code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Demo 01: Describing your geometry
# + active=""
# In TIGRE the geometry is stored in a class.
#
# --------------------------------------------------------------------------
# --------------------------------------------------------------------------
# This file is part of the TIGRE Toolbox
#
# Copyright (c) 2015, University of Bath and
# CERN-European Organization for Nuclear Research
# All rights reserved.
#
# License: Open Source under BSD.
# See the full license at
# https://github.com/CERN/TIGRE/license.txt
#
# Contact: <EMAIL>
# Codes: https://github.com/CERN/TIGRE/
# --------------------------------------------------------------------------
# Coded by: MATLAB (original code): <NAME>
# PYTHON : <NAME>,<NAME>
#
# To see a demo of what the geometry paramterers should look like, do as follows:
# -
from tigre import geometry
geo = geometry.TIGREParameters(high_quality=False)
print(geo)
# + active=""
# Geometry definition
#
# Detector plane, behind
# |-----------------------------|
# | |
# | |
# | |
# Centered | |
# at O A V +--------+ |
# | / /| |
# A Z | / / |*D |
# | | +--------+ | |
# | | | | | |
# | | | *O | + |
# *--->y | | | / |
# / | | |/ |
# V X | +--------+ U |
# .--------------------->-------|
#
# *S
# -
# We recommend using the template below and defining you're class as such:
#
# +
from __future__ import division
import numpy as np
class TIGREParameters:
def __init__(self, high_quality=True):
if high_quality:
# VARIABLE DESCRIPTION UNITS
# -------------------------------------------------------------------------------------
self.DSD = 1536 # Distance Source Detector (mm)
self.DSO = 1000 # Distance Source Origin (mm)
# Detector parameters
self.nDetector = np.array((512, 512)) # number of pixels (px)
self.dDetector = np.array((0.8, 0.8)) # size of each pixel (mm)
self.sDetector = self.nDetector * self.dDetector # total size of the detector (mm)
# Image parameters
self.nVoxel = np.array((256, 256, 256)) # number of voxels (vx)
self.sVoxel = np.array((256, 256, 256)) # total size of the image (mm)
self.dVoxel = self.sVoxel/self.nVoxel # size of each voxel (mm)
# Offsets
self.offOrigin = np.array((0, 0, 0)) # Offset of image from origin (mm)
self.offDetector = np.array((0, 0)) # Offset of Detector (mm)
# Auxiliary
self.accuracy = 0.5 # Accuracy of FWD proj (vx/sample)
# Mode
self.mode = 'cone' # parallel, cone ...
else:
# VARIABLE DESCRIPTION UNITS
# -------------------------------------------------------------------------------------
self.DSD = 1536 # Distance Source Detector (mm)
self.DSO = 1000 # Distance Source Origin (mm)
# Detector parameters
self.nDetector = np.array((128, 128)) # number of pixels (px)
self.dDetector = np.array((0.8, 0.8))*4 # size of each pixel (mm)
self.sDetector = self.nDetector * self.dDetector # total size of the detector (mm)
# Image parameters
self.nVoxel = np.array((64, 64 , 64)) # number of voxels (vx)
self.sVoxel = np.array((256, 256, 256)) # total size of the image (mm)
self.dVoxel = self.sVoxel / self.nVoxel # size of each voxel (mm)
# Offsets
self.offOrigin = np.array((0, 0, 0)) # Offset of image from origin (mm)
self.offDetector = np.array((0, 0)) # Offset of Detector (mm)
# Auxiliary
self.accuracy = 0.5 # Accuracy of FWD proj (vx/sample)
# Mode
self.mode=None # parallel, cone ...
self.filter=None
| Python/tigre_demo_file/d01_create_geometry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
#import os
import pathlib as pl
from datetime import datetime
import matplotlib.pyplot as plot
from pandas import DataFrame, Series
# read .xlsx files under ./data directory into datatables.
#os.listdir()
datapath=pl.Path("./data")
#datapath=pl.Path.cwd()
file_list=[]
DataFrame_list=[]
for x in datapath.glob("*"):
for y in x.glob("*[0-9].xlsx"):
f=pd.ExcelFile(y)
t=f.parse(f.sheet_names[0])
DataFrame_list.append(t)
file_list.append(y.name)
dfs=[]
for x in DataFrame_list:
x.columns=x.iloc[1]
x=x[x.时间.apply(lambda x: x.__class__==datetime)]
x=x[x.时间.apply(lambda x: x.minute % 3==0)]
x.时间=x.时间.apply(lambda y: datetime(y.year,y.month,y.day,y.hour,y.minute,y.second))
dfs.append(x)
for x in dfs:
print(len(x))
x0=dfs[0];x2=dfs[2];x3=dfs[3]
for i in range(0,6):
for j in range(13,20):
print(i,j)
print((dfs[i].时间.values==dfs[j].时间.values).all())
#t0=x0.时间.apply(lambda y: datetime(y.year,y.month,y.day,y.hour,y.minute,y.second))
#t2=x2.时间.apply(lambda y: datetime(y.year,y.month,y.day,y.hour,y.minute,y.second))
#[t for t in t0.values if t not in t2.values]
#[t for t in t2.values if t not in t0.values]
for df in dfs:
df.时间=df.时间.apply(lambda y: datetime(y.year,y.month,y.day,y.hour,y.minute,y.second))
dfs1=dfs[:8]+dfs[13:20]
i =[t in dfs1[0].时间.values for t in dfs1[6].时间.values]
dfs1[6]=dfs1[6][i]
dfs1[7]=dfs1[7][i]
i =[t in dfs1[6].时间.values for t in dfs1[0].时间.values]
for j in range(0,6):
dfs1[j]=dfs1[j][i]
for j in range(9,15):
dfs1[j]=dfs1[j][i]
for j in range(len(dfs1)):
dfs1[j]=dfs1[j].set_index("时间")
#dfs1[0] = dfs1[0][t in dfs1[2].时间.values for t in dfs1[0].时间.values]
tr_data=dfs1[0].iloc[:,0].apply(float)
tr_data=DataFrame(tr_data)
for i in range(1,len(dfs1)):
tr_data=tr_data.join(dfs1[i].iloc[:,0].apply(float))
tr_data1=dfs1[0].iloc[:,1].apply(float)
tr_data1=DataFrame(tr_data1)
for i in range(1,len(dfs1)):
tr_data1=tr_data1.join(dfs1[i].iloc[:,1].apply(float))
# +
tr_data2=dfs1[0].iloc[:,2].apply(float)
tr_data2=DataFrame(tr_data2)
for i in range(1,len(dfs1)):
tr_data2=tr_data2.join(dfs1[i].iloc[:,2].apply(float))
# -
for x in tr_data:
print(x)
corMat=DataFrame(tr_data1.corr())
plot.pcolor(corMat)
plot.show()
tr_data1.to_csv("avg.csv")
# +
###### network from keras for SHDKY data simulation ###########
from keras.models import Sequential
from keras.layers import Dense, Activation,Input
import keras
tr=(tr_data1-tr_data1.min())/(tr_data1.max()-tr_data1.min())
model = Sequential()
model.add(Dense(14, input_dim=14,init="normal"))
model.add(Activation('sigmoid'))
model.add(Dense(7, activation='sigmoid',init="normal"))
model.add(Dense(1, activation='linear',init="normal"))
model.compile(loss='mean_squared_error',
optimizer=keras.optimizers.SGD(lr=0.2))
Y=np.array(tr.iloc[:,0:1])
X=np.array(tr.iloc[:,1:15])
# -
model.fit(X, Y, epochs=30,batch_size=10,
shuffle=True,verbose=2,validation_split=0.2)
model.predict(X)
Y
(model.predict(X)-Y)/Y
model.save('SH_NNmodel.h5')
from keras.models import load_model
model = load_model('SH_NNmodel.h5')
for x in dfs1:
for y in x:
if y.find("Avg[I 有效值 A]") != -1:
print(y)
tr_data2=DataFrame()
for x in dfs1:
for y in x.columns:
if y.find("Avg[I 有效值 A]") != -1:
tr_data2[y]=x[y].apply(float)
tr_data2
tr_data2.to_csv('Iavg.csv')
| SHDKY/.ipynb_checkpoints/SHDKY-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Ingest
#
# In this notebook, we read a wiki text snippet and save the data to an S3 bucket
# +
import os
import warnings
from dotenv import load_dotenv, find_dotenv
import boto3
from datasets import load_dataset
warnings.filterwarnings("ignore")
load_dotenv(find_dotenv())
# + tags=[]
## Create a .env file on your local with the correct configs
s3_endpoint_url = os.getenv("S3_ENDPOINT")
s3_access_key = os.getenv("S3_ACCESS_KEY")
s3_secret_key = os.getenv("S3_SECRET_KEY")
s3_bucket = os.getenv("S3_BUCKET")
# -
# Create an S3 client
s3 = boto3.client(
service_name="s3",
aws_access_key_id=s3_access_key,
aws_secret_access_key=s3_secret_key,
endpoint_url=s3_endpoint_url,
)
# +
# # ! jupyter labextension install @jupyter-widgets/jupyterlab-manager
# -
dataset = load_dataset('wikitext', 'wikitext-2-v1')
dataset
text = dataset['train']['text'][0:50]
text = list(filter(None, text))
text
# ## Upload to S3
file_path = '../data/raw/wiki.txt'
with open(file_path, 'w') as file:
for item in text:
file.write('%s\n' % item)
key = 'op1-pipelines/wiki.txt'
s3.upload_file(Bucket=s3_bucket, Key=key, Filename=str(file_path))
| notebooks/ingest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp models.error_dist
# -
# # Error Distributions
# > distribution of logT - mean in AFT models
#
# In order to get the distribution of $T$ we can use change of variable theorem:
# $$
# \begin{aligned}
# \xi &= \log T - \mu\\
# \frac{d\xi}{dT} &= \frac{1}{T}\\
# p(T) &= p(\log(T) - \mu|\theta)\frac{d\xi}{dT}
# \end{aligned}
# $$
# +
#export
from functools import partial
import torch
import torch.nn as nn
import torch.nn.functional as F
# +
# hide
import matplotlib.pyplot as plt
import numpy as np
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
# ## Gumbel Distributed Error
#
# Suppose that the error is [Gumbel](https://en.wikipedia.org/wiki/Gumbel_distribution) distributed, $\xi_i\sim Gumbel(1)$.
#
# $$
# \begin{aligned}
# p(\xi) &= \exp(-\xi - \exp(-\xi))\\
# p(T) &= \exp(-(\log T - \mu) - \exp(-(\log T - \mu)))\times\frac{1}{T}\\
# p(T) &= \frac{1}{T}\exp\left(\mu-\frac{1}{T}\exp(\mu)\right)\times\frac{1}{T}\\
# p(T) &\propto \left(\frac{1}{T}\right)^2\exp\left(-\frac{1}{T}\exp(\mu)\right)
# \end{aligned}
# $$
# Therefore, $T$ is [Inverse Gamma](https://en.wikipedia.org/wiki/Inverse-gamma_distribution) distributed, such that $T\sim IG(1, \exp(\mu))$. The survival function in this case is (by using the identities that $\Gamma(1) = 1$ and $\Gamma(1,x) = \exp(-x)$:
# $$
# \begin{aligned}
# p(T>t) &= 1 - \frac{\Gamma(1, \exp(\mu)/t)}{\Gamma(1)}\\
# p(T>t) &= 1 - \exp\left(-\frac{\exp(\mu)}{t}\right)
# \end{aligned}
# $$
# export
def get_distribution(dist:str):
"""
Get the logpdf and logcdf of a given torch distribution
"""
dist = getattr(torch.distributions, dist.title())
if not isinstance(dist.support, torch.distributions.constraints._Real):
raise Exception("Distribution needs support over ALL real values.")
dist = partial(dist, loc=0.0)
def dist_logpdf(ξ, σ):
return dist(scale=σ).log_prob(ξ)
def dist_logicdf(ξ, σ):
"""
log of inverse cumulative distribution function
"""
return torch.log(1 - dist(scale=σ).cdf(ξ))
return dist_logpdf, dist_logicdf
_, logicdf = get_distribution("Gumbel")
μs = np.log(np.arange(1, 5))
t = np.linspace(1e-3, 10)
for μ in μs:
logT = np.log(t)
ξ = torch.Tensor(logT - μ)
S = torch.exp(logicdf(torch.Tensor(ξ), 1))
plt.plot(t, S, label=f'μ = log {int(np.exp(μ))}')
plt.legend()
plt.grid()
plt.show()
# hide
from nbdev.export import *
notebook2script()
| 65_AFT_error_distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: urbanroute
# language: python
# name: urbanroute
# ---
# +
"""Find the least cost path from source to target by minimising air pollution."""
from typing import Optional, Tuple
import logging
import geopandas as gpd
import osmnx as ox
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.colors as colors
import networkx as nx
import math
from cleanair.databases.queries import AirQualityResultQuery
from cleanair.loggers import get_logger
from shapely.geometry import Polygon
from routex.types import Node
from urbanroute.geospatial import update_cost, ellipse_bounding_box
from urbanroute.queries import HexGridQuery
def main( # pylint: disable=too-many-arguments
secretfile: str = "/home/james/clean-air-infrastructure/.secrets/db_secrets_ad.json",
instance_id: str = "d5e691ef9a1f2e86743f614806319d93e30709fe179dfb27e7b99b9b967c8737",
sourceLat: float = 51.4929,
sourceLong: float = -0.1215,
start_time: Optional[str] = "2020-01-24T09:00:00",
targetLat: float = 51.4929,
targetLong: float = -0.2215,
upto_time: Optional[str] = "2020-01-24T10:00:00",
verbose: Optional[bool] = False,
):
"""
secretfile: Path to the database secretfile.
instance_id: Id of the air quality trained model.
sourceLat: latitude of the source point.
sourceLong: longitude of the source point.
targetLat: latitude of the target point.
targetLong: longitude of the target point.
"""
source = (sourceLat, sourceLong)
target = (targetLat, targetLong)
logger = get_logger("Shortest path entrypoint")
if verbose:
logger.level = logging.DEBUG
# TODO change this to a AirQualityResultQuery
result_query = HexGridQuery(secretfile=secretfile)
logger.info("Querying results from an air quality model")
result_sql = result_query.query_results(
instance_id,
join_hexgrid=True,
output_type="sql",
start_time=start_time,
upto_time=upto_time,
)
logger.debug(result_sql)
gdf = gpd.GeoDataFrame.from_postgis(
result_sql, result_query.dbcnxn.engine, crs=4326
)
gdf = gdf.rename(columns=dict(geom="geometry"))
gdf.crs = "EPSG:4326"
# gdf = gpd.GeoDataFrame(result_df, crs=4326, geometry="geom")
logger.info("%s rows in hexgrid results", len(gdf))
if source is not None and target is not None:
# use bounding box of surrounding ellipse to limit graph size
box = ellipse_bounding_box((source[1], source[0]), (target[1], target[0]))
G: nx.MultiDiGraph = ox.graph_from_bbox(box[0], box[1], box[2], box[3])
# snap source and target to the graph
newSource = ox.distance.get_nearest_node(G, source)
newTarget = ox.distance.get_nearest_node(G, target)
logger.info(
"%s nodes and %s edges in graph.", G.number_of_nodes(), G.number_of_edges()
)
logger.info("Mapping air quality predictions to the road network.")
G = update_cost(G, gdf, cost_attr="NO2_mean", weight_attr="length")
logger.debug("Printing basic stats for the graph:")
logger.debug(ox.stats.basic_stats(G))
for i, (u, v, k, data) in enumerate(G.edges(keys=True, data=True)):
if i > 10:
break
print(u, v, k, data)
ev = [
G.get_edge_data(edge[0], edge[1], edge[2])["NO2_mean"] ** (1 / 2)
for edge in G.edges
]
norm = colors.Normalize(vmin=min(ev), vmax=max(ev))
cmap = cm.ScalarMappable(norm=norm, cmap=cm.inferno)
ec = [cmap.to_rgba(cl) for cl in ev]
fig, ax = ox.plot_graph_route(
G,
nx.dijkstra_path(G, newSource, newTarget, weight="NO2_mean"),
edge_color=ec,
)
main()
# -
| notebooks/visualisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import metrics
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.neighbors import KNeighborsRegressor
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix, mean_squared_error
from sklearn.tree import export_graphviz
from IPython.display import Image
# -
# Cochise county
cochise = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/cochise.csv")
# Get X and y
X = cochise.drop(columns = 'sierra_vista_hmi')
y = cochise.sierra_vista_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = cochise.corr()
correlation['sierra_vista_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Sierra Vista hmi and Cochise county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Coconino county
coconino = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/coconino.csv")
# Get X and y
X = coconino.drop(columns = 'flagstaff_hmi')
y = coconino.flagstaff_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = coconino.corr()
correlation['flagstaff_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Flagstaff hmi and Coconino county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Gila county
gila = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/gila.csv")
# Get X and y
X = gila.drop(columns = 'payson_hmi')
y = gila.payson_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = gila.corr()
correlation['payson_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Payson hmi and Gila county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Graham
graham = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/graham.csv")
# Get X and y
X = graham.drop(columns = 'safford_hmi')
y = graham.safford_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = graham.corr()
correlation['safford_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Safford hmi and Graham county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Maricopa county
maricopa = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/maricopa.csv")
# Get X and y
X = maricopa.drop(columns = 'phoenix_hmi')
y = maricopa.phoenix_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = maricopa.corr()
correlation['phoenix_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Phoenix hmi and Maricopa county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Mohave county
mohave = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/mohave.csv")
# Get X and y
X = mohave.drop(columns = 'lake_havasu_city_hmi')
y = mohave.lake_havasu_city_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = mohave.corr()
correlation['lake_havasu_city_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Lake Havasu City hmi and Mohave county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Pima county
navajo = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/navajo.csv")
# Get X and y
X = navajo.drop(columns = 'show_low_hmi')
y = navajo.show_low_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = navajo.corr()
correlation['show_low_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Show Low hmi and Navajo county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Pima county
pima = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/pima.csv")
# Get X and y
X = pima.drop(columns = 'tucson_hmi')
y = pima.tucson_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = pima.corr()
correlation['tucson_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Tucson hmi and Pima county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Santa Cruz county
santa_cruz = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/santa_cruz.csv")
# Get X and y
X = santa_cruz.drop(columns = 'nogales_hmi')
y = santa_cruz.nogales_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = santa_cruz.corr()
correlation['nogales_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Nogales hmi and Santa Cruz county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Yavapai county
yavapai = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/yavapai.csv")
# Get X and y
X = yavapai.drop(columns = 'prescott_hmi')
y = yavapai.prescott_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = yavapai.corr()
correlation['prescott_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Prescott hmi and Yavapai county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# -
# Yuma county
yuma = pd.read_csv("C:/Users/edoar/Documents/School_Documents/Fall_2021/Capstone/random_forest_files/yuma.csv")
# Get X and y
X = yuma.drop(columns = 'yuma_hmi')
y = yuma.yuma_hmi
# Split data
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# Find correlation of target
correlation = yuma.corr()
correlation['yuma_hmi'].sort_values(ascending=False)
# +
# Graph Heatmap
plt.figure(figsize = (16, 12))
plt.title('Correlation Heatmap of Yuma hmi and Yuma county sources', fontsize = 20)
a = sns.heatmap(correlation, square = True, annot = True)
a.set_xticklabels(a.get_xticklabels(), rotation = 45)
a.set_yticklabels(a.get_yticklabels(), rotation = 45)
plt.show()
# +
# Fit decision tree
d_tree = DecisionTreeClassifier()
d_tree.fit(X_train, y_train)
pred = d_tree.predict(X_test)
print("Accuracy:", metrics.accuracy_score(y_test, pred))
| models/Heatmap/heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Tutorial-IllinoisGRMHD: Configuration files
#
# ## Authors: <NAME> & <NAME>
#
# <font color='red'>**This module is currently under development**</font>
#
# ## In this tutorial module we explain the files necessary to configure `IllinoisGRMHD` so that it can be properly used by the Einstein Toolkit
#
# ### Required and recommended citations:
#
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>., and <NAME>. IllinoisGRMHD: an open-source, user-friendly GRMHD code for dynamical spacetimes. Class. Quantum Grav. 32 (2015) 175009. ([arxiv:1501.07276](http://arxiv.org/abs/1501.07276)).
# * **(Required)** <NAME>., <NAME>., <NAME>., <NAME>. Primitive Variable Solvers for Conservative General Relativistic Magnetohydrodynamics. Astrophysical Journal, 641, 626 (2006) ([astro-ph/0512420](https://arxiv.org/abs/astro-ph/0512420)).
# * **(Recommended)** <NAME>., <NAME>., <NAME>. An efficient shock-capturing central-type scheme for multidimensional relativistic flows - II. Magnetohydrodynamics. A&A 400 (2) 397-413 (2003). DOI: 10.1051/0004-6361:20021641 ([astro-ph/0210618](https://arxiv.org/abs/astro-ph/0210618)).
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This module is organized as follows
#
# 0. [Step 0](#src_dir): **Source directory creation**
# 1. [Step 1](#introduction): **Introduction**
# 1. [Step 2](#make_code_defn): **`make.code.defn`**
# 1. [Step 3](#configuration__ccl): **`configuration.ccl`**
# 1. [Step 4](#interface__ccl): **`interface.ccl`**
# 1. [Step 5](#param__ccl): **`param.ccl`**
# 1. [Step 6](#schedule__ccl): **`schedule.ccl`**
# 1. [Step 7](#code_validation__txt): **`code_validation.txt`**
# 1. [Step 8](#code_validation): **Code validation**
# 1. [Step 8.a](#code_validation__make_code_defn): *`make.code.defn`*
# 1. [Step 8.b](#code_validation__configuration__ccl): *`configuration.ccl`*
# 1. [Step 8.c](#code_validation__interface__ccl): *`interface.ccl`*
# 1. [Step 8.d](#code_validation__param__ccl): *`param.ccl`*
# 1. [Step 8.e](#code_validation__schedule__ccl): *`schedule.ccl`*
# 1. [Step 8.f](#code_validation__code_validation__txt): *`code_validation.txt`*
# 1. [Step 9](#latex_pdf_output): **Output this notebook to $\LaTeX$-formatted PDF file**
# <a id='src_dir'></a>
#
# # Step 0: Source directory creation \[Back to [top](#toc)\]
# $$\label{src_dir}$$
#
# We will now use the [cmdline_helper.py NRPy+ module](Tutorial-Tutorial-cmdline_helper.ipynb) to create the source directory within the `IllinoisGRMHD` NRPy+ directory, if it does not exist yet.
# +
# Step 0: Creation of the IllinoisGRMHD source directory
# Step 0a: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..","..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Step 0b: Load up cmdline_helper and create the directory
import cmdline_helper as cmd
IGM_src_dir_path = os.path.join("..","src")
cmd.mkdir(IGM_src_dir_path)
# Step 0c: For this tutorial module we will also need IllinoisGRMHD's main directory path
IGM_main_dir_path = ".."
# Step 0d: Create the output file path
outfile_path__make_code_defn = os.path.join(IGM_src_dir_path, "make.code.defn")
outfile_path__configuration__ccl = os.path.join(IGM_main_dir_path,"configuration.ccl")
outfile_path__interface__ccl = os.path.join(IGM_main_dir_path,"interface.ccl")
outfile_path__param__ccl = os.path.join(IGM_main_dir_path,"param.ccl")
outfile_path__schedule__ccl = os.path.join(IGM_main_dir_path,"schedule.ccl")
outfile_path__code_validation__txt = "code_validation.txt"
# -
# <a id='introduction'></a>
#
# # Step 1: Introduction \[Back to [top](#toc)\]
# $$\label{introduction}$$
# <a id='make_code_defn'></a>
#
# # Step 2: `make.code.defn` \[Back to [top](#toc)\]
# $$\label{make_code_defn}$$
# +
# %%writefile $outfile_path__make_code_defn
# Main make.code.defn file for thorn IllinoisGRMHD
# Source files in this directory
SRCS = InitSymBound.C MoL_registration.C \
\
postpostinitial__set_symmetries__copy_timelevels.C \
\
convert_ADM_to_BSSN__enforce_detgtij_eq_1__and_compute_gtupij.C \
\
driver_evaluate_MHD_rhs.C \
compute_B_and_Bstagger_from_A.C \
driver_conserv_to_prims.C \
symmetry__set_gzs_staggered_gfs.C \
\
outer_boundaries.C
# -
# <a id='configuration__ccl'></a>
#
# # Step 3: `configuration.ccl` \[Back to [top](#toc)\]
# $$\label{configuration__ccl}$$
# +
# %%writefile $outfile_path__configuration__ccl
# Configuration definition for thorn IllinoisGRMHD
REQUIRES EOS_Omni Boundary CartGrid3D SpaceMask
PROVIDES IllinoisGRMHD
{
}
# -
# <a id='interface__ccl'></a>
#
# # Step 4: `interface.ccl` \[Back to [top](#toc)\]
# $$\label{interface__ccl}$$
# +
# %%writefile $outfile_path__interface__ccl
# Interface definition for thorn IllinoisGRMHD
implements: IllinoisGRMHD
inherits: ADMBase, Boundary, SpaceMask, Tmunubase, HydroBase, grid
includes header: IllinoisGRMHD_headers.h in IllinoisGRMHD_headers.h
USES INCLUDE: Symmetry.h
public:
#vvvvvvvv EVOLVED VARIABLES vvvvvvvv#
#cctk_real grmhd_conservatives type = GF TAGS='prolongation="Lag3"' Timelevels=3
cctk_real grmhd_conservatives type = GF Timelevels=3
{
rho_star,tau,mhd_st_x,mhd_st_y,mhd_st_z # Note that st = Stilde, as mhd_st_i = \tilde{S}_i.
} "Evolved mhd variables"
# These variables are semi-staggered:
# Ax is defined on the semi-staggered grid (i,j+1/2,k+1/2)
# WARNING: WILL NOT WORK PROPERLY WITHOUT SEMI-STAGGERED PROLONGATION/RESTRICTION:
cctk_real em_Ax type = GF Timelevels=3 tags='Prolongation="STAGGER011"'
{
Ax
} "x-component of the vector potential, evolved when constrained_transport_scheme==3"
# Ay is defined on the semi-staggered grid (i+1/2,j,k+1/2)
# WARNING: WILL NOT WORK PROPERLY WITHOUT SEMI-STAGGERED PROLONGATION/RESTRICTION:
cctk_real em_Ay type = GF Timelevels=3 tags='Prolongation="STAGGER101"'
{
Ay
} "y-component of the vector potential, evolved when constrained_transport_scheme==3"
# WARNING: WILL NOT WORK PROPERLY WITHOUT SEMI-STAGGERED PROLONGATION/RESTRICTION:
# Az is defined on the semi-staggered grid (i+1/2,j+1/2,k)
cctk_real em_Az type = GF Timelevels=3 tags='Prolongation="STAGGER110"'
{
Az
} "z-component of the vector potential, evolved when constrained_transport_scheme==3"
# psi6phi is defined on the staggered grid (i+1/2,j+1/2,k+1/2)
# WARNING: WILL NOT WORK PROPERLY WITHOUT FULLY-STAGGERED PROLONGATION/RESTRICTION:
#
cctk_real em_psi6phi type = GF Timelevels=3 tags='Prolongation="STAGGER111"'
{
psi6phi
} "sqrt{gamma} Phi, where Phi is the em scalar potential"
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv PRIMITIVE VARIABLES vvvvvvv#
# TODO: split into groups with well defined symmetry properties: (rho_b, P, u0), (vx,vy,vz)
cctk_real grmhd_primitives_allbutBi type = GF TAGS='InterpNumTimelevels=1 prolongation="none"'
{
rho_b,P,vx,vy,vz
} "Primitive variables density, pressure, and components of three velocity v^i. Note that v^i is defined in terms of 4-velocity as: v^i = u^i/u^0. Note that this definition differs from the Valencia formalism."
# It is useful to split Bi from Bi_stagger, since we're generally only interested in outputting Bi for diagnostics
cctk_real grmhd_primitives_Bi type = GF TAGS='InterpNumTimelevels=1 prolongation="none"'
{
Bx,By,Bz
} "B-field components defined at vertices."
cctk_real grmhd_primitives_Bi_stagger type = GF TAGS='InterpNumTimelevels=1 prolongation="none"'
{
Bx_stagger,By_stagger,Bz_stagger
} "B-field components defined at staggered points [Bx_stagger at (i+1/2,j,k),By_stagger at (i,j+1/2,k),Bz_stagger at (i,j,k+1/2)]."
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv BSSN-based quantities, computed from ADM quantities.v vvvvvvv#
cctk_real BSSN_quantities type = GF TAGS='prolongation="none" Checkpoint="no"'
{
gtxx,gtxy,gtxz,gtyy,gtyz,gtzz,gtupxx,gtupxy,gtupxz,gtupyy,gtupyz,gtupzz,phi_bssn,psi_bssn,lapm1
} "BSSN quantities, computed from ADM quantities"
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
private:
#vvvvvvv DIAGNOSTIC GRIDFUNCTIONS vvvvvvv#
cctk_real diagnostic_gfs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
failure_checker
} "Gridfunction to track conservative-to-primitives solver fixes. Beware that this gridfunction is overwritten at each RK substep."
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv TEMPORARY VARIABLES FOR RECONSTRUCTION vvvvvvv#
cctk_real grmhd_primitives_reconstructed_temps type = GF TAGS='prolongation="none" Checkpoint="no"'
{
ftilde_gf,temporary,
rho_br,Pr,vxr,vyr,vzr,Bxr,Byr,Bzr,Bx_staggerr,By_staggerr,Bz_staggerr,
rho_bl,Pl,vxl,vyl,vzl,Bxl,Byl,Bzl,Bx_staggerl,By_staggerl,Bz_staggerl,
vxrr,vxrl,vyrr,vyrl,vzrr,vzrl,vxlr,vxll,vylr,vyll,vzlr,vzll
} "Temporary variables used for primitives reconstruction"
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv RHS VARIABLES vvvvvvv#
cctk_real grmhd_conservatives_rhs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
rho_star_rhs,tau_rhs,st_x_rhs,st_y_rhs,st_z_rhs
} "Storage for the right-hand side of the partial_t rho_star, partial_t tau, and partial_t tilde{S}_i equations. Needed for MoL timestepping."
cctk_real em_Ax_rhs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
Ax_rhs
} "Storage for the right-hand side of the partial_t A_x equation. Needed for MoL timestepping."
cctk_real em_Ay_rhs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
Ay_rhs
} "Storage for the right-hand side of the partial_t A_y equation. Needed for MoL timestepping."
cctk_real em_Az_rhs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
Az_rhs
} "Storage for the right-hand side of the partial_t A_z equation. Needed for MoL timestepping."
cctk_real em_psi6phi_rhs type = GF TAGS='prolongation="none" Checkpoint="no"'
{
psi6phi_rhs
} "Storage for the right-hand side of the partial_t (psi^6 Phi) equation. Needed for MoL timestepping."
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv TEMPORARY VARIABLES USEFUL FOR A-FIELD EVOLUTION vvvvvvv#
cctk_real grmhd_cmin_cmax_temps type = GF TAGS='prolongation="none" Checkpoint="no"'
{
cmin_x,cmax_x,
cmin_y,cmax_y,
cmin_z,cmax_z
} "Store min and max characteristic speeds in all three directions."
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv TEMPORARY VARIABLES USEFUL FOR FLUX COMPUTATION vvvvvvv#
cctk_real grmhd_flux_temps type = GF TAGS='prolongation="none" Checkpoint="no"'
{
rho_star_flux,tau_flux,st_x_flux,st_y_flux,st_z_flux
} "Temporary variables for storing the flux terms of tilde{S}_i."
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
#vvvvvvv T^{\mu \nu}, stored to avoid expensive recomputation vvvvvvv#
cctk_real TUPmunu type = GF TAGS='prolongation="none" Checkpoint="no"'
{
TUPtt,TUPtx,TUPty,TUPtz,TUPxx,TUPxy,TUPxz,TUPyy,TUPyz,TUPzz
} "T^{mu nu}, stored to avoid expensive recomputation"
#^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^#
###########################################################################
####################################################
### Functions provided by MoL for registration ###
####################################################
CCTK_INT FUNCTION MoLRegisterEvolved(CCTK_INT IN EvolvedIndex, \
CCTK_INT IN RHSIndex)
CCTK_INT FUNCTION MoLRegisterEvolvedSlow(CCTK_INT IN EvolvedIndex, \
CCTK_INT IN RHSIndexSlow)
CCTK_INT FUNCTION MoLRegisterConstrained(CCTK_INT IN ConstrainedIndex)
CCTK_INT FUNCTION MoLRegisterEvolvedGroup(CCTK_INT IN EvolvedIndex, \
CCTK_INT IN RHSIndex)
CCTK_INT FUNCTION MoLRegisterEvolvedGroupSlow(CCTK_INT IN EvolvedIndex, \
CCTK_INT IN RHSIndexSlow)
CCTK_INT FUNCTION MoLRegisterConstrainedGroup(CCTK_INT IN ConstrainedIndex)
CCTK_INT FUNCTION MoLRegisterSaveAndRestoreGroup(CCTK_INT IN SandRIndex)
USES FUNCTION MoLRegisterEvolved
USES FUNCTION MoLRegisterEvolvedSlow
USES FUNCTION MoLRegisterConstrained
USES FUNCTION MoLRegisterEvolvedGroup
USES FUNCTION MoLRegisterEvolvedGroupSlow
USES FUNCTION MoLRegisterConstrainedGroup
USES FUNCTION MoLRegisterSaveAndRestoreGroup
#########################################
### Aliased functions from Boundary ###
#########################################
CCTK_INT FUNCTION Boundary_SelectVarForBC(CCTK_POINTER_TO_CONST IN GH, \
CCTK_INT IN faces, CCTK_INT IN boundary_width, CCTK_INT IN table_handle, \
CCTK_STRING IN var_name, CCTK_STRING IN bc_name)
CCTK_INT FUNCTION Boundary_SelectGroupForBC(CCTK_POINTER_TO_CONST IN GH, \
CCTK_INT IN faces, CCTK_INT IN boundary_width, CCTK_INT IN table_handle, \
CCTK_STRING IN group_name, CCTK_STRING IN bc_name)
USES FUNCTION Boundary_SelectVarForBC
USES FUNCTION Boundary_SelectGroupForBC
###########################################################################
#########################################
### Aliased functions from Carpet ###
#########################################
CCTK_INT FUNCTION \
GetRefinementLevel \
(CCTK_POINTER_TO_CONST IN cctkGH)
USES FUNCTION GetRefinementLevel
# -
# <a id='param__ccl'></a>
#
# # Step 5: `param.ccl` \[Back to [top](#toc)\]
# $$\label{param__ccl}$$
# +
# %%writefile $outfile_path__param__ccl
# Parameter definitions for thorn IllinoisGRMHD
shares: ADMBase
USES CCTK_INT lapse_timelevels
USES CCTK_INT shift_timelevels
USES CCTK_INT metric_timelevels
#########################################################
restricted:
#########################################################
# Verbosity Level
KEYWORD verbose "Determines how much evolution information is output" STEERABLE=ALWAYS
{
"no" :: "Complete silence"
"essential" :: "Essential health monitoring of the GRMHD evolution: Information about conservative-to-primitive fixes, etc."
"essential+iteration output" :: "Outputs health monitoring information, plus a record of which RK iteration. Very useful for backtracing a crash."
} "essential+iteration output"
#########################################################
#########################################################
# SPEED LIMIT: Set maximum relativistic gamma factor
#
REAL GAMMA_SPEED_LIMIT "Maximum relativistic gamma factor."
{
1:* :: "Positive > 1, though you'll likely have troubles far above 10."
} 10.0
#########################################################
#########################################################
# CONSERV TO PRIMS PARAMETERS
# FIXME: Enable this parameter! IllinoisGRMHD is currently hard-coded to tau_stildefix_enable=2.
#INT tau_stildefix_enable "tau<0 fix in primitive_vars_hybrid2 to reduce number of Font fixes, especially in puncture+matter evolutions" STEERABLE=ALWAYS
#{
# 0:3 :: "zero (disable), one (enable everywhere), or two (enable only where Psi6 > Psi6threshold [i.e., inside the horizon, where B's are set to zero], or three (kludge: set B=0 if tau<0 inside horizon))"
#} 0
BOOLEAN update_Tmunu "Update Tmunu, for RHS of Einstein's equations?" STEERABLE=ALWAYS
{
} "yes"
############################
# Limiters on tau and rho_b:
REAL tau_atm "Floor value on the energy variable tau (cf. tau_stildefix_enable). Given the variety of systems this code may encounter, there *is no reasonable default*. Effectively the current (enormous) value should disable the tau_atm floor. Please set this in your initial data thorn, and reset at will during evolutions." STEERABLE=ALWAYS
{
0:* :: "Positive"
} 1e100
REAL rho_b_atm "Floor value on the baryonic rest mass density rho_b (atmosphere). Given the variety of systems this code may encounter, there *is no reasonable default*. Your run will die unless you override this default value in your initial data thorn." STEERABLE=ALWAYS
{
*:* :: "Allow for negative values. This enables us to debug the code and verify if rho_b_atm is properly set."
} 1e200
REAL rho_b_max "Ceiling value on the baryonic rest mass density rho_b. The enormous value effectively disables this ceiling by default. It can be quite useful after a black hole has accreted a lot of mass, leading to enormous densities inside the BH. To enable this trick, set rho_b_max in your initial data thorn! You are welcome to change this parameter mid-run (after restarting from a checkpoint)." STEERABLE=ALWAYS
{
0:* :: "Note that you will have problems unless rho_b_atm<rho_b_max"
} 1e300
############################
INT conserv_to_prims_debug "0: no, 1: yes" STEERABLE=ALWAYS
{
0:1 :: "zero (no) or one (yes)"
} 0
REAL Psi6threshold "Where Psi^6 > Psi6threshold, we assume we're inside the horizon in the primitives solver, and certain limits are relaxed or imposed" STEERABLE=ALWAYS
{
*:* :: "Can set to anything"
} 1e100
#########################################################
#########################################################
# EQUATION OF STATE PARAMS, LOOK FOR MORE IN interface.ccl!
INT neos "number of parameters in EOS table. If you want to increase from the default max value, you MUST also set eos_params_arrays1 and eos_params_arrays2 in interface.ccl to be consistent!"
{
1:10 :: "Any integer between 1 and 10"
} 1
REAL Gamma_th "thermal gamma parameter"
{
0:* :: "Physical values"
-1 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1
REAL K_ppoly_tab0 "Also known as k_tab[0], this is the polytropic constant for the lowest density piece of the piecewise polytrope. All other k_tab EOS array elements are set from user-defined rho_tab EOS array elements and by enforcing continuity in the equation of state."
{
0:* :: "Physical values"
-1 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1
REAL rho_ppoly_tab_in[10] "Set polytropic rho parameters"
{
0.0:* :: "after this time (inclusively)"
-1.0 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1.0
REAL Gamma_ppoly_tab_in[11] "Set polytropic rho parameters"
{
0.0:* :: "after this time (inclusively)"
-1.0 :: "forbidden value to make sure it is explicitly set in the parfile"
} -1.0
#########################################################
#########################################################
# OUTER BOUNDARY CONDITION CHOICE
KEYWORD Matter_BC "Chosen Matter boundary condition"
{
"outflow" :: "Outflow boundary conditions"
"frozen" :: "Frozen boundaries"
} "outflow"
KEYWORD EM_BC "EM field boundary condition"
{
"copy" :: "Copy data from nearest boundary point"
"frozen" :: "Frozen boundaries"
} "copy"
#########################################################
#########################################################
# SYMMETRY BOUNDARY PARAMS. Needed for handling staggered gridfunctions.
KEYWORD Symmetry "Currently only no symmetry supported, though work has begun in adding equatorial-symmetry support. FIXME: Extend ET symmetry interface to support symmetries on staggered gridfunctions"
{
"none" :: "no symmetry, full 3d domain"
} "none"
REAL Sym_Bz "In-progress equatorial symmetry support: Symmetry parameter across z axis for magnetic fields = +/- 1"
{
-1.0:1.0 :: "Set to +1 or -1."
} 1.0
#########################################################
###############################################################################################
private:
#########################################################
# EVOLUTION PARAMS
REAL damp_lorenz "Damping factor for the generalized Lorenz gauge. Has units of 1/length = 1/M. Typically set this parameter to 1.5/(maximum Delta t on AMR grids)." STEERABLE=ALWAYS
{
*:* :: "any real"
} 0.0
#########################################################
# -
# <a id='schedule__ccl'></a>
#
# # Step 6: `schedule.ccl` \[Back to [top](#toc)\]
# $$\label{schedule__ccl}$$
# +
# %%writefile $outfile_path__schedule__ccl
# Scheduler setup for IllinoisGRMHD
STORAGE: ADMBase::metric[metric_timelevels], ADMBase::curv[metric_timelevels], ADMBase::lapse[lapse_timelevels], ADMBase::shift[shift_timelevels]
STORAGE: IllinoisGRMHD::BSSN_quantities
STORAGE: grmhd_conservatives[3],em_Ax[3],em_Ay[3],em_Az[3],em_psi6phi[3]
STORAGE: grmhd_primitives_allbutBi,grmhd_primitives_Bi,grmhd_primitives_Bi_stagger,grmhd_primitives_reconstructed_temps,grmhd_conservatives_rhs,em_Ax_rhs,em_Ay_rhs,em_Az_rhs,em_psi6phi_rhs,grmhd_cmin_cmax_temps,grmhd_flux_temps,TUPmunu,diagnostic_gfs
####################
# RUN INITIALLY ONLY
schedule IllinoisGRMHD_RegisterVars in MoL_Register after BSSN_RegisterVars after lapse_RegisterVars
{
LANG: C
OPTIONS: META
} "Register evolved, rhs variables in IllinoisGRMHD for MoL"
# Tells the symmetry thorn how to apply symmetries on each gridfunction:
schedule IllinoisGRMHD_InitSymBound at BASEGRID after Lapse_InitSymBound
{
LANG: C
} "Schedule symmetries"
####################
####################
# POSTPOSTINITIAL
# HydroBase_Con2Prim in CCTK_POSTPOSTINITIAL sets conserv to prim then
# outer boundaries (OBs, which are technically disabled). The post OB
# SYNCs actually reprolongate the conservative variables, making cons
# and prims INCONSISTENT. So here we redo the con2prim, avoiding the
# SYNC afterward, then copy the result to other timelevels"
schedule GROUP IllinoisGRMHD_PostPostInitial at CCTK_POSTPOSTINITIAL before MoL_PostStep after HydroBase_Con2Prim
{
} "HydroBase_Con2Prim in CCTK_POSTPOSTINITIAL sets conserv to prim then outer boundaries (OBs, which are technically disabled). The post OB SYNCs actually reprolongate the conservative variables, making cons and prims INCONSISTENT. So here we redo the con2prim, avoiding the SYNC afterward, then copy the result to other timelevels"
schedule IllinoisGRMHD_InitSymBound in IllinoisGRMHD_PostPostInitial as postid before compute_b
{
SYNC: grmhd_conservatives,em_Ax,em_Ay,em_Az,em_psi6phi
LANG: C
} "Schedule symmetries -- Actually just a placeholder function to ensure prolongations / processor syncs are done BEFORE outer boundaries are updated."
# Easiest primitives to solve for: B^i
schedule IllinoisGRMHD_compute_B_and_Bstagger_from_A in IllinoisGRMHD_PostPostInitial as compute_b after postid after empostid after lapsepostid
{
# This is strictly a processor sync, as prolongation is disabled for all primitives & B^i's.
SYNC: grmhd_primitives_Bi,grmhd_primitives_Bi_stagger # FIXME: Are both SYNC's necessary?
LANG: C
} "Compute B and B_stagger from A SYNC: grmhd_primitives_Bi,grmhd_primitives_Bi_stagger"
# Nontrivial primitives solve, for P,rho_b,vx,vy,vz:
schedule IllinoisGRMHD_conserv_to_prims in IllinoisGRMHD_PostPostInitial after compute_b
{
LANG: C
} "Compute primitive variables from conservatives. This is non-trivial, requiring a Newton-Raphson root-finder."
# Copy data to other timelevels.
schedule IllinoisGRMHD_PostPostInitial_Set_Symmetries__Copy_Timelevels in IllinoisGRMHD_PostPostInitial as mhdpostid after compute_b after p2c
{
LANG: C
} "Compute post-initialdata quantities"
####################
####################
# RHS EVALUATION
schedule IllinoisGRMHD_driver_evaluate_MHD_rhs in MoL_CalcRHS as IllinoisGRMHD_RHS_eval after bssn_rhs after shift_rhs
{
LANG: C
} "Evaluate RHSs of GR Hydro & GRMHD equations"
####################
############################################################
# COMPUTE B FROM A & RE-SOLVE FOR PRIMITIVES
# After a full timestep, there are two types of boundaries that need filling:
# (A) Outer boundaries (on coarsest level)
# (B) AMR grid refinement boundaries
# (A) OUTER BOUNDARY STEPS:
# ( 0) Synchronize (prolongate/restrict) all evolved variables
# ( 1) Apply outer boundary conditions (BCs) on A_{\mu}
# ( 2) Compute B^i from A_i everywhere, synchronize (processor sync) B^i
# ( 3) Call con2prim to get consistent primitives {P,rho_b,vx,vy,vz} and conservatives at all points (if no restriction, really only need interior)
# ( 4) Apply outer BCs on {P,rho_b,vx,vy,vz}, recompute conservatives.
# (B) AMR GRID REFINEMENT BOUNDARY STEPS:
# Same as steps 0,2,3 above. Just need if() statements in steps 1,4 to prevent "outer boundaries" being updated
# Problem: all the sync's in outer boundary updates might just overwrite prolongated values.
############################################################
schedule IllinoisGRMHD_InitSymBound in HydroBase_Boundaries as Boundary_SYNCs before compute_B_postrestrict
{
SYNC: grmhd_conservatives,em_Ax,em_Ay,em_Az,em_psi6phi
LANG: C
} "Schedule symmetries -- Actually just a placeholder function to ensure prolongations / processor syncs are done BEFORE outer boundaries are updated."
schedule IllinoisGRMHD_outer_boundaries_on_A_mu in HydroBase_Boundaries after Boundary_SYNCs before mhd_conserv2prims_postrestrict
{
LANG: C
} "Apply linear extrapolation BCs on A_{mu}, so that BCs are flat on B^i."
# Easiest primitives to solve for: B^i.
# Note however that B^i depends on derivatives of A_{\mu}, so a SYNC is necessary on B^i.
schedule IllinoisGRMHD_compute_B_and_Bstagger_from_A in HydroBase_Boundaries after outer_boundaries_on_A_mu
{
# This is strictly a processor sync, as prolongation is disabled for all B^i's.
SYNC: grmhd_primitives_Bi,grmhd_primitives_Bi_stagger # FIXME: Are both SYNC's necessary?
LANG: C
} "Compute B and B_stagger from A, SYNC: grmhd_primitives_Bi,grmhd_primitives_Bi_stagger"
# Nontrivial primitives solve, for P,rho_b,vx,vy,vz.
schedule IllinoisGRMHD_conserv_to_prims in AddToTmunu after compute_B_and_Bstagger_from_A
{
LANG: C
} "Compute primitive variables from conservatives. This is non-trivial, requiring a Newton-Raphson root-finder."
schedule IllinoisGRMHD_outer_boundaries_on_P_rho_b_vx_vy_vz in AddToTmunu after IllinoisGRMHD_conserv_to_prims
{
# We must sync {P,rho_b,vx,vy,vz} here.
SYNC: grmhd_primitives_allbutBi
LANG: C
} "Apply outflow-only, flat BCs on {P,rho_b,vx,vy,vz}. Outflow only == velocities pointed inward from outer boundary are set to zero."
##########################################################
# -
# <a id='code_validation__txt'></a>
#
# # Step 7: `code_validation.txt` \[Back to [top](#toc)\]
# $$\label{code_validation__txt}$$
# +
# %%writefile $outfile_path__code_validation__txt
0 0 0.129285232345409 0
6 0.375 0.127243949890016 0
12 0.75 0.12663194218958 9.99201e-16
18 1.125 0.126538236778999 0
24 1.5 0.126693696091085 0
30 1.875 0.126354095699745 0
36 2.25 0.125536381948334 0
42 2.625 0.124535850791511 0
48 3 0.123592701331336 -1.9984e-15
54 3.375 0.122798174115381 6.00908e-15
60 3.75 0.122119289857302 5.20001e-14
66 4.125 0.121490429374158 -7.219e-12
72 4.5 0.120874029334284 9.062e-12
78 4.875 0.120259717466328 8.073e-12
84 5.25 0.11967641900387 6.773e-12
90 5.625 0.119123915533179 3.9644e-11
96 6 0.118609688443716 8.7028e-11
102 6.375 0.118142085593448 1.0356e-10
108 6.75 0.117724554127494 -2.1806e-11
114 7.125 0.117368613539 -2.98919e-10
120 7.5 0.117103537221293 -6.55126e-10
126 7.875 0.116916242747612 -8.58055e-10
132 8.25 0.116788537512057 -6.40583e-10
138 8.625 0.116760604471516 1.2299e-10
144 9 0.116800459472318 1.17986e-09
150 9.375 0.116876502360487 2.30931e-09
156 9.75 0.117001584159343 3.22793e-09
162 10.125 0.117152245129722 3.70731e-09
168 10.5 0.117321427488885 3.80249e-09
174 10.875 0.117504914930401 3.18643e-09
180 11.25 0.117677337706038 1.53467e-09
186 11.625 0.11783535130534 -1.0388e-09
192 12 0.11797817411705 -4.09727e-09
198 12.375 0.118099716148388 -7.36175e-09
204 12.75 0.118197132134204 -1.11993e-08
210 13.125 0.118268160681997 -1.54798e-08
216 13.5 0.118308938021455 -1.879e-08
222 13.875 0.118311817073909 -1.9184e-08
228 14.25 0.118266030948925 -1.56794e-08
234 14.625 0.118178894932027 -8.96657e-09
240 15 0.118048178792271 -8.64983e-10
# -
# <a id='code_validation'></a>
#
# # Step 8: Code validation \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# <a id='code_validation__make_code_defn'></a>
#
# ## Step 8.a: `make.code.defn` \[Back to [top](#toc)\]
# $$\label{code_validation__make_code_defn}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/src/make.code.defn"
original_IGM_file_name = "make.code.defn-original"
original_IGM_file_path = os.path.join(IGM_src_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__make_code_defn = !diff $original_IGM_file_path $outfile_path__make_code_defn
if Validation__make_code_defn == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for make.code.defn: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for make.code.defn: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__make_code_defn:
print(diff_line)
# -
# <a id='code_validation__configuration__ccl'></a>
#
# ## Step 8.b: `configuration.ccl` \[Back to [top](#toc)\]
# $$\label{code_validation__configuration__ccl}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/configuration.ccl"
original_IGM_file_name = "configuration-original.ccl"
original_IGM_file_path = os.path.join(IGM_main_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__configuration__ccl = !diff $original_IGM_file_path $outfile_path__configuration__ccl
if Validation__configuration__ccl == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for configuration.ccl: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for configuration.ccl: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__configuration__ccl:
print(diff_line)
# -
# <a id='code_validation__interface__ccl'></a>
#
# ## Step 8.c: `interface.ccl` \[Back to [top](#toc)\]
# $$\label{code_validation__interface__ccl}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/interface.ccl"
original_IGM_file_name = "interface-original.ccl"
original_IGM_file_path = os.path.join(IGM_main_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__interface__ccl = !diff $original_IGM_file_path $outfile_path__interface__ccl
if Validation__interface__ccl == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for interface.ccl: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for interface.ccl: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__interface__ccl:
print(diff_line)
# -
# <a id='code_validation__param__ccl'></a>
#
# ## Step 8.d: `param.ccl` \[Back to [top](#toc)\]
# $$\label{code_validation__param__ccl}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/param.ccl"
original_IGM_file_name = "param-original.ccl"
original_IGM_file_path = os.path.join(IGM_main_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__param__ccl = !diff $original_IGM_file_path $outfile_path__param__ccl
if Validation__param__ccl == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for param.ccl: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for param.ccl: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__param__ccl:
print(diff_line)
# -
# <a id='code_validation__schedule__ccl'></a>
#
# ## Step 8.e: `schedule.ccl` \[Back to [top](#toc)\]
# $$\label{code_validation__schedule__ccl}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/schedule.ccl"
original_IGM_file_name = "schedule-original.ccl"
original_IGM_file_path = os.path.join(IGM_main_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__schedule__ccl = !diff $original_IGM_file_path $outfile_path__schedule__ccl
if Validation__schedule__ccl == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for schedule.ccl: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for schedule.ccl: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__schedule__ccl:
print(diff_line)
# -
# <a id='code_validation__code_validation__txt'></a>
#
# ## Step 8.f: `code_validation.txt` \[Back to [top](#toc)\]
# $$\label{code_validation__code_validation__txt}$$
#
# First we download the original `IllinoisGRMHD` source code and then compare it to the source code generated by this tutorial notebook.
# +
# Verify if the code generated by this tutorial module
# matches the original IllinoisGRMHD source code
# First download the original IllinoisGRMHD source code
import urllib
from os import path
original_IGM_file_url = "https://bitbucket.org/zach_etienne/wvuthorns/raw/5611b2f0b17135538c9d9d17c7da062abe0401b6/IllinoisGRMHD/code_validation.txt"
original_IGM_file_name = "code_validation-original.txt"
original_IGM_file_path = os.path.join(IGM_main_dir_path,original_IGM_file_name)
# Then download the original IllinoisGRMHD source code
# We try it here in a couple of ways in an attempt to keep
# the code more portable
try:
original_IGM_file_code = urllib.request.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
try:
original_IGM_file_code = urllib.urlopen(original_IGM_file_url).read().decode("utf-8")
# Write down the file the original IllinoisGRMHD source code
with open(original_IGM_file_path,"w") as file:
file.write(original_IGM_file_code)
except:
# If all else fails, hope wget does the job
# !wget -O $original_IGM_file_path $original_IGM_file_url
# Perform validation
# Validation__code_validation__txt = !diff $original_IGM_file_path $outfile_path__code_validation__txt
if Validation__code_validation__txt == []:
# If the validation passes, we do not need to store the original IGM source code file
# !rm $original_IGM_file_path
print("Validation test for code_validation.txt: PASSED!")
else:
# If the validation fails, we keep the original IGM source code file
print("Validation test for code_validation.txt: FAILED!")
# We also print out the difference between the code generated
# in this tutorial module and the original IGM source code
print("Diff:")
for diff_line in Validation__code_validation__txt:
print(diff_line)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 9: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-IllinoisGRMHD__Configuration_files.pdf](Tutorial-IllinoisGRMHD__Configuration_files.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means).
latex_nrpy_style_path = os.path.join(nrpy_dir_path,"latex_nrpy_style.tplx")
# #!jupyter nbconvert --to latex --template $latex_nrpy_style_path --log-level='WARN' Tutorial-IllinoisGRMHD__Configuration_files.ipynb
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Configuration_files.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Configuration_files.tex
# #!pdflatex -interaction=batchmode Tutorial-IllinoisGRMHD__Configuration_files.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
| IllinoisGRMHD/doc/Tutorial-IllinoisGRMHD__Configuration_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
################################################################################
## Code adapted from demo_caiman_cnmf_3D as imported from github 21/11/2018
## https://github.com/flatironinstitute/CaImAn
################################################################################
import cde_cell_functions as cc
# Import relevant packages
#===============================================================================
import imp
import logging
import matplotlib.pyplot as plt
import numpy as np
import os
import psutil
import shutil
from scipy.ndimage.filters import gaussian_filter
import scipy.sparse
import sys
import re
from skimage.external.tifffile import imread
from skimage import io
import warnings
from IPython.display import clear_output
import copy
from importlib import reload
from PIL import Image
import datetime
# Caiman setup
#-------------------------------------------------------------------------------
import caiman as cm
import caiman.source_extraction.cnmf as cnmf
from caiman.utils.visualization import nb_view_patches3d
from caiman.source_extraction.cnmf import params as params
from caiman.components_evaluation import evaluate_components, estimate_components_quality_auto
from caiman.motion_correction import MotionCorrect
from caiman.cluster import setup_cluster
from caiman.paths import caiman_datadir
from caiman.utils.visualization import inspect_correlation_pnr
# Jupyter specific autoreloading for external functions (in case changes are made)
# # %load_ext autoreload
# # %autoreload
# +
# Housekeeping
#===============================================================================
# Module flags
display_movie = False # play movie of tifs that are loaded
save_results = False # flag to save results or not
# Define folder locations
#-------------------------------------------------------------------------------
reload(cc)
Fbase = '/Users/roschkoenig/Dropbox/Research/1812 Critical Dynamics Epilepsy'
Fsave = '/Users/roschkoenig/Dropbox/Research/1812 Critical Dynamics Epilepsy Data'
Fscripts = Fbase + os.sep + '03 - Cell detection'
Fdata = '/Volumes/ALBERS/1812 Critical Dynamics in Epilepsy'
Zfish = cc.cde_cell_fishspec(Fdata, 'RM')
# -
Fish = Zfish[-2:]
Fish[0]["Name"]
# # Split images up into planes
for z in Fish:
print('----------------------------------------------------------------------------')
print('Currently working on ' + z["Name"])
print('----------------------------------------------------------------------------')
cc.cde_cell_planesave(Fdata, z, mxpf = 7000)
# # Run actual cell segmentation
imp.reload(cc),
Pfish = cc.cde_cell_fishspec(Fdata, 'PL')
f = 3
# +
for f in range(len(Pfish)):
for c in range(len(Pfish[f]["Cond"])):
planes = Pfish[f]["Cond"][c]["Plane"]
Estimates = []
for p in range(len(planes)-1): # That last plane is a bitch
fname = planes[p]["Tpaths"]
fr = 4 # frame rate (Hz)
decay_time = 0.5 # approximate length of transient event in seconds
gSig = (4,4) # expected half size of neurons
p = 1 # order of AR indicator dynamics
min_SNR = 1 # minimum SNR for accepting new components
rval_thr = 0.90 # correlation threshold for new component inclusion
ds_factor = 1 # spatial downsampling factor (increases speed but may lose some fine structure)
gnb = 2 # number of background components
gSig = tuple(np.ceil(np.array(gSig)/ds_factor).astype('int')) # recompute gSig if downsampling is involved
mot_corr = True # flag for online motion correction
pw_rigid = False # flag for pw-rigid motion correction (slower but potentially more accurate)
max_shifts_online = np.ceil(10./ds_factor).astype('int') # maximum allowed shift during motion correction
sniper_mode = True # flag using a CNN to detect new neurons (o/w space correlation is used)
init_batch = 200 # number of frames for initialization (presumably from the first file)
expected_comps = 500 # maximum number of expected components used for memory pre-allocation (exaggerate here)
dist_shape_update = True # flag for updating shapes in a distributed way
min_num_trial = 10 # number of candidate components per frame
K = 2 # initial number of components
epochs = 2 # number of passes over the data
show_movie = False # show the movie with the results as the data gets processed
params_dict = {'fnames': fname,
'fr': fr,
'decay_time': decay_time,
'gSig': gSig,
'p': p,
'min_SNR': min_SNR,
'rval_thr': rval_thr,
'ds_factor': ds_factor,
'nb': gnb,
'motion_correct': mot_corr,
'init_batch': init_batch,
'init_method': 'bare',
'normalize': True,
'expected_comps': expected_comps,
'sniper_mode': sniper_mode,
'dist_shape_update' : dist_shape_update,
'min_num_trial': min_num_trial,
'K': K,
'epochs': epochs,
'max_shifts_online': max_shifts_online,
'pw_rigid': pw_rigid,
'show_movie': show_movie}
opts = cnmf.params.CNMFParams(params_dict=params_dict)
clear_output()
print('-----------------------------------------------------------------------')
print('Currently processing condition ' + Pfish[f]["Cond"][c]["Name"])
print('> Plane ' + str(p) + ' of ' + str(len(planes)))
print('-----------------------------------------------------------------------')
cmn = cnmf.online_cnmf.OnACID(params=opts)
cmn.fit_online()
Estimates.append({'Spatial':cmn.estimates.A,'Temporal':cmn.estimates.C,'Background':cmn.estimates.b})
Pfish[f]["Cond"][c].update({"CMN":Estimates})
# Save everyhting into folder
#---------------------------------------------------------------------------------
Fcmn = Fsave + os.sep + 'Analysis' + os.sep + 'CMN' + os.sep + Pfish[f]["Name"]
if not os.path.exists(Fcmn): os.makedirs(Fcmn)
for c in range(len(Pfish[f]["Cond"])):
Fccond = Fcmn + os.sep + Pfish[f]["Cond"][c]["Name"]
if not os.path.exists(Fccond):
os.makedirs(Fccond)
for p in range(len(Pfish[f]["Cond"][c]["CMN"])):
scipy.io.savemat(Fccond + os.sep + Pfish[f]["Name"] + '_P' + str(p).zfill(2), Pfish[f]["Cond"][c]["CMN"][p])
# +
# Save everyhting into folder
#---------------------------------------------------------------------------------
Fcmn = Fsave + os.sep + 'Analysis' + os.sep + 'CMN' + os.sep + Pfish[f]["Name"]
if not os.path.exists(Fcmn): os.makedirs(Fcmn)
for c in range(len(Pfish[f]["Cond"])):
Fccond = Fcmn + os.sep + Pfish[f]["Cond"][c]["Name"]
if not os.path.exists(Fccond):
os.makedirs(Fccond)
for p in range(len(Pfish[f]["Cond"][c]["CMN"])):
scipy.io.savemat(Fccond + os.sep + Pfish[f]["Name"] + '_P' + str(p).zfill(2), Pfish[f]["Cond"][c]["CMN"][p])
# -
Estimates = Secure_Estimates[0:9]
Pfish[f]["Cond"][0].update({"CMN":Estimates})
Pfish[0]['Cond'][0]['Path']
| 03 - Cell detection/cde_cell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/monocongo/model_learn/blob/master/notebooks/model_learn.ipynb)
# + [markdown] id="74l7lcFQk4kT" colab_type="text"
# ## Setup
#
# + [markdown] id="ixh2Tyl1FHaj" colab_type="text"
# In this first cell we''ll load the necessary libraries and setup some logging and display options.
# + id="JaCENoitkiXK" colab_type="code" colab={}
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.2f}'.format
# + [markdown] id="6tSwOT2bsUNM" colab_type="text"
# ## Pull data files from Google Drive
#
# + [markdown] id="6dljBwZdE_Es" colab_type="text"
# Install PyDrive which will be used to access Google Drive and kick off the process to authorize the notebook running in the Google Colaboratory environment to touch our Drive files. When this cell executes it'll provide a link to authenticate into a Google Drive account to instatiate a PyDrive client. The Drive account that is selected should be one which has access to our all variables dataset file that we'll use for training/testing our model.
# + id="SqSqvIptsV-t" colab_type="code" colab={}
# !pip install -U -q PyDrive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + [markdown] id="eyG5gCxjseUQ" colab_type="text"
# Next we'll create dataset files within the Google Colaboratory environment corresponding to the flow and time tendency dataset files located on our Google Drive.
#
# + id="1LdCggflswj7" colab_type="code" colab={}
filename_h0 = 'fv091x180L26_dry_HS.cam.h0.2000-12-27-00000.nc'
filename_h1 = 'fv091x180L26_dry_HS.cam.h1.2000-12-27-00000.nc'
id_h0 = '1vptBPguIMU4FrvkC91xAd7_wOxoylBW0'
id_h1 = '1ru8gmDKv8qPZGnfaTP2Sqv8Fsps48vAX'
file_h0 = drive.CreateFile({'id': id_h0}) # creates a file in the Colab env using the ID for file <filename_h0>
file_h1 = drive.CreateFile({'id': id_h1}) # creates a file in the Colab env using the ID for file <filename_h1>
file_h0.GetContentFile(filename_h0) # gets the file's contents and saves it as a local file named <filename_h0>
file_h1.GetContentFile(filename_h1) # gets the file's contents and saves it as a local file named <filename_h1>
# + [markdown] id="y0gBz25Glf-3" colab_type="text"
# Next we'll load our flow variables and time tendency forcings datasets into Xarray Dataset objects.
# + id="_cC_-nNSlWIO" colab_type="code" colab={}
# !pip install -U -q xarray
# !pip install -U -q netCDF4
import xarray as xr
data_h0 = xr.open_dataset(filename_h0)
data_h1 = xr.open_dataset(filename_h1)
# + [markdown] id="fLH3YQ2azUce" colab_type="text"
# ## Define the features and configure feature columns
#
# + [markdown] id="sh36g2stEz-l" colab_type="text"
# In TensorFlow, we indicate a feature's data type using a construct called a feature column. Feature columns store only a description of the feature data; they do not contain the feature data itself. As features we'll use the following flow variables:
#
# * U (west-east (zonal) wind, m/s)
# * V (south-north (meridional) wind, m/s)
# * T (temperature, K)
# * PS (surface pressure, Pa)
#
# We'll take the flow variables dataset and trim out all but the above variables, and use this as the data source for features.
#
# The variables correspond to Numpy arrays, and we'll use the shapes of the variable arrays as the shapes of the corresponding feature columns.
#
# + id="Pv3DemYvmTh1" colab_type="code" colab={}
# Define the input features as PS, T, U, and V.
# remove all non-feature variables and unrelated coordinate variables from the DataSet, in order to trim the memory footprint.
feature_vars = ['PS', 'T', 'U', 'V']
feature_coord_vars = ['time', 'lev', 'lat', 'lon']
for var in data_h0.variables:
if (var not in feature_vars) and (var not in feature_coord_vars):
data_h0 = data_h0.drop(var)
features = data_h0
# Configure numeric feature columns for the input features.
feature_columns = []
for var in feature_vars:
feature_columns.append(tf.feature_column.numeric_column(var, shape=features[var].shape))
# + [markdown] id="zoAKA7O6se2m" colab_type="text"
# Display the flow variables (features) DataSet.
# + id="7pk5TNd4sjVL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="fe96129b-6df1-4e07-ddfc-754cae9812cd"
features
# + id="cIEYStXorUUp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="a5c5e2a7-d4ca-4b93-897b-d3ecd2d6900b"
feature_columns
# + [markdown] id="ADi7KQHtBKMf" colab_type="text"
# ## Define the targets (labels)
#
# + [markdown] id="VjEZ2K5HEF6t" colab_type="text"
# Time tendency forcings are the targets (labels) that our model should learn to predict.
#
# * PTTEND (time tendency of the temperature)
# * PUTEND (time tendency of the zonal wind)
# * PVTEND (time tendency of the meridional wind)
#
# We'll take the time tendency forcings dataset and trim out all other variables so we can use this as the data source for targets.
# + id="hN19tItj6hrN" colab_type="code" colab={}
# Define the targets (labels) as PTTEND, PUTEND, and PVTEND.
# Remove all non-target variables and unrelated coordinate variables from the DataSet, in order to trim the memory footprint.
target_vars = ['PTTEND', 'PUTEND', 'PVTEND']
target_coord_vars = ['time', 'lev', 'lat', 'lon']
for var in data_h1.variables:
if (var not in target_vars) and (var not in target_coord_vars):
data_h1 = data_h1.drop(var)
targets = data_h1
# + [markdown] id="bUXvCOIgsqx1" colab_type="text"
# Display the time tendency forcings (targets/labels) dataset.
# + id="DM8R4w3fszNv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="fbf6b4d6-30ca-4d60-b8c4-070e5edcd4dd"
targets
# + [markdown] id="7YNrGbEeNN2c" colab_type="text"
# Confirm the compatability of our features and targets datasets, in terms of dimensions and coordinates, to provide an initial sanity check.
# + id="P5RsmkV9IvZQ" colab_type="code" colab={}
if features.dims != targets.dims:
print("WARNING: Unequal dimensions")
else:
for coord in features.coords:
if not (features.coords[coord] == targets.coords[coord]).all():
print("WARNING: Unequal {} coordinates".format(coord))
# + [markdown] id="J-yXL6dZMWjP" colab_type="text"
# ## Split the data into training, validation, and testing datasets
# + [markdown] id="ApCinhR8__SL" colab_type="text"
# We'll initially split the dataset into training, validation, and testing datasets with 50% for training and 25% each for validation and testing. We'll use the longitude dimension to split since it has 180 points and divides evenly by four. We get every other longitude starting at the first longitude to get 50% of the dataset for training, then every fourth longitude starting at the second longitude to get 25% of the dataset for validation, and every fourth longitude starting at the fourth longitude to get 25% of the dataset for testing.
# + id="Q6waMx-cMg71" colab_type="code" colab={}
lon_range_training = list(range(0, features.dims['lon'], 2))
lon_range_validation = list(range(1, features.dims['lon'], 4))
lon_range_testing = list(range(3, features.dims['lon'], 4))
features_training = features.isel(lon=lon_range_training)
features_validation = features.isel(lon=lon_range_validation)
features_testing = features.isel(lon=lon_range_testing)
targets_training = targets.isel(lon=lon_range_training)
targets_validation = targets.isel(lon=lon_range_validation)
targets_testing = targets.isel(lon=lon_range_testing)
# + [markdown] id="cHEqkPkjCPDm" colab_type="text"
# ## Create the neural network
#
# + [markdown] id="nUHTM6yaDzZV" colab_type="text"
# Next, we'll instantiate and configure a neural network using TensorFlow's [DNNRegressor](https://www.tensorflow.org/api_docs/python/tf/estimator/DNNRegressor) class. We'll train this model using the GradientDescentOptimizer, which implements Mini-Batch Stochastic Gradient Descent (SGD). The learning_rate argument controls the size of the gradient step.
#
# NOTE: To be safe, we also apply gradient clipping to our optimizer via `clip_gradients_by_norm`. Gradient clipping ensures the magnitude of the gradients do not become too large during training, which can cause gradient descent to fail.
#
# We use `hidden_units`to define the structure of the NN. The `hidden_units` argument provides a list of ints, where each int corresponds to a hidden layer and indicates the number of nodes in it. For example, consider the following assignment:
#
# `hidden_units=[3, 10]`
#
# The preceding assignment specifies a neural net with two hidden layers:
#
# The first hidden layer contains 3 nodes.
# The second hidden layer contains 10 nodes.
# If we wanted to add more layers, we'd add more ints to the list. For example, `hidden_units=[10, 20, 30, 40]` would create four layers with ten, twenty, thirty, and forty units, respectively.
#
# By default, all hidden layers will use ReLu activation and will be fully connected.
# + id="Cmvlnh4uC9SS" colab_type="code" colab={}
# Use gradient descent as the optimizer for training the model.
gd_optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.0000001)
gd_optimizer = tf.contrib.estimator.clip_gradients_by_norm(gd_optimizer, 5.0)
# Use two hidden layers with 3 and 10 nodes each.
hidden_units=[3, 4]
# Instantiate the neural network.
dnn_regressor = tf.estimator.DNNRegressor(feature_columns=feature_columns,
hidden_units=hidden_units,
optimizer=gd_optimizer)
# + [markdown] id="L1CRW1a0Ds1C" colab_type="text"
# ## Define the input function
# + [markdown] id="rhyUxyMoF0wQ" colab_type="text"
# To import our weather data into our DNNRegressor, we need to define an input function, which instructs TensorFlow how to preprocess the data, as well as how to batch, shuffle, and repeat it during model training.
#
# First, we'll convert our xarray feature data into a dict of NumPy arrays. We can then use the TensorFlow Dataset API to construct a dataset object from our data, and then break our data into batches of `batch_size`, to be repeated for the specified number of epochs (`num_epochs`).
#
# NOTE: When the default value of `num_epochs=None` is passed to `repeat()`, the input data will be repeated indefinitely.
#
# Next, if `shuffle` is set to True, we'll shuffle the data so that it's passed to the model randomly during training. The `buffer_size` argument specifies the size of the dataset from which shuffle will randomly sample.
#
# Finally, our input function constructs an iterator for the dataset and returns the next batch of data.
# + id="-ZiWqTxvGNJO" colab_type="code" colab={}
from tensorflow.python.data import Dataset
def get_input(features,
targets,
batch_size=1,
shuffle=True,
num_epochs=None):
"""
Extracts a batch of elements from a dataset.
Args:
features: xarray Dataset of features
targets: xarray Dataset of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated.
None == repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert xarray data into a dict of numpy arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features, targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(buffer_size=10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# Create input functions. Wrap get_input() in a lambda so we
# can pass in features and targets as arguments.
input_training = lambda: get_input(features_training,
targets_training,
batch_size=10)
predict_input_training = lambda: get_input(features_training,
targets_training,
num_epochs=1,
shuffle=False)
predict_input_validation = lambda: get_input(features_validation,
targets_validation,
num_epochs=1,
shuffle=False)
# + [markdown] id="oqJj8vtMIbt5" colab_type="text"
# ## Train and evaluate the model
# + [markdown] id="jQTKlqSHIgVh" colab_type="text"
# We can now call `train()` on our `dnn_regressor` to train the model. We'll loop over a number of periods and on each loop we'll train the model, use it to make predictions, and compute the RMSE of the loss for both training and validation datasets.
# + id="iwfgGEY-I1pR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="0471a7f1-211b-4412-dd69-61bb8ca2386c"
print("Training model...")
print("RMSE (on training data):")
training_rmse = []
validation_rmse = []
steps = 500
periods = 20
steps_per_period = steps / periods
# Train the model inside a loop so that we can periodically assess loss metrics.
for period in range (0, periods):
# Train the model, starting from the prior state.
dnn_regressor.train(input_fn=input_training,
steps=steps_per_period)
# Take a break and compute predictions, converting to numpy arrays.
training_predictions = dnn_regressor.predict(input_fn=predict_input_training)
training_predictions = np.array([item['predictions'][0] for item in training_predictions])
validation_predictions = dnn_regressor.predict(input_fn=predict_input_validation)
validation_predictions = np.array([item['predictions'][0] for item in validation_predictions])
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, targets_training))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, targets_validation))
# Print the current loss.
print(" period %02d : %0.2f" % (period, training_root_mean_squared_error))
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print("Model training finished.")
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
print("Final RMSE (on training data): %0.2f" % training_root_mean_squared_error)
print("Final RMSE (on validation data): %0.2f" % validation_root_mean_squared_error)
# + id="wVzN6_fWZDJn" colab_type="code" colab={}
| notebooks/model_learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext nb_black
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import numpy as np
import sqlite3
import statsmodels.api as sm
from statsmodels.stats.outliers_influence import variance_inflation_factor
from statsmodels.tools.eval_measures import mse, rmse
# sklearn/XGboost imports
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.feature_selection import SelectKBest, f_regression
from sklearn.linear_model import LogisticRegression, LassoCV, RidgeCV, ElasticNetCV
from sklearn.neighbors import KNeighborsRegressor
from sklearn.metrics import (
confusion_matrix,
mean_absolute_error,
mean_squared_error,
classification_report,
)
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from xgboost import XGBRegressor
import warnings
warnings.filterwarnings("ignore")
# -
# cd
# cd Desktop
with sqlite3.connect("database.sqlite") as con:
matches = pd.read_sql_query("SELECT * from Match", con)
team_attributes = pd.read_sql_query("SELECT * from Team_Attributes", con)
"""
Datasets I care about:
Matches
team_attributes
I need to figure out how to combine players and the teams. If i had more time/better understanding, I would combine. For now,
I'll just look at teams
"""
# # Gonna check win loss split to make sure this data isn't super lopsided
won = matches["home_team_goal"] > matches["away_team_goal"]
loss = matches["home_team_goal"] < matches["away_team_goal"]
tie = matches["home_team_goal"] == matches["away_team_goal"]
tie
# 51.1%
won.value_counts()
# 69.5%
tie.value_counts()
# 71.2%
print(loss.value_counts())
# # Well it seems that the distributions aren't too bad. I can stick with the team attributes for now.
team_attributes.info()
for col in matches.columns:
print(col)
# # Going through and dropping columns with an absolute value coorelation of over 50%. Also using my judgement that they aren't as similar as almost all goalkeeper skills
matches_corr = matches.corr()
matches_corr = matches_corr.reset_index().melt("index")
matches_corr["ABS"] = matches_corr["value"].abs()
matches_corr = matches_corr.sort_values(by="ABS", ascending=False)
matches_corr = matches_corr[matches_corr["index"] != matches_corr["variable"]]
matches_corr.head(90)
# country and league ID aren't that useful so I'm gonna drop those. No need so the correlations aren't important
ting = matches.drop(
columns=[
"id",
"country_id",
"league_id",
"B365H",
"B365D",
"B365A",
"BWH",
"BWD",
"BWA",
"IWH",
"IWD",
"IWA",
"LBH",
"LBD",
"LBA",
"PSH",
"PSD",
"PSA",
"WHH",
"WHD",
"WHA",
"SJH",
"SJD",
"SJA",
"VCH",
"VCD",
"VCA",
"GBH",
"GBD",
"GBA",
"BSH",
"BSD",
"BSA",
"home_player_X1",
"home_player_X2",
"home_player_X3",
"home_player_X4",
"home_player_X5",
"home_player_X6",
"home_player_X7",
"home_player_X8",
"home_player_X9",
"home_player_X10",
"home_player_X11",
"away_player_X1",
"away_player_X2",
"away_player_X3",
"away_player_X4",
"away_player_X5",
"away_player_X6",
"away_player_X7",
"away_player_X8",
"away_player_X9",
"away_player_X10",
"away_player_X11",
"home_player_Y1",
"home_player_Y2",
"home_player_Y3",
"home_player_Y4",
"home_player_Y5",
"home_player_Y6",
"home_player_Y7",
"home_player_Y8",
"home_player_Y9",
"home_player_Y10",
"home_player_Y11",
"away_player_Y1",
"away_player_Y2",
"away_player_Y3",
"away_player_Y4",
"away_player_Y5",
"away_player_Y6",
"away_player_Y7",
"away_player_Y8",
"away_player_Y9",
"away_player_Y10",
"away_player_Y11",
"home_player_1",
"home_player_2",
"home_player_3",
"home_player_4",
"home_player_5",
"home_player_6",
"home_player_7",
"home_player_8",
"home_player_9",
"home_player_10",
"home_player_11",
"away_player_1",
"away_player_2",
"away_player_3",
"away_player_4",
"away_player_5",
"away_player_6",
"away_player_7",
"away_player_8",
"away_player_9",
"away_player_10",
"away_player_11",
]
)
ting = ting.corr().reset_index().melt("index")
ting["ABS"] = ting["value"].abs()
ting = ting.sort_values(by="ABS", ascending=False)
ting = ting[ting["index"] != ting["variable"]]
ting.head(30)
# # now that i've seen what matches look like, time to look at team attributes to see if there is anything I should drop before dropping.
fig, ax = plt.subplots(figsize=(30, 30))
sns.heatmap(team_attributes.corr(), vmin=-1, vmax=1, annot=True, ax=ax)
plt.show()
# An argument could be made for pressure and width being similar, but that isn't necessarily true so I'm not dropping it
TA_corr = team_attributes.corr()
TA_corr = TA_corr.reset_index().melt("index")
TA_corr["ABS"] = TA_corr["value"].abs()
TA_corr = TA_corr.sort_values(by="ABS", ascending=False)
TA_corr = TA_corr[TA_corr["index"] != TA_corr["variable"]]
TA_corr.head(20)
# # So after looking at the columns, I have deceided to drop the following columns due to clear leack of coorelation(player names/birthday/etc.), redundancy with other statistics (the bottom letters are betting site stats and I'm not presenting to Michael Jordan or Gretsky's wife so ignoring those as well).
drop_col=[
"id",
"country_id",
"league_id",
"B365H",
"B365D",
"B365A",
"BWH",
"BWD",
"BWA",
"IWH",
"IWD",
"IWA",
"LBH",
"LBD",
"LBA",
"PSH",
"PSD",
"PSA",
"WHH",
"WHD",
"WHA",
"SJH",
"SJD",
"SJA",
"VCH",
"VCD",
"VCA",
"GBH",
"GBD",
"GBA",
"BSH",
"BSD",
"BSA",
"home_player_X1",
"home_player_X2",
"home_player_X3",
"home_player_X4",
"home_player_X5",
"home_player_X6",
"home_player_X7",
"home_player_X8",
"home_player_X9",
"home_player_X10",
"home_player_X11",
"away_player_X1",
"away_player_X2",
"away_player_X3",
"away_player_X4",
"away_player_X5",
"away_player_X6",
"away_player_X7",
"away_player_X8",
"away_player_X9",
"away_player_X10",
"away_player_X11",
"home_player_Y1",
"home_player_Y2",
"home_player_Y3",
"home_player_Y4",
"home_player_Y5",
"home_player_Y6",
"home_player_Y7",
"home_player_Y8",
"home_player_Y9",
"home_player_Y10",
"home_player_Y11",
"away_player_Y1",
"away_player_Y2",
"away_player_Y3",
"away_player_Y4",
"away_player_Y5",
"away_player_Y6",
"away_player_Y7",
"away_player_Y8",
"away_player_Y9",
"away_player_Y10",
"away_player_Y11",
"home_player_1",
"home_player_2",
"home_player_3",
"home_player_4",
"home_player_5",
"home_player_6",
"home_player_7",
"home_player_8",
"home_player_9",
"home_player_10",
"home_player_11",
"away_player_1",
"away_player_2",
"away_player_3",
"away_player_4",
"away_player_5",
"away_player_6",
"away_player_7",
"away_player_8",
"away_player_9",
"away_player_10",
"away_player_11",
]
# # Now it's time to work on joining the stuff to one dataframe
mat_drop = matches.drop(columns=drop_col)
mat_drop
type(team_attributes["date"])
type(mat_drop["date"])
mat_drop["date"].unique()
team_attributes["date"].unique()
years = mat_drop["date"]
years[0][0:4]
mat_drop["date"] = pd.to_datetime(mat_drop["date"])
mat_drop["year"] = mat_drop["date"].dt.year
team_attributes["date"] = pd.to_datetime(team_attributes["date"])
team_attributes["year"] = team_attributes["date"].dt.year
# +
x = mat_drop.drop(
columns=[
"date",
"season",
"stage",
"goal",
"shoton",
"shotoff",
"foulcommit",
"card",
"cross",
"corner",
"possession",
]
)
x
# -
y = team_attributes.drop(columns=["id", "team_fifa_api_id", "date"])
y.sort_values(by="year", ascending=False)
home_teams = pd.DataFrame(
data=mat_drop,
columns=["match_api_id", "home_team_api_id", "home_team_goal", "year"],
)
home_teams = home_teams.rename(columns={"home_team_api_id": "team_api_id"})
home_teams
away_teams = pd.DataFrame(
data=mat_drop,
columns=["match_api_id", "away_team_api_id", "away_team_goal", "year"],
)
away_teams = away_teams.rename(columns={"away_team_api_id": "team_api_id"})
away_teams
away_teams = away_teams.merge(team_attributes, on=["team_api_id", "year"])
away_teams
mat_drop[mat_drop["match_api_id"] == 665626]
team_attributes[team_attributes["team_api_id"] == 8342]
# # Now that I'm sure my merge merged the data frames correctly, I'm going to do the same for the home team, check for nulls and clean both up
home_teams = home_teams.merge(team_attributes, on=["team_api_id", "year"])
home_teams
# dang, why so many NA values
away_teams.isna().mean()
away_teams["buildUpPlayDribbling"].value_counts()
away_teams["buildUpPlayDribblingClass"].value_counts()
# # 66% is a ton of missing data. Rather than trying to fill in the missing data, I'm going to drop it since it would be more fake than real. It might lead to discoveries but that is something that could be explored in another passthrough. Make sure hometeams only has that issue then deal with it the same way too
home_teams.isna().mean()
home_teams = home_teams.drop(columns="buildUpPlayDribbling")
away_teams = away_teams.drop(columns="buildUpPlayDribbling")
# Let's explore these class columns values to see what they're about
away_teams.info()
away_teams
home_teams
obj_cols = away_teams.select_dtypes("object").columns
obj_cols
# a lot of these look p different. I need dummies to help me compare
away_teams["defenceAggressionClass"].value_counts()
away_teams = pd.get_dummies(away_teams, columns=obj_cols, drop_first=True)
away_teams
home_teams = pd.get_dummies(home_teams, columns=obj_cols, drop_first=True)
home_teams
home_teams.info()
away_teams.info()
# # The difference of two games I don't think is super important in the big scheme of things
away_teams = away_teams.add_prefix("Away ")
home_teams = home_teams.add_prefix("Home ")
away_teams = away_teams.rename(
columns={"Away date": "date", "Away match_api_id": "match_api_id"}
)
home_teams = home_teams.rename(
columns={"Home date": "date", "Home match_api_id": "match_api_id"}
)
test = away_teams.merge(home_teams, on=["match_api_id"])
test
test[test["match_api_id"] == 665626]["Home team_api_id"]
matches[matches["match_api_id"] == 665626]
# # After confirming a few rows, I'm going to rename test, check, nulls, then make a goals column to start predicting goals. Negative goals means the away team has more. Positive means the home team has more
# AYYYYYYYY we have no NaNs! Let's start looking at reducing some features to make this even more selective
soccer = test.copy()
soccer.isna().mean().sort_values(ascending=False)
for col in soccer.columns:
print(col)
soccer["goals"] = soccer["Home home_team_goal"] - soccer["Away away_team_goal"]
soccer["goals"].value_counts()
# pretty normal. Outliers shouldn't mess me up too much
plt.hist(soccer["goals"])
plt.show()
for col in soccer:
print(col)
soccer = soccer.drop(
columns=[
"date_x",
"date_y",
"Home year",
"Away away_team_goal",
"Home home_team_goal",
]
)
soccer.info()
# # Alright. Check VIF for multicoliniarity, then gonna check features noise levels
# +
# Gonna keep this here in case I need to mess with stuff again
def print_vif(x):
"""Utility for checking multicollinearity assumption
:param x: input features to check using VIF. This is assumed to be a pandas.DataFrame
:return: nothing is returned the VIFs are printed as a pandas series
"""
# Silence numpy FutureWarning about .ptp
with warnings.catch_warnings():
warnings.simplefilter("ignore")
x = sm.add_constant(x)
vifs = []
for i in range(x.shape[1]):
vif = variance_inflation_factor(x.values, i)
vifs.append(vif)
print("VIF results\n-------------------------------")
print(pd.Series(vifs, index=x.columns))
print("-------------------------------\n")
# -
X = soccer.drop(columns="goals")
y = soccer["goals"]
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=69
)
# # Now I'll actually check OLS and VIF
# +
# dang, so overfit
X_train_const = sm.add_constant(X_train)
lm = sm.OLS(y_train, X_train_const).fit()
lm.summary()
# -
aways = soccer.columns.str.contains("Away")
true = [
False,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
True,
]
len(true)
home_soccer = soccer.drop(columns=soccer.columns[1:35])
# +
# Now we're gonna run models with both only home data and all data to see if the away team data does indeed lead to better predictions
HX = home_soccer.drop(columns="goals")
hy = home_soccer["goals"]
HX_train, HX_test, hy_train, hy_test = train_test_split(
HX, hy, test_size=0.2, random_state=69
)
# +
# I mean it isn't as overfit which is nice... Going to see what
HX_train_const = sm.add_constant(HX_train)
lm = sm.OLS(hy_train, HX_train_const).fit()
lm.summary()
# -
lr_bsl = LogisticRegression()
lr_fitted = lr_bsl.fit(HX_train, hy_train)
# +
# That is a pretty trash score ngl Let's try it with the regular ones
Htrain_score = lr_fitted.score(HX_train, hy_train)
Htest_score = lr_fitted.score(HX_test, hy_test)
print(f"train_score: {Htrain_score}")
print(f"test_score: {Htest_score}")
# +
r_fitted = lr_bsl.fit(X_train, y_train)
train_score = lr_fitted.score(X_train, y_train)
test_score = lr_fitted.score(X_test, y_test)
print(f"train_score: {train_score}")
print(f"test_score: {test_score}")
print(0.26253768155659085 - 0.2622636338722938)
# +
y_preds = r_fitted.predict(X_test)
plt.scatter(y_test, y_preds)
plt.plot(y_test, y_test, color="red")
plt.xlabel("true values")
plt.ylabel("predicted values")
plt.title("Charges: true and predicted values")
plt.show()
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds)))
print("Root mean squared error of the prediction is: {}".format(rmse(y_test, y_preds)))
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds) / y_test)) * 100
)
)
# -
# # Well it seems like logistic regression is a pretty trash model.
# # I'm gonna check a few more models
# +
# Check regularization models
# +
# lasso seems pretty overfit imo
alphas = [np.power(10.0, p) for p in np.arange(-10, 40, 1)]
lasso_cv = LassoCV(alphas=alphas, cv=5)
lasso_cv.fit(HX_train, hy_train)
# We are making predictions here
y_preds_train = lasso_cv.predict(HX_train)
y_preds_test = lasso_cv.predict(HX_test)
print("Best alpha value is: {}".format(lasso_cv.alpha_))
print(
"R-squared of the model in training set is: {}".format(
lasso_cv.score(HX_train, hy_train)
)
)
print("-----Test set statistics-----")
print(
"R-squared of the model in test set is: {}".format(lasso_cv.score(HX_test, hy_test))
)
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(hy_test, y_preds_test)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds_test)))
print(
"Root mean squared error of the prediction is: {}".format(
rmse(hy_test, y_preds_test)
)
)
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((hy_test - y_preds_test) / y_test)) * 100
)
)
# +
ridge_cv = RidgeCV(alphas=alphas, cv=5)
ridge_cv.fit(X_train, y_train)
# We are making predictions here
y_preds_train = ridge_cv.predict(X_train)
y_preds_test = ridge_cv.predict(X_test)
print("Best alpha value is: {}".format(ridge_cv.alpha_))
print(
"R-squared of the model in training set is: {}".format(
ridge_cv.score(X_train, y_train)
)
)
print("-----Test set statistics-----")
print(
"R-squared of the model in test set is: {}".format(ridge_cv.score(X_test, y_test))
)
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds_test)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds_test)))
print(
"Root mean squared error of the prediction is: {}".format(
rmse(y_test, y_preds_test)
)
)
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds_test) / y_test)) * 100
)
)
# +
elasticnet_cv = ElasticNetCV(alphas=alphas, cv=5)
elasticnet_cv.fit(X_train, y_train)
# We are making predictions here
y_preds_train = elasticnet_cv.predict(X_train)
y_preds_test = elasticnet_cv.predict(X_test)
print("Best alpha value is: {}".format(elasticnet_cv.alpha_))
print(
"R-squared of the model in training set is: {}".format(
elasticnet_cv.score(X_train, y_train)
)
)
print("-----Test set statistics-----")
print(
"R-squared of the model in test set is: {}".format(
elasticnet_cv.score(X_test, y_test)
)
)
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds_test)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds_test)))
print(
"Root mean squared error of the prediction is: {}".format(
rmse(y_test, y_preds_test)
)
)
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds_test) / y_test)) * 100
)
)
# +
y_preds = lasso_cv.predict(HX_test)
plt.scatter(hy_test, y_preds)
plt.plot(hy_test, hy_test, color="red")
plt.xlabel("true values")
plt.ylabel("predicted values")
plt.title("Charges: true and predicted values")
plt.show()
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds)))
print("Root mean squared error of the prediction is: {}".format(rmse(y_test, y_preds)))
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds) / y_test)) * 100
)
)
# -
# # These are all pretty bad. Let's go on with Gradient boosting to see if that's better?
XGB = XGBRegressor()
# +
n_trees = 1000
learning_rate = 2 / n_trees
grid = {
"gbt__subsample": [0.75, 1.0],
"gbt__max_features": [0.75, 1.0],
"gbt__max_depth": [2, 3],
"gbt__n_estimators": [n_trees],
"gbt__learning_rate": [learning_rate, 0.001, 0.2],
}
XGB_cv = GridSearchCV(XGB, grid, verbose=1, cv=2)
XGB_cv.fit(X_train, y_train)
print(XGB_cv.best_params_)
# +
train_score = XGB_cv.score(X_train, y_train)
test_score = XGB_cv.score(X_test, y_test)
print(f"train_score: {train_score}")
print(f"test_score: {test_score}")
# +
y_preds = XGB_cv.predict(X_test)
plt.scatter(y_test, y_preds)
plt.plot(y_test, y_test, color="red")
plt.xlabel("true values")
plt.ylabel("predicted values")
plt.title("Charges: true and predicted values")
plt.show()
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds)))
print("Root mean squared error of the prediction is: {}".format(rmse(y_test, y_preds)))
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds) / y_test)) * 100
)
)
# +
y_pred = XGB_cv.predict(X_test)
y_pred = y_pred.round()
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
# # Well the gradient boosted regressor seems pretty good. Want to see if I can improve the model
select = SelectKBest(k=5).fit(X_train, y_train)
features = pd.DataFrame({"Feature": list(X_train.columns), "Scores": select.scores_})
features.sort_values(by="Scores", ascending=False).head(30)
to_drop = features.sort_values(by="Scores", ascending=False).tail(15)
to_drop.reset_index(drop=True)
to_be = to_drop["Feature"].tolist()
to_be
new_soccer = soccer.drop(columns=to_be)
new_soccer
# +
n_trees = 1000
learning_rate = 2 / n_trees
grid = {
"gbt__subsample": [0.75, 1.0],
"gbt__max_features": [0.75, 1.0],
"gbt__max_depth": [2, 3],
"gbt__n_estimators": [n_trees],
"gbt__learning_rate": [learning_rate, 0.001, 0.2],
}
XGB_cv = GridSearchCV(XGB, grid, verbose=1, cv=2)
XGB_cv.fit(X_train, y_train)
print(XGB_cv.best_params_)
# +
train_score = XGB_cv.score(X_train, y_train)
test_score = XGB_cv.score(X_test, y_test)
print(f"train_score: {train_score}")
print(f"test_score: {test_score}")
# -
# # ok that's absolute trash. No dropping those columns but both have pretty bad overfitting. I'm gonna try logging
X = new_soccer.drop(columns="goals")
y = new_soccer["goals"]
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=69
)
preprocessing = ColumnTransformer(
[("scale_nums", StandardScaler(), X.columns),], remainder="passthrough"
)
pipeline = Pipeline([("preprocessing", preprocessing), ("gbt", XGBRegressor()),])
# +
n_trees = 1000
learning_rate = 2 / n_trees
grid = {
"gbt__subsample": [0.75, 1.0],
"gbt__max_features": [0.75, 1.0],
"gbt__max_depth": [2, 3],
"gbt__n_estimators": [n_trees],
"gbt__learning_rate": [learning_rate, 0.001, 0.2],
}
pipeline_cv = GridSearchCV(pipeline, grid, verbose=1, cv=2)
pipeline_cv.fit(X_train, y_train)
print(pipeline_cv.best_params_)
# +
train_score = pipeline_cv.score(X_train, y_train)
test_score = pipeline_cv.score(X_test, y_test)
print(f"train_score: {train_score}")
print(f"test_score: {test_score}")
# +
importance_df = pd.DataFrame()
importance_df["feat"] = X_train.columns
importance_df["importance"] = pipeline_cv.best_estimator_["gbt"].feature_importances_
importance_df = importance_df.sort_values("importance", ascending=False)
importance_df.head()
# +
y_preds = pipeline_cv.predict(X_test)
plt.scatter(y_test, y_preds)
plt.plot(y_test, y_test, color="red")
plt.xlabel("true values")
plt.ylabel("predicted values")
plt.title("Charges: true and predicted values")
plt.show()
print(
"Mean absolute error of the prediction is: {}".format(
mean_absolute_error(y_test, y_preds)
)
)
print("Mean squared error of the prediction is: {}".format(mse(y_test, y_preds)))
print("Root mean squared error of the prediction is: {}".format(rmse(y_test, y_preds)))
print(
"Mean absolute percentage error of the prediction is: {}".format(
np.mean(np.abs((y_test - y_preds) / y_test)) * 100
)
)
# +
y_pred = pipeline_cv.predict(X_test)
y_pred = y_pred.round()
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
# # Last model I'm going to be using is Knearestneighbors. If this model is trash then I'm just going to end it there
# +
# Preprocessing aka column transforming
preprocessing = ColumnTransformer(
[("scale_nums", StandardScaler(), X.columns),], remainder="passthrough"
)
# Modeling pipeline (which will include the preprocessing pipeline)
pipeline = Pipeline(
[
("preprocess", preprocessing),
("feat_select", SelectKBest(f_regression)),
("regress", KNeighborsRegressor()),
]
)
grid = {
"feat_select__k": range(49, 50),
"regress__n_neighbors": range(49, 50),
"regress__weights": ["uniform", "distance"],
}
pipeline_cv = GridSearchCV(pipeline, grid, verbose=1)
pipeline_cv.fit(X_train, y_train)
pipeline_cv.best_params_
# +
train_score = pipeline_cv.score(X_train, y_train)
test_score = pipeline_cv.score(X_test, y_test)
print(f"train_score: {train_score}")
print(f"test_score: {test_score}")
# -
# # I have run every model I can think of that would be applicable, but the test and train accuracy are absolutely terrible. I don't think this data is good at predicting goals given the spread of points
y.value_counts()
print(
4628
/ (
3862
+ 2862
+ 2557
+ 1483
+ 1182
+ 590
+ 532
+ 214
+ 176
+ 62
+ 17
+ 16
+ 7
+ 5
+ 2
+ 1
+ 1
)
)
# +
X_train_const = sm.add_constant(X_train)
lm = sm.OLS(y_train, X_train_const).fit()
lm.summary()
# -
| .ipynb_checkpoints/Assisted AI challenge-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # USA Resiliece Opportunity Mapping Data Processing
#
# The data that we have selected for the platform can be seen here:
# https://docs.google.com/spreadsheets/d/1xEHzYpkWgZTu_DNXw3YDk3LNI7iVEDfP/edit#gid=1856162673&fvid=1697691070
#
# ## Table of Contents
# - ## [Python libraries](#libraries)
# - ## [Utils](#utils)
# - ## [List of datasets and indicators](#datasets)
# - ## [Read vector data](#vector_data)
# - **[Sates](#states)**
# - **[County](#county)**
# - **[Census tract](#tract)**
# - ## [Preprocess carto data](#carto_data)
# <a id='libraries'></a>
# ## Python libraries
from matplotlib import pyplot as plt
import LMIPy as lmi
import geopandas as gpd
import shapely.wkt
import pandas as pd
import requests
import zipfile
import json
import io
import os
# <a id='utils'></a>
# ## Utils
#import functions form notebook
# %run './preprocessing_main_functions.ipynb'
# <a id='datasets'></a>
# ## List of datasets and indicators
# **Datasets**
# +
datasets = {
'gee':{
'extreme_heat_days':'4077d297-c3d4-4302-8e2b-2a1701a24d8d',
'extreme_precipitation_days':'6a818d38-2361-4c9a-bd9e-c0cd3493d5ce',
'landslide_susceptibility':'ea2db3a6-49c8-4d41-a2ab-758eb6fe4bc0',
'earthquake_frequency_and_distribution':'b3ebc10d-9de8-4ee6-870d-1d049e8e2a99',
'volcano_frequency_and_distribution':'74ebf4fe-6afb-46aa-a820-96fcae9a7e03',
},
'carto':{
'riverine_flood_risk':'5152c286-53c1-4583-9519-816a6e41889d',
'costal_flood_risk':'5152c286-53c1-4583-9519-816a6e41889d',
'drought_risk':'5152c286-53c1-4583-9519-816a6e41889d',
},
'others':{
'Wildfires': 'https://www.nature.com/articles/s41597-019-0312-2',
'CDC Social Vulnerability Index':'https://svi.cdc.gov/Documents/Data/2018_SVI_Data/SVI2018Documentation-508.pdf',
}
}
datasets = pd.DataFrame(datasets)
datasets
# -
# **Indicators**
# +
indicators = {'Name':['riverine flood risk', 'coastal flood risk', 'extreme heat days', 'extreme precipitation days',
'drought risk', 'landslide susceptibility', 'earthquake frequency and distribution',
'volcano frequency and distribution', 'wildfires'] +
['socioeconomic status', 'percentage of persons below poverty', 'percentage of civilian (age 16+) unemployed', 'per capita income', 'percentage of persons with no high school diploma (age 25+)',
'household composition & disability','percentage of persons aged 65 and older', 'percentage of persons aged 17 and younger', 'percentage of civilian with a disability', 'percentage of single parent households with children under 18',
'minority status & language', 'minority (all persons except white, non-Hispanic)', 'speaks english “less than well”',
'housing type & transportation', 'multi-unit structures', 'mobile homes', 'crowding', 'no vehicle', 'group quarters'],
'Slug':['rfr', 'cfr', 'ehd', 'epd','drr', 'lss', 'efd','vfd', 'wlf'] +
['ses', 'pov', 'uep', 'pci', 'hsd', 'hcd', 'a6o', 'a1y', 'pcd', 'sph', 'msl', 'mnt', 'sen', 'htt', 'mus', 'mhm', 'cwd', 'vhc', 'gqt'],
'Category': ['climate risk'] * 9 + ['vulnerability'] * 19,
'Group': list(range(0, 9)) + [9, 9.1,9.2,9.3,9.4] + [10, 10.1,10.2,10.3,10.4] + [11, 11.1,11.2] + [12, 12.1,12.2,12.3,12.4,12.5]}
indicators = pd.DataFrame(indicators)
#indicators.to_csv('../data/indicators_list.csv')
indicators
# +
indicators = {'group': ['climate risk'] * 9 + ['vulnerability'] * 5,
'indicator':['riverine flood risk', 'coastal flood risk', 'extreme heat days', 'extreme precipitation days',
'drought risk', 'landslide susceptibility', 'earthquake frequency and distribution',
'volcano frequency and distribution', 'wildfires'] +
['population', 'community resilience estimate', 'unemployment', 'median household income', 'poverty'],
'slug':['rfr', 'cfr', 'ehd', 'epd', 'drr', 'lss', 'efd','vfd', 'wlf'] + ['pop', 'cre', 'uep', 'mhi', 'pvt'],
'state_labels':
[['Data not available', 'Less than 1 in 1k', '1 in 1k to 2 in 1k', '2 in 1k to 6 in 1k', '6 in 1k to 1 in 100', 'More than 1 in 100'],
['Data not available', 'Less than 9 in 1M', '9 in 1M to 7 in 100k', '7 in 100k to 3 in 10k', '3 in 10k to 2 in 1k', 'More than 2 in 1k'],
'',
'',
['Data not available', '0.0 to 0.2', '0.2 to 0.4', '0.4 to 0.6', '0.6 to 0.8', '0.8 to 1.0'],
['Slight', 'Low', 'Moderate', 'High', 'Severe'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['Less than 30 kha', '30 kha to 280 kha', '280 kha to 1 Mha', '1 Mha to 3 Mha', 'More than 3 Mha'],
['Less than 1.5M', '1.5M to 3M', '3M to 6M', '6M to 9M', 'More than 9M'],
['Less than 22.6%', '22.6% to 24.2%', '24.2% to 25.7%', '25.7% to 27.5%', '27.5% or higher'],
['Less than 2.9%', '2.9% to 3.3%', '3.3% to 3.6%', '3.6% to 4.1%', '4.1% or higher'],
['Less than $54k', '$54k to $58k', '$58k to $62k', '$62k to $73k', '$73k or higher'],
['Less than 10.6%', '10.6% to 11.9%', '11.9% to 13.1%', '13.1% to 15.2%', '15.2% or higher']],
'county_labels':
[['Data not available', 'Less than 1 in 1k', '1 in 1k to 2 in 1k', '2 in 1k to 6 in 1k', '6 in 1k to 1 in 100', 'More than 1 in 100'],
['Data not available', 'Less than 9 in 1M', '9 in 1M to 7 in 100k', '7 in 100k to 3 in 10k', '3 in 10k to 2 in 1k', 'More than 2 in 1k'],
'',
'',
['Data not available', '0.0 to 0.2', '0.2 to 0.4', '0.4 to 0.6', '0.6 to 0.8', '0.8 to 1.0'],
['Slight', 'Low', 'Moderate', 'High', 'Severe'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['Less than 1 kha', '1 kha to 4 kha', '4 kha to 10 kha', '10 kha to 35 kha', 'More than 35 kha'],
['Less than 9k', '9k to 20k', '20k to 40k', '40k to 95k', 'More than 95k'],
['Less than 21.5%', '21.5% to 24.3%', '24.3% to 26.9%', '26.9% to 30.0%', '30.0% or higher'],
['Data not available', 'Less than 2.9%', '2.9% to 3.4%', '3.4% to 4.0%', '4.0% to 4.9%', '4.9% or higher'],
['Data not available', 'Less than $42k', '$42k to $48k', '$48k to $53k', '$53k to $61k', '$61k or higher'],
['Data not available', 'Less than 10.2%', '10.2% to 12.8%', '12.8% to 15.5%', '15.5% to 19.6%', '19.6% or higher']],
'title':
['Riverine flood risk',
'Coastal flood risk',
'Extreme heat days',
'Extreme precipitation days',
'Drought risk',
'Landslide susceptibility',
'Earthquake hazards - frequency (deciles)',
'Volcano hazards - frequency (deciles)',
'Total hectares burned (2001-2017)',
'Population',
'Percentage of residents with 3+ risk factors',
'Percentage of people in unemployment',
'Median household income',
'Percentage of people in poverty'],
'description':
['Riverine flood risk measures the percentage of population expected to be affected by Riverine flooding in an average year, accounting for existing flood-protection standards. Flood risk is assessed using hazard (inundation caused by river overflow), exposure (population in flood zone), and vulnerability. The existing level of flood protection is also incorporated into the risk calculation. It is important to note that this indicator represents flood risk not in terms of maximum possible impact but rather as average annual impact. The impacts from infrequent, extreme flood years are averaged with more common, less newsworthy flood years to produce the “expected annual affected population.” Higher values indicate that a greater proportion of the population is expected to be impacted by Riverine floods on average.',
'Coastal flood risk measures the percentage of the population expected to be affected by coastal flooding in an average year, accounting for existing flood protection standards. Flood risk is assessed using hazard (inundation caused by storm surge), exposure (population in flood zone), and vulnerability. The existing level of flood protection is also incorporated into the risk calculation. It is important to note that this indicator represents flood risk not in terms of maximum possible impact but rather as average annual impact. The impacts from infrequent, extreme flood years are averaged with more common, less newsworthy flood years to produce the “expected annual affected population.” Higher values indicate that a greater proportion of the population is expected to be impacted by coastal floods on average.',
'Extreme heat days represent the annual average count of days with heat greater than the 99th percentile of the baseline, as derived from the Localized Constructed Analogs (LOCA). LOCA is a technique for downscaling climate model projections of the future climate. The method estimates finer-scale climate detail using the systematic historical effects of topography on local weather patterns. The LOCA downscaled climate projections provide temperature and precipitation on pixels that are six kilometers (3.7 miles) across. The data are daily, covering the period 1950-2100 for 32 global climate models. LOCA attempts to better preserve extreme hot days and heavy rain events than the previous generation of downscaling approaches. While previous downscaling techniques typically formed the downscaled model day using a weighted average of 30 similar historical days, LOCA looks locally around each point of interest to find the one best matching day. The data spans from Mexico through southern Canada and can be used to assess climate impacts across much of North America, including the entire conterminous U.S. The work represents a collaboration across many groups, including SIO, the California Energy Commission, the U.S. Army Corps of Engineers, the U.S. Geological Survey, the Bureau of Reclamation, the NOAA Regional Integrated Sciences and Assessments (RISA) program, the Climate Analytics Group, the nonprofit group Climate Central, Lawrence Livermore National Laboratory, NASA Ames Research Center, Santa Clara University, the Cooperative Institute for Research in Environmental Sciences at the University of Colorado Boulder, and the Southwest Climate Science Center.',
'Extreme precipitation days represent the annual average count of days with precipitation greater than the 99th percentile of the baseline, as derived from the Localized Constructed Analogs (LOCA). LOCA is a technique for downscaling climate model projections of the future climate. The method estimates finer-scale climate detail using the systematic historical effects of topography on local weather patterns. The LOCA downscaled climate projections provide temperature and precipitation on pixels that are six kilometers (3.7 miles) across. The data are daily, covering the period 1950-2100 for 32 global climate models. LOCA attempts to better preserve extreme hot days and heavy rain events than the previous generation of downscaling approaches. While previous downscaling techniques typically formed the downscaled model day using a weighted average of 30 similar historical days, LOCA looks locally around each point of interest to find the one best matching day. The data spans from Mexico through southern Canada and can be used to assess climate impacts across much of North America, including the entire conterminous U.S. The work represents a collaboration across many groups, including SIO, the California Energy Commission, the U.S. Army Corps of Engineers, the U.S. Geological Survey, the Bureau of Reclamation, the NOAA Regional Integrated Sciences and Assessments (RISA) program, the Climate Analytics Group, the nonprofit group Climate Central, Lawrence Livermore National Laboratory, NASA Ames Research Center, Santa Clara University, the Cooperative Institute for Research in Environmental Sciences at the University of Colorado Boulder, and the Southwest Climate Science Center.',
'Drought risk measures where droughts are likely to occur, the population and assets exposed, and the vulnerability of the population and assets to adverse effects. Higher values indicate higher risk of drought.',
'The Landslide Susceptibility Map, created by scientists at the National Aeronautics and Space Administration (NASA) and published in 2017 at 1 km resolution, improves upon past landslide susceptibility maps by incorporating the most up-to-date data. NASA scientists evaluated landslide susceptibility on slope from the Shuttle Radar Topography Mission and forest loss from a Landsat-based record of forest change compiled by a University of Maryland team. They also included data on key factors including the presence of roads from OpenStreetMap, the strength of bedrock and soils, and the locations of faults from the Geological Map of the World, 3rd ed., by <NAME> (2009). They generated the map using a heuristic fuzzy approach that generated the possibility of landslides and validated it through landslide inventories.',
'The Earthquake Hazard Frequency and Distribution data set is created by the World Bank Group, the Columbia University Center for Hazards and Risk Research (Columbia CHRR), and the Columbia University Earth Institute Center for International Earth Science Information Network (CIESIN). The data are released in a 2.5 minute grid utilizing Advanced National Seismic System (ANSS) Earthquake Catalog data of actual earthquake events exceeding 4.5 on the Richter scale during the time period 1976 through 2002.',
'The Volcano Hazard Frequency and Distribution data are created by the World Bank Group, the Columbia University Center for Hazards and Risk Research (Columbia CHRR), and the Columbia University Earth Institute Center for International Earth Science Information Network (CIESIN). The data are released in a 2.5 minute gridded data set based on the National Geophysical Data Center (NGDC) Volcano Database spanning the period 1979 through 2000. This database includes nearly 4,000 volcanic events categorized as moderate or above (values 2 through 8) according to the Volcano Explosivity Index (VEI). Most volcanoes are georeferenced to the nearest 10th or 100th of a degree, with a few to the nearest 1,000th of a degree.',
'Total hectares burned (2001-2017) has been obtained from the Global Wildfire Database for GWIS, An individual fire event focused database. Post processing of MCD64A1 providing geometries of final fire perimeters including initial and final date and the corresponding daily active areas for each fire. This dataset uses MCD64A1 Collection 6 MODIS Burned Area Product v6 and has a coverage from 2000 to 2018.',
'The Population data set represents the 7/1/2019 residents total population estimates for states and counties. This population estimate has been obtained from the CO-EST2019-all data which shows the annual resident population estimates, estimated components of resident population change and rates of the components of resident population change for states and counties. The CO-EST2019 data set has been created by the U.S. Census Bureau, Population Division and was released in March 2020.',
'Percentage of residents with 3+ risk factors identifies populations at states and county level with high risk of recovering from a disaster. This information is obtained from the Community Resilience data set that measures the capacity of individuals and households within a community to absorb, endure and recover from the impact of a disaster. The Community Resilience Estimates are experimental estimates produced using information on individuals and households from the 2018 American Community Survey (ACS) and the Census Bureau’s Population Estimates Program as well as publicly available health condition rates from the National Health Interview Survey (NHIS). The following risk factors are used: age 65 and above; low-income household; single caregiver householder; household communication barrier; employment status; disability status; physical crowding; lack of health insurance; respiratory disease; heart disease; and diabetes.',
'Percentage of people in unemployment shows the 2019 percentage of people in unemployenment for states and counties. The data set has been obtained from the U.S. Census Bureaus Small Area Income and Poverty Estimate (SAIPE) program. The SAIPE program produces single-year model-based estimates of income and poverty for all U.S. states and counties as well as estimates of school-age children in poverty for all 13,000+ school districts.',
'Median household income shows the 2019 estimates of Median household income for states and counties. The data set has been obtained from the U.S. Census Bureaus Small Area Income and Poverty Estimate (SAIPE) program. The SAIPE program produces single-year model-based estimates of income and poverty for all U.S. states and counties as well as estimates of school-age children in poverty for all 13,000+ school districts.',
'Percentage of people in poverty data set shows the 2018 estimated percentage of people of all ages in poverty for states and counties. This estimates have been obtained from the U.S. Census Bureaus Small Area Income and Poverty Estimate (SAIPE) program. The SAIPE program produces single-year model-based estimates of income and poverty for all U.S. states and counties as well as estimates of school-age children in poverty for all 13,000+ school districts.'],
'source':
['https://www.wri.org/publication/aqueduct-30',
'https://www.wri.org/publication/aqueduct-30',
'http://loca.ucsd.edu/',
'http://loca.ucsd.edu/',
'https://www.wri.org/publication/aqueduct-30',
'https://pmm.nasa.gov/applications/global-landslide-model',
'http://sedac.ciesin.columbia.edu/data/set/ndh-earthquake-frequency-distribution',
'http://sedac.ciesin.columbia.edu/data/set/ndh-volcano-hazard-frequency-distribution',
'https://doi.pangaea.de/10.1594/PANGAEA.895835',
'Population',
'Percentage of residents with 3+ risk factors',
'Percentage of people in unemployment',
'Median household income',
'Percentage of people in poverty']}
indicators = pd.DataFrame(indicators)
indicators
# +
indicators = {'group': ['climate risk'] * 9 + ['vulnerability'] * 19,
'indicator':['riverine flood risk', 'coastal flood risk', 'extreme heat days', 'extreme precipitation days',
'drought risk', 'landslide susceptibility', 'earthquake frequency and distribution',
'volcano frequency and distribution', 'wildfires'] +
['socioeconomic status', 'below poverty', 'unemployed', 'income', 'no high school diploma',
'household composition & disability','aged 65 or older', 'aged 17 or younger', 'civilian with a Disability', 'single-parent households',
'minority status & language', 'minority', 'speaks english “less than well”',
'housing type & transportation', 'multi-unit structures', 'mobile homes', 'crowding', 'no vehicle', 'group quarters'],
'slug':['rfr', 'cfr', 'ehd', 'epd','drr', 'lss', 'efd','vfd', 'wlf'] +
['ses', 'pov', 'uep', 'pci', 'hsd', 'hcd', 'a6o', 'a1y', 'pcd', 'sph', 'msl', 'mnt', 'sen', 'htt', 'mus', 'mhm', 'cwd', 'vhc', 'gqt'],
'state_labels':
[['Data not available', 'Less than 1 in 1k', '1 in 1k to 2 in 1k', '2 in 1k to 6 in 1k', '6 in 1k to 1 in 100', 'More than 1 in 100'],
['Data not available', 'Less than 9 in 1M', '9 in 1M to 7 in 100k', '7 in 100k to 3 in 10k', '3 in 10k to 2 in 1k', 'More than 2 in 1k'],
['Data not available', 'Less than 11 days', '11 to 13 days', '13 to 15 days', '15 to 17 days', 'More than 17 days'],
['Data not available', 'Less than 1.07 days', '1.07 to 1.11 days', '1.11 to 1.16 days', '1.16 to 1.19 days', 'More than 1.19 days'],
['Data not available', '0.0 to 0.2', '0.2 to 0.4', '0.4 to 0.6', '0.6 to 0.8', '0.8 to 1.0'],
['Slight', 'Low', 'Moderate', 'High', 'Severe'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['Less than 30 kha', '30 kha to 280 kha', '280 kha to 1 Mha', '1 Mha to 3 Mha', 'More than 3 Mha'],
['0.6 to 1.3th', '1.3 to 2.0th', '2.0 to 2.3th', '2.3 to 2.4th', '2.4 to 3.2th'],
['Less than 10.6%', '10.6% to 12.1%', '12.1% to 13.8%', '13.8% to 15.6%', '15.6% or higher'],
['Less than 2.2%', '2.2% to 2.7%', '2.7% to 3.0%', '3.0% to 3.2%', '3.2% or higher'],
['Less than $24k', '$24k to $27k', '$27k to $29k', '$29k to $33k', '$33k or higher'],
['Less than 5.5%', '5.5% to 6.6%', '6.6% to 7.5%', '7.5% to 9.2%', '9.2% or higher'],
['0.9 to 1.3th', '1.3 to 1.9th', '1.9 to 2.2th', '2.2 to 2.6th', '2.6 to 3.2th'],
['Less than 14.6%', '14.6% to 15.0%', '15.0% to 16.1%', '16.1% to 16.7%', '16.7% or higher'],
['Less than 21.1%', '21.1% to 22.4%', '22.4% to 23.1%', '23.1% to 23.8%', '23.8% or higher'],
['Less than 11.3%', '11.3% to 12.1%', '12.1% to 13.3%', '13.3% to 14.4%', '14.4% or higher'],
['Less than 3.0%', '3.0% to 3.2%', '3.2% to 3.4%', '3.4% to 3.6%', '3.6% or higher'],
['0.0 to 0.5th', '0.5 to 0.9th', '0.9 to 1.1th', '1.1 to 1.5th', '1.5 to 1.9th'],
['Less than 17.8%', '17.8% to 24.0%', '24.0% to 34.3%', '34.3% to 44.1%', '44.1% or higher'],
['Less than 1.0%', '1.0% to 1.5%', '1.5% to 2.3%', '2.3% to 4.1%', '4.1% or higher'],
['1.4 to 2.1th', '2.1 to 2.4th', '2.4 to 2.6th', '2.6 to 2.8th', '2.8 to 3.8th'],
['Less than 3.2%', '3.2% to 3.9%', '3.9% to 5.0%', '5.0% to 6.3%', '6.3% or higher'],
['Less than 1.3%', '1.3% to 2.1%', '2.1% to 3.6%', '3.6% to 5.3%', '5.3% or higher'],
['Less than 0.7%', '0.7% to 0.8%', '0.8% to 0.9%', '0.9% to 1.3%', '1.3% or higher'],
['Less than 2.1%', '2.1% to 2.4%', '2.4% to 2.7%', '2.7% to 3.3%', '3.3% or higher'],
['Less than 2.2%', '2.2% to 2.5%', '2.5% to 2.7%', '2.7% to 3.2%', '3.2% or higher']],
'county_labels':
[['Data not available', 'Less than 1 in 1k', '1 in 1k to 2 in 1k', '2 in 1k to 6 in 1k', '6 in 1k to 1 in 100', 'More than 1 in 100'],
['Data not available', 'Less than 9 in 1M', '9 in 1M to 7 in 100k', '7 in 100k to 3 in 10k', '3 in 10k to 2 in 1k', 'More than 2 in 1k'],
['Data not available', 'Less than 12 days', '12 to 14 days', '14 to 15 days', '15 to 18 days', 'More than 18 days'],
['Data not available', 'Less than 1.06 days', '1.06 to 1.10 days', '1.10 to 1.14 days', '1.14 to 1.18 days', 'More than 1.18 days'],
['Data not available', '0.0 to 0.2', '0.2 to 0.4', '0.4 to 0.6', '0.6 to 0.8', '0.8 to 1.0'],
['Slight', 'Low', 'Moderate', 'High', 'Severe'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['Less than 1 kha', '1 kha to 4 kha', '4 kha to 10 kha', '10 kha to 35 kha', 'More than 35 kha'],
['Data not available', '0.1 to 1.0th', '1.0 to 1.6th', '1.6 to 2.3th', '2.3 to 3.0th', '3.0 to 4.0th'],
['Data not available', 'Less than 10.1%', '10.1% to 13.3%', '13.3% to 16.2%', '16.2% to 20.2%', '20.2% or higher'],
['Data not available', 'Less than 3.6%', '3.6% to 4.9%', '4.9% to 6.0%', '6.0% to 7.6%', '7.6% or higher'],
['Data not available', 'Less than $22k', '$22k to $25k', '$25k to $28k', '$28k to $31k', '$31k or higher'],
['Less than 8.0%', '8.0% to 10.7%', '10.7% to 13.9%', '13.9% to 18.7%', '18.7% or higher'],
['0.1 to 1.6th', '1.6 to 1.9th', '1.9 to 2.1th', '2.1 to 2.4th', '2.4 to 3.6th'],
['Less than 14.8%', '14.8% to 17.1%', '17.1% to 19.1%', '19.1% to 21.5%', '21.5% or higher'],
['Less than 19.8%', '19.8% to 21.7%', '21.7% to 23.0%', '23.0% to 24.6%', '24.6% or higher'],
['Less than 12.2%', '12.2% to 14.4%', '14.4% to 16.6%', '16.6% to 19.4%', '19.4% or higher'],
['Less than 6.2%', '6.2% to 7.6%', '7.6% to 8.7%', '8.7% to 10.3%', '10.3% or higher'],
['0.0 to 0.4th', '0.4 to 0.8th', '0.8 to 1.1th', '1.1 to 1.5th', '1.5 to 2.0th'],
['Less than 6.2%', '6.2% to 11.9%', '11.9% to 22.6%', '22.6% to 39.8%', '39.8% or higher'],
['Less than 0.3%', '0.3% to 0.6%', '0.6% to 1.1%', '1.1% to 2.5%', '2.5% or higher'],
['0.2 to 1.8th', '1.8 to 2.3th', '2.3 to 2.7th', '2.7 to 3.1th', '3.1 to 4.4th'],
['Less than 1.2%', '1.2% to 2.2%', '2.2% to 3.8%', '3.8% to 7.3%', '7.3% or higher'],
['Less than 4.4%', '4.4% to 8.5%', '8.5% to 13.4%', '13.4% to 20.8%', '20.8% or higher'],
['Less than 1.1%', '1.1% to 1.7%', '1.7% to 2.3%', '2.3% to 3.2%', '3.2% or higher'],
['Less than 3.9%', '3.9% to 5.1%', '5.1% to 6.3%', '6.3% to 8.1%', '8.1% or higher'],
['Less than 1.1%', '1.1% to 1.6%', '1.6% to 2.6%', '2.6% to 4.8%', '4.8% or higher']],
'tract_labels':
[['Data not available', 'Less than 1 in 1k', '1 in 1k to 2 in 1k', '2 in 1k to 6 in 1k', '6 in 1k to 1 in 100', 'More than 1 in 100'],
['Data not available', 'Less than 9 in 1M', '9 in 1M to 7 in 100k', '7 in 100k to 3 in 10k', '3 in 10k to 2 in 1k', 'More than 2 in 1k'],
['Data not available', 'Less than 10 days', '10 to 13 days', '13 to 14 days', '14 to 17 days', 'More than 17 days'],
['Data not available', 'Less than 1.04 days', '1.04 to 1.10 days', '1.10 to 1.16 days', '1.16 to 1.20 days', 'More than 1.20 days'],
['Data not available', '0.0 to 0.2', '0.2 to 0.4', '0.4 to 0.6', '0.6 to 0.8', '0.8 to 1.0'],
['Slight', 'Low', 'Moderate', 'High', 'Severe'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['0 to 2th', '2 to 4th', '4 to 6th', '6 to 8th', '8 to 10th'],
['Less than 140 ha', '140 ha to 540 ha', '540 ha to 2 kha', '2 kha to 6 kha', 'More than 6 kha'],
['Data not available', '0.0 to 1.0th', '1.0 to 1.6th', '1.6 to 2.3th', '2.3 to 3.0th', '3.0 to 4.0th'],
['Data not available', 'Less than 5.6%', '5.6% to 9.7%', '9.7% to 14.9%', '14.9% to 23.4%', '23.4% or higher'],
['Data not available', 'Less than 3.0%', '3.0% to 4.5%', '4.5% to 6.2%', '6.2% to 8.9%', '8.9% or higher'],
['Data not available', 'Less than $20k', '$20k to $26k', '$26k to $32k', '$32k to $41k', '$41k or higher'],
['Data not available', 'Less than 4.6%', '4.6% to 8.1%', '8.1% to 12.6%', '12.6% to 20.1%', '20.1% or higher'],
['Data not available', '0.0 to 1.5th', '1.5 to 1.9th', '1.9 to 2.2th', '2.2 to 2.5 th', '2.5 to 3.8th'],
['Data not available', 'Less than 10.0%', '10.0% to 13.6%', '13.6% to 16.9%', '16.9% to 20.7%', '20.7% or higher'],
['Less than 17.6%', '17.6% to 20.9%', '20.9% to 23.6%', '23.6% to 27.1%', '27.1% or higher'],
['Data not available', 'Less than 8.5%', '8.5% to 11.2%', '11.2% to 14.0%', '14.0% to 17.8%', '17.8% or higher'],
['Less than 4.3%', '4.3% to 6.7%', '6.7% to 9.4%', '9.4% to 13.7%', '13.7% or higher'],
['Data not available', '0.0 to 0.4th', '0.4 to 0.8th', '0.8 to 1.1th', '1.1 to 1.5th', '1.5 to 2.0th'],
['Less than 9.9%', '9.9% to 21.8%', '21.8% to 39.6%', '39.6% to 70.1%', '70.1% or higher'],
['Less than 0.6%', '0.6% to 1.4%', '1.4% to 3.1%', '3.1% to 7.9%', '7.9% or higher'],
['Data not available', '0.0 to 1.6th', '1.6 to 2.1th', '2.1 to 2.5th', '2.5 to 3.0th', '3.0 to 4.6th'],
['Less than 1.8%', '1.8% to 5.4%', '5.4% to 11.9%', '11.9% to 25.2%', '25.2% or higher'],
['Data not available', 'Less than 1.0%', '1.0% to 3.1%', '3.1% to 8.7%', '8.7% to 18.7%', '18.7% or higher'],
['Less than 1.0%', '1.0% to 1.9%', '1.9% to 3.3%', '3.3% to 6.2%', '6.2% or higher'],
['Data not available', 'Less than 2.2%', '2.2% to 4.2%', '4.2% to 7.2%', '7.2% to 13.4%', '13.4% or higher'],
['Less than 0.3%', '0.3% to 0.6%', '0.6% to 1.6%', '1.6% to 3.8%', '3.8% or higher']],
'title':
['Riverine flood risk',
'Coastal flood risk',
'Projected Change in Extreme Heat Days (2030)',
'Projected Change in Extreme Precipitation Days (2030)',
'Drought risk',
'Landslide susceptibility',
'Earthquake hazards - frequency (deciles)',
'Volcano hazards - frequency (deciles)',
'Total hectares burned (2001-2017)']+
['Percentile ranking for Socioeconomic Status', 'percentage of persons below poverty', 'percentage of civilian (age 16+) unemployed', 'per capita income', 'percentage of persons with no high school diploma (age 25+)',
'Percentile ranking for Household & Disability','percentage of persons aged 65 and older', 'percentage of persons aged 17 and younger', 'percentage of civilian noninstitutionalized population with a disability', 'percentage of single parent households with children under 18',
'Percentile ranking for Minority Status & Language', 'percentage minority (all persons except white, non-Hispanic)', 'percentage of persons (age 5+) who speak English "less than well"',
'Percentile ranking for Housing Type & Transportation', 'percentage of housing in structures with 10 or more units estimate', 'percentage of mobile homes', 'percentage of occupied housing units with more people than rooms', 'percentage of households with no vehicle available', 'percentage of persons in institutionalized group quarters'],
'description':
['Riverine flood risk measures the percentage of population expected to be affected by Riverine flooding in an average year, accounting for existing flood-protection standards. Flood risk is assessed using hazard (inundation caused by river overflow), exposure (population in flood zone), and vulnerability. The existing level of flood protection is also incorporated into the risk calculation. It is important to note that this indicator represents flood risk not in terms of maximum possible impact but rather as average annual impact. The impacts from infrequent, extreme flood years are averaged with more common, less newsworthy flood years to produce the “expected annual affected population.” Higher values indicate that a greater proportion of the population is expected to be impacted by Riverine floods on average.',
'Coastal flood risk measures the percentage of the population expected to be affected by coastal flooding in an average year, accounting for existing flood protection standards. Flood risk is assessed using hazard (inundation caused by storm surge), exposure (population in flood zone), and vulnerability. The existing level of flood protection is also incorporated into the risk calculation. It is important to note that this indicator represents flood risk not in terms of maximum possible impact but rather as average annual impact. The impacts from infrequent, extreme flood years are averaged with more common, less newsworthy flood years to produce the “expected annual affected population.” Higher values indicate that a greater proportion of the population is expected to be impacted by coastal floods on average.',
'The Projected Change in Extreme Heat Days in the United States (U.S.) dataset shows the change in the average number of days of extreme heat at 2030, compared to a baseline time period of 1960-1990. An extreme heat day is a day where the maximum temperature is greater than the 99th percentile maximum temperature during the baseline period. The data shown represents a 31-year average, centered around the indicated year. The number of extreme heat days in 2030 is actually an average of the annual number of extreme heat days between the years 2015 and 2045. Temperature projections are based on the future greenhouse gas emission rates determined by the Intergovernmental Panel on Climate Change’s (IPCC’s) Representative Concentration Pathways (RCP) 8.5. RCP 8.5 is a hypothetical scenario where there is no decrease in greenhouse gas emission rates within the 21st century. Positive values indicate that the number of extreme heat days is increasing, while negative values indicate that the number is decreasing.',
'The Projected Change in Extreme Precipitation in the United States (U.S.) dataset shows the change in annual average days of extreme precipitation at 2030, compared to a baseline time period of 1960-1990. An extreme precipitation day is defined as a day where precipitation is greater than the 99th percentile precipitation compared to the baseline period. The data shown represents a 31-year average, centered around the indicated year. The number of extreme precipitation days in 2030 is actually an average of the annual number of extreme precipitation days between the years 2015 and 2045. Precipitation projections are based on the future greenhouse gas emission rates determined by the Intergovernmental Panel on Climate Change’s (IPCC’s) Representative Concentration Pathways (RCP) 8.5. RCP 8.5 is a hypothetical scenario where there is no decrease in greenhouse gas emission rates within the 21st century. Days of extreme precipitation are divided by the baseline average to calculate the projected change. Values greater than 1 indicate that extreme precipitation is increasing, while values less than one indicate that the number is decreasing.',
'Drought risk measures where droughts are likely to occur, the population and assets exposed, and the vulnerability of the population and assets to adverse effects. Higher values indicate higher risk of drought.',
'The Landslide Susceptibility Map, created by scientists at the National Aeronautics and Space Administration (NASA) and published in 2017 at 1 km resolution, improves upon past landslide susceptibility maps by incorporating the most up-to-date data. NASA scientists evaluated landslide susceptibility on slope from the Shuttle Radar Topography Mission and forest loss from a Landsat-based record of forest change compiled by a University of Maryland team. They also included data on key factors including the presence of roads from OpenStreetMap, the strength of bedrock and soils, and the locations of faults from the Geological Map of the World, 3rd ed., by <NAME> (2009). They generated the map using a heuristic fuzzy approach that generated the possibility of landslides and validated it through landslide inventories.',
'The Earthquake Hazard Frequency and Distribution data set is created by the World Bank Group, the Columbia University Center for Hazards and Risk Research (Columbia CHRR), and the Columbia University Earth Institute Center for International Earth Science Information Network (CIESIN). The data are released in a 2.5 minute grid utilizing Advanced National Seismic System (ANSS) Earthquake Catalog data of actual earthquake events exceeding 4.5 on the Richter scale during the time period 1976 through 2002.',
'The Volcano Hazard Frequency and Distribution data are created by the World Bank Group, the Columbia University Center for Hazards and Risk Research (Columbia CHRR), and the Columbia University Earth Institute Center for International Earth Science Information Network (CIESIN). The data are released in a 2.5 minute gridded data set based on the National Geophysical Data Center (NGDC) Volcano Database spanning the period 1979 through 2000. This database includes nearly 4,000 volcanic events categorized as moderate or above (values 2 through 8) according to the Volcano Explosivity Index (VEI). Most volcanoes are georeferenced to the nearest 10th or 100th of a degree, with a few to the nearest 1,000th of a degree.',
'Total hectares burned (2001-2017) has been obtained from the Global Wildfire Database for GWIS, An individual fire event focused database. Post processing of MCD64A1 providing geometries of final fire perimeters including initial and final date and the corresponding daily active areas for each fire. This dataset uses MCD64A1 Collection 6 MODIS Burned Area Product v6 and has a coverage from 2000 to 2018.',
'Socioeconomic status shows the 2014-2018 ACS 5-year estimates of the percentile ranking for socioeconomic theme summary for every county and census tract. Theme rankings are obtained by summing the percentiles of the social factors comprising each theme. The summed percentiles for each theme are then ordered to determine the theme-specific percentile rankings. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Below poverty social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons below poverty for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Unemployed social factor shows the 2014-2018 ACS 5-year estimates of the percentage of civilian (age 16+) unemployed for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Income social factor shows the 2014-2018 ACS 5-year estimates of the per capita income for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'No high school diploma social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons with no high school diploma (age 25+) for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Household composition and disability shows the 2014-2018 ACS 5-year estimates of the percentile ranking for household composition theme summary for every county and census tract. Theme rankings are obtained by summing the percentiles of the social factors comprising each theme. The summed percentiles for each theme are then ordered to determine the theme-specific percentile rankings. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Aged 65 or older social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons aged 65 and older for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Aged 17 or younger social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons aged 17 and younger for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Civilian with a disability social factor shows the 2014-2018 ACS 5-year estimates of the percentage of civilian noninstitutionalized population with a disability for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Single-parent households social factor shows the 2014-2018 ACS 5-year estimates of the percentage of single parent households with children under 18 for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Minority status and language shows the 2014-2018 ACS 5-year estimates of the percentile ranking for minority status and language theme summary for every county and census tract. Theme rankings are obtained by summing the percentiles of the social factors comprising each theme. The summed percentiles for each theme are then ordered to determine the theme-specific percentile rankings. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Minority social factor shows the 2014-2018 ACS 5-year estimates of the percentage minority (all persons except white, non-Hispanic) for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Speaks english "less than well" social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons (age 5+) who speak English "less than well" for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Housing type and transportation shows the 2014-2018 ACS 5-year estimates of the percentile ranking for housing type and transportation theme summary for every county and census tract. Theme rankings are obtained by summing the percentiles of the social factors comprising each theme. The summed percentiles for each theme are then ordered to determine the theme-specific percentile rankings. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Multi-unit structures social factor shows the 2014-2018 ACS 5-year estimates of the percentage of housing in structures with 10 or more units for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Mobile homes social factor shows the 2014-2018 ACS 5-year estimates of the percentage of mobile homes for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Crowding social factor shows the 2014-2018 ACS 5-year estimates of the percentage of occupied housing units with more people than rooms for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'No vehicle social factor shows the 2014-2018 ACS 5-year estimates of the percentage of households with no vehicle available for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.',
'Group quarters social factor shows the 2014-2018 ACS 5-year estimates of the percentage of persons in institutionalized group quarters for every county and census tract. This estimates belongs to the CDC Social Vulnerability Index (CDC SVI) and are created and maintained by the Geospatial Research, Analysis, and Services Program (GRASP). CDC SVI uses U.S. Census data to determine the social vulnerability of every census tract. The CDC SVI ranks each tract on 15 social factors and groups them into four related themes.'],
'source':
['https://www.wri.org/publication/aqueduct-30',
'https://www.wri.org/publication/aqueduct-30',
'http://loca.ucsd.edu/',
'http://loca.ucsd.edu/',
'https://www.wri.org/publication/aqueduct-30',
'https://pmm.nasa.gov/applications/global-landslide-model',
'http://sedac.ciesin.columbia.edu/data/set/ndh-earthquake-frequency-distribution',
'http://sedac.ciesin.columbia.edu/data/set/ndh-volcano-hazard-frequency-distribution',
'https://doi.pangaea.de/10.1594/PANGAEA.895835']+ ['https://www.atsdr.cdc.gov/placeandhealth/svi/data_documentation_download.html'] * 19
}
indicators = pd.DataFrame(indicators)
indicators
# -
# **Save as `csv`**
indicators.to_csv('../data/indicators_list.csv')
# ***
# <a id='vector_data'></a>
# ## Read vector data
# <a id='states'></a>
# **`States`:**
# +
url = 'https://www2.census.gov/geo/tiger/GENZ2019/shp/cb_2019_us_state_500k.zip'
local_path = '../data/cb_2019_us_state_500k'
if not os.path.isdir(local_path):
os.mkdir(local_path)
print('Downloading shapefile...')
