysum
import numpy
arr = numpy.random.random((300, 300))
sum_array(arr)
# plain = %timeit -o sum_array(arr)
# # Let's get started
from numba import jit
# ## As a function call
sum_array_numba = jit()(sum_array)
# What's up with the weird double `()`s? We'll cover that in a little bit.
sum_array_numba(arr)
# jitted = %timeit -o sum_array_numba(arr)
plain.best / jitted.best
# ## (more commonly) As a decorator
@jit
def sum_array(inp):
I, J = inp.shape
mysum = 0
for i in range(I):
for j in range(J):
mysum += inp[i, j]
return mysum
sum_array(arr)
# %timeit sum_array(arr)
# ## How does this compare to NumPy?
# %timeit arr.sum()
# ## When does `numba` compile things?
# The first time you call the function.
# ## [Your turn!](./exercises/02.Intro.to.JIT.exercises.ipynb#JIT-Exercise)
| notebooks/02.Intro.to.jit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wel51x/DS-Unit-3-Sprint-2-SQL-and-Databases/blob/master/module3-nosql-and-document-oriented-databases/LS_DS2_MongoDB_Playground.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HIuHUD9N0q66" colab_type="text"
# # MongoDB with PyMongo
#
# LSDS Unit 3 Sprint 2 Module 3
#
# Some resources:
#
# https://docs.atlas.mongodb.com/getting-started/
#
# https://api.mongodb.com/python/current/
#
# HN Discussion on MongoDB versus PostgreSQL/SQLite: https://news.ycombinator.com/item?id=19158854
# + id="i5Xl7DJ30mWo" colab_type="code" outputId="673fceeb-93b1-4a60-9c9a-d254c9293e77" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !curl ipecho.net/plain
# + id="4Mn5zOkzE35E" colab_type="code" outputId="239c9dc6-bd1c-4019-ad91-744084e77326" colab={"base_uri": "https://localhost:8080/", "height": 54}
# !pip install pymongo
# + id="YhMw_j93E9Jx" colab_type="code" colab={}
import pymongo
# + id="jQER5Y6pDJLt" colab_type="code" colab={}
client = pymongo.MongoClient("mongodb://<USER>:<PASSWORD>@cluster0-shard-00-00-gydux.mongodb.net:27017,cluster0-shard-00-01-gydux.mongodb.net:27017,cluster0-shard-00-02-gydux.mongodb.net:27017/test?ssl=true&replicaSet=Cluster0-shard-0&authSource=admin&retryWrites=true")
db = client.test
# + id="Vtm0Q256IrVI" colab_type="code" outputId="4a0c6229-6878-4c71-ba6e-f93e79b60865" colab={"base_uri": "https://localhost:8080/", "height": 69}
client.nodes
# + id="KaNyhbXpF0hV" colab_type="code" outputId="6f35cbe5-9178-4881-ffb5-c3bab24e4d52" colab={"base_uri": "https://localhost:8080/", "height": 714}
help(db.test.insert_one)
# + id="ySdb830-H-Jx" colab_type="code" outputId="825f3ff8-0f0e-4e9f-e611-7f675b336a2d" colab={"base_uri": "https://localhost:8080/", "height": 34}
dmitriys_doc = {'favorite animal!!!': 'narwhal'}
db.test.insert_one(dmitriys_doc)
# + id="8PjiIieCJUaj" colab_type="code" outputId="ba780187-04ff-43c9-d51e-31c0aedcfbb4" colab={"base_uri": "https://localhost:8080/", "height": 34}
shilpas_doc = {'favorite_colors': ['black', 'white', 'red']}
db.test.insert_one(shilpas_doc)
# + id="NaI78qhKKu90" colab_type="code" outputId="738bd06c-0837-41cf-9461-878fbbfb2663" colab={"base_uri": "https://localhost:8080/", "height": 1158}
db.test.insert_one(dmitriys_doc) # Inserting again
# + id="AY-ioMnwLkvg" colab_type="code" colab={}
results = db.test.find()
# + id="wVUpc11bL2lD" colab_type="code" outputId="8aa70436-6dea-40f5-eece-775c8220e2ee" colab={"base_uri": "https://localhost:8080/", "height": 34}
results
# + id="k_kePY6JL7bh" colab_type="code" colab={}
results_list = list(results)
# + id="T2gWrJioL_Oh" colab_type="code" outputId="598caddb-3a56-4f9c-cff6-48afc13b1297" colab={"base_uri": "https://localhost:8080/", "height": 86}
results_list
# + id="-zqfOyeoMACJ" colab_type="code" outputId="96cbf079-1eee-4ef3-d7ec-eb9aea538f79" colab={"base_uri": "https://localhost:8080/", "height": 52}
db.test.find_one(shilpas_doc)
# + id="HD5aD8rOMVKQ" colab_type="code" outputId="ae9cff3f-8f77-404b-9d97-7711d5e1b515" colab={"base_uri": "https://localhost:8080/", "height": 52}
shilpas_doc
# + id="9p9KslhjMXe8" colab_type="code" outputId="5b98c4cd-9122-4323-ce7c-61d21ad2ad90" colab={"base_uri": "https://localhost:8080/", "height": 54}
dmitriys_doc
# + id="9S9q7fwbMayq" colab_type="code" colab={}
shilpas_doc_orig = {'favorite_colors': ['black', 'white', 'red']}
# + id="bipS9FpdMxj7" colab_type="code" outputId="aeb4c52e-aa9e-482c-e8e7-84de024d1c9c" colab={"base_uri": "https://localhost:8080/", "height": 34}
shilpas_doc_orig
# + id="bbH_wvi9M0Kw" colab_type="code" outputId="7c90f706-e268-40c2-9edf-1629bf10c005" colab={"base_uri": "https://localhost:8080/", "height": 52}
list(db.test.find(shilpas_doc_orig))
# + id="Z4yfA2I0M36f" colab_type="code" outputId="83b8a6b3-f9fb-4fc3-dd4a-4a6713283fab" colab={"base_uri": "https://localhost:8080/", "height": 34}
db.test.insert_one(shilpas_doc_orig)
# + id="BUKhSEK3NHh3" colab_type="code" outputId="0b7a3a0a-494a-490a-d4c7-765c2e214ecb" colab={"base_uri": "https://localhost:8080/", "height": 86}
list(db.test.find({'favorite_colors': ['black', 'white', 'red']}))
# + id="43t0zKLINKNg" colab_type="code" outputId="46f61f5a-dfac-49dc-ac89-f36ed068b6db" colab={"base_uri": "https://localhost:8080/", "height": 34}
macs_doc = {'favorite_colors': ['red', 'white', 'blue'],
'favorite animal!!!': 'dog'}
db.test.insert_one(macs_doc)
# + id="4SYBMSpoNuMI" colab_type="code" outputId="e6288ca3-8028-4b02-bd4c-78bc1e7daa0e" colab={"base_uri": "https://localhost:8080/", "height": 3163}
help(db.test.find)
# + id="gMUfdgdKNw6V" colab_type="code" outputId="ad9e0193-0ffb-4a50-dc5e-bdb5f965b050" colab={"base_uri": "https://localhost:8080/", "height": 104}
list(db.test.find({'favorite_colors': ['red', 'white', 'blue']}))
# + id="8BNmAjNcOIRk" colab_type="code" outputId="a4f70a72-2ecc-4cae-c91f-a67a38839bbe" colab={"base_uri": "https://localhost:8080/", "height": 550}
list(db.test.find({'favorite_colors'}))
# + id="M1upAlVNOqzT" colab_type="code" outputId="40168fe2-d1b8-4dea-c599-7f10a0d78d00" colab={"base_uri": "https://localhost:8080/", "height": 34}
rpg_character = (1, "<NAME>", 10, 3, 0, 0, 0)
db.test.insert_one({'rpg_character': rpg_character})
# + id="jWzRUs22P_ht" colab_type="code" outputId="9dfec45d-fea6-4991-987d-d3d87238bf97" colab={"base_uri": "https://localhost:8080/", "height": 52}
db.test.find_one({'rpg_character': rpg_character})
# + id="gWzc2pyHQI8Y" colab_type="code" outputId="606423fd-54b0-488f-9238-f776f8de571b" colab={"base_uri": "https://localhost:8080/", "height": 34}
# for character in characters...
db.test.insert_one({
'sql_id': rpg_character[0],
'name': rpg_character[1],
'hp': rpg_character[2]
})
# + id="k-o18mK_QZZ2" colab_type="code" outputId="f072e30c-8f71-4dc7-993b-23c574fb1142" colab={"base_uri": "https://localhost:8080/", "height": 312}
list(db.test.find())
# + id="4v_h4qU3Qa9T" colab_type="code" colab={}
| module3-nosql-and-document-oriented-databases/LS_DS2_MongoDB_Playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danilodioliveira/Python_Projects/blob/main/rock_paper_scissors.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 181} id="mKS-Vpmbj1iy" outputId="47437518-fed5-40be-9de0-cadb509a1803"
import random
def play():
user = input(" 'r' for rock, 'p', for paper, 's' for scissors\n")
computer = random.choice(['r', 'p', 's'])
print(f'Player: {user} and Computer: {computer}')
if user == computer:
print('It\'s a tie')
play()
if is_win(user, computer):
return 'You won!'
return 'You lost!'
def is_win( player, opponent):
if (player == 'r' and opponent == 's') or (player == 's' and opponent == 'p') or (player == 'p' and opponent == 'r'):
return True
print('Hi! let\'s play Rock, Paper and Scissors')
play()
# + id="pk5ASR-VzSes"
| rock_paper_scissors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Read in the data
# +
import pandas as pd
import numpy as np
import re
import matplotlib.pyplot as plt
data_files = [
"ap_2010.csv",
"class_size.csv",
"demographics.csv",
"graduation.csv",
"hs_directory.csv",
"sat_results.csv"
]
data = {}
for file in data_files:
data[file[:-4]] = pd.read_csv('schools/{}'.format(file))
# -
print(data['sat_results'].head())
# # Read in the surveys
# +
all_survey = pd.read_csv('schools/survey_all.txt', delimiter = '\t', encoding = 'windows - 1252')
d75_survey = pd.read_csv('schools/survey_d75.txt', delimiter = '\t', encoding = 'windows - 1252')
survey = pd.concat([all_survey, d75_survey], axis = 0)
print(survey.head())
survey["DBN"] = survey["dbn"]
survey_fields = [
"DBN",
"rr_s",
"rr_t",
"rr_p",
"N_s",
"N_t",
"N_p",
"saf_p_11",
"com_p_11",
"eng_p_11",
"aca_p_11",
"saf_t_11",
"com_t_11",
"eng_t_11",
"aca_t_11",
"saf_s_11",
"com_s_11",
"eng_s_11",
"aca_s_11",
"saf_tot_11",
"com_tot_11",
"eng_tot_11",
"aca_tot_11",
]
survey = survey.loc[:,survey_fields]
data["survey"] = survey
# -
# # Add DBN columns
# +
hs_directory = data['hs_directory']
class_size = data['class_size']
hs_directory['DBN'] = hs_directory['dbn']
class_size['padded_csd'] = class_size['CSD'].apply(lambda x: str(x).zfill(2))
class_size['DBN'] = class_size['padded_csd'] + class_size['SCHOOL CODE']
print(class_size.head())
data['hs_directory'] = hs_directory
data['class_size'] = class_size
# -
# # Convert columns to numeric
# +
cols = ['SAT Math Avg. Score', 'SAT Critical Reading Avg. Score', 'SAT Writing Avg. Score']
for c in cols:
data["sat_results"][c] = pd.to_numeric(data["sat_results"][c], errors="coerce")
data['sat_results']['sat_score'] = data['sat_results'][cols[0]] + data['sat_results'][cols[1]] + data['sat_results'][cols[2]]
data['sat_results']['sat_score'].isnull().sum()
# +
def find_lat(loc):
coords = re.findall("\(.+, .+\)", loc)
return coords[0].split(",")[0].replace("(", "")
def find_lon(loc):
coords = re.findall("\(.+, .+\)", loc)
return coords[0].split(",")[1].replace(")", "").strip()
data["hs_directory"]["lat"] = data["hs_directory"]["Location 1"].apply(find_lat)
data["hs_directory"]["lon"] = data["hs_directory"]["Location 1"].apply(find_lon)
data["hs_directory"]["lat"] = pd.to_numeric(data["hs_directory"]["lat"], errors="coerce")
data["hs_directory"]["lon"] = pd.to_numeric(data["hs_directory"]["lon"], errors="coerce")
print(data["hs_directory"][["lat", 'lon']].head())
# -
# # Condense datasets
# +
class_size = data["class_size"]
class_size = class_size[class_size["GRADE "] == "09-12"]
class_size = class_size[class_size["PROGRAM TYPE"] == "GEN ED"]
class_size = class_size.groupby("DBN").agg(np.mean)
class_size.reset_index(inplace=True)
data["class_size"] = class_size
data["class_size"].head()
# +
demographics = data['demographics']
demographics = demographics[demographics['schoolyear'] == 20112012]
data['demographics'] = demographics
data['demographics'].head()
# +
graduation = data['graduation']
graduation = graduation[(graduation['Cohort'] == '2006') & (graduation['Demographic'] == 'Total Cohort')]
data['graduation'] = graduation
graduation.head()
# -
# # Convert AP scores to numeric
cols = ['AP Test Takers ', 'Total Exams Taken', 'Number of Exams with scores 3 4 or 5']
ap_2010 = data['ap_2010']
for name in cols:
ap_2010[name] = pd.to_numeric(ap_2010[name], errors = 'coerce')
data['ap_2010'] = ap_2010
data['ap_2010'].head()
# # Combine the datasets
# +
combined = data["sat_results"]
combined = combined.merge(data['ap_2010'], on = 'DBN', how = 'left')
combined = combined.merge(data['graduation'], on = 'DBN', how = 'left')
combined = combined.merge(data['class_size'], on = 'DBN', how = 'inner')
combined = combined.merge(data['demographics'], on = 'DBN', how = 'inner')
combined = combined.merge(data['survey'], on = 'DBN', how = 'inner')
combined = combined.merge(data['hs_directory'], on = 'DBN', how = 'inner')
print('Shape of the dataframe: {}'.format(combined.shape))
combined.head()
# -
mean = combined.mean()
combined = combined.fillna(mean)
combined = combined.fillna(0)
# # Add a school district column for mapping
combined['school_dist'] = combined['DBN'].apply(lambda x: x[:2])
combined['school_dist'].head()
# # Find correlations
correlations = combined.corr()
correlations = correlations["sat_score"]
print(correlations)
