code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/muthonioprah/Football-Match-Prediction-Results/blob/main/Moringa_Data_Science_Core_W6_Independent_Project_2022_2_Oprah_Muthoni_Python_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Nbx0CK9IczOG"
# # **Introduction**
# You have been recruited as a football analyst in a company - Mchezopesa Ltd and tasked to accomplish the task below.
#
# A prediction result of a game between team 1 and team 2, based on who's home and who's away, and on whether or not the game is friendly (include rank in your training).
#
# You have two possible approaches (as shown below) given the datasets that will be provided
#
# Input: Home team, Away team, Tournament type (World cup, Friendly, Other)
#
# Approach 1: Polynomial approach
#
# What to train given:
#
# Rank of home team Rank of away team Tournament type Model 1: Predict how many goals the home team scores.
#
# Model 2: Predict how many goals the away team scores.
#
# Approach 2: Logistic approach
#
# Feature Engineering: Figure out from the home team’s perspective if the game is a Win, Lose or Draw (W, L, D)
#
# #**Research Question**
#
# Predict how many goals the home team scores.
#
# Predict how many goals the away team scores.
#
# Figure out from the home team’s perspective if the game is a Win, Lose or Draw (W, L, D)
#
# #**Defining the metric for success**
#
# A models that successfully predicts how many goals the home team scores, how many goals the away team scores and determine from the home team’s perspective if the game is a Win, Lose or Draw (W, L, D)
# + [markdown] id="o9RKe2iR2jT-"
# # **Data Understanding**
# + id="E1LBbjraZzxr"
#importing the relevant libraries
import pandas as pd
import numpy as np
# + id="O6ohobMEeJ4z"
#reading the csv documents
ranking = pd.read_csv('/content/fifa_ranking.csv')
results = pd.read_csv('/content/results.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 392} id="fEmaNNJKeJ0a" outputId="5d1aca18-73ae-450c-c56f-89cffc46c879"
#studying the datasets
#we start with the ranking datset
print(ranking.shape)
print("****************************************************************************")
ranking.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="DUvloiPMnl1f" outputId="dcb7009e-469c-4601-eb81-c4804d16adff"
#viewing the results datset
print(results.shape)
print("****************************************************************************")
results.head()
# + colab={"base_uri": "https://localhost:8080/"} id="sbPAL_1-eJt_" outputId="879cdfcd-49ec-47a8-ecdb-11f01b42b4e3"
#displaying the dataset information
ranking.info()
print("****************************************************************************")
print(results.info())
# + colab={"base_uri": "https://localhost:8080/", "height": 364} id="Gi5RqUF-eJnu" outputId="426222c4-ea8f-4f3f-f200-8a8edd4885bc"
#showing the descriptive statisics for this first dataset
ranking.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="ohyRRklxnGw4" outputId="beaf7fb3-4d5e-4c7f-d373-d859fb6e8ce3"
#descriptive statistics
results.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="fb0w0PoWeJhe" outputId="2f8db3f1-b317-41ff-ca7b-ea23da2af5b9"
#finding the null values
print(ranking.isnull().any())
print("***********************************************")
print(results.isnull().any())
# + [markdown] id="0wp5z751oUU0"
# Both datasets have no null values
# + colab={"base_uri": "https://localhost:8080/"} id="qnBHWVWieJe3" outputId="ac382959-51f7-4670-ede9-7ee4a429f9e5"
#checking for duplicate values
print("Sum of duplicated records in the ranking datset is " + str(ranking.duplicated().sum()))
print("Sum of duplicated records in the results datset is " + str(results.duplicated().sum()))
# + [markdown] id="RQtyTqrAn-4C"
#
#
# > The results dataset has no duplicate values
#
#
#
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="j4CV49yAeJb3" outputId="489126e6-9088-48a1-c11b-8ed0b0675469"
#viewing the exact duplicated records in the ranking dataset
dup_rank = ranking[ranking.duplicated()]
dup_rank
# + [markdown] id="uEGS169yvUcn"
# ## **Data Cleaning**
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="kL5fE_oCeJYm" outputId="ee461485-05dc-459a-c967-613830f44c08"
#dropping the columns thatwont be usedin this analysis
colz = ['country_abrv', 'total_points','previous_points', 'rank_change', 'cur_year_avg', 'cur_year_avg_weighted', 'last_year_avg', 'last_year_avg_weighted',
'two_year_ago_avg', 'two_year_ago_weighted', 'three_year_ago_avg','three_year_ago_weighted', 'confederation']
ranking_colz = ranking.drop(columns=colz)
ranking_colz.head()
# + colab={"base_uri": "https://localhost:8080/"} id="bMoebikGeJVO" outputId="6f62ebf0-e5f9-47ea-cff8-97ba80d04d03"
#checking the shape of the new dataframe
ranking_colz.shape
# + id="8XP4Q_vDt7RU" colab={"base_uri": "https://localhost:8080/"} outputId="0a7c7a35-f01c-43d4-b96c-f72ec04c4071"
#changing the column with the datesto datetime format
anking_colz['rank_date'] = pd.to_datetime(ranking_colz['rank_date'], format='%Y-%m-%d')
ranking_colz.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="i5R8lf2hQy6x" outputId="f5fe20ac-c6b8-4a46-9195-b20d416b3b9f"
#changing the column with the datesto datetime format
results['date'] = pd.to_datetime(results['date'], format='%Y-%m-%d')
results.dtypes
# + id="TLlxcCE-t7OG" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="6e05e88f-50a7-48b8-fafa-4f5b9e965f2b"
#the two datasets have the date column in common s the will be merged on that axis
#we first extract the year from the date time column
ranking_colz['year'] = ranking_colz['rank_date'].dt.year
ranking_colz.head()
# + id="j9xjXV39t7Kv" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="1432ab0e-a89c-46ed-fc88-9a9fdf8b4785"
#do the same for the results dataframe
results['year'] = results['date'].dt.year
results.tail()
# + id="ml2A5spXt7HP"
#rename columns in the ranking table for uniformity
ranking_colz.rename(columns = {'country_full' : 'country'}, inplace = True)
# + id="DTVTUeWht7D7" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="7fea6bb0-4752-4492-a438-0957f6d418c9"
#merging the 2 datasets based on year and home team
df = pd.merge(results,ranking_colz, left_on=['home_team', 'year'], right_on=['home_team', 'year'], how='inner')
df.head()
# + id="zOL0GVLYt62R" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="30a27388-b34f-4652-b39f-6da6534bfad4"
# Then with the away teams' ranking
df1 = pd.merge(results,ranking_colz, left_on=['away_team', 'year'], right_on=['home_team', 'year'], how='inner')
df1.head(5)
# + id="juvJHqL6t6yp" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="2ef8e322-7bb6-44bb-fdb0-64a125dec113"
#to merge both home and away
merged = pd.merge(df, df1, how='inner', left_on=['date','home_team', 'away_team','home_score','away_score'], right_on=['date','country','away_team',
'home_score', 'away_score'])
merged.head()
# + id="70M8gaLNt6uh" colab={"base_uri": "https://localhost:8080/"} outputId="e177eb8b-e723-4939-e61a-db99950984ae"
# Checking the shape of the fully merged dataframe.
merged.shape
# + id="AnY0lc2Gt6qi" colab={"base_uri": "https://localhost:8080/"} outputId="eaf3a557-7ef3-4566-c814-573d7104859f"
#checing for duplicated records
erged.duplicated().sum()
# + id="dC89OX9gt6nd" colab={"base_uri": "https://localhost:8080/"} outputId="7df6132a-7316-41c0-eab6-6921158bd8b6"
#dropping duplicated records
merged.drop_duplicates(keep= 'first', inplace= True)
merged.shape
# + id="BaFI6NlVt6ju" colab={"base_uri": "https://localhost:8080/"} outputId="bf99aab7-129f-4318-f782-6a87b260f8f4"
#check for null records
merged.isnull().sum()
# + [markdown] id="gJSrP8wfbsJq"
# There are no null records
# + id="W8cxWv9ct6f9" colab={"base_uri": "https://localhost:8080/", "height": 259} outputId="d338822b-189c-481a-8424-2960aad35eb4"
#dropping the similar columns from merged dataset
dff = merged.drop(columns=['rank_date_x', 'year_x'])
dff.rename(columns={'rank_x':'home_team_rank', 'rank_y':'away_team_rank'},inplace=True)
dff.head(3)
# + id="Z2S8TxNelTvJ"
# using a function to create the status column based on Home team
def my_function(x):
if x['home_score'] == x['away_score']:
status = 'Draw'
elif x['home_score'] > x['away_score']:
status = 'Win'
else:
status = 'loss'
return status
dff['score'] = dff.apply(my_function, axis=1)
# + [markdown] id="TnOEyvgM2Y9T"
# # **Univariate Analysis**
# + id="aY9XdlKft6cY" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="63f8509a-c641-48c0-f042-3e2000453028"
# We will check for outliers in our numerical columns
import matplotlib.pyplot as plt
col= ['home_score', 'away_score','home_team_rank', 'away_team_rank']
for i in col:
fig = plt.figure(figsize = (10,7))
plt.boxplot(dff[i])
plt.title(i)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="WGZSG7C_bB0m" outputId="80104753-2cbc-47e1-e5c6-b9884b09e994"
# Frequency Table on distribution of tournaments
dff.tournament_x.value_counts().head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="IU7dVJVVbBsP" outputId="603c43aa-7bd2-458d-d64a-d0305285851e"
# Histograms of our numerical values
import seaborn as sns
for i in col:
plt.figure(figsize = (10,3), dpi=100)
sns.displot(dff[i], kde = True, color = 'aqua')
plt.title(i, color = 'Black')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 366} id="2QeflVrLbBwo" outputId="de9f3cc8-0f5e-4831-e4b6-14eed822e430"
# Bar chart on Status
plt.figure(figsize = (10,5))
dff.groupby('score').size().plot(kind='bar', color='black')
plt.title('Status')
plt.ylabel('Total')
plt.xlabel('score')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="31ctIkAcbBoL" outputId="f981657e-5c96-40bb-97a3-9fe606d18a05"
# Checking the descriptive statistics
dff.describe()
# + [markdown] id="1uY6eiaV2RRH"
# # **Bivariate Aalysis**
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="8Bgy14w_bBie" outputId="e88a171c-b63c-4b27-815d-d7ddc568f918"
# Home teams with the most wins
win= dff[(dff['score']== 'Win')]
win1 = win.groupby(['home_team'])['score'].size().sort_values(ascending=True).head()
win1.plot(kind = 'bar', title= 'Top 5 Home Team with the most wins', ylabel = 'Wins' , xlabel = 'Country' , figsize =(10,6) , color = 'violet')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="SXjrAF_ibBca" outputId="6d401977-5320-4cd6-ae93-6989f964b7b8"
# Home teams with the most draws
draws= dff[(dff['score']== 'Draw')]
draw1 = draws.groupby(['home_team'])['score'].size().sort_values(ascending= True).head()
draw1.plot(kind = 'bar', title= 'Top 5 Home Team with the most Draws', ylabel = 'Draws' , xlabel = 'Country' , figsize =(10,6) , color = 'purple')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 466} id="qL_tZhQSbBXh" outputId="2e4942cd-13eb-40e9-9195-717bf5731fe6"
# Home teams with the most lossess
loss =dff[(dff['score']== 'loss')]
loss1 = loss.groupby(['home_team'])['score'].size().sort_values(ascending= True).head()
loss1.plot(kind = 'bar', title= 'Top 5 Home Team with the most losses', ylabel = 'loss' , xlabel = 'Country' , figsize =(10,6) , color = 'pink')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="pO-iDvtAfefw" outputId="0cb75493-060b-4b7f-ff05-b0925f8081c5"
#Most Played Tournament
tournament= dff['tournament_x'].value_counts()
print(tournament)
# + [markdown] id="tIj6B_RL2I3g"
# # **Multivariate Analysis**
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="zUi2swHpbBTo" outputId="896ff1aa-068e-4e58-86c8-9dc22a9060d0"
# Asses correlation between our variables
dff.corr()
# + colab={"base_uri": "https://localhost:8080/", "height": 672} id="E4ZaJ9TWbBQL" outputId="db12a759-ad73-4c12-90c0-e434fe22cfa7"
# Plot a heatmap
fig, heat = plt.subplots(figsize = (10,10))
heat = sns.heatmap(dff.corr(), annot = True, cmap = 'coolwarm')
# + colab={"base_uri": "https://localhost:8080/", "height": 761} id="LVHxi55BfIaB" outputId="4ad79db3-9fa4-474a-a5d2-0cef5531d692"
# Plotting a pairplot of all the numerical values
plt.figure(figsize = (7, 5), dpi=100)
sns.pairplot(dff[col])
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="OnQS17iRfIOW" outputId="e4c0d0fa-6c84-44c7-c017-ea453d36a0cf"
# Predict the number of goals scored by the away team having away_score as the dependent variable
# We'll drop columns that are irrelevant in predicting the goals scored by away team
independent = dff.drop(columns=['date', 'home_team', 'away_team', 'away_score', 'tournament_x'])
corr = independent.corr()
corr
# + id="BvjVdbXffICn"
# Import libraries
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
# + [markdown] id="EUiXgLCmjIcR"
# # **Logistic regression**
# + colab={"base_uri": "https://localhost:8080/", "height": 270} id="xt3NPf6uisBa" outputId="86829e4b-00e9-42bc-cfb4-cd6d3ae29aa9"
#Applying label encoding to our dataset
from sklearn.preprocessing import LabelEncoder
dff1= dff.apply(LabelEncoder().fit_transform)
dff1.head()
# + id="bpEWDO1buQnK"
X = dff1.drop(['home_score', 'away_score'], axis = 1)
y = dff1['home_score']
# + id="bv5wOdz8uBAZ"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = .2, random_state=0)
# + colab={"base_uri": "https://localhost:8080/"} id="4lpXFf4VirwI" outputId="6809dbe5-48a4-4816-a183-620e7755e90a"
# Using our model to make a prediction
from sklearn.linear_model import LinearRegression, LogisticRegression
model = LogisticRegression()
model = model.fit(X_train, y_train)
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="BlkiAALFjops" outputId="84b5f923-6023-472c-d9eb-7729c2619110"
#comparing the values from the dataset to those predicted
predict = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
predict
# + colab={"base_uri": "https://localhost:8080/"} id="w4_t4OstjomX" outputId="53b15e59-a975-40a7-da83-68a37e6a1630"
#checking the model score
model.score(X_test, y_test)
# + [markdown] id="4a1U5w8XvWu5"
# # **Using Cross Validation**
# + id="icgKf9i0vjTn"
# dependent variable as y
from sklearn.model_selection import KFold, LeaveOneOut
X = dff1.drop(['home_score', 'away_score'], axis = 1).values
y = dff1['home_score'].values
# specifying the number of folds
folds = KFold(n_splits = 5)