r = requests.get(url)
z = zipfile.ZipFile(io.BytesIO(r.content))
print("Done")
z.extractall(path=local_path) # extract to folder
filenames = [y for y in sorted(z.namelist()) for ending in ['dbf', 'prj', 'shp', 'shx'] if y.endswith(ending)]
print(filenames)
state = gpd.read_file(local_path+ '/' + 'cb_2019_us_state_500k.shp')
state = state.set_crs(epsg=4326, allow_override=True)
state = state.to_crs("EPSG:4326")
state.sort_values('GEOID', inplace=True)
# Remove Territories of the United States
state = state.iloc[:51]
state = state[['GEOID', 'STATEFP', 'NAME', 'geometry']]
state.columns = map(str.lower, state.columns)
print("Shape of the dataframe: {}".format(state.shape))
print("Projection of dataframe: {}".format(state.crs))
# -
# **Add bbox**
state['bbox'] = state['geometry'].apply(lambda x: list(x.bounds))
# move level column
cols = state.columns.tolist()
state = state[cols[:len(cols)-2] + [cols[len(cols)-1]] + [cols[-2]]]
state = state.astype({'bbox': str})
state.head()
state
fig, ax = plt.subplots(figsize=(20, 10))
state.plot(ax=ax, edgecolor='k', facecolor='w')
plt.xlim(-180, -65)
# Save as `GeoJSON`
state.to_file('../data/state.json',driver='GeoJSON')
# <a id='county'></a>
# **`County`:**
# +
url = 'https://www2.census.gov/geo/tiger/GENZ2019/shp/cb_2019_us_county_500k.zip'
local_path = '../data/cb_2019_us_county_500k'
if not os.path.isdir(local_path):
os.mkdir(local_path)
print('Downloading shapefile...')
r = requests.get(url)
z = zipfile.ZipFile(io.BytesIO(r.content))
print("Done")
z.extractall(path=local_path) # extract to folder
filenames = [y for y in sorted(z.namelist()) for ending in ['dbf', 'prj', 'shp', 'shx'] if y.endswith(ending)]
print(filenames)
county = gpd.read_file(local_path+ '/' + 'cb_2019_us_county_500k.shp')
county = county.set_crs(epsg=4326, allow_override=True)
county = county.to_crs("EPSG:4326")
county.sort_values('GEOID', inplace=True)
# Remove Territories of the United States
county = county[~county['STATEFP'].isin(['60','66','69', '72', '78'])]
county = county[['GEOID', 'STATEFP', 'COUNTYFP', 'NAME', 'geometry']]
county.columns = map(str.lower, county.columns)
print("Shape of the dataframe: {}".format(county.shape))
print("Projection of dataframe: {}".format(county.crs))
# -
county
# **Add bbox**
county['bbox'] = county['geometry'].apply(lambda x: list(x.bounds))
# move level column
cols = county.columns.tolist()
county = county[cols[:len(cols)-2] + [cols[len(cols)-1]] + [cols[-2]]]
county = county.astype({'bbox': str})
county.head()
fig, ax = plt.subplots(figsize=(20, 10))
county.plot(ax=ax, edgecolor='k', facecolor='w')
plt.xlim(-180, -60)
# Save as `GeoJSON`
county.to_file('../data/county.json',driver='GeoJSON')
# <a id='tract'></a>
# **`Census tract`:**
# +
url = 'https://www2.census.gov/geo/tiger/GENZ2019/shp/cb_2019_us_tract_500k.zip'
local_path = '../data/cb_2019_us_tract_500k'
if not os.path.isdir(local_path):
os.mkdir(local_path)
print('Downloading shapefile...')
r = requests.get(url)
z = zipfile.ZipFile(io.BytesIO(r.content))
print("Done")
z.extractall(path=local_path) # extract to folder
filenames = [y for y in sorted(z.namelist()) for ending in ['dbf', 'prj', 'shp', 'shx'] if y.endswith(ending)]
print(filenames)
tract = gpd.read_file(local_path+ '/' + 'cb_2019_us_tract_500k.shp')
tract = tract.set_crs(epsg=4326, allow_override=True)
tract = tract.to_crs("EPSG:4326")
tract.sort_values('GEOID', inplace=True)
# Remove Territories of the United States
tract = tract[~tract['STATEFP'].isin(['60','66','69', '72', '78'])]
tract = tract[['GEOID', 'STATEFP', 'COUNTYFP', 'TRACTCE', 'NAME', 'geometry']]
tract.columns = map(str.lower, tract.columns)
print("Shape of the dataframe: {}".format(tract.shape))
print("Projection of dataframe: {}".format(tract.crs))
# -
tract
# **Add bbox**
tract['bbox'] = tract['geometry'].apply(lambda x: list(x.bounds))
# move level column
cols = tract.columns.tolist()
tract = tract[cols[:len(cols)-2] + [cols[len(cols)-1]] + [cols[-2]]]
tract = tract.astype({'bbox': str})
tract.head()
fig, ax = plt.subplots(figsize=(20, 10))
tract.plot(ax=ax, edgecolor='k', facecolor='w')
plt.xlim(-180, -60)
# Save as `GeoJSON`
tract.to_file('../data/tract.json',driver='GeoJSON')
# ### Save State and County names
# +
df = pd.merge(county[['geoid', 'statefp', 'name']], state[['statefp', 'name']], on='statefp', how='left')
df.rename(columns={'name_x': 'county', 'name_y': 'state'}, inplace=True)
d = (df.groupby('state')
.apply(lambda x: list(x['county']))
.to_dict())
with open('../data/state_county_names.json', 'w') as fp:
json.dump(d, fp)
# -
# ***
# <a id='carto_data'></a>
# ## Preprocess carto data
# ### Read data
# the four layers are provided from the same dataset
# search for the dataset
ds = lmi.Dataset(id_hash=datasets['carto']['riverine_flood_risk'])
ds
#get the layer from the dataset
l = ds.layers[0]
l
# ### Use raw values
# +
#build the sql to select the 4 attributes at the same time
account = l.attributes['layerConfig']['account']
query = "SELECT s.aq30_id, s.gid_1, s.pfaf_id, s.string_id, s.drr_label, s.drr_cat, s.drr_raw, s.drr_score, s.cfr_label, s.cfr_cat, s.cfr_raw, s.cfr_score, s.rfr_label, s.rfr_cat, s.rfr_raw, s.rfr_score, r.the_geom \
FROM water_risk_indicators_annual_all s \
LEFT JOIN y2018m12d06_rh_master_shape_v01 r on s.aq30_id=r.aq30_id \
WHERE s.pfaf_id != -9999 and s.gid_1 != '-9999' and r.aqid != -9999 and s.gid_1 like 'USA%'"
df_input = df_from_carto(account, query)
df_input = df_input[['rfr_raw', 'cfr_raw', 'drr_raw', 'geometry']]
df_input.rename(columns={'rfr_raw': 'rfr', 'cfr_raw': 'cfr', 'drr_raw': 'drr'}, inplace=True)
df_input = df_input[['rfr', 'cfr', 'drr', 'geometry']]
# Reproject
df_input = df_input.set_crs(epsg=4326, allow_override=True)
df_input = df_input.to_crs("EPSG:4326")
df_input.head()
# -
# **Check the inputs**
# +
#check the outputs
name = 'drr'
fig, ax = plt.subplots(figsize=(20, 10))
df_input.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# #### Intersect the geometries:
for n, df in enumerate([state, county]):
df_tmp = gpd.overlay(df, df_input, how='intersection')
df_tmp['level'] = n+1
# move level column
cols = df_tmp.columns.tolist()
df_tmp = df_tmp[cols[-1:] + cols[:-1]]
if n == 0:
df_out = df_tmp
else:
df_out = pd.concat([df_out, df_tmp])
# #### Compute area of each geometry
df_out['area'] = df_out.area
df_out.head()
# #### Weighted mean
# +
# Weighted mean
df_tmp1 = weighted_mean(df_out[df_out['level'] == 1], columns=['rfr', 'cfr', 'drr'], weight_column='area', groupby_on='geoid')
# Merge with geometries
df_tmp1 = pd.merge(state, df_tmp1, on='geoid', how='left')
df_tmp1['level'] = 1
cols = df_tmp1.columns.tolist()
df_tmp1 = df_tmp1[cols[-1:] + cols[:-1]]
# Weighted mean
df_tmp2 = weighted_mean(df_out[df_out['level'] == 2], columns=['rfr', 'cfr', 'drr'], weight_column='area', groupby_on='geoid')
# Merge with geometries
df_tmp2 = pd.merge(county, df_tmp2, on='geoid', how='left')
df_tmp2['level'] = 2
cols = df_tmp2.columns.tolist()
df_tmp2 = df_tmp2[cols[-1:] + cols[:-1]]
df_carto = pd.concat([df_tmp1, df_tmp2])
df_carto.head()
# -
# **Export temporal json data with carto processing**
#export temporal json data with carto processing
df_out.to_file(
'../data/carto_data.json',
driver='GeoJSON')
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'drr'
df = df_carto[df_carto['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# `Counties`:
# +
name = 'drr'
df = df_carto[df_carto['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# #### Compute scores category and ranges:
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
df_tmp1 = compute_score_category_label(df_carto[df_carto['level'] == 1], columns=['rfr', 'cfr', 'bws', 'drr'], q=q, scores=scores)
df_tmp2 = compute_score_category_label(df_carto[df_carto['level'] == 2], columns=['rfr', 'cfr', 'bws', 'drr'], q=q, scores=scores)
df_out_carto = pd.concat([df_tmp1, df_tmp2])
# -
df_out_carto.sort_values(['level','geoid'], inplace=True)
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'drr_cat'
df = df_out_carto[df_out_carto['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap='RdYlBu_r')
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
name = 'drr_cat'
df = df_out_carto[df_out_carto['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ### Use scores
# +
#build the sql to select the 4 attributes at the same time
account = l.attributes['layerConfig']['account']
query = "SELECT s.aq30_id, s.gid_1, s.pfaf_id, s.string_id, s.drr_label, s.drr_cat, s.drr_raw, s.drr_score, s.cfr_label, s.cfr_cat, s.cfr_raw, s.cfr_score, s.rfr_label, s.rfr_cat, s.rfr_raw, s.rfr_score, r.the_geom \
FROM water_risk_indicators_annual_all s \
LEFT JOIN y2018m12d06_rh_master_shape_v01 r on s.aq30_id=r.aq30_id \
WHERE s.pfaf_id != -9999 and s.gid_1 != '-9999' and r.aqid != -9999 and s.gid_1 like 'USA%'"
df_input = df_from_carto(account, query)
df_input = df_input[['rfr_score', 'cfr_score', 'drr_score', 'geometry']]
# Reproject
df_input = df_input.set_crs(epsg=4326, allow_override=True)
df_input = df_input.to_crs("EPSG:4326")
df_input.head()
# -
# #### Intersect the geometries:
for n, df in enumerate([state, county, tract]):
df_tmp = gpd.overlay(df, df_input, how='intersection')
df_tmp['level'] = n+1
# move level column
cols = df_tmp.columns.tolist()
df_tmp = df_tmp[cols[-1:] + cols[:-1]]
if n == 0:
df_out = df_tmp
else:
df_out = pd.concat([df_out, df_tmp])
# #### Compute area of each geometry
df_out['area'] = df_out.area
df_out.head()
# #### Weighted mean
# +
for n, df in enumerate([state, county, tract]):
# Weighted mean
df_tmp = weighted_mean(df_out[df_out['level'] == n+1], columns=['rfr_score', 'cfr_score', 'drr_score'], weight_column='area', groupby_on='geoid')
# Merge with geometries
df_tmp = pd.merge(df, df_tmp, on='geoid', how='left')
df_tmp['level'] = n+1
cols = df_tmp.columns.tolist()
df_tmp = df_tmp[cols[-1:] + cols[:-1]]
if n == 0:
df_carto = df_tmp
else:
df_carto = pd.concat([df_carto, df_tmp])
df_carto
# -
# #### Compute category and ranges:
# +
for indicator in ['rfr', 'cfr', 'drr']:
df_carto[indicator] = df_carto[f'{indicator}_score']
for n in df_carto['level'].unique():
df_tmp = compute_category_label(df_carto[df_carto['level'] == n], columns=['rfr', 'cfr', 'drr'])
if n == 1:
df_out_carto = df_tmp
else:
df_out_carto = pd.concat([df_out_carto, df_tmp])
df_out_carto
# -
df_out_carto.sort_values(['level','geoid'], inplace=True)
df_out_carto.to_csv('../data/carto_indicators.csv')
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'rfr_cat'
df = df_out_carto[df_out_carto['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap='RdYlBu_r')
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
name = 'rfr_cat'
df = df_out_carto[df_out_carto['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Census tracts`:
# +
#check the outputs
name = 'rfr_cat'
df = df_out_carto[df_out_carto['level'] == 3].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
df_out_carto.to_csv('../data/carto_indicators.csv')
#export temporal json data with carto processing
df_out_cat.to_file(
'../data/carto_data_cat.json',
driver='GeoJSON')
# ## 3. Preprocess Wildfire data:
# ### Read data
wildfires = gpd.read_file('../data/wildfires/wildfires.shp')
wildfires.head()
# ### Intersect the geometries:
# +
for n, df in enumerate([state, county, tract]):
number_fires, mean_fire_size, total_fire_size, per_total_fire_size = rtree_intersect(wildfires, df)
df_tmp = df.copy()
df_tmp['number_wlf'] = number_fires
df_tmp['mean_size_wlf'] = mean_fire_size
df_tmp['total_size_wlf'] = total_fire_size
df_tmp['per_total_size_wlf'] = per_total_fire_size
df_tmp['level'] = n+1
# move level column
cols = df_tmp.columns.tolist()
df_tmp = df_tmp[cols[-1:] + cols[:-1]]
if n == 0:
df_out = df_tmp
else:
df_out = pd.concat([df_out, df_tmp])
df_out.head()
# -
df_out
df_out = df_out.rename(columns={'total_size_wlf': 'wlf'})
df_out = df_out[['level', 'geoid', 'statefp', 'name', 'bbox', 'geometry', 'countyfp', 'wlf']]
# **Check the outputs**
#
# `States`:
# +
#check the outputs
df = df_out[df_out['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot('total_wlf_per', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
df = df_out[df_out['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot('total_wlf_per', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# `Census tracts`:
# +
#check the outputs
df = df_out[df_out['level'] == 3].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot('total_wlf_per', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# #### Compute scores category and ranges:
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
for n in df_carto['level'].unique():
df_tmp = compute_score_category_label(df_out[df_out['level'] == n], columns=['wlf'], q=q, scores=scores)
if n == 1:
df_out_wlf = df_tmp
else:
df_out_wlf = pd.concat([df_out_wlf, df_tmp])
df_out_wlf
# -
df_out_wlf.sort_values(['level','geoid'], inplace=True)
df_out_wlf.to_csv('../data/wlf_indicators.csv')
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'wlf_cat'
df = df_out_wlf[df_out_wlf['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Counties`
# +
#check the outputs
name = 'wlf_cat'
df = df_out_wlf[df_out_wlf['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Census tracts`
# +
#check the outputs
name = 'wlf_cat'
df = df_out_wlf[df_out_wlf['level'] == 3].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ## 4. GEE data preprocessing:
# ### Read `xarray.Dataset` from `TIFF` in GCS
file_names = {"efd": "dis.004_Earthquake_Frequency_and_Distribution.tif",
"lss": "dis.007_Landslide_Susceptibility_replacement.tif",
"vfd": "dis.008_Volcano_Frequency_and_Distribution.tif",
"ehd": "cli.056a_Projected_Change_in_Extreme_Heat_Days.tif",
"epd": "cli.059a_Projected_Change_in_Extreme_Precipitation_Days.tif"}
datasets = {}
base_url = 'https://storage.googleapis.com/us-resilience-map/datasets/'
for name, file_name in file_names.items():
print(f"Read xarray.Dataset from {file_name}")
url = f'{base_url}{file_name}'
xda = xr.open_rasterio(url).squeeze().drop("band")
if name == 'ehd':
# Replace all values equal to 0 with np.nan
xda = xda.where(xda != 0.)
else:
# Replace all values equal to -9999 with np.nan
xda = xda.where(xda != -9999.)
# Rename coordinates
xda = xda.rename({'x': 'lon', 'y': 'lat'})
# Convert the DataArray to a Dataset.
ds = xda.to_dataset(name=name)
datasets[name] = ds
# **Check data**
#
# `Landslide Susceptibility`:
fig, ax = plt.subplots(figsize=(20, 10))
ax.imshow(datasets['lss']['lss'].values)
fig, ax = plt.subplots(figsize=(20, 10))
ax.imshow(datasets['epd']['epd'].values)
# ### Zonal statistics
# **Create the data mask by rasterizing the vector data**
# +
masks = {'state': state,
'county': county,
'tract': tract}
for n, dataset in enumerate(datasets.items()):
ds_name = dataset[0]
ds = dataset[1]
print(f"Creating data masks in {ds_name}")
for mask_name, df in masks.items():
df = masks[mask_name]
df = df.reset_index().drop(columns='index').reset_index()
print(f"Creating the data mask by rasterizing the vector data for {mask_name}")
# Create the data mask by rasterizing the vector data
ds[mask_name], index = create_ds_mask(df, ds, name, lon_name='lon', lat_name='lat')
# Compute the mean value inside each geometry
print(f"Computing the mean value inside each geometry")
grouped_ds = ds.groupby(ds[mask_name])
grid_mean = grouped_ds.mean()
df_mean = grid_mean.to_dataframe()
df_mean = df_mean[[ds_name]].reset_index().rename(columns={mask_name: 'index'})
# Get centroid value for geometries smaller than mean cell size
print(f"Getting centroid value for geometries smaller than mean cell size")
if len(index) > 0:
df_others = df[df['index'].isin(index)][['index', 'geometry']].copy()
df_others['centroid'] = df['geometry'].apply(lambda x: list(x.centroid.coords)[0])
values = []
for centroid in tqdm(df_others['centroid']):
values.append(float(ds[ds_name].interp(lat=centroid[1], lon=centroid[0]).values))
df_others[ds_name] = values
df_mean = pd.concat([df_mean,df_others[['index', ds_name]]]).sort_values('index')
# Merge values with geometry GeoDataFrame
if mask_name == 'state':
if n == 0:
df_tmp1 = pd.merge(df, df_mean, on='index', how='left')
df_tmp1['level'] = 1
cols = df_tmp1.columns.tolist()
df_tmp1 = df_tmp1[cols[-1:] + cols[:-1]]
else:
df_tmp1[ds_name] = list(df_mean[ds_name])
elif mask_name == 'county':
if n == 0:
df_tmp2 = pd.merge(df, df_mean, on='index', how='left')
df_tmp2['level'] = 2
cols = df_tmp2.columns.tolist()
df_tmp2 = df_tmp2[cols[-1:] + cols[:-1]]
else:
df_tmp = pd.merge(df, df_mean, on='index', how='left')
df_tmp2[ds_name] = list(df_tmp[ds_name])
else:
if n == 0:
df_tmp3 = pd.merge(df, df_mean, on='index', how='left')
df_tmp3['level'] = 3
cols = df_tmp3.columns.tolist()
df_tmp3 = df_tmp3[cols[-1:] + cols[:-1]]
else:
df_tmp = pd.merge(df, df_mean, on='index', how='left')
df_tmp3[ds_name] = list(df_tmp[ds_name])
df_out = pd.concat([df_tmp1, df_tmp2, df_tmp3])
# -
df_out.head()
# **Check the outputs**
#
# `States`:
# +
name = 'ehd'
df = df_out[df_out['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# `Counties`:
# +
df = df_out[df_out['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# `Census tract`:
# +
df = df_out[df_out['level'] == 3].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# -
# #### Compute scores category and labels:
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
for n in df_out['level'].unique():
df_tmp = compute_score_category_label(df_out[df_out['level'] == n], columns=['lss', 'ehd', 'epd'], q=q, scores=scores)
if n == 1:
df_out_score = df_tmp
else:
df_out_score = pd.concat([df_out_score, df_tmp])
df_out_score
# -
# #### Compute decile category and labels:
# +
df_out['efd'] = df_out['efd'].fillna(value=0)
df_out['vfd'] = df_out['vfd'].fillna(value=0)
df_out = df_out[['level', 'index', 'geoid', 'statefp', 'countyfp', 'name', 'bbox', 'geometry', 'efd', 'vfd']].copy()
for n in df_out['level'].unique():
df_tmp = compute_decile_category_label(df_out[df_out['level'] == n], columns=['efd', 'vfd'])
if n == 1:
df_out_decile = df_tmp
else:
df_out_decile = pd.concat([df_out_decile, df_tmp])
df_out_decile
# -
df_out_decile.rename(columns={'efd_decile': 'efd_score', 'vfd_decile': 'vfd_score'}, inplace=True)
df_out_gee = pd.merge(df_out_score, df_out_decile.drop(columns=['level', 'index', 'statefp', 'countyfp', 'name', 'bbox', 'geometry']), on='geoid', how='left')
df_out_gee.sort_values(['level', 'geoid'], inplace=True)
df_out_gee.to_csv('../data/gee_indicators.csv')
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'efd_cat'
df = df_out_gee[df_out_gee['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
name = 'efd_cat'
df = df_out_gee[df_out_gee['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ## 5. Other datasets:
# ### 5.1. Population data:
# Population data has been obtained from the following link: https://www2.census.gov/programs-surveys/popest/datasets/2010-2019/counties/totals/co-est2019-alldata.pdf
url = "https://www2.census.gov/programs-surveys/popest/datasets/2010-2019/counties/totals/co-est2019-alldata.csv"
pop_state_county = pd.read_csv(url,encoding='ISO-8859-1')
pop_state_county = pop_state_county[['STNAME','COUNTY','STATE', 'CTYNAME','POPESTIMATE2019']]
pop_state_county = pop_state_county.astype({'POPESTIMATE2019':'int32'})
pop_state_county.columns = map(str.lower, pop_state_county.columns)
pop_state_county.rename(columns={'popestimate2019':'pop'}, inplace=True)
pop_state_county.head()
# #### Population by `State`
state.head()
# **Get population data by state**
pop_state = pop_state_county[pop_state_county['county'] == 0]
pop_state = pop_state[['state', 'stname', 'pop']]
pop_state = pop_state.reset_index().drop(columns='index').reset_index()
pop_state.head()
# **Merge state with the population dataset**
pop_l1 = pd.merge(state.astype({'statefp':'int32'}), pop_state[['state', 'pop']], left_on='statefp', right_on='state', how='left')
pop_l1 = gpd.GeoDataFrame(pop_l1)
pop_l1 = pop_l1[['geoid', 'statefp', 'name', 'pop', 'geometry']]
pop_l1['level'] = 1
pop_l1.head()
# **Check the outputs**
fig, ax = plt.subplots(figsize=(20, 10))
pop_l1.plot('pop', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# #### Population by `County`
county.head()
# **Get population data by county**
pop_county = pop_state_county[pop_state_county['county'] != 0]
pop_county.head()
# **Merge county with the population dataset**
pop_l2 = pd.merge(county.astype({'statefp':'int32', 'countyfp':'int32'}), pop_county[['stname', 'state', 'county', 'pop']].astype({'state':'object', 'county':'object'}), left_on=['statefp','countyfp'], right_on=['state','county'], how='left')
pop_l2 = gpd.GeoDataFrame(pop_l2)
pop_l2 = pop_l2[['geoid', 'countyfp', 'name', 'pop', 'geometry']]
pop_l2['level'] = 2
pop_l2.head()
# **Check the outputs**
fig, ax = plt.subplots(figsize=(20, 10))
pop_l2.plot('pop', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
df_pop = pd.concat([pop_l1, pop_l2])
df_pop
# #### Compute scores category and ranges:
df_out = df_pop[['geoid', 'name', 'level', 'geometry', 'pop']]
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
df_tmp1 = compute_score_category_label(df_out[df_out['level'] == 1], columns=['pop'], q=q, scores=scores)
df_tmp2 = compute_score_category_label(df_out[df_out['level'] == 2], columns=['pop'], q=q, scores=scores)
df_out_pop = pd.concat([df_tmp1, df_tmp2])
# -
df_out_pop.sort_values(['level','geoid'], inplace=True)
df_out_pop.head()
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'pop_cat'
df = df_out_pop[df_out_pop['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Counties`
# +
#check the outputs
name = 'pop_cat'
df = df_out_pop[df_out_pop['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ### 5.2. [Poverty, Unemployment, and Median Household Income](https://www.ers.usda.gov/data-products/county-level-data-sets/):
# Poverty, Unemployment, and Median Household Income data has been obtained from the following link: https://www.ers.usda.gov/data-products/county-level-data-sets/download-data/
#
# **Variables:**
# - Unemployment_rate_2019: Unemployment rate, 2019
# - Median_Household_Income_2018: Estimate of Median household Income, 2018 ($)
url = "https://www.ers.usda.gov/webdocs/DataFiles/48747/Unemployment.xls?v=7657.3"
mhi_state_county = pd.read_excel(url, sheet_name='Unemployment Med HH Income', skiprows = range(0, 7), usecols=['FIPStxt', 'Stabr', 'area_name', 'Unemployment_rate_2019', 'Median_Household_Income_2018'])
mhi_state_county.columns = map(str.lower, mhi_state_county.columns)
mhi_state_county.rename(columns={'unemployment_rate_2019':'uep', 'median_household_income_2018':'mhi'}, inplace=True)
# Drop US
mhi_state_county.drop(index=0, inplace=True)
# Reset Index
mhi_state_county = mhi_state_county.reset_index().drop(columns='index')
url = "https://www.ers.usda.gov/webdocs/DataFiles/48747/PovertyEstimates.xls?v=5138.5"
ppp_state_county = pd.read_excel(url, sheet_name='Poverty Data 2018', skiprows = range(0, 4), usecols=['FIPStxt', 'Stabr', 'Area_name', 'PCTPOVALL_2018'])
ppp_state_county.columns = map(str.lower, ppp_state_county.columns)
ppp_state_county.rename(columns={'pctpovall_2018':'pvt'}, inplace=True)
# Drop US
ppp_state_county.drop(index=0, inplace=True)
# Reset Index
ppp_state_county = ppp_state_county.reset_index().drop(columns='index')
mhi_state_county = pd.merge(mhi_state_county, ppp_state_county[['fipstxt', 'pvt']], on='fipstxt', how='left')
mhi_state_county.head()
# #### Poverty, Unemployment, and Median Household Income by `State`
# **Get population data by state**
mhi_state = mhi_state_county[pd.Series([s.find(',') for s in list(mhi_state_county['area_name'])]) == -1]
mhi_state.drop(index=331, inplace=True)
mhi_state = mhi_state.reset_index().drop(columns='index')
mhi_state['statefp'] = mhi_state['fipstxt'].apply(lambda x: int(x/1000))
mhi_state.head()
# **Merge state with the population dataset**
mhi_l1 = pd.merge(state.astype({'statefp':'int32'}), mhi_state[['statefp', 'uep', 'mhi', 'pvt']], on='statefp', how='left')
mhi_l1 = gpd.GeoDataFrame(mhi_l1)
mhi_l1 = mhi_l1[['geoid', 'statefp', 'name', 'uep', 'mhi', 'pvt', 'geometry']]
mhi_l1['level'] = 1
mhi_l1.head()
# **Check the outputs**
fig, ax = plt.subplots(figsize=(20, 10))
mhi_l1.plot('mhi', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# #### Poverty, Unemployment, and Median Household Income by `County`
# **Get population data by county**
mhi_county = mhi_state_county[pd.Series([s.find(',') for s in list(mhi_state_county['area_name'])]) != -1]
mhi_county = mhi_county.reset_index().drop(columns='index')
mhi_county.head()
# **Merge county with the population dataset**
mhi_l2 = pd.merge(county.astype({'geoid':'int32'}), mhi_county[['fipstxt', 'uep', 'mhi', 'pvt']], left_on='geoid', right_on='fipstxt', how='left')
mhi_l2 = gpd.GeoDataFrame(mhi_l2)
mhi_l2 = mhi_l2[['geoid', 'countyfp', 'name', 'uep', 'mhi', 'pvt', 'geometry']]
mhi_l2['level'] = 2
mhi_l2.head()
# **Check the outputs**
fig, ax = plt.subplots(figsize=(20, 10))
mhi_l2.plot('mhi', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
df_mhi = pd.concat([mhi_l1, mhi_l2])
df_mhi
# #### Compute scores category and ranges:
df_out = df_mhi[['geoid', 'name', 'level', 'geometry', 'uep', 'mhi', 'pvt']]
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
df_tmp1 = compute_score_category_label(df_out[df_out['level'] == 1], columns=['uep', 'mhi', 'pvt'], q=q, scores=scores)
df_tmp2 = compute_score_category_label(df_out[df_out['level'] == 2], columns=['uep', 'mhi', 'pvt'], q=q, scores=scores)
df_out_mhi = pd.concat([df_tmp1, df_tmp2])
# -
df_out_mhi.sort_values(['level','geoid'], inplace=True)
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'mhi_cat'
df = df_out_mhi[df_out_mhi['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
name = 'mhi_cat'
df = df_out_mhi[df_out_mhi['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ### 5.3. [Community Resilience Estimate data](https://www.census.gov/data/experimental-data-products/community-resilience-estimates.html):
# Community Resilience Estimate data has been obtained from the following link: https://www2.census.gov/data/experimental-data-products/community-resilience-estimates/2020/technical-document.pdf
url = "https://www2.census.gov/data/experimental-data-products/community-resilience-estimates/2020/cre-2018-a11.csv"
cre_county = pd.read_csv(url,encoding='ISO-8859-1')
cre_county = cre_county[cre_county['tract'] == 0]
cre_county = cre_county[['stname','county','state', 'ctname','rfgrp', 'prednum', 'prednum_moe', 'predrt', 'predrt_moe', 'popuni']]
cre_county.head()
len(cre_county)
# We take only as an indicator the percentage of people at 3+ Risk Factors
cre_county = cre_county[cre_county['rfgrp'] == '3PLRF']
cre_county.head()
# **Community Resilience Estimate by `County`**
county.head()
#NOTE: the merge is not matching the data properly
#I've changed the merge on by two attributes (state and county)
cre_l2 = pd.merge(county.astype({'countyfp': int, 'statefp': int}), cre_county[['county','state', 'prednum', 'predrt', 'popuni']], left_on=['countyfp','statefp'], right_on=['county','state'], how='left')
cre_l2 = cre_l2[['geoid', 'countyfp', 'name', 'predrt', 'geometry']]
cre_l2['level'] = 2
cre_l2.rename(columns={'predrt': 'cre'}, inplace=True)
cre_l2.head()
# `predrt` (Percentage of Residents in County with 3+ Risk Factors) es el indicador que queremos usar
#
# **Check outputs:**
fig, ax = plt.subplots(figsize=(20, 10))
cre_l2.plot('cre', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
# **Community Resilience Estimate by `State`**
# Para calcular el `predrt` por stado agregar el `prednum` (Number of Residents in County with 3+ Risk Factors) y el `popuni` (Number of Residents in County) y sacar los porcentages.
# #### Agregate by state:
cre_state = cre_county[['state', 'prednum', 'popuni']].groupby(['state']).sum().reset_index()
cre_state['predrt'] = cre_state['prednum']/cre_state['popuni']*100
cre_state.head()
cre_l1 = pd.merge(state.astype({'statefp': int}), cre_state[['state', 'predrt']], left_on='statefp', right_on='state', how='left')
cre_l1 = cre_l1[['geoid', 'statefp', 'name', 'predrt', 'geometry']]
cre_l1['level'] = 1
cre_l1.rename(columns={'predrt': 'cre'}, inplace=True)
cre_l1.head()
# **Check outputs:**
fig, ax = plt.subplots(figsize=(20, 10))
cre_l1.plot('cre', ax=ax, cmap="RdYlBu_r", scheme='quantiles', legend=True)
plt.xlim(-180, -65)
df_cre = pd.concat([cre_l1, cre_l2])
df_cre
# #### Compute scores category and ranges:
df_out = df_cre[['geoid', 'name', 'level', 'geometry', 'cre']]
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
df_tmp1 = compute_score_category_label(df_out[df_out['level'] == 1], columns=['cre'], q=q, scores=scores)
df_tmp2 = compute_score_category_label(df_out[df_out['level'] == 2], columns=['cre'], q=q, scores=scores)
df_out_cre = pd.concat([df_tmp1, df_tmp2])
# -
df_out_cre.sort_values(['level','geoid'], inplace=True)
# **Check the outputs**
#
# `States`:
# +
#check the outputs
name = 'cre_cat'
df = df_out_cre[df_out_cre['level'] == 1].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# `Counties`:
# +
#check the outputs
name = 'cre_cat'
df = df_out_cre[df_out_cre['level'] == 2].copy()
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
# ### 5.4. [Economic damage from climate change](http://www.impactlab.org/research/estimating-economic-damage-from-climate-change-in-the-united-states/):
df = pd.read_csv('http://www.globalpolicy.science/s/county_damages_by_sector.csv')
# **Economic damage by `County`**
ecd_l1 = pd.merge(left=county.astype({'geoid': int}), right=df.drop(columns=['State Code', 'State Code', 'County Name']).rename(columns={'County FIPS code': 'geoid'}), how='left', on='geoid')
ecd_l1.head()
# **Economic damage by `State`**
# Seleccionar los indicadores de interes y agregarlos
# ### 5.5. [CDC Social Vulnerability Index data](https://www.atsdr.cdc.gov/placeandhealth/svi/data_documentation_download.html):
# CDC SVI Documentation: https://svi.cdc.gov/Documents/Data/2018_SVI_Data/SVI2018Documentation-508.pdf
# +
indicators = {'Name':['riverine flood risk', 'coastal flood risk', 'extreme heat days', 'extreme precipitation days',
'drought risk', 'landslide susceptibility', 'earthquake frequency and distribution',
'volcano frequency and distribution', 'wildfires'] +
['socioeconomic status', 'percentage of persons below poverty', 'percentage of civilian (age 16+) unemployed', 'per capita income', 'percentage of persons with no high school diploma (age 25+)',
'household composition & disability','percentage of persons aged 65 and older', 'percentage of persons aged 17 and younger', 'percentage of civilian with a disability', 'percentage of single parent households with children under 18',
'minority status & language', 'minority (all persons except white, non-Hispanic)', 'speaks english “less than well”',
'housing type & transportation', 'multi-unit structures', 'mobile homes', 'crowding', 'no vehicle', 'group quarters'],
'Slug':['rfr', 'cfr', 'ehd', 'epd','drr', 'lss', 'efd','vfd', 'wlf'] +
['ses', 'pov', 'uep', 'pci', 'hsd', 'hcd', 'a6o', 'a1y', 'pcd', 'sph', 'msl', 'mnt', 'sen', 'htt', 'mus', 'mhm', 'cwd', 'vhc', 'gqt'],
'Category': ['climate risk'] * 9 + ['vulnerability'] * 19,
'Group': list(range(0, 9)) + [9, 9.1,9.2,9.3,9.4] + [10, 10.1,10.2,10.3,10.4] + [11, 11.1,11.2] + [12, 12.1,12.2,12.3,12.4,12.5]}
indicators = pd.DataFrame(indicators)
indicators.to_csv('../data/indicators_list.csv')
indicators
# +
indicators = {'group': ['climate risk'] * 9 + ['vulnerability'] * 19,
'indicator':['riverine flood risk', 'coastal flood risk', 'extreme heat days', 'extreme precipitation days',
'drought risk', 'landslide susceptibility', 'earthquake frequency and distribution',
'volcano frequency and distribution', 'wildfires'] +
['socioeconomic status', 'percentage of persons below poverty', 'percentage of civilian (age 16+) unemployed', 'per capita income', 'percentage of persons with no high school diploma (age 25+)',
'household composition & disability','percentage of persons aged 65 and older', 'percentage of persons aged 17 and younger', 'percentage of civilian with a disability', 'percentage of single parent households with children under 18',
'minority status & language', 'minority (all persons except white, non-Hispanic)', 'speaks english “less than well”',
'housing type & transportation', 'multi-unit structures', 'mobile homes', 'crowding', 'no vehicle', 'group quarters'],
'slug':['rfr', 'cfr', 'ehd', 'epd','drr', 'lss', 'efd','vfd', 'wlf'] +
['ses', 'pov', 'uep', 'pci', 'hsd', 'hcd', 'a6o', 'a1y', 'pcd', 'sph', 'msl', 'mnt', 'sen', 'htt', 'mus', 'mhm', 'cwd', 'vhc', 'gqt']}
indicators = pd.DataFrame(indicators)
indicators
# -
# `States`
# +
indicators = ['E_TOTPOP', 'E_POV', 'E_UNEMP', 'E_PCI', 'E_NOHSDP', 'E_AGE65', 'E_AGE17', 'E_DISABL', 'E_SNGPNT', 'E_MINRTY',
'E_LIMENG', 'E_MUNIT', 'E_MOBILE', 'E_CROWD', 'E_NOVEH', 'E_GROUPQ']
svi_indicator = {
'SPL_THEME1': 'ses',
'EP_POV': 'pov',
'EP_UNEMP': 'uep',
'EP_PCI': 'pci',
'EP_NOHSDP': 'hsd',
'SPL_THEME2': 'hcd',
'EP_AGE65': 'a6o',
'EP_AGE17': 'a1y',
'EP_DISABL': 'pcd',
'EP_SNGPNT': 'sph',
'SPL_THEME3': 'msl',
'EP_MINRTY': 'mnt',
'EP_LIMENG': 'sen',
'SPL_THEME4': 'htt',
'EP_MUNIT': 'mus',
'EP_MOBILE': 'mhm',
'EP_CROWD': 'cwd',
'EP_NOVEH': 'vhc',
'EP_GROUPQ': 'gqt'}
THEME_indicator = {'SPL_THEME1': ['EPL_POV', 'EPL_UNEMP','EPL_PCI', 'EPL_NOHSDP'],
'SPL_THEME2': ['EPL_AGE65', 'EPL_AGE17', 'EPL_DISABL', 'EPL_SNGPNT'],
'SPL_THEME3': ['EPL_MINRTY', 'EPL_LIMENG'],
'SPL_THEME4': ['EPL_MUNIT', 'EPL_MOBILE','EPL_CROWD', 'EPL_NOVEH', 'EPL_GROUPQ']}
# -
svi_state = pd.read_csv('https://svi.cdc.gov/Documents/Data/2018_SVI_Data/CSV/SVI2018_US_COUNTY.csv')
svi_state = svi_state[['ST'] + indicators]
svi_state = svi_state.sort_values('ST')
svi_state = svi_state.replace(-999, np.nan)
svi_state.head()
# +
indicators.remove('E_PCI')
e_svi_state = svi_state[['ST']+['E_PCI']].groupby('ST').mean().reset_index()
ep_svi_state = svi_state[['ST']+ indicators].groupby('ST').sum().reset_index()
colm = 'E_PCI'
e_svi_state[colm.replace('E_', 'EP_')] = e_svi_state[colm]
e_svi_state.drop(columns=colm, inplace=True)
indicators.remove('E_TOTPOP')
for colm in indicators:
ep_svi_state[colm.replace('E_', 'EP_')] = ep_svi_state[colm]/ep_svi_state['E_TOTPOP']*100
ep_svi_state.drop(columns=colm, inplace=True)
ep_svi_state.drop(columns='E_TOTPOP', inplace=True)
svi_state = pd.merge(e_svi_state, ep_svi_state, on='ST', how='left')
svi_state.head()
# -
def compute_percentile(df, columns):
df = df.copy()
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,0.2,0.4,0.6,0.8,1]
for column in columns:
df.sort_values(column, inplace=True)
s = df[column]
f, quantiles =quantile_interp_function(s,q,scores)
# Add percentile
df[column.replace('EP', 'EPL')] = df[column].apply(f)
return df
svi_state = compute_percentile(svi_state, [colm.replace('E_', 'EP_') for colm in ['E_PCI'] + indicators])
svi_state.head()
# +
for theme in THEME_indicator.keys():
svi_state[theme] = svi_state[THEME_indicator[theme]].sum(axis=1)
svi_state.drop(columns=[colm.replace('E_', 'EPL_') for colm in ['E_PCI'] + indicators], inplace=True)
svi_state = svi_state.rename(columns=svi_indicator)
svi_state = svi_state.rename(columns={'ST': 'geoid'})
svi_state.columns = map(str.lower, svi_state.columns)
# add level
svi_state['level'] = 1
# add name
svi_state = pd.merge(state[['geoid', 'name']].astype({'geoid': int}), svi_state, on='geoid', how='left')
svi_state.sort_values('geoid', inplace=True)
svi_state = svi_state[['geoid', 'name', 'level'] + list(svi_indicator.values())]
svi_state.head()
# -
# `County`
svi_indicator = {
'SPL_THEME1': 'ses',
'EP_POV': 'pov',
'EP_UNEMP': 'uep',
'E_PCI': 'pci',
'EP_NOHSDP': 'hsd',
'SPL_THEME2': 'hcd',
'EP_AGE65': 'a6o',
'EP_AGE17': 'a1y',
'EP_DISABL': 'pcd',
'EP_SNGPNT': 'sph',
'SPL_THEME3': 'msl',
'EP_MINRTY': 'mnt',
'EP_LIMENG': 'sen',
'SPL_THEME4': 'htt',
'EP_MUNIT': 'mus',
'EP_MOBILE': 'mhm',
'EP_CROWD': 'cwd',
'EP_NOVEH': 'vhc',
'EP_GROUPQ': 'gqt'}
svi_county = pd.read_csv('https://svi.cdc.gov/Documents/Data/2018_SVI_Data/CSV/SVI2018_US_COUNTY.csv')
svi_county = svi_county[['ST','FIPS']+list(svi_indicator.keys())]
svi_county = svi_county.rename(columns=svi_indicator)
svi_county = svi_county.rename(columns={'FIPS': 'geoid'})
svi_county.columns = map(str.lower, svi_county.columns)
svi_county = svi_county.replace(-999, np.nan)
svi_county['level'] = 2
# add name
svi_county = pd.merge(county[['geoid', 'name']].astype({'geoid': int}), svi_county, on='geoid', how='left')
svi_county.sort_values('geoid', inplace=True)
svi_county = svi_county[['geoid', 'name', 'level'] + list(svi_indicator.values())]
svi_county.head()
# `Census tract`
svi_tract = pd.read_csv('https://svi.cdc.gov/Documents/Data/2018_SVI_Data/CSV/SVI2018_US.csv')
svi_tract = svi_tract[['ST','FIPS']+list(svi_indicator.keys())]
svi_tract = svi_tract.rename(columns=svi_indicator)
svi_tract = svi_tract.rename(columns={'FIPS': 'geoid'})
svi_tract.columns = map(str.lower, svi_tract.columns)
svi_tract = svi_tract.replace(-999, np.nan)
svi_tract['level'] = 3
# add name
svi_tract = pd.merge(tract[['geoid', 'name']].astype({'geoid': int}), svi_tract, on='geoid', how='left')
svi_tract = svi_tract[['geoid', 'name', 'level'] + list(svi_indicator.values())]
svi_tract.sort_values('geoid', inplace=True)
svi_tract.head()
df_svi = pd.concat([svi_state, svi_county, svi_tract])
df_svi
# +
q = [0,0.2,0.4,0.6,0.8,1]
scores = [0,1,2,3,4,5]
for n in df_svi['level'].unique():
df_tmp = compute_score_category_label(df_svi[df_svi['level'] == n], columns=list(svi_indicator.values()), q=q, scores=scores)
if n == 1:
df_out_svi = df_tmp
else:
df_out_svi = pd.concat([df_out_svi, df_tmp])
df_out_svi
# -
df_out_svi.sort_values(['level', 'geoid'], inplace=True)
df_out_svi.to_csv('../data/svi_indicators.csv')
# **Check the outputs**
#
# `States`:
#check the outputs
name = 'pov_cat'
df = df_out_svi[df_out_svi['level'] == 1].copy()
df = gpd.GeoDataFrame(pd.merge(df, state[['geoid', 'geometry']].astype({'geoid': int}), on='geoid', how='left'))
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap='RdYlBu_r')
plt.xlim(-180, -65)
# `Counties`:
#check the outputs
name = 'pov_cat'
df = df_out_svi[df_out_svi['level'] == 2].copy()
df = gpd.GeoDataFrame(pd.merge(df, county[['geoid', 'geometry']].astype({'geoid': int}), on='geoid', how='left'))
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap='RdYlBu_r')
plt.xlim(-180, -65)
# `Census tracts`:
#check the outputs
name = 'pov_cat'
df = df_out_svi[df_out_svi['level'] == 3].copy()
df = gpd.GeoDataFrame(pd.merge(df, tract[['geoid', 'geometry']].astype({'geoid': int}), on='geoid', how='left'))
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap='RdYlBu_r')
plt.xlim(-180, -65)
# ## 6. Merge all datasets:
# +
svi_indicator = {
'SPL_THEME1': 'ses',
'EP_POV': 'pov',
'EP_UNEMP': 'uep',
'EP_PCI': 'pci',
'EP_NOHSDP': 'hsd',
'SPL_THEME2': 'hcd',
'EP_AGE65': 'a6o',
'EP_AGE17': 'a1y',
'EP_DISABL': 'pcd',
'EP_SNGPNT': 'sph',
'SPL_THEME3': 'msl',
'EP_MINRTY': 'mnt',
'EP_LIMENG': 'sen',
'SPL_THEME4': 'htt',
'EP_MUNIT': 'mus',
'EP_MOBILE': 'mhm',
'EP_CROWD': 'cwd',
'EP_NOVEH': 'vhc',
'EP_GROUPQ': 'gqt'}
indicartos = ['rfr', 'cfr', 'ehd', 'epd', 'drr', 'lss', 'efd','vfd', 'wlf']+list(svi_indicator.values())
# -
df_carto = pd.read_csv('../data/carto_indicators.csv').drop(columns=['Unnamed: 0', 'statefp', 'countyfp', 'tractce', 'bbox', 'geometry'])
df_gee = pd.read_csv('../data/gee_indicators.csv').drop(columns=['Unnamed: 0', 'index', 'statefp', 'countyfp', 'bbox', 'geometry'])
df_wlf = pd.read_csv('../data/wlf_indicators.csv').drop(columns=['Unnamed: 0', 'statefp', 'countyfp', 'bbox', 'geometry'])
df_svi = pd.read_csv('../data/svi_indicators.csv').drop(columns='Unnamed: 0')
#df_all = pd.merge(df_svi, df_carto, on=['geoid', 'level', 'name'], how='left')
#df_all = pd.merge(df_all, df_gee, on=['geoid', 'level', 'name'], how='left')
#df_all = pd.merge(df_all, df_wlf, on=['geoid', 'level', 'name'], how='left')
df_all = pd.merge(df_svi, df_carto, on=['geoid', 'level'], how='left')
df_all = pd.merge(df_all, df_gee, on=['geoid', 'level'], how='left')
df_all = pd.merge(df_all, df_wlf, on=['geoid', 'level'], how='left')
df_all.drop(columns='name_y', inplace=True)
df_all.rename(columns={'name_x': 'name'}, inplace=True)
# + jupyter={"outputs_hidden": true}
df_all
# -
all_indicators = []
for indicator in indicartos:
all_indicators.append([indicator, f'{indicator}_score', f'{indicator}_cat', f'{indicator}_range'])
all_indicators = [item for sublist in all_indicators for item in sublist]
df_all = df_all[['geoid', 'name', 'level']+all_indicators]
# +
state['geoid_int'] = state['geoid']
county['geoid_int'] = county['geoid']
tract['geoid_int'] = tract['geoid']
df_all = pd.concat([pd.merge(state[['geoid', 'geoid_int']].astype({'geoid_int':int}), df_all[df_all['level'] == 1], left_on='geoid_int', right_on='geoid', how='left')\
.drop(columns=['geoid_int', 'geoid_y']).rename(columns={'geoid_x': 'geoid'}),
pd.merge(county[['geoid', 'geoid_int']].astype({'geoid_int':int}), df_all[df_all['level'] == 2], left_on='geoid_int', right_on='geoid', how='left')\
.drop(columns=['geoid_int', 'geoid_y']).rename(columns={'geoid_x': 'geoid'}),
pd.merge(tract[['geoid', 'geoid_int']].astype({'geoid_int':int}), df_all[df_all['level'] == 3], left_on='geoid_int', right_on='geoid', how='left')\
.drop(columns=['geoid_int', 'geoid_y']).rename(columns={'geoid_x': 'geoid'})])
# -
df_all
# Save as `csv`
df_all.to_csv("../data/indicators.csv")
df = pd.read_csv("../data/indicators.csv")
df
df = pd.read_csv("../data/indicators.csv")
df.drop(columns='Unnamed: 0', inplace=True)
df = df[df['level'] == 3]
df = pd.merge(df,tract.astype({'geoid': int}), how='left', on='geoid')
df = gpd.GeoDataFrame(df)
df.head()
# +
#check the outputs
name = 'rfr_score'
fig, ax = plt.subplots(figsize=(20, 10))
df.plot(name, ax=ax, cmap="RdYlBu_r")
plt.xlim(-180, -65)
# -
indicators
| processing/USA_Resilience_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2: Exploring sources of bias in data
# ## Bias in data, demystified
#
# **All datasets have limitations. Any of these limitations can be a potential source of bias.** In the context of data, a "bias" is really just a divergence between what a dataset is supposed to capture about the world--according to the *dataset designer's intention*, or according to the *end-user's expectations*--and what's actually represented in the data.
#
# So biases in data are often highly *contextual*, which can make them subtle and hard to spot. Similarly, it's hard to predict ahead of time what the *consequences* of those biases might be, because it depends on what the data is being used for.
# Nevertheless some sources of "bias" in data show up over and over again. Some of these are:
# - **duplicated data:** the same data shows up multiple times
# - **incomplete data:** some data is missing from the dataset
# - **misleading data:** it looks like a piece of data means one thing, but it actually means something different
# - **unrepresentative data:** the dataset doesn't represent the population it's gathered from OR the population it's intended to model
#
#
# Any of these sources of bias, unless their properly documented (with a data statement, etc) can have unintended consequences: they cause a researcher who is using the data to reach incorrect conclusions, or cause a machine learning model that is trained on that dataset to make classification errors.
#
# The nature or scale of these consequences can be hard to predict. That's why any time you prepare to use data that you didn't gather yourself, it pays to spend some time exploring the dataset, identifying limitations, and thinking critically about how these limitations might affect your analysis or your machine learning model.
# ## Introduction to assignment 2
# For this assignment, you'll be working with the Wikipedia Talk Corpus. You have already created a data statement for this dataset in class, so you're an 'expert' on it compared to most people! For more background on the Wikipedia Talk Corpus, please review the background information section of the [A2 assignment sheet](https://docs.google.com/document/d/1JZ9IubfxpCCKjv4v1jRLLw--IDM34lafcvg9V5-CaAs/edit?usp=sharing).
#
# **In assignment 1, you learned how to process and analyze a dataset created by someone else.** In that assignment, we took the dataset at face value: we assumed that there were no major errors, no missing data, nothing that would skew our results. We assumed that the dataset really did accurately capture all the bike and pedestrian traffic on the Burke-Gilman trail over the specified period of time.
#
# **In assignment 2, we'll also be analyzing a dataset created by someone else,** but this time *we won't assume that the data is complete and correct.* Instead, we will try to identify ways in which the data might be WRONG, and form hypotheses about how the limitations we discover might make it an unsuitable, or at least a potentially problematic, training dataset for a general-purpose hostile speech detector.
#
# ***In part 1 of the assignment,*** you will load one type of Wikipedia Talk Corpus data--the demographics of the crowdworkers who labelled the comments--into your copy of this Jupyter Notebook, and we will walk through a series of data processing steps, with the goal of ending up with a complete and accurate set of data about all of those crowdworkers. In the process, we will discover and discuss several limitations of the dataset.
#
# ***In part 2 of the assignment,*** you will generate some basic descriptive statistics about the demographics of the crowdworkers described in that dataset. If you are comfortable in Python, you can perform that analysis in this Notebook. If you aren't as comfortable in Python (yet!), you can perform the analysis in Google Sheets (just be sure to link to that Google Sheet from this notebook, and set the permissions so that your instructors can view it!).
#
# ***In part 3 of the assignment,*** you will answer some additional research questions about this dataset. Some of these questions can be answered without writing additional code or analyzing additional data; other questions will require you to combine this dataset another dataset in the corpus. You will have the option to choose whether you want to answer code or no-code questions.
#
# Whether you choose code or no-code question for part 3, you will need to write your responses to the question within this notebook, and submit a link to this notebook for grading.
#
# Part 3 also contains some optional "challenge" questions that you can investigate if you want to (hint: any one of these challenge questions could be the focus of a final course project).
#
#
#
# ***Note:*** *Since the purpose of this class is to get you comfortable thinking critically (like a researcher), rather than programming perfectly, you won't be graded on your code. You'll be graded on how well you DOCUMENT YOUR PROCESS and REFLECT on the implications of your findings. If you still aren't sure what that means, ask your instructor or TA!*
# ## Part 1: Cleaning and analyzing the annotator demographic data
#
# For the first part of this assignment, we're going to prepare one set of Wikipedia Talk data--the annotator demographics files--for analysis. In the process we'll perform a few "sanity checks" to make sure we understand what the data means, and know any limitations.
#
# This sort of "[data wrangling](https://en.wikipedia.org/wiki/Data_wrangling)" is a critical, if sometimes tedious, first step for any quantitative research project.
# ### 1.1 Load the data into the notebook
#
# According to the documetation, the worker demographic data for the Wikipedia Talk Corpus is spread across three files:
# - ``toxicity_worker_demographics.tsv``
# - ``aggression_worker_demographics.tsv``
# - ``attack_worker_demographics.tsv``
#
# We will need to combine the data in these three files to come up with our canonical list of workers.
#
# First we'll load each of the annotator datafiles into our Notebook and save them into data structures that's easy to work with. In this case, I'm choosing to save each of these files as a list-of-dictionaries, since that's fairly standard, and it makes it easy to check your work as you go.
#
# By the way: ``.tsv`` stands for "tab-separated values", and it means that this file is organized into rows and columns, like a spreadsheet, and the data values for each column are separated by "tab" characters.
#import the csv module, a little code toolkit for working with spreadsheet-style data files
import csv
# The function below will load in a tab-separated (.tsv) file and convert it into a lists-of-dictionaries.
#
# If you don't have much experience with Python, this (and some of the other code in this notebook) might be hard to understand. That's okay! For now, it's most important that you know what it does.
#
# If you have a ***lot*** of experience with Python, the code in this notebook might seem really, really primitive. That's also okay! Remember: in this course we're primarily interested in data, not code. Code is just one of the many tools we use to ask and answer questions about data.
def prepare_datasets(file_path):
"""
Accepts: path to a tab-separated plaintext file
Returns: a list containing a dictionary for every row in the file,
with the file column headers as keys
"""
with open(file_path) as infile:
reader = csv.DictReader(infile, delimiter='\t')
list_of_dicts = [dict(r) for r in reader]
return list_of_dicts
# ### 1.2 Identifying duplicated datafiles
# Let's load our three .tsv files into Python and store them as three variables with relevant names, so that we know which is which.
#
# Once we've created these three lists-of-dicts, we will do two things to check our work so far:
# - we will print the first annotator's demographic data (list index ``[0]``) so that we know what the format looks like
# - we will print the length of each list (the ``len`` function), to see how many rows is in each file. Each row should correspond to one crowdworker/annotator.
#
# ***Note:*** for the cell below to run, your version of these datafiles and folders will need have the same names as the ones below, and your version of this Notebook will need to be stored in the same directory as the three folders that hold the datafiles.
# +
#load the data from the flat files into three lists-of-dictionaries
toxicity_annotators = prepare_datasets("Wikipedia_Talk_Labels_Toxicity_4563973/toxicity_worker_demographics.tsv")
print(toxicity_annotators[0])
print(len(toxicity_annotators))
attack_annotators = prepare_datasets("Wikipedia_Talk_Labels_Personal_Attacks_4054689/attack_worker_demographics.tsv")
print(attack_annotators[0])
print(len(attack_annotators))
aggression_annotators = prepare_datasets("Wikipedia_Talk_Labels_Personal_Attacks_4054689/attack_worker_demographics.tsv")
print(aggression_annotators[0])
print(len(aggression_annotators))
# -
# Interesting! This tells us a few things that we didn't know before:
# 1. it looks like the demographic data matches what's listed in [the schema](https://meta.wikimedia.org/wiki/Research:Detox/Data_Release#Schema_for_{attack/aggression/toxicity}_worker_demographics.tsv), which is great!
# 2. it looks like the "toxicity" dataset was annotated by a lot more people (3,591) than the "attack" or "aggression" datasets (2,190)
# 3. ``aggression_worker_demographics.tsv`` and ``attack_worker_demographics.tsv`` seem to contain the same number of workers, and the worker at the beginning of each list has the same ID and demographic data.
# Let's dig a little deeper into finding #3. Are the ***last*** entries in both of these lists also identical?
print(attack_annotators[-1]) # "-1" tells Python to find the last item in any list
print(aggression_annotators[-1])
# Yes, the last rows in these two lists are also identical!
#
# And in fact if you had opened the two lists in a text editor or spreadsheet program, you would find that the *aggression and attack .tsv files contain exactly the same data.* By the way, it doesn't say anywhere in the dataset documentation that these two files are identical!
#
# That brings us to our first lesson about bias: watch out for duplicate data!
#
# Consider: *What would have happened if we had just combined these three files and then analyzed the worker demographics? What mistaken conclusions might we have drawn from that?*
#
# Fortunately, now that we know that there is duplicate data we can work around it. Since two files are identical, we only need to use one of them. So from now on, we will ignore ``aggression_annotators`` entirely.
#
# Since want to remember that ``attack_annotators`` really refers to both "attack" and "aggression" annotators, we can just rename the variable we're using to store that dataset.
attack_aggro_annotators = attack_annotators
# Okay, that looks good. Now we can continue getting our data ready for analysis--while keeping an eye out for additional duplicate data and other "gotchas"!
# ### 1.3 Understanding what the properties of your data really mean
#
# Whenever you are working with data that you didn't create, it's very useful to perform some basic sanity-checking to make sure the data actually means what you think it means.
#
# For example, take the ``worker_id`` field in our datasets.
#
# The [schema](https://meta.wikimedia.org/wiki/Research:Detox/Data_Release#Schema_for_{attack/aggression/toxicity}_worker_demographics.tsv) says that the ``worker_id`` field contains an "anonymized crowd-worker id" and that this ID is meant to join the worker demographics datafiles with the annotator comments datafiles, so that if we wanted find all of the comments that worker "85" (from above) labelled, we could find her by looking for that ID in each row of ``toxicity_annotated_comments.tsv``.
#
# So far, so good. But since we want to combine the toxicity and attack + aggression annotator demographics data into a single dataset, we probably want to know...
#
# 1. did any of the values for ``worker_id`` appear in both datasets?
# 2. if so, do they correspond to the same person (or at least, a person with matching gender, age group, etc.)?
# ### 1.4 Checking for duplicate worker IDs
#
# If we want to see if worker_id means the same thing across datasets, the first step is to see if any of the same worker_ids exist across the two datasets.
#
# To check this, let's first pull all the worker IDs out of each dataset and combine them into a single list. Then we can check that list to see if it contains any duplicate values. If it does, we know that there is at least 1 value for ``worker_id`` that appears in both datasets.
#
# **Note:** *We're assuming that there are no duplicate values for ``worker_id`` within each dataset. (There aren't, I checked). It's a pretty safe assumption though, because worker_id is intended to be a unique key that links the ``worker_demographics.tsv`` files and the ``annotated_comments.tsv`` datasets. Can you explain why it would be an issue if there were duplicate values for ``worker_id`` within an individual dataset?*
# +
#pull the worker ids out of the individual files
tox_w_ids = [item['worker_id'] for item in toxicity_annotators]
aa_w_ids = [item['worker_id'] for item in attack_aggro_annotators]
#create a new list to hold all the ids
all_w_ids = tox_w_ids + aa_w_ids
# #combine the two worker_id lists into the new list
# all_w_ids.extend(tox_w_ids)
# all_w_ids.extend(aa_w_ids)
# -
#how many worker ids do we have, total?
#this number should match the total count of toxicity_annotators and attack_aggro_annotators (3591 + 2190 = 5781)
print(len(all_w_ids))
#what does our new list look like? Let's print the first five values
print(all_w_ids[0:5])
# Below is our duplicate-checker function. You pass it a list of values, and it will return "True" if it finds at least 1 duplicate value in that list. Can you figure out how it works?
def determine_dupes(w_ids):
set_of_ids = set()
found_a_dupe = False
for w in w_ids:
if w in set_of_ids:
found_a_dupe = True
else:
set_of_ids.add(w)
return found_a_dupe
# +
#we'll call determine_dupes to check for duplicates in the combined worker id list
has_dupes = determine_dupes(all_w_ids)
#if this prints 'True', that means we found at least one duplicate ID
print(has_dupes)
# -
# Hmmm... So it looks like there's at least 1 duplicate! Okay, we'll need to do some additional verification before we can decide what to do with that information.
#
# The next thing we'll do is check how many duplicates there are. We'll write a short script that reads through ``all_w_ids`` and every time it finds a value that appears more than once, it adds that value to a new list.
#
# Can you figure out how the script below works?
# +
#create an empty list to hold any duplicate worker_id values we find
dupes = []
#look through the data, if you encounter any value more than once, add it to our 'dupes' list
for w in all_w_ids:
if all_w_ids.count(w) > 1:
dupes.append(w)
#how many values were added to 'dupes'?
print(len(dupes))
#how many worker_ids are present twice in the dataset?
#can you explain why we are dividing the length of "dupes" by 2 in order to answer that question?
print(len(dupes)/2)
# -
# Huh, so it looks like 1,850 of the ``worker_ids`` in our merged dataset are duplicates! That's a lot of duplication, since our list was only 5,781 rows in the first place, including these dupes!
#
# We will definitely need to account for these duplicates before we start analyzing worker demographics.
# - If the duplicate ``worker_id``s have different demographic metadata, we will assume they are different people with the same ID.
# - If the metadata is identical, then we know we can safely remove the duplicates
# - If some metadata is identical and some is different... well, let's hope that's not the case!
# ### 1.5 Check for duplicate worker demographic metadata
# Let's see if it's just the value for ``worker_id`` that is duplicated across the two datasets (meaning that the these duplicate ids correspond to different workers with different demographics), or if ``worker_id`` really corresponds to the same people across ``toxicity_annotators`` and ``attack_annotators``.
# To do this we'll perform some spot checks, meaning we'll visually compare the demographic data of workers with duplicate IDs, to see if they look like the same worker, or not. Running random 'spot checks' is a really common and useful way of finding systematic issues with your data without having to check every entry.
#
#
# First, we'll extract 10 random ids from our dupe set. Using a random sample (rather than just looking at the first 10 rows, for instance) helps us be more confident that any patterns we see are really there.
#handy Python library that lets you select things randomly from a list
import random
# +
#convert our 'dupes' list into a set
#bonus: can you explain why we created "dupeset" rather than just grabbing 10 random values from "dupes"?
#hint: in Python, a 'set' is like a list that can only contain a single instance of any value
dupeset = set(dupes)
#store our random sample of dupes in its own list
dupe_sample = random.sample(dupeset, 10)
#print to confirm everything looks how we expect it to...
print(dupe_sample)
# -
# Now, we'll use this ``dupe_sample`` list that we created to pull the corresponding worker demographics from each of our two datasets, using the function below. See if you can figure out how the function works!
# +
def worker_id_lookup(annotator_list, dupe_id):
"""
Accepts: a list of dictionaries
& a list of known duplicate values for
the key 'worker_id' in those dictionaries
If a duplicate value for worker_id is found,
print the complete dictionary of demographic data for that worker
"""
for a in annotator_list:
if a['worker_id'] == dupe_id:
print(a)
#loop through the duplicate sample list and call our worker_id_lookup function
#to check each dataset for corresponding worker demographic data
for d in dupe_sample:
worker_id_lookup(toxicity_annotators, d)
worker_id_lookup(attack_aggro_annotators, d)
# -
# Aha! It looks like few if any of these ``worker_id`` values correspond to the same demographic data info across the two datasets.
#
# It's possible that some of the same people labelled both datasets and were assigned different IDs each time. But there's no way for us to determine that with the data we have.
#
# So for now we will assume that these datasets were labelled by two entirely different sets of crowdworkers. Which means we can combine these two datasets into one, with no duplication to mess up our analysis.
# ### 1.6 Final preparation of the dataset
#
# Before we start our analysis of worker demographics, let's do two more things:
#
# 1. Since we know now that worker ID isn't unique, let's give each worker in our new, combined dataset a **truly** unique ID.
# 2. While we're at it, let's also add a new field to each worker's demographic dictionary that lists which dataset the worker worked on (toxicity or attack/aggressive).
def add_dataset_id(list_of_dicts, dataset_ref):
"""
Accepts: a list of dictionaries & a strig value for "dataset"
Returns: that list of dictionaries, with the new "dataset" key and the specified value
"""
for w in list_of_dicts:
w.update({"dataset" : dataset_ref})
return list_of_dicts
#update our worker demographic datasets with the new key and value
toxicity_annotators = add_dataset_id(toxicity_annotators, "toxicity")
attack_aggro_annotators = add_dataset_id(attack_aggro_annotators, "attack and aggression")
#did it work?
print(toxicity_annotators[0])
print(attack_aggro_annotators[0])
# Great. Now we'll use the function below to...
#
# 1. combine the two datasets into one
# 2. assign a truly unique id to each worker
#
# It doesn't matter what this ID is, as long as its unique within the dataset. We'll use sequential IDs, starting at "1" and going up from there.
def finalize_dataset(list1, list2):
#combine the two lists into a new one
combined_list = list1 + list2
#initialize our counter variable
new_id = 1
for w in combined_list:
#add the new sequential id field, and populate with the current value of 'counter'
w.update({"unique_id" : str(new_id)})
#increment the counter variable by 1, so that the next ID will be one number higher
new_id = new_id + 1
return combined_list
# +
#create our new combined list with sequential IDs
all_annotators = finalize_dataset(toxicity_annotators, attack_aggro_annotators)
#print the list length and a sample, for sanity-checking purposes
print(len(all_annotators))
print(all_annotators[0])
# -
# It worked! Now we can FINALLY start to analyze this dataset, and find out more about the workers who labelled the Wikipedia Talk Corpus.
# ## Part 2: Analyzing worker demographics
# Now that we have our worker demographic data de-duplicated, combined and uniquely identified, we can start using it to ask and answer research questions!
#
# You are welcome to do this next steps in Python, here in this notebook. But if you aren't super comfortable with Python, you can run the cell below to export this dataset to a .csv file called ``a2_all_annotator_demographics.csv`` which you can open in Google Sheets and do the analysis there.
#
# Note: if you choose Google Sheets, please title your Google Sheet "A2 worker demographics", share it with your instructor and TA ("can view" or "can edit") and paste a link to it in a Markdown cell below.
with open('a2_all_annotator_demographics.csv', 'w', encoding='utf-8') as f:
writer = csv.writer(f)
#write a header row
writer.writerow(('unique_id',
'worker_id',
'gender',
'english_first_language',
'age_group',
'education',
'dataset',))
#loop through our dataset and write it to the file, row by row
for a in all_annotators:
writer.writerow((a['unique_id'], a['worker_id'], a['gender'], a['english_first_language'], a['age_group'], a['education'], a['dataset']))
# ### 2.1 Questions 1-4: compute descriptive statistics
#
# Please create tables or bar charts that answer the following questions:
#
# - **Q1.** What is the gender distribution of these workers?
# - **Q2.** What is the distribution of the workers by first language?
# - **Q3.** What is the distribution of these workers by age group?
# - **Q4.** what is the distribution of these workers by education level?
# *POST YOUR ANSWERS TO QUESTIONS 1-4 HERE.*
#
# Use both Markdown and code-formatted cells.
# ### 2.2 Questions 5-8
#
# Each of the next four questions requires a different kinds of answer.
#
# Questions 5 and 6 have a "correct" answer, but we'll still give you credit for the "incorrect" answer as long as you document your process well (in Markdown, in this notebook). These questions are intended to get you thinking about how even small errors in data processing can create bias in your data.
#
# Question 6 requires you to re-run some (but not all) of the data processing steps you performed above. Do this by copying the code into NEW cells below this line, and make sure to document each step using Markdown cells or inline (#) comments. You will probably want to change the names of the variables too.
#
# - **Q5.** Analyze all_annotators in Sheets or Jupyter: Do any of the fields in this dataset have missing data? If so, which fields, and what % of the rows contain missing values?
# - **Q6.** Build a version of the all_annotators dataset *without removing duplicates*, then re-run the summary statistics from questions 1-4. How have these summary statistics changed?
#
# Questions 7 and 8 don't have one "correct" answer. These questions are intended to get you thinking about how using this dataset to train a machine learning model might lead to bias in the way that model performs when used as intended. You will need to provide a written answer to each of these questions in 3-4 full sentences. Consider what you know about the workers themselves, the task they were given, and about the purpose of the Perspective API.
#
# Question 7 requires you to do some research of your own (hint: use Google). Question 8 doesn't require any new analysis or research, just critical thinking.
#
# - **Q7.** In general, how do the demographics of these workers compare to those of English-speaking internet users overall? Why would it matter if the worker demographics don't match the demographics of the intended end users of the Perspective API?
#
# - **Q8.** Given what you now know about the Wikipedia Talk Corpus, what issues might arise if it was used to train a machine learning-driven "hostile speech detector" that could be used on any website or social media platform?
#
# *POST YOUR ANSWERS TO QUESTIONS 5-8 HERE.*
#
# Use both Markdown and code-formatted cells.
# ## Part 3: Digging deeper
#
# ### 3.1 Questions 9-16
#
# Below is a list of additional questions that you should be able to answer, based on what you've done today.
#
# ***You may answer EITHER the "code" questions (Q9-Q12) OR the "no code" questions (Q13-Q16).***
#
# If you don't feel very comfortable with Python yet, choose the "no code" questions.
#
# - ***For the "code" questions*** you will need to load ``toxicity_annotations.tsv`` into Python and join that dataset with ``all_annotators`` on ``worker_id``.
# - ***For the "no code" questions*** you will need to load the file ``toxicity_labelled_comments_3k_sample.csv`` into Google Sheets. This file contains the text of 3000 comments that were annotated for toxicity, as well as the toxicity score that each worker gave to these comments. The file is a random sample taken from the ~1.5 million comments contained in ``toxicity_annotated_comments.tsv``
#
# *Note:* answering question 13-16 below ***will require reading some comments that contain offensive speech!*** If you do not want to be exposed to offensive speech, either answer the "code" questions instead, or reach out to your Instructor or TA and ask for an alternate activity.
#
# #### Code questions
# - **Q9.** What % of *workers* who labelled the toxicity dataset do we have demographic data for?
# - **Q10.** What % of the *comments* in the toxicity dataset were labelled by male vs. female-identified workers?
# - **Q11.** What % of the *comments* in the toxicity dataset were labelled by people for whom English is NOT their first language?
# - **Q12.** Based on your findings from questions 9-11, how might this dataset present a biased view of "toxicity"? How would you expect such biases to impact how the Perspective API performs?
#
# #### No-code questions
#
# Before you answer the questions below, read through at least 20 comments from ``toxicity_labelled_comments_3k_sample.csv`` that were labelled "toxic" (-1 or -2), 20 that were labelled "non-toxic" (1 or 2), and 20 that are labelled "neutral" (0). Note in your spreadsheet whether you agree or disagree with the labeller's judgement.
#
#
# - **Q13.** Pick 2-3 examples of comments where you disagreed with the labeller about the toxicity of a comment. Why did you disagree? Why do you think that the labeller might have labelled these comments the way they did?
# - **Q14.** Pick 2-3 examples of comments where you don't understand what the commenter was saying, and therefore had a hard time classifying as toxic or non-toxic. What additional context or information would you need in order to be confident in your judgement about the toxicity of this comment?
# - **Q15.** Read through the [instructions and labelling options](https://github.com/ewulczyn/wiki-detox/blob/master/src/modeling/toxicity_question.png) given to the crowdworkers. How could the way these instructions were written have made it difficult for crowdworkers to accurately label comments as toxic or non-toxic? If you were going to run a labelling campaign like this one yourself, how would you change these instructions or labelling options to help the crowdworkers make more accurate or consistent judgements about toxicity?
# - **Q16.** Based on your findings from questions 13-15, how might this dataset present a biased view of "toxicity"? How would you expect such biases to impact how the Perspective API performs as a general-purpose hostile speech detector?
# *POST YOUR ANSWERS TO QUESTIONS 9-16 HERE.*
#
# Use both Markdown and code-formatted cells.
# ### 3.2 Challenge questions (optional)
#
# The questions below are optional; you don't need to answer them to receive full credit for this assignment.
#
# If you'd like to explore additional analyses of this dataset, here are a few questions to get you started!
#
# - ***Challenge #1.*** Are female-identified workers more likely to label a comment as "toxic" than male identified workers?
# - ***Challenge #2.*** Are workers with a higher level of education more consistent in their "toxicity" labelling--in other words, do they tend to agree with other labellers more often? (remember that, according to the documentation, every comment was labelled by at least 10 crowdworkers).
# - ***Challenge #3.*** What are the words most frequently associated with toxic comments? (perhaps focus on comments where most or all of the workers agree are toxic)
# - ***Challenge #4.*** What are the most polarizing comments--comments that some workers consider toxic, and others consider non-toxic? What about these comments made them so polarizing, or hard to classify?
# - ***Challenge #5.*** Answer some or all of the homework questions above for one the other datasets ("attack" and "aggressive"). How are the potential sources of bias for these datasets the same, or different, than for the toxicity dataset? Would you "trust" a machine learning model based on these datasets more or less than one trained on the "toxicity" dataset? Why?
# *OPTIONAL: POST YOUR ANSWERS TO THE CHALLENGE QUESTIONS HERE.*
#
# Use both Markdown and code-formatted cells.
| hcde410-a2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''tf1.13'': conda)'
# name: python3710jvsc74a57bd0c1abdd2cb8e2bb7bc61f0f837085c52fe48b926da48959e95678065ce71dcf54
# ---
# +
#-*- encoding: utf-8 -*-
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import argparse
import os.path
import sys
import numpy as np
import tensorflow as tf
import input_data
import models
# +
def data_stats(train_data, val_data, test_data):
"""mean and std_dev
Args:
train_data: (36923, 490)
val_data: (4445, 490)
test_data: (4890, 490)
Return: (mean, std_dev)
Result:
mean: -3.975149608704592, 220.81257374779565
std_dev: 0.8934739293234528
"""
print(train_data.shape, val_data.shape, test_data.shape)
all_data = np.concatenate((train_data, val_data, test_data), axis=0)
std_dev = 255. / (all_data.max() - all_data.min())
# mean_ = all_data.mean()
mean_ = 255. * all_data.min() / (all_data.min() - all_data.max())
return (mean_, std_dev)
def fp32_to_uint8(r):
# method 1
# s = (r.max() - r.min()) / 255.
# z = 255. - r.max() / s
# q = r / s + z
# method 2
std_dev = 0.8934739293234528
mean_ = 220.81257374779565
q = r / std_dev + mean_
q = q.astype(np.uint8)
return q
def calc(interpreter, input_data, label):
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
# print(input_details)
# print(output_details)
# Test model on random input data.
# input_shape = input_details[0]['shape']
# input_data = np.array(np.random.random_sample(input_shape), dtype=np.float32)
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
# print(output_data)
# print(label)
return output_data
# -
wanted_words = 'yes,no,up,down,left,right,on,off,stop,go'
sample_rate = 16000
clip_duration_ms = 1000
model_architecture = 'mobilenet-v3'
dct_coefficient_count = 10
batch_size = 1
window_size_ms = 40
window_stride_ms = 20
model_size_info = [4, 16, 10, 4, 2, 2, 16, 3, 3, 1, 1, 2, 32, 3, 3, 1, 1, 2, 32, 5, 5, 1, 1, 2]
data_url = 'http://download.tensorflow.org/data/speech_commands_v0.02.tar.gz'
data_dir = '/tmp/speech_dataset/'
silence_percentage = 10.0
unknown_percentage = 10.0
testing_percentage = 10
validation_percentage = 10
# +
tf.logging.set_verbosity(tf.logging.INFO)
sess = tf.InteractiveSession()
words_list = input_data.prepare_words_list(wanted_words.split(','))
model_settings = models.prepare_model_settings(
len(words_list), sample_rate, clip_duration_ms, window_size_ms,
window_stride_ms, dct_coefficient_count)
audio_processor = input_data.AudioProcessor(
data_url, data_dir, silence_percentage,
unknown_percentage,
wanted_words.split(','), validation_percentage,
testing_percentage, model_settings)
label_count = model_settings['label_count']
fingerprint_size = model_settings['fingerprint_size']
fingerprint_input = tf.placeholder(
tf.float32, [None, fingerprint_size], name='fingerprint_input')
logits = models.create_model(
fingerprint_input,
model_settings,
model_architecture,
model_size_info,
is_training=False)
ground_truth_input = tf.placeholder(
tf.float32, [None, label_count], name='groundtruth_input')
predicted_indices = tf.argmax(logits, 1)
expected_indices = tf.argmax(ground_truth_input, 1)
correct_prediction = tf.equal(predicted_indices, expected_indices)
confusion_matrix = tf.confusion_matrix(
expected_indices, predicted_indices, num_classes=label_count)
evaluation_step = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# +
from decimal import Decimal
class Transformer_bd(object):
def __init__(self, precision):
self.pre = precision
self.shift_num = None
def bTod(self, n):
'''
把一个带小数的二进制数n转换成十进制
小数点后面保留pre位小数
'''
string_number1 = str(n) #number1 表示二进制数,number2表示十进制数
decimal = 0 #小数部分化成二进制后的值
flag = False
for i in string_number1: #判断是否含小数部分
if i == '.':
flag = True
break
if flag: #若二进制数含有小数部分
string_integer, string_decimal = string_number1.split('.') #分离整数部分和小数部分
for i in range(len(string_decimal)):
decimal += 2**(-i-1)*int(string_decimal[i]) #小数部分化成二进制
number2 = int(str(int(string_integer, 2))) + decimal
return round(number2, self.pre)
else: #若二进制数只有整数部分
return int(string_number1, 2)#若只有整数部分 直接一行代码二进制转十进制 python还是骚
def dTob(self, n, shift=False):
'''
把十进制的浮点数n转换成二进制
小数点后面保留pre位小数
'''
string_number1 = str(n) #number1 表示十进制数,number2表示二进制数
flag = False
for i in string_number1: #判断是否含小数部分
if i == '.':
flag = True
break
if flag:
string_integer, string_decimal = string_number1.split('.') #分离整数部分和小数部分
integer = int(string_integer)
decimal = Decimal(str(n)) - integer
l1 = [0,1]
l2 = []
decimal_convert = ""
while True:
if integer == 0: break
x,y = divmod(integer, 2) #x为商,y为余数
l2.append(y)
integer = x
string_integer = ''.join([str(j) for j in l2[::-1]]) #整数部分转换成二进制
i = 0
while decimal != 0 and i < self.pre:
result = int(decimal * 2)
decimal = decimal * 2 - result
decimal_convert = decimal_convert + str(result)
i = i + 1
string_number2 = string_integer + '.' + decimal_convert
# return float(string_number2)
# 这里进行移位修改
if string_integer == '':
string_integer = '0'
if shift == True:
string_number3 = decimal_convert.lstrip('0')
lshift_num = len(decimal_convert) - len(string_number3) # 左移的位数
self.shift_num = lshift_num
return string_integer + '.' + string_number3
else:
return string_integer + '.' + decimal_convert
else: #若十进制只有整数部分
l1 = [0,1]
l2 = []
while True:
if n == 0: break
x,y = divmod(n, 2) #x为商,y为余数
l2.append(y)
n = x
string_number = ''.join([str(j) for j in l2[::-1]])
# return int(string_number)
return string_number
def right_shift(self, s):
'''右移二进制字符串'''
point_pos = s.find('.') # point_pos的值就说明小数点前面有多少个数字
ss = s[:point_pos] + s[point_pos + 1:]
if self.shift_num > point_pos:
zero_fill = self.shift_num - point_pos # 需要补0的个数
for _ in range(zero_fill):
ss = '0' + ss
return '0.' + ss
else:
insert_pos = point_pos - self.shift_num
ssl = list(ss)
ssl.insert(insert_pos, '.')
ss = ''.join(ssl)
if insert_pos == 0:
ss = '0' + ss # 在头部补零
return ss
def lshift_decimal(self, num):
'''得到左移后的十进制'''
num = self.dTob(num, shift=True)
num = self.bTod(num)
return num
def rshift_decimal(self, num):
'''得到右移后的十进制'''
num = self.dTob(num, shift=False)
num = self.right_shift(num)
num = self.bTod(num)
return num
# +
a = 150
num = 0.001
print('a * b =', a*num)
b = Transformer_bd(32)
num2 = b.lshift_decimal(num)
print(num2)
print('before right shift, a * num2 =', a*num2)
print()
aft_shift = b.rshift_decimal(num2)
print(aft_shift)
num_aft = a * aft_shift
print('after right shift, a * b =', num_aft)
# +
# tflite_path = 'test_log/mobilenetv3_quant_gen/symmetric_8bit_mean220_std0.89.lite'
tflite_path = 'test_log/mobilenetv3_quant_gen/layers_lite_model/inverted_residual_1_depthwise.lite'
# tflite_path = 'test_log/mobilenetv3_quant_gen/layers_lite_model/inverted_residual_3_depthwise.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/stem_conv.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_1_expansion.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_1_depthwise.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_1_projection.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_1_add.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_2_expansion.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_2_depthwise.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_2_projection.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_3_expansion.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_3_depthwise.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_3_projection.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/inverted_residual_3_add.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/AvgPool.lite'
# tflite_path = 'test_log/mobilenetv3_quant_eval/layers_lite_model/Conv2D.lite'
# Boss
# tflite_path = 'test_log/mobilenetv3_quant_eval/uint8input_8bit_calc_mean220_std0.89.lite'
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path=tflite_path)
# interpreter = tf.lite.Interpreter(model_path="tflite_factory/swiftnet-uint8.lite")
interpreter.allocate_tensors()
# -
def manual_int(input_data):
################## stem conv ##################
# print('stem conv')
new_data = input_data.reshape(-1, 49, 10, 1)
new_data = tf.convert_to_tensor(new_data, tf.float32, name='input')
new_data = new_data - 221.
s_iwr = tf.constant(0.0011018913937732577 / 0.16148914396762848, tf.float32)
s_iwr = tf.cast(s_iwr, tf.float32)
weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_stem_conv_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_stem_conv_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
weight = tf.convert_to_tensor(weight, tf.float32, name='weight')
weight = weight - 132.
weight = tf.transpose(weight, perm=[1,2,0,3])
bias = tf.convert_to_tensor(bias, tf.float32, name='bias')
# print(weight)
# print(bias)
output = tf.nn.depthwise_conv2d(new_data, # 张量输入
filter=weight, # 卷积核参数
strides=[1,2,2,1], # 步长参数
padding="SAME", # 卷积方式
data_format=None, # 数据格式,与步长参数配合,决定移动方式
name='stem_conv') # 名字,用于tensorboard图形显示时使用
output = tf.add(output, bias, name='add')
output *= s_iwr
output += 0.0035
output = tf.nn.relu(output)
output_uint8 = tf.math.round(output, name='round')
output_uint8 = tf.cast(output_uint8, tf.uint8, name='uint8')
add_2 = tf.identity(output_uint8) # 给之后的做加法
# print()
# ################## inverted residual 1 expansion ##################
# print('inverted residual 1 expansion')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.003049441846087575 / 0.27361148595809937, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_expansion_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_expansion_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 146
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.0074
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 1 depthwise ##################
# print('inverted residual 1 depthwise')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.0045695919543504715 / 0.12676289677619934, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_depthwise_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_depthwise_depthwise_conv_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 127
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.depthwise_conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.0301
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 1 projection ##################
# print('inverted residual 1 projection')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.0009397256653755903 / 0.16901935636997223, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_projection_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_1_projection_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 101
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.00052
# output = output * s_iwr + 133
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# add_1 = tf.identity(output_uint8)
# print()
# ################## inverted residual 1 add ##################
# add_1 = tf.cast(add_1, tf.float32)
# add_2 = tf.cast(add_2, tf.float32)
# add_1 = tf.constant(0.16901935636997223, tf.float32) * (add_1 - 133)
# add_2 = tf.constant(0.16148914396762848, tf.float32) * add_2
# output_result = tf.add(add_1, add_2)
# output = output_result / tf.constant(0.24699252843856812, tf.float32) + 89
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# ################## inverted residual 2 expansion ##################
# print('inverted residual 2 expansion')
# new_data = tf.cast(output_uint8, tf.float32)
# new_data -= 89
# s_iwr = tf.constant(0.0020657109562307596 / 0.09814818948507309, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_expansion_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_expansion_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 149
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.01062
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 2 depthwise ##################
# print('inverted residual 2 depthwise')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.0014443636173382401 / 0.062810979783535, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_depthwise_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_depthwise_depthwise_conv_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 120
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.depthwise_conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.0153
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 2 projection ##################
# print('inverted residual 2 projection')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.00040918667218647897 / 0.0929793044924736, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_projection_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_2_projection_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 148
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# output = output * s_iwr + 138
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# add_2 = tf.identity(output_uint8)
# print()
# ################## inverted residual 3 expansion ##################
# print('inverted residual 3 expansion')
# new_data = tf.cast(output_uint8, tf.float32)
# new_data -= 138
# s_iwr = tf.constant(0.0005567758926190436 / 0.07842949777841568, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_expansion_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_expansion_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 137
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# # output += 0.00113
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 3 depthwise ##################
# print('inverted residual 3 depthwise')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.013642110861837864 / 0.05131378769874573, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_depthwise_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_depthwise_depthwise_conv_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 79
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.depthwise_conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# output *= s_iwr
# output = tf.nn.relu(output)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
# ################## inverted residual 3 projection ##################
# print('inverted residual 3 projection')
# new_data = tf.cast(output_uint8, tf.float32)
# s_iwr = tf.constant(0.0008600406581535935 / 0.20826007425785065, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_projection_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_inverted_residual_3_projection_conv_Conv2D_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 125
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# output = output * s_iwr + 133
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# add_1 = tf.identity(output_uint8)
# print()
# ################## inverted residual 3 add ##################
# add_1 = tf.cast(add_1, tf.float32)
# add_2 = tf.cast(add_2, tf.float32)
# add_1 = tf.constant(0.20826007425785065, tf.float32) * (add_1 - 133)
# add_2 = tf.constant(0.0929793044924736, tf.float32) * (add_2 - 138)
# output_result = tf.add(add_1, add_2)
# output_uint8 = output_result / tf.constant(0.21021947264671326, tf.float32) + 131
# output_uint8 = tf.math.round(output_uint8)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# ################## AvgPool ##################
# # method 1
# # new_data = tf.cast(output_uint8, tf.float32)
# # new_data = 0.21021947264671326 * (new_data - 131)
# # output = tf.nn.avg_pool(new_data,
# # ksize=[1,25,5,1],
# # strides=[1,25,5,1],
# # padding='VALID')
# # output = output / 0.21021947264671326 + 131
# # output_uint8 = tf.math.round(output)
# # output_uint8 = tf.cast(output, tf.uint8)
# # method 2 (简化版本,发现scale和zero_point完全可以消除)
# new_data = tf.cast(output_uint8, tf.float32)
# output = tf.nn.avg_pool(new_data,
# ksize=[1,25,5,1],
# strides=[1,25,5,1],
# padding='VALID')
# # output -= 0.0041
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# ################## Conv2D ##################
# print('Conv2D')
# new_data = tf.cast(output_uint8, tf.float32)
# new_data -= 131
# s_iwr = tf.constant(0.0033858332317322493 / 0.1784215271472931, tf.float16)
# s_iwr = tf.cast(s_iwr, tf.float32)
# weight = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_fc_conv_weights_quant_FakeQuantWithMinMaxVars.npy')
# bias = np.load('test_log/mobilenetv3_quant_eval/weight/MBNetV3-CNN_fc_conv_Conv2D_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
# weight = tf.convert_to_tensor(weight, tf.float32)
# weight -= 143
# weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
# bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
# output = tf.nn.conv2d(new_data, # 张量输入
# filter=weight, # 卷积核参数
# strides=[1,1,1,1], # 步长参数
# padding="SAME", # 卷积方式
# data_format=None) # 数据格式,与步长参数配合,决定移动方式
# output = output + bias
# output = output * s_iwr + 129
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
# add_1 = tf.identity(output_uint8)
# print()
# ################## Reshape ##################
# output_uint8 = tf.squeeze(output_uint8, axis=[1,2])
# ################## Softmax ##################
# new_data = tf.cast(output_uint8, tf.float32)
# new_data = tf.constant(0.1784215271472931, tf.float32) * (new_data - 129)
# output = tf.nn.softmax(new_data)
# output = output / tf.constant(0.00390625, tf.float32)
# output_uint8 = tf.math.round(output)
# output_uint8 = tf.cast(output_uint8, tf.uint8)
################## running ##################
return output_uint8.eval(), output.eval()
def debug(output_uint8, add_2):
add_2 = np.load('test_log/mobilenetv3_quant_eval/debug/add_2.npy')
################## inverted residual 1 add ##################
add_1 = tf.cast(output_uint8, tf.float32)
add_2 = tf.convert_to_tensor(add_2, tf.float32)
add_1_scale = tf.constant(0.16901935636997223, tf.float32)
add_2_scale = tf.constant(0.16148914396762848, tf.float32)
add_1 = add_1_scale * (add_1 - 133)
add_2 = add_2_scale * add_2
# low = tf.ones_like(add_1) * -22.47957420349121
# high = tf.ones_like(add_1) * 20.620361328125
# add_1 = tf.where(add_1 < -22.47957420349121, low, add_1)
# add_1 = tf.where(add_1 > 20.620361328125, high, add_1)
# high = tf.ones_like(add_2) * 20.620361328125
# add_2 = tf.where(add_2 > 20.620361328125, high, add_2)
output_result = tf.math.add(add_1, add_2)
output_scale = tf.constant(0.24699252843856812, tf.float32)
output = output_result / output_scale + 89
output_uint8 = tf.math.round(output)
# output_uint8 = tf.math.floor(output)
output_uint8 = tf.cast(output_uint8, tf.uint8)
return output_uint8.eval(), output.eval()
def manual_real(input_data, add_2):
################## inverted residual 1 depthwise ##################
# print('inverted residual 1 depthwise')
new_data = tf.cast(input_data, tf.float32)
new_data -= 128
s_iwr = tf.constant(0.00785899069160223 / 0.23841151595115662, tf.float32)
s_iwr = tf.cast(s_iwr, tf.float32)
weight = np.load('test_log/mobilenetv3_quant_gen/weight/MBNetV3-CNN_inverted_residual_1_depthwise_weights_quant_FakeQuantWithMinMaxVars.npy')
bias = np.load('test_log/mobilenetv3_quant_gen/weight/MBNetV3-CNN_inverted_residual_1_depthwise_depthwise_conv_Fold_bias.npy')
# print(weight.dtype, weight.shape)
# print(bias.dtype, bias.shape)
weight = tf.convert_to_tensor(weight, tf.float32)
weight -= 128
weight = tf.transpose(weight, perm=[1,2,3,0])
# print(weight)
bias = tf.convert_to_tensor(bias, tf.float32)
# print(bias)
output = tf.nn.depthwise_conv2d(new_data, # 张量输入
filter=weight, # 卷积核参数
strides=[1,1,1,1], # 步长参数
padding="SAME", # 卷积方式
data_format=None) # 数据格式,与步长参数配合,决定移动方式
output = output + bias
# output += 0.0301
output = output * s_iwr
output = tf.nn.relu(output)
output += 128
output_uint8 = tf.math.round(output)
mask = tf.ones_like(output_uint8) * 255
output_uint8 = tf.where(output_uint8 > 255, mask, output_uint8)
output_uint8 = tf.cast(output_uint8, tf.uint8)
# print()
return output_uint8.eval(), output.eval()
# + tags=[]
# test set
set_size = audio_processor.set_size('testing')
tf.logging.info('set_size=%d', set_size)
total_accuracy = 0
for i in range(0, set_size, batch_size):
test_fingerprints, test_ground_truth = audio_processor.get_data(
batch_size, i, model_settings, 0.0, 0.0, 0, 'testing', sess)
# print(test_fingerprints.shape) # (batch_size 490)
# print(test_ground_truth.shape) # (batch_size, 12)
test_fingerprints = fp32_to_uint8(test_fingerprints)
# output_simulate, output_ = manual_int(test_fingerprints)
# Load TFLite model and allocate tensors.
interpreter2 = tf.lite.Interpreter(model_path='test_log/mobilenetv3_quant_gen/layers_lite_model/inverted_residual_1_expansion.lite')
# interpreter = tf.lite.Interpreter(model_path="tflite_factory/swiftnet-uint8.lite")
interpreter2.allocate_tensors()
# Load TFLite model and allocate tensors.
interpreter3 = tf.lite.Interpreter(model_path='test_log/mobilenetv3_quant_gen/layers_lite_model/inverted_residual_2_projection.lite')
# interpreter = tf.lite.Interpreter(model_path="tflite_factory/swiftnet-uint8.lite")
interpreter3.allocate_tensors()
output_uint8 = calc(interpreter2, test_fingerprints, test_ground_truth)
add_2 = calc(interpreter3, test_fingerprints, test_ground_truth)
output_simulate, output_ = manual_real(output_uint8, add_2)
output_real = calc(interpreter, test_fingerprints, test_ground_truth)
# output_simulate, output_ = debug(output_real)
# real_output
# np.save('test_log/mobilenetv3_quant_eval/debug/add_2.npy', output_real)
# np.save('test_log/mobilenetv3_quant_eval/debug/output_real.npy', output_real)
# output_real = np.load('test_log/mobilenetv3_quant_eval/debug/output_real.npy')
# print(test_fingerprints.shape)
# print(output_simulate.shape)
# print(output_real.shape)
# print(output_simulate)
# print(output_simulate.max(), output_real.max())
# print(output_simulate.min(), output_real.min())
# print(np.count_nonzero(output_real == 128))
# print(np.count_nonzero(output_simulate <= 128))
# print(output_real)
# print(test_ground_truth)
sys.exit(0)
neq = output_simulate != output_real
print(output_[neq])
print(output_real[neq])
print('add', sorted(output_real[neq] - output_[neq]))
# print('sub', sorted(output_[neq] - output_real[neq]))
eq = tf.equal(output_real, output_simulate)
mask = tf.cast(tf.zeros_like(eq), tf.bool)
neq = tf.reduce_sum(tf.cast(tf.equal(eq, mask), tf.int32))
print(sess.run(neq), '/', sess.run(tf.size(eq)))
# sys.exit(0)
'''
############################### get all data mean and std_dev ###############################
training_fingerprints, training_ground_truth = audio_processor.get_data(
-1, 0, model_settings, 0.0, 0.0, 0, 'training', sess)
validation_fingerprints, validation_ground_truth = audio_processor.get_data(
-1, 0, model_settings, 0.0, 0.0, 0, 'validation', sess)
testing_fingerprints, testing_ground_truth = audio_processor.get_data(
-1, 0, model_settings, 0.0, 0.0, 0, 'testing', sess)
mean_, std_dev = data_stats(training_fingerprints, validation_fingerprints, testing_fingerprints)
print(mean_, std_dev)
'''
# +
inputs = np.ones((1,3,3,2))
inputs = tf.convert_to_tensor(inputs, tf.float32)
print(inputs)
weights = np.ones((2,2,2,3))
weights = weights.transpose(3,0,1,2)
weights[0,:,:,:] *= 1
weights[1,:,:,:] *= 2
weights[2,:,:,:] *= 3
weights = weights.transpose(1,2,3,0)
weights = tf.convert_to_tensor(weights, tf.float32)
weights = tf.transpose(weights, perm=[1,2,0,3])
print(weights)
output = tf.nn.depthwise_conv2d(inputs, # 张量输入
filter=weights, # 卷积核参数
strides=[1,1,1,1], # 步长参数
padding="SAME", # 卷积方式
data_format=None) # 数据格式,与步长参数配合,决定移动方式
a = output.eval()
print(a.shape)
print(a.transpose(0,3,1,2))
# +
import numpy as np
new_data = np.random.rand(1,29,9,64)
weight = np.random.rand(5,5,64,1)
new_data = tf.convert_to_tensor(new_data, tf.float32)
weight = tf.convert_to_tensor(weight, tf.float32)
output = tf.nn.depthwise_conv2d(new_data, # 张量输入
filter=weight, # 卷积核参数
strides=[1,1,1,1], # 步长参数
padding="VALID", # 卷积方式
data_format=None, # 数据格式,与步长参数配合,决定移动方式
name='stem_conv') # 名字,用于tensorboard图形显示时使用
with tf.Session().as_default():
print(output.eval().shape)
# +
import numpy as np
def decimal2md(x, d):
m = int(np.round(x * d))
return(m, d)
bias_scale = [0.0008852639002725482, 0.0035931775346398354, 0.00785899069160223, 0.0014689048985019326, 0.0015524440677836537, 0.0028435662388801575, 0.001141879241913557, 0.0007087105768732727, 0.009289528243243694, 0.0015117411967366934, 0.004092711955308914]
result_sacale = [0.20100615918636322, 0.42823609709739685, 0.23841151595115662, 0.1732778549194336, 0.21222199499607086, 0.15781369805335999, 0.12740808725357056, 0.1111915186047554, 0.11338130384683609, 0.19232141971588135, 0.17540767788887024]
add = [0.1732778549194336, 0.20100615918636322, 0.26455792784690857, 0.19232141971588135, 0.12740808725357056, 0.20970593392848969]
# x = 0.0008852639002725482
min_loss = 1.
record_m = 0
record_d = 1
for d in range(10, 10000):
m, d = decimal2md(x, d)
loss = abs(m * 1. / d - x)
if loss < min_loss:
min_loss = loss
record_m = m
record_d = d
print(x)
print(record_m * 1. / record_d, record_m, record_d)
# +
import numpy as np
bias_scale = np.array([0.0008852639002725482, 0.0035931775346398354, 0.00785899069160223, 0.0014689048985019326, 0.0015524440677836537, 0.0028435662388801575, 0.001141879241913557, 0.0007087105768732727, 0.009289528243243694, 0.0015117411967366934, 0.004092711955308914])
result_sacale = np.array([0.20100615918636322, 0.42823609709739685, 0.23841151595115662, 0.1732778549194336, 0.21222199499607086, 0.15781369805335999, 0.12740808725357056, 0.1111915186047554, 0.11338130384683609, 0.19232141971588135, 0.17540767788887024])
add_scale = np.array([0.1732778549194336, 0.20100615918636322, 0.26455792784690857, 0.19232141971588135, 0.12740808725357056, 0.20970593392848969])
scale = bias_scale / result_sacale
scale = np.round(scale * 2**10).astype(np.uint16)
add_scale = np.round(add_scale * 2**10).astype(np.uint16)
print(scale, add_scale)
print(np.concatenate((scale, add_scale), axis=0))
# +
'''修改传输顺序,更改为先把一个输出通道按列扫描传输完'''
import numpy as np
weight = np.array(list(range(24)))
weight = weight.reshape(1,2,3,4)
print(weight)
print()
weight_transpose = weight.transpose(0,3,1,2)
print(weight_transpose)
print()
print(weight_transpose.reshape(-1))
# -
| test_tflite_intermediate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# - ブレンディングやってみる
# # Import everything I need :)
import glob
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import mean_absolute_error
# # All function in this notebook
def kaggle_metric(df, preds):
df["prediction"] = preds
maes = []
for t in df.type.unique():
y_true = df[df.type==t].scalar_coupling_constant.values
y_pred = df[df.type==t].prediction.values
mae = np.log(mean_absolute_error(y_true, y_pred))
maes.append(mae)
return np.mean(maes)
# # Preparation
nb = 67
train = pd.read_csv('./../input/champs-scalar-coupling/train.csv')[['type', 'scalar_coupling_constant']]
type_train = train.type
# ## my submission, oof data
path_dir = './../output/'
# <br>
# <br>
# score が -1.0 以下のファイルパスを取り出す
output_paths = sorted(glob.glob(path_dir + '*-1.*.csv'))
output_paths
# <br>
# <br>
# oof path list
# +
oof_paths = []
for path in output_paths:
if 'oof' in path:
oof_paths.append(path)
print(len(oof_paths))
oof_paths
# -
# <br>
# <br>
# submission path list
sub_paths = []
for path in output_paths:
if 'submission' in path:
sub_paths.append(path)
print(len(sub_paths))
sub_paths
# <br>
# <br>
# data_paths
#
# - これらをブレンドする
# +
data_paths = [
'./../output/nb54_{}_random_forest_regressor_-1.45569.csv',
'./../output/nb57_{}_lasso_-1.07263.csv',
'./../output/nb60_{}_lgb_-1.5330660525700779.csv',
'./../output/nb63_{}_ridge_-1.37017.csv',
]
details = [data_paths[i].split('_{}_')[1] for i in range(len(data_paths))]
details
# -
# <br>
# <br>
# load data
# +
# submission
sub_dfs = []
for i, path in enumerate(data_paths):
sub_dfs.append(pd.read_csv(path.format('submission')).drop(['id'], axis=1))
print(f'submission{i}: {path.format("submission")}')
print('-'*80)
oof_dfs = []
for i, path in enumerate(data_paths):
df = pd.read_csv(path.format('oof'))
oof_dfs.append(df)
print(f'oof{i}: {path.format("oof")}')
# -
# <br>
# <br>
# concat
concat_oof = pd.concat(oof_dfs, axis=1)
concat_sub = pd.concat(sub_dfs, axis=1)
# # Blending
# +
median_oof = concat_oof.median(axis=1).values
mean_oof = concat_oof.mean(axis=1).values
median_sub = concat_sub.median(axis=1).values
mean_sub = concat_sub.mean(axis=1).values
# -
# # score
median_oof = kaggle_metric(train, median_oof)
print(f'median: {median_oof}')
mean_oof = kaggle_metric(train, mean_oof)
print(f'mean: {mean_oof}')
# + active=""
# save_score = mean_oof
# save_data = mean_sub
# save_path = f'nb{nb}_blend_{save_score}.csv'
#
# sample_sub = pd.read_csv('./../input/champs-scalar-coupling/sample_submission.csv')
# sample_sub['scalar_coupling_constant'] = save_data
# sample_sub.to_csv(save_path, index=False)
# -
# # analysis each type
types = np.unique(type_train)
result_df = pd.DataFrame({'type': types})
type_scores = []
for i_type, type_ in enumerate(types):
print(f'\n ----- {type_} -----')
idx = type_train==type_
y_true = train['scalar_coupling_constant'][idx]
oof_scores = []
for i_oof in range(len(oof_dfs)):
pred = oof_dfs[i_oof][idx]['oof']
score = np.log(mean_absolute_error(y_true, pred))
print(f'score: {score} \t {details[i_oof]}')
oof_scores.append(score)
type_scores.append(oof_scores)
best_idx = np.argmin(oof_scores)
print(f'---> best score is {details[best_idx]}')
type_scores = np.array(type_scores)
type_scores = pd.DataFrame(type_scores, columns=details)
result_df = pd.concat([result_df, pd.DataFrame(type_scores)], axis=1)
display(result_df)
result_df.plot.bar(x='type', figsize=(10,7))
| src/67_blending_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (Inquire venv)
# language: python
# name: inquire_venv
# ---
# # Loading Libraries
from sentence_transformers import SentenceTransformer
model = SentenceTransformer('bert-base-nli-mean-tokens')
# +
import numpy as np
np.set_printoptions(suppress=True) #prevent numpy exponential
model.encode(["my mother is sick and i have financial difficulties. I am divorced also"])
# + colab={} colab_type="code" id="hsZvic2YxnTz"
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedShuffleSplit
import sys
import os
import pandas as pd
import numpy as np
from datetime import datetime
import matplotlib.pyplot as plt
from sklearn.utils.multiclass import unique_labels
from sklearn.metrics import f1_score,confusion_matrix,classification_report,accuracy_score
import logging
logging.basicConfig(stream=sys.stdout, level=logging.ERROR)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_colwidth', 1000)
# -
# ## Utils functions
def create_examples_prediction(df):
"""Creates examples for the training and dev sets."""
examples = []
for index, row in df.iterrows():
#labels = row[LABEL_HOT_VECTOR].strip('][').split(', ')
#labels = [float(x) for x in labels]
labels = list(row[label_list_text])
examples.append(labels)
return pd.DataFrame(examples)
def f(x):
n = 2 # index of the second proability to get labeled
index = np.argsort(x.values.flatten().tolist())[-n:][0]
print(f"index is {index}")
label = label_list_text[index]
print(f"label is {label}")
return label
def get_test_experiment_df(test):
test_predictions = [x[0]['probabilities'] for x in zip(getListPrediction(in_sentences=list(test[DATA_COLUMN])))]
test_live_labels = np.array(test_predictions).argmax(axis=1)
test['Predicted label'] = [label_list_text[x] for x in test_live_labels] # appending the labels to the dataframe
probabilities_df_live = pd.DataFrame(test_predictions) # creating a proabilities dataset
probabilities_df_live.columns = [x + " Predicted"for x in label_list_text] # naming the columns
probabilities_df_live['Predicted label 2'] = probabilities_df_live.apply(lambda x:f(x),axis=1)
#print(test)
#label_df = create_examples_prediction(test)
#label_df.columns = label_list_text
#label_df['label 2'] = label_df.apply(lambda x:f(x),axis=1)
test.reset_index(inplace=True,drop=True) # resetting index
experiment_df = pd.concat([test,probabilities_df_live],axis=1, ignore_index=False)
experiment_df = experiment_df.reindex(sorted(experiment_df.columns), axis=1)
return test,experiment_df
def getListPrediction(in_sentences):
#1
input_examples = [InputExample(guid="", text_a = x, text_b = None, labels = [0]*len(label_list)) for x in in_sentences] # here, "" is just a dummy label
#2
input_features = convert_examples_to_features(input_examples, MAX_SEQ_LENGTH, tokenizer)
#3
predict_input_fn = input_fn_builder(features=input_features, seq_length=MAX_SEQ_LENGTH, is_training=False, drop_remainder=False)
print(input_features[0].input_ids)
#4
predictions = estimator.predict(input_fn=predict_input_fn,yield_single_examples=True)
return predictions
# +
is_normalize_active=False
def get_confusion_matrix(y_test,predicted,labels):
class_names=labels
# plotting confusion matrix
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plot_confusion_matrix(y_test, predicted, classes=class_names,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plot_confusion_matrix(y_test, predicted, classes=class_names, normalize=True,
title='Normalized confusion matrix')
plt.show()
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
classes =classes
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
#print("Normalized confusion matrix")
else:
test =1
#print('Confusion matrix, without normalization')
fig, ax = plt.subplots()
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
#ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
#fig.tight_layout()
return ax
def plot_matrix(cm,classes,title=None,cmap=plt.cm.Reds):
fig, ax = plt.subplots()
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
#ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f'
thresh = cm.max() / 2.5
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
#fig.tight_layout()
return ax
# -
# # Loading the data
def data_prep_bert(df,test_size):
#print("Filling missing values")
#df[DATA_COLUMN] = df[DATA_COLUMN].fillna('_NA_')
print("Splitting dataframe with shape {} into training and test datasets".format(df.shape))
X_train, X_test = train_test_split(df, test_size=test_size, random_state=2018,stratify = df[LABEL_COLUMN_RAW])
return X_train, X_test
def open_dataset(NAME,mapping_index,excluded_categories):
df = pd.read_csv(PATH+NAME+'.csv',sep =',')
#df[LABEL_COLUMN_RAW] = df[LABEL_COLUMN_RAW].fillna("Other")
df = df[df['is_stressor'] == 1]
df = df[df[LABEL_COLUMN_RAW] != 'Not Stressful']
#df.columns = [LABEL_COLUMN_RAW,'Severity',DATA_COLUMN,'Source']
if excluded_categories is not None:
for category in excluded_categories:
df = df[df[LABEL_COLUMN_RAW] !=category]
label_list=[]
label_list_final =[]
if(mapping_index is None):
df[LABEL_COLUMN_RAW] = df[LABEL_COLUMN_RAW].astype('category')
df[LABEL_COLUMN], mapping_index = pd.Series(df[LABEL_COLUMN_RAW]).factorize() #uses pandas factorize() to convert to numerical index
else:
df[LABEL_COLUMN] = df[LABEL_COLUMN_RAW].apply(lambda x: mapping_index.get_loc(x))
label_list_final = [None] * len(mapping_index.categories)
label_list_number = [None] * len(mapping_index.categories)
for index,ele in enumerate(list(mapping_index.categories)):
lindex = mapping_index.get_loc(ele)
label_list_number[lindex] = lindex
label_list_final[lindex] = ele
frequency_dict = df[LABEL_COLUMN_RAW].value_counts().to_dict()
df["class_freq"] = df[LABEL_COLUMN_RAW].apply(lambda x: frequency_dict[x])
return df,mapping_index,label_list_number,label_list_final
# # Require user changes > Start Here
# ### Experiment Name
PATH = './datasets/'
TODAY_DATE = "01_05_2020/"
EXPERIMENT_NAME = 'main_turk_analysis_of_5_turkers_popbots_test_live_10votes'
EXPERIMENTS_PATH = PATH + 'experiments/'+TODAY_DATE+EXPERIMENT_NAME
if not os.path.exists(PATH + 'experiments/'+TODAY_DATE):
os.mkdir(PATH + 'experiments/'+TODAY_DATE)
if not os.path.exists(EXPERIMENTS_PATH):
os.mkdir(EXPERIMENTS_PATH)
# ### Model Hyperparameters
# + colab={} colab_type="code" id="OjwJ4bTeWXD8"
# Compute train and warmup steps from batch size
# These hyperparameters are copied from this colab notebook (https://colab.sandbox.google.com/github/tensorflow/tpu/blob/master/tools/colab/bert_finetuning_with_cloud_tpus.ipynb)
BATCH_SIZE = 32
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 3.0
# Warmup is a period of time where hte learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 1000
SAVE_SUMMARY_STEPS = 100
# We'll set sequences to be at most 32 tokens long.
MAX_SEQ_LENGTH = 32
OUTPUT_DIR = './models/'+EXPERIMENT_NAME+ '/' #_01_04_2020/
##use downloaded model, change path accordingly
BERT_VOCAB= './bert_model/uncased_L-12_H-768_A-12/vocab.txt'
BERT_INIT_CHKPNT = './bert_model/uncased_L-12_H-768_A-12/bert_model.ckpt'
BERT_CONFIG = './bert_model/uncased_L-12_H-768_A-12/bert_config.json'
# +
DATASET_NAME = '2020-04-29-MainTurkAggregation-5-Turkers_v0_Sorted'
DATA_COLUMN = 'Input.text'
LABEL_COLUMN_RAW = 'top_label'#'Answer.Label'
LABEL_COLUMN = 'label_numeric'
MTURK_NAME = 'mTurk_synthetic'
LIVE_NAME = 'popbots_live'
LABEL_HOT_VECTOR = 'label_conf'
#dataset,mapping_index,label_list, label_list_text = open_dataset('mturk900balanced',None)
EXCLUDED_CATEGORIES = None #['Other'] #None # # if nothing to exclude put None, THIS ALWAYS MUST BE A LIST
mapping_dict = {'Other': 0, 'Everyday Decision Making': 1, 'Work': 2, 'Social Relationships': 3, 'Financial Problem': 4, 'Emotional Turmoil': 5, 'Health, Fatigue, or Physical Pain': 6, 'School': 7, 'Family Issues': 8}#,'Not Stressful':9}
mapping_index = pd.CategoricalIndex([key for key,value in mapping_dict.items()])
dataset,mapping_index,label_list, label_list_text = open_dataset(DATASET_NAME,mapping_index,EXCLUDED_CATEGORIES)
#dataset = dataset[dataset['is_stressor'] == 1]
test_on_mturk_and_popbots_live = False # include live data in training + include mturk in testing
if test_on_mturk_and_popbots_live:
mturk = dataset[dataset['Source']== MTURK_NAME]
live = dataset[dataset['Source']== LIVE_NAME]
live = live.sample(frac=1).reset_index(drop=True) # shuffle live
PERCENTAGE_LIVE_TEST = 50
TEST_PERCENTAGE = len(live)/((100/PERCENTAGE_LIVE_TEST)*len(mturk)) # given to set the percentage of mturk used as test set to have 50/50
print(f"Test percentage is {TEST_PERCENTAGE}")
train,test = data_prep_bert(mturk,TEST_PERCENTAGE) # test size from mturk
train = train.append(live.loc[0:int((1-(PERCENTAGE_LIVE_TEST/100))*len(live))]) # taking 1/2 of that dataset for training
test = test.append(live.loc[int(len(live)*(1-(PERCENTAGE_LIVE_TEST/100))):int(len(live))]) # taking 1/2 of live dataset for testing
else:
# or taking live only for testing
train,test = dataset[dataset['Source']== MTURK_NAME],dataset[dataset['Source']== LIVE_NAME]
train = train[train['is_stressor'] == 1] # remove only non stressor from train
#print(f"Dataset has {len(dataset)} training examples")
print(f"Normal label list is {label_list}")
print(f"The labels text is {label_list_text}")
#Export train test to csv
#train.to_csv(PATH+'900_CSV_SPLITTED/train.csv')
#test.to_csv(PATH+'900_CSV_SPLITTED/test.csv')
# -
dataset.head(1)
# +
df_columns = ['category', 'nb_sentence','nb_sentence_sampled','mean_distinct_word_nb','mean_distinc_word_per_sentence', 'sd','sd per sentence', '95conf_int' ]
count_results = pd.DataFrame(columns = df_columns)
boostrap_number = 50
for category in label_list_text:
len_word_distinct_word = 0
len_word_distinct_word_list = []
for i in range(boostrap_number):
category_df = dataset[dataset[LABEL_COLUMN_RAW] == category].sample(n=38)
category_df_unsampled = dataset[dataset[LABEL_COLUMN_RAW] == category]
results = set()
category_df[DATA_COLUMN].str.lower().str.split().apply(results.update)
len_word_distinct_word += len(list(results))
len_word_distinct_word_list.append(len(list(results)))
count_results = count_results.append({'category':category,
'nb_sentence':len(category_df_unsampled),
'nb_sentence_sampled':len(category_df),'mean_distinct_word_nb':len_word_distinct_word/boostrap_number,
'mean_distinc_word_per_sentence':len_word_distinct_word/boostrap_number/len(category_df),'sd':0,
'list_distinct':len_word_distinct_word_list},
ignore_index=True)
# -
def return_conf_interval(stats):
alpha = 0.95
p = ((1.0-alpha)/2.0) * 100
lower = max(0.0, np.percentile(stats, p))
p = (alpha+((1.0-alpha)/2.0)) * 100
upper = min(1.0, np.percentile(stats, p))
return ["{0:.4f}".format(lower),"{0:.4f}".format(upper)]
# +
count_results['sd']= count_results['list_distinct'].apply(lambda x:np.std(np.array(x), axis=0))
count_results['sd per sentence']= count_results['sd']/38
count_results['95conf_int']= count_results['list_distinct'].apply(lambda x:return_conf_interval([x/38 for x in x]))
count_results[df_columns].sort_values(by=['mean_distinc_word_per_sentence'])
# -
# +
np.set_printoptions(threshold=np.inf,suppress=True)
df_columns = ['category', 'nb_sentence','mean_sd','sd_mean_sd','95conf_int']
count_results = pd.DataFrame(columns = df_columns)
boostrap_number = 1
for category in label_list_text:
boostrap_sd = []
all_mean = []
for i in range(boostrap_number):
category_df = dataset[dataset[LABEL_COLUMN_RAW] == category].sample(n=38)
category_df_unsampled = dataset[dataset[LABEL_COLUMN_RAW] == category]
category_df['embedding'] = model.encode(category_df[DATA_COLUMN].values)
category_df['embedding'] = category_df['embedding'].apply(lambda x: np.array(x))
average_mean = np.mean(np.array(category_df['embedding'].values),axis=0) # vector of 768 dim
average_sd = np.std(np.array(category_df['embedding'].values),axis=0) # vector of 768 dim
overall_sd = np.std(average_sd) # overall sd
boostrap_sd.append(overall_sd)
all_mean.append(average_mean)
print(np.array(all_mean).shape)
mean_all_mean = np.mean(np.array(all_mean),axis=0)
print(np.array(mean_all_mean).shape)
print(category)
print(list(mean_all_mean))
count_results = count_results.append({'category':category,\
'nb_sentence':len(category_df_unsampled),\
'nb_sentence_sampled':len(category_df),
'mean_sd':float(overall_sd),
'all_sd':boostrap_sd,
'mean_vector':mean_all_mean},ignore_index=True)
# +
count_results['sd_mean_sd']= count_results['all_sd'].apply(lambda x:np.std(np.array(x), axis=0))
count_results['95conf_int']= count_results['all_sd'].apply(lambda x:return_conf_interval([x for x in x]))
count_results[df_columns].sort_values(by=['mean_sd'])
# -
count_results['mean_vector'] = count_results['mean_vector'].apply(lambda x: np.array(x).T)
lis = [vector for vector in np.array(count_results['mean_vector'])]
# +
from sklearn.metrics.pairwise import cosine_similarity as cs
ortho = pd.DataFrame(cs(lis,lis))
ortho.index =label_list_text
ortho.columns = label_list_text
ortho
# -
plot_matrix(cm=np.array(ortho),title="Cosine similarity matrix",classes=label_list_text)
def cosine_similarity(u, v):
"""
Cosine similarity reflects the degree of similariy between u and v
Arguments:
u -- a word vector of shape (n,)
v -- a word vector of shape (n,)
Returns:
cosine_similarity -- the cosine similarity between u and v defined by the formula above.
"""
distance = 0.0
### START CODE HERE ###
# Compute the dot product between u and v (≈1 line)
dot = np.dot(u,v)
# Compute the L2 norm of u (≈1 line)
norm_u = np.linalg.norm(u,2,axis=0)
# Compute the L2 norm of v (≈1 line)
norm_v = np.linalg.norm(v,2,axis=0)
# Compute the cosine similarity defined by formula (1) (≈1 line)
cosine_similarity = dot/np.multiply(norm_u,norm_v)
### END CODE HERE ###
return cosine_similarity
dataset
# +
from itertools import combinations
#dataset = dataset[dataset['second_label'].isna()]
np.set_printoptions(threshold=np.inf,suppress=True)
df_columns = ['category', 'nb_sentence','TC_score_mean','TC_score_sd','TC_score_se']
count_results = pd.DataFrame(columns = df_columns)
boostrap_number = 5
embedding_clusters = []
for category in label_list_text:
boostrap_coherence = []
all_mean = []
for i in range(boostrap_number):
category_df = dataset[dataset[LABEL_COLUMN_RAW] == category].sample(n=38)
category_df_unsampled = dataset[dataset[LABEL_COLUMN_RAW] == category]
category_df['embedding'] = model.encode(category_df[DATA_COLUMN].values)
category_df['embedding'] = category_df['embedding'].apply(lambda x: np.array(x))
embedding_clusters.append(list(category_df['embedding']))
# Compute coherence per pair of words
pair_scores = []
for pair in combinations(category_df['embedding'], 2):
pair_scores.append(cosine_similarity(pair[0], pair[1]))
# get the mean over all pairs in this topic
topic_score = sum(pair_scores) / len(pair_scores)
boostrap_coherence.append(topic_score)
#total_coherence += topic_score
# get the mean score across all topics
mean_coherence = np.mean(boostrap_coherence)
sd_coherence= np.std(boostrap_coherence)
sde = np.std(boostrap_coherence)/np.sqrt(38)
count_results = count_results.append({'category':category,\
'nb_sentence':len(category_df_unsampled),\
'nb_sentence_sampled':len(category_df),
'TC_score_mean':mean_coherence,'TC_score_sd':sd_coherence,'TC_score_se':sde},ignore_index=True)
# -
count_results.sort_values(by='TC_score_mean')
def compute_TC_W2V(w2v_lookup, topics_words):
'''
Compute TC_W2V for the topics of a model using the w2v_lookup.