# # Plotting survey correlations
# Remove DBN since it's a unique identifier, not a useful numerical value for correlation.
# survey_fields.remove("DBN")
survey_fields
# %matplotlib inline
correlations[survey_fields].plot.bar()
# ## Investigating safety scores
combined.plot.scatter('saf_s_11', 'sat_score')
# +
from mpl_toolkits.basemap import Basemap
import matplotlib.pyplot as plt
dist_avrg = combined.groupby('school_dist').agg(np.mean)
dist_avrg.reset_index(inplace = True)
fig,ax = plt.subplots(figsize = (10,10))
m = Basemap(
projection='merc',
llcrnrlat=40.496044,
urcrnrlat=40.915256,
llcrnrlon=-74.255735,
urcrnrlon=-73.700272,
resolution='h'
)
m.drawmapboundary(fill_color='#85A6D9')
m.drawcoastlines(color='#6D5F47', linewidth=.4)
m.drawrivers(color='#6D5F47', linewidth=.4)
m.fillcontinents(color='white',lake_color='#85A6D9')
lon = dist_avrg['lon'].tolist()
lat = dist_avrg['lat'].tolist()
m.scatter(lon, lat, s = 50, zorder = 2, latlon = True, c = dist_avrg['saf_s_11'],
cmap = 'summer')
m.colorbar(location='bottom', label='Safety Score')
plt.title('Average safety score for each school district in NY', weight = 'bold')
plt.show()
# -
# ## Investigating racial differences in SAT scores
# +
cols = [
'white_per',
'asian_per',
'black_per',
'hispanic_per'
]
correlations[cols].plot.bar(color = 'grey')
plt.tick_params(left = False, top = False, right = False, bottom = False)
plt.axhline(0)
plt.text(-0.25, 0.65, 'white per')
plt.text(0.75, 0.6, 'asian per')
plt.text(1.75, - 0.35, 'black per')
plt.text(2.7, - 0.45, 'hispanic per')
plt.xticks([])
plt.title('Racial percentage correlated with SAT score', weight = 'bold')
plt.box(False)
# -
# ## Exploring schools with low SAT scores and high values for hispanic percentage
combined.plot.scatter('hispanic_per', 'sat_score')
combined.loc[combined['hispanic_per'] > 95, "SCHOOL NAME"]
combined[(combined["hispanic_per"] < 10) & (combined["sat_score"] > 1800)]["SCHOOL NAME"]
# ## Investigating gender differences in SAT scores
gender_fields = ["male_per", "female_per"]
combined.corr()["sat_score"][gender_fields].plot.bar(color = 'lightblue')
plt.tick_params(left = False, top = False, right = False, bottom = False)
plt.axhline(0, color = 'black')
plt.xticks([])
plt.title('Gender correlated with SAT score', weight = 'bold')
plt.text(-0.08, -0.13, 'male', fontsize = 12)
plt.text(0.9, 0.12, 'female', fontsize = 12)
plt.box(False)
plt.show()
combined.plot.scatter('female_per', 'sat_score')
combined.loc[(combined["female_per"] > 60) & (combined["sat_score"] > 1700), "SCHOOL NAME"]
# ## Advanced Placement (AP) exams investigation
# In the U.S., high school students take Advanced Placement (AP) exams to earn college credit. There are AP exams for many different subjects.
#
# It makes sense that the number of students at a school who took AP exams would be highly correlated with the school's SAT scores. Let's explore this relationship.
combined['ap_per'] = combined['AP Test Takers ']/combined['total_enrollment']
combined.plot.scatter('ap_per', 'sat_score')
| NYC_schools/Schools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task 3 - Missing Values
import pandas as pd
import numpy as np
df = pd.read_csv('../csv_output/pre_process_1.csv')
df
df.info()
# ***taking care of missing values***
df.isna()
df.Age.value_counts(dropna=False)
# ***filling constant value***
df.Age
df.Age.fillna(df.Age.mean())
# ***forward fill method***
df.Age.fillna(method='ffill')
# ***backward fill method***
df.Salary.fillna(method='backfill')
# ***dropping the row of NaN***
df.dropna().reset_index().drop('index', axis=1)
| tutorials/pandas/Task 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import progressbar
# +
# honest network delay over next n blocks.
def vectorDelayHonest(ps, es, init_endorsers, delay_priority, delay_endorse):
return (60 * len(ps)
+ delay_priority * sum(ps)
+ sum([delay_endorse * max(init_endorsers - e, 0) for e in es]))
# attacking network delay over next n blocks.
def vectorDelayAttacker(ps, es, init_endorsers, delay_priority, delay_endorse):
return (60 * len(ps)
+ delay_priority * sum(ps)
+ sum([delay_endorse * max(init_endorsers - e, 0) for e in es[1:]]))
# efficient sample generation
def getAH(alpha):
x = np.random.geometric(1-alpha)
if x == 1:
h = 0
a = np.random.geometric(alpha)
else:
a = 0
h = x - 1
return [a, h]
# -
def getProbReorg(alpha, length, init_endorsers, delay_priority, delay_endorse, sample_size = int(1e5)):
bar = progressbar.ProgressBar()
feasible_count = 0
for _ in range(sample_size):
aVals = []
hVals = []
for i in range(length):
a, h = getAH(alpha)
aVals.append(a)
hVals.append(h)
eVals = np.random.binomial(32, alpha, size = length)
honest_delay = vectorDelayHonest(hVals, 32 - eVals, init_endorsers, delay_priority, delay_endorse)
selfish_delay = vectorDelayAttacker(aVals, eVals, init_endorsers, delay_priority, delay_endorse)
if selfish_delay <= honest_delay:
feasible_count += 1
return feasible_count / sample_size
getProbReorg(alpha = 0.45, length = 20)
resultsFinal2 = np.zeros((20, 4))
x = np.arange(0.3, 0.5, 0.01)
lengths = [20,35,55,80]
bar = progressbar.ProgressBar()
for i in bar(range(len(x))):
for j in range(len(lengths)):
resultsFinal2[i,j] = getProbReorg(
x[i],
lengths[j],
init_endorsers=10,
delay_endorse=40,
delay_priority=60,
sample_size=int(1e4)
)
# +
f, axarr = plt.subplots(ncols = 2, figsize=(14,6))
x = np.arange(0.3, 0.5, 0.01)
lengths = [20,35,55,80]
markers = ['^', 's', 'o', 'd']
for i in range(len(lengths)):
# plotting the markers
axarr[0].plot(x, resultsFinal2[:,i], 'k--', marker=markers[i], fillstyle='none', linewidth=0.15,
label=r'length-{}'.format(lengths[i]))
axarr[1].plot(x, resultsFinal2[:,i], 'k--', marker=markers[i], fillstyle='none', linewidth=0.15,
label=r'length-{}'.format(lengths[i]))
axarr[1].axhline(1/float(24*60), color='blue', linewidth = 1, alpha=1, linestyle='--', label='daily')
axarr[1].axhline(1/float(24*60*365), color='green', linewidth = 1, alpha=1, linestyle='--', label='yearly')
for ax in axarr:
ax.legend()
ax.set_xlabel(r'Attacker stake', size=15)
ax.set_ylabel(r'Probability of feasible attack', size=15)
ax.set_xticks(np.arange(0.33, 0.5, 0.02))
ax.set_xlim(0.33, 0.5)
axarr[1].set_yscale('log')
axarr[0].set_title('Linear Axis', size=15)
axarr[1].set_title('Log Axis - Confidence Intervals', size=15)
axarr[1].set_ylim(float(2e-11), float(1))
plt.show()
| new_params.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from collections import Counter
s = 'hackerhappy'
t = 'hackerrank'
ns = len(s)
nt = len(t)
k = 9
list(s)
list(t)
i = 0
while s[i]==t[i]:
i+=1
i
import math
n = math.sqrt(48)
int(n)
4**2
def appendAndDelete(s, t, k):
i = 0
while s[i]==t[i]:
i+=1
print(i)
print((len(s)-i)+(len(t)-i))
if k==(len(s)-i)+(len(t)-i):
print("Yes")
else:
print("No")
a, b = 17, 24
counter = 1
# find out first square integer in the range
start =0
for i in range(a, b+1):
if int(math.sqrt(i))==math.sqrt(i):
start = i
first_sqr = int(math.sqrt(i))
break
while (first_sqr+1)**2 <=b:
first_sqr+=1
counter+=1
counter
c = 0
for _ in range(2):
c+=1
c
a = [1,2,3]
a.pop(2)
a
a = list(map(int, input().split()))
len(a)
def cutTheSticks(arr):
min_value = min(arr)
max_value = max(arr)
print(min_value, max_value)
if min_value==max_value:
return len(arr)
else:
res = []
while min_value<max_value:
n = len(arr)
res.append(n)
min_value = min(arr)
max_value = max(arr)
# print(arr, n, min_value, max_value)
i = 0
while i<=n-1:
arr[i] = arr[i] - min_value
if arr[i]==0:
arr.pop(i)
i-=1
n = len(arr)
i+=1
return res
cutTheSticks(a)
a = [2,3,4,3,2,4,1]
while len(a)!=0:
a = [x for x in a if x!=min(a)]
print(a)
len('3333')
list('2222')
t = [1,2,3,4,5,6,7,8]
t[4:7]
def gridSearch(G, P):
for i in range(R):
for j in range(C):
if G[i][j]==P[0][0] and (C-j)>=c and (R-i)>=r:
# print('---')
counter = 0
for k,u in zip(range(i,i+r),range(r)):
# print(G[k][j:j+c], P[u], i,i+r, r)
if G[k][j:j+c]==P[u]:
counter+=1
if counter==r:
return 'YES'
return 'NO'
C
RC = input().split()
R = int(RC[0])
C = int(RC[1])
G = []
for _ in range(R):
G_item = input()
G.append(G_item)
rc = input().split()
r = int(rc[0])
c = int(rc[1])
P = []
for _ in range(r):
P_item = input()
P.append(P_item)
G[4][3:7]
P
gridSearch(G, P)
for i in range(3,5):
print(i)
for i, j in zip(range(0,3), range(3,6)):
print(i, j)
def surfaceArea(A):
# find out largest number and dips for each row
R = []
C = []
counter = 0
for i in range(H):
temp = []
temp1 = []
for j in range(W):
temp.append(A[i][j])
# temp1.append(A[j][i])
# print(i, j, W)
# if (j>0 and j<W-1):
# if A[i][j]<A[i][j-1] and A[i][j]<A[i][j+1]:
# counter+=(A[i][j-1]-A[i][j])+(A[i][j+1]-A[i][j])
R.append(max(temp))
# C.append(max(temp1))
for i in range(H):
for j in range(W):
if i>0 and i<H-1 and A[i][j]<A[i-1][j] and A[i][j]<A[i+1][j]:
print((A[i-1][j]-A[i][j]),(A[i+1][j]-A[i][j]))
counter+=(min(A[i-1][j],A[i+1][j])-A[i][j])*2
print(W*H*2, R, C, counter)
return W*H*2+sum(R)*2+sum(C)*2+counter
HW = input().split()
H = int(HW[0])
W = int(HW[1])
A = []
for _ in range(H):
A.append(list(map(int, input().rstrip().split())))
A
surfaceArea(A)
sum([51, 32, 28, 49, 28, 21, 98, 56, 99, 77])*2
8+14+84+1078+99+99
21+4+7+77+42+43
1204+99*2
t =[[1,2,3,4,5],[5,6,7,4,5],[10,11,12,1,1]]
for i in range(3):
for j in range(5):
if (i==0 and j==0) or (i==0 and j==5-1) or (i==3-1 and j):
print(t[i][j])
t = [1,2,3,4,5,6,7]
n = len(t)
r = 4
t
t[r:]+t[:r]
d = {i+1:v+1 for i,v in enumerate(range(4))}
d
for i in range(1,5+1):
print(i)
d1 = list(d.values())
d1
list(d.values())
def absolutePermutation(n, k):
d = {i+1:v+1 for i,v in enumerate(range(n))}
c = 0
for i in range(1, n+1):
val = i+k
if val!=d[i]:
if val>n:
return -1
else:
d[i]=val
d[val]=i
c+=2
if c>n/2:
break
return list(d.values())
absolutePermutation(4,2)
def list_cal(x, y):
return [abs(i-j) for i,j in zip(x, y)]
'..'.isupper()
letters = 'abcdefghijklmnopqrstuvwxyz'
d_lower = {i+1:v for i,v in enumerate(letters)}
d_lower
def marsExploration(s):
counter=0
for i in range(len(s)-2):
if i%3==0:
if s[i]!='S' or s[i+1]!='O' or s[i+2]!='S':
print([s[i]=='S',s[i+1]=='O',s[i+2]=='S'])
counter+=3-sum([s[i]=='S',s[i+1]=='O',s[i+2]=='S'])
return counter
s = 'rhackerrank'
word = 'hackerrank'
s.find(word[0],2)
def hackerrankInString(s):
word = 'hackerrank'
valid = 'YES'
for i in range(len(word)):
x = s.find(word[i],i)
if x!=-1:
x = s.find(word[i],i)
else:
valid = 'NO'
break
return valid
'We promptly judged antique ivory buckles for the next prize'.lower().replace(' ','')
s='abccddde'
s
from collections import Counter
d = {v:i+1 for i,v in enumerate('abcdefghijklmnopqrstuvwxyz')}
d
ord('z')-ord('a')+1
# +
s_d = dict(Counter(s))
dd = dict(Counter(s))
for k in s_d:
dd[k] = dd[k]*d[k]
dd
# -
s_d
for i in q:
for
s
# +
d={}
weight = 0
for i in range(len(s)):
if i==0 or s[i]!=s[i-1]:
weight = ord(s[i])-ord('a')+1
else:
weight=weight+ord(s[i])-ord('a')+1
d[weight]=1
d
# -
s = 'abccddde'
res ={}
weight = 0
for i in range(len(s)):
if s[i]==s[i-1] and i>0:
weight += d[s[i]]
res[weight] = 1
else:
weight=0
weight = d[s[i]]
res[d[s[i]]]=1
res
from time import time
st_time = time()
d = {}
for i in range(1,50000000):
d[i] = 1
print(time()-st_time)
s = '99100101102'
len(s)
# +
n = len(s)
res = []
for i in range(1, n//2+1):
k = 0
total=''
first = s[:i]
while len(total)<n:
current = str(int(s[:i])+k)
total += current
k+=1
res.append(total)
res
# -
from collections import Counter
arr = ['abcdde', 'baccd', 'eeabg']
res = []
for i in range(len(arr)):
res+=list(set(arr[i]))
d =dict(Counter(res))
k = 0
total = ''
first = s[:i]
while len(total)<n:
current = str(int(s[:i])+k)
total += current
print(total)
k+=1
| scratch_script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Subset Distributions
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from sklearn.neighbors import KNeighborsClassifier
from mlxtend.feature_selection import SequentialFeatureSelector as SFS
from wakeful import log_munger, metrics, virus_total, pipelining, preprocessing
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from bat.dataframe_to_matrix import DataFrameToMatrix
# %matplotlib inline
# #### dnscat2
# %ls data/*.h5
df_train = log_munger.hdf5_to_df('dnscat2_2017_12_31_conn_train', './data')
df_test = log_munger.hdf5_to_df('dnscat2_2017_12_31_conn_test', './data')
print('train value counts:\n', df_train.label.value_counts())
print('test value counts:\n', df_test.label.value_counts())
'label' in df_train.columns, 'label' in df_test.columns
y_train = df_train.pop('label')
X_train = df_train
to_matrix = DataFrameToMatrix()
X_train_mat = to_matrix.fit_transform(X_train)
y_test = df_test.pop('label')
X_test = df_test
to_matrix = DataFrameToMatrix()
X_test_mat = to_matrix.fit_transform(X_test)
# ### PCA Feature Reduction
pca = PCA(n_components=6)
pca.fit(X_train_mat)
print(sum(pca.explained_variance_ratio_), '=', pca.explained_variance_ratio_)
print(pca.n_components_)
X_test_mat_pca = pca.fit_transform(X_test_mat)
X_test_mat_pca.shape
# ### SequentialFeature Reduction
df_train.columns
df_test.columns
pipelining.feature_selection_pipeline(train_df=df_train, test_df=df_test)
sns.pairplot(df_train_conn_dnscat2, hue='label')
# ### Distributions
# %ls ./data
keys = [
('iodine-forwarded-2017-12-31-conn-test', 'iodine-forwarded-2017-12-31-conn-train'),
('iodine-raw-2017-12-31-conn-test', 'iodine-raw-2017-12-31-conn-train'),
('dnscat2-2017-12-31-conn-test', 'dnscat2-2017-12-31-conn-train'),
('iodine-forwarded-2017-12-31-dns-test', 'iodine-forwarded-2017-12-31-dns-train'),
('iodine-raw-2017-12-31-dns-test', 'iodine-raw-2017-12-31-dns-train'),
('dnscat2-2017-12-31-dns-test', 'dnscat2-2017-12-31-dns-train'),]
train_key = 'iodine-forwarded-2017-12-31-conn-train'
test_key = 'iodine-forwarded-2017-12-31-conn-test'
data_dir='./data'
train_df = log_munger.hdf5_to_df(key, data_dir)
test_df = log_munger.hdf5_to_df(key, data_dir)
df.columns
pipelining.feature_selection_pipeline(train_df=train_df, test_df=test_df)
df = df[['local_orig', 'local_resp', 'orig_ip_bytes', 'pcr', 'label']]
df = df.dropna(axis=0, how='any')
sns.pairplot(df, hue='label')
| clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Gtk7R7enf9WO"
# Alunos:
# * <NAME> - DRE: 114197959
# * <NAME> - DRE: 117154510
#
#
# Fontes:
# 1. [Hill Climbing Algorithm | Hill Climbing in Artificial Intelligence | Data Science Tutorial | Edureka](https://www.youtube.com/watch?v=_ThdIOA9Lbk)
# 2. [Algoritmos de otimização: Hill Climbing e Simulated Annealing](https://medium.com/data-hackers/algoritmos-de-otimiza%C3%A7%C3%A3o-hill-climbing-e-simulated-annealing-3803061f66f0)
#
#
# + id="s5H19KtJmXgK"
from random import randint
import sys
from random import uniform
from math import exp
# + [markdown] id="y6Dp4Q1OFXL5"
#
#
# > IMPLEMENTAÇÃO
#
#
# 1. Defina uma função que dado o tamanho do tabuleiro N, retorna um tabuleiro N×N com N rainhas. O tabuleiro deve ser gerado de maneira aleatoria.