# We now assess models based on the folds we created.
RMSES = [] # An array of RMSEs to keep track of the RSME of each model
count = 1 # starting point # helps to keep track of the model number in training
# + colab={"base_uri": "https://localhost:8080/"} id="tmhSC-XXvmUg" outputId="dc90eea1-ed7b-4cc9-d140-3bba4c6e35f3"
for train_index, test_index in folds.split(X):
print('\nTraining model ' + str(count))
# Setting up the train and test based on the split determined by KFold
# With 10 folds we split our data into training and test sets
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
# fitting a Logistic regression model
logreg = LogisticRegression()
logreg=logreg.fit(X_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="fOLmtmDpvmOX" outputId="afcba4e7-59dc-4b99-9882-757f865ef298"
from sklearn import metrics
from sklearn.metrics import confusion_matrix
# Assess the accuracy of the model
y_pred = logreg.predict(X_test)
# Calculating the RMSES of each model
# Appending each RMSE into the list earlier created
rmse_value = np.sqrt(metrics.mean_squared_error(y_test, y_pred))
RMSES.append(rmse_value)
# printing each model RMSE
print('Model ' + str(count) + ' Root Mean Squared Error:',rmse_value)
count = count + 1
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="eWnGoldJvmKd" outputId="dacc8205-f222-4359-8b14-04a4bd2d7e88"
pred = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
pred
# + colab={"base_uri": "https://localhost:8080/"} id="rSL0bFoXvmF1" outputId="9f7c2bf4-34a1-4144-bcbc-c2f4a4d2db59"
#checking the model score
logreg.score(X_test, y_test)
# + [markdown] id="qZb_Q6_80njC"
# # **Challenging the solution**
# A more accurate model can be used to predict the outcome
#
# # **Reviewing the Solution**
#
# Follow up questions
#
# a). Did we have the right data?
# Yes, the data we had enabled us to create a strong model.
#
# b). Do we need other data to answer our question?
# No more data is needed to answer our question.
#
# c). Did we have the right question?
# Yes, the question we had allowed us to create the relevant model.
| Moringa_Data_Science_Core_W6_Independent_Project_2022_2_Oprah_Muthoni_Python_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p36)
# language: python
# name: conda_pytorch_p36
# ---
# %load_ext autoreload
# %autoreload 2
# +
import sys
sys.path.append('../src/')
from __future__ import print_function
import time
import os
import numpy as np
import matplotlib.pyplot as plt
import nltk
import pickle
import pandas as pd
import argparse
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
from torch.autograd import Variable
from utils.Vocabulary import Vocabulary
from utils.ImageDataloader import get_image_dataloader, ImageDataset
from models.ImageCaptioner import ImageCaptioner
# -
images_path = os.environ['HOME'] + '/Database/coco/images/'
captions_path = os.environ['HOME'] + '/programs/cocoapi/annotations/coco_captions.csv'
models_path = '../models/'
batch_size = 64
coco_set = 2014
load_features = True
preload = True
base_model='resnet152' # 'vgg16' # 'resnet152'
embedding_size = 2048 # 25088 # 2048
load_captions = True
vocab_path = '../data/processed/coco_vocab.pkl'
print ("Loading validation data...\r", end="")
val_loader = get_image_dataloader('val',coco_set,
images_path,
vocab_path, captions_path,
batch_size,
embedding_size=embedding_size,
load_features=load_features,
load_captions=load_captions,
model=base_model,
preload=preload)
val_loader.dataset.mode = 'val'
print ("Loading validation data...Done")
vocab_size = val_loader.dataset.get_vocab_size()
start_id = val_loader.dataset.get_idx()[val_loader.dataset.vocab.start_word]
end_id = val_loader.dataset.get_idx()[val_loader.dataset.vocab.end_word]
max_caption_length = val_loader.dataset.max_len
embed_size = 256
hidden_size = 512
rnn_type = 'gru'
# +
captioner = ImageCaptioner(embedding_size, embed_size,
hidden_size, vocab_size,
max_caption_length,
start_id, end_id,
rnn_type='gru')
if torch.cuda.is_available():
captioner.cuda()
# -
model_path = '../models/'
#model_path += 'image_caption-model11-20-0.1226-4.64.pkl'
model_path += 'image_caption-model11-20-0.1226-4.64.pkl'
# +
checkpoint = torch.load(model_path)
captioner.load_state_dict(checkpoint['params'])
captioner.eval()
# +
val_bleu = 0.0
beam_size = 0
for val_id, val_batch in enumerate(val_loader):
idxs, im_embeddings, caption_embeddings = val_batch
if torch.cuda.is_available():
im_embeddings = im_embeddings.cuda()
caption_embeddings = caption_embeddings.cuda()
# Get ground truth captions
refs = val_loader.dataset.get_references(idxs.numpy())
preds = captioner.predict(im_embeddings, beam_size=beam_size)
# Calculate bleu loss per sample in batch
# Sum and add length normalized sum to val_loss
batch_bleu = 0.0
for pred_id in range(len(preds)):
pred = preds[pred_id].cpu().numpy().astype(int)
pred_embed = val_loader.dataset.vocab.decode(pred, clean=True)
batch_bleu += val_loader.dataset.vocab.evaluate(refs[pred_id], pred_embed)
val_bleu += (batch_bleu/len(preds))
# Get training statistics
stats = "Validation step [%d/%d], Bleu: %.4f" \
% (val_id, val_loader.dataset.get_seq_len(),
batch_bleu/len(preds))
print("\r" + stats, end="")
sys.stdout.flush()
if val_id % 250 == 0:
print('\r' + stats)
val_bleu /= val_loader.dataset.get_seq_len()
print ("\nValidation -- bleu: %.4f" % (val_bleu))
# +
val_bleu = 0.0
beam_size = 3
for val_id, val_batch in enumerate(val_loader):
idxs, im_embeddings, caption_embeddings = val_batch
if torch.cuda.is_available():
im_embeddings = im_embeddings.cuda()
caption_embeddings = caption_embeddings.cuda()
# Get ground truth captions
refs = val_loader.dataset.get_references(idxs.numpy())
preds = captioner.predict(im_embeddings, beam_size=beam_size)
# Calculate bleu loss per sample in batch
# Sum and add length normalized sum to val_loss
batch_bleu = 0.0
for pred_id in range(len(preds)):
pred = preds[pred_id].cpu().numpy().astype(int)
pred_embed = val_loader.dataset.vocab.decode(pred, clean=True)
batch_bleu += val_loader.dataset.vocab.evaluate(refs[pred_id], pred_embed)
val_bleu += (batch_bleu/len(preds))
# Get training statistics
stats = "Validation step [%d/%d], Bleu: %.4f" \
% (val_id, val_loader.dataset.get_seq_len(),
batch_bleu/len(preds))
print("\r" + stats, end="")
sys.stdout.flush()
if val_id % 250 == 0:
print('\r' + stats)
val_bleu /= val_loader.dataset.get_seq_len()
print ("\nValidation -- bleu: %.4f" % (val_bleu))
# -
# +
val_bleu = 0.0
beam_size = 5
for val_id, val_batch in enumerate(val_loader):
idxs, im_embeddings, caption_embeddings = val_batch
if torch.cuda.is_available():
im_embeddings = im_embeddings.cuda()
caption_embeddings = caption_embeddings.cuda()
# Get ground truth captions
refs = val_loader.dataset.get_references(idxs.numpy())
preds = captioner.predict(im_embeddings, beam_size=beam_size)
# Calculate bleu loss per sample in batch
# Sum and add length normalized sum to val_loss
batch_bleu = 0.0
for pred_id in range(len(preds)):
pred = preds[pred_id].cpu().numpy().astype(int)
pred_embed = val_loader.dataset.vocab.decode(pred, clean=True)
batch_bleu += val_loader.dataset.vocab.evaluate(refs[pred_id], pred_embed)
val_bleu += (batch_bleu/len(preds))
# Get training statistics
stats = "Validation step [%d/%d], Bleu: %.4f" \
% (val_id, val_loader.dataset.get_seq_len(),
batch_bleu/len(preds))
print("\r" + stats, end="")
sys.stdout.flush()
if val_id % 250 == 0:
print('\r' + stats)
val_bleu /= val_loader.dataset.get_seq_len()
print ("\nValidation -- bleu: %.4f" % (val_bleu))
| notebooks/Image-Captioner-Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 7
#
# ## Video 31: Floating Storage
# **Python for the Energy Industry**
#
# One use of the CargoTimeSeries endpoint is to study how floating storage levels change over time. We start with our usual setup:
#
# [Cargo Time Series documentation.](https://vortechsa.github.io/python-sdk/endpoints/cargo_timeseries/)
#
# +
# initial imports
import pandas as pd
import numpy as np
from datetime import datetime
from dateutil.relativedelta import relativedelta
import vortexasdk as v
# The cargo unit for the time series (barrels)
TS_UNIT = 'b'
# The granularity of the time series
TS_FREQ = 'day'
# datetimes to access last 7 weeks of data
now = datetime.utcnow()
seven_weeks_ago = now - relativedelta(weeks=7)
# -
# Let's look at how global levels of floating storage of crude have varied over the last 7 weeks. There will be enough data to look at variation on a daily basis. We also add a requirement to only include floating storage that lasted longer than 14 days.
#
# *Note: a cargo is defined by Vortexa as becoming floating storage after it has been stationary for 7 days with no STS transfers. So a cargo that has been in floating storage for 14 days has been stationary for 21 days. 'Long-term' floating storage is FS for > 30 days.*
# +
# Find crude ID
crude = [p.id for p in v.Products().search('crude').to_list() if p.name=='Crude']
assert len(crude) == 1
search_result = v.CargoTimeSeries().search(
timeseries_frequency=TS_FREQ,
timeseries_unit=TS_UNIT,
filter_products=crude,
filter_time_min=seven_weeks_ago,
filter_time_max=now,
# Filter for cargo in floating storage
filter_activity="storing_state",
# Only get floating storage that lasted longer than 21 days
timeseries_activity_time_span_min=1000 * 60 * 60 * 24 * 14,
)
df_floating_storage = search_result.to_df()
df_floating_storage = df_floating_storage.rename(columns = {'key': 'date', 'value': 'barrels'})
# -
df_floating_storage.head()
# We can make a plot of how the total quantity in barrels varies over time:
ax = df_floating_storage.plot(x='date',y='barrels',legend=False,figsize=(8,4))
ax.set_ylabel('floating storage (barrels)')
# Let's say we now want to compare the time variation of floating storage of each category of crude. To do this, we can loop over each category and do a separate search that filters on that category only.
#
# To start, we get the IDs of each category of crude:
# +
medium_sour = [p.id for p in v.Products().search('medium-sour').to_list() if p.name=='Medium-Sour']
light_sour = [p.id for p in v.Products().search('light-sour').to_list() if p.name=='Light-Sour']
heavy_sour = [p.id for p in v.Products().search('heavy-sour').to_list() if p.name=='Heavy-Sour']
medium_sweet = [p.id for p in v.Products().search('medium-sweet').to_list() if p.name=='Medium-Sweet']
light_sweet = [p.id for p in v.Products().search('light-sweet').to_list() if p.name=='Light-Sweet']
heavy_sweet = [p.id for p in v.Products().search('heavy-sweet').to_list() if p.name=='Heavy-Sweet']
assert len(medium_sour) == 1
assert len(light_sour) == 1
assert len(heavy_sour) == 1
assert len(medium_sweet) == 1
assert len(light_sweet) == 1
assert len(heavy_sweet) == 1
# -
# To make these more convenient to work with, we make a dictionary for the ID of each category:
crude_dict = {
'medium-sour': medium_sour[0],
'light-sour': light_sour[0],
'heavy-sour': heavy_sour[0],
'medium-sweet': medium_sweet[0],
'light-sweet': light_sweet[0],
'heavy-sweet': heavy_sweet[0],
}
# We then do a loop over the categories in the dictionary. For each search, we produce a DataFrame, and then add the `values` column to our `df_floating_storage` DataFrame.
for category in crude_dict:
search_result = v.CargoTimeSeries().search(
timeseries_frequency=TS_FREQ,
timeseries_unit=TS_UNIT,
# filter only on one category of crude
filter_products=crude_dict[category],
filter_time_min=seven_weeks_ago,
filter_time_max=now,
# Filter for cargo in floating storage
filter_activity="storing_state",
# Only get floating storage that lasted longer than 21 days
timeseries_activity_time_span_min=1000 * 60 * 60 * 24 * 14,
)
df_cat = search_result.to_df()
df_floating_storage[category] = df_cat['value']
df_floating_storage.head()
# We now have the data we need to make a plot comparing each of these categories:
ax = df_floating_storage.plot(x='date',y=list(crude_dict.keys()),figsize=(8,4))
ax.set_ylabel('floating storage (barrels)')
# ### Exercise
#
# Compare and plot the floating storage of crude in different geographic regions over the last 7 weeks.
| docs/examples/academy/31. Floating Storage Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Chapter 3 – Kaggle Titantic dataset**
#
# *The task is to predict which passengers survived the Titanic shipwreck.*
#
# *In this challenge, we are asked to build a predictive model that answers the question: “what sorts of people were more likely to survive?” using passenger data (ie name, age, gender, socio-economic class, etc).*
#
# *This notebook contains all the code for tackling the titanic dataset on Kaggle*
# # Setup
# First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
# +
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "titantic"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
# -
# # Get the data
# Forget this step for now. Its seems this can work well if you download public and personal datasets, but I'm struggling to download the competition dataset. Currently it downloads them but it only downloads the zip file. Then some magic commands would need to be run to unzip them which wouldn't be too difficult
# +
import os
import kaggle
TITANIC_PATH = os.path.join("datasets", "titanic")
kaggle.api.authenticate()
def fetch_titanic_data(titanic_path=TITANIC_PATH):
if not os.path.isdir(titanic_path):
os.makedirs(titanic_path)
# +
fetch_titanic_data()
kaggle.api.authenticate()
kaggle.api.competition_download_files('titanic', path=TITANIC_PATH)
from zipfile import ZipFile
file_name = '/titanic.zip'
with ZipFile(TITANIC_PATH + file_name, 'r') as zip:
# printing all the contents of the zip file
zip.printdir()
# extracting all the files
print('Extracting all the files now...')
zip.extractall(path=TITANIC_PATH)
print('Done!')
# +
import pandas as pd
def load_titanic_data(filename, titanic_path=TITANIC_PATH):
csv_path = os.path.join(titanic_path, filename)
return pd.read_csv(csv_path)
# -
train_data = load_titanic_data(filename="train.csv")
test_data = load_titanic_data(filename="test.csv")
# The data is already split into a training set and a test set. However, the test data does not contain the labels: your goal is to train the best model you can using the training data, then make your predictions on the test data and upload them to Kaggle to see your final score.