TC_W2V is calculated for all possible pairs of words in the topic and then averaged with the mean for that topic.
The total TC_W2V for the model is the mean over all topics.
'''
total_coherence = 0.0
for topic_index in range(len(topics_words)):
# Compute coherence per pair of words
pair_scores = []
for pair in combinations(topics_words[topic_index], 2):
try:
pair_scores.append(w2v_lookup.similarity(pair[0], pair[1]))
except KeyError as e:
# If word is not in the word2vec model then as score 0.5
print(e)
pair_scores.append(0.5)
# get the mean over all pairs in this topic
topic_score = sum(pair_scores) / len(pair_scores)
total_coherence += topic_score
# get the mean score across all topics
return total_coherence / len(topics_words)
tsne_model_en_2d = TSNE(perplexity=15, n_components=2, init='pca', n_iter=3500, random_state=32)
embedding_clusters = np.array(embedding_clusters)
n, m, k = embedding_clusters.shape
embeddings_en_2d = np.array(tsne_model_en_2d.fit_transform(embedding_clusters.reshape(n * m, k))).reshape(n, m, 2)
# +
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
# %matplotlib inline
def tsne_plot_similar_words(labels, embedding_clusters, word_clusters, a=0.7):
plt.figure(figsize=(16, 9))
colors = cm.rainbow(np.linspace(0, 1, len(labels)))
for label, embeddings, color in zip(labels, embedding_clusters, colors):
x = embeddings[:,0]
y = embeddings[:,1]
plt.scatter(x, y, c=color, alpha=a, label=label)
#for i, word in enumerate(words):
# plt.annotate(word, alpha=0.5, xy=(x[i], y[i]), xytext=(5, 2),
# textcoords='offset points', ha='right', va='bottom', size=8)
plt.legend(loc=4)
plt.grid(True)
#plt.savefig("f/г.png", format='png', dpi=150, bbox_inches='tight')
plt.show()
tsne_plot_similar_words(label_list_text, embeddings_en_2d, word_clusters=None)
# -
# ### Train set and test set analysis
def print_dataset_info(train,test):
print(f"Train size {len(train)} with {len(train[train['Source']== LIVE_NAME])} from Popbots and {len(train[train['Source']== MTURK_NAME])} from mturk")
print(f"Test size {len(test)} with {len(test[test['Source']== LIVE_NAME])} from Popbots and {len(test[test['Source']== MTURK_NAME])} from mturk")
print('\nTraining distribution:')
print(pd.pivot_table(train[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=True)
print('\nTesting distribution:')
print(pd.pivot_table(test[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=True)
len(test)
train = train.sample(frac=1).reset_index(drop=True) #reshuffle everything
test = test.sample(frac=1).reset_index(drop=True)
print('\nAll dataset distribution:')
print(pd.pivot_table(dataset[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=T
print_dataset_info(train,test)
# ### Step to reduce the most dominant categories and balance the dataset
# +
sampling_cutoff = 100 # all the categories which had less than 100 example won't be sampled down
total_training_size = 1501
REVERSE_FREQ = 'Max_reverse_sampling_chance'
train[REVERSE_FREQ] = train['class_freq'].apply(lambda x: (max(train['class_freq'])/x))
sampling_boolean = (train['Source'] != LIVE_NAME) & (train['class_freq'].astype(float) > sampling_cutoff)
train_to_be_balanced = train[sampling_boolean]
train_not_resampled = train[~sampling_boolean]
train_temp = train_to_be_balanced.sample(n=(total_training_size-len(train_not_resampled)), weights=REVERSE_FREQ, random_state=2020)
train = pd.concat([train_temp,train_not_resampled])
# -
print_dataset_info(train,test)
mapping_index
train.to_csv(EXPERIMENTS_PATH+'/TRAIN_'+DATASET_NAME+'.csv')
test.to_csv(EXPERIMENTS_PATH+'/TEST_'+DATASET_NAME+'.csv')
| bert-pipeline/Dataset_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rtazV2N5pELO" colab_type="text"
# # Step1
# + [markdown] id="yec7Qt08pHqo" colab_type="text"
# ### Target
#
# 1. Get the setup right
# 2. Read MNIST dataset, set train test split and create Data Loader
# 3. Get the summary statistics for the data
# 4. Set initial transforms and apply transformation to the train and test set separately
# 5. Get the basic neural net architecture skeleton right. We will try and avoid changing the skeleton later
# 6. Set basic training and test loop
#
# ### Results
#
# 1. Parameters: 992,800
# 2. Best Training Accuracy: 99.88
# 3. Best Test Accuracy: 99.14
#
# ### Analysis
#
# 1. Very heavy model for such an easy problem. Lots of parameters. Have to reduce the number of parameters in the next step
# 2. Test accuracy is way below the target accuracy
# 3. Model is overfitting
# + [markdown] id="aO-7t1Y7-hV4" colab_type="text"
# # Import Libraries
# Import the necessary packages
# + id="8kH16rnZ7wt_" colab_type="code" colab={}
from __future__ import print_function
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torchsummary import summary
import matplotlib.pyplot as plt
from tqdm import tqdm
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
# + [markdown] id="oQciFYo2B1mO" colab_type="text"
# # Read Dataset and Create Train/Test Split
# + id="_4A84rlfDA23" colab_type="code" colab={}
train = datasets.MNIST('./data', train=True, download=True)
test = datasets.MNIST('./data', train=False, download=True)
# + [markdown] id="-TFjoFekE_va" colab_type="text"
# # Data Statistics and Visualization
#
# Let us check some of the summary statistics for our data. It will help us to set initial transform. We can also visualize some sample images to get an idea of what transforms we can apply later.
# + id="hWZPPo3yEHDW" colab_type="code" colab={}
# Function to get summary statistics
def get_summary_stats(sample_data, data_label="Sample Data"):
"""
Function to get summary statistics from a given data
Args
----
sample_data: Tensor
data_label: str
Returns
-------
None
"""
sample = sample_data.data
sample = sample.numpy()
# Normalizing data between 0 and 1
sample = sample/255
print(f'Summary Statistics for {data_label}')
print(' - Numpy Shape:', sample.shape)
print(' - Tensor Shape:', sample.size)
print(' - min:', np.min(sample))
print(' - max:', np.max(sample))
print(' - mean:', np.mean(sample))
print(' - std:', np.std(sample))
print(' - var:', np.var(sample))
# + id="CQFIL7-ixtKt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 487} outputId="bb91fbf4-7cf2-4488-856c-f6dc687307a5"
combined_data = torch.cat([train.data, test.data])
get_summary_stats(train.data, data_label="Training Dataset")
print("\n")
get_summary_stats(test.data, data_label="Test Dataset")
print("\n")
get_summary_stats(combined_data, data_label="Complete Dataset")
# + id="RQlcjBWQ1FWz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="152d9844-0799-4acd-e2a4-5f2941cd61f5"
fig, ax = plt.subplots(figsize = (14, 5))
count_plot = sns.countplot(pd.Series(train.targets.numpy()), color="#3274a1", ax = ax)
ax.set_xlabel("Class")
ax.set_ylabel("Count of class")
ax.set_title("Countplot for classes in training set")
# + [markdown] id="_NTCBwsn47hf" colab_type="text"
# ### Inference
# The classes are almost balanced. No need for any class imbalance tricks
# + [markdown] id="_WlLQMDX6ISg" colab_type="text"
# Now lets visualize some of the images
# + id="smqff6lJ46PI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="93de5d2e-8056-455d-f5c7-46125b5f7058"
figure = plt.figure(figsize=(10, 6))
num_of_images = 60
for index in range(1, num_of_images + 1):
plt.subplot(6, 10, index)
plt.axis('off')
plt.imshow(train.data.numpy()[index], cmap='gray_r')
#Deleting the combined data since its of no use any longer
del combined_data
# + [markdown] id="WaXMkAQt6oZW" colab_type="text"
# # Setting Transforms
# Now we can set transforms from the summary statistics. We will be applying the following transformations:
# * ToTensor() : It converts PIL image or numpy array to FloatTensor in the range of [0.0, 1.0]
# * Normalize: It normalizes each channel of an input given a mean and std deviation. We have already ascertained the mean and std deviation of the dataset to be 0.1309 ad 0.3084
# + id="YtssFUKb-jqx" colab_type="code" colab={}
# Train Phase transformations
train_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1309,), (0.3084,))
])
# Test Phase transformations
test_transforms = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1309,), (0.3084,))
])
# + id="eF6Wu_Ns9SzG" colab_type="code" colab={}
#Read data and apply the above transforms
train = datasets.MNIST('./data', train=True, download=True, transform=train_transforms)
test = datasets.MNIST('./data', train=False, download=True, transform=test_transforms)
# + [markdown] id="qgldp_3-Dn0c" colab_type="text"
# # Set Dataloader Arguments & Create Test/Train Dataloaders
#
# + id="C8OLDR79DrHG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="ed05eb5f-b7b2-4a3c-f5bf-cabf623085cb"
# Set the manual seed
SEED = 1
# Initialize batch size
bs = 128
# Check for availability of cuda
cuda = torch.cuda.is_available()
print(" Is CUDA Available ?\n", cuda)
# For reproducibility
torch.manual_seed(SEED)
if cuda:
torch.cuda.manual_seed(SEED)
# Setting the dataloader arguments
dataloader_args = dict(shuffle=True, batch_size=bs, num_workers=4, pin_memory=True) if cuda else dict(shuffle=True, batch_size=64)
# train dataloader
train_loader = torch.utils.data.DataLoader(train, **dataloader_args)
# test dataloader
test_loader = torch.utils.data.DataLoader(test, **dataloader_args)
# + [markdown] id="ubQL3H6RJL3h" colab_type="text"
# # The model architecture
# Now we can setup or initial model skeleton
# + id="7FXQlB9kH1ov" colab_type="code" colab={}
class Net(nn.Module):
def __init__(self):
"""
Initializes all the model layers
"""
#Inheriting from the nn.Module class
super(Net, self).__init__()
#Convolution Block 1
self.conv1 = nn.Sequential(
nn.Conv2d(in_channels=1, out_channels=32, kernel_size=3, bias=False, padding=0),
nn.ReLU()
) #Input: 28X28X1 | Output: 26X26X32 | RF: 3
#Convolution Block 2
self.conv2 = nn.Sequential(
nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, bias=False, padding=0),
nn.ReLU()
) #Input: 26X26X32 | Output: 24X24X64 | RF: 5
#Convolution Block 3
self.conv3 = nn.Sequential(
nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, bias=False, padding=0),
nn.ReLU()
) #Input: 24X24X64 | Output: 22X22X128 | RF: 7
#Transition Block 1
self.pool1 = nn.MaxPool2d(2, 2) #Input: 22X28X128 | Output: 11X11X128 | RF: 8
self.conv4 = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=32, kernel_size=1, bias=False),
nn.ReLU()
) #Input: 11X11X128 | Output: 11X11X32 | RF: 8
#Convolution Block 5
self.conv5 = nn.Sequential(
nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, bias=False, padding=0),
nn.ReLU()
) #Input: 11X11X32 | Output: 9X9X64 | RF: 12
#Convolution Block 6
self.conv6 = nn.Sequential(
nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, bias=False, padding=0),
nn.ReLU()
) #Input: 9X9X64 | Output: 7X7X128 | RF: 16
#Output Block
self.conv7 = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=128, kernel_size=7, bias=False, padding=0),
nn.ReLU()
) #Input: 7X7X128 | Output: 1X1X128 | RF: 28
self.conv8 = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=10, kernel_size=1, bias=False, padding=0),
) #Input: 1X1X128 | Output: 1X1X10 | RF: 28
def forward(self, x):
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.pool1(x)
x = self.conv4(x)
x = self.conv5(x)
x = self.conv6(x)
x = self.conv7(x)
x = self.conv8(x)
x = x.view(-1, 10)
return F.log_softmax(x, dim=-1)
# + [markdown] id="M3-vp8X9LCWo" colab_type="text"
# # Model Params
# We can now check the model summary and see the total number of parameters
# + id="5skB97zIJQQe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 521} outputId="30b9795c-ee15-4341-f5bc-143a788a00f6"
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu") #Use cuda if available else use cpu
print(device)
model = Net().to(device) #Sending the model to cuda if available otherwise use cpu
summary(model, input_size=(1, 28, 28))
# + [markdown] id="1__x_SbrL7z3" colab_type="text"
# # Training and Testing
#
# All right, so we have 9.9M params, and that's too many, we know that. But the purpose of this notebook is to set things right for our future experiments.
#
# Looking at logs can be boring, so we'll introduce **tqdm** progressbar to get cool looking logs.
#
# Let's write train and test functions
# + id="fbkF2nN_LYIb" colab_type="code" colab={}
train_losses = []
test_losses = []
train_acc = []
test_acc = []
def train(model, device, train_loader, optimizer, epoch):
model.train()
pbar = tqdm(train_loader)
correct = 0
processed = 0
for batch_idx, (data, target) in enumerate(pbar):
# get samples
data, target = data.to(device), target.to(device)
# Init
optimizer.zero_grad()
# In PyTorch, we need to set the gradients to zero before starting to do backpropragation because PyTorch accumulates the gradients on subsequent backward passes.
# Because of this, when you start your training loop, ideally you should zero out the gradients so that you do the parameter update correctly.
# Predict
y_pred = model(data)
# Calculate loss
loss = F.nll_loss(y_pred, target)
train_losses.append(loss)
# Backpropagation
loss.backward()
optimizer.step()
# Update pbar-tqdm
pred = y_pred.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
processed += len(data)
train_acc.append(100*correct/processed)
pbar.set_description(desc=f'Loss={loss.item():0.4f} Batch_ID={batch_idx} Accuracy={train_acc[-1]:.2f}')
def test(model, device, test_loader):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item() # sum up batch loss
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
test_losses.append(test_loss)
test_acc.append(100. * correct / len(test_loader.dataset))
print(f'\nTest set: Average loss: {test_loss:.4f}, Accuracy: {correct}/{len(test_loader.dataset)} ({test_acc[-1]:.2f}%)\n')
# + [markdown] id="drokW8wWODKq" colab_type="text"
# # Let's Train and test our model
# + id="xMCFxeAKOB53" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d21800e9-dd5e-46ca-d5d8-00dd5cfe26a0"
model = Net().to(device)
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
EPOCHS = 15
for epoch in range(EPOCHS):
print("EPOCH:", epoch+1)
train(model, device, train_loader, optimizer, epoch)
test(model, device, test_loader)
# + id="87RaqGSEOWDe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 624} outputId="39c89af7-8000-46c1-96e5-1637e4deeee0"
fig, axs = plt.subplots(2,2,figsize=(15,10))
axs[0, 0].plot(train_losses)
axs[0, 0].set_title("Training Loss")
axs[1, 0].plot(train_acc)
axs[1, 0].set_title("Training Accuracy")
axs[0, 1].plot(test_losses)
axs[0, 1].set_title("Test Loss")
axs[1, 1].plot(test_acc)
axs[1, 1].set_title("Test Accuracy")
# + id="odozjbIvY12p" colab_type="code" colab={}
| Week5/Step1.ipynb |
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% some housekeeping stuff
register_graphics_toolkit("gnuplot");
available_graphics_toolkits();
graphics_toolkit("gnuplot")
clear
% end of housekeeping
% # ionic strength effect on pH of acetic acid
%
% see notes for description
% +
%plot -s 600,500 -f 'svg'
logmu=-6:0.2:-1; mu=10.^logmu; AT=0.01; Ka=10^-4.75; sizeH=900; sizeAc=450;
%zero ionic strength
a=1; b=Ka; c=-Ka*AT; t=roots([a b c]); t(imag(t)==0); t=t(t>0); pHzero=-log10(t);
for i=1:length(logmu)
loggammaH=-0.51*sqrt(mu(i))/(1+((sizeH/350)*sqrt(mu(i)))); gammaH=10^loggammaH;
loggammaAc=-0.51*sqrt(mu(i))/(1+((sizeAc/350)*sqrt(mu(i)))); gammaAc=10^loggammaAc;
Kaprime=Ka/(gammaH*gammaAc);
a=1; b=Kaprime; c=-Kaprime*AT;
t=roots([a b c]);
t(imag(t)==0); %sets any imaginary roots to zero
t=t(t>0); %t=positive real roots
pH(i)=-log10(t*gammaH);
end
plot(logmu,pH,'ko','markersize',4,'markerfacecolor','b')
set(gca,'linewidth',1.5)
xlabel('log(\mu)'); ylabel('pH')
hold on
plot(logmu,pHzero*ones(size(logmu)),'k--','linewidth',2)
| pH_acetic_acid_versus_ionicstrength.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Create subsets à 10000 spectra for full sample runs
#
# ## Author(s): <NAME> (SB, WG4)
#
# ### History:
# 180926 SB Created
import numpy as np
import astropy.io.fits as pyfits
sobject_data = pyfits.getdata('sobject_iraf_53_2MASS_GaiaDR2_WISE_PanSTARRSDR1_BailerJones_K2seis.fits',1)
# ## Apply quality cut: PLX available and FLAG_GUESS <= 8
# +
print('initial set: '+str(len(sobject_data)))
quality_cut = np.isfinite(sobject_data['parallax']) & (sobject_data['flag_guess'] <= 8)
sobject_data = sobject_data[quality_cut]
u1, sobject_data_index = np.unique(sobject_data['sobject_id'], return_index=True)
sobject_data = sobject_data[sobject_data_index]
print('set after quality cut: '+str(len(sobject_data)))
# -
# ## Create subsets after sorting by effective temperature
sobject_data = np.sort(sobject_data,order='teff_guess')
for each_subset in range(len(sobject_data)/10000+1):
subset = sobject_data[each_subset*10000:np.min([(each_subset+1)*10000,len(sobject_data)])]
np.savetxt('10k_subsets/GALAH_10k_'+str(each_subset)+'_lbol',zip(['10k_'+str(each_subset)+'_lbol' for x in range(len(subset))],subset['s',['DR3' for x in range(len(subset))]),fmt='%s')
| input/create_random_10k_sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datojen haku ja esikäsittely
from set_path import set_path
mainpath, path = set_path('areadata')
from read_and_prepare_data import read_and_prepare_data
stat, post, kunta_stat, vaalidata = read_and_prepare_data(path)
from selected_cols import selected_cols
numeric_features, categorical_features = selected_cols(largeset=True, parties=False)
# +
from sklearn.feature_selection import mutual_info_classif
import pandas as pd
from draw_and_create_clusters import create_kmeans_clusters
from prepare_and_scale_data import prepare_and_scale_data
from create_prediction import select_kbest
from select_columns_and_clean_data import select_columns_and_clean_data
from draw_and_create_clusters import draw_pca, drawTSNE, display_scree_plot, display_circles, display_parallel_coordinates_centroids, display_factorial_planes
from delete_outliers import delete_outliers
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# Load the TensorBoard notebook extension
# %load_ext tensorboard
pd.options.display.max_colwidth = 100
# +
from create_target_columns import create_target_columns
from create_neuro_prediction import create_neuro_prediction
list_of_partiest = ['VIHR', 'KOK', 'SDP', 'KD', 'KESK', 'RKP', 'PS', 'VAS']
target_col_start = 'Äänet yhteensä lkm'
target = create_target_columns(list_of_partiest, target_col_start)
data, test, model, hist, log_path = create_neuro_prediction(stat, stat, target, mainpath, numeric_features=numeric_features, categorical_features=categorical_features, scaled=True, test_size = 0.2, Skfold=False)
# -
from create_neuro_prediction import plot_history
plot_history(hist)
from show_election_result import show_election_result
show_election_result(data, vaalidata, target_col_start, list_of_partiest)
# +
# #%tensorboard --logdir=log_path
| create_political_neuro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Load the data
# -
import pandas as pd
data = pd.read_csv('./data.csv')
# +
## Clean the data
# -
data.columns
data.drop(['sqft_living','sqft_lot','waterfront','view','condition','sqft_above','sqft_basement','street','city','statezip','country'],axis=1,inplace=True)
data.drop('date',axis=1,inplace=True)
data.head()
# +
## Feature Enginnering
# -
def fe(data,col):
print(len(data))
max_no = data[col].quantile(0.99)
min_no = data[col].quantile(0.05)
data = data[data[col] > min_no]
data = data[data[col] < max_no]
print(len(data))
return data
for col in list(data.columns):
print(col)
data = fe(data,'price')
data.head()
X = data.drop('price',axis=1)
y = data['price']
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.25)
len(X_train),len(X_test)
# +
## Modelling
# -
import torch
import torch.nn as nn
import torch.optim as optim
class BaseLine_Model(nn.Module):
def __init__(self,input_shape,output_shape):
super().__init__()
self.fc1 = nn.Linear(input_shape,32)
self.fc2 = nn.Linear(32,64)
self.fc3 = nn.Linear(64,128)
self.fc4 = nn.Linear(128,64)
self.fc5 = nn.Linear(64,output_shape)
def forward(self,X):
preds = self.fc1(X)
preds = self.fc2(preds)
preds = self.fc3(preds)
preds = self.fc4(preds)
preds = self.fc5(preds)
return preds
EPOCHS = 100
import wandb
BATCH_SIZE = 32
PROJECT_NAME = 'House-Price-Pred'
from tqdm import tqdm
device = torch.device('cuda')
model = BaseLine_Model(5,1).to(device)
criterion = nn.MSELoss()
optimizer = optim.Adam(model.parameters(),lr=0.1)
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if pred[0] == y[i]:
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
preds.to(device)
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if pred[0] == y[i]:
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
preds.to(device)
y.to(device)
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if pred[0] == y[i]:
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
def get_loss(criterion,X,y,model):
preds = model(X.float().to(device))
preds.to(device)
y.to(device)
criterion.to(device)
loss = criterion(preds,y)
return loss.item()
def get_accuracy(X,y,model):
correct = 0
total = 0
for i in range(len(X)):
pred = model(X[i].float().to(device))
pred.to(device)
if pred[0] == y[i]:
correct += 1
total += 1
if correct == 0:
correct += 1
return round(correct/total,3)
import numpy as np
X_train = torch.from_numpy(np.array(X_train))
y_train = torch.from_numpy(np.array(y_train))
X_test = torch.from_numpy(np.array(X_test))
y_test = torch.from_numpy(np.array(y_test))
get_accuracy(X_test,y_test,model)
wandb.init(project=PROJECT_NAME,name='baseline')
for _ in tqdm(range(EPOCHS)):
for i in range(0,len(X_train),BATCH_SIZE):
X_batch = X_train[i:i+BATCH_SIZE].to(device)
y_batch = y_train[i:i+BATCH_SIZE].to(device)
model.to(device)
preds = model(X_batch.float())
preds.to(device)
loss = criterion(preds.float(),y_batch.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
wandb.log({'loss':loss.item(),'val_loss':get_loss(criterion,X_test,y_test,model),'accuracy':get_accuracy(X_train,y_train,model),'val_accuracy':get_accuracy(X_test,y_test,model)})
| wandb/run-20210519_201451-3ab5hkba/tmp/code/_session_history.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + nbpresent={"id": "f5fc4534-488a-4e54-8e4c-e97a86dc9388"} slideshow={"slide_type": "skip"}
# %pylab inline
pylab.rcParams['figure.figsize'] = (16.0, 8.0)
# + [markdown] nbpresent={"id": "e1323dc1-41cf-40d4-8250-9dab630288a5"} slideshow={"slide_type": "slide"}
# # Adaptive determination of Monte Carlo trials
# + [markdown] nbpresent={"id": "411f678c-474e-4890-a4c1-c7c8a3b40bab"} slideshow={"slide_type": "subslide"}
# The Monte Carlo outcome is based on **random** draws from the joint probability distribution associated with the input quantities. Thus, the outcome and every statistics derived are **random**.
# + [markdown] nbpresent={"id": "395824a3-46fd-4e80-85c2-6f28f82d6393"} slideshow={"slide_type": "subslide"}
# ### Exercise 5.1
#
# For the model function
# $$ Y = f(X_1,X_2,X_3) = X_1 + X_2 + X_3 $$
# with independent input quantities for which knowledge is encoded as
#
# - $X_1$: Gamma distribution with scale parameter $a=1.5$
#
# - $X_2$: normal distribution with $\mu=1.3$ and $\sigma=0.1$
#
# - $X_3$: t-distribution with location parameter $0.8$ and scale parameter $0.3$ and with 5 degrees of freedom
#
# carry out a Monte Carlo simulation with 1000 runs. Repeat this simulation 100 times using a for-loop. Calculate and store the estimates $y$ for each simulation run and compare the different outcomes.
# + nbpresent={"id": "6f133a6d-1ac8-412b-a3af-8bbd10d381d6"} slideshow={"slide_type": "skip"}
from scipy.stats import gamma, norm, t
draws = 1000
repeats = 100
y_mean = zeros(repeats)
y_unc = zeros(repeats)
for k in range(repeats):
X1 = gamma.rvs(1.5, size=draws)
X2 = norm.rvs(loc=1.3, scale=0.1, size=draws)
X3 = t.rvs(5, loc=0.8, scale=0.3, size=draws)
Y = X1 + X2 + X3
y_mean[k] = mean(Y)
y_unc[k] = std(Y)
figure(1)
subplot(121)
hist(y_mean)
subplot(122)
hist(y_unc);
# + [markdown] nbpresent={"id": "8e94c66a-8598-46b6-8ecf-dd1a3c6d28c2"} slideshow={"slide_type": "slide"}
# ## Adaptive Monte Carlo method
# + [markdown] nbpresent={"id": "a4ea3da2-36b8-4b28-8c6f-929864f0272d"} slideshow={"slide_type": "subslide"}
# The randomness of the Monte Carlo outcomes cannot be avoided. However, the variation between runs decreases with an increasing number of Monte Carlo simulations. The aim is thus to adaptively decide on the number of Monte Carlo trials based on
#
# * a prescribed numerical tolerance
#
# * at a chosen level of confidence
# + [markdown] nbpresent={"id": "94e88390-ae8c-400f-943d-626a869e5dbc"} slideshow={"slide_type": "subslide"}
# #### Stein's method
#
# From Wübbeler et al. (doi: http://iopscience.iop.org/0026-1394/47/3/023):
#
# Let $y_1, y_2, \ldots$ be a sequence of values drawn independentyl from a Gaussian distribution with unknown expecation $\mu$ and variance $\sigma^2$.
# The aim is to determine a rule that terminates this sequence such that $\bar{y}(h)$, being the average of the sequence terminated at $h$, satisfies that the interval
# $$ [\bar{y}(h)-\delta, \bar{y}(h)+\delta] $$
# is a confidence interval for $\mu$ at confidence level $1-\alpha$.
# + [markdown] nbpresent={"id": "1fd959e0-6d89-494f-82cb-3c23604b5525"} slideshow={"slide_type": "subslide"}
# 1) Draw an initial number $h_1>1$ of samples and calculate
# $$ s_y^2(h_1) = \frac{1}{h-1} \sum_{i=1}^{h_1} (y_i - \bar{y}(h_1))^2 $$
#
# 2) Calculate the number $h_2$ of additional values as
# $$ h_2 = \max \left( floor({\frac{s_y^2(h_1)(t_{h_1-1,1-\alpha/2})^2}{\delta^2}})-h_1+1,0 \right) $$
# + [markdown] nbpresent={"id": "11faae09-3d8d-4b6f-9b4e-04d22ab0c5a0"} slideshow={"slide_type": "subslide"}
# #### Application to Monte Carlo simulations
#
# We consider Monte Carlo simulations block-wise. That is, we choose a modest number of Monte Carlo trials, e.g. 1000, and consider a Monte Carlo simulation with that number of trials as one block. Each block has a block mean, standard deviation (uncertainty), etc.
# + [markdown] nbpresent={"id": "8076a57b-5c40-4348-ad22-12e39a47b7d0"} slideshow={"slide_type": "fragment"}
# With $h_1$ being the number of such blocks and $y_1,y_2,\ldots$ a selected outcome of each block (e.g. the mean, variance, interval boundaries, etc.) Stein's method can be applied to calculate the additionally required number of blocks.
# + [markdown] nbpresent={"id": "e0946270-2b4b-4438-a1b1-a3a39488b59c"} slideshow={"slide_type": "subslide"}
# **Reminder**
# The deviation $\delta$ can be calculated from a prescribed number of significant digits as follows:
#
# - Write the number of interest in the form $ z = c \times 10^l$ with $c$ having the chosen number of digits.
#
# - Calculate the numerical tolerance as $\delta = \frac{1}{2} 10^l$
# + [markdown] nbpresent={"id": "8ba4d8a4-8e0e-480a-b8e9-f524ee82251a"} slideshow={"slide_type": "subslide"}
# ### Exercise 5.2
#
# Repeat Exercise 5.1 using Stein's method, starting with an initial number of $h_1 = 10$ repetitions. Calculate $h_2$ such that a numerical tolerance of 2 digits is achieved with a 95% level of confidence.
# + nbpresent={"id": "6f49734d-9983-402b-b6ef-64f07de8b332"} slideshow={"slide_type": "skip"}
from scipy.stats import gamma, norm, t
rst = random.RandomState(1)
h1 = 10
y_mean = zeros(h1)
y_unc = zeros(h1)
for k in range(h1):
X1 = gamma.rvs(1.5, size=draws)
X2 = norm.rvs(loc=1.3, scale=0.1, size=draws)
X3 = t.rvs(5, loc=0.8, scale=0.3, size=draws)
Y = X1 + X2 + X3
y_mean[k] = mean(Y)
y_unc[k] = std(Y)
# -
delta = 0.005
alpha = 0.05
h2 = int(max( floor(y_mean.var()*t(h1-1).ppf(1-alpha/2)**2/delta**2) - h1+1, 0 ))
print(h2)
# + [markdown] nbpresent={"id": "e9e465ca-a013-420c-a268-60c02aaed12a"}
# The confidence level for the achieved accuracy is a frequentist measure. Therefore, in order to verify the achieved confidence, we repeat the adaptive Monte Carlo method and assess the long run success.
# + nbpresent={"id": "7b8de154-c2b1-4ad2-9404-d667f4505bb1"}
# validate the level of confidence
reruns = 1000
y = zeros(reruns)
MCruns = 1000
h1 = 10
# + [markdown] nbpresent={"id": "f8bca1de-65fa-4009-80ce-ceb83285bfba"}
# The results of the adaptive Monte Carlo method are still random. The spread of calculated mean values, however, is below the chosen tolerance with the prescribed level of confidence.
| .ipynb_checkpoints/06 Adaptive determination of Monte Carlo trials-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# ## Define a function and its integral
def dfdx(x, f):
return x**2 + x
def f_int(x, C):
return (x**3)/3. + 0.5*x**2 + C
# ## Define 2nd order RK method
# +
def rk2_core(x_i, f_i, h, g):
# Advance f by a step h
# Half step
x_ipoh = x_i + 0.5*h #ipoh = i plus a half
f_ipoh = f_i + 0.5*h*g(x_i, f_i)
# Full step
f_ipo = f_i + h*g(x_ipoh, f_ipoh)
return f_ipo #ipo = i plus one :o
#I think here we're running the derivative on the right or left end of the function
#so we can find the halfstep later on...
# -
# ## Define a wrapper routine for RK2
def rk2(dfdx, a, b, f_a, N):
#dfdx = derivative wrapper routine x
#a = lower bound
#b = upper bound
#f_a = boundary condition at a
#N = number of steps
# Define our steps
x = np.linspace(a, b, N)
# A single step size
h = x[1] - x[0]
# An array to hold f
f = np.zeros(N, dtype=float)
f[0] = f_a # Value of f at a
# Evolve f along x
for i in range(1, N): #our value at 0 is f[0]=f_a
f[i] = rk2_core(x[i-1], f[i-1], h, dfdx)
return x, f #We return both so we can follow how each changes
#with respect to the other
# A wrapper routine is ...?
# ## Define the 4th order RK method
def rk4_core(x_i, f_i, h, g):
# Define x at 1/2 step
x_ipoh = x_i + 0.5*h
# Define x at 1 step
x_ipo = x_i +h
# Advance f by a step h
k_1 = h*g(x_i, f_i)
k_2 = h*g(x_ipoh, f_i + 0.5*k_1)
k_3 = h*g(x_ipoh, f_i + 0.5*k_2)
k_4 = h*g(x_ipo, f_i + k_3)
f_ipo = f_i + (k_1 + 2*k_2 + 2*k_3 + k_4)/6.
return f_ipo
# ## Define a wrapper for RK4
def rk4(dfdx, a, b, f_a, N):
#dfdx = derivative wrapper routine x
#a = lower bound
#b = upper bound
#f_a = boundary condition at a
#N = number of steps
# Define our steps
x = np.linspace(a, b, N)
# A single step size
h = x[1] - x[0]
# An array to hold f
f = np.zeros(N, dtype=float)
f[0] = f_a # Value of f at a
# Evolve f along x
for i in range(1, N): #our value at 0 is f[0]=f_a
f[i] = rk4_core(x[i-1], f[i-1], h, dfdx)
return x, f
# ...
# Here we decided to take 4 steps and we got a really good approximation (to 10 to the -16, which is as close as the computer can caluclate due to the bit size). Even if we had taken a smaller amount of steps, it would have been still good.
#
# This means that, even though we don't know how many steps we need specifically, we don't need that many in general.
#
# In sinusoidal functions, steps must be smaller than the period of the function
# ...
# ## Perform the integration
a = 0.0
b = 1.0
f_a = 0.0
N = 10
x_2, f_2 = rk2(dfdx, a, b, f_a, N)
x_4, f_4 = rk4(dfdx, a, b, f_a, N)
x = x_2.copy() #Remember: creating a copy of x_2 array, called 'x', so we can edit it freely
plt.plot(x_2, f_2, label = 'RK2')
plt.plot(x_4, f_4, label = 'RK4')
plt.plot(x, f_int(x, f_a), 'o', label='Analytic')
plt.legend(frameon=False)
# ## Plot against the error
# +
#Gotta add other stuff
a = 0.0
b = 1.0
f_a = 1.0
N = 10
x_2, f_2 = rk2(dfdx, a, b, f_a, N)
x_4, f_4 = rk4(dfdx, a, b, f_a, N)
x = x_2.copy() #Remember: creating a copy of x_2 array, called 'x', so we can edit it freely
f_analytic = f_int(x, f_a)
error_2 = (f_2 - f_analytic)/f_analytic
error_4 = (f_4 - f_analytic)/f_analytic
plt.plot(x_2, error_2, label = 'RK2')
plt.plot(x_4, error_4, label = 'RK4')
#plt.ylim(-1.e-3, 1.0e-4)
#plt.plot(x, f_int(x, f_a), 'o', label='Analytic')
plt.legend(frameon=False)
# -
| runge_kutta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''ekole'': conda)'
# name: python3
# ---
# +
import torch
import torchvision
import torch.nn as nn
import torch.nn.functional as F
import math
import torch.optim as optim
import torchvision.transforms as transforms
import numpy as np
import matplotlib.pyplot as plt
torch.set_printoptions(linewidth=120)
torch.set_grad_enabled(True)
# +
import tarfile
tar= tarfile.open('D:/Dev Projects/AI_Projects/ocr_recognition_model/EnglishFnt.tgz')
tar.extractall('./EnglishFnt')
tar.close()
# +
#Applying Transforms
dataset= torchvision.datasets.ImageFolder(
root= './EnglishFnt/English/Fnt',
transform= transforms.Compose(
[
transforms.Resize((48,48)),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
)
# +
#create fxn to split dataset
def split_data(dts, batch_size, test_split=0.3):
shuffle_dataset= True
random_seed= 42
dataset_size=len(dts)
indices= list(range(dataset_size))
split=int(np.floor(test_split*dataset_size))
if shuffle_dataset:
np.random.seed(random_seed)
np.random.shuffle(indices)
train_indices, test_indices=indices[split:], indices[:split]
test_size=len(test_indices)
indices= list(range(test_size))
split=int(np.floor(0.5*test_size))
if shuffle_dataset:
np.random.seed(random_seed)
np.random.shuffle(indices)
val_indices, test_indices= indices[split:], indices[:split]
#data samplers and loaders
train_sampler=torch.utils.data.SubsetRandomSampler(train_indices)
test_sampler= torch.utils.data.SubsetRandomSampler(test_indices)
val_sampler= torch.utils.data.SubsetRandomSampler(val_indices)
train_loader=torch.utils.data.DataLoader(dts,batch_size, sampler=train_sampler)
val_loader=torch.utils.data.DataLoader(dts, batch_size, sampler=val_sampler)
test_loader= torch.utils.data.DataLoader(dts,batch_size, sampler=test_sampler)
return train_loader, test_loader, val_loader
# -
batch_size=36
train_loader,test_loader,val_loader= split_data(dataset, batch_size,test_split=0.3)
# +
#Definint the neural network
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv1=nn.Conv2d(3,16,3)
self.conv2=nn.Conv2d(16,32,3)
self.conv3=nn.Conv2d(32,64,3)
self.fc1=nn.Linear(64*9*9,62)
self.max_pool=nn.MaxPool2d(2,2, ceil_mode=True)
self.dropout= nn.Dropout(0.2)
self.conv_bn1=nn.BatchNorm2d(48,3)
self.conv_bn2= nn.BatchNorm2d(16)
self.conv_bn3= nn.BatchNorm2d(32)
self.conv_bn4= nn.BatchNorm2d(64)
def forward(self, x):
x=F.relu(self.conv1(x))
x=self.max_pool(x)
x=self.conv_bn2(x)
x=F.relu(self.conv2(x))
x=self.max_pool(x)
x=self.conv_bn3(x)
x=F.relu(self.conv3(x))
x=self.conv_bn4(x)
x=x.view(-1,64*9*9)
x=self.dropout(x)
x=self.fc1(x)
return x
# +
#One Hot encoding
def one_hot_encode(lables, pred_size):
encoded=torch.zeros(len(lables), pred_size)
y=0
for x in lables:
encoded[y][x]=1
y+=1
return encoded
# +
#Defining the loss and optimiser
class LossFxn(torch.autograd.Function):
@staticmethod
def forward(ctx, pred, lables):
y=one_hot_encode(lables, len(pred[0]))
y=y.cpu()
ctx.save_for_backward(y, pred)
loss=-y*torch.log(pred)
loss=loss.sum()/len(lables)
return loss
@staticmethod
def backward(ctx,grad_output):
y, pred=ctx.saved_tensors
grad_input=(-y/pred)-y
grad_input= grad_input/len(pred)
return grad_input, grad_output
# -
class loss_cell(torch.nn.Module):
def __init__(self):
super(loss_cell, self).__init__()
def forward(self, pred, lables):
y=one_hot_encode(lables, len(pred[0]))
y=y.cpu()
loss=-y*torch.log(pred)
loss=loss.sum()/len(lables)
return loss
# +
neural_net=CNN()
use_cuda=True
if use_cuda and torch.cuda.is_available():
neural_net.cuda()
optimiser= optim.SGD(neural_net.parameters(), lr=0.001, momentum=0.9)
epoch=0
max_epoch=3
end=False
myloss=loss_cell()
while epoch< max_epoch and not end:
epoch+=1
total_loss=0
total_correct=0
total_val=0
total_train=0
for dataset in (train_loader):
images, lables=dataset
if use_cuda and torch.cuda.is_available():
images=images.cuda()
lables=lables.cuda()
pred=neural_net(images)
pred=F.softmax(pred)
loss=myloss(pred, lables)
total_loss+=loss.item()
total_train+=len(pred)
optimiser.zero_grad()
loss.backward()
optimiser.step()
total_correct +=pred.argmax(dim=1).eq(lables).sum()
train_acc= (total_correct*1.0)/total_train
print("Epoch: ", epoch, "Training accu:", train_acc, "Train Loss:", total_loss*1.0/len(train_loader))
if total_correct*1.0/total_train>=0.98:
end=True
total_loss=0
val_total_correct=0
for batch in (val_loader):
images, lables=batch
if use_cuda and torch.cuda.is_available():
images=images.cuda()
lables=lables.cuda()
pred=neural_net(images)
loss=F.cross_entropy(pred, lables)
total_loss+=loss.item()
total_val+=len(pred)
val_total_correct+=pred.argmax(dim=1).eq(lables).sum()
val_acc= (val_total_correct*1.0)/total_val
print("Epoch: ", epoch,"Val Acc: ", val_acc,"Val Loss:", total_loss*1.0/len(val_loader))
torch.cuda.empty_cache()
else:
neural_net.cpu()
optimiser= optim.SGD(neural_net.parameters(), lr=0.001, momentum=0.9)
epoch=0
max_epoch=10
end=False
myloss=loss_cell()
while epoch< max_epoch and not end:
epoch+=1
total_loss=0
total_correct=0
total_val=0
total_train=0
for dataset in (train_loader):
images, lables=dataset
images=images.cpu()
lables=lables.cpu()
pred=neural_net(images)
pred=F.softmax(pred)
loss=myloss(pred, lables)
total_loss+=loss.item()
total_train+=len(pred)
optimiser.zero_grad()
loss.backward()
optimiser.step()
total_correct +=pred.argmax(dim=1).eq(lables).sum()
train_acc= (total_correct*1.0)/total_train
print("Epoch: ", epoch, "Training accu:", train_acc, "Train Loss:", total_loss*1.0/len(train_loader))
if total_correct*1.0/total_train>=0.98:
end=True
total_loss=0
val_total_correct=0
for batch in (val_loader):
images, lables=batch
images=images.cpu()
lables=lables.cpu()
pred=neural_net(images)
loss=F.cross_entropy(pred, lables)
total_loss+=loss.item()
total_val+=len(pred)
val_total_correct+=pred.argmax(dim=1).eq(lables).sum()
val_acc= (val_total_correct*1.0)/total_val
print("Epoch: ", epoch,"Val Acc: ", val_acc,"Val Loss:", total_loss*1.0/len(val_loader))
# +
test_total_correct=0
total_test=0
x=0
for batch in (test_loader):
images, lables=batch
if use_cuda and torch.cuda.is_available():
images=images.cuda()
lables=lables.cuda()
else:
images=images.cpu()
lables=lables.cpu()
pred= neural_net(images)
total_test+=len(pred)
x+=1
test_total_correct+=pred.argmax(dim=1).eq(lables).sum()
print("Test Acc:", test_total_correct*1.0/total_test)
# +
path="model.pth"
torch.save(neural_net.state_dict(), path)
# +
local_machine=True
def load_model(path):
if local_machine:
checkpoint= torch.load(path, map_location='cpu')
else:
checkpoint= torch.load(path)
model=neural_net
for params in model.parameters():
params.requires_grad= False
return model
# -
model= load_model("model.pth")
# +
from PIL import Image
def process_image(image):
img_transforms=transforms.Compose(
[
transforms.Resize((48,48)),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))])
image=img_transforms(Image.open(image))
return image
# -
def display_img(image, ax=None, title=None):
if ax is None:
fig,ax=plt.subplots()
image=image.numpy().transpose((1,2,0))
mean= np.array([0.5])
std=np.array([0.5])
image= std*image+mean
ax.display_img(image)
return ax
img= process_image('./EnglishFnt/English/Fnt/Sample001/img001-00007.png')
display_img(img)
# +
def transcribe(image_path, neural_net):
image_data= process_image(image_path)
model=load_model(path)
model_p= model.eval()
inputs= Variable(image_data.unsqueeze(0))
output=model_p(inputs)
return output
| ocr_recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import Cell_BLAST as cb
import utils
os.environ["CUDA_VISIBLE_DEVICES"] = utils.pick_gpu_lowest_memory()
cb.config.RANDOM_SEED = 0
cb.config.N_JOBS = 4
fixed_model_kwargs = dict(
latent_dim=10, cat_dim=20,
epoch=500, patience=20
)
cb.__version__
# ---
# # Drosophila
# ## Ariss
ariss = cb.data.ExprDataSet.read_dataset("../../Datasets/data/Ariss/data.h5")
utils.peek(ariss, "build/eye_disc/Ariss")
ariss.obs.head()
ariss_model = cb.directi.fit_DIRECTi(
ariss, ariss.uns["seurat_genes"],
**fixed_model_kwargs
)
ariss.latent = ariss_model.inference(ariss)
ax = ariss.visualize_latent("cell_type1", scatter_kws=dict(rasterized=True))
ax.get_figure().savefig("build/eye_disc/Ariss/cell_type1.svg", dpi=utils.DPI, bbox_inches="tight")
ax = ariss.visualize_latent("cell_ontology_class", scatter_kws=dict(rasterized=True))
ax.get_figure().savefig("build/eye_disc/Ariss/cell_ontology_class.svg", dpi=utils.DPI, bbox_inches="tight")
ariss.write_dataset("build/eye_disc/Ariss/Ariss.h5")
# %%capture capio
ariss_models = [ariss_model]
for i in range(1, cb.config.N_JOBS):
print("==== Model %d ====" % i)
ariss_models.append(cb.directi.fit_DIRECTi(
ariss, ariss.uns["seurat_genes"],
**fixed_model_kwargs,
random_seed=i
))
ariss_blast = cb.blast.BLAST(ariss_models, ariss)
ariss_blast.save("build/eye_disc/Ariss")
with open("build/eye_disc/Ariss/stdout.txt", "w") as f:
f.write(capio.stdout)
with open("build/eye_disc/Ariss/stderr.txt", "w") as f:
f.write(capio.stderr)
utils.self_projection(ariss_blast, "build/eye_disc/Ariss")
# %%writefile "build/eye_disc/Ariss/predictable.txt"
cell_ontology_class
cell_type1
| Notebooks/Database/eye_disc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
from sklearn import linear_model
from matplotlib import pyplot as plt
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA,TruncatedSVD
from sklearn.datasets import load_boston
boston = load_boston()
df_x = pd.DataFrame(boston.data, columns=boston.feature_names)
df_y = pd.DataFrame(boston.target)
reg = linear_model.LinearRegression()
x_train, x_test, y_train, y_test = train_test_split(df_x,df_y,test_size=0.2, random_state=4)
reg.fit(x_train,y_train)
reg.score(x_test,y_test)
df_x.head()
pca = PCA(n_components=10, whiten='True')
x = pca.fit(df_x).transform(df_x)
pca.explained_variance_
reg = linear_model.LinearRegression()
x_train, x_test, y_train, y_test = train_test_split(x,df_y,test_size=0.2, random_state=4)
reg.fit(x_train,y_train)
reg.score(x_test,y_test)
svd = TruncatedSVD(n_components = 10)
x = svd.fit(df_x).transform(df_x)
reg = linear_model.LinearRegression()
x_train, x_test, y_train, y_test = train_test_split(x,df_y,test_size=0.2, random_state=4)
reg.fit(x_train,y_train)
reg.score(x_test,y_test)
df_x.corr()
data=pd.read_csv('mnist.csv')
data.head()
df_x = data.iloc[:,1:]
df_y = data.iloc[:,0]
x_train, x_test, y_train, y_test = train_test_split(df_x,df_y,test_size=0.2, random_state=4)
rf = RandomForestClassifier(n_estimators = 50)
rf.fit(x_train,y_train)
pred = rf.predict(x_test)
s = y_test.values
count = 0
for i in range(len(pred)):
if pred[i]==s[i]:
count = count + 1
count/float(len(pred))
pca = PCA(n_components=25, whiten='True')
x = pca.fit(df_x).transform(df_x)
x_train, x_test, y_train, y_test = train_test_split(x,df_y,test_size=0.2, random_state=4)
rf = RandomForestClassifier(n_estimators = 50)
rf.fit(x_train,y_train)
pred = rf.predict(x_test)
s = y_test.values
count = 0
for i in range(len(pred)):
if pred[i]==s[i]:
count = count + 1
count/float(len(pred))
pca.explained_variance_
pca = PCA(n_components=2, whiten='True')
x = pca.fit(df_x).transform(df_x)
x_train, x_test, y_train, y_test = train_test_split(x,df_y,test_size=0.2, random_state=4)
rf = RandomForestClassifier(n_estimators = 50)
rf.fit(x_train,y_train)
pred = rf.predict(x_test)
s = y_test.values
count = 0
for i in range(len(pred)):
if pred[i]==s[i]:
count = count + 1
count/float(len(pred))
y = df_y.values
for i in range(5000):
if y[i]==0:
plt.scatter(x[i,1],x[i,0],c='r')
else:
plt.scatter(x[i,1],x[i,0],c='g')
plt.show()
| Data_Science/Dim. Reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
def function_for_roots(x):
a = 1.01
b = -3.04
c = 2.07
return a*x**2 + b*x + c #finding the roots of ax^2 + bx + c
# Function to check if initial values are valid
# Define bisection root finding search
# +
def check_initial_values(f, x_min, x_max, tol):
#check initial guesses
y_min = f(x_min)
y_max = f(x_max)
#check that x_min and x_max contain a 0 crossing
if(y_min*y_max>=0.0):
print("no zero crossing found in the range = ",x_min,x_max)
s = "f(%f) = %f, f(%f) = %f" % (x_min,y_min,x_max,y_max)
print(s)
return 0
#if x_min is root, then return flag == 1
if(np.fabs(y_min)<tol):
return 1
#if x_max is root, return flag == 2
if(np.fabs(y_max)<tol):
return 2
#if we reach this point, the bracket is valid, and we will return 3
return 3
def bisection_root_finding(f, x_min_start, x_max_start, tol):
x_min = x_min_start
x_max = x_max_start
x_mid = 0.0
y_min = f(x_min)
y_max = f(x_max)
y_mid = 0.0
imax = 10000
i = 0
#checking initial values
flag = check_initial_values(f,x_min,x_max,tol)
if(flag==0):
print("Error in bisection root finding().")
raise ValueError('Initial values invalid',x_min,x_max)
elif(flag==1):
return x_min
elif(flag==2):
return x_max
flag = 1
while(flag):
x_mid = 0.5*(x_min+x_max)
y_mid = f(x_mid)
if(np.fabs(y_mid)<tol):
flag = 0
else:
#xmid isnt a root
#if product of fxn @ midpoint at 1 end of points is>0, replace end point
if(f(x_min)*f(x_mid)>0):
x_min = x_mid
else:
x_max = x_mid
print(x_min,f(x_min),x_max,f(x_max))
#count iteration
i += 1
if(i>=imax):
print("Exceeded max # of iterations = ",i)
s = "Min bracket f(%f) = %f" % (x_min,f(x_min))
print(s)
s = "Max bracket f(%f) = %f" % (x_max,f(x_max))
print(s)
s = "Mid bracket f(%f) = %f" % (x_mid,f(x_mid))
print(s)
raise StopIteration('Stopping iterations after ',i)
return x_mid, i
# -
# conduct the search
# +
x_min = 0.0
x_max = 1.5
tolerance = 1.0e-6
print(x_min,function_for_roots(x_min))
print(x_max,function_for_roots(x_max))
x_root, i_root = bisection_root_finding(function_for_roots,x_min,x_max,tolerance)
y_root = function_for_roots(x_root)
s = "Root found with y(%f) = %f" % (x_root,y_root)
print(s)
print(i_root)
# +
x=np.linspace(0,3,1000)
plt.plot(x,function_for_roots(x))
plt.xlim(0,3,)
plt.ylim(-0.5,2.1,)
plt.axhline(0, c='red')
plt.plot(x_root,i_root)
# -
| hw-3.ipynb |
# +
# Author: <NAME>
"""
Figure 11.16 and 11.17 in the book "Probabilistic Machine Learning: An Introduction by <NAME>"
Dependencies: spams(pip install spams), group-lasso(pip install group-lasso)
Illustration of group lasso:
To show the effectiveness of group lasso, in this code we demonstrate:
a)Actual Data b)Vanilla Lasso c)Group lasso(L2 norm) d)Group Lasso(L infinity norm)
on signal which is piecewise gaussian and on signal which is piecewise constant
we apply the regression methods to the linear model - y = XW + ε and estimate and plot W
(X)Data: 1024(rows) x 4096(dimensions)
(W)Coefficients : 4096(dimensions)x1(coefficient for the corresponding row)
(ε)Noise(simulated via N(0,1e-4)): 4096(dimensions) x 1(Noise for the corresponding row)
(y)Target Variable: 1024(rows) x 1(dimension)
##### Debiasing step #####
Lasso Regression estimator is prone to biasing
Large coefficients are shrunk towards zero
This is why lasso stands for “least absolute selection and shrinkage operator”
A simple solution to the biased estimate problem, known as debiasing, is to use a two-stage
estimation process: we first estimate the support of the weight vector (i.e., identify which elements
are non-zero) using lasso; we then re-estimate the chosen coefficients using least squares.
Sec. 11.5.3. in the book "Probabilistic Machine Learning: An Introduction by <NAME>"
for more information
"""
import numpy as np
import matplotlib.pyplot as plt
import math
import scipy.linalg
try:
from group_lasso import GroupLasso
except ModuleNotFoundError:
# %pip install group_lasso
from group_lasso import GroupLasso
try:
from sklearn import linear_model
except ModuleNotFoundError:
# %pip install scikit-learn
from sklearn import linear_model
from sklearn.metrics import mean_squared_error
try:
import spams
except ModuleNotFoundError:
# %pip install spams
import spams
from scipy.linalg import lstsq
import warnings
warnings.simplefilter(action="ignore", category=FutureWarning)
np.random.seed(0)
def generate_data(signal_type):
"""
Generate X, Y and ε for the linear model y = XW + ε
"""
dim = 2**12
rows = 2**10
n_active = 8
n_groups = 64
size_groups = dim / n_groups
# Selecting 8 groups randomly
rand_perm = np.random.permutation(n_groups)
actives = rand_perm[:n_active]
groups = np.ceil(np.transpose(np.arange(dim) + 1) / size_groups) # Group number for each column
# Generating W actual
W = np.zeros((dim, 1))
if signal_type == "piecewise_gaussian":
for i in range(n_active):
W[groups == actives[i]] = np.random.randn(len(W[groups == actives[i]]), 1)
elif signal_type == "piecewise_constant":
for i in range(n_active):
W[groups == actives[i]] = np.ones((len(W[groups == actives[i]]), 1))
X = np.random.randn(rows, dim)
sigma = 0.02
Y = np.dot(X, W) + sigma * np.random.randn(rows, 1) # y = XW + ε
return X, Y, W, groups
def groupLasso_demo(signal_type, fig_start):
X, Y, W_actual, groups = generate_data(signal_type)
# Plotting the actual W
plt.figure(0 + fig_start)
plt.plot(W_actual)
plt.title("Original (D = 4096, number groups = 64, active groups = 8)")
plt.savefig("W_actual_{}.png".format(signal_type), dpi=300)
##### Applying Lasso Regression #####
# L1 norm is the sum of absolute values of coefficients
lasso_reg = linear_model.Lasso(alpha=0.5)
lasso_reg.fit(X, Y)
W_lasso_reg = lasso_reg.coef_
##### Debiasing step #####
ba = np.argwhere(W_lasso_reg != 0) # Finding where the coefficients are not zero
X_debiased = X[:, ba]
W_lasso_reg_debiased = np.linalg.lstsq(
X_debiased[:, :, 0], Y
) # Re-estimate the chosen coefficients using least squares
W_lasso_reg_debiased_2 = np.zeros((4096))
W_lasso_reg_debiased_2[ba] = W_lasso_reg_debiased[0]
lasso_reg_mse = mean_squared_error(W_actual, W_lasso_reg_debiased_2)
plt.figure(1 + fig_start)
plt.plot(W_lasso_reg_debiased_2)
plt.title("Standard L1 (debiased 1, regularization param(L1 = 0.5), MSE = {:.4f})".format(lasso_reg_mse))
plt.savefig("W_lasso_reg_{}.png".format(signal_type), dpi=300)
##### Applying Group Lasso L2 regression #####
# L2 norm is the square root of sum of squares of coefficients
# PNLL(W) = NLL(W) + regularization_parameter * Σ(groups)L2-norm
group_lassoL2_reg = GroupLasso(
groups=groups,
group_reg=3,
l1_reg=1,
frobenius_lipschitz=True,
scale_reg="inverse_group_size",
subsampling_scheme=1,
supress_warning=True,
n_iter=1000,
tol=1e-3,
)
group_lassoL2_reg.fit(X, Y)
W_groupLassoL2_reg = group_lassoL2_reg.coef_
##### Debiasing step #####
ba = np.argwhere(W_groupLassoL2_reg != 0) # Finding where the coefficients are not zero
X_debiased = X[:, ba]
W_group_lassoL2_reg_debiased = np.linalg.lstsq(
X_debiased[:, :, 0], Y
) # Re-estimate the chosen coefficients using least squares
W_group_lassoL2_reg_debiased_2 = np.zeros((4096))
W_group_lassoL2_reg_debiased_2[ba] = W_group_lassoL2_reg_debiased[0]
groupLassoL2_mse = mean_squared_error(W_actual, W_group_lassoL2_reg_debiased_2)
plt.figure(2 + fig_start)
plt.plot(W_group_lassoL2_reg_debiased_2)
plt.title("Block-L2 (debiased 1, regularization param(L2 = 3, L1=1), MSE = {:.4f})".format(groupLassoL2_mse))
plt.savefig("W_groupLassoL2_reg_{}.png".format(signal_type), dpi=300)
##### Applying Group Lasso Linf regression #####
# To use spams library, it is necessary to convert data to fortran normalized arrays
# visit http://spams-devel.gforge.inria.fr/ for the documentation of spams library
# Linf is the supremum of all the coeifficients
# PNLL(W) = NLL(W) + regularization_parameter * Σ(groups)Linf-norm
X_normalized = np.asfortranarray(X - np.tile(np.mean(X, 0), (X.shape[0], 1)), dtype=float)
X_normalized = spams.normalize(X_normalized)
Y_normalized = np.asfortranarray(Y - np.tile(np.mean(Y, 0), (Y.shape[0], 1)), dtype=float)
Y_normalized = spams.normalize(Y_normalized)
groups_modified = np.concatenate([[i] for i in groups]).reshape(-1, 1)
W_initial = np.zeros((X_normalized.shape[1], Y_normalized.shape[1]), dtype=float, order="F")
param = {
"numThreads": -1,
"verbose": True,
"lambda2": 3,
"lambda1": 1,
"max_it": 500,
"L0": 0.1,
"tol": 1e-2,
"intercept": False,
"pos": False,
"loss": "square",
}
param["regul"] = "group-lasso-linf"
param2 = param.copy()
param["size_group"] = 64
param2["groups"] = groups_modified
(W_groupLassoLinf_reg, optim_info) = spams.fistaFlat(Y_normalized, X_normalized, W_initial, True, **param)
##### Debiasing step #####
ba = np.argwhere(W_groupLassoLinf_reg != 0) # Finding where the coefficients are not zero
X_debiased = X[:, ba[:, 0]]
W_groupLassoLinf_reg_debiased = np.linalg.lstsq(
X_debiased, Y
) # Re-estimate the chosen coefficients using least squares
W_group_lassoLinf_reg_debiased_2 = np.zeros((4096))
W_group_lassoLinf_reg_debiased_2[ba] = W_groupLassoLinf_reg_debiased[0]
groupLassoLinf_mse = mean_squared_error(W_actual, W_group_lassoLinf_reg_debiased_2)
plt.figure(3 + fig_start)
axes = plt.gca()
plt.plot(W_group_lassoLinf_reg_debiased_2)
plt.title("Block-Linf (debiased 1, regularization param(L2 = 3, L1=1), MSE = {:.4f})".format(groupLassoLinf_mse))
plt.savefig("W_groupLassoLinf_reg_{}.png".format(signal_type), dpi=300)
plt.show()
def main():
groupLasso_demo("piecewise_gaussian", fig_start=0)
groupLasso_demo("piecewise_constant", fig_start=4)
if __name__ == "__main__":
main()
| notebooks/book1/11/groupLassoDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (PPI-env)
# language: python
# name: ppi-env
# ---
# +
import pandas as pd
import numpy as np
import os
import re
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import requests
import urllib
import re
from io import StringIO
import gzip
from ftplib import FTP
import random
import math
import copy
import pickle as pkl
from toolbox import *
# %matplotlib inline
# +
cfg = load_cfg()
logVersions = load_LogVersions()
# -
# ---
# **For figures**
from figures_toolbox import *
# +
mpl.rcParams.update(mpl.rcParamsDefault)
sns.set(
context='paper',
style='ticks',
)
# %matplotlib inline
# -
mpl.rcParams.update(performancePlot_style)
# # Load data
# ## IntAct
# **Using only interactions with scores above a certain threshold (filtered)**
intact = pd.read_pickle(
os.path.join(cfg['outputPreprocessingIntAct'],
"intact_filteredScore_v{}.pkl".format(logVersions['IntAct']['preprocessed']['filtered'])
)
)
glance(intact)
allIDs_intact = pd.concat([intact.uniprotID_A,intact.uniprotID_B])
# **Also loading unfiltered set for sanity checks**
intact_all = pd.read_pickle(
os.path.join(cfg['outputPreprocessingIntAct'],
"intact_allScores_v{}.pkl".format(logVersions['IntAct']['preprocessed']['all'])
)
)
glance(intact_all)
# ## UniProt
uniprotIDs = pd.read_csv(
os.path.join(cfg['rawDataUniProt'],
"uniprot_allProteins_Human_v{}.pkl".format(logVersions['UniProt']['rawData'])),
header=None,
names=['uniprotID']
)
glance(uniprotIDs)
# ## Hubs
# +
path0 = os.path.join(
cfg['outputPreprocessingIntAct'],
"listHubs_20p_v{}.pkl".format(logVersions['IntAct']['preprocessed']['all'])
)
with open(path0, 'rb') as f:
list_hubs20 = pkl.load(f)
glance(list_hubs20)
# -
# # EDA
# +
foo = set(allIDs_intact)
baar = len(uniprotIDs)
print(f"Number of interactions: {len(intact):,}")
print("Number of unique proteins: {:,} / {:,} ({:.2%})".format(len(foo), baar, len(foo)/baar))
# +
exportIt = True
valCounts = allIDs_intact.value_counts()
print('\nNumber of interactions per protein in IntAct (cleaned)\n')
fig, ax = plt.subplots(figsize=(10,6))
ax.plot(
range(len(valCounts)), valCounts.values,
color=palette_redon[0],
lw=1.5,
)
ax.set_xlabel('')
ax.set_ylabel('Number of interactions')
ax.set_xlim(left=-200)
ax.set_ylim(bottom=-20)
ax.spines['left'].set_position(("outward", 10))
ax.spines['bottom'].set_position(("outward", 10))
ax.spines['left'].set_bounds(0, 800)
ax.spines['bottom'].set_bounds(0, 12000)
ax.grid(which='major', linewidth=.7, linestyle=':')
# print(ax.get_yticks())
plt.tight_layout()
if exportIt:
export_figure(fig, f"nInteractions_per_protein_IntAct_2")
# -
def EDA_interactionsPerProtein(df, list_hubs, onlyHub=False):
listID = pd.concat([intact.uniprotID_A,intact.uniprotID_B])
if not onlyHub:
print("-- Percentiles of number of interactions per protein:")
foo = list(np.linspace(0.1,0.9,9))+list(np.linspace(0.91,0.99,9))
print(listID.value_counts().describe(percentiles = foo))
print()
print("-- Bigger hubs:")
print(listID.value_counts()[:10])
print()
print(f'-- Official list of hubs: {len(list_hubs):,} proteins')
print()
bar = df.loc[df.uniprotID_A.isin(list_hubs)|df.uniprotID_B.isin(list_hubs)]
print(f"Contribution of the hubs (at least one protein): {len(bar):,} interactions out of {len(df):,} ({len(bar)/len(df):.2%})")
baar = df.loc[df.uniprotID_A.isin(list_hubs)&df.uniprotID_B.isin(list_hubs)]
print(f"... both proteins: {len(baar):,} interactions out of {len(df):,} ({len(baar)/len(df):.2%})")
print(f"... and {len(df)-len(bar):,} interactions with no hubs at all")
print()
EDA_interactionsPerProtein(df=intact, list_hubs=list_hubs20)
# ---
# Quick experience with uniform sampling
# +
foo = sorted(list(set(pd.concat([intact_all.uniprotID_A,intact_all.uniprotID_B]))))
bar = sampleNIPs(
IDs2sample=foo,
seed=4,
targetSampleSize=len(intact),
referenceIntAct=intact_all,
)
glance(bar)
# -
EDA_interactionsPerProtein(df=bar, list_hubs=list_hubs20, onlyHub=True)
a = len(list_hubs20)
b = len(set(allIDs_intact))
b = len(set(pd.concat([intact_all.uniprotID_A,intact_all.uniprotID_B])))
print(a, b)
print(a/b)
# # Create positive set
seed_GS = 864
# **List of proteins in the positive set**
proteinList = sorted(list(set(allIDs_intact)))
glance(proteinList)
# +
PPI = intact[['uniprotID_A','uniprotID_B']].copy()
PPI['isInteraction'] = 1
glance(PPI)
# -
# ---
# **EDA**
EDA_predSet(
PPI,
list_hubs=list_hubs20,
PosNeg=False,
trainSet=None,
Overlap=False
)
# ## Split positive train/test1/test2
PPI2 = PPI.copy()
# ----
# **Sample proteins to save for test**
# +
hubsHere = sorted(list(set(list_hubs20) & set(proteinList)))
nonHubsHere = sorted(list(set(proteinList) - set(list_hubs20)))
print(f"In this PPI set, we have {len(hubsHere):,} hubs and {len(nonHubsHere):,} non-hubs ({len(proteinList):,} total)")
assert len(proteinList) == len(hubsHere)+len(nonHubsHere)
# +
random.seed(seed_GS)
testProteins_rate = 0.13
test_hubs = random.sample(hubsHere, int(testProteins_rate*len(hubsHere)))
test_nonHubs = random.sample(nonHubsHere, int(testProteins_rate*len(nonHubsHere)))
test_proteins = test_hubs + test_nonHubs
glance(test_proteins)
# -
print(f"Total set of proteins: {len(proteinList):,}")
print(f"... {len(test_proteins):,} for no/partial overlap ({len(test_proteins)/len(proteinList):.2%})")
# ---
# **Set aside `test` PPIs**
# +
PPI2['trainTest'] = ''
glance(PPI2)
# +
AisTestProt = PPI2.uniprotID_A.isin(test_proteins).astype(int)
BisTestProt = PPI2.uniprotID_B.isin(test_proteins).astype(int)
testProtType = AisTestProt + BisTestProt
print(testProtType.value_counts())
# +
PPI2.loc[testProtType >= 1, 'trainTest'] = 'test'
print(PPI2.trainTest.value_counts())
# -
# ---
# **Set aside interactions for complete overlap**
# +
random.seed(seed_GS + 10)
CO_rate = 0.1
foo = PPI2.loc[PPI2.trainTest == '']
testCO = random.sample(list(foo.index), int(CO_rate*len(foo)))
glance(testCO)
# +
PPI2.loc[testCO,'trainTest'] = 'test'
print(PPI2.trainTest.value_counts())
# -
# ---
# **EDA**
foo = PPI2.trainTest.value_counts()
for x in ['','test']:
if x == '':
x2 = 'train'
else:
x2=x
print(f"{x2}: {foo[x]:,} PPIs ({foo[x]/len(PPI2):.2%})")
print('## Train set')
EDA_predSet(
PPI2.loc[PPI2.trainTest == ''],
list_hubs=list_hubs20,
trainSet = PPI2.loc[PPI2.trainTest == ''],
PosNeg=False,
Overlap=False
)
print('## Test set')
outEDA0 = EDA_predSet(
PPI2.loc[PPI2.trainTest == 'test'],
list_hubs=list_hubs20,
trainSet = PPI2.loc[PPI2.trainTest == ''],
PosNeg=False,
out=True,
# Overlap=False
)
# ---
# **Split between test1 and test2**
PPI3 = PPI2.copy()
# +
random.seed(seed_GS + 20)
foo = PPI3.loc[PPI3.trainTest == 'test']
test1_idx = random.sample(list(foo.index), int(0.5*len(foo)))
test2_idx = list(set(foo.index) - set(test1_idx))
glance(test1_idx)
glance(test2_idx)
assert len(set(test1_idx) & set(test2_idx)) == 0
# +
PPI3.loc[test1_idx,'trainTest'] = 'test1'
PPI3.loc[test2_idx,'trainTest'] = 'test2'
print(PPI3.trainTest.value_counts())
# -
# ---
# **EDA**
print('## Test1')
EDA_predSet(
PPI3.loc[PPI3.trainTest == 'test1'],
list_hubs=list_hubs20,
trainSet = PPI3.loc[PPI3.trainTest == ''],
PosNeg=False,
# Overlap=False
)
print('## Test2')
EDA_predSet(
PPI3.loc[PPI3.trainTest == 'test2'],
list_hubs=list_hubs20,
trainSet = PPI3.loc[PPI3.trainTest == ''],
PosNeg=False,
# Overlap=False
)
# ---
# **Allocate `train`**
PPI3.loc[PPI3.trainTest == '','trainTest'] = 'train'
foo = PPI3.trainTest.value_counts()
for x in ['train','test1','test2']:
print(f"{x}: {foo[x]:,} PPIs ({foo[x]/len(PPI3):.2%})")
PPI3.trainTest.value_counts()['train']
# ## Small experiment: showing overlap when splitting train/test randomly
glance(PPI3)
# +
random.seed(24)
test_idx = np.random.choice(
list(PPI3.index),
size=int(0.3*len(PPI3)),
replace=False,
)
train_idx = list(set(PPI3.index) - set(test_idx))
assert len(set(train_idx) & set(test_idx)) == 0
outEDA = EDA_predSet(
PPI3.loc[test_idx],
list_hubs=list_hubs20,
trainSet = PPI3.loc[train_idx],
PosNeg=False,
out=True
)
# foo_train = PPI3.loc[]
# glance(test_idx)
# glance(train_idx)
# -
# ---
# **Figure manuscript**
# +
exportIt = True
fig, ax = plt.subplots(figsize=(3.5,3.5))
size=.25
labels = ['no overlap','partial overlap','total overlap']
palette = [palette_hiroshige[i] for i in [5,7,9]]
sizes = []
for label in labels:
sizes.append(outEDA[label][1])
my_pie, texts, pct_txts = ax.pie(
sizes, radius=1,
labels=labels,
autopct='%1.1f%%',
colors = palette,
labeldistance=None,
pctdistance=.88,
textprops={'fontsize': 8},
startangle=90,
frame=True,
counterclock=False,
wedgeprops=dict(width=size, edgecolor='w')
)
# Moves the 0.1 out of the chart
# https://stackoverflow.com/questions/60228476/conditionally-moving-the-position-of-a-single-data-label-in-a-pie-chart?noredirect=1&lq=1
for patch, txt in zip(my_pie, pct_txts):
if (patch.theta2 - patch.theta1) <= 5:
# the angle at which the text is normally located
angle = (patch.theta2 + patch.theta1) / 2.
# new distance to the pie center
x = patch.r * 1.1 * np.cos(angle * np.pi / 180)
y = patch.r * 1.1 * np.sin(angle * np.pi / 180)
# move text to new position
txt.set_position((x, y))
txt.set_color('black')
pct_txts[-1].set_color('white')
pct_txts[-2].set_color('white')
sizes = []
for label in labels:
sizes.append(outEDA0[label][1])
my_pie, texts, pct_txts = ax.pie(
sizes, radius=1-size,
labels=labels,
autopct='%1.1f%%',
textprops={'fontsize': 8},
colors = palette,
labeldistance=None,
pctdistance=.82,
startangle=90,
counterclock=False,
wedgeprops=dict(width=size, edgecolor='w')
)
pct_txts[-1].set_color('white')
pct_txts[-2].set_color('white')
fig.subplots_adjust(top=1, bottom=0, right=1, left=0, hspace=0, wspace=0)
ax.margins(0,0)
ax.set(aspect="equal")
plt.legend(labels, loc='center', frameon=False,
fontsize=8.5
)
plt.tight_layout()
plt.show()
if exportIt:
export_figure(fig, 'pie_overlap_by_strategy_2')
# -
# # Sample NIPs
PPI_final = PPI3.copy()
glance(PPI_final)
# ## `train`
#
# - Weighted sampling
# - 50% positive
# +
PPI_train = PPI_final.loc[PPI_final.trainTest == 'train']
allIDs_PPI_train = pd.concat([PPI_train.uniprotID_A,PPI_train.uniprotID_B])
NIPs_train = sampleNIPs(
IDs2sample=allIDs_PPI_train.to_list(),
seed=seed_GS+30,
targetSampleSize=len(PPI_train),
referenceIntAct=intact_all
)
NIPs_train['isInteraction'] = 0
NIPs_train['trainTest'] = 'train'
# -
glance(NIPs_train)
# ## `test1`
#
# - uniform sampling
# - 50% positive (which is harder than it seems to keep 50% positive in each category!)
# +
PPI_test1 = PPI_final.loc[PPI_final.trainTest == 'test1']
glance(PPI_test1)
glance(test_proteins)
# +
_, _, isInTrain = find_overlapStatus(
df = PPI_test1,
trainSet = PPI_train
)
isInTrain.value_counts()
# +
NIPs_temp = NIPs_train.copy()
# No overlap
NIPs_test1_no = sampleNIPs(
IDs2sample=test_proteins,
seed=seed_GS+50,
targetSampleSize=isInTrain.value_counts()[0],
referenceIntAct=intact_all,
otherRefDf=NIPs_temp[['uniprotID_A','uniprotID_B']]
)
NIPs_test1_no['isInteraction'] = 0
NIPs_test1_no['trainTest'] = 'test1'
NIPs_temp = NIPs_temp.append(NIPs_test1_no)
NIPs_test1 = NIPs_test1_no
print()
# Complete overlap
NIPs_test1_co = sampleNIPs(
IDs2sample=sorted(list(set(allIDs_PPI_train))),
seed=seed_GS+60,
targetSampleSize=isInTrain.value_counts()[2],
referenceIntAct=intact_all,
otherRefDf=NIPs_temp[['uniprotID_A','uniprotID_B']]
)
NIPs_test1_co['isInteraction'] = 0
NIPs_test1_co['trainTest'] = 'test1'
NIPs_temp = NIPs_temp.append(NIPs_test1_co)
NIPs_test1 = NIPs_test1.append(NIPs_test1_co)
print()
# Partial overlap
NIPs_test1_po = sampleNIPs(
IDs2sample=(
test_proteins,
sorted(list(set(allIDs_PPI_train)))
),
seed=seed_GS+70,
targetSampleSize=isInTrain.value_counts()[1],
referenceIntAct=intact_all,
otherRefDf=NIPs_temp[['uniprotID_A','uniprotID_B']]
)
NIPs_test1_po['isInteraction'] = 0
NIPs_test1_po['trainTest'] = 'test1'
NIPs_temp = NIPs_temp.append(NIPs_test1_po)
NIPs_test1 = NIPs_test1.append(NIPs_test1_po)
# -
glance(NIPs_test1)
# +
## Sanity checks
assert len(NIPs_test1.loc[NIPs_test1.duplicated(subset=['uniprotID_A','uniprotID_B'], keep=False)]) == 0
# -
# ## `test2`
# - uniform sampling
# - 10 times more negative examples
PPI_test2 = PPI_final.loc[PPI_final.trainTest == 'test2']
glance(PPI_test2)
# +
NIPs_temp = NIPs_train.append(NIPs_test1).copy()
NIPs_test2 = sampleNIPs(
IDs2sample=proteinList,
seed=seed_GS+80,
targetSampleSize=len(PPI_test2)*10,
referenceIntAct=intact_all,
otherRefDf=NIPs_temp[['uniprotID_A','uniprotID_B']]
)
NIPs_test2['isInteraction'] = 0
NIPs_test2['trainTest'] = 'test2'
# +
## Sanity checks
assert len(NIPs_test2.loc[NIPs_test2.duplicated(subset=['uniprotID_A','uniprotID_B'], keep=False)]) == 0
# -
# # Aggregate GS
# +
# GS = PPI_final.append(NIPs_train).append(NIPs_test1)
GS = PPI_final.append(NIPs_train).append(NIPs_test1).append(NIPs_test2)
glance(GS)
# -
GS.tail()
# +
## Sanity checks
assert len(GS.loc[GS.duplicated(subset=['uniprotID_A','uniprotID_B'], keep=False)]) == 0
# -
# ---
# **EDA**
print('## Train\n')
EDA_predSet(
GS.loc[GS.trainTest == 'train'],
list_hubs=list_hubs20,
trainSet = GS.loc[GS.trainTest == 'train'],
Overlap=False
)
# +
print('## Test1\n')
EDA_predSet(
GS.loc[GS.trainTest == 'test1'],
list_hubs=list_hubs20,
trainSet = GS.loc[GS.trainTest == 'train'],
)
# +
print('## Test2\n')
EDA_predSet(
GS.loc[GS.trainTest == 'test2'],
list_hubs=list_hubs20,
trainSet = PPI3.loc[PPI3.trainTest == 'train'],
)
# -
for x in ['train','test1', 'test2']:
for y in [0,1]:
foo = GS.loc[(GS.isInteraction == y)&(GS.trainTest == x)]
allIDs = pd.concat([foo.uniprotID_A, foo.uniprotID_B])
valCounts = allIDs.value_counts()
plt.plot(
range(len(valCounts)), valCounts.values,
label=f"Label: {y}"
)
plt.xlabel('')
plt.ylabel('Number of interactions')
plt.title(f'Number of interactions per protein - {x}')
plt.yscale('log')
plt.legend()
plt.show();
# +
for x in [
# 'train',
'test1',
'test2'
]:
foo = GS.loc[(GS.isInteraction == 0)&(GS.trainTest == x)]
allIDs = pd.concat([foo.uniprotID_A, foo.uniprotID_B])
valCounts = allIDs.value_counts()
plt.plot(
range(len(valCounts)), valCounts.values,
label=x
)
plt.xlabel('')
plt.ylabel('Number of interactions')
plt.title(f'Number of interactions per protein in the negative set')
# plt.yscale('log')
plt.legend()
plt.show();
# -
GS.trainTest.value_counts()
# ## Add Similarity Measures
#
# Requires 60GB of memory
to_enrich = GS
# +
GS_SM = AddSimilarityMeasures(to_enrich, cfg=cfg, logVersions=logVersions)
glance(GS_SM)
print("success")
# -
# ## Add sequence
glance(GS_SM)
# +
sequenceData = pd.read_pickle(
os.path.join(
cfg['outputPreprocessingUniprot'],
f"sequenceData_v{logVersions['UniProt']['rawData']}--{logVersions['UniProt']['preprocessed']}.pkl"
)
)
glance(sequenceData)
# +
sequenceData.columns = ['uniprotID_A', 'sequence_A']
GS_SM_seq = GS_SM.merge(
sequenceData,
how = 'left',
on = "uniprotID_A"
)
sequenceData.columns = ['uniprotID_B', 'sequence_B']
GS_SM_seq = GS_SM_seq.merge(
sequenceData,
how = 'left',
on = "uniprotID_B"
)
glance(GS_SM_seq)
assert GS_SM_seq.isna().sum()['sequence_A'] + GS_SM_seq.isna().sum()['sequence_B'] == 0
# -
# # Export
#
# v1 is the first version (16/11/2021)
# - v1.0 uses 864 as `seed_GS`
glance(GS)
glance(GS_SM)
glance(GS_SM_seq)
versionGS = '1-0'
# +
logVersions['otherGoldStandard']['benchmarkingGS'] = versionGS
dump_LogVersions(logVersions)
# -
# ---
# **Export as pkl files**
# +
to_export = GS
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# +
to_export = GS_SM
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_v{logVersions['featuresEngineering']['similarityMeasure']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# +
to_export = GS_SM_seq
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_sequence_v{logVersions['featuresEngineering']['similarityMeasure']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# -
# ---
# **Export as csv**
# +
to_export = GS
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
# +
to_export = GS_SM
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_v{logVersions['featuresEngineering']['similarityMeasure']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
# +
to_export = GS_SM_seq
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_sequence_v{logVersions['featuresEngineering']['similarityMeasure']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
# -
# # Alternate test set with weighting sampling for Hubs investigation
# - PPI_final already done, just need to aggregate `test1` and `test2`
# - NIPs_train already done
# - NIPs_test to be done
# ## Select alternative test set
# ----
# **Group `test1` and `test2`**
PPI_alt = PPI_final.copy()
glance(PPI_alt)
PPI_alt.loc[PPI_alt.trainTest != 'train','trainTest'] = 'test'
# Sanity checks
assert PPI_final.trainTest.value_counts()['train'] == PPI_alt.trainTest.value_counts()['train']
assert (PPI_final.trainTest.value_counts()['test1']+PPI_final.trainTest.value_counts()['test2']) == PPI_alt.trainTest.value_counts()['test']
# ---
# **Sample NIPs for test**
# +
PPI_test_alt = PPI_alt.loc[PPI_alt.trainTest == 'test']
allIDs_PPI_test_alt = pd.concat([PPI_test_alt.uniprotID_A,PPI_test_alt.uniprotID_B])
NIPs_test_alt = sampleNIPs(
IDs2sample=allIDs_PPI_test_alt.to_list(),
seed=seed_GS+100,
targetSampleSize=len(PPI_test_alt),
referenceIntAct=intact_all,
otherRefDf=NIPs_train[['uniprotID_A','uniprotID_B']]
)
NIPs_test_alt['isInteraction'] = 0
NIPs_test_alt['trainTest'] = 'test'
print()
glance(NIPs_test_alt)
# -
# ---
# **Aggregate `GS_alt`**
GS_alt = PPI_alt.append(NIPs_train).append(NIPs_test_alt)
glance(GS_alt)
# +
## Sanity checks
assert len(GS_alt.loc[GS_alt.duplicated(subset=['uniprotID_A','uniprotID_B'], keep=False)]) == 0
# -
# ---
# **EDA**
# +
print('## Test\n')
EDA_predSet(
GS_alt.loc[GS_alt.trainTest == 'test'],
list_hubs=list_hubs20,
trainSet = GS_alt.loc[GS_alt.trainTest == 'train'],
)
# -
# ## Add Similarity Measures
#
# Requires 60GB of memory
# +
to_enrich = GS_alt
GS_SM_alt = AddSimilarityMeasures(to_enrich, cfg=cfg, logVersions=logVersions)
glance(GS_SM_alt)
print("success")
# -
# ## Add sequence
glance(GS_SM_alt)
# +
sequenceData = pd.read_pickle(
os.path.join(
cfg['outputPreprocessingUniprot'],
f"sequenceData_v{logVersions['UniProt']['rawData']}--{logVersions['UniProt']['preprocessed']}.pkl"
)
)
glance(sequenceData)
# +
sequenceData.columns = ['uniprotID_A', 'sequence_A']
GS_SM_alt_seq = GS_SM_alt.merge(
sequenceData,
how = 'left',
on = "uniprotID_A"
)
sequenceData.columns = ['uniprotID_B', 'sequence_B']
GS_SM_alt_seq = GS_SM_alt_seq.merge(
sequenceData,
how = 'left',
on = "uniprotID_B"
)
glance(GS_SM_alt_seq)
assert GS_SM_alt_seq.isna().sum()['sequence_A'] + GS_SM_alt_seq.isna().sum()['sequence_B'] == 0
# -
# ## Export
# **to pickle**
# +
to_export = GS_alt
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# +
to_export = GS_SM_alt
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_v{logVersions['featuresEngineering']['similarityMeasure']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# +
to_export = GS_SM_alt_seq
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_sequence_v{logVersions['featuresEngineering']['similarityMeasure']}.pkl"
)
print(export_path)
with open(export_path, 'wb') as f:
pkl.dump(to_export, f, protocol=pkl.HIGHEST_PROTOCOL)
# -
# ---
# **to csv**
# +
to_export = GS_alt
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
# +
to_export = GS_SM_alt
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_v{logVersions['featuresEngineering']['similarityMeasure']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
# +
to_export = GS_SM_alt_seq
export_path = os.path.join(
cfg['outputGoldStandard'],
f"benchmarkingGS_4hubs_v{logVersions['otherGoldStandard']['benchmarkingGS']}_similarityMeasure_sequence_v{logVersions['featuresEngineering']['similarityMeasure']}.csv"
)
print(export_path)
to_export.to_csv(export_path, sep='|', index=False)
| 2. Gold standard/Human gold standard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import spacy, nltk, gensim
from nltk.stem import WordNetLemmatizer
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
nltk.download('wordnet')
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.corpus import stopwords
from tqdm import tqdm
# -
# Reading the data
data_recipe = pd.read_json('data/full_format_recipes.json')
data_recipe.head()
data_recipe.shape
data_recipe.info()
# Let's drop the column 'desc', since we are not going to use it
data_recipe = data_recipe.drop('desc',axis=1)
# Let's also remove the rows where the column 'rating' has value 0
data_recipe = data_recipe[data_recipe.rating>0]
# Checking the number of null values for each column
data_recipe.isnull().sum()
# Let's drop all rows with at least one missing value
data_recipe = data_recipe.dropna()
# Let's drop also duplicate recipes
# +
# removing possible whitespaces on both ends
data_recipe.title = data_recipe.title.apply(lambda x: x.strip())
data_recipe = data_recipe.drop_duplicates(subset='title')
# -
# We are left with 13001 rows
data_recipe.shape
# Now, let's have a look at the distributions of the following features in our data: calories, sodium, protein and fat.
plt.figure(figsize=(15,7))
boxplot = data_recipe.boxplot(column=['calories', 'sodium', 'protein', 'fat']);
boxplot.set_title('Boxplot for calories, sodium, protein and fat')
boxplot.set_ylabel('Value')
boxplot.set_xlabel('Feature')
boxplot.set_yscale('log')
# As we can see from the plot shown above, there are outliers some of which are abnormal values (e.g. calorie values of more than 10^7). Let's remove those values.
# +
data_recipe = data_recipe[data_recipe.calories<2000]
data_recipe = data_recipe[data_recipe.sodium<3000]
data_recipe = data_recipe[data_recipe.protein<200]
data_recipe = data_recipe[data_recipe.fat<300]
# The threshold for each column is chosen based on the boxplot and the knowledge about reasonable ranges
# for each nutritional fact
# resetting index
data_recipe = data_recipe.reset_index(drop=True)
# -
# Finally, we are left with 12466 rows.
data_recipe.shape
# An example of a recipe
data_recipe.iloc[0]['ingredients']
# Second example of a recipe
data_recipe.iloc[1]['ingredients']
# Let's make the list of strings a single joined string
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: ' '.join(x))
data_recipe.iloc[0]['ingredients']
data_recipe.iloc[1]['ingredients']
# Let's now make all words lowercase
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: x.lower())
# Defining the units and some common words that need to be removed from recipes
unwanted_words = ['c', 'quarts', 'tbs', 'oz.', 'pinch', 'half', 'coarse', 't', 'liters', 'bunch', 'lb', 'T.',
'glass', 'room', 'cube', 'sprigs', 'round', 'basket', 'gal.', 'pints', 'package', 'teaspoons',
'T', 'bag', 'kilograms', 'tbsp.', 'pot', 'lb.', 'dashes', 'filets', 'jar', 'sprig', 'grill',
'dash', 'specialty', 'slice', 'temperature', 'pan', 'oz', 'kg.', 'large', 'pound', 'touch',
'yam', 'clove', 'box', 'chunk', 'gallon', 'qt.', 'lid', 'qts.', 'kg', 'mail', 'g.', 'fluid ounce',
'cups', 'qts', 'lengthwise', 't.', 'order', 'ml.', 'grocer', 'skillet', 'tsp', 'milligrams',
'milligram', 'kilogram', 'filet', 'gal', 'small', 'handful', 'thick', 'spray', 'cut', 'pt.',
'spoon', 'milliliters', 'shop', 'key', 'pint', 'mg.', 'fl. oz.', 'crosswise', 'fl oz', 'touches',
'liter', 'gr.', 'c.', 'milliliter', 'scoops', 'thin', 'l.', 'supermarket', 'cans', 'medium',
'tsp.', 'ounce', 'g', 'tablespoons', 'tbs.', 'accompaniment', 'market', 'sheet', 'center',
'gallons', 'dice', 'can', 'handfuls', 'tablespoon', 'teaspoon', 'tbsp', 'ml', 'equipment',
'pinches', 'part', 'gr', 'cloves', 'qt', 'ounces', 'grams', 'gram', 'cup', 'stick', 'sticks',
'envelope', 'scoop', 'flake', 'mg', 'l', 'pt', 'pounds', 'fluid ounces', 'quart', 'food',
'microwave', 'piece', 'inch', 'layer', 'top', 'granny', 'triangle', 'note']
# In order to extract ingredients from the recipes we are going to apply the following pipeline:
#
# 1. Keep only nouns using nltk
# 2. Remove our defined unwanted words
# 3. Do lemmatisation using nltk
# 4. Keep only unique words in the final list
# First we need to do tokenization
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: word_tokenize(x))
# Attaching a part of speech tag to each word
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: nltk.pos_tag(x))
# Keeping only nouns
data_recipe.ingredients = data_recipe.ingredients.apply(
lambda x: [word for word, pos in x if (pos == 'NN' or pos == 'NNP' or pos == 'NNS' or pos == 'NNPS')])
# Removing our defined unwanted words
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: [w for w in x if w not in unwanted_words])
# Lemmatisation step
wnl = WordNetLemmatizer()
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: [wnl.lemmatize(token) for token in x])
# Removing duplicate words from the final list
# +
# stripping
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: [each.strip('*/-.,+') for each in x])
data_recipe.ingredients = data_recipe.ingredients.apply(lambda x: list(set(x)))
# -
# The result for the first example after applying the pipeline
data_recipe.iloc[0]['ingredients']
# The result for the second example after applying the pipeline
data_recipe.iloc[1]['ingredients']
# They are not perfect but seem good enough :)
# Let's look at the distribution of the number of ingredients in recipes
ingredients = data_recipe.ingredients.copy()
l = np.array([len(elem) for elem in ingredients])
plt.figure(figsize=(15,7))
plt.hist(l,bins=65)
plt.title('Distribution of the length of recipes')
plt.xlabel('Number of ingredients in a recipe')
plt.ylabel('Count')
plt.show()
# As we can see from the plot, we have a right-skewed normal distribution.
# Given this data, we want to construct a weighted graph with nodes represented by the recipes and connect them based on the common ingredients they share. As a similarity metric between two recipes, we will use Jaccard similarity index which will serve as the weights for the graph.
# Since our data is large, we want to keep only a subset of it. Out of the available 12466 recipes, we will take only 5000 based on random sampling.
# +
np.random.seed(42)
node_size = 5000
indicies_to_take = np.random.permutation(data_recipe.shape[0])[:node_size]
# -
# Selecting the rows corresponding to the chosen 5000 recipes
new_data_recipe = data_recipe.loc[indicies_to_take]
# Now let's build our jaccard similarity matrix
# +
jaccard_similarity_matrix = np.zeros((node_size,node_size))
new_ingredients = new_data_recipe.ingredients.values
for i in tqdm(range(node_size)):
for j in range(i,node_size):
jaccard_similarity_matrix[i,j] = 1 - nltk.jaccard_distance(set(new_ingredients[i]), set(new_ingredients[j]))
jaccard_similarity_matrix[j,i] = jaccard_similarity_matrix[i,j]
# -
# Setting diagonal entries 0
np.fill_diagonal(jaccard_similarity_matrix,0)
# Let's take a look at the distribution of the obtained Jaccard similarity values.
plt.figure(figsize=(15,7))
plt.hist(jaccard_similarity_matrix.flatten(),bins=100)
plt.title('Distribution of the Jaccard similarity values')
plt.xlabel('Jaccard similarity index')
plt.ylabel('Count')
plt.xticks(np.linspace(0,1,11))
plt.show()
print('Number of edges: {}'.format(int(len(jaccard_similarity_matrix[jaccard_similarity_matrix>0])/2)))
# As we can see, we have enourmous number of edges. Most of them are weakly connected, so let's remove those edges.
# +
threshold = 0.2
jaccard_similarity_matrix[jaccard_similarity_matrix<threshold] = 0
# -
print('Number of edges now: {}'.format(int(len(jaccard_similarity_matrix[jaccard_similarity_matrix>0])/2)))
# Saving the adjacency matrix as a numpy array
np.save('adjacency_matrix.npy',jaccard_similarity_matrix)
# Saving the chosen subset of the processed data
new_data_recipe.to_json('processed_data.json')
| Data Processing and Graph Construction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from violet.hw6 import Patient
example = Patient("mark", ["fever", "cough"])
example.add_test("blood_count", [123,456,789])
example.add_test("pressure", [534,23,76])
example.tests
example.has_covid()
import pandas as pd
from violet.model import Model
from violet.preprocessor import Preprocessor
from violet.transform import Transform, Standardize, Polynomial
from violet.split import SplitClass
name = "sample_diabetes_mellitus_data.csv"
example = SplitClass(name)
train_data,test_data=example.train_test(0.1)
data_prep = Preprocessor(train_data)
data_prep.clean_nan(['age', 'gender', 'ethnicity'])
data_prep.fill_nan(['height', 'weight'])
numeric_data = test_data.select_dtypes(include = ['float', 'int']).dropna()
test_standardize = Standardize(numeric_data)
answ_standardize = test_standardize.change()
test_polynomial = Polynomial(numeric_data)
answ_polynomial = test_polynomial.change(2)
features = ['age', 'height', 'weight', 'aids', 'cirrhosis', 'hepatic_failure',
'immunosuppression', 'leukemia', 'lymphoma', 'solid_tumor_with_metastasis']
target = ['diabetes_mellitus']
test_model = Model(features, target, 500)
test_model.train(train_data)
test_model.predict(test_data.dropna())
| diabetes/.ipynb_checkpoints/question7-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print('Hello the first code')
my_name = 'ibrahim'
my_name
my_name.capitalize
my_name.capitalize()
my_name_capitalized = my_name.capitalize()
my_name_capitalized
my_fullname = '<NAME>'
my_fullname.split()
my_fullname.split()[0]
my_fullname.split()[1]
my_fullname.upper()
# # shift+TAB help kutusunu açar.
'ibrahim' * 10 * 10
# # LIST subject
my_list = [31,44,13,67,5]
my_list.append(7) #listenin sonuna 7 ekler
my_list
# echo my_list.reverse()
my_list
# ## nested list
new_list = [1,4,'a',[3,'c']] #liste içerisinde yeni bir liste eklenebilir
nested_list = new_list[3]
nested_list
new_list[3][1]
new_list[2:]
new_list[:2]
# # Dictionary
my_dictionary = {'key':'value'}
my_dictionary['key']
my_fitness_dictionary = {'key':'value', 'run':100, 'swim':200}
my_dictionary2 = {'key':'value', 'list':[2,3,5], 'key_dict':{3:5}}
my_dictionary2['key_dict'][3] #sözlük içerisindeki key_dict sözlüğündeki 3'ün karşılığını getirir
# # SET type
my_list
# +
#listede tekrar eden olmadığı için set dediğinde hepsini gösterir ama tekrar eden eklersek duplikeyi göstermez
# -
new_my_list = [7,7,5,67,13,44,31,5,31]
my_set_1 = set(new_my_list)
my_set_1
type(my_set_1)
# # TUPLE type
my_tuple = ('a',1,'b') #immutable
# +
#liste içerisindeki veriler değiştirilemez. Set veya List'ten bu yönde ayrılır.
# -
my_tuple_1 = ('a',1,3,5,1,1,1,'a','b')
my_tuple_1.count(1) #tuple içerisindeki 1'lerin adedi
| 01_Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JeffMboya/Computer-Guess-Name/blob/main/Style_transfer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="NyftRTSMuwue" outputId="2c8f53ae-cbf7-4117-bd77-2e2629625e88"
# @title Import and configure modules
import functools
import os
from matplotlib import gridspec
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
print("TF Version: ", tf.__version__)
print("TF Hub version: ", hub.__version__)
print("Eager mode enabled: ", tf.executing_eagerly())
print("GPU available: ", tf.config.list_physical_devices('GPU'))
# + id="y5Cc3dInrAVs"
# @title Define image loading and visualization functions { display-mode: "form" }
def crop_center(image):
"""Returns a cropped square image."""
shape = image.shape
new_shape = min(shape[1], shape[2])
offset_y = max(shape[1] - shape[2], 0) // 2
offset_x = max(shape[2] - shape[1], 0) // 2
image = tf.image.crop_to_bounding_box(
image, offset_y, offset_x, new_shape, new_shape)
return image
@functools.lru_cache(maxsize=None)
def load_image(image_url, image_size=(256, 256), preserve_aspect_ratio=True):
"""Loads and preprocesses images."""
# Cache image file locally.
image_path = tf.keras.utils.get_file(os.path.basename(image_url)[-128:], image_url)
# Load and convert to float32 numpy array, add batch dimension, and normalize to range [0, 1].
img = tf.io.decode_image(
tf.io.read_file(image_path),
channels=3, dtype=tf.float32)[tf.newaxis, ...]
img = crop_center(img)
img = tf.image.resize(img, image_size, preserve_aspect_ratio=True)
return img
def show_n(images, titles=('',)):
n = len(images)
image_sizes = [image.shape[1] for image in images]
w = (image_sizes[0] * 6) // 320
plt.figure(figsize=(w * n, w))
gs = gridspec.GridSpec(1, n, width_ratios=image_sizes)
for i in range(n):
plt.subplot(gs[i])
plt.imshow(images[i][0], aspect='equal')
plt.axis('off')
plt.title(titles[i] if len(titles) > i else '')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 479} id="384lQ_lqo4Kt" outputId="1edaa01e-ce43-4dc5-d921-b2f38b9f272d"
# @title Load example images { display-mode: "form" }
content_image_url = 'https://upload.wikimedia.org/wikipedia/commons/5/5f/Violinist_playing.jpg' # @param {type:"string"}
style_image_url = 'https://i.pinimg.com/originals/49/56/b2/4956b2aa75c4b2036d5d6261b7bb022c.jpg' # @param {type:"string"}
output_image_size = 384 # @param {type:"integer"}
# The content image size can be arbitrary.
content_img_size = (output_image_size, output_image_size)
# The style prediction model was trained with image size 256 and it's the
# recommended image size for the style image (though, other sizes work as
# well but will lead to different results).
style_img_size = (256, 256) # Recommended to keep it at 256.
content_image = load_image(content_image_url, content_img_size)
style_image = load_image(style_image_url, style_img_size)
style_image = tf.nn.avg_pool(style_image, ksize=[3,3], strides=[1,1], padding='SAME')
show_n([content_image, style_image], ['Content image', 'Style image'])
# + id="una6zq6Go9B8"
# Load TF Hub module.
hub_handle = 'https://tfhub.dev/google/magenta/arbitrary-image-stylization-v1-256/2'
hub_module = hub.load(hub_handle)
# + id="rmgg5xlxo_sF"
outputs = hub_module(content_image, style_image)
stylized_image = outputs[0]
# + id="bbz3KM8PpDCh"
# Stylize content image with given style image.
# This is pretty fast within a few milliseconds on a GPU.
outputs = hub_module(tf.constant(content_image), tf.constant(style_image))
stylized_image = outputs[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="dhgxvLZopFzG" outputId="56d88494-55c6-48bb-bfef-cf678100c554"
# Visualize input images and the generated stylized image.
show_n([stylized_image])
| Style_transfer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import useful standard modules. Note that geopandas is not included standard with Anaconda.
# +
# Some fairly standard modules
import os, csv, lzma
import numpy as np
import matplotlib.pyplot as plt
# The geopandas module does not come standard with anaconda,
# so you'll need to run the anaconda prompt as an administrator
# and install it via "conda install -c conda-forge geopandas".
# That installation will include pyproj and shapely automatically.
# These are useful modules for plotting geospatial data.
import geopandas as gpd
import pyproj
import shapely.geometry
# These modules are useful for tracking where modules are
# imported from, e.g., to check we're using our local edited
# versions of open_cp scripts.
import sys
import inspect
import importlib
# In order to use our local edited versions of open_cp
# scripts, we insert the parent directory of the current
# file ("..") at the start of our sys.path here.
sys.path.insert(0, os.path.abspath(".."))
# Elements from PredictCode's custom "open_cp" package
import open_cp.sources.chicago as chicago
import open_cp.sources.ukpolice as ukpolice
# Confirm we're using local versions of code
print(inspect.getfile(chicago))
print(inspect.getfile(ukpolice))
# -
# Declare location of data directory for Chicago data.
datadir = os.path.join("..", "..", "Data")
chicago.set_data_directory(datadir)
# Read data from CSV or CSV.XZ file, view first couple rows.
# +
def get_csv_data(shortfilename="chicago.csv"):
filename = os.path.join(datadir, shortfilename)
if shortfilename.endswith(".csv.xz"):
with lzma.open(filename, "rt") as f:
yield from csv.reader(f)
elif shortfilename.endswith(".csv"):
with open(filename, "rt") as f:
yield from csv.reader(f)
else:
yield None
rows = get_csv_data("chicago.csv")
print(next(rows))
print(next(rows))
# -
# Obtain polygon shapely object for South side of Chicago via custom "get_side" function.
polygon = chicago.get_side("South")
polygon
# Declare geopandas GeoDataFrame object named "South Side", set its geometry equal to the shapely polygon we obtained from the Chicago data, and set its Coordinate Reference System (CRS) to epsg:2790.
frame = gpd.GeoDataFrame({"name":["South Side"]})
frame.geometry = [polygon]
frame.crs = {"init":"epsg:2790"}
frame
# Save geopandas GeoDataFrame object as a "file" that can be reloaded later (actually a directory containing a set of corresponding files).
#
# Note that this currently generates a warning about part of the fiona module being deprecated.
frame.to_file("SouthSide")
# Obtain default burglary data from Chicago data directory. Takes the form of custom object TimedPoints.
points = chicago.default_burglary_data()
print(type(points))
# View some initial features: size of data, earliest and latest times, size of bounding box for spatial data, and the aspect ratio of that bounding box.
print("Number of timestamps: " + str(len(points.timestamps)))
print("Earliest time: " + str(points.time_range[0]))
print("Latest time: " + str(points.time_range[1]))
bbox = points.bounding_box
print("X coord range:", bbox.xmin, bbox.xmax)
print("Y coord range:", bbox.ymin, bbox.ymax)
print("Aspect ratio: " + str(bbox.aspect_ratio))
# Plot points
_, ax = plt.subplots(figsize=(10,10 * bbox.aspect_ratio))
ax.scatter(points.coords[0], points.coords[1], alpha=0.1, marker="o", s=1)
# Focus on region of downtown.
# +
mask = ( (points.xcoords >= 355000) & (points.xcoords <= 365000) &
(points.ycoords >= 575000) & (points.ycoords <= 585000) )
downtown = points[mask]
bbox = downtown.bounding_box
print("X coord range:", bbox.xmin, bbox.xmax)
print("Y coord range:", bbox.ymin, bbox.ymax)
_, ax = plt.subplots(figsize=(5, 5 * bbox.aspect_ratio))
ax.scatter(downtown.coords[0], downtown.coords[1], alpha=0.1, marker="o", s=1)
# -
# # UK Data
#
# This example uses data from January 2017, West Yorkshire. The csv file is expected to be titled "ukpolice.csv" and located in the current directory. (Not the data_dir as in the Chicago data.)
points = ukpolice.default_burglary_data()
len(points.timestamps)
bbox = points.bounding_box
_, ax = plt.subplots(figsize=(10, 10 * bbox.aspect_ratio))
ax.scatter(points.xcoords, points.ycoords, s=10, alpha=0.2)
import open_cp
projected_points = open_cp.data.points_from_lon_lat(points, epsg=7405)
bbox = projected_points.bounding_box
_, ax = plt.subplots(figsize=(10, 10 * bbox.aspect_ratio))
ax.scatter(projected_points.xcoords, projected_points.ycoords, s=10, alpha=0.2)
| sandbox/TryingExampleData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# # Quantum Key Distribution
# -
# ## 1. Introduction
#
# When Alice and Bob want to communicate a secret message (such as Bob’s online banking details) over an insecure channel (such as the internet), it is essential to encrypt the message. Since cryptography is a large area and almost all of it is outside the scope of this textbook, we will have to believe that Alice and Bob having a secret key that no one else knows is useful and allows them to communicate using symmetric-key cryptography.
#
# If Alice and Bob want to use Eve’s classical communication channel to share their key, it is impossible to tell if Eve has made a copy of this key for herself- they must place complete trust in Eve that she is not listening. If, however, Eve provides a quantum communication channel, Alice and Bob no longer need to trust Eve at all- they will know if she tries to read Bob’s message before it gets to Alice.
#
# For some readers, it may be useful to give an idea of how a quantum channel may be physically implemented. An example of a classical channel could be a telephone line; we send electric signals through the line that represent our message (or bits). A proposed example of a quantum communication channel could be some kind of fiber-optic cable, through which we can send individual photons (particles of light). Photons have a property called _polarisation,_ and this polarisation can be one of two states. We can use this to represent a qubit.
#
#
# ## 2. Protocol Overview
#
# The protocol makes use of the fact that measuring a qubit can change its state. If Alice sends Bob a qubit, and an eavesdropper (Eve) tries to measure it before Bob does, there is a chance that Eve’s measurement will change the state of the qubit and Bob will not receive the qubit state Alice sent.
# + tags=["thebelab-init"]
from qiskit import QuantumCircuit, Aer, transpile
from qiskit.visualization import plot_histogram, plot_bloch_multivector
from numpy.random import randint
import numpy as np
# -
# If Alice prepares a qubit in the state $|+\rangle$ (`0` in the $X$-basis), and Bob measures it in the $X$-basis, Bob is sure to measure `0`:
# +
qc = QuantumCircuit(1,1)
# Alice prepares qubit in state |+>
qc.h(0)
qc.barrier()
# Alice now sends the qubit to Bob
# who measures it in the X-basis
qc.h(0)
qc.measure(0,0)
# Draw and simulate circuit
display(qc.draw())
aer_sim = Aer.get_backend('aer_simulator')
job = aer_sim.run(qc)
plot_histogram(job.result().get_counts())
# -
# But if Eve tries to measure this qubit in the $Z$-basis before it reaches Bob, she will change the qubit's state from $|+\rangle$ to either $|0\rangle$ or $|1\rangle$, and Bob is no longer certain to measure `0`:
# +
qc = QuantumCircuit(1,1)
# Alice prepares qubit in state |+>
qc.h(0)
# Alice now sends the qubit to Bob
# but Eve intercepts and tries to read it
qc.measure(0, 0)
qc.barrier()
# Eve then passes this on to Bob
# who measures it in the X-basis
qc.h(0)
qc.measure(0,0)
# Draw and simulate circuit
display(qc.draw())
aer_sim = Aer.get_backend('aer_simulator')
job = aer_sim.run(qc)
plot_histogram(job.result().get_counts())
# -
# We can see here that Bob now has a 50% chance of measuring `1`, and if he does, he and Alice will know there is something wrong with their channel.
#
# The quantum key distribution protocol involves repeating this process enough times that an eavesdropper has a negligible chance of getting away with this interception. It is roughly as follows:
#
# **- Step 1**
#
# Alice chooses a string of random bits, e.g.:
#
# `1000101011010100`
#
# And a random choice of basis for each bit:
#
# `ZZXZXXXZXZXXXXXX`
#
# Alice keeps these two pieces of information private to herself.
#
# **- Step 2**
#
# Alice then encodes each bit onto a string of qubits using the basis she chose; this means each qubit is in one of the states $|0\rangle$, $|1\rangle$, $|+\rangle$ or $|-\rangle$, chosen at random. In this case, the string of qubits would look like this:
#
# $$ |1\rangle|0\rangle|+\rangle|0\rangle|-\rangle|+\rangle|-\rangle|0\rangle|-\rangle|1\rangle|+\rangle|-\rangle|+\rangle|-\rangle|+\rangle|+\rangle
# $$
#
# This is the message she sends to Bob.
#
# **- Step 3**
#
# Bob then measures each qubit at random, for example, he might use the bases:
#
# `XZZZXZXZXZXZZZXZ`
#
# And Bob keeps the measurement results private.
#
# **- Step 4**
#
# Bob and Alice then publicly share which basis they used for each qubit. If Bob measured a qubit in the same basis Alice prepared it in, they use this to form part of their shared secret key, otherwise they discard the information for that bit.
#
# **- Step 5**
#
# Finally, Bob and Alice share a random sample of their keys, and if the samples match, they can be sure (to a small margin of error) that their transmission is successful.
#
# ## 3. Qiskit Example: Without Interception
#
# Let’s first see how the protocol works when no one is listening in, then we can see how Alice and Bob are able to detect an eavesdropper. As always, let's start by importing everything we need:
# To generate pseudo-random keys, we will use the `randint` function from numpy. To make sure you can reproduce the results on this page, we will set the seed to 0:
np.random.seed(seed=0)
# We will call the length of Alice's initial message `n`. In this example, Alice will send a message 100 qubits long:
n = 100
# ### 3.1 Step 1:
#
# Alice generates her random set of bits:
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
print(alice_bits)
# At the moment, the set of bits '`alice_bits`' is only known to Alice. We will keep track of what information is only known to Alice, what information is only known to Bob, and what has been sent over Eve's channel in a table like this:
#
# | Alice's Knowledge |Over Eve's Channel| Bob's Knowledge |
# |:-----------------:|:----------------:|:---------------:|
# | alice_bits | | |
#
# ### 3.2 Step 2:
#
# Alice chooses to encode each bit on qubit in the $X$ or $Z$-basis at random, and stores the choice for each qubit in `alice_bases`. In this case, a `0` means "prepare in the $Z$-basis", and a `1` means "prepare in the $X$-basis":
# +
np.random.seed(seed=0)
n = 100
## Step 1
#Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
print(alice_bases)
# -
# Alice also keeps this knowledge private:
#
# | Alice's Knowledge |Over Eve's Channel| Bob's Knowledge |
# |:-----------------:|:----------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
#
# The function `encode_message` below, creates a list of `QuantumCircuit`s, each representing a single qubit in Alice's message:
# + tags=["thebelab-init"]
def encode_message(bits, bases):
message = []
for i in range(n):
qc = QuantumCircuit(1,1)
if bases[i] == 0: # Prepare qubit in Z-basis
if bits[i] == 0:
pass
else:
qc.x(0)
else: # Prepare qubit in X-basis
if bits[i] == 0:
qc.h(0)
else:
qc.x(0)
qc.h(0)
qc.barrier()
message.append(qc)
return message
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
# -
# We can see that the first bit in `alices_bits` is `0`, and the basis she encodes this in is the $X$-basis (represented by `1`):
print('bit = %i' % alice_bits[0])
print('basis = %i' % alice_bases[0])
# And if we view the first circuit in `message` (representing the first qubit in Alice's message), we can verify that Alice has prepared a qubit in the state $|+\rangle$:
message[0].draw()
# As another example, we can see that the fourth bit in `alice_bits` is `1`, and it is encoded in the $Z$-basis, Alice prepares the corresponding qubit in the state $|1\rangle$:
print('bit = %i' % alice_bits[4])
print('basis = %i' % alice_bases[4])
message[4].draw()
# This message of qubits is then sent to Bob over Eve's quantum channel:
#
# | Alice's Knowledge |Over Eve's Channel| Bob's Knowledge |
# |:-----------------:|:----------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
#
# ### 3.3 Step 3:
#
# Bob then measures each qubit in the $X$ or $Z$-basis at random and stores this information:
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Step 3
# Decide which basis to measure in:
bob_bases = randint(2, size=n)
print(bob_bases)
# -
# `bob_bases` stores Bob's choice for which basis he measures each qubit in.
#
# | Alice's Knowledge |Over Eve's Channel| Bob's Knowledge |
# |:-----------------:|:----------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
#
# Below, the function `measure_message` applies the corresponding measurement and simulates the result of measuring each qubit. We store the measurement results in `bob_results`.
# + tags=["thebelab-init"]
def measure_message(message, bases):
backend = Aer.get_backend('aer_simulator')
measurements = []
for q in range(n):
if bases[q] == 0: # measuring in Z-basis
message[q].measure(0,0)
if bases[q] == 1: # measuring in X-basis
message[q].h(0)
message[q].measure(0,0)
aer_sim = Aer.get_backend('aer_simulator')
result = aer_sim.run(message[q], shots=1, memory=True).result()
measured_bit = int(result.get_memory()[0])
measurements.append(measured_bit)
return measurements
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Step 3
# Decide which basis to measure in:
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
# -
# We can see that the circuit in `message[0]` (representing the 0th qubit) has had an $X$-measurement added to it by Bob:
message[0].draw()
# Since Bob has by chance chosen to measure in the same basis Alice encoded the qubit in, Bob is guaranteed to get the result `0`. For the 6th qubit (shown below), Bob's random choice of measurement is not the same as Alice's, and Bob's result has only a 50% chance of matching Alices'.
message[6].draw()
print(bob_results)
# Bob keeps his results private.
#
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
#
# ### 3.4 Step 4:
#
# After this, Alice reveals (through Eve's channel) which qubits were encoded in which basis:
#
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
#
# And Bob reveals which basis he measured each qubit in:
#
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
# | bob_bases | bob_bases | |
#
# If Bob happened to measure a bit in the same basis Alice prepared it in, this means the entry in `bob_results` will match the corresponding entry in `alice_bits`, and they can use that bit as part of their key. If they measured in different bases, Bob's result is random, and they both throw that entry away. Here is a function `remove_garbage` that does this for us:
# + tags=["thebelab-init"]
def remove_garbage(a_bases, b_bases, bits):
good_bits = []
for q in range(n):
if a_bases[q] == b_bases[q]:
# If both used the same basis, add
# this to the list of 'good' bits
good_bits.append(bits[q])
return good_bits
# -
# Alice and Bob both discard the useless bits, and use the remaining bits to form their secret keys:
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Step 3
# Decide which basis to measure in:
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
## Step 4
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
print(alice_key)
# -
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
# | bob_bases | bob_bases | |
# | alice_key | | |
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Step 3
# Decide which basis to measure in:
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
## Step 4
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
bob_key = remove_garbage(alice_bases, bob_bases, bob_results)
print(bob_key)
# -
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
# | bob_bases | bob_bases | |
# | alice_key | | bob_key |
# ### 3.5 Step 5:
#
# Finally, Bob and Alice compare a random selection of the bits in their keys to make sure the protocol has worked correctly:
# + tags=["thebelab-init"]
def sample_bits(bits, selection):
sample = []
for i in selection:
# use np.mod to make sure the
# bit we sample is always in
# the list range
i = np.mod(i, len(bits))
# pop(i) removes the element of the
# list at index 'i'
sample.append(bits.pop(i))
return sample
# -
# Alice and Bob both broadcast these publicly, and remove them from their keys as they are no longer secret:
# +
np.random.seed(seed=0)
n = 100
## Step 1
# Alice generates bits
alice_bits = randint(2, size=n)
## Step 2
# Create an array to tell us which qubits
# are encoded in which bases
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Step 3
# Decide which basis to measure in:
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
## Step 4
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
bob_key = remove_garbage(alice_bases, bob_bases, bob_results)
## Step 5
sample_size = 15
bit_selection = randint(n, size=sample_size)
bob_sample = sample_bits(bob_key, bit_selection)
print(" bob_sample = " + str(bob_sample))
alice_sample = sample_bits(alice_key, bit_selection)
print("alice_sample = "+ str(alice_sample))
# -
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
# | bob_bases | bob_bases | |
# | alice_key | | bob_key |
# | bob_sample | bob_sample | bob_sample |
# | alice_sample | alice_sample | alice_sample |
# If the protocol has worked correctly without interference, their samples should match:
bob_sample == alice_sample
# If their samples match, it means (with high probability) `alice_key == bob_key`. They now share a secret key they can use to encrypt their messages!
#
# | Alice's Knowledge | Over Eve's Channel | Bob's Knowledge |
# |:-----------------:|:------------------:|:---------------:|
# | alice_bits | | |
# | alice_bases | | |
# | message | message | message |
# | | | bob_bases |
# | | | bob_results |
# | | alice_bases | alice_bases |
# | bob_bases | bob_bases | |
# | alice_key | | bob_key |
# | bob_sample | bob_sample | bob_sample |
# | alice_sample | alice_sample | alice_sample |
# | shared_key | | shared_key |
print(bob_key)
print(alice_key)
print("key length = %i" % len(alice_key))
# ## 4. Qiskit Example: *With* Interception
#
# Let’s now see how Alice and Bob can tell if Eve has been trying to listen in on their quantum message. We repeat the same steps as without interference, but before Bob receives his qubits, Eve will try and extract some information from them. Let's set a different seed so we get a specific set of reproducible 'random' results:
np.random.seed(seed=3)
# ### 4.1 Step 1:
#
# Alice generates her set of random bits:
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
print(alice_bits)
# ### 4.2 Step 2:
#
# Alice encodes these in the $Z$ and $X$-bases at random, and sends these to Bob through Eve's quantum channel:
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
## Step 2
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
print(alice_bases)
# In this case, the first qubit in Alice's message is in the state $|+\rangle$:
message[0].draw()
# ### Interception!
#
# Oh no! Eve intercepts the message as it passes through her channel. She tries to measure the qubits in a random selection of bases, in the same way Bob will later.
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
## Step 2
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Interception!!
eve_bases = randint(2, size=n)
intercepted_message = measure_message(message, eve_bases)
print(intercepted_message)
# We can see the case of qubit 0 below; Eve's random choice of basis is not the same as Alice's, and this will change the qubit state from $|+\rangle$, to a random state in the $Z$-basis, with 50% probability of $|0\rangle$ or $|1\rangle$:
message[0].draw()
# ### 4.3 Step 3:
#
# Eve then passes on the qubits to Bob, who measures them at random. In this case, Bob chose (by chance) to measure in the same basis Alice prepared the qubit in. Without interception, Bob would be guaranteed to measure `0`, but because Eve tried to read the message he now has a 50% chance of measuring `1` instead.
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
## Step 2
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Interception!!
eve_bases = randint(2, size=n)
intercepted_message = measure_message(message, eve_bases)
## Step 3
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
message[0].draw()
# ### 4.4 Step 4:
#
# Bob and Alice reveal their basis choices, and discard the useless bits:
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
## Step 2
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Interception!!
eve_bases = randint(2, size=n)
intercepted_message = measure_message(message, eve_bases)
## Step 3
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
## Step 4
bob_key = remove_garbage(alice_bases, bob_bases, bob_results)
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
# ### 4.5 Step 5:
#
# Bob and Alice compare the same random selection of their keys to see if the qubits were intercepted:
np.random.seed(seed=3)
## Step 1
alice_bits = randint(2, size=n)
## Step 2
alice_bases = randint(2, size=n)
message = encode_message(alice_bits, alice_bases)
## Interception!!
eve_bases = randint(2, size=n)
intercepted_message = measure_message(message, eve_bases)
## Step 3
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
## Step 4
bob_key = remove_garbage(alice_bases, bob_bases, bob_results)
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
## Step 5
sample_size = 15
bit_selection = randint(n, size=sample_size)
bob_sample = sample_bits(bob_key, bit_selection)
print(" bob_sample = " + str(bob_sample))
alice_sample = sample_bits(alice_key, bit_selection)
print("alice_sample = "+ str(alice_sample))
bob_sample == alice_sample
# Oh no! Bob's key and Alice's key do not match. We know this is because Eve tried to read the message between steps 2 and 3, and changed the qubits' states. For all Alice and Bob know, this could be due to noise in the channel, but either way they must throw away all their results and try again- Eve's interception attempt has failed.
#
# ## 5. Risk Analysis
#
# For this type of interception, in which Eve measures all the qubits, there is a small chance that Bob and Alice's samples could match, and Alice sends her vulnerable message through Eve's channel. Let's calculate that chance and see how risky quantum key distribution is.
#
# - For Alice and Bob to use a qubit's result, they must both have chosen the same basis. If Eve chooses this basis too, she will successfully intercept this bit without introducing any error. There is a 50% chance of this happening.
# - If Eve chooses the *wrong* basis, i.e. a different basis to Alice and Bob, there is still a 50% chance Bob will measure the value Alice was trying to send. In this case, the interception also goes undetected.
# - But if Eve chooses the *wrong* basis, i.e. a different basis to Alice and Bob, there is a 50% chance Bob will not measure the value Alice was trying to send, and this *will* introduce an error into their keys.
#
# 
#
# If Alice and Bob compare 1 bit from their keys, the probability the bits will match is $0.75$, and if so they will not notice Eve's interception. If they measure 2 bits, there is a $0.75^2 = 0.5625$ chance of the interception not being noticed. We can see that the probability of Eve going undetected can be calculated from the number of bits ($x$) Alice and Bob chose to compare:
#
# $$ P(\text{undetected}) = 0.75^x $$
#
# If we decide to compare 15 bits as we did above, there is a 1.3% chance Eve will be undetected. If this is too risky for us, we could compare 50 bits instead, and have a 0.00006% chance of being spied upon unknowingly.
#
# You can retry the protocol again by running the cell below. Try changing `sample_size` to something low and see how easy it is for Eve to intercept Alice and Bob's keys.
# +
n = 100
# Step 1
alice_bits = randint(2, size=n)
alice_bases = randint(2, size=n)
# Step 2
message = encode_message(alice_bits, alice_bases)
# Interception!
eve_bases = randint(2, size=n)
intercepted_message = measure_message(message, eve_bases)
# Step 3
bob_bases = randint(2, size=n)
bob_results = measure_message(message, bob_bases)
# Step 4
bob_key = remove_garbage(alice_bases, bob_bases, bob_results)
alice_key = remove_garbage(alice_bases, bob_bases, alice_bits)
# Step 5
sample_size = 15 # Change this to something lower and see if
# Eve can intercept the message without Alice
# and Bob finding out
bit_selection = randint(n, size=sample_size)
bob_sample = sample_bits(bob_key, bit_selection)
alice_sample = sample_bits(alice_key, bit_selection)
if bob_sample != alice_sample:
print("Eve's interference was detected.")
else:
print("Eve went undetected!")
# -
import qiskit.tools.jupyter
# %qiskit_version_table
| notebooks/ch-algorithms/quantum-key-distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
data = """
0.794,2.41
0.828,2.39
0.889,2.31
0.917,2.25
0.964,2.00
0.998,1.92
1.10,1.80
1.21,1.61
1.31,1.47
1.36,1.42
1.43,1.33
1.55,1.26
1.61,1.22
1.69,1.18
1.90,1.11
1.96,1.08
2.15,1.04
2.30,1.02
2.47,1.03
2.64,1.04
2.83,1.06
2.92,1.07
3.37,1.11
3.86,1.19
3.94,1.19
4.21,1.23
4.43,1.27
4.80,1.35
5.19,1.43
5.48,1.50
5.79,1.55
6.09,1.62
6.28,1.65
6.55,1.73
6.73,1.77
7.17,1.85
7.42,1.91
7.69,1.96
7.99,2.00
8.44,2.07
8.98,2.19
9.48,2.25
10.1,2.28
10.9,2.24
12.1,2.14
12.9,2.10
13.8,2.06
14.4,2.04
15.1,2.03
16.4,1.98
18.9,1.92
21.6,1.87
23.5,1.83
24.2,1.82
26.5,1.77
28.3,1.76
30.7,1.72
32.6,1.70
34.2,1.68
36.5,1.65
37.7,1.63
39.8,1.60
41.5,1.60
43.4,1.59
44.6,1.57
46.3,1.54
49.3,1.53
51.8,1.52
53.5,1.50
"""
data = """
0.6012,1.998
0.6299,2.007
0.6548,2.011
0.7273,2.020
0.8047,2.025
0.8799,2.039
0.9474,2.039
1.052,2.039
1.137,2.039
1.239,2.062
1.381,2.062
1.534,2.062
1.691,2.053
1.871,2.048
2.038,2.048
2.152,2.048
2.362,2.030
2.494,2.030
2.603,2.016
2.825,2.016
3.077,2.011
3.249,2.002
3.512,1.998
3.825,1.971
4.102,1.944
4.539,1.914
5.061,1.862
5.470,1.821
5.958,1.740
6.440,1.659
6.907,1.593
7.070,1.533
7.293,1.445
7.293,1.375
7.582,1.305
7.914,1.261
8.454,1.219
8.755,1.168
8.927,1.137
9.245,1.096
9.574,1.043
9.877,0.9699
10.11,0.9227
10.35,0.8739
10.72,0.8314
10.97,0.7982
11.19,0.7698
11.45,0.7491
11.45,0.7160
11.54,0.6905
11.76,0.6539
12.00,0.6364
12.37,0.6510
12.57,0.6689
12.77,0.7031
13.17,0.7390
13.27,0.7928
13.53,0.8485
13.59,0.8899
13.85,0.9290
14.23,0.9633
14.68,1.001
15.27,1.043
16.50,1.079
17.83,1.106
19.88,1.121
22.43,1.147
25.30,1.160
27.78,1.168
29.22,1.170
30.97,1.178
34.27,1.186
36.75,1.181
39.11,1.195
41.78,1.205
44.81,1.222
48.25,1.239
51.74,1.239
54.21,1.241
57.25,1.247
60.92,1.250
64.83,1.247
67.40,1.256
"""
# +
import numpy as np
import matplotlib.pyplot as plt
KcCd = [[float(j) for j in i.split(",")] for i in data.split("\n") if i]
K,Cd = np.c_[KcCd].T
plt.plot(K,Cd)
plt.show()
# +
def poly1d(x,a):
y = 0
for i in range(0,a.size,1):
y+=a[i]*x**i
return y
def cm_kc(kc,beta):
a = np.r_[ 1.36741509e+01, -8.42861085e+01, 2.64036089e+02, -4.77831612e+02,
5.61871443e+02 , -4.57394697e+02, 2.68110607e+02 , -1.16137971e+02,
3.78175431e+01 , -9.35265180e+00 , 1.76423150e+00 , -2.53388070e-01,
2.74679857e-02 , -2.20736173e-03 , 1.27362928e-04, -4.98508748e-06,
1.18461756e-07, -1.28959671e-09]
b = np.r_[ -2.94244141e+02 , 1.09857912e+02, -1.83362700e+01 , 1.81417151e+00,
-1.18448186e-01 , 5.37606571e-03, -1.73958299e-04 , 4.04478076e-06,
-6.71033926e-08 , 7.75122027e-10 , -5.92174081e-12 , 2.68868435e-14,
-5.49128236e-17]
if (kc<=10.5):
cm = 0
for i in range(0,a.size):
cm+=a[i]*pow(kc,i)
elif (kc >10.5 and kc < 15):
k = np.log10(1.2/0.6)/np.log10(2400/14000)
cm = 1.2 * pow(beta/2400,k)
print(kc,beta,k,cm)
elif (kc >= 15):
cm = 0
for i in range(0,b.size):
cm+=b[i]*pow(kc,i)
return cm
def spline_cm(o,n,m,dn,dm,K,Cd):
id1 = (K>=n) & (K<=m)
K=K[id1]
Cd=Cd[id1]
f = np.polyfit(K,Cd,o)
print(f[::-1])
p = np.poly1d(f)
K1 = np.linspace(dn,dm,500)
Cd1 = p(K1)
#Cd2 = np.c_[[poly1d(k,f[::-1]) for k in K]]
plt.plot(K,Cd,'k.-')
Cd1[Cd1>=Cd.max()]=Cd.max()
Cd1[Cd1<=Cd.min()]=Cd.min()
plt.plot(K1[:],Cd1[:],)
#plt.plot(K,Cd2)
plt.title(o)
def get_cm_splines():
plt.figure(figsize=(20,10))
plt.locator_params(nbins=60)
spline_cm(17, K.min(), 10.5, K.min(), 10.5, K,Cd)
spline_cm(12, 15, K.max(), 15, K.max(), K,Cd)
plt.show()
#plt.savefig("Cm(Kc) - Spline.svg")
cm = np.zeros(K.size)
for i in range(K.size):
cm[i] = cm_kc(K[i])
plt.plot(K,cm)
plt.show()
# -
print(f)
print(p)
| MPI_CUDA/Untitled000.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from datetime import datetime, timedelta
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(42)
# # Portfolio Planner
#
# In this activity, you will use the iedfinance api to grab historical data for a 60/40 portfolio using `SPY` to represent the stock portion and `AGG` to represent the bonds.
from iexfinance.stocks import get_historical_data
from iexfinance.refdata import get_symbols
import iexfinance as iex
# # Data Collection
#
# In this step, you will need to use the IEX api to fetch closing prices for the `SPY` and `AGG` tickers. Save the results as a pandas DataFrame
# +
list_of_tickers = ["SPY", "AGG"]
# YOUR CODE HERE
# Set start and end datetimes of 1 year, between now and 365 days ago.
end_date = datetime.now()
start_date = end_date + timedelta(-365)
# Get 1 year's worth of historical data for AAPL
df = get_historical_data(list_of_tickers, start_date, end_date, output_format='pandas')
df.head()
# -
# # Use the `drop` function with the `level` parameter to drop extra columns in the multi-index DataFrame
# df.drop(columns=['open', 'high', 'low', 'volume'], level=1, inplace=True)
# df.head()
# Calculate the daily roi for the stocks
# YOUR CODE HERE
# Use the `drop` function with the `level` parameter to drop extra columns in the multi-index DataFrame
df.drop(columns=['open', 'high', 'low', 'volume'], level=1, inplace=True)
df.head()
# Use the `pct_change` function to calculate daily returns of `SYP` and `AGG`.
daily_returns = df.pct_change()
daily_returns.head()
# +
# Use the `mean` function to calculate the mean of daily returns for `SPY` and `AGG`, respectively
#C = daily_returns.mean()['SPY']['close']
avg_daily_return_spy = daily_returns.mean()['SPY']['close']
avg_daily_return_agg = daily_returns.mean()['AGG']['close']
avg_daily_return_agg
# -
#AVG daily return 'SPY'
avg_daily_return_spy
#AVG daily return 'AGG'
avg_daily_return_agg
# Calculate volatility AKA standard deviation
std_dev_daily_return_spy = daily_returns.std()['SPY']['close']
std_dev_daily_return_agg = daily_returns.std()['AGG']['close']
std_dev_daily_return_agg
# +
# Save the last day's closing price
spy_last_price = df['SPY']['close'][-1]
agg_last_price = df['AGG']['close'][-1]
# -
# Setup the Monte Carlo Parameters
num_simulations = 100
number_records = 252 * 30
monte_carlo = pd.DataFrame() #simulated_price_df = pd.DataFrame()
portfolio_cumulative_returns = pd.DataFrame()
# +
# Run the Monte Carlo Simulation
# Initialize empty DataFrame to hold simulated prices for each simulation
for n in range(num_simulations):
# Initialize the simulated prices list with the last closing price of `JNJ` and `MU`
simulated_spy_prices = [spy_last_price]
simulated_agg_prices = [agg_last_price]
# Simulate the returns for 252 days
for i in range(number_records):
# Calculate the simulated price using the last price within the list
simulated_spy_price = simulated_spy_prices[-1] * (1 + np.random.normal(avg_daily_return_spy, std_dev_daily_return_spy))
simulated_agg_price = simulated_agg_prices[-1] * (1 + np.random.normal(avg_daily_return_agg, std_dev_daily_return_agg))
# Append the simulated price to the list
simulated_spy_prices.append(simulated_spy_price)
simulated_agg_prices.append(simulated_agg_price)
# Append a simulated prices of each simulation to DataFrame
simulated_price_df["SPY prices"] = pd.Series(simulated_spy_prices)
simulated_price_df["AGG prices"] = pd.Series(simulated_agg_prices)
# Calculate the daily returns of simulated prices
simulated_daily_returns = simulated_price_df.pct_change()
# Set the portfolio weights (60% JNJ; 40% MU)
weights = [0.60, 0.40]
# Use the `dot` function with the weights to multiply weights with each column's simulated daily returns
portfolio_daily_returns = simulated_daily_returns.dot(weights)
# Calculate the normalized, cumulative return series
portfolio_cumulative_returns[n] = (1 + portfolio_daily_returns.fillna(0)).cumprod()
# Print records from the DataFrame
portfolio_cumulative_returns.head()
# +
# Visualize the Simulation
plot_title = f"{n+1} Simulations of Cumulative Portfolio Return Trajectories Over the Next 30 Years"
portfolio_cumulative_returns.plot(legend=None, title=plot_title)
# -
# Select the last row for the cumulative returns (cumulative returns at 30 years)
# YOUR CODE HERE
ending_cumulative_return_30 = portfolio_cumulative_returns.iloc[252*30, :]
ending_cumulative_return_30.head()
# Select the last row for the cumulative returns (cumulative returns at 20 years)
ending_cumulative_return_20 = portfolio_cumulative_returns.iloc[252*20, :]
ending_cumulative_return_20.head()
# Display the 90% confidence interval for the ending returns @ 30 yrs
confidence_interval = ending_cumulative_return_30.quantile(q=[0.05, 0.95])
confidence_interval
# +
# Visualize the distribution of the ending returns
# Use the `plot` function to create a probability distribution histogram of simulated ending prices
# with markings for a 90% confidence interval
plt.figure();
ending_cumulative_return_30.plot(kind='hist', density=True, bins=[0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
plt.axvline(confidence_interval.iloc[0], color='r')
plt.axvline(confidence_interval.iloc[1], color='r')
S
# -
# ---
# # Retirement Analysis
#
# In this section, you will use the monte carlo model to answer the following retirement planning questions:
#
# 1. What are the expected cumulative returns at 30 years for the 10th, 50th, and 90th percentiles?
# 2. Given an initial investment of `$20,000`, what is the expected portfolio return in dollars at the 10th, 50th, and 90th percentiles?