#
# 4. Defina uma função que dado um tabuleiro qualquer, retorna a avaliação deste tabuleiro (numero de ataques entre as rainhas).
#
#
#
# + id="1mJxlJaOfzW8"
#criar tabuleiro de uma lista de tamanho N (N = numero de rainhas)
#cada elemento da lista poderá ir de zero a N
def criar_Tabuleiro(num_de_Rainhas):
tabuleiro = []
for i in range(0, num_de_Rainhas):
tabuleiro.append(randint(1, num_de_Rainhas))
print("O tabuleiro foi criado:")
return tabuleiro
# Para cada rainha no tabuleiro, busca quantos ataques são possíveis na horizontal
def buscarNumeroDeAtaquesNaHorizontal(tabuleiro):
numeroDeAtaques = 0
for posicaoRainha in range (0,len(tabuleiro)):
for possivelAtaqueNaDireita in range (posicaoRainha,len(tabuleiro)):
if possivelAtaqueNaDireita == posicaoRainha:
continue
elif tabuleiro[posicaoRainha] == tabuleiro[possivelAtaqueNaDireita]:
numeroDeAtaques += 1
break
return numeroDeAtaques
# Analisa o tabuleiro no sentido nordeste de cada rainha, contando o número de ataques possíveis no diagonal
def buscarNumeroDeAtaquesNoNordeste(tabuleiro):
numeroDeAtaques = 0
for posicaoRainha in range (0,len(tabuleiro)):
mapeiaDiagonal = tabuleiro[posicaoRainha]
for posicaoHorizontal in range (posicaoRainha + 1, len(tabuleiro)):
mapeiaDiagonal -= 1
if tabuleiro[posicaoHorizontal] == mapeiaDiagonal:
numeroDeAtaques += 1
break
return numeroDeAtaques
# Analisa o tabuleiro no sentido sudeste de cada rainha, contando o número de ataques possíveis nos diagonal
def buscarNumeroDeAtaquesNoSudeste(tabuleiro):
numeroDeAtaques = 0
for posicaoRainha in range (0,len(tabuleiro)):
mapeiaDiagonal = tabuleiro[posicaoRainha]
for posicaoHorizontal in range (posicaoRainha + 1, len(tabuleiro)):
mapeiaDiagonal += 1
if tabuleiro[posicaoHorizontal] == mapeiaDiagonal:
numeroDeAtaques += 1
break
return numeroDeAtaques
def buscarNumeroDeAtaquesNaVertical(tabuleiro):
#TO-DO
ataque_vertical = buscarNumeroDeAtaquesNoNordeste(tabuleiro) + buscarNumeroDeAtaquesNoSudeste(tabuleiro)
return ataque_vertical
# Como a heurística é baseada em números de ataques. Esta função retorna quantos ataques são possíveis no tabuleiro
def num_Ataques_No_Tabuleiro(tabuleiro):
num_de_ataques = buscarNumeroDeAtaquesNaHorizontal(tabuleiro) + buscarNumeroDeAtaquesNaVertical(tabuleiro)
print(num_de_ataques)
return num_de_ataques
#8,16,32,64
#criar_Tabuleiro(8)
# + [markdown] id="3wRd2wAqF1Fv"
# 2. Defina uma função que dado um tabuleiro qualquer, retorna todos os seus vizinhos.
# 3. Defina uma função que dado um tabulleiro qualquer, retorna um de seus vizinhos. A escolha do vizinho a ser retornado pela função deve ser aleatoria.
# + id="RVh2a9JBsYx_"
#Entre um conjunto de tabuleiro, retorna o tabuleiro com o melhor heuristica
def busca_primeiro_melhorVizinho(tabuleiro, vizinhos):
melhor_Vizinho = tabuleiro
#uma busca de ataques
melhor_Heuristica = num_Ataques_No_Tabuleiro(tabuleiro)
for vizinho in vizinhos:
heuristica_Vizinho = num_Ataques_No_Tabuleiro(vizinho)
if heuristica_Vizinho < melhor_Heuristica:
melhor_Heuristica = heuristica_Vizinho
melhor_Vizinho = vizinho
break
print("O primeiro melhor vizinho encontrado foi:")
print(melhor_Vizinho)
print("Com o heuristica de valor: ")
print(melhor_Heuristica)
return [melhor_Vizinho, melhor_Heuristica, len(vizinhos)]
# Busca e retornar todos os vizinhos do tabuleiro em questão
def recuperaVizinhosDeTabulero(tabuleiro):
vizinhos_De_Tabuleiro = []
for identificadorDaRainha in range (0, len(tabuleiro)):
for posicaoDaRainha in range (1, len(tabuleiro)+1):
if posicaoDaRainha == tabuleiro[identificadorDaRainha] :
continue
vizinho = tabuleiro.copy()
vizinho[identificadorDaRainha] = posicaoDaRainha
vizinhos_De_Tabuleiro.append(vizinho)
return vizinhos_De_Tabuleiro
# + [markdown] id="_IWnqc2oG6Jh"
# # Hill Climbing
# 1. Implementação Hill Climbing
#
# + id="Uu_FHsuCoUrY" colab={"base_uri": "https://localhost:8080/"} outputId="91463e67-a53f-4019-819c-e8739c7e4277"
#Simula o problema Usando Hillclimbing
def problema_N_Rainha_com_HillClimbing(num_de_Rainhas):
tabuleiro = criar_Tabuleiro(num_de_Rainhas)
tabuleiro_Atual = tabuleiro
heuristicaAtual = sys.maxsize
cont_heuristica_Repetida = 0
cont_Tabuleiro_Gerados = 0
print("----------------------------------------------------")
while(True):
melhor_vizinho_comHeuristica = busca_primeiro_melhorVizinho(tabuleiro_Atual, recuperaVizinhosDeTabulero(tabuleiro_Atual))
print("melhor_vizinho_comHeuristica")
print(melhor_vizinho_comHeuristica)
cont_Tabuleiro_Gerados += melhor_vizinho_comHeuristica[2]
if heuristicaAtual == melhor_vizinho_comHeuristica[1]:
cont_heuristica_Repetida += 1
if heuristicaAtual > melhor_vizinho_comHeuristica[1]:
cont_heuristica_Repetida = 0
tabuleiro_Atual = melhor_vizinho_comHeuristica[0]
heuristicaAtual = melhor_vizinho_comHeuristica[1]
if cont_heuristica_Repetida > 3:
print("foi encontrado um ombro, terminamos a execução...")
break
if heuristicaAtual == 0:
print("Solução Encontrada!!")
print("A solução do problema será:")
print(tabuleiro_Atual)
break
print("==================================================")
return [tabuleiro_Atual, heuristicaAtual, cont_Tabuleiro_Gerados]
problema_N_Rainha_com_HillClimbing(4)
#problema_N_Rainha_com_HillClimbing(8)
#problema_N_Rainha_com_HillClimbing(16)
#problema_N_Rainha_com_HillClimbing(32)
# + id="fWLJoR538QRr"
#Retorna um vizinho alearório dado um tabuleiro
def busca_Vizinho_Aleatorio(tabuleiro):
#TO-DO: (done)
vizinhos = recuperaVizinhosDeTabulero(tabuleiro)
vizinhoAleatorio = vizinhos[randint(0,len(tabuleiro))]
print("O vizinho aleatório retornado será:")
print(vizinhoAleatorio)
return [vizinhoAleatorio, len(vizinhos)]
# + [markdown] id="FM_221P9IXz8"
# # Implementação Simulated Annealing
# + id="hAC6lxmto6MN"
#Simula o problema usando Simulated Annealing
def problema_N_Rainha_com_Simulted_Annealing(num_de_Rainhas, iter_Max, temp_Inicial, alpha):
tabuleiro = criar_Tabuleiro(num_de_Rainhas)
tabuleiro_Atual = tabuleiro
solucao = tabuleiro_Atual
temp_Atual = temp_Inicial
cont_vizinho_gerados = 0
for i in range(1, iter_Max):
if(temp_Atual <= 0):
break
vizinho_Aleatorio = busca_Vizinho_Aleatorio(solucao)
heuristica_Vizinho_Aleatorio = num_Ataques_No_Tabuleiro(vizinho_Aleatorio[0])
diferenca_Custo = heuristica_Vizinho_Aleatorio - num_Ataques_No_Tabuleiro(tabuleiro_Atual)
cont_vizinho_gerados += vizinho_Aleatorio[1]
if(diferenca_Custo < 0):
tabuleiro_Atual = vizinho_Aleatorio[0]
if(num_Ataques_No_Tabuleiro(vizinho_Aleatorio[0]) <= num_Ataques_No_Tabuleiro(solucao)):
solucao = vizinho_Aleatorio[0]
if(heuristica_Vizinho_Aleatorio == 0):
break
else:
if(uniform(0,1) < exp(-(diferenca_Custo / temp_Atual))):
tabuleiro_Atual = vizinho_Aleatorio[0]
temp_Atual = temp_Atual * alpha
print("-------------------------------------------------")
print("A soluçao encontrada foi:")
print(solucao)
print("Com a heuristica: ")
print(num_Ataques_No_Tabuleiro(solucao))
print("=================================================")
return [solucao, cont_vizinho_gerados]
problema_N_Rainha_com_Simulted_Annealing(4, 20, 100, 0.1)
#problema_N_Rainha_com_Simulted_Annealing(8, 10, 50, 0.5)
#problema_N_Rainha_com_Simulted_Annealing(16, 10, 50, 0.1)
| Trabalho/HillClimbing_AnnealingSimulated/trabalho_hillclimbing_Annealing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Installing packages: `feedeR`, `rtweet`, `RCurl`, `xml2`, `RJSONIO`, `RSQLite`, `stringr`
# **1.1 Establish an SSH connection with X11 tunnel to `user.palmetto.clemson.edu`: **
#
# - For Linux machines, you can use the default command line terminal
# - For Mac machines, you need to make sure that XQuartz is installed before using the default command line terminal
# - For Windows machines, the recommended approach is to download and install [MobaXterm](http://mobaxterm.mobatek.net/)
#
# Additional documentations can be found at:
#
# - [Logging on to Palmetto using MobaXterm for Windows](https://www.palmetto.clemson.edu/palmetto/userguide_basic_usage.html)
# - [How to run graphical appliction](https://www.palmetto.clemson.edu/palmetto/userguide_howto_run_graphical_applications.html)
# **1.2. Request a temporary node with X11 tunnel for setting the required R packages **
#
# Once you are logged into palmetto, prior to getting a new node, you need to have an environmental variable set up to run at start. This is done via the following command:
#
# ```
# # echo 'export CONDA_ENVS_PATH=/usr/local/share/jupyterhub/env' >> .bashrc
# ```
#
# Next, request a node for two hours:
#
# ```
# clear
# qsub -I -X -l walltime=02:00:00
# ```
# **1.3. Open the correct R distribution that is used for JupyterHub's R notebooks **
#
# ```
# module load anaconda3/4.2.0
# source activate R
# R
# ```
# **1.4. Install packages:**
#
# Inside the R prompt, run the following command
#
# ```
# package_list <- c('feedeR','rtweet','RCurl','xml2','RJSONIO','RSQLite','stringr')
# install.packages(package_list,repos='http://cran.cnr.berkeley.edu/')
# ```
#
# - If this is the first time that you run R from inside Palmetto, R will ask for a non-root installation directory for future packages. Accept the suggested path that looks similar to the following: ‘/home/YOUR_USER_NAME/R/x86_64-pc-linux-gnu-library/3.3’
# - If for some reasons the Berkely mirror URL does not work, you can also try using the folowing URLS for repos:
# - http://cran.stat.ucla.edu/
# - http://mirror.las.iastate.edu/CRAN/
# - http://cran.mtu.edu/
# - Other mirror URLs can be found at https://cran.r-project.org/mirrors.html
# - You can test that all packages are installed and usable by load them all and check session information to see the loaded packages under header *other attached packages*
#
# ```
# library(feedeR)
# library(rtweet)
# library(RCurl)
# library(xml2)
# library(RJSONIO)
# library(RSQLite)
# library(stringr)
# sessionInfo()
# ```
# <img src="./figures/r_jupyter.png">
# ## 2. Setup `rtweet` for streaming Twitter data
#
# - This section is based on instructions provided at https://mkearney.github.io/rtweet/articles/auth.html
# - The terminal connecting to Palmetto with X11 tunneling from step 1 should be kept open.
# ** 2.1. Creating Twitter account **
#
# - Sign up for a Twitter account at https://twitter.com
# - Make sure that your account has an associated phone number. This is required to make your Twitter account into a developer account (being able to create app)
# ** 2.2. Creating Twitter app **
#
# - Go to https://apps.twitter.com and sign in with your Twitter account
# - Create an application:
# - Application names are unique, you will need to pick a different name from R_Workshop_Clemson
# - Some descriptions are required, it is just to describe what your application will do
# - The website is required, but you do not have to provide a specific website. You only need to provide a place holder URL that is in the correct format.