# Let's take a peek at the top few rows of the training set:
train_data.head()
# The attributes have the following meaning:
#
# * **Survived**: that's the target, 0 means the passenger did not survive, while 1 means he/she survived.
# * **Pclass**: passenger class.
# * **Name**, **Sex**, **Age**: self-explanatory
# * **SibSp**: how many siblings & spouses of the passenger aboard the Titanic.
# * **Parch**: how many children & parents of the passenger aboard the Titanic.
# * **Ticket**: ticket id
# * **Fare**: price paid (in pounds)
# * **Cabin**: passenger's cabin number
# * **Embarked**: where the passenger embarked the Titanic
# Let's get more info to see how much data is missing:
titanic.info()
# Ok the **Age**, **Cabin** and **Embarked** attributes are sometimes null (less than 891 non-null), especially the **Cabin** (77% are null). We will ignore the **Cabin** for now and focus on the rest. The **Age** attribute has about 19% null values, so we will need to decide what to do with them. Replacing null values with the median age seems reasonable.
#
# The **Name** and **Ticket** attributes may have some value, but they will be a bit tricky to convert into useful numbers that a model can consume. So for now, we will ignore them.
#
# Let's take a look at the numerical attributes:
titanic.describe()
# * Only 38% **Survived**. :( That's close enough to 40%, so accuracy will be a reasonable metric to evaluate our model.
# * The mean **Fare** was 32.20 pounds, which does not seem so expensive (but it was probably a lot of money back then).
# * The mean **Age** was less than 30 years old.
# Let's check that the target is indeed 0 or 1:
train_data["Survived"].value_counts()
# Now let's take a quick look at all the categorical attributes:
train_data["Name"].value_counts()
train_data["Sex"].value_counts()
train_data["Ticket"].value_counts()
train_data["Cabin"].value_counts()
train_data["Embarked"].value_counts()
# The Embarked attribute tells us where the passenger embarked: C=Cherbourg, Q=Queenstown, S=Southampton.
# %matplotlib inline
import matplotlib.pyplot as plt
titanic.hist(bins=50, figsize=(20,15))
save_fig("attribute_histogram_plots")
plt.show()
# # Discover and Visualise the data to gain insights
# ## Looking for correlations
# +
titanic = train_data.copy()
corr_matrix = titanic.corr()
corr_matrix["Survived"].sort_values(ascending=False)
# -
corr_matrix.columns.to_list()
# +
from pandas.plotting import scatter_matrix
attributes = ['Survived', 'Pclass', 'Age', 'SibSp', 'Parch', 'Fare']
scatter_matrix(train_data[attributes], figsize=(12,8))
# -
# # Prepare the data for Machine Learning algorithms
# In preparation for ML we need to consider the following:
# * scaling features so ML algorithms perform better
# * dropping useless features that only create noise
# * keeping more related features can check their correlation with the target label
# * creating new features
# * numerically encodoing categorical features
#
# We will perform these data transform tasks using pipelines so they can be easily re-used for inference on the test set, and also final deployment if required.
# Although for development if may not be required. Instead could manually perform these steps. Bowen mentioned he creates a custom class for these, which he normally can just import in future projects.
# first create a copy of the data
titanic = train_data.drop("Survived", axis=1)
titanic_labels = train_data["Survived"].copy()
# +
from sklearn.base import BaseEstimator, TransformerMixin
class DataFrameSelector(BaseEstimator, TransformerMixin):
def __init__(self, attribute_names):
self.attribute_names = attribute_names
def fit(self, X, y=None):
return self
def transform(self, X):
return X[self.attribute_names]
# Inspired from stackoverflow.com/questions/25239958
class MostFrequentImputer(BaseEstimator, TransformerMixin):
def fit(self, X, y=None):
self.most_frequent_ = pd.Series([X[c].value_counts().index[0] for c in X],
index=X.columns)
return self
def transform(self, X, y=None):
return X.fillna(self.most_frequent_)
# +
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler # The chapter solutions did not use this
num_pipeline = Pipeline([
('select_numeric', DataFrameSelector(['Pclass', 'Age', 'SibSp', 'Parch', 'Fare'])),
('imputer', SimpleImputer(strategy="median"))
])
cat_pipeline = Pipeline([
('select_cat', DataFrameSelector(cat_attributes_keep)),
('imputer', SimpleImputer(strategy="most_frequent")),
('cat_encoder', OneHotEncoder())
])
# +
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
num_attributes = titanic.select_dtypes(exclude=['object']).columns.to_list()
cat_attributes = titanic.select_dtypes(include=['object']).columns.to_list()
cat_attributes_keep = ['Sex', 'Embarked']
full_pipeline = ColumnTransformer([
('num', num_pipeline, num_attributes),
('cat', cat_pipeline, cat_attributes)
])
# -
X_train = full_pipeline.fit_transform(titanic)
y_train = titanic_labels
# # Select and train a model
# Now we're ready to train a classifier, let's start with linearRegressor
# +
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X_train, y_train)
# -
# Great our model is trained, let's use it to make predictions on the test set
X_test = full_pipeline.transform(test_data)
y_pred = lin_reg.predict(X_test)
# And now we could just build a CSV file with these predictions (respecting the format excepted by Kaggle), then upload it and hope for the best. But wait! We can do better than hope. Why don't we use cross-validation to have an idea of how good our model is?
# +
from sklearn.model_selection import cross_val_score
lin_reg_scores = cross_val_score(lin_reg, X_train, y_train, cv=10)
lin_reg_scores.mean()
# +
from sklearn.svm import SVC
svm_clf = SVC(gamma="auto")
svm_clf.fit(X_train, y_train)
# +
from sklearn.model_selection import cross_val_score
svm_scores = cross_val_score(svm_clf, X_train, y_train, cv=10)
svm_scores.mean()
# +
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(n_estimators=100, random_state=42)
forest_scores = cross_val_score(forest_clf, X_train, y_train, cv=10)
forest_scores.mean()
# -
plt.figure(figsize=(8, 4))
plt.plot([1]*10, svm_scores, ".")
plt.plot([2]*10, forest_scores, ".")
plt.boxplot([svm_scores, forest_scores], labels=("SVM","Random Forest"))
plt.ylabel("Accuracy", fontsize=14)
plt.show()
| 03_kaggle_titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # miRNA-Seq
#
# (2019.04.02)
#
# ## Only valid miRNAs
#
# On the paper [Evolutionary history of plant microRNAs](https://doi.org/10.1016/j.tplants.2013.11.008), miRBase is scan for valid miRNAs. For a miRNA to be considered valid...
#
# * miRNA sequence must have high complementarity to opposing arm (>= 15 nt)
# * It should be observed an precision on 5' cleavage
# * Little heterogeneity in the sequence matching to the miRNA precursor. If not, this is a siRNA, not a miRNA.
# * The miRNA* should be present
#
# The list of valid *Vitis vinifera* miRNAs is provided in the addtional files. Here, I will scan my miRNAs list, for the valid miRNAs, and remove the non validated ones.
import pandas
# ### Importing list of valid miRNAs
#
# Note: This list does not contain only valid miRNAs. It contains the indication if the miRNA is valid or not! miRNA should pass on miRNA* or Structure to be considered valid.
# +
valid_mirnas = pandas.read_csv('valid_mirnas.tsv',
sep = '\t'
)
valid_mirnas.head()
# -
valid_mirnas.shape
valid_mirnas_pass = valid_mirnas[(valid_mirnas['miRNA*'] == '✓')
| (valid_mirnas['Structure'] == '✓')
]
valid_mirnas_pass.shape
# Only 4 miRNAs should be removed from our lists... To be honest, may be we do nor even got those.
valid_mirnas_pass_list = valid_mirnas_pass['ID'].tolist()
valid_mirnas_all_list = valid_mirnas['ID'].tolist()
# ### Importing miRNAs from miRBase
#
# #### Hairpins
# +
mirbase_hairpins_counts = pandas.read_csv('../2019_miRNA_sequencing/mirbase_hairpins_counts.tsv',
sep = '\t'
)
mirbase_hairpins_counts.head()
# -
mirbase_hairpins_counts.shape
# Below is the list of miRNAs that were not considered valid. This does not mean that the miRNAs are invalid, only that the miRNAs were not considered valid (but they may have not even be tested)
mirbase_hairpins_counts[~mirbase_hairpins_counts['miRNA'].isin(valid_mirnas_pass_list)]
# +
mirbase_hairpins_counts_non_valid = mirbase_hairpins_counts[~mirbase_hairpins_counts['miRNA'].isin(valid_mirnas_pass_list)]['miRNA'].tolist()
[mirna for mirna in mirbase_hairpins_counts_non_valid if mirna in valid_mirnas_all_list]
# -
# The miRNAs listed above, is the list of miRNAs that are present in the analysis provided on the paper and on our results, but did not pass. The other miRNAs present on the table, were just not checked along the paper.
#
# Instead of checkig the valid miRNAs, as this will not consider the miRNAs that were not processed on the paper, we should remove directly the non valid miRNAs.
valid_mirnas_nopass = valid_mirnas[(valid_mirnas['miRNA*'] == '✗')
& (valid_mirnas['Structure'] == '✗')
]
valid_mirnas_nopass.shape
valid_mirnas_nopass_list = valid_mirnas_nopass['ID'].tolist()
mirbase_hairpins_counts_pass = mirbase_hairpins_counts[~mirbase_hairpins_counts['miRNA'].isin(valid_mirnas_nopass_list)]
mirbase_hairpins_counts_pass.head()
# Those were the valid or non processed miRNAs.
# +
mirbase_hairpins_de = pandas.read_csv('../2019_miRNA_sequencing/mirbase_hairpins_diffexpression.tsv',
sep = '\t'
)
mirbase_hairpins_de.head()
# -
mirbase_hairpins_de_pass = mirbase_hairpins_de[~mirbase_hairpins_de['miRNA'].isin(valid_mirnas_nopass_list)]
mirbase_hairpins_de_pass.head()
# Next, are the no pass!
mirbase_hairpins_de[mirbase_hairpins_de['miRNA'].isin(valid_mirnas_nopass_list)].head()
# ### Saving the new lists to new files
# +
mirbase_hairpins_counts_pass.to_csv('mirbase_hairpins_counts_pass.tsv',
sep = '\t'
)
mirbase_hairpins_de_pass.to_csv('mirbase_hairpins_diffexpression_pass.tsv',
sep = '\t'
)
| jupyter_notebooks/valid_mirnas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ur8xi4C7S06n"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="JAPoU8Sm5E6e"
# <table align="left">
#
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/vertex-ai-samples/tree/master/notebooks/official/matching_engine/two-tower-model-introduction.ipynb"">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/vertex-ai-samples/tree/master/notebooks/official/matching_engine/two-tower-model-introduction.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# + [markdown] id="tvgnzT1CKxrO"
# ## Overview
#
# This tutorial demonstrates how to use the Two-Tower built-in algorithm on the Vertex AI platform.
#
# Two-tower models learn to represent two items of various types (such as user profiles, search queries, web documents, answer passages, or images) in the same vector space, so that similar or related items are close to each other. These two items are referred to as the query and candidate object, since when paired with a nearest neighbor search service such as Vertex Matching Engine, the two-tower model can retrieve candidate objects related to an input query object. These objects are encoded by a query and candidate encoder (the two "towers") respectively, which are trained on pairs of relevant items. This built-in algorithm exports trained query and candidate encoders as model artifacts, which can be deployed in Vertex Prediction for usage in a recommendation system.
#
# ### Dataset
#
# This tutorial uses the `movielens_100k sample dataset` in the public bucket `gs://cloud-samples-data/vertex-ai/matching-engine/two-tower`, which was generated from the [MovieLens movie rating dataset](https://grouplens.org/datasets/movielens/100k/). For simplicity, the data for this tutorial only includes the user id feature for users, and the movie id and movie title features for movies. In this example, the user is the query object and the movie is the candidate object, and each training example in the dataset contains a user and a movie they rated (we only include positive ratings in the dataset). The two-tower model will embed the user and the movie in the same embedding space, so that given a user, the model will recommend movies it thinks the user will like.
#
# ### Objective
#
# In this notebook, you will learn how to run the two-tower model.
# The tutorial covers the following steps:
# 1. **Setup**: Importing the required libraries and setting your global variables.
# 2. **Configure parameters**: Setting the appropriate parameter values for the training job.
# 3. **Train on Vertex Training**: Submitting a training job.
# 4. **Deploy on Vertex Prediction**: Importing and deploying the trained model to a callable endpoint.
# 5. **Predict**: Calling the deployed endpoint using online or batch prediction.
# 6. **Hyperparameter tuning**: Running a hyperparameter tuning job.
# 7. **Cleaning up**: Deleting resources created by this tutorial.
#
#
# ### Costs
#
#
# This tutorial uses billable components of Google Cloud:
#
# * Vertex AI
# * Cloud Storage
#
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="ze4-nDLfK4pw"
# ### Set up your local development environment
#
# **If you are using Colab or Google Cloud Notebooks**, your environment already meets
# all the requirements to run this notebook. You can skip this step.
# + [markdown] id="gCuSR8GkAgzl"
# **Otherwise**, make sure your environment meets this notebook's requirements.
# You need the following:
#
# * The Google Cloud SDK
# * Git
# * Python 3
# * virtualenv
# * Jupyter notebook running in a virtual environment with Python 3
#
# The Google Cloud guide to [Setting up a Python development
# environment](https://cloud.google.com/python/setup) and the [Jupyter
# installation guide](https://jupyter.org/install) provide detailed instructions
# for meeting these requirements. The following steps provide a condensed set of
# instructions:
#
# 1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
#
# 1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
#
# 1. [Install
# virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
# and create a virtual environment that uses Python 3. Activate the virtual environment.
#
# 1. To install Jupyter, run `pip3 install jupyter` on the
# command-line in a terminal shell.
#
# 1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
#
# 1. Open this notebook in the Jupyter Notebook Dashboard.
# + [markdown] id="i7EUnXsZhAGF"
# ### Install additional packages
#
# + id="2b4ef9b72d43"
import os
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# Google Cloud Notebook requires dependencies to be installed with '--user'
USER_FLAG = ""
if IS_GOOGLE_CLOUD_NOTEBOOK:
USER_FLAG = "--user"
# + id="wyy5Lbnzg5fi"
# ! pip3 install {USER_FLAG} --upgrade tensorflow
# ! pip3 install {USER_FLAG} --upgrade google-cloud-aiplatform tensorboard-plugin-profile
# ! gcloud components update --quiet
# + [markdown] id="hhq5zEbGg0XX"
# ### Restart the kernel
#
# After you install the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="EzrelQZ22IZj"
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="lWEdiXsJg0XY"
# ## Before you begin
# + [markdown] id="BF1j6f9HApxa"
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
#
# 1. [Enable the Vertex AI API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com).
#
# 1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
#
# 1. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + [markdown] id="WReHDGG5g0XY"
# #### Set your project ID
#
# **If you do not know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="oM1iC_MfAts1"
import os
PROJECT_ID = ""
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
shell_output=! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
# + [markdown] id="qJYoRfYng0XZ"
# Otherwise, set your project ID here.
# + id="riG_qUokg0XZ"
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# ! gcloud config set project {PROJECT_ID}
# + [markdown] id="06571eb4063b"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
# + id="697568e92bd6"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="dr--iN2kAylZ"
# ### Authenticate your Google Cloud account
#
# **If you are using Google Cloud Notebooks**, your environment is already
# authenticated. Skip this step.
# + [markdown] id="sBCra4QMA2wR"
# **If you are using Colab**, run the cell below and follow the instructions
# when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# 1. In the Cloud Console, go to the [**Create service account key**
# page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
#
# 2. Click **Create service account**.
#
# 3. In the **Service account name** field, enter a name, and
# click **Create**.
#
# 4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
# into the filter box, and select
# **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# 5. Click *Create*. A JSON file that contains your key downloads to your
# local environment.
#
# 6. Enter the path to your service account key as the
# `GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
# + id="PyQmSRbKA8r-"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# The Google Cloud Notebook product has specific requirements
IS_GOOGLE_CLOUD_NOTEBOOK = os.path.exists("/opt/deeplearning/metadata/env_version")
# If on Google Cloud Notebooks, then don't execute this code
if not IS_GOOGLE_CLOUD_NOTEBOOK:
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="zgPO1eR3CYjk"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# Before you submit a training job for the two-tower model, you need to upload your training data and schema to Cloud Storage. Vertex AI trains the model using this input data. In this tutorial, the Two-Tower built-in algorithm also saves the trained model that results from your job in the same bucket. Using this model artifact, you can then create Vertex AI model and endpoint resources in order to serve online predictions.