# 3. Given the current projected annual income from the Plaid analysis, will a 4% withdraw rate from the retirement portfolio meet or exceed that value at the 10th percentile?
# 4. How would a 50% increase in the initial investment amount affect the 4% retirement withdrawal?
# ### What are the expected cumulative returns at 30 years for the 10th, 50th, and 90th percentiles?
# YOUR CODE HERE
print(f"Expected cumulative portfolio return at 30 years for the 10th percentile is {round(np.percentile(ending_cumulative_return_30,10),2)}")
print(f"Expected cumulative portfolio return at 30 years for the 50th percentile is {round(np.percentile(ending_cumulative_return_30,50),2)}")
print(f"Expected cumulative portfolio return at 30 years for the 90th percentile is {round(np.percentile(ending_cumulative_return_30,90),2)}")
# ### Given an initial investment of `$20,000`, what is the expected portfolio return in dollars at the 10th, 50th, and 90th percentiles?
# +
# YOUR CODE HERE
initial_investment = 20000
end_exp_trn = 20000 * ending_cumulative_return_30
print(f"Expected portfolio return in dollars at the 10th percentile is ${(np.percentile(end_exp_trn,10))}")
print(f"Expected portfolio return in dollars at the 50th percentile is ${(np.percentile(end_exp_trn,50))}")
print(f"Expected portfolio return in dollars at the 90th percentile is ${(np.percentile(end_exp_trn,90))}")
# -
# ### Given the current projected annual income from the Plaid analysis, will a 4% withdraw rate from the retirement portfolio meet or exceed that value at the 10th percentile?
#
# Note: This is effectively saying that 90% of the expected returns will be greater than the return at the 10th percentile, so this can help measure the uncertainty about having enough funds at retirement
# YOUR CODE HERE
# ### How would a 50% increase in the initial investment amount affect the 4% retirement withdrawal?
# +
# YOUR CODE HERE
# -
# ### Optional Challenge
#
# In this section, you will calculate and plot the cumulative returns for the median and 90% confidence intervals. This plot shows the expected cumulative returns for any given day between the first day and the last day of investment.
# +
# YOUR CODE HERE
| portfolio_planner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Location Allocation problem
# *This notebook illustrates methods to solve the location-allocation problem of a supply chain network.
# *Use the virtual environment logproj_distribution.yml to run this notebook.*
# ***
# <NAME> 2020
# ### Import packages
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display, HTML
# %% append functions path
import sys; sys.path.insert(0, '..') #add the above level with the package
# -
# ### Set AS IS scenario parameters
# +
# all cost are expressed using tuples where the first element is a mean, and the second a standard deviation of the cost
# per year
# variable costs
direct_labor_cost_asis_dist=(1, 0.5)
logistic_cost_asis_dist=(0.3, 0.1)
operational_cost_asis_dist=(0.4, 0.05)
# fixed costs
depreciation_cost_as_is_dist = (10000,100)
fixed_cost_asis_dist = (1e5,1e3)
productivity_asis = 1e6 *np.ones(10)
# -
# ### Set TO-BE scenario parameters
# +
#The tobe scenario uses lists with as many elements as plants in the TOBE network
#investment costs
investment_cost_plant_1 = (0.5e6,1e2)
investment_cost_plant_2 = (1e6,1e2)
investment_cost_tobe_dist = [investment_cost_plant_1, investment_cost_plant_2]
#logistic costs
logistic_cost_tobe_plant_1=(0.4, 0.05)
logistic_cost_tobe_plant_2=(0.3,0.05)
logistic_cost_tobe_dist = [logistic_cost_tobe_plant_1 , logistic_cost_tobe_plant_2]
#operational cost
operational_cost_tobe_dist_plant_1=(0.3,0.3)
operational_cost_tobe_dist_plant_2=(0.1,0.1)
operational_cost_tobe_dist = [operational_cost_tobe_dist_plant_1, operational_cost_tobe_dist_plant_2]
#direct labour costs
direct_labor_cost_tobe_dist_plant_1=(0.9,0.1)
direct_labor_cost_tobe_dist_plant_2=(0.9,0.5)
direct_labor_cost_tobe_dist=[direct_labor_cost_tobe_dist_plant_1,direct_labor_cost_tobe_dist_plant_2]
#fixed costs
fixed_cost_tobe_dist_plant_1 = (8e4,1e3)
fixed_cost_tobe_dist_plant_2 = (7e4,2e3)
fixed_cost_tobe_dist=[fixed_cost_tobe_dist_plant_1, fixed_cost_tobe_dist_plant_2]
#depreciation costs
depreciation_cost_tobe_dist_plant_1 = (5e4,0)
depreciation_cost_tobe_dist_plant_2 = (6e4, 0)
depreciation_cost_tobe_dist = [depreciation_cost_tobe_dist_plant_1, depreciation_cost_tobe_dist_plant_2]
#############################################################################################
######################################## produttivita' ######################################
#############################################################################################
productivity_plant_1 = [400000, 450000, 0.5e6, 0.5e6, 0.5e6, 0.5e6, 0.5e6, 0.55e6, 0.6e6, 0.6e6]
productivity_plant_2 = [400000, 450000, 0.5e6, 0.5e6, 0.5e6, 0.5e6, 0.5e6, 0.55e6, 0.6e6, 0.6e6]
productivity_tobe = [productivity_plant_1, productivity_plant_2]
# -
# ### Set simulation parameters
years = 10
num_iter = 100
# ### Import cost model definition
# +
from logproj.P6_placement.locationAllocationProblem import totalCostASIS, totalCostTOBE
# -
# ### Import simulation engines
from logproj.P6_placement.locationAllocationProblem import runSimulation, runSimulationNoVariance
# ### Run montecarlo simulation
#Run montecarlo simulation
fig_result = runSimulation(num_iter,
years,
productivity_asis,
logistic_cost_asis_dist,
operational_cost_asis_dist,
direct_labor_cost_asis_dist,
depreciation_cost_as_is_dist,
fixed_cost_asis_dist,
productivity_tobe,
logistic_cost_tobe_dist,
operational_cost_tobe_dist,
direct_labor_cost_tobe_dist,
depreciation_cost_tobe_dist,
investment_cost_tobe_dist,
fixed_cost_tobe_dist)
fig_result.show()
# ### Run static simulation
# +
#Run static simulation
fig_result, df_results = runSimulationNoVariance(num_iter,
years,
productivity_asis,
logistic_cost_asis_dist,
operational_cost_asis_dist,
direct_labor_cost_asis_dist,
depreciation_cost_as_is_dist,
fixed_cost_asis_dist,
productivity_tobe,
logistic_cost_tobe_dist,
operational_cost_tobe_dist,
direct_labor_cost_tobe_dist,
depreciation_cost_tobe_dist,
investment_cost_tobe_dist,
fixed_cost_tobe_dist)
fig_result.show()
for key in df_results.keys():
print(key)
display(HTML(df_results[key].to_html()))
| examples/DIST_04 Location-Allocation problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # French Wikipedia Newcomer Welcoming Experiment Power Analysis
# [<NAME>](https://github.com/natematias)
# May/June 2019
#
# Some components of this are drawn from [github.com/natematias/poweranalysis-onlinebehavior](https://github.com/natematias/poweranalysis-onlinebehavior).
#
# Eventually, this power analysis code will ask a series of questions of [historical data prepared by <NAME>](https://docs.google.com/document/d/1zj-yIR7s7-MWEk3u9O1kQTfL0d6bF0Yz1H_CV-n1T4U/edit?usp=drive_web&ouid=117701977297551627494) and produce a series of answers used for power analysis and study design in CivilServant's research with Wikipedians on [the effects of welcoming new Wikipedians](https://meta.wikimedia.org/wiki/CivilServant%27s_Wikimedia_studies/Testing_French_Wikipedia%27s_welcome_message)
# * The experiment plan is on Overleaf: (TBD) **Experiment Plan: Welcoming Newcomers on Wikipedia
#
# This analysis will define and report the following:
#
# * Assumptions about minimum observable treatment effects for each DV
# * Reports on the statistical power, bias, and type S error rate for all possible estimators, given the above assumptions
# * Data-driven decisions:
# * Decisions about the final set of measures to use
# * Decisions about the randomization procedure
# * Decisions about the final estimators to use
# * Decisions about the sample size to specify for the experiment
# * Decisions about any stop rules to use in the experiment
# # Load Libraries
# +
options("scipen"=9, "digits"=4)
library(dplyr)
library(MASS)
library(ggplot2)
library(rlang)
library(gmodels)
library(tidyverse)
library(viridis)
library(fabricatr)
library(estimatr)
library(DeclareDesign)
library(blockTools)
library(beepr)
## Installed DeclareDesign 0.13 using the following command:
# install.packages("DeclareDesign", dependencies = TRUE,
# repos = c("http://R.declaredesign.org", "https://cloud.r-project.org"))
library(survminer)
library(survival)
## ^^ documentation: https://cran.r-project.org/web/packages/survminer/vignettes/Informative_Survival_Plots.html
## DOCUMENTATION AT: https://cran.r-project.org/web/packages/DeclareDesign/DeclareDesign.pdf
cbPalette <- c("#999999", "#E69F00", "#56B4E9", "#009E73", "#F0E442", "#0072B2", "#D55E00", "#CC79A7")
options(repr.plot.width=7, repr.plot.height=3.5)
sessionInfo()
# -
# # Load Power Analysis Dataframes and Review the Data
data.path <- "~/Tresors/CivilServant/projects/wikipedia-integration/fr-newcomer-study/datasets/"
fr.power.df <- read.csv(file.path(data.path, "french_power-analysis_dataset_sim_date_20180307_v1.csv"))
fr.power.df$user_registration <- as.Date(fr.power.df$user_registration)
fr.power.df$lang <- "fr"
simulated.treatment.date <- as.Date("20180306", "%Y%M%D")
colnames(fr.power.df)
# # Summarize Data
# +
day.range <- as.integer(max(fr.power.df$user_registration) - min(fr.power.df$user_registration))
num.accounts <- nrow(fr.power.df)
print(paste("Out of", nrow(fr.power.df), "newcomer accounts"))
print(paste("At a rate of", as.integer(num.accounts / day.range), "per day"))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, has_email=="True")) /
nrow(fr.power.df) * 100), "% have email addresses", sep=""))
cat("\n")
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_1_activation_namespace_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a namespace '0' edit after 1 day", sep=""))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_2_activation_namespace_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a namespace '0' edit after 2 days", sep=""))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_7_activation_namespace_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a namespace '0' edit after 7 days", sep=""))
cat("\n")
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_1_activation_namespace_non_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a non-0 namespace edit after 1 day", sep=""))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_2_activation_namespace_non_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a non-0 namespace edit after 2 days", sep=""))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, day_7_activation_namespace_non_0_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% made a non-0 namespace edit after 7 days", sep=""))
cat("\n")
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, response_rate_nouveaux_allposts_post_treatment>0)) /
nrow(fr.power.df) * 100), "% posted to the forum nouveaux within 7 days after registration", sep=""))
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, response_rate_nouveaux_newquestion_post_treatment>0)) /
nrow(fr.power.df) * 100), "% posted a new question to the forum nouveaux within 7 days after registration", sep=""))
cat("\n")
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, response_rate_personalization_7_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% contacted the inviting editor within 7 days after registration", sep=""))
cat("\n")
print(paste(sprintf("%0.1f",
nrow(subset(fr.power.df, response_rate_draft_7_post_treatment=="True")) /
nrow(fr.power.df) * 100), "% created a draft article within 7 days after registration", sep=""))
# -
## CREATE 0 / 1 variables out of the above variables
fr.power.df$bin_day_1_activation_namespace_0_post_treatment <- as.integer(fr.power.df$day_1_activation_namespace_0_post_treatment == "True")
fr.power.df$bin_day_2_activation_namespace_0_post_treatment <- as.integer(fr.power.df$day_2_activation_namespace_0_post_treatment == "True")
fr.power.df$bin_day_7_activation_namespace_0_post_treatment <- as.integer(fr.power.df$day_7_activation_namespace_0_post_treatment == "True")
fr.power.df$bin_day_1_activation_namespace_non_0_post_treatment <- as.integer(fr.power.df$day_1_activation_namespace_non_0_post_treatment == "True")
fr.power.df$bin_day_2_activation_namespace_non_0_post_treatment <- as.integer(fr.power.df$day_2_activation_namespace_non_0_post_treatment == "True")
fr.power.df$bin_day_7_activation_namespace_non_0_post_treatment <- as.integer(fr.power.df$day_7_activation_namespace_non_0_post_treatment == "True")
fr.power.df$bin_response_rate_nouveaux_allposts_post_treatment <- as.integer(fr.power.df$response_rate_nouveaux_allposts_post_treatment > 0 )
fr.power.df$bin_response_rate_nouveaux_newquestion_post_treatment <- as.integer(fr.power.df$response_rate_nouveaux_newquestion_post_treatment > 0 )
fr.power.df$bin_response_rate_personalization_7_post_treatment <- as.integer(fr.power.df$response_rate_personalization_7_post_treatment == "True")
fr.power.df$bin_response_rate_draft_7_post_treatment <- as.integer(fr.power.df$response_rate_draft_7_post_treatment == "True")
fr.power.df$bin_day_7_any_action <-
(fr.power.df$bin_day_7_activation_namespace_0_post_treatment +
fr.power.df$bin_day_7_activation_namespace_non_0_post_treatment +
fr.power.df$bin_response_rate_personalization_7_post_treatment +
fr.power.df$bin_response_rate_draft_7_post_treatment) > 0
summary(fr.power.df$has_gender)
# ### Summarize Labor Hours over 90 Days
# +
print("Labor Hours 90 Days After Treatment")
summary(fr.power.df$labour_hours_90_post_treatment)
hist(log1p(fr.power.df$labour_hours_90_post_treatment))
print("log1p Labor Hours 90 Days After Treatment")
summary(log1p(fr.power.df$labour_hours_90_post_treatment))
# -
labor.hour.90.dist <- fitdistr(as.integer(fr.power.df$labour_hours_90_post_treatment), densfun="negative binomial")
labor.hour.90.dist
# # Configure Power Analysis
source.path <- "~/CivilServant-Wikipedia-Analysis/power-analysis"
source(file.path(source.path, "general-power-analysis-utils.R"))
current.treat.any.action.7.day.rate <- nrow(subset(fr.power.df, bin_day_7_any_action > 0 )) / nrow(fr.power.df)
current.treat.any.action.7.day.rate
# +
## overall scenario:
## current treatment is slightly better than control
## alt treatment is slightly better than current treatment
pa.config <- data.frame(
pa.label = "fr.experiment",
n.max = 100000,
n.min = 30000,
## ANY ACTION AFTER 7 DAYS
## since we are working from the treatment condition
## we imagine the control and other treatments from this point
aa7.ctl <- current.treat.any.action.7.day.rate - 0.01,
aa7.current.treat <- current.treat.any.action.7.day.rate,
aa7.new.treat <- current.treat.any.action.7.day.rate + 0.01,
## LABOR HOURS
lh.current.treat.mu = labor.hour.90.dist$estimate[['mu']],
lh.current.treat.theta = labor.hour.90.dist$estimate[['size']],
lh.ctl.effect.irr = exp(-0.1053), # 0.9 incidence rate ratio
lh.new.treat.effect.irr = exp(0.0953) # 1.1 incidence rate ratio
)
# -
diagnose.experiment <- function( n.size, cdf, sims.count = 500, bootstrap.sims.count = 500){
design <- declare_population(N = n.size) +
declare_potential_outcomes(
## ANY ACTION AFTER 7 DAYS
AA7_Z_0 = rbinom(n=n.size, 1, cdf$aa7.ctl),
AA7_Z_1 = rbinom(n=n.size, 1, cdf$aa7.current.treat),
AA7_Z_2 = rbinom(n=n.size, 1, cdf$aa7.new.treat),
# LABOR HOURS AFTER 90 DAYS
LH90_Z_0 = rnegbin(n=N, mu = cdf$lh.current.treat.mu + mu.diff.from.mu.irr(
cdf$lh.current.treat.mu,
cdf$lh.ctl.effect.irr),
theta = cdf$lh.current.treat.theta),
LH90_Z_1 = rnegbin(n=N, mu = cdf$lh.current.treat.mu,
theta = cdf$lh.current.treat.theta),
LH90_Z_2 = rnegbin(n=N, mu = cdf$lh.current.treat.mu + mu.diff.from.mu.irr(
cdf$lh.current.treat.mu,
cdf$lh.new.treat.effect.irr),
theta = cdf$lh.current.treat.theta)
) +
declare_assignment(num_arms = 3,
conditions = (c("0", "1", "2"))) +
## we compare 0 to 1 in this simulation because 1 is the baseline
declare_estimand(ate_LH90_0_1 = log(cdf$lh.ctl.effect.irr)) +
declare_estimand(ate_LH90_2_1 = log(cdf$lh.new.treat.effect.irr)) +
declare_estimand(ate_AA7_1_0_dm = aa7.current.treat - aa7.ctl) +
declare_estimand(ate_AA7_2_0_dm = aa7.new.treat - aa7.ctl) +
declare_estimand(ate_AA7_2_1_dm = aa7.new.treat - aa7.current.treat) +
declare_reveal(outcome_variables=c("LH90", "AA7")) +
declare_estimator(estimand = "ate_LH90_0_1", label="NB LH90 0_1", handler = tidy_estimator(function(data){
data$Z <- factor(data$Z, levels = c("1", "0", "2"))
m <- glm.nb(formula = LH90 ~ Z, data)
out <- subset(tidy(m), term == "Z0")
transform(out,
conf.low = estimate - 1.96*std.error,
conf.high = estimate + 1.96*std.error
)
})) +
declare_estimator(estimand = "ate_LH90_2_1", label="NB LH90 2_1", handler = tidy_estimator(function(data){
data$Z <- factor(data$Z, levels = c("1", "0", "2"))
m <- glm.nb(formula = LH90 ~ Z, data)
out <- subset(tidy(m), term == "Z2")
transform(out,
conf.low = estimate - 1.96*std.error,
conf.high = estimate + 1.96*std.error
)})) +
declare_estimator(AA7 ~ Z, condition1="0", condition2="1", estimand = "ate_AA7_1_0_dm", label="DM AA7 1_0") +
declare_estimator(AA7 ~ Z, condition1="0", condition2="2", estimand = "ate_AA7_2_0_dm", label="DM AA7 2_0") +
declare_estimator(AA7 ~ Z, condition1="1", condition2="2", estimand = "ate_AA7_2_1_dm", label="DM AA7 2_1")
# declare_estimator(estimand = "ate_AA7_2_1_dm", label="DM AA7 2_1", handler = tidy_estimator(function(data){
# data$Z <- factor(data$Z, levels = c("1", "0", "2"))
# m <- difference_in_means(formula = AA7 ~ Z, data)
# out <- subset(tidy(m), term == "Z0")
# }))
diagnosis <- diagnose_design(design, sims = sims.count,
bootstrap_sims = bootstrap.sims.count)
diagnosis
}
# # Conduct Power Analysis (Iterative)
# +
#diagnose.experiment(100,pa.config, sims.count = 100)
# -
interval = 5000
power.iterate.df <- iterate.for.power(pa.config,
diagnosis.method=diagnose.experiment,
iteration.interval = interval)
ggplot(power.iterate.df, aes(n, power, color=estimator_label)) +
## CHART SUBSTANCE
geom_line() +
geom_point() +
## LABELS AND COSMETICS
geom_hline(yintercept=0.8, size=0.25) +
theme_bw(base_size = 12, base_family = "Helvetica") +
theme(axis.text.x = element_text(angle=45, hjust = 1)) +
scale_y_continuous(breaks = seq(0,1,0.1), limits = c(0,1), labels=scales::percent) +
scale_x_continuous(breaks = seq(pa.config$n.min,pa.config$n.max,interval)) +
scale_color_viridis(discrete=TRUE) +
xlab("sample size") +
ggtitle("Statistical Power Associated with Estimators")
# # Diagnose Two Arm Experiment
diagnose.two.arm.experiment <- function( n.size, cdf, sims.count = 500, bootstrap.sims.count = 500){
design <- declare_population(N = n.size) +
declare_potential_outcomes(
## ANY ACTION AFTER 7 DAYS
AA7_Z_0 = rbinom(n=n.size, 1, cdf$aa7.ctl),
AA7_Z_1 = rbinom(n=n.size, 1, cdf$aa7.current.treat),
# LABOR HOURS AFTER 90 DAYS
LH90_Z_0 = rnegbin(n=N, mu = cdf$lh.current.treat.mu + mu.diff.from.mu.irr(
cdf$lh.current.treat.mu,
cdf$lh.ctl.effect.irr),
theta = cdf$lh.current.treat.theta),
LH90_Z_1 = rnegbin(n=N, mu = cdf$lh.current.treat.mu,
theta = cdf$lh.current.treat.theta)
) +
declare_assignment(num_arms = 2,
conditions = (c("0", "1"))) +
## we compare 0 to 1 in this simulation because 1 is the baseline
declare_estimand(ate_LH90_0_1 = log(cdf$lh.ctl.effect.irr)) +
# declare_estimand(ate_LH90_2_1 = log(cdf$lh.new.treat.effect.irr)) +
declare_estimand(ate_AA7_1_0_dm = aa7.current.treat - aa7.ctl) +
# declare_estimand(ate_AA7_2_0_dm = aa7.new.treat - aa7.ctl) +
# declare_estimand(ate_AA7_2_1_dm = aa7.new.treat - aa7.current.treat) +
declare_reveal(outcome_variables=c("LH90", "AA7")) +
declare_estimator(estimand = "ate_LH90_0_1", label="NB LH90 0_1", handler = tidy_estimator(function(data){
data$Z <- factor(data$Z, levels = c("1", "0"))
m <- glm.nb(formula = LH90 ~ Z, data)
out <- subset(tidy(m), term == "Z0")
transform(out,
conf.low = estimate - 1.96*std.error,
conf.high = estimate + 1.96*std.error
)
})) +
# declare_estimator(estimand = "ate_LH90_2_1", label="NB LH90 2_1", handler = tidy_estimator(function(data){
# data$Z <- factor(data$Z, levels = c("1", "0", "2"))
# m <- glm.nb(formula = LH90 ~ Z, data)
# out <- subset(tidy(m), term == "Z2")
# transform(out,
# conf.low = estimate - 1.96*std.error,
# conf.high = estimate + 1.96*std.error
# )})) +
declare_estimator(AA7 ~ Z, condition1="0", condition2="1", estimand = "ate_AA7_1_0_dm", label="DM AA7 1_0")
# declare_estimator(AA7 ~ Z, condition1="0", condition2="2", estimand = "ate_AA7_2_0_dm", label="DM AA7 2_0") +
# declare_estimator(AA7 ~ Z, condition1="1", condition2="2", estimand = "ate_AA7_2_1_dm", label="DM AA7 2_1")
# declare_estimator(estimand = "ate_AA7_2_1_dm", label="DM AA7 2_1", handler = tidy_estimator(function(data){
# data$Z <- factor(data$Z, levels = c("1", "0", "2"))
# m <- difference_in_means(formula = AA7 ~ Z, data)
# out <- subset(tidy(m), term == "Z0")
# }))
diagnosis <- diagnose_design(design, sims = sims.count,
bootstrap_sims = bootstrap.sims.count)
diagnosis
}
interval = 5000
two.arm.power.iterate.df <- iterate.for.power(pa.config,
diagnosis.method=diagnose.two.arm.experiment,
iteration.interval = interval)
ggplot(two.arm.power.iterate.df, aes(n, power, color=estimator_label)) +
## CHART SUBSTANCE
geom_line() +
geom_point() +
## LABELS AND COSMETICS
geom_hline(yintercept=0.8, size=0.25) +
theme_bw(base_size = 12, base_family = "Helvetica") +
theme(axis.text.x = element_text(angle=45, hjust = 1)) +
scale_y_continuous(breaks = seq(0,1,0.1), limits = c(0,1), labels=scales::percent) +
scale_x_continuous(breaks = seq(pa.config$n.min,pa.config$n.max,interval)) +
scale_color_viridis(discrete=TRUE) +
xlab("sample size") +
ggtitle("Statistical Power Associated with Estimators")
# # Diagnose Two Arm Stop Rule
# +
## overall scenario:
## current treatment is slightly better than control
## alt treatment is slightly better than current treatment
pa.stoprule.config <- data.frame(
pa.label = "fr.experiment",
n.max = 30000,
n.min = 10000,
## ANY ACTION AFTER 7 DAYS
## since we are working from the treatment condition
## we imagine the control and other treatments from this point
aa7.ctl <- current.treat.any.action.7.day.rate - 0.05,
aa7.current.treat <- current.treat.any.action.7.day.rate,
aa7.new.treat <- current.treat.any.action.7.day.rate + 0.05,
## LABOR HOURS
lh.current.treat.mu = labor.hour.90.dist$estimate[['mu']],
lh.current.treat.theta = labor.hour.90.dist$estimate[['size']],
lh.ctl.effect.irr = exp(-0.1053), # 0.9 incidence rate ratio
lh.new.treat.effect.irr = exp(0.0953) # 1.1 incidence rate ratio
)
# -
interval = 5000
two.arm.power.stoprule.iterate.df <- iterate.for.power(pa.stoprule.config,
diagnosis.method=diagnose.two.arm.experiment,
iteration.interval = interval)
interval = 5000
two.arm.power.stoprule.iterate.df.sub.3k <- iterate.for.power(pa.stoprule.config,
diagnosis.method=diagnose.two.arm.experiment,
iteration.interval = interval)
# +
total.two.arm.power.stoprule.iterate.df <- rbind(two.arm.power.stoprule.iterate.df, two.arm.power.stoprule.iterate.df.sub.3k)
ggplot(total.two.arm.power.stoprule.iterate.df, aes(n, power, color=estimator_label)) +
## CHART SUBSTANCE
geom_line() +
geom_point() +
## LABELS AND COSMETICS
geom_hline(yintercept=0.8, size=0.25) +
theme_bw(base_size = 12, base_family = "Helvetica") +
theme(axis.text.x = element_text(angle=45, hjust = 1)) +
scale_y_continuous(breaks = seq(0,1,0.1), limits = c(0,1), labels=scales::percent) +
scale_x_continuous(breaks = seq(pa.stoprule.config$n.min,pa.config$n.max,interval)) +
scale_color_viridis(discrete=TRUE) +
xlab("sample size") +
ggtitle("Statistical Power Associated with Estimators")
# -
# # Proposal for French Wikipedia Newcomer Study
# * We propose a two-arm study involving:
# * 60,000 participants
# * (with 950 participants per day, this is 63 days)
# * a stop rule to check partway through if withdrawing the newcomer message causes any kind of harm
# * 17.5% day-one activation rate
| power-analysis/fr-newcomer-study-2019/fr-newcomer-power-analysis-spring-2019.R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature engineering
# ## Brief
# 选择kaggle上一个经典的[泰坦尼克号之灾问题](https://www.kaggle.com/c/titanic)作为数据源,做一些特征工程的练习,同时用logistics model作为baseline,并且复习一些decision tree。
#
# ## Description
# > One of the reasons that the shipwreck led to such loss of life was that there were not enough lifeboats for the passengers and crew. Although there was some element of luck involved in surviving the sinking, some groups of people were more likely to survive than others, such as women, children, and the upper-class.
#
# > In this challenge, we ask you to complete the analysis of what sorts of people were likely to survive. In particular, we ask you to apply the tools of machine learning to predict which passengers survived the tragedy.
# ## 开始撸
# ### 引入需要的包
import numpy as np#科学计算
import pandas as pd#数据分析
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
from sklearn import cross_validation
from sklearn.metrics import accuracy_score
# ### 读入数据
#
dataset = pd.read_csv('../input/kaggle_titanic_data/train.csv',header=0)
# ## 先要分析数据,做特征工程
# ### 读入内存后,先看一眼数据长啥样
dataset.head(10)
# ### 看一下,各字段的数据类型
print(dataset.dtypes)
# ### 各字段的含义
# PassengerId => 乘客ID
#
# Pclass => 乘客等级(1/2/3等舱位)
#
# Name => 乘客姓名
#
# Sex => 性别
#
# Age => 年龄
#
# SibSp => 堂兄弟/妹个数
#
# Parch => 父母与小孩个数
#
# Ticket => 船票信息
#
# Fare => 票价
#
# Cabin => 客舱
#
# Embarked => 登船港口
#
# Survived => 是否存活
# ### 看一下数据的规模
print(dataset.shape)
# **<font color=red>总结一下:总共有891行,12列数据</font>**<br>
# **<font color=red>看一下,有没有脏数据,看看有没有缺省值</font>**<br>
dataset.count()
# 总数据条数应该是891行,发现Age,Cabin,Embarked这三个column是空的,对于缺省的数据,要采取一部分策略填充缺省值。
# ### 统计一下,粗略的看一眼数据的分布情况
dataset.describe()
# **<font color=red>mean字段告诉我们,有0.383838的人最后获救了,2/3等舱的人数比1等舱要多,平均乘客年龄大概是29.7岁。</font>**<br>
# **<font color=red>然并卵,画画图看看吧</font>**<br>
dataset.head()
# ## 数据分析
# +
mpl.rcParams['axes.titlesize'] = 8
mpl.rcParams['xtick.labelsize'] = 8
mpl.rcParams['ytick.labelsize'] = 8
mpl.rcParams['axes.labelsize'] = 10
mpl.rcParams['xtick.major.size'] = 0
mpl.rcParams['ytick.major.size'] = 0
# 创建一个 10 * 10 点(point)的图,并设置分辨率为 80
fig = plt.figure(figsize=(10,10), dpi=100)
# Customer survived distribution
# 在整张图上加入一个子图,131的意思是在一个1行3列的子图中的第一张
ax = plt.subplot2grid((3, 4), (0, 0), colspan=1)
#ax = fig.add_subplot(421)
xticks = np.arange(2)*2
# Series.value_counts返回的是该Series对象中独一无二的元素的个数
survived = dataset.Survived.value_counts()
bar_width = 1
# 画柱状图,横轴是两类标签的位置,纵轴是人数,定义柱的宽度,同时设置柱的边缘为透明
bars = ax.bar(xticks, survived, width=bar_width, edgecolor='none')
# 设置y轴的标题
ax.set_ylabel('Number of survived')
# x轴每个标签的具体位置,设置为每个柱的中央
ax.set_xticks(xticks+bar_width/2-0.5)
# 设置每个标签的名字
ax.set_xticklabels(['Notsurvived', 'Survived'])
# 设置x轴的范围
ax.set_xlim([bar_width/2-1, 3.5-bar_width/2])
# 设置标题
plt.title('Distribution of survived')
# 给每个bar分配指定的颜色
colors = ['#7199cf', '#4fc4aa', '#e1a7a2']
for bar, color in zip(bars, colors):
bar.set_color(color)
# Customer pclass distribution
#ax = fig.add_subplot(4,2,2)
ax = plt.subplot2grid((3, 4), (0, 2), colspan=1)
xticks = np.arange(3)*2
pclass = dataset.Pclass.value_counts()
bar_width = 1
bars = ax.bar(xticks, pclass, width=bar_width, edgecolor='none')
ax.set_ylabel('Number of custormer')
ax.set_xticks(xticks+bar_width/2-0.5)
ax.set_xticklabels(['3', '1', '2'])
plt.title('Distribution of pclass')
colors = ['#7199cf', '#4fc4aa', '#e1a7a2']
for bar, color in zip(bars, colors):
bar.set_color(color)
# Age-Survived distribution
#ax = fig.add_subplot(412)
ax = plt.subplot2grid((3, 4), (1, 0), colspan=1)
plt.scatter(dataset.Survived, dataset.Age)
ax.set_ylim([0, 90])
plt.title('Distribution of Age-Survived')
ax.set_ylabel('age')
# 设置标题
plt.title('Distribution of survived')
# 给每个bar分配指定的颜色
colors = ['#7199cf', '#4fc4aa', '#e1a7a2']
for bar, color in zip(bars, colors):
bar.set_color(color)
# Age-pclass distribution
ax = plt.subplot2grid((3, 4), (1, 2), colspan=3)
dataset.Age[dataset.Pclass == 1].plot(kind='kde')
dataset.Age[dataset.Pclass == 2].plot(kind='kde')
dataset.Age[dataset.Pclass == 3].plot(kind='kde')
plt.title('Distribution of Age-pclass')
ax.set_ylabel('Number of survived')
ax.set_xlabel('age')
ax = plt.subplot2grid((3, 4), (2, 0), colspan=3)
dataset.Embarked.value_counts().plot(kind='bar')
plt.title('Number of people boarding at boarding ports')
plt.ylabel('number of customer')
# show
plt.show()
# -
# **<font color=red>看图说话</font>**<br>
# 在图上可以看出来:
#
# 被救的人300多点,不到半数;
#
# 3等舱乘客灰常多;遇难和获救的人年龄似乎跨度都很广;
#
# 3个不同的舱年龄总体趋势似乎也一致,2/3等舱乘客20岁多点的人最多,1等舱40岁左右的最多;
# **<font color=red>具体的还得具体统计分析</font>**<br>
# ### 属性与获救结果的关联统计
# **<font color=red>仓位与获救情况分析</font>**<br>
fig2 = plt.figure()
mpl.rcParams['axes.titlesize'] = 12
fig2.set(alpha=0.2)
not_survived = dataset.Pclass[dataset.Survived == 0].value_counts()
survived = dataset.Pclass[dataset.Survived == 1].value_counts()
df = pd.DataFrame({'survived' : survived, 'not_survived' : not_survived})
df.plot(kind='bar', stacked=True)
plt.title('Relationship between plcass and survive')
plt.xlabel('Plcass')
plt.ylabel('count')
plt.show()
# **<font color=red>可以看到仓位越好,获救的概率越大,头等舱大于2等舱,2等舱大于3等舱</font>**<br>
#
# **<font color=red>分析一下性别与获救的关系</font>**<br>
fig3 = plt.figure()
mpl.rcParams['axes.titlesize'] = 12
mpl.rcParams['xtick.labelsize'] = 12
fig3.set(alpha=0.2)
not_survived = dataset.Sex[dataset.Survived == 0].value_counts()
survived = dataset.Sex[dataset.Survived == 1].value_counts()
df = pd.DataFrame({'survived' : survived, 'not_survived' : not_survived})
df.plot(kind='bar', stacked=True)
plt.title('Relationship between Sex and survive')
plt.xlabel('Sex')
plt.ylabel('count')
plt.show()
# **<font color=red>发现女性(female)明显获救的比例比男性获救的比例高。</font>**<br>
#
# **<font color=red>下面综合分析一下,女性Sex,仓位Pclass,与survive三个属性之间的关系</font>**<br>
# +
mpl.rcParams['xtick.labelsize'] = 9
mpl.rcParams['ytick.labelsize'] = 12
mpl.rcParams['axes.titlesize'] = 11
fig4 = plt.figure(figsize=(10,10), dpi=100)
'''
fig4 = plt.figure()
ax1 = fig4.add_subplot(141)
df1 = dataset.Survived[dataset.Pclass == 1][dataset.Sex == 'female'].value_counts()
df1.plot(kind='bar', label="female highclass", color='#FA2479')
ax1.set_xticklabels(["S", "NS"], rotation=0)
plt.title('pclass1')
'''
def plot_sub(fig, row, col, pclass, sex, title, pcolor='#FA2479'):
ax = plt.subplot2grid((3, 5), (row, col), colspan=1)
df = dataset.Survived[dataset.Pclass == pclass][dataset.Sex == sex].value_counts()
df.plot(kind='bar', color=pcolor)
ax.set_xticklabels(["S", "NS"], rotation=0)
plt.title(title)
plot_sub(fig4, 0, 0, 1, 'female', 'pclass1 female')
plot_sub(fig4, 0, 2, 2, 'female', 'pclass2 female', pcolor='blue')
plot_sub(fig4, 0, 4, 3, 'female', 'pclass3 female', pcolor='green')
plot_sub(fig4, 2, 0, 1, 'male', 'pclass1 male')
plot_sub(fig4, 2, 2, 2, 'male', 'pclass2 male', pcolor='blue')
plot_sub(fig4, 2, 4,3, 'male', 'pclass3 male', pcolor='green')
plt.show()
# -
fig5 = plt.figure()
S = dataset.Embarked[dataset.Survived == 1].value_counts()
N = dataset.Embarked[dataset.Survived == 0].value_counts()
df = pd.DataFrame({'Survived' : S, 'NotSurvived' : N})
df.plot(kind='bar', stacked=True)
plt.show()
# **<font color=red>貌似C登船口幸存者比例较高</font>**<br>
fig5 = plt.figure()
S = dataset.SibSp[dataset.Survived == 1].value_counts()
N = dataset.SibSp[dataset.Survived == 0].value_counts()
df = pd.DataFrame({'Survived' : S, 'NotSurvived' : N})
df.plot(kind='bar', stacked=True)
plt.show()
# **<font color=red>亲戚关系没看出来,留作备选特征,Ticket应该是标准的特征,先放弃。看一下Cabin客舱</font>**<br>
#dataset.info()
dataset.head()
dataset.Cabin.value_counts()
# 先这么放着了。下面做数据预处理
#
# ## 数据预处理
# ### 先补全缺失特征
#
# **<font color=red>处理缺失特征的四个基本原则</font>**<br>
# > 1.如果缺值的样本占总数比例极高,我们可能就直接舍弃了,作为特征加入的话,可能反倒带入noise,影响最后的结果了
#
# > 2.如果缺值的样本适中,而该属性非连续值特征属性(比如说类目属性),那就把NaN作为一个新类别,加到类别特征中
#
# > 3.如果缺值的样本适中,而该属性为连续值特征属性,有时候我们会考虑给定一个step(比如这里的age,我们可以考虑每隔2/3岁为一个步长),然后把它离散化,之后把NaN作为一个type加到属性类目中。
#
# > 4.有些情况下,缺失的值个数并不是特别多,那我们也可以试着根据已有的值,拟合一下数据,补充上。
#
#
# **<font color=red>可以参考特征处理章节</font>**<br>
dataset.info()
# **<font color=red>看到Age,Cabin,Embarked这三个特征存在缺省值。</font>**<br>
#
#
# * Age和Embarked准备用现有数据拟合上;
# * Cabin准备将缺失和不缺失的数据分为两类;
#
#
# ### 补全Age先
# **<font color=red>用sklearn 补全Age</font>**<br>
# +
from sklearn.ensemble import RandomForestRegressor
import copy
def set_missing_ages(df):
df_tmp = copy.deepcopy(df)
data = df_tmp[['Age', 'Pclass', 'Parch', 'SibSp', 'Fare']]
know_age = data[data.Age.notnull()].as_matrix()
unknow_age = data[data.Age.isnull()].as_matrix()
y = know_age[: , 0]
X = know_age[: , 1:]
regr = RandomForestRegressor(random_state=0, n_estimators=2000, n_jobs=-1)
regr.fit(X, y)
predictAge = regr.predict(unknow_age[:, 1:])
df_tmp.loc[(dataset.Age.isnull()), 'Age'] = predictAge
return df_tmp, regr
def set_Cabin(df):
df_tmp = copy.deepcopy(df)
df_tmp.loc[(dataset.Cabin.isnull()), 'Cabin'] = 'NO'
df_tmp.loc[(dataset.Cabin.notnull()), 'Cabin'] = 'YES'
return df_tmp
dataset2, regr = set_missing_ages(dataset)
dataset2 = set_Cabin(dataset2)
# -
dataset2.info()
#
# **<font color=red>Okay,现在把Age,Cabin两值的缺省处理完了,再看看数据</font>**<br>
dataset2.head(10)
# ### One-Hot Encoding
# 在实际的机器学习的应用任务中,特征有时候并不总是连续值,有可能是一些分类值,如性别可分为“male”和“female”。在机器学习任务中,对于这样的特征,通常我们需要对其进行特征数字化。
#
# 在这个问题中,我们要把Sex、Cabin、Pclass和Embarked这4个特征数字化。
#
# Cabin原本一个属性维度,因为其取值可以是[‘yes’,’no’],而将其平展开为’Cabin_yes’,’Cabin_no’两个属性。
# * 原本Cabin取值为yes的,在此处的”Cabin_yes”下取值为1,在”Cabin_no”下取值为0;
# * 原本Cabin取值为no的,在此处的”Cabin_yes”下取值为0,在”Cabin_no”下取值为1;
#
cabin_dummies = pd.get_dummies(dataset2['Cabin'], prefix='Cabin')
sex_dummies = pd.get_dummies(dataset2['Sex'], prefix='Sex')
pclass_dummies = pd.get_dummies(dataset2['Pclass'], prefix='Pclass')
embarked_dummies = pd.get_dummies(dataset2['Embarked'], prefix='Embarked')
dataset3 = copy.deepcopy(dataset2)
dataset3 = pd.concat([dataset3, cabin_dummies, sex_dummies, pclass_dummies, embarked_dummies], axis=1)
dataset3.drop(['Name','Cabin', 'Sex', 'Pclass', 'Embarked', 'Ticket'], axis=1, inplace=True)
dataset3.head()
# 这回数据处理的有点样了,不过Age和Fare特征还没处理好,Age和Fare两个属性的数值幅度变化很大,这里我们做归一化,把这连个变量scaling(缩放)到[-1,1]之间。
import sklearn.preprocessing as preprocessing
scaler = preprocessing.StandardScaler()
age_scale_param = scaler.fit(dataset3.Age)
dataset3['Age_scaled'] = scaler.fit_transform(dataset3['Age'], age_scale_param)
fare_scale_param = scaler.fit(dataset3['Fare'])
dataset3['Fare_scaled'] = scaler.fit_transform(dataset3['Fare'], fare_scale_param)
dataset3.head(10)
# **<font color=red>Okay,特征工程做到这里基本差不多了</font>**<br>
#
# ## 建模
# ### 用sklean的logistics regression做个baseline
#
# 把需要的feature字段取出来,转成numpy格式,使用scikit-learn中的LogisticRegression建模。
# +
from sklearn import linear_model
train_df = dataset3.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
train_np = train_df.as_matrix()
X = train_np[:, 1:]
y = train_np[:, 0]
lr_regr = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
lr_regr.fit(X, y)
# -
# ### 按照上面的步骤把测试集数据也做特征工程
# +
data_test = pd.read_csv("../input/kaggle_titanic_data/test.csv")
data_test.loc[ (data_test.Fare.isnull()), 'Fare' ] = 0
# 接着我们对test_data做和train_data中一致的特征变换
# 首先用同样的RandomForestRegressor模型填上丢失的年龄
tmp_df = data_test[['Age','Fare', 'Parch', 'SibSp', 'Pclass']]
null_age = tmp_df[data_test.Age.isnull()].as_matrix()
# 根据特征属性X预测年龄并补上
X = null_age[:, 1:]
predictedAges = regr.predict(X)
data_test.loc[ (data_test.Age.isnull()), 'Age' ] = predictedAges
data_test = set_Cabin(data_test)
dummies_Cabin = pd.get_dummies(data_test['Cabin'], prefix= 'Cabin')
dummies_Embarked = pd.get_dummies(data_test['Embarked'], prefix= 'Embarked')
dummies_Sex = pd.get_dummies(data_test['Sex'], prefix= 'Sex')
dummies_Pclass = pd.get_dummies(data_test['Pclass'], prefix= 'Pclass')
df_test = pd.concat([data_test, dummies_Cabin, dummies_Embarked, dummies_Sex, dummies_Pclass], axis=1)
df_test.drop(['Pclass', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked'], axis=1, inplace=True)
df_test['Age_scaled'] = scaler.fit_transform(df_test['Age'], age_scale_param)
df_test['Fare_scaled'] = scaler.fit_transform(df_test['Fare'], fare_scale_param)
# -
# **<font color=red>运行模型预测测试集数据</font>**<br>
test = df_test.filter(regex='Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
predictions = lr_regr.predict(test)
print(type(predictions))
print(predictions)
# **<font color=red>保存预测结果</font>**<br>
result = pd.DataFrame({'PassengerId':data_test['PassengerId'].as_matrix(), 'Survived':predictions.astype(np.int32)})
result.to_csv("../output/Titanic_data/logistic_regression_predictions.csv", index=False)
# **<font color=red>在kaggle上make a submission一下baseline结果</font>**<br>
from IPython.display import Image
Image(filename="D805EAA4-5FCA-4CB1-B788-BEF90B34D9F3.png")
# 在baseline之后,继续挖掘其它属性。
# 1. Name和Ticket两个属性被舍弃了,因为几乎每一条记录都是一个完全不同的值,我们并没有找到很直接的处理方式;
# 2. 年龄的拟合本身也未必是一件非常靠谱的事情,而且老人和小孩在救援过程中必然是受照顾的对象,所以年龄属性按区段分属性也许是好的选择
tp = pd.DataFrame({'col': list(train_df.columns[1:]) , 'val': list(lr_regr.coef_.T)})
print(tp)
# 分析一下:
# 1. Sex特征:是Sex_female的系数是1.95,Sex_male的系数是-0.677,说明女性获救的几率很高;
# 2. Cabin特征:有统计记录的获救几率高,这个特征有待继续挖掘;
# 3. Pclass特征:Pclass_1一等舱的乘客获救几率高,Pclass_3获救几率降低;
# 4. Embarked特征:显示Embarked_S登船口上的乘客,幸存可能性低,与我们前期观察到的特征不符,可能考虑去掉Embarked特征试试;
# 5. Age特征:呈现负相关,似乎年龄越小获救率越高,需继续挖掘;
# 6. Fare特征:呈微弱正相关,票越贵获救几率越高?需继续观察;
# ## 交叉验证(cross validation)
# 不能每做一次调整就make a submission,然后根据结果来判定这次调整的好坏;
#
# 这么做cross validation:把train.csv分成两部分,一部分用于训练需要的模型,另外一部分数据上看预测算法的效果。
#
# 我们用scikit-learn的cross_validation来帮我们完成小数据集上的这个工作。
#
# 先简单看看cross validation情况下的打分
# **<font color=red>用留出法把训练集数据按照7:3划分,然后训练模型,在结果集上预测。预测结果与正确结果对比,找到bad case。然后肉眼看bad case。</font>**<br>
from sklearn import cross_validation
# 分割数据,按照 训练数据:cv数据 = 7:3的比例
split_train, split_cv = cross_validation.train_test_split(dataset3, test_size=0.3, random_state=0)
train_df2 = split_train.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
# 生成模型
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
clf.fit(train_df2.as_matrix()[:,1:], train_df2.as_matrix()[:,0])
# 对cross validation数据进行预测
cv_df = split_cv.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass_.*')
predictions = clf.predict(cv_df.as_matrix()[:,1:])
origin_data_train = pd.read_csv("../input/kaggle_titanic_data/train.csv")
bad_cases = origin_data_train.loc[origin_data_train['PassengerId'].isin(split_cv[predictions != cv_df.as_matrix()[:,0]]['PassengerId'].values)]
bad_cases
# 1. Age属性不使用现在的拟合方式,而是根据名称中的『Mr』『Mrs』『Miss』等的平均值进行填充。
# 2. Age不做成一个连续值属性,而是使用一个步长进行离散化,变成离散的类目feature。
# 3. Cabin再细化一些,对于有记录的Cabin属性,我们将其分为前面的字母部分(我猜是位置和船层之类的信息) 和 后面的数字部分(应该是房间号,有意思的事情是,如果你仔细看看原始数据,你会发现,这个值大的情况下,似乎获救的可能性高一些)。
# 4. Pclass和Sex俩太重要了,我们试着用它们去组出一个组合属性来试试,这也是另外一种程度的细化。
# 5. 单加一个Child字段,Age<=12的,设为1,其余为0(你去看看数据,确实小盆友优先程度很高啊)
# 6. 如果名字里面有『Mrs』,而Parch>1的,我们猜测她可能是一个母亲,应该获救的概率也会提高,因此可以多加一个Mother字段,此种情况下设为1,其余情况下设为0
# 7. 登船港口可以考虑先去掉试试(Q和C本来就没权重,S有点诡异)
# 8. 把堂兄弟/兄妹 和 Parch 还有自己 个数加在一起组一个Family_size字段(考虑到大家族可能对最后的结果有影响)
# 9. Name是一个我们一直没有触碰的属性,我们可以做一些简单的处理,比如说男性中带某些字眼的(‘Capt’, ‘Don’, ‘Major’, ‘Sir’)可以统一到一个Title,女性也一样。
# ## Learning curve
#
# ### 画Learning curve
# +
from sklearn.model_selection import learning_curve
def plot_learning_curve(estimator, X, y, ylim=None, cv=3, train_sizes=np.linspace(.05, 1., 20), plot=True):
"""
画出data在某模型上的learning curve.
参数解释
----------
estimator : 你用的分类器。
title : 表格的标题。
X : 输入的feature,numpy类型
y : 输入的target vector
ylim : tuple格式的(ymin, ymax), 设定图像中纵坐标的最低点和最高点
cv : 做cross-validation的时候,数据分成的份数,其中一份作为cv集,其余n-1份作为training(默认为3份)
n_jobs : 并行的的任务数(默认1)
"""
train_sizes, train_scores, test_scores = learning_curve(estimator, X, y, train_sizes=train_sizes, cv=cv, verbose=0)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
if plot:
plt.figure()
plt.title('title')
#if ylim is not None:
# plt.ylim(*ylim)
plt.ylim(0.90,0.76)
plt.xlabel("The number of train samples")
plt.ylabel("Scores")
plt.gca().invert_yaxis()
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std, train_scores_mean + train_scores_std,
alpha=0.1, color="b")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std, test_scores_mean + test_scores_std,
alpha=0.1, color="r")
plt.plot(train_sizes, train_scores_mean, 'o-', color="b", label="Train socres")
plt.plot(train_sizes, test_scores_mean, 'o-', color="r", label="valid scores")
plt.legend(loc="best")
plt.draw()
plt.show()
plt.gca().invert_yaxis()
midpoint = ((train_scores_mean[-1] + train_scores_std[-1]) + (test_scores_mean[-1] - test_scores_std[-1])) / 2
diff = (train_scores_mean[-1] + train_scores_std[-1]) - (test_scores_mean[-1] - test_scores_std[-1])
return midpoint, diff
plot_learning_curve(lr_regr, X, y)
# -
# ## 模型融合
# ### bagging
# 每次次取训练集的一个subset,做训练,这样,虽然用的是同一个机器学习算法,但是得到的模型却是不一样的;同时,因为没有任何一份子数据集是全的,因此即使出现过拟合,也是在子训练集上出现过拟合,而不是全体数据上,这样做一个融合,可能对最后的结果有一定的帮助.
# +
from sklearn.ensemble import BaggingRegressor
train_df = dataset3.filter(regex='Survived|Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*|Mother|Child|Family|Title')
train_np = train_df.as_matrix()
# y即Survival结果
y = train_np[:, 0]
# X即特征属性值
X = train_np[:, 1:]
# fit到BaggingRegressor之中
clf = linear_model.LogisticRegression(C=1.0, penalty='l1', tol=1e-6)
bagging_clf = BaggingRegressor(clf, n_estimators=20, max_samples=0.8, max_features=1.0, bootstrap=True, bootstrap_features=False, n_jobs=-1)
bagging_clf.fit(X, y)
test = df_test.filter(regex='Age_.*|SibSp|Parch|Fare_.*|Cabin_.*|Embarked_.*|Sex_.*|Pclass.*|Mother|Child|Family|Title')
predictions = bagging_clf.predict(test)
result = pd.DataFrame({'PassengerId':data_test['PassengerId'].as_matrix(), 'Survived':predictions.astype(np.int32)})
result.to_csv("../output/Titanic_data/logistic_regression_bagging_predictions.csv", index=False)
# -
# ## 总结
# 还有部分特征没有继续挖掘:
# 1. Age属性不使用现在的拟合方式,而是根据名称中的『Mr』『Mrs』『Miss』等的平均值进行填充。
# 2. Age不做成一个连续值属性,而是使用一个步长进行离散化,变成离散的类目feature。
# 3. Cabin再细化一些,对于有记录的Cabin属性,我们将其分为前面的字母部分(我猜是位置和船层之类的信息) 和 后面的数字部分(应该是房间号,有意思的事情是,如果你仔细看看原始数据,你会发现,这个值大的情况下,似乎获救的可能性高一些)。
# 4. Pclass和Sex俩太重要了,我们试着用它们去组出一个组合属性来试试,这也是另外一种程度的细化。
# 5. 单加一个Child字段,Age<=12的,设为1,其余为0(你去看看数据,确实小盆友优先程度很高啊)
# 6. 如果名字里面有『Mrs』,而Parch>1的,我们猜测她可能是一个母亲,应该获救的概率也会提高,因此可以多加一个Mother字段,此种情况下设为1,其余情况下设为0
# 7. 登船港口可以考虑先去掉试试(Q和C本来就没权重,S有点诡异)
# 8. 把堂兄弟/兄妹 和 Parch 还有自己 个数加在一起组一个Family_size字段(考虑到大家族可能对最后的结果有影响)
# 9. Name是一个我们一直没有触碰的属性,我们可以做一些简单的处理,比如说男性中带某些字眼的(‘Capt’, ‘Don’, ‘Major’, ‘Sir’)可以统一到一个Title,女性也一样。
| practice/code/feature_engineering/.ipynb_checkpoints/feature_engineering-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Gauss-Hermite Quadrature
# ## Efficient numerical integration method with weight function exp(-x^2)
# ## You need this for implementing Kennedy's method
# There are two versions:
# * [Probabilists’ Gauss-Hermite module](https://docs.scipy.org/doc/numpy-1.13.0/reference/routines.polynomials.hermite_e.html):
# integration weight is the standard normal PDF: exp(-x^2/2)
#
# * [Physicists’ Gauss-Hermite module](https://docs.scipy.org/doc/numpy-1.13.0/reference/routines.polynomials.hermite.html):
# integration weight is exp(-x^2)
#
# We mostly use __Probabilists’ Gauss-Hermite module__. You still need to devide the weight by sqrt(2*pi) for use with normal PDF
import numpy as np
import numpy.polynomial as nppoly
import scipy
import scipy.stats as ss
import scipy.special as spsp
import matplotlib.pyplot as plt
const = 1/np.sqrt(2.0*np.pi)
z, w = nppoly.hermite_e.hermegauss(deg=20)
w = w*const
print(z)
print(w)
pdf = ss.norm.pdf(z)
plt.plot(z, np.log(w))
plt.plot(z, np.log(pdf))
plt.grid()
plt.show()
# ## Exact integration of polynomials with degree upto _2*deg-1_
z, w = nppoly.hermite_e.hermegauss(deg=3)
w = w*const
sum(w)
# Let's test on the moments of normal distribution
deg = np.array([2,4,6,8,10,12,14])
moments = [sum(z**2 * w), sum(z**4 * w), sum(z**6 * w), sum(z**8 * w), sum(z**10 * w), sum(z**12 * w), sum(z**14 * w)]
print(moments)
# luckily we know the exact answer: (2*deg-1)!!
spsp.factorial2([1,3,5,7,9,11,13])
# Find out upto which degree integration is correct
deg[np.abs(moments - spsp.factorial2([1,3,5,7,9,11,13])) < 0.1 ]
# # Overall GHQ is very accurate for integrating smooth functions
# Let's test on Geometric Brownian Motion:
#
# $ S_T = S_0 exp\left(\sigma\sqrt{T} z - \frac12 \sigma^2 T\right)$
spot = 100
texp = 2
vol = 0.2
z = np.linspace(-5,5,10)
price = spot * np.exp(vol*np.sqrt(texp)*z - 0.5*vol*vol*texp)
print(price)
# Let's check the expectation of the prices are same as 100 (assuming 0 interest rate)
z, w = nppoly.hermite_e.hermegauss(deg=10)
w = w*const
price = spot * np.exp(vol*np.sqrt(texp)*z - 0.5*vol*vol*texp)
price_mean = sum(price * w)
price_mean - 100
plt.plot(price, w, 'o-')
plt.grid()
plt.show()
# ## Gauss-general Laguerre quadrature.
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.special.roots_genlaguerre.html
# +
scale = 1
n_quad = 10
x, w = spsp.roots_genlaguerre(n_quad, alpha=2)
x *= scale
w /= w.sum()
x, w
# -
# ## Gauss-Legendre quadrature.
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.special.roots_legendre.html
# +
scale = 1
n_quad = 10
x, w = spsp.roots_legendre(n_quad)
x *= scale
w /= w.sum()
x, w
# -
# # Gauss-Jacobi Quadrature
#
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.special.roots_jacobi.html
# +
scale = 1
n_quad = 10
x, w = spsp.roots_jacobi(n_quad, alpha=1, beta=1.5)
x *= scale
w /= w.sum()
x, w
# -
| py/HW3/Demo_GHQ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + papermill={"duration": 13.193777, "end_time": "2021-12-05T05:26:07.020684", "exception": false, "start_time": "2021-12-05T05:25:53.826907", "status": "completed"} tags=[]
pip install langdetect
# + papermill={"duration": 9.026077, "end_time": "2021-12-05T05:26:16.066330", "exception": false, "start_time": "2021-12-05T05:26:07.040253", "status": "completed"} tags=[]
pip install bpemb
# + papermill={"duration": 1.776296, "end_time": "2021-12-05T05:26:17.864298", "exception": false, "start_time": "2021-12-05T05:26:16.088002", "status": "completed"} tags=[]
import numpy as np
from nltk.corpus import stopwords
import pandas as pd
import re
import nltk
from tqdm import tqdm
import string
from collections import Counter
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import TweetTokenizer
from langdetect import detect
import emoji
from bpemb import BPEmb
# + papermill={"duration": 0.126798, "end_time": "2021-12-05T05:26:18.012342", "exception": false, "start_time": "2021-12-05T05:26:17.885544", "status": "completed"} tags=[]
# Read DataFrame From Local Path
# Change the path to local path where you store kaggle dataset twcs.csv
df = pd.read_csv("../input/customer-support-on-twitter/twcs/twcs.csv")
df
# + papermill={"duration": 0.03048, "end_time": "2021-12-05T05:26:18.064710", "exception": false, "start_time": "2021-12-05T05:26:18.034230", "status": "completed"} tags=[]
# Create function to filter out text that are not written in English
def keepEnglish(d):
mask = []
for i,doc in enumerate(d["text"]) :
try:
if doc and doc and 'en' == detect(doc) :
mask.append(True)
else:
mask.append(False)
except Exception:
mask.append(False)
return mask
# + papermill={"duration": 58.183228, "end_time": "2021-12-05T05:27:16.270476", "exception": false, "start_time": "2021-12-05T05:26:18.087248", "status": "completed"} tags=[]
# Convert to lower case and only keep English words
mask = keepEnglish(df)
df = df[mask]
df['text'] = df["text"].str.lower()
df["text"]
# + papermill={"duration": 0.057484, "end_time": "2021-12-05T05:27:16.350660", "exception": false, "start_time": "2021-12-05T05:27:16.293176", "status": "completed"} tags=[]
# Only Keep First asked questions and their reponses
df_q= df[pd.isnull(df.in_response_to_tweet_id) & df.inbound]
df_qa = pd.merge(df_q, df, left_on='tweet_id', right_on='in_response_to_tweet_id')
qa_tweets = df_qa[["text_x","text_y"]]
qa_tweets.columns = ["questions","answers"]
qa_tweets
# + papermill={"duration": 4.987127, "end_time": "2021-12-05T05:27:21.360945", "exception": false, "start_time": "2021-12-05T05:27:16.373818", "status": "completed"} tags=[]
bpemb_en = BPEmb(lang="en", dim=100)
def clean_tweet(tweet):
# Clean Tweet, remove emoji, symbols and tokenize
lemmatizer = WordNetLemmatizer()
english_stopwords = set(stopwords.words("english"))
digit_chr = re.compile("^[a-zA-Z0-9@_!#$%^&*()<>?/\|}{~:,.]*$")
tweet = re.sub(r'https?:\/\/.*[\r\n]*', '', tweet)
tweet = re.sub(r'#', '', tweet)
tweet = re.sub(r'\$\w*', '', tweet)
tweet = re.sub(r'^RT[\s]+', '', tweet)
tokenizer = TweetTokenizer(preserve_case=False, strip_handles=True, reduce_len=True)
tweet_tokens = tokenizer.tokenize(tweet)
tweets_cleaned = []
for word in tweet_tokens:
# lemmatize each word and then check pattern before making new sentence
lemma_word = lemmatizer.lemmatize(word)
check = digit_chr.match(word)
if word not in emoji.UNICODE_EMOJI['en'] and check:
tweets_cleaned.append(lemma_word)
#Create new sentence based on old sentence, apply bpe for new sentence
tweet_clean = " ".join(tweets_cleaned)
tweet_bpe = bpemb_en.encode(tweet_clean)
return tweet_bpe
# User counter to create vocab and calculate frequency
def tweet_counter(df):
count = Counter()
for tweet in df.values:
for word in tweet:
count[word] += 1
return count
# create a index list given a sentence and vocab
def index_tweet(vocab,tweet_list):
indice = [vocab[x] for x in tweet_list]
return indice
# + papermill={"duration": 0.051821, "end_time": "2021-12-05T05:27:21.442580", "exception": false, "start_time": "2021-12-05T05:27:21.390759", "status": "completed"} tags=[]
# Drop duplicated questions
qa_tweets.drop_duplicates('questions', inplace=True)
qa_tweets
# + papermill={"duration": 4.58733, "end_time": "2021-12-05T05:27:26.059843", "exception": false, "start_time": "2021-12-05T05:27:21.472513", "status": "completed"} tags=[]
# Create token df after clean tweet
qa_token = pd.DataFrame(columns = ["question_token", "answer_token"])
qa_token["question_token"] = qa_tweets["questions"].apply(lambda x: clean_tweet(x))
qa_token["answer_token"] = qa_tweets["answers"].apply(lambda x: clean_tweet(x))
qa_token
# + papermill={"duration": 0.072468, "end_time": "2021-12-05T05:27:26.163422", "exception": false, "start_time": "2021-12-05T05:27:26.090954", "status": "completed"} tags=[]
# Further checking about empty questions or answers
qa_token.reset_index(drop=True, inplace = True)
print("Empty questions:")
for i, d in enumerate(tqdm(qa_token['question_token'].values.tolist())):
if d == []:
qa_token.drop(i, inplace=True)
for i, d in enumerate(tqdm(qa_token['question_token'].values.tolist())):
if d == []:
print(i)
qa_token.reset_index(drop=True, inplace = True)
print("")
print("Empty Answers:")
for i, d in enumerate(tqdm(qa_token['answer_token'].values.tolist())):
if d == []:
qa_token.drop(i, inplace=True)
for i, d in enumerate(tqdm(qa_token['answer_token'].values.tolist())):
if d == []:
print(i)
# + papermill={"duration": 0.076806, "end_time": "2021-12-05T05:27:26.275311", "exception": false, "start_time": "2021-12-05T05:27:26.198505", "status": "completed"} tags=[]
# Counter and vocab for question and answers
cnt_q = tweet_counter(qa_token["question_token"])
cnt_a = tweet_counter(qa_token["answer_token"])
vocab_ql = sorted(list(cnt_q))
vocab_al = sorted(list(cnt_a))
# + papermill={"duration": 0.049635, "end_time": "2021-12-05T05:27:26.359780", "exception": false, "start_time": "2021-12-05T05:27:26.310145", "status": "completed"} tags=[]
# Create final vocab and index lists
vocab = set(vocab_ql).union(set(vocab_al))
vocab = sorted(list(vocab))
vocab_dict = {}
for i, word in enumerate(vocab):
vocab_dict[word] = i
print(len(vocab))
# + papermill={"duration": 0.079537, "end_time": "2021-12-05T05:27:26.475727", "exception": false, "start_time": "2021-12-05T05:27:26.396190", "status": "completed"} tags=[]
# Create index list for each sentence for questions and answers
qa_index = pd.DataFrame(columns = ["question_index", "answer_index"])
qa_index["question_index"] = qa_token["question_token"].apply(lambda x: index_tweet(vocab_dict,x))
qa_index["answer_index"] = qa_token["answer_token"].apply(lambda x: index_tweet(vocab_dict,x))
qa_index
# + papermill={"duration": 0.079081, "end_time": "2021-12-05T05:27:26.590488", "exception": false, "start_time": "2021-12-05T05:27:26.511407", "status": "completed"} tags=[]
#Save data to csv and txt files
qa_index.to_csv("index.csv", index=False)
with open('vocab.txt', 'w') as f:
for item in vocab_dict:
f.write("%s\n" % item)
| data/preprocess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import pickle
import cv2
from lesson_functions import *
import glob
from scipy.ndimage.measurements import label
# %matplotlib inline
dist_pickle = pickle.load( open("svc_pickle.p", "rb" ) )
svc = dist_pickle["svc"]
X_scaler = dist_pickle["scaler"]
orient = dist_pickle["orient"]
pix_per_cell = dist_pickle["pix_per_cell"]
cell_per_block = dist_pickle["cell_per_block"]
spatial_size = dist_pickle["spatial_size"]
hist_bins = dist_pickle["hist_bins"]
# +
def add_heat(heatmap, bbox_list):
# Iterate through list of bboxes
for box in bbox_list:
# Add += 1 for all pixels inside each bbox
# Assuming each "box" takes the form ((x1, y1), (x2, y2))
heatmap[box[0][1]:box[1][1], box[0][0]:box[1][0]] += 1
# Return updated heatmap
return heatmap
def apply_threshold(heatmap, threshold):
# Zero out pixels below the threshold
heatmap[heatmap <= threshold] = 0
# Return thresholded map
return heatmap
def draw_labeled_bboxes(img, labels):
# Iterate through all detected cars
for car_number in range(1, labels[1]+1):
# Find pixels with each car_number label value
nonzero = (labels[0] == car_number).nonzero()
# Identify x and y values of those pixels
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Define a bounding box based on min/max x and y
bbox = ((np.min(nonzerox), np.min(nonzeroy)), (np.max(nonzerox), np.max(nonzeroy)))
# Draw the box on the image
cv2.rectangle(img, bbox[0], bbox[1], (0,0,255), 6)
# Return the image
return img
# -
def find_cars(img, ystart, ystop, scale, svc, X_scaler, orient, pix_per_cell, cell_per_block,
spatial_size, hist_bins,cells_per_step = 2,hog_gray = False):
draw_img = np.copy(img)
img = img.astype(np.float32)/255
heatmap = np.zeros_like(img[:,:,0]).astype(np.float)
img_tosearch = img[ystart:ystop,:,:]
ctrans_tosearch = cv2.cvtColor(img_tosearch, cv2.COLOR_RGB2YCrCb)
if scale != 1:
imshape = ctrans_tosearch.shape
ctrans_tosearch = cv2.resize(ctrans_tosearch, (np.int(imshape[1]/scale), np.int(imshape[0]/scale)))
ch1 = ctrans_tosearch[:,:,0]
ch2 = ctrans_tosearch[:,:,1]
ch3 = ctrans_tosearch[:,:,2]
# Define blocks and steps as above
nxblocks = (ch1.shape[1] // pix_per_cell) - cell_per_block + 1
nyblocks = (ch1.shape[0] // pix_per_cell) - cell_per_block + 1
#nxblocks = (ch1.shape[1] // pix_per_cell) + 1
#nyblocks = (ch1.shape[0] // pix_per_cell) + 1
nfeat_per_block = orient*cell_per_block**2
# 64 was the orginal sampling rate, with 8 cells and 8 pix per cell
window = 64
nblocks_per_window = (window // pix_per_cell) - cell_per_block + 1
#nblocks_per_window = (window // pix_per_cell)+ 1
#cells_per_step = 2 # Instead of overlap, define how many cells to step
nxsteps = (nxblocks - nblocks_per_window) // cells_per_step + 1
nysteps = (nyblocks - nblocks_per_window) // cells_per_step + 1
if hog_gray == True:
ctrans_tosearch_rgb = cv2.cvtColor(ctrans_tosearch,cv2.COLOR_YCrCb2RGB)
ctrans_tosearch_gray = cv2.cvtColor(ctrans_tosearch,cv2.COLOR_RGB2GRAY)
hog = get_hog_features(ctrans_tosearch_gray, orient, pix_per_cell, cell_per_block, feature_vec=False)
else:
# Compute individual channel HOG features for the entire image
hog1 = get_hog_features(ch1, orient, pix_per_cell, cell_per_block, feature_vec=False)
hog2 = get_hog_features(ch2, orient, pix_per_cell, cell_per_block, feature_vec=False)
hog3 = get_hog_features(ch3, orient, pix_per_cell, cell_per_block, feature_vec=False)
hot_windows = []
windows = []
for xb in range(nxsteps):
for yb in range(nysteps):
ypos = yb*cells_per_step
xpos = xb*cells_per_step
if hog_gray == True:
hog_features = hog[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
else:
# Extract HOG for this patch
hog_feat1 = hog1[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_feat2 = hog2[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_feat3 = hog3[ypos:ypos+nblocks_per_window, xpos:xpos+nblocks_per_window].ravel()
hog_features = np.hstack((hog_feat1, hog_feat2, hog_feat3))
xleft = xpos*pix_per_cell
ytop = ypos*pix_per_cell
# Extract the image patch
subimg = cv2.resize(ctrans_tosearch[ytop:ytop+window, xleft:xleft+window], (64,64))
# Get color features
spatial_features = bin_spatial(subimg, size=spatial_size)
hist_features = color_hist(subimg, nbins=hist_bins)
# Scale features and make a prediction
test_features = X_scaler.transform(np.hstack((spatial_features, hist_features, hog_features)).reshape(1, -1))
#test_features = X_scaler.transform(np.hstack((shape_feat, hist_feat)).reshape(1, -1))
test_prediction = svc.predict(test_features)
xbox_left = np.int(xleft*scale)
ytop_draw = np.int(ytop*scale)
win_draw = np.int(window*scale)
windows.append([(xbox_left, ytop_draw+ystart),(xbox_left+win_draw,ytop_draw+win_draw+ystart)])
if test_prediction == 1:
hot_windows.append([(xbox_left, ytop_draw+ystart),(xbox_left+win_draw,ytop_draw+win_draw+ystart)])
#cv2.rectangle(draw_img,(xbox_left, ytop_draw+ystart),(xbox_left+win_draw,ytop_draw+win_draw+ystart),(0,0,255),6)
heatmap = add_heat(heatmap,hot_windows)
return hot_windows,heatmap,windows
def visualize(fig,rows,cols,images,titles):
for i,img in enumerate(images):
plt.subplot(rows,cols,i+1)
plt.title(titles[i])
img_dims = len(img.shape)
if img_dims<3:
plt.imshow(img,cmap='hot')
plt.title(titles[i])
plt.axis('off')
else:
plt.imshow(img)
plt.title(titles[i])
plt.axis('off')
img_names = glob.glob('./test_images/test*.jpg')
img_names
# +
ystart = 400
ystop = 656
scale = 1.5
img_names = glob.glob('./test_images/*.jpg')
#img = mpimg.imread('./test_images/test5.jpg')
#print('pix_per_cell:',pix_per_cell)
#print('cell_per_block:',cell_per_block)
#print(img.shape)
out_images = []
titles = []
for img_name in img_names:
img = cv2.imread(img_name)
img = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
hot_windows,heatmap,windows = find_cars(img, ystart, ystop, scale, svc, X_scaler,
orient, pix_per_cell, cell_per_block, spatial_size, hist_bins,hog_gray = True)
for window in hot_windows:
cv2.rectangle(img,window[0],window[1],(0,0,255),6)
out_images.append(img)
titles.append(img_name.split('\\')[1])
out_images.append(heatmap)
titles.append(img_name.split('\\')[1]+' heatmap')
heatmap = apply_threshold(heatmap,2)
labels = label(heatmap)
img_labeled = draw_labeled_bboxes(mpimg.imread(img_name),labels)
out_images.append(img_labeled)
titles.append(img_name.split('\\')[1])
'''
fig,ax = plt.subplots(1,figsize=(15, 10))
fig.tight_layout()
ax.imshow(img)
plt.show()
'''
# +
fig = plt.figure(figsize=(15,70))
visualize(fig,16,3,out_images,titles)
# -
def process_image(image):
ystart = 400
ystop = 720
scale = 1.5
#img = cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
hot_windows,heatmap = find_cars(image, ystart, ystop, scale, svc, X_scaler,
orient, pix_per_cell, cell_per_block, spatial_size, hist_bins)
heatmap = apply_threshold(heatmap,2)
labels = label(heatmap)
output_image = draw_labeled_bboxes(np.copy(image),labels)
return output_image
# +
from moviepy.editor import VideoFileClip
from IPython.display import HTML
write_output = 'project_video_output_1.mp4'
clip1 = VideoFileClip('project_video.mp4')
#clip1 = VideoFileClip('test_video.mp4')
write_clip = clip1.fl_image(process_image)
# %time write_clip.write_videofile(write_output, audio=False)
# -
| Test_files/SubSampling.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: scala-2.13
// language: scala
// name: scala-2.13
// ---
// ## Quadrangulations and Edge paths
//
// This is a demo of code due to <NAME> and <NAME>
import $cp.bin.`superficial-59af60d121.jar`
import superficial._
import SphereComplex._
import Triangles._
import NonPosQuad._
import EdgePath._
import Quadrangulation._
import StandardSurface._
import SvgPlot._
import almond.interpreter.api._
def showPlot(c: TwoComplex) = kernel.publish.display(DisplayData.svg(SvgPlot.plotComplex(c).toString))
showPlot(doubleBigon)
showPlot(quadrangulate(doubleBigon)._1)
showPlot(doubleTriangle)
showPlot(quadrangulate(doubleTriangle)._1)
val genus2 = new StandardSurface(2)
val quad = quadrangulate(genus2)._1
showPlot(genus2)
showPlot(quad)
quad.edges
val eds = quad.edges.toVector
eds.map(_.initial)
eds(0)
def randomPath = turnPathToEdgePath(eds(0), Vector(1,2,3,4,5,6), quad.asInstanceOf[NonPosQuad])
def randomPath = turnPathToEdgePath(eds(0), Vector(1,2,3,4,5,6))
| CATG2020/notebooks/.ipynb_checkpoints/QuadsEdgePaths-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="mGXI745KCCba" colab_type="text"
# # NLP Essay Organization Scorer
#
# This project seeks to use NLP tools on a set of middle and high school essays to classify them based on their level of organization. The data was provided in a Kaggle competition by the Hewlett Foundation seven years ago. The essays were hand-graded by more than one rater for standardized testing. I will be using spacy to tokenize the essays of interest and clustering and supervised learning models to attempt to predict their organization scores.
# + id="0qG3XC1rNJ1O" colab_type="code" outputId="d5fdb3e8-6004-429d-c2e1-eddc26df65c1" colab={"base_uri": "https://localhost:8080/", "height": 126}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="IfAUbUajNNTA" colab_type="code" outputId="6e27659a-70f7-46b7-95ab-5f1418934ae3" colab={"base_uri": "https://localhost:8080/", "height": 35}
# %cd '/content/gdrive/My Drive/Python'
# + id="KgfBC1xtCCbf" colab_type="code" outputId="0c4ef4fc-8279-454e-d307-13c6f4f616ec" colab={"base_uri": "https://localhost:8080/", "height": 179}
# import all the necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import re
import spacy
from sklearn.linear_model import LogisticRegression
from sklearn.cluster import estimate_bandwidth
from sklearn.cluster import KMeans, MeanShift, SpectralClustering
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.model_selection import train_test_split, GridSearchCV
# !python -m spacy download en
# + id="wa299JDmCCbn" colab_type="code" outputId="59d62bdc-42c9-469b-ef00-093174e6b8e3" colab={"base_uri": "https://localhost:8080/", "height": 370}
# Read in our first ten essays from the json file we have saved them to
data = pd.read_csv('training_set_rel3.tsv',
delimiter='\t',
encoding='latin_1',
error_bad_lines=False,
warn_bad_lines=True)
# Determine how many essays there are by set
data.groupby('essay_set').count()
# + id="WVPtjHC9CCcO" colab_type="code" outputId="b208310f-59dd-4fbc-aaf9-b6f7676fe677" colab={"base_uri": "https://localhost:8080/", "height": 395}
# Get just the essays in the sets that have a distinct organization score
org_score_data = data[data['essay_set'].isin([7,8])]
# Essays 7 & 8 have organization scores at trait 2,
# rename them to something simpler
org_score_data.rename(columns={'rater1_trait2': 'org1',
'rater2_trait2': 'org2',
'rater3_trait2': 'org3'}, inplace=True)
org_score_data['mean_org_score'] = 0
org_score_data = org_score_data.loc[:, ['essay_id',
'essay_set',
'essay',
'org1',
'org2',
'mean_org_score']]
# Calculate the mean org score
for i in range(len(org_score_data)):
org_score_data.iloc[i, -1] = np.mean([org_score_data.iloc[i, -2],
org_score_data.iloc[i, -3]])
org_score_data.head()
# + id="IXrCx51Lx7It" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 200} outputId="b221dc62-3dc3-495b-8fc8-5fa7fd2971f5"
# Fix punctuation errors that interfere with sentence splitting
for i in range(len(org_score_data.essay)):
org_score_data.iloc[i, 2] = re.sub(r'(?<=[\.\,\?\!])(?=[^\s])',
' ',
org_score_data.iloc[i, 2])
org_score_data.head()
# + id="5PBtG3z2CCc8" colab_type="code" colab={}
# Restructure the data into rows for each sentence in the essay
sentence_data = []
nlp = spacy.load('en')
for i in range(len(org_score_data)):
essay_id = org_score_data.essay_id.tolist()[i]
essay_set = org_score_data.essay_set.tolist()[i]
org_1 = org_score_data.org1.tolist()[i]
org_2 = org_score_data.org2.tolist()[i]
org_score = org_score_data.mean_org_score.tolist()[i]
sentences = list(nlp(org_score_data.essay.tolist()[i]).sents)
for j in range(len(sentences)):
sentence_data.append([essay_id, essay_set, j, sentences[j], org_1, org_2, org_score])
# + id="GOtdGHuOCCdI" colab_type="code" outputId="2fed9f91-bb0c-4399-b47e-b2eea602ae5f" colab={"base_uri": "https://localhost:8080/", "height": 200}
# Create the sentence-wise data frame
sent_df = pd.DataFrame(sentence_data, columns=['essay_id', 'essay_set', 'sentence_num',
'sentence', 'org_1', 'org_2', 'org_score'])
sent_df.head()
# + id="Mq3GCMpHduex" colab_type="code" outputId="1f7949da-11aa-4bf3-db81-9b51f59abcaf" colab={"base_uri": "https://localhost:8080/", "height": 413}
seven_df = sent_df.loc[sent_df['essay_set']==7]
eight_df = sent_df.loc[sent_df['essay_set']==8]
seven_df['org_score'] = seven_df['org_score'].apply(lambda x: int((x + 1) * 3) -2)
eight_df['org_score'] = eight_df['org_score'] * 2
sent_df = pd.concat([seven_df, eight_df])
sent_df['org_score'] = sent_df['org_score'].astype('int64')
sent_df.head()
# + id="NcID1TWKhJtm" colab_type="code" colab={}
sent_df['raw_score'] = 0
# + id="1tgM4BCNjiWS" colab_type="code" outputId="d2733a41-7ac3-410d-e06e-602a5661563c" colab={"base_uri": "https://localhost:8080/", "height": 479}
raw_score_dict = {}
cluster_dict = {}
for e in sent_df['essay_id'].unique():
cluster_dict[e] = {}
essay = sent_df.loc[sent_df['essay_id']==e]
sn = range(essay.sentence_num.tolist()[-1])
raw_org_score = []
for i in sn:
sent_score = np.sum([essay.sentence.tolist()[i].similarity(essay.sentence.tolist()[j]) / (np.log(abs(j-i))+1) for j in sn if j != i])
raw_org_score.append(sent_score)
cluster_dict[e].update({str(i)+str(j): essay.sentence.tolist()[i].similarity(essay.sentence.tolist()[j]) for j in sn if j != i and abs(j-i) < 6})
raw_score_dict[e] = np.mean(raw_org_score)
if e % 100 == 0:
print(raw_score_dict[e])
# + id="8gIiJuKM6kO_" colab_type="code" outputId="b7889e1e-6421-4c97-d3ff-39ac7d504985" colab={"base_uri": "https://localhost:8080/", "height": 249}
cluster_df = pd.DataFrame(cluster_dict)
cluster_df.fillna(cluster_df.mean(), inplace=True)
cluster_df = cluster_df.T
cluster_df.head()
# + id="wLXGkORVyqCo" colab_type="code" outputId="9400b8ae-b961-403e-df91-5e1d8509d288" colab={"base_uri": "https://localhost:8080/", "height": 200}
sent_df['raw_score'] = sent_df['essay_id'].replace(raw_score_dict, inplace=False)
sent_df.tail()
# + id="Zp80DN0BXZyh" colab_type="code" colab={}
score_df = sent_df.groupby('essay_id')[['org_score', 'raw_score']].mean()
# + id="4DU5rXI8Q5f-" colab_type="code" outputId="df131337-67de-4370-bbe3-010ec2ff2a51" colab={"base_uri": "https://localhost:8080/", "height": 426}
cluster_indices = np.where(cluster_df.isna())
drop_indices = [x for x in np.unique(cluster_indices[0])]
drop_values = [cluster_df.index.values.tolist()[i] for i in drop_indices]
drop_values
# + id="Q3bAZYAsMuzK" colab_type="code" colab={}
cluster_df = cluster_df.drop(drop_values, axis=0)
score_df = score_df.drop(drop_values, axis=0)
# + id="zzmUVyjl_puc" colab_type="code" colab={}
cluster_train, cluster_test, cluster_labels_train, cluster_labels_test = train_test_split(cluster_df, score_df['org_score'])
kmc = KMeans(n_clusters=11).fit(cluster_train)
# + id="HNQ_AxHSJ8IZ" colab_type="code" outputId="e4e3e580-c25d-43a4-d033-074bce35a3c5" colab={"base_uri": "https://localhost:8080/", "height": 513}
cm = pd.crosstab(kmc.labels_+1, cluster_labels_train)
plt.figure(figsize=(12,8))
sns.heatmap(cm)
plt.title('KMeans Clusters by Organization Scores')
plt.ylabel('Clusters')
plt.show()
# + id="JWGK9yMEWkSE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="0ecb7377-4e29-4b2f-89e3-5e3f707a5665"
cm
# + id="P9rysWFY86DB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a4066e23-e6b1-42b4-d00f-8e1b2288841a"
rc_dict = {8:6, 5:7, 4:9, 1:4, 7:3, 2:5, 0:8, 10:1}
acc = np.sum([cm.iloc[k, v] for k,v in rc_dict.items()]) / np.sum(cm.sum())
print('Clustering model train accuracy: {}'.format(acc))
# + id="MZDpzXRrakH7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="190d0286-33a2-4d1f-b59b-bc101870a22c"
cluster_pred = kmc.predict(cluster_test)
cluster_test_cm = pd.crosstab(cluster_pred, cluster_labels_test)
cluster_test_cm
# + id="834_XNu6aoEV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3a442a06-db9c-40db-a916-7da45daa2ed2"
clrtst_rc_dict = {1:4, 8:5, 4:2, 5:1, 7:3, 9:7, 0:6}
clrtst_acc = np.sum([cluster_test_cm.iloc[k, v]
for k,v in clrtst_rc_dict.items()]) / np.sum(cluster_test_cm.sum())
print('Clustering model test accuracy: {}'.format(clrtst_acc))
# + id="qRffmVYqqLHL" colab_type="code" colab={}
bw = estimate_bandwidth(cluster_train)
ms = MeanShift(bandwidth=bw).fit(cluster_train)
# + id="JE8Ml7iAqMA4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 838} outputId="c96004bc-e326-496a-dd28-2710829bda59"
ms_pred = ms.predict(cluster_test)
ms_test_cm = pd.crosstab(ms_pred, cluster_labels_test)
ms_test_cm
# + id="5rtgl3PHqMXL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="08cd8f11-c362-4a7d-b743-a4d6718aac0c"
ms_test_cm.iloc[0, 4] / np.sum(ms_test_cm.sum())
# + id="bNqDW0WnqNyB" colab_type="code" colab={}
sc = SpectralClustering(n_clusters=11).fit(cluster_train)
# + id="-iXHunnmqOUX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 412} outputId="c9dd0f40-03c0-46e2-f84c-a2e0ede05336"
sc_pred = sc.fit_predict(cluster_test)
sc_test_cm = pd.crosstab(sc_pred, cluster_labels_test)
sc_test_cm
# + id="RJ2QXXjDqOlN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4a6fa1c2-d705-4400-ba56-33b4355b3302"
160 / np.sum(sc_test_cm.sum())
# + id="46bUsyzlMZY6" colab_type="code" outputId="d8550f71-f39e-42f7-db78-0265db801d08" colab={"base_uri": "https://localhost:8080/", "height": 70}
lgr = LogisticRegression(solver='lbfgs', C=0.2, multi_class='multinomial')
X_train, X_test, y_train, y_test = train_test_split(
np.array(score_df['raw_score']).reshape(-1, 1),
score_df['org_score'], test_size=0.3)
lgr.fit(X_train, y_train)
pred = lgr.predict(X_test)
lgr_cm = pd.crosstab(pred, y_test)
lgr.score(X_train, y_train)
# + id="w1o9wSqFOsQq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c898793f-7325-4ff3-f0e3-d2a1a2ca8d93"
lgr_acc = []
for row in lgr_cm.index.tolist():
lgr_acc.append(lgr_cm.loc[row, row])
lgr_acc = np.sum(lgr_acc) / np.sum(lgr_cm.sum())
lgr_acc
# + id="_irSa4b4knk1" colab_type="code" outputId="69fa7405-23a0-41f8-c496-879f7297ed6d" colab={"base_uri": "https://localhost:8080/", "height": 35}
rfc = RandomForestClassifier(n_estimators=100, criterion='entropy')
rfc.fit(X_train, y_train)
pred = rfc.predict(X_test)
rfc_cm = pd.crosstab(pred, y_test)
rfc.score(X_train, y_train)
# + id="E8CCytjaQKNE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="64b63319-017a-4dd1-ae62-a2e0ab21b22c"
rfc_acc = []
for row in rfc_cm.index.tolist():
if row in rfc_cm.columns.tolist():
rfc_acc.append(rfc_cm.loc[row, row])
rfc_acc = np.sum(rfc_acc) / np.sum(rfc_cm.sum())
rfc_acc
# + id="v6whxHU7nhY7" colab_type="code" outputId="2684fa9a-0354-483a-88a7-f3b7ed585de1" colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.ensemble import GradientBoostingClassifier
gbc = GradientBoostingClassifier(n_estimators=100)
gbc.fit(X_train, y_train)
pred = gbc.predict(X_test)
gbc_cm = pd.crosstab(pred, y_test)
gbc.score(X_train, y_train)
# + id="W5zUZptDWOBu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e51a8cb5-f133-4ce3-9fb7-e090eadb9d5e"
gbc_acc = []
for row in gbc_cm.index.tolist():
if row in gbc_cm.columns.tolist():
gbc_acc.append(gbc_cm.loc[row, row])
gbc_acc = np.sum(gbc_acc) / np.sum(gbc_cm.sum())
gbc_acc
# + [markdown] id="TOIjsIDlEf8r" colab_type="text"
# # Write-up
#
# ## Corpus
#
# My area of interest is in using NLP to help lighten the work load for teacher. With that in mind, I wanted to select a corpus of essays that I could attempt to model a grade for. Since scraping a large set of essays was beyond the scope and timeframe I had for this project, I decided to hone my first phase on a corpus of essays that was already collected, organized, and labelled from an old Kaggle competition.
#
# This essay dataset came from the Hewlett Foundation Automated Essay Scoring Competition from 7 years ago. The data had named entities replaced with an anonymous tag (e.g. @LOCATION1) for the purposes of reducing bias in scoring of the essays in the standardized tests. There were 8 sets of essays based on different prompts and from different standardized tests, but I chose to focus on two sets (sets 7 & 8) that had component scores for organization.
#
# ## Hypothesis
#
# From my experience as a writer and a teacher, I hypothesized that the sentences of an essay would follow a pattern of similarity. That is, the first sentence should be most like the second sentence, less like the third sentence, and even less like the fourth sentence. If one were to visualize this as a heatmap of sentence similarity, It should look like a gradual dimming of diagonals with the brightest spots on the index diagonal (1,1; 2,2; 3,3; etc.) because a sentence is exactly similar to itself, but adjacent sentences should be nearly as bright and sentences furthest from each other should be dimmest (having the least correlation). I sought to quantify this hypothesis in a scoring function that I would then use to score essays and feed those scores into my model.
#
# One caveat for the generalizability of this model: all of these essays are single paragraph essays writtent by 7th or 10th grade students. To adapt this model for a mutli-paragraph essay, I would need to account for the fact that each paragraph is a self-contained entity where supporting details should be more closely tied to the topic sentence than a closer-by-distance sentence in the next paragraph. And all topic sentences should be closely related to the thesis statement, regardless of their distance.
#
# ## Scoring Function
# I knew I wanted to take the similarity between sentences to determine how organized the essay was, but I also knew I need to account for the distance between sentences. For each sentence, I decided to give the sentence a score based on the sum of its similarities to all other sentences divided by the natural log of its distance from those sentences. This ensured that its similarity to more distant sentences counted, but not quite as much as its similarity to nearby sentences. Finally, the essay received an organization score that was the mean of all the sentence-level organization scores. This allowed for the essays not to be penalized based on their length. For future iterations with mutli-paragraph essays, I would like to make a weight based on the average length of paragraphs compared to the total length of the essay.
# + id="moiNO0XfgxMc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="e36607f3-7bec-4877-e139-898d29dba4f0"
hypothesis = [[1, 0.85, 0.7, 0.55, 0.4, 0.3],
[0.85, 1, 0.85, 0.7, 0.55, 0.4],
[0.7, 0.85, 1, 0.85, 0.7, 0.55],
[0.55, 0.7, 0.85, 1, 0.85, 0.7],
[0.4, 0.55, 0.7, 0.85, 1, 0.85],
[0.3, 0.4, 0.55, 0.7, 0.85, 1]]
plt.figure(figsize=(8,6))
sns.heatmap(hypothesis)
plt.title('Hypothetical Distribution of Sentence Similarity')
plt.show()
# + [markdown] id="VAFC9KSqioOz" colab_type="text"
# ## Preparing the Data for Modeling
# ### Decoding the Essays
# The dataset was encoded in a way that made it difficult to properly decode with the python data science toolkit. Some bytes were left encoded, but I felt that there were not so many as to interfere significantly with the similarity scoring of sentences.
#
# ### Cleaning the Text
# I considered running spell check and other automated text cleaning functions to ensure that words were properly spelled and could be matched with their counterpart words in other sentences more readily. The scope and timeframe of this project did not allow for an in-depth exploration into whether those spell check methods would have made a difference, so I have relegated them to future iterations of this project. I have, however, found all instances of punctuation being immediately followed by a word instead of a space and added in that space to facilitate sentence delineation.
#
# ### Tokenization
# There were several models I considered for tokenization of the sentences in the essays before comparison for similarity. For the sake of building the simplest model first, I chose the native attribute `doc.sents` for a doc tokenized by spacy. This attribute has a built-in method `sent.similiarity(text)` that allows for easy extraction of similarity scores between sentences. In future iterations, I intend to compare this sentence tokenization method to comparable methods from gensim's word2vec and doc2vec.
#
# ### Feature Engineering
# In order to get features for my supervised learning model, I needed to implement the scoring function that I outlined above. I iterated over the sentences, taking the similarity scores with all the other sentences, divided by the natural logarithm of their distance, and summed those to create a sentence score. Then I took the mean of all sentence scores to create the raw organization score.
#
# For the labels, I needed to get the average score of the two raters, but my final labels needed to be integers. Also, essay set 7 and set 8 were graded on two different scales (0-4, 1-6). First, I split the data frame based on the essay set, then I added an offset to the zero-indexed mean score, multiplied by 3, and subtracted another offset to get values between 1 and 10. I multiplied the mean score of the other essay set by 2 to get values between 2 and 12.
#
# For the clustering model features, I took the similarity scores of every sentence pair in the essay that was within 6 sentences of each other. The 6 sentence maximum distance was chosen as an arbitrary stopping point to maintain the relevance of the features and prevent the feature engineering algorithm from running too long. To fill the null features for essays shorter than the longest essay, I imputed the mean. This assumes that if the student were asked to continue writing until they reached the maximum number of sentences in the essay set, they would continue to write in a style as organized as they had to the point they stopped.
#
# ## Clustering
# ### K-Means
# K-Means clustering assumes clusters that are isotropic, have similar variance, and roughly equal observations. These assumptions are not met by the sentence similarity data and lead to a poor model that over classifies the dominant classes and neglects the other. This is not the best clustering model to benchmark my supervised models against
# + id="H3lJK82AZ44E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="60eda658-0c20-4905-9e3c-a1660441e2fc"
plt.figure(figsize=(12,8))
sns.heatmap(cluster_test_cm)
plt.title('KMeans Clusters by Organization Scores')
plt.ylabel('Clusters')
plt.show()
# + [markdown] id="En6I3vg-XVub" colab_type="text"
# ### Mean Shift
# Mean Shift clusters got granular with the clustering beyond the majority class. It determined that there should be way more clusters than there are possible organization scores, but all of the clusters except one were relatively empty. This was not a good choice for a clustering model.
# + id="cElI1No2qLnD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="afa0cea3-58c6-41f8-ed53-c060afbaade0"
ms_cm = pd.crosstab(ms.labels_, cluster_labels_train)
plt.figure(figsize=(12,8))
sns.heatmap(ms_cm)
plt.title('Mean Shift Clusters by Organization Scores')
plt.ylabel('Clusters')
plt.show()
# + [markdown] id="mbgl1C3CXjco" colab_type="text"
# ### Spectral Clustering
#
# Spectral Clustering performed almost as terribly as Mean Shift clustering, except for the fact that I told it how many clusters to look for. Both models were able to get near 30% accuracy just by predicting the majority class, but they were not in the ballpark on detecting any of the other classes. Despite not meeting the model assumptions, K-Means was still the best performing clustering model for classes other then the dominant class.
# + id="9vp7HNt1qOD8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="115e3c3b-0100-4f7d-99f7-4d88f725c743"
sc_cm = pd.crosstab(sc.labels_, cluster_labels_train)
plt.figure(figsize=(12,8))
sns.heatmap(sc_cm)
plt.title('Spectral Clusters by Organization Scores')
plt.ylabel('Clusters')
plt.show()
# + [markdown] id="r6wuoFnyXofK" colab_type="text"
# ## Supervised Models
# ###Logisitic Regression
# The multinomial logistic regression model assumes low multi-collinearity and feature values that map one-to-one on labels. While the first assumption is met by the presence of only one feature, the second assumption is demonstrably broken as visualized in the scatter plot below . As shown in the scatter plot, the majority of essays are scored 6-8. The heatmap below shows that the logistic regression model gets most of its 39% accuracy from predicting the two majority classes. It makes an attempt at predicting the score 4, which I would interpret as the most dominant class to be tightly clustered on a range of lower scores. This would not be my model of choice under these circumstances.
# + id="N8uqune7Ix8K" colab_type="code" outputId="7afe606b-96c3-432e-897c-8be024c8003c" colab={"base_uri": "https://localhost:8080/", "height": 338}
sns.set_style('darkgrid')
plt.figure(figsize=(10,5))
sns.scatterplot(x='org_score', y='raw_score', data=score_df)
plt.show()
# + id="ZmYgyNf1faP9" colab_type="code" outputId="cc0803f1-767e-415d-eecb-f2db5d9ae130" colab={"base_uri": "https://localhost:8080/", "height": 513}
plt.figure(figsize=(12,8))
sns.heatmap(lgr_cm)
plt.title('Logistic Regression Predictions by Organization Scores')
plt.ylabel('Predicted Scores')
plt.show()
# + [markdown] id="Pw8Fq8pncgFe" colab_type="text"
# ### Random Forest Classifier
# The Random Forest model severely overfit the training data (99+%), but achieved just 29% accuracy on the test set. Of the supervised learning classifiers, though, it would be my model of choice under the current circumstances. This model tried to classify some of the minority classes. The heatmap below resembles the heatmap for the K-Means clustering model, but more disperse. I believe that with a more finely tuned scoring function and some more features derived from sound reasoning, the random forest classifier would be able to outperform the clustering model.
# + id="-gjybc5GliYi" colab_type="code" outputId="14562f09-2711-448b-c64a-bbbf929a7fbc" colab={"base_uri": "https://localhost:8080/", "height": 513}
plt.figure(figsize=(12,8))
sns.heatmap(rfc_cm)
plt.title('Random Forest Predictions by Organization Scores')
plt.ylabel('Predicted Scores')
plt.show()
# + [markdown] id="pV6ErBL-fzE1" colab_type="text"
# ### Gradient Boosting Classifier
# The Gradient Boosting model did not overfit as much as the Random Forest model, and it performed better in test accuracy, but it did so by sacrificing minority classes for the majority classes. I will continue compare it to the other classifiers as I iterate over this project with new features.
# + id="emxAd_3GUnFF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="7fb38f36-2291-4f4f-fd31-9b15728f9956"
plt.figure(figsize=(12,8))
sns.heatmap(gbc_cm)
plt.title('Gradient Boosting Predictions by Organization Scores')
plt.ylabel('Predicted Scores')
plt.show()
# + [markdown] id="1Y8AUtwlje3D" colab_type="text"
# ## Conclusion and Next Steps
#
# The first iteration of the organization grading model did not meet my expectations. However, there is much room for improvement within the project.
#
# ### Text Data
# I could improve on the project by collecting more labelled data and cleaning the data that I have. The current data needs to be explored to see what other errors are hiding in the essays aside from the ones I have seen in the head and tails of the data frames (undecoded bytes, rectangle where apostrophes should be, spaces in the middle of words.) I intend to use different spell checkers that are available to find words that match known misspellings in the vocabulary and try permutations of surrounding characters for words that do not exist within the vocabulary.
#
# ### Features
#
# #### A New Scoring Function
# The scoring function that I created changed a few times during the course of this project and will likely change many more before I am satisfied with its performance and the reasoning behind it. The first, simple tweak will be comparing the performance of the function if sentence similarities are divided by the distance instead of the natural logarithm of the distance. Still, I want there to be more features within the feature set and will continue to think about whether one number for the essay organization score is sufficient or whether there should be more scores included within the set.
#
# #### Other Text Features
# I always intended for the next iteration of this project to include a comparison of different models for tokenizing sentences. Some of those models will include reducing the sentence to its lemmas before creating the sentence vector. I want to explore reasonable ways in which parts of speech, dependencies, and named entities could play a role in the feature set. Named entities will require some extra legwork since all named entities have been anonymized in the essay sets I was using up to this point. Finally, some text summarization can be used to determine a thesis for the essay and a topic sentence for each paragraph, to begin to determine how closely linked all sentences in a paragraph are to their topic sentence and how well the topic sentences relate to the thesis.
#
# ### Models
# I will do research on the models that have the best performance for this type of problem and the range of parameters that work best. Then, I will perform GridSearchCV to determine the best model with the best parameters. I will also look into pretrained neural networks that already have word embeddings to use for transfer learning on the featureset.
# + id="CrCyQaXknJK4" colab_type="code" colab={}
| UnsupervisedLearning/EssayFeedback/NLP_Organization_Scorer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
import numpy as np
from utils.data_composer import feature_engineering
import neptune.new as neptune
import torch
# # 1. Load all data
# +
with open("data.json", "r") as f:
data = json.load(f)
# data is a list of 1, grab the core data inside
core_data = json.loads(data[0])
# Transform dataframe
df = pd.DataFrame(core_data)
# -
# Okay, so remember, at the previous stage of cleaning data (Part 1), we know that some data samples are invalid on some of their columns. When predicting genders, it's very important to tell the model to not rely on that invalid feature.
# The way to do that (in our practice) is to fill data that by a "middle value", this could be the mean or the median of that particular column. In my practice, I use median because it is robust to outlier.
#
# But first let's time them to null first
# +
# Replace all invalid value with null
# Replace invalid coupon by null (< 0 and > 1)
def nullize_invalid_coupon(value):
if value > 1 or value < 0:
return np.nan
else:
# Else keep
return value
# Replace
def nullize_zero_revenue(value):
if value <= 0:
return np.nan
else:
return value
# Turn invalid coupon to np.nan
df.loc[:,"coupon_discount_applied"] = df.loc[:,"coupon_discount_applied"].apply(nullize_invalid_coupon)
# Turn invalid revenue to np.nan
df.loc[:,"revenue"] = df.loc[:,"revenue"].apply(nullize_zero_revenue)
# -
# Run through pre-processor to get useful features
df = feature_engineering(df)
# +
# Get engineered data
feature_df = df.iloc[:,33:]
# Also append column "devices" and "coupon_discount_applied" into
feature_df = pd.concat([feature_df, df.loc[:,["coupon_discount_applied","devices","customer_id"]]],axis=1)
# partial labels
with open("partial_labels.csv","r") as f:
partial_labels_df = pd.read_csv(f)
# -
partial_labels_df.rename(columns={"Unnamed: 0":"df_index"}, inplace=True)
partial_labels_df
# # Normalize
import joblib
scaler = joblib.load("robust_scaler.pkl")
# Apply scaler on first 33 features
scaled_feature_df = feature_df.copy()
scaled_feature_df.iloc[:,:33] = scaler.transform(feature_df.iloc[:,:33])
# Here we'll handle missing values in a way that will confuse the model, to let it rely on other features to predict genders. So we fill this with median value
scaled_feature_df.columns
# # Clean data for prediction
# +
# add label
full_df = pd.merge(scaled_feature_df, partial_labels_df, how="left", left_on=scaled_feature_df.index, right_on="df_index")
# drop duplicate columns
full_df.drop(['customer_id_y','df_index'], axis=1, inplace=True)
full_df.rename(columns={'customer_id_x':"customer_id"}, inplace=True)
# Rename to denote our current self-labeled
full_df = full_df.rename(columns={"female_flag":"pseudo_female_flag"})
# Replace nan with median value to confuse the model on the feature
coupon_median = full_df["coupon_discount_applied"].median()
full_df["coupon_discount_applied"] = full_df["coupon_discount_applied"].fillna(coupon_median)
median_revenue_per_order = full_df["revenue_per_order"].median()
full_df["revenue_per_order"] = full_df["revenue_per_order"].fillna(median_revenue_per_order)
median_revenue_per_item = full_df["revenue_per_items"].median()
full_df["revenue_per_items"] = full_df["revenue_per_items"].fillna(median_revenue_per_item)
# "2" denotes unlabeled class
full_df.pseudo_female_flag = full_df.pseudo_female_flag.fillna(2)
# -
X = full_df.iloc[:,:33].to_numpy()
Y = full_df.loc[:,"pseudo_female_flag"].to_numpy()
# # Visualized Embedding features
# +
import optuna
import plotly
from sklearn.mixture import GaussianMixture
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from models.embeddingnet import EmbeddingNet
import torch
from collections import OrderedDict
from sklearn.metrics import v_measure_score
# -
# # Search for best dropout models
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
# +
def run_random_trial(trial, min_val, max_val):
# Suggest dropout rate
dropout_rate = round(trial.suggest_float("dropout_rate",min_val,max_val,step=0.05),2)
embedding_model = EmbeddingNet(input_dim = 33, dropout=dropout_rate)
# load embedder weight
ckpt_path = f"outputs/weights_dropout_{dropout_rate}.ckpt"
checkpoint = torch.load(ckpt_path, map_location=lambda storage, loc: storage)
state_dict = checkpoint["state_dict"]
state_dict = OrderedDict([(k.replace("embeddingnet.",""), v) for k, v in state_dict.items()])
embedding_model.load_state_dict(state_dict)
embedding_model.eval()
embedded_X = embedding_model.forward(torch.Tensor(X))
embedded_X = embedded_X.detach().numpy()
# Fit with GMM
#pred = GaussianMixture(n_components=2, random_state=0).fit_predict(embedded_X.detach().numpy())
pred = KMeans(n_clusters=2, random_state=0).fit_predict(embedded_X)
# Constraint with V measure
mask_pseudo_label = np.logical_or(Y == 1, Y == 0)
return embedded_X, pred, mask_pseudo_label
def objective_v_measure(trial):
embedded_X, pred, mask_pseudo_label = run_random_trial(trial, min_val=0.0, max_val=1.0)
v_measure = v_measure_score(Y[mask_pseudo_label],pred[mask_pseudo_label])
return v_measure
def objective_silhouette(trial):
embedded_X, pred, mask_pseudo_label = run_random_trial(trial, min_val=0.0, max_val=0.5)
# because silhouette score too expensive to compute, scale quadratically with n
# sample a small one to measure
embedded_X_sample, _, pred_sample, _ = train_test_split(embedded_X, pred,stratify=pred, train_size = N_SILHOUTTE_SAMPLES)
score = silhouette_score(embedded_X_sample, pred_sample)
return score
# -
# # Study V-measure as dropout choice
# +
import neptune.new as neptune
import neptune.new.integrations.optuna as optuna_utils
from joblib import parallel_backend
N_JOBS = 12
# connect your script to Neptune
run = neptune.init(project='patricknewyen/gfg-challenge',
api_token='<KEY>',
name = "search_dropout",
tags = ["optuna","EmbeddingNet","dropout","v_measure"])
neptune_callback = optuna_utils.NeptuneCallback(run) # skip chart because failed plotly import
study_v_measure = optuna.create_study(direction="maximize")
with parallel_backend('threading', n_jobs=N_JOBS):
study_v_measure.optimize(objective_v_measure, n_trials=100,n_jobs=N_JOBS, callbacks=[neptune_callback])
# -
optuna.visualization.plot_slice(study_v_measure, target_name="V measure")
# # Study Silhouette score as dropout choice
# +
V_MEASURE_THRESH = 0.95
# because silhouette score too expensive to compute, sample then compute
N_SILHOUTTE_SAMPLES = 10000
# +
import neptune.new as neptune
import neptune.new.integrations.optuna as optuna_utils
from joblib import parallel_backend
N_JOBS = 12
# connect your script to Neptune
run = neptune.init(project='patricknewyen/gfg-challenge',
api_token='<KEY>',
name = "search_dropout",
tags = ["optuna", "SuperTiny","dropout","silhouette"])
neptune_callback = optuna_utils.NeptuneCallback(run) # skip chart because failed plotly import
study_silhouette = optuna.create_study(direction="maximize")
with parallel_backend('threading', n_jobs=N_JOBS):
study_silhouette.optimize(objective_silhouette, n_trials=100,n_jobs=N_JOBS, callbacks=[neptune_callback])
# -
| Search for genders - Part 4 - Choose best model (EmbeddingNet).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neutral Network
# ## Objectives:
# Building Convolutional Neutral Networks (CNN) to:
# * Learn and train based on the matrix of features by building convolutional neutral network layers
# * Use feature detectors (filters - e.g., sharpen, blur, edge detect) to find features in the images by convolving with input images to build feature maps
# * Apply ReLu (Rectifier linear Unit) to break up non-linearity (images are highly non-linear)
# * Apply MaxPooling to complete downsampling the feature maps to form Pooled Feature Maps
# * Flattening the Pooled Feature Maps which forms the input layer
# * Apply stochastic gradient descent to minimise the lose function
# * Complete backpropagation to adjusts the weights
# * Complete parameter tuning if necessary
#
# ## Steps:
# 1) Visualise the dataset, transform the image and label data to be the correct dimensions (shape) and complete normalisation ready for CNN
#
# 2) Initialise CNN
#
# 3) Add first CNN layer with input shape of 28x28 pixels as our sample size, creating 32 feature maps using a feature detector with a kernel size of 3x3 matrix.
#
# 4) Add second CNN layer to improve model's accuracy
#
# 5) Complete MaxPooling, Regularization and Flattening
#
# 6) Add output layer with 10 nodes (for classifying digit 0 to 9), using the softmax activation function.
#
# 6) Apply stochastic gradient descent to achieve a set of optimal weights
#
# 7) Evaluate the model, visualise the analysis results
#
# Scenario: To build a classification model for a digit recongition system
#
# ### Dataset:
#
# MNIST: http://yann.lecun.com/exdb/mnist/
#
# The MNIST database of handwritten digits, available from the link above, has a training set of 60,000 examples, and a test set of 10,000 examples. It is a subset of a larger set available from NIST. The digits have been size-normalized and centered in a fixed-size image.
import numpy as np
np.random.seed(123) # for reproducible results
from scipy import misc
import tensorflow as tf
import h5py
import matplotlib.pyplot as plt
# %matplotlib inline
#import mnist dataset
from keras.datasets import mnist
from keras.utils import np_utils
#import keras libraries
from keras.models import Sequential #feedforward CNN
from keras.layers import Dense, Dropout, Activation, Flatten #core layers
from keras.layers import Convolution2D, MaxPooling2D #CNN layers
#Setting variables for MNIST image dimensions
mnist_image_height = 28
mnist_image_width = 28
#Import train and test sets of MNIST data
(X_img_train, y_img_train), (X_img_test, y_img_test) = mnist.load_data()
#Inspect the downloaded data
print("Shape of training dataset (depth,rows,columns): {}".format((X_img_train.shape)))
print("Shape of test dataset (depth,rows,columns): {}".format((X_img_test.shape)))
# ## Visualise one image
plt.figure()
plt.imshow(X_img_train[1], cmap='gray')
print("Label for image: {}".format(y_img_train[1]))
# ## Visualise the first digit class
fig, ax = plt.subplots(nrows=2, ncols=5, sharex=True, sharey=True, figsize=(10,4))
ax = ax.flatten()
for i in range(10):
img = X_img_train[y_img_train == i][0].reshape(28, 28)
ax[i].imshow(img, cmap='Greys', interpolation='nearest')
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
plt.show()
# ## Visualise 30 different versions of a digit
fig, ax = plt.subplots(nrows=5, ncols=6, sharex=True, sharey=True, figsize=(10,5))
ax = ax.flatten()
for i in range(30):
img = X_img_train[y_img_train == 7][i].reshape(28, 28)
ax[i].imshow(img, cmap='Greys', interpolation='nearest')
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
plt.show()
# ## Preprocess input images
# * Theano backend requires RGB to be explictly stated, i.e., Greyscale = 1, colour = 3
# * First, Reshape the input data (depth, rows, height) to (depth, rows, height, Greyscale)
X_train = X_img_train.reshape(X_img_train.shape[0], 28, 28, 1)
X_test = X_img_test.reshape(X_img_test.shape[0], 28, 28, 1)
#Transform data type to float32, and normalise values to the range [0,1]
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
# ## Preprocess image labels
# inspect y_train: it is a 1-D array
print(y_img_train.shape)
# ### Inspecting the first 10 labels. We can see the digit numerical values not the digit labels
print (y_img_train[:10])
# ### Transforming 1-D array of digit values to 10-D matrices of categorical digit labels
Y_train = np_utils.to_categorical(y_img_train, 10)
Y_test = np_utils.to_categorical(y_img_test, 10)
# ### Inspect the shape, ensure it is in the correct form
print(Y_train.shape)
# ## Initialising the CNN as sequence of layers
model = Sequential()
# ## Adding the first convolution layer
# * Convolution (maintain the spatial structure)
# * Apply ReLu (Rectifier linear Unit) to break up non-linearity (images are highly non-linear)
# * filters: number of filters is equalled to the number of feature maps we want to create, in this example, we create 32 x feature maps
# * kernel_size: no. of rows and columns for the feature dectector, in this example, we create 3x3 matrix as our feature detector
# * input_shape: convert input images into 2D or 3D array, in this example, our input images are greyscale images, so we convert to 2D array of 28 pixels by 28 pixels
# * for 3D images, then we set to 3 instead of 1 to create 3D array
# +
model.add(Convolution2D(32, (3, 3), activation='relu', input_shape=(28,28,1))) # (rows,columns,greyscale)
# Inspect the shape of the model
print(model.output_shape)
# -
# ## Adding the second convolution layer
# * Apply MaxPooling to complete downsampling the feature maps to form Pooled Feature Maps.
# * Reducing the number of parameters in our model by sliding a 2x2 pooling filter across the previous layer.
# * pool_size: halve the input in both spatial dimension
# * Flattening the Pooled Feature Maps which forms the input layer
model.add(Convolution2D(32, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25)) #Regularizing the model to prevent overfitting
model.add(Flatten())#Flattening
# ## Fully connected the layers
# * connect the input layer, 128 is the total number of o/p nodes for that layer, common practice is 128 or 2^(x) for hidden layers
# * Add output layer with 10 nodes (for classifying digit 0 to 9), using the softmax activation function.
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax')) #connect the output layer, 10 = 0 to 9 digits
# ## Applying Stochastic Gradient Descent
# * Adam is a variant of SGD for its effeciency and the coresponding loss function to be optimsed in order to achieve a set of optimal weights of the CNN.
# * The loss function for adam SGD is Logarithmic Loss.
# * For a binary classification outcome, the loss = binary_crossentropy.
# * For a categorical outcome, the loss = categorical_crossentrophy.
# * During each Epoch (observations) training, after all weights have been updated, accuracy metric is used to improve the model.
model.compile(optimizer = 'adam',loss='categorical_crossentropy', metrics=['accuracy'])
#Callback
from keras.callbacks import History
histories = History()
#FItting the CNN to the Training set
### Run a batch size of 32 observations before all the weights are updated.
model.fit(X_train, Y_train, batch_size=32, epochs=10, verbose=1, validation_data = (X_test, Y_test), callbacks = [histories])
# ## Evaluating the model
score = model.evaluate(X_test, Y_test, verbose=0)
print('\nThe {0} function of the test set is: {1:0.3}'.format(model.metrics_names[0],score[0]))
print('The {0} of the test set is: {1:0.3%}'.format(model.metrics_names[1],score[1]))
score = model.evaluate(X_train, Y_train, verbose=0)
print('\nThe {0} function of the training set is: {1:0.3}'.format(model.metrics_names[0],score[0]))
print('The {0} of the training set is: {1:0.3%}'.format(model.metrics_names[1],score[1]))
# summarize history for accuracy
plt.plot(histories.history['acc'])
plt.plot(histories.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
plt.show()
# summarize history for loss
plt.plot(histories.history['loss'])
plt.plot(histories.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
# ## Making the predictions and visualising the results
# * Showing a sample of 36 test images and the corresponding results
# * T: Ground Truth
# * P: Predicted Result
# Step 9 - CNN Model Prediction
y_pred = model.predict(X_test)
# +
miscl_img = X_test[y_img_test != np.argmax(y_pred)][10:46]
actual_labels = y_img_test[y_img_test != np.argmax(y_pred)][10:46]
predicted_labels = y_pred[y_img_test != np.argmax(y_pred)][10:46]
fig, ax = plt.subplots(nrows=6, ncols=6, sharex=True, sharey=True, figsize=(13,7))
ax = ax.flatten()
for i in range (36):
img = miscl_img[i].reshape(28, 28)
ax[i].imshow(img, cmap='Greys', interpolation='nearest')
ax[i].set_title('{}) T: {} P: {}' .format(i+1, actual_labels[i], np.argmax(predicted_labels[i])))
ax[0].set_xticks([])
ax[0].set_yticks([])
plt.tight_layout()
plt.show()
# -
| cnn_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qiskit import *
# Version
print(qiskit.__qiskit_version__)
from qiskit import IBMQ
from qiskit.tools.visualization import plot_histogram
from qiskit.tools.monitor import job_monitor
from qiskit.ignis.mitigation.measurement import (complete_meas_cal, CompleteMeasFitter)
# %matplotlib inline
# +
# API Token
from dotenv import load_dotenv
import os
load_dotenv()
API_TOKEN = os.getenv("IBM_API_TOKEN")
IBMQ.save_account(API_TOKEN, overwrite=True)
# -
IBMQ.load_account()
# +
nqubits = 3
ncbits = 3
circuit = QuantumCircuit(nqubits, ncbits)
# -
circuit.h(0)
circuit.cx(0,1)
circuit.cx(1,2)
circuit.measure(range(nqubits), range(ncbits))
circuit.draw(output='mpl');
# Run simulator
simulator = simulator = Aer.get_backend('qasm_simulator')
result = execute(experiments=circuit, backend=simulator, shots=1024).result()
counts = result.get_counts(circuit)
plot_histogram(counts);
# Execute on quantum computer
provider = IBMQ.get_provider('ibm-q')
qcomp = provider.get_backend('ibmq_manila')
# Execute circut on quantum computer
job = execute(experiments=circuit, backend=qcomp, shots=1024)
job_monitor(job)
# Plot result
qresult = job.result()
counts = qresult.get_counts(circuit)
plot_histogram(counts);
# +
# Noise mitigation
cal_circuits, state_labels = complete_meas_cal(qr=circuit.qregs[0], circlabel='measerrormitigationcal')
calib_job = execute(experiments=cal_circuits,
backend=qcomp,
shots=1024,
optimization_level=0)
job_monitor(calib_job)
# Plot result
calib_result = calib_job.result()
# -
meas_fitter = CompleteMeasFitter(calib_result, state_labels)
meas_fitter.plot_calibration()
meas_filter = meas_fitter.filter
mitigated_result = meas_filter.apply(qresult)
mitigated_counts = mitigated_result.get_counts(circuit)
plot_histogram([counts, mitigated_counts], legend=['qcomp, noisy', 'qcomp, mitigated']);
| noise_mitigation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### import
# +
import sys
import warnings
warnings.filterwarnings('ignore')
sys.path.append('/home/aleks/git/azmarabou')
print(sys.path)
# +
from maraboupy import DnCSolver
from maraboupy import DnC
import numpy as np
from multiprocessing import Process, Pipe
import os
# -
#network_name = "../../resources/nnet/acasxu/ACASXU_experimental_v2a_1_7" # 1s
#network_name = "../../resources/nnet/acasxu/ACASXU_experimental_v2a_2_9" # 2s
#network_name = ".../../resources/nnet/acasxu/ACASXU_experimental_v2a_2_6" # 10s
#network_name = "./acas/ACASXU_run2a_2_5_batch_2000" # 30s
#network_name = "./acas/ACASXU_run2a_5_2_batch_2000" # 45s
#network_name = "./acas/ACASXU_run2a_5_3_batch_2000" # 60s
network_name = "../../resources/nnet/acasxu/ACASXU_experimental_v2a_5_1" # 300s
property_path = "../../resources/properties/acas_property_3.txt"
# +
# Arguments and initiate the solver
num_workers = 3
initial_splits = 0
online_split = 2
init_to = 5
to_factor = 1.5
splitting_strategy = 3
solver = DnCSolver.DnCSolver(network_name, property_path, num_workers,
initial_splits, online_split, init_to, to_factor,
splitting_strategy)
# -
# Initial split of the input region
parent_conn, child_conn = Pipe()
p = Process(target=DnC.getSubProblems, args=(solver, child_conn))
p.start()
sub_queries = parent_conn.recv()
p.join()
# Solve the created subqueries
solver.solve(sub_queries)
| maraboupy/examples/DNCDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# Kaggle Basic Data Exploration Tutorial and Exercise
import pandas as pd # Import pandas library for data analysis
import numpy as np # Import numpy library for linear algebra (Unnecessary for this exercise)
melbourne_file_path = 'data/melb_data.csv' # Set file path of the data.
melbourne_data = pd.read_csv(melbourne_file_path) # Read the data and store in a data frame.
# -
melbourne_data.shape # Number of rows and columns in the data. NB: Rows: Observations, Columns: Variables
melbourne_data.columns # List of the column headers in the data.
melbourne_data.head() # Print first 5 rows of data from the dataframe.
melbourne_data.isnull().sum() # Identify missing data values from the dataframe.
# +
melbourne_data.describe() # Print a summary of the dataframe.