# - The Callback URL must be http://127.0.0.1:1410
# - Once the Twitter app is created, you will be able to click on the app's name on the front page of https://apps.twitter.com to go to the Application Management page. Select the **Keys and Access Tokens** tab to see your access tokens. You will need the **Consumer Key (API Key)** and **Consumer Secret (API Secret)** strings as shown in this tab for the next steps.
# <img src="./figures/twitter_app_creation.png">
# ** 2.3. Setting up Twitter security token for R's rtweet package: **
#
# - Continue in the same command line terminal from step 1, type in the following R codes
#
# ```
# appname <- YOUR_APP_NAME
# key <- YOUR_CONSUMER_KEY
# secret <- YOUR_CONSUMER_SECRET
# twitter_token <- create_token(app = appname,consumer_key = key,consumer_secret = secret)
# ```
#
# - After these R commands are executed, a Firefox browser will pop up asking you to sign in and authenticate the access token, and the R environment will print the following lines while waiting for the authentication:
#
# ```
# Waiting for authentication in browser...
# Press Esc/Ctrl + C to abort
# ```
#
# - Click on the blue **Authorize App** button to confirm the authorization. This will take you to a web page that has the lines *Authentication complete. Please close this page and return to R*. You can now close the Firefox browser. The R environment will print out the line *Authentication complete* and escape from waiting mode into the normal R prompt.
# <img src="./figures/twitter_app_authorization.png">
# ** 2.4. Saving Twitter security token for future usage: **
#
# The `twitter_token` variable should be saved to a file after step 2.3 is completed so that it can be reused later. The process is as follows:
#
# - Continuing in the same R command line terminal as step 2.3, execute the followings:
#
# ```
# home_directory <- path.expand("~/")
# file_name <- file.path(home_directory, "twitter_token.rds")
# saveRDS(twitter_token, file = file_name)
# cat(paste0("TWITTER_PAT=", file_name),file = file.path(home_directory, ".Renviron"),append = TRUE)
# ```
#
# - The above codes will save the `twitter_toke` variable into the `twitter_token.rds` file stored inside your home directory. Next, it creates an environment variable called `TWITTER_PAT` that points to this file and store the environment variable in the default `.Renviron` file that will be loaded by R whenever R is started.
# - It is possible to customize the path to the token file (maybe save it to a specific directory) and `TWITTER_PAT`
# - When tne `rtweet` library is loaded, `TWITTER_PAT` will be read and all subsequent streaming calls to Twitter will be validated automatically.
| jupyter/0_preparations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Administrative Descriptive Stats Preparation
# After appending travel time information to each populated place in an administrative center we can prepare any number of descriptive stats. Given the quantity of data in question these are best prepared with Dask Dataframes. This notebook separates out the descriptive stats preparations,
# +
import dask
import coiled
from dask.distributed import Client, LocalCluster, Lock
from dask.utils import SerializableLock
import dask.dataframe as dd
import pandas as pd
import geopandas as gpd
import spatialpandas as sp
import dask_geopandas as dg
import rioxarray as rx
import xarray as xr
import re
import os
from dask_control import *
from raster_ex import *
from spatialpandas.geometry import (
PointArray, MultiPointArray, LineArray,
MultiLineArray, PolygonArray, MultiPolygonArray
)
import numpy as np
from datetime import date
# + [markdown] tags=[]
# ## Setup
# -
# **Files and variables**
# Today's date
today = date.today().strftime("%d%m%y")
# Directories
# + tags=[]
geo_dir = r'P:\PAK\GEO'
data_dir = r'../../data'
dest_dir = r'destinations'
acc_dir = r'access'
tab_dir = r'tabular'
# -
POINTS_URL = r'tabular/df_pixels_final-*.csv'
# Projections
# + tags=[]
# change this to whatever the desired output projection is
DEST_CRS = 'EPSG:32642'
dcrs_int = int(re.findall('[0-9]+',DEST_CRS)[0])
dcrs_int
# -
# **Initiate Dask Client**
client=get_dask_client(cluster_type='local',n_workers=4,processes=True,threads_per_worker=8)
client
# + [markdown] tags=[]
# ## Loading data
# -
# Pixel level data
# + tags=[]
# df_pixels = dd.read_parquet(os.path.join(data_dir,acc_dir,'tabular/df_pixels_final.parquet'),
# na_values = ' ',
# blocksize=100e6)\
# .sort_values('Adm2_Code')\
# .set_index('ix', drop=True, sorted=False)
# -
df_pixels = dd.read_csv(os.path.join(data_dir,acc_dir,POINTS_URL),header=0,
na_values = ' ',
blocksize=100e6)\
.sort_values('Adm2_Code')\
.set_index('ix', drop=True, sorted=False)
df_pixels
# persist as well will be conducting many operations on this dataset and want to branch off from it
df_pixels = df_pixels.persist()
# Reference rasters
rasters = {}
rlimit = 9999
r_ct = 0
for file in os.listdir(os.path.join(data_dir,acc_dir,'current/master')):
#if file.endswith("_COG.tif"):
if file.startswith("Current_"):
acc_rast = re.search(r'Current_(.*?).tif',os.path.basename(file)).group(1)
rasters[acc_rast] = f"{data_dir}/{acc_dir}/current/master/{file}"
r_ct = r_ct + 1
if r_ct >= rlimit:
break
# +
# rasters
# -
# Use reference rasters to ID columns to extract
access_cols = [acc_col for acc_col in rasters]
access_cols[::10]
# ## Aggregation code
# ### Binning by travel time
# Filter down data
# +
# Make a dataframe with just the columns we'll be using
# Need to change range numbers if we add columns
dft = dd.multi.concat([df_pixels[["POP","ADM2_EN","ADM3_EN","Adm2_Code","Adm3_Code"]],df_pixels[access_cols]],axis=1)
# -
dft = dft.compute()
# Travel time ranges
# +
tt_bins = [0, 0.5, 1, 2, 4, 8, 16, 10000]
tt_bin_labels = ["0 - 30 minutes", "31 - 60 minutes", "1 - 2 hours", "2 - 4 hours", "4 - 8 hours", "8 - 16 hours", "16+ hours"]
# rename dict
tt_rename_dct = {
1 : "0 - 30 minutes",
2 : "31 - 60 minutes",
3 : "1 - 2 hours",
4 : "2 - 4 hours",
5 : "4 - 8 hours",
6 : "8 - 16 hours",
7 : "16+ hours"}
# +
# # Alternately calculate using PLSM TT breaks for direct comparison with survey data
# tt_bins = [0, 0.249999, 0.49999, 0.749999, 0.999999, 10000]
# tt_bin_labels = ["0 - 14 minutes", "15 - 29 minutes", "30 - 44 minutes", "45 - 59 hours", "60+ minutes"]
# # rename dict
# tt_rename_dct = {
# 1 : "0 - 14 minutes",
# 2 : "15 - 29 minutes",
# 3 : "30 - 44 minutes",
# 4 : "45 - 59 hours",
# 5 : "60+ minutes"}
# -
# Functions
def long_per_indicator(df,indicator,adm_col):
indic_label = indicator.replace('_COG','')
# pivot the data for just that indicator, with the column VALUES = the population value for that pixel
pop_total = df.pivot_table(index = adm_col, columns=indicator,
values = 'POP', aggfunc = 'sum', fill_value = 0)
# divide by the rowsum to get the % of population falling in each travel category, per admin area
pop_pct = pop_total.div(np.nansum(pop_total,axis=1),axis=0)
# create labels
pop_total['indicator'] = indic_label
pop_pct['indicator'] = indic_label
# remove the multi-index, compress in long format with the adm and indicator as labels, and then change labels/sort
pop_pct = pop_pct.reset_index()\
.melt(id_vars=[adm_col,'indicator'])\
.rename({indicator:'travel_time_range','value':'pop_pct'},axis=1)
pop_total = pop_total.reset_index()\
.melt(id_vars=[adm_col,'indicator'])\
.rename({indicator:'travel_time_range','value':'pop_total'},axis=1)
long_indic = pd.concat([pop_pct,pop_total[['pop_total']]],axis=1,ignore_index=False)
return long_indic
# Replace values with bin number (via numpy), then rename bin numbers using dict created above
# %timeit
dft[access_cols] = pd.DataFrame(np.digitize(dft[access_cols], bins=tt_bins),columns=dft[access_cols].columns, index=dft.index)
# %timeit
dft[access_cols] = dft[access_cols].apply(lambda x: x.replace(tt_rename_dct))
dft
# For each indicator, pivot data by administrative unit, calculate the pct of total population per travel time bin, and reshape the data into a long format.</br>Then merge all these reshaped long tables into one master table
# +
long_data_lst_adm2 = []
long_data_lst_adm3 = []
for i in access_cols:
long_i_adm2 = long_per_indicator(dft,i,'Adm2_Code')
long_i_adm3 = long_per_indicator(dft,i,'Adm3_Code')
long_data_lst_adm2.append(long_i_adm2)
long_data_lst_adm3.append(long_i_adm3)
# -
tt_bin_labels
# +
# concatenate
long_acc_indicators_adm2 = pd.concat(long_data_lst_adm2,ignore_index=True)
long_acc_indicators_adm3 = pd.concat(long_data_lst_adm3,ignore_index=True)
# convert tt ranges to categorical and order appropriately
long_acc_indicators_adm2['travel_time_range'] = long_acc_indicators_adm2.travel_time_range.astype('category').cat.set_categories(tt_bin_labels)
long_acc_indicators_adm3['travel_time_range'] = long_acc_indicators_adm3.travel_time_range.astype('category').cat.set_categories(tt_bin_labels)
# order as desired
long_acc_indicators_adm2 = long_acc_indicators_adm2.sort_values(['Adm2_Code','indicator','travel_time_range']).reset_index(drop=True)
long_acc_indicators_adm3 = long_acc_indicators_adm3.sort_values(['Adm3_Code','indicator','travel_time_range']).reset_index(drop=True)
# -
long_acc_indicators_adm2
# Calculate cumulative sums per indicator and the Adm2_Code for Adm3 datasets
# +
long_acc_indicators_adm2['pop_pct_csum'] = long_acc_indicators_adm2.groupby(['Adm2_Code','indicator'])['pop_pct'].cumsum(axis=0)
long_acc_indicators_adm3['pop_pct_csum'] = long_acc_indicators_adm3.groupby(['Adm3_Code','indicator'])['pop_pct'].cumsum(axis=0)
long_acc_indicators_adm2['pop_total_csum'] = long_acc_indicators_adm2.groupby(['Adm2_Code','indicator'])['pop_total'].cumsum(axis=0)
long_acc_indicators_adm3['pop_total_csum'] = long_acc_indicators_adm3.groupby(['Adm3_Code','indicator'])['pop_total'].cumsum(axis=0)
# -
long_acc_indicators_adm3['Adm2_Code'] = long_acc_indicators_adm3['Adm3_Code'].str[:5]
long_acc_indicators_adm2
long_acc_indicators_adm3
# Export final long data
long_acc_indicators_adm2.to_csv(os.path.join(data_dir,tab_dir,f'final//adm2_acc_indicators_long_{today}.csv'))
long_acc_indicators_adm3.to_csv(os.path.join(data_dir,tab_dir,f'final//adm3_acc_indicators_long_{today}.csv'))
# + [markdown] tags=[]
# ## Compute standard deviation access values, weighted by pixel population
# +
# Make a dataframe with just the columns we'll be using
# Need to change range numbers if we add columns
dfsd = dd.multi.concat([df_pixels[["POP","ADM2_EN","ADM3_EN","Adm2_Code","Adm3_Code","adm2_pop","adm3_pop","wt_adm_2","wt_adm_3"]],df_pixels[access_cols]],axis=1)
# -
dfsd = dfsd.compute()
# +
# # Get Pops by Adm2_Code
# adm2_pop = dfsd.groupby('Adm2_Code')['POP'].sum().to_frame("adm2_pop")
# # Get Pops by CBS Ward
# adm3_pop = dfsd.groupby('Adm3_Code')['POP'].sum().to_frame("adm3_pop")
# +
# # Merge the Pops into Ref DF
# df_pixels = dd.merge(df_pixels, adm2_pop, how = 'left', left_on="Adm2_Code", right_index=True)
# df_pixels = dd.merge(df_pixels, adm3_pop, how = 'left', left_on="Adm3_Code", right_index=True)
# #points = points.persist()
# df_pixels = df_pixels.persist()
# +
# # Calculate the population weight of each pixel within its enclosing admin area -- e.g. 10 pixel population for a 100 population admin - 0.1 weight
# df_pixels['wt_adm_2'] = (df_pixels['POP'] / df_pixels['adm2_pop'])
# df_pixels['wt_adm_3'] = (df_pixels['POP'] / df_pixels['adm3_pop'])
# +
adm2_avg_cols = []
adm3_avg_cols = []
for rkey in rasters:
hrs_col = f"{rkey}"
avg_col_adm_2 = f"{rkey}_std_adm2"
avg_col_adm_3 = f"{rkey}_std_adm3"
dfsd[avg_col_adm_2] = dfsd[hrs_col] * dfsd['wt_adm_2']
dfsd[avg_col_adm_3] = dfsd[hrs_col] * dfsd['wt_adm_3']
adm2_avg_cols.append(avg_col_adm_2)
adm3_avg_cols.append(avg_col_adm_3)
# -
print(",".join(adm2_avg_cols))
print(",".join(adm3_avg_cols))
adm2_sd_final = dfsd.groupby(['Adm2_Code'])[adm2_avg_cols].std().reset_index()
adm3_sd_final = dfsd.groupby(['Adm3_Code'])[adm3_avg_cols].std().reset_index()
# + [markdown] tags=[]
# #### Export
# -
# Export names
adm2_sd_output = "../../data/tabular/processed/adm2_sd_211015.csv"
adm3_sd_output = "../../data/tabular/processed/adm3_sd_211015.csv"
# Actually export
# %time
adm2_sd_final.to_csv(adm2_sd_output)
adm3_sd_final.to_csv(adm3_sd_output)
# + [markdown] tags=[]
# ### Compute mean access values, weighted by pixel population
# +
# Get Pops by Adm2_Code
adm2_pop = df_pixels.groupby('Adm2_Code')['POP'].sum().to_frame("adm2_pop")
# Get Pops by CBS Ward
adm3_pop = df_pixels.groupby('Adm3_Code')['POP'].sum().to_frame("adm3_pop")
# +
# Merge the Pops into Ref DF
df_pixels = dd.merge(df_pixels, adm2_pop, how = 'left', left_on="Adm2_Code", right_index=True)
df_pixels = dd.merge(df_pixels, adm3_pop, how = 'left', left_on="Adm3_Code", right_index=True)
#points = points.persist()
df_pixels = df_pixels.persist()
# -
# Calculate the population weight of each pixel within its enclosing admin area -- e.g. 10 pixel population for a 100 population admin - 0.1 weight
df_pixels['wt_adm_2'] = (df_pixels['POP'] / df_pixels['adm2_pop'])
df_pixels['wt_adm_3'] = (df_pixels['POP'] / df_pixels['adm3_pop'])
# +
adm2_avg_cols = []
adm3_avg_cols = []
for rkey in rasters:
hrs_col = f"{rkey}"
avg_col_adm_2 = f"{rkey}_avg_adm2"
avg_col_adm_3 = f"{rkey}_avg_adm3"
df_pixels[avg_col_adm_2] = df_pixels[hrs_col] * df_pixels['wt_adm_2']
df_pixels[avg_col_adm_3] = df_pixels[hrs_col] * df_pixels['wt_adm_3']
adm2_avg_cols.append(avg_col_adm_2)
adm3_avg_cols.append(avg_col_adm_3)
# -
print(",".join(adm2_avg_cols))
print(",".join(adm3_avg_cols))
adm2_final = df_pixels.groupby(['Adm2_Code'])[adm2_avg_cols].sum().reset_index()
adm3_final = df_pixels.groupby(['Adm3_Code'])[adm3_avg_cols].sum().reset_index()
# #### Export
# Export names
adm2_output = "../../data/outputs/adm2_final.csv"
adm3_output = "../../data/outputs/adm3_final.csv"
df_pixels_out = "../../data/outputs/df_pixels_final.csv"
# Actually export
# %time
adm2_final.to_csv(adm2_output, single_file=True)
adm3_final.to_csv(adm3_output, single_file=True)
client.close()
| notebooks/Access_Modeling/Step 3 - Point Access Data Aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ## Datatypes and attributes
# Numpy's main datatype is ndarray
a1 = np.array([1,2,3,4,5])
a1
type(a1)
a2 = np.array([[1,2,3.0],
[4,5.6,6],
[7,8.8,9]])
a3 = np.array([[[1,2,3],[4,5,6]],[[5,5,5],[6,6,6]]])
a2
a3
a1.shape
a2.shape
a3.shape
a1.ndim, a2.ndim, a3.ndim
a1.dtype, a2.dtype, a3.dtype
a1.size, a2.size, a3.size
type(a1), type(a2), type(a3)
# Create A DataFrame from a NumPy array
import pandas as pd
df = pd.DataFrame(a2)
df
# ## 2. Create NumPy Arrays
sample_array = np.array([1, 2, 3])
sample_array
sample_array.dtype
ones = np.ones((2,3))
ones
range_array = np.arange(0,10,2)
range_array
random_array = np.random.randint(0,10,size=(3,5))
random_array
# ## Viewing array and matrices
np.unique(random_array)
# ## 4. Manipulating and comparing Arrays
# ### Arithmic
a1
ones = np.ones(5)
a1 + ones
a1 = a1[:3]
a1
a2
a1 * a2
np.log(a1)
np.exp(a1)
a2 * a3
a2
a2.reshape(1,3,3)
a3
a2.reshape(3,3,1) * a3
# ### Aggregation
# Aggregation = performing the same operations on a number of things
sum(a1)
np.sum(a1)
# Use Python's methods on python's datatypes and numpy methods on numpy arrays
massive_array = np.random.rand(100000)
massive_array[:10]
# %timeit sum(massive_array) # Python's method sum()
# %timeit np.sum(massive_array) # NumPy's method np.sum()
# Variance = measure of the average degree to which each number is different to the mean
# High Variance = wider range of numbers
# Low Variance = lower range of numbers
np.var(a1)
# Standard Deviation = measure of how spread out are a group of numbers from the mean
np.std(a1)
np.sqrt(np.var(a1))
high_var_arr = np.array([1,100,200,300,4000,5000])
low_var_arr = np.array([2,4,6,8,10])
np.var(high_var_arr), np.var(low_var_arr)
np.std(high_var_arr), np.std(low_var_arr)
np.mean(high_var_arr), np.mean(low_var_arr)
# %matplotlib inline
import matplotlib.pyplot as plt
plt.hist(high_var_arr)
plt.show()
plt.hist(low_var_arr)
plt.show()
a2
a2 = a2[:2]
a2.shape
a3.shape
a3 = np.array([[[5,5,5],[6,6,6],[3,2,3]],[[7,7,7],[9,8,8],[1,2,4]]])
a3
a3.shape
a2 * a3
a2.reshape(2,3,1)
a2.reshape(2,3,1).shape
a3.shape
a2_reshape = a2.reshape(2,3,1)
a2_reshape * a3
a2.shape
a2.T
a2.T.shape
# ## Dot Product
np.random.seed(0)
mat1 = np.random.randint(10,size=(5,3))
mat2 = np.random.randint(10,size=(5,3))
mat1
mat2
# Element-wise multiplication (Hadamard Product)
mat1 * mat2
mat1.shape, mat2.shape
mat2.T
mat1
mat3 = np.dot(mat1,mat2.T)
mat3
mat3.shape
# ## Dot Product (Nut Butter Example)
np.random.seed(0)
# Sales for the week
sales_amount = np.random.randint(20, size=(5, 3))
sales_amount
# weekly sales Data Frame
weekly_sales = pd.DataFrame(sales_amount,
index=["Mon", "Tues", "Wed", "Thurs", "Fri"],
columns=["Almond Butter", "Peanut Butter", "Cashew Butter"])
weekly_sales
# Create prices array
prices = np.array([10, 8, 12])
prices = prices.reshape(1,3)
prices
butter_prices = pd.DataFrame(prices,
index=["Prices"],
columns=["Almond Butter", "Peanut Butter", "Cashew Butter"])
butter_prices
prices
sales_amount.T
total_sales = prices.dot(sales_amount.T)
total_sales
daily_sales = butter_prices.dot(weekly_sales.T)
daily_sales
weekly_sales["Total ($)"] = daily_sales.T
weekly_sales
# ## Comparing operators
a1 < a2
a1 <= a2
a1
a2
# ## 5. Sorting Arrays
random_array = np.random.randint(10, size=(3,5))
random_array
np.sort(random_array)
np.argsort(random_array)
random_array
np.argmax(random_array, axis=1)
# ## 6. Practical Example - NumPy in Action!!!
# <img src="numpy-images/panda.png" width="400px"/>
# +
# Turn an image to a NumPy array
from matplotlib.image import imread
panda = imread("numpy-images/panda.png")
type(panda), panda.dtype
# -
panda
panda.size, panda.shape, panda.ndim
# <img src="numpy-images/car-photo.png" />
car = imread("numpy-images/car-photo.png")
car.size, car.shape, car.ndim, type(car), car.dtype
# <img src="numpy-images/dog-photo.png"/>
dog = imread("numpy-images/dog-photo.png")
dog.size, dog.shape, dog.ndim, type(dog), dog.dtype
| Introduction-to-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## <center>Computer Science Intensive Course - MindX</center>
# 
# # <center>LAB 12. THUẬT TOÁN TÌM ĐƯỜNG (2)</center>
# run this cell FIRST
# %run test_cases_12.ipynb
# ## Bài 1. Kích Thước Tầng
#
# Kích thước của một tầng trong cây nhị phân được định nghĩa là số node trên tầng đó.
#
# **Yêu cầu**: Cho một cây nhị phân có không quá 1000 node và một số nguyên *k*. Hãy tìm kích thước của tầng thứ *k*, biết các tầng được đánh số từ 0, bắt đầu từ node gốc.
#
# **Input**: Một cây nhị phân chứa số nguyên và một số nguyên *-10<sup>6</sup> < k < 10<sup>6</sup>*.
# **Output**: Một số nguyên là kích thước của tầng *k*. Trả về 0 nếu cây không chứa tầng thứ *k*.
# **Ví dụ**:
#
# - Input: <code>root</code>, 2
# - Output: 3
# - Giải thích: Ở tầng thứ hai có 3 node là [1, 6, 14]
#
# 
# +
class Node:
def __init__(self, data):
self.data = data
self.left = None
self.right = None
root = Node(8)
root.left, root.right = Node(3), Node(10)
root.left.left, root.left.right = Node(1), Node(6)
root.left.right.left, root.left.right.right = Node(4), Node(7)
root.right.right = Node(14)
root.right.right.left = Node(13)
# -
def find_level_size(root, level):
pass
# !!! DO NOT MODIFY THIS CELL
# Check result on test cases
test1(find_level_size)
# ## Bài 2. Thoát Khỏi Mê Cung
#
# Ở bài trước, ta đã dùng DFS để tạo mê cung. Tuy nhiên, thuật toán DFS không đảm bảo ta tìm được đường ngắn nhất để thoát khỏi mê cung.
#
# Do đó, ta sẽ dùng thuật toán BFS để tìm đường ngắn nhất:
# - Bắt đầu duyệt mê cung từ ô trên cùng bên trái.
# - Ở mỗi bước duyệt, kiểm tra xem có thể đi từ ô hiện tại sang các ô cùng cạnh hay không bằng cách kiểm tra các mảng <code>vertical</code> và <code>horizontal</code>. Khi sang một ô mới, lưu ô liền trước nó để truy vấn lại đường đi.
# - Kết thúc duyệt khi tìm thấy ô dưới cùng bên phải. Do mê cung ta tạo đảm bảo luôn có lối thoát nên không cần xử lý trường hợp không tìm thấy đường ra.
#
# **Ví dụ** về đường đi ngắn nhất để thoát khỏi mê cung:
# 
#
# Hàm <code>plot_maze_with_path()</code> để vẽ mê cung từ <code>vertical</code>, <code>horizontal</code> và vẽ đường đi từ <code>path</code> đã được viết sẵn.
# +
import matplotlib.pyplot as plt
plt.style.use('default')
def plot_maze_with_path(vertical, horizontal, path=None, before=None, fig_height=8):
# init height & width
height = len(vertical)
width = len(vertical[0])
# init figure
fig = plt.figure(figsize=(fig_height*2, fig_height))
fig.patch.set_visible(False)
# draw maze borders
for row in range(height):
for col in range(width):
if vertical[row][col]:
plt.plot((col, col), (row, row+1), color='white')
if horizontal[row][col]:
plt.plot((col, col+1), (row, row), color='white')
# draw surrounding borders on the right & bottom
plt.plot((width, width), (0, height-1), color='white')
plt.plot((0, width), (height, height), color='white')
# styling the plot
ax = plt.gca()
ax.set_facecolor((0, 0, 0))
ax.set_ylim(ax.get_ylim()[::-1])
plt.xticks([])
plt.yticks([])
# add arrows
plt.arrow(0, 0.5, 0.8, 0, width=0.07, length_includes_head=True, color='white')
plt.arrow(width-0.8, height-0.5, 0.8, 0, width=0.07, length_includes_head=True, color='white')
# plot path
if path != None:
last_step = (0, -0.5)
for step in path + [(height-1, width-0.5)]:
plt.plot((last_step[1]+0.5, step[1]+0.5), (last_step[0]+0.5, step[0]+0.5), color='cyan', linestyle='-.')
last_step = step
# plot every paths
if before != None:
for row in range(height):
for col in range(width):
if before[row][col] != None:
last_step = before[row][col]
plt.plot((last_step[1]+0.5, col+0.5), (last_step[0]+0.5, row+0.5), color='yellow', linestyle='-.')
plt.show()
# -
# Các mảng <code>vertical</code> và <code>horizontal</code> được định nghĩa như bài trước.
#
# Ta cần tìm mảng <code>path</code> chứa đường đi ngắn nhất để thoát khỏi mê cung. Mỗi phần tử trong mảng là một tuple chứa tọa độ của một ô trên đường đi, được lưu theo dạng (<code>row</code>, <code>column</code>).
# +
vertical = [
[False, True, False, False, True, False, False, False],
[True, True, False, False, False, True, False, True],
[True, False, False, False, False, True, True, False],
[True, False, False, False, False, False, True, False]]
horizontal = [
[True, True, True, True, True, True, True, True],
[False, False, True, True, False, True, True, False],
[False, True, True, True, True, False, False, False],
[True, True, True, True, False, False, True, False]]
path = [(0, 0), (1, 0), (2, 0), (2, 1), (2, 2), (2, 3), (2, 4), (3, 4), (3, 5), (2, 5), (1, 5), (1, 6), (2, 6), (2, 7), (3, 7)]
plot_maze_with_path(vertical, horizontal, path=path, fig_height=4)
# -
# **Yêu cầu**: Hãy hiện thực hàm <code>find_path()</code> nhận vào <code>vertical</code>, <code>horizontal</code> và trả về đường đi <code>path</code> theo định dạng như trên.