#
# Set the name of your Cloud Storage bucket below. It must be unique across all
# Cloud Storage buckets.
#
# You may also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Make sure to [choose a region where Vertex AI services are
# available](https://cloud.google.com/vertex-ai/docs/general/locations#available_regions). You may
# not use a Multi-Regional Storage bucket for training with Vertex AI.
# + id="MzGDU7TWdts_"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
REGION = "us-central1" # @param {type:"string"}
# + id="cf221059d072"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="-EcIXiGsCePi"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="NIq7R4HZCfIc"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="ucvCsknMCims"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="vhOb7YnwClBb"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="XoEqT2Y4DJmf"
# ### Import libraries and define constants
# + id="pRUOFELefqf1"
import json
import os
import re
import time
from google.cloud import aiplatform
# %load_ext tensorboard
# + [markdown] id="ixC92jeHQMxk"
# ## Configure parameters
# + [markdown] id="GgsNm8aim0Ym"
# The following table shows parameters that are common to all Vertex Training jobs created using the `gcloud ai custom-jobs create` command. See the [official documentation](https://cloud.google.com/sdk/gcloud/reference/ai/custom-jobs/create) for all the possible arguments.
#
# | Parameter | Data type | Description | Required |
# |--|--|--|--|
# | `display-name` | string | Name of the job. | Yes |
# | `worker-pool-spec` | string | Comma-separated list of arguments specifying a worker pool configuration (see below). | Yes |
# | `region` | string | Region to submit the job to. | No |
#
# The `worker-pool-spec` flag can be specified multiple times, one for each worker pool. The following table shows the arguments used to specify a worker pool.
#
# | Parameter | Data type | Description | Required |
# |--|--|--|--|
# | `machine-type` | string | Machine type for the pool. See the [official documentation](https://cloud.google.com/vertex-ai/docs/training/configure-compute) for supported machines. | Yes |
# | `replica-count` | int | The number of replicas of the machine in the pool. | No |
# | `container-image-uri` | string | Docker image to run on each worker. | No |
# + [markdown] id="0MvQ22Sbm8lh"
# The following table shows the parameters for the two-tower model training job:
#
# | Parameter | Data type | Description | Required |
# |--|--|--|--|
# | `training_data_path` | string | Cloud Storage pattern where training data is stored. | Yes |
# | `input_schema_path` | string | Cloud Storage path where the JSON input schema is stored. | Yes |
# | `input_file_format` | string | The file format of input. Currently supports `jsonl` and `tfrecord`. | No - default is `jsonl`. |
# | `job_dir` | string | Cloud Storage directory where the model output files will be stored. | Yes |
# | `eval_data_path` | string | Cloud Storage pattern where eval data is stored. | No |
# | `candidate_data_path` | string | Cloud Storage pattern where candidate data is stored. Only used for top_k_categorical_accuracy metrics. If not set, it's generated from training/eval data. | No |
# | `train_batch_size` | int | Batch size for training. | No - Default is 100. |
# | `eval_batch_size` | int | Batch size for evaluation. | No - Default is 100. |
# | `eval_split` | float | Split fraction to use for the evaluation dataset, if `eval_data_path` is not provided. | No - Default is 0.2 |
# | `optimizer` | string | Training optimizer. Lowercase string name of any TF2.3 Keras optimizer is supported ('sgd', 'nadam', 'ftrl', etc.). See [TensorFlow documentation](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers). | No - Default is 'adagrad'. |
# | `learning_rate` | float | Learning rate for training. | No - Default is the default learning rate of the specified optimizer. |
# | `momentum` | float | Momentum for optimizer, if specified. | No - Default is the default momentum value for the specified optimizer. |
# | `metrics` | string | Metrics used to evaluate the model. Can be either `auc`, `top_k_categorical_accuracy` or `precision_at_1`. | No - Default is `auc`. |
# | `num_epochs` | int | Number of epochs for training. | No - Default is 10. |
# | `num_hidden_layers` | int | Number of hidden layers. | No |
# | `num_nodes_hidden_layer{index}` | int | Num of nodes in hidden layer {index}. The range of index is 1 to 20. | No |
# | `output_dim` | int | The output embedding dimension for each encoder tower of the two-tower model. | No - Default is 64. |
# | `training_steps_per_epoch` | int | Number of steps per epoch to run the training for. Only needed if you are using more than 1 machine or using a master machine with more than 1 gpu. | No - Default is None. |
# | `eval_steps_per_epoch` | int | Number of steps per epoch to run the evaluation for. Only needed if you are using more than 1 machine or using a master machine with more than 1 gpu. | No - Default is None. |
# | `gpu_memory_alloc` | int | Amount of memory allocated per GPU (in MB). | No - Default is no limit. |
# + id="2sEfn2ZVnI_s"
DATASET_NAME = "movielens_100k" # Change to your dataset name.
# Change to your data and schema paths. These are paths to the movielens_100k
# sample data.
TRAINING_DATA_PATH = f"gs://cloud-samples-data/vertex-ai/matching-engine/two-tower/{DATASET_NAME}/training_data/*"
INPUT_SCHEMA_PATH = f"gs://cloud-samples-data/vertex-ai/matching-engine/two-tower/{DATASET_NAME}/input_schema.json"
# URI of the two-tower training Docker image.
LEARNER_IMAGE_URI = "us-docker.pkg.dev/vertex-ai-restricted/builtin-algorithm/two-tower"
# Change to your output location.
OUTPUT_DIR = f"{BUCKET_NAME}/experiment/output"
TRAIN_BATCH_SIZE = 100 # Batch size for training.
NUM_EPOCHS = 3 # Number of epochs for training.
print(f"Dataset name: {DATASET_NAME}")
print(f"Training data path: {TRAINING_DATA_PATH}")
print(f"Input schema path: {INPUT_SCHEMA_PATH}")
print(f"Output directory: {OUTPUT_DIR}")
print(f"Train batch size: {TRAIN_BATCH_SIZE}")
print(f"Number of epochs: {NUM_EPOCHS}")
# + [markdown] id="upLZ8kcankwj"
# ## Train on Vertex Training
# + [markdown] id="6M_O_L55nwQ0"
# Submit the two-tower training job to Vertex Training. The following command uses a single CPU machine for training. When using single node training, `training_steps_per_epoch` and `eval_steps_per_epoch` do not need to be set.
# + id="1gXZRq80nl2S"
learning_job_name = f"two_tower_cpu_{DATASET_NAME}_{TIMESTAMP}"
CREATION_LOG = ! gcloud ai custom-jobs create \
--display-name={learning_job_name} \
--worker-pool-spec=machine-type=n1-standard-8,replica-count=1,container-image-uri={LEARNER_IMAGE_URI} \
--region={REGION} \
--args=--training_data_path={TRAINING_DATA_PATH} \
--args=--input_schema_path={INPUT_SCHEMA_PATH} \
--args=--job-dir={OUTPUT_DIR} \
--args=--train_batch_size={TRAIN_BATCH_SIZE} \
--args=--num_epochs={NUM_EPOCHS}
print(CREATION_LOG)
# + [markdown] id="RaIkcFT2n4_U"
# If you want to train using GPUs, you need to write configuration to a YAML file:
# + id="nAod1hbSn5yw"
learning_job_name = f"two_tower_gpu_{DATASET_NAME}_{TIMESTAMP}"
config = f"""workerPoolSpecs:
-
machineSpec:
machineType: n1-highmem-4
acceleratorType: NVIDIA_TESLA_K80
acceleratorCount: 1
replicaCount: 1
containerSpec:
imageUri: {LEARNER_IMAGE_URI}
args:
- --training_data_path={TRAINING_DATA_PATH}
- --input_schema_path={INPUT_SCHEMA_PATH}
- --job-dir={OUTPUT_DIR}
- --training_steps_per_epoch=1500
- --eval_steps_per_epoch=1500
"""
# !echo $'{config}' > ./config.yaml
CREATION_LOG = ! gcloud ai custom-jobs create \
--display-name={learning_job_name} \
--region={REGION} \
--config=config.yaml
print(CREATION_LOG)
# + [markdown] id="94tmU59YrKfe"
# If you want to use TFRecord input file format, you can try the following command:
# + id="8wZbRgUhrLD0"
TRAINING_DATA_PATH = f"gs://cloud-samples-data/vertex-ai/matching-engine/two-tower/{DATASET_NAME}/tfrecord/*"
learning_job_name = f"two_tower_cpu_tfrecord_{DATASET_NAME}_{TIMESTAMP}"
CREATION_LOG = ! gcloud ai custom-jobs create \
--display-name={learning_job_name} \
--worker-pool-spec=machine-type=n1-standard-8,replica-count=1,container-image-uri={LEARNER_IMAGE_URI} \
--region={REGION} \
--args=--training_data_path={TRAINING_DATA_PATH} \
--args=--input_schema_path={INPUT_SCHEMA_PATH} \
--args=--job-dir={OUTPUT_DIR} \
--args=--train_batch_size={TRAIN_BATCH_SIZE} \
--args=--num_epochs={NUM_EPOCHS} \
--args=--input_file_format=tfrecord
print(CREATION_LOG)
# + [markdown] id="yceUSlyWrWes"
# After the job is submitted successfully, you can view its details and logs:
# + id="XXDC7F_8rXWM"
JOB_ID = re.search(r"(?<=/customJobs/)\d+", CREATION_LOG[1]).group(0)
print(JOB_ID)
# + id="NkbwWCEMoQcy"
# View the job's configuration and state.
STATE = "state: JOB_STATE_PENDING"
while STATE not in ["state: JOB_STATE_SUCCEEDED", "state: JOB_STATE_FAILED"]:
DESCRIPTION = ! gcloud ai custom-jobs describe {JOB_ID} --region={REGION}
STATE = DESCRIPTION[-2]
print(STATE)
time.sleep(60)
# + [markdown] id="wgs0qV_Nr-RN"
# When the training starts, you can view the logs in TensorBoard. Colab users can use the TensorBoard widget below:
# + id="8SweSrkhr_DP"
TENSORBOARD_DIR = os.path.join(OUTPUT_DIR, "tensorboard")
# %tensorboard --logdir {TENSORBOARD_DIR}
# + [markdown] id="RzCrdxgAsGll"
# For Google CLoud Notebooks users, the TensorBoard widget above won't work. We recommend you to launch TensorBoard through the Cloud Shell.
#
# 1. In your Cloud Shell, launch Tensorboard on port 8080:
#
# ```
# export TENSORBOARD_DIR=gs://xxxxx/tensorboard
# tensorboard --logdir=${TENSORBOARD_DIR} --port=8080
# ```
#
# 2. Click the "Web Preview" button at the top-right of the Cloud Shell window (looks like an eye in a rectangle).
#
# 3. Select "Preview on port 8080". This should launch the TensorBoard webpage in a new tab in your browser.
#
# After the job finishes successfully, you can view the output directory:
# + id="mFPLfY4gsK1V"
# ! gsutil ls {OUTPUT_DIR}
# + [markdown] id="ZhY0h8ijsPlP"
# ## Deploy on Vertex Prediction
# + [markdown] id="6oquDjRgsS2V"
# ### Import the model
# + [markdown] id="gidmXBWysaeP"
# Our training job will export two TF SavedModels under `gs://<job_dir>/query_model` and `gs://<job_dir>/candidate_model`. These exported models can be used for online or batch prediction in Vertex Prediction. First, import the query (or candidate) model:
# + id="yEFd3Og_sbdm"
# The following imports the query (user) encoder model.
MODEL_TYPE = "query"
# Use the following instead to import the candidate (movie) encoder model.
# MODEL_TYPE = 'candidate'
DISPLAY_NAME = f"{DATASET_NAME}_{MODEL_TYPE}" # The display name of the model.
MODEL_NAME = f"{MODEL_TYPE}_model" # Used by the deployment container.
# + id="GCrhxf7GsdZS"
aiplatform.init(
project=PROJECT_ID,
location=REGION,
staging_bucket=BUCKET_NAME,
)
model = aiplatform.Model.upload(
display_name=DISPLAY_NAME,
artifact_uri=OUTPUT_DIR,
serving_container_image_uri="us-central1-docker.pkg.dev/cloud-ml-algos/two-tower/deploy",
serving_container_health_route=f"/v1/models/{MODEL_NAME}",
serving_container_predict_route=f"/v1/models/{MODEL_NAME}:predict",
serving_container_environment_variables={
"MODEL_BASE_PATH": "$(AIP_STORAGE_URI)",
"MODEL_NAME": MODEL_NAME,
},
)
# + [markdown] id="0x-pZJzUsh22"
# ### Deploy the model
# + [markdown] id="QJECrcTdskix"
# After importing the model, you must deploy it to an endpoint so that you can get online predictions. More information about this process can be found in the [official documentation](https://cloud.google.com/vertex-ai/docs/predictions/deploy-model-api).
# + id="jB3yT5xassCt"
# ! gcloud ai models list --region={REGION} --filter={DISPLAY_NAME}
# + [markdown] id="zkJh2rmysu2M"
# Create a model endpoint:
# + id="er94Wp82sxYW"
endpoint = aiplatform.Endpoint.create(display_name=DATASET_NAME)
# + [markdown] id="PICvm8PhqtMw"
# Deploy model to the endpoint
# + id="OUEF7Yces4uD"
model.deploy(
endpoint=endpoint,
machine_type="n1-standard-4",
traffic_split={"0": 100},
deployed_model_display_name=DISPLAY_NAME,
)
# + [markdown] id="Y_LMW1rjtMM6"
# ## Predict
# + [markdown] id="QAkHJlY7tOmu"
# Now that you have deployed the query/candidate encoder model on Vertex Prediction, you can call the model to calculate embeddings for live data. There are two methods of getting predictions, online and batch, which are shown below.