# The count, shows how many rows have non-missing values.
# The mean is the average.
# The std is the standard deviation (How numerically spread out the values are).
# min, 25%, 50%, 75% and max (self explanatory).
# -
| Kaggle_ML_Tutorials/IntroMachineLearning/BDEtutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split,cross_val_score,GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier,GradientBoostingClassifier,AdaBoostClassifier
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from xgboost import XGBClassifier
from sklearn.preprocessing import StandardScaler,MinMaxScaler
from sklearn.naive_bayes import GaussianNB
from imblearn.under_sampling import NearMiss
from keras.models import Sequential
from keras.layers import Dense
from sklearn import metrics
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from pandas_profiling import ProfileReport
data=pd.read_csv("train_ctrUa4K.csv")
data
pd.options.display.float_format = '{:,.0f}'.format
data['Dependents']=data.Dependents.map({'0':'zero','1':'one','2':'two','3+':'three_or_more'})
data['Credit_History']=data.Credit_History.map({0:'zero',1:'one'})
data['Loan_Amount_Term']=data.Loan_Amount_Term.map({12:'one',36:'three',60:'five',84:'seven',120:'ten',180:'fifteen',240:'twenty',300:'twentyfive',360:'thirty',480:'forty'})
for column in ('Gender','Married','Dependents','Self_Employed','Credit_History','Loan_Amount_Term','Property_Area','Education'):
data[column].fillna(data[column].mode()[0],inplace=True)
for column in ('LoanAmount','CoapplicantIncome','ApplicantIncome'):
data[column].fillna(data[column].mean(),inplace=True)
data.isna().sum()
data['Education'] = data['Education'].str.replace(' ','_')
data['Loan_Status']=data.Loan_Status.map({'Y':0,'N':1})
Y=data['Loan_Status'].values
data.drop(['Loan_Status'],axis=1,inplace=True)
X=data[data.iloc[:,1:13].columns]
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.33, stratify=Y)
# +
from sklearn.feature_extraction.text import CountVectorizer
print("="*50,"Gender","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Gender'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Gender_ohe = vectorizer.transform(X_train['Gender'].values)
X_test_Gender_ohe = vectorizer.transform(X_test['Gender'].values)
print("After vectorizations")
print(X_train_Gender_ohe.shape, y_train.shape)
print(X_test_Gender_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Married","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Married'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Married_ohe = vectorizer.transform(X_train['Married'].values)
X_test_Married_ohe = vectorizer.transform(X_test['Married'].values)
print("After vectorizations")
print(X_train_Married_ohe.shape, y_train.shape)
print(X_test_Married_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Dependents","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Dependents'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Dependents_ohe = vectorizer.transform(X_train['Dependents'].values)
X_test_Dependents_ohe = vectorizer.transform(X_test['Dependents'].values)
print("After vectorizations")
print(X_train_Dependents_ohe.shape, y_train.shape)
print(X_test_Dependents_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Education","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Education'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Education_ohe = vectorizer.transform(X_train['Education'].values)
X_test_Education_ohe = vectorizer.transform(X_test['Education'].values)
print("After vectorizations")
print(X_train_Education_ohe.shape, y_train.shape)
print(X_test_Education_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Self_Employed","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Self_Employed'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Self_Employed_ohe = vectorizer.transform(X_train['Self_Employed'].values)
X_test_Self_Employed_ohe = vectorizer.transform(X_test['Self_Employed'].values)
print("After vectorizations")
print(X_train_Self_Employed_ohe.shape, y_train.shape)
print(X_test_Self_Employed_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Property_Area","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Property_Area'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Property_Area_ohe = vectorizer.transform(X_train['Property_Area'].values)
X_test_Property_Area_ohe = vectorizer.transform(X_test['Property_Area'].values)
print("After vectorizations")
print(X_train_Property_Area_ohe.shape, y_train.shape)
print(X_test_Property_Area_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Loan_Amount_Term","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Loan_Amount_Term'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Loan_Amount_Term_ohe = vectorizer.transform(X_train['Loan_Amount_Term'].values)
X_test_Loan_Amount_Term_ohe = vectorizer.transform(X_test['Loan_Amount_Term'].values)
print("After vectorizations")
print(X_train_Loan_Amount_Term_ohe.shape, y_train.shape)
print(X_test_Loan_Amount_Term_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
print("="*50,"Credit_History","="*50)
vectorizer = CountVectorizer()
vectorizer.fit(X_train['Credit_History'].values) # fit has to happen only on train data
# we use the fitted CountVectorizer to convert the text to vector
X_train_Credit_History_ohe = vectorizer.transform(X_train['Credit_History'].values)
X_test_Credit_History_ohe = vectorizer.transform(X_test['Credit_History'].values)
print("After vectorizations")
print(X_train_Credit_History_ohe.shape, y_train.shape)
print(X_test_Credit_History_ohe.shape, y_test.shape)
print(vectorizer.get_feature_names())
print()
# +
from sklearn.preprocessing import Normalizer
print("="*50,"LoanAmount","="*50)
normalizer = Normalizer()
normalizer.fit(X_train['LoanAmount'].values.reshape(1,-1))
X_train_LoanAmount_norm = normalizer.transform(X_train['LoanAmount'].values.reshape(1,-1))
X_test_LoanAmount_norm = normalizer.transform(X_test['LoanAmount'].values.reshape(1,-1))
X_train_LoanAmount_norm = X_train_LoanAmount_norm.reshape(-1,1)
X_test_LoanAmount_norm = X_test_LoanAmount_norm.reshape(-1,1)
print("After vectorizations")
print(X_train_LoanAmount_norm.shape, y_train.shape)
print(X_test_LoanAmount_norm.shape, y_test.shape)
print()
print("="*50,"ApplicantIncome","="*50)
normalizer = Normalizer()
normalizer.fit(X_train['ApplicantIncome'].values.reshape(1,-1))
X_train_ApplicantIncome_norm = normalizer.transform(X_train['ApplicantIncome'].values.reshape(1,-1))
X_test_ApplicantIncome_norm = normalizer.transform(X_test['ApplicantIncome'].values.reshape(1,-1))
X_train_ApplicantIncome_norm = X_train_ApplicantIncome_norm.reshape(-1,1)
X_test_ApplicantIncome_norm = X_test_ApplicantIncome_norm.reshape(-1,1)
print("After vectorizations")
print(X_train_ApplicantIncome_norm.shape, y_train.shape)
print(X_test_ApplicantIncome_norm.shape, y_test.shape)
print()
print("="*50,"CoapplicantIncome","="*50)
normalizer = Normalizer()
normalizer.fit(X_train['CoapplicantIncome'].values.reshape(1,-1))
X_train_CoapplicantIncome_norm = normalizer.transform(X_train['CoapplicantIncome'].values.reshape(1,-1))
X_test_CoapplicantIncome_norm = normalizer.transform(X_test['CoapplicantIncome'].values.reshape(1,-1))
X_train_CoapplicantIncome_norm = X_train_CoapplicantIncome_norm.reshape(-1,1)
X_test_CoapplicantIncome_norm = X_test_CoapplicantIncome_norm.reshape(-1,1)
print("After vectorizations")
print(X_train_CoapplicantIncome_norm.shape, y_train.shape)
print(X_test_CoapplicantIncome_norm.shape, y_test.shape)
# +
from scipy.sparse import hstack
X_tr = hstack((X_train_Gender_ohe, X_train_Married_ohe, X_train_Dependents_ohe,X_train_Education_ohe,X_train_Self_Employed_ohe,X_train_Property_Area_ohe,X_train_Loan_Amount_Term_ohe,X_train_Credit_History_ohe,X_train_LoanAmount_norm,X_train_ApplicantIncome_norm,X_train_CoapplicantIncome_norm)).tocsr()
X_te = hstack((X_test_Gender_ohe, X_test_Married_ohe, X_test_Dependents_ohe,X_test_Education_ohe,X_test_Self_Employed_ohe,X_test_Property_Area_ohe,X_test_Loan_Amount_Term_ohe,X_test_Credit_History_ohe,X_test_LoanAmount_norm,X_test_ApplicantIncome_norm,X_test_CoapplicantIncome_norm)).tocsr()
print("Final Data matrix")
print(X_tr.shape, y_train.shape)
print(X_te.shape, y_test.shape)
print("="*125)
# -
# # Logistic Regression
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000] }
classifier = GridSearchCV(LogisticRegression(), param_grid,cv=10,scoring='roc_auc',return_train_score=True)
classifier.fit(X_tr, y_train)
# +
results_tf = pd.DataFrame.from_dict(classifier.cv_results_)
results_tf = results_tf.sort_values(['param_C'])
train_auc= results_tf['mean_train_score']
train_auc_std= results_tf['std_train_score']
cv_auc = results_tf['mean_test_score']
cv_auc_std= results_tf['std_test_score']
A = results_tf['param_C']
plt.plot(A, train_auc, label='Train AUC')
plt.plot(A, cv_auc, label='CV AUC')
plt.scatter(A, train_auc, label='Train AUC points')
plt.scatter(A, cv_auc, label='CV AUC points')
plt.xscale('log')
plt.legend()
plt.xlabel("C: hyperparameter")
plt.ylabel("AUC")
plt.title("Hyper parameter Vs AUC plot")
plt.grid()
plt.show()
# -
best_param=classifier.best_params_
print("Best Hyperparameter: ",best_param)
p_C=best_param['C']
# +
from sklearn.metrics import roc_curve, auc
Log_model = LogisticRegression(C=p_C)
Log_model.fit(X_tr, y_train)
y_train_pred = Log_model.predict_proba(X_tr)
y_test_pred = Log_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
#Computing AUC_Score with best parameter
AUC_Score_test_LOG=metrics.roc_auc_score(y_test,y_test_pred[:,1])
print('AUC_Score on test data: ',AUC_Score_test_LOG)
AUC_Score_train_LOG=metrics.roc_auc_score(y_train,y_train_pred[:,1])
print('AUC_Score on train data: ',AUC_Score_train_LOG)
#y_test_predict=predict_with_best_t(y_test_pred[:,1], best_t)
y_test_predict=Log_model.predict(X_te)
print("Recall for logistic regression model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for logistic regression model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for logistic regression model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for logistic regression model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for logistic regression model:",metrics.log_loss(y_test,y_test_predict))
importance = Log_model.coef_[0]
importance
importances = Log_model.coef_[0] #array with importances of each feature
ind = np.arange(0, X_tr.shape[1]) #create an index array, with the number of features
#only keep features whose importance is greater than 0
X_tr_features_to_keep = X_tr[:,ind[importances > 0.01]]
X_te_features_to_keep = X_te[:,ind[importances > 0.01]]
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000] }
classifier = GridSearchCV(LogisticRegression(), param_grid,cv=3,scoring='roc_auc',return_train_score=True)
classifier.fit(X_tr_features_to_keep, y_train)
best_param=classifier.best_params_
print("Best Hyperparameter: ",best_param)
p_C=best_param['C']
# +
results_tf = pd.DataFrame.from_dict(classifier.cv_results_)
results_tf = results_tf.sort_values(['param_C'])
train_auc= results_tf['mean_train_score']
train_auc_std= results_tf['std_train_score']
cv_auc = results_tf['mean_test_score']
cv_auc_std= results_tf['std_test_score']
A = results_tf['param_C']
plt.plot(A, train_auc, label='Train AUC')
plt.plot(A, cv_auc, label='CV AUC')
plt.scatter(A, train_auc, label='Train AUC points')
plt.scatter(A, cv_auc, label='CV AUC points')
plt.xscale('log')
plt.legend()
plt.xlabel("C: hyperparameter")
plt.ylabel("AUC")
plt.title("Hyper parameter Vs AUC plot")
plt.grid()
plt.show()
# +
from sklearn.metrics import roc_curve, auc
Log_model = LogisticRegression(C=p_C)
Log_model.fit(X_tr_features_to_keep, y_train)
y_train_pred = Log_model.predict_proba(X_tr_features_to_keep)
y_test_pred = Log_model.predict_proba(X_te_features_to_keep)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=Log_model.predict(X_te_features_to_keep)
print("Recall for logistic regression model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for logistic regression model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for logistic regression model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for logistic regression model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for logistic regression model:",metrics.log_loss(y_test,y_test_predict))
# # Decision Tree Model
min_sample_leaf_val=[1,2,3,4,5,6,7,8,9,10]
criterion_val=['entropy','gini']
max_depth=[1,2,3,4,5,6,7,8,9,10]
min_samples_split=[10,100,150,200,250]
param_grid = {'max_depth':max_depth,'criterion':criterion_val,'min_samples_leaf':min_sample_leaf_val,'min_samples_split':min_samples_split}
DT_model=DecisionTreeClassifier()
clf = GridSearchCV(estimator=DT_model, param_grid=param_grid, cv=3)
clf.fit(X_tr,y_train)
best_param=clf.best_params_
print("Best Hyperparameter: ",best_param)
max_depth_DT=best_param['max_depth']
min_samples_split_DT=best_param['min_samples_split']
min_samples_leaf_DT=best_param['min_samples_leaf']
criterion_DT=best_param['criterion']
# +
from sklearn.metrics import roc_curve, auc
DT_model= DecisionTreeClassifier(max_depth=max_depth_DT,min_samples_leaf=min_samples_leaf_DT,criterion=criterion_DT,min_samples_split=min_samples_split_DT)
#DT = DecisionTreeClassifier(max_depth=50,min_samples_split=5)
DT_model.fit(X_tr, y_train)
y_train_pred = DT_model.predict_proba(X_tr)
y_test_pred = DT_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
DT_pred=DT_model.predict(X_te)
print("Recall for decision tree model:",metrics.recall_score(y_test,DT_pred))
print("Precision for decision tree model:",metrics.precision_score(y_test,DT_pred))
print("Accuracy for decision tree model:",metrics.accuracy_score(y_test,DT_pred))
print("F-score for decision tree model:",metrics.f1_score(y_test,DT_pred))
print("Log-loss for decision tree model:",metrics.log_loss(y_test,DT_pred))
importances = DT_model.feature_importances_
print(importances)#array with importances of each feature
ind = np.arange(0, X_tr.shape[1]) #create an index array, with the number of features
#only keep features whose importance is greater than 0
X_tr_features_to_keep = X_tr[:,ind[importances > 0]]
X_te_features_to_keep = X_te[:,ind[importances > 0]]
"""from sklearn.linear_model import LogisticRegression
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000] }
classifier = GridSearchCV(LogisticRegression(), param_grid,cv=3,scoring='roc_auc',return_train_score=True)
classifier.fit(X_tr_features_to_keep, y_train)"""
"""best_param=classifier.best_params_
print("Best Hyperparameter: ",best_param)
p_C=best_param['C']"""
"""results_tf = pd.DataFrame.from_dict(classifier.cv_results_)
results_tf = results_tf.sort_values(['param_C'])
train_auc= results_tf['mean_train_score']
train_auc_std= results_tf['std_train_score']
cv_auc = results_tf['mean_test_score']
cv_auc_std= results_tf['std_test_score']
A = results_tf['param_C']
plt.plot(A, train_auc, label='Train AUC')
plt.plot(A, cv_auc, label='CV AUC')
plt.scatter(A, train_auc, label='Train AUC points')
plt.scatter(A, cv_auc, label='CV AUC points')
plt.xscale('log')
plt.legend()
plt.xlabel("C: hyperparameter")
plt.ylabel("AUC")
plt.title("Hyper parameter Vs AUC plot")
plt.grid()
plt.show()"""
"""from sklearn.metrics import roc_curve, auc
DT = LogisticRegression(C=p_C)
DT.fit(X_tr_features_to_keep, y_train)
y_train_pred = DT.predict_proba(X_tr_features_to_keep)
y_test_pred = DT.predict_proba(X_te_features_to_keep)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()"""
"""y_test_predict=DT.predict(X_te_features_to_keep)
print("Recall for logistic regression model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for logistic regression model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for logistic regression model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for logistic regression model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for logistic regression model:",metrics.log_loss(y_test,y_test_predict))"""
# # NaiveBayesModel
from sklearn.naive_bayes import MultinomialNB
NB = MultinomialNB()
param_grid = {'alpha': [0.00001,0.0005, 0.0001,0.005,0.001,0.05,0.01,0.1,0.5,1,5,10,50,100],'class_prior': [None,[0.5,0.5], [0.1,0.9],[0.2,0.8]]}
clf = GridSearchCV(NB, param_grid=param_grid, cv=3, scoring='roc_auc',return_train_score=True)
clf.fit(X_tr, y_train)
# +
results = pd.DataFrame.from_dict(clf.cv_results_)
results = results.sort_values(['param_alpha'])
train_auc= results['mean_train_score']
train_auc_std= results['std_train_score']
cv_auc = results['mean_test_score']
cv_auc_std= results['std_test_score']
A = results['param_alpha']
plt.plot(A, train_auc, label='Train AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
# plt.gca().fill_between(K, train_auc - train_auc_std,train_auc + train_auc_std,alpha=0.2,color='darkblue')
plt.plot(A, cv_auc, label='CV AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
# plt.gca().fill_between(K, cv_auc - cv_auc_std,cv_auc + cv_auc_std,alpha=0.2,color='darkorange')
plt.scatter(A, train_auc, label='Train AUC points')
plt.scatter(A, cv_auc, label='CV AUC points')
plt.xscale('log')
plt.legend()
plt.xlabel("Alpha: hyperparameter")
plt.ylabel("AUC")
plt.title("Hyper parameter Vs AUC plot")
plt.grid()
plt.show()
# -
best_param=clf.best_params_
print("Best Hyperparameter: ",best_param)
Alpha_BoW=best_param['alpha']
Class_Prior_BoW=best_param['class_prior']
# +
from sklearn.metrics import roc_curve, auc
NB = MultinomialNB(alpha=best_param['alpha'],class_prior=best_param['class_prior'])
NB.fit(X_tr, y_train)
# roc_auc_score(y_true, y_score) the 2nd parameter should be probability estimates of the positive class
# not the predicted outputs
y_train_pred = NB.predict_proba(X_tr)
y_test_pred = NB.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=NB.predict(X_te)
print("Recall for Naive Bayes model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for Naive Bayes model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for Naive Bayes model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for Naive Bayes model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for Naive Bayes model:",metrics.log_loss(y_test,y_test_predict))
# # KNN Model
n_neighbors_val=[5,10,20,30,40,50]
KNN_model = KNeighborsClassifier()
param_grid={'n_neighbors':n_neighbors_val}
clf=GridSearchCV(estimator=KNN_model,param_grid=param_grid,cv=5,scoring='roc_auc',return_train_score=True)
clf.fit(X_tr,y_train)
# +
results = pd.DataFrame.from_dict(clf.cv_results_)
results = results.sort_values(['param_n_neighbors'])
train_auc= results['mean_train_score']
train_auc_std= results['std_train_score']
cv_auc = results['mean_test_score']
cv_auc_std= results['std_test_score']
A = results['param_n_neighbors']
plt.plot(A, train_auc, label='Train AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
# plt.gca().fill_between(K, train_auc - train_auc_std,train_auc + train_auc_std,alpha=0.2,color='darkblue')
plt.plot(A, cv_auc, label='CV AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
# plt.gca().fill_between(K, cv_auc - cv_auc_std,cv_auc + cv_auc_std,alpha=0.2,color='darkorange')
plt.scatter(A, train_auc, label='Train AUC points')
plt.scatter(A, cv_auc, label='CV AUC points')
plt.xscale('log')
plt.legend()
plt.xlabel("Neighbor: hyperparameter")
plt.ylabel("AUC")
plt.title("Hyper parameter Vs AUC plot")
plt.grid()
plt.show()
# -
best_param=clf.best_params_
print("Best Hyperparameter: ",best_param)
Neighbor=best_param['n_neighbors']
# +
from sklearn.metrics import roc_curve, auc
Knn = KNeighborsClassifier(n_neighbors=best_param['n_neighbors'])
Knn.fit(X_tr, y_train)
# roc_auc_score(y_true, y_score) the 2nd parameter should be probability estimates of the positive class
# not the predicted outputs
y_train_pred = Knn.predict_proba(X_tr)
y_test_pred = Knn.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=Knn.predict(X_te)
print("Recall for KNN model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for KNN model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for KNN model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for KNN model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for KNN model:",metrics.log_loss(y_test,y_test_predict))
from sklearn.inspection import permutation_importance
results = permutation_importance(Knn,X_tr.toarray(), y_train, scoring='accuracy')
importance = results.importances_mean
print(importance)
# # Random Forest Model
n_estimator_val = [100,150,300,500,1000]
n_sample_leaf_val = [1,2,3,4,5,6]
max_feature_val=["auto","sqrt",None,0.9]
param_grid = {'n_estimators': n_estimator_val, 'min_samples_leaf' : n_sample_leaf_val,'max_features':max_feature_val}
RF_model=RandomForestClassifier()
grid_search_RF = GridSearchCV(estimator = RF_model,param_grid=param_grid, cv=3,scoring='roc_auc',return_train_score=True)
grid_search_RF.fit(X_tr, y_train)
best_param=grid_search_RF.best_params_
print("Best Hyperparameter: ",best_param)
# +
from sklearn.metrics import roc_curve, auc
RF_model= RandomForestClassifier(n_estimators=best_param['n_estimators'],min_samples_leaf=best_param['min_samples_leaf'],max_features=best_param['max_features'])
#DT = DecisionTreeClassifier(max_depth=50,min_samples_split=5)
RF_model.fit(X_tr, y_train)
y_train_pred = RF_model.predict_proba(X_tr)
y_test_pred = RF_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=RF_model.predict(X_te)
print("Recall for Random Forest model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for Random Forest model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for Random Forest model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for Random Forest model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for Random Forest model:",metrics.log_loss(y_test,y_test_predict))
importances = RF_model.feature_importances_
print(importances)#array with importances of each feature
ind = np.arange(0, X_tr.shape[1]) #create an index array, with the number of features
#only keep features whose importance is greater than 0
X_tr_features_to_keep = X_tr[:,ind[importances > 0]]
X_te_features_to_keep = X_te[:,ind[importances > 0]]
# # XGBoost
n_estimators=[150,200,500,1000,1500,2000]
max_features=[1,2,3]
max_depth=[1,2,3,4,5,6,7,8,9,10]
gammas = [0.001, 0.01, 0.1, 1]
learning_rate_val=[0.01,0.1,1,10,100]
param_grid = {'n_estimators': n_estimators,'max_features':max_features,'max_depth':max_depth,'gamma':gammas}
grid_search_xg = GridSearchCV(XGBClassifier(learning_rate=0.01), param_grid, cv=3)
grid_search_xg.fit(X_tr,y_train)
best_param=grid_search_xg.best_params_
print("Best Hyperparameter: ",best_param)
# +
from sklearn.metrics import roc_curve, auc
XGB_model= XGBClassifier(learning_rate=0.01,n_estimators=best_param['n_estimators'],max_features=best_param['max_features'],max_depth=best_param['max_depth'],gammas=best_param['gamma'])
#DT = DecisionTreeClassifier(max_depth=50,min_samples_split=5)
XGB_model.fit(X_tr, y_train)
y_train_pred = XGB_model.predict_proba(X_tr)
y_test_pred = XGB_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=XGB_model.predict(X_te)
print("Recall for XGBoost model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for XGBoost model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for XGBoost model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for XGBoost model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for XGBoost model:",metrics.log_loss(y_test,y_test_predict))
# # GradientBoosting
n_estimators=[150,200,500,1000,1500,2000]
max_features=[1,2,3]
max_depth=[1,2,3,4,5,6,7,8,9,10]
param_grid = {'n_estimators': n_estimators,'max_features':max_features,'max_depth':max_depth}
grid_search_gbm = GridSearchCV(GradientBoostingClassifier(learning_rate= 0.01), param_grid, cv=3)
grid_search_gbm.fit(X_tr,y_train)
best_param=grid_search_gbm.best_params_
print("Best Hyperparameter: ",best_param)
# +
from sklearn.metrics import roc_curve, auc
GRAD_model= GradientBoostingClassifier(learning_rate=0.01,n_estimators=best_param['n_estimators'],max_features=best_param['max_features'],max_depth=best_param['max_depth'])
#DT = DecisionTreeClassifier(max_depth=50,min_samples_split=5)
GRAD_model.fit(X_tr, y_train)
y_train_pred = GRAD_model.predict_proba(X_tr)
y_test_pred = GRAD_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=GRAD_model.predict(X_te)
print("Recall for Gradient model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for Gradient model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for Gradient model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for Gradient model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for Gradient model:",metrics.log_loss(y_test,y_test_predict))
# # SVM Model
Cs = [0.001, 0.01, 0.1, 1, 10]
gammas = [0.001, 0.01, 0.1, 1]
param_grid = {'C': Cs, 'gamma' : gammas}
grid_search_svm = GridSearchCV(SVC(kernel='rbf'), param_grid, cv=5)
grid_search_svm.fit(X_tr, y_train)
best_param=grid_search_svm.best_params_
print("Best Hyperparameter: ",best_param)
# +
from sklearn.metrics import roc_curve, auc
SVM_model= SVC(kernel='rbf',C=best_param['C'],gamma=best_param['gamma'],probability=True)
#DT = DecisionTreeClassifier(max_depth=50,min_samples_split=5)
SVM_model.fit(X_tr, y_train)
y_train_pred = SVM_model.predict_proba(X_tr)
y_test_pred = SVM_model.predict_proba(X_te)
train_fpr, train_tpr, tr_thresholds = roc_curve(y_train, y_train_pred[:,1])
test_fpr, test_tpr, te_thresholds = roc_curve(y_test, y_test_pred[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC ="+str(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC ="+str(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title("AUC ROC Curve")
plt.grid()
plt.show()
# -
y_test_predict=SVM_model.predict(X_te)
print("Recall for SVM model:",metrics.recall_score(y_test,y_test_predict))
print("Precision for SVM model:",metrics.precision_score(y_test,y_test_predict))
print("Accuracy for SVM model:",metrics.accuracy_score(y_test,y_test_predict))
print("F-score for SVM model:",metrics.f1_score(y_test,y_test_predict))
print("Log-loss for SVM model:",metrics.log_loss(y_test,y_test_predict))
| Vectoriser_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="3f0liPhfn83D"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="iK5ZtnDFoHKD"
# preprocessing
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV
import pandas_profiling as pp
# + id="9a9n2paloRDL"
# models
from sklearn.linear_model import LogisticRegression, Perceptron, RidgeClassifier, SGDClassifier
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier, ExtraTreesClassifier
from sklearn.ensemble import BaggingClassifier, AdaBoostClassifier, VotingClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.tree import DecisionTreeClassifier
from sklearn import metrics
import xgboost as xgb
from xgboost import XGBClassifier
import lightgbm as lgb
from lightgbm import LGBMClassifier
# + id="2gBulj9MoWCS"
# NN models
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras import optimizers
from keras.wrappers.scikit_learn import KerasClassifier
from keras.callbacks import EarlyStopping, ModelCheckpoint
# + id="JbTQbyljoalZ"
# model tuning
from hyperopt import STATUS_OK, Trials, fmin, hp, tpe, space_eval
# import warnings filter
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
# + id="FuscW9Ewoeof"
data = pd.read_csv("/content/sample_data/column_2C_weka.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="t3NpKPV4oo1X" outputId="4c2a424f-3b1b-4756-b269-e599b9c1ef14"
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="po12oMzkorxY" outputId="af5dda17-6d12-48e9-866d-ce1aeed255f3"
data.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="vpwsz37Totej" outputId="fd22978a-cab8-4185-c59d-f0710ee72a64"
data.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="eoACa-DsovlI" outputId="20570cb7-e552-4840-b5f2-9fac6d8bbcad"
data.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="R0O8AcnjoxMy" outputId="be995791-8a48-4ba3-ea51-d1eab804e082"
data.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="vnIY3-rbozSi" outputId="aece631d-87c8-4119-c31c-865e64d29f8c"
data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="2TAtazXEo1je" outputId="10aff7fe-960a-4702-ac48-19500c0210a2"
data.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 456} id="h07dbEHBpA95" outputId="cc18d4e4-48e0-4dd5-ebe9-d18818e8af83"
data.drop(columns=['degree_spondylolisthesis'])
# + [markdown] id="rMlYzIv0pbxi"
# EDA
# + colab={"base_uri": "https://localhost:8080/"} id="hW8mDZejqAEm" outputId="3d6db122-ae8c-4f03-eb45-90fd4883cafc"
# !pip install pandas
# + [markdown] id="glioV8TlqMy4"
# # Preparing to modeling
#
# Encoding categorical features
# + colab={"base_uri": "https://localhost:8080/"} id="Dq-P9gAPpEHm" outputId="6904230a-74c0-41fd-cb57-5f7ba34e6a7c"
# Determination categorical features
numerics = ['int8', 'int16', 'int32', 'int64', 'float16', 'float32', 'float64']
categorical_columns = []
features = data.columns.values.tolist()
for col in features:
if data[col].dtype in numerics: continue
categorical_columns.append(col)
categorical_columns
# + id="AQsRhw3qpHUo"
# Encoding categorical features
for col in categorical_columns:
if col in data.columns:
le = LabelEncoder()
le.fit(list(data[col].astype(str).values))
data[col] = le.transform(list(data[col].astype(str).values))
# + id="Krx7fUCPqZlz"
target_name = 'class'
data_target = data[target_name]
data = data.drop([target_name], axis=1)
# + id="PjsMXXj_qdMJ"
train, test, target, target_test = train_test_split(data, data_target, test_size=0.3, random_state=1)
# + colab={"base_uri": "https://localhost:8080/"} id="F3chvn8-qgjj" outputId="e259e870-4f19-4a06-d63c-a841704977d8"
print(target.shape)
print(target_test.shape)
print(train.shape)
print(test.shape)
# + [markdown] id="OX7RiAHKq52A"
# Creation of training and validation sets
# + id="SM1NG29kqyhv"
#%% split training set to validation set
Xtrain, Xval, Ztrain, Zval = train_test_split(train, target, test_size=0.3, random_state=1)
# + [markdown] id="2BYSWMtrrCJ2"
# Tuning models and test for all features
# + [markdown] id="DcuPifXtrIYj"
# # Logistic Regression
# + colab={"base_uri": "https://localhost:8080/"} id="bjtM-M5Oq9zo" outputId="8b945469-5daf-4747-b4a6-d3836f81cb0e"
# Logistic Regression
logreg = LogisticRegression()
logreg.fit(train, target)
acc_log = round(logreg.score(train, target) * 100, 2)
acc_log
# + colab={"base_uri": "https://localhost:8080/"} id="dSgvfK5ErPCR" outputId="db1d0948-ac74-4f1f-c6d5-e91b7dffd1cf"
acc_test_log = round(logreg.score(test, target_test) * 100, 2)
acc_test_log
# + [markdown] id="EtEcO2u2rbEv"
# # Support Vector Machines
# + colab={"base_uri": "https://localhost:8080/"} id="zWEAgeHZrSgP" outputId="9528c7b8-f0b4-451e-9430-2e188eadd839"
svc = SVC()
svc.fit(train, target)
acc_svc = round(svc.score(train, target) * 100, 2)
acc_svc
# + colab={"base_uri": "https://localhost:8080/"} id="1ri0-Vzmrdi6" outputId="6a5b1a38-7ccb-48f5-aade-089e600e26af"
acc_test_svc = round(svc.score(test, target_test) * 100, 2)
acc_test_svc
# + [markdown] id="_N4bo5KCrtlh"
# # Linear SVC
# + colab={"base_uri": "https://localhost:8080/"} id="w56T_haerkGJ" outputId="daab7143-ea65-40ec-a88f-daf47f654509"
linear_svc = LinearSVC(dual=False) # dual=False when n_samples > n_features.
linear_svc.fit(train, target)
acc_linear_svc = round(linear_svc.score(train, target) * 100, 2)
acc_linear_svc
# + colab={"base_uri": "https://localhost:8080/"} id="Tn6V0agDrwIw" outputId="85247722-afbc-442d-d234-a9506164ec59"
acc_test_linear_svc = round(linear_svc.score(test, target_test) * 100, 2)
acc_test_linear_svc
# + [markdown] id="US7dKb2Vr8hd"
# # k-Nearest Neighbors algorithm
# + colab={"base_uri": "https://localhost:8080/"} id="A3-K1CKgr0Rt" outputId="09f033cf-a90b-4a5f-8a4b-a94d91f73ca1"
knn = GridSearchCV(estimator=KNeighborsClassifier(), param_grid={'n_neighbors': [2, 3]}, cv=10).fit(train, target)
acc_knn = round(knn.score(train, target) * 100, 2)
print(acc_knn, knn.best_params_)
# + colab={"base_uri": "https://localhost:8080/"} id="dyl-tnn2r-u0" outputId="f00e744f-9e61-4603-ab9a-a3853ef7ace6"
acc_test_knn = round(knn.score(test, target_test) * 100, 2)
acc_test_knn
# + [markdown] id="TG0qwuF0sJz8"
# # Gaussian Naive Bayes
# + colab={"base_uri": "https://localhost:8080/"} id="OzbxLH2vsCjR" outputId="5a089730-4df8-4f87-f7bd-74113a54d744"
gaussian = GaussianNB()
gaussian.fit(train, target)
acc_gaussian = round(gaussian.score(train, target) * 100, 2)
acc_gaussian
# + colab={"base_uri": "https://localhost:8080/"} id="01vx60L_sNUm" outputId="6f1962be-279f-4b92-8d4e-ee5f87777d52"
acc_test_gaussian = round(gaussian.score(test, target_test) * 100, 2)
acc_test_gaussian
# + [markdown] id="xKaaB6K3sXB8"
# # Perceptron
# + colab={"base_uri": "https://localhost:8080/"} id="OQLuSvjasQyd" outputId="02bde35f-414f-418c-8208-2ab8e2810657"
perceptron = Perceptron()
perceptron.fit(train, target)
acc_perceptron = round(perceptron.score(train, target) * 100, 2)
acc_perceptron
# + colab={"base_uri": "https://localhost:8080/"} id="GBve5bKLsZm4" outputId="1f9637db-decb-4d9e-c7ef-eb92b03bccfd"
acc_test_perceptron = round(perceptron.score(test, target_test) * 100, 2)
acc_test_perceptron
# + [markdown] id="UqNObXL5sjSc"
# # Stochastic Gradient Descent
# + colab={"base_uri": "https://localhost:8080/"} id="gAOnzbowscyb" outputId="54a38788-bfa6-4fbd-eb82-cbc71dd375d3"
sgd = SGDClassifier()
sgd.fit(train, target)
acc_sgd = round(sgd.score(train, target) * 100, 2)
acc_sgd
# + colab={"base_uri": "https://localhost:8080/"} id="F0lDHPPeslIo" outputId="e3e27e9a-1e9f-49f6-cadb-eafafa674989"
acc_test_sgd = round(perceptron.score(test, target_test) * 100, 2)
acc_test_sgd
# + [markdown] id="l3JxH6kBsxfj"
# # Decision Tree Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="TEthED4yspnK" outputId="2e5da4a4-e258-4e30-b120-897e43a2cd4b"
decision_tree = DecisionTreeClassifier()
decision_tree.fit(train, target)
acc_decision_tree = round(decision_tree.score(train, target) * 100, 2)
acc_decision_tree
# + colab={"base_uri": "https://localhost:8080/"} id="oeYSM3ZBszOk" outputId="c5d485a5-215b-4cf6-e444-4edb42ab775d"
acc_test_decision_tree = round(decision_tree.score(test, target_test) * 100, 2)
acc_test_decision_tree
# + [markdown] id="qoC_2cvDs9jM"
# # Random Forest
# + colab={"base_uri": "https://localhost:8080/"} id="A9AoMU0Vs2iE" outputId="30f90d88-f0de-4e31-b035-e927f24b9ebe"
random_forest = GridSearchCV(estimator=RandomForestClassifier(), param_grid={'n_estimators': [100, 300]}, cv=5).fit(train, target)
random_forest.fit(train, target)
acc_random_forest = round(random_forest.score(train, target) * 100, 2)
print(acc_random_forest,random_forest.best_params_)
# + colab={"base_uri": "https://localhost:8080/"} id="ee-VFZb8tCBc" outputId="46917290-0757-412d-c54c-d9c5fba51d91"
acc_test_random_forest = round(random_forest.score(test, target_test) * 100, 2)
acc_test_random_forest
# + [markdown] id="6kH6qSN3tPkP"
# XGB
# + colab={"base_uri": "https://localhost:8080/"} id="DIKqxgSHtGMY" outputId="e78c70e6-55b2-41c7-9048-56fda173d409"
def hyperopt_xgb_score(params):
clf = XGBClassifier(**params)
current_score = cross_val_score(clf, train, target, cv=10).mean()
print(current_score, params)
return current_score
space_xgb = {
'learning_rate': hp.quniform('learning_rate', 0, 0.05, 0.0001),
'n_estimators': hp.choice('n_estimators', range(100, 1000)),
'eta': hp.quniform('eta', 0.025, 0.5, 0.005),
'max_depth': hp.choice('max_depth', np.arange(2, 12, dtype=int)),
'min_child_weight': hp.quniform('min_child_weight', 1, 9, 0.025),
'subsample': hp.quniform('subsample', 0.5, 1, 0.005),
'gamma': hp.quniform('gamma', 0.5, 1, 0.005),
'colsample_bytree': hp.quniform('colsample_bytree', 0.5, 1, 0.005),
'eval_metric': 'auc',
'objective': 'binary:logistic',
'booster': 'gbtree',
'tree_method': 'exact',
'silent': 1,
'missing': None
}
best = fmin(fn=hyperopt_xgb_score, space=space_xgb, algo=tpe.suggest, max_evals=10)
print('best:')
print(best)
# + colab={"base_uri": "https://localhost:8080/"} id="RGZAJ03EtScq" outputId="a95fbdd7-5d9c-42db-8b46-be5a4a564efc"
params = space_eval(space_xgb, best)
params
# + colab={"base_uri": "https://localhost:8080/"} id="OoPsPSlXtYHx" outputId="031833a8-6060-4d68-a20b-eb9e6647bdb7"
XGB_Classifier = XGBClassifier(**params)
XGB_Classifier.fit(train, target)
acc_XGB_Classifier = round(XGB_Classifier.score(train, target) * 100, 2)
acc_XGB_Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="yKT90uottbxv" outputId="fe4878e5-0699-49f0-eccb-03fffbc3f82d"
acc_test_XGB_Classifier = round(XGB_Classifier.score(test, target_test) * 100, 2)
acc_test_XGB_Classifier
# + id="Su2bLewzteXx"
fig = plt.figure(figsize = (15,15))
axes = fig.add_subplot(111)
xgb.plot_importance(XGB_Classifier,ax = axes,height =0.5)
plt.show();
plt.close()
# + [markdown] id="7IqagcbBttmX"
# LGBM Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="9MWauFmZtotW" outputId="23d0462e-922e-4504-bd06-ffdb4b4465d2"
def hyperopt_lgb_score(params):
clf = LGBMClassifier(**params)
current_score = cross_val_score(clf, train, target, cv=10).mean()
print(current_score, params)
return current_score
space_lgb = {
'learning_rate': hp.quniform('learning_rate', 0, 0.05, 0.0001),
'n_estimators': hp.choice('n_estimators', range(100, 1000)),
'max_depth': hp.choice('max_depth', np.arange(2, 12, dtype=int)),
'num_leaves': hp.choice('num_leaves', 2*np.arange(2, 2**11, dtype=int)),
'min_child_weight': hp.quniform('min_child_weight', 1, 9, 0.025),
'colsample_bytree': hp.quniform('colsample_bytree', 0.5, 1, 0.005),
'objective': 'binary',
'boosting_type': 'gbdt',
}
best = fmin(fn=hyperopt_lgb_score, space=space_lgb, algo=tpe.suggest, max_evals=10)
print('best:')
print(best)
# + colab={"base_uri": "https://localhost:8080/"} id="HaL6GO3Wtw02" outputId="20b87d7b-ddea-44af-8a64-aa91760d5fd9"
params = space_eval(space_lgb, best)
params
# + colab={"base_uri": "https://localhost:8080/"} id="W4_iT_KEt3Bt" outputId="a6ee56c3-3ade-4b69-a758-878d55f70856"
LGB_Classifier = LGBMClassifier(**params)
LGB_Classifier.fit(train, target)
acc_LGB_Classifier = round(LGB_Classifier.score(train, target) * 100, 2)
acc_LGB_Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="sOpO4lvst6eB" outputId="8d223d73-7a45-4ede-faf4-56df92a8c2fb"
acc_test_LGB_Classifier = round(LGB_Classifier.score(test, target_test) * 100, 2)
acc_test_LGB_Classifier
# + id="uJ5fEa3Jt9B0"
fig = plt.figure(figsize = (15,15))
axes = fig.add_subplot(111)
lgb.plot_importance(LGB_Classifier,ax = axes,height = 0.5)
plt.show();
plt.close()
# + [markdown] id="m0xVzlDFuMf9"
# GradientBoosting
# + colab={"base_uri": "https://localhost:8080/"} id="8KkO-8rmuAId" outputId="6a5d4d16-900d-44fb-b646-6a29bf9d47b2"
def hyperopt_gb_score(params):
clf = GradientBoostingClassifier(**params)
current_score = cross_val_score(clf, train, target, cv=10).mean()
print(current_score, params)
return current_score
space_gb = {
'n_estimators': hp.choice('n_estimators', range(100, 1000)),
'max_depth': hp.choice('max_depth', np.arange(2, 10, dtype=int))
}
best = fmin(fn=hyperopt_gb_score, space=space_gb, algo=tpe.suggest, max_evals=10)
print('best:')
print(best)
# + colab={"base_uri": "https://localhost:8080/"} id="Yto6aXQwuOZB" outputId="0c6f83f6-7388-4e00-9f30-10dad5392e65"
params = space_eval(space_gb, best)
params
# + colab={"base_uri": "https://localhost:8080/"} id="tcv79WPhuS4F" outputId="c4258f1d-f55c-429a-be78-6b25b5bcac85"
# Gradient Boosting Classifier
gradient_boosting = GradientBoostingClassifier(**params)
gradient_boosting.fit(train, target)
acc_gradient_boosting = round(gradient_boosting.score(train, target) * 100, 2)
acc_gradient_boosting
# + colab={"base_uri": "https://localhost:8080/"} id="ejp-aDGDuWcP" outputId="96a00007-1fba-4b47-8ecf-8e8eb5c11f12"
acc_test_gradient_boosting = round(gradient_boosting.score(test, target_test) * 100, 2)
acc_test_gradient_boosting
# + [markdown] id="wviSMUX1ug9U"
# # Ridge Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="J8EqtwSquZIK" outputId="4f8dc025-1a34-45bd-e47f-2f02655f4334"
ridge_classifier = RidgeClassifier()
ridge_classifier.fit(train, target)
acc_ridge_classifier = round(ridge_classifier.score(train, target) * 100, 2)
acc_ridge_classifier
# + colab={"base_uri": "https://localhost:8080/"} id="zpq_zAGVuimf" outputId="09e719c0-17e9-4d43-be55-cf48aa98e7bf"
acc_test_ridge_classifier = round(ridge_classifier.score(test, target_test) * 100, 2)
acc_test_ridge_classifier
# + [markdown] id="xFPiqFChus6a"
# # Bagging Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="xWrNo0iXulVm" outputId="75942b7e-5b33-44ee-9a3a-2b3eebe91ac0"
bagging_classifier = BaggingClassifier()
bagging_classifier.fit(train, target)
Y_pred = bagging_classifier.predict(test).astype(int)
acc_bagging_classifier = round(bagging_classifier.score(train, target) * 100, 2)
acc_bagging_classifier
# + colab={"base_uri": "https://localhost:8080/"} id="l7WEhtGxuvq8" outputId="a92774cb-6fb1-4af4-aad3-30a6634a0e28"
acc_test_bagging_classifier = round(bagging_classifier.score(test, target_test) * 100, 2)
acc_test_bagging_classifier
# + [markdown] id="BRcKzFQSu7xx"
# ExtraTreesClassifier
# + colab={"base_uri": "https://localhost:8080/"} id="A_GmBz-ruzn-" outputId="eaf1b6d4-ce22-4108-ab9e-affe7f6127e2"
def hyperopt_etc_score(params):
clf = ExtraTreesClassifier(**params)
current_score = cross_val_score(clf, train, target, cv=10).mean()
print(current_score, params)
return current_score
space_etc = {
'n_estimators': hp.choice('n_estimators', range(100, 1000)),
'max_features': hp.choice('max_features', np.arange(2, 17, dtype=int)),
'min_samples_leaf': hp.choice('min_samples_leaf', np.arange(1, 5, dtype=int)),
'max_depth': hp.choice('max_depth', np.arange(2, 12, dtype=int)),
'max_features': None # for small number of features
}
best = fmin(fn=hyperopt_etc_score, space=space_etc, algo=tpe.suggest, max_evals=10)
print('best:')
print(best)
# + colab={"base_uri": "https://localhost:8080/"} id="qXINR44yu90a" outputId="195a0ad0-15e1-41a5-9bee-73bf46eb8b78"
params = space_eval(space_etc, best)
params
# + colab={"base_uri": "https://localhost:8080/"} id="zLBVTWBTvCEU" outputId="7508eab5-bf3f-4106-8502-00d23cef3c8a"
# Extra Trees Classifier
extra_trees_classifier = ExtraTreesClassifier(**params)
extra_trees_classifier.fit(train, target)
acc_etc = round(extra_trees_classifier.score(train, target) * 100, 2)
acc_etc
# + colab={"base_uri": "https://localhost:8080/"} id="Cz3chsWcvErN" outputId="e3dfe985-2512-4116-afd2-0273b719343d"
acc_test_etc = round(extra_trees_classifier.score(test, target_test) * 100, 2)
acc_test_etc
# + [markdown] id="FVj9-yTZvesw"
# NN
# + id="9DiTm5vVvH0p"
def build_ann(optimizer='adam'):
# Initializing the ANN
ann = Sequential()
# Adding the input layer and the first hidden layer of the ANN with dropout
ann.add(Dense(units=32, kernel_initializer='glorot_uniform', activation='relu', input_shape=(len(train.columns),)))
# Add other layers, it is not necessary to pass the shape because there is a layer before
ann.add(Dense(units=64, kernel_initializer='glorot_uniform', activation='relu'))
ann.add(Dropout(rate=0.5))
ann.add(Dense(units=64, kernel_initializer='glorot_uniform', activation='relu'))
ann.add(Dropout(rate=0.5))
# Adding the output layer
ann.add(Dense(units=1, kernel_initializer='glorot_uniform', activation='sigmoid'))
# Compiling the ANN
ann.compile(optimizer=optimizer, loss='binary_crossentropy', metrics=['accuracy'])
return ann
# + colab={"base_uri": "https://localhost:8080/"} id="7XoR7R4NvgO8" outputId="d59efcb9-d366-48ea-d15e-282fe75f2380"
opt = optimizers.Adam(lr=0.001)
ann = build_ann(opt)
# Training the ANN
history = ann.fit(Xtrain, Ztrain, batch_size=16, epochs=100, validation_data=(Xval, Zval))
# + colab={"base_uri": "https://localhost:8080/"} id="VmQaqqwpvj1A" outputId="8b548bbf-a83b-4aba-a1c2-d0190fb0b894"
# Predicting the Train set results
ann_prediction = ann.predict(train)
ann_prediction = (ann_prediction > 0.5)*1 # convert probabilities to binary output
# Compute error between predicted data and true response and display it in confusion matrix
acc_ann1 = round(metrics.accuracy_score(target, ann_prediction) * 100, 2)
acc_ann1
# + colab={"base_uri": "https://localhost:8080/"} id="bxMlBeP0vqmw" outputId="ff89a6ca-c53b-4923-bc4b-31f82602dd33"
# Predicting the Test set results
ann_prediction_test = ann.predict(test)
ann_prediction_test = (ann_prediction_test > 0.5)*1 # convert probabilities to binary output
# Compute error between predicted data and true response and display it in confusion matrix
acc_test_ann1 = round(metrics.accuracy_score(target_test, ann_prediction_test) * 100, 2)
acc_test_ann1
# + [markdown] id="FPyiiyNWvzHD"
# Neural Network 2
# + colab={"base_uri": "https://localhost:8080/"} id="njf4gfZsvuUi" outputId="c3b9479b-7753-455d-884e-75a1f0ae3566"
# Model
model = Sequential()
model.add(Dense(16, input_dim = train.shape[1], activation = 'relu'))
model.add(Dropout(0.3))
model.add(Dense(64, activation = 'relu'))
model.add(Dropout(0.3))
model.add(Dense(32, activation = 'relu'))
model.add(Dense(1, activation = 'sigmoid'))
model.summary()
# + id="oP_vVqIuv2nz"
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="NBRNkWItwoGk" outputId="83953585-cd61-4716-e4e3-fd791253fbf3"
es = EarlyStopping(monitor='val_accuracy', patience=20, mode='max')
hist = model.fit(train, target, batch_size=64, validation_data=(Xval, Zval),
epochs=500, verbose=1, callbacks=[es])
# + colab={"base_uri": "https://localhost:8080/"} id="jZjVbawBwq9g" outputId="7db80688-af95-4188-c9bd-eaf5d63d37fc"
plt.plot(hist.history['accuracy'], label='acc')
plt.plot(hist.history['val_accuracy'], label='val_acc')
# plt.plot(hist.history['acc'], label='acc')
# plt.plot(hist.history['val_acc'], label='val_acc')
plt.ylim((0, 1))
plt.legend()
# + colab={"base_uri": "https://localhost:8080/"} id="qxS7kMOTwxNQ" outputId="36d95783-f045-4e49-e1c3-5821f26bd772"
# Predicting the Train set results
nn_prediction = model.predict(train)
nn_prediction = (nn_prediction > 0.5)*1 # convert probabilities to binary output
# Compute error between predicted data and true response
acc_ann2 = round(metrics.accuracy_score(target, nn_prediction) * 100, 2)
acc_ann2
# + colab={"base_uri": "https://localhost:8080/"} id="3HXLupX9w4we" outputId="798af099-b489-4fc9-8096-40cd6553ee47"
# Predicting the Test set results
nn_prediction_test = model.predict(test)
nn_prediction_test = (nn_prediction_test > 0.5)*1 # convert probabilities to binary output
# Compute error between predicted data and true response
acc_test_ann2 = round(metrics.accuracy_score(target_test, nn_prediction_test) * 100, 2)
acc_test_ann2
# + [markdown] id="RFq7szeSqD_O"
# Voting_Classifier(hars_Voting)
# + colab={"base_uri": "https://localhost:8080/"} id="pgaXsPSBxA-T" outputId="1b904b8e-95e8-4375-8c1f-aa1cb779fe31"
Voting_Classifier_hard = VotingClassifier(estimators=[('lr', logreg), ('rf', random_forest), ('gbc', gradient_boosting)], voting='hard')
for clf, label in zip([logreg, random_forest, gradient_boosting, Voting_Classifier_hard],
['Logistic Regression', 'Random Forest', 'Gradient Boosting Classifier', 'Ensemble']):
scores = cross_val_score(clf, train, target, cv=10, scoring='accuracy')
print("Accuracy: %0.2f (+/- %0.2f) [%s]" % (scores.mean(), scores.std(), label))
# + colab={"base_uri": "https://localhost:8080/"} id="_B83cJ0hqL9b" outputId="58984fa7-41fd-40d3-dba3-0589abba499b"
Voting_Classifier_hard.fit(train, target)
acc_VC_hard = round(Voting_Classifier_hard.score(train, target) * 100, 2)
acc_VC_hard
# + colab={"base_uri": "https://localhost:8080/"} id="CVP9IeWMqM-N" outputId="8bd205e2-1d0b-4bcc-ec52-0c27654c4254"
acc_test_VC_hard = round(Voting_Classifier_hard.score(test, target_test) * 100, 2)
acc_test_VC_hard
# + [markdown] id="6AcCuawjqVUy"
# VotingClassifier (soft voting)
# + colab={"base_uri": "https://localhost:8080/"} id="MDk8CmdQqQ_j" outputId="0c6056d8-0819-4ce8-fb7a-a08a36ae0315"
eclf = VotingClassifier(estimators=[('lr', logreg), ('rf', random_forest), ('gbc', gradient_boosting)], voting='soft')
params = {'lr__C': [1.0, 100.0], 'gbc__learning_rate': [0.05, 1]}
Voting_Classifier_soft = GridSearchCV(estimator=eclf, param_grid=params, cv=5)
Voting_Classifier_soft.fit(train, target)
acc_VC_soft = round(Voting_Classifier_soft.score(train, target) * 100, 2)
acc_VC_soft
# + colab={"base_uri": "https://localhost:8080/"} id="We3EqGpGqZVB" outputId="f76d1b82-d930-4dbb-c1e1-aaf0c4f4904d"
acc_test_VC_soft = round(Voting_Classifier_soft.score(test, target_test) * 100, 2)
acc_test_VC_soft
# + [markdown] id="tlw-aTljqq-l"
# AdaBoost Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="1-eL246WqcF6" outputId="f41cdad5-7f27-4a5e-f62f-b9de2f93d4af"
def hyperopt_ab_score(params):
clf = AdaBoostClassifier(**params)
current_score = cross_val_score(clf, train, target, cv=10).mean()
print(current_score, params)
return current_score
space_ab = {
'n_estimators': hp.choice('n_estimators', range(50, 1000)),
'learning_rate': hp.quniform('learning_rate', 0, 0.05, 0.0001)
}
best = fmin(fn=hyperopt_ab_score, space=space_ab, algo=tpe.suggest, max_evals=10)
print('best:')
print(best)
# + colab={"base_uri": "https://localhost:8080/"} id="02SVgCcSqvR5" outputId="767c0a6c-117f-4ceb-c20a-e534e95e5ce3"
params = space_eval(space_ab, best)
params
# + colab={"base_uri": "https://localhost:8080/"} id="nAJuMif_qyeu" outputId="d3096640-5752-44b4-9b16-7563499a133c"
# AdaBoost Classifier
Ada_Boost = AdaBoostClassifier(**params)
Ada_Boost.fit(train, target)
Ada_Boost.score(train, target)
acc_AdaBoost = round(Ada_Boost.score(train, target) * 100, 2)
acc_AdaBoost
# + colab={"base_uri": "https://localhost:8080/"} id="RvT8oObpq1bI" outputId="8d7689bc-7694-4770-f31f-26f73a263252"
acc_test_AdaBoost = round(Ada_Boost.score(test, target_test) * 100, 2)
acc_test_AdaBoost
# + [markdown] id="gKUFoTfOq8_N"
# Models evaluation
# + id="YcpNAZcSq42M"
models = pd.DataFrame({
'Model': ['Logistic Regression', 'Support Vector Machines', 'Linear SVC', 'k-Nearest Neighbors', 'Naive Bayes',
'Perceptron', 'Stochastic Gradient Decent',
'Decision Tree Classifier', 'Random Forest', 'XGBClassifier', 'LGBMClassifier',
'GradientBoostingClassifier', 'RidgeClassifier', 'BaggingClassifier', 'ExtraTreesClassifier',
'Neural Network 1', 'Neural Network 2',
'VotingClassifier-hard voiting', 'VotingClassifier-soft voting',
'AdaBoostClassifier'],
'Score_train': [acc_log, acc_svc, acc_linear_svc, acc_knn, acc_gaussian,
acc_perceptron, acc_sgd,
acc_decision_tree, acc_random_forest, acc_XGB_Classifier, acc_LGB_Classifier,
acc_gradient_boosting, acc_ridge_classifier, acc_bagging_classifier, acc_etc,
acc_ann1, acc_ann2,
acc_VC_hard, acc_VC_soft,
acc_AdaBoost],
'Score_test': [acc_test_log, acc_test_svc, acc_test_linear_svc, acc_test_knn, acc_test_gaussian,
acc_test_perceptron, acc_test_sgd,
acc_test_decision_tree, acc_test_random_forest, acc_test_XGB_Classifier, acc_test_LGB_Classifier,
acc_test_gradient_boosting, acc_test_ridge_classifier, acc_test_bagging_classifier, acc_test_etc,
acc_test_ann1, acc_test_ann2,
acc_test_VC_hard, acc_test_VC_soft,
acc_test_AdaBoost]
})
# + colab={"base_uri": "https://localhost:8080/", "height": 669} id="_5rGg1XdrDOi" outputId="b41bfd92-4295-4917-e5da-0f0567f7a09a"
models.sort_values(by=['Score_train', 'Score_test'], ascending=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 669} id="5C0_eExprG5g" outputId="04bf1d60-29ee-4d20-af51-a933709a74e0"
models.sort_values(by=['Score_test', 'Score_train'], ascending=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 669} id="YZDMVbKHrKCY" outputId="2528166a-8fc6-4ebf-f9c0-192be2ddb5a6"
models['Score_diff'] = abs(models['Score_train'] - models['Score_test'])
models.sort_values(by=['Score_diff'], ascending=True)
# + id="2bHHz4zZrNg4"
# Plot
import matplotlib.pyplot as plt
plt.figure(figsize=[25,6])
xx = models['Model']
plt.tick_params(labelsize=14)
plt.plot(xx, models['Score_train'], label = 'Score_train')
plt.plot(xx, models['Score_test'], label = 'Score_test')
plt.legend()
plt.title('Score of 20 popular models for train and test datasets')
plt.xlabel('Models')
plt.ylabel('Score, %')
plt.xticks(xx, rotation='vertical')
plt.savefig('graph.png')
plt.show()
# + id="amdn3iknrRIm"
| Pipeline/popular_models_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="2HZFjYitOZyx"
import os
import numpy as np
import cv2
import pickle
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.model_selection import train_test_split
# + id="6x-LdZLxfwaU"
from google.colab.patches import cv2_imshow
#/content/drive/MyDrive/data/data.pickle
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="FgGnTDn00YXe" outputId="41121eba-1354-4cd2-d551-e8d74ccbbef8"
tf.test.gpu_device_name()
# + colab={"base_uri": "https://localhost:8080/"} id="1snRiSmL0p4B" outputId="818f5c78-d012-4fc6-8126-064154bf04e6"
# !ln -sf /opt/bin/nvidia-smi /usr/bin/nvidia-smi
# !pip install gputil
# !pip install psutil
# !pip install humanize
import psutil
import humanize
import os
import GPUtil as GPU
GPUs = GPU.getGPUs()
# XXX: only one GPU on Colab and isn’t guaranteed
gpu = GPUs[0]
def printm():
process = psutil.Process(os.getpid())
print("Gen RAM Free: " + humanize.naturalsize( psutil.virtual_memory().available ), " | Proc size: " + humanize.naturalsize( process.memory_info().rss))
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
printm()
# + id="rZZvDUM7zJdb"
i=int(1)
# + colab={"base_uri": "https://localhost:8080/"} id="wCDmNFtPaPLs" outputId="f8180c99-93ac-43e0-ddbc-2f81be13d06f"
data_dir = '/content/drive/MyDrive/archiveofflower/flowers/flowers'
categories = ['daisy','dandelion','rose','sunflower','tulip']
data = []
def make_data():
for category in categories:
global i
print(i)
i=i+1;
path = os.path.join(data_dir,category)
label = categories.index(category)
for img_name in os.listdir(path):
image_path = os.path.join(path,img_name)
image = cv2.imread(image_path)
try:
image = cv2.cvtColor(image,cv2.COLOR_BGR2RGB)
image = cv2.resize(image,(224,224))
image = np.array(image,dtype=np.float32)
data.append([image,label])
except Exception as e:
pass
#pik = open('data.pickle','wb')
with open('/content/drive/MyDrive/data/data.pickle', 'wb') as f:
pickle.dump(data, f)
#pickle.dump(data,pik)
f.close()
make_data()
# + id="k07qjLZagigk"
def load_data():
np.random.shuffle(data)
feature = []
labels = []
for img,label in data:
feature.append(img)
labels.append(label)
feature = np.array(feature,dtype=np.float32)
labels = np.array(labels)
feature = feature/255.0
return [feature,labels]
# + id="MV8ZF3BO3qrC"
(features,labels) = load_data()
X_train, X_test, y_train, y_test = train_test_split(features,labels,test_size=0.1)
categories = ['daisy','dandelion','rose','sunflower','tulip']
# + colab={"base_uri": "https://localhost:8080/"} id="xK-rddkXu4CF" outputId="c1dc9295-b7e8-4b55-8d34-63e3a01f4066"
X_train.shape
# + id="qN7AODCt5hIr"
input_layer = tf.keras.layers.Input([224,224,3])
conv1 = tf.keras.layers.Conv2D(filters = 32,kernel_size=(5,5),padding='Same',activation='relu')(input_layer)
pool1 = tf.keras.layers.MaxPooling2D(pool_size=(2,2))(conv1)
conv2 = tf.keras.layers.Conv2D(filters = 64,kernel_size=(3,3),padding='Same',activation='relu')(pool1)
pool2 = tf.keras.layers.MaxPooling2D(pool_size=(2,2),strides=(2,2))(conv2)
conv3 = tf.keras.layers.Conv2D(filters = 96,kernel_size=(3,3),padding='Same',activation='relu')(pool2)
pool3 = tf.keras.layers.MaxPooling2D(pool_size=(2,2),strides=(2,2))(conv3)
conv4 = tf.keras.layers.Conv2D(filters = 96,kernel_size=(3,3),padding='Same',activation='relu')(pool3)
pool4 = tf.keras.layers.MaxPooling2D(pool_size=(2,2),strides=(2,2))(conv4)
flt1 = tf.keras.layers.Flatten()(pool4)
dn1 = tf.keras.layers.Dense(512,activation='relu')(flt1)
out = tf.keras.layers.Dense(5,activation='softmax')(dn1)
model = tf.keras.Model(input_layer,out)
# + id="6nEpSDYH_Of4"
model.compile(optimizer='adam',loss='sparse_categorical_crossentropy',metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="gQ9-I0BmxdVz" outputId="3d640175-a5f0-48be-9567-a719b5eb958b"
model.fit(X_train,y_train,batch_size=100,epochs=10)
# + id="Hr1uypSQyINr"
model.save('my_model.h5')
# + colab={"base_uri": "https://localhost:8080/"} id="4eFzHo37yo1d" outputId="016bf981-9091-4025-8668-4cd08dc91dc4"
model.evaluate(X_test,y_test,verbose=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 884} id="QT7nZoWZyv71" outputId="7e07246c-6fde-420a-81fe-b0d5a8c685d3"
prediction = model.predict(X_test)
plt.figure(figsize=(9,9))
for i in range(9):
plt.subplot(3,3,i+1)
plt.imshow(X_test[i])
plt.xlabel('actual : '+categories[y_test[i]]+'\n'+'Predicted: \n'+
categories[np.argmax(prediction[i])])
print('\n')
plt.xticks([])
plt.show()
# + id="1c70LxHb3tfn"
| PIAIC Assignments/Deep Learning Assignments Set/Flowers_Recongnition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Measurement simulation
# A way to simulate data from measurements of a specific quantum state.
#
# Start with standard imports:
import matplotlib.pyplot as plt
from numpy import sqrt,pi,cos,sin,arange,random,real,imag
from qutip import *
# %matplotlib inline
# Define several standard states, these are photon polarization states:
H = Qobj([[1],[0]])
V = Qobj([[0],[1]])
P45 = Qobj([[1/sqrt(2)],[1/sqrt(2)]])
M45 = Qobj([[1/sqrt(2)],[-1/sqrt(2)]])
R = Qobj([[1/sqrt(2)],[-1j/sqrt(2)]])
L = Qobj([[1/sqrt(2)],[1j/sqrt(2)]])
# Define the Phv measurement operator:
Phv = H*H.dag() - V*V.dag()
Phv
# Define a quantum state: $$|\psi\rangle = \frac{1}{\sqrt{5}} |H\rangle + \frac{2}{\sqrt{5}} |V\rangle$$
psi = 1/sqrt(5)*H + 2/sqrt(5)*V
psi
# The function to generate a mock data set:
def simulateData(state,oper,size=10000):
"""Generate a simulated data set given a state and measurement operator.
state -> the prepared state
oper -> the measurement operator
Example:
H = Qobj([[1],[0]])
V = Qobj([[0],[1]])
psi = 1/sqrt(5)*H + 2/sqrt(5)*V
Phv = H*H.dag() - V*V.dag()
data = simulateData(psi,Phv)
will generate 10000 values in the data array that obey the probability defined in the state.
"""
A = basis(2,0)
B = basis(2,1)
allowed_results = [r.data.data[0] for r in [A.dag()*oper*A, B.dag()*oper*B]]
probability_amps = [qo.data.data[0] for qo in [A.dag()*state, B.dag()*state]]
pvals = [abs(pa.conjugate()*pa) for pa in probability_amps]
data = random.choice(allowed_results,size=size,p=pvals)
return data
data = simulateData(psi,Phv)
print("Variance: ",data.var())
print("Mean: ",data.mean())
plt.hist(real(data))
| Simulating measurements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jrhumberto/cd/blob/main/003_NLP.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="WN0t7zMhnlR7"
# Fontes:
# - https://github.com/aasouzaconsult/Cientista-de-Dados/tree/master/NLP%20-%20Classifica%C3%A7%C3%A3o%20de%20Not%C3%ADcias%20Curtas%20PTB
# - https://medium.com/blog-do-zouza/classifica%C3%A7%C3%A3o-de-not%C3%ADcias-utilizando-machine-learning-b25ff63ea51f
# + id="-NMzrCabnh7M"
from time import time
from tabulate import tabulate
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import gensim
import pickle
from gensim.models.word2vec import Word2Vec
from gensim.models import FastText
from collections import Counter, defaultdict
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.multiclass import OneVsRestClassifier
from sklearn.pipeline import Pipeline
from sklearn.svm import SVC
from sklearn import tree
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, classification_report, confusion_matrix
from sklearn.metrics import average_precision_score
from sklearn import metrics
from sklearn.preprocessing import label_binarize
from sklearn.ensemble import RandomForestClassifier
# + id="UofhD3QEqfkZ"
class E2V_AVG(object):
def __init__(self, word2vec):
self.w2v = word2vec
self.dimensao = 300
def fit(self, X, y):
return self
def transform(self, X):
return np.array([
np.mean([self.w2v[word] for word in words if word in self.w2v] or [np.zeros(self.dimensao)], axis=0)
for words in X
])
# + id="1VT-ScVtqfoe"
# ReferÊncia (SOUZA, 2019)
class E2V_IDF(object):
def __init__(self, word2vec):
self.w2v = word2vec
self.wIDF = None # IDF da palavra na colecao
self.dimensao = 300
def fit(self, X, y):
tfidf = TfidfVectorizer(analyzer=lambda x: x)
tfidf.fit(X)
maximo_idf = max(tfidf.idf_) # Uma palavra que nunca foi vista (rara) então o IDF padrão é o máximo de idfs conhecidos (exemplo: 9.2525763918954524)
self.wIDF = defaultdict(
lambda: maximo_idf,
[(word, tfidf.idf_[i]) for word, i in tfidf.vocabulary_.items()])
return self
# Gera um vetor de 300 dimensões, para cada documento, com a média dos vetores (embeddings) dos termos * IDF, contidos no documento.
def transform(self, X):
return np.array([
np.mean([self.w2v[word] * self.wIDF[word] for word in words if word in self.w2v] or [np.zeros(self.dimensao)], axis=0)
for words in X
])
# + id="c-WKZe2CqftH"
# Arquivo com nóticias curtas em Português do site G1
X = pickle.load(open('/data/z6News_X.ipy', 'rb'))
# Arquivo com o rótulos das notícias
y = pickle.load(open('/data/z6News_y.ipy', 'rb'))
# Essa fonte de dados é própria e esta disponível aqui no GitHub na Pasta: data
# - Podem utilizar, bastando referenciar o autor: SOUZA, 2019 (descrito na seção Referências)
# + id="px1dSdeiqfwv"
# Tranformando em Array
X, y = np.array(X), np.array(y)
# + id="aqAAHo4Cqf0K"
print ("Total de Notícias - G1: %s" % len(y))
# + id="wcAxddBrqf4D"
model = Word2Vec(X, size=300, window=5, sg=1, workers=4)
w2v = {w: vec for w, vec in zip(model.wv.index2word, model.wv.vectors)}
# + id="5EndL7Qlqf71"
# + id="Pc84VF_DqgAa"
| 003_NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # Symbolic Fuzzing
#
# One of the problems with traditional methods of fuzzing is that they fail to exercise all the possible behaviors that a system can have, especially when the input space is large. Quite often the execution of a specific branch of execution may happen only with very specific inputs, which could represent an extremely small fraction of the input space. The traditional fuzzing methods relies on chance to produce inputs they need. However, relying on randomness to generate values that we want is a bad idea when the space to be explored is huge. For example, a function that accepts a string, even if one only considers the first $10$ characters, already has $2^{80}$ possible inputs. If one is looking for a specific string, random generation of values will take a few thousand years even in one of the super computers.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# In the [chapter on concolic testing](ConcolicFuzzer.ipynb), we have seen how _concolic tracing_ can offer a way out. We saw how concolic tracing can be implemented using direct information flows using the Python interpreter. However, there are two problems with this approach.
# * The first is that concolic tracing relies on the existence of sample inputs. What if one has no sample inputs?
# * Second, direct information flows could be unreliable if the program has indirect information flows such as those based on control flow.
#
# In both cases, _static code analysis_ can bridge the gap. However, that raises the question: Can we determine the complete behavior of the program by examining it statically, and check if it behaves unexpectedly under some (unknown) input or result in an unexpected output?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# _Symbolic execution_ is one of the ways that we can reason about the behavior of a program without executing it. A program is a computation that can be treated as a system of equations that obtains the output values from the given inputs. Executing the program symbolically -- that is, solving these mathematically -- along with any specified objective such as covering a particular branch or obtaining a particular output will get us inputs that can accomplish this task.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "fragment"}
# In this chapter, we investigate how symbolic execution can be implemented, and how it can be used to obtain interesting values for fuzzing.
# + slideshow={"slide_type": "skip"}
from bookutils import YouTubeVideo
YouTubeVideo('RLQ_ORBezkk')
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# **Prerequisites**
#
# * You should understand how to use [type annotations](https://docs.python.org/3/library/typing.html) in Python.
# * A working knowledge of [SMT solvers](https://en.wikipedia.org/wiki/Satisfiability_modulo_theories), especially [Z3](https://github.com/Z3Prover/z3) is useful.
# * You should have read the [chapter on coverage](Coverage.ipynb).
# * A familiarity with [chapter on concolic fuzzing](ConcolicFuzzer.ipynb) would be helpful.
# + [markdown] jp-MarkdownHeadingCollapsed=true slideshow={"slide_type": "skip"} tags=[]
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from fuzzingbook.SymbolicFuzzer import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter provides an implementation of a symbolic fuzzing engine `SymbolicFuzzer`. The fuzzer uses symbolic execution to exhaustively explore paths in the program to a limited depth, and generate inputs that will reach these paths.
#
# As an example, consider the function `gcd()`, computing the greatest common divisor of `a` and `b`:
#
# ```python
# def gcd(a: int, b: int) -> int:
# if a < b:
# c: int = a # type: ignore
# a = b
# b = c
#
# while b != 0:
# c: int = a # type: ignore
# a = b
# b = c % b
#
# return a
# ```
# To explore `gcd()`, the fuzzer can be used as follows, producing values for arguments that cover different paths in `gcd()` (including multiple times of loop iterations):
#
# ```python
# >>> gcd_fuzzer = SymbolicFuzzer(gcd, max_tries=10, max_iter=10, max_depth=10)
# >>> for i in range(10):
# >>> args = gcd_fuzzer.fuzz()
# >>> print(args)
# {'a': 5, 'b': 3}
# {'a': 1, 'b': 4}
# {'a': 4, 'b': 5}
# {'a': 6, 'b': -7}
# {'a': 3, 'b': 4}
# {'a': 1, 'b': 1}
# {'a': 13, 'b': 7}
# {'a': 2, 'b': 4}
# {'a': 6, 'b': 6}
# {'a': 9, 'b': 8}
#
# ```
# Note that the variable values returned by `fuzz()` are Z3 _symbolic_ values; to convert them to Python numbers, use their method `as_long()`:
#
# ```python
# >>> for i in range(10):
# >>> args = gcd_fuzzer.fuzz()
# >>> a = args['a'].as_long()
# >>> b = args['b'].as_long()
# >>> d = gcd(a, b)
# >>> print(f"gcd({a}, {b}) = {d}")
# gcd(0, 5) = 5
# gcd(-1, 0) = -1
# gcd(14, 13) = 1
# gcd(0, 14) = 14
# gcd(14, 15) = 1
# gcd(15, 15) = 15
# gcd(2, 3) = 1
# gcd(16, 0) = 16
# gcd(16, -1) = -1
# gcd(-1, 1) = -1
#
# ```
# The symbolic fuzzer is subject to a number of constraints. First, it requires that the function to be fuzzed has correct type annotations, including all local variables. Second, it solves loops by unrolling them, but only for a fixed amount.
#
# For programs without loops and variable reassignments, the `SimpleSymbolicFuzzer` is a faster, but more limited alternative.
#
# 
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Obtaining Path Conditions for Coverage
# + [markdown] slideshow={"slide_type": "subslide"}
# In the chapter on [parsing and recombining inputs](SearchBasedFuzzer.ipynb), we saw how difficult it was to generate inputs for `process_vehicle()` -- a simple function that accepts a string. The solution given there was to rely on preexisting sample inputs. However, this solution is inadequate as it assumes the existence of sample inputs. What if there are no sample inputs at hand?
#
# For a simpler example, let us consider the following triangle function (which we already have seen in the [chapter on concolic fuzzing](ConcolicFuzzer.ipynb)). Can we generate inputs to cover all the paths?
#
# *Note.* We use type annotations to denote the argument types of programs. The [chapter on discovering dynamic invariants](DynamicInvariants.ipynb) will discuss how these types can be inferred automatically.
# + slideshow={"slide_type": "subslide"}
def check_triangle(a: int, b: int, c: int) -> str:
if a == b:
if a == c:
if b == c:
return "Equilateral"
else:
return "Isosceles"
else:
return "Isosceles"
else:
if b != c:
if a == c:
return "Isosceles"
else:
return "Scalene"
else:
return "Isosceles"
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The Control Flow Graph
# + [markdown] slideshow={"slide_type": "fragment"}
# The control flow graph of this function can be represented as follows:
# + slideshow={"slide_type": "skip"}
import bookutils
# + slideshow={"slide_type": "skip"}
import inspect
# + slideshow={"slide_type": "skip"}
from ControlFlow import PyCFG, to_graph, gen_cfg
# + slideshow={"slide_type": "fragment"}
def show_cfg(fn, **kwargs):
return to_graph(gen_cfg(inspect.getsource(fn)), **kwargs)
# + slideshow={"slide_type": "fragment"}
show_cfg(check_triangle)
# + [markdown] slideshow={"slide_type": "fragment"}
# The possible execution paths traced by the program can be represented as follows, with the numbers indicating the specific line numbers executed.
# + slideshow={"slide_type": "subslide"}
paths = {
'<path 1>': ([1, 2, 3, 4, 5], 'Equilateral'),
'<path 2>': ([1, 2, 3, 4, 7], 'Isosceles'),
'<path 3>': ([1, 2, 3, 9], 'Isosceles'),
'<path 4>': ([1, 2, 11, 12, 13], 'Isosceles'),
'<path 5>': ([1, 2, 11, 12, 15], 'Scalene'),
'<path 6>': ([1, 2, 11, 17], 'Isosceles'),
}
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider the `<path 1>`. To trace this path, we need to execute the following statements in order.
# + [markdown] slideshow={"slide_type": "subslide"}
# ```python
# 1: check_triangle(a, b, c)
# 2: if (a == b) -> True
# 3: if (a == c) -> True
# 4: if (b == c) -> True
# 5: return 'Equilateral'
# ```
# + [markdown] slideshow={"slide_type": "fragment"}
# That is, any execution that traces this path has to start with values for `a`, `b`, and `c` that obeys the constraints in line numbers `2: (a == b)` evaluates to `True`, `3: (a == c)` evaluates to `True`, and `4: (b == c)` evaluates to `True`. Can we generate inputs such that these constraints are satisfied?
# + [markdown] slideshow={"slide_type": "fragment"}
# We have seen from the [chapter on concolic fuzzing](ConcolicFuzzer.ipynb) how one can use an SMT solver such as Z3 to obtain a solution.
# + slideshow={"slide_type": "skip"}
import z3 # type: ignore
# + slideshow={"slide_type": "subslide"}
z3_ver = z3.get_version()
print(z3_ver)
# + slideshow={"slide_type": "fragment"}
assert z3_ver >= (4, 8, 6, 0), "Please check z3 version"
# + [markdown] slideshow={"slide_type": "fragment"}
# What kind of symbolic variables do we need? We can obtain that information from the type annotations of the function.
# + slideshow={"slide_type": "fragment"}
def get_annotations(fn):
sig = inspect.signature(fn)
return ([(i.name, i.annotation)
for i in sig.parameters.values()], sig.return_annotation)
# + slideshow={"slide_type": "fragment"}
params, ret = get_annotations(check_triangle)
params, ret
# + [markdown] slideshow={"slide_type": "fragment"}
# We create symbolic variables to represent each of the parameters
# + slideshow={"slide_type": "subslide"}
SYM_VARS = {
int: (
z3.Int, z3.IntVal), float: (
z3.Real, z3.RealVal), str: (
z3.String, z3.StringVal)}
# + slideshow={"slide_type": "fragment"}
def get_symbolicparams(fn):
params, ret = get_annotations(fn)
return [SYM_VARS[typ][0](name)
for name, typ in params], SYM_VARS[ret][0]('__return__')
# + slideshow={"slide_type": "fragment"}
(a, b, c), r = get_symbolicparams(check_triangle)
a, b, c, r
# + [markdown] slideshow={"slide_type": "fragment"}
# We can now ask *z3* to solve the set of equations for us as follows.
# + slideshow={"slide_type": "subslide"}
z3.solve(a == b, a == c, b == c)
# + [markdown] slideshow={"slide_type": "fragment"}
# Here we find the first problem in our program. Our program seems to not check whether the sides are greater than zero. (Real-world triangles all have sides with a positive length.) Assume for now that we do not have that restriction. Does our program correctly follow the path described?
#
# We can use the `ArcCoverage` from the [chapter on concolic fuzzing](ConcolicFuzzer.ipynb) as a tracer to visualize that information as below.
# + slideshow={"slide_type": "skip"}
from ConcolicFuzzer import ArcCoverage # minor dependency
# + [markdown] slideshow={"slide_type": "fragment"}
# First, we recover the trace.
# + slideshow={"slide_type": "subslide"}
with ArcCoverage() as cov:
assert check_triangle(0, 0, 0) == 'Equilateral'
cov._trace, cov.arcs()
# + [markdown] slideshow={"slide_type": "fragment"}
# We can now determine the path taken.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The CFG with Path Taken
# + slideshow={"slide_type": "fragment"}
show_cfg(check_triangle, arcs=cov.arcs())
# + [markdown] slideshow={"slide_type": "fragment"}
# As you can see, the path taken is `<path 1>`.
# + [markdown] slideshow={"slide_type": "fragment"}
# Similarly, for solving `<path 2>` we need to simply invert the condition at <line 2>:
# + slideshow={"slide_type": "fragment"}
z3.solve(a == b, a == c, z3.Not(b == c))
# + [markdown] slideshow={"slide_type": "fragment"}
# The symbolic execution suggests that there is no solution. A moment's reflection will convince us that it is indeed true. Let us proceed with the other paths. The `<path 3>` can be obtained by inverting the condition at `<line 4>`.
# + slideshow={"slide_type": "subslide"}
z3.solve(a == b, z3.Not(a == c))
# + slideshow={"slide_type": "fragment"}
with ArcCoverage() as cov:
assert check_triangle(1, 1, 0) == 'Isosceles'
[i for fn, i in cov._trace if fn == 'check_triangle']
# + slideshow={"slide_type": "fragment"}
paths['<path 3>']
# + [markdown] slideshow={"slide_type": "fragment"}
# How about path <4>?
# + slideshow={"slide_type": "fragment"}
z3.solve(z3.Not(a == b), b != c, a == c)
# + [markdown] slideshow={"slide_type": "subslide"}
# As we mentioned earlier, our program does not account for sides with zero or negative length. We can modify our program to check for zero and negative input. However, do we always have to make sure that every function has to account for all possible inputs? It is possible that the `check_triangle` is not directly exposed to the user, and it is called from another function that already guarantees that the inputs would be positive. In the [chapter on dynamic invariants](DynamicInvariants.ipynb), we will show how to discover such preconditions and post conditions.
#
# We can easily add such a precondition here.
# + slideshow={"slide_type": "fragment"}
pre_condition = z3.And(a > 0, b > 0, c > 0)
# + slideshow={"slide_type": "fragment"}
z3.solve(pre_condition, z3.Not(a == b), b != c, a == c)
# + slideshow={"slide_type": "subslide"}
with ArcCoverage() as cov:
assert check_triangle(1, 2, 1) == 'Isosceles'
[i for fn, i in cov._trace if fn == 'check_triangle']
# + slideshow={"slide_type": "fragment"}
paths['<path 4>']
# + [markdown] slideshow={"slide_type": "fragment"}
# Continuing to path <5>:
# + slideshow={"slide_type": "fragment"}
z3.solve(pre_condition, z3.Not(a == b), b != c, z3.Not(a == c))
# + [markdown] slideshow={"slide_type": "fragment"}
# And indeed it is a *Scalene* triangle.
# + slideshow={"slide_type": "fragment"}
with ArcCoverage() as cov:
assert check_triangle(3, 1, 2) == 'Scalene'
# + slideshow={"slide_type": "fragment"}
paths['<path 5>']
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, for `<path 6>` the procedure is similar.
# + slideshow={"slide_type": "fragment"}
z3.solve(pre_condition, z3.Not(a == b), z3.Not(b != c))
# + slideshow={"slide_type": "fragment"}
with ArcCoverage() as cov:
assert check_triangle(2, 1, 1) == 'Isosceles'
[i for fn, i in cov._trace if fn == 'check_triangle']
# + slideshow={"slide_type": "fragment"}
paths['<path 6>']
# + [markdown] slideshow={"slide_type": "fragment"}
# What if we wanted another solution? We can simply ask the solver to solve again, and not give us the same values.
# + slideshow={"slide_type": "fragment"}
seen = [z3.And(a == 2, b == 1, c == 1)]
# + slideshow={"slide_type": "subslide"}
z3.solve(pre_condition, z3.Not(z3.Or(seen)), z3.Not(a == b), z3.Not(b != c))
# + slideshow={"slide_type": "fragment"}
seen.append(z3.And(a == 1, b == 2, c == 2))
# + slideshow={"slide_type": "fragment"}
z3.solve(pre_condition, z3.Not(z3.Or(seen)), z3.Not(a == b), z3.Not(b != c))
# + [markdown] slideshow={"slide_type": "fragment"}
# That is, using simple symbolic computation, we were able to easily see that (1) some of the paths are not reachable, and (2) some of the conditions were insufficient -- we needed preconditions. What about the total coverage obtained?
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Visualizing the Coverage
# + [markdown] slideshow={"slide_type": "fragment"}
# Visualizing the statement coverage can be accomplished as below.
# + slideshow={"slide_type": "fragment"}
class VisualizedArcCoverage(ArcCoverage):
def show_coverage(self, fn):
src = fn if isinstance(fn, str) else inspect.getsource(fn)
covered = set([lineno for method, lineno in self._trace])
for i, s in enumerate(src.split('\n')):
print('%s %2d: %s' % ('#' if i + 1 in covered else ' ', i + 1, s))
# + [markdown] slideshow={"slide_type": "fragment"}
# We run all the inputs obtained under the coverage tracer.
# + slideshow={"slide_type": "fragment"}
with VisualizedArcCoverage() as cov:
assert check_triangle(0, 0, 0) == 'Equilateral'
assert check_triangle(1, 1, 0) == 'Isosceles'
assert check_triangle(1, 2, 1) == 'Isosceles'
assert check_triangle(3, 1, 2) == 'Scalene'
assert check_triangle(2, 1, 1) == 'Isosceles'
# + slideshow={"slide_type": "subslide"}
cov.show_coverage(check_triangle)
# + [markdown] slideshow={"slide_type": "subslide"}
# The coverage is as expected. The generated values does seem to cover all code that can be covered.
#
# We have seen how to reason about each path through the program. Can we combine them together to produce a single expression that represents the program behavior? This is what we will discuss next.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Function Summaries
# + [markdown] slideshow={"slide_type": "fragment"}
# Consider this equation for determining absolute value.
# + slideshow={"slide_type": "fragment"}
def abs_value(x: float) -> float:
if x < 0:
v: float = -x # type: ignore
else:
v: float = x # type: ignore
return v
# + slideshow={"slide_type": "fragment"}
show_cfg(abs_value)
# + [markdown] slideshow={"slide_type": "fragment"}
# What can we say about the value of `v` at `line: 5`? Let us trace and see. First, we have variable `x` at `line: 1`.
# + slideshow={"slide_type": "fragment"}
(x,), r = get_symbolicparams(abs_value)
# + [markdown] slideshow={"slide_type": "fragment"}
# At `line: 2`, we face a bifurcation in the possible paths. Hence, we produce two paths with corresponding constraints.
# + slideshow={"slide_type": "subslide"}
l2_T = x < 0
l2_F = z3.Not(x < 0)
# + [markdown] slideshow={"slide_type": "fragment"}
# For `line: 3`, we only need to consider the `If` path. However, we have an assignment. So we use a new variable here. The type _float_ is indicated in the source, and its equivalent *z3* type is _Real_.
# + slideshow={"slide_type": "fragment"}
v_0 = z3.Real('v_0')
l3 = z3.And(l2_T, v_0 == -x)
# + [markdown] slideshow={"slide_type": "fragment"}
# Similarly, for `line: 5`, we have an assignment. (Can we reuse the variable `v_0` from before?)
# + slideshow={"slide_type": "fragment"}
v_1 = z3.Real('v_1')
l5 = z3.And(l2_F, v_1 == x)
# + [markdown] slideshow={"slide_type": "fragment"}
# When we come to `line: 6`, we see that we have *two* input streams. We have a choice. We can either keep each path separate as we did previously.
# + slideshow={"slide_type": "subslide"}
v = z3.Real('v')
for s in [z3.And(l3, v == v_0), z3.And(l5, v == v_1)]:
z3.solve(x != 0, s)
# + [markdown] slideshow={"slide_type": "fragment"}
# Or, we can combine them together and produce a single predicate at `line: 6`.
# + slideshow={"slide_type": "fragment"}
v = z3.Real('v')
l6 = z3.Or(z3.And(l3, v == v_0), z3.And(l5, v == v_1))
z3.solve(l6)
# + [markdown] slideshow={"slide_type": "subslide"}
# **Note.** Merging two incoming streams of execution can be non-trivial, especially when the execution paths are traversed multiple times (E.g. loops and recursion). For those interested, lookup [inferring loop invariants](https://www.st.cs.uni-saarland.de/publications/details/galeotti-hvc-2014/).
# + [markdown] slideshow={"slide_type": "fragment"}
# We can get this to produce any number of solutions for `abs()` as below.
# + slideshow={"slide_type": "subslide"}
s = z3.Solver()
s.add(l6)
for i in range(5):
if s.check() == z3.sat:
m = s.model()
x_val = m[x]
print(m)
else:
print('no solution')
break
s.add(z3.Not(x == x_val))
s
# + [markdown] slideshow={"slide_type": "subslide"}
# The solver is not particularly random. So we need to help it a bit to produce values on the negative range.
# + slideshow={"slide_type": "subslide"}
s.add(x < 0)
for i in range(5):
if s.check() == z3.sat:
m = s.model()
x_val = m[x]
print(m)
else:
print('no solution')
break
s.add(z3.Not(x == x_val))
# + slideshow={"slide_type": "subslide"}
s
# + [markdown] slideshow={"slide_type": "subslide"}
# Note that the single expression produced at `line: 6` is essentially a summary for `abs_value()`.
# + slideshow={"slide_type": "fragment"}
abs_value_summary = l6
abs_value_summary
# + [markdown] slideshow={"slide_type": "fragment"}
# The *z3* solver can be used to simplify the predicates where possible.
# + slideshow={"slide_type": "fragment"}
z3.simplify(l6)
# + [markdown] slideshow={"slide_type": "subslide"}
# One can use this summary rather than trace into `abs_value()` when `abs_value()` is used elsewhere. However, that presents us with a problem. It is possible that the same function may be called multiple times. In this case, using the same variables will lead to collision. One way to avoid that is to *prefix* some call specific value to the variables.
#
# **Note:** The SMT 2.0 standard allows one to define functions (*macros* in SMT parlance) directly. For example, the `abs-value` will be defined as follows:
#
# ```lisp
# (define-fun abs-value ((x Int)) Int
# (if (> x 0)
# x
# (* -1 x)))
# ```
#
# Or equivalantly, (especially if `abs-value` is recursively defined)
#
# ```lisp
# (declare-fun abs-value (Int) Int)
# (assert (forall ((x Int))
# (= (abs-value x)
# (if (> x 0)
# x
# (* -1 x)))))
# ```
# One can then say
# ```
# (> (abs-value x) (abs-value y))
# ```
#
# Unfortunately, the z3py project does not expose this facility in Python. Hence we have to use the `prefix_vars()` hack.
# + slideshow={"slide_type": "skip"}
import ast
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `prefix_vars()` modifies the variables in an expression such that the variables are prefixed with a given value.
# + slideshow={"slide_type": "subslide"}
def prefix_vars(astnode, prefix):
if isinstance(astnode, ast.BoolOp):
return ast.BoolOp(astnode.op,
[prefix_vars(i, prefix) for i in astnode.values], [])
elif isinstance(astnode, ast.BinOp):
return ast.BinOp(
prefix_vars(astnode.left, prefix), astnode.op,
prefix_vars(astnode.right, prefix))
elif isinstance(astnode, ast.UnaryOp):
return ast.UnaryOp(astnode.op, prefix_vars(astnode.operand, prefix))
elif isinstance(astnode, ast.Call):
return ast.Call(prefix_vars(astnode.func, prefix),
[prefix_vars(i, prefix) for i in astnode.args],
astnode.keywords)
elif isinstance(astnode, ast.Compare):
return ast.Compare(
prefix_vars(astnode.left, prefix), astnode.ops,
[prefix_vars(i, prefix) for i in astnode.comparators])
elif isinstance(astnode, ast.Name):
if astnode.id in {'And', 'Or', 'Not'}:
return ast.Name('z3.%s' % (astnode.id), astnode.ctx)
else:
return ast.Name('%s%s' % (prefix, astnode.id), astnode.ctx)
elif isinstance(astnode, ast.Return):
return ast.Return(prefix_vars(astnode.value, env))
else:
return astnode
# + [markdown] slideshow={"slide_type": "subslide"}
# For applying `prefix_vars()` one needs the _abstract syntax tree_ (AST) of the Python expression involved. We obtain this by invoking `ast.parse()`:
# + slideshow={"slide_type": "fragment"}
xy_ast = ast.parse('x+y')
# + [markdown] slideshow={"slide_type": "fragment"}
# We can visualize the resulting tree as follows:
# + slideshow={"slide_type": "skip"}
from bookutils import rich_output
# + slideshow={"slide_type": "fragment"}
if rich_output():
# Normally, this will do
from showast import show_ast
else:
def show_ast(tree):
ast.dump(tree, indent=4)
# + slideshow={"slide_type": "fragment"}
show_ast(xy_ast)
# + [markdown] slideshow={"slide_type": "subslide"}
# What the visualization does _not_ show, though, is that when parsing Python source code, the resulting AST comes wrapped in a `Module` by default:
# + slideshow={"slide_type": "fragment"}
xy_ast
# + [markdown] slideshow={"slide_type": "fragment"}
# And to access the expression (`Expr`), we need to access the first child of that "module":
# + slideshow={"slide_type": "fragment"}
xy_ast.body[0]
# + [markdown] slideshow={"slide_type": "fragment"}
# The actual expression is within that `Expr` object:
# + slideshow={"slide_type": "fragment"}
xy_ast.body[0].value # type: ignore
# + [markdown] slideshow={"slide_type": "fragment"}
# Hence, for easier manipulation of an expression AST, we define a function `get_expression()` which unwraps it and returns the AST representation of the expression inside.
# + slideshow={"slide_type": "subslide"}
def get_expression(src):
return ast.parse(src).body[0].value
# + [markdown] slideshow={"slide_type": "fragment"}
# It is used as follows:
# + slideshow={"slide_type": "fragment"}
e = get_expression('x+y')
e
# + [markdown] slideshow={"slide_type": "fragment"}
# The function `to_src()` allows us to *unparse* an expression.
# + slideshow={"slide_type": "fragment"}
def to_src(astnode):
return ast.unparse(astnode).strip()
# + [markdown] slideshow={"slide_type": "fragment"}
# It is used as follows:
# + slideshow={"slide_type": "fragment"}
to_src(e)
# + [markdown] slideshow={"slide_type": "fragment"}
# We can combine both pieces to produce a prefixed expression. Let us prefix all variables with `x1_`:
# + slideshow={"slide_type": "subslide"}
abs_value_summary_ast = get_expression(str(abs_value_summary))
print(to_src(prefix_vars(abs_value_summary_ast, 'x1_')))
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Get Names and Types of Variables Used
# + [markdown] slideshow={"slide_type": "fragment"}
# What about the declarations used? Given that we have all equations in *Z3*, we can retrieve this information directly. We define `z3_names_and_types()` that takes in a *Z3* expression, and extracts the variable definitions required.
# + slideshow={"slide_type": "subslide"}
def z3_names_and_types(z3_ast):
hm = {}
children = z3_ast.children()
if children:
for c in children:
hm.update(z3_names_and_types(c))
else:
# HACK.. How else to distinguish literals and vars?
if (str(z3_ast.decl()) != str(z3_ast.sort())):
hm["%s" % str(z3_ast.decl())] = 'z3.%s' % str(z3_ast.sort())
else:
pass
return hm
# + slideshow={"slide_type": "subslide"}
abs_value_declarations = z3_names_and_types(abs_value_summary)
abs_value_declarations
# + [markdown] slideshow={"slide_type": "fragment"}
# However, `z3_names_and_types()` is limited in that it requires the *Z3* AST to operate. Hence, we also define `used_identifiers()` that can extract identifiers directly from the string representation of any Python expression, (including *Z3* constraints). One trade-off here is that we lose track of the type information. But we will see how to recover that later.
# + slideshow={"slide_type": "subslide"}
def used_identifiers(src):
def names(astnode):
lst = []
if isinstance(astnode, ast.BoolOp):
for i in astnode.values:
lst.extend(names(i))
elif isinstance(astnode, ast.BinOp):
lst.extend(names(astnode.left))
lst.extend(names(astnode.right))
elif isinstance(astnode, ast.UnaryOp):
lst.extend(names(astnode.operand))
elif isinstance(astnode, ast.Call):
for i in astnode.args:
lst.extend(names(i))
elif isinstance(astnode, ast.Compare):
lst.extend(names(astnode.left))
for i in astnode.comparators:
lst.extend(names(i))
elif isinstance(astnode, ast.Name):
lst.append(astnode.id)
elif isinstance(astnode, ast.Expr):
lst.extend(names(astnode.value))
elif isinstance(astnode, (ast.Num, ast.Str, ast.Tuple, ast.NameConstant)):
pass
elif isinstance(astnode, ast.Assign):
for t in astnode.targets:
lst.extend(names(t))
lst.extend(names(astnode.value))
elif isinstance(astnode, ast.Module):
for b in astnode.body:
lst.extend(names(b))
else:
raise Exception(str(astnode))
return list(set(lst))
return names(ast.parse(src))
# + slideshow={"slide_type": "subslide"}
used_identifiers(str(abs_value_summary))
# + [markdown] slideshow={"slide_type": "fragment"}
# We can now register the function summary `abs_value` for later use.
# + slideshow={"slide_type": "fragment"}
function_summaries = {}
function_summaries['abs_value'] = {
'predicate': str(abs_value_summary),
'vars': abs_value_declarations}
# + [markdown] slideshow={"slide_type": "fragment"}
# As we mentioned previously, we do not want to rely on *Z3* to extract the type information. A better alternative is to let the user specify the type information as annotations, and extract this information from the program. We will see next how this can be achieved.
#
# First, we convert the *Python type to Z3 type* map to its string equivalent.
# + slideshow={"slide_type": "subslide"}
SYM_VARS_STR = {
k.__name__: ("z3.%s" % v1.__name__, "z3.%s" % v2.__name__)
for k, (v1, v2) in SYM_VARS.items()
}
SYM_VARS_STR
# + [markdown] slideshow={"slide_type": "fragment"}
# We also define a convenience method `translate_to_z3_name()` for accessing the *Z3* type for symbolic variables.
# + slideshow={"slide_type": "fragment"}
def translate_to_z3_name(v):
return SYM_VARS_STR[v][0]
# + [markdown] slideshow={"slide_type": "subslide"}
# We now define the method `declarations()` that extracts variables used in Python _statements_. The idea is to look for augmented assignments that contain annotated type information. These are collected and returned.
#
# If there are `call` nodes, they represent function calls. The used variables in these function calls are recovered from the corresponding function summaries.
# + slideshow={"slide_type": "subslide"}
def declarations(astnode, hm=None):
if hm is None:
hm = {}
if isinstance(astnode, ast.Module):
for b in astnode.body:
declarations(b, hm)
elif isinstance(astnode, ast.FunctionDef):
# hm[astnode.name + '__return__'] = \
# translate_to_z3_name(astnode.returns.id)
for a in astnode.args.args:
hm[a.arg] = translate_to_z3_name(a.annotation.id)
for b in astnode.body:
declarations(b, hm)
elif isinstance(astnode, ast.Call):
# get declarations from the function summary.
n = astnode.function
assert isinstance(n, ast.Name) # for now.
name = n.id
hm.update(dict(function_summaries[name]['vars']))
elif isinstance(astnode, ast.AnnAssign):
assert isinstance(astnode.target, ast.Name)
hm[astnode.target.id] = translate_to_z3_name(astnode.annotation.id)
elif isinstance(astnode, ast.Assign):
# verify it is already defined
for t in astnode.targets:
assert isinstance(t, ast.Name)
assert t.id in hm
elif isinstance(astnode, ast.AugAssign):
assert isinstance(astnode.target, ast.Name)
assert astnode.target.id in hm
elif isinstance(astnode, (ast.If, ast.For, ast.While)):
for b in astnode.body:
declarations(b, hm)
for b in astnode.orelse:
declarations(b, hm)
elif isinstance(astnode, ast.Return):
pass
else:
raise Exception(str(astnode))
return hm
# + [markdown] slideshow={"slide_type": "subslide"}
# With this, we can now extract the variables used in an expression.
# + slideshow={"slide_type": "fragment"}
declarations(ast.parse('s: int = 3\np: float = 4.0\ns += 1'))
# + [markdown] slideshow={"slide_type": "fragment"}
# We wrap `declarations()` in the method `used_vars()` that operates directly on function objects.
# + slideshow={"slide_type": "fragment"}
def used_vars(fn):
return declarations(ast.parse(inspect.getsource(fn)))
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is how it can be used:
# + slideshow={"slide_type": "fragment"}
used_vars(check_triangle)
# + slideshow={"slide_type": "fragment"}
used_vars(abs_value)
# + [markdown] slideshow={"slide_type": "subslide"}
# Given the extracted variables and their *Z3* types, we need a way to reinstantiate them when needed. We define `define_symbolic_vars()` that translates these descriptions to a form that can be directly `exec()`ed.
# + slideshow={"slide_type": "fragment"}
def define_symbolic_vars(fn_vars, prefix):
sym_var_dec = ', '.join([prefix + n for n in fn_vars])
sym_var_def = ', '.join(["%s('%s%s')" % (t, prefix, n)
for n, t in fn_vars.items()])
return "%s = %s" % (sym_var_dec, sym_var_def)
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is how it can be used:
# + slideshow={"slide_type": "fragment"}
define_symbolic_vars(abs_value_declarations, '')
# + [markdown] slideshow={"slide_type": "fragment"}
# We next define `gen_fn_summary()` that returns a function summary in instantiable form using *Z3*.
# + slideshow={"slide_type": "subslide"}
def gen_fn_summary(prefix, fn):
summary = function_summaries[fn.__name__]['predicate']
fn_vars = function_summaries[fn.__name__]['vars']
decl = define_symbolic_vars(fn_vars, prefix)
summary_ast = get_expression(summary)
return decl, to_src(prefix_vars(summary_ast, prefix))
# + [markdown] slideshow={"slide_type": "fragment"}
# Here is how it can be used:
# + slideshow={"slide_type": "fragment"}
gen_fn_summary('a_', abs_value)
# + slideshow={"slide_type": "fragment"}
gen_fn_summary('b_', abs_value)
# + [markdown] slideshow={"slide_type": "fragment"}
# How do we use our function summaries? Here is a function `abs_max()` that uses `abs_value()`.
# + slideshow={"slide_type": "subslide"}
def abs_max(a: float, b: float):
a1: float = abs_value(a)
b1: float = abs_value(b)
if a1 > b1:
c: float = a1 # type: ignore
else:
c: float = b1 # type: ignore
return c
# + [markdown] slideshow={"slide_type": "fragment"}
# To trace this function symbolically, we first define the two variables `a` and `b`.
# + slideshow={"slide_type": "fragment"}
a = z3.Real('a')
b = z3.Real('b')
# + [markdown] slideshow={"slide_type": "fragment"}
# The `line: 2` contains definition for `a1`, which we define as a symbolic variable.
# + slideshow={"slide_type": "fragment"}
a1 = z3.Real('a1')
# + [markdown] slideshow={"slide_type": "subslide"}
# We also need to call `abs_value()`, which is accomplished as follows. Since this is the first call to `abs_value()`, we use `abs1` as the prefix.
# + slideshow={"slide_type": "fragment"}
d, v = gen_fn_summary('abs1_', abs_value)
d, v
# + [markdown] slideshow={"slide_type": "fragment"}
# We also need to equate the resulting value (`<prefix>_v`) to the symbolic variable `a1` we defined earlier.
# + slideshow={"slide_type": "fragment"}
l2_src = "l2 = z3.And(a == abs1_x, a1 == abs1_v, %s)" % v
l2_src
# + [markdown] slideshow={"slide_type": "fragment"}
# Applying both declaration and the assignment.
# + slideshow={"slide_type": "fragment"}
exec(d)
exec(l2_src)
# + slideshow={"slide_type": "subslide"}
l2 # type: ignore
# + [markdown] slideshow={"slide_type": "fragment"}
# We need to do the same for `line: 3`, but with `abs2` as the prefix.
# + slideshow={"slide_type": "fragment"}
b1 = z3.Real('b1')
d, v = gen_fn_summary('abs2_', abs_value)
l3_src = "l3_ = z3.And(b == abs2_x, b1 == abs2_v, %s)" % v
exec(d)
exec(l3_src)
# + slideshow={"slide_type": "subslide"}
l3_ # type: ignore
# + [markdown] slideshow={"slide_type": "fragment"}
# To get the true set of predicates at `line: 3`, we need to add the predicates from `line: 2`.
# + slideshow={"slide_type": "fragment"}
l3 = z3.And(l2, l3_) # type: ignore
# + slideshow={"slide_type": "subslide"}
l3
# + [markdown] slideshow={"slide_type": "fragment"}
# This equation can be simplified a bit using z3.
# + slideshow={"slide_type": "subslide"}
z3.simplify(l3)
# + [markdown] slideshow={"slide_type": "subslide"}
# Coming to `line: 4`, we have a condition.
# + slideshow={"slide_type": "fragment"}
l4_cond = a1 > b1
l4 = z3.And(l3, l4_cond)
# + [markdown] slideshow={"slide_type": "fragment"}
# For `line: 5`, we define the symbolic variable `c_0` assuming we took the *IF* branch.
# + slideshow={"slide_type": "fragment"}
c_0 = z3.Real('c_0')
l5 = z3.And(l4, c_0 == a1)
# + [markdown] slideshow={"slide_type": "fragment"}
# For `line: 6`, the *ELSE* branch was taken. So we invert that condition.
# + slideshow={"slide_type": "fragment"}
l6 = z3.And(l3, z3.Not(l4_cond))
# + [markdown] slideshow={"slide_type": "fragment"}
# For `line: 7`, we define `c_1`.
# + slideshow={"slide_type": "fragment"}
c_1 = z3.Real('c_1')
l7 = z3.And(l6, c_1 == b1)
# + slideshow={"slide_type": "subslide"}
s1 = z3.Solver()
s1.add(l5)
s1.check()
# + slideshow={"slide_type": "fragment"}
m1 = s1.model()
sorted([(d, m1[d]) for d in m1.decls() if not d.name(
).startswith('abs')], key=lambda x: x[0].name())
# + slideshow={"slide_type": "fragment"}
s2 = z3.Solver()
s2.add(l7)
s2.check()
# + slideshow={"slide_type": "subslide"}
m2 = s2.model()
sorted([(d, m2[d]) for d in m2.decls() if not d.name(
).startswith('abs')], key=lambda x: x[0].name())
# + [markdown] slideshow={"slide_type": "fragment"}
# What we really want to do is to automate this process, because doing this by hand is tedious and error prone. Essentially, we want the ability to extract *all paths* in the program, and symbolically execute each path, which will generate the inputs required to cover all reachable portions of the program.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Simple Symbolic Fuzzing
# + [markdown] slideshow={"slide_type": "subslide"}
# We define a simple *symbolic fuzzer* that can generate input values *symbolically* with the following assumptions:
#
# * There are no loops in the program
# * The function is self contained.
# * No recursion.
# * No reassignments for variables.
#
# The key idea is as follows: We traverse through the control flow graph from the entry point, and generate all possible paths to a given depth. Then we collect constraints that we encountered along the path, and generate inputs that will traverse the program up to that point.
# + [markdown] slideshow={"slide_type": "subslide"}
# We build our fuzzer based on the class `Fuzzer`.
# + slideshow={"slide_type": "skip"}
from Fuzzer import Fuzzer
# + [markdown] slideshow={"slide_type": "fragment"}
# We start by extracting the control flow graph of the function passed. We also provide a hook for child classes to do their processing.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(Fuzzer):
"""Simple symbolic fuzzer"""
def __init__(self, fn, **kwargs):
"""Constructor.
`fn` is the function to be fuzzed.
Possible keyword parameters:
* `max_depth` - the depth to which one should attempt
to trace the execution (default 100)
* `max_tries` - the maximum number of attempts
we will try to produce a value before giving up (default 100)
* `max_iter` - the number of iterations we will attempt (default 100).
"""
self.fn_name = fn.__name__
py_cfg = PyCFG()
py_cfg.gen_cfg(inspect.getsource(fn))
self.fnenter, self.fnexit = py_cfg.functions[self.fn_name]
self.used_variables = used_vars(fn)
self.fn_args = list(inspect.signature(fn).parameters)
self.z3 = z3.Solver()
self.paths = None
self.last_path = None
self.options(kwargs)
self.process()
def process(self):
... # to be defined later
# + [markdown] slideshow={"slide_type": "subslide"}
# We need a few variables to control how much we are willing to traverse.
# + [markdown] slideshow={"slide_type": "fragment"}
# `MAX_DEPTH` is the depth to which one should attempt to trace the execution.
# + slideshow={"slide_type": "fragment"}
MAX_DEPTH = 100
# + [markdown] slideshow={"slide_type": "fragment"}
# `MAX_TRIES` is the maximum number of attempts we will try to produce a value before giving up.
# + slideshow={"slide_type": "fragment"}
MAX_TRIES = 100
# + [markdown] slideshow={"slide_type": "fragment"}
# `MAX_ITER` is the number of iterations we will attempt.
# + slideshow={"slide_type": "fragment"}
MAX_ITER = 100
# + [markdown] slideshow={"slide_type": "fragment"}
# The `options()` method sets these parameters in the fuzzing class.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def options(self, kwargs):
self.max_depth = kwargs.get('max_depth', MAX_DEPTH)
self.max_tries = kwargs.get('max_tries', MAX_TRIES)
self.max_iter = kwargs.get('max_iter', MAX_ITER)
self._options = kwargs
# + [markdown] slideshow={"slide_type": "fragment"}
# The initialization generates a control flow graph and hooks it to `fnenter` and `fnexit`.
# + slideshow={"slide_type": "fragment"}
symfz_ct = SimpleSymbolicFuzzer(check_triangle)
# + slideshow={"slide_type": "fragment"}
symfz_ct.fnenter, symfz_ct.fnexit
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Generating All Possible Paths
# We can use the procedure `get_all_paths()` starting from `fnenter` to recursively retrieve all paths in the function.
#
# The idea is as follows: Start with the function entry point `fnenter`, and recursively follow the children using the CFG. At any node there is a branching, there would be multiple children. On other nodes there would be only one child. Let us say a node had $n$ children. Such a node would result in $n$ paths. We attach the current node to the head of each paths, and return all paths thus generated.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def get_all_paths(self, fenter, depth=0):
if depth > self.max_depth:
raise Exception('Maximum depth exceeded')
if not fenter.children:
return [[(0, fenter)]]
fnpaths = []
for idx, child in enumerate(fenter.children):
child_paths = self.get_all_paths(child, depth + 1)
for path in child_paths:
# In a conditional branch, idx is 0 for IF, and 1 for Else
fnpaths.append([(idx, fenter)] + path)
return fnpaths
# + [markdown] slideshow={"slide_type": "fragment"}
# This can be used as follows.
# + slideshow={"slide_type": "subslide"}
symfz_ct = SimpleSymbolicFuzzer(check_triangle)
all_paths = symfz_ct.get_all_paths(symfz_ct.fnenter)
# + slideshow={"slide_type": "fragment"}
len(all_paths)
# + slideshow={"slide_type": "fragment"}
all_paths[1]
# + [markdown] slideshow={"slide_type": "fragment"}
# We hook `get_all_paths()` to initialization as below.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def process(self):
self.paths = self.get_all_paths(self.fnenter)
self.last_path = len(self.paths)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Extracting All Constraints
#
# For any given path, we define a function `extract_constraints()` to extract the constraints such that they are executable directly with *Z3*. The `idx` represents the particular branch that was taken. Hence, if the `False` branch was taken in a conditional, we attach a negation of the conditional.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def extract_constraints(self, path):
predicates = []
for (idx, elt) in path:
if isinstance(elt.ast_node, ast.AnnAssign):
if elt.ast_node.target.id in {'_if', '_while'}:
s = to_src(elt.ast_node.annotation)
predicates.append(("%s" if idx == 0 else "z3.Not(%s)") % s)
elif isinstance(elt.ast_node.annotation, ast.Call):
assert elt.ast_node.annotation.func.id == self.fn_name
else:
node = elt.ast_node
t = ast.Compare(node.target, [ast.Eq()], [node.value])
predicates.append(to_src(t))
elif isinstance(elt.ast_node, ast.Assign):
node = elt.ast_node
t = ast.Compare(node.targets[0], [ast.Eq()], [node.value])
predicates.append(to_src(t))
else:
pass
return predicates
# + slideshow={"slide_type": "subslide"}
symfz_ct = SimpleSymbolicFuzzer(check_triangle)
all_paths = symfz_ct.get_all_paths(symfz_ct.fnenter)
symfz_ct.extract_constraints(all_paths[0])
# + slideshow={"slide_type": "fragment"}
constraints = symfz_ct.extract_constraints(all_paths[1])
constraints
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fuzzing with Simple Symbolic Fuzzer
#
# To actually generate solutions, we define `fuzz()`. For that, we need to first extract all paths. Then choose a particular path, and extract the constraints in that path, which is then solved using *z3*.
# + slideshow={"slide_type": "skip"}
from contextlib import contextmanager
# + [markdown] slideshow={"slide_type": "fragment"}
# First we create a checkpoint for our current solver so that we can check a predicate, and rollback if necessary.
# + slideshow={"slide_type": "fragment"}
@contextmanager
def checkpoint(z3solver):
z3solver.push()
yield z3solver
z3solver.pop()
# + [markdown] slideshow={"slide_type": "subslide"}
# The `use_path()` function extracts constraints for a single function, applies it to our current solver (under a checkpoint), and returns the results if some solutions can be found.
# If solutions were found, we also make sure that we never reuse those solutions.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def solve_path_constraint(self, path):
# re-initializing does not seem problematic.
# a = z3.Int('a').get_id() remains the same.
constraints = self.extract_constraints(path)
decl = define_symbolic_vars(self.used_variables, '')
exec(decl)
solutions = {}
with checkpoint(self.z3):
st = 'self.z3.add(%s)' % ', '.join(constraints)
eval(st)
if self.z3.check() != z3.sat:
return {}
m = self.z3.model()
solutions = {d.name(): m[d] for d in m.decls()}
my_args = {k: solutions.get(k, None) for k in self.fn_args}
predicate = 'z3.And(%s)' % ','.join(
["%s == %s" % (k, v) for k, v in my_args.items()])
eval('self.z3.add(z3.Not(%s))' % predicate)
return my_args
# + [markdown] slideshow={"slide_type": "subslide"}
# We define `get_path()` that retrieves the current path and updates the path used.
# + slideshow={"slide_type": "fragment"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def get_next_path(self):
self.last_path -= 1
if self.last_path == -1:
self.last_path = len(self.paths) - 1
return self.paths[self.last_path]
# + [markdown] slideshow={"slide_type": "fragment"}
# The `fuzz()` method simply solves each path in order.
# + slideshow={"slide_type": "subslide"}
class SimpleSymbolicFuzzer(SimpleSymbolicFuzzer):
def fuzz(self):
"""Produce one solution for each path.
Returns a mapping of variable names to (symbolic) Z3 values."""
for i in range(self.max_tries):
res = self.solve_path_constraint(self.get_next_path())
if res:
return res
return {}
# + [markdown] slideshow={"slide_type": "fragment"}
# The fuzzer can be used as follows. Note that we need to convert the symbolic variables returned to Python numbers, using `as_long()`:
# + slideshow={"slide_type": "subslide"}
a, b, c = None, None, None
symfz_ct = SimpleSymbolicFuzzer(check_triangle)
for i in range(1, 10):
args = symfz_ct.fuzz()
res = check_triangle(args['a'].as_long(),
args['b'].as_long(),
args['c'].as_long())
print(args, "result:", res)
# + [markdown] slideshow={"slide_type": "subslide"}
# For symbolic fractions, we access their numerators and denominators:
# + slideshow={"slide_type": "subslide"}
symfz_av = SimpleSymbolicFuzzer(abs_value)
for i in range(1, 10):
args = symfz_av.fuzz()
abs_res = abs_value(args['x'].numerator_as_long() /
args['x'].denominator_as_long())
print(args, "result:", abs_res)
# + [markdown] slideshow={"slide_type": "subslide"}
# The _SimpleSymbolicFuzzer_ seems to work well for the _simple_ programs we checked above.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Problems with the Simple Fuzzer
#
# As we mentioned earlier, the `SimpleSymbolicFuzzer` cannot yet deal with variable reassignments. Further, it also fails to account for any loops. For example, consider the following program.
# + slideshow={"slide_type": "subslide"}
def gcd(a: int, b: int) -> int:
if a < b:
c: int = a # type: ignore
a = b
b = c
while b != 0:
c: int = a # type: ignore
a = b
b = c % b
return a
# + slideshow={"slide_type": "fragment"}
show_cfg(gcd)
# + slideshow={"slide_type": "skip"}
from ExpectError import ExpectError
# + slideshow={"slide_type": "subslide"}
with ExpectError():
symfz_gcd = SimpleSymbolicFuzzer(gcd, max_depth=1000, max_iter=10)
for i in range(1, 100):
r = symfz_gcd.fuzz()
v = gcd(r['a'].as_long(), r['b'].as_long())
print(r, v)
# + [markdown] slideshow={"slide_type": "subslide"}
# The problem here is that our *SimpleSymbolicFuzzer* has no concept of loops and variable reassignments. We will see how to fix this shortcoming next.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Advanced Symbolic Fuzzing
# + [markdown] slideshow={"slide_type": "fragment"}
# We next define `SymbolicFuzzer` that can deal with reassignments and *unrolling of loops*.
# + slideshow={"slide_type": "fragment"}
class SymbolicFuzzer(SimpleSymbolicFuzzer):
"""Symbolic fuzzing with reassignments and loop unrolling"""
def options(self, kwargs):
super().options(kwargs)
# + [markdown] slideshow={"slide_type": "fragment"}
# Once we allow reassignments and loop unrolling, we have to deal with what to call the new variables generated. This is what we will tackle next.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Dealing with Reassignments
#
# We want to rename all variables present in an expression such that the variables are annotated with their usage count. This makes it possible to determine variable reassignments. To do that, we define the `rename_variables()` function that, when given an `env` that contains the current usage index of different variables, renames the variables in the passed in AST node with the annotations, and returns a copy with the modifications. Note that we can't use [NodeTransformer](https://docs.python.org/3/library/ast.html#ast.NodeTransformer) here as it would modify the AST.
#
# That is, if the expression is `env[v] == 1`, `v` is renamed to `_v_1`
# + slideshow={"slide_type": "subslide"}
def rename_variables(astnode, env):
if isinstance(astnode, ast.BoolOp):
fn = 'z3.And' if isinstance(astnode.op, ast.And) else 'z3.Or'
return ast.Call(
ast.Name(fn, None),
[rename_variables(i, env) for i in astnode.values], [])
elif isinstance(astnode, ast.BinOp):
return ast.BinOp(
rename_variables(astnode.left, env), astnode.op,
rename_variables(astnode.right, env))
elif isinstance(astnode, ast.UnaryOp):
if isinstance(astnode.op, ast.Not):
return ast.Call(
ast.Name('z3.Not', None),
[rename_variables(astnode.operand, env)], [])
else:
return ast.UnaryOp(astnode.op,
rename_variables(astnode.operand, env))
elif isinstance(astnode, ast.Call):
return ast.Call(astnode.func,
[rename_variables(i, env) for i in astnode.args],
astnode.keywords)
elif isinstance(astnode, ast.Compare):
return ast.Compare(
rename_variables(astnode.left, env), astnode.ops,
[rename_variables(i, env) for i in astnode.comparators])
elif isinstance(astnode, ast.Name):
if astnode.id not in env:
env[astnode.id] = 0
num = env[astnode.id]
return ast.Name('_%s_%d' % (astnode.id, num), astnode.ctx)
elif isinstance(astnode, ast.Return):
return ast.Return(rename_variables(astnode.value, env))
else:
return astnode
# + [markdown] slideshow={"slide_type": "subslide"}
# To verify that it works as intended, we start with an environment.
# + slideshow={"slide_type": "fragment"}
env = {'x': 1}
# + slideshow={"slide_type": "fragment"}
ba = get_expression('x == 1 and y == 2')
type(ba)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(ba, env)) == 'z3.And(_x_1 == 1, _y_0 == 2)'
# + slideshow={"slide_type": "fragment"}
bo = get_expression('x == 1 or y == 2')
type(bo.op)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(bo, env)) == 'z3.Or(_x_1 == 1, _y_0 == 2)'
# + slideshow={"slide_type": "fragment"}
b = get_expression('x + y')
type(b)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(b, env)) == '_x_1 + _y_0'
# + slideshow={"slide_type": "subslide"}
u = get_expression('-y')
type(u)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(u, env)) == '-_y_0'
# + slideshow={"slide_type": "fragment"}
un = get_expression('not y')
type(un.op)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(un, env)) == 'z3.Not(_y_0)'
# + slideshow={"slide_type": "fragment"}
c = get_expression('x == y')
type(c)
# + slideshow={"slide_type": "fragment"}
assert to_src(rename_variables(c, env)) == '_x_1 == _y_0'
# + slideshow={"slide_type": "fragment"}
f = get_expression('fn(x,y)')
type(f)
# + slideshow={"slide_type": "subslide"}
assert to_src(rename_variables(f, env)) == 'fn(_x_1, _y_0)'
# + slideshow={"slide_type": "fragment"}
env
# + [markdown] slideshow={"slide_type": "fragment"}
# Next, we want to process the CFG, and correctly transform the paths.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Tracking Assignments
#
# For keeping track of assignments in the CFG, We define a data structure `PNode` that stores the current CFG node.
# + slideshow={"slide_type": "fragment"}
class PNode:
def __init__(self, idx, cfgnode, parent=None, order=0, seen=None):
self.seen = {} if seen is None else seen
self.max_iter = MAX_ITER
self.idx, self.cfgnode, self.parent, self.order = idx, cfgnode, parent, order
def __repr__(self):
return "PNode:%d[%s order:%d]" % (self.idx, str(self.cfgnode),
self.order)
# + [markdown] slideshow={"slide_type": "fragment"}
# Defining a new `PNode` is done as follows.
# + slideshow={"slide_type": "subslide"}
cfg = PyCFG()
cfg.gen_cfg(inspect.getsource(gcd))
gcd_fnenter, _ = cfg.functions['gcd']
# + slideshow={"slide_type": "fragment"}
PNode(0, gcd_fnenter)
# + [markdown] slideshow={"slide_type": "fragment"}
# The `copy()` method generates a copy for the child's keep, indicating which path was taken (with `order` of the child).
# + slideshow={"slide_type": "fragment"}
class PNode(PNode):
def copy(self, order):
p = PNode(self.idx, self.cfgnode, self.parent, order, self.seen)
assert p.order == order
return p
# + [markdown] slideshow={"slide_type": "fragment"}
# Using the copy operation.
# + slideshow={"slide_type": "fragment"}
PNode(0, gcd_fnenter).copy(1)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Stepwise Exploration of Paths
#
# A problem we had with our `SimpleSymbolicFuzzer` is that it explored a path to completion before attempting another. However, this is non-optimal. One may want to explore the graph in a more step-wise manner, expanding every possible execution one step at a time.
#
# Hence, we define `explore()` which explores the children of a node if any, one step at a time. If done exhaustively, this will generate all paths from a starting node until no more children are left. We made `PNode` to a container class so that this iteration can be driven from outside, and stopped if say a maximum iteration is complete, or certain paths need to be prioritized.
# + slideshow={"slide_type": "subslide"}
class PNode(PNode):
def explore(self):
ret = []
for (i, n) in enumerate(self.cfgnode.children):
key = "[%d]%s" % (self.<KEY>
ccount = self.seen.get(key, 0)
if ccount > self.max_iter:
continue # drop this child
self.seen[key] = ccount + 1
pn = PNode(self.idx + 1, n, self.copy(i), seen=self.seen)
ret.append(pn)
return ret
# + [markdown] slideshow={"slide_type": "fragment"}
# We can use `explore()` as follows.
# + slideshow={"slide_type": "fragment"}
PNode(0, gcd_fnenter).explore()
# + slideshow={"slide_type": "subslide"}
PNode(0, gcd_fnenter).explore()[0].explore()
# + [markdown] slideshow={"slide_type": "fragment"}
# The method `get_path_to_root()` recursively goes up through child->parent chain retrieving the complete chain to the topmost parent.
# + code_folding=[] slideshow={"slide_type": "fragment"}
class PNode(PNode):
def get_path_to_root(self):
path = []
n = self
while n:
path.append(n)
n = n.parent
return list(reversed(path))
# + slideshow={"slide_type": "subslide"}
p = PNode(0, gcd_fnenter)
[s.get_path_to_root() for s in p.explore()[0].explore()[0].explore()[0].explore()]
# + [markdown] slideshow={"slide_type": "fragment"}
# The string representation of the node is in `z3` solvable form.
# + slideshow={"slide_type": "fragment"}
class PNode(PNode):
def __str__(self):
path = self.get_path_to_root()
ssa_path = to_single_assignment_predicates(path)
return ', '.join([to_src(p) for p in ssa_path])
# + [markdown] slideshow={"slide_type": "fragment"}
# However, before using it, we need to take care of variable renaming so that reassignments can work.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Renaming Used Variables
#
# We need to rename used variables. Any variable `v = xxx` should be renamed to `_v_0` and any later assignment such as `v = v + 1` should be transformed to `_v_1 = _v_0 + 1` and later conditionals such as `v == x` should be transformed to `(_v_1 == _x_0)`. The method `to_single_assignment_predicates()` does this for a given path.
# + slideshow={"slide_type": "subslide"}
def to_single_assignment_predicates(path):
env = {}
new_path = []
for i, node in enumerate(path):
ast_node = node.cfgnode.ast_node
new_node = None
if isinstance(ast_node, ast.AnnAssign) and ast_node.target.id in {
'exit'}:
new_node = None
elif isinstance(ast_node, ast.AnnAssign) and ast_node.target.id in {'enter'}:
args = [
ast.parse(
"%s == _%s_0" %
(a.id, a.id)).body[0].value for a in ast_node.annotation.args]
new_node = ast.Call(ast.Name('z3.And', None), args, [])
elif isinstance(ast_node, ast.AnnAssign) and ast_node.target.id in {'_if', '_while'}:
new_node = rename_variables(ast_node.annotation, env)
if node.order != 0:
assert node.order == 1
new_node = ast.Call(ast.Name('z3.Not', None), [new_node], [])
elif isinstance(ast_node, ast.AnnAssign):
assigned = ast_node.target.id
val = [rename_variables(ast_node.value, env)]
env[assigned] = 0 if assigned not in env else env[assigned] + 1
target = ast.Name('_%s_%d' %
(ast_node.target.id, env[assigned]), None)
new_node = ast.Expr(ast.Compare(target, [ast.Eq()], val))
elif isinstance(ast_node, ast.Assign):
assigned = ast_node.targets[0].id
val = [rename_variables(ast_node.value, env)]
env[assigned] = 0 if assigned not in env else env[assigned] + 1
target = ast.Name('_%s_%d' %
(ast_node.targets[0].id, env[assigned]), None)
new_node = ast.Expr(ast.Compare(target, [ast.Eq()], val))
elif isinstance(ast_node, (ast.Return, ast.Pass)):
new_node = None
else:
s = "NI %s %s" % (type(ast_node), ast_node.target.id)
raise Exception(s)
new_path.append(new_node)
return new_path
# + [markdown] slideshow={"slide_type": "subslide"}
# Here is how it can be used:
# + slideshow={"slide_type": "fragment"}
p = PNode(0, gcd_fnenter)
path = p.explore()[0].explore()[0].explore()[0].get_path_to_root()
spath = to_single_assignment_predicates(path)
# + slideshow={"slide_type": "fragment"}
[to_src(s) for s in spath]
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Check Before You Loop
#
# One of the ways in which the *concolic* execution simplifies *symbolic* execution is in the treatment of loops. Rather than trying to determine an invariant for a loop, we simply _unroll_ the loops a number of times until we hit the `MAX_DEPTH` limit.
# However, not all loops will need to be unrolled until `MAX_DEPTH` is reached. Some of them may exit before. Hence, it is necessary to check whether the given set of constraints can be satisfied before continuing to explore further.
# + slideshow={"slide_type": "subslide"}
def identifiers_with_types(identifiers, defined):
with_types = dict(defined)
for i in identifiers:
if i[0] == '_':
nxt = i[1:].find('_', 1)
name = i[1:nxt + 1]
assert name in defined
typ = defined[name]
with_types[i] = typ
return with_types
# + [markdown] slideshow={"slide_type": "fragment"}
# The `extract_constraints()` generates the `z3` constraints from a path. The main work is done by `to_single_assignment_predicates()`. The `extract_constraints()` then converts the AST to source.
# + slideshow={"slide_type": "subslide"}
class SymbolicFuzzer(SymbolicFuzzer):
def extract_constraints(self, path):
return [to_src(p) for p in to_single_assignment_predicates(path) if p]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Solving Path Constraints
#
# We now update our `solve_path_constraint()` method to take into account the new identifiers created during reassignments.
# + slideshow={"slide_type": "subslide"}
class SymbolicFuzzer(SymbolicFuzzer):
def solve_path_constraint(self, path):
# re-initializing does not seem problematic.
# a = z3.Int('a').get_id() remains the same.
constraints = self.extract_constraints(path)
identifiers = [
c for i in constraints for c in used_identifiers(i)] # <- changes
with_types = identifiers_with_types(
identifiers, self.used_variables) # <- changes
decl = define_symbolic_vars(with_types, '')
exec(decl)
solutions = {}
with checkpoint(self.z3):
st = 'self.z3.add(%s)' % ', '.join(constraints)
eval(st)
if self.z3.check() != z3.sat:
return {}
m = self.z3.model()
solutions = {d.name(): m[d] for d in m.decls()}
my_args = {k: solutions.get(k, None) for k in self.fn_args}
predicate = 'z3.And(%s)' % ','.join(
["%s == %s" % (k, v) for k, v in my_args.items()])
eval('self.z3.add(z3.Not(%s))' % predicate)
return my_args
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Generating All Paths
# + [markdown] slideshow={"slide_type": "fragment"}
# The `get_all_paths()` is now similarly updated so that we unroll loops only to a specified height. It is also converted to an iterative exploration style so that we explore the CFG in a breadth first manner.
# + slideshow={"slide_type": "subslide"}
class SymbolicFuzzer(SymbolicFuzzer):
def get_all_paths(self, fenter):
path_lst = [PNode(0, fenter)]
completed = []
for i in range(self.max_iter):
new_paths = [PNode(0, fenter)]
for path in path_lst:
# explore each path once
if path.cfgnode.children:
np = path.explore()
for p in np:
if path.idx > self.max_depth:
break
new_paths.append(p)
else:
completed.append(path)
path_lst = new_paths
return completed + path_lst
# + [markdown] slideshow={"slide_type": "subslide"}
# We can now obtain all paths using our advanced symbolic fuzzer as follows.
# + slideshow={"slide_type": "fragment"}
asymfz_gcd = SymbolicFuzzer(
gcd, max_iter=10, max_tries=10, max_depth=10)
all_paths = asymfz_gcd.get_all_paths(asymfz_gcd.fnenter)
# + slideshow={"slide_type": "fragment"}
len(all_paths)
# + slideshow={"slide_type": "subslide"}
all_paths[37].get_path_to_root()
# + [markdown] slideshow={"slide_type": "fragment"}
# We can also list the predicates in each path.
# + slideshow={"slide_type": "subslide"}
for s in to_single_assignment_predicates(all_paths[37].get_path_to_root()):
if s is not None:
print(to_src(s))
# + slideshow={"slide_type": "subslide"}
constraints = asymfz_gcd.extract_constraints(all_paths[37].get_path_to_root())
# + slideshow={"slide_type": "fragment"}
constraints
# + [markdown] slideshow={"slide_type": "subslide"}
# The constraints printed out demonstrates that our approach for renaming variables was successful. We need only one more piece to complete the puzzle. Our path is still a `PNode`. We need to modify `get_next_path()` so that we return the corresponding predicate chain.