def find_path(vertical, horizontal):
# YOUR CODE HERE
return path
# Kết quả mong đợi như bên dưới:
path = find_path(vertical, horizontal)
plot_maze_with_path(vertical, horizontal, path=path, fig_height=4)
vertical = [[False, True, False, True, False, False, False, False, False, True, False, False, False, False, False, False, False, False, False, False, True, False, False, False, False, False, False, False, False, True, False, False, True, False, True, False, False, False, True, False], [True, True, False, False, False, True, False, True, False, False, False, True, False, True, False, True, True, False, False, True, False, False, False, True, False, True, False, False, True, True, False, True, False, True, True, True, True, True, False, True], [True, False, False, False, False, True, True, False, True, False, False, True, True, True, False, True, True, True, False, True, False, False, False, True, False, True, True, False, True, False, True, False, True, True, False, False, True, False, True, False], [True, False, True, False, True, False, True, True, False, True, True, False, True, False, True, True, True, False, True, False, True, True, False, False, True, True, False, True, False, False, True, True, True, True, False, True, True, False, True, False], [True, False, False, True, False, False, False, True, True, False, False, False, False, True, False, False, True, True, False, True, True, False, True, False, True, False, True, False, False, True, False, True, True, True, True, False, True, False, False, True], [True, True, False, False, False, False, True, True, False, True, False, True, True, False, True, False, False, False, True, False, True, True, True, False, False, False, True, True, False, False, True, False, True, True, True, False, True, False, False, True], [True, True, False, False, False, True, False, False, False, False, False, True, True, True, False, True, False, True, False, True, False, True, False, True, False, True, True, False, True, True, True, True, False, False, True, False, False, False, False, False], [True, False, False, False, True, True, False, False, True, False, True, True, False, True, False, False, True, False, False, True, True, False, True, True, False, True, False, True, True, False, False, True, True, False, True, False, False, False, False, True], [True, False, False, False, False, True, True, True, True, True, False, False, True, True, False, True, False, False, True, True, False, True, True, False, False, True, False, True, False, False, False, False, True, True, False, True, False, True, False, False], [True, True, False, False, True, False, True, True, False, True, True, False, False, True, True, True, False, True, True, False, True, False, True, False, False, True, False, True, True, False, False, True, True, False, True, True, False, True, False, False], [True, True, False, False, True, False, False, True, False, True, False, False, True, True, True, False, True, True, True, True, True, False, False, True, False, False, True, False, True, False, True, False, False, False, True, True, False, True, False, True], [True, False, False, False, False, False, True, True, False, False, True, False, False, True, True, False, False, True, True, True, False, True, True, True, False, False, False, True, False, False, False, False, False, True, False, True, False, True, False, True], [True, False, True, True, False, True, False, True, False, False, True, False, True, False, False, False, False, True, True, False, False, True, False, True, False, True, True, True, False, False, False, True, False, True, True, True, False, False, True, True], [True, True, True, True, False, False, True, True, False, True, False, True, False, True, False, True, False, False, True, True, False, True, True, False, True, True, True, False, True, False, True, False, True, False, True, False, False, True, False, True], [True, False, True, False, True, False, False, False, True, True, True, True, False, True, True, False, True, False, False, True, True, True, True, True, True, True, False, False, False, True, False, False, True, False, False, True, False, False, True, False], [True, True, False, True, False, True, False, True, True, True, False, False, False, False, True, False, True, True, True, False, False, True, True, False, True, False, True, False, True, False, True, False, True, False, True, False, False, True, False, False], [True, False, True, False, True, False, True, False, True, False, True, False, False, False, False, True, True, True, True, True, True, False, True, True, False, False, True, True, True, False, True, True, False, False, True, False, True, True, False, False], [True, True, False, True, False, True, False, True, True, True, False, False, True, False, True, False, True, False, True, False, True, True, False, True, True, False, False, False, True, True, False, False, False, True, False, True, False, True, False, False], [True, False, True, False, False, False, False, True, False, True, False, True, True, True, False, True, False, True, True, False, True, True, True, False, True, False, False, False, True, False, True, False, True, False, False, False, False, True, False, True], [True, True, False, False, False, False, True, False, False, False, True, False, True, False, True, False, False, False, False, True, False, False, True, False, False, False, False, False, False, True, False, True, False, False, False, False, True, False, False, False]]
horizontal = [[True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True, True], [False, False, True, False, True, True, True, False, True, False, True, True, True, False, True, False, True, True, True, False, True, True, True, False, True, False, True, True, False, True, False, False, True, False, False, True, False, False, False, False], [False, True, True, True, True, False, False, True, True, True, True, False, False, True, False, False, False, True, False, True, True, True, False, True, True, False, True, False, False, False, True, True, False, False, False, False, False, True, True, False], [True, True, True, True, False, False, True, False, False, True, False, False, False, True, False, False, True, False, True, False, True, True, True, False, False, True, False, True, True, True, False, False, False, True, True, False, True, False, True, True], [False, True, False, False, True, True, False, True, True, False, True, True, True, False, True, False, False, True, False, True, False, False, True, False, False, False, True, True, False, False, True, False, False, False, False, True, False, True, False, False], [False, True, True, True, True, False, False, False, False, False, True, False, True, True, True, True, False, False, True, False, False, True, False, True, True, True, False, True, True, True, False, False, False, False, True, False, False, True, True, False], [False, False, True, True, True, True, False, True, True, True, False, False, False, False, False, True, True, True, False, False, False, False, True, True, True, False, False, True, False, False, False, True, False, False, False, True, True, True, True, False], [False, True, True, True, False, True, True, True, True, True, False, True, False, True, True, False, False, True, True, False, True, False, False, False, True, False, True, False, False, True, False, False, True, True, True, True, True, True, True, False], [True, True, True, True, False, False, True, False, False, False, True, False, True, False, True, True, True, True, False, False, True, False, False, True, False, True, False, False, True, True, True, False, False, False, True, False, True, False, True, False], [False, True, True, False, True, False, False, False, False, True, True, True, False, False, False, False, True, False, False, True, False, False, True, True, False, False, True, False, True, True, True, False, False, True, False, True, False, True, True, True], [False, False, True, True, False, True, False, True, True, False, False, True, False, False, False, True, False, False, True, False, True, True, True, False, True, True, False, True, False, True, False, True, True, False, False, False, True, False, True, False], [False, True, True, True, True, True, False, False, True, False, True, True, False, False, False, True, False, False, False, False, False, True, False, True, True, True, True, False, True, False, True, True, True, True, False, True, False, True, False, False], [False, True, True, True, False, True, False, True, True, True, False, True, True, False, True, True, True, False, False, True, True, False, False, False, True, True, False, True, True, True, True, True, True, False, True, False, True, True, False, False], [True, False, False, False, True, False, True, False, True, False, True, False, False, True, True, True, True, False, False, False, True, False, True, False, False, False, False, False, True, True, False, False, False, False, False, True, True, False, True, False], [False, False, False, True, True, True, False, False, False, True, False, False, True, False, False, False, True, True, True, False, False, False, False, True, False, False, True, True, False, False, True, True, True, True, True, False, True, True, False, False], [False, True, False, False, False, True, True, False, False, False, False, True, True, False, True, True, False, False, False, True, False, False, False, False, False, True, False, True, False, True, True, False, False, True, False, True, False, True, True, False], [False, True, True, True, True, False, False, True, False, True, True, True, True, True, False, False, False, False, True, False, True, False, False, True, True, False, True, False, True, False, False, True, True, False, True, True, False, False, True, True], [True, False, False, False, False, True, True, False, False, False, True, True, False, True, True, False, True, False, False, False, False, True, True, False, True, True, False, False, False, True, False, True, False, True, False, False, False, True, True, False], [False, True, True, True, True, False, True, False, True, True, True, False, False, False, True, False, False, False, True, True, False, False, False, False, False, True, True, True, False, True, True, True, True, True, True, True, True, False, True, True], [False, False, False, True, True, True, False, True, False, False, False, False, False, True, False, True, True, True, False, False, False, False, True, True, True, True, True, False, True, False, False, False, False, True, True, True, False, True, False, False]]
path = find_path(vertical, horizontal)
plot_maze_with_path(vertical, horizontal, path=path)
| Lab12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# !pip install albumentations > /dev/null
# !pip install -U segmentation-models
# !pip install -U efficientnet
import numpy as np
import pandas as pd
import gc
import keras
import matplotlib.pyplot as plt
plt.style.use('seaborn-white')
import seaborn as sns
sns.set_style("white")
from sklearn.model_selection import train_test_split,StratifiedKFold
from skimage.transform import resize
import tensorflow as tf
import keras.backend as K
from keras.losses import binary_crossentropy
from keras.preprocessing.image import load_img
from keras import Model
from keras.callbacks import ModelCheckpoint
from keras.layers import Input, Conv2D, Conv2DTranspose, MaxPooling2D, concatenate, Dropout,BatchNormalization
from keras.layers import Conv2D, Concatenate, MaxPooling2D
from keras.layers import UpSampling2D, Dropout, BatchNormalization
from tqdm import tqdm_notebook
from keras import initializers
from keras import regularizers
from keras import constraints
from keras.utils import conv_utils
from keras.utils.data_utils import get_file
from keras.engine.topology import get_source_inputs
from keras.engine import InputSpec
from keras import backend as K
from keras.layers import LeakyReLU
from keras.layers import ZeroPadding2D
from keras.losses import binary_crossentropy
import keras.callbacks as callbacks
from keras.callbacks import Callback
from keras.applications.xception import Xception
from keras.layers import multiply
from keras import optimizers
from keras.legacy import interfaces
from keras.utils.generic_utils import get_custom_objects
import segmentation_models as sm
from keras.engine.topology import Input
from keras.engine.training import Model
from keras.layers.convolutional import Conv2D, UpSampling2D, Conv2DTranspose
from keras.layers.core import Activation, SpatialDropout2D
from keras.layers.merge import concatenate
from keras.layers.normalization import BatchNormalization
from keras.layers.pooling import MaxPooling2D
from keras.layers import Input,Dropout,BatchNormalization,Activation,Add
from keras.regularizers import l2
from keras.layers.core import Dense, Lambda
from keras.layers.merge import concatenate, add
from keras.layers import GlobalAveragePooling2D, Reshape, Dense, multiply, Permute
from keras.optimizers import SGD,Adam
from keras.models import Model,load_model
from keras.preprocessing.image import ImageDataGenerator
import glob
import shutil
import os
import random
from PIL import Image
import cv2
from random import shuffle
seed = 10
np.random.seed(seed)
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
# %matplotlib inline
# -
# ## Creating Dataframe containing file_path, mask percentage and corresponding label(pneumothorax or no pneumothorax)
# +
all_mask_fn = glob.glob('/kaggle/input/siimacr-pneumothorax-segmentation-data-512/masks/*')
mask_df = pd.DataFrame()
mask_df['file_names'] = all_mask_fn
mask_df['mask_percentage'] = 0
mask_df.set_index('file_names',inplace=True)
for fn in all_mask_fn:
mask_df.loc[fn,'mask_percentage'] = np.array(Image.open(fn)).sum()/(512*512*255) #255 is bcz img range is 255
mask_df.reset_index(inplace=True)
sns.distplot(mask_df.mask_percentage)
mask_df['labels'] = 0
mask_df.loc[mask_df.mask_percentage>0,'labels'] = 1
# -
# ## Train-test splitting of 85%-15%
train_df,val_df = train_test_split(mask_df,test_size = 0.15,stratify = mask_df.labels,random_state = 100)
print('No. of train files:', len(train_df))
print('No. of val files:', len(val_df))
train_filepath = train_df['file_names'].tolist()
val_filepath = val_df['file_names'].tolist()
train_im_path = 'train'
train_mask_path = 'masks'