# + [markdown] id="rwiJRfJRtQ1V"
# ### Online prediction
# + [markdown] id="THz5Gn5ftTsm"
# [Online prediction](https://cloud.google.com/vertex-ai/docs/predictions/online-predictions-custom-models) is used to synchronously query a model on a small batch of instances with minimal latency. The following function calls the deployed Vertex Prediction model endpoint using Vertex SDK for Python:
# + [markdown] id="bFVnzRzltfDa"
# The input data you want predictions on should be provided as a stringified JSON in the `data` field. Note that you should also provide a unique `key` field (of type str) for each input instance so that you can associate each output embedding with its corresponding input.
# + id="E8Wt7wYgtg_f"
# Input items for the query model:
input_items = [
{"data": '{"user_id": ["1"]}', "key": "key1"},
{"data": '{"user_id": ["2"]}', "key": "key2"},
]
# Input items for the candidate model:
# input_items = [{
# 'data' : '{"movie_id": ["1"], "movie_title": ["fake title"]}',
# 'key': 'key1'
# }]
encodings = endpoint.predict(input_items)
print(f"Number of encodings: {len(encodings.predictions)}")
print(encodings.predictions[0]["encoding"])
# + [markdown] id="1k-_XJzlthfP"
# You can also do online prediction using the gcloud CLI, as shown below:
# + id="Tn5L9V0utkpA"
request = json.dumps({"instances": input_items})
with open("request.json", "w") as writer:
writer.write(f"{request}\n")
ENDPOINT_ID = endpoint.resource_name
# ! gcloud ai endpoints predict {ENDPOINT_ID} \
# --region={REGION} \
# --json-request=request.json
# + [markdown] id="ocLE_U6ftnA3"
# ### Batch prediction
# + [markdown] id="U__JdxfPto-_"
# [Batch prediction](https://cloud.google.com/vertex-ai/docs/predictions/batch-predictions) is used to asynchronously make predictions on a batch of input data. This is recommended if you have a large input size and do not need an immediate response, such as getting embeddings for candidate objects in order to create an index for a nearest neighbor search service such as [Vertex Matching Engine](https://cloud.google.com/vertex-ai/docs/matching-engine/overview).
#
# The input data needs to be on Cloud Storage and in JSONL format. You can use the sample query object file provided below. Like with online prediction, it's recommended to have the `key` field so that you can associate each output embedding with its corresponding input.
# + id="ar13RZ4VtquX"
QUERY_SAMPLE_PATH = f"gs://cloud-samples-data/vertex-ai/matching-engine/two-tower/{DATASET_NAME}/query_sample.jsonl"
# ! gsutil cat {QUERY_SAMPLE_PATH}
# + [markdown] id="Z4f5yEmatu8P"
# The following function calls the deployed Vertex Prediction model using the sample query object input file. Note that it uses the model resource directly and doesn't require a deployed endpoint. Once you start the job, you can track its status on the [Cloud Console](https://console.cloud.google.com/vertex-ai/batch-predictions).
# + id="EYEOOPS8txYY"
model.batch_predict(
job_display_name=f"batch_predict_{DISPLAY_NAME}",
gcs_source=[QUERY_SAMPLE_PATH],
gcs_destination_prefix=OUTPUT_DIR,
machine_type="n1-standard-4",
starting_replica_count=1,
)
# + [markdown] id="zImnlP2Yt6Sv"
# ## Hyperparameter tuning
# + [markdown] id="I2nRxtLTt8xn"
# After successfully training your model, deploying it, and calling it to make predictions, you may want to optimize the hyperparameters used during training to improve your model's accuracy and performance. See the Vertex AI documentation for an [overview of hyperparameter tuning](https://cloud.google.com/vertex-ai/docs/training/hyperparameter-tuning-overview) and [how to use it in your Vertex Training jobs](https://cloud.google.com/vertex-ai/docs/training/using-hyperparameter-tuning).
#
# For this example, the following command runs a Vertex AI hyperparameter tuning job with 8 trials that attempts to maximize the validation AUC metric. The hyperparameters it optimizes are the number of hidden layers, the size of the hidden layers, and the learning rate.
# + id="z_ea5wfjt_XD"
PARALLEL_TRIAL_COUNT = 4
MAX_TRIAL_COUNT = 8
METRIC = "val_auc"
hyper_tune_job_name = f"hyper_tune_{DATASET_NAME}_{TIMESTAMP}"
config = json.dumps(
{
"displayName": hyper_tune_job_name,
"studySpec": {
"metrics": [{"metricId": METRIC, "goal": "MAXIMIZE"}],
"parameters": [
{
"parameterId": "num_hidden_layers",
"scaleType": "UNIT_LINEAR_SCALE",
"integerValueSpec": {"minValue": 0, "maxValue": 2},
"conditionalParameterSpecs": [
{
"parameterSpec": {
"parameterId": "num_nodes_hidden_layer1",
"scaleType": "UNIT_LOG_SCALE",
"integerValueSpec": {"minValue": 1, "maxValue": 128},
},
"parentIntValues": {"values": [1, 2]},
},
{
"parameterSpec": {
"parameterId": "num_nodes_hidden_layer2",
"scaleType": "UNIT_LOG_SCALE",
"integerValueSpec": {"minValue": 1, "maxValue": 128},
},
"parentIntValues": {"values": [2]},
},
],
},
{
"parameterId": "learning_rate",
"scaleType": "UNIT_LOG_SCALE",
"doubleValueSpec": {"minValue": 0.0001, "maxValue": 1.0},
},
],
"algorithm": "ALGORITHM_UNSPECIFIED",
},
"maxTrialCount": MAX_TRIAL_COUNT,
"parallelTrialCount": PARALLEL_TRIAL_COUNT,
"maxFailedTrialCount": 3,
"trialJobSpec": {
"workerPoolSpecs": [
{
"machineSpec": {
"machineType": "n1-standard-4",
},
"replicaCount": 1,
"containerSpec": {
"imageUri": LEARNER_IMAGE_URI,
"args": [
f"--training_data_path={TRAINING_DATA_PATH}",
f"--input_schema_path={INPUT_SCHEMA_PATH}",
f"--job-dir={OUTPUT_DIR}",
],
},
}
]
},
}
)
# ! curl -X POST -H "Authorization: Bearer "$(gcloud auth print-access-token) \
# -H "Content-Type: application/json; charset=utf-8" \
# -d '{config}' https://us-central1-aiplatform.googleapis.com/v1/projects/{PROJECT_ID}/locations/{REGION}/hyperparameterTuningJobs
# + [markdown] id="TpV-iwP9qw9c"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
# + id="sx_vKniMq9ZX"
# Delete endpoint resource
endpoint.delete(force=True)
# Delete model resource
model.delete()
# Delete Cloud Storage objects that were created
# ! gsutil -m rm -r $OUTPUT_DIR
| notebooks/official/matching_engine/two-tower-model-introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Examine tidal constituents - M2/K1 elevation
#
# M2/K1 tidal currents are also examined at the VENUS nodes.
#
# Inference for P1 and K2 is not applied
#
# Comparing background visc = 1e-4, 1e-5, and 1e-6.
#
# Simulations had no weather, isoneutral lateral mixing.
# +
# imports
# %matplotlib inline
import matplotlib.pylab as plt
import numpy as np
import netCDF4 as nc
from salishsea_tools import tidetools
from salishsea_tools import viz_tools
from salishsea_tools import bathy_tools
import pandas as pd
import datetime
from dateutil import tz
import os
import angles
from salishsea_tools import ellipse
from __future__ import division
# -
# # Run Details
# +
# pathname for data
path = '/data/nsoontie/MEOPAR/SalishSea/results/tides/'
#the runs we want to analyze
runs = ['visc1e-4', 'visc1e-5', 'visc1e-6']
# grid
grid = nc.Dataset('/ocean/imachuca/MEOPAR/NEMO-forcing/grid/bathy_meter_SalishSea2.nc')
bathy, X, Y = tidetools.get_bathy_data(grid)
# -
# # Observations
# +
filename = '/data/nsoontie/MEOPAR/analysis/compare_tides/obs_tidal_wlev_const_all.csv'
harm_obs = pd.read_csv(filename,sep=';',header=0)
harm_obs = harm_obs.rename(columns={'Site': 'site', 'Lat': 'lat', 'Lon': 'lon',
'M2 amp': 'M2_amp', 'M2 phase (deg UT)': 'M2_pha',
'K1 amp': 'K1_amp', 'K1 phase (deg UT)': 'K1_pha'})
# -
# This is a list of observations that we can compare with our model output. Now we have a struc object called harm_obs that contains the data printed above.
# +
filename = '/data/nsoontie/MEOPAR/analysis/Idalia/other_constituents.csv'
harm_other = pd.read_csv(filename,sep=',',header=0)
harm_other = harm_other.rename(columns={'Site': 'site', 'Lat': 'lat', 'Lon': 'lon',
'O1 amp': 'O1_amp', 'O1 phase (deg UT)': 'O1_pha',
'P1 amp': 'P1_amp', 'P1 phase (deg UT)': 'P1_pha',
'Q1 amp': 'Q1_amp', 'Q1 phase (deg UT)': 'Q1_pha',
'S2 amp': 'S2_amp', 'S2 phase (deg UT)': 'S2_pha',
'N2 amp': 'N2_amp', 'N2 phase (deg UT)': 'N2_pha',
'K2 amp': 'K2_amp', 'K2 phase (deg UT)': 'K2_pha'})
# -
# # Model
# We don't have model output at all of the above locations. The model outputs are listed below. There is a location.nc file in the run directory for each of the stations listed below.
#
# For some reason, the iodef file I used had fewer stations than our previous analaysis...
stations_obs = ['Port Renfrew','Sheringham Point', 'Sooke','Pedder Bay','Esquimalt',
'Victoria','Clover Point','Finnerty Cove','Fulford Harbour',
'Tumbo Channel','Patos Island','Whaler Bay', 'Tsawwassen',
'Sandheads', 'Point Grey','Point Atkinson','Gibsons Landing',
'Halfmoon Bay','Irvines Landing','Powell River', 'Lund',
'Twin Islets','Campbell River','Maude Island E', 'Nymphe Cove',
'Seymour Narrows','Brown Bay','Chatham Point','Kelsey Bay','Yorke Island']
# +
fig,ax=plt.subplots(1, 1, figsize=(8, 10))
ax.pcolormesh(X,Y,bathy,cmap='winter_r')
numsta=len(stations_obs)
for stn in range(numsta):
location = stations_obs[stn]
lon=-harm_obs.lon[harm_obs.site==location]
lat=harm_obs.lat[harm_obs.site==location]
ax.plot(lon,lat,'.k',label=location)
ax.annotate(stn, xy = (lon,lat), xytext = (5,5),ha = 'right', va = 'bottom',
textcoords = 'offset points')
print stn, location
ax.axis([-126.1,-122,47,51])
# -
# # Tidal Elevation Harmonics
# We need a way of determing the amplitude and phase of the main constituents. We will do this using some functions in tidetools.
#
# First, define the nodal corrections used by nemo.
#from ocean.output
# Simulation reftime October 26, 2002
NodalCorr = {
'K1': {
'ft': 1.050578,
'uvt': 296.314842},
'M2': {
'ft': 0.987843,
'uvt': 245.888564},
'O1': {
'ft': 1.081364,
'uvt': 312.950020},
'S2': {
'ft': 1.000000,
'uvt': 0.000000},
'P1': {
'ft': 1.000000,
'uvt': 55.794860},
'N2': {
'ft': 0.987843,
'uvt': 353.570277},
'Q1': {
'ft': 1.081364,
'uvt': 60.631733},
'K2': {
'ft': 1.114095,
'uvt': 52.129248},
'reftime': datetime.datetime(2002, 10, 26, tzinfo=tz.tzutc())
}
nconst=8
# Do the harmonic analysis using Muriel's functions
# +
elev ={}
for run in runs:
elev[run] = {}
for stn in stations_obs:
stn_mod = stn.replace(" ", "")
f = nc.Dataset(os.path.join(path, run,'{}.nc'.format(stn_mod)))
ssh = f.variables['sossheig'][:]
time = f.variables['time_counter'][:]/3600.
tide_fit = tidetools.fittit(ssh,time,nconst)
#apply nodal corrections
for const in tide_fit:
tide_fit[const]['phase'] = tide_fit[const]['phase'] +NodalCorr[const]['uvt']
tide_fit[const]['amp'] = tide_fit[const]['amp'] / NodalCorr[const]['ft']
tide_fit[const]['amp'], tide_fit[const]['phase'] = tidetools.convention_pha_amp(
tide_fit[const]['amp'], tide_fit[const]['phase'])
elev[run][stn] = tide_fit
# -
# Function for calculating complex difference.
def complex_diff(Ao,go,Am,gm):
#calculates complex differences between observations and model
#Ao, go - amplitude and phase from observations
#Am, gm - amplitude and phase from model
D = np.sqrt((Ao*np.cos(np.pi*go/180)-Am*np.cos(np.pi*gm/180))**2 +
(Ao*np.sin(np.pi*go/180)-Am*np.sin(np.pi*gm/180))**2)
return D
# Compare between simulations
def compare_elevation(const, runs):
"""compares the amplitude, phase and complex differences of const
for several simulations listed in runs"""
fig,axs=plt.subplots(1,3,figsize=(15,3))
for run in runs:
model_amps =[];
model_phases =[];
cdiffs = []
for n, stn in enumerate(stations_obs):
#Observations
try:
obs_amp = harm_obs[harm_obs.site==stn]['{}_amp'.format(const)].values[0]/100
obs_phase = harm_obs[harm_obs.site==stn]['{}_pha'.format(const)].values[0]
except KeyError:
try:
obs_amp = harm_other[harm_other.site==stn]['{}_amp'.format(const)].values[0]/100
obs_phase = harm_other[harm_other.site==stn]['{}_pha'.format(const)].values[0]
except IndexError:
obs_amp = float('NaN')
obs_phase= float('NaN')
#Model
model_amp = elev[run][stn][const]['amp'][0][0]
model_phase = angles.normalize(elev[run][stn][const]['phase'][0][0])
model_amps.append(model_amp/obs_amp)
model_phases.append(model_phase-obs_phase)
#Calculate complex difference
cdiffs.append(complex_diff(obs_amp, obs_phase, model_amp, model_phase))
#plotting
ax=axs[0]
ax.plot(np.arange(numsta), model_amps,'-o',label=run)
ax.set_ylabel('Amplitude Ratio (model/obs)')
ax.set_title('{} amplitude'.format(const))
ax=axs[1]
ax.plot(np.arange(numsta), model_phases,'-o', label=run)
ax.set_ylabel('Phase difference (model-obs, deg)')
ax.set_title('{} phase'.format(const))
ax=axs[2]
ax.plot(np.arange(numsta), cdiffs,'-o', label=run)
ax.set_ylabel('Complex difference [m]')
ax.set_title('{} complex differences'.format(const))
print '{} mean complex diff: {} Exluding North {}'.format(run, np.nanmean(cdiffs), np.nanmean(cdiffs[0:25]))
#Exluding North means does not include anything Seymour Narrows and north of that
for ax in axs:
ax.grid()
ax.legend(loc=0)
ax.set_xlabel('Station')
return fig
# ### M2
fig = compare_elevation('M2', runs)
# ###K1
# Almost no change in M2/K1 elevation harmonics
fig = compare_elevation('K1', runs)
# ###S2
fig = compare_elevation('S2', runs)
# ###O1
fig = compare_elevation('O1', runs)
# ##Summary
#
# * The background vertical viscosity has almost no effect on the tidal elevation.
# * In terms of complex differences, there is a slight improvement in the semi-diurnals and a slight degradation of the diurnals. But the changes in the mean complex difference over all stations is typically less than one mm.