# + slideshow={"slide_type": "fragment"}
class SymbolicFuzzer(SymbolicFuzzer):
def get_next_path(self):
self.last_path -= 1
if self.last_path == -1:
self.last_path = len(self.paths) - 1
return self.paths[self.last_path].get_path_to_root()
# + [markdown] slideshow={"slide_type": "fragment"}
# We will see next how to use our fuzzer for fuzzing.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fuzzing with Advanced Symbolic Fuzzer
#
# We use our advanced symbolic fuzzer on *gcd* to generate plausible inputs.
# + slideshow={"slide_type": "subslide"}
asymfz_gcd = SymbolicFuzzer(
gcd, max_tries=10, max_iter=10, max_depth=10)
data = []
for i in range(10):
r = asymfz_gcd.fuzz()
data.append((r['a'].as_long(), r['b'].as_long()))
v = gcd(*data[-1])
print(r, "result:", repr(v))
# + [markdown] slideshow={"slide_type": "subslide"}
# The outputs look reasonable. However, what is the coverage obtained?
# + slideshow={"slide_type": "fragment"}
with VisualizedArcCoverage() as cov:
for a, b in data:
gcd(a, b)
# + slideshow={"slide_type": "subslide"}
cov.show_coverage(gcd)
# + slideshow={"slide_type": "subslide"}
show_cfg(gcd, arcs=cov.arcs())
# + [markdown] slideshow={"slide_type": "fragment"}
# Indeed both branch and statement coverage visualization seems to indicate that we achieved complete coverage.
# How do we make use of our fuzzer in practice? We explore a small case study of a program to solve the roots of a quadratic equation.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Example: Roots of a Quadratic Equation
# Here is the famous equation for finding the roots of a quadratic equation.
# + slideshow={"slide_type": "skip"}
from typing import Tuple
# + slideshow={"slide_type": "subslide"}
def roots(a: float, b: float, c: float) -> Tuple[float, float]:
d: float = b * b - 4 * a * c
ax: float = 0.5 * d
bx: float = 0
while (ax - bx) > 0.1:
bx = 0.5 * (ax + d / ax)
ax = bx
s: float = bx
a2: float = 2 * a
ba2: float = b / a2
return -ba2 + s / a2, -ba2 - s / a2
# + [markdown] slideshow={"slide_type": "subslide"}
# Does the program look correct? Let us investigate if the program is reasonable. But before that, we need a helper
# function `sym_to_float()` to convert symbolic values to floating point.
# + slideshow={"slide_type": "fragment"}
def sym_to_float(v):
if v is None:
return math.inf
elif isinstance(v, z3.IntNumRef):
return v.as_long()
return v.numerator_as_long() / v.denominator_as_long()
# + [markdown] slideshow={"slide_type": "fragment"}
# Now we are ready to fuzz.
# + slideshow={"slide_type": "subslide"}
asymfz_roots = SymbolicFuzzer(
roots,
max_tries=10,
max_iter=10,
max_depth=10)
# + slideshow={"slide_type": "subslide"}
with ExpectError():
for i in range(100):
r = asymfz_roots.fuzz()
print(r)
d = [sym_to_float(r[i]) for i in ['a', 'b', 'c']]
v = roots(*d)
print(d, v)
# + [markdown] slideshow={"slide_type": "subslide"}
# We have a `ZeroDivisionError`. Can we eliminate it?
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Roots - Check Before Divide
# + slideshow={"slide_type": "subslide"}
def roots2(a: float, b: float, c: float) -> Tuple[float, float]:
d: float = b * b - 4 * a * c
xa: float = 0.5 * d
xb: float = 0
while (xa - xb) > 0.1:
xb = 0.5 * (xa + d / xa)
xa = xb
s: float = xb
if a == 0:
return -c / b, -c / b # only one solution
a2: float = 2 * a
ba2: float = b / a2
return -ba2 + s / a2, -ba2 - s / a2
# + slideshow={"slide_type": "subslide"}
asymfz_roots = SymbolicFuzzer(
roots2,
max_tries=10,
max_iter=10,
max_depth=10)
# + slideshow={"slide_type": "subslide"}
with ExpectError():
for i in range(1000):
r = asymfz_roots.fuzz()
d = [sym_to_float(r[i]) for i in ['a', 'b', 'c']]
v = roots2(*d)
#print(d, v)
# + [markdown] slideshow={"slide_type": "fragment"}
# Apparently, our fix was incomplete. Let us try again.
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Roots - Eliminating the Zero Division Error
# + slideshow={"slide_type": "skip"}
import math
# + slideshow={"slide_type": "subslide"}
def roots3(a: float, b: float, c: float) -> Tuple[float, float]:
d: float = b * b - 4 * a * c
xa: float = 0.5 * d
xb: float = 0
while (xa - xb) > 0.1:
xb = 0.5 * (xa + d / xa)
xa = xb
s: float = xb
if a == 0:
if b == 0:
return math.inf, math.inf
return -c / b, -c / b # only one solution
a2: float = 2 * a
ba2: float = b / a2
return -ba2 + s / a2, -ba2 - s / a2
# + slideshow={"slide_type": "subslide"}
asymfz_roots = SymbolicFuzzer(
roots3,
max_tries=10,
max_iter=10,
max_depth=10)
# + slideshow={"slide_type": "subslide"}
for i in range(10):
r = asymfz_roots.fuzz()
print(r)
d = [sym_to_float(r[i]) for i in ['a', 'b', 'c']]
v = roots3(*d)
print(d, v)
# + [markdown] slideshow={"slide_type": "subslide"}
# With this, we have demonstrated that we can use our *SymbolicFuzzer* to fuzz programs, and it can aid in identifying problems in code.
# + [markdown] slideshow={"slide_type": "slide"} tags=[]
# ## Limitations
#
# There is an evident error in the `roots3()` function. We are not checking for negative roots. However, the symbolic execution does not seem to have detected it. Why are we not able to detect the problem of negative roots? Because we stop execution at a predetermined depth without throwing an error. That is, our symbolic execution is wide but shallow. One of the ways this limitation can be overcome is by relying on [concolic execution](ConcolicFuzzer.ipynb), that allows one to go deeper than pure symbolic execution.
# + [markdown] jp-MarkdownHeadingCollapsed=true slideshow={"slide_type": "fragment"} tags=[]
# A second problem is that symbolic execution is necessarily computation intensive. This means that specification based fuzzers are often able to generate a much larger set of inputs, and consecutively more coverage on programs that do not check for magic bytes, such that they provide a reasonable gradient of exploration.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Synopsis
# + [markdown] slideshow={"slide_type": "fragment"}
# This chapter provides an implementation of a symbolic fuzzing engine `SymbolicFuzzer`. The fuzzer uses symbolic execution to exhaustively explore paths in the program to a limited depth, and generate inputs that will reach these paths.
# + [markdown] slideshow={"slide_type": "fragment"}
# As an example, consider the function `gcd()`, computing the greatest common divisor of `a` and `b`:
# + slideshow={"slide_type": "fragment"}
# ignore
from bookutils import print_content
# + slideshow={"slide_type": "subslide"}
# ignore
print_content(inspect.getsource(gcd), '.py')
# + [markdown] slideshow={"slide_type": "subslide"}
# To explore `gcd()`, the fuzzer can be used as follows, producing values for arguments that cover different paths in `gcd()` (including multiple times of loop iterations):
# + slideshow={"slide_type": "subslide"}
gcd_fuzzer = SymbolicFuzzer(gcd, max_tries=10, max_iter=10, max_depth=10)
for i in range(10):
args = gcd_fuzzer.fuzz()
print(args)
# + [markdown] slideshow={"slide_type": "subslide"}
# Note that the variable values returned by `fuzz()` are Z3 _symbolic_ values; to convert them to Python numbers, use their method `as_long()`:
# + slideshow={"slide_type": "subslide"}
for i in range(10):
args = gcd_fuzzer.fuzz()
a = args['a'].as_long()
b = args['b'].as_long()
d = gcd(a, b)
print(f"gcd({a}, {b}) = {d}")
# + [markdown] slideshow={"slide_type": "subslide"}
# The symbolic fuzzer is subject to a number of constraints. First, it requires that the function to be fuzzed has correct type annotations, including all local variables. Second, it solves loops by unrolling them, but only for a fixed amount.
# + [markdown] slideshow={"slide_type": "fragment"}
# For programs without loops and variable reassignments, the `SimpleSymbolicFuzzer` is a faster, but more limited alternative.
# + slideshow={"slide_type": "fragment"}
# ignore
from ClassDiagram import display_class_hierarchy
display_class_hierarchy(SymbolicFuzzer)
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * One can use symbolic execution to augment the inputs that explore all characteristics of a program.
# * Symbolic execution can be broad but shallow.
# * Symbolic execution is well suited for programs that rely on specific values to be present in the input, however, its utility decreases when such values are not present, and the input space represents a gradient in terms of coverage.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# * [Search based fuzzing](SearchBasedFuzzer.ipynb) can often be an acceptable middle ground when random fuzzing does not provide sufficient results, but symbolic fuzzing is too heavyweight.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# Symbolic execution of programs was originally described by King \cite{king1976symbolic} in 1976. It is used extensively in vulnerability analysis of software, especially binary programs. Some of the well known symbolic execution tools include *KLEE* \cite{KLEE}, *angr* \cite{wang2017angr}, *Driller* \cite{stephens2016driller}, and *SAGE* \cite{godefroid2012sage}. The best known symbolic execution environment for Python is CHEF \cite{bucur2014prototyping} which does symbolic execution by modifying the interpreter.
#
# The Z3 solver we use in this chapter was developed at Microsoft Research under the lead of <NAME> and <NAME> \cite{z3}. It is one of the most popular solvers.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 1: Extending Symbolic Fuzzer to use function summaries
#
# We showed in the first section how function summaries may be produced. Can you extend the `SymbolicFuzzer` to use function summaries when needed?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** _None yet available._
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise 2: Statically checking if a loop should be unrolled further
#
# We examined how loops would be unrolled during exploration to a fixed depth. However, not all loops need to be unrolled completely. Some of the loops may contain only a constant number of iterations. For example, consider the loop below.
# + slideshow={"slide_type": "fragment"}
i = 0
while i < 10:
i += 1
# + [markdown] slideshow={"slide_type": "fragment"}
# This loop needs to be unrolled exactly $10$ times. For such cases, can you implement a method `can_be_satisfied()` which is invoked as below, to only unroll further if the path condition can be satisfied.
# + slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
class SymbolicFuzzer(SymbolicFuzzer):
def get_all_paths(self, fenter):
path_lst = [PNode(0, fenter)]
completed = []
for i in range(self.max_iter):
new_paths = [PNode(0, fenter)]
for path in path_lst:
# explore each path once
if path.cfgnode.children:
np = path.explore()
for p in np:
if path.idx > self.max_depth:
break
if self.can_be_satisfied(p):
new_paths.append(p)
else:
break
else:
completed.append(path)
path_lst = new_paths
return completed + path_lst
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# **Solution.** Here is a solution.
# + slideshow={"slide_type": "skip"} solution2="hidden"
class SymbolicFuzzer(SymbolicFuzzer):
def can_be_satisfied(self, p):
s2 = self.extract_constraints(p.get_path_to_root())
s = z3.Solver()
identifiers = [c for i in s2 for c in used_identifiers(i)]
with_types = identifiers_with_types(identifiers, self.used_variables)
decl = define_symbolic_vars(with_types, '')
exec(decl)
exec("s.add(z3.And(%s))" % ','.join(s2), globals(), locals())
return s.check() == z3.sat
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# With this implementation, new conditions are appended to paths if and only if the paths are still satisfiable after incorporating the condition.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercise 3: Implementing a Concolic Fuzzer
#
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
# We have seen in the chapter on [concolic fuzzing](ConcolicFuzzer.ipynb) how to trace a function concolically using information flow. However, this is somewhat sub-optimal as the constraints can get dropped when the information flow is indirect (as in control flow based information flow). Can you implement concolic tracing using the infrastructure we built for symbolic execution?
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# **Solution.** Here is a possible solution.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# In *concolic execution*, we rely on a seed input to guide our symbolic execution. We collect the line numbers that our seed input traces, and feed it to the symbolic execution such that in the `explore` step, only the child node that correspond to the seed input execution path is chosen. This allows us to collect the complete set of constraints along a *representative path*. Once we have it, we can choose any particular predicate and invert it to explore the program execution paths near the representative path.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We modify our original `ArcCoverage` to provide *all* line numbers that the program traversed.
# + slideshow={"slide_type": "skip"} solution2="hidden"
class TrackingArcCoverage(ArcCoverage):
def offsets_from_entry(self, fn):
zero = self._trace[0][1] - 1
return [l - zero for (f, l) in self._trace if f == fn]
# + slideshow={"slide_type": "skip"} solution2="hidden"
with TrackingArcCoverage() as cov:
roots3(1, 1, 1)
# + slideshow={"slide_type": "skip"} solution2="hidden"
cov.offsets_from_entry('roots3')
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# The `ConcolicTracer` first extracts the program trace on a seed input.
# + slideshow={"slide_type": "skip"} solution2="hidden"
class ConcolicTracer(SymbolicFuzzer):
def __init__(self, fn, fnargs, **kwargs):
with TrackingArcCoverage() as cov:
fn(*fnargs)
self.lines = cov.offsets_from_entry(fn.__name__)
self.current_line = 0
super().__init__(fn, **kwargs)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# The method `get_all_paths()` now tries to follow the seed execution path.
# + slideshow={"slide_type": "skip"} solution2="hidden"
class ConcolicTracer(ConcolicTracer):
def get_all_paths(self, fenter):
assert fenter.ast_node.lineno == self.lines[self.current_line]
self.current_line += 1
last_node = PNode(0, fenter)
while last_node and self.current_line < len(self.lines):
if last_node.cfgnode.children:
np = last_node.explore()
for p in np:
if self.lines[self.current_line] == p.cfgnode.ast_node.lineno:
self.current_line += 1
last_node = p
break
else:
last_node = None
break
else:
break
assert len(self.lines) == self.current_line
return [last_node]
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We are now ready to concolicaly trace our execution.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# #### Tracing the Execution Concolicaly
# + slideshow={"slide_type": "fragment"} solution2="hidden"
acfz_roots = ConcolicTracer(
roots3,
fnargs=[1, 1, 1],
max_tries=10,
max_iter=10,
max_depth=10)
# + slideshow={"slide_type": "subslide"} solution2="hidden"
acfz_roots.paths[0].get_path_to_root()
# + slideshow={"slide_type": "subslide"} solution2="hidden"
print(cov.offsets_from_entry('roots3'))
print([i.cfgnode.ast_node.lineno for i in acfz_roots.paths[0].get_path_to_root()])
print(acfz_roots.lines)
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# As can be seen above, we recovered the trace information correctly.
# Next, we extract the constraints as usual.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
constraints = acfz_roots.extract_constraints(
acfz_roots.paths[0].get_path_to_root())
# + slideshow={"slide_type": "subslide"} solution2="hidden"
constraints
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# Next, we change our constraints to symbolic variables and solve them.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
identifiers = [c for i in constraints for c in used_identifiers(i)]
with_types = identifiers_with_types(identifiers, acfz_roots.used_variables)
decl = define_symbolic_vars(with_types, '')
exec(decl)
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# We are ready solve our constraints. However, before that, here is a question for you.
# *Should it result in exactly the same arguments?*
# + slideshow={"slide_type": "subslide"} solution2="hidden"
eval('z3.solve(%s)' % ','.join(constraints))
# + slideshow={"slide_type": "subslide"} solution2="hidden"
acfz_roots.fuzz()
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden"
# Did they take the same path?
# + slideshow={"slide_type": "fragment"} solution2="hidden"
with ArcCoverage() as cov:
roots(1, 1, 1)
show_cfg(roots, arcs=cov.arcs())
# + slideshow={"slide_type": "fragment"} solution2="hidden"
with ArcCoverage() as cov:
roots(-1, 0, 0)
show_cfg(roots, arcs=cov.arcs())
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# Indeed, even though the arguments were different, the path traced is exactly the same.
#
# As we saw in the chapter on [concolic fuzzing](ConcolicFuzzer.ipynb), concolic tracing has another use, namely that we can use it to explore nearby paths. We will see how to do that next.
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# #### Exploring Nearby Paths
#
# We collected the following constraints.
# + slideshow={"slide_type": "fragment"} solution2="hidden"
constraints
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden"
# We can explore nearby paths by negating the conditionals starting from the very last. (A question for the student: Why do we want to start negating from the very last?)
# + slideshow={"slide_type": "fragment"} solution2="hidden"
new_constraints = constraints[0:4] + ['z3.Not(%s)' % constraints[4]]
# + slideshow={"slide_type": "fragment"} solution2="hidden"
new_constraints
# + slideshow={"slide_type": "subslide"} solution2="hidden"
eval('z3.solve(%s)' % ','.join(new_constraints))
# + slideshow={"slide_type": "fragment"} solution2="hidden"
with ArcCoverage() as cov:
roots3(1, 0, -11 / 20)
show_cfg(roots3, arcs=cov.arcs())
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# Indeed, the path traced is now different. One can continue this procedure to the necessary number of times to explore all nearby paths to the execution.
#
# Can you incorporate this exploration into the concolic fuzzer?
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** _None yet available._
# + [markdown] slideshow={"slide_type": "slide"}
# ## Compatibility
# + [markdown] slideshow={"slide_type": "fragment"}
# Earlier versions of this chapter used the name `AdvancedSymbolicFuzzer` for `SymbolicFuzzer`.
# + slideshow={"slide_type": "fragment"}
AdvancedSymbolicFuzzer = SymbolicFuzzer
| docs/notebooks/SymbolicFuzzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from statistics import stdev
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# -
# datasets = ["cancer", "card", "diabetes", "gene", "glass", "heart", "horse", "mushroom", "soybean", "thyroid"]
datasets = ["cancer", "card", "gene", "glass", "heart", "horse", "mushroom", "soybean", "thyroid"]
multi = []
one = []
for dataset in datasets:
df = pd.read_csv(f"../../log/prelim_out_rep/out_rep_multi_{dataset}.txt")
df["dataset"] = dataset
multi.append(df)
df = pd.read_csv(f"../../log/prelim_out_rep/out_rep_one_{dataset}.txt")
df["dataset"] = dataset
one.append(df)
multi = pd.concat(multi)
one = pd.concat(one)
multimeans = pd.pivot_table(multi, index="dataset")
multistds = pd.pivot_table(multi, index="dataset", aggfunc=stdev)
onemeans = pd.pivot_table(one, index="dataset")
onestds = pd.pivot_table(one, index="dataset", aggfunc=stdev)
nclass = [datasets[i] + "\n(" + str(multimeans.nout[i]) + ")" for i in range(len(datasets))]
spc = np.arange(len(datasets))
w = 0.45
errstyle = dict(elinewidth=1, capsize=5)
errstyletr = dict(elinewidth=1, capsize=5, alpha=0.4)
fig = plt.figure(figsize=(14, 9))
plt.rcParams.update({"font.size": 13})
plt.bar(spc, multimeans.ftest, width=w, label="One-hot (test)", edgecolor="k", yerr=multistds.ftest, error_kw=errstyle)
plt.bar(spc, multimeans.ftrain, width=w, label="One-hot (train)", edgecolor="k", yerr=multistds.ftrain, alpha=0.1, error_kw=errstyletr)
plt.bar(spc + w, onemeans.ftest, width=w, label="Label (test)", edgecolor="k", yerr=onestds.ftest, error_kw=errstyle)
plt.bar(spc + w, onemeans.ftrain, width=w, label="Label (train)", edgecolor="k", yerr=onestds.ftrain, alpha=0.1, error_kw=errstyletr)
plt.legend(loc="lower left")
plt.xticks(spc + w / 2, nclass)
plt.xlabel("Dataset (#classes)", fontsize=18)
plt.ylabel("$F_1$-score (mean $\pm$ stdev)", fontsize=18)
plt.title("One-hot vs label output encoding", fontsize=24)
# fig.savefig("output_representation_f1.png")
| dataproc/prelim/output_representation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Writing Reusable Code with Classes and Functions
#
# ### <NAME>
#
# ### PHY321: Classical Mechanics
#
# ### February 16, 2022
# * Many pieces of code will be reused throughout the course of this class...copy and paste is not sustainable
# * Better practice: write one good piece of code in a way that can be used everywhere
# * Functions and Classes
# ## 1. Start with a code you have written
#
# * Euler's Method for an object in freefall with drag (Week 3 Notes)
# +
# Common imports
import numpy as np
import pandas as pd
from math import *
import matplotlib.pyplot as plt
import os
g = 9.80655 #m/s^2
D = 0.00245 #m/s
DeltaT = 0.1
#set up arrays
tfinal = 0.5
n = ceil(tfinal/DeltaT)
# define scaling constant vT
vT = sqrt(g/D)
# set up arrays for t, a, v, and y and we can compare our results with analytical ones
t = np.zeros(n)
a = np.zeros(n)
v = np.zeros(n)
y = np.zeros(n)
yanalytic = np.zeros(n)
# Initial conditions
v[0] = 0.0 #m/s
y[0] = 10.0 #m
yanalytic[0] = y[0]
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
data = {'t[s]': t,
'y[m]': y-yanalytic,
'v[m/s]': v,
'a[m/s^2]': a
}
NewData = pd.DataFrame(data)
display(NewData)
#finally we plot the data
fig, axs = plt.subplots(3, 1)
axs[0].plot(t, y, t, yanalytic)
axs[0].set_xlim(0, tfinal)
axs[0].set_ylabel('y and exact')
axs[1].plot(t, v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(t, a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
plt.show()
# -
# ## 2. Identify Pieces of the Code that You Reuse (Or Could Reuse)
# +
g = 9.80655 #m/s^2
D = 0.00245 #m/s
# define scaling constant vT
vT = sqrt(g/D)
###############################################
# SETTING UP TIME
DeltaT = 0.1
#set up arrays
tfinal = 0.5
n = ceil(tfinal/DeltaT)
t = np.zeros(n)
###############################################
###############################################
# SETTING UP THE INITIAL ARRAYS
# set up arrays for a, v, and y and we can compare our results with analytical ones
a = np.zeros(n)
v = np.zeros(n)
y = np.zeros(n)
yanalytic = np.zeros(n)
# Initial conditions
v[0] = 0.0 #m/s
y[0] = 10.0 #m
yanalytic[0] = y[0]
###############################################
###############################################
# EULER'S METHOD
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
###############################################
###############################################
# DISPSLAY THE DATA
data = {'t[s]': t,
'y[m]': y-yanalytic,
'v[m/s]': v,
'a[m/s^2]': a
}
NewData = pd.DataFrame(data)
display(NewData)
###############################################
###############################################
# GRAPH THE DATA
#finally we plot the data
fig, axs = plt.subplots(3, 1)
axs[0].plot(t, y, t, yanalytic)
axs[0].set_xlim(0, tfinal)
axs[0].set_ylabel('y and exact')
axs[1].plot(t, v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(t, a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
plt.show()
###############################################
# -
# ## 3. For Each Piece of Code You Identified, Try to Write a Function
# * Think about what the arguments fo the function need to be and what needs to be returned
###############################################
# EULER'S METHOD
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
###############################################
# * Arguments: n, DeltaT, t, a, v, y, yanalytic, g, D, vT
# * Returns: t, a, v, y, yanalytic
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, vT):
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
return t, a, v, y, yanalytic
###############################################
# +
###############################################
# SETTING UP TIME
def set_up_time (DeltaT, tfinal):
n = ceil(tfinal/DeltaT)
t = np.zeros(n)
return n, t
###############################################
###############################################
# SETTING UP THE INITIAL ARRAYS
def set_up_initial_arrays (n, v_0, y_0):
# set up arrays for t, a, v, and y and we can compare our results with analytical ones
a = np.zeros(n)
v = np.zeros(n)
y = np.zeros(n)
yanalytic = np.zeros(n)
# Initial conditions
v[0] = v_0 #m/s
y[0] = y_0 #m
yanalytic[0] = y[0]
return a, v, y, yanalytic
###############################################
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, vT):
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i]
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT))
if ( y[i+1] < 0.0):
break
a[n-1] = -g + D*v[n-1]*v[n-1]
return t, a, v, y, yanalytic
###############################################
###############################################
# DISPSLAY THE DATA
def display_data (t, a, v, y, yanalytic):
data = {'t[s]': t,
'y[m]': y-yanalytic,
'v[m/s]': v,
'a[m/s^2]': a
}
NewData = pd.DataFrame(data)
display(NewData)
###############################################
###############################################
# GRAPH THE DATA
#finally we plot the data
def graph_data (t, a, v, y, yanalytic):
fig, axs = plt.subplots(3, 1)
axs[0].plot(t, y, t, yanalytic)
axs[0].set_xlim(0, tfinal)
axs[0].set_ylabel('y and exact')
axs[1].plot(t, v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(t, a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
plt.show()
###############################################
# -
g = 9.80655 #m/s^2
D = 0.00245 #m/s
# define scaling constant vT
vT = sqrt(g/D)
n,t = set_up_time (DeltaT=0.1, tfinal=0.5)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D, vT=vT)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
# ## 4. Identify Places Where You Can Make the Code More General
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, vT):
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i] ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT)) ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
if ( y[i+1] < 0.0): ## WHAT IF WE DON'T WANT TO STOP HERE??
break
a[n-1] = -g + D*v[n-1]*v[n-1] ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
return t, a, v, y, yanalytic
###############################################
# Fixing the extra argument is fairly simple, but the equations for acceleration and the analytical solution will be harder.
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, vT, min_position): # Note the extra argument
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = -g + D*v[i]*v[i] ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = y[0]-(vT*vT/g)*log(cosh(g*t[i+1]/vT)) ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
if ( y[i+1] < min_position):
break
a[n-1] = -g + D*v[n-1]*v[n-1] ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
return t, a, v, y, yanalytic
###############################################
# ## 5. See If You Can Make the Code More General by Passing Functions
def freefall_acceleration (g, D, v):
return -g + D*v*v
def freefall_yanalytic (g, D, y_0, t, v_0=0):
vT = sqrt(g/D)
return y_0-(vT*vT/g)*log(cosh(g*t/vT))
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, min_position, acceleration_eq, yanalytic_eq):
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = acceleration_eq (g, D, v[i]) ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = yanalytic_eq(g, D, y[0], t[i+1], v[0]) ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
if ( y[i+1] < min_position):
break
a[n-1] = acceleration_eq (g, D, v[n-1]) ## THIS EQUATION IS SPECIFICALLY FOR FREEFALL
return t, a, v, y, yanalytic
###############################################
n,t = set_up_time (DeltaT=0.1, tfinal=0.5)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D,min_position=0.0,\
acceleration_eq=freefall_acceleration, yanalytic_eq=freefall_yanalytic)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
def freefall_no_drag_acceleration (g, D, v): #Arguments must be in the same order
return -g
def freefall_no_drag_yanalytic (g, D, y_0, t, v_0):
return -0.5*g*t**2 + v_0*t + y_0
n,t = set_up_time (DeltaT=0.1, tfinal=0.5)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D, min_position=0.0,\
acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
# ## 6. Go Through Your Code And Identify Places Where There May Be User Errors
# What if the time was much larger?
n,t = set_up_time (DeltaT=0.1, tfinal=10.0)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D, min_position=0.0,\
acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
###############################################
# EULER'S METHOD
def euler (n, DeltaT, t, a, v, y, yanalytic, g, D, min_position, acceleration_eq, yanalytic_eq):
i_stop = n
# Start integrating using Euler's method
for i in range(n-1):
# expression for acceleration
a[i] = acceleration_eq (g, D, v[i])
# update velocity and position
y[i+1] = y[i] + DeltaT*v[i]
v[i+1] = v[i] + DeltaT*a[i]
# update time to next time step and compute analytical answer
t[i+1] = t[i] + DeltaT
yanalytic[i+1] = yanalytic_eq(g, D, y[0], t[i+1], v[0])
if ( y[i+1] < min_position):
stop_i = i+2
break
if stop_i != n:
t = t[0:stop_i]
a = a[0:stop_i]
v = v[0:stop_i]
y = y[0:stop_i]
yanalytic = yanalytic[0:stop_i]
a[-1] = acceleration_eq (g, D, v[-1])
return t, a, v, y, yanalytic
###############################################
n,t = set_up_time (DeltaT=0.1, tfinal=100.0)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D, min_position=0.0,\
acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
###############################################
# SETTING UP TIME
def set_up_time (DeltaT, tfinal):
n = ceil(tfinal/DeltaT)
t = np.zeros(n)
return n, t
###############################################
# What if we don't pass numbers?
set_up_time('words', 'words')
###############################################
# SETTING UP TIME
def set_up_time (DeltaT, tfinal):
assert isinstance(DeltaT, float) or isinstance(DeltaT, int)
assert isinstance(tfinal, float) or isinstance(tfinal, int)
n = ceil(tfinal/DeltaT)
t = np.zeros(n)
return n, t
###############################################
set_up_time('words', 'words')
# ## 7. Prevent So Many Arguments And Returns By Making a Class (Optional)
n,t = set_up_time (DeltaT=0.1, tfinal=100.0)
a, v, y, yanalytic = set_up_initial_arrays (n=n, v_0=10, y_0=0)
t, a, v, y, yanalytic = euler (n=n, DeltaT=0.1, t=t, a=a, v=v, y=y, yanalytic=yanalytic, g=g, D=D, min_position=0.0,\
acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
display_data(t, a, v, y, yanalytic)
graph_data(t, a, v, y, yanalytic)
# When making a class, any variable used in more than one place becomes class level so it can be used everywhere!
class ClassicalMechanicsSolvers ():
def __init__(self, g, D):
self.g = g
self.D = D
###############################################
# SETTING UP TIME
def set_up_time (self, DeltaT, tfinal):
self.DeltaT = DeltaT
self.n = ceil(tfinal/DeltaT)
self.t = np.zeros(n)
###############################################
###############################################
# SETTING UP THE INITIAL ARRAYS
def set_up_initial_arrays (self, v_0, y_0): ## No longer need n as an argument
# set up arrays for t, a, v, and y and we can compare our results with analytical ones
self.a = np.zeros(self.n)
self.v = np.zeros(self.n)
self.y = np.zeros(self.n)
self.yanalytic = np.zeros(self.n)
# Initial conditions
self.v[0] = v_0 #m/s
self.y[0] = y_0 #m
self.yanalytic[0] = y[0]
###############################################
###############################################
# EULER'S METHOD
def euler (self, min_position, acceleration_eq, yanalytic_eq):
i_stop = self.n
# Start integrating using Euler's method
for i in range(self.n-1):
# expression for acceleration
self.a[i] = acceleration_eq (self.g, self.D, self.v[i])
# update velocity and position
self.y[i+1] = self.y[i] + self.DeltaT*self.v[i]
self.v[i+1] = self.v[i] + self.DeltaT*self.a[i]
# update time to next time step and compute analytical answer
self.t[i+1] = self.t[i] + self.DeltaT
self.yanalytic[i+1] = yanalytic_eq(self.g, self.D, self.y[0], self.t[i+1], self.v[0])
if ( self.y[i+1] < min_position):
stop_i = i+2
break
if stop_i != self.n:
self.t = self.t[0:stop_i]
self.a = self.a[0:stop_i]
self.v = self.v[0:stop_i]
self.y = self.y[0:stop_i]
self.yanalytic = self.yanalytic[0:stop_i]
self.a[-1] = acceleration_eq (self.g, self.D, self.v[-1])
###############################################
###############################################
# DISPSLAY THE DATA
def display_data (self):
data = {'t[s]': self.t,
'y[m]': self.y-self.yanalytic,
'v[m/s]': self.v,
'a[m/s^2]': self.a
}
NewData = pd.DataFrame(data)
display(NewData.head())
###############################################
###############################################
# GRAPH THE DATA
#finally we plot the data
def graph_data (self):
fig, axs = plt.subplots(3, 1)
axs[0].plot(self.t, self.y, self.t, self.yanalytic)
axs[0].set_ylabel('y and exact')
axs[1].plot(self.t, self.v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(self.t, self.a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
plt.show()
###############################################
g = 9.80655 #m/s^2
D = 0.00245 #m/s
cm = ClassicalMechanicsSolvers (g, D)
cm.set_up_time (DeltaT=0.01, tfinal=100.0)
cm.set_up_initial_arrays (v_0=10, y_0=0)
cm.euler (min_position=0.0, acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
cm.display_data()
cm.graph_data()
# ## 8. See If There Are Ways to Reduce The Number of Function Calls (Optional)
# +
g = 9.80655 #m/s^2
D = 0.00245 #m/s
cm = ClassicalMechanicsSolvers (g, D)
################################################
# THESE ARE ALWAYS CALLED
cm.set_up_time (DeltaT=0.01, tfinal=100.0)
cm.set_up_initial_arrays (v_0=10, y_0=0)
################################################
cm.euler (min_position=0.0, acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
cm.display_data()
cm.graph_data()
# -
class ClassicalMechanicsSolvers ():
def __init__(self, g, D, DeltaT, tfinal, v_0, y_0):
self.g = g
self.D = D
self.set_up_time (DeltaT, tfinal)
self.set_up_initial_arrays (v_0, y_0)
###############################################
# SETTING UP TIME
def set_up_time (self, DeltaT, tfinal):
self.DeltaT = DeltaT
self.n = ceil(tfinal/DeltaT)
self.t = np.zeros(self.n)
###############################################
###############################################
# SETTING UP THE INITIAL ARRAYS
def set_up_initial_arrays (self, v_0, y_0): ## No longer need n as an argument
# set up arrays for t, a, v, and y and we can compare our results with analytical ones
self.a = np.zeros(self.n)
self.v = np.zeros(self.n)
self.y = np.zeros(self.n)
self.yanalytic = np.zeros(self.n)
# Initial conditions
self.v[0] = v_0 #m/s
self.y[0] = y_0 #m
self.yanalytic[0] = self.y[0]
###############################################
###############################################
# EULER'S METHOD
def euler (self, min_position, acceleration_eq, yanalytic_eq):
i_stop = self.n
# Start integrating using Euler's method
for i in range(self.n-1):
# expression for acceleration
self.a[i] = acceleration_eq (self.g, self.D, self.v[i])
# update velocity and position
self.y[i+1] = self.y[i] + self.DeltaT*self.v[i]
self.v[i+1] = self.v[i] + self.DeltaT*self.a[i]
# update time to next time step and compute analytical answer
self.t[i+1] = self.t[i] + self.DeltaT
self.yanalytic[i+1] = yanalytic_eq(self.g, self.D, self.y[0], self.t[i+1], self.v[0])
if ( self.y[i+1] < min_position):
stop_i = i+2
break
if stop_i != self.n:
self.t = self.t[0:stop_i]
self.a = self.a[0:stop_i]
self.v = self.v[0:stop_i]
self.y = self.y[0:stop_i]
self.yanalytic = self.yanalytic[0:stop_i]
self.a[-1] = acceleration_eq (self.g, self.D, self.v[-1])
###############################################
###############################################
# DISPSLAY THE DATA
def display_data (self):
data = {'t[s]': self.t,
'y[m]': self.y-self.yanalytic,
'v[m/s]': self.v,
'a[m/s^2]': self.a
}
NewData = pd.DataFrame(data)
display(NewData.head())
###############################################
###############################################
# GRAPH THE DATA
#finally we plot the data
def graph_data (self):
fig, axs = plt.subplots(3, 1)
axs[0].plot(self.t, self.y, self.t, self.yanalytic)
axs[0].set_ylabel('y and exact')
axs[1].plot(self.t, self.v)
axs[1].set_ylabel('v[m/s]')
axs[2].plot(self.t, self.a)
axs[2].set_xlabel('time[s]')
axs[2].set_ylabel('a[m/s^2]')
fig.tight_layout()
plt.show()
###############################################
g = 9.80655 #m/s^2
D = 0.00245 #m/s
DeltaT = 0.01
tfinal = 100
v_0 = 10
y_0 = 0
cm = ClassicalMechanicsSolvers(g, D, DeltaT, tfinal, v_0, y_0)
cm.euler (min_position=0.0, acceleration_eq=freefall_no_drag_acceleration, yanalytic_eq=freefall_no_drag_yanalytic)
cm.display_data()
cm.graph_data()
# ## 9. Think About Other Functionality You Can Add To The Class
#
# * Are there any other variables we should have in the initial arguments to make it more general?
# * Can you make your class work for 2D data as well or is it limited to 1D? Would it just be better to make another class to analyze 2D data?
# * Can you add in other solvers?
# * Should the function for acceleration and the analytical solution be included inside the class or not? Will you use them other places?
#
# **Bonus for Homework 6**: Submit a solvers class which implements Euler's, Euler-Cromer, and Velocity-Verlet as possible solvers (10 pts).
# ## 10. Using Your Code In Other Notebooks
| doc/pub/week7/ipynb/WritingReusableCode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sklearn as sk
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix
from sklearn import preprocessing as pp
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import NearMiss
from sklearn.model_selection import GridSearchCV
import pickle
import sqlite3
df = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/production_dataframe.csv')
df
x_user = df[df['user'] == 50].copy()
x_user.drop(columns=['city', 'continent_city'], inplace=True)
x_user.drop(columns=['rank'], inplace=True)
y_user = x_user['rank']
df_ready = df.drop(df[df.user == 50].index)
def get_df(table_name):
try:
conn = sqlite3.connect('/Users/tristannisbet/Documents/travel_app/places.db')
except Exception as e:
print('Error durring connection: ', str(e))
sql = """select * from {}""".format(table_name)
df = pd.read_sql_query(sql, conn)
return df
# +
# Sim scores
train_set2 = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_train_sim.csv')
test_set2 = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_test_sim.csv')
y_train_set = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_train_sim.csv')
y_test_set = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_test_sim.csv')
train_set2
# -
all_data_raw = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/user_city_raw.csv')
all_data_sim = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/user_city_sim.csv')
# +
x_trainfull = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_train_full.csv')
x_testfull = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_test_full.csv')
y_trainfull = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_train_full.csv')
y_testfull = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_test_full.csv')
# -
x_trainfull.set_index(['user', 'label_id'], inplace=True)
y_trainfull.set_index(['user', 'label_id'], inplace=True)
y_testfull.set_index(['user', 'label_id'], inplace=True)
x_testfull.set_index(['user', 'label_id'], inplace=True)
x_trainfull
all_data_sim
y_trainfull = pd.Series(y_trainfull['rank_y'].values, index=y_trainfull.index)
all_data
y_train_set.set_index(['user', 'city_id'], inplace=True)
train_set2.set_index(['user', 'city_id'], inplace=True)
y_test_set.set_index(['user', 'city_id'], inplace=True)
test_set2.set_index(['user', 'city_id'], inplace=True)
train_set2.drop(columns=['top'], inplace=True)
test_set2.drop(columns=['top'], inplace=True)
# +
# Raw attraction and food scores
x_test = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_test.csv')
y_test = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_test.csv')
x_train = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/x_train.csv')
y_train = pd.read_csv('/Users/tristannisbet/Documents/SM/Dataframe/new/y_train.csv')
x_train
# -
y_train.set_index(['user', 'label_id'], inplace=True)
x_train.set_index(['user', 'label_id'], inplace=True)
y_test.set_index(['user', 'label_id'], inplace=True)
x_test.set_index(['user', 'label_id'], inplace=True)
# ## Fake user data
# +
x_test_fake = pd.read_csv("/Users/tristannisbet/Documents/SM/Dataframe/new/x_testfake.csv")
y_test_fake = pd.read_csv("/Users/tristannisbet/Documents/SM/Dataframe/new/y_testfake.csv")
x_train_fake = pd.read_csv("/Users/tristannisbet/Documents/SM/Dataframe/new/x_trainfake.csv")
y_train_fake = pd.read_csv("/Users/tristannisbet/Documents/SM/Dataframe/new/y_trainfake.csv")
# -
x_test_fake.set_index(['user', 'city_id'], inplace=True)
y_test_fake.set_index(['user', 'city_id'], inplace=True)
x_train_fake.set_index(['user', 'city_id'], inplace=True)
y_train_fake.set_index(['user', 'city_id'], inplace=True)
y_test_fake[y_test_fake['rank'] == 3]
def rank_from_col(x):
if x['rank'] == 5:
return 1
elif x['rank'] == 4:
return 1
elif x['rank'] == 3:
return 1
elif x['rank'] == 2:
return 1
elif x['rank'] == 1:
return 1
else:
return 0
# +
#y_test_fake['top'] = y_test_fake.apply(rank_from_col, axis=1)
y_train_fake['top'] = y_train_fake.apply(rank_from_col, axis=1)
# -
y_train_fake[y_train_fake['rank'] == 5]
y_train_fake.drop(columns=['rank'], inplace=True)
y_test_fake.drop(columns=['rank'], inplace=True)
# ## Production Test
ready = df_ready.drop(columns=['city'])
ready.set_index(['user', 'city_id'], inplace=True)
x = ready.drop(columns=['rank'])
y = ready['rank']
y
x.drop(columns=['continent_city'], inplace=True)
# ### Model Building
# +
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(random_state=12, max_depth=25)
clf.fit(x, y)
predictions_rf = clf.predict(x)
probs_rf = clf.predict_proba(x)
# -
x
x_user.set_index(['user', 'city_id'], inplace=True)
# +
pickle.dump(clf, open('model_test2.pkl','wb'))
model = pickle.load(open('model_test2.pkl','rb'))
predicts = model.predict(x_user)
probs = model.predict_proba(x_user)
# -
probs
x_user
y_trainfull = pd.Series(y_trainfull['rank_y'].values, index=y_trainfull.index)
print(classification_report(y_user, predicts))
print(confusion_matrix(y_user,predicts))
probs
from sklearn.metrics import mean_squared_error
#print('RMSE on train data: ', mean_squared_error(y_trainfull, )**(0.5))
print('RMSE on test data: ', mean_squared_error(y_testfull, predictions_rf)**(0.5))
import matplotlib.pyplot as plt
plt.figure(figsize=(10,7))
feat_importances = pd.Series(clf.feature_importances_, index = x_trainfull.columns)
feat_importances.nlargest(7).plot(kind='barh');
x_testfull.index.get_level_values('user').unique()
x_user = x_testfull.xs(55, level='user', drop_level=False).copy()
y_user = y_testfull.xs(55, level='user', drop_level=False).copy()
# ### Accuracy Score Metric
results5 = x_testfull.copy()
results = x_user.copy()
# This will add the probability and the predicted values from the model
def split_array(prob, x_df, predict):
df = x_df.copy()
lst1 = list(prob[:, 0])
lst2 = list(prob[:, 1])
predict_lst = list(predict)
df['prob_0'] = pd.Series(lst1, index=df.index)
df['prob_1'] = pd.Series(lst2, index=df.index)
df['predicted'] = pd.Series(predict_lst, index=df.index)
return df
results
okk = split_array(probs, results, predicts)
okk
ok.sort_values('prob_1', ascending=False)
cities = get_df('cities')
cities
# +
def add_top_city(results_df, y_test_set):
# adds actual top ranked cities
results_df_top = pd.merge(right=y_test_set, left=results_df, right_index=True, left_index=True)
return results_df_top
def add_city_name(results_df, city_df):
results_df.reset_index(level=1, inplace=True)
city_dict = dict(zip(city_df['city_id'], city_df['city']))
results_df['city'] = results_df['label_id'].map(city_dict)
results_df.set_index('label_id', append=True, inplace=True)
return results_df
# -
df.columns
ok2 = add_city_name(okk, df)
ok2
ok3 = add_top_city(ok2, y_user)
ok3
ok3.sort_values('prob_1', ascending=False)
# +
# filter only continent = 4 and rank_y != 0
# Choose and print the top 3 cities
# -
y_user
| notebooks/model_ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip list
# +
from selenium import webdriver
import os
chromediver_path = 'C:\\Users\demetoir_desktop\\PycharmProjects\\kaggle-MLtools\\chromedriver_win32'
driver = webdriver.Chrome(os.path.join(chromediver_path, 'chromedriver'))
# driver = webdriver.PhantomJS(os.path.join('.', 'phantomjs-2.1.1-windows', 'bin', 'phantomjs'))
base_url = 'http://www.ppomppu.co.kr/zboard/zboard.php?id=freeboard&no=5836128'
driver.get(base_url)
# +
ase_url = 'http://www.ppomppu.co.kr/zboard/zboard.php?id=freeboard&no=5836128'
base_url = 'http://www.ppomppu.co.kr/zboard/view.php?id=faq&no=2'
show_comment_page = """javascript:go_page( "faq", 2, 1 ); return false;"""
go_comment_page_n = """javascript:go_page( "faq", 2, {} ); return false;"""
url, querys = base_url.split('?')
url, querys
# -
query_list = querys.split('&')
query_list
query_dict = {}
for s in query_list:
name, val = s.split('=')
query_dict[name] = val
query_dict
| jupyter/selenium_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import climlab
alb = 0.25
# State variables (Air and surface temperature)
state = climlab.column_state(num_lev=20)
# Parent model process
rcm = climlab.TimeDependentProcess(state=state)
# Fixed relative humidity
h2o = climlab.radiation.ManabeWaterVapor(state=state)
# Couple water vapor to radiation
rad = climlab.radiation.RRTMG(state=state, specific_humidity=h2o.q, albedo=alb)
# Convective adjustment
conv = climlab.convection.ConvectiveAdjustment(state=state, adj_lapse_rate=6.5)
# Couple everything together
rcm.add_subprocess('Radiation', rad)
rcm.add_subprocess('WaterVapor', h2o)
rcm.add_subprocess('Convection', conv)
# Run the model
rcm.integrate_years(5)
# Check for energy balance
print(rcm.ASR - rcm.OLR)
# +
alb = 1.
# State variables (Air and surface temperature)
state = climlab.column_state(num_lev=30)
# Parent model process
rcm = climlab.TimeDependentProcess(state=state)
# Fixed relative humidity
h2o = climlab.radiation.ManabeWaterVapor(state=state)
# Couple water vapor to radiation
rad = climlab.radiation.RRTMG(state=state, specific_humidity=h2o.q, albedo=alb)
rcm.add_subprocess('Radiation', rad)
rcm.compute()
rcmx = rcm.to_xarray(rcm.diagnostics)
# -
rcmx
| notebooks/01_climlab_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="PlNF-_DjfgtK"
#
# # **Install libraries**
# + colab={"base_uri": "https://localhost:8080/"} id="04jwVrdr9sdb" outputId="a002cff5-7bc6-43ac-90f9-0252450d40f5"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="BGiskWK4_6b-" outputId="d50478df-cc7b-463c-d6b9-f68734bbb7ab"
# !pip install datasets tqdm pandas
# + colab={"base_uri": "https://localhost:8080/"} id="_d8OEpLA5Aii" outputId="fbc0bb8a-6d6c-404c-c33a-244d23a8ee2b"
# !pip install sentencepiece
# + colab={"base_uri": "https://localhost:8080/"} id="CjKf77Z7EYpk" outputId="eb91f4fc-4763-4528-b8fc-a31f6ae493e3"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/"} id="oaGb9Nk2sc7v" outputId="dab86df3-3652-4a1c-b263-8aef1656cf1f"
# !pip install wandb
# + id="0OaSKNC3AvaX"
import pandas as pd
from datasets import load_dataset
from tqdm import tqdm
# + colab={"base_uri": "https://localhost:8080/"} id="Pn-G_eUwFovO" outputId="9f10ba64-a488-4848-b655-ccb7dc89c1f7"
# Check we have a GPU and check the memory size of the GPU
# !nvidia-smi
# + [markdown] id="H0n55Ex1Bl2k"
# # **Import packages**
# + colab={"base_uri": "https://localhost:8080/"} id="ht8Fu9U2-IRG" outputId="ed2d0670-cabf-4869-f47d-84217c28c263"
import argparse
import glob
import os
import json
import time
import logging
import random
import re
from itertools import chain
from string import punctuation
import nltk
nltk.download('punkt')
from nltk.tokenize import sent_tokenize
import pandas as pd
import numpy as np
import torch
from torch.utils.data import Dataset, DataLoader
from transformers import (
AdamW,
T5ForConditionalGeneration,
T5Tokenizer,
get_linear_schedule_with_warmup
)
# + [markdown] id="Ykds8V47B1XT"
# # **Set a seed**
# + id="p03e0mY13jdV"
import random
import numpy as np
import torch
import datasets
# + id="CyrYjMFREUCn"
def set_seed(seed):
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
set_seed(42)
# + [markdown] id="l7YZhzbl88AE"
#
#
# ```
# # This is formatted as code
# ```
#
# # ***C4-200M dataset***
# + id="bYqzCt4mfkoc"
pd.set_option('display.max_colwidth', None)
# + colab={"base_uri": "https://localhost:8080/"} id="SWQZdyVlfLL_" outputId="f4c1f085-3e59-443d-c38d-0879146260ea"
df = pd.read_csv('/content/drive/MyDrive/c4_200m/c4_200m_550k.csv')
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="8AcvjhlVfPe2" outputId="5ad86c9b-97d2-407c-dcdc-e0a0097ff14d"
df.head()
# + id="6ANFcc6SE0p2"
from transformers import (
T5ForConditionalGeneration, T5Tokenizer,
Seq2SeqTrainingArguments, Seq2SeqTrainer, DataCollatorForSeq2Seq
)
from torch.utils.data import Dataset, DataLoader
# + id="35po7q9CE57P" colab={"base_uri": "https://localhost:8080/", "height": 113, "referenced_widgets": ["362a0db21dc14770b470ed10b85a0bf5", "65c1ed7dd4f64a0c9dfdc58527a40489", "d7b577a45c02458e8e2f766a71a7ad72", "919ee86b19b740e8b1570a70cc2e0ba4", "62877730538342f19934ff9cd4ba8548", "3cb2c936c0df4c9aac652754396b0248", "ba1d94e63dac45728e8411ae99016849", "f3aedbb9691b4165816cc8c6fb92f7e2", "49efa2311d4c43caa402bcd30e378ad2", "<KEY>", "<KEY>", "2bb78851ed64436499ed9317cbba7e6b", "4e5ec4ea3a3e44138d1391a692843529", "<KEY>", "c79abf5f210b4bad94168e67e1a6e131", "<KEY>", "d2ef0a9f9ff24dbdaddbcb23ad49d6a3", "<KEY>", "<KEY>", "0b69945bc05a490484f75cfe4c69ee36", "<KEY>", "a2f5a6caaa0f419d88893b54ebd2ca56", "<KEY>", "c633a9d9ab7141328511168f8879710e", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "d7e16c482e0340509bb75709e25c6195", "8cb1265204a34152b0865eaf7d9b98e1", "c5af34257313455d8dba68c774f18955", "<KEY>"]} outputId="5c5fd27a-9977-414a-c9bd-7167922654e4"
model_name = 't5-base'
tokenizer = T5Tokenizer.from_pretrained(model_name)
model = T5ForConditionalGeneration.from_pretrained(model_name)
# + id="a75jy7E1igUi"
def calc_token_len(example):
return len(tokenizer(example).input_ids)
# + id="UyMhxBbjYN-u" colab={"base_uri": "https://localhost:8080/"} outputId="1f3010fa-ff39-4acf-ed36-da722c8d5f3d"
from sklearn.model_selection import train_test_split
train_df, test_df = train_test_split(df, test_size=0.10, shuffle=True)
train_df.shape, test_df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="uruzAQuujAeA" outputId="40602509-b32e-43cc-f383-a689a9e4ba92"
test_df['input_token_len'] = test_df['input'].apply(calc_token_len)
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="oNvjbMF3jWkW" outputId="4fb7903d-ebc1-445c-ebe2-10273278b357"
test_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="Qy3eyFHxjZWm" outputId="504d7119-92c2-4744-ad8e-d7cc0501f39f"
test_df['input_token_len'].describe()
# + [markdown] id="5hz7cdWnr_OO"
# ### We will use a token length of 64 since it will cover the vast majority of examples
# + id="rSvsHw6HeHrl"
from datasets import Dataset
train_dataset = Dataset.from_pandas(train_df)
test_dataset = Dataset.from_pandas(test_df)
# + colab={"base_uri": "https://localhost:8080/"} id="XfRhXHBGoiiU" outputId="3c39aaab-2c7d-4e78-af3e-b50a0a491c12"
test_dataset
# + [markdown] id="onckS0qK4kTF"
# ### Load the Dataset
# + id="hDjTyQH75OKH"
from torch.utils.data import Dataset, DataLoader
class GrammarDataset(Dataset):
def __init__(self, dataset, tokenizer,print_text=False):
self.dataset = dataset
self.pad_to_max_length = False
self.tokenizer = tokenizer
self.print_text = print_text
self.max_len = 64
def __len__(self):
return len(self.dataset)
def tokenize_data(self, example):
input_, target_ = example['input'], example['output']
# tokenize inputs
tokenized_inputs = tokenizer(input_, pad_to_max_length=self.pad_to_max_length,
max_length=self.max_len,
return_attention_mask=True)
tokenized_targets = tokenizer(target_, pad_to_max_length=self.pad_to_max_length,
max_length=self.max_len,
return_attention_mask=True)
inputs={"input_ids": tokenized_inputs['input_ids'],
"attention_mask": tokenized_inputs['attention_mask'],
"labels": tokenized_targets['input_ids']
}
return inputs
def __getitem__(self, index):
inputs = self.tokenize_data(self.dataset[index])
if self.print_text:
for k in inputs.keys():
print(k, len(inputs[k]))
return inputs
# + colab={"base_uri": "https://localhost:8080/"} id="AlkkfaIS64lZ" outputId="67954e1d-89f6-4ff5-c577-eb270103d3e4"
dataset = GrammarDataset(test_dataset, tokenizer, True)
print(dataset[121])
# + [markdown] id="17tTZNRU9Nbd"
# ### Define Evaluator
# + colab={"base_uri": "https://localhost:8080/"} id="dAYzYHBR9Pj2" outputId="5b40481e-b4a5-420b-9f09-697c02a4d1b5"
# !pip install rouge_score
# + id="ojPxcaxr9P_o" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["b59067fb92d64f03bcc1e0ac0373f159", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "3316912ff6bd47f980f746368545808a", "063911b025f3455c9a13fff36913f469", "cee0a4e66ddb4e4a8a19c5a1a4ec62e7", "<KEY>"]} outputId="758621eb-857f-431d-dbd0-5c850c1df3f4"
from datasets import load_metric
rouge_metric = load_metric("rouge")
# + [markdown] id="tlkEgSrq9lxJ"
# ### Train Model
# + id="5x4_f3P79oCK"
data_collator = DataCollatorForSeq2Seq(tokenizer, model=model, padding='longest', return_tensors='pt')
# + id="rc-YLKSK9pkA"
# defining training related arguments
batch_size = 16
args = Seq2SeqTrainingArguments(output_dir="/content/drive/MyDrive/c4_200m/weights",
evaluation_strategy="steps",
per_device_train_batch_size=batch_size,
per_device_eval_batch_size=batch_size,
learning_rate=2e-5,
num_train_epochs=1,
weight_decay=0.01,
save_total_limit=2,
predict_with_generate=True,
fp16 = True,
gradient_accumulation_steps = 6,
eval_steps = 500,
save_steps = 500,
load_best_model_at_end=True,
logging_dir="/logs",
report_to="wandb")
# + colab={"base_uri": "https://localhost:8080/"} id="YutXc8Q1-DG2" outputId="ccc17962-fd5e-45eb-dbee-b10d9342d7cf"
import nltk
nltk.download('punkt')
import numpy as np
def compute_metrics(eval_pred):
predictions, labels = eval_pred
decoded_preds = tokenizer.batch_decode(predictions, skip_special_tokens=True)
# Replace -100 in the labels as we can't decode them.
labels = np.where(labels != -100, labels, tokenizer.pad_token_id)
decoded_labels = tokenizer.batch_decode(labels, skip_special_tokens=True)
# Rouge expects a newline after each sentence
decoded_preds = ["\n".join(nltk.sent_tokenize(pred.strip())) for pred in decoded_preds]
decoded_labels = ["\n".join(nltk.sent_tokenize(label.strip())) for label in decoded_labels]
result = rouge_metric.compute(predictions=decoded_preds, references=decoded_labels, use_stemmer=True)
# Extract a few results
result = {key: value.mid.fmeasure * 100 for key, value in result.items()}
# Add mean generated length
prediction_lens = [np.count_nonzero(pred != tokenizer.pad_token_id) for pred in predictions]
result["gen_len"] = np.mean(prediction_lens)
return {k: round(v, 4) for k, v in result.items()}
# + colab={"base_uri": "https://localhost:8080/"} id="fyOdMJQW-FFJ" outputId="2580b1d1-bc96-4efb-8923-5d1842f951be"
# defining trainer using 🤗
trainer = Seq2SeqTrainer(model=model,
args=args,
train_dataset= GrammarDataset(train_dataset, tokenizer),
eval_dataset=GrammarDataset(test_dataset, tokenizer),
tokenizer=tokenizer,
data_collator=data_collator,
compute_metrics=compute_metrics)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="a_JO3oUR-G10" outputId="f2f98e53-d82a-4582-fbbd-141a4659cf16"
trainer.train()
# + colab={"base_uri": "https://localhost:8080/"} id="yIkeGbykntXp" outputId="7b1619c6-f79c-401c-f26b-755cf5be965a"
trainer.save_model('t5_gec_model')
# + colab={"base_uri": "https://localhost:8080/"} id="uvoTQ_e-nyqi" outputId="e41e0917-2bcd-479e-da32-35b725922a09"
# !zip -r 't5_gec_model.zip' 't5_gec_model'
# + id="ZkutuP4Tn3j1"
# !mv t5_gec_model.zip /content/drive/MyDrive/c4_200m
# + [markdown] id="DjfvFEgEiXgR"
# I have uploaded this model to HuggingFace Model Zoo and we can run inference using it
# + [markdown] id="mQzJE6ybKPaR"
# ## Testing
# + id="lqDyDu5GKOrg"
import torch
from transformers import T5Tokenizer, T5ForConditionalGeneration
model_name = 'deep-learning-analytics/GrammarCorrector'
torch_device = 'cuda' if torch.cuda.is_available() else 'cpu'
tokenizer = T5Tokenizer.from_pretrained(model_name)
model = T5ForConditionalGeneration.from_pretrained(model_name).to(torch_device)
def correct_grammar(input_text,num_return_sequences):
batch = tokenizer([input_text],truncation=True,padding='max_length',max_length=64, return_tensors="pt").to(torch_device)
translated = model.generate(**batch,max_length=64,num_beams=4, num_return_sequences=num_return_sequences, temperature=1.5)
tgt_text = tokenizer.batch_decode(translated, skip_special_tokens=True)
return tgt_text
# + colab={"base_uri": "https://localhost:8080/"} id="mMm8PGpMhYxl" outputId="095043df-98dc-4307-906e-8d930480cee3"
text = 'He are moving here.'
print(correct_grammar(text, num_return_sequences=2))
# + id="8qLMT5DyvsKT" colab={"base_uri": "https://localhost:8080/"} outputId="0ec27408-8b9a-49ad-9a7f-29a187256526"
text = 'Cat drinked milk'
print(correct_grammar(text, num_return_sequences=1))
# + id="AX_peD3xh-fH"
| GrammarCorrector/T5_Grammar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from random import randrange
# +
class Dice(object):
roll_count = 0
rolls = {1:0, 2:0, 3:0, 4:0, 5:0, 6:0}
def reset(self):
self.rolls = {1:0, 2:0, 3:0, 4:0, 5:0, 6:0}
self.roll_count = 0
def rollOnce(self):
val = randrange(1,7)
if val in self.rolls:
self.rolls[val] += 1
self.roll_count += 1
def rollMany(self, num_rolls):
for i in range(num_rolls):
self.rollOnce()
def printResults(self):
print(f'Rolls: {self.roll_count}')
for (val, cnt) in self.rolls.items():
if self.roll_count > 0:
pct = round(cnt / self.roll_count * 100, 1)
else:
pct = "-"
print(f'Roll {val}: {cnt} ({pct}%)')
def rollPrint(self, num_rolls):
self.rollMany(num_rolls)
self.printResults()
dice = Dice()
# -
dice.rollPrint(10000)
dice.reset()
dice.rollPrint(100000)
| rollDice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nlp_gpu)
# language: python
# name: nlp_gpu
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # BiDAF Model Deep Dive on AzureML
# 
# This notebook demonstrates a deep dive into a popular question-answering (QA) model, Bi-Directional Attention Flow (BiDAF). We use [AllenNLP](https://allennlp.org/), an open-source NLP research library built on top of PyTorch, to train the BiDAF model from scratch on the [SQuAD](https://rajpurkar.github.io/SQuAD-explorer/) dataset, using Azure Machine Learning ([AzureML](https://azure.microsoft.com/en-us/services/machine-learning-service/)).
#
# The following capabilities are highlighted in this notebook:
# - AmlCompute
# - Datastore
# - Logging
# - AllenNLP library
# ## Table of Contents
# 1. [Introduction](#1.-Introduction)
# * 1.1 [SQuAD Dataset](#1.1-SQuAD-Dataset)
# * 1.2 [BiDAF Model](#1.2-BiDAF-Model)
# * 1.3 [AllenNLP](#1.3-AllenNLP)
# 2. [AzureML Setup](#2.-AzureML-Setup)
# * 2.1 [Link to or create a `Workspace`](#2.1-Link-to-or-create-a-Workspace)
# * 2.2 [Set up an `Experiment` and Logging](#2.2-Set-up-an-Experiment-and-Logging)
# * 2.3 [Link `AmlCompute` compute target](#2.3-Link-AmlCompute-Compute-Target)
# * 2.4 [Upload Files to `Datastore`](#2.4-Upload-Files-to-Datastore)
# 3. [Prepare Training Script](#3.-Prepare-Training-Script)
# 4. [Create a PyTorch Estimator](#4.-Create-a-PyTorch-Estimator)
# 5. [Submit a Job](#5.-Submit-a-Job)
# 6. [Inspect Results of Run](#6.-Inspect-Results-of-Run)
# * 6.1 [Evaluation on SQuAD](#6.1-Evaluation-on-SQuAD)
# * 6.2 [Try the Best Model](#6.2-Try-the-Best-Model)
# ## 1. Introduction
# ### 1.1 SQuAD Dataset
# The [SQuAD](https://rajpurkar.github.io/SQuAD-explorer/) dataset was released in 2016 and has become a benchmarking dataset for machine comprehension tasks. It contains a set of more than 100,000 question-context tuples along with their answers, extracted from Wikipedia articles. 90,000 of the question-context tuples make up the training set and the remaining 10,000 compose the development set. The answers are spans in the context (given passage) and are evaluated against human-labeled answers. Two metrics are used for evaluation: Exact Match (EM) and F1 score.
# ### 1.2 BiDAF Model
# The [BiDAF](https://www.semanticscholar.org/paper/Bidirectional-Attention-Flow-for-Machine-Seo-Kembhavi/007ab5528b3bd310a80d553cccad4b78dc496b02
# ) model achieved state-of-the-art performance on the SQuAD dataset in 2017 and is a well-respected, performant baseline for QA. The BiDAF network is a "hierarchical multi-stage architecture for modeling representations of the context at different levels of granularity. BiDAF includes character-level, word-level, and phrase-level embeddings, and uses bi-directional attention flow to allow for query-aware context representations".
#
# The network contains six different layers, as described by [Seo et al, 2017](https://www.semanticscholar.org/paper/Bidirectional-Attention-Flow-for-Machine-Seo-Kembhavi/007ab5528b3bd310a80d553cccad4b78dc496b02):
#
# 1. **Character Embedding Layer**: character-level CNNs to embed each word
# 2. **Word Embedding Layer**: word embeddings using pre-trained GloVe word vectors
# 3. **Phrase Embedding Layer**: LSTM on top of the previous layers to model the temporal interactions between words
# 4. **Attention Flow Layer**: Fuses information from the context and query words. Unlike previous models, "the attention flow layer is not used to summarize the query and context into a single feature vectors. Instead, the attention vectors at each time step, along with embeddings from previous layers, are allowed to flow through to the subsequent modeling layers", reducing information loss.
# 5. **Modeling Layer**: produces a matrix of contextual information about the word with respect to the entire context paragraph and query
# 6. **Output Layer**: predicts the start and end indices of the phrase in the paragraph
#
# The following figure displays the architecture of the BiDAF network.
# 
# ### 1.3 AllenNLP
# The notebook demonstrates how to use the BiDAF implementation provided by [AllenNLP](https://www.semanticscholar.org/paper/A-Deep-Semantic-Natural-Language-Processing-Gardner-Grus/a5502187140cdd98d76ae711973dbcdaf1fef46d), an open-source NLP research library built on top of PyTorch. AllenNLP is a product of the Allen Institute for Artifical Intelligence and is used widely across differnet universities and top companies (including Facebook Research and Amazon Alexa). They maintain a robust and active [Github repository](https://github.com/allenai/allennlp) as well as a [website](https://allennlp.org/) with documentation and demos. Their model is a reimplementation of the original BiDAF model and they report a higher EM score and faster training times than the original BiDAF system (68.3 EM score versus 67.7 and 10x speedup, taking ~4 hours on a p2.xlarge). The AllenNLP library is mainly designed for use through the command line (and most tutorials use this method), but can also be used programatically.
# The AllenNLP library focuses on the creation of NLP pipelines with easily interchangable building blocks. The general pipeline steps are as follows:
# - DatasetReader: defines how to extract information from your data and convert it into Instance objects that will be used by the model
# - Iterator: takes the instances produced by the DatasetReader and batches them for training
# - Model
# - Trainer: trains the model and records metrics
# - Predictor: takes raw strings and produces predictions
#
# Each step is loosely-coupled, making it easy to swap different options for each step. While it is possible to construct your own AllenNLP objects (see this [tutorial](https://mlexplained.com/2019/01/30/an-in-depth-tutorial-to-allennlp-from-basics-to-elmo-and-bert/) for a great deep-dive into constructing your own AllenNLP pipeline), the easiest way is to utilize the JSON-like parameter constructor methods provided by most AllenNLP objects. For example, rather than
#
# ```
# lstm = PytorchSeq2SeqWrapper(torch.nn.LSTM(EMBEDDING_DIM, HIDDEN_DIM, batch_first=True))
# ```
# we can use
#
# ```
# lstm_params = Params({
# "type": "lstm",
# "input_size": EMBEDDING_DIM,
# "hidden_size": HIDDEN_DIM
# })
#
# lstm = Seq2SeqEncoder.from_params(lstm_params)
# ```
# This provides two advantages:
# 1. Experiments can be declaratively specified in a separate [configuration file](https://github.com/allenai/allennlp/blob/master/tutorials/tagger/README.md#using-config-files)
# 2. Experiments can be easily changed with no coding, rather just changing the entry in the config file
# **AllenNLP Resources:**
#
# The following resources are recommended for understanding how the AllenNLP library works and being able to implement your own models and pipelines
#
# - Information about the provided AllenNLP models: https://allennlp.org/models
# - Using configuration files: https://github.com/allenai/allennlp/blob/master/tutorials/tagger/README.md#using-config-files
# - In-depth discussion of each AllenNLP object used and how to construct your own specialized ones: https://mlexplained.com/2019/01/30/an-in-depth-tutorial-to-allennlp-from-basics-to-elmo-and-bert/
# - AllenNLPs Part-of-Speech-Tagging tutorial showcasing how to use their methods programatically: https://allennlp.org/tutorials
# - Short AllenNLP programatic tutorial: https://github.com/titipata/allennlp-tutorial/blob/master/allennlp_tutorial.ipynb
# +
# Imports
import sys
import os
import shutil
sys.path.append("../../")
import json
from urllib.request import urlretrieve
import scrapbook as sb
#import utils
from utils_nlp.common.timer import Timer
from utils_nlp.azureml import azureml_utils
import azureml as aml
from azureml.core import Datastore, Experiment
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.exceptions import ComputeTargetException
from azureml.train.dnn import PyTorch
from azureml.widgets import RunDetails
from azureml.core.conda_dependencies import CondaDependencies
from azureml.exceptions import ComputeTargetException
from allennlp.predictors import Predictor
print("System version: {}".format(sys.version))
print("Azure ML SDK Version:", aml.core.VERSION)
# + tags=["parameters"]
PROJECT_FOLDER = "./bidaf-question-answering"
SQUAD_FOLDER = "./squad"
BIDAF_CONFIG_PATH = "."
LOGS_FOLDER = '.'
NUM_EPOCHS = 25
PIP_PACKAGES = [
"allennlp==0.8.4",
"azureml-sdk==1.0.48",
"https://github.com/explosion/spacy-models/releases/download/en_core_web_sm-2.1.0/en_core_web_sm-2.1.0.tar.gz",
]
CONDA_PACKAGES = ["jsonnet", "cmake", "regex", "pytorch", "torchvision"]
config_path = (
"./.azureml"
) # Path to the directory containing config.json with azureml credentials
# Azure resources
subscription_id = "YOUR_SUBSCRIPTION_ID"
resource_group = "YOUR_RESOURCE_GROUP_NAME"
workspace_name = "YOUR_WORKSPACE_NAME"
workspace_region = "YOUR_WORKSPACE_REGION" #Possible values eastus, eastus2 and so on.
# -
# ## 2. AzureML Setup
# Now, we set up the necessary components for running this as an AzureML experiment
# 1. Create or link to an existing `Workspace`
# 2. Set up an `Experiment` with `logging`
# 3. Create or attach existing `AmlCompute`
# 4. Upload our data to a `Datastore`
# ### 2.1 Link to or create a Workspace
# The following cell looks to set up the connection to your [Azure Machine Learning service Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace). You can choose to connect to an existing workspace or create a new one.
#
# **To access an existing workspace:**
# 1. If you have a `config.json` file, you do not need to provide the workspace information; you will only need to update the `config_path` variable that is defined above which contains the file.
# 2. Otherwise, you will need to supply the following:
# * The name of your workspace
# * Your subscription id
# * The resource group name
#
# **To create a new workspace:**
#
# Set the following information:
# * A name for your workspace
# * Your subscription id
# * The resource group name
# * [Azure region](https://azure.microsoft.com/en-us/global-infrastructure/regions/) to create the workspace in, such as `eastus2`.
#
# This will automatically create a new resource group for you in the region provided if a resource group with the name given does not already exist.
ws = azureml_utils.get_or_create_workspace(
config_path=config_path,
subscription_id=subscription_id,
resource_group=resource_group,
workspace_name=workspace_name,
workspace_region=workspace_region,
)
print(
"Workspace name: " + ws.name,
"Azure region: " + ws.location,
"Subscription id: " + ws.subscription_id,
"Resource group: " + ws.resource_group,
sep="\n",
)
# ### 2.2 Set up an Experiment and Logging
# Next, we set up an `Experiment` named bidaf-question-answering, add logging capabilities, and create a local folder that will be the source directory for the AzureML run.
# +
# Make a folder for the project
os.makedirs(PROJECT_FOLDER, exist_ok=True)
# Set up an experiment
experiment_name = "NLP-QA-BiDAF-deepdive"
experiment = Experiment(ws, experiment_name)
# Add logging to our experiment
run = experiment.start_logging(snapshot_directory=PROJECT_FOLDER)
# -
# ### 2.3 Link AmlCompute Compute Target
#
# We need to link a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training our model (see [compute options](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#supported-compute-targets) for explanation of the different options). We will use an [AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) target and link to an existing target (if the cluster_name exists) or create a STANDARD_NC6 GPU cluster (autoscales from 0 to 4 nodes) in this example. Creating a new AmlComputes takes approximately 5 minutes.
#
# As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
# choose your cluster
cluster_name = "gpu-test"
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print("Found existing compute target.")
except ComputeTargetException:
print("Creating a new compute target...")
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_NC6", max_nodes=4
)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# use get_status() to get a detailed status for the current AmlCompute.
print(compute_target.get_status().serialize())
# -
# ### 2.4 Upload Files to Datastore
# This step uploads our local files to a `Datastore` so that the data is accessible from the remote compute target. A DataStore is backed either by a Azure File Storage (default option) or Azure Blob Storage ([how to decide between these options](https://docs.microsoft.com/en-us/azure/storage/common/storage-decide-blobs-files-disks)) and data is made accessible by mounting or copying data to the compute target.
# First, we download the SQuAD data files and save to a folder called squad.
# +
os.makedirs(SQUAD_FOLDER, exist_ok=True) # make squad folder locally
urlretrieve(
"https://allennlp.s3.amazonaws.com/datasets/squad/squad-train-v1.1.json",
filename=SQUAD_FOLDER+"/squad_train.json",
)
urlretrieve(
"https://allennlp.s3.amazonaws.com/datasets/squad/squad-dev-v1.1.json",
filename=SQUAD_FOLDER+"/squad_dev.json",
)
# -
# We also copy our AllenNLP configuration file (bidaf_config.json) into this squad folder so that it can be uploaded to the `Datastore` and accessed during training. As described in [Section 1.3](#1.3-AllenNLP), this configuration files allows us to easily specify the parameters for instantiating AllenNLP objects. This file contains a dictionary of dictionaries. The top level contains 4 main keys: dataset_reader, model, iterator, and trainer (plus keys for train_data_path, validation_data_path, and evaluate_on_test). If you notice carefully from [Section 1.3](#1.3-AllenNLP), these correspond to the AllenNLP object building blocks. Each of these keys map to a dictionary of parameters. For instance, the trainer dictionary contains keys to specify the number of epochs, learning rate scheduler, optimizer, etc. The parameter settings provided here are the ones suggested by AllenNLP for the BiDAF model; however, below we demonstrate how to override these parameters without having to change this configuration file directly.
shutil.copy(BIDAF_CONFIG_PATH+'/bidaf_config.json', SQUAD_FOLDER)
# Now we upload both the SQuAD data files as well as the configuration file to the datastore. `ws.datastores` lists all options for datastores and `ds.account_name` gets the name of the datastore that can be used to find it in the Azure portal. Once we have selected the appropriate datastore, we use the `upload()` method to upload all files from the squad local folder to a folder on the datastore called squad_data.
# +
# Select a specific datastore or you can call ws.get_default_datastore()
datastore_name = "workspacefilestore"
ds = ws.datastores[datastore_name]
# Upload files in squad data folder to the datastore
ds.upload(
src_dir=SQUAD_FOLDER, target_path="squad_data", overwrite=True, show_progress=True
)
# -
# ## 3. Prepare Training Script
# Here, we create a simple training script that uses AllenNLP's `train_model_from_file()` function containing the following parameters:
# - parameter_filename (str) : A json parameter file specifying an AllenNLP experiment
# - serialization_dir (str): The directory in which to save results and logs
# - overrides (str): A JSON string that we will use to override values in the input parameter file
# - file_friendly_logging (bool, optional): If True, we make our output more friendly to saved model files
# - recover (bool, optional): If True, we will try to recover a training run from an existing serialization
#
# Our training script parameters are: the location of the data folder, name of the configuration file, and JSON string with any overrides for the configuration file. See the [documentation](https://github.com/allenai/allennlp/blob/9a13ab570025a0c1659986009d2abddb2e652020/allennlp/commands/train.py) on AllenNLP `train_model_from_file()` function for more details.
# +
# %%writefile $PROJECT_FOLDER/train.py
import torch
import argparse
import os
import shutil
from allennlp.common import Params
from allennlp.commands.train import train_model_from_file
def main():
# get command-line arguments
parser = argparse.ArgumentParser()
parser.add_argument('--data_folder', type=str,
help='Folder where data is stored')
parser.add_argument('--config_name', type=str,
help='Name of json configuration file')
parser.add_argument('--overrides', type=str,
help='Override parameters on config file')
args = parser.parse_args()
squad_folder = os.path.join(args.data_folder, "squad_data")
serialization_folder = "./logs" #save to the run logs folder
#delete log file if it already exists
if os.path.isdir(serialization_folder):
shutil.rmtree(serialization_folder)
train_model_from_file(parameter_filename = os.path.join(squad_folder, args.config_name),
overrides = args.overrides,
serialization_dir = serialization_folder,
file_friendly_logging = True,
recover = False)
if __name__ == "__main__":
main()
# -
# ## 4. Create a PyTorch Estimator
#
# AllenNLP is built on PyTorch, so we will use the AzureML SDK's PyTorch estimator to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, see [How to Train Pytorch Models on AzureML](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch). First we set up a .yml file with the necessary dependencies.
# +
myenv = CondaDependencies.create(
conda_packages= CONDA_PACKAGES,
pip_packages= PIP_PACKAGES,
python_version="3.6.8",
)
myenv.add_channel("conda-forge")
myenv.add_channel("pytorch")
conda_env_file_name = "bidafenv.yml"
myenv.save_to_file(PROJECT_FOLDER, conda_env_file_name)
# -
# We next define any parameters in the configuration file that we want to override for this specific training run. We demonstrate overriding the num_epochs parameter to perform 25 epochs (rather than 20 epochs as set in bidaf_config.json). The AllenNLP training function expects that overrides are a JSON string, so we convert our dictionary into a JSON string before passing it in as an argument to our training script.
overrides = {"trainer":{'num_epochs': NUM_EPOCHS}}
overrides = json.dumps(overrides)
# Define the parameters to pass to the training script, the project folder, compute target, conda dependencies file, and the name of the training script. Notice that we set `use_gpu` equal to True.
# +
script_params = {
"--data_folder": ds.as_mount(),
"--config_name": "bidaf_config.json",
"--overrides": overrides,
}
estimator = PyTorch(
source_directory=PROJECT_FOLDER,
script_params=script_params,
compute_target=compute_target,
entry_script="train.py",
use_gpu=True,
conda_dependencies_file="bidafenv.yml",
)
# -
# ## 5. Submit a Job
# Submit the estimator object to run your experiment. Results can be monitored using a Jupyter widget. The widget and run are asynchronous and update every 10-15 seconds until job completion.
run = experiment.submit(estimator)
print(run)
RunDetails(run).show()
#wait for the run to complete before continuing in the notebook
run.wait_for_completion()
# **Cancel the Job**
#
# Interrupting/restarting the Jupyter kernel will not properly cancel the run, which can lead to wasted compute resources. To avoid this, we recommend explicitly canceling a run with the following code:
# +
#run.cancel()
# -
# ## 6. Inspect Results of Run
# AllenNLP's training saves all intermediate and final results to the serialization_dir (defined in train.py). In order to inspect the results as well as use the trained model, we will download the files from the run logs using the `download_files()` command.
run.download_files(prefix="./logs", output_directory=LOGS_FOLDER)
# ### 6.1 Evaluation on SQuAD
# The metrics.json file contains the final metrics. We can load this file and extract the final SQuAD dev set EM score (key is 'best_validation_em'). AllenNLP reports an EM score of 68.3, so depending on the parameters specified in your config file, expect a score in that range.
# +
with open(LOGS_FOLDER+"/logs/metrics.json") as f:
metrics = json.load(f)
sb.glue("validation_EM", metrics["best_validation_em"])
metrics["best_validation_em"]
# -
# ### 6.2 Try the Best Model
# In order to use our model, we need to create an AllenNLP [Predictor](https://github.com/allenai/allennlp/blob/master/allennlp/predictors/predictor.py) object. We instantiate this object from an archive path. An archive comprises a Model and its experimental configuration file. After training a model, the archive is saved to the serialization_dir (whose path is set in train.py).
model = Predictor.from_path(LOGS_FOLDER+"/logs")
# The Predictor object allows us to directly pass in a question and passage (behind the scenes it converts this to Instance objects using the DatasetReader). We define an example passage/question, call the model's `predict()` function, and finally extract the `best_span_str` attribute which contains the answer to our query.
# +
passage = "Machine Comprehension (MC), answering questions about a given context, \
requires modeling complex interactions between the context and the query. Recently,\
attention mechanisms have been successfully extended to MC. Typically these mechanisms\
use attention to summarize the query and context into a single vector, couple \
attentions temporally, and often form a uni-directional attention. In this paper \
we introduce the Bi-Directional Attention Flow (BIDAF) network, a multi-stage \
hierarchical process that represents the context at different levels of granularity \
and uses a bi-directional attention flow mechanism to achieve a query-aware context \
representation without early summarization. Our experimental evaluations show that \
our model achieves the state-of-the-art results in Stanford QA (SQuAD) and \
CNN/DailyMail Cloze Test datasets."
question = "What dataset does BIDAF achieve state-of-the-art results on?"
# -
result = model.predict(question, passage)["best_span_str"]
result
| examples/question_answering/bidaf_aml_deep_dive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/danielsaggau/causal-dyna-fair/blob/master/delayed_impact.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xCgMlCrWqmcB"
# # Delayed Impact
#
# This notebook explores causal inference in the context of decision making with dynamics. As our starting point, we use the lending simulator introduced in [Delayed Impact of Fair Machine Learning](https://arxiv.org/abs/1803.04383).
#
# To connect this simulator with causal inference, we exploit it's representation as a structural causal model introduced in [Causal Modeling for Fairness in Dynamical Systems](https://arxiv.org/abs/1909.09141). All of the interventions we consider, including the "credit bureau" intervention, were introduced by Creager et al. For more details, see
#
# 1. <NAME>, et al. "Causal Modeling for Fairness in Dynamical Systems." International Conference on Machine Learning. 2020.
#
# 2. <NAME>, et al. "Delayed Impact of Fair Machine Learning." International Conference on Machine Learning. 2018.
# + id="fgbT2xlYqmcG"
# %load_ext autoreload
# %autoreload 2
# !pip install whynot
# + id="W7yGrNc8qmcI"
import matplotlib.pylab as plt
from tqdm.auto import tqdm
import whynot as wn
import whynot.traceable_numpy as np
from whynot.simulators.delayed_impact.simulator import INV_CDFS, GROUP_SIZE_RATIO
# + [markdown] id="IEccnXfzqmcJ"
# ## Constructing the experiment
# + [markdown] id="zRo4O5ljqmcJ"
# For illustrative purposes, we show how to construct the credit bureau intervention from [Creager et al.](https://arxiv.org/abs/1909.09141) in WhyNot. The user writes a handful of small functions, each determining a different aspect of the causal experiment.
# + [markdown] id="IO3q7qhMqmcK"
# ### Sample initial states
# Each state in the delayed impact model consists of an agent parameterized by a binary "group" indicator and a credit score.
# We sample group membership and credit scores based on historical FICO data.
# + id="ZPR07tKxqmcL"
def sample_initial_states(rng):
group = int(rng.uniform() < GROUP_SIZE_RATIO[1])
# Compute credit score via inverse CDF trick
score = INV_CDFS[group](rng.uniform())
return wn.delayed_impact.State(group=group, credit_score=score)
# + [markdown] id="bqBtjjNLqmcM"
# ### Set up simulator configuration
# The simulator is parameterized by lending thresholds for each group $\tau_0$ and $\tau_1$, which we instantiate as `parameters` to allow the user to vary them
# during experiments.
# + id="vn7G46wgqmcN"
@wn.parameter(
name="threshold_g0", default=550, description="Lending threshold for group 0")
@wn.parameter(
name="threshold_g1", default=550, description="Lending threshold for group 1")
def construct_config(threshold_g0, threshold_g1):
"""Return the experimental config for runs without intervention"""
return wn.delayed_impact.Config(
start_time=0, end_time=1, threshold_g0=threshold_g0, threshold_g1=threshold_g1
)
# + [markdown] id="DiZarN8hqmcO"
# ### Define outcome measurement
# The outcome measurement we measure is the change in an agent's credit score from time step 0 to time step 1, along with the profit
# the bank earns for this individual.
# + id="6dF8XW-UqmcO"
def extract_outcomes(run):
# Recall states are individuals in this model
agent_t0 = run.states[0]
agent_t1 = run.states[1]
return [agent_t1.credit_score - agent_t0.credit_score, agent_t1.profits]
# + [markdown] id="eJViUqQKqmcP"
# ### Define the intervention
#
# The intervention we consider is the implementation of a credit bureau that treats intermediates between the individual and the lender
# and reports credit scores of $\min(\text{score}, 600)$, as discussed in [Creager et al.](https://arxiv.org/abs/1909.09141).
# + id="Izrh-3WxqmcQ"
def creditscore_threshold(score):
"""Alternate credit bureau scoring policy."""
return max(score, 600)
def intervention():
return wn.delayed_impact.Intervention(credit_scorer=creditscore_threshold, time=0)
# + [markdown] id="u7h-kAI-qmcQ"
# ### Put the components together
# We put each of these components in a `DynamicsExperiment` object to create an the `CreditBureauExperiment`. Importantly, we assume all agents in the model
# have the credit bureau intervention, so they are "treated" in the causal inference sense with probability 1.0
# + id="BIkSLz51qmcR"
CreditBureauExperiment = wn.DynamicsExperiment(
name="CreditBureauExperiment",
description="Intervention on the credit scoring mechanism.",
simulator=wn.delayed_impact,
simulator_config=construct_config,
intervention=intervention,
state_sampler=sample_initial_states,
propensity_scorer=1.0,
outcome_extractor=extract_outcomes,
covariate_builder=lambda run: run.initial_state.group,
)
# + [markdown] id="YwfIq8SvqmcS"
# ## Running the experiment
#
# Compute average credit score changes for the minority group, as well as institutional profits, as the credit score threshold for the minority varies from 300 to 800.
# Throughout, the majority threshold is fixed.
# + colab={"base_uri": "https://localhost:8080/", "height": 65, "referenced_widgets": ["65be138ed04249e1bdaf6eda53fe6282", "4a3801350bb34970955e2fff2e03e9a7", "11bbff26e3404fb7a3ebfab825ad90e3", "cc33b363f92e48c0bb017a4bd5894b58", "9b3e7c1f877e48d48c60a7f200ea8a71", "94717e1b0d1e43b3a4538afdc54bc70c", "6c6c16f4e6014080b95b8e902f2bee29", "365e94f2ee9f436ba1eb32a51e04a14e"]} id="1lziZj-1qmcT" outputId="3a5c1778-83a8-49ce-9ba9-ead3f821308e"
minority_thresholds = list(range(300, 800, 10))
average_min_score_changes = []
average_inst_profits = []
for tau_0 in tqdm(minority_thresholds):
# Run the experiment to generate the dataset
dataset = CreditBureauExperiment.run(threshold_g0=tau_0, num_samples=1000, parallelize=True)
# Only consider score changes for the minority group
minority_locs = dataset.covariates[:, 0] == 0
minority_treated_locs = minority_locs & (dataset.treatments == 1)
score_changes = dataset.outcomes[:, 0]
minority_changes = score_changes[minority_treated_locs]
average_min_score_changes.append(np.mean(minority_changes))
# Report profits over the entire group
ind_profits = dataset.outcomes[:, 1][dataset.treatments == 1]
average_inst_profits.append(np.mean(ind_profits))
# + [markdown] id="Uxd_4DtEqmcW"
# ## Visualizing the results
# + colab={"base_uri": "https://localhost:8080/", "height": 387} id="a8gPYKaqqmcW" outputId="c8690bff-fb09-46c9-b0dc-7f0d5566555e"
_, axs = plt.subplots(1, 2, figsize=(12, 6))
axs[0].plot(minority_thresholds, average_min_score_changes, label="Average Minority\n Score Change")
axs[0].plot([600] * 20, np.linspace(-60, 20, 20), label="Credit Bureau Cutoff", linestyle="--", color="black")
axs[0].legend()
axs[0].set_xlabel("Minority Threshold")
axs[0].set_ylabel("Score Change")
axs[1].plot(minority_thresholds, average_inst_profits, label="Average Profit")
axs[1].plot([600] * 20, np.linspace(0, 0.5, 20), label="Credit Bureau Cutoff", linestyle="--", color="black")
axs[1].set_xlabel("Minority Threshold")
axs[1].set_ylabel("Institutional Profit")
axs[1].legend();
# + [markdown] id="b13fCZ0AqmcX"
# # Treatment effect estimation with confounding
#
# Now, we slightly modify the CreditBureau example to show how WhyNot can also be used to address causal inference questions. Rather than exposing all agents to the credit bureau intervention, we imagine
# members of the minority group are more likely to receive the credit bureau intervention (75% chance) compared to members of the majority group (20% chance). Using this data, we wish to estimate the population-level causal effect of the credit intervention.
#
# + [markdown] id="J2K8K8hlqmcY"
# ## Constructing the experiment
#
# We slightly modify the `CreditBureauExperiment` to include confounded treatment assignment by changing the ``propensity_scorer`` function.
# + id="yJy4D6xpqmcZ"
def propensity_scorer(untreated_run):
"""Assign minority to treatment 75% of the time, compared to 25% for the majority"""
return 0.75 * (untreated_run.initial_state.group == 0) + 0.2 * (untreated_run.initial_state.group == 1)
# + id="tTy7Fm1SqmcZ"
BiasedCreditBureauExperiment = wn.DynamicsExperiment(
name="BiasedCreditBureauExperiment",
description="Intervention on the credit scoring mechanism, with treatment bias.",
simulator=wn.delayed_impact,
simulator_config=construct_config,
intervention=intervention,
state_sampler=sample_initial_states,
propensity_scorer=propensity_scorer,
outcome_extractor=extract_outcomes,
# Only covariate is group membership, which is a confounder for this experiment.
covariate_builder=lambda run: [run.initial_state.group, run.initial_state.credit_score]
)
# + [markdown] id="twFjXvQdqmca"
# ## Running the experiment and generating causal graphs
# + [markdown] id="sOwT69Fxqmcb"
# Run the experiment to generate an observational dataset. Using `causal_graph=True` generates the causal graph associated with the experiment.
# + colab={"base_uri": "https://localhost:8080/", "height": 65, "referenced_widgets": ["f684dab2b98f4949b90b690acbfd8e4b", "85714fb395804d86b764506c2b6cddd0", "f1fa560579704677a836cdd0e0583fd4", "a1e6de0ecfe04b07ace6d8ee58cde4b1", "d6dd75e05a5b4928ad17f1d6f423f29a", "<KEY>", "128c18a7e2df41288ae06ab2f730920c", "65f950634f934eb8ab3c61a5e546f9a5"]} id="Ri850RZTqmcb" outputId="1a1284d5-cdad-4de0-b094-1c4a9ab22ede"
dataset = BiasedCreditBureauExperiment.run(num_samples=1000, causal_graph=True, show_progress=True)
# + [markdown] id="-v6htTf3qmcc"
# ## Estimating treatment effects
# + colab={"base_uri": "https://localhost:8080/"} id="kiDuNDGrqmcd" outputId="76d77b42-897f-418e-a40a-9bf6e12f90aa"
covariates, treatment, outcome = dataset.covariates, dataset.treatments, dataset.outcomes
score_changes = dataset.outcomes[:, 0]
inference_result = wn.algorithms.ols.estimate_treatment_effect(
covariates, treatment, score_changes)
print("Estimated ATE: {:.2f} ({:.2f}, {:.2f})".format(inference_result.ate, *inference_result.ci))
print("True ATE: {:.2f}".format(np.mean(dataset.true_effects[:, 0])))
# + [markdown] id="Kcb2idFnqmcd"
# ## Inspecting the causal graph
#
# The causal graph is a `networkx.Digraph` that can easily be connected to graphical methods for estimating treatment effects, e.g. [DoWhy](https://github.com/microsoft/dowhy).
# For a complete example, see [here](https://github.com/zykls/whynot/blob/master/examples/causal_inference/graphical_methods.ipynb).
# + colab={"base_uri": "https://localhost:8080/"} id="kGtI5AZqqmce" outputId="bbb3ade8-65e5-4cd2-9b83-385a504f7882"
graph = dataset.causal_graph
print("## NODES ##")
for node in graph.nodes:
print(node)
print("\n## EDGES ##\n")
for edge in graph.edges:
print(edge)
| delayed_impact.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# ## Load dataset
dataset = sns.load_dataset("anscombe")
# ## Show samples
dataset.head()
# ## Plot data
subsets = ['I', 'II', 'III', 'IV']
plt.scatter(dataset.x, dataset.y, c=dataset['dataset'].apply(subsets.index), cmap='Accent')
# ## Some statistics
pd.DataFrame([np.mean(dataset[dataset['dataset']==ds].iloc[:, 1:], axis = 0) for ds in subsets])
pd.DataFrame([np.var(dataset[dataset['dataset']==ds].iloc[:, 1:], axis = 0) for ds in subsets])
# ## Linear Regression
regplot = lambda ds: sns.regplot(dataset[dataset['dataset']==ds].x, dataset[dataset['dataset']==ds].y)
[regplot(ds) for ds in subsets]
regplot('I')
regplot('II')
regplot('III')
regplot('IV')
| ai2hw1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# graded = 9/9
#
#
# # Homework assignment #3
#
# These problem sets focus on using the Beautiful Soup library to scrape web pages.
#
# ## Problem Set #1: Basic scraping
#
# I've made a web page for you to scrape. It's available [here](http://static.decontextualize.com/widgets2016.html). The page concerns the catalog of a famous [widget](http://en.wikipedia.org/wiki/Widget) company. You'll be answering several questions about this web page. In the cell below, I've written some code so that you end up with a variable called `html_str` that contains the HTML source code of the page, and a variable `document` that stores a Beautiful Soup object.
from bs4 import BeautifulSoup
from urllib.request import urlopen
html_str = urlopen("http://static.decontextualize.com/widgets2016.html").read()
document = BeautifulSoup(html_str, "html.parser")
# Now, in the cell below, use Beautiful Soup to write an expression that evaluates to the number of `<h3>` tags contained in `widgets2016.html`.
# +
h3_tags = document.find_all('h3')
h3_tags_count = 0
for tag in h3_tags:
h3_tags_count = h3_tags_count + 1
print(h3_tags_count)
# -
# Now, in the cell below, write an expression or series of statements that displays the telephone number beneath the "Widget Catalog" header.
# +
#inspecting webpace with help of developer tools -- shows infomation is stored in an a tag that has the class 'tel'
a_tags = document.find_all('a', {'class':'tel'})
for tag in a_tags:
print(tag.string)
#Does not return the same: [tag.string for tag in a_tags]
# -
# In the cell below, use Beautiful Soup to write some code that prints the names of all the widgets on the page. After your code has executed, `widget_names` should evaluate to a list that looks like this (though not necessarily in this order):
#
# ```
# Skinner Widget
# Widget For Furtiveness
# Widget For Strawman
# Jittery Widget
# Silver Widget
# Divided Widget
# Manicurist Widget
# Infinite Widget
# Yellow-Tipped Widget
# Unshakable Widget
# Self-Knowledge Widget
# Widget For Cinema
# ```
# +
search_table = document.find_all('table',{'class': 'widgetlist'})
#print(search_table)
tables_content = [table('td', {'class':'wname'}) for table in search_table]
#print(tables_content)
for table in tables_content:
for single_table in table:
print(single_table.string)
# -
# ## Problem set #2: Widget dictionaries
#
# For this problem set, we'll continue to use the HTML page from the previous problem set. In the cell below, I've made an empty list and assigned it to a variable called `widgets`. Write code that populates this list with dictionaries, one dictionary per widget in the source file. The keys of each dictionary should be `partno`, `wname`, `price`, and `quantity`, and the value for each of the keys should be the value for the corresponding column for each row. After executing the cell, your list should look something like this:
#
# ```
# [{'partno': 'C1-9476',
# 'price': '$2.70',
# 'quantity': u'512',
# 'wname': 'Skinner Widget'},
# {'partno': 'JDJ-32/V',
# 'price': '$9.36',
# 'quantity': '967',
# 'wname': u'Widget For Furtiveness'},
# ...several items omitted...
# {'partno': '5B-941/F',
# 'price': '$13.26',
# 'quantity': '919',
# 'wname': 'Widget For Cinema'}]
# ```
#
# And this expression:
#
# widgets[5]['partno']
#
# ... should evaluate to:
#
# LH-74/O
#
# +
widgets = []
#STEP 1: Find all tr tags, because that's what tds are grouped by
for tr_tags in document.find_all('tr', {'class': 'winfo'}):
#STEP 2: For each tr_tag in tr_tags, make a dict of its td
tr_dict ={}
for td_tags in tr_tags.find_all('td'):
td_tags_class = td_tags['class']
for tag in td_tags_class:
tr_dict[tag] = td_tags.string
#STEP3: add dicts to list
widgets.append(tr_dict)
widgets
#widgets[5]['partno']
# -
# In the cell below, duplicate your code from the previous question. Modify the code to ensure that the values for `price` and `quantity` in each dictionary are floating-point numbers and integers, respectively. I.e., after executing the cell, your code should display something like this:
#
# [{'partno': 'C1-9476',
# 'price': 2.7,
# 'quantity': 512,
# 'widgetname': 'Skinner Widget'},
# {'partno': 'JDJ-32/V',
# 'price': 9.36,
# 'quantity': 967,
# 'widgetname': 'Widget For Furtiveness'},
# ... some items omitted ...
# {'partno': '5B-941/F',
# 'price': 13.26,
# 'quantity': 919,
# 'widgetname': 'Widget For Cinema'}]
#
# (Hint: Use the `float()` and `int()` functions. You may need to use string slices to convert the `price` field to a floating-point number.)
#had to rename variables as it kept printing the ones from the cell above...
widgetsN = []
for trN_tags in document.find_all('tr', {'class': 'winfo'}):
trN_dict ={}
for tdN_tags in trN_tags.find_all('td'):
tdN_tags_class = tdN_tags['class']
for tagN in tdN_tags_class:
if tagN == 'price':
sliced_tag_string = tdN_tags.string[1:]
trN_dict[tagN] = float(sliced_tag_string)
elif tagN == 'quantity':
trN_dict[tagN] = int(tdN_tags.string)
else:
trN_dict[tagN] = tdN_tags.string
widgetsN.append(trN_dict)
widgetsN
# Great! I hope you're having fun. In the cell below, write an expression or series of statements that uses the `widgets` list created in the cell above to calculate the total number of widgets that the factory has in its warehouse.
#
# Expected output: `7928`
widget_quantity_list = [element['quantity'] for element in widgetsN]
sum(widget_quantity_list)
# In the cell below, write some Python code that prints the names of widgets whose price is above $9.30.
#
# Expected output:
#
# ```
# Widget For Furtiveness
# Jittery Widget
# Silver Widget
# Infinite Widget
# Widget For Cinema
# ```
for widget in widgetsN:
if widget['price'] > 9.30:
print(widget['wname'])
# ## Problem set #3: Sibling rivalries
#
# In the following problem set, you will yet again be working with the data in `widgets2016.html`. In order to accomplish the tasks in this problem set, you'll need to learn about Beautiful Soup's `.find_next_sibling()` method. Here's some information about that method, cribbed from the notes:
#
# Often, the tags we're looking for don't have a distinguishing characteristic, like a class attribute, that allows us to find them using `.find()` and `.find_all()`, and the tags also aren't in a parent-child relationship. This can be tricky! For example, take the following HTML snippet, (which I've assigned to a string called `example_html`):
example_html = """
<h2>Camembert</h2>
<p>A soft cheese made in the Camembert region of France.</p>
<h2>Cheddar</h2>
<p>A yellow cheese made in the Cheddar region of... France, probably, idk whatevs.</p>
"""
# If our task was to create a dictionary that maps the name of the cheese to the description that follows in the `<p>` tag directly afterward, we'd be out of luck. Fortunately, Beautiful Soup has a `.find_next_sibling()` method, which allows us to search for the next tag that is a sibling of the tag you're calling it on (i.e., the two tags share a parent), that also matches particular criteria. So, for example, to accomplish the task outlined above:
# +
example_doc = BeautifulSoup(example_html, "html.parser")
cheese_dict = {}
for h2_tag in example_doc.find_all('h2'):
cheese_name = h2_tag.string
cheese_desc_tag = h2_tag.find_next_sibling('p')
cheese_dict[cheese_name] = cheese_desc_tag.string
cheese_dict
# -
# With that knowledge in mind, let's go back to our widgets. In the cell below, write code that uses Beautiful Soup, and in particular the `.find_next_sibling()` method, to print the part numbers of the widgets that are in the table *just beneath* the header "Hallowed Widgets."
#
# Expected output:
#
# ```
# MZ-556/B
# QV-730
# T1-9731
# 5B-941/F
# ```
for h3_tags in document.find_all('h3'):
if h3_tags.string == 'Hallowed widgets':
hallowed_table = h3_tags.find_next_sibling('table')
for element in hallowed_table.find_all('td', {'class':'partno'}):
print(element.string)
# Okay, now, the final task. If you can accomplish this, you are truly an expert web scraper. I'll have little web scraper certificates made up and I'll give you one, if you manage to do this thing. And I know you can do it!
#
# In the cell below, I've created a variable `category_counts` and assigned to it an empty dictionary. Write code to populate this dictionary so that its keys are "categories" of widgets (e.g., the contents of the `<h3>` tags on the page: "Forensic Widgets", "Mood widgets", "Hallowed Widgets") and the value for each key is the number of widgets that occur in that category. I.e., after your code has been executed, the dictionary `category_counts` should look like this:
#
# ```
# {'Forensic Widgets': 3,
# 'Hallowed widgets': 4,
# 'Mood widgets': 2,
# 'Wondrous widgets': 3}
# ```
# +
category_counts = {}
for x_tags in document.find_all('h3'):
x_table = x_tags.find_next_sibling('table')
tr_info_tags = x_table.find_all('tr', {'class':'winfo'})
category_counts[x_tags.string] = len(tr_info_tags)
category_counts
# -
# Congratulations! You're done.
| data-databases-homework/Homework_3_Gruen_graded.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
EXPORT_PATH = r"/home/aadc/share/images/"
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from litdrive.zeromq.server import ZmqServer
IMAGE_HEIGHT = 960
IMAGE_WIDTH = 1280
# %matplotlib notebook
# -
def process(image):
a = np.array(image)
mpl.image.imsave('/home/aadc/share/images/pic.png', a)
print(image)
view.set_data(image)
plt.draw()
plt.pause(.1)
#raise IOError
# open a server for the filter
zmq = ZmqServer("tcp://*:5555", [("front", IMAGE_HEIGHT, IMAGE_WIDTH)])
# +
# get an empty interactive view for the image
plt.ion()
plt.figure(figsize=(10, 10))
view = plt.imshow(np.ones((IMAGE_HEIGHT, IMAGE_WIDTH, 3)))
try:
zmq.connect()
zmq.run(process)
finally:
zmq.disconnect()
# -
| src/aadcUserPython/litdrive/camera_stream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: repo
# language: python
# name: repo
# ---
# # $\newcommand{\MODp}{\texttt{MOD}_{p}}\newcommand{\MOD}[1]{\texttt{MOD}_{#1}}$2-State $\texttt{MOD}_{p}$ Automata
# The most basic type of automata which can recognise the $\MODp$ language is a 2-State automaton. A 2-state automaton testing a 3 symbol word against $\MOD{7}$ is given in the figure below.
#
# <center><img src="./figures/fig1.svg" width="500px"/></center>
#
# A two state automaton can be created via `qfa.TwoState`. The circuit above is constructed as an example
from qfa import TwoState
TwoState(p=7, word_length=3).draw(plot_barriers=False)
| notebooks/01-two-state.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/The-ML-Hero/DeepLearningBasedDescriptivePaperCorrection/blob/main/DeepLearningBasedDescriptivePaperCorrection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="D8vs-LUjs-_G" outputId="5dc174f8-ae0d-4175-dd07-5a6c248f3fc4"
# !pip install pdf2image
# + colab={"base_uri": "https://localhost:8080/"} id="PDu0nIUzu2Nb" outputId="20e2de43-1c84-4e24-e8ac-2ba4e53bdf35"
# !pip install -U sentence-transformers
# + colab={"base_uri": "https://localhost:8080/"} id="8o10hyzuxZS0" outputId="7e79a0c0-2410-4ab3-f807-b53e3101247e"
!!apt-get install poppler-utils
# + id="BKip3KRyzYOI"
import os
import tempfile
from pdf2image import convert_from_path
from PIL import Image
def convert_pdf(file_path, output_path):
# save temp image files in temp dir, delete them after we are finished
with tempfile.TemporaryDirectory() as temp_dir:
# convert pdf to multiple image
images = convert_from_path(file_path, output_folder=temp_dir)
# save images to temporary directory
temp_images = []
for i in range(len(images)):
image_path = f'{temp_dir}/{i}.jpg'
images[i].save(image_path, 'JPEG')
temp_images.append(image_path)
# read images into pillow.Image
imgs = list(map(Image.open, temp_images))
# find minimum width of images
min_img_width = min(i.width for i in imgs)
# find total height of all images
total_height = 0
for i, img in enumerate(imgs):
total_height += imgs[i].height
# create new image object with width and total height
merged_image = Image.new(imgs[0].mode, (min_img_width, total_height))
# paste images together one by one
y = 0
for img in imgs:
merged_image.paste(img, (0, y))
y += img.height
# save merged image
merged_image.save(output_path)
return output_path
# + colab={"base_uri": "https://localhost:8080/", "height": 918} id="xQWFFnJz1IuH" outputId="2169c3b3-31a6-478a-f903-854fe717108a"
i = 0
for files in os.listdir('/content/drive/MyDrive/Essay'):
i += 1
convert_pdf(f'/content/drive/MyDrive/Essay/{files}',f'/content/EassyPngs/Multiple{i}.png')
print(f'Wrote {i} File')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="my2Fx-UIzbIM" outputId="8b0b5064-2ba4-4b0a-81c8-f265648bc531"
convert_pdf('/content/drive/MyDrive/Essay/2021_01_06 7_14 pm Office Lens.pdf','mutiplte.png')
# + colab={"base_uri": "https://localhost:8080/"} id="j0UIiKAwu38b" outputId="0703777f-9070-4362-9970-cb09e71e79ec"
from sentence_transformers import SentenceTransformer
model = SentenceTransformer('paraphrase-distilroberta-base-v1')
# + colab={"base_uri": "https://localhost:8080/"} id="idxek6wmzs6A" outputId="6dc071fe-41c8-4292-c048-ec0f75412e62"
# !pip install pyngrok
# + id="EX-m89VTBVCz"
import contextualSpellCheck
import spacy
## We require NER to identify if it is PERSON
## also require parser because we use Token.sent for context
nlp = spacy.load("en_core_web_sm")
contextualSpellCheck.add_to_pipe(nlp)
doc = nlp('I was playin guitar in my garden')
doc._.outcome_spellCheck
# + colab={"base_uri": "https://localhost:8080/"} id="slYX9ov1u7Q4" outputId="368bc7e9-6144-4f65-d9ca-a0c8b5205241"
from sentence_transformers import SentenceTransformer, util
model = SentenceTransformer('stsb-distilbert-base')
# Two lists of sentences
sentences1 = """The process of formation of hybrid layer is hybridization
This layer forms following initial demineralization of dental sujace with an
acidic conditioner. Exposing a collagens fibril microporosites that subsequently become diffures with low viscosity monomers. This one in
whiten resin adhesive system. interlocks micromechanically with dental collagenic termed hybrid layer Cor sone
Base layer of hybrid Gone is characterised by a gradual transition to underlying altered deutin.
Hybrid layer as stres breaken (or) stress reliever with young mode of 3 Gpa
Top layer of hybrid gone is amorphous electron dense phase."""
sentences2 = """COMPOSITES: THE PAST, PRESENT AND THE FUTURE
INTRODUCTION
Restorative dental materials date back as far as dentistry. When all or part of a tooth is
missing, replacing it with something is important for the patient to regain full function, not to
mention esthetics. The dental materials of the past were rarely tooth colored, so dental
restorations stood out in a patient’s mouth regardless of the clinician’s skill. Thus, the
invention almost 60 years back, these aesthetic materials have come a long way and have
witnessed lot of changes in their development. Dental composites are technique sensitive to
use, while producing highly functional, and highly esthetic outcomes.
DISCUSSION OF PROPERTIES:
The Past
The journey started way back in early 1950s with discovery of “Acid Etching” by <NAME>.
Buonocore and then successive developments of resin monomers bisphenol A-glycidyl
methacrylate (BisGMA) by <NAME>. The primitive composites though being crude
had been an attempt to have a material which resembles tooth structure. By composition they
had resin matrix of BisGMA and UDMA (urethane dimethacrylate) monomers bonded with
silica fillers ranging from 50 micron to 0.5 mm using a silane coupling agent.
RESIN MATRIX: contains BIS-GMA, TEGDMA, UDMA
• Addition of TEGDMA enables the addition of sufficient filler but TEGDMA addition
can result in shrinkage stress.
• Modifications of bisphenol-A-based dimethacrylate systems have included the use of
pendant bulky (aromatic) constituents (Ge et al., 2005) as well as pendant alkyl.
• Recent developments regarding public perceptions of bisphenol-A toxicity may have
a strong influence on steering future monomer development efforts toward bisphenol-A
alternatives.
Fillers– Quartz or glasses, amorphous silica {0.1 – 100 µm},30- 70 vol % or 50 –85%
weight. In chronological developments the filler particle size was reduced from macro filler
to micro fillers where the size of particles was just few micro meters and then came the
hybrids a combination of micro and mini fillers; thus, improvising the strength and handling
properties. Initially to increase the filler ratio fumed silica was being added in different forms
like prepolymerized or agglomerated and sintered agglomerated particles.
The present and future
With the advent of nano era the filler particle size went down to as low as few nanometers
thereby enabling us to have very high filler loading for use in both anterior and posterior
region.Thus it got easy to develop exclusive posterior composites having very high
compressive strength and low wear rate. Addition of fillers increased compressive strength,
tensile strength, stiffness, abrasion resistance, hardness reduced wear, polymerization
shrinkage, thermal expansion and contraction, water sorption.
Polymerization shrinkage - During the curing of composites as the monomer used to get
converted to polymer there was a high stress buildup within the restoration thereby pulling it
from the tooth surface and causing post-operative sensitivity, secondary caries and marginal
discoloration and often but not the least fracture of the restoration. It was documented to be
as high as approximately 10 percent. Recent modifications with the introduction of epoxy
based silorane system which claimed to reduce the shrinkage stress by opening of an oxirane
ring during the epoxide curing reaction.
SHADING MATCHING IN COMPOSITE RESTORATION
The shade matching can be done two ways:
1. Manual, Visual Shade Selection Techniques
2. Automatic, Instrumental Shade Selection Techniques
1.Manual, Visual Shade Selection Techniques:
The most frequently used technique in the shade matching of teeth to restorative materials is
done manually and visually with dental shade guides.
Recommendations for selection:
• Teeth to be matched must be clean.
• Remove bright colors from the field of view.
• Tooth shade should be determined in daylight or under standardized daylight lamps (not
operation lamps).
• View at the patient eye level.
• Evaluate shade under multiple light sources.
• Make shade matching at the beginning of treatment before the teeth begin to dehydrate.
• Shade matching should be made quickly to avoid eye fatigue (5-7) seconds. The observer
can look at a blue or grey card to rest eyes.
The selected shades may be verified by placing a small unetched button of composite on the
buccal surface, light curing for 5 seconds, then comparing it to the natural cervical and
occlusal hues. The composite buttons can then be removed with an explorer tip.
2.Automatic, Instrumental Shade Selection Techniques:
These shade matching instruments were introduced to the dental profession to overcome the
limitations and inconsistencies of the manual, visual shade matching systems.
• Colorimeters
• Spectrophotometers
• digital imaging devices.
MANIPULATION OF COMPOSITES:
Rubber dam isolation is critical to prevent moisture interference or contamination with the
intricate adhesive process.
The Class II posterior direct resin restoration is the most challenging due to the operative
intricacy of the proximal contact.
First appropriate metal matrices or proximal contact formers should be selected.
When using a matrix band, the marginal ridge heights of the adjacent teeth must be observed.
Overcontouring of the marginal ridges will result in subsequent over contouring of the entire
restoration.
Once the matrix band is properly placed, the adhesive process can be initiated and completed
according to manufacturer’s specifications. Initiation of the incremental buildup begins with
the application of a flowable resin to the base of the preparation. An explorer tip should be
used to manipulate a thin layer evenly across the pulpal floor and proximal walls.
Additionally, flowable resin can be drawn along the margins of the proximal box and light
cured.
The general anatomy and morphology of the final restoration is reflected in the primary
anatomy of the dentinal resin. Any characterization, tints, or fossa colors the patient desires
can be added after the dentin buildup. Microcannula tips can be used to place color in precise
amounts.
Any additional polishing can be performed with a narrow fine diamond abrasive strip in the
embrasure area. The 2-mm-wide strip can be passed through the contact area apical to the
gingival margin.
Composite polishing cups and points are used to polish the previously adjusted areas only,
using light, intermittent touches to prevent loss of anatomy and surface morphology.
SURVIVABILITY OF COMPOSITE RESTORATION:
The overall findings suggest that at least 60% of resin composite restorations will last more
than 10 years when proper materials are applied correctly.
The mean annual failure rate is 2.9% for resin‐composite restorations and 1.6% for
amalgams. For resin‐composite restorations, secondary caries was the most common reason
for replacement (73.9%), followed by loss (8.0%), fracture (5.3%), and marginal defects
(2.4%).
Studies have shown that young age of the patient, high previous caries experience, deep
cavities, and saucer‐shaped preparation technique as predisposing to shorter longevity of
resin‐composite restorations.
CONCLUSION
It is emphasized here that methods be developed to completely cure the resin monomer and at
same time to have closer sintering of filler particles thus increasing the strength of the
material and ensuring its longevity in the oral cavity of the patient."""
def remove_ascii(input):
return ''.join([i if ord(i) < 128 else ' ' for i in input])
sentences2 = remove_ascii(sentences2)
#Compute embedding for both lists
embeddings1 = model.encode(sentences1, convert_to_tensor=True)
embeddings2 = model.encode(sentences2, convert_to_tensor=True)
#Compute cosine-similarits
cosine_scores = util.pytorch_cos_sim(embeddings1, embeddings2)
cosine_scores[0][0]
# + id="Xlm8Xknv7Kz9"
sentences2 = """COMPOSITES: THE PAST, PRESENT AND THE FUTURE
INTRODUCTION
Restorative dental materials date back as far as dentistry. When all or part of a tooth is
missing, replacing it with something is important for the patient to regain full function, not to
mention esthetics. The dental materials of the past were rarely tooth colored, so dental
restorations stood out in a patient’s mouth regardless of the clinician’s skill. Thus, the
invention almost 60 years back, these aesthetic materials have come a long way and have
witnessed lot of changes in their development. Dental composites are technique sensitive to
use, while producing highly functional, and highly esthetic outcomes.
DISCUSSION OF PROPERTIES:
The Past
The journey started way back in early 1950s with discovery of “Acid Etching” by <NAME>.
Buonocore and then successive developments of resin monomers bisphenol A-glycidyl
methacrylate (BisGMA) by <NAME>. The primitive composites though being crude
had been an attempt to have a material which resembles tooth structure. By composition they
had resin matrix of BisGMA and UDMA (urethane dimethacrylate) monomers bonded with
silica fillers ranging from 50 micron to 0.5 mm using a silane coupling agent.
RESIN MATRIX: contains BIS-GMA, TEGDMA, UDMA
• Addition of TEGDMA enables the addition of sufficient filler but TEGDMA addition
can result in shrinkage stress.
• Modifications of bisphenol-A-based dimethacrylate systems have included the use of
pendant bulky (aromatic) constituents (Ge et al., 2005) as well as pendant alkyl.
• Recent developments regarding public perceptions of bisphenol-A toxicity may have
a strong influence on steering future monomer development efforts toward bisphenol-A
alternatives.
Fillers– Quartz or glasses, amorphous silica {0.1 – 100 µm},30- 70 vol % or 50 –85%
weight. In chronological developments the filler particle size was reduced from macro filler
to micro fillers where the size of particles was just few micro meters and then came the
hybrids a combination of micro and mini fillers; thus, improvising the strength and handling
properties. Initially to increase the filler ratio fumed silica was being added in different forms
like prepolymerized or agglomerated and sintered agglomerated particles.
The present and future
With the advent of nano era the filler particle size went down to as low as few nanometers
thereby enabling us to have very high filler loading for use in both anterior and posterior
region.Thus it got easy to develop exclusive posterior composites having very high
compressive strength and low wear rate. Addition of fillers increased compressive strength,
tensile strength, stiffness, abrasion resistance, hardness reduced wear, polymerization
shrinkage, thermal expansion and contraction, water sorption.
Polymerization shrinkage - During the curing of composites as the monomer used to get
converted to polymer there was a high stress buildup within the restoration thereby pulling it
from the tooth surface and causing post-operative sensitivity, secondary caries and marginal
discoloration and often but not the least fracture of the restoration. It was documented to be
as high as approximately 10 percent. Recent modifications with the introduction of epoxy
based silorane system which claimed to reduce the shrinkage stress by opening of an oxirane
ring during the epoxide curing reaction.
SHADING MATCHING IN COMPOSITE RESTORATION
The shade matching can be done two ways:
1. Manual, Visual Shade Selection Techniques
2. Automatic, Instrumental Shade Selection Techniques
1.Manual, Visual Shade Selection Techniques:
The most frequently used technique in the shade matching of teeth to restorative materials is
done manually and visually with dental shade guides.
Recommendations for selection:
• Teeth to be matched must be clean.
• Remove bright colors from the field of view.
• Tooth shade should be determined in daylight or under standardized daylight lamps (not
operation lamps).
• View at the patient eye level.
• Evaluate shade under multiple light sources.
• Make shade matching at the beginning of treatment before the teeth begin to dehydrate.
• Shade matching should be made quickly to avoid eye fatigue (5-7) seconds. The observer
can look at a blue or grey card to rest eyes.
The selected shades may be verified by placing a small unetched button of composite on the
buccal surface, light curing for 5 seconds, then comparing it to the natural cervical and
occlusal hues. The composite buttons can then be removed with an explorer tip.
2.Automatic, Instrumental Shade Selection Techniques:
These shade matching instruments were introduced to the dental profession to overcome the
limitations and inconsistencies of the manual, visual shade matching systems.
• Colorimeters
• Spectrophotometers
• digital imaging devices.
MANIPULATION OF COMPOSITES:
Rubber dam isolation is critical to prevent moisture interference or contamination with the
intricate adhesive process.
The Class II posterior direct resin restoration is the most challenging due to the operative
intricacy of the proximal contact.
First appropriate metal matrices or proximal contact formers should be selected.
When using a matrix band, the marginal ridge heights of the adjacent teeth must be observed.
Overcontouring of the marginal ridges will result in subsequent over contouring of the entire
restoration.
Once the matrix band is properly placed, the adhesive process can be initiated and completed
according to manufacturer’s specifications. Initiation of the incremental buildup begins with
the application of a flowable resin to the base of the preparation. An explorer tip should be
used to manipulate a thin layer evenly across the pulpal floor and proximal walls.
Additionally, flowable resin can be drawn along the margins of the proximal box and light
cured.
The general anatomy and morphology of the final restoration is reflected in the primary
anatomy of the dentinal resin. Any characterization, tints, or fossa colors the patient desires
can be added after the dentin buildup. Microcannula tips can be used to place color in precise
amounts.
Any additional polishing can be performed with a narrow fine diamond abrasive strip in the
embrasure area. The 2-mm-wide strip can be passed through the contact area apical to the
gingival margin.
Composite polishing cups and points are used to polish the previously adjusted areas only,
using light, intermittent touches to prevent loss of anatomy and surface morphology.
SURVIVABILITY OF COMPOSITE RESTORATION:
The overall findings suggest that at least 60% of resin composite restorations will last more
than 10 years when proper materials are applied correctly.
The mean annual failure rate is 2.9% for resin‐composite restorations and 1.6% for
amalgams. For resin‐composite restorations, secondary caries was the most common reason
for replacement (73.9%), followed by loss (8.0%), fracture (5.3%), and marginal defects
(2.4%).
Studies have shown that young age of the patient, high previous caries experience, deep
cavities, and saucer‐shaped preparation technique as predisposing to shorter longevity of
resin‐composite restorations.
CONCLUSION
It is emphasized here that methods be developed to completely cure the resin monomer and at
same time to have closer sintering of filler particles thus increasing the strength of the
material and ensuring its longevity in the oral cavity of the patient."""
# + colab={"base_uri": "https://localhost:8080/", "height": 171} id="aqmUH4BA8KG_" outputId="0066496a-881b-4294-91ec-f81a168bb8bc"
# + id="bGGp7kPJ0hCe"
"""HYBRID LAYER:
Definition:
The structure formed in the dental hard tissue by demineralization of the surface and the sub-
surface followed by infiltration of monomers and subsequent polymerization. – Nakabayashi, 1982
It is a process which creates a molecular level interface between dentin and composite resin.
Zones of hybrid layer:
Top Layer: Loosely arranged collagen fibrils directed towards adhesive resin.
Middle layer: Collagen fibrils separated by electrolucent spaces represent areas in which
hydroxyapatite crystals have been replaced by resin due to hybridization
Base layer: Partially demineralised dentin.
The formation of hybrid layer is an integral part of dentin bonding. The quality of hybrid layer formed
decides the strength of resin dentin interface. The thicker and more uniform the hybrid layer, better is
the bond strength.
Hybrid layer formation:
A dentin bonding agent is a low viscosity unfilled or semifilled resin for easy penetration and
formation of hybrid layer. When bonding agent is applied, part of it penetrates into the collagen
network, known as intertubular penetration and rest of it penetrates into dentinal tubules called
intertubular penetration. In intertubular penetration, it polymerises with primar monomers forming a
hybrid layer/ reinforced layer.
Tubule wall hybridization:
The extension of the hybrid wall into tubule area. It allows hermetically sealing the pulp
dentinal complex against microleakage.
Lateral Tubule hybridization:
Formation of tiny hybrid layer into the walls of lateral tubule branches.
Reverse hybrid layer:
The formation of reverse hybrid layer by application of NaOCl after acid etching. This
procedure removes the exposed collagen and solubilises the fibrils down into underlying mineral
matrix to create microporosities within the mineral phase.
Ghost hybrid layer:
Aluminum oxide air abrasion resulted in partial removal of original hybrid layer, followed by
the formation of new ghost like hybrid layer. It also modifies the adhesive layer interface."""
# + id="GZglvv9_u9rq"
# !pip install contextualSpellCheck
# + colab={"base_uri": "https://localhost:8080/", "height": 92} id="ksrJ5A2UyvIZ" outputId="e0972c08-227e-44fa-b351-903844f7b43d"
import contextualSpellCheck
import spacy
## We require NER to identify if it is PERSON
## also require parser because we use Token.sent for context
nlp = spacy.load("en_core_web_sm")
contextualSpellCheck.add_to_pipe(nlp)
doc = nlp('I was playin guitar in my garden')
doc._.outcome_spellCheck
# + colab={"base_uri": "https://localhost:8080/"} id="AF8BHU6ly47W" outputId="856b27a4-747e-42b4-9acc-e0bd5ee710a5"
# !ngrok authtoken 1goWTmF64bbpZ7lUDQwXboXtK9o_4rV1HnTNwsQYmxFyPNGJu
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="T34IMd6d2FF5" outputId="e130d42f-4777-420b-f3a5-c9b178631a6e"
# !pip install streamlit
# + id="HZDTSrqwdt6d"
import os
# + id="ZnRnh2PSduuI"
import warnings
# + id="xOf7ZfGmned-"
# !export PYTHONWARNINGS="ignore"
# + colab={"base_uri": "https://localhost:8080/"} id="7fYrSTLQ4Q1Y" outputId="bd4031c6-69c5-4cbb-e2ea-eb599cff185f"
# !pip install contextualSpellCheck
# + colab={"base_uri": "https://localhost:8080/"} id="hoVIwriv2QQI" outputId="7a566a4c-d11a-44bf-e894-ae0a543b6658"
#from __future__ import print_function
#import streamlit as st
from PIL import Image
from difflib import SequenceMatcher as sm
import time
from time import sleep
import os
import contextualSpellCheck
import spacy
import cv2
import numpy as np
import httplib2
import io
from apiclient import discovery
from oauth2client import client
from oauth2client import tools
from oauth2client.file import Storage
from apiclient.http import MediaFileUpload, MediaIoBaseDownload
from sentence_transformers import SentenceTransformer, util
import random
import pandas as pd
start = time.time()
# stsb-roberta-large
epsilon = 15
model = SentenceTransformer('stsb-distilbert-base')
nlp = spacy.load("en_core_web_sm")
contextualSpellCheck.add_to_pipe(nlp)
SCOPES = 'https://www.googleapis.com/auth/drive'
CLIENT_SECRET_FILE = 'client_secrets.json'
APPLICATION_NAME = 'Drive API Python Quickstart'
def get_credentials():
"""Gets valid user credentials from storage.
If nothing has been stored, or if the stored credentials are invalid,
the OAuth2 flow is completed to obtain the new credentials.
Returns:
Credentials, the obtained credential.
"""
credential_path = os.path.join("./", 'drive-python-quickstart.json')
store = Storage(credential_path)
credentials = store.get()
if not credentials or credentials.invalid:
flow = client.flow_from_clientsecrets(CLIENT_SECRET_FILE, SCOPES)
flow.user_agent = APPLICATION_NAME
if flags:
credentials = tools.run_flow(flow, store, flags)
else: # Needed only for compatibility with Python 2.6
credentials = tools.run(flow, store)
print('Storing credentials to ' + credential_path)
return credentials
def main(imgfile):
credentials = get_credentials()
http = credentials.authorize(httplib2.Http())
service = discovery.build('drive', 'v3', http=http)
# Image with texts (png, jpg, bmp, gif, pdf)
imgfile = imgfile
txtfile = 'output.txt' # Text file outputted by OCR
mime = 'application/vnd.google-apps.document'
res = service.files().create(
body={
'name': imgfile,
'mimeType': mime
},
media_body=MediaFileUpload(imgfile, mimetype=mime, resumable=True)
).execute()
downloader = MediaIoBaseDownload(
io.FileIO(txtfile, 'wb'),
service.files().export_media(fileId=res['id'], mimeType="text/plain")
)
done = False
while done is False:
status, done = downloader.next_chunk()
service.files().delete(fileId=res['id']).execute()
print("Done.")
MAIN_DIR = '/content/LowMark'
for files in os.listdir(MAIN_DIR):
if files != ".ipynb_checkpoints":
print("Getting List of Files")
main(f'{MAIN_DIR}/{files}')
print("Recognizing Text")
def open_context():
context = open("output.txt", "r").read()
return ''.join([i if ord(i) < 128 else ' ' for i in context]).replace('_','').replace('DOO','')
context = open_context()
#context = nlp(context)
# context_suggest = context._.suggestions_spellCheck
# if context_suggest != {}:
# print(context)
# context = str(context._.outcome_spellCheck)
print("Preprocessing the Output")
# print(context)
ans = sentences2
print("Getting the Embeddings")
RealAnsEmbeddings = model.encode(ans, convert_to_tensor=True)
AnsEmbeddings = model.encode(context, convert_to_tensor=True)
print("Calculating the Marks...")
cosine_scores = util.pytorch_cos_sim(AnsEmbeddings, RealAnsEmbeddings)
cosine_scores = float(cosine_scores[0][0])
cosine_scores *= 100
cosine_scores = int(cosine_scores)
def final_mark(score):
if score + epsilon >= 98:
return 95
elif score <= 70:
return score + random.randint(11,15)
else:
return score + epsilon
# def marks(scr):
# if scr <= 80:
# return scr + random.randint(7,9)
# else:
# return scr
# def marks_final_pass(scr):
# if scr <= 65:
# return scr + random.randint(15,17)
# elif scr <= 80:
# return scr + random.randint(7,9)
# else:
# return scr
if cosine_scores <= 58:
cosine_scores_final = final_mark(cosine_scores)
print(f'The Mark For {files} is {cosine_scores_final}')
else:
print(f'The Mark For {files} is {cosine_scores}')
# cosine_scores = marks(cosine_scores)
# cosine_scores = marks_final_pass(cosine_scores)
end = time.time()
print(f"Total Time Took Is {end-start}")
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="VGCRNI4JW4zo" outputId="deb6bc8c-b4ab-4c35-d0c8-b498c14b9df5"
main(f'/content/drive/MyDrive/EssayPreprocessed/Composite Materials- JA.Evangeline -24.pdf')
def open_context():
context = open("output.txt", "r").read()
return ''.join([i if ord(i) < 128 else ' ' for i in context]).replace('_','').replace('DOO','')
context = open_context()
context = nlp(context)
context = str(context._.outcome_spellCheck)
print("Preprocessing the Output")
# print(context)
ans = sentences2
print("Getting the Embeddings")
RealAnsEmbeddings = model.encode(ans, convert_to_tensor=True)
AnsEmbeddings = model.encode(context, convert_to_tensor=True)
print("Calculating the Marks...")
cosine_scores = util.pytorch_cos_sim(AnsEmbeddings, RealAnsEmbeddings)
cosine_scores = float(cosine_scores[0][0])
cosine_scores *= 100
cosine_scores = int(cosine_scores)
def final_mark(score):
if score + epsilon >= 98:
return 95
elif score <= 70:
return score + random.randint(9,11)
else:
return score + epsilon
def marks(scr):
if scr <= 80:
return scr + random.randint(7,9)
else:
return scr
def marks_final_pass(scr):
if scr <= 65:
return scr + random.randint(15,17)
elif scr <= 80:
return scr + random.randint(7,9)
else:
return scr
cosine_scores = final_mark(cosine_scores)
cosine_scores = marks(cosine_scores)
cosine_scores = marks_final_pass(cosine_scores)
print(f'The Mark is {cosine_scores}')
# + colab={"base_uri": "https://localhost:8080/"} id="WRAZNpQIRuDL" outputId="fd897d5f-c1eb-44b1-ff78-1b3426131675"
for files in os.listdir('/content/drive/MyDrive/EssayPreprocessed'):
print(files)
# + colab={"base_uri": "https://localhost:8080/"} id="TppaQ2jaIGWn" outputId="fe62f700-86ec-4914-a671-8ed158fc4211"
print(df_temp)
# + colab={"base_uri": "https://localhost:8080/"} id="rRgOONMZ4faf" outputId="2e76fe7a-28ff-461f-e852-57a79652dfae"
# !python drive.py --noauth_local_webserver
# + id="REVWb3z3rMXj"
import random
# + id="AUIq3wqQrMRG"
random.randint()
# + id="-roFwYSciGs0"
img_test = cv2.imread('/content/drive/MyDrive/AutomaticExamCorrection/UnMarked/IMG-20210105-WA0002 (2).jpg')
# + colab={"base_uri": "https://localhost:8080/"} id="Nvy5vQWTkWOk" outputId="005317a8-08dc-4682-b32c-9975c6037615"
img_test.shape
# + colab={"base_uri": "https://localhost:8080/"} id="VMBHvCn4k0zq" outputId="81d1f29f-7f7b-48a1-fd72-98528997f20b"
img_test_Re = cv2.resize(img_test,(0,0),None,0.,0.95)
img_test_Re.shape
# + id="7EeSSpUceEvA"
import cv2
# + id="PGvSBbV13W0c"
streamlit run your_script.py
# + colab={"base_uri": "https://localhost:8080/"} id="dFvqEc4F4XIB" outputId="44aaf1eb-afd2-4a51-fe6d-1f39b4f6909a"
# %%writefile drive.py
from __future__ import print_function
import httplib2
import os
import io
from apiclient import discovery
from oauth2client import client
from oauth2client import tools
from oauth2client.file import Storage
from apiclient.http import MediaFileUpload, MediaIoBaseDownload
try:
import argparse
flags = argparse.ArgumentParser(parents=[tools.argparser]).parse_args()
except ImportError:
flags = None
# If modifying these scopes, delete your previously saved credentials
# at ~/.credentials/drive-python-quickstart.json
SCOPES = 'https://www.googleapis.com/auth/drive'
CLIENT_SECRET_FILE = '/content/client_secrets.json'
APPLICATION_NAME = 'Drive API Python Quickstart'
def get_credentials():
"""Gets valid user credentials from storage.