img_size = 512
# ## Adding Augmentations
from albumentations import (
Compose, HorizontalFlip, CLAHE, HueSaturationValue,
RandomBrightness, RandomContrast, RandomGamma,OneOf,
ToFloat, ShiftScaleRotate,GridDistortion, ElasticTransform, JpegCompression, HueSaturationValue,
RGBShift, RandomBrightness, RandomContrast, Blur, MotionBlur, MedianBlur, GaussNoise,CenterCrop,
IAAAdditiveGaussianNoise,GaussNoise,OpticalDistortion,RandomSizedCrop
)
train_augment = Compose([
HorizontalFlip(p = 0.5),
ShiftScaleRotate(p = 0.5),
#ElasticTransform(alpha=120, sigma=120 * 0.05, alpha_affine=120 * 0.03,p = 0.5),
OneOf([
RandomContrast(),
RandomGamma(),
RandomBrightness(),
], p=0.3),
RandomSizedCrop(min_max_height=(176, 256), height=512, width=512,p=0.25),
ToFloat()
])
# ## DataGenerator Class
class DataGenerator(keras.utils.Sequence):
def __init__(self,filepath = train_filepath,train_im_path = train_im_path,train_mask_path = train_mask_path,
augmentations = None,img_size = img_size,batch_size = 64,nchannels = 3,shuffle = True):
self.train_im_paths = list(filepath)
self.train_im_path = train_im_path
self.train_mask_path = train_mask_path
self.img_size = img_size
self.batch_size = batch_size
self.nchannels = nchannels
self.shuffle = shuffle
self.augmentations = augmentations
self.on_epoch_end()
def __len__(self):
return int(np.ceil(len(self.train_im_paths)/ self.batch_size))
def __getitem__(self,index):
indexes = self.indexes[index * self.batch_size : min((index + 1) * self.batch_size, len(self.train_im_paths))]
list_im_ids = [self.train_im_paths[i] for i in indexes]
X,y = self.data_generation(list_im_ids)
if(self.augmentations is None):
return np.array(X,dtype = 'float32'),np.array(y) / 255
im,mask = [],[]
for x,y in zip(X,y):
augmented = self.augmentations(image = x,mask = y)
im.append(augmented['image'])
mask.append(augmented['mask'])
return np.array(im,dtype = 'float32'),np.array(mask) / 255
def on_epoch_end(self):
self.indexes = np.arange(len(self.train_im_paths))
if(self.shuffle):
np.random.shuffle(self.indexes)
def data_generation(self,list_im_ids):
X = np.empty((len(list_im_ids),self.img_size,self.img_size,self.nchannels))
y = np.empty((len(list_im_ids),self.img_size,self.img_size,1))
for i,mask_path in enumerate(list_im_ids):
#print(mask_path)
mask = np.array(Image.open(mask_path))
#plt.imshow(mask)
img_path = mask_path.replace(self.train_mask_path,self.train_im_path)
img = cv2.imread(img_path)
if(len(img.shape) == 2):
img = np.repeat(img[...,np.newaxis],3,2)
# plt.imshow(img,cmap = 'bone')
X[i,] = cv2.resize(img,(self.img_size,self.img_size))
y[i,] = cv2.resize(mask,(self.img_size,self.img_size))[...,np.newaxis]
y[y > 0] = 255
return np.uint8(X),np.uint8(y)
# ## Testing the Generator
# +
a = DataGenerator(batch_size=64,shuffle=False,augmentations=train_augment)
images,masks = a.__getitem__(0)
max_images = 64
grid_width = 16
grid_height = int(max_images / grid_width)
fig, axs = plt.subplots(grid_height, grid_width, figsize=(grid_width,grid_height))
for i,(im, mask) in enumerate(zip(images,masks)):
ax = axs[int(i / grid_width), i % grid_width]
ax.imshow(im.squeeze(), cmap="bone")
ax.imshow(mask.squeeze(), alpha=0.5, cmap="Reds")
ax.axis('off')
plt.suptitle("Chest X-rays, Masks")
# -
# ## EfficientUNet model code
# +
from efficientnet.keras import EfficientNetB4
def UEfficientNet(input_shape=(None, None, 3),dropout_rate=0.1):
backbone = EfficientNetB4(weights='imagenet',
include_top=False,
input_shape=input_shape)
input = backbone.input
start_neurons = 8
conv4 = backbone.layers[342].output
conv4 = LeakyReLU(alpha=0.1)(conv4)
pool4 = MaxPooling2D((2, 2))(conv4)
pool4 = Dropout(dropout_rate)(pool4)
# Middle
convm = Conv2D(start_neurons * 32, (3, 3), activation=None, padding="same",name='conv_middle')(pool4)
convm = residual_block(convm,start_neurons * 32)
convm = residual_block(convm,start_neurons * 32)
convm = LeakyReLU(alpha=0.1)(convm)
deconv4 = Conv2DTranspose(start_neurons * 16, (3, 3), strides=(2, 2), padding="same")(convm)
deconv4_up1 = Conv2DTranspose(start_neurons * 16, (3, 3), strides=(2, 2), padding="same")(deconv4)
deconv4_up2 = Conv2DTranspose(start_neurons * 16, (3, 3), strides=(2, 2), padding="same")(deconv4_up1)
deconv4_up3 = Conv2DTranspose(start_neurons * 16, (3, 3), strides=(2, 2), padding="same")(deconv4_up2)
uconv4 = concatenate([deconv4, conv4])
uconv4 = Dropout(dropout_rate)(uconv4)
uconv4 = Conv2D(start_neurons * 16, (3, 3), activation=None, padding="same")(uconv4)
uconv4 = residual_block(uconv4,start_neurons * 16)
# uconv4 = residual_block(uconv4,start_neurons * 16)
uconv4 = LeakyReLU(alpha=0.1)(uconv4) #conv1_2
deconv3 = Conv2DTranspose(start_neurons * 8, (3, 3), strides=(2, 2), padding="same")(uconv4)
deconv3_up1 = Conv2DTranspose(start_neurons * 8, (3, 3), strides=(2, 2), padding="same")(deconv3)
deconv3_up2 = Conv2DTranspose(start_neurons * 8, (3, 3), strides=(2, 2), padding="same")(deconv3_up1)
conv3 = backbone.layers[154].output
uconv3 = concatenate([deconv3,deconv4_up1, conv3])
uconv3 = Dropout(dropout_rate)(uconv3)
uconv3 = Conv2D(start_neurons * 8, (3, 3), activation=None, padding="same")(uconv3)
uconv3 = residual_block(uconv3,start_neurons * 8)
# uconv3 = residual_block(uconv3,start_neurons * 8)
uconv3 = LeakyReLU(alpha=0.1)(uconv3)
deconv2 = Conv2DTranspose(start_neurons * 4, (3, 3), strides=(2, 2), padding="same")(uconv3)
deconv2_up1 = Conv2DTranspose(start_neurons * 4, (3, 3), strides=(2, 2), padding="same")(deconv2)
conv2 = backbone.layers[92].output
uconv2 = concatenate([deconv2,deconv3_up1,deconv4_up2, conv2])
uconv2 = Dropout(0.1)(uconv2)
uconv2 = Conv2D(start_neurons * 4, (3, 3), activation=None, padding="same")(uconv2)
uconv2 = residual_block(uconv2,start_neurons * 4)
# uconv2 = residual_block(uconv2,start_neurons * 4)
uconv2 = LeakyReLU(alpha=0.1)(uconv2)
deconv1 = Conv2DTranspose(start_neurons * 2, (3, 3), strides=(2, 2), padding="same")(uconv2)
conv1 = backbone.layers[30].output
uconv1 = concatenate([deconv1,deconv2_up1,deconv3_up2,deconv4_up3, conv1])
uconv1 = Dropout(0.1)(uconv1)
uconv1 = Conv2D(start_neurons * 2, (3, 3), activation=None, padding="same")(uconv1)
uconv1 = residual_block(uconv1,start_neurons * 2)
# uconv1 = residual_block(uconv1,start_neurons * 2)
uconv1 = LeakyReLU(alpha=0.1)(uconv1)
uconv0 = Conv2DTranspose(start_neurons * 1, (3, 3), strides=(2, 2), padding="same")(uconv1)
uconv0 = Dropout(0.1)(uconv0)
uconv0 = Conv2D(start_neurons * 1, (3, 3), activation=None, padding="same")(uconv0)
uconv0 = residual_block(uconv0,start_neurons * 1)
# uconv0 = residual_block(uconv0,start_neurons * 1)
uconv0 = LeakyReLU(alpha=0.1)(uconv0)
uconv0 = Dropout(dropout_rate/2)(uconv0)
output_layer = Conv2D(1, (1,1), padding="same", activation="sigmoid")(uconv0)
model = Model(input, output_layer)
model.name = 'u-efficient'
return model
# -
model = sm.Unet('efficientnetb4', input_shape=(img_size,img_size,3), encoder_weights='imagenet',decoder_block_type='transpose')
model.summary()
# ## Stochatic Weight Averaging Class
class SWA(keras.callbacks.Callback):
def __init__(self, filepath, swa_epoch):
super(SWA, self).__init__()
self.filepath = filepath
self.swa_epoch = swa_epoch
def on_train_begin(self, logs=None):
self.nb_epoch = self.params['epochs']
print('Stochastic weight averaging selected for last {} epochs.'
.format(self.nb_epoch - self.swa_epoch))
def on_epoch_end(self, epoch, logs=None):
if epoch == self.swa_epoch:
self.swa_weights = self.model.get_weights()
elif epoch > self.swa_epoch:
for i in range(len(self.swa_weights)):
self.swa_weights[i] = (self.swa_weights[i] *
(epoch - self.swa_epoch) + self.model.get_weights()[i])/((epoch - self.swa_epoch) + 1)
else:
pass
def on_train_end(self, logs=None):
self.model.set_weights(self.swa_weights)
print('Final model parameters set to stochastic weight average.')
self.model.save_weights(self.filepath)
print('Final stochastic averaged weights saved to file.')
# ## Cosine Annealing Learning Rate Class
# +
class SnapshotCallbackBuilder:
def __init__(self, nb_epochs, nb_snapshots, init_lr=0.01):
self.T = nb_epochs
self.M = nb_snapshots
self.alpha_zero = init_lr
def get_callbacks(self, model_prefix='Model'):
callback_list = [
callbacks.ModelCheckpoint("./keras.model",monitor='val_iou_metric',
mode = 'max', save_best_only=True, verbose=1),
swa,
callbacks.LearningRateScheduler(schedule=self._cosine_anneal_schedule)
]
return callback_list
def _cosine_anneal_schedule(self, t):
cos_inner = np.pi * (t % (self.T // self.M)) # t - 1 is used when t has 1-based indexing.
cos_inner /= self.T // self.M
cos_out = np.cos(cos_inner) + 1
return float(self.alpha_zero / 2 * cos_out)
# -
# ## IOU Evaluation Metric
def get_iou_vector(A,B):
batch_size = A.shape[0]
metric = 0.0
for i in range(batch_size):
t,p = A[i],B[i]
#print(t.dtype)
p = tf.dtypes.cast(p, tf.float32)
intersection = np.sum(t * p)
true = np.sum(t)
pred = np.sum(p)
if(true == 0):
metric += (pred == 0)
union = true + pred - intersection
iou = intersection / union
iou = np.floor(max(0,(iou - 0.45) * 20)) / 10
metric += iou
return metric / batch_size
def iou_metric(label,pred):
return tf.py_function(get_iou_vector,[label,pred > 0.5],tf.float64)
# ## Dice Coefficient and Dice loss function
def dice_coeff(y_true,y_pred):
y_true_f = K.flatten(y_true)
y_pred_f = K.cast(K.greater(K.flatten(y_pred),0.5),'float32')
intersection = K.sum(y_true_f * y_pred_f)
dice_coeff = (intersection * 2) / (K.sum(y_true_f) + K.sum(y_pred_f))
return dice_coeff
def dice_loss(y_true,y_pred):
smooth = 1
y_true_f = K.flatten(y_true)
y_pred_f = K.flatten(y_pred)
intersection = K.sum(y_true_f * y_pred_f)
dice_coeff = (intersection * 2 + smooth) / (K.sum(y_true_f) + K.sum(y_pred_f) + smooth)
return 1 - dice_coeff
def bce_dice_loss(y_true,y_pred):
return binary_crossentropy(y_true,y_pred) + dice_loss(y_true,y_pred)
def bce_logdice_loss(y_true,y_pred):
return binary_crossentropy(y_true,y_pred) - K.log(1. - dice_loss(y_true,y_pred))
# ## Compiling the model
#model.compile(loss = sm.losses.bce_jaccard_loss,optimizer = SGD(learning_rate = 0.0001, momentum=0.0, nesterov=False),metrics = [sm.metrics.iou_score])
model.compile(loss=bce_dice_loss, optimizer='adam', metrics=[iou_metric])
# ## Train Code
train_filepath = train_df['file_names'].tolist()
val_filepath = val_df['file_names'].tolist()
train_im_path = 'train'
train_mask_path = 'masks'
epochs = 50
snapshot = SnapshotCallbackBuilder(nb_epochs = epochs,nb_snapshots = 1, init_lr = 1e-3)
swa = SWA('./keras_swa.model',epochs - 3)
batch_size = 8
train_generator = DataGenerator(filepath = train_filepath, augmentations=train_augment, batch_size = batch_size)
val_generator = DataGenerator(filepath = val_filepath, augmentations=train_augment, batch_size = batch_size)
history = model.fit_generator(train_generator,validation_data = val_generator,epochs = epochs,callbacks = snapshot.get_callbacks())
# ## Plotting
# +
plt.figure(figsize=(16,4))
plt.subplot(1,2,1)
plt.plot(history.history['iou_metric'][1:])
plt.plot(history.history['val_iou_metric'][1:])
plt.ylabel('iou')
plt.xlabel('epoch')
plt.legend(['train','Validation'], loc='lower right')
plt.title('model IOU')
plt.savefig('iou.png')
plt.subplot(1,2,2)
plt.plot(history.history['loss'][1:])
plt.plot(history.history['val_loss'][1:])
plt.ylabel('val_loss')
plt.xlabel('epoch')
plt.legend(['train','Validation'], loc='upper right')
plt.title('model loss')
plt.savefig('loss.png')
gc.collect()
| efficientnetunet-for-pneumothorax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.1 (''.stroke'': venv)'
# name: pythonjvsc74a57bd0f95739c3243ee941eb965455c083e867fa1156c770d63e1e75cd982cc778ae6b
# ---
# +
import numpy as np
import pandas as pd
import math
PLOT_VISUALIZATIONS = True
RANDOM_SEED = 1243
TEST_SIZE = 0.1
# +
import sys
from importlib import reload
customPackages = ["visualizations", "modelling"]
for pkg in customPackages:
reload(sys.modules[pkg]) if pkg in sys.modules else __import__(pkg)
vis = sys.modules["visualizations"].Visualizations()
mod = sys.modules["modelling"].Modelling(vis=vis)
# -
rawData = pd.read_csv("data/healthcare-dataset-stroke-data.csv")
print(rawData.shape)
rawData.head()
# ## Data cleansing steps
#
# 1. Drop id column
# 2. Convert all categorical columns into 1-hot vectors (gender, ever_married, work_type, Residence_type, smoking_status)
# 3. Fill NaN values
# 4. Normalize numerical columns (age, avg_glucose_level, bmi)
TARGET_COLUMN = "stroke"
rawData.drop(columns="id", inplace=True)
print("There are only %d rows with Gender = Other. Dropping these to reduce noise!" % rawData[rawData["gender"] == "Other"].shape[0])
rawData = rawData[rawData["gender"] != "Other"]
# +
rawData["ever_married"] = rawData["ever_married"].apply(lambda x: "married" if x == "Yes" else "not_married")
binaryColumns = ["gender", "Residence_type", "ever_married"]
for col in binaryColumns:
trueValue = rawData[col].unique()[0]
rawData["is_" + trueValue.lower()] = rawData[col].apply(lambda x: 1 if x == trueValue else 0)
rawData.drop(columns=binaryColumns, inplace=True)
uniqueCounts = rawData.apply(lambda x: len(np.unique(x)), axis=0)
binaryColumns = np.array(uniqueCounts[uniqueCounts == 2].index)
binaryColumns = binaryColumns[binaryColumns != TARGET_COLUMN]
# +
print("There are %d True/False or binary columns in the dataset." % len(binaryColumns))
print(binaryColumns)
if PLOT_VISUALIZATIONS:
vis.plotMultipleCountPlots(data=rawData, columns=binaryColumns, figSize=(10,8), plotTitle="Binary Features")
# +
numericColumns = list(rawData.columns[rawData.dtypes == float])
print("There are %d numeric columns in the dataset." % len(numericColumns))
print(numericColumns)
if PLOT_VISUALIZATIONS:
vis.plotHistograms(rawData[numericColumns], figSize=(12,10))
# +
categoryColumns = list(rawData.columns[rawData.dtypes == object])
print("There are %d True/False or binary columns in the dataset." % len(categoryColumns))
print(categoryColumns)
if PLOT_VISUALIZATIONS:
vis.plotMultipleCountPlots(data=rawData, columns=categoryColumns, figSize=(6,8), plotTitle="Categorical Features")
# -
print("There are %d null values in the raw data." % (rawData.isnull().values.sum()))
rawData.isnull().sum(axis = 0)
rawData["age_bucket"] = rawData["age"].apply(lambda x: 20 * math.floor(x / 20))
if PLOT_VISUALIZATIONS:
vis.plotBoxPlot(data=rawData, x="age_bucket", y="bmi", hue="is_male", figSize=(10,6), plotTitle="BMI Variation w/ Age, Gender")