# # Tidal Currents
# Do the harmonic analysis using Muriel's functions
# +
curr = {}
nodes = ['east', 'central', 'delta']
for run in runs:
curr[run] = {}
for node in nodes:
f = nc.Dataset(os.path.join(path,run,'VENUS_{}_gridded.nc'.format(node)))
u = f.variables['vozocrtx'][:]
v = f.variables['vomecrty'][:]
dep = f.variables['depthu'][:]
t = f.variables['time_counter'][:]/3600.
u_rot,v_rot = ellipse.prepare_vel(u, v)
params = ellipse.get_params(u_rot, v_rot, t, nconst, tidecorr = NodalCorr)
curr[run][node] = params
# -
# ##Comparisons
# ### Observations
#
# taken from /home/nsoontie/Documents/Muriel/Mark_Currents_Nodes.png
codar ={'east' : {
'M2':
{
'Semi-Major Axis': 0.138,
'Semi-Minor Axis': -0.037,
'Inclination': 134,
'Phase': 282},
'K1':
{
'Semi-Major Axis': 0.077,
'Semi-Minor Axis': -0.009,
'Inclination': 129,
'Phase': 187}
},
'central' : {
'M2': {
'Semi-Major Axis': 0.113,
'Semi-Minor Axis': 0.008,
'Inclination': 126,
'Phase': 273},
'K1':
{
'Semi-Major Axis': 0.074,
'Semi-Minor Axis': 0.006,
'Inclination': 134,
'Phase': 131}
}
}
def compare_currents(const, runs, eparams, units, xlims,nodes):
"""compares tidal currents of a constituent near the surface for simulations in runs
The ellipse parameters in eparams are compared
Comparisons carried out at locations listed in nodes
xlims are the limits of the x axis associated with eparams
units are the units associated with eparams"""
fig,axs = plt.subplots(len(nodes),len(eparams),figsize=(5*len(eparams), 4.5*len(nodes)))
row=0
for eparam, xlim, unit, in zip(eparams, xlims, units):
for node, ax in zip(nodes, axs[:,row]):
for run in runs:
ax.plot(curr[run][node][const][eparam][:,0,0], dep, label=run)
ax.plot(codar[node][const][eparam],0,'o',label='observations')
ax.legend(loc=0)
ax.set_title(node)
ax.grid()
ax.set_ylim([50,0])
ax.set_xlim(xlim)
ax.set_xlabel('{} [{}]'.format(eparam,unit))
ax.set_ylabel('Depth [m]')
row=row+1
return fig
# ###M2
const='M2'
eparams = ['Semi-Major Axis', 'Semi-Minor Axis', 'Inclination', 'Phase']
units = ['m/s', 'm/s', 'deg ccw of East', 'deg GMT']
xlims = [[0,.5], [-.3,.3], [80,180] , [250, 360]]
nodes= ['central', 'east']
fig = compare_currents(const, runs, eparams, units, xlims, nodes)
# **Note:** The observations were analyzed over a much longer time period and a different time of year.
# * Surface layer is very much affected by the background viscsoity. At depth, the profiles are almost identical.
# * Central
# * surface major and minor axis decreased with lower viscosity (closer to obs)
# * East
# * surface major and minor axis increased with lower viscsoity (farther from obs)
# * large vertical gradients above 10m in both the major and minor axis
#
#
#
# Why are 1e-5 and 1e-6 so similar but 1e-4 is different. Look into Stephanie's suggestions... maybe 1e-4 is just too high and the gls never kicks in here... I don't know if I saved these values in this sim
#
# Could also look into decreasing the background diffusivity, depending on the value of the diff in the surface layer.
# ##K1
# +
const='K1'
eparams = ['Semi-Major Axis', 'Semi-Minor Axis', 'Inclination', 'Phase']
units = ['m/s', 'm/s', 'deg ccw of East', 'deg GMT']
xlims = [[0,.2], [-.1,.1], [80,180] , [130, 270]]
fig = compare_currents(const, runs, eparams, units, xlims, nodes)
# -
# * Surface layer looks noisier than M2, especially with 1e-6.
# * Note: Axis limits are smaller than the the M2 plots
# * Central
# * decrease in semi-major, increase in semi-minor
# * large changes in the inclination and phase (1e-6). Should the inclination really be that much different from the inclination at depth? Observations have 112 (depth averaged) and 134 (surface). But remember there can be errors with the compass of the node...
# * Observed phase is 188 (depth averaged) and 131 (CODAR)
# * East
# * Semi-minor axis increase but is still very small.
# #Summary
#
# * It is likely that our background vertical viscosity is too high. Next, I will look at our values in these regions for the 1e-4 and 1e-6 cases (and maybe 1e-5 if I have one). I think I have some deep water renewal cases that I could use.
# * Modifying the backgound vertical viscosity has a big effect on the tidal currents near the surface (upper 20m). There is not much impact at depth.
# * Decreasing the background viscosity does not, necessarily, bring us closer to the observations.
# * The M2 major/minor at central decreased, which is closer to obs
# * The M2 major/minor at east increased, which is further from obs
# BUT keep in mind that the observations were analyzed over a different length of time (10-11 vs one month in the model) so we should be careful with this comparison. Our tidal run is in April and uses climatological river so the fresh water influence is stronger than the observations which were anlayzed over 10-11 months (according to an email from Mark but it might be worthwhile to double check). Regardless, the trend is that smaller background viscosity = larger M2 surface (0.5m) currents at east and smaller M2 surface currents at central. What does this mean for across strait velcoities?
# * Question - The observed drifter tracks did not appear to be tidal. So should we also be looking at the 'mean' current in our analysis and its sensitivity to the vertical viscosity?
| Nancy/tides/Tides and background vertical viscosity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from chimera import runCommand as rc # use 'rc' as shorthand for runCommand
from chimera import replyobj # for emitting status messages
# change to folder with data files
os.chdir("/Users/pett/data")
# gather the names of .pdb files in the folder
file_names = [fn for fn in os.listdir(".") if fn.endswith(".pdb")]
# loop through the files, opening, processing, and closing each in turn
for fn in file_names:
replyobj.status("Processing " + fn) # show what file we're working on
rc("open " + fn)
rc("align ligand ~ligand") # put ligand in front of remainder of molecule
rc("focus ligand") # center/zoom ligand
rc("surf") # surface receptor
rc("preset apply publication 1") # make everything look nice
rc("surftransp 15") # make the surface a little bit see-through
# save image to a file that ends in .png rather than .pdb
png_name = fn[:-3] + "png"
rc("copy file " + png_name + " supersample 3")
rc("close all")
# uncommenting the line below will cause Chimera to exit when the script is done
#rc("stop now")
# note that indentation is significant in Python; the fact that
# the above command is exdented means that it is executed after
# the loop completes, whereas the indented commands that
# preceded it are executed as part of the loop.
# -
| ChimeraAttempt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Processing With Neural Networks
# > Convolutional neural networks use the data that is represented in images to learn. In this chapter, we will probe data in images, and we will learn how to use Keras to train a neural network to classify objects that appear in images. This is the Summary of lecture "Image Processing with Keras in Python", via datacamp.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Datacamp, Tensorflow-Keras, Vision, Deep_Learning]
# - image: images/fashion_mnist.png
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
# ## Introducing convolutional neural networks
# ### Images as data: visualizations
# To display image data, you will rely on Python's [Matplotlib](https://matplotlib.org/) library, and specifically use matplotlib's `pyplot` sub-module, that contains many plotting commands. Some of these commands allow you to display the content of images stored in arrays.
# +
from skimage import transform
# Load the image
data = plt.imread('./dataset/bricks.png')
data = transform.resize(data, (159, 240))
# Display the image
plt.imshow(data);
# -
# ### Images as data: changing images
# To modify an image, you can modify the existing numbers in the array. In a color image, you can change the values in one of the color channels without affecting the other colors, by indexing on the last dimension of the array.
# +
# Set the red channel in this part of the image to 1
data[:10, :10, 0] = 1
# Set the green channel in this part of the image to 0
data[:10, :10, 1] = 0
# Set the blue channel in this part of the image to 0
data[:10, :10, 2] = 0
# Visualize the result
plt.imshow(data);
# -
# ## Classifying images
#
# ### Using one-hot encoding to represent images
# Neural networks expect the labels of classes in a dataset to be organized in a one-hot encoded manner: each row in the array contains zeros in all columns, except the column corresponding to a unique label, which is set to 1.
#
# The fashion dataset contains three categories:
#
# - Shirts
# - Dresses
# - Shoes
#
# In this exercise, you will create a one-hot encoding of a small sample of these labels.
#
#
#
# +
labels = ['shoe', 'shirt', 'shoe', 'shirt', 'dress', 'dress', 'dress']
# The number of image categories
n_categories = 3
# The unique values of categories in the data
categories = np.array(['shirt', 'dress', 'shoe'])
# Initalize ohe_labels as all zeros
ohe_labels = np.zeros((len(labels), n_categories))
# Loop over the labels
for ii in range(len(labels)):
# Find the location of this label in the categories variables
jj = np.where(categories == labels[ii])
# Set the corresponding zero to one
ohe_labels[ii, jj] = 1
# -
# ### Evaluating a classifier
# To evaluate a classifier, we need to test it on images that were not used during training. This is called "cross-validation": a prediction of the class (e.g., t-shirt, dress or shoe) is made from each of the test images, and these predictions are compared with the true labels of these images.
#
#
# +
test_labels = np.array([[0., 0., 1.], [0., 1., 0.], [0., 0., 1.], [0., 1., 0.],
[0., 0., 1.], [0., 0., 1.], [0., 0., 1.], [0., 1., 0.]])
predictions = np.array([[0., 0., 1.], [0., 1., 0.], [0., 0., 1.], [1., 0., 0.],
[0., 0., 1.], [1., 0., 0.], [0., 0., 1.], [0., 1., 0.]])
# +
# Calculate the number of correct predictions
number_correct = (test_labels * predictions).sum()
print(number_correct)
# Calculate the proportion of correct predictions
proportion_correct = number_correct / predictions.shape[0]
print(proportion_correct)
# -
# ## Classification with Keras
#
# ### Build a neural network
# We will use the Keras library to create neural networks and to train these neural networks to classify images. These models will all be of the `Sequential` type, meaning that the outputs of one layer are provided as inputs only to the next layer.
#
# In this exercise, you will create a neural network with `Dense` layers, meaning that each unit in each layer is connected to all of the units in the previous layer. For example, each unit in the first layer is connected to all of the pixels in the input images. The `Dense` layer object receives as arguments the number of units in that layer, and the activation function for the units. For the first layer in the network, it also receives an `input_shape` keyword argument.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
# Initializes a sequential model
model = Sequential()
# First layer
model.add(Dense(10, activation='relu', input_shape=(784, )))
# Second layer
model.add(Dense(10, activation='relu'))
# Output layer
model.add(Dense(3, activation='softmax'))
model.summary()
# -
# ### Compile a neural network
# Once you have constructed a model in Keras, the model needs to be compiled before you can fit it to data. This means that you need to specify the optimizer that will be used to fit the model and the loss function that will be used in optimization. Optionally, you can also specify a list of metrics that the model will keep track of. For example, if you want to know the classification accuracy, you will provide the list `['accuracy']` to the `metrics` keyword argument.
# Compile the model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# ### Fitting a neural network model to clothing data
# In this exercise, you will fit the fully connected neural network that you constructed in the previous exercise to image data. The training data is provided as two variables: `train_data` that contains the pixel data for 50 images of the three clothing classes and `train_labels`, which contains one-hot encoded representations of the labels for each one of these 50 images. Transform the data into the network's expected input and then fit the model on training data and training labels.
# +
(train_data, train_labels), (test_data, test_labels) = tf.keras.datasets.fashion_mnist.load_data()
train_data = train_data[(train_labels >= 0) & (train_labels < 3)][0:50].reshape(-1, 28, 28, 1)
train_labels = train_labels[(train_labels >= 0) & (train_labels < 3)][0:50]
train_labels = pd.get_dummies(train_labels).to_numpy()
test_data = test_data[(test_labels >= 0) & (test_labels < 3)][0:10].reshape(-1, 28, 28, 1)
test_labels = test_labels[(test_labels >= 0) & (test_labels < 3)][0:10]
test_labels = pd.get_dummies(test_labels).to_numpy()
# +
# Reshape the data to two-dimensional array
train_data = train_data.reshape((50, 784))
# Fit the model
model.fit(train_data, train_labels, validation_split=0.2, epochs=20, verbose=False);
# -
# ### Cross-validation for neural network evaluation
# To evaluate the model, we use a separate test data-set. As in the train data, the images in the test data also need to be reshaped before they can be provided to the fully-connected network because the network expects one column per pixel in the input.