If nothing has been stored, or if the stored credentials are invalid,
the OAuth2 flow is completed to obtain the new credentials.
Returns:
Credentials, the obtained credential.
"""
credential_path = os.path.join("./", 'drive-python-quickstart.json')
store = Storage(credential_path)
credentials = store.get()
if not credentials or credentials.invalid:
flow = client.flow_from_clientsecrets(CLIENT_SECRET_FILE, SCOPES)
flow.user_agent = APPLICATION_NAME
if flags:
credentials = tools.run_flow(flow, store, flags)
else: # Needed only for compatibility with Python 2.6
credentials = tools.run(flow, store)
print('Storing credentials to ' + credential_path)
return credentials
def main():
credentials = get_credentials()
http = credentials.authorize(httplib2.Http())
service = discovery.build('drive', 'v3', http=http)
imgfile = '/content/Scanned Documents.pdf' # Image with texts (png, jpg, bmp, gif, pdf)
txtfile = 'output.txt' # Text file outputted by OCR
mime = 'application/vnd.google-apps.document'
res = service.files().create(
body={
'name': imgfile,
'mimeType': mime
},
media_body=MediaFileUpload(imgfile, mimetype=mime, resumable=True)
).execute()
downloader = MediaIoBaseDownload(
io.FileIO(txtfile, 'wb'),
service.files().export_media(fileId=res['id'], mimeType="text/plain")
)
done = False
while done is False:
status, done = downloader.next_chunk()
service.files().delete(fileId=res['id']).execute()
print("Done.")
if __name__ == '__main__':
main()
# + colab={"base_uri": "https://localhost:8080/"} id="Y-bI1K474cLh" outputId="6ae39d84-5195-423d-f861-354c003464b6"
# !python drive.py --noauth_local_webserver
# + colab={"base_uri": "https://localhost:8080/"} id="stReITeC2xGC" outputId="0a568b32-82e7-4ad1-e877-cbcd30ad7f53"
# !ngrok http 8501
# + colab={"base_uri": "https://localhost:8080/"} id="S6haK1AC2xnf" outputId="06d01ed0-92c3-4cb3-a1a8-fc79af6f2f5d"
from pyngrok import ngrok
# Open a HTTP tunnel on the default port 80
public_url = ngrok.connect(port = '8501')
public_url
# + id="vqSOM1EjLy4p"
epsilon = 15
def final_mark(score):
if score + epsilon >= 98:
return 95
else:
return score + epsilon
# + colab={"base_uri": "https://localhost:8080/"} id="P1rIdGAwL2ws" outputId="388cc9de-446e-4cfe-b2d8-32012e01093a"
final_mark(87)
# + colab={"base_uri": "https://localhost:8080/"} id="aN6_lcm12CG2" outputId="c9bed31f-4377-454f-e0db-f2438a2652e2"
# !streamlit run /content/app.py --server.port 80
# + id="a8_HufMZCGPK" colab={"base_uri": "https://localhost:8080/", "height": 299, "referenced_widgets": ["cf92f9e79b4e4a4c957f2cb1d63acc16", "cd18734a3bde43d0bfecdfc937b9fe7e", "9dc7b7fcab50495ebe253210ce8880c5", "1b179c5a66a84b99832239e1c09bc049", "5e016406553b4417909a0b73328afd0d", "4a2977714b2c4e4e982fb48f3d15604d", "544d83cc7cd94b2f8adfb240af395b24", "ce4e87facde24977ad7b9ff2000556d6", "503615d8f9cc488281bcf4f1e37014e3", "b45cbbdeb2a143efb33c3c53459ec5f4", "0373ab74fd4a4e64a022affa0b6fa0f1", "774319eeff1f42e0a1b0399b9896470f", "ae611ca3cef048048ffa6084934310b7", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8da967f910fa4a679b7ed9c0ab14cbff", "<KEY>", "<KEY>", "507f9878855346ef8a0cc3a8d031554f", "<KEY>", "<KEY>", "e0843c0cf73c4e2d9fc948e35c30e37b", "<KEY>", "<KEY>", "cfcee3f8ccad4fcc88ce6e4cfacfc550", "<KEY>", "<KEY>", "3694c81916c34ae6a79b82138a0c1ea0", "ac3468c1704849e0ac773738e9c1d19b", "<KEY>", "<KEY>", "<KEY>", "251af6961f4a4f6cbbe23c985ee7a327", "d75795837bc84e72859683ccca2e058a", "63a16c0ec3c84b1dbe85dd173cf40dc5", "847839d89d434eb0afe8f6de7637f296", "d782591968a14e7c8f6e4ea376ebb7b8"]} outputId="219289c6-84d7-4fb7-f471-947e36b352f4"
from transformers import pipeline
text_generator = pipeline("text-generation")
print(text_generator("I'm Adithya and I live in NewYork, for breakfast I eat", max_length=50, do_sample=False))
# + id="jGqt7MNlcrEH"
| DeepLearningBasedDescriptivePaperCorrection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hackerton
# language: python
# name: hackerton
# ---
# +
import xml.etree.ElementTree as ET
import os
import glob
import shutil
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import copy
import cv2
from PIL import Image, ImageDraw
# -
# 원본이미지 폴더
basepath = '/home/ssac26/Downloads/wego/ai_test2'
os.path.dirname(basepath)
# txt변환후 이미지와 매칭되면 이동시킬 폴더
copy_path = os.path.join(os.path.dirname(basepath), os.path.basename(basepath)+'_copy')
copy_path
os.mkdir(copy_path)
bfolders = os.listdir(basepath)
bfolders
# custom annotation format to yolov4
def convert(box, size):
dw = 1./(size[0])
dh = 1./(size[1])
x = (box[0] + box[2])/2.0
y = (box[1] + box[3])/2.0
w = box[2] - box[0]
h = box[3] - box[1]
x = x*dw
w = w*dw
y = y*dh
h = h*dh
return (x,y,w,h)
def getimgdir(dir_path):
image_list = []
for i in glob.glob(dir_path + '/*.jpg'):
image_list.append(i)
return image_list
def gettextdir(dir_path):
label_list = []
for i in glob.glob(dir_path + '/*.txt'):
label_list.append(i)
return label_list
copy_path
# +
# parsing from xml, to txt
def parsing(path):
new_folder = os.path.join(copy_path, os.path.basename(path))
os.makedirs(new_folder) # make copy folder loc
#print(new_folder)
filelist = os.listdir(path)
for file in filelist:
if file.endswith('xml'):
#print(file)
xmlpath = os.path.join(path, file)
shutil.copy(xmlpath, new_folder)
doc = ET.parse(xmlpath)
root = doc.getroot()
for i in root.iter('image'):
# cou = 0
for j in i.iter('box'):
if j.attrib['label']=='person':
picname = i.attrib['name'].split('.')[0]
bbox_x1 = float(j.attrib['xtl'])
bbox_y1 = float(j.attrib['ytl'])
bbox_x2 = float(j.attrib['xbr'])
bbox_y2 = float(j.attrib['ybr'])
#print(convert([bbox_x1, bbox_y1, bbox_x2, bbox_y2],(1920,1080)))
xx, yy, xx2, yy2 = convert([bbox_x1, bbox_y1, bbox_x2, bbox_y2],(1920,1080))
#print(i.attrib['name'])
#print(i)
#print(j.attrib)
#print(xx, yy, xx2, yy2)
annotxt = os.path.join(path, picname) + '.txt'
#print(annotxt)
#print('osexist:',os.path.isfile(annotxt))
folder = os.listdir(path)
if os.path.basename(annotxt) in folder:
out_file = open(annotxt, 'a' ,encoding='UTF8')
#print()
out_file.write(f'{0} {round(xx,6)} {round(yy,6)} {round(xx2,6)} {round(yy2,6)}\n')
#print('addmode')
else:
out_file = open(os.path.join(path, picname) + '.txt', 'w' ,encoding='UTF8')
out_file.write(f'{0} {round(xx,6)} {round(yy,6)} {round(xx2,6)} {round(yy2,6)}\n')
#print('maketxt')
# cou += 1
# print(cou)
# -
copy_path
# +
def file_filtering(path):
img_1 = getimgdir(path)
text_1 = gettextdir(path)
addlist = []
new_folder = os.path.join(copy_path, os.path.basename(path))
# os.makedirs(new_folder)
for i in img_1:
full_fname = os.path.basename(i)
split_fname = os.path.splitext(full_fname)
#print(split_fname[0])
#print(i)
for j in text_1:
if split_fname[0] == os.path.splitext(os.path.basename(j))[0]: # 일치한 파일 827
addlist.append(split_fname[0])
#print(i)
#print(j)
shutil.copy(i, new_folder)
shutil.move(j, new_folder)
return len(addlist)
# +
#폴더의 모든파일을 읽는 함수
def read_all_file(path):
file_list = []
output = os.listdir(path)
for i in output:
file_list.append(path+'/'+i)
return file_list
# -
bbox_folders = os.listdir(basepath)
# +
for bbox_folder in bbox_folders:
workfolder = os.path.join(basepath, bbox_folder)
print(workfolder)
parsing(workfolder)
file_filtering(workfolder)
workfolder_copy = read_all_file(os.path.join(copy_path, os.path.basename(workfolder)))
txtcount = 0
jpgcount = 0
for i in workfolder_copy:
if 'txt' in i:
txtcount +=1
if 'jpg' in i:
jpgcount += 1
print('txt counts:',txtcount)
print('jpg counts:',jpgcount)
# -
| bbox_parsing_xml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pandas import read_csv
from datetime import datetime
from math import sqrt
from numpy import concatenate
from matplotlib import pyplot
from pandas import read_csv
from pandas import DataFrame
from pandas import concat
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_squared_error
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
# load and process data
def parse(x):
return datetime.strptime(x, '%Y %m %d %H')
dataset = read_csv('data.csv', parse_dates = [['year', 'month', 'day', 'hour']], index_col=0, date_parser=parse)
dataset.drop('No', axis=1, inplace=True)
dataset.columns = ['pollution', 'dew', 'temp', 'press', 'wnd_dir', 'wnd_spd', 'snow', 'rain']
dataset.index.name = 'date'
dataset['pollution'].fillna(0, inplace=True)
dataset = dataset[24:]
print("||"*40)
print("** DATA PROCESSING COMPLETED **")
print(dataset.head(5))
print("||"*40)
dataset.to_csv('pollution.csv')
# generating dataset plot
from pandas import read_csv
from matplotlib import pyplot
dataset = read_csv('pollution.csv', header=0, index_col=0)
values = dataset.values
groups = [0, 1, 2, 3, 5, 6, 7]
i = 1
pyplot.figure()
for group in groups:
pyplot.subplot(len(groups), 1, i)
pyplot.plot(values[:, group],'k')
pyplot.title(dataset.columns[group], y=0.5, loc='right')
i += 1
pyplot.show()
# Lets normalize all features, and remove the weather variables for the hour to be predicted.
import pandas as pd
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
def s_to_super(data, n_in=1, n_out=1, dropnan=True):
n_vars = 1 if type(data) is list else data.shape[1]
df = pd.DataFrame(data)
cols, names = list(), list()
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [('var%d(t-%d)' % (j+1, i)) for j in range(n_vars)]
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [('var%d(t)' % (j+1)) for j in range(n_vars)]
else:
names += [('var%d(t+%d)' % (j+1, i)) for j in range(n_vars)]
agg = pd.concat(cols, axis=1)
agg.columns = names
if dropnan:
agg.dropna(inplace=True)
return agg
# load dataset
dataset = read_csv('pollution.csv', header=0, index_col=0)
values = dataset.values
encoder = preprocessing.LabelEncoder()
values[:,4] = encoder.fit_transform(values[:,4])
values = values.astype('float32')
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
reframed = s_to_super(scaled, 1, 1)
# drop columns we don't want to predict
reframed.drop(reframed.columns[[9,10,11,12,13,14,15]], axis=1, inplace=True)
print("** NOT REQUIRED DATA COLUMNS DROPPED **")
print("||"*40)
# split data into training and testing, futher splitting the train and test sets into i/p and o/p variables
# reshaped data further into 3D formate expected by LSTMs
values = reframed.values
n_train_hours = 365 * 24
train = values[:n_train_hours, :]
test = values[n_train_hours:, :]
# split into input and outputs
train_X, train_y = train[:, :-1], train[:, -1]
test_X, test_y = test[:, :-1], test[:, -1]
train_X = train_X.reshape((train_X.shape[0], 1, train_X.shape[1]))
test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
print("** DATA SPLITTING COMPLETED **")
print(" Training data shape X, y => ",train_X.shape, train_y.shape," Testing data shape X, y => ", test_X.shape, test_y.shape)
print("||"*40)
# defining LSTM with 50 neurons in first hidden layer and 1 neuron in the o/p layer
# using the MAE loss function and Adma version of stochastic gradient descent
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers import Dense, Dropout
model = Sequential()
# 50 neurons in first hidden layer
model.add(LSTM(50, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dropout(0.3))
model.add(Dense(1,kernel_initializer='normal', activation='sigmoid'))
model.compile(loss='mae', optimizer='adam')
history = model.fit(train_X, train_y, epochs=50, batch_size=72, validation_data=(test_X, test_y), verbose=2, shuffle=False)
# tracking history for plots
pyplot.plot(history.history['loss'], 'b', label='training history')
pyplot.plot(history.history['val_loss'], 'r',label='testing history')
pyplot.title("Train and Test Loss for the LSTM")
pyplot.legend()
pyplot.show()
# evaluating model
# make a prediction
from math import sqrt
from numpy import concatenate
yhat = model.predict(test_X)
test_X = test_X.reshape((test_X.shape[0], test_X.shape[2]))
inv_yhat = concatenate((yhat, test_X[:, 1:]), axis=1)
inv_yhat = scaler.inverse_transform(inv_yhat)
inv_yhat = inv_yhat[:,0]
inv_y = scaler.inverse_transform(test_X)
inv_y = inv_y[:,0]
# calculate RMSE
rmse = sqrt(mean_squared_error(inv_y, inv_yhat))
print('Test RMSE: %.3f' % rmse)
| forecasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
input_patterns=np.load('input_patterns.npy')
weights=np.load('weights.npy')
# +
def update_theta(theta_BCM,post_syn_patterns):
dt=0.001
BCM_target = 2.0
tau_BCM=0.01
#post_syn_patterns += dt*(-1*post_syn_patterns + np.dot(W,update_rates(pre_syn_patterns)))
#Ex= post_syn_patterns*(0.5 * (np.sign(post_syn_patterns) + 1))
theta_BCM+=dt*(-theta_BCM+(post_syn_patterns*(post_syn_patterns/BCM_target)))/tau_BCM
#print('theta_BCM',theta_BCM)
return theta_BCM
def update_w(W,pre_syn_patterns,post_syn_patterns,theta_BCM):
dt=0.01
tau_w=0.01
W_max=1.0
W+=dt*(pre_syn_patterns*np.dot(post_syn_patterns,(post_syn_patterns-theta_BCM)))/tau_w
W = W*(0.5 * (np.sign(W) + 1))
# bounding weights below max value
W[W>W_max] = W_max
return W
def update_rates(x):
#rates = x
r_0 = 1.0
r_max = 20.0
x[x<=0] = r_0*np.tanh(x[x<=0]/r_0)
x[x>0] = (r_max-r_0)*np.tanh(x[x>0]/(r_max-r_0))
return x
def one_timestep(W,pre_syn_patterns,post_syn_patterns,theta_BCM):
dt=0.001
#new_W=update_w(W,pre_syn_patterns,post_syn_patterns,theta_BCM)
#new_theta_BCM=update_theta(theta_BCM,W,post_syn_patterns)
new_post_syn_patterns =post_syn_patterns+(-1*post_syn_patterns + np.dot(W,update_rates(pre_syn_patterns)))
new_post_syn_patterns= new_post_syn_patterns*(0.5 * (np.sign(new_post_syn_patterns) + 1))
new_theta_BCM=update_theta(theta_BCM,post_syn_patterns)
#print('prim',theta_BCM)
new_W=update_w(W,pre_syn_patterns,post_syn_patterns,theta_BCM)
#theta_BCM=update_theta(theta_BCM,W,post_syn_patterns)
#return post_syn_patterns,new_W,new_theta_BCM
return new_post_syn_patterns,new_W,new_theta_BCM
def run_sim(W,pre_syn_patterns):
post_syn_patterns=np.array([0.0])
T=10000
Ws=[]
theta_BCM=np.array([0.5])
xs=[]
thrs=[]
for i in range(T):
post_syn_patterns, W, theta_BCM=one_timestep(W,pre_syn_patterns[:,i],post_syn_patterns,theta_BCM)
#print('double',theta_BCM)
#print(post_syn_patterns)
#print(W)
Ws.append(W)
xs.append(post_syn_patterns)
thrs.append(theta_BCM[0])
xs=np.array(xs)
return xs, Ws, thrs
# -
W=weights.copy()
print(W)
pre_syn_patterns=input_patterns
xs,Ws,thrs=run_sim(W,pre_syn_patterns)
plt.plot(xs)
plt.plot(thrs)
print(thrs)
print(xs)
Ws=np.array(Ws)
print(Ws.shape)
plt.plot(Ws.T[2])
| WorkInProgress2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="SJQanHSPPo-L"
#Import Libraries
import numpy as np
import pandas as pd
import nltk
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="kPmX6ukIWeB7" outputId="1269c99a-30d5-45bd-8c51-584323003c9b"
sample_string = "This is a pretty good movie"
#Punkt is a tokenizer present in NLTK
nltk.download('punkt')
tokens = nltk.tokenize.word_tokenize(sample_string)
print(tokens)
#We can also manually tokenize
split_words = [words for words in sample_string.split()]
print(split_words)
# + colab={"base_uri": "https://localhost:8080/"} id="BJJElyd-XHvC" outputId="96ef51e7-a8fa-4ad2-e08b-cfd8f95264eb"
#A corpus is a large and structured set of machine-readable texts that have been produced in a natural communicative setting.
#Stopwords are words that dont add much meaning to the sentence, that is, if they are removed then the sentence still manages
#pass on most if not all of its meaning
from nltk.corpus import stopwords
nltk.download('stopwords')
#Creating an array of stopwords present in the English language
stopwords = np.array(stopwords.words('english'))
print(stopwords)
print(stopwords.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="Jc7QG9g0ZCEB" outputId="bb402aab-0a8e-4e8c-bb22-e44ac8f30b7e"
#Separating the stopwords from the useful words
useful_words = [x for x in split_words if x not in stopwords]
print(useful_words)
# + colab={"base_uri": "https://localhost:8080/"} id="bZhdmf_rZgpw" outputId="248c429f-f6d9-4fef-eb6b-892329f4b011"
#Stemming is the process of keeping the root words from a list of similar words
#It may or may not retain any meaning
#For example, 'ban', 'banana', 'bankruptcy', 'banner' all start with 'ban-' but they are not related to each other at all
from nltk.stem import PorterStemmer
ps = PorterStemmer()
test_set = ['ban', 'banana', 'banner', 'bankruptcy']
stemmed = [ps.stem(words) for words in test_set]
#Stemming changes all words to lower case
print(stemmed)
# + id="b72ksDRIa9Qf" colab={"base_uri": "https://localhost:8080/", "height": 684} outputId="c5c89515-d98e-4888-8186-aec30f8d511b"
#Importing tweet data
data = pd.read_csv('https://github.com/psantheus/TwitterSentimentAnalysisNLTK/blob/main/data.csv')
data.info()
data.head(15)
# + id="7JfcvMJje_Py"
import re
def remove_patterns(pattern, text):
#Finds all matching strings that match the regex pattern provided
occur = re.findall(pattern, text)
#Substitutes every such pattern that was found with a empty string
#and returns the string after every removal
for match in occur:
text = re.sub(match, "", text)
return text
# + id="znb5BbKTkPAu" colab={"base_uri": "https://localhost:8080/", "height": 701} outputId="d0d1aca3-256a-4c19-e8cb-d9aaae07cd55"
#Adds a new column containing the cleaned data to the original dataframe
data['cleaned'] = [remove_patterns('@[\w]*', sentence) for sentence in data['tweet']]
data.info()
data.head(15)
# + id="DFNEx8-rlTTs" colab={"base_uri": "https://localhost:8080/", "height": 514} outputId="839004fe-7bac-42ee-ae74-6a2ebb217891"
#Further processing, ^ implies 'except' when within regex sets, so it removes everything
#except 'a-z', 'A-Z' and '#' which constitutes hashtags together with the alphabets
data['cleaned'] = data['cleaned'].str.replace("[^a-zA-Z#]", " ")
data.head(15)
# + colab={"base_uri": "https://localhost:8080/", "height": 514} id="lteiGWSqmlQ7" outputId="6b61eb0a-87d8-4a2a-9301-1f456c52bd40"
#Here, lambda expressions basically work on every item, x is the item,
#and then the operation is applied on x
#Tokenizing
data['cleaned'] = data['cleaned'].apply(lambda x : x.split())
#Removing stopwords
data['cleaned'] = data['cleaned'].apply(lambda x : [words for words in x if words not in stopwords])
#Stemming
data['cleaned'] = data['cleaned'].apply(lambda x : [ps.stem(words) for words in x])
#Converting each item back to sentence
data['cleaned'] = data['cleaned'].apply(lambda x : ' '.join(x))
#Showing current data
data.head(15)
# + colab={"base_uri": "https://localhost:8080/", "height": 648} id="nLJyo4EfjYpY" outputId="d098e387-4cd2-42d7-9092-00282bca15e9"
#Count Vectorizer basically performs a count of every word present as a whole,
#It maps each word to a label and then counts the number of times the labeled words appear
#In each sentence, throughout the document as a whole
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(max_df=0.8, min_df=2, max_features=1000, stop_words='english')
vectorized_data = vectorizer.fit_transform(data['cleaned'])
vectorized_data = pd.DataFrame(vectorized_data.todense())
vectorized_data.info()
vectorized_data.head(15)
# + id="kUrZdiVajfZ5"
#Test train splitting of data
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(vectorized_data, data['label'], test_size = 0.3)
# + id="1rgy98kVkdQp" colab={"base_uri": "https://localhost:8080/"} outputId="b4acefa5-cc2d-49f1-c9ca-5f752d09303d"
#Logistic Regression
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
LRClf = LogisticRegression(solver = 'lbfgs').fit(X_train, Y_train)
probab_est = LRClf.predict_proba(X_test)
print(probab_est.shape)
#accuracy_score returns the accuracy by comparing Y_predicted with Y_true
predictions = LRClf.predict(X_test)
accuracy = accuracy_score(predictions, Y_test)
print(accuracy)
#LogisticRegression.score() method does the exact same thing except in one line
print(LRClf.score(X_test, Y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="6BsAjT3BsYRU" outputId="82413d89-e0b1-401f-c055-f52b3bff9c34"
#f1_score is used when classes are largely unbalanced, in this case number of happy tweets
#largely outnumbers the number of sad tweets
from sklearn.metrics import f1_score
print(np.unique(predictions, return_counts=True))
f1_accuracy = f1_score(predictions, Y_test)
print(f1_accuracy)
# + colab={"base_uri": "https://localhost:8080/"} id="gAEjEDzBnorW" outputId="cc35db6f-6d55-46d3-b056-12a7c302f99c"
#SVM
from sklearn import svm
SVMClf = svm.SVC().fit(X_train, Y_train)
predictions = SVMClf.predict(X_test)
accuracy = accuracy_score(predictions, Y_test)
print(accuracy)
#f1 score from SVM
f1_accuracy = f1_score(predictions, Y_test)
print(f1_accuracy)
# + colab={"base_uri": "https://localhost:8080/"} id="m6gJM8d9wkfq" outputId="11195729-ebbf-4fa8-dc48-3eb89135e891"
#TfidfVectorizer provides the same results as CountVectorizer followed by TfidfTranformer
#Tf stands for Term-frequency
#Idf stands for Inverse document frequency
#The picture at https://miro.medium.com/max/1050/1*qQgnyPLDIkUmeZKN2_ZWbQ.png explains the concept in a nutshell
#In TfidfVectorizer we consider overall document weightage of a word. It helps us in dealing with most frequent words.
#Using it we can penalize them. TfidfVectorizer weights the word counts by a measure of how often they appear in the documents.
#If a word appears in every document, it is not so unique hence its actual affect on the meaning must be low, but not zero
#Using TfidfVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
tfidfvectorizer = TfidfVectorizer(max_df=0.8, min_df=2, max_features=1000, stop_words='english')
tfidfvectorized_data = tfidfvectorizer.fit_transform(data['cleaned'])
tfidfvectorized_data = pd.DataFrame(tfidfvectorized_data.todense())
X_train, X_test, Y_train, Y_test = train_test_split(tfidfvectorized_data, data['label'], test_size = 0.3)
#SVM using default kernel
TfidfSVMClf = svm.SVC().fit(X_train, Y_train)
predictions = TfidfSVMClf.predict(X_test)
accuracy = accuracy_score(predictions, Y_test)
print(accuracy)
#f1 score from SVM
f1_accuracy = f1_score(predictions, Y_test)
print(f1_accuracy)
#SVM with linear kernel
LinKernSVMClf = svm.SVC(kernel='linear').fit(X_train, Y_train)
predictions = LinKernSVMClf.predict(X_test)
accuracy = accuracy_score(predictions, Y_test)
print(accuracy)
#f1 score from SVM
f1_accuracy = f1_score(predictions, Y_test)
print(f1_accuracy)
| Twitter_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example: Redshift-space Galaxy Power Spectrum
from molino import GalaxyCatalog
from pyspectrum import pyspectrum as pySpec
# -- plotting --
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
# # read Molino galaxy catalogs
# Lets read in 3 N-body relations and 2 HOD realizations at the `Mnu_p` cosmology, *first in real-space*
nbodys = range(3)
hods = range(2)
# +
# GalaxyCatalog?
# -
xyzs = GalaxyCatalog(
'Mnu_p', # specify the cosmological and HOD parameters
i_nbody=nbodys, # n-body realizations
i_hod=hods, # HOD realizations
apply_rsd=False # don't apply HOD
)
print(len(xyzs))
print(xyzs[0].shape)
# ## Now lets measure their power spectrum using the `pySpectrum` package
pks = []
for xyz in xyzs:
pks.append(pySpec.Pk_periodic(xyz.T, Lbox=1000., Ngrid=360, silent=False))
fig = plt.figure(figsize=(6,6))
sub = fig.add_subplot(111)
for i_nbody in range(len(nbodys)):
for i_hod in range(len(hods)):
_pk = pks[2*i_nbody+i_hod]
sub.plot(_pk['k'], _pk['p0k'], c='C%i' % i_nbody, ls=['-', ':'][i_hod])
sub.set_ylabel('real-space $P(k)$', fontsize=25)
sub.set_yscale('log')
sub.set_xlabel('$k$', fontsize=25)
sub.set_xlim([3e-3, 1.])
sub.set_xscale('log')
# # now in redshift-space
# RSD applied along the `z`-axis
xyzs = GalaxyCatalog(
'Mnu_p',
i_nbody=nbodys,
i_hod=hods,
apply_rsd='z' # RSD along the 'z' direction (you can also put 'x' and 'y')
)
pks = []
for xyz in xyzs:
pks.append(pySpec.Pk_periodic_rsd(xyz.T, Lbox=1000., Ngrid=360, silent=True))
# +
fig = plt.figure(figsize=(12,6))
sub = fig.add_subplot(121)
for i_nbody in range(len(nbodys)):
for i_hod in range(len(hods)):
_pk = pks[2*i_nbody+i_hod]
sub.plot(_pk['k'], _pk['p0k'], c='C%i' % i_nbody, ls=['-', ':'][i_hod])
sub.set_ylabel('$P_0(k)$', fontsize=25)
sub.set_yscale('log')
sub.set_xlabel('$k$', fontsize=25)
sub.set_xlim([3e-3, 1.])
sub.set_xscale('log')
sub = fig.add_subplot(122)
for i_nbody in range(len(nbodys)):
for i_hod in range(len(hods)):
_pk = pks[2*i_nbody+i_hod]
sub.plot(_pk['k'], _pk['p2k'], c='C%i' % i_nbody, ls=['-', ':'][i_hod])
sub.set_ylabel('$P_2(k)$', fontsize=25)
sub.set_yscale('log')
sub.set_xlabel('$k$', fontsize=25)
sub.set_xlim([3e-3, 1.])
sub.set_xscale('log')
# -
| docs/eg_powerspectrum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Word Alignment Assignment__
#
# Your task is to learn word alignments for the data provided with this Python Notebook.
#
# Start by running the 'train' function below and implementing the assertions which will fail. Then consider the following improvements to the baseline model:
# * Is the TranslationModel parameterized efficiently?
# * What form of PriorModel would help here? (Currently the PriorModel is uniform.)
# * How could you use a Hidden Markov Model to model word alignment indices? (There's an implementation of simple HMM below to help you start.)
# * How could you initialize more complex models from simpler ones?
# * How could you model words that are not aligned to anything?
#
# Grades will be assigned as follows*:
#
# AER below on blinds | Grade
# ----------|-------------
# 0.5 - 0.6 | 1
# 0.4 - 0.5 | 2
# 0.35 - 0.4 | 3
# 0.3 - 0.35 | 4
# 0.25 - 0.3 | 5
#
# You should save the notebook with the final scores for 'dev' and 'test' test sets.
#
# *__Note__: Students who submitted a version of this assignment last year will have a 0.05 AER handicap, i.e to get a grade of 5, they will need to get an AER below 0.25.
#
# +
# This cell contains the generative models that you may want to use for word alignment.
# Currently only the TranslationModel is at all functional.
import numpy as np
from collections import defaultdict
from copy import deepcopy
class TranslationModel:
"Models conditional distribution over trg words given a src word."
def __init__(self, src_corpus, trg_corpus, identity_matrix, hard_align=False):
self.identity_matrix = identity_matrix
self.num_unique_src_tokens = identity_matrix.shape[0]
self.num_unique_trg_tokens = identity_matrix.shape[1]
self._trg_given_src_probs = np.ones((self.num_unique_src_tokens,
self.num_unique_trg_tokens)) / self.num_unique_trg_tokens
self._src_trg_counts = np.zeros((self.num_unique_src_tokens, self.num_unique_trg_tokens))
self.hard_align = hard_align
def get_params(self):
return self._trg_given_src_probs
def get_conditional_prob(self, src_token, trg_token):
"Return the conditional probability of trg_token given src_token."
return self._trg_given_src_probs[src_token][trg_token]
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
"Returns matrix with t[i][j] = p(f_j|e_i)."
return self._trg_given_src_probs[np.ix_(src_tokens, trg_tokens)]
def collect_statistics(self, src_tokens, trg_tokens, posterior_matrix, hmm=False):
"Accumulate counts of translations from: posterior_matrix[j][i] = p(a_j=i|e, f)"
# assert posterior_matrix.shape == (len(trg_tokens), len(src_tokens))
# assert False, "Implement collection of statistics here."
self._src_trg_counts[np.ix_(src_tokens, trg_tokens)] += posterior_matrix
def recompute_parameters(self):
"Reestimate parameters and reset counters."
# assert False, "Implement reestimation of parameters from counters here."
self._trg_given_src_probs = self._src_trg_counts / np.sum(self._src_trg_counts, axis=1, keepdims=True)
self._src_trg_counts = np.zeros((self.num_unique_src_tokens, self.num_unique_trg_tokens))
if self.hard_align:
self._trg_given_src_probs[self.identity_matrix.row, self.identity_matrix.col] = 1.0
class PriorModel:
"Models the prior probability of an alignment given only the sentence lengths and token indices."
def __init__(self, src_corpus, trg_corpus):
"Add counters and parameters here for more sophisticated models."
self._distance_counts = {}
self._distance_probs = {}
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
src_length = len(src_tokens)
trg_length = len(trg_tokens)
return np.ones((src_length, trg_length)) * 1.0 / src_length
def get_prior_prob(self, src_index, trg_index, src_length, trg_length):
"Returns a uniform prior probability."
return 1.0 / src_length
def collect_statistics(self, src_length, trg_length, posterior_matrix):
"Extract the necessary statistics from this matrix if needed."
pass
def recompute_parameters(self):
"Reestimate the parameters and reset counters."
pass
class ComplexPriorModel:
"Models the prior probability of an alignment given the sentence lengths and token indices."
def __init__(self, src_corpus, trg_corpus, use_null=False,
src_phi=0.5, trg_phi=0.5, src_null_index=0, trg_null_index=0):
"Add counters and parameters here for more sophisticated models."
self.num_src_indices = np.max(list(map(len, src_corpus)))
self.num_trg_indices = np.max(list(map(len, trg_corpus)))
self._distance_counts = defaultdict(lambda:
np.zeros((self.num_src_indices,
self.num_trg_indices)))
self._distance_probs = defaultdict(lambda:
np.ones((self.num_src_indices,
self.num_trg_indices)) / self.num_trg_indices)
self.src_phi = src_phi
self.trg_phi = trg_phi
self.src_null_index = src_null_index
self.trg_null_index = trg_null_index
self.use_null = use_null
def get_prior_prob(self, src_index, trg_index, src_length, trg_length):
"Returns a uniform prior probability."
return self._distance_probs[(src_length, trg_length)][src_index, trg_index]
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
src_length = len(src_tokens)
trg_length = len(trg_tokens)
return (self._distance_probs[(src_length, trg_length)]
[np.ix_(np.arange(src_length), np.arange(trg_length))])
def collect_statistics(self, src_tokens, trg_tokens, posterior_matrix):
"Extract the necessary statistics from this matrix if needed."
src_length = len(src_tokens)
trg_length = len(trg_tokens)
src_indices = np.arange(src_length)
trg_indices = np.arange(trg_length)
(self._distance_counts[(src_length, trg_length)]
[np.ix_(src_indices, trg_indices)]) += posterior_matrix
def recompute_parameters(self):
"Reestimate the parameters and reset counters."
for key in self._distance_counts:
denoms = np.sum(self._distance_counts[key], axis=0, keepdims=True)
self._distance_probs[key] = self._distance_counts[key] / denoms
if self.use_null:
self._distance_probs[key][self.src_null_index, :] *= self.src_phi
self._distance_probs[key][:self.src_null_index, :] *= (1 - self.src_phi)
self._distance_probs[key][(self.src_null_index + 1):, :] *= (1 - self.src_phi)
self._distance_probs[key][:, self.trg_null_index] *= self.trg_phi
self._distance_probs[key][:, :self.trg_null_index] *= (1 - self.trg_phi)
self._distance_probs[key][:, (self.trg_null_index + 1):] *= (1 - self.trg_phi)
self._distance_counts[key] = np.zeros((self.num_src_indices, self.num_trg_indices))
class ImprovedComplexPriorModel:
"Models the prior probability of an alignment given the sentence lengths and token indices."
def __init__(self, src_corpus, trg_corpus, num_indices=10,
use_null=False, src_phi=0.5, trg_phi=0.5, src_null_index=0, trg_null_index=0):
"Add counters and parameters here for more sophisticated models."
self.num_src_indices = num_indices
self.num_trg_indices = num_indices
self._distance_counts = np.zeros((self.num_src_indices, self.num_trg_indices))
self._distance_probs = np.ones((self.num_src_indices,
self.num_trg_indices)) / self.num_trg_indices
self.src_phi = src_phi
self.trg_phi = trg_phi
self.src_null_index = src_null_index
self.trg_null_index = trg_null_index
self.use_null = use_null
def get_prior_prob(self, src_index, trg_index, src_length, trg_length):
"Returns a uniform prior probability."
return self._distance_probs[int(trg_index / trg_length * self.num_trg_indices),
int(src_index / src_length * self.num_src_indices)]
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
src_length = len(src_tokens)
trg_length = len(trg_tokens)
squeezed_src_indices = np.array(list(map(lambda x: int(x / src_length * self.num_src_indices),
np.arange(src_length))))
squeezed_trg_indices = np.array(list(map(lambda x: int(x / trg_length * self.num_trg_indices),
np.arange(trg_length))))
return self._distance_probs[np.ix_(squeezed_src_indices, squeezed_trg_indices)]
def collect_statistics(self, src_tokens, trg_tokens, posterior_matrix):
"Extract the necessary statistics from this matrix if needed."
src_length = len(src_tokens)
trg_length = len(trg_tokens)
squeezed_src_indices = np.array(list(map(lambda x: int(x / src_length * self.num_src_indices),
np.arange(src_length))))
squeezed_trg_indices = np.array(list(map(lambda x: int(x / trg_length * self.num_trg_indices),
np.arange(trg_length))))
self._distance_counts[np.ix_(squeezed_src_indices, squeezed_trg_indices)] += posterior_matrix
def recompute_parameters(self):
"Reestimate the parameters and reset counters."
denoms = np.sum(self._distance_counts, axis=0, keepdims=True)
self._distance_probs = self._distance_counts / denoms
if self.use_null:
self._distance_probs[self.src_null_index, :] *= self.src_phi
self._distance_probs[:self.src_null_index, :] *= (1 - self.src_phi)
self._distance_probs[(self.src_null_index + 1):, :] *= (1 - self.src_phi)
self._distance_probs[:, self.trg_null_index] *= self.trg_phi
self._distance_probs[:, :self.trg_null_index] *= (1 - self.trg_phi)
self._distance_probs[:, (self.trg_null_index + 1):] *= (1 - self.trg_phi)
self._distance_counts = np.zeros((self.num_src_indices, self.num_trg_indices))
class TransitionModel:
"Models the prior probability of an alignment conditioned on previous alignment."
def __init__(self, src_corpus, trg_corpus):
"Add counters and parameters here for more sophisticated models."
self.num_src_indices = np.max(list(map(len, src_corpus)))
self.alignment_probs_given_prev = dict()
self.alignment_counts = dict()
def get_parameters_for_sentence_pair(self, src_tokens, trg_tokens):
"Retrieve the parameters for this sentence pair: A[k, i] = p(a_{j} = i|a_{j-1} = k)"
src_length = len(src_tokens)
trg_length = len(trg_tokens)
if src_length not in self.alignment_probs_given_prev:
self.alignment_probs_given_prev[src_length] = np.ones((src_length, src_length)) / src_length
return self.alignment_probs_given_prev[src_length]
def collect_statistics(self, src_tokens, trg_tokens, bigram_posteriors):
"Extract statistics from the bigram posterior[i][j]: p(a_{t-1} = i, a_{t} = j| e, f)"
src_length = len(src_tokens)
trg_length = len(trg_tokens)
if src_length not in self.alignment_counts:
self.alignment_counts[src_length] = np.zeros((src_length, src_length))
self.alignment_counts[src_length] += np.sum(bigram_posteriors, axis=2)
def recompute_parameters(self):
"Recompute the transition matrix"
for length in self.alignment_counts:
denoms = np.sum(self.alignment_counts[length], axis=0, keepdims=True)
self.alignment_probs_given_prev[length] = self.alignment_counts[length] / denoms
self.alignment_counts[length] = np.zeros((length, length))
# +
# This cell contains the framework for training and evaluating a model using EM.
from utils import read_parallel_corpus, extract_test_set_alignments, score_alignments
from itertools import starmap
from math import log
from scipy.sparse import coo_matrix
import editdistance
import multiprocessing
import os
import functools
def infer_posteriors(src_tokens, trg_tokens, prior_model, translation_model, hmm=False):
"Compute the posterior probability p(a_j=i | f, e) for each target token f_j given e and f."
# HINT: An HMM will require more complex statistics over the hidden alignments.
P = prior_model.get_parameters_for_sentence_pair(src_tokens, trg_tokens)
T = translation_model.get_parameters_for_sentence_pair(src_tokens, trg_tokens) # t[i][j] = P(f_j|e_i)
# assert False, "Compute the posterior distribution over src indices for each trg word."
if hmm:
initial_distribution = np.ones(len(src_tokens)) / len(src_tokens)
bigram_posterior_matrix = np.zeros((len(src_tokens), len(src_tokens), len(trg_tokens)))
unigram_posterior_matrix = np.zeros((len(trg_tokens), len(src_tokens)))
alpha, beta, sentence_marginal_log_likelihood = forward_backward(initial_distribution, P, T)
unigram_posterior_matrix = alpha * beta
denoms = np.sum(unigram_posterior_matrix, axis=0, keepdims=True)
unigram_posterior_matrix /= denoms
bigram_posterior_matrix = (alpha[:, None, :-1] * P[:, :, None] *
beta[None, :, 1:] * T[None, :, 1:])
denoms = np.sum(bigram_posterior_matrix, axis=(0, 1), keepdims=True)
bigram_posterior_matrix /= denoms
return unigram_posterior_matrix, bigram_posterior_matrix, sentence_marginal_log_likelihood
posterior_matrix = P * T
denoms = np.sum(posterior_matrix, axis=0, keepdims=True)
posterior_matrix /= denoms
sentence_marginal_log_likelihood = np.sum(np.log(denoms))
return posterior_matrix, sentence_marginal_log_likelihood
def collect_expected_statistics(src_corpus, trg_corpus, prior_model, translation_model, hmm=False):
"E-step: infer posterior distribution over each sentence pair and collect statistics."
corpus_log_likelihood = 0.0
for src_tokens, trg_tokens in zip(src_corpus, trg_corpus):
# Infer posterior
if hmm:
unigram_posteriors, bigram_posteriors, log_likelihood = infer_posteriors(
src_tokens, trg_tokens, prior_model, translation_model, hmm=hmm)
prior_model.collect_statistics(src_tokens, trg_tokens, bigram_posteriors)
translation_model.collect_statistics(src_tokens, trg_tokens, unigram_posteriors)
else:
posteriors, log_likelihood = infer_posteriors(src_tokens, trg_tokens, prior_model,
translation_model, hmm=hmm)
# Collect statistics in each model.
prior_model.collect_statistics(src_tokens, trg_tokens, posteriors)
translation_model.collect_statistics(src_tokens, trg_tokens, posteriors)
# Update log prob
corpus_log_likelihood += log_likelihood
return corpus_log_likelihood
def estimate_models(src_corpus, trg_corpus, prior_model, translation_model,
num_iterations, hmm=False, use_null=False,
src_null_index=0, trg_null_index=0):
"Estimate models iteratively using EM."
for iteration in range(num_iterations):
# E-step
corpus_log_likelihood = collect_expected_statistics(src_corpus, trg_corpus,
prior_model, translation_model, hmm=hmm)
# M-step
prior_model.recompute_parameters()
translation_model.recompute_parameters()
if iteration > 0:
print("corpus log likelihood: %1.3f" % corpus_log_likelihood)
aligned_corpus = align_corpus(src_corpus, trg_corpus,
prior_model, translation_model, hmm=hmm,
use_null=use_null, src_null_index=src_null_index,
trg_null_index=trg_null_index)
evaluate(extract_test_set_alignments(aligned_corpus))
return prior_model, translation_model
def get_alignments_from_posterior(posteriors, hmm=False, use_null=False,
src_null_index=0, trg_null_index=0):
"Returns the MAP alignment for each target word given the posteriors."
# HINT: If you implement an HMM, you may want to implement a better algorithm here.
alignments = {}
for trg_index, src_index in enumerate(np.argmax(posteriors, 0)):
if src_index == src_null_index or trg_index == trg_null_index:
continue
if use_null:
src_index -= 1
trg_index -= 1
if trg_index not in alignments:
alignments[trg_index] = {}
alignments[trg_index][src_index] = '*'
return alignments
def align_corpus(src_corpus, trg_corpus, prior_model, translation_model, hmm=False,
use_null=False, src_null_index=0, trg_null_index=0):
"Align each sentence pair in the corpus in turn."
aligned_corpus = []
for src_tokens, trg_tokens in zip(src_corpus, trg_corpus):
if hmm:
posteriors, _, _, = infer_posteriors(src_tokens, trg_tokens, prior_model,
translation_model, hmm=hmm)
else:
posteriors, _ = infer_posteriors(src_tokens, trg_tokens, prior_model,
translation_model, hmm=hmm)
alignments = get_alignments_from_posterior(posteriors, hmm=hmm, use_null=use_null,
src_null_index=src_null_index,
trg_null_index=trg_null_index)
aligned_corpus.append((src_tokens, trg_tokens, alignments))
return aligned_corpus
def initialize_models(src_corpus, trg_corpus, identity_matrix, translation_model_cls,
prior_model_cls, translation_model_=None, prior_model_=None,
hard_align=False, **prior_params):
prior_model = (prior_model_cls(src_corpus, trg_corpus, **prior_params)
if prior_model_ is None else prior_model_)
translation_model = (translation_model_cls(src_corpus, trg_corpus, identity_matrix, hard_align)
if translation_model_ is None else translation_model_)
return prior_model, translation_model
def load_lemmas(filenames):
word_to_lemma = {}
for filename in filenames:
with open(filename) as fin:
for line in fin:
lemma, word = line.strip().split()
word_to_lemma[word] = lemma
return word_to_lemma
def normalize_corpus(corpus, use_null=False, null_token="<null>",
use_lemmas=False, lemmas_files=[], use_hashing=False, num_buckets=3000):
if use_lemmas:
word_to_lemma = load_lemmas(lemmas_files)
corpus = [list(map(lambda word: word_to_lemma.get(word.lower(), word.lower()), tokens))
for tokens in corpus]
unique_tokens = sorted(set(token for tokens in corpus for token in tokens))
if use_null:
unique_tokens = [null_token] + unique_tokens
token_to_idx = {token: idx for idx, token in enumerate(unique_tokens)}
null_index = token_to_idx.get(null_token, None)
normalized_corpus = []
for tokens in corpus:
token_indices = [token_to_idx[token] for token in tokens]
if use_hashing:
offset = 1 if use_null else 0
token_indices = [offset + (hash(token) % num_buckets) for token in tokens]
else:
token_indices = [token_to_idx[token] for token in tokens]
if use_null:
token_indices = [null_index] + token_indices
normalized_corpus.append(token_indices)
return normalized_corpus, unique_tokens, null_index
def calc_trg_indices(src_data, unique_trg_tokens, use_editdistance,
use_hashing, num_buckets, use_null):
trg_indices = []
src_idx, src_token = src_data
offset = 1 if use_null else 0
if use_hashing:
trg_tokens_with_indices = map(lambda token: (offset + (hash(token) % num_buckets), token),
unique_trg_tokens)
else:
trg_tokens_with_indices = enumerate(unique_trg_tokens)
for trg_idx, trg_token in trg_tokens_with_indices:
if (src_token == trg_token or
(use_editdistance and
(editdistance.eval(src_token, trg_token) / len(src_token)) < 0.2)):
trg_indices.append(trg_idx)
return trg_indices, src_idx, src_token
def calc_identity_matrix(unique_src_tokens, unique_trg_tokens, use_editdistance,
use_hashing, num_buckets, use_null):
iis = []
js = []
values = []
offset = 1 if use_null else 0
with multiprocessing.Pool(8) as pool:
map_func = functools.partial(calc_trg_indices,
unique_trg_tokens=unique_trg_tokens,
use_editdistance=use_editdistance,
use_hashing=use_hashing,
num_buckets=num_buckets,
use_null=use_null)
if use_hashing:
src_tokens_with_indices = map(lambda token: (offset + (hash(token) % num_buckets), token),
unique_src_tokens)
else:
src_tokens_with_indices = enumerate(unique_src_tokens)
for trg_indices, src_idx, src_token in pool.imap(map_func, src_tokens_with_indices):
iis.extend([src_idx] * len(trg_indices))
js.extend(trg_indices)
values.extend([1.0] * len(trg_indices))
if use_hashing:
shape = (offset + num_buckets, offset + num_buckets)
else:
shape = (len(unique_src_tokens), len(unique_trg_tokens))
return coo_matrix((values, (iis, js)), shape=shape)
def normalize(src_corpus, trg_corpus, use_null=False,
src_null_token="<src_null>", trg_null_token="<trg_null>",
use_editdistance=False, use_lemmas=False, lemmas_folder="lemmatization-lists",
use_hashing=False, num_buckets=3000):
# assert False, "Apply some normalization here to reduce the numbers of parameters."
(normalized_src,
unique_src_tokens,
src_null_index) = normalize_corpus(src_corpus, use_null, src_null_token,
use_lemmas, [os.path.join(lemmas_folder,
"lemmatization-en.txt")],
use_hashing, num_buckets)
(normalized_trg,
unique_trg_tokens,
trg_null_index) = normalize_corpus(trg_corpus, use_null, trg_null_token,
use_lemmas, [os.path.join(lemmas_folder,
"lemmatization-sl.txt"),
os.path.join(lemmas_folder,
"lemmatization-sk.txt"),
os.path.join(lemmas_folder,
"lemmatization-cs.txt")],
use_hashing, num_buckets)
identity_matrix = calc_identity_matrix(unique_src_tokens, unique_trg_tokens,
use_editdistance, use_hashing, num_buckets, use_null)
return normalized_src, normalized_trg, identity_matrix, src_null_index, trg_null_index
def train(num_iterations, translation_model_cls=TranslationModel, prior_model_cls=PriorModel,
translation_model=None, prior_model=None, hmm=False, hard_align=False,
src_null_token="<src_null>", trg_null_token="<trg_null>", use_editdistance=False,
use_lemmas=False, lemmas_folder="lemmatization-lists",
use_hashing=False, num_buckets=3000, **prior_params):
src_corpus, trg_corpus, _ = read_parallel_corpus('en-cs.all')
use_null = prior_params.get("use_null", False)
if translation_model is not None:
use_editdistance = False
(src_corpus, trg_corpus, identity_matrix,
src_null_index, trg_null_index) = normalize(src_corpus, trg_corpus,
use_null, src_null_token, trg_null_token,
use_editdistance, use_lemmas, lemmas_folder,
use_hashing, num_buckets)
if use_null and not hmm and prior_model_cls != PriorModel:
prior_params["src_null_index"] = src_null_index
prior_params["trg_null_index"] = trg_null_index
if use_null and (hmm or prior_model_cls == PriorModel):
del prior_params["use_null"]
prior_model, translation_model = initialize_models(src_corpus, trg_corpus, identity_matrix,
translation_model_cls, prior_model_cls,
translation_model, prior_model, hard_align,
**prior_params)
prior_model, translation_model = estimate_models(src_corpus, trg_corpus, prior_model,
translation_model, num_iterations,
hmm=hmm, use_null=use_null,
src_null_index=src_null_index,
trg_null_index=trg_null_index)
aligned_corpus = align_corpus(src_corpus, trg_corpus, prior_model, translation_model,
hmm=hmm, use_null=use_null,
src_null_index=src_null_index, trg_null_index=trg_null_index)
return extract_test_set_alignments(aligned_corpus), translation_model, prior_model
def evaluate(candidate_alignments):
src_dev, trg_dev, wa_dev = read_parallel_corpus('en-cs-wa.dev', has_alignments=True)
src_test, trg_test, wa_test = read_parallel_corpus('en-cs-wa.test', has_alignments=True)
print('dev: recall %1.3f; precision %1.3f; aer %1.3f' % score_alignments(wa_dev, candidate_alignments['dev']))
print('test: recall %1.3f; precision %1.3f; aer %1.3f' % score_alignments(wa_test, candidate_alignments['test']))
# -
# # Experimenting with different models
# Let's start with a simple IBM Model 1:
test_alignments, _, _ = train(5)
evaluate(test_alignments)
# Now we will add hard-alignment explicitly setting alignment probability of identical tokens to 1.
test_alignments, _, _ = train(5, hard_align=True)
evaluate(test_alignments)
# Hard alignment lowered our AER by 0.01. We will use it in all later experiments.
# Now let's try IBM Model 2 with a prior that depends on word positions in a sentence.
test_alignments, _, _ = train(5, prior_model_cls=ComplexPriorModel, hard_align=True)
evaluate(test_alignments)
# More complex prior certainly improved our AER.
# Now we pretrain translation model with IBM Model 1 for 2 epochs and then
# train IBM Model 2 using pretrained translation model.
_, translation_model1, _ = train(2, hard_align=True)
test_alignments, _, _ = train(5, prior_model_cls=ComplexPriorModel,
translation_model=translation_model1)
evaluate(test_alignments)
# Pretrained model produces better AER than a model without pretraining.
# For now our ComplexPrior depended on sentence lengths. We remove that dependency in ImprovedComplexPrior by using only relative position of the word in a sentence: $relative\_pos = \frac{word\_index}{sentence\_length}$. To simplify things, we introduce buckets, each of which will be responsible for one area of a sentence then we will use mapping from a relative position in a sentence to a bucket.
# We can calculate bucket numbers from relative positions as follows: $$bucket\_number = \lfloor{relative\_pos \cdot num\_buckets}\rfloor$$
#
# For example in a sentence "Quick brown | fox jumps | over the | lazy dog" where bucket borders are depicted using '|' token, the word 'jumps' has index 3 and therefore its relative position is $\frac{3}{8} = 0.375$ and its bucket number is $\lfloor0.375 * 4\rfloor = 1$
#
# In our improved prior we use bucket indices instead of word indices, that way we reduce the number of parameters in our model.
test_alignments, _, _ = train(5, prior_model_cls=ImprovedComplexPriorModel,
hard_align=True)
evaluate(test_alignments)
# Let's try to pretrain IBM Model 2 with improved prior using Model 1.
_, translation_model1, _ = train(2, hard_align=True)
test_alignments, _, _ = train(5, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1)
evaluate(test_alignments)
# Now, let's use HMM to train our alignments
test_alignments, _, _ = train(10, prior_model_cls=TransitionModel, hmm=True,
hard_align=True)
evaluate(test_alignments)
# As we can see, HMM starts to diverge after 5-6 iterations, so it's no use to train it longer.
# Trying to add pretraining with Model 1 to HMM
_, translation_model1, _ = train(2, hard_align=True)
test_alignments, _, _ = train(6, prior_model_cls=TransitionModel,
translation_model=translation_model1,
hmm=True)
evaluate(test_alignments)
# Let's try to optimise parameters of Model 2 with Model 1 pretraining:
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True)
test_alignments, _, _ = train(15, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1)
evaluate(test_alignments)
# -
# Let's increase the number of buckets (default was 10)
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True)
test_alignments, _, _ = train(15, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=15)
evaluate(test_alignments)
# -
_, translation_model1, _ = train(10, hard_align=True)
test_alignments, _, _ = train(15, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20)
evaluate(test_alignments)
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True)
test_alignments, _, _ = train(15, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=25)
evaluate(test_alignments)
# -
# All versions of Model 2 with pretraining start to diverge after 5-6 iterations, so there is no point in training it further. Model with num_buckets=20 gives the best AER
# Now let's use the best chained pretraining model to pretrain HMM
_, translation_model1, _ = train(10, hard_align=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20)
test_alignments, _, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2, hmm=True)
evaluate(test_alignments)
# # Adding data normalization
# Now that we've experimented with different models, we can improve them further by modifying our data.
# ### Using NULL
# We start with adding NULL tokens to source and target sentences, so that our models have the option of not aligning the word anywhere.
test_alignments, _, _ = train(5, hard_align=True, use_null=True)
evaluate(test_alignments)
# + jupyter={"outputs_hidden": true}
test_alignments, _, _ = train(5, prior_model_cls=ComplexPriorModel, hard_align=True,
use_null=True)
evaluate(test_alignments)
# + jupyter={"outputs_hidden": true}
test_alignments, _, _ = train(5, prior_model_cls=ImprovedComplexPriorModel, hard_align=True,
use_null=True, num_indices=20)
evaluate(test_alignments)
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True, use_null=True)
test_alignments, _, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True)
evaluate(test_alignments)
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True, use_null=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True)
evaluate(test_alignments)
# -
# Null tokens improved AER, but not by much.
# ### Using lemmas and edidistance
# Now we reduce number of different words in our corpora by mapping them to their lowercase lemmas. Also we improve hard-alignment by setting alignment probability to 1 when tokens have small editdistance compared to the source word length (for example $\frac{edit\_distance}{source\_word\_length} < 0.2$)
# Only using editdistance and nulls:
_, translation_model1, _ = train(10, hard_align=True, use_null=True, use_editdistance=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True)
evaluate(test_alignments)
# Adding lemmas from [this repo](https://github.com/michmech/lemmatization-lists) (needs to be cloned and placed alongside this notebook). Using english lemmas for english and Czech, Slovak and Slovene lemmas for Czech.
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True, use_null=True, use_editdistance=True,
use_lemmas=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True, use_lemmas=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True, use_lemmas=True)
evaluate(test_alignments)
# -
# ### Adding hashing
# To further decrease the number of parameters we simplify things by mapping words to indices using hash function: $word\_idx = hash(word)\ \%\ num\_buckets$.
# The final model has num_buckets=3000.
_, translation_model1, _ = train(10, hard_align=True, use_null=True, use_editdistance=True,
use_lemmas=True, use_hashing=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True, use_lemmas=True, use_hashing=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
# Hashing didn't help to decrease AER, now let's try to do several runs of HMM using fresh TransitionModels, because Translation Model will keep improving and Transition model won't diverge.
# +
_, translation_model1, _ = train(10, hard_align=True, use_null=True, use_editdistance=True,
use_lemmas=True, use_hashing=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True, use_lemmas=True, use_hashing=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model_hmm,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model_hmm,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
evaluate(test_alignments)
# -
# Let's see how far we can push this:
# + jupyter={"outputs_hidden": true}
_, translation_model1, _ = train(10, hard_align=True, use_null=True, use_editdistance=True,
use_lemmas=True, use_hashing=True)
_, translation_model2, _ = train(6, prior_model_cls=ImprovedComplexPriorModel,
translation_model=translation_model1,
num_indices=20, use_null=True, use_lemmas=True, use_hashing=True)
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model2,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
num_repetitions = 5
for i in range(num_repetitions):
test_alignments, translation_model_hmm, _ = train(5, prior_model_cls=TransitionModel,
translation_model=translation_model_hmm,
hmm=True, use_null=True, use_lemmas=True,
use_hashing=True)
evaluate(test_alignments)
# -
# Well, repeating HMM over and over didn't improve the AER significantly, so I guess the best model is the previous one (with only 3 repetitions of HMM)
# +
# Discrete HMM with scaling. You may want to use this if you decide to implement an HMM.
# The parameters for this HMM will still need to be provided by the models above.
def forward(pi, A, O):
S, T = O.shape
alpha = np.zeros((S, T))
scaling_factors = np.zeros(T)
# base case
alpha[:, 0] = pi * O[:, 0]
scaling_factors[0] = np.sum(alpha[:, 0])
alpha[:, 0] /= scaling_factors[0]
# recursive case
for t in range(1, T):
alpha[:, t] = np.dot(alpha[:, t-1], A[:, :]) * O[:, t]
# Normalize at each step to prevent underflow.
scaling_factors[t] = np.sum(alpha[:, t])
alpha[:, t] /= scaling_factors[t]
return (alpha, scaling_factors)
def backward(pi, A, O, forward_scaling_factors):
S, T = O.shape
beta = np.zeros((S, T))
# base case
beta[:, T-1] = 1 / forward_scaling_factors[T-1]
# recursive case
for t in range(T-2, -1, -1):
beta[:, t] = np.sum(beta[:, t+1] * A[:, :] * O[:, t+1], 1) / forward_scaling_factors[t]
return beta
def forward_backward(pi, A, O):
alpha, forward_scaling_factors = forward(pi, A, O)
beta = backward(pi, A, O, forward_scaling_factors)
return alpha, beta, np.sum(np.log(forward_scaling_factors))
# -
| week7_mt/homework/word_alignment_assignment-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("..")
from optimus import Optimus
# Create optimus
op = Optimus(master="local", app_name= "optimus", verbose = True)
# -
# # Mysql
# Put your db credentials here
db = op.connect(
driver="mysql",
host="172.16.58.3",
database= "optimus",
user= "test",
password = "<PASSWORD>")
db.tables()
# +
# db.execute("SHOW KEYS FROM test_data WHERE key_name = 'PRIMARY'")
# -
db.execute("SELECT * FROM test_data").ext.display()
db.execute("SELECT * FROM test_data", partition_column ="id", table_name = "test_data").ext.display()
db.table_to_df("test_data", partition_column ="id").ext.display()
df = db.table_to_df("test_data", limit=None)
db.tables_names_to_json()
# # Postgres
# Put your db credentials here
db = op.connect(
driver="postgresql",
host="172.16.58.3",
database= "optimus",
user= "testuser",
password = "<PASSWORD>")
db.tables()
db.table_to_df("test_data").table()
db.tables_names_to_json()
# ## MSSQL
# Put your db credentials here
db = op.connect(
driver="sqlserver",
host="172.16.58.3",
database= "optimus",
user= "test",
password = "<PASSWORD>")
db.tables()
db.table_to_df("test_data").table()
db.tables_names_to_json()
# ## Redshit
# Put your db credentials here
db = op.connect(
driver="redshift",
host="172.16.58.3",
database= "optimus",
user= "testuser",
password = "<PASSWORD>")
db.tables()
db.table_to_df("test_data").table()
# ## Oracle
# Put your db credentials here
db = op.connect(
driver="oracle",
host="172.16.58.3",
database= "optimus",
user= "testuser",
password = "<PASSWORD>")
# ## SQLlite
# Put your db credentials here
db = op.connect(
driver="sqlite",
host="chinook.db",
database= "employes",
user= "testuser",
password = "<PASSWORD>")
db.tables()
db.table_to_df("albums",limit="all").table()
db.tables_names_to_json()
# ## Redis
df = op.load.csv("https://raw.githubusercontent.com/ironmussa/Optimus/master/examples/data/foo.csv", sep=",", header='true', infer_schema='true', charset="UTF-8", null_value="None")
df.table()
# Put your db credentials here
db = op.connect(
driver="redis",
host="172.16.58.3",
port = 6379,
database= 1,
password = "")
db.df_to_table(df, "hola1", redis_primary_key="id")
# +
# https://stackoverflow.com/questions/56707978/how-to-write-from-a-pyspark-dstream-to-redis
db.table_to_df(0)
# -
| examples/jdbc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model
# ### Imports and plotting preferences
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import scipy.stats as stats
import seaborn as sns
import statsmodels.api as sm
from numpy.random import gamma, lognormal, normal
from statsmodels.sandbox.regression.predstd import wls_prediction_std
from tqdm.notebook import tqdm
from graphs import load_data, occupancy_arrays
mpl.rc('font', family = 'serif', size = 15)
mpl.rcParams['xtick.labelsize'] = 14
mpl.rcParams['ytick.labelsize'] = 14
# -
save_dir = os.path.join(os.pardir, 'figs')
if not os.path.exists(save_dir):
os.makedirs(save_dir)
# Global preferences
n_obs = 11 # UPDATE THIS
n_future = 14
n_total = n_obs + n_future
# ## Case number extrapolation
# ### Local plotting functions
def ci_plot(y, x_pred, y_pred, ci_l, ci_u, r,
regions, dates, iter, ylabel='% ICU bed occupancy', obs=True, pct=False, y_max=2500):
# plt.figure(figsize=(15, 15))
ax = plt.subplot(331+iter)
ax.grid(True)
ax.set_xticks(np.arange(0, len(x_pred), 3))
plt.fill(np.concatenate([x_pred, x_pred[::-1]]), np.concatenate([ci_l, ci_u[::-1]]),
alpha=0.5, fc='b', ec='None', label='95% CI')
plt.plot(x_pred, y_pred, 'b-', label='Fit (OLS)')
if obs:
plt.plot(np.arange(n_obs), y[:n_obs], 'r.', markersize=10, label='Observed')
plt.text(15, 100, f'R$^{2}$={r:.2f}', bbox=dict(boxstyle="round", ec=(1., 0.5, 0.5), fc=(1., 0.8, 0.8)))
else:
y_max = 100 if pct else 1000
plt.plot([n_obs, n_obs], [0, y_max], 'r')
plt.text(n_obs, y_max/2, 'Prediction\n window\n$\longrightarrow$', color='r')
plt.xlim(0, x_pred.max())
plt.ylim(0, y_max)
plt.xticks(range(0, n_total, 3), dates[::3]) if iter in [4, 5, 6] else ax.set_xticklabels([])
plt.setp(ax.xaxis.get_majorticklabels(), rotation=45)
if iter in [0, 3, 6]: plt.ylabel(ylabel)
plt.title(regions[iter])
if iter==0:
plt.legend(loc='upper left')
def regional_predictions(X, X_pred, Y, regions, dates, ylabel, log=True):
fig = plt.figure(figsize=(15, 15))
mean_arr = np.zeros((7, len(X_pred)))
std_arr = np.zeros((7, len(X_pred)))
e = []
for i in range(len(regions)):
y = Y[i]
if log:
y = np.log(y)
mod = sm.OLS(y, X)
res = mod.fit()
e += [res.params[1]]
y_pred = res.predict(X_pred)
_, _, std_u = wls_prediction_std(res, exog=X_pred, alpha=1-0.6827) # 1 s.d.
_, ci_l, ci_u = wls_prediction_std(res, exog=X_pred, alpha=1-0.95) # 95% CI
# Store
mean_arr[i] = y_pred
std_arr[i] = std_u - y_pred
if log:
y_pred = np.exp(y_pred)
ci_l = np.exp(ci_l)
ci_u = np.exp(ci_u)
# Plot
ci_plot(Y[i], X_pred[:, 1], y_pred, ci_l, ci_u, res.rsquared,
regions, dates, iter=i, ylabel=ylabel)
return mean_arr, std_arr, e
# ### Callable functions for website
X, X_pred, cum_cases, regions, dates = load_data()
new_cases = cum_cases[:, 1:] - cum_cases[:, :-1]
df = pd.DataFrame(new_cases.transpose(), columns=list(regions))
corr = df.corr()
mask = np.triu(np.ones_like(corr, dtype=np.bool), 1)
f, ax = plt.subplots(figsize=(11, 9))
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(corr, mask=mask, cmap=cmap, vmin=0.0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
_ = plt.xticks(rotation=45, ha='right')
plt.savefig(os.path.join(save_dir, 'regional_correlation.pdf'))
df = pd.DataFrame(cum_cases.transpose(), columns=list(regions))
df.plot(marker='o', figsize=(8, 8))
plt.plot([n_obs-1, n_obs-1], [0, df.max().max()+10], 'k:', zorder=0)
plt.xlim(-0.1, n_obs+1.1)
plt.ylim(0, df.max().max()+10)
support = np.concatenate((np.arange(n_obs), [n_obs+0.5, n_obs+2]))
plt.xticks(support, dates[:n_obs]+[dates[n_obs+6]]+[dates[n_obs+13]], rotation=45)
plt.ylabel('Cumulative COVID-19 patients')
plt.text(1, 600, 'Observed', size=20, rotation=45)
plt.text(n_obs-0.5, 600, 'Prediction\n window', size=20, rotation=45)
plt.savefig(os.path.join(save_dir, 'windows.pdf'))
means, stds, exponents = regional_predictions(X, X_pred, cum_cases, regions, dates, ylabel='Cumulative COVID-19 patients', log=False)
plt.savefig(os.path.join(save_dir, 'new_patients_linear_fit.pdf'))
log_means, log_stds, exponents = regional_predictions(X, X_pred, cum_cases, regions, dates, ylabel='Cumulative COVID-19 patients', log=True)
exponents = np.array(exponents)
print(exponents)
plt.savefig(os.path.join(save_dir, 'new_patients_log-linear_fit.pdf'))
# ### LOS
beds = pd.read_csv(os.path.join(os.pardir, 'data', 'model', 'ICU_beds_region.csv'))['n_beds (2019)'].values
death_and_icu_info = pd.read_csv(os.path.join(os.pardir, 'data', 'model', 'hospitalisation_and_fatalities.csv'))
cfr = death_and_icu_info['Mortality Rate']
pct_need_icu = death_and_icu_info['Critical Care Needs Rate']
mu, sig = occupancy_arrays(log_means, log_stds, exponents, pct_need_icu,
icu_delay_normal_loc=2.0, los_gamma_shape=8.0, log=True)
fig = plt.figure(figsize=(15, 15))
for i in range(len(regions)):
ci_plot(new_cases[i], X_pred[:, 1], mu[i], mu[i]-1.96*sig[i], mu[i]+1.96*sig[i], None,
regions, dates, i, ylabel='New COVID-19 patients in ICU', obs=False)
plt.savefig(os.path.join(save_dir, 'covid_icu_patients.pdf'))
avg_occ = mu / beds[:, np.newaxis] * 100
std_occ = sig / beds[:, np.newaxis] * 100
fig = plt.figure(figsize=(15, 15))
for i in range(len(regions)):
ci_plot(new_cases[i], X_pred[:, 1], avg_occ[i], avg_occ[i]-1.96*std_occ[i], avg_occ[i]+1.96*std_occ[i], None,
regions, dates, i, ylabel='% ICU occupancy', obs=False, pct=True)
plt.savefig(os.path.join(save_dir, 'pct_covid_icu_occupancy.pdf'))
| extrapolation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# ## Introduction
#
# This page describes the technical design of Intake-esm, motivation behind the project, and components of the package.
# ## Why Intake-esm?
# Project efforts such as [CMIP](https://www.wcrp-climate.org/wgcm-cmip), and [CESM Large Ensembe](http://www.cesm.ucar.edu/projects/community-projects/LENS/) produce a huge of amount data persisted in multiple NetCDF files. Finding, investigating, loading these files into data array containers such as `xarray` can be a difficult task because the number of files a user may be interested in can be large. `Intake-esm` was written to make it easy to seamlessly find, investigate, load, and disseminate earth system data holdings produced by these projects.
#
# `Intake-esm` solves a set of problems:
#
# - It eliminates the need for the user to know specific locations (file path) of their data set of interest.
# - It allows the user to specify simple spec to define data sources and build collection catalogs for these data sources.
# - It loads data sets into data array containers such as `xarray`, and gets out of your way.
# - It allows reproduciblity, and data provenance.
#
#
# Intake-esm supports data holdings from the following projects:
#
# - [CMIP](./cmip5.ipynb): Coupled Model Intercomparison Project ([phase 5](https://esgf-node.llnl.gov/projects/cmip5/) and [phase 6](https://esgf-node.llnl.gov/projects/cmip6/))
# - [CESM](./cesm.ipynb): [Community Earth System Model Large Ensemble (LENS), and Decadal Prediction Large Ensemble (DPLE)](http://www.cesm.ucar.edu/projects/community-projects/)
# - [MPI-GE](./mpige.ipynb): [The Max Planck Institute for Meteorology (MPI-M) Grand Ensemble (MPI-GE)](https://www.mpimet.mpg.de/en/grand-ensemble/)
# - [GMET](./gmet.ipynb): [The Gridded Meteorological Ensemble Tool data](https://ncar.github.io/hydrology/models/GMET)
# - [ERA5](./era5.ipynb): [ECWMF ERA5 Reanalysis dataset stored on NCAR's GLADE](https://rda.ucar.edu/datasets/ds630.0/#!description) in ``/glade/collections/rda/data/ds630.0``
# ## Concepts
# `Intake-esm` extends functionality provided by `intake`. `Intake-esm` is built out of four core concepts:
#
#
# - **Collection**: An object that represents a reference to a data holding such as a collection of CESM Large ensemble model output.
#
# - **Data Source**: An object that represents a reference to a data source. Data source objects have methods for loading the data into `xarray` containers namely `xarray` datasets.
#
# - **Catalog**: A collection of catalog entries, each of which defines a data source. Like in `intake`, catalog objects can be created from local YAML definitions by some driver that knows how to query a data collection.
#
# - **Catalog Entry**: A named data source. The catalog entry includes metadata about the source, as well as the name of the driver and arguments. Arguments can be parameterized, allowing one entry to return different subsets of data depending on the user request.
| docs/source/notebooks/overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="QKxW4D6DHCyx"
# # Advanced Certification in AIML
# ## A Program by IIIT-H and TalentSprint
# + [markdown] colab_type="text" id="OzkHDYHGZnyC"
# ### Learning Objectives:
#
# At the end of the experiment, you will be able to:
#
# * Preprocessing text data
# * Representation of text document using Bag of Words
# * Understand Bag of Words represented text data with K-nearest neighbours
# + [markdown] colab_type="text" id="Fzskman3ZuWC"
# ### Dataset
# In this experiment we use the 20 newsgroup dataset
#
# **Description**
#
# This dataset is a collection of approximately 20,000 newsgroup documents, partitioned across 20 different newsgroups. That is there are approximately one thousand documents taken from each of the following newsgroups:
#
# alt.athesim
# comp.graphics
# comp.os.ms-windows.misc
# comp.sys.ibm.pc.hardware
# comp.sys.mac.hardware
# comp.windows.x
# misc.forsale
# rec.autos
# rec.motorcycles
# rec.sport.baseball
# rec.sport.hockey
# sci.crypt
# sci.electronics
# sci.med
# sci.space
# soc.religion.christian
# talk.politics.guns
# talk.politics.mideast
# talk.politics.misc
# talk.religion.misc
#
# The dataset consists **Usenet** posts--essentially an email sent by subscribers to that newsgroup. They typically contain quotes from previous posts as well as cross posts i.e. a few posts may be sent to more than once in a newsgroup.
#
# Each newsgroup is stored in a subdirectory, with each post stored as a separate file.
#
# Data source to this experiment : http://archive.ics.uci.edu/ml/datasets/Twenty+Newsgroups
# + [markdown] colab_type="text" id="vW1Vu2adZ0oO"
# ### Domain Information
# A newsgroup, despite the name, has nothing to do with news. It is what we would call today a mailing list or a discussion forum. *Usenet* is a distributed discussion system designed and developed in 1979 and deployed in 1980.
#
# Members joined newsgroups of interest to them and made *posts* to them. Posts are very similar to email -- in later years, newsgroups became mailing lists and people posted via email.
# + [markdown] colab_type="text" id="bo3iVdSkZ5gb"
# The problem that we are attempting is "Text classification". This is a broadly defined task which is common to many services and products: for example, gmail classifies an incoming mail into different sections such as Updates, Forums etc
#
# + [markdown] colab_type="text" id="a0muqcxoZ6fL"
# ### Bag of Words (BoW)
#
# * The bag-of-words is a simple to understand representation of documents and words. As you are aware it makes use of the one-hot representation of each word based on the vocabulary and the document is represented as a sum of the BoW vectors of all the words in the document
#
# #### Challenges
#
# * The dimension of each vector representing a word is the number of words in the vocabulary. So we definitely will encounter the *curse of dimensionality*
# * Bag of words representation doesn’t consider the semantic relation between words.
# * Nor does it capture the grammar of the language--parts of speech etc.,
# + [markdown] colab_type="text" id="FfgFJFcCHCy2"
# #### Keywords
#
# * Numpy
# * Collections
# * Gensim
# * Bag-of-Words (Word Frequency, Pre-Processing)
# * Bag-of-Words representation
# + [markdown] colab_type="text" id="xw5JkrNeHCy3"
# #### Expected Time : 60 min
# + [markdown] colab_type="text" id="YuhepSBE20co"
# ### Setup Steps
# + colab={} colab_type="code" id="K8-lZrNaHCy8"
# Importing required Packages
import pickle
import re
import operator
from collections import defaultdict
import matplotlib.pyplot as plt
import numpy as np
import math
import collections
import gensim
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" executionInfo={"elapsed": 926, "status": "ok", "timestamp": 1581572141375, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="iy0iuGsTHCzA" outputId="f226cbd3-173a-405d-f769-86c57f93aa09"
# Loading the dataset
dataset = pickle.load(open('AIML_DS_NEWSGROUPS_PICKELFILE.pkl','rb'))
print(type(dataset))
print(dataset.keys())
# + [markdown] colab_type="text" id="I2k1kcF_HCy0"
# To get a sense of our data, let us first start by counting the frequencies of the target classes in our news articles in the training set.
# + colab={"base_uri": "https://localhost:8080/", "height": 391} colab_type="code" executionInfo={"elapsed": 862, "status": "ok", "timestamp": 1581572156022, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="h8T5v8vGHCzC" outputId="c6c5a5de-74a4-4d65-c95c-b735db180d67"
# Print frequencies of dataset
print("Class : count")
print("--------------")
number_of_documents = 0
for key in dataset:
print(key, ':', len(dataset[key]))
# + [markdown] colab_type="text" id="Cl74CoG8HCzE"
# Next, let us split our dataset which consists of about 1000 samples per class, into training and test sets. We use about 95% samples from each class in the training set, and the remaining in the test set.
#
#
#
#
#
# As a mental exercise you should try reasoning about why is it important to ensure a nearly equal distribution of classes in your training and test sets.
# + colab={} colab_type="code" id="xgUy5WyFHCzF"
train_set = {}
test_set = {}
new_dataset = {}
# Clean dataset for text encoding issues :- Very useful when dealing with non-unicode characters
for key in dataset:
new_dataset[key] = [[i.decode('utf-8', errors='replace').lower() for i in f] for f in dataset[key]]
# Break dataset into 95-5 split for training and testing
n_train = 0
n_test = 0
for k in new_dataset:
split = int(0.95*len(new_dataset[k]))
train_set[k] = new_dataset[k][0:split]
test_set[k] = new_dataset[k][split:]
n_train += len(train_set[k])
n_test += len(test_set[k])
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 833, "status": "ok", "timestamp": 1581572387049, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="4dv0_hjvDqtA" outputId="2a668c77-6a2f-4288-82f8-7dbc971ba3bf"
len(new_dataset.values())
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 856, "status": "ok", "timestamp": 1581572299515, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="JK0Wz9ApDgOp" outputId="de8428cb-ad75-4a79-f025-7c404a71c16c"
split
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" executionInfo={"elapsed": 1041, "status": "ok", "timestamp": 1581572274603, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="GpbwjKUaDWoT" outputId="9fbd1fab-942a-41af-a133-2e7ccacc1ca6"
train_set.keys()
# + [markdown] colab_type="text" id="jvwvqYpJHCzH"
# ## 1. Bag-of-Words
#
# Let us begin our journey into text classification with one of the simplest but most commonly used feature representations for news documents - Bag-of-Words.
#
# As you might have realized, machine learning algorithms need good feature representations of different inputs. Concretely, we would like to represent each news article $D$ in terms of a feature vector $V$, which can be used for classification. Feature vector $V$ is made up of the number of occurences of each word in the vocabulary.
#
# Let us begin by counting the number of occurences of every word in the news documents in the training set.
# + [markdown] colab_type="text" id="zcoPXdxBHCzI"
# ### 1.1 Word frequency
# + [markdown] colab_type="text" id="iuHlXAyFHCzJ"
# Let us try understanding the kind of words that appear frequently, and those that occur rarely. We now count the frequencies of words:
# + colab={"base_uri": "https://localhost:8080/", "height": 408} colab_type="code" executionInfo={"elapsed": 3597, "status": "ok", "timestamp": 1581572416619, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="HXR_X4GSHCzK" outputId="23691060-53b3-41c3-f173-58e536af080c"
# Initialize a dictionary to store frequencies of words.
# Key:Value === Word:Count
frequency = defaultdict(int)
for key in train_set:
for f in train_set[key]:
# Find all words which consist only of capital and lowercase characters and are between length of 2-9.
# We ignore all special characters such as !.$ and words containing numbers
words = re.findall(r'(\b[A-Za-z][a-z]{2,9}\b)', ' '.join(f))
for word in words:
frequency[word] += 1
sorted_words = sorted(frequency.items(), key=operator.itemgetter(1), reverse=True)
print("Top-10 most frequent words:")
for word in sorted_words[:10]:
print(word)
print('----------------------------')
print("10 least frequent words:")
for word in sorted_words[-10:]:
print(word)
# + [markdown] colab_type="text" id="eUCZWOwhHCzP"
# Next, we attempt to plot a histogram of the counts of various words in descending order.
#
# Could you comment about the relationship between the frequency of the most frequent word to the second frequent word?
# And what about the third most frequent word?
#
# (Hint - Check the relative frequencies of the first, second and third most frequent words)
#
# (After answering, you can visit https://en.wikipedia.org/wiki/Zipf%27s_law for further Reading)
# + colab={"base_uri": "https://localhost:8080/", "height": 623} colab_type="code" executionInfo={"elapsed": 2338, "status": "ok", "timestamp": 1581572533176, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="KV-PAkMBHCzQ" outputId="acdf0452-cd25-403f-e905-fdb665d29190"
# %matplotlib inline
fig = plt.figure()
fig.set_size_inches(20,10)
plt.bar(range(len(sorted_words[:100])), [v for k, v in sorted_words[:100]] , align='center')
plt.xticks(range(len(sorted_words[:100])), [k for k, v in sorted_words[:100]])
locs, labels = plt.xticks()
plt.setp(labels, rotation=90)
plt.show()
# + [markdown] colab_type="text" id="ZZDlJ5cyHCzT"
# ### 1.2 Pre-processing to remove most and least frequent words
#
# We can see that different words appear with different frequencies.
#
# The most common words appear in almost all documents. Hence, for a classification task, having information about those words' frequencies does not mater much since they appear frequently in every type of document. To get a good feature representation, we eliminate them since they do not add too much value.
#
# Additionally, notice how the least frequent words appear so rarely that they might not be useful either.
#
# Let us pre-process our news articles now to remove the most frequent and least frequent words by thresholding their counts:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1025, "status": "ok", "timestamp": 1581572578170, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="XSPqj3pZHCzT" outputId="8cc64b26-f9ba-4202-f15f-886cf9c36571"
valid_words = defaultdict(int)
print('Number of words before preprocessing:', len(sorted_words))
# Ignore the 25 most frequent words, and the words which appear less than 100 times
ignore_most_frequent = 25
freq_thresh = 100
feature_number = 0
for word, word_frequency in sorted_words[ignore_most_frequent:]:
if word_frequency > freq_thresh:
valid_words[word] = feature_number
feature_number += 1
print('Number of words after preprocessing:', len(valid_words))
word_vector_size = len(valid_words)
# + [markdown] colab_type="text" id="SEggZhtmHCzV"
# ### 1.3 Bag-of-Words representation
#
# The simplest way to represent a document $D$ as a vector $V$ would be to now count the relevant words in the document.
#
# For each document, make a vector of the count of each of the words in the vocabulary (excluding the words removed in the previous step - the "stopwords").
# + colab={} colab_type="code" id="iJKDjIjJHCzV"
def convert_to_BoW(dataset, number_of_documents):
bow_representation = np.zeros((number_of_documents, word_vector_size))
labels = np.zeros((number_of_documents, 1))
i = 0
for label, class_name in enumerate(dataset):
# For each file
for f in dataset[class_name]:
# Read all text in file
text = ' '.join(f).split(' ')
# For each word
for word in text:
if word in valid_words:
bow_representation[i, valid_words[word]] += 1
# Label of document
labels[i] = label
# Increment document counter
i += 1
return bow_representation, labels
# Convert the dataset into their bag of words representation treating train and test separately
train_bow_set, train_bow_labels = convert_to_BoW(train_set, n_train)
test_bow_set, test_bow_labels = convert_to_BoW(test_set, n_test)
# + [markdown] colab_type="text" id="P4bTBjw_HCzX"
# ### 1.4 Document classification using Bag-of-Words
#
# For the test documents, use your favorite distance metric (Cosine, Eucilidean, etc.) to find similar news articles from your training set and classify using kNN.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 25310, "status": "ok", "timestamp": 1581572722528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="LoBjz1NSDJME" outputId="cce9e731-3049-4f2f-ba44-a231cd250256"
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=5, metric='euclidean')
model.fit(train_bow_set, train_bow_labels)
# + [markdown] colab_type="text" id="xbGnpvY6HCzc"
# Computing accuracy for the bag-of-words features on the full test set:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 171727, "status": "ok", "timestamp": 1581572904664, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="4CaiubSSDZoI" outputId="3160e552-280d-4fa3-88a5-868a296c8358"
model.score(test_bow_set, test_bow_labels) # This cell may take some time to finish its execution
# + [markdown] colab_type="text" id="_gytBI972w7-"
# ### Not happy with the accuracy above? Stay tuned for other representations (for same/similar algorithms) as opposed to 'Bag Of words', and compare the accuracies.
# + [markdown] colab_type="text" id="dv58hpDAHCzk"
# ### Ungraded Exercise 1
#
# The frequency thresholds represents the minimum frequency a word must have to be considered relevant. Experiment with the following values of frequency threshold in your preprocessing step from section 1.2. Re-run all the codes with the new set of valid words and check your accuracies. Use the following values:
#
# `freq_thresh` =
# * 10
# * 1000
#
# Report the accuracies using bag of words features
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1022, "status": "ok", "timestamp": 1581574083810, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "08508186513102229355"}, "user_tz": -330} id="vugpX4XBHCzk" outputId="3aea0f13-b93e-4300-a574-31b22b75fdcd"
valid_words = defaultdict(int)
print('Number of words before preprocessing:', len(sorted_words))
# Ignore the 25 most frequent words, and the words which appear less than 100 times
ignore_most_frequent = 25
freq_thresh = 1000
feature_number = 0
for word, word_frequency in sorted_words[ignore_most_frequent:]:
if word_frequency > freq_thresh:
valid_words[word] = feature_number
feature_number += 1
print('Number of words after preprocessing:', len(valid_words))
word_vector_size = len(valid_words)
# + colab={} colab_type="code" id="wYBx7l9qGJ5z"
def convert_to_BoW(dataset, number_of_documents):
bow_representation = np.zeros((number_of_documents, word_vector_size))
labels = np.zeros((number_of_documents, 1))
i = 0
for label, class_name in enumerate(dataset):
# For each file
for f in dataset[class_name]:
# Read all text in file
text = ' '.join(f).split(' ')
# For each word
for word in text:
if word in valid_words:
bow_representation[i, valid_words[word]] += 1
# Label of document
labels[i] = label
# Increment document counter
i += 1
return bow_representation, labels
# + colab={} colab_type="code" id="bbhCvlfMGM9L"
# Convert the dataset into their bag of words representation treating train and test separately
train_bow_set, train_bow_labels = convert_to_BoW(train_set, n_train)
test_bow_set, test_bow_labels = convert_to_BoW(test_set, n_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 3406, "status": "ok", "timestamp": 1581574358307, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="_r5OzWfEGSJa" outputId="e4230829-16d5-43c4-926d-7e5741a68db9"
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=5, metric='euclidean')
model.fit(train_bow_set, train_bow_labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 875, "status": "ok", "timestamp": 1581574364615, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="SP_7089JGZS0" outputId="3c924caf-2f30-4036-e4f0-ddd227e22c24"
train_bow_set[:5]
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 1000, "status": "ok", "timestamp": 1581574365938, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="H2_G5oKeG0Tq" outputId="e3dc9b8a-f42f-47b0-ec5f-d9713ea0514b"
train_bow_labels[:5]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 20477, "status": "ok", "timestamp": 1581574387981, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="Q_jqJZXxG5fa" outputId="e52881ba-d8ef-4d7c-d0e3-a4e2cf82067e"
model.score(test_bow_set, test_bow_labels)
# + colab={} colab_type="code" id="B9YBp1PXHGvK"
#freq_thresh - 25 | score - 0.4378947368421053
#freq_thresh - 10 | score - 0.42736842105263156
#freq_thresh - 1000 | score - 0.3463157894736842
# + [markdown] colab_type="text" id="FjteJMM2HCzn"
# ### Ungraded Exercise 2
#
# To classify news articles into their 20 news groups, experiment with the following parameter choices.
#
# * K-NN
# ** K : 10, 50
#
# Report the accuracies using bag of words features.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 4544, "status": "ok", "timestamp": 1581574760664, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="LVmy22wSHCzo" outputId="62293115-9545-4aae-9982-f33cf5cdba48"
valid_words = defaultdict(int)
print('Number of words before preprocessing:', len(sorted_words))
# Ignore the 25 most frequent words, and the words which appear less than 100 times
ignore_most_frequent = 25
freq_thresh = 25
feature_number = 0
for word, word_frequency in sorted_words[ignore_most_frequent:]:
if word_frequency > freq_thresh:
valid_words[word] = feature_number
feature_number += 1
print('Number of words after preprocessing:', len(valid_words))
word_vector_size = len(valid_words)
def convert_to_BoW(dataset, number_of_documents):
bow_representation = np.zeros((number_of_documents, word_vector_size))
labels = np.zeros((number_of_documents, 1))
i = 0
for label, class_name in enumerate(dataset):
# For each file
for f in dataset[class_name]:
# Read all text in file
text = ' '.join(f).split(' ')
# For each word
for word in text:
if word in valid_words:
bow_representation[i, valid_words[word]] += 1
# Label of document
labels[i] = label
# Increment document counter
i += 1
return bow_representation, labels
# Convert the dataset into their bag of words representation treating train and test separately
train_bow_set, train_bow_labels = convert_to_BoW(train_set, n_train)
test_bow_set, test_bow_labels = convert_to_BoW(test_set, n_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 49997, "status": "ok", "timestamp": 1581575421264, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="zWaVhzRNM4aP" outputId="3012520e-3db9-43b1-f97b-cbc8f279ad9e"
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=50, metric='euclidean')
model.fit(train_bow_set, train_bow_labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 454911, "status": "ok", "timestamp": 1581576068867, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mB71PsBYLhuVqmFbtN-6NzmoyqKEkdtw1RZyZYeLw=s64", "userId": "08508186513102229355"}, "user_tz": -330} id="2Pb0P0vgMvLn" outputId="149a6def-8ac6-44f3-8a99-564fef5dc61b"
model.score(test_bow_set, test_bow_labels)
# + colab={} colab_type="code" id="GRNiqRlPPI8R"
#KNN - 10 : 0.3873684210526316
#KNN - 50 : 0.25157894736842107
# + [markdown] colab_type="text" id="-f2KsA0nHCzr"
# ### Summary
# + [markdown] colab_type="text" id="xmNjJflBHCzs"
# Form the above experiment we can observe that the output of the bags of words would be a vector for each individual document. These documents will be parsed through different algorithms to extract the features that are used to classify the text.
| BowMlpMfcc/001_BOW_20newsgroup_C.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Anchor explanations for ImageNet
import tensorflow as tf
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from tensorflow.keras.applications.inception_v3 import InceptionV3, preprocess_input, decode_predictions
from alibi.datasets import fetch_imagenet
from alibi.explainers import AnchorImage
# ### Load InceptionV3 model pre-trained on ImageNet
model = InceptionV3(weights='imagenet')
# ### Download and preprocess some images from ImageNet
# The *fetch_imagenet* function takes as arguments any of the [1000 ImageNet categories](https://github.com/SeldonIO/alibi/tree/master/alibi/data/imagenet_class_names_to_id.json) as well as the number of images to return and the target size of the image.
category = 'Persian cat'
image_shape = (299, 299, 3)
data, labels = fetch_imagenet(category, nb_images=10, target_size=image_shape[:2], seed=2, return_X_y=True)
print('Images shape: {}'.format(data.shape))
# Apply image preprocessing, make predictions and map predictions back to categories. The output label is a tuple which consists of the class name, description and the prediction probability.
images = preprocess_input(data)
preds = model.predict(images)
label = decode_predictions(preds, top=3)
print(label[0])
# ### Define prediction function
predict_fn = lambda x: model.predict(x)
# ### Initialize anchor image explainer
#
# The segmentation function will be used to generate superpixels. It is important to have meaningful superpixels in order to generate a useful explanation. Please check scikit-image's [segmentation methods](http://scikit-image.org/docs/dev/api/skimage.segmentation.html) (*felzenszwalb*, *slic* and *quickshift* built in the explainer) for more information.
#
# In the example, the pixels not in the proposed anchor will take the average value of their superpixel. Another option is to superimpose the pixel values from other images which can be passed as a numpy array to the *images_background* argument.
segmentation_fn = 'slic'
kwargs = {'n_segments': 15, 'compactness': 20, 'sigma': .5}
explainer = AnchorImage(predict_fn, image_shape, segmentation_fn=segmentation_fn,
segmentation_kwargs=kwargs, images_background=None)
# ### Explain a prediction
#
# The explanation of the below image returns a mask with the superpixels that constitute the anchor.
i = 0
plt.imshow(data[i]);
# The *threshold*, *p_sample* and *tau* parameters are also key to generate a sensible explanation and ensure fast enough convergence. The *threshold* defines the minimum fraction of samples for a candidate anchor that need to lead to the same prediction as the original instance. While a higher threshold gives more confidence in the anchor, it also leads to longer computation time. *p_sample* determines the fraction of superpixels that are changed to either the average value of the superpixel or the pixel value for the superimposed image. The pixels in the proposed anchors are of course unchanged. The parameter *tau* determines when we assume convergence. A bigger value for *tau* means faster convergence but also looser anchor restrictions.
image = images[i]
np.random.seed(0)
explanation = explainer.explain(image, threshold=.95, p_sample=.5, tau=0.25)
# Superpixels in the anchor:
plt.imshow(explanation.anchor);
# A visualization of all the superpixels:
plt.imshow(explanation.segments);
| examples/anchor_image_imagenet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from flame_db.gen_insert_data import *
from flame_db.FLAME_db_algorithm import *
from flame_db.matching_helpers import *
from flame_db.utils import *
import unittest
import pandas as pd
import os
import sys
def check_statistics(res_post_new):
ATE_ = ATE_db(res_post_new)
ATT_ = ATT_db(res_post_new)
if type(ATE_) == np.nan:
print("ATE: " + str(ATE_))
return True
if type(ATT_) == np.nan:
print("ATT:" + str(ATT_))
return True
return False
p = 20
TE = 5
gen_data_db(n = 1000,p = 2, TE = TE)
data,weight_array = gen_data_db(n = 1000,p = p, TE = TE)
holdout,weight_array = gen_data_db(n = 500,p = p, TE = TE)
#Connect to the database
select_db = "postgreSQL" # Select the database you are using
database_name='tmp' # database name
host = 'localhost' #host ='vcm-17819.vm.duke.edu' # "127.0.0.1"
port = "5432"
user="newuser"
password= "<PASSWORD>"
conn = connect_db(database_name, user, password, host, port)
insert_data_to_db("test_df_t", # The name of your table containing the dataset to be matched
data,
treatment_column_name= "treated",
outcome_column_name= 'outcome',conn = conn)
df = holdout.copy()
df.loc[0,'treated'] = 4
res_post_new1 = FLAME_db(input_data = "test_df_t", # The name of your table containing the dataset to be matched
holdout_data = df, # holdout set
C = 0.1,
conn = conn,
matching_option = 0,
verbose = 3,
k = 0
)
# +
import numpy as np
import pandas as pd
from flame_db.gen_insert_data import *
from flame_db.FLAME_db_algorithm import *
from flame_db.matching_helpers import *
from flame_db.utils import *
import unittest
import pandas as pd
import os
import sys
def check_statistics(res_post_new):
ATE_ = ATE_db(res_post_new)
ATT_ = ATT_db(res_post_new)
if type(ATE_) == np.nan:
print("ATE: " + str(ATE_))
return True
if type(ATT_) == np.nan:
print("ATT:" + str(ATT_))
return True
return False
p = 20
TE = 5
gen_data_db(n = 100,p = 2, TE = TE)
data,weight_array = gen_data_db(n = 1000,p = p, TE = TE)
holdout,weight_array = gen_data_db(n = 500,p = p, TE = TE)
# Select the database you are using
database_name='tmp' # database name
host ='localhost' # "127.0.0.1"
port = "5432"
user="newuser"
password= "<PASSWORD>"
conn = connect_db(database_name, user, password, host, port, select_db = "MySQL")
#Insert the data into database
insert_data_to_db("test_df101", # The name of your table containing the dataset to be matched
data,
treatment_column_name= "treated",
outcome_column_name= 'outcome',conn = conn)
res_post_new1 = FLAME_db(input_data = "test_df100", # The name of your table containing the dataset to be matched
holdout_data = holdout, # holdout set
treatment_column_name= "treated",
outcome_column_name= 'outcome',
adaptive_weights = 'ridge',
C = 0.1,
conn = conn,
matching_option = 0,
verbose = 3,
k = 0
)
# -
pip uninstall mysql-connector-python--yes
pip install mysql-connector-python
pip install mysql-connector
# +
import numpy as np
import pandas as pd
from flame_db.gen_insert_data import *
from flame_db.FLAME_db_algorithm import *
from flame_db.matching_helpers import *
from flame_db.utils import *
import unittest
import pandas as pd
import os
import sys
# #Generate toy dataset
# p = 20
# TE = 5
# data,weight_array = gen_data_db(n = 5000,p = p, TE = TE)
# holdout,weight_array = gen_data_db(n = 500,p = p, TE = TE)
# #Connect to the database
# select_db = "postgreSQL" # Select the database you are using
# database_name='tmp' # database name
# host ='vcm-17819.vm.duke.edu' # "127.0.0.1"
# port = "5432"
# user="newuser"
# password= "<PASSWORD>"
# conn = connect_db(database_name, user, password, host, port)
# #Insert the data into database
# insert_data_to_db("test_df", # The name of your table containing the dataset to be matched
# data,
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',conn = conn)
def check_statistics(res_post_new):
ATE_ = ATE_db(res_post_new)
ATT_ = ATT_db(res_post_new)
if type(ATE_) == np.nan:
print("ATE: " + str(ATE_))
return True
if type(ATT_) == np.nan:
print("ATT:" + str(ATT_))
return True
return False
# -
class TestFlame_db(unittest.TestCase):
def test_weights(self):
is_corrct = 1
try:
for verbose in [0,1,2,3]:
for matching_option in [0,1,2,3]:
#Test fixed weights
for adaptive_weights in ['ridge', 'decisiontree',False]:
res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
holdout_data = holdout, # holdout set
treatment_column_name= "Treated",
outcome_column_name= 'outcome123',
C = 0.1,
conn = conn,
matching_option = matching_option,
adaptive_weights = adaptive_weights,
weight_array = weight_array,
verbose = verbose,
k = 0
)
if check_statistics(res_post_new):
is_corrct = 0
except (KeyError, ValueError):
is_corrct = 0
self.assertEqual(1, is_corrct,
msg='Error when test weights')
def test_stop_iterations(self):
is_corrct = 1
try:
for early_stop_iterations in [2,3]:
res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
holdout_data = holdout, # holdout set
treatment_column_name= "Treated",
outcome_column_name= 'outcome123',
C = 0.1,
conn = conn,
matching_option = 0,
adaptive_weights = 'decisiontree',
verbose = 1,
k = 0,
early_stop_iterations = early_stop_iterations
)
if check_statistics(res_post_new):
is_corrct = 0
except (KeyError, ValueError):
is_corrct = 0
self.assertEqual(1, is_corrct,
msg='Error when test stop_iterations')
def test_missing_datasets(self):
is_corrct = 1
try:
for missing_data_replace in [0,1,2]:
for missing_holdout_replace in [0,1]:
holdout_miss = holdout.copy()
m,n = holdout_miss.shape
for i in range(int(m/100)):
for j in [0,int(n/2)]:
holdout_miss.iloc[i,j] = np.nan
res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
holdout_data = holdout_miss, # holdout set
treatment_column_name= "Treated",
outcome_column_name= 'outcome123',
C = 0,
conn = conn,
matching_option = 1,
adaptive_weights = 'decisiontree',
verbose = 1,
missing_data_replace = missing_data_replace,
missing_holdout_replace = missing_holdout_replace)
if check_statistics(res_post_new):
is_corrct = 0
except (KeyError, ValueError):
is_corrct = 0
self.assertEqual(1, is_corrct,
msg='Error when test missing datasets')
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
t = TestFlame_db()
t.test_missing_datasets()
t.test_stop_iterations()
# +
p = 20
TE = 5
data,weight_array = gen_data_db(n = 5000,p = p, TE = TE)
holdout,weight_array = gen_data_db(n = 500,p = p, TE = TE)
holdout = pd.read_csv("holdout.csv")
data = pd.read_csv("data.csv")
#Connect to the database
select_db = "postgreSQL" # Select the database you are using
database_name='tmp' # database name
host ='vcm-17819.vm.duke.edu' # "127.0.0.1"
port = "5432"
user="newuser"
password= "<PASSWORD>"
conn = connect_db(database_name, user, password, host, port)
#Insert the data into database
insert_data_to_db("test_df", # The name of your table containing the dataset to be matched
data,
treatment_column_name= "Treated",
outcome_column_name= 'outcome123',conn = conn)
for verbose in [2]:
for matching_option in [0]:
#Test fixed weights
for adaptive_weights in ['ridge']:
res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
holdout_data = holdout, # holdout set
treatment_column_name= "Treated",
outcome_column_name= 'outcome123',
C = 0.1,
conn = conn,
matching_option = matching_option,
adaptive_weights = adaptive_weights,
weight_array = weight_array,
verbose = verbose,
k = 0
)
# -
res_post_new[0][0]['num_control'].sum() + res_post_new[0][0]['num_treated'].sum()
res_post_new[0][1]['num_control'].sum() + res_post_new[0][1]['num_treated'].sum()
res_post_new[0][2]['num_control'].sum() + res_post_new[0][2]['num_treated'].sum()
# +
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0.1,
# conn = conn,
# matching_option = 0,
# adaptive_weights = 'ridge',
# weight_array = weight_array,
# verbose = 2,
# k = 0
# )
# -
holdout
holdout.to_csv("holdout.csv",index = None)
data.to_csv("data.csv",index = None)
# +
# t = TestFlame_db()
# t.test_weights()
# t.test_stop_iterations()
# t.test_missing_datasets()
# +
# #**********************************Test**********************************#
# for C in [0.0, 0.2,0.6]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = C,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'ridge',
# verbose = verbose,
# k = 0
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for matching_option in [0,1,2,3]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0.0,
# conn = conn,
# matching_option = matching_option,
# adaptive_weights = 'ridge',
# verbose = 1,
# k = 0
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for alpha in [0.1,0.8]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# k = 0,
# alpha = alpha,
# early_stop_iterations = 2
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for max_depth in [8,9]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# k = 0,
# max_depth = max_depth,
# early_stop_iterations = 2
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for k in [0,2,4]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# k = k
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for ratio in [0.01,0.1]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# ratio = ratio
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for early_stop_un_c_frac in [0.2,0.5]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# early_stop_un_c_frac = early_stop_un_c_frac
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for early_stop_un_t_frac in [0.2,0.5]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# early_stop_un_t_frac = early_stop_un_t_frac
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for early_stop_pe in [3,5]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# early_stop_pe = early_stop_pe
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for early_stop_pe_frac in [0.5,1]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# early_stop_pe_frac = early_stop_pe_frac
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for missing_data_replace in [0,1,2]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# missing_data_replace = missing_data_replace
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# for missing_holdout_replace in [0,1]:
# res_post_new = FLAME_db(input_data = "test_df", # The name of your table containing the dataset to be matched
# holdout_data = holdout, # holdout set
# treatment_column_name= "Treated",
# outcome_column_name= 'outcome123',
# C = 0,
# conn = conn,
# matching_option = 1,
# adaptive_weights = 'decisiontree',
# verbose = 1,
# missing_holdout_replace = missing_holdout_replace
# )
# print(ATE_db(res_post_new))
# print(ATT_db(res_post_new))
# -
class Test_exceptions(unittest.TestCase):
def test_false_dataset(self):
def broken_false_dataset():
df, true_TE = generate_uniform_given_importance(num_control=1000, num_treated=1000)
holdout, true_TE = generate_uniform_given_importance(num_control=1000, num_treated=1000)
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(False)
with self.assertRaises(Exception) as false_dataset:
broken_false_dataset()
self.assertTrue("Need to specify either csv file name or pandas data "\
"frame in parameter 'input_data'" in str(false_dataset.exception))
def test_false_early_stop_un_t_frac(self):
def broken_early_stop_un_t_frac():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(early_stop_un_t_frac = -1)
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as early_stop_un_t_frac:
broken_early_stop_un_t_frac()
self.assertTrue('The value provided for the early stopping critera '\
'of proportion of unmatched treatment units needs to '\
'be between 0.0 and 1.0' in str(early_stop_un_t_frac.exception))
def test_false_early_stop_un_c_frac(self):
def broken_early_stop_un_c_frac():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(early_stop_un_c_frac = -1)
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as early_stop_un_c_frac:
broken_early_stop_un_c_frac()
self.assertTrue('The value provided for the early stopping critera '\
'of proportion of unmatched control units needs to '\
'be between 0.0 and 1.0' in str(early_stop_un_c_frac.exception))
def test_false_early_stop_iterations(self):
def broken_early_stop_iterations():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(early_stop_iterations = True)
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as early_stop_iterations:
broken_early_stop_iterations()
self.assertTrue('The value provided for early_stop_iteration needs '\
'to be an integer number of iterations, or False if '\
'not stopping early based on the number of iterations' in str(early_stop_iterations.exception))
def test_false_early_stop_pe_frac(self):
def broken_early_stop_pe_frac():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(early_stop_pe_frac = 123)
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as early_stop_pe_frac:
broken_early_stop_pe_frac()
self.assertTrue('The value provided for the early stopping critera of'\
' PE needs to be between 0.0 and 1.0' in str(early_stop_pe_frac.exception))
def test_false_weight_array_type(self):
def broken_weight_array_type():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(adaptive_weights = False)
model.fit(holdout_data=df, weight_array = np.array([1,2,3,4,5]))
output = model.predict(df)
with self.assertRaises(Exception) as weight_array_type:
broken_weight_array_type()
self.assertTrue('Invalid input error. A weight array of type'\
'array needs to be provided when the'\
'parameter adaptive_weights == True' in str(weight_array_type.exception))
def test_false_weight_array_len(self):
def broken_weight_array_len():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(adaptive_weights = False)
model.fit(holdout_data=df, weight_array = [1])
output = model.predict(df)
with self.assertRaises(Exception) as weight_array_len:
broken_weight_array_len()
self.assertTrue('Invalid input error. Weight array size not equal'\
' to number of columns in dataframe' in str(weight_array_len.exception))
def test_false_weight_array_sum(self):
def broken_weight_array_sum():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(adaptive_weights = False)
model.fit(holdout_data=df, weight_array = [1,1,1,1])
output = model.predict(df)
with self.assertRaises(Exception) as weight_array_sum:
broken_weight_array_sum()
self.assertTrue('Invalid input error. Weight array values must '\
'sum to 1.0' in str(weight_array_sum.exception))
def test_false_alpha(self):
def broken_alpha():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(adaptive_weights = 'ridge',alpha = -10)
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as alpha:
broken_alpha()
self.assertTrue('Invalid input error. The alpha needs to be '\
'positive for ridge regressions.' in str(alpha.exception))
def test_false_adaptive_weights(self):
def broken_adaptive_weights():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
model = matching.FLAME(adaptive_weights = 'safdsaf')
model.fit(holdout_data=df)
output = model.predict(df)
with self.assertRaises(Exception) as adaptive_weights:
broken_adaptive_weights()
self.assertTrue("Invalid input error. The acceptable values for "\
"the adaptive_weights parameter are 'ridge', "\
"'decisiontree', 'decisiontreeCV', or 'ridgeCV'. Additionally, "\
"adaptive-weights may be 'False' along "\
"with a weight array" in str(adaptive_weights.exception))
def test_false_data_len(self):
def broken_data_len():
df, true_TE = generate_uniform_given_importance(num_control=1000, num_treated=1000,
num_cov=7, min_val=0,
max_val=3, covar_importance=[4,3,2,1,0,0,0])
holdout, true_TE = generate_uniform_given_importance()
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df)
with self.assertRaises(Exception) as data_len:
broken_data_len()
self.assertTrue('Invalid input error. The holdout and main '\
'dataset must have the same number of columns' in str(data_len.exception))
def test_false_column_match(self):
def broken_column_match():
df, true_TE = generate_uniform_given_importance()
holdout, true_TE = generate_uniform_given_importance()
set_ = holdout.columns
set_ = list(set_)
set_[0] = 'dasfadf'
holdout.columns = set_
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df)
with self.assertRaises(Exception) as column_match:
broken_column_match()
self.assertTrue('Invalid input error. The holdout and main '\
'dataset must have the same columns' in str(column_match.exception))
def test_false_C(self):
def broken_C():
df, true_TE = generate_uniform_given_importance()
holdout, true_TE = generate_uniform_given_importance()
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df,C = -1)
with self.assertRaises(Exception) as C:
broken_C()
self.assertTrue('The C, or the hyperparameter to trade-off between'\
' balancing factor and predictive error must be '\
' nonnegative. 'in str(C.exception))
def test_false_missing_data_replace(self):
def broken_missing_data_replace():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100,
num_cov=7, min_val=0,
max_val=3, covar_importance=[4,3,2,1,0,0,0])
holdout, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100,
num_cov=7, min_val=0,
max_val=3, covar_importance=[4,3,2,1,0,0,0])
covar_importance = np.array([4,3,2,1,0,0,0])
weight_array = covar_importance/covar_importance.sum()
model = matching.FLAME(missing_data_replace = 2, adaptive_weights =False)
model.fit(holdout_data=holdout,weight_array = list(weight_array))
output = model.predict(df)
with self.assertRaises(Exception) as missing_data_replace:
broken_missing_data_replace()
self.assertTrue('Invalid input error. We do not support missing data '\
'handing in the fixed weights version of algorithms'in str(missing_data_replace.exception))
def test_false_treatment_column_name(self):
def broken_treatment_column_name():
df, true_TE = generate_uniform_given_importance()
holdout, true_TE = generate_uniform_given_importance()
model = matching.FLAME()
model.fit(holdout_data=holdout,treatment_column_name = "sadfdag")
output = model.predict(df)
with self.assertRaises(Exception) as treatment_column_name:
broken_treatment_column_name()
self.assertTrue('Invalid input error. Treatment column name does not'\
' exist' in str(treatment_column_name.exception))
def test_false_outcome_column_name(self):
def broken_outcome_column_name():
df, true_TE = generate_uniform_given_importance()
holdout, true_TE = generate_uniform_given_importance()
model = matching.FLAME()
model.fit(holdout_data=holdout,outcome_column_name = "sadfdag")
output = model.predict(df)
with self.assertRaises(Exception) as outcome_column_name:
broken_outcome_column_name()
self.assertTrue('Invalid input error. Outcome column name does not'\
' exist' in str(outcome_column_name.exception))
def test_false_treatment_column_name_value(self):
def broken_treatment_column_name_value():
df, true_TE = generate_uniform_given_importance()
holdout, true_TE = generate_uniform_given_importance()
df.loc[0,'treated'] = 4
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df)
with self.assertRaises(Exception) as treatment_column_name_value:
broken_treatment_column_name_value()
self.assertTrue('Invalid input error. All rows in the treatment '\
'column must have either a 0 or a 1 value.' in str(treatment_column_name_value.exception))
def test_false_data_type(self):
def broken_data_type():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
holdout = df.copy()
df.iloc[0,0] = 's'
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df)
with self.assertRaises(Exception) as _data_type:
broken_data_type()
self.assertTrue('Invalid input error on matching dataset. Ensure all inputs asides from '\
'the outcome column are integers, and if missing' \
' values exist, ensure they are handled.' in str(_data_type.exception))
def test_false_holdout_type(self):
def broken_holdout_type():
df, true_TE = generate_uniform_given_importance(num_control=100, num_treated=100)
holdout = df.copy()
holdout.iloc[0,0] = 's'
model = matching.FLAME()
model.fit(holdout_data=holdout)
output = model.predict(df)
with self.assertRaises(Exception) as holdout_type:
broken_holdout_type()
self.assertTrue('Invalid input error on holdout dataset. Ensure all inputs asides from '\
'the outcome column are integers, and if missing' \
' values exist, ensure they are handled.' in str(holdout_type.exception))
def test_false_ATE_input(self):
def broken_ATE_input():
ATE(1)
with self.assertRaises(Exception) as ATE_input:
broken_ATE_input()
self.assertTrue("The matching_object input parameter needs to be "\
"of type DAME or FLAME" in str(ATE_input.exception))
def test_false_ATE_input_model(self):
def broken_ATE_input_model():
model = matching.FLAME()
ATE(model)
with self.assertRaises(Exception) as ATE_input_model:
broken_ATE_input_model()
self.assertTrue("This function can be only called after a match has "\
"been formed using the .fit() and .predict() functions" in str(ATE_input_model.exception))
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# name: python3
# ---
# ## Canada - Admissions of Permanent Residents by Country of Citizenship and Immigration Category, January 2015 - December 2020
# IRCC also provide the immigration statistics grouped by the immigration category (ie., FSW, CEC, PNP, etc.) for each individual country of citizenship. The information is stored in a compelex table with multiple level of indexing. This notebook demonstrates how to clean up such data and draw some insight using Pandas.
#
# By analyzing the admission data, we try to answer the following questions:
#
# - Question 1: What are the most popupar immigration category for people with citizenship of China?
# - Question 2: What are the trend for these categories over time?
import pandas as pd
import os
import xlrd
import matplotlib.pyplot as plt
import numpy as np
from clean_col import clean_col # User defined function
from clean_indx import cleaner # User defined function
from multiplot import multiplot # User defined function
# ### Loading and examing the raw data
# Loading raw data download from IRCC website
file = os.path.join(os.getcwd(),'ircc_m_pradmiss_0013_e.xls')
# Read data and remove header & footer
data = pd.read_excel(file,skiprow=4,header=[4],skipfooter=5)
data.head()
# The table contains two parts, the 1st part inclues details for each individual countries, and the 2nd part inclues the summary.
#
# Because of the multi-level indexing and merged cells, there are many NaN values created in the index columns. Simply using the **index_col** argment does not work out due to the abnormal design of the table. Therefore, a custom funtion is design to clean up the data.
# +
#Separate the table into two parts
indv = data.iloc[0:3981,:]
summ = data.iloc[3982:,:]
indv = cleaner(indv) # Clean index with the cleaner function
col_names = [clean_col(i) for i in indv.columns] # Clean column names with the clean_col function
indv.columns = col_names
#Convert all missing value '--' to 0
indv = indv.applymap(lambda x: 0 if x == '--' else int(x))
indv.head()
# -
# ### Checking the data after cleaning
#
# To validate the data cleaning process, a few queries are performed. The data is intact.
#
indv.loc['Australia - Total','Economic - Total','Worker Program']
# ### Analyze Data
#
# Question 1: What are the most popupar immigration category for people with citizenship of China?
# Get the information for China
China = indv.loc['China, People\'s Republic of - Total'][['Total, 2015','Total, 2016','Total, 2017','Total, 2018','Total, 2019','Total, 2020']]
China
# +
#Since there are too many sub-categories, the Program_Lv1 is used to generate the pie chart
Economic = China.loc['Economic - Total','Economic - Total','Economic - Total'].values[0]
Sponsored_Family = China.loc['Sponsored Family - Total','Sponsored Family - Total','Sponsored Family - Total'].values[0]
Refugee_Protected_Person = China.loc['Resettled Refugee & Protected Person in Canada - Total','Resettled Refugee & Protected Person in Canada - Total','Resettled Refugee & Protected Person in Canada - Total'].values[0]
Other = China.loc['All Other Immigration - Total','All Other Immigration - Total','All Other Immigration - Total'].values[0]
pie = [sum(Economic),sum(Refugee_Protected_Person),sum(Sponsored_Family),sum(Other)]
plt.pie(x=pie, autopct="%.1f%%", explode=[0.05]*4, labels=['Economic','Refugee','Family','Other'], pctdistance=0.5)
plt.title("Immigration by Category, China\n", fontsize=14);
# -
#
# Question 2: What are the trend for these categories over time?
# +
trend = pd.DataFrame([Economic,Sponsored_Family,Refugee_Protected_Person,Other],columns=['2015','2016','2017','2017','2019','2020'],
index = ['Economic','Family','Refugee','Other'])
plt.figure(figsize=(8,6))
plt.ylabel('Number of People')
multiplot(trend)
# -
| Project_IRCC/By_category.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
import sklearn.preprocessing
from sklearn.model_selection import train_test_split
import pandas as pd
# # Setup
# +
np.random.seed(123)
x = stats.skewnorm(7).rvs(1500) * 10 + 100
x = x.reshape(-1, 1)
plt.hist(x, bins=25,ec='black')
print('Here is a histogram of the dataset we will be working with.')
# -
x_train_and_validate, x_test = train_test_split(x, random_state=123)
x_train, x_validate = train_test_split(x_train_and_validate)
# ## Min-Max Scaling
#
# Min-max scaling is a linear scaling method that transforms our features such that the range is between 0 and 1.
# +
scaler = sklearn.preprocessing.MinMaxScaler()
# Note that we only call .fit with the training data,
# but we use .transform to apply the scaling to all the data splits.
scaler.fit(x_train)
x_train_scaled = scaler.transform(x_train)
x_validate_scaled = scaler.transform(x_validate)
x_test_scaled = scaler.transform(x_test)
plt.figure(figsize=(13, 6))
plt.subplot(121)
plt.hist(x_train, bins=25, ec='black')
plt.title('Original')
plt.subplot(122)
plt.hist(x_train_scaled, bins=25, ec='black')
plt.title('Scaled')
# -
# ## Standard Scaler
#
# Standardization is a linear transformation of our data such that is looks like the standard normal distribution. That is, it will have a mean of 0 and a standard deviation of 1.
# Sometimes this is split into two operations:
#
# scaling is dividing each data point by the standard deviation. This causes the resulting dataset to have a standard deviation of 1.
# centering is subtracting the mean from each data point. This causes the resulting dataset to have a mean of 0.
#
# +
scaler = sklearn.preprocessing.StandardScaler()
# Note that we only call .fit with the training data,
# but we use .transform to apply the scaling to all the data splits.
scaler.fit(x_train)
x_train_scaled = scaler.transform(x_train)
x_validate_scaled = scaler.transform(x_validate)
x_test_scaled = scaler.transform(x_test)
plt.figure(figsize=(13, 6))
plt.subplot(121)
plt.hist(x_train, bins=25, ec='black')
plt.title('Original')
plt.subplot(122)
plt.hist(x_train_scaled, bins=25, ec='black')
plt.title('Scaled')
# -
# ## RobustScaler
#
# A robust scaler is another linear transformation that follows the same idea as the standard scaler but uses parameters that are more robust to outliers.
# +
scaler = sklearn.preprocessing.RobustScaler()
# Note that we only call .fit with the training data,
# but we use .transform to apply the scaling to all the data splits.
scaler.fit(x_train)
x_train_scaled = scaler.transform(x_train)
x_validate_scaled = scaler.transform(x_validate)
x_test_scaled = scaler.transform(x_test)
plt.figure(figsize=(13, 6))
plt.subplot(121)
plt.hist(x_train, bins=25, ec='black')
plt.title('Original')
plt.subplot(122)
plt.hist(x_train_scaled, bins=25, ec='black')
plt.title('Scaled')
# +
#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# -
import env
| scaling_lesson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies
import requests
url = "http://www.omdbapi.com/?apikey=trilogy&t="
# -
# Who was the director of the movie Aliens?
movie = requests.get(url + "Aliens").json()
print(f'The director of Aliens was {movie["Director"]}.')
# What was the movie Gladiator rated?
movie = requests.get(url + "Gladiator").json()
print(f'The rating of Gladiator was {movie["Rated"]}.')
# What year was 50 First Dates released?
movie = requests.get(url + "50 First Dates").json()
print(f'The movie 50 First Dates was released in {movie["Year"]}.')
# Who wrote Moana?
movie = requests.get(url + "Moana").json()
print(f'Moana was written by {movie["Writer"]}.')
# What was the plot of the movie Sing?
movie = requests.get(url + "Sing").json()
print(f'The plot of Sing was: {movie["Plot"]}')
| 1/Activities/08-Stu_MovieQuestions/Solved/Stu_MovieQuestions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: zenv
# language: python
# name: zenv
# ---
# Based off of the efficientnet in Timm: https://github.com/rwightman/pytorch-image-models/blob/master/timm/models/efficientnet.py we will reporpose this architecture for our use with 1-dimension sequence data
import torch
import numpy as np
import pandas as pd
import os
import h5py
from exabiome.nn.loader import read_dataset, LazySeqDataset
import argparse
from torch.utils.data import Dataset, DataLoader
import torch.nn as nn
import torch.nn.functional as F
path = '/global/homes/a/azaidi/ar122_r202.toy.input.h5'
hparams = argparse.Namespace(**{'load': False,
'window': 4096,
'step': 4096,
'classify': True,
'tgt_tax_lvl': "phylum",
'fwd_only': True})
chunks = LazySeqDataset(hparams, path=path, keep_open=True)
len(chunks)
def old_pad_seq(seq):
if(len(seq) < 4096):
padded = torch.zeros(4096)
padded[:len(seq)] = seq
return padded
else:
return seq
class taxon_ds(Dataset):
def __init__(self, chunks, transform=None):
self.chunks = chunks
self.transform = transform
def __len__(self):
return len(self.chunks)
def __getitem__(self, idx):
x = chunks[idx][1]
if self.transform:
x = self.transform(x)
y = chunks[idx][2]
return (x.unsqueeze(0), y)
# %time
ds = taxon_ds(chunks, old_pad_seq)
dl = DataLoader(ds, batch_size=16, shuffle=True)
len(dl)
batch = next(iter(dl))
batch[0].shape, batch[1].shape
# # An Efficientnet has basically three parts:
# **(0) Base (Feet) --> (1) Body --> (2) Head**
#
# Within these three parts -- we are **mainly** only using three tools/units of computation:
#
# (0) Conv1d: https://pytorch.org/docs/stable/generated/torch.nn.Conv1d.html <br>
# (1) BatchNorm1d: https://pytorch.org/docs/stable/generated/torch.nn.BatchNorm1d.html <br>
# (2) SiLU: https://pytorch.org/docs/stable/generated/torch.nn.SiLU.html <br>
#
# *There are a few other items that are added as well, that we will see below
#
# <br>**Base** (feet):<br>
# 0) Conv1d --> 1) BatchNorm1d --> 2) SiLU
#
# **Head**: <br>
# (0) Conv1d --> (1) BatchNorm1d --> (2) SiLU --> (3) SelectAdaptivePool1d --> (4) Linear
#
# *the base & head are relatively straightforward -- we'll implement both below:*
def get_conv_bn(in_ch=1, out_ch=2, ks=2, stride=2, padding=None):
return nn.Sequential(
nn.Conv1d(in_channels = in_ch, out_channels = out_ch,
kernel_size = ks, stride = stride,
padding=padding, bias=False),
nn.BatchNorm1d(num_features = out_ch)
)
def conv(in_ch, out_ch, ks, stride, padding=0, activation=False):
res = get_conv_bn(in_ch, out_ch, ks, stride, padding)
if activation:
res = nn.Sequential(res, nn.SiLU(inplace=True))
return res
# Let's make a function to add the SiLU layer
conv(1,2,3,4, activation=True)
# This was the old base layer fxn
def get_base_layer(in_chans=1, out_chans=32, ks=3, stride=2, padding=1):
return nn.Sequential(
nn.Conv1d(in_channels= in_chans, out_channels= out_chans,
kernel_size= ks, stride= stride,
padding=padding, bias=False),
nn.BatchNorm1d(num_features = out_chans),
nn.SiLU(inplace=True))
# Now we can just use our conv function to replace that + this will be the building block for the entire model
def get_base_layer(in_ch=1, out_ch=32, ks=3, stride=2, padding=1):
return conv(in_ch=in_ch, out_ch=out_ch, ks=ks,
stride=stride, padding=padding, activation=True)
get_base_layer()(batch[0]).shape
# The head had a bit more going on, but we can still simplify it
def get_head_layer(in_chans=320, out_chans=1280, ks=1, stride=1,
avg_out_feats=10, lin_out_feats=1):
return nn.Sequential(
nn.Conv1d(in_channels= in_chans, out_channels= out_chans,
kernel_size= ks, stride= stride, bias=False),
nn.BatchNorm1d(num_features = out_chans),
nn.SiLU(inplace=True),
nn.AdaptiveAvgPool1d(output_size=avg_out_feats),
nn.Linear(in_features=avg_out_feats, out_features=lin_out_feats))
get_head_layer()
def get_head_layer(in_chans=320, out_chans=1280, ks=1, stride=1,
avg_out_feats=200, lin_out_feats=1):
return nn.Sequential(
conv(in_chans, out_chans, ks, stride, activation=True),
nn.AdaptiveAvgPool1d(output_size=avg_out_feats),
nn.Linear(in_features=avg_out_feats, out_features=lin_out_feats))
get_head_layer()
# Not too much shorter, but better nonetheless
# **Body**:<br>
# (0) DepthwiseSeparableConv <br>
# (1) InvertedResidual (two in a row) <br>
# (2) InvertedResidual (two in a row) <br>
# (3) InvertedResidual (three in a row) <br>
# (4) InvertedResidual (three in a row) <br>
# (5) InvertedResidual (three in a row) <br>
# (6) InvertedResidual (one) <br>
#
# *ok so what are these layers in the body?*
# # DepthwiseSeperable:
# (0) Conv1d <br>
# (1) BatchNorm1d <br>
# (2) SiLU <br>
# (3) **Squeeze Excite**<br>
# (4) Conv1d <br>
# (5) BatchNorm1d <br>
# (6) Identity <br>
# # InvertedResidual:
# (0) Conv1d <br>
# (1) BatchNorm1d <br>
# (2) SiLU <br>
# (3) Conv1d <br>
# (4) BatchNorm1d <br>
# (5) SiLU <br>
# (6) **Squeeze Excite**<br>
# (7) Conv1d <br>
# (8) BatchNorm1d <br>
# **"Squeeze Excite" = Conv1d --> SiLU --> Conv1d**
#
# Let's define our squeeze excite function -- since we have two conv layers, let's use tuples for our parameters for now -- the parameters in the paper are much more structured for the squueze excite layer, but we will keep this optionality in place (for now)
#
# In the paper the squueze excite takes the number of filters from 240 --> 10 --> 240. This would be easier to encode into the function below, but would make it harder to tweak these values
def get_sq_ex(in_ch= (1,1), out_ch= (2,2), ks= (2,2), stride= (2,2)):
return nn.Sequential(
nn.Conv1d(in_channels= in_ch[0], out_channels= out_ch[0],
kernel_size= ks[0], stride= stride[0]),
nn.SiLU(),
nn.Conv1d(in_channels= in_ch[1], out_channels= out_ch[1],
kernel_size= ks[1], stride= stride[1])
)
#uncomment to confirm the above function works
get_sq_ex()
# The above functions have simplified our work to produce the desired layers -- we have everything we need to create both the layer types in our models body
#
# **DepthwiseSeperable**: <br>
# (0) conv<br>
# (1) get_sq_ex <br>
# (2) conv <br>
# (3) Identity <br>
#
# **InvertedResidual**: <br>
# (0) conv <br>
# (1) conv <br>
# (2) get_sq_ex <br>
# (3) conv <br>
# A squeeze-excite unit compresses the number of channels down and then expands it back to the original amount
def get_dep_sep(in_ch, out_ch, ks=3, mid_ch=8):
return nn.Sequential(
conv(in_ch=in_ch, out_ch=in_ch, ks=ks, stride=1, activation=True),
get_sq_ex(in_ch=(in_ch, mid_ch),
out_ch=(mid_ch, in_ch)),
conv(in_ch=in_ch, out_ch=out_ch, ks=1, stride=1),
nn.Identity()
)
get_dep_sep(32, 16)
#let's just make sure things are moving forward with our depthwise seperable layer
model = nn.Sequential(
get_base_layer(),
get_dep_sep(32, 16))
model(batch[0]).shape
get_sq_ex((96,4),(4,96))
nn.Sequential(
model,
conv(16,96,1,1, activation=True),
conv(96,96, ks=3,stride=2,padding=1, activation=True),
get_sq_ex((96,4), (4,96), ks=(1,1), stride=(1,1)),
conv(96, 24, 1,1)
)(batch[0]).shape
# (1) The first conv layer in the inverted residuals ALWAYS has a kernel size of 1 and stride of 1, with no padding.
#
# (2) For the second conv layer, the kernel size, stride and padding can be different in each layer
#
# (3) The squeeze excite layer always has a kernel size of 1 and stride of 1
#
# (4) The last conv layer always has a stride of 1 and kernel size of 1
def get_inv_res(in_ch, mid_ch, out_ch, sq_ch=4, ks=1, stride=1, padding=1):
return nn.Sequential(
conv(in_ch=in_ch, out_ch=mid_ch, ks=1, stride=1, activation=True),
conv(in_ch=mid_ch, out_ch=mid_ch, ks=ks, stride=stride,
padding=padding, activation=True),
get_sq_ex((mid_ch,sq_ch), (sq_ch, mid_ch) ,ks=(1,1), stride=(1,1)),
conv(in_ch=mid_ch, out_ch=out_ch, ks=1, stride=1)
)
nn.Sequential(
model,
get_inv_res(16,96,24, 4, ks=3, stride=2,padding=1))(batch[0]).shape
nn.Sequential(
get_base_layer(),
get_dep_sep(in_ch=32,out_ch=16),
get_inv_res(in_ch=16, mid_ch=96, out_ch= 24,
sq_ch=4, ks=3, stride=2, padding=1)
)(batch[0]).shape
# Looks like things are working! :)
| effnet_redux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Illustration of converting a ABFv1 format ephys file to NWBv2
# This notebook goes through an example of converting a single [Axon Binary Format version 1 file (ABFv1)](https://swharden.com/pyabf/abf1-file-format.md.html) into a a single [Neurodata Without Borders version 2 file (NWBv2)](https://nwb-schema.readthedocs.io/en/latest/format.html#intracellular-electrophysiology).