y = rawData[TARGET_COLUMN]
X = rawData.drop(columns=TARGET_COLUMN)
X_train, X_test, y_train, y_test = mod.splitData(X, y, testSize=TEST_SIZE, randomSeed=RANDOM_SEED, resetIndex=True)
X_train_preproc, encoderModels, imputerModel, scalerModel = mod.preprocData(X=X_train, categoryColumns=categoryColumns, numericColumns=numericColumns)
X_train_preproc.head()
X_test_preproc, _, _, _ = mod.preprocData(X=X_test, categoryColumns=categoryColumns, numericColumns=numericColumns, \
encoderModels=encoderModels, imputerModel=imputerModel, scalerModel=scalerModel)
| dataExploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import PuLP modeler functions
from pulp import *
# A list of all the roll lengths is created
LenOpts = ["5","7","9"]
# A dictionary of the demand for each roll length is created
rollDemand = {"5":150,
"7":200,
"9":300}
# A list of all the patterns is created
PatternNames = ["A","B","C"]
# Creates a list of the number of rolls in each pattern for each different roll length
patterns = [#A B C
[0,2,2],# 5
[1,1,0],# 7
[1,0,1] # 9
]
# -
# The cost of each 20cm long sponge roll used
cost = 1
# The pattern data is made into a dictionary
patterns = makeDict([LenOpts,PatternNames],patterns,0)
patterns
# The problem variables of the number of each pattern to make are created
vars = LpVariable.dicts("Patt",PatternNames,0,None,LpInteger)
vars
# The variable 'prob' is created
prob = LpProblem("Cutting Stock Problem",LpMinimize)
# The objective function is entered: the total number of large rolls used * the fixed cost of each
prob += lpSum([vars[i]*cost for i in PatternNames]),"Production Cost"
# The demand minimum constraint is entered
for i in LenOpts:
prob += lpSum([vars[j]*patterns[i][j] for j in PatternNames])>=rollDemand[i],"Ensuring enough %s cm rolls"%i
# +
# The problem data is written to an .lp file
prob.writeLP("SpongeRollProblem.lp")
# The problem is solved using PuLP's choice of Solver
prob.solve()
# The status of the solution is printed to the screen
print("Status:", LpStatus[prob.status])
# Each of the variables is printed with it's resolved optimum value
for v in prob.variables():
print(v.name, "=", v.varValue)
# The optimised objective function value is printed to the screen
print("Production Costs = ", value(prob.objective))
| PULP/tutorial from youtube/SpongeRollProblem1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/holps-7/PDC-weekly/blob/master/PDC_lab4(1.0).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="KGNjj2lHv4gW" colab_type="code" colab={}
#<NAME> 18BCE2030
code = """
#include <omp.h>
#include <stdio.h>
#include <stdlib.h>
#define RNG_MOD 0x80000000
int state;
int rng_int(void);
double rng_doub(double range);
int main() {
int i, ctr;
unsigned int n;
double x, y, pi;
n = 1<<30;
ctr = 0;
#pragma omp threadprivate(state)
#pragma omp parallel private(x, y) reduction(+:ctr)
{
state = 25234 + 17 * omp_get_thread_num();
#pragma omp for
for (i = 0; i <= n; i++) {
x = (double)rng_doub(1.0);
y = (double)rng_doub(1.0);
if (x*x + y*y <= 1) ctr++;
}
}
pi = 4.0*ctr / n;
printf("Pi: %lf", pi);
return 0;
}
int rng_int(void) {
return (state = (state * 1103515245 + 12345) & 0x7fffffff);
}
double rng_doub(double range) {
return ((double)rng_int()) / (((double)RNG_MOD)/range);
}
"""
# + id="dbLfFWzhv6l0" colab_type="code" colab={}
text_file = open("code.c", "w")
text_file.write(code)
text_file.close()
# + id="lNG_3GVUv8Xi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="5b4c299b-628a-4444-c21a-183f26a8bbc3"
# %env OMP_NUM_THREADS=10
# !gcc -o hello -fopenmp code.c
# !./hello
# + id="E5McoI2sv-MA" colab_type="code" colab={}
| PDC_lab4(1.0).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('base')
# language: python
# name: python3
# ---
# # COLOR DETECTION
# +
import cv2
import numpy as np
# Creating a trackbar window
def empty(a):
pass
# FUNCTION TO STACK IMAGES
def stackImages(scale, imgArray):
rows = len(imgArray)
cols = len(imgArray[0])
rowsAvailable = isinstance(imgArray[0], list)
width = imgArray[0][0].shape[1]
height = imgArray[0][0].shape[0]
if rowsAvailable:
for x in range (0, rows):
for y in range (0, cols):
if imgArray[x][y].shape[:2] == imgArray[0][0].shape[:2]:
imgArray[x][y] = cv2.resize(imgArray[x][y], (0,0), None, scale, scale)
else:
imgArray[x][y] = cv2.resize(imgArray[x][y], (imgArray[0][0].shape[1], imgArray[0][0].shape[0]), None, scale, scale)
if len(imgArray[x][y].shape) == 2:imgArray[x][y]= cv2.cvtColor(imgArray[x][y], cv2.COLOR_GRAY2BGR)
imageBlank = np.zeros((height, width, 3), np.uint8)
hor = [imageBlank]*rows
hor_con = [imageBlank]*rows
for x in range(0, rows):
hor[x] = np.hstack(imgArray[x])
ver = np.vstack(hor)
else:
for x in range(0, rows):
if imgArray[x].shape[:2] == imgArray[0].shape[:2]:
imgArray[x] = cv2.resize(imgArray[x], (0,0), None, scale, scale)
else:
imgArray[x] = cv2.resize(imgArray[x], (imgArray[0].shape[1], imgArray[0].shape[0]), None, scale, scale)
if len(imgArray[x].shape) == 2: imgArray[x] = cv2.cvtColor(imgArray[x], cv2.COLOR_GRAY2BGR)
hor = np.hstack(imgArray)
ver = hor
return ver
path = 'resources/trust.png'
cv2.namedWindow('Trackbars')
cv2.resizeWindow('Trackbars', 640,240)
cv2.createTrackbar('Hue Min', "Trackbars", 0,179, empty) # where first argument is the trackbarName, second argument is the windowName and
# the third argument is the value i.e form 0 to 179
cv2.createTrackbar('Hue Max', "Trackbars", 0,179, empty)
cv2.createTrackbar('Sat Min', "Trackbars", 0,255, empty)
cv2.createTrackbar('Sat Max', "Trackbars", 51,255, empty)
cv2.createTrackbar('Val Min', "Trackbars", 56,255, empty)
cv2.createTrackbar('Val Max', "Trackbars", 255,255, empty)
while True:
img = cv2.imread(path)
imgHSV = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
h_min = cv2.getTrackbarPos('Hue Min', 'Trackbars')
h_max = cv2.getTrackbarPos('Hue Max', 'Trackbars')
s_min = cv2.getTrackbarPos('Sat Min', 'Trackbars')
s_max = cv2.getTrackbarPos('Sat Max', 'Trackbars')
v_min = cv2.getTrackbarPos('Val Min', 'Trackbars')
v_max = cv2.getTrackbarPos('Val Max', 'Trackbars')
print(h_min, h_max, s_min, s_max, v_min, v_max)
lower = np.array([h_min, s_min, v_min])
upper = np.array([h_max, s_max, v_max])
mask = cv2.inRange(imgHSV,lower,upper)
imgResult = cv2.bitwise_and(img, img, mask=mask)
#cv2.imshow('Original', img)
#cv2.imshow('Image', imgHSV)
#cv2.imshow('Mask', mask)
#cv2.imshow('Result', imgResult)
imgStack = stackImages(0.6, ([img, imgHSV], [mask, imgResult]))
cv2.imshow('Stacked Images', imgStack)
cv2.waitKey(1)
# -
| basics6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import json
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from multiprocessing import cpu_count
from utils_plot import rmse, plot_comparisons, plot_feature_importances
import predictor
import optimizer
from itertools import combinations
from copy import deepcopy
# %matplotlib inline
seed = 100
np.random.seed(seed)
# -
# # 1. Cross-Validation Tuning and Train
# ## Import data and preprocess
#
# This dataset contains all the literature data + 87 experimental datapoints
# +
with open('./full_data.json', 'r') as file:
data = pd.read_json(json.load(file), orient='table')
data
# -
# ## CV Tuning with XGBoost
# +
inputs = np.asarray(data.loc[:,'Units of A':'Mp'])
labels = np.asarray(data['Cloud Point'])
scaler_regressor = MinMaxScaler()
inputs = scaler_regressor.fit_transform(inputs)
inputs_train, inputs_valid, labels_train, labels_valid = train_test_split(
inputs, labels, test_size=0.1, random_state=seed)
# -
XGB_Options = {
'cv': 10,
'scoring': 'neg_mean_squared_error',
'seed': seed,
'max_depth': np.arange(2,14,2),
'min_child_weight': np.arange(1,8,1),
'n_estimators': np.arange(10,80,5),
'gamma': np.arange(0.05,0.45,0.05),
'colsample_bytree': np.arange(0.60, 0.95, 0.05),
'subsample': np.arange(0.60, 0.95, 0.05),
'reg_alpha': [1e-5, 1e-2, 0.1, 0.5, 1, 5, 10], #alpha
'reg_lambda': [1e-5, 1e-2, 0.1, 0.5, 1, 5, 10],#lambda
'learning_rate': np.arange(0.025,0.150,0.025),
'scaler': scaler_regressor,
'n_jobs': 4,
'verbose': 1
}
trained_regressor = predictor.XGB_Regressor(options=XGB_Options)
trained_regressor.fit(inputs_train, labels_train)
tuned_regressor = trained_regressor.regressor
# +
n_splits = 3
np.random.seed(seed)
targets = []
results = []
for n in range(n_splits):
inputs_train, inputs_test, labels_train, labels_test = train_test_split(inputs, labels, test_size=0.1)
tuned_regressor.fit(inputs_train, labels_train)
pred_train = tuned_regressor.predict(inputs_train)
pred_test = tuned_regressor.predict(inputs_test)
results.append({'train': pred_train, 'test': pred_test, 'name': 'Split {}'.format(n+1)})
targets.append({'train': labels_train, 'test': labels_test})
# -
plot_comparisons(targets, results)
# # 2. Train NN as Secondary Validation
# +
import tensorflow as tf
tf.reset_default_graph()
np.random.seed(seed)
tf.set_random_seed(seed)
sess = tf.Session()
# +
NUM_NN = 10
regressors = []
for n in range(NUM_NN):
options = {'input_shape': [6, ], 'target_shape': [1, ],
'n_layers': 2, 'layer_widths': [64, 128],
'name': 'nn_{}'.format(n), 'session': sess}
regressors.append(predictor.NeuralNetRegressor(options=options))
ensb_regressor = predictor.EnsembleRegressor(regressors, scaler_regressor)
# -
options={'monitor': False,
'reinitialize': True,
'n_iter': 10000}
list_of_options = [options,]*NUM_NN
ensb_regressor.fit(inputs_train, labels_train.reshape(-1, 1), list_of_options)
# +
predicted_train = ensb_regressor.predict_mean(inputs_train)
predicted_validation = ensb_regressor.predict_mean(inputs_valid)
train_rmse = rmse(predicted_train, labels_train)
validation_rmse = rmse(predicted_validation, labels_valid)
print('Train RMSE: ', train_rmse, 'validation RMSE: ', validation_rmse)
# -
# ### Train all data
# +
ensb_regressor.fit(inputs, labels.reshape(-1,1), list_of_options)
predicted_nn = ensb_regressor.predict_mean(inputs)
std_nn = np.sqrt(ensb_regressor.predict_covariance(inputs))
rmse_nn = rmse(predicted_nn, labels)
# -
# # 3. Inverse Design
# ## Train model with tuned parameters on the full dataset
trained_regressor.regressor.fit(inputs, labels)
predicted = trained_regressor.predict(inputs)
print('Train RMSE: ', rmse(predicted, labels))
# ## Constraint and Target Specifications
#
# We solve the problem
# $$
# \min_{x} \mathrm{objective}(x, CP)
# $$
# subject to
# $$
# \mathrm{constraints}(x) \geq 0
# $$
#
# Some soft constraints are written into the objective function as a regularizer, and some hard constraints are moved to the selection criterion function, which must evaluates to true before we consider the solution admissiable. This is to maximize efficiency of the particle swarm method.
# +
lb = np.array([0]*6)
ub = np.array([203, 187, 43, 96, 0.1, 23196])
def objective(x, target_CP, regressor):
'''
loss function:
The loss function contains 2 parts
1. 0.5*(predicted_CP(x) - target_CP)**2
2. coef * regularizer
'''
def regularizer(x):
'''
Regularizer (or soft constraints):
We want to drive the system towards
binary systems involving A,B,C,D
Here, we use the simple penalizer
\sum_{(i,j,k) \in (A,B,C,D)} i*j*k
For task 1:
We also want to minimize A
'''
reg = 0.0
for c in combinations(x[:4], 3):
reg += np.prod(c)**(1.0/3.0)
reg += x[0] # minimize A
return reg
return 0.5 * (regressor.predict_transform(x) - target_CP)**2 \
+ 1e-1 * regularizer(x)
def constraints(x, *args):
''' hard constraints '''
Final_M = np.dot(x[:4], [99.13, 113.16, 111.14, 113.16])
cons_Mp_ub = [Final_M * 1.2 - x[5]]
cons_Mp_lb = [x[5] - Final_M * 0.8]
return np.asarray(cons_Mp_ub + cons_Mp_lb)
def selection_criteria(x):
"""
Selection criteria:
Desired solution x should evaluates to all
non-negative
1. Minimum non-zero component must be at least 10%
of the max component. (note: non-zero is defined
to be <= ZERO_CUTOFF, purely for numerical stability)
2. Allow for at most ternary systems
"""
ZERO_CUTOFF = 0.1
# Criterion 1
xcomps = x[:4]
x_non_zero = xcomps[xcomps > ZERO_CUTOFF]
x_max = xcomps.max()
x_min = x_non_zero.min()
comp_condition = [x_min - 0.1*x_max]
# Criterion 2
num_non_zero = np.sum(x[:4] > ZERO_CUTOFF)
nonzero_condition = [3.1 - num_non_zero]
# return np.asarray([1.0])
return all([c >= 0 for c in comp_condition + nonzero_condition])
# -
# ## Optimze via Particle Swarm Optimizer
#
# The optimisation options are found using a systematic parameter study that we omit here for brevity
# +
opt = optimizer.PSO_Optimizer(
regressor=trained_regressor,
scaler=scaler_regressor,
objective=objective,
constraints=constraints,
selection_criteria=selection_criteria
)
optimisation_options = { 'criteria': 0.1,
'max_rounds': 200,
'n_solutions': 50,
'lb':lb, 'ub':ub,
'opt_vars': [ 'Units of A', 'Units of B', 'Units of C', 'Units of D', 'Units of E', 'Mp'],
'swarmsize': 400, 'omega':0.1, 'phip':0.9, 'phig': 0.8, 'maxiter': 100,
'debug_flag': False,
'nprocessors': 12
}
# -
# ### 1. Target = 37$^{\circ}$C
optimisation_options['target_CP'] = 37
results37, runtime = opt.optimisation_parallel(optimisation_options)
results37
# ### 2. Target = 45$^{\circ}$C
optimisation_options['target_CP'] = 45
results45, runtime = opt.optimisation_parallel(optimisation_options)
results45
# ### 3. Target = 60$^{\circ}$C
# +
optimisation_options['target_CP'] = 60
results60, runtime = opt.optimisation_parallel(optimisation_options)
results60
# -
# ### 4. Target = 80$^{\circ}$C
optimisation_options['target_CP'] = 80
results80, runtime = opt.optimisation_parallel(optimisation_options)
results80
# ## NN validation of the XGB results
def NN_Mean_Std(input_df, scaler, ensb_reg):
data = deepcopy(input_df)
inputs = scaler_regressor.transform(data.iloc[:, :6 ].values)
mean = ensb_regressor.predict_mean(inputs)
cov = ensb_regressor.predict_covariance(inputs)
std = np.sqrt(cov.squeeze())
data['Mean_Pred_NN'] = mean
data['Std_Error_NN'] = std
data['%Error_Mean_Pred_NN'] = 100*(data.Mean_Pred_NN - data.target_CP)/data.target_CP
return data
NN_validation37 = NN_Mean_Std(results37, scaler_regressor, ensb_regressor)
NN_validation37
NN_validation45 = NN_Mean_Std(results45, scaler_regressor, ensb_regressor)
NN_validation45
NN_validation60 = NN_Mean_Std(results60, scaler_regressor, ensb_regressor)
NN_validation60
NN_validation80 = NN_Mean_Std(results80, scaler_regressor, ensb_regressor)
NN_validation80
| CloudPoint-MachineLearning/inverse_design.ipynb |