# +
# Reshape test data
test_data = test_data.reshape((10, 784))
# Evaluate the model
model.evaluate(test_data, test_labels)
| _notebooks/2020-08-03-01-Image-Processing-With-Neural-Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
import sys
import os
import re
import collections
import itertools
import bcolz
import pickle
import numpy as np
import pandas as pd
import gc
import random
import smart_open
import h5py
import csv
import tensorflow as tf
import gensim
import string
import datetime as dt
from tqdm import tqdm_notebook as tqdm
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('seaborn-poster')
import seaborn as sns
random_state_number = 967898
# -
# # Code
# + [markdown] heading_collapsed=true
# ## Day 1: Inverse Captcha
# + hidden=true
# ! cat day1_input.txt
# + hidden=true
input_data = None
with open("day1_input.txt") as f:
input_data = f.read().strip().split()[0]
# + [markdown] hidden=true
# ### part1
# + [markdown] hidden=true
# The captcha requires you to review a sequence of digits (your puzzle input) and
#
# find the sum of all digits that match the next digit in the list. The list is circular,
#
# so the digit after the last digit is the first digit in the list.
# + hidden=true
def get_captcha11(s):
ssum = 0
for i1, i2 in zip(s, s[1:]+s[0]):
if i1 == i2:
ssum += int(i1)
return ssum
# + hidden=true
def get_captcha12(s):
n = len(s)
ssum = int(s[0]) if s[0] == s[-1] else 0
i1 = 0
while i1 < n-1:
if s[i1] == s[i1+1]:
ssum += int(s[i1])
i1+=1
return ssum
# + hidden=true
assert get_captcha12("1122") == 3
assert get_captcha12("1111") == 4
assert get_captcha12("1234") == 0
assert get_captcha12("91212129") == 9
# + hidden=true
get_captcha12(input_data)
# + [markdown] hidden=true
# ### part2
# + [markdown] hidden=true
# Now, instead of considering the next digit, it wants you to consider the digit halfway around
#
# the circular list. That is, if your list contains 10 items, only include a digit in your sum
#
# if the digit 10/2 = 5 steps forward matches it. Fortunately, your list has an even number of elements.
# + hidden=true
def get_captcha21(s):
n = len(s)
ssum = 0
i1 = 0
halfway_circle = n//2
while i1 < n:
if s[i1] == s[(i1+halfway_circle)%n]:
ssum += int(s[i1])
i1+=1
return ssum
# + hidden=true
assert get_captcha21("1212") == 6
assert get_captcha21("1221") == 0
assert get_captcha21("123425") == 4
assert get_captcha21("123123") == 12
assert get_captcha21("12131415") == 4
# + hidden=true
get_captcha21(input_data)
# + [markdown] heading_collapsed=true
# ## Day 2: Corruption Checksum
# + hidden=true
# ! cat day2_input.txt
# + hidden=true
input_data = []
with open("day2_input.txt") as f:
for line in f.read().split("\n"):
input_data += [list(map(int,line.split("\t")))] if line else []
# + [markdown] hidden=true
# ### part 1
# + [markdown] hidden=true
# For each row, determine the difference between the largest value and the
#
# smallest value; the checksum is the sum of all of these differences.
# + hidden=true
def get_checksum1(ll):
return sum([max(l) - min(l) for l in ll])
# + hidden=true
assert get_checksum1([[5,1,9,5],[7,5,3],[2,4,6,8]]) == 18
# + hidden=true
get_checksum1(input_data)
# + [markdown] hidden=true
# ### part 2
# + [markdown] hidden=true
# the goal is to find the only two numbers in each row where one evenly
#
# divides the other - that is, where the result of the division operation is
#
# a whole number. They would like you to find those numbers on each line,
#
# divide them, and add up each line's result.
# + hidden=true
def get_checksum2(ll):
csum = 0
for l in ll:
ht = {}
for i,a in enumerate(l):
for b in l[i:]:
if a == b:
continue
if a%b == 0:
csum += a//b
break
elif b%a == 0:
csum += b//a
break
return csum
# + hidden=true
assert get_checksum2([[5,9,2,8],[9,4,7,3],[3,8,6,5]]) == 9
# + hidden=true
get_checksum2(input_data)
# -
# ## Day 3: Spiral Memory
# ### part 1
# +
import math
def get_md_spiral_mem(n):
# this number marks the odd square end from where the counting starts.
# this counting is used to estimate the catesian cor ordinate which
# can be used to find the manhattan distance
ref = math.floor(math.sqrt(n))
# the spiral starts after the squared odd number
# if this is even reduce by 1 to get this place for reference
ref = ref - 1 if ref%2==0 else ref
# the number can be on either of the 4 sides when spiraling
# find difference from the squared odd number reference
diff = n - ref**2
# since all are odd number squares
# and the side considers the full side including
# the number that starts on the next side
side_size = ref+1
print("------", ref, diff, side_size)
# lets estimate the absolutre value of thier
# cartesian co ordinates. Since the reference is taken from the odd
# square numbers, handle them properly, this part is not implemented
# also this can be done pretty simply using comprehension,
# but this way the code is understandable
x,y = None, None
if n <= ref**2 + side_size:
# right side
x = side_size//2
d = ref**2 + side_size - n
print("----right",d)
y = (side_size//2) - d
elif n <= ref**2 + 2*side_size:
# top side
y = side_size//2
d = ref**2 + 2*side_size - n
print("----top",d)
x = (side_size//2) - d
elif n <= ref**2 + 3*side_size:
# left side
x = side_size//2
d = ref**2 + 3*side_size - n
print("----left",d)
y = (side_size//2) - d
else:
# bottom side
y = side_size//2
d = ref**2 + 4*side_size - n
print("----bottom",d)
x = (side_size//2) - d
print("-----",x,y)
return x + y
# -
get_md_spiral_mem(347991)
# ### part 2 (incomplete)
# + [markdown] heading_collapsed=true
# ## Day 4: High-Entropy Passphrases
# + hidden=true
# ! cat day4_input.txt
# + hidden=true
input_data = []
with open("day4_input.txt") as f:
for line in f.read().split("\n"):
input_data += [list(line.split())] if line else []
# + [markdown] hidden=true
# ### part 1
# + hidden=true
def get_valid1(list_of_strings):
valids = 0
for l in list_of_strings:
valids += 1 if len(set(l)) == len(l) else 0
return valids
# + hidden=true
assert get_valid([['aa','bb','cc','dd','ee']]) == 1
assert get_valid([['aa','bb','cc','dd','aaa']]) == 1
assert get_valid([['aa','bb','cc','dd','aa']]) == 0
# + hidden=true
get_valid(input_data)
# + [markdown] hidden=true
# ### part 2
# + hidden=true
import collections
def get_valid2(list_of_strings):
valids = 0
for l in list_of_strings:
counts = [collections.Counter(s) for s in l]
valids += 1 if all([c1 != c2 for i,c1 in enumerate(counts) for c2 in counts[i+1:]]) else 0
return valids
# + hidden=true
assert get_valid2([["abcde","fghij"]]) == 1
assert get_valid2([["abcde","xyz","ecdab"]]) == 0
assert get_valid2([["a","ab","abc","abd","abf","abj"]]) == 1
assert get_valid2([["iiii","oiii","ooii","oooi","oooo"]]) == 1
assert get_valid2([["oiii","ioii","iioi","iiio"]]) == 0
# + hidden=true
get_valid2(input_data)
# + [markdown] heading_collapsed=true
# ## Day 5: A Maze of Twisty Trampolines, All Alike
# + hidden=true
# ! cat day5_input.txt
# + hidden=true
input_data = []
with open("day5_input.txt") as f:
input_data = list(map(int,f.read().strip().split("\n")))
# + [markdown] hidden=true
# ### part 1
# + hidden=true
def when_out1(list_of_nos):
l = list_of_nos[:]
current_index = 0
steps = 0
while current_index <= len(l)-1 and current_index >= 0:
# print("----",steps, l,current_index, current_index <= len(l)-1, current_index >= 0)
steps += 1
forward_index_by = l[current_index]
l[current_index] += 1
current_index += forward_index_by
return steps
# + hidden=true
assert(when_out1([0,3,0,1,-3])) == 5
# + hidden=true
when_out1(input_data)
# + [markdown] hidden=true
# ### part 2
# + hidden=true
def when_out2(list_of_nos):
l = list_of_nos[:]
current_index = 0
steps = 0
while current_index <= len(l)-1 and current_index >= 0:
# print("----",steps, l,current_index, current_index <= len(l)-1, current_index >= 0)
steps += 1
forward_index_by = l[current_index]
if forward_index_by >= 3:
l[current_index] -= 1
else:
l[current_index] += 1
current_index += forward_index_by
return steps
# + hidden=true
assert(when_out2([0,3,0,1,-3])) == 10
# + hidden=true
when_out2(input_data)
# + [markdown] heading_collapsed=true
# ## Day 6: Memory Reallocation
# + hidden=true
# ! cat day6_input.txt
# + hidden=true
input_data = []
with open("day6_input.txt") as f:
input_data = list(map(int,f.read().strip().split()))
# + [markdown] hidden=true
# ### part 1
# + hidden=true
import numpy as np
def cycling_reallocated_buffers1(buffers):
l = np.array(buffers)
n = len(l)
diviup_by = n-1
visited_buff_configs = set()
buffer_config = "-".join(list(map(str,l)))
step = 1
while buffer_config not in visited_buff_configs:
visited_buff_configs.add(buffer_config)
max_index = np.argmax(l)
divided = l[max_index]//diviup_by
remainder = l[max_index]%diviup_by
if divided == 0:
for i in range(1,l[max_index]+1):
l[(max_index+i)%n] += 1
l[max_index] = 0
else:
l += divided
l[max_index] = 0
l[max_index] += remainder
buffer_config = "-".join(list(map(str,l)))
step += 1
return step - 1
# + hidden=true
assert cycling_reallocated_buffers1([0,2,7,0]) == 5
# + hidden=true
cycling_reallocated_buffers1(input_data)
# + [markdown] hidden=true
# ### part 2
# + hidden=true
import numpy as np
def cycling_reallocated_buffers2(buffers):
l = np.array(buffers)
n = len(l)
diviup_by = n-1
visited_buff_configs = {}
buffer_config = "-".join(list(map(str,l)))
step = 1
while buffer_config not in visited_buff_configs:
visited_buff_configs[buffer_config] = step
max_index = np.argmax(l)
divided = l[max_index]//diviup_by
remainder = l[max_index]%diviup_by
if divided == 0:
for i in range(1,l[max_index]+1):
l[(max_index+i)%n] += 1
l[max_index] = 0
else:
l += divided
l[max_index] = 0
l[max_index] += remainder
buffer_config = "-".join(list(map(str,l)))
step += 1
return step - visited_buff_configs[buffer_config]
# + hidden=true
assert cycling_reallocated_buffers2([0,2,7,0]) == 4
# + hidden=true
cycling_reallocated_buffers2(input_data)
# -
# ## Day 7: Recursive Circus
# ! cat day7_input.txt
# +
input_data_adj_list = {}
input_data_values = {}
with open("day7_input.txt") as f:
for line in f.read().strip().split("\n"):
m = re.match(r"([a-z]+)\s+\((\d+)\)\s*[->\s]*(.*)",line)
input_data_values[m.group(1)] = int(m.group(2))
input_data_adj_list[m.group(1)] = m.group(3).split(", ") if m.group(3) else []
# input_data_adj_list
# input_data_values
# -
# ### part 1
def find_source1(adj_list):
# hash table to track incoming edges
nodes = {node:0 for node in adj_list.keys()}
for from_node, to_nodes in adj_list.items():
for to_node in to_nodes:
nodes[to_node] += 1
return [node for node,val in nodes.items() if val == 0][0]
# +
sample = """pbga (66)
xhth (57)
ebii (61)
havc (66)
ktlj (57)
fwft (72) -> ktlj, cntj, xhth
qoyq (66)
padx (45) -> pbga, havc, qoyq
tknk (41) -> ugml, padx, fwft
jptl (61)
ugml (68) -> gyxo, ebii, jptl
gyxo (61)
cntj (57)
"""
sample_values = {}
sample_adj_list = {}
for line in sample.strip().split("\n"):
m = re.match(r"([a-z]+)\s+\((\d+)\)\s*[->\s]*(.*)",line)
sample_values[m.group(1)] = int(m.group(2))
sample_adj_list[m.group(1)] = m.group(3).split(", ") if m.group(3) else []
assert find_source1(sample_adj_list) == "tknk"
# -
find_source1(input_data_adj_list)
# ### part 2 (incomplete)
def find_unbalanced_weight(adj_list, values):
# get sum from all the leaf nodes that map to a single parent
for from_node, to_nodes in adj_list.items():
# finding the leaf nodes
if len(to_nodes) > 0 and all([len(adj_list[to_node]) == 0 for to_node in to_nodes]):
print("----",from_node,values[from_node]+sum([values[to_node] for to_node in to_nodes]))
find_unbalanced_weight(sample_adj_list, sample_values)
[(to_nodes,input_data_adj_list[to_nodes]) for to_nodes in input_data_adj_list["okzkfw"]]
find_unbalanced_weight(input_data_adj_list, input_data_values)
# ## Day 8: I Heard You Like Registers
# ! cat day8_input.txt
# +
input_data_registers = {}
input_inc_dec = []
input_cond = []
with open("day8_input.txt") as f:
for line in f.read().strip().split("\n"):
m = re.match(r"(([a-z]+)\s[a-z]+\s[-]*\d+)\sif\s(([a-z]+)\s[!><=]+\s[-]*\d+)",line)
input_data_registers[m.group(2)] = 0
input_data_registers[m.group(4)] = 0
input_inc_dec += [m.group(1)]
input_cond += [m.group(3)]
len(input_data_registers),input_data_registers
# -
# ### part 1
# +
import operator
def get_operator_fn(op):
return {
'<' : operator.lt,
'<=' : operator.le,
'==' : operator.eq,
'!=' : operator.ne,
'>=' : operator.ge,
'>' : operator.gt,
'inc': operator.add,
'dec': operator.sub
}[op]
def find_largest_reg1(data_registers, inc_dec, cond):
for i in range(len(inc_dec)):
cond_l = cond[i].split(" ")
if get_operator_fn(cond_l[1])(data_registers[cond_l[0]], int(cond_l[2])):
inc_dec_l = inc_dec[i].split(" ")
data_registers[inc_dec_l[0]] = get_operator_fn(inc_dec_l[1])(data_registers[inc_dec_l[0]], int(inc_dec_l[2]))
return max(data_registers.items(), key=operator.itemgetter(1))[1]
# -
sample = """b inc 5 if a > 1
a inc 1 if b < 5
c dec -10 if a >= 1
c inc -20 if c == 10
"""
sample_data_registers = {}
sample_inc_dec = []
sample_cond = []
for line in sample.strip().split("\n"):
m = re.match(r"(([a-z]+)\s[a-z]+\s[-]*\d+)\sif\s(([a-z]+)\s[!><=]+\s[-]*\d+)",line)
sample_data_registers[m.group(2)] = 0
input_data_registers[m.group(4)] = 0
sample_inc_dec += [m.group(1)]
sample_cond += [m.group(3)]
assert find_largest_reg1(sample_data_registers, sample_inc_dec, sample_cond) == 1
find_largest_reg1(input_data_registers, input_inc_dec, input_cond)
# ### part 2
def find_largest_reg2(data_registers, inc_dec, cond):
max_value = -float('inf')
for i in range(len(inc_dec)):
cond_l = cond[i].split(" ")
if get_operator_fn(cond_l[1])(data_registers[cond_l[0]], int(cond_l[2])):
inc_dec_l = inc_dec[i].split(" ")