#
# TODO: Stitch together multiple ABFv1 files collected from the same single cell (like for different stimulus protocols) into a single NWBv2 file.
# ## Clone and import repository for ABFv1 to NWBv2 conversion developed by Young and adapted slightly by Shreejoy
#
# The git clone command commented out below pulls the ABF and NWB code repo from github
# #! git clone https://github.com/stripathy/NWB.git # uncomment this if you don't have the repo
from NWB.ABF1Converter import ABF1Converter ### import the converter class
# ## Define a function that calls the ABFv1 converter class and sets inputs and outputs
#
# The below file takes a single ABFv1 file, a bunch of arguments, and then outputs a single NWBv2 file in a form that can be directly parsed by IPFX tools.
# +
# import functions
import os
import sys
import glob
import argparse
def abf_to_nwb(inputPath, outFolder, outputMetadata, acquisitionChannelName, stimulusChannelName, overwrite,
responseGain = 1.0, stimulusGain = 1.0, responseOffset = 0.0, clampMode = 1):
"""
Sample file handling script for NWB conversion.
Takes the path to the ABF v1 file as the first command line argument and writes the corresponding NWB file
to the folder specified by the second command line argument.
NWB Files organized by cell, with assumption that each abf file corresponds to each cell
"""
if not os.path.exists(inputPath):
raise ValueError(f"The file or folder {inputPath} does not exist.")
if not os.path.exists(outFolder):
raise ValueError(f"The file or folder {outFolder} does not exist.")
# Collect ABF files from the specified directory.
if os.path.isfile(inputPath):
files = [inputPath]
elif os.path.isdir(inputPath):
files = glob.glob(inputPath + "/*.abf")
else:
raise ValueError(f"Invalid path {inputPath}: input must be a path to a file or a directory")
if len(files) == 0:
raise ValueError(f"Invalid path {inputPath} does not contain any ABF files.")
for inputFile in files:
fileName = os.path.basename(inputFile)
root, _ = os.path.splitext(fileName)
print(f"Converting {fileName}...")
# Generate name for new NWB file
outFile = os.path.join(outFolder, root + ".nwb")
if os.path.exists(outFile):
if overwrite:
os.unlink(outFile)
else:
raise ValueError(f"The file {outFile} already exists.")
# Enter each ABF file into the converter script. The additional arguments are meant for command line operations.
conv = ABF1Converter(inputFile, outFile,
acquisitionChannelName=acquisitionChannelName, stimulusChannelName=stimulusChannelName,
responseGain = responseGain,
stimulusGain = stimulusGain,
responseOffset = responseOffset, clampMode = clampMode
)
conv.convert()
if outputMetadata:
conv._outputMetadata()
# -
# ## Import metadata from Valiante Lab datasets compiled by Shreejoy
# +
import pandas as pd
import pyabf
# read final csv that has the ouput of metadata gathering process
csv_meta_save_path = 'example_datasets/valiante_lab/cell_final_raw_meta_df.csv'
cell_final_raw_meta_df = pd.read_csv(open(csv_meta_save_path, 'rb'))
# an example abf file in the example datasets folder
curr_file = '13d02049.abf'
abf_file_path = file_rel_path + curr_file
abf = pyabf.ABF(abf_file_path)
#stim_abf = pyabf.ABF(stim_file_path) # for some files we're using stim traces from a different file
# -
# ## Get metadata variables that are needed for conversion
#
# Shreejoy comment: these calls are pretty ugly and probably super inefficient
# It would be nice to convert these parameter lists to a dictionary or something
# +
# get the row for the cell from the csv file that we just imported
meta_row = cell_final_raw_meta_df.loc[cell_final_raw_meta_df['cell_id'] == curr_file]
num_sweeps = int(meta_row['num_sweeps'].values[0])
stim_channel_num = int(meta_row['stim_chan'].values[0])
response_chan_num = int(meta_row['resp_chan'].values[0])
acq_channel_name = abf.adcNames[response_chan_num]
if stim_name == 'sweepC':
stim_chan_name = abf.dacNames[stim_channel_num]
else:
stim_chan_name = abf.adcNames[stim_channel_num]
stim_gain = meta_row['stim_gain'].values[0]
if stim_gain == 1000:
stim_gain = 1.0
response_gain = meta_row['resp_gain'].values[0]
start_time = meta_row['stim_start_time'].values[0]
end_time = meta_row['stim_end_time'].values[0]
resp_sampling_rate = meta_row['resp_sampling_rate'].values[0]
resp_offset = meta_row['resp_offset'].values[0]
responseGain = response_gain
responseOffset = resp_offset
stimulusGain = stim_gain
clampMode = 1 # this is current clamp
# -
# ## Perform the ABF to NWB conversion!
# +
# sets the folder for output files
outFolder = 'example_datasets/valiante_lab/'
outputMetadata = True
abf_to_nwb(inputPath=abf_file_path,
outFolder=outFolder,
outputMetadata=False,
acquisitionChannelName=acq_channel_name,
stimulusChannelName=stim_chan_name,
overwrite=True,
responseGain = responseGain,
stimulusGain = stimulusGain,
responseOffset = responseOffset, clampMode = clampMode)
# -
# ## Illustrate loading of converted NWB dataset into IPFX
from ipfx.dataset import hbg_nwb_data
from ipfx.data_set_features import extract_data_set_features
from ipfx.dataset.create import create_ephys_data_set
import ipfx
# +
nwb_file_name = outFolder + '13d02049.nwb'
print(nwb_file_name)
data_set = create_ephys_data_set(nwb_file=nwb_file_name) ##loads the NWB as an HBG dataset, Equal to the nwb1.0 -> AIBS dataset
#data_set = hbg_nwb_data.HBGNWBData(nwb_file=nwb_file_name, ontology = ipfx.) ##loads the NWB as an HBG dataset, Equal to the nwb1.0 -> AIBS dataset
# -
# this doesn't work? why?
pd.DataFrame(data_set.extract_sweep_stim_info())
# ## Illustrate plotting of a sweep with IPFX tools
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from ipfx.sweep import Sweep, SweepSet
sweep_plot_index = 13
curr_sweep = data_set.sweep_set([0, 13]).sweeps[1]
t = curr_sweep.t
v = curr_sweep.v
i = curr_sweep.i
fig, axes = plt.subplots(2, 1, sharex=True)
axes[0].plot(t, v)
axes[0].set_xlim(0, 2)
axes[0].set_ylabel("mV")
axes[1].plot(t, i, c="orange")
axes[1].set_ylabel("pA")
sns.despine()
# -
| ABFv1_to_NWB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Reference: <br>
# https://www.tensorflow.org/tutorials/keras/regression?hl=ja <br>
# https://archive.ics.uci.edu/ml/datasets/glass+identification
# +
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
# -
dataset_path = keras.utils.get_file("glass.data", "https://archive.ics.uci.edu/ml/machine-learning-databases/glass/glass.data")
# +
column_names = ['Id number','RI','Na','Mg','Al','Si', 'K', 'Ca','Ba', 'Fe', 'Type of glass']
raw_dataset = pd.read_csv(dataset_path, names = column_names,na_values = "?", comment = '\t',sep=",", skipinitialspace=True)
dataset = raw_dataset.copy()
print(dataset.shape)
dataset.head()
# -
u = dataset['Type of glass'].nunique()
print(u)
dataset.to_csv('glass.csv')
| [1]_Dataset_preparation/Glass_dataset_preparation_UCI_ML_repository.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# #### Simulate with exponential interarrival time
# * Counting processs N(1) follows poisson distribution.
# * Let $Y = N(t) + 1$.
# * Let $t_n$ denotes the occurance time of the n-th event
# * Number of events that occurs befor time 1 follows poisson distribution: $Y = min(n\ge1; t_n>1)$
# * Get poisson as $N (1) = Y − 1$
# * Recall that exponential distribution can be simulated as $-(1/\alpha)*log(1 - U_i)$ or $-(1/\alpha)*log(U_i)$
#
# <img src="figs/poisson.png" alt="Drawing" style="height: 110px;"/>
#
# ref: http://www.columbia.edu/~ks20/4404-Sigman/4404-Notes-ITM.pdf
def poisson(alpha = 5):
uni = np.random.rand(alpha * 100)
expo = - (1 / alpha) * np.log(uni)
s = 0
N = 0
while s < 1:
s += expo[N]
N += 1
return N - 1
# #### Observation
# * When lambda = 1, Poisson reduces to Exponential distribution.
# * When lambda is large, Poisson looks like normal.
# +
lamb = 3 # count / unit of time
m = [poisson(lamb) for _ in range(10000)]
plt.hist(m, density=True)
mean_empirical = np.mean(m)
var_empirical = np.var(m)
mean_analytical = lamb
var_analytical = lamb
print("""
mean empirical: %.2f
mean analytical: %.2f
variance empirical: %.2f
variance analytical: %.2f
"""%(mean_empirical, mean_analytical,
var_empirical, var_analytical))
plt.show()
# -
def plot_poisson(m, ax):
mean_empirical = np.mean(m)
std_empirical = np.std(m)
normal = np.random.normal(lamb, np.sqrt(lamb), len(m))
ax.hist(m, density=True, bins=20, alpha=0.5)
ax.hist(normal, density=True, bins=20, alpha=0.5)
ax.legend(["poisson lambda=%i"%lamb, "normal mean=%i"%lamb])
# +
plt.figure(figsize=(15, 4))
ax1 = plt.subplot(1, 3, 1)
m = [poisson(3) for _ in range(10000)]
plot_poisson(m, ax=ax1)
ax2 = plt.subplot(1, 3, 2)
m = [poisson(10) for _ in range(10000)]
plot_poisson(m, ax=ax2)
ax3 = plt.subplot(1, 3, 3)
m = [poisson(100) for _ in range(10000)]
plot_poisson(m, ax=ax3)
# -
# ##### Using numpy
# +
plt.figure(figsize=(15, 4))
ax1 = plt.subplot(1, 3, 1)
m = np.random.poisson(3, size=10000)
plot_poisson(m, ax=ax1)
ax2 = plt.subplot(1, 3, 2)
m = np.random.poisson(10, size=10000)
plot_poisson(m, ax=ax2)
ax3 = plt.subplot(1, 3, 3)
m = np.random.poisson(100, size=10000)
plot_poisson(m, ax=ax3)
# +
from scipy.stats import poisson
rv = poisson(mu=2)
rv.pmf(0)
| distribution/poisson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1-4.3 Intro Python
# ## Conditionals
# - **`if`, `else`, `pass`**
# - Conditionals using Boolean String Methods
# - Comparison operators
# - **String comparisons**
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# - **control code flow with `if`... `else` conditional logic**
# - using Boolean string methods (`.isupper(), .isalpha(), startswith()...`)
# - using comparison (`>, <, >=, <=, ==, !=`)
# - **using Strings in comparisons**
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concept</B></font>
# ## String Comparisons
# - ### Strings can be equal `==` or unequal `!=`
# - ### Strings can be greater than `>` or less than `<`
# - ### alphabetically `"A"` is less than `"B"`
# - ### lower case `"a"` is greater than upper case `"A"`
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# review and run code
"hello" < "Hello"
# review and run code
"Aardvark" > "Zebra"
# review and run code
'student' != 'Student'
# review and run code
print("'student' >= 'Student' is", 'student' >= 'Student')
print("'student' != 'Student' is", 'student' != 'Student')
# review and run code
"Hello " + "World!" == "Hello World!"
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
# ## String Comparisons
msg = "Hello"
# [ ] print the True/False results of testing if msg string equals "Hello" string
print(msg=="Hello")
# +
greeting = "Hello"
# [ ] get input for variable named msg, and ask user to 'Say "Hello"'
msg=input('Say "Hello": ')
# [ ] print the results of testing if msg string equals greeting string
print(msg==greeting)
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concept</B></font>
# ## Conditionals: String comparisons with `if`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/d66365b5-03fa-4d0d-a455-5adba8b8fb1b/Unit1_Section4.3-string-compare-if.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/d66365b5-03fa-4d0d-a455-5adba8b8fb1b/Unit1_Section4.3-string-compare-if.vtt","srclang":"en","kind":"subtitles","label":"english"}])
#
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# +
# [ ] review and run code
msg = "Save the notebook"
if msg.lower() == "save the notebook":
print("message as expected")
else:
print("message not as expected")
# +
# [ ] review and run code
msg = "Save the notebook"
prediction = "save the notebook"
if msg.lower() == prediction.lower():
print("message as expected")
else:
print("message not as expected")
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
# ## Conditionals: comparison operators with if
#
# +
# [ ] get input for a variable, answer, and ask user 'What is 8 + 13? : '
answer=input("What is 8 + 13? :")
# [ ] print messages for correct answer "21" or incorrect answer using if/else
if answer=='21':
print("You got it right!!!")
else:
print("Might want to check that addition")
# note: input returns a "string"
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# ## Program: True False Quiz Function
# Call the tf_quiz function with 2 arguments
# - T/F question string
# - answer key string like "T"
#
# Return a string: "correct" or incorrect"
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/3805cc48-f5c9-4ec8-86ad-1e1db45788e4/Unit1_Section4.3-TF-quiz.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/3805cc48-f5c9-4ec8-86ad-1e1db45788e4/Unit1_Section4.3-TF-quiz.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### Define and use `tf_quiz()` function
# - **`tf_quiz()`** has **2 parameters** which are both string arguments
# - **`question`**: a string containg a T/F question like "Should save your notebook after edit?(T/F): "
# - **`correct_ans`**: a string indicating the *correct answer*, either **"T"** or **"F"**
# - **`tf_quiz()`** returns a string: "correct" or "incorrect"
# - Test tf_quiz(): **create a T/F question** (*or several!*) to **call tf_quiz()**
#
# [ ] Create the program, run tests
def tf_quiz(question,correct_ans):
answer=input(question)
if answer.lower()==correct_ans.lower():
return "correct"
else:
return "incorrect"
quiz_result=tf_quiz("A megabyte is smaller than a gigabyte", "F")
print("Your answer was",quiz_result)
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Absolute Beginner/Module_3_3_Absolute_Beginner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Depression detection
# This notebook shows a prototype on how using a public dataset and a pretrained model it can be possible to detect depression from images of personne.
# +
import warnings
warnings.filterwarnings('ignore')
from mtcnn import MTCNN
import torch
from models import BaselineCNN
from utils import get_emotion_from_image, device
from train import MODEL_WEIGHTS_PATH
# %matplotlib inline
# -
# ### Load the different models
model = BaselineCNN() # init face emotion recongition model
detector = MTCNN() # init face detector model
model.load_state_dict(torch.load(MODEL_WEIGHTS_PATH))
model.eval()
model = model.to(device) # Move the model to GPU is availible
# ## Evaluation on random images from google image
get_emotion_from_image("data/happy.jpg", model, detector)
print("=====================")
get_emotion_from_image("data/angry.jpg", model, detector)
print("=====================")
get_emotion_from_image("data/surprise.jpg", model, detector)
print("=====================")
get_emotion_from_image("data/happy2.jpg", model, detector)
# ## Trying on images of depressed people
# When evaluated on images of depressed people found on google image, we can see that the category are mostly negative.
get_emotion_from_image("data/depressed.png", model, detector)
print("=====================")
get_emotion_from_image("data/depressed2.jpg", model, detector)
print("=====================")
get_emotion_from_image("data/depressed3.jpg", model, detector)
| pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import re
from sklearn.metrics import cohen_kappa_score
import itertools
from typing import Dict, List
import numpy as np
import pandas as pd
from tqdm import tqdm
from enum import Enum
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (15, 10)
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.sans-serif'] = 'cm'
import seaborn as sns
sns.set()
sns.set(font_scale=2) # crazy big
# -
DATA_PATH = os.path.join('..', 'Data', 'Keep')
def read_data():
doc_annotations = {}
text_lengths = {}
authors_set = set()
for author_folder in os.listdir(DATA_PATH):
authors_set.add(author_folder)
full_path = os.path.join(DATA_PATH, author_folder)
ann_files = list(filter(lambda x: x.endswith('.ann'), os.listdir(full_path)))
for filename in ann_files:
doc_name = os.path.splitext(filename)[0]
if doc_name not in doc_annotations.keys():
doc_annotations[doc_name] = {}
if author_folder in doc_annotations[doc_name].keys():
raise Exception(f'Author "{author_folder}" has duplicated annotation for document "{doc_name}"')
with open(os.path.join(full_path, filename), 'r') as file_handler:
doc_annotations[doc_name][author_folder] = file_handler.read()
if doc_name not in text_lengths.keys():
with open(os.path.join(full_path, f'{doc_name}.txt'), 'r') as txt_file_handler:
text_lengths[doc_name] = len(txt_file_handler.read())
return doc_annotations, text_lengths, list(sorted(authors_set))
doc_annotations, text_lengths, authors_set = read_data()
valid_annotations = {
k: v for k, v in doc_annotations.items()
if len(v.keys()) > 1
}
class Annotation:
def __init__(self, key: str, entity: str, start_pos: int, end_pos: int):
self.key = key
self.entity = entity
self.start_pos = start_pos
self.end_pos = end_pos
def __str__(self):
return f'<{self.key}-{self.entity}-[{self.start_pos}:{self.end_pos}]>'
class AnnotationType(Enum):
Unknown = 0
Base = 1
PersonGender = 2
PersonLegalStatus = 3
PersonRole = 4
OrganizationNotary = 5
OrganizationBeneficiary = 5
def get_annotation_type(annotation_str: str) -> AnnotationType:
annotation_parts = annotation_str.split('\t')
if re.search('([T]{1}[0-9]+)', annotation_parts[0]):
return AnnotationType.Base
if not re.search('([A]{1}[0-9]+)', annotation_parts[0]):
# print(annotation_str)
return AnnotationType.Unknown
if annotation_parts[1].startswith('Gender'):
return AnnotationType.PersonGender
if annotation_parts[1].startswith('LegalStatus'):
return AnnotationType.PersonLegalStatus
if annotation_parts[1].startswith('Role'):
return AnnotationType.PersonRole
if annotation_parts[1].startswith('Notary'):
return AnnotationType.OrganizationNotary
if annotation_parts[1].startswith('Beneficiary'):
return AnnotationType.OrganizationBeneficiary
raise Exception(f'Unknown sub-entity annotation: {annotation_str}')
def get_base_entity_positions(annotation_parts: List[str]):
split_annotation = annotation_parts[0].split(' ')
current_annotations_parts = ' '.join(split_annotation[1:]).split(';')
start = int(current_annotations_parts[0].split(' ')[0])
end = int(current_annotations_parts[-1].split(' ')[-1])
return (start, end)
# +
invalid_labels = ['DuplicatePage', 'TranscriptionError_Document']
class DocumentAnnotation:
def __init__(self, doc_key: str, annotations_str: str, text_length: int, labels_to_use: List[str], annotation_type: AnnotationType):
self.doc_key = doc_key
self.text_length = text_length
self.is_valid=True
self.annotations = self._parse_annotations(annotations_str, labels_to_use, annotation_type)
def is_empty(self) -> bool:
return len(self.annotations) == 0
def _parse_annotations(self, annotations_str: str, labels_to_use: List[str], annotation_type: AnnotationType):
result = []
annotations = annotations_str.split('\n')
entity_positions = {}
for annotation in annotations:
annotation_parts = annotation.split('\t')
current_annotation_type = get_annotation_type(annotation)
if current_annotation_type == AnnotationType.Unknown:
continue
entity_id = annotation_parts[0]
if current_annotation_type == AnnotationType.Base:
main_entity_id = annotation_parts[0]
entity_positions[main_entity_id] = get_base_entity_positions(annotation_parts[1:])
if current_annotation_type != annotation_type:
continue
if len(annotation_parts) < 2: print(annotation_parts)
split_annotation = annotation_parts[1].split(' ')
label = split_annotation[0]
if current_annotation_type != AnnotationType.Base:
main_entity_id = split_annotation[1]
if label in invalid_labels:
continue
if labels_to_use is not None and label not in labels_to_use:
continue
result.append(
Annotation(
key=entity_id,
entity=label,
start_pos=entity_positions[main_entity_id][0],
end_pos=entity_positions[main_entity_id][1]))
return result
# -
def parse_annotations(valid_annotations, text_lengths: Dict[str, int], labels_to_use: List[str], annotation_type: AnnotationType):
parsed_annotations = {
doc_key: {
author: DocumentAnnotation(doc_key, annotations, text_lengths[doc_key], labels_to_use, annotation_type)
for author, annotations in annotations_per_author.items()
}
for doc_key, annotations_per_author in valid_annotations.items()
}
# Remove invalid annotations
parsed_annotations = {
doc_key: {
author: doc_annotation
for author, doc_annotation in annotations_per_author.items()
if doc_annotation.is_valid and not doc_annotation.is_empty()
}
for doc_key, annotations_per_author in parsed_annotations.items()
}
# Remove documents where we are left with only one annotation
parsed_annotations = {
doc_key: annotations_per_author
for doc_key, annotations_per_author in parsed_annotations.items()
if len(annotations_per_author.keys()) > 1
}
return parsed_annotations
def annotations_overlap(annotation1: Annotation, annotation2: Annotation, offset_chars: int, match_entity: bool) -> bool:
# if the two annotations do not even overlap with one character, we return false
if (annotation1.start_pos > annotation2.end_pos or
annotation1.end_pos < annotation2.start_pos):
return False
if match_entity and (annotation1.entity != annotation2.entity):
return False
out_of_boundary_chars = abs(annotation1.start_pos - annotation2.start_pos) + abs(annotation1.end_pos - annotation2.end_pos)
result = out_of_boundary_chars <= offset_chars
return result
def print_mapped_annotations(mapped_annotations: dict):
for ann1, ann2 in mapped_annotations.items():
print(f'{ann1.key} <{ann1.start_pos}-{ann1.end_pos}> ---', end='')
if ann2 is None:
print('NONE')
else:
print(f'{ann2.key} <{ann2.start_pos}-{ann2.end_pos}>')
def get_overlapping_annotation(annotation_to_compare: Annotation, annotations: List[Annotation], keys_to_skip: List[str], offset_chars: int, match_entity: bool) -> Annotation:
overlaps = []
for annotation2 in annotations:
if annotation_to_compare.start_pos > annotation2.end_pos:
continue
if annotation2.key in keys_to_skip:
continue
if annotations_overlap(annotation_to_compare, annotation2, offset_chars, match_entity):
overlaps.append(annotation2)
if len(overlaps) == 0:
return None
for overlap in overlaps:
if overlap.entity == annotation_to_compare.entity:
return overlap
return overlaps[0]
def calculate_entity_overlap(doc_annotation1: DocumentAnnotation, doc_annotation2: DocumentAnnotation, offset_chars: int, debug: bool = False):
assert (doc_annotation1.doc_key == doc_annotation2.doc_key)
mapped_annotations = {}
used_counter_annotations = set()
empty_positions = [1 for _ in range(0, doc_annotation1.text_length)]
for annotation in doc_annotation1.annotations:
for i in range(annotation.start_pos, annotation.end_pos):
empty_positions[i] = 0
# Perform iteration using strict overlap matching
for annotation in doc_annotation1.annotations:
overlapping_annotation = get_overlapping_annotation(annotation, doc_annotation2.annotations, used_counter_annotations, offset_chars, match_entity=True)
if overlapping_annotation is not None:
used_counter_annotations.add(overlapping_annotation.key)
mapped_annotations[annotation] = overlapping_annotation
# Perform iteration using loose overlap matching
for annotation in doc_annotation1.annotations:
if annotation in mapped_annotations.keys():
continue
overlapping_annotation = get_overlapping_annotation(annotation, doc_annotation2.annotations, used_counter_annotations, offset_chars, match_entity=False)
if overlapping_annotation is not None:
used_counter_annotations.add(overlapping_annotation.key)
mapped_annotations[annotation] = overlapping_annotation
if debug:
print_mapped_annotations(mapped_annotations)
annotation_maps = [ x.entity for x in mapped_annotations.keys() ]
counter_annotations = [ x.entity if x is not None else 'O' for x in mapped_annotations.values() ]
for annotation2 in doc_annotation2.annotations:
for i in range(annotation2.start_pos, annotation2.end_pos):
empty_positions[i] = 0
if annotation2.key in used_counter_annotations:
continue
annotation_maps.append('O')
counter_annotations.append(annotation2.entity)
free_positions = sum(empty_positions)
for _ in range(free_positions):
annotation_maps.append('O')
counter_annotations.append('O')
if annotation_maps == counter_annotations:
result = 1
else:
result = cohen_kappa_score(annotation_maps, counter_annotations)
return result
def create_comparison_matrix(parsed_annotations, offset_chars: int, authors_set: set, authors:List[str]=None):
comparisons = {
author_1 : {
author_2: []
for author_2 in authors_set
} for author_1 in authors_set
}
for _, annotations in parsed_annotations.items():
for author_1, author_2 in itertools.product(authors_set, authors_set):
if author_1 in annotations.keys() and author_2 in annotations.keys():
kappa_score = calculate_entity_overlap(annotations[author_1], annotations[author_2], offset_chars)
comparisons[author_1][author_2].append(kappa_score)
for author in authors_set:
comparisons[author][author] = [1]
return comparisons
def print_comparison_matrix(comparisons, authors_set: List[str], offset_chars: int, labels_to_use: List[str]):
print('Result')
print(f'- offset characters: {offset_chars}')
print(f'- used labels: {", ".join(labels_to_use)}')
print('----------------------------------------------')
print('\t', end='')
print('\t'.join(authors_set))
for author_1 in authors_set:
print(author_1, end='\t')
for author_2 in authors_set:
print(round(np.mean(comparisons[author_1][author_2]), 2), end='\t')
print()
def plot_comparison_lines(values_per_offset, authors_set):
sns.set_palette(sns.color_palette('husl', 15))
offset_chars = list(values_per_offset.keys())
passed_authors = []
for author1 in authors_set:
for author2 in authors_set:
if author1 == author2: continue
if (author1, author2) in passed_authors or (author2, author1) in passed_authors:
continue
y = [np.mean(x[author1][author2]) for x in values_per_offset.values() if author1 in x.keys() and author2 in x[author1].keys()]
if all(x is None for x in y):
continue
label = f'{author1}, {author2}'
sns.lineplot(x=offset_chars, y=y, label=label)
passed_authors.append((author1, author2))
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.show()
def create_axes():
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
return ax
def plot_confidence_intervals(values_per_offset, authors_set, ax=None, label=None, show: bool = True):
if ax is None:
ax = create_axes()
sns.set_palette('tab10')
offset_chars = list(values_per_offset.keys())
y_values = [[] for _ in offset_chars]
passed_authors = []
for author1 in authors_set:
for author2 in authors_set:
if author1 == author2: continue
if (author1, author2) in passed_authors or (author2, author1) in passed_authors:
continue
pair_label = f'{author1}, {author2}'
y = [[pair_label, offset, np.mean(x[author1][author2])] for offset, x in values_per_offset.items() if author1 in x.keys() and author2 in x[author1].keys() and len(x[author1][author2]) > 0]
if any(x is None for x in y):
continue
y_values.extend(y)
df = pd.DataFrame(y_values,columns=['pairs', 'offsets', 'values'])
sns.lineplot(data=df, x='offsets', y='values', ax=ax, label=label)
if show:
plt.show()
return ax
def calculate_comparisons(valid_annotations, text_lengths: Dict[str, int], authors_set: List[str], offset_chars: List[int], labels_to_use: List[str], annotation_type: AnnotationType = AnnotationType.Base):
parsed_annotations = parse_annotations(valid_annotations, text_lengths, labels_to_use, annotation_type)
values_per_offset = {
offset_char: create_comparison_matrix(parsed_annotations, offset_char, authors_set)
for offset_char in tqdm(offset_chars)
}
return values_per_offset
# +
offset_chars = list(range(0, 51))
labels_to_use=['Person', 'Place', 'Organization']
# -
values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=labels_to_use)
def filter_values(values_per_offset, specific_author: str):
specific_values_per_offset = {
offset: {
author_1: {
author_2: values
for author_2, values in values_per_author.items()
if author_1 == specific_author or author_2 == specific_author
}
for author_1, values_per_author in values_per_authors.items()
}
for offset, values_per_authors in values_per_offset.items()
}
return specific_values_per_offset
plot_comparison_lines(values_per_offset, authors_set)
bert_values_per_offset = filter_values(values_per_offset, specific_author='Bert')
plot_comparison_lines(bert_values_per_offset, authors_set)
plot_confidence_intervals(values_per_offset, authors_set)
plot_confidence_intervals(bert_values_per_offset, authors_set)
gender_values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=None, annotation_type=AnnotationType.PersonGender)
legal_status_values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=None, annotation_type=AnnotationType.PersonLegalStatus)
role_values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=None, annotation_type=AnnotationType.PersonRole)
notary_values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=None, annotation_type=AnnotationType.OrganizationNotary)
beneficiary_values_per_offset = calculate_comparisons(valid_annotations, text_lengths, authors_set, offset_chars=offset_chars, labels_to_use=None, annotation_type=AnnotationType.OrganizationBeneficiary)
# +
ax = create_axes()
ax = plot_confidence_intervals(values_per_offset, authors_set, ax=ax, label='Main', show=False)
ax = plot_confidence_intervals(gender_values_per_offset, authors_set, ax=ax, label='Person - Gender', show=False)
ax = plot_confidence_intervals(legal_status_values_per_offset, authors_set, ax=ax, label='Person - Legal Status', show=False)
ax = plot_confidence_intervals(role_values_per_offset, authors_set, ax=ax, label='Person - Role')
# +
ax = create_axes()
ax = plot_confidence_intervals(values_per_offset, authors_set, ax=ax, label='Main', show=False)
ax = plot_confidence_intervals(gender_values_per_offset, authors_set, ax=ax, label='Person - Gender', show=False)
ax = plot_confidence_intervals(legal_status_values_per_offset, authors_set, ax=ax, label='Person - Legal Status', show=False)
ax = plot_confidence_intervals(role_values_per_offset, authors_set, ax=ax, label='Person - Role', show=False)
ax = plot_confidence_intervals(notary_values_per_offset, authors_set, ax=ax, label='Organization - Notary', show=False)
ax = plot_confidence_intervals(beneficiary_values_per_offset, authors_set, ax=ax, label='Organization - Beneficiary')
| notebooks/.ipynb_checkpoints/evaluation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "4"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. Obtain and review raw data
# <p>One day, my old running friend and I were chatting about our running styles, training habits, and achievements, when I suddenly realized that I could take an in-depth analytical look at my training. I have been using a popular GPS fitness tracker called <a href="https://runkeeper.com/">Runkeeper</a> for years and decided it was time to analyze my running data to see how I was doing.</p>
# <p>Since 2012, I've been using the Runkeeper app, and it's great. One key feature: its excellent data export. Anyone who has a smartphone can download the app and analyze their data like we will in this notebook.</p>
# <p><img src="https://assets.datacamp.com/production/project_727/img/runner_in_blue.jpg" alt="Runner in blue" title="Explore world, explore your data!"></p>
# <p>After logging your run, the first step is to export the data from Runkeeper (which I've done already). Then import the data and start exploring to find potential problems. After that, create data cleaning strategies to fix the issues. Finally, analyze and visualize the clean time-series data.</p>
# <p>I exported seven years worth of my training data, from 2012 through 2018. The data is a CSV file where each row is a single training activity. Let's load and inspect it.</p>
# + dc={"key": "4"} tags=["sample_code"]
# Import pandas
import pandas as pd
# Define file containing dataset
runkeeper_file = 'datasets/cardioActivities.csv'
# Create DataFrame with parse_dates and index_col parameters
df_activities = pd.read_csv(runkeeper_file,parse_dates=True,index_col="Date")
# First look at exported data: select sample of 3 random rows
display(df_activities.sample(3))
# Print DataFrame summary
df_activities.info()
# + dc={"key": "12"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. Data preprocessing
# <p>Lucky for us, the column names Runkeeper provides are informative, and we don't need to rename any columns.</p>
# <p>But, we do notice missing values using the <code>info()</code> method. What are the reasons for these missing values? It depends. Some heart rate information is missing because I didn't always use a cardio sensor. In the case of the <code>Notes</code> column, it is an optional field that I sometimes left blank. Also, I only used the <code>Route Name</code> column once, and never used the <code>Friend's Tagged</code> column.</p>
# <p>We'll fill in missing values in the heart rate column to avoid misleading results later, but right now, our first data preprocessing steps will be to:</p>
# <ul>
# <li>Remove columns not useful for our analysis.</li>
# <li>Replace the "Other" activity type to "Unicycling" because that was always the "Other" activity.</li>
# <li>Count missing values.</li>
# </ul>
# + dc={"key": "12"} tags=["sample_code"]
# Define list of columns to be deleted
cols_to_drop = ['Friend\'s Tagged','Route Name','GPX File','Activity Id','Calories Burned', 'Notes']
# Delete unnecessary columns
df_activities.drop(columns=cols_to_drop,inplace=True)
# Count types of training activities
display(df_activities["Type"].value_counts())
# Rename 'Other' type to 'Unicycling'
df_activities['Type'] = df_activities['Type'].str.replace('Other','Unicycling')
# Count missing values for each column
df_activities.isnull().sum()
# + dc={"key": "19"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Dealing with missing values
# <p>As we can see from the last output, there are 214 missing entries for my average heart rate.</p>
# <p>We can't go back in time to get those data, but we can fill in the missing values with an average value. This process is called <em>mean imputation</em>. When imputing the mean to fill in missing data, we need to consider that the average heart rate varies for different activities (e.g., walking vs. running). We'll filter the DataFrames by activity type (<code>Type</code>) and calculate each activity's mean heart rate, then fill in the missing values with those means.</p>
# + dc={"key": "19"} tags=["sample_code"]
# Calculate sample means for heart rate for each training activity type
avg_hr_run = df_activities[df_activities['Type'] == 'Running']['Average Heart Rate (bpm)'].mean()
avg_hr_cycle = df_activities[df_activities['Type'] == 'Cycling']['Average Heart Rate (bpm)'].mean()
# Split whole DataFrame into several, specific for different activities
df_run = df_activities[df_activities['Type'] == 'Running'].copy()
df_walk = df_activities[df_activities['Type'] == 'Walking'].copy()
df_cycle = df_activities[df_activities['Type'] == 'Cycling'].copy()
# Filling missing values with counted means
df_walk['Average Heart Rate (bpm)'].fillna(110, inplace=True)
df_run['Average Heart Rate (bpm)'].fillna(int(avg_hr_run), inplace=True)
df_cycle['Average Heart Rate (bpm)'].fillna(int(avg_hr_cycle), inplace=True)
# Count missing values for each column in running data
df_run.isnull().sum()
# + dc={"key": "26"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Plot running data
# <p>Now we can create our first plot! As we found earlier, most of the activities in my data were running (459 of them to be exact). There are only 29, 18, and two instances for cycling, walking, and unicycling, respectively. So for now, let's focus on plotting the different running metrics.</p>
# <p>An excellent first visualization is a figure with four subplots, one for each running metric (each numerical column). Each subplot will have a different y-axis, which is explained in each legend. The x-axis, <code>Date</code>, is shared among all subplots.</p>
# + dc={"key": "26"} tags=["sample_code"]
# %matplotlib inline
# Import matplotlib, set style and ignore warning
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
plt.style.use('ggplot')
warnings.filterwarnings(
action='ignore', module='matplotlib.figure', category=UserWarning,
message=('This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.')
)
# Prepare data subsetting period from 2013 till 2018
runs_subset_2013_2018 = df_run[(df_run.index>"2013-01-01") &\
(df_run.index<"2018-12-31")]
# Create, plot and customize in one step
runs_subset_2013_2018.plot(subplots=True,
sharex=False,
figsize=(12,16),
linestyle='none',
marker='o',
markersize=3,
)
# Show plot
plt.show()
# + dc={"key": "33"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. Running statistics
# <p>No doubt, running helps people stay mentally and physically healthy and productive at any age. And it is great fun! When runners talk to each other about their hobby, we not only discuss our results, but we also discuss different training strategies. </p>
# <p>You'll know you're with a group of runners if you commonly hear questions like:</p>
# <ul>
# <li>What is your average distance?</li>
# <li>How fast do you run?</li>
# <li>Do you measure your heart rate?</li>
# <li>How often do you train?</li>
# </ul>
# <p>Let's find the answers to these questions in my data. If you look back at plots in Task 4, you can see the answer to, <em>Do you measure your heart rate?</em> Before 2015: no. To look at the averages, let's only use the data from 2015 through 2018.</p>
# <p>In pandas, the <code>resample()</code> method is similar to the <code>groupby()</code> method - with <code>resample()</code> you group by a specific time span. We'll use <code>resample()</code> to group the time series data by a sampling period and apply several methods to each sampling period. In our case, we'll resample annually and weekly.</p>
# + dc={"key": "33"} tags=["sample_code"]
# Prepare running data for the last 4 years
runs_subset_2015_2018 = df_run[(df_run.index>"2015-01-01") &\
(df_run.index<"2018-12-31")]
# Calculate annual statistics
print('How my average run looks in last 4 years:')
display(runs_subset_2015_2018.resample('A').mean())
# Calculate weekly statistics
print('Weekly averages of last 4 years:')
display(runs_subset_2015_2018.resample('W').mean().mean())
# Mean weekly counts
weekly_counts_average = runs_subset_2015_2018['Distance (km)'].resample('W').count()\
.mean()
print('How many trainings per week I had on average:', weekly_counts_average)
# + dc={"key": "40"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. Visualization with averages
# <p>Let's plot the long term averages of my distance run and my heart rate with their raw data to visually compare the averages to each training session. Again, we'll use the data from 2015 through 2018.</p>
# <p>In this task, we will use <code>matplotlib</code> functionality for plot creation and customization.</p>
# + dc={"key": "40"} tags=["sample_code"]
# Prepare data
runs_subset_2015_2018 = df_run['2018':'2015']
runs_distance = runs_subset_2015_2018['Distance (km)']
runs_hr = runs_subset_2015_2018['Average Heart Rate (bpm)']
# Create plot
fig, (ax1, ax2) = plt.subplots(2,sharex=True,figsize=(12,8))
# Plot and customize first subplot
runs_distance.plot(ax=ax1)
ax1.set(ylabel='Distance (km)', title='Historical data with averages')
ax1.axhline(runs_distance.mean(), color='blue', linewidth=1, linestyle='-.')
# Plot and customize second subplot
runs_hr.plot(ax=ax2, color='gray')
ax2.set(xlabel='Date', ylabel='Average Heart Rate (bpm)')
ax2.axhline(runs_hr.mean(), color='blue', linewidth=1, linestyle='-.')
# Show plot
plt.show()
# + dc={"key": "47"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. Did I reach my goals?
# <p>To motivate myself to run regularly, I set a target goal of running 1000 km per year. Let's visualize my annual running distance (km) from 2013 through 2018 to see if I reached my goal each year. Only stars in the green region indicate success.</p>
# + dc={"key": "47"} tags=["sample_code"]
# Prepare data
df_run_dist_annual = df_run["2018":"2013"]["Distance (km)"].resample("A").sum()
# Create plot
fig = plt.figure(figsize=(8,5))
# Plot and customize
ax = df_run_dist_annual.plot(marker='*', markersize=14, linewidth=0, color='blue')
ax.set(ylim=[0, 1210],
xlim=['2012','2019'],
ylabel='Distance (km)',
xlabel='Years',
title='Annual totals for distance')
ax.axhspan(1000, 1210, color='green', alpha=0.4)
ax.axhspan(800, 1000, color='yellow', alpha=0.3)
ax.axhspan(0, 800, color='red', alpha=0.2)
# Show plot
plt.show()
# + dc={"key": "54"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. Am I progressing?
# <p>Let's dive a little deeper into the data to answer a tricky question: am I progressing in terms of my running skills? </p>
# <p>To answer this question, we'll decompose my weekly distance run and visually compare it to the raw data. A red trend line will represent the weekly distance run.</p>
# <p>We are going to use <code>statsmodels</code> library to decompose the weekly trend.</p>
# + dc={"key": "54"} tags=["sample_code"]
# Import required library
import statsmodels.api as sm
# Prepare data
df_run_dist_wkly = df_run["2018":"2013"]["Distance (km)"].resample("W").bfill()
decomposed = sm.tsa.seasonal_decompose(df_run_dist_wkly, extrapolate_trend=1, freq=52)
# Create plot
fig = plt.figure(figsize=(12,5))
# Plot and customize
ax = decomposed.trend.plot(label='Trend', linewidth=2)
ax = decomposed.observed.plot(label='Observed', linewidth=0.5)
ax.legend()
ax.set_title('Running distance trend')
# Show plot
plt.show()
# + dc={"key": "61"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. Training intensity
# <p>Heart rate is a popular metric used to measure training intensity. Depending on age and fitness level, heart rates are grouped into different zones that people can target depending on training goals. A target heart rate during moderate-intensity activities is about 50-70% of maximum heart rate, while during vigorous physical activity it’s about 70-85% of maximum.</p>
# <p>We'll create a distribution plot of my heart rate data by training intensity. It will be a visual presentation for the number of activities from predefined training zones. </p>
# + dc={"key": "61"} tags=["sample_code"]
# Prepare data
hr_zones = [100, 125, 133, 142, 151, 173]
zone_names = ['Easy', 'Moderate', 'Hard', 'Very hard', 'Maximal']
zone_colors = ['green', 'yellow', 'orange', 'tomato', 'red']
df_run_hr_all = df_run["2018":"2015-03-01"]["Average Heart Rate (bpm)"]
# Create plot
fig, ax = plt.subplots(figsize=(8,5))
# Plot and customize
n, bins, patches = ax.hist(df_run_hr_all, bins=hr_zones, alpha=0.5)
for i in range(0, len(patches)):
patches[i].set_facecolor(zone_colors[i])
ax.set(title='Distribution of HR', ylabel='Number of runs')
ax.xaxis.set(ticks=hr_zones)
ax.set_xticklabels(labels=zone_names,rotation=-30,ha='left')
# Show plot
plt.show()
# + dc={"key": "68"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 10. Detailed summary report
# <p>With all this data cleaning, analysis, and visualization, let's create detailed summary tables of my training. </p>
# <p>To do this, we'll create two tables. The first table will be a summary of the distance (km) and climb (m) variables for each training activity. The second table will list the summary statistics for the average speed (km/hr), climb (m), and distance (km) variables for each training activity.</p>
# + dc={"key": "68"} tags=["sample_code"]
# Concatenating three DataFrames
df_run_walk_cycle = (df_run.append(df_walk).append(df_cycle)).\
sort_index(ascending=False)
dist_climb_cols, speed_col = ['Distance (km)', 'Climb (m)'], ['Average Speed (km/h)']
# Calculating total distance and climb in each type of activities
df_totals = df_run_walk_cycle.groupby('Type')[dist_climb_cols].sum()
print('Totals for different training types:')
display(df_totals)
# Calculating summary statistics for each type of activities
df_summary = df_run_walk_cycle.groupby('Type')[dist_climb_cols + speed_col].describe()
# Combine totals with summary
for i in dist_climb_cols:
df_summary[i, 'total'] = df_totals[i]
print('Summary statistics for different training types:')
df_summary.stack()
# + dc={"key": "75"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 11. Fun facts
# <p>To wrap up, let’s pick some fun facts out of the summary tables and solve the last exercise.</p>
# <p>These data (my running history) represent 6 years, 2 months and 21 days. And I remember how many running shoes I went through–7.</p>
# <pre><code>FUN FACTS
# - Average distance: 11.38 km
# - Longest distance: 38.32 km
# - Highest climb: 982 m
# - Total climb: 57,278 m
# - Total number of km run: 5,224 km
# - Total runs: 459
# - Number of running shoes gone through: 7 pairs
# </code></pre>
# <p>The story of <NAME> is well known–the man, who for no particular reason decided to go for a "little run." His epic run duration was 3 years, 2 months and 14 days (1169 days). In the picture you can see Forrest’s route of 24,700 km. </p>
# <pre><code>FORREST RUN FACTS
# - Average distance: 21.13 km
# - Total number of km run: 24,700 km
# - Total runs: 1169
# - Number of running shoes gone through: ...
# </code></pre>
# <p>Assuming Forest and I go through running shoes at the same rate, figure out how many pairs of shoes Forrest needed for his run.</p>
# <p><img src="https://assets.datacamp.com/production/project_727/img/Forrest_Gump_running_route.png" alt="Forrest's route" title="Little run of Forrest Gump"></p>
# + dc={"key": "75"} tags=["sample_code"]
# Count average shoes per lifetime (as km per pair) using our fun facts
average_shoes_lifetime = 5224/7
# Count number of shoes for Forrest's run distance
shoes_for_forrest_run = 24700/average_shoes_lifetime
print('Forrest Gump would need {} pairs of shoes!'.format(shoes_for_forrest_run))
| Python Projects/Analyze Your Runkeeper Fitness Data/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="dLsS-BYExZ81"
# ### Mounting Colab To Drive
# + id="Qyg7jcQxmn2o" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611691159956, "user_tz": 360, "elapsed": 58101, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="d52e100e-c4ce-4726-bd0d-f45a0131d8b6"
from google.colab import drive
drive.mount('/content/drive/')
# + [markdown] id="sRgrlX2SxQ5q"
# ### Importing the necessary libraries
# + id="OkGT9L_18T3s" executionInfo={"status": "ok", "timestamp": 1611691160314, "user_tz": 360, "elapsed": 58456, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="EJvNATVx7e5R"
# ### Information About The Datasets
# + [markdown] id="vRqYixy57y6i"
#
# + [markdown] id="lNiRML65xnWS"
# # Reading The Datasets With Pandas
# + [markdown] id="lSFgJmPSx0kN"
# ### 1) spg-report2019 dataset
# + id="SHlpCJ_bbJGt" colab={"base_uri": "https://localhost:8080/", "height": 428} executionInfo={"status": "ok", "timestamp": 1611691163209, "user_tz": 360, "elapsed": 61319, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="566eaac2-e34e-4af1-b4ac-86fa683b5b7e"
schools_df = pd.read_csv('/content/drive/Shared drives/Data Science for All - Empowerment/education-data-analysis-project/spg-report2019_final.csv')
schools_df.head()
# + [markdown] id="_LbdCdS_qXcO"
# Printing the basic information about the spg-report2019 dataset
# + colab={"base_uri": "https://localhost:8080/"} id="nJLdXNPh4xLL" executionInfo={"status": "ok", "timestamp": 1611691163209, "user_tz": 360, "elapsed": 61311, "user": {"displayName": "<NAME>ham", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="b878779d-3e03-4cf8-8262-f0446248ff62"
schools_df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="UBqWA6UpXE7t" executionInfo={"status": "ok", "timestamp": 1611691163210, "user_tz": 360, "elapsed": 61306, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="02ebd8a2-3b07-45cb-b53a-2b0926f1d690"
schools_df.info()
# + [markdown] id="5axTbMTKq1nS"
# Checking null values for spg-report2019 dataset
# + id="FDxWlejIXv26" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611691163211, "user_tz": 360, "elapsed": 61300, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="86a9251f-1281-4985-fe8d-ed2189b87888"
schools_df.isna().sum()
# + [markdown] id="F7oGceNGyHDT"
# 2) 2019-2020 SAT Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 979} id="w5IBAnx0J6Ed" executionInfo={"status": "ok", "timestamp": 1611691164114, "user_tz": 360, "elapsed": 62196, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="ecd1700e-ce8c-497a-8568-22ad3f5de263"
sat_df = pd.read_csv('/content/drive/Shared drives/Data Science for All - Empowerment/education-data-analysis-project/satleasch2020_09212020.csv')
sat_df.head(30)
# + [markdown] id="ImE87LzJqqAy"
# Printing the basic information about the SAT dataset
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="iVuNqzn13wsZ" executionInfo={"status": "ok", "timestamp": 1611691164115, "user_tz": 360, "elapsed": 62190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="6b636ccf-f16f-4cbd-e5a8-21acaeb951f2"
sat_df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="VljaTP_e3ssj" executionInfo={"status": "ok", "timestamp": 1611691164116, "user_tz": 360, "elapsed": 62184, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="30c98704-1523-47a0-c22f-2632437ad614"
sat_df.info()
# + [markdown] id="t88IE9rvq-XF"
# Checking null values for SAT dataset
# + colab={"base_uri": "https://localhost:8080/"} id="H3Qb_W4G30VV" executionInfo={"status": "ok", "timestamp": 1611691164117, "user_tz": 360, "elapsed": 62179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="24627ed5-4463-41d8-eca1-691047da2df1"
sat_df.isna().sum()
# + [markdown] id="5OltjEU6J1C0"
# 3) 2019-2020 ACT Dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 752} id="lWki1OrKOzEo" executionInfo={"status": "ok", "timestamp": 1611691168119, "user_tz": 360, "elapsed": 66174, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="8044e4ae-5a6b-40e3-d70c-a678f52bff80"
act_df = pd.read_excel('/content/drive/Shared drives/Data Science for All - Empowerment/education-data-analysis-project/actgrads2019-2020_10122020.xlsx')
act_df.head(15)
# + [markdown] id="vMCkyxPFqxKL"
# Printing the basic information about the ACT dataset
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="fG7C4f474MT8" executionInfo={"status": "ok", "timestamp": 1611691168120, "user_tz": 360, "elapsed": 66167, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="02dbdb32-a4d4-4fd1-8283-b7045a7355c0"
act_df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="OGf_Nk-a4QBU" executionInfo={"status": "ok", "timestamp": 1611691168120, "user_tz": 360, "elapsed": 66160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="9351b457-ee16-4ce8-9fe8-f726c79393d6"
act_df.info()
# + [markdown] id="n8nJHD7trB67"
# Checking null values for ACT dataset
# + colab={"base_uri": "https://localhost:8080/"} id="I9tWku2e4XQw" executionInfo={"status": "ok", "timestamp": 1611691168121, "user_tz": 360, "elapsed": 66154, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="1ef810d8-0a45-4b65-88a4-68fd4437a0de"
act_df.isna().sum()
# + [markdown] id="2Py5j7IUu5Rg"
# # Data Cleaning
# + [markdown] id="L8iMQnQyL1qM"
# ### ACT Dataset Cleaning
# + [markdown] id="fVZZJWG_L_6z"
# Removing top 3 rows and the last row
#
# + id="xwMNY5l5L_ki" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1611691168122, "user_tz": 360, "elapsed": 66148, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="5e5e7154-2e97-4b7d-bf41-06157a677316"
act_df = act_df.iloc[5:765]
act_df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="mkw4h5ipLyCs" executionInfo={"status": "ok", "timestamp": 1611691168123, "user_tz": 360, "elapsed": 66143, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="63b2b667-331b-42e4-c522-dbdc5abb7166"
act_df.head()
# + [markdown] id="OrI90Llc8WWR"
# Preparing Strings To Numeric Values
# + id="H952MB4w-64m" executionInfo={"status": "ok", "timestamp": 1611691168124, "user_tz": 360, "elapsed": 66142, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
act_df = act_df.applymap(lambda x: str(x).replace('>95', '95'))
act_df = act_df.applymap(lambda x: str(x).replace('<5', '5'))
act_df = act_df.applymap(lambda x: str(x).replace('*', '0'))
act_df = act_df.applymap(lambda x: str(x).replace('nan', '999'))
act_df = act_df.applymap(lambda x: str(x).replace(' .', '0'))
# + [markdown] id="E-pvW8coNMyp"
# Changing data types of columns to float
# + id="UvqNA9ixALnX" executionInfo={"status": "ok", "timestamp": 1611691168124, "user_tz": 360, "elapsed": 66140, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
columns_to_float = [' # tested', 'Average Composite Score', 'Average English Score','% Met English Benchmark', 'Average Math Score', '% Met Math Benchmark',
'Average Reading Score', '% Met Reading Benchmark', 'Average Science Score ', '% Met Science Benchmark', '% Met All Four Benchmarks']
for column in columns_to_float:
act_df[column] = pd.to_numeric(act_df[column], errors='coerce')
# + id="zx_8p_7gCYtu" executionInfo={"status": "ok", "timestamp": 1611691168125, "user_tz": 360, "elapsed": 66140, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
# + [markdown] id="HkPIuyWN03e2"
# ### SAT Dataset Cleaning
# + id="0VUzUCo93VDD" executionInfo={"status": "ok", "timestamp": 1611691168126, "user_tz": 360, "elapsed": 66139, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
sat_df = sat_df.iloc[2:]
# + id="ARMrK19e1w9L" executionInfo={"status": "ok", "timestamp": 1611691168126, "user_tz": 360, "elapsed": 66137, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
sat_df['# Tested'] = sat_df['# Tested'].apply(lambda x: str(x).replace('<10', '10'))
sat_df['# Tested'] = sat_df['# Tested'].apply(lambda x: str(x).replace('nan', '0'))
sat_df['# Tested'] = sat_df['# Tested'].apply(lambda x: str(x).replace('*', '0'))
# + colab={"base_uri": "https://localhost:8080/"} id="6R-yyXh9GFzH" executionInfo={"status": "ok", "timestamp": 1611691168127, "user_tz": 360, "elapsed": 66132, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="1744bf28-cc5e-4b3d-b9ca-68e2242013c5"
sat_df['# Tested'].unique()
# + id="BH2--3CJ3pJW" executionInfo={"status": "ok", "timestamp": 1611691168127, "user_tz": 360, "elapsed": 66130, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
# sat_df['# Tested'] = pd.to_numeric(sat_df['# Tested'], errors='coerce')
# + colab={"base_uri": "https://localhost:8080/"} id="NDE_5DuBe2v_" executionInfo={"status": "ok", "timestamp": 1611691168127, "user_tz": 360, "elapsed": 66126, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="de2b1d31-2be7-4de4-b0f9-d0ad7a02d741"
sat_df.columns
# + id="ZA7W_fL_1IXW" executionInfo={"status": "ok", "timestamp": 1611691168128, "user_tz": 360, "elapsed": 66120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
sat_df = sat_df.drop('Unnamed: 3', axis = 1)
sat_df = sat_df.drop('Unnamed: 4', axis = 1)
# + id="j9psc95gelnu" executionInfo={"status": "ok", "timestamp": 1611691168130, "user_tz": 360, "elapsed": 66120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
columns_to_float = ['# Tested', '% Tested', 'Total', 'ERW', 'Math']
for column in columns_to_float:
sat_df[column] = pd.to_numeric(sat_df[column], errors='coerce')
# + [markdown] id="8PrJELsivDJ9"
# ### Schools_df dataset cleaning
# + [markdown] id="o6ZrW-sxTuXg"
# Dropping unnecessary columns
# + id="9HHCNmx-TktS" executionInfo={"status": "ok", "timestamp": 1611691168130, "user_tz": 360, "elapsed": 66118, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
columns_to_drop = ['asm_option', 'k2_feeder', 'use_3yr', 'use_alt_weight', 'scgs_score', 'reporting_year',
'rd_spg_grade', 'rd_spg_score', 'rdgs_ach_score', 'rd_eg_score', 'rd_eg_status', 'rd_eg_index',
'ma_spg_grade', 'ma_spg_score', 'mags_ach_score', 'ma_eg_score', 'ma_eg_status', 'ma_eg_index']
schools_df = schools_df.drop(columns_to_drop, axis = 1)
# + [markdown] id="bzsJqQOdBewF"
# Dropping unnecessary rows from grade_span column
#
# We focus on the high schools so we drop the other schools from grade_span column. After droping elementary and middle schools, we have 6600 rows.
#
#
# + id="lzx6DguGArQr" executionInfo={"status": "ok", "timestamp": 1611691168131, "user_tz": 360, "elapsed": 66117, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
grade_span_to_drop = ['01-05','01-08','02-03','02-04','02-05','03-05','03-08','04-05','04-06','04-08',
'04-09','05-06','05-08','05-09','06-06','06-07','06-08','06-09','06-10','07-08',
'09-09','09-10','09-11','0K-01','0K-02','0K-03','0K-04','0K-05','0K-06','0K-07',
'0K-08','0K-09','0K-11','PK-01','PK-02','PK-03','PK-04','PK-05','PK-06','PK-07',
'PK-08','PK-09','PK-0K','UN-GR']
schools_df = schools_df[~schools_df['grade_span'].isin(grade_span_to_drop)]
# + [markdown] id="ABoLpxVpsMuk"
# After dropping we have 6600 rows.
# + colab={"base_uri": "https://localhost:8080/"} id="xt81Ig0kKlvb" executionInfo={"status": "ok", "timestamp": 1611691168132, "user_tz": 360, "elapsed": 66112, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="53ce06e7-4a3f-48fc-d774-a68415a13bf0"
schools_df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="esQ9zolSLJPk" executionInfo={"status": "ok", "timestamp": 1611691168132, "user_tz": 360, "elapsed": 66106, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="655ead3f-cbff-48a5-8b04-2c1a6a44879b"
schools_df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="bErp3XRRLeoo" executionInfo={"status": "ok", "timestamp": 1611691168133, "user_tz": 360, "elapsed": 66101, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="bcb35ff2-5946-4c7e-c010-dd2b10cd5ce6"
schools_df.isna().sum()
# + id="lszoQlnRVXpO" executionInfo={"status": "ok", "timestamp": 1611691168133, "user_tz": 360, "elapsed": 66100, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
columns_to_numeric = ['elp_score', 'cgrs_score', 'bi_score', 'awa_score', 'mcr_score']
for column in columns_to_numeric:
schools_df[column] = schools_df[column].apply(lambda x: str(x).replace('<5', '5'))
schools_df[column] = schools_df[column].apply(lambda x: str(x).replace('nan', '999'))
# + id="sLOThklWVFC-" executionInfo={"status": "ok", "timestamp": 1611691168134, "user_tz": 360, "elapsed": 66099, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
for column in columns_to_numeric:
schools_df[column] = pd.to_numeric(schools_df[column], errors='coerce')
# + id="vm2DHTmPxIw2" executionInfo={"status": "ok", "timestamp": 1611691168135, "user_tz": 360, "elapsed": 66098, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
schools_df['title_1'] = schools_df['title_1'].apply(lambda x: str(x).replace('nan', 'N'))
schools_df['spg_grade'] = schools_df['spg_grade'].apply(lambda x: str(x).replace('nan', 'I'))
schools_df['spg_score'] = schools_df['spg_score'].apply(lambda x: str(x).replace('nan', '999'))
schools_df['ach_score'] = schools_df['ach_score'].apply(lambda x: str(x).replace('nan', '999'))
schools_df['eg_score'] = schools_df['eg_score'].apply(lambda x: str(x).replace('nan', '999'))
schools_df['eg_status'] = schools_df['eg_status'].apply(lambda x: str(x).replace('nan', 'insuff_data'))
schools_df['eg_index'] = schools_df['eg_index'].apply(lambda x: str(x).replace('nan', '50.'))
# + [markdown] id="_lHcou5191wo"
# Making A Dataframe Without Demografic Subgroups
# + [markdown] id="b7fZyWuDsdLu"
# When we exclude the subgroups we have only the unique schools. This school_df_all can be joined with zip code information to discover the unique properties of the schools. We have 660 schools after eliminating the subgroups.
# + id="QScS_u33OP7h" executionInfo={"status": "ok", "timestamp": 1611691168135, "user_tz": 360, "elapsed": 66097, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
schools_df_all = schools_df[schools_df['subgroup'].isin(['ALL'])]
# + colab={"base_uri": "https://localhost:8080/"} id="qLjAMP_SPERe" executionInfo={"status": "ok", "timestamp": 1611691168136, "user_tz": 360, "elapsed": 66092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="dba54548-7667-41c3-be76-97ea53b3cb63"
schools_df_all.shape
# + colab={"base_uri": "https://localhost:8080/"} id="Q03fSsqzPLUB" executionInfo={"status": "ok", "timestamp": 1611691168137, "user_tz": 360, "elapsed": 66087, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="ae45991b-f57b-4ad9-eae7-3818f8f3cd6a"
schools_df_all.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="JToxweSTPrGN" executionInfo={"status": "ok", "timestamp": 1611691168137, "user_tz": 360, "elapsed": 66081, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="04b7a22a-fbbb-4e5b-8487-0c94962267d9"
schools_df_all.info()
# + [markdown] id="NjCm-z7jppoh"
# **bold text**## Basic EDA
# + [markdown] id="XSxfhWjXsscP"
# ### Schools_df EDA
# + [markdown] id="-0WuzBKwlehv"
# ### Univariate EDA
# + [markdown] id="gx6VG_SDq3ic"
# Schools_df
# + id="TowAbQWdi_Vz" executionInfo={"status": "ok", "timestamp": 1611691168140, "user_tz": 360, "elapsed": 66082, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
# for column in list(schools_df_all.columns):
# schools_df_all[column].hist()
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="9-JBQx-GjwZG" executionInfo={"status": "ok", "timestamp": 1611691168141, "user_tz": 360, "elapsed": 66077, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="778805f9-ea79-471c-cfdf-b86443660959"
plt.figure(figsize=(22,8))
schools_df_all.sbe_region.hist(bins=50, xrot=70);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="qJ-fPZRVlq09" executionInfo={"status": "ok", "timestamp": 1611691168141, "user_tz": 360, "elapsed": 66070, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="02ac1553-ba2a-4be1-d3a8-00e12e79166b"
plt.figure(figsize=(22,8))
schools_df_all.grade_span.hist(bins=50);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="HrzZ6m7tmEHJ" executionInfo={"status": "ok", "timestamp": 1611691168142, "user_tz": 360, "elapsed": 66064, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="c6034645-b984-4d0d-a0d3-7594c77672a5"
plt.figure(figsize=(22,8))
schools_df_all.title_1.hist(bins=50);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="kDXXMMKBmUVo" executionInfo={"status": "ok", "timestamp": 1611691169073, "user_tz": 360, "elapsed": 66989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="694b0bbe-d4d8-47ed-96b9-e141e9720217"
plt.figure(figsize=(22,8))
schools_df_all.missed_days.hist(bins=100);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="EZnoZi1pm0I-" executionInfo={"status": "ok", "timestamp": 1611691169074, "user_tz": 360, "elapsed": 66983, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="d84c95e6-5691-4810-bf9c-74d35578ce61"
plt.figure(figsize=(22,8))
schools_df.subgroup.hist(bins=100);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="3XRaAqTXnK79" executionInfo={"status": "ok", "timestamp": 1611691169075, "user_tz": 360, "elapsed": 66978, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="4ca47f1b-9a21-4bd7-96b9-ada5305ce63c"
plt.figure(figsize=(22,8))
schools_df_all.spg_grade.hist(bins=100);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="yldpNw8anXgw" executionInfo={"status": "ok", "timestamp": 1611691171023, "user_tz": 360, "elapsed": 68919, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="3eba8112-7326-467f-f500-2708d3e63397"
plt.figure(figsize=(22, 10))
schools_df_all.spg_score.hist(bins=60,xrot=80,xlabelsize=14, ylabelsize=20);
# + id="SIHajuK45JcS" executionInfo={"status": "ok", "timestamp": 1611691171024, "user_tz": 360, "elapsed": 68919, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="AFhM_vv4nz9n" executionInfo={"status": "ok", "timestamp": 1611691175414, "user_tz": 360, "elapsed": 73303, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="91489a77-f2f3-4cc0-e93f-6dfc0ac8218a"
plt.figure(figsize=(50,20))
schools_df_all.ach_score.hist(bins=100, xrot=90, xlabelsize=11, ylabelsize=30);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="SDC4KtN7oM3E" executionInfo={"status": "ok", "timestamp": 1611691183173, "user_tz": 360, "elapsed": 81055, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="df099ba9-29b3-443d-8990-18dfa1d23233"
plt.figure(figsize=(50, 20))
schools_df_all.eg_score.hist(bins=100, xrot=90);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="Efcww8bDoYgm" executionInfo={"status": "ok", "timestamp": 1611691183865, "user_tz": 360, "elapsed": 81741, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="8676dd09-73b6-4943-b6fb-78a099e310bb"
plt.figure(figsize=(22,8))
schools_df_all.eg_status.hist(bins=100, xrot=70, xlabelsize=14, ylabelsize=14);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="wrvylsXKqcj4" executionInfo={"status": "ok", "timestamp": 1611691183866, "user_tz": 360, "elapsed": 81736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="27a99b67-3fab-4b2d-dca1-0642d0b5540e"
plt.figure(figsize=(22,8))
schools_df_all.elp_score.hist(bins=50, xrot=90, xlabelsize=14, ylabelsize=14);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="74I88dm5osAz" executionInfo={"status": "ok", "timestamp": 1611691183866, "user_tz": 360, "elapsed": 81730, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="22feb6e3-48b8-4b95-ba0f-fa8a52336854"
plt.figure(figsize=(22,8))
schools_df_all.cgrs_score.hist(bins=50, xrot=90, xlabelsize=14, ylabelsize=14);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="UmMkAWZcqfC-" executionInfo={"status": "ok", "timestamp": 1611691183867, "user_tz": 360, "elapsed": 81725, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="05e8ae49-7b84-47ff-bb04-e44fd2c03917"
plt.figure(figsize=(22,8))
schools_df_all.bi_score.hist(bins=50, xrot=90, xlabelsize=14, ylabelsize=14);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="H8HOhD5GqmOK" executionInfo={"status": "ok", "timestamp": 1611691183867, "user_tz": 360, "elapsed": 81718, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="12da3bf9-35dc-46cc-ee9f-0f38197e8d74"
plt.figure(figsize=(22,8))
schools_df_all.awa_score.hist(bins=50, xrot=90, xlabelsize=14, ylabelsize=14);
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="tUinbI9uquvR" executionInfo={"status": "ok", "timestamp": 1611691183868, "user_tz": 360, "elapsed": 81713, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="aafa98a3-b6f1-4830-e0ad-fc532fb303a3"
plt.figure(figsize=(22,8))
schools_df_all.mcr_score.hist(bins=50, xrot=90, xlabelsize=14, ylabelsize=14);
# + [markdown] id="inRUI7vWs9GX"
# ### Act_df EDA
# + id="42CB2d12tLMf" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691183868, "user_tz": 360, "elapsed": 81706, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="16c89c46-cbc0-4815-f315-2d63a1a0c184"
act_df
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="NJkShNkfBvdx" executionInfo={"status": "ok", "timestamp": 1611691183869, "user_tz": 360, "elapsed": 81701, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="aabf6734-57a3-4c02-e8fd-e6132dcef05f"
plt.figure(figsize=(30,12))
act_df['Unnamed: 0'].hist(bins=50, xrot=90, xlabelsize=12, ylabelsize=30);
# + [markdown] id="tf_IC2g2tCjf"
#
#
#
#
#
#
# ss
#
#
#
#
# ### Sat_df EDA
# + id="8Ec22-abtWOt" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691183870, "user_tz": 360, "elapsed": 81695, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="7b0f276d-c06b-4985-cd25-710c59d95928"
sat_df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="32WAO0kXxz-R" executionInfo={"status": "ok", "timestamp": 1611691183870, "user_tz": 360, "elapsed": 81689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="133eab3d-b225-481a-b718-bda206e36022"
sat_df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="0P0oHHoLyugB" executionInfo={"status": "ok", "timestamp": 1611691184646, "user_tz": 360, "elapsed": 82459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="cd855994-e55f-4aa2-ab6e-b9478bde3e56"
plt.figure(figsize=(50, 20))
sat_df['Unnamed: 0'].hist(xrot=80);
# + id="vRuk9DF5Ei5G" executionInfo={"status": "ok", "timestamp": 1611691184647, "user_tz": 360, "elapsed": 82458, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
sat_df['% Tested'] = sat_df['% Tested'].dropna();
# + id="EU6GaR_QzWA0" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691185114, "user_tz": 360, "elapsed": 82919, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="84479ae6-3cec-420c-8610-a88f760c7650"
plt.figure(figsize=(50,20))
sat_df['Total'].hist(xrot=80, bins=20);
# + id="05ouSZ8u2RB1" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691185563, "user_tz": 360, "elapsed": 83362, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="828022c5-f23d-4010-b02c-8c12fcf353a5"
plt.figure(figsize=(50,20))
sat_df['ERW'].hist(xrot=80, bins=20);
# + id="iKL5Gb942Wtj" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691187940, "user_tz": 360, "elapsed": 85732, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="ea302985-2f96-4444-ab96-03ac18fb5671"
plt.figure(figsize=(100,40))
sat_df['Math'].hist(xrot=80, bins=20);
# + [markdown] id="tJWU0oDERUhK"
# ### Multivariate analysis
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="uosSL3X6S5g9" executionInfo={"status": "ok", "timestamp": 1611691187941, "user_tz": 360, "elapsed": 85727, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="b66e0ecf-dc57-44da-c090-ce4ea26c539b"
act_df.corr()
# + id="Wp62cGwnmGaE" colab={"base_uri": "https://localhost:8080/", "height": 0} executionInfo={"status": "ok", "timestamp": 1611691189796, "user_tz": 360, "elapsed": 87575, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="98352152-c70e-4398-e4bb-1fc28627d365"
sns.set(font_scale=1.4)
plt.figure(figsize=(20, 20))
ax = sns.heatmap(act_df.corr(), center=0, linewidths=.5, annot=True)
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="Ymv_FqV_gQkp" executionInfo={"status": "ok", "timestamp": 1611691190322, "user_tz": 360, "elapsed": 88095, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="e7e623f6-456b-416c-9aae-7d59c227e972"
sns.set(font_scale=1.4)
plt.figure(figsize=(20, 20))
ax = sns.heatmap(sat_df.corr(), center=0, linewidths=.5, annot=True,)
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="8FwCB8GLTGzc" executionInfo={"status": "ok", "timestamp": 1611691190830, "user_tz": 360, "elapsed": 88596, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}} outputId="4e857039-3143-42ca-b7de-a431bfa6bbfa"
sns.set(font_scale=1.4)
g= sns.jointplot(x='% Met All Four Benchmarks', y='% Met English Benchmark', data=act_df, height=9)
g.fig.subplots_adjust(top=0.9)
g.fig.suptitle('English vs All Scatterplot', fontsize=16)
plt.show()
# + id="PQsGwLRNu96P" executionInfo={"status": "ok", "timestamp": 1611691190831, "user_tz": 360, "elapsed": 88595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiaHIjgVpmiQbnL8UAZdVqhH0xKplkClF46OUuafg=s64", "userId": "16259157206279310513"}}
| Team69-Education-data-analysis-project_Jan_26_IndepthEDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data structures
#
# For a refresher on object-oriented programming, see [Object-oriented programming](https://github.com/parrt/msds501/blob/master/notes/OO.ipynb).
# ## A simple set implementation
#
# Sets in Python can be specified with set notation:
s = {1,3,2,9}
# Or with by creating a `set` object and assigning it to a variable then manually adding elements:
s = set()
s.add(1)
s.add(3)
# We can build our own set object implementation by creating a class definition:
class MySet:
def __init__(self):
self.elements = []
def add(self, x):
if x not in self.elements:
self.elements.append(x)
s = MySet()
s.add(3) # same as MySet.add(a,3)
s.add(3)
s.add(2)
s.add('cat')
s.elements
from lolviz import *
objviz(s)
# **Question**: How expensive is it to add an element to a set with this implementation?
# ### Exercise
#
# Add a method called `hasmember()` that returns true or false according to whether parameter `x` is a member of the set.
class MySet:
def __init__(self):
self.elements = []
def add(self, x):
if x not in self.elements:
self.elements.append(x)
def hasmember(self, x):
return x in self.elements
s = MySet()
s.add(3) # same as MySet.add(a,3)
s.add(3)
s.add(2)
s.add('cat')
s.hasmember(3), s.hasmember(99)
# ## Linked lists -- the gateway drug
#
# We've studied arrays/lists that are built into Python but they are not always the best kind of list to use. Sometimes, we are inserting and deleting things from the head or middle of the list. If we do this in lists made up of contiguous cells in memory, we have to move a lot of cells around to make room for a new element or to close a hole made by a deletion. Most importantly, linked lists are the degenerate form of a general object graph. So, it makes sense to start with the simple versions and move up to general graphs.
#
# Linked lists allow us to efficiently insert and remove things anywhere we want, at the cost of more memory.
#
# A linked list associates a `next` pointer with each `value`. We call these things *nodes* and here's a simple implementation for node objects:
class LLNode:
def __init__(self, value, next=None):
self.value = value
self.next = next
head = LLNode('tombu')
callsviz(varnames='head')
head = LLNode('parrt', head)
callsviz(varnames='head')
head = LLNode("xue", head)
callsviz(varnames='head')
# ## Walk list
#
# To walk a list, we use the notion of a *cursor*, which we can think of as a finger that moves along a data structure from node to node. We initialize the cursor to point to the first node of the list, the head, and then walk the cursor through the list via the `next` fields:
p = head
while p is not None:
print(p.value)
p = p.next
# **Question**: How fast can we walk the linked list?
# ### Exercise
#
# Modify the walking code so that it lives in a method of `LLNode` called `exists(self, x)` that looks for a node with value `x` starting at `self`. If we test with `head.exists('parrt')` then `self` would be our global `head` variable. Have the function return true if `x` exists in the list, else return false. You can test it with:
#
# ```python
# head = LLNode('tombu')
# head = LLNode('parrt', head)
# head = LLNode("xue", head)
# head.exists('parrt'), head.exists('part')
# ```
# +
class LLNode:
def __init__(self, value, next=None):
self.value = value
self.next = next
def exists(self, x):
p = self # start looking at this node
while p is not None:
if x==p.value:
return True
p = p.next
return False
head = LLNode('tombu')
head = LLNode('parrt', head)
head = LLNode("xue", head)
head.exists('parrt'), head.exists('part')
# -
# ## Insertion at head
#
# If we want to insert an element at the front of a linked list, we create a node to hold the value and set its `next` pointer to point to the old `head`. Then we have the `head` variable point at the new node. Here is the sequence.
# **Create new node**
x = LLNode('mary')
callviz(varnames=['head','x'])
# **Make next field of new node point to head**
x.next = head
callviz(varnames=['head','x'])
# **Make head point at new node**
head = x
callviz(varnames=['head','x'])
# ## Deletion of node
# to delete xue, make previous node skip over xue
xue = head.next
callviz(varnames=['head','x','xue'])
head.next = xue.next
callviz(varnames=['head','x'])
# Notice that `xue` still points at the node but we are going to ignore that variable from now on. Moving from the head of the list, we still cannot see the node with `'xue'` in it.
head.next = xue.next
callviz(varnames=['head','x','xue'])
# ### Exercise
#
# Get a pointer to the node with value `tombu` and then delete it from the list using the same technique we just saw.
before_tombu = head.next
callviz(varnames=['head','x','before_tombu'])
before_tombu.next = None
callviz(varnames=['head','x','before_tombu'])
# ## Binary trees
#
# The tree data structure is one of the most important in computer science and is extremely common in data science as well. Decision trees, which form the core of gradient boosting machines and random forests (machine learning algorithms), are naturally represented as trees in memory. When we process HTML and XML files, those are generally represented by trees. For example:
#
# <img align="right" src="figures/xml-tree.png" width="200"></td>
# ```xml
# <bookstore>
# <book category="cooking">
# <title lang="en">Everyday Italian</title>
# <author><NAME></author>
# <year>2005</year>
# <price>30.00</price>
# </book>
# <book category="web">
# <title lang="en">Learning XML</title>
# <author><NAME></author>
# <year>2003</year>
# <price>39.95</price>
# </book>
# </bookstore>
# ```
#
# We're going to look at a simple kind of tree that has at most two children: a *binary tree*. A node that has no children is called a *leaf* and non-leaves are called *internal nodes*.
#
# In general, trees with $n$ nodes have $n-1$ edges. Each node has a single incoming edge and the root has none.
#
# Nodes have *parents* and *children* and *siblings* (at the same level).
#
# Sometimes nodes have links back to their parents for programming convenience reasons. That would make it a graph not a tree but we still consider it a tree.
class Tree:
def __init__(self, value, left=None, right=None):
self.value = value
self.left = left
self.right = right
root = Tree('parrt')
root.left = Tree('mary')
root.right = Tree('april')
treeviz(root)
root = Tree('parrt', Tree('mary'), Tree('april'))
treeviz(root)
root = Tree('parrt', None, Tree('mary'))
treeviz(root)
# +
root = Tree('parrt')
mary = Tree('mary')
april = Tree('april')
jim = Tree('jim')
sri = Tree('sri')
mike = Tree('mike')
root.left = mary
root.right = april
mary.left = jim
mary.right = mike
april.right = sri
treeviz(root)
# -
# ### Exercise
# Create a class definition for `NTree` that allows arbitrary numbers of children. (Use a list for field `children` rather than `left` and `right`.) The constructor should init an empty children list. Test your code using:
#
# ```python
# from lolviz import objviz
# root2 = NTree('parrt')
# mary = NTree('mary')
# april = NTree('april')
# jim = NTree('jim')
# sri = NTree('sri')
# mike = NTree('mike')
#
# root2.addchild(mary)
# root2.addchild(jim)
# root2.addchild(sri)
# sri.addchild(mike)
# sri.addchild(april)
#
# objviz(root2)
# ```
# #### Solution
class NTree:
def __init__(self, value):
self.value = value
self.children = []
def addchild(self, child):
if isinstance(child, NTree):
self.children.append(child)
# +
root2 = NTree('parrt')
mary = NTree('mary')
april = NTree('april')
jim = NTree('jim')
sri = NTree('sri')
mike = NTree('mike')
root2.addchild(mary)
root2.addchild(jim)
root2.addchild(sri)
sri.addchild(mike)
sri.addchild(april)
objviz(root2)
# -
# ### Walking trees
#
# Walking a tree is a matter of moving a cursor like we did with the linked lists above. The goal is to visit each node in the tree. We start out by having the cursor point at the root of the tree and then walk downwards until we hit leaves, and then we come back up and try other alternatives.
#
# A good physical analogy: imagine a person (cursor) from HR needing to speak (visit) each person in a company starting with the president/CEO. Here's a sample org chart:
#
# <img src="figures/orgchart.png" width="200">
# The general visit algorithm starting at node `p` is meet with `p` then visit each direct report. Then visit all of their direct reports, one level of the tree at a time. The node visitation sequence would be A,B,C,F,H,J,... This is a *breadth-first search* of the tree and easy to describe but a bit more work to implement that a *depth-first search*. Depth first means visiting a person then visit their first direct report and that person's direct report etc... until you reach a leaf node. Then back up a level and move to next direct report. That visitation sequence is A,B,C,D,E,F,G,H,I,J,K,L.
#
# If you'd like to start at node B, not A, what is the procedure? The same, of course. So visiting A means, say, printing `A` then visiting B. Visiting B means visiting C, and when that completes, visiting F, etc... The key is that the procedure for visiting a node is exactly the same regardless of which node you start with. This is generally true for any self-similar data structure like a tree.
# Another easy way to think about binary tree visitation in particular is positioning yourself in a room with a bunch of doors as choices. Each door leads to other rooms, which might also have doors leading to other rooms. We can think of a room as a node and doors as pointers to other nodes. Each room is identical and has 0, 1, or 2 doors (for a binary tree). At the root node we might see two choices and, to explore all nodes, we can visit each door in turn. Let's go left:
#
# <img src="figures/left-door.png" width="100">
#
# After exploring all possible rooms by taking the left door, we come all the way back out to the root room and try the next alternative on the right:
#
# <img src="figures/right-door.png" width="100">
# Algorithmically what were doing in each room is
#
# ```
# procedure visit room:
# if left door exists, visit rooms accessible through left door
# if right door exists, visit rooms accessible through right door
# ```
#
# Or in code notation:
#
# ```python
# def visit(room):
# if room.left: visit(room.left)
# if room.right: visit(room.right)
# ```
#
# This mechanism works from any room. Imagine waking up and finding yourself in a room with two doors. You have no idea whether you are at the root or somewhere in the middle of a labyrinth (maze) of rooms.
#
# This approach is called *backtracking*.
#
# Let's code this up but make a regular function not a method of the tree class to keep things simple. Let's look at that tree again:
treeviz(root)
# +
def walk(t):
"Depth-first walk of binary tree"
if t is None: return
# if t.left is None: callsviz(varnames=['t']).view()
print(t.value) # "visit" or process this node
walk(t.left) # walk into the left door
walk(t.right) # after visiting all those, enter right door
walk(root)
# -
# That is a *recursive* function, meaning that `walk` calls itself. It's really no different than the recurrence relations we use in mathematics, such as the gradient descent recurrence:
#
# $x_{t+1} = x_t - \eta f'(x_t)$
#
# Variable $x$ is a function of previous incarnations of itself.
# +
def fact(n):
print(f"fact({n})")
if n==0: return 1
return n * fact(n-1)
fact(10)
# -
# Don't let the recursion scare you, just pretend that you are calling a different function or that you are calling the same function except that it is known to be correct. We call that the "recursive leap of faith." (See [Fundamentals of Recursion](https://www2.cs.duke.edu/courses/cps006/spring03/forbes/inclass/recursion.pdf),Although that one is using C++ not Python.)
#
# As the old joke goes: "*To truly understand recursion, you must first understand recursion.*"
#
# The order in which we reach (enter/exit) each node during the search is always the same for a given search strategy, such as depth first search. Here is a visualization from Wikipedia:
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d4/Sorted_binary_tree_preorder.svg/440px-Sorted_binary_tree_preorder.svg.png" width="250">
#
# We always try to go as deep as possible before exploring siblings.
#
# Now, notice the black dots on the traversal. That signifies processing or "visiting" a node and in this case is done before visiting the children. When we process a node and then it's children, we call that a *preorder traversal*. If we process a node after walking the children, we call it a *post-order traversal*:
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/9/9d/Sorted_binary_tree_postorder.svg/440px-Sorted_binary_tree_postorder.svg.png" width="250">
#
# In code, that just means switching the processing step two after the walk of the children:
# +
def walk(t):
if t is None: return
walk(t.left)
walk(t.right)
print(t.value) # process after visiting children
walk(root)
# -
# In both cases we are performing a *depth-first walk* of the tree, which means that we are immediately seeking the leaves rather than siblings. A depth first walk scans down all of the left child fields of the nodes until it hits a leaf and then goes back up a level, looking for children at that level.
#
# In contrast, a *breadth-first walk* processes all children before looking at grandchildren. This is a less common walk but, for our tree, would be the sequence parrt, mary, april, jim, mike, sri. In a sense, breadth first processes one level of the tree at a time:
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d1/Sorted_binary_tree_breadth-first_traversal.svg/440px-Sorted_binary_tree_breadth-first_traversal.svg.png" width="250">
# ### Exercise
#
# Alter the depth-first recursive tree walk above to sum the values in a binary tree. Have `walk()` return the sum of a node's value and all it childrens' values. Test with:
#
# ```python
# a = Tree(3)
# b = Tree(5)
# c = Tree(10)
# d = Tree(9)
# e = Tree(4)
# f = Tree(1)
#
# a.left = b
# a.right = c
# b.left = d
# b.right = e
# e.right = f
# treeviz(a)
#
# print(walk(a), walk(b), walk(c))
# ```
# +
class Tree:
def __init__(self, value, left=None, right=None):
self.value = value
self.left = left
self.right = right
def walk(t:Tree) -> int:
if t is None: return 0
return t.value + walk(t.left) + walk(t.right)
a = Tree(3)
b = Tree(5)
c = Tree(10)
d = Tree(9)
e = Tree(4)
f = Tree(1)
a.left = b
a.right = c
b.left = d
b.right = e
e.right = f
treeviz(a)
print(walk(a), walk(b), walk(c))
# -
# ## Graphs
#
# Trees are actually a subset of the class of directed, acyclic graphs. If we remove the acyclic restriction and the restriction that nodes have a single incoming edge, we get a general, directed graph. These are also extremely common in computer science and are used to represent graphs of users in a social network, locations on a map, or a graph of webpages, which is how Google does page ranking.
#
# ### graphviz
#
# You might find it useful to display graphs visually and [graphviz](https://www.graphviz.org/) is an excellent way to do that. Here's an example
# +
import graphviz as gv
gv.Source("""
digraph G {
node [shape=box penwidth="0.6" margin="0.0" fontname="Helvetica" fontsize=10]
edge [arrowsize=.4 penwidth="0.6"]
rankdir=LR;
ranksep=.25;
cat->dog
dog->cat
dog->horse
dog->zebra
horse->zebra
zebra->llama
}
""")
# -
# Once again, it's very convenient to represent a node in this graph as an object, which means we need a class definition:
class GNode:
def __init__(self, value):
self.value = value
self.edges = [] # outgoing edges
def connect(self, other):
self.edges.append(other)
# +
cat = GNode('cat')
dog = GNode('dog')
horse = GNode('horse')
zebra = GNode('zebra')
llama = GNode('llama')
cat.connect(dog)
dog.connect(cat)
dog.connect(horse)
dog.connect(zebra)
horse.connect(zebra)
zebra.connect(llama)
# -
objviz(cat)
# ### Walking graphs
#
# Walking a graph (depth-first) is just like walking a tree in that we use backtracking to try all possible branches out of every node until we have reached all reachable nodes. When we run into a dead end, we back up to the most recently available on visited path and try that. That's how you get from the entrance to the exit of a maze.
#
# <img src="figures/maze.jpg" width="300">
#
# The only difference between walking a tree and walking a graph is that we have to watch out for cycles when walking a graph, so that we don't get stuck in an infinite loop. We leave a trail of breadcrumbs or candies or string to help us keep track of where we have visited and where we have not. If we run into our trail, we have hit a *cycle* and must also backtrack to avoid an infinite loop. This is a [depth first search](https://en.wikipedia.org/wiki/Tree_traversal#Depth-first_search).
#
# Here's a nice [visualization website for graph walking](http://algoanim.ide.sk/index.php?page=showanim&id=47).
#
# <a href="http://algoanim.ide.sk/index.php?page=showanim&id=47)"><img src="figures/graph-dfs-icon.png" width="300"></a>
#
# In code, here is how we perform a depth-frist search on a graph:
# +
def walk(g, visited):
"Depth-first walk of a graph"
if g is None or g in visited: return
visited.add(g) # mark as visited
print(g.value) # process before visiting outgoing edges
for node in g.edges:
walk(node, visited) # walk all outgoing edge targets
walk(cat, set())
# -
# Where we start the walk of the graph matters:
walk(llama, set())
walk(horse, set())
# # Operator overloading
# (Note: We *overload* operators but *override* methods in a subclass definition)
#
# Python allows class definitions to implement functions that are called when standard operator symbols such as `+` and `/` are applied to objects of that type. This is extremely useful for mathematical libraries such as numpy, but is often abused. Note that you could redefine subtraction to be multiplication when someone used the `-` sign. (Yikes!)
#
# Here's an extension to `Point` that supports `+` for `Point` addition:
# +
import numpy as np
class Point:
def __init__(self, x, y):
self.x = x
self.y = y
def distance(self, other):
return np.sqrt( (self.x - other.x)**2 + (self.y - other.y)**2 )
def __add__(self,other):
x = self.x + other.x
y = self.y + other.y
return Point(x,y)
def __str__(self):
return f"({self.x},{self.y})"
# -
p = Point(3,4)
q = Point(5,6)
print(p, q)
print(p + q) # calls p.__add__(q) or Point.__add__(p,q)
print(Point.__add__(p,q))
# ## Exercise
#
# Add a method to implement the `-` subtraction operator for `Point` so that the following code works:
#
# ```python
# p = Point(5,4)
# q = Point(1,5)
# print(p, q)
# print(p - q)
# ```
# +
import numpy as np
class Point:
def __init__(self, x, y):
self.x = x
self.y = y
def distance(self, other):
return np.sqrt( (self.x - other.x)**2 + (self.y - other.y)**2 )
def __add__(self,other):
x = self.x + other.x
y = self.y + other.y
return Point(x,y)
def __sub__(self,other):
x = self.x - other.x
y = self.y - other.y
return Point(x,y)
def __str__(self):
return f"({self.x},{self.y})"
p = Point(5,4)
q = Point(1,5)
print(p, q)
print(p - q)
| notes/datastructures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src='https://raw.githubusercontent.com/autonomio/hyperio/master/logo.png' width=250px>
# ## This notebook is an adaptation of the Breath Cancer Example with Functional Model
# ## Overview
#
# There are four steps to setting up an experiment with Talos:
#
# 1) Imports and data
#
# 2) Creating the Keras model
#
# 3) Defining the Parameter Space Boundaries
#
# 4) Running the Experiment
# ## 1. The Required Inputs and Data
# +
import talos as ta
import wrangle as wr
import pandas as pd
from keras.models import Sequential, Model
from keras.layers import Dropout, Dense, Input
from keras.optimizers import Adam, Nadam
from keras.activations import relu, elu, sigmoid
from keras.losses import binary_crossentropy
# +
# then we load the dataset
x, y = ta.datasets.breast_cancer()
# and normalize every feature to mean 0, std 1
x = wr.mean_zero(pd.DataFrame(x)).values
# -
# ## 2. Creating the Keras Model
# first we have to make sure to input data and params into the function
def breast_cancer_model(x_train, y_train, x_val, y_val, params):
inputs = Input(shape=(x_train.shape[1],))
layer = Dense(params['first_neuron'], activation=params['activation'],
kernel_initializer=params['kernel_initializer'])(inputs)
layer = Dropout(params['dropout'])(layer)
outputs = Dense(1, activation=params['last_activation'],
kernel_initializer=params['kernel_initializer'])(layer)
model = Model(inputs, outputs)
model.compile(loss=params['losses'],
optimizer=params['optimizer'](),
metrics=['acc'])
history = model.fit(x_train, y_train,
validation_data=[x_val, y_val],
batch_size=params['batch_size'],
epochs=params['epochs'],
verbose=0)
return history, model
# ## 3. Defining the Parameter Space Boundary
# then we can go ahead and set the parameter space
p = {'first_neuron':[9, 10, 11],
'batch_size': [30],
'epochs': [100],
'dropout': [0],
'kernel_initializer': ['uniform','normal'],
'optimizer': [Nadam, Adam],
'losses': [binary_crossentropy],
'activation':[relu, elu],
'last_activation': [sigmoid]}
# ## 4. Starting the Experiment
# and run the experiment
t = ta.Scan(x=x,
y=y,
model=breast_cancer_model,
grid_downsample=1,
params=p,
dataset_name='breast_cancer',
experiment_no='1',
functional_model=True)
r = ta.Reporting("breast_cancer_1.csv")
| examples/Functional Model Hyperparameter Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
# Load files
fileNames = {
"recogidas" : {"fileName" : "Recogidas.csv", "sep" : ";"},
"elementos" : {"fileName" : "elementos.csv", "sep" : ";"},
"posiciones" : {"fileName": "posiciones.csv", "sep" : "\t"},
"vehiculos" : {"fileName": "vehículos.csv", "sep" : ";"},
"x_ele_tra" : {"fileName": "x_ele_tra.csv", "sep" : ";"},
"categoria" : {"fileName": "categoria.csv", "sep" : ";"},
"transponders" : {"fileName": "transponders.csv", "sep" : ";"},
"x_cat_ele" : {"fileName": "x_cat_ele.csv", "sep" : ";"}
}
basePath = "/Users/guillermoblascojimenez/Documents/codeProjects/datathon_sant_cugat/dataset/"
for v in fileNames.values():
v["fileName"] = basePath + v["fileName"]
loadDataFrame = lambda(f) : pd.read_csv(filepath_or_buffer =f['fileName'], header=0, sep=f['sep'])
dfs = {k : loadDataFrame(v) for k,v in fileNames.iteritems()}
dfs['posiciones'][['POS_LATITUD','POS_LONGITUD']] = dfs['posiciones'][['POS_LATITUD','POS_LONGITUD']].applymap(lambda x: x.replace(',','.'))
dfs['posiciones'][['POS_LATITUD','POS_LONGITUD']] = dfs['posiciones'][['POS_LATITUD','POS_LONGITUD']].astype(float)
# +
# Clean files
# remove null columns
countNotNullsForCol = lambda df, col: len(df[df[col].notnull()].index)
countNotNullsForCol(dfs['vehiculos'], 'VEH_CALCA')
for k,v in dfs.iteritems():
for c in v.columns:
if countNotNullsForCol(v,c) is 0:
v.drop(c, 1, inplace=True)
# -
dfs['categoria'].columns
dfs['elementos']
pd.merge(dfs['elementos'], dfs['categoria'], how='left', left_on='ELE_ICO_ID', right_on='CAT_ICO_ID')
dfs['posiciones'][dfs['posiciones']['POS_VEH_ID'] == 77][['POS_LATITUD','POS_LONGITUD','POS_FECHAHORA']].to_csv("/Users/guillermoblascojimenez/Documents/codeProjects/datathon_sant_cugat/dataset/ruta_77.csv")
dfs['posiciones'][['POS_LATITUD','POS_LONGITUD']].applymap(lambda x: x.replace(',','.'))
import numpy as np
i = 0
j = 100
def process(i,j, K=0.00000000000001):
points = dfs['posiciones'][['POS_ID','POS_LONGITUD','POS_LATITUD']]
#LON = 2
#LAT = 41
#px = points['POS_LONGITUD'] - LON
#f = lambda lon, lat: lambda row: (row['POS_LONGITUD'] - lon) + (row['POS_LATITUD'] - lat)
#points.apply(f(LON,LAT),1)
points2 = points[:]
points2.columns = ['POS_ID_2','POS_LONGITUD_2', 'POS_LATITUD_2']
points = points[i:j]
points[0] = points2[0] = 0
m = pd.merge(points, points2, how='outer', on=0)
m['d'] = np.power(m['POS_LONGITUD'] - m['POS_LONGITUD_2'],2) + np.power(m['POS_LATITUD'] - m['POS_LATITUD_2'],2)
m['not_itself'] = m['POS_ID'] != m['POS_ID_2']
m['edge_candidate'] = m['d'] < K
m['edge'] = m.apply(lambda x: x['not_itself'] and x['edge_candidate'], axis=1)
m = m[m['edge']][['POS_ID','POS_LONGITUD','POS_LATITUD','POS_ID_2','POS_LONGITUD_2','POS_LATITUD_2']]
fileName = basePath + 'edges_' + str(i) + '_' + str(j) + ".csv"
m.to_csv(fileName)
print(len(m.index))
# +
process(0,100)
#for i in range(0,len(dfs['posiciones'].index),1000):
# print i
# process(i,i+1000)
# -
| ipython/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from torchvision.datasets import DatasetFolder
import torchvision.transforms as transform
import PIL as Image
import torch.nn as nn
import numpy as np
import torch
# + pycharm={"name": "#%%\n"}
# loader = lambda x : Image.open(x)
# + pycharm={"name": "#%%\n"}
# train_tfm = transform.Compose([
# transform.RandomCrop([128,128]),
# transform.RandomHorizontalFlip(0.5),
# transform.ColorJitter(brightness=0.5),
# transform.RandomAffine(degrees=20, translate=(0.2,0.2), scale=(0.7,1.3)),
# transform.ToTensor()
# ])
#
# test_tfm = transform.Compose([
# transform.RandomCrop([128, 128]),
# transform.ToTensor()
# ])
# + pycharm={"name": "#%%\n"}
# train_path = '../data/hw3/training/labeled'
# train_set1 = DatasetFolder(train_path,)
# + pycharm={"name": "#%%\n"}
input = torch.randn((3, 4, 5, 6))
sum0 = torch.sum(input, dim=3)
print(sum0)
sm = nn.Softmax(dim=3)
output = sm(input)
sum = torch.sum(output, dim=3)
print(sum)
# + pycharm={"name": "#%%\n"}
| hw3_CNN_Classifiction/CodingTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 + Spark 2.x + SystemML
# language: python
# name: pyspark3_2.x
# ---
# # Deep Learning Image Classification using Apache SystemML
#
# This notebook demonstrates how to train a deep learning model on SystemML for the classic [MNIST](http://yann.lecun.com/exdb/mnist/) problem of mapping images of single digit numbers to their corresponding numeric representations, using a classic [LeNet](http://yann.lecun.com/exdb/lenet/)-like convolutional neural network model. See [Neural Networks and Deep Learning](http://neuralnetworksanddeeplearning.com/chap6.html) for more information on neural networks and deep learning.
#
# The downloaded MNIST dataset contains labeled images of handwritten digits, where each example is a 28x28 pixel image of grayscale values in the range [0,255] stretched out as 784 pixels, and each label is one of 10 possible digits in [0,9]. We download 60,000 training examples, and 10,000 test examples, where the images and labels are stored in separate matrices. We then train a SystemML LeNet-like convolutional neural network (i.e. "convnet", "CNN") model. The resulting trained model has an accuracy of 98.6% on the test dataset.
#
# 1. [Download the MNIST data](#download_data)
# 1. [Train a CNN classifier for MNIST handwritten digits](#train)
# 1. [Detect handwritten Digits](#predict)
# <div style="text-align:center" markdown="1">
# 
# Mapping images of numbers to numbers
# </div>
# ### Note: This notebook is supported with SystemML 0.14.0 and above.
# !pip show systemml
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn import datasets
from sklearn.cross_validation import train_test_split # module deprecated in 0.18
#from sklearn.model_selection import train_test_split # use this module for >=0.18
from sklearn import metrics
from systemml import MLContext, dml
# -
ml = MLContext(sc)
print("Spark Version: {}".format(sc.version))
print("SystemML Version: {}".format(ml.version()))
print("SystemML Built-Time: {}".format(ml.buildTime()))
# <a id="download_data"></a>
# ## Download the MNIST data
#
# Download the [MNIST data from the MLData repository](http://mldata.org/repository/data/viewslug/mnist-original/), and then split and save.
# +
mnist = datasets.fetch_mldata("MNIST Original")
print("MNIST data features: {}".format(mnist.data.shape))
print("MNIST data labels: {}".format(mnist.target.shape))
X_train, X_test, y_train, y_test = train_test_split(
mnist.data, mnist.target.astype(np.uint8).reshape(-1, 1),
test_size = 10000)
print("Training images, labels: {}, {}".format(X_train.shape, y_train.shape))
print("Testing images, labels: {}, {}".format(X_test.shape, y_test.shape))
print("Each image is: {0:d}x{0:d} pixels".format(int(np.sqrt(X_train.shape[1]))))
# -
# ### Note: The following command is not required for code above SystemML 0.14 (master branch dated 05/15/2017 or later).
# !svn --force export https://github.com/apache/systemml/trunk/scripts/nn
# <a id="train"></a>
# ## Train a LeNet-like CNN classifier on the training data
# <div style="text-align:center" markdown="1">
# 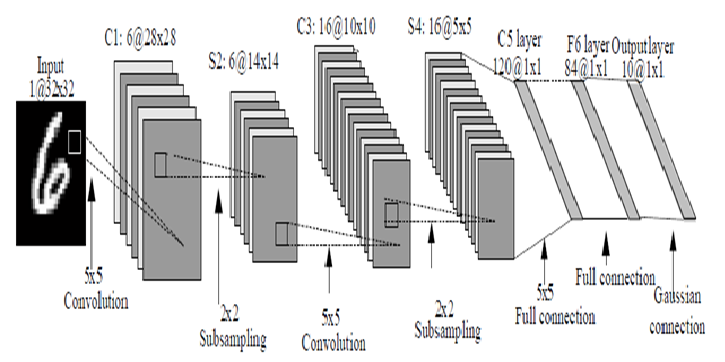
# MNIST digit recognition – LeNet architecture
# </div>
# ### Train a LeNet-like CNN model using SystemML
# +
script = """
source("nn/examples/mnist_lenet.dml") as mnist_lenet
# Scale images to [-1,1], and one-hot encode the labels
images = (images / 255) * 2 - 1
n = nrow(images)
labels = table(seq(1, n), labels+1, n, 10)
# Split into training (55,000 examples) and validation (5,000 examples)
X = images[5001:nrow(images),]
X_val = images[1:5000,]
y = labels[5001:nrow(images),]
y_val = labels[1:5000,]
# Train the model to produce weights & biases.
[W1, b1, W2, b2, W3, b3, W4, b4] = mnist_lenet::train(X, y, X_val, y_val, C, Hin, Win, epochs)
"""
out = ('W1', 'b1', 'W2', 'b2', 'W3', 'b3', 'W4', 'b4')
prog = (dml(script).input(images=X_train, labels=y_train, epochs=1, C=1, Hin=28, Win=28)
.output(*out))
W1, b1, W2, b2, W3, b3, W4, b4 = ml.execute(prog).get(*out)
# -
# Use the trained model to make predictions for the test data, and evaluate the quality of the predictions.
# +
script_predict = """
source("nn/examples/mnist_lenet.dml") as mnist_lenet
# Scale images to [-1,1]
X_test = (X_test / 255) * 2 - 1
# Predict
y_prob = mnist_lenet::predict(X_test, C, Hin, Win, W1, b1, W2, b2, W3, b3, W4, b4)
y_pred = rowIndexMax(y_prob) - 1
"""
prog = (dml(script_predict).input(X_test=X_test, C=1, Hin=28, Win=28, W1=W1, b1=b1,
W2=W2, b2=b2, W3=W3, b3=b3, W4=W4, b4=b4)
.output("y_pred"))
y_pred = ml.execute(prog).get("y_pred").toNumPy()
# -
print(metrics.accuracy_score(y_test, y_pred))
print(metrics.classification_report(y_test, y_pred))
# <a id="predict"></a>
# ## Detect handwritten digits
# Define a function that randomly selects a test image, displays the image, and scores it.
# +
img_size = int(np.sqrt(X_test.shape[1]))
def displayImage(i):
image = (X_test[i]).reshape(img_size, img_size).astype(np.uint8)
imgplot = plt.imshow(image, cmap='gray')
# -
def predictImage(i):
image = X_test[i].reshape(1, -1)
out = "y_pred"
prog = (dml(script_predict).input(X_test=image, C=1, Hin=28, Win=28, W1=W1, b1=b1,
W2=W2, b2=b2, W3=W3, b3=b3, W4=W4, b4=b4)
.output(out))
pred = int(ml.execute(prog).get(out).toNumPy())
return pred
# +
i = np.random.randint(len(X_test))
p = predictImage(i)
print("Image {}\nPredicted digit: {}\nActual digit: {}\nResult: {}".format(
i, p, int(y_test[i]), p == int(y_test[i])))
displayImage(i)
# -
pd.set_option('display.max_columns', 28)
pd.DataFrame((X_test[i]).reshape(img_size, img_size), dtype='uint')
| samples/jupyter-notebooks/Deep_Learning_Image_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_pickle("../grab-ai-safety-data/df_full.pickle")
np.round(df.describe())
# +
mask = df["bookingid"] != df["bookingid"].shift(1)
df["change_in_second"] = df["second"].diff().fillna(0)
df["change_in_bearing"] = np.abs(df["bearing"].diff().fillna(0))
df["change_in_speed"] = np.abs(df["speed"].diff().fillna(0))
df["change_in_acx"] = df["acceleration_x"].diff().fillna(0)
df["change_in_acy"] = df["acceleration_y"].diff().fillna(0)
df["change_in_acz"] = df["acceleration_z"].diff().fillna(0)
df["change_in_gyx"] = df["gyro_x"].diff().fillna(0)
df["change_in_gyy"] = df["gyro_y"].diff().fillna(0)
df["change_in_gyz"] = df["gyro_z"].diff().fillna(0)
df.loc[ mask, [
"change_in_second",
"change_in_bearing",
"change_in_speed",
"change_in_acx",
"change_in_acy",
"change_in_acz",
"change_in_gyx",
"change_in_gyy",
"change_in_gyz"
]] = 0
# -
# ## Duration of trip
# - Quite an obvious difference in trip durations after removing the outliers. Seems that shorter trips tend to be labelled safe.
df.groupby("bookingid").agg(
{
"second" : np.max,
"label" : np.max
}
).groupby("label").agg(
{
"second" : [lambda x: np.percentile(x, q=50), np.mean]
}
)
df_time = df.groupby("bookingid").agg(
{
"second" : np.max,
"label" : np.max
}
)
df_time.sort_values("second")
# ## Distance of trip
# - There is a weird entry with a change in time at the last entry
# - Looks like there are 5 trips with ridiculous trip durations
df_dist = df.copy()
df_dist["distance_covered"] = df_dist["change_in_second"] * df_dist["speed"]
df_dist.groupby("bookingid").agg(
{
"distance_covered" : np.sum,
"label" : np.max
}
).groupby("label").mean()
df_dist.loc[df_dist["bookingid"] == 1503238553722, :][list(df.columns)]["second"].max()
duration_check = df_dist.groupby("bookingid").agg(
{
"second" : np.max,
"label" : np.max
}
)
duration_check.loc[duration_check["label"] == 0, :] \
.groupby("second") \
.count().sort_values("second", ascending=False).reset_index()["second"][1]
# + [markdown] heading_collapsed=true
# ## Acceleration
# - Let's try euclidean first to see if that works
# - Mean of bookings, mean of label: no significant difference ~1% higher only
# - Max of the bookings, mean of label: ~12% higher for dangerous trips
# + hidden=true
df_acc = df.copy()
df_acc["acceleration"] = np.sqrt(
(df_acc["acceleration_x"] ** 2) + (df_acc["acceleration_y"] ** 2) + (df_acc["acceleration_z"] ** 2)
)
test = df_acc.groupby("bookingid").agg(
{
"acceleration" : [np.max, np.mean],
"label" : [np.max]
}
)
test.columns = test.columns.get_level_values(0)
test.groupby("label").agg("mean")
# + [markdown] heading_collapsed=true
# ## Gyro
# - Same technique with gyro to see
# - Max of bookings, mean of label: ~47% higher for dangerous trips
# - Mean of bookings, mean of label: ~36% higher for dangerous trips
# + hidden=true
df_gyro = df.copy()
df_gyro["gyro"] = np.sqrt(
(df_gyro["gyro_x"] ** 2) + (df_gyro["gyro_y"] ** 2) + (df_gyro["gyro_z"] ** 2)
)
test = df_gyro.groupby("bookingid").agg(
{
"gyro" : [np.max, np.mean],
"label" : [np.max]
}
)
test.columns = test.columns.get_level_values(0)
test.groupby("label").agg("mean")
# + [markdown] heading_collapsed=true
# ## Speed
# - More variations for speed
# - Max of bookings, mean of label: 5% higher for dangerous trips
# - Mean of bookings, mean of label: 10% lower for dangerous trips
# + hidden=true
df_spd = df.copy()
test = df_spd.groupby("bookingid").agg(
{
"speed" : [np.max, np.mean],
"label" : [np.max]
}
)
test.columns = test.columns.get_level_values(0)
test.groupby("label").agg("mean")
# + [markdown] heading_collapsed=true
# ## Bearing
# - Change in bearing?
# + hidden=true
| .ipynb_checkpoints/2. Safety Challenge - Exploratory Data Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="SKvGdvJC7g0Y"
# !pip install otter-grader==1.1.6
from google.colab import drive
drive.mount('/content/gdrive')
# %cd /content/gdrive/MyDrive/'Colab Notebooks'/Colab-data-8/lab/lab08
# + deletable=false editable=false id="i24jVwfU7aBQ"
# Initialize Otter
import otter
grader = otter.Notebook()
# + [markdown] id="IkYxch0O7aBR"
# # Lab 8: Normal Distribution and Variance of Sample Means
#
# Welcome to Lab 8!
#
# In today's lab, we will learn about [the variance of sample means](https://www.inferentialthinking.com/chapters/14/5/variability-of-the-sample-mean.html) as well as [the normal distribution](https://www.inferentialthinking.com/chapters/14/3/SD_and_the_Normal_Curve.html).
# + id="UaP-Xcgz7aBT"
# Run this cell, but please don't change it.
# These lines import the Numpy and Datascience modules.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
# These lines load the tests.
from client.api.notebook import Notebook
# + [markdown] id="QnXj3O2M7aBU"
# # 1. Normal Distributions
#
# When we visualize the distribution of a sample, we are often interested in the mean and the standard deviation of the sample (for the rest of this lab, we will abbreviate “standard deviation” as “SD”). These two summary statistics can give us a bird’s eye view of the distribution - by letting us know where the distribution sits on the number line and how spread out it is, respectively.
# + [markdown] id="PLA2yNOs7aBW"
# We want to check if the data is linearly related, so we should look at the data.
# + [markdown] deletable=false editable=false id="FNLCUpCZ7aBX"
# **Question 1.1.** The next cell loads the table `births` from lecture, which is a large random sample of US births and includes information about mother-child pairs.
#
# Plot the distribution of mother’s ages from the table. Don’t change the last line, which will plot the mean of the sample on the distribution itself.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# -->
# + deletable=false id="SbOb6kFM7aBY"
births = Table.read_table('baby.csv')
...
# Do not change this line
plt.scatter(np.mean(births.column("Maternal Age")), 0, color='red', s=50);
# + [markdown] id="iKn8Rgop7aBZ"
# From the plot above, we can see that the mean is the center of gravity or balance point of the distribution. If you cut the distribution out of cardboard, and then placed your finger at the mean, the distribution would perfectly balance on your finger. Since the distribution above is right skewed (which means it has a long right tail), we know that the mean of the distribution is larger than the median, which is the “halfway” point of the data. Conversely, if the distribution had been left skewed, we know the mean would be smaller than the median.
# + [markdown] deletable=false editable=false id="NK5wpU4K7aBb"
# **Question 1.2.** Run the following cell to compare the mean (red) and median (green) of the distribution of mothers ages.
#
# <!--
# BEGIN QUESTION
# name: q1_2
# -->
# + id="uQKUde2v7aBc"
births.hist("Maternal Age")
plt.scatter(np.mean(births.column("Maternal Age")), 0, color='red', s=50);
plt.scatter(np.median(births.column("Maternal Age")), 0, color='green', s=50);
# + [markdown] id="B0czoYJn7aBd"
# We are also interested in the standard deviation of mother’s ages. The SD gives us a sense of how variable mothers' ages are around the average mothers' age. If the SD is large, then the mothers' heights should spread over a large range from the mean. If the SD is small, then the mothers' heights should be tightly clustered around the average mother height.
#
# **The SD of an array is defined as the root mean square of deviations (differences) from average**.
#
# Fun fact! σ (Greek letter sigma) is used to represent the SD and μ (Greek letter mu) is used for the mean.
# + [markdown] deletable=false editable=false id="wRk-Vheh7aBe"
# **Question 1.3.** Run the cell below to see the width of one SD (blue) from the sample mean (red) plotted on the histogram of maternal ages.
#
# <!--
# BEGIN QUESTION
# name: q1_3
# -->
# + for_assignment_type="solution" id="MfvSQuF57aBf"
age_mean = ...
age_sd = ...
births.hist("Maternal Age")
plt.scatter(age_mean, 0, color='red', s=50);
plt.scatter(age_mean+age_sd, 0, marker='^', color='blue', s=50);
plt.scatter(age_mean-age_sd, 0, marker='^', color='blue', s=50);
# + [markdown] id="HYoefaGp7aBf"
# In this histogram, the standard deviation is not easy to identify just by looking at the graph.
#
# However, the distributions of some variables allow us to easily spot the standard deviation on the plot. For example, if a sample follows a *normal distribution*, the standard deviation is easily spotted at the point of inflection (the point where the curve begins to change the direction of its curvature) of the distribution.
# + [markdown] deletable=false editable=false id="_KPAVpbs7aBg"
# **Question 1.4.** Fill in the following code to examine the distribution of maternal heights, which is roughly normally distributed. We’ll plot the standard deviation on the histogram, as before - notice where one standard deviation (blue) away from the mean (red) falls on the plot.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# -->
# + id="A3_dTmdf7aBh"
height_mean = ...
height_sd = ...
births.hist("Maternal Height", bins=np.arange(55,75,1))
plt.scatter((height_mean), 0, color='red', s=50);
plt.scatter(height_mean+height_sd, 0, marker='^', color='blue', s=50);
plt.scatter(height_mean-height_sd, 0, marker='^', color='blue', s=50);
# + [markdown] id="GZ1loSVY7aBj"
# We don’t always know how a variable will be distributed, and making assumptions about whether or not a variable will follow a normal distribution is dangerous. However, the Central Limit Theorem defines one distribution that always follows a normal distribution. The distribution of the *sums* and *means* of many large random samples drawn with replacement from a single distribution (regardless of the distribution’s original shape) will be normally distributed. Remember that the Central Limit Theorem refers to the distribution of a *statistic* calculated from a distribution, not the distribution of the original sample or population. If this is confusing, ask a TA!
#
# The next section will explore distributions of sample means, and you will see how the standard deviation of these distributions depends on sample sizes.
# + [markdown] id="FCMNkVM_7aBk"
# # 2. Variability of the Sample Mean
#
# By the [Central Limit Theorem](https://www.inferentialthinking.com/chapters/14/4/Central_Limit_Theorem.html), the probability distribution of the mean of a large random sample is roughly normal. The bell curve is centered at the population mean. Some of the sample means are higher and some are lower, but the deviations from the population mean are roughly symmetric on either side, as we have seen repeatedly. Formally, probability theory shows that the sample mean is an **unbiased estimate** of the population mean.
#
# In our simulations, we also noticed that the means of larger samples tend to be more tightly clustered around the population mean than means of smaller samples. In this section, we will quantify the [variability of the sample mean](https://www.inferentialthinking.com/chapters/14/5/Variability_of_the_Sample_Mean.html) and develop a relation between the variability and the sample size.
#
# Let's take a look at the salaries of employees of the City of San Francisco in 2014. The mean salary reported by the city government was about $75,463.92.
#
# *Note: If you get stuck on any part of this lab, please refer to [chapter 14 of the textbook](https://www.inferentialthinking.com/chapters/14/Why_the_Mean_Matters.html).*
# + id="rfPsX7yp7aBl"
salaries = Table.read_table('sf_salaries_2014.csv').select("salary")
salaries
# + id="H5Hq-QBT7aBm"
salary_mean = np.mean(salaries.column('salary'))
print('Mean salary of San Francisco city employees in 2014: ', salary_mean)
# + id="5lGkTiSz7aBm"
salaries.hist('salary', bins=np.arange(0, 300000+10000*2, 10000))
plt.scatter(salary_mean, 0, marker='^', color='red', s=100);
plt.title('2014 salaries of city of SF employees');
# + [markdown] id="XKE2pAKk7aBn"
# Clearly, the population does not follow a normal distribution. Keep that in mind as we progress through these exercises.
#
# Let's take random samples *with replacement* and look at the probability distribution of the sample mean. As usual, we will use simulation to get an empirical approximation to this distribution.
# + [markdown] deletable=false editable=false id="z2yMo3fB7aBn"
# **Question 2.1.** Define a function `one_sample_mean`. Its arguments should be `table` (the name of a table), `label` (the label of the column containing the variable), and `sample size`(the number of employees in the sample). It should sample with replacement from the table and
# return the mean of the `label` column of the sample.
#
# <!--
# BEGIN QUESTION
# name: q2_1
# -->
# + id="4FJvd6xL7aBo"
def one_sample_mean(table, label, sample_size):
new_sample = ...
new_sample_mean = ...
...
# + deletable=false editable=false id="BJ0gsmUj7aBo"
grader.check("q2_1")
# + [markdown] deletable=false editable=false id="2y3G7Ctz7aBp"
# **Question 2.2.** Use `one_sample_mean` to define a function `simulate_sample_mean`. The arguments are the name of the table, the label of the column containing the variable, the sample size, and the number of simulations.
#
# The function should sample with replacement from the table and calculate the mean of each sample. It should save the sample means in an array called `means`. The remaining code in the function displays an empirical histogram of the sample means.
#
# <!--
# BEGIN QUESTION
# name: q2_2
# -->
# + deletable=false id="ngt3XoAm7aBp"
"""Empirical distribution of random sample means"""
def simulate_sample_mean(table, label, sample_size, repetitions):
means = make_array()
for i in np.arange(repetitions):
new_sample_mean = ...
means = ...
sample_means = Table().with_column('Sample Means', means)
# Display empirical histogram and print all relevant quantities – don't change this!
sample_means.hist(bins=20)
plt.xlabel('Sample Means')
plt.title('Sample Size ' + str(sample_size))
print("Sample size: ", sample_size)
print("Population mean:", np.mean(table.column(label)))
print("Average of sample means: ", np.mean(means))
print("Population SD:", np.std(table.column(label)))
print("SD of sample means:", np.std(means))
return np.std(means)
# + [markdown] id="7nmy82qc7aBq"
# Verify with your neighbor or TA that you've implemented the function above correctly. If you haven't implemented it correctly, the rest of the lab won't work properly, so this step is crucial.
# + [markdown] id="jV24bjEN7aBr"
# In the following cell, we will create a sample of size 100 from `salaries` and graph it using our new `simulate_sample_mean` function.
#
# *Hint: You should see a distribution similar to something we've been talking about. If not, check your function*
# + id="w-GORR7d7aBr"
simulate_sample_mean(salaries, 'salary', 100, 10000)
plt.xlim(50000, 100000);
# + [markdown] deletable=false editable=false id="Uq0eA9v77aBs"
# **Question 2.3.** Simulate two sample means, one for a sample of 400 salaries and one for a sample of 625 salaries. In each case, perform 10,000 repetitions. Don't worry about the `plots.xlim` line – it just makes sure that all of the plots have the same x-axis.
#
# <!--
# BEGIN QUESTION
# name: q2_3
# -->
# + for_assignment_type="solution" id="WfVImY3g7aBs"
simulate_sample_mean(..., ..., ..., ...)
plt.xlim(50000, 100000);
plt.show();
print('\n')
simulate_sample_mean(..., ..., ..., ...)
plt.xlim(50000, 100000);
plt.show();
# + [markdown] deletable=false editable=false id="EujWRKKQ7aBs"
# **Question 2.4.** Assign `q2_4` to an array of numbers corresponding to true statement(s) about the plots from 2.3.
#
# 1. We see the Central Limit Theorem (CLT) in action because the distributions of the sample means are bell-shaped.
# 2. We see the Law of Averages in action because the distributions of the sample means look like the distribution of the population.
# 3. One of the conditions for CLT is that we have to draw a small random sample with replacement from the population.
# 4. One of the conditions for CLT is that we have to draw a large random sample with replacement from the population.
# 5. One of the conditions for CLT is that the population must be normally distributed.
# 6. Both plots in 2.3 are roughly centered around the population mean.
# 7. Both plots in 2.3 are roughly centered around the mean of a particular sample.
# 8. The distribution of sample means for sample size 625 has less variability than the distribution of sample means for sample size 400.
# 9. The distribution of sample means for sample size 625 has more variability than the distribution of sample means for sample size 400.
#
# <!--
# BEGIN QUESTION
# name: q2_4
# -->
# + for_assignment_type="solution" id="b7txklE47aBt"
q2_4 = ...
# + deletable=false editable=false id="taRrMc-g7aBt"
grader.check("q2_4")
# + [markdown] id="LocDqW1o7aBu"
# Below, we'll look at what happens when we take an increasing number of resamples of a fixed sample size. Notice what number in the code changes, and what stays the same. How does the distribution of the resampled means change?
# + id="w4F0Hs_Z7aBu"
simulate_sample_mean(salaries, 'salary', 100, 500)
plt.xlim(50000, 100000);
# + id="ati76Dux7aBu"
simulate_sample_mean(salaries, 'salary', 100, 1000)
plt.xlim(50000, 100000);
# + id="NhaEXOFG7aBv"
simulate_sample_mean(salaries, 'salary', 100, 5000)
plt.xlim(50000, 100000);
# + id="LXvjoV7V7aBv"
simulate_sample_mean(salaries, 'salary', 100, 10000)
plt.xlim(50000, 100000);
# + [markdown] id="dMIa6LHj7aBv"
# What did you notice about the distributions of sample means in the four histograms above? Discuss with your neighbors. If you're unsure of your conclusion, ask your TA.
# + [markdown] deletable=false editable=false id="q3dQLjNs7aBv"
# **Question 2.5.** Assign the variable `SD_of_sample_means` to the integer corresponding to your answer to the following question:
#
# When I increase the number of resamples that I take, for a fixed sample size, the SD of my sample means will...
#
# 1. Increase
# 2. Decrease
# 3. Stay about the same
# 4. <NAME>
#
#
# <!--
# BEGIN QUESTION
# name: q2_5
# -->
# + deletable=false id="sQxoHclA7aBw"
SD_of_sample_means = ...
# + deletable=false editable=false id="_9agP7QW7aBw"
grader.check("q2_5")
# + [markdown] deletable=false editable=false id="qBe4_uk37aBw"
# **Question 2.6.** Let's think about how the relationships between population SD, sample SD, and SD of sample means change with varying sample size. Which of the following is true? Assign the variable `pop_vs_sample` to an array of integer(s) that correspond to true statement(s).
#
# 1. Sample SD gets smaller with increasing sample size.
# 2. Sample SD gets larger with increasing sample size.
# 3. Sample SD becomes more consistent with population SD with increasing sample size.
# 4. SD of sample means gets smaller with increasing sample size.
# 5. SD of sample means gets larger with increasing sample size.
# 6. SD of sample means stays the same with increasing sample size.
#
# <!--
# BEGIN QUESTION
# name: q2_6
# -->
# + deletable=false id="kmkbyn2p7aBx"
pop_vs_sample = ...
# + deletable=false editable=false id="7ltu6AME7aBx"
grader.check("q2_6")
# + [markdown] id="CsqBMPox7aBx"
# Run the following three cells multiple times and examine how the sample SD and the SD of sample means change with sample size.
#
# The first histogram is of the sample; the second histogram is the distribution of sample means with that particular sample size. Adjust the bins as necessary.
# + id="tb8SqBjF7aBx"
sample_10 = salaries.sample(10)
sample_10.hist("salary")
plt.title('Distribution of salary for sample size 10')
print("Sample SD: ", np.std(sample_10.column("salary")))
simulate_sample_mean(salaries, 'salary', 10, 1000)
plt.xlim(5,120000);
plt.ylim(0, .0001);
plt.title('Distribution of sample means for sample size 10');
# + id="Dt0VmScs7aBy"
sample_200 = salaries.sample(200)
sample_200.hist("salary")
plt.title('Distribution of salary for sample size 200')
print("Sample SD: ", np.std(sample_200.column("salary")))
simulate_sample_mean(salaries, 'salary', 200, 1000)
plt.xlim(5,100000)
plt.ylim(0, .00015);
plt.title('Distribution of sample means for sample size 200');
# + id="vLWq-fi27aBy"
sample_1000 = salaries.sample(1000)
sample_1000.hist("salary")
plt.title('Distribution of salary for sample size 1000')
print("Sample SD: ", np.std(sample_1000.column("salary")))
simulate_sample_mean(salaries, 'salary', 1000, 1000)
plt.xlim(5,100000)
plt.ylim(0, .00025);
plt.title('Distribution of sample means for sample size 1000');
# + [markdown] id="fl1BD2Kq7aBy"
# You should notice that the distribution of means gets narrower and spikier, and that the distribution of the sample increasingly looks like the distribution of the population as we get to larger sample sizes.
#
# Let's illustrate these trends. Below, you will see how the sample SD changes with respect to sample size (N). The blue line is the population SD.
# + id="Y9ZrbL1c7aBy"
# Don't change this cell, just run it!
pop_sd = np.std(salaries.column('salary'))
sample_sds = make_array()
sample_sizes = make_array()
for i in np.arange(10, 500, 10):
sample_sds = np.append(sample_sds, [np.std(salaries.sample(i).column("salary")) for d in np.arange(100)])
sample_sizes = np.append(sample_sizes, np.ones(100) * i)
Table().with_columns("Sample SD", sample_sds, "N", sample_sizes).scatter("N", "Sample SD")
matplotlib.pyplot.axhline(y=pop_sd, color='blue', linestyle='-');
# + [markdown] id="iYphIt6D7aBz"
# The next cell shows how the SD of the sample means changes relative to the sample size (N).
# + id="dEr5Qphe7aBz"
# Don't change this cell, just run it!
def sample_means(sample_size):
means = make_array()
for i in np.arange(1000):
sample = salaries.sample(sample_size).column('salary')
means = np.append(means, np.mean(sample))
return np.std(means)
sample_mean_SDs = make_array()
for i in np.arange(50, 1000, 100):
sample_mean_SDs = np.append(sample_mean_SDs, sample_means(i))
Table().with_columns("SD of sample means", sample_mean_SDs, "Sample Size", np.arange(50, 1000, 100))\
.plot("Sample Size", "SD of sample means")
# + [markdown] id="HovTnQmC7aBz"
# From these two plots, we can see that the SD of our *sample* approaches the SD of our population as our sample size increases, but the SD of our *sample means* (in other words, the variability of the sample mean) decreases as our sample size increases.
# + [markdown] deletable=false editable=false id="An8Gf7lo7aBz"
# **Question 2.7.** Is there a relationship between the sample size and the standard deviation of the sample mean? Assign `q2_7` to the number corresponding to the statement that answers this question.
#
# 1. The SD of the sample means is inversely proportional to the square root of sample size.
# 2. The SD of the sample means is directly proportional to the square root of sample size.
#
# <!--
# BEGIN QUESTION
# name: q2_7
# -->
# + id="PoMvqgjp7aB0"
q2_7 = ...
# + deletable=false editable=false id="E69aUzC77aB0"
grader.check("q2_7")
# + [markdown] id="8h2mXN6t7aB0"
# Throughout this lab, we have been taking many random samples from a population. However, all of these principles hold for bootstrapped resamples from a single sample. If your original sample is relatively large, all of your re-samples will also be relatively large, and so the SD of resampled means will be relatively small.
#
# In order to change the variability of your sample mean, you’d have to change the size of the original sample from which you are taking bootstrapped resamples.
# + [markdown] id="_n7tr5lI7aB0"
# That's it! You've completed Lab 8. There weren't many tests, but there were a lot of points at which you should've stopped and understood exactly what was going on. Consult the textbook or ask your TA if you have any other questions!
#
# Be sure to
# - **run all the tests** (the next cell has a shortcut for that),
# - **Save and Checkpoint** from the `File` menu,
# - **run the last cell to submit your work**,
# - and ask one of the staff members to check you off.
# + [markdown] deletable=false editable=false id="kRiu94mi7aB1"
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false id="c2uMCVVw7aB1"
grader.check_all()
# + [markdown] deletable=false editable=false id="NiItmKW27aB1"
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
# + deletable=false editable=false id="lyKaoVsc7aB1"
# Save your notebook first, then run this cell to export your submission.
grader.export(pdf=False)
# + [markdown] id="h4TC4ErJ7aB2"
#
| lab/lab08/lab08.ipynb |