data_registers[inc_dec_l[0]] = get_operator_fn(inc_dec_l[1])(data_registers[inc_dec_l[0]], int(inc_dec_l[2]))
max_value = max(max_value, data_registers[inc_dec_l[0]])
return max_value
assert find_largest_reg2(sample_data_registers, sample_inc_dec, sample_cond) == 10
find_largest_reg2(input_data_registers, input_inc_dec, input_cond)
# ## Day 9: Stream Processing
# ! cat day9_input.txt
input_data = None
with open("day9_input.txt") as f:
input_data = f.read().strip()
# ### part 1
# +
def update_cancelled_chars(s):
n = len(s)
updated_s = ''
i = 0
while i < n:
if s[i] == '!':
i += 2
else:
updated_s += s[i]
i += 1
return updated_s
def score_stream1(s):
matching_braces = {'{':'}', '<':'>'}
updated_s = update_cancelled_chars(s)
# print("----",updated_s)
n = len(updated_s)
# some corner cases to add the first character
# are not verified
# push and pop at the end
stack = []
score = 0
current_stream_score = 0
seen_garbage_start = False
for c in updated_s:
if c not in ['{','}','<','>']:
continue
if c == '<':
seen_garbage_start = True
if c == '>':
seen_garbage_start = False
if seen_garbage_start is False:
if c == '{':
current_stream_score += 1
stack += [('{', current_stream_score)]
score += current_stream_score
elif c == matching_braces[stack[-1][0]]:
stack.pop()
current_stream_score -= 1
# print("----",score)
return score
# -
assert update_cancelled_chars("<!!!>>") == "<>"
assert update_cancelled_chars("<{o\"i!a,<{i<a>") == "<{o\"i,<{i<a>"
assert update_cancelled_chars("<!!>") == "<>"
assert update_cancelled_chars("<{!>}>") == "<{}>"
assert score_stream1("{}") == 1
assert score_stream1("{{{}}}") == 6
assert score_stream1("{{},{}}") == 5
assert score_stream1("{{{},{},{{}}}}") == 16
assert score_stream1("{<a>,<a>,<a>,<a>}") == 1
assert score_stream1("{{<ab>},{<ab>},{<ab>},{<ab>}}") == 9
assert score_stream1("{{<!!>},{<!!>},{<!!>},{<!!>}}") == 9
assert score_stream1("{{<a!>},{<a!>},{<a!>},{<ab>}}") == 3
score_stream1(input_data)
# ### part 2
def score_stream2(s):
matching_braces = {'{':'}', '<':'>'}
updated_s = update_cancelled_chars(s)
n = len(updated_s)
garbage_count = 0
seen_garbage_start = False
for c in updated_s:
if seen_garbage_start is True:
garbage_count += 1
if c == '<':
seen_garbage_start = True
if c == '>':
seen_garbage_start = False
garbage_count -= 1
return garbage_count
assert score_stream2("<>") == 0
assert score_stream2("<random characters>") == 17
assert score_stream2("<<<<>") == 3
assert score_stream2("<{!>}>") == 2
assert score_stream2("<!!>") == 0
assert score_stream2("<!!!>>") == 0
assert score_stream2("<{o\"i!a,<{i<a>") == 10
score_stream2(input_data)
# ## Day 10: Knot Hash
# ! cat day10_input.txt
input_data = None
with open('day10_input.txt') as f:
input_data = f.read().strip()
input_data
# ### part 1
# +
def perform_one_round_knot_hash(l,lengths,ci=0,skip_size=0):
n = len(l)
# ci current index
for length in lengths:
#print("ci",ci,"skip",skip_size,"length",length,l)
if ci+length > n:
length_in_end = ci+length-n
new_l = (l[ci:ci+length] + l[:length_in_end])[::-1]
# print("new_l",new_l)
# print(l[ci:ci+length],new_l[:-length_in_end])
# print(l[:length_in_end],new_l[-length_in_end:])
l[ci:ci+length] = new_l[:-length_in_end]
l[:length_in_end] = new_l[-length_in_end:]
else:
l[ci:ci+length] = l[ci:ci+length][::-1]
ci = (ci + length + skip_size)%n
skip_size += 1
# print("ci",ci,l)
# print("======")
return (l, ci, skip_size)
def howmany_knothashes(input_chars, lengths):
l, _, _ = perform_one_round_knot_hash(input_chars, lengths)
return l[0]*l[1]
# -
assert howmany_knothashes(list(range(5)),[3,4,1,5]) == 12
howmany_knothashes(list(range(256)),list(map(int,input_data.split(","))))
# ### part 2
def knot_hash(input_string, rounds=64):
suffix = [17, 31, 73, 47, 23]
lengths = [ord(ch) for ch in input_string] + suffix
l = list(range(256))
# round 1
l, ci, skip_size = perform_one_round_knot_hash(l, lengths)
for r in range(rounds-1):
l, ci, skip_size = perform_one_round_knot_hash(l, lengths, ci, skip_size)
import functools
si,ei = 0,16
dense_hash = ""
for i in range(16):
xored = functools.reduce(lambda x,y: x^y, l[si:ei])
dense_hash += '{:02x}'.format(xored)
si += 16
ei += 16
return dense_hash
assert knot_hash("") == "a2582a3a0e66e6e86e3812dcb672a272"
assert knot_hash("AoC 2017") == "33efeb34ea91902bb2f59c9920caa6cd"
assert knot_hash("1,2,3") == "3efbe78a8d82f29979031a4aa0b16a9d"
assert knot_hash("1,2,4") == "63960835bcdc130f0b66d7ff4f6a5a8e"
knot_hash(input_data)
len(bin(0xa2582a3a0e66e6e86e3812dcb672a272))-2
# ## Day 11: Hex Ed
# ! cat day11_input.txt
input_data = None
with open("day11_input.txt") as f:
input_data = f.read().strip().split(",")
len(input_data)
from IPython.display import Image
Image(filename='/home/bicepjai/Projects/mypuzzles/adventofcode/2017/images/hex_grid.png')
# https://www.redblobgames.com/grids/hexagons/
# ### part 1
# +
# hex grid directions in cube co ordinates (x,y,z)
dir_map = {"n" :(0,1,-1), "ne":(1,0,-1), "nw":(-1,1,0),
"s" :(0,-1,1), "se":(1,-1,0), "sw":(-1,0,1)}
add_2cube_cord = lambda xyz1,xyz2:(xyz1[0]+xyz2[0],xyz1[1]+xyz2[1],xyz1[2]+xyz2[2])
distance_from_origin = lambda xyz: max(abs(xyz[0]), abs(xyz[1]), abs(xyz[2]))
def shortest_distance_to_hexgrid1(path):
# starting at origin
cco = (0,0,0) #current_co_ordinate
for d in path:
cco = add_2cube_cord(cco, dir_map[d])
# shortest distance from origin
return distance_from_origin(cco)
# -
assert shortest_distance_to_hexgrid1(["ne","ne","ne"]) == 3
assert shortest_distance_to_hexgrid1(["ne","ne","sw","sw"]) == 0
assert shortest_distance_to_hexgrid1(["ne","ne","s","s"]) == 2
assert shortest_distance_to_hexgrid1(["se","sw","se","sw","sw"]) == 3
shortest_distance_to_hexgrid(input_data)
# ### part 2
def shortest_distance_to_hexgrid2(path):
# starting at origin
cco = (0,0,0) #current_co_ordinate
max_distance = 0
for d in path:
cco = add_2cube_cord(cco, dir_map[d])
max_distance = max(max_distance, distance_from_origin(cco))
return max_distance
shortest_distance_to_hexgrid2(input_data)
# ### for fun
def get_path_cords(path):
# starting at origin
path_cords = []
cco = (0,0,0) #current_co_ordinate
for d in path:
path_cords += [cco]
cco = add_2cube_cord(cco, dir_map[d])
return path_cords
# +
path_cords = get_path_cords(input_data)
fig,axes = plt.subplots(nrows=1,ncols=3)
fig.set_size_inches(18,8)
axes[0].set(title="pathxy")
axes[0].plot([t[0] for t in path_cords],[t[1] for t in path_cords])
axes[1].set(title="pathyz")
axes[1].plot([t[1] for t in path_cords],[t[2] for t in path_cords])
axes[2].set(title="pathzx")
axes[2].plot([t[2] for t in path_cords],[t[0] for t in path_cords])
plt.show()
# -
# ## Day 12: Digital Plumber
# ! cat day12_input.txt
# forming adjaceny list as provided
input_data = {}
with open("day12_input.txt") as f:
for line in f.read().strip().split("\n"):
m = re.match(r"(\d+)\s*<->(.*)",line)
input_data[int(m.group(1).strip())] = [int(n.strip()) for n in m.group(2).strip().split(",")]
len(input_data)
# +
def connected_componenets_dfs1(adj_list):
# perform depth first search to count the components
# connected to the node 0
num_connected = 0
visited = set()
def helper_dfs(node):
nonlocal num_connected, visited, adj_list
if node not in visited:
visited.add(node)
num_connected += 1
for child in adj_list[node]:
helper_dfs(child)
helper_dfs(0)
return num_connected
def connected_componenets_dfs2(adj_list):
# perform depth first search to count the components
# connected to the node 0
num_connected = 0
visited = set()
def helper_dfs(node):
nonlocal num_connected, visited, adj_list
visited.add(node)
num_connected += 1
for child in adj_list[node]:
if child not in visited:
helper_dfs(child)
helper_dfs(0)
return num_connected
# +
def connected_componenets_bfs1(adj_list):
# perform breadth first search to count the components
# connected to the node 0
num_connected = 0
visited = set()
q = [0] # enqueue->append dequeue->pop(0)
while len(q) > 0:
node = q.pop(0)
if node not in visited:
visited.add(node)
num_connected += 1
for child in adj_list[node]:
q += [child]
return num_connected
def connected_componenets_bfs2(adj_list):
# perform breadth first search to count the components
# connected to the node 0
num_connected = 0
visited = set()
q = [0] # enqueue->append dequeue->pop(0)
while len(q) > 0:
node = q.pop(0)
for child in adj_list[node]:
if child not in visited:
visited.add(child)
num_connected += 1
q += [child]
return num_connected
# -
sample_input="""0 <-> 2
1 <-> 1
2 <-> 0, 3, 4
3 <-> 2, 4
4 <-> 2, 3, 6
5 <-> 6
6 <-> 4, 5
"""
sample_data = {}
for line in sample_input.strip().split("\n"):
m = re.match(r"(\d+)\s*<->(.*)",line)
sample_data[int(m.group(1).strip())] = [int(n.strip()) for n in m.group(2).strip().split(",")]
assert connected_componenets_dfs1(sample_data) == connected_componenets_dfs2(sample_data)
assert connected_componenets_dfs1(sample_data) == connected_componenets_bfs1(sample_data)
assert connected_componenets_bfs1(sample_data) == connected_componenets_bfs2(sample_data)
assert connected_componenets_dfs1(sample_data) == connected_componenets_bfs2(sample_data)
assert connected_componenets_dfs1(input_data) == connected_componenets_dfs2(input_data)
assert connected_componenets_dfs1(input_data) == connected_componenets_bfs1(input_data)
assert connected_componenets_bfs1(input_data) == connected_componenets_bfs2(input_data)
assert connected_componenets_dfs1(input_data) == connected_componenets_bfs2(input_data)
connected_componenets_bfs2(input_data)
# ### part 2
def nof_connected_componenets_bfs(adj_list):
num_groups = 0
visited = set()
# lets go thru all the nodes
# and form connected compenents
for root in adj_list:
if root not in visited:
num_groups += 1
q = [root]
while len(q) > 0:
node = q.pop(0)
if node not in visited:
visited.add(node)
for child in adj_list[node]:
q += [child]
return num_groups
assert nof_connected_componenets_bfs(sample_data) == 2
nof_connected_componenets_bfs(input_data)
# ## Day 13: Packet Scanners
# ! cat day13_input.txt
input_data = {}
with open("day13_input.txt") as f:
for line in f.read().strip().split("\n"):
k,v = (p for p in line.split(":"))
input_data[int(k)] = int(v)
len(input_data)
# ### part 1
# Brute Force Solution where we are updating all the scanners as per the time step
# +
def update_scanner_depths(scanner_pos,scanner_depths):
for d in scanner_depths:
scanner_pos[d][0] += scanner_pos[d][1]
if (scanner_pos[d][0] == scanner_depths[d]-1 or scanner_pos[d][0] == 0):
scanner_pos[d][1] *= -1
def scanner_severity_bf(scanner_depths):
"""
given scanner depths find serverity
"""
# dict holding the current depth and the inc/dec flag
# once it reached bottom or top flag multiplies by -1, thus rotating
scanner_pos = {k:[0,1] for k in scanner_depths.keys()}
# since we always get caught first
caughts = []
# going thru each pico second thru each scanner
# if scanner find us in place, we are not caught
# cur_pos is same as current time
total_time_range = max(scanner_depths.keys()) + 1
for cur_time in range(total_time_range):
# when the cur_pos that indicates the
# scanner caught us, its index will be top one i.e, 0
if cur_time in scanner_depths.keys() and scanner_pos[cur_time][0] == 0:
caughts += [(cur_time, scanner_depths[cur_time])]
# update scanner depths
update_scanner_depths(scanner_pos,scanner_depths)
return sum([a*b for a,b in caughts])
# -
#
# one could notice that we are interested only in the 0th position not others, lets say the depth is 3.
# the channel would be tracking as
#
# depth 3
#
# time 0 1 2 3 4 5 6 7 8 9
#
# position 0 1 2 1 0 1 2 1 0 1
#
# one could notice the 0s are always occuring at timesteps that are multiple of 4.
# can we generalize this pattern
#
# depth: 2
#
# time 0 1 2 3 4 5 6 7 8 9
#
# position 0 1 0 1 0 1 0 1 0 1
#
# depth: 4
#
# time 0 1 2 3 4 5 6 7 8 9
#
# position 0 1 2 3 2 1 0 1 2 3
#
# its (depth - 1)*2
#
def scanner_severity(scanner_depths):
severity = 0
# here sc_id is also time
for sc_id_time, depth in scanner_depths.items():
if sc_id_time % ((depth - 1)*2) == 0:
severity += (sc_id_time * depth)
return severity
sample_depths = {0:3, 1:2, 4:4, 6:4}
assert scanner_severity(sample_depths) == 24
scanner_severity(input_data)
# ### part 2
def escaped_scanners_bf(scanner_depths):
caught = True
delay_time = 0
while caught:
# dict holding the current depth and the inc/dec flag
# once it reached bottom or top flag multiplies by -1, thus rotating
scanner_pos = {k:[0,1] for k in scanner_depths.keys()}
caught = False
# run for the delayed time
for i in range(delay_time):
update_scanner_depths(scanner_pos,scanner_depths)
# go thru the scanners now
total_time_range = max(scanner_depths.keys()) + 1
for cur_time in range(total_time_range):
# when the cur_pos that indicates the
# scanner caught us, its index will be top one i.e, 0
# print("---b4 ",cur_time+10,scanner_pos)
if cur_time in scanner_depths.keys() and scanner_pos[cur_time][0] == 0:
# print("---caught---",cur_time,scanner_pos[cur_time])
caught = True
delay_time += 1
break
# update scanner depths
update_scanner_depths(scanner_pos,scanner_depths)
# print("---aftr",cur_time+10,scanner_pos)
# successfully ran thru without getting caught
# print("=====================",delay_time)
if not caught:
break
return delay_time
def escaped_scanners(scanner_depths):
delay_time = 0
caught = True
while caught:
caught = False
for sc_id_time,depth in scanner_depths.items():
if (sc_id_time + delay_time) % ((depth-1)*2) == 0:
caught = True
delay_time += 1
break
if not caught:
break
return delay_time
assert escaped_scanners_bf(sample_depths) == 10
assert escaped_scanners(sample_depths) == 10
escaped_scanners(input_data)
(1,1,"u")[:2]
# ## Day 14: Disk Defragmentation
input_data = 'ffayrhll'
sample_data = 'flqrgnkx'
# ### part 1
# +
bit_map = {
0:0, 1:1, 2:1, 3:2,
4:1, 5:2, 6:2, 7:3,
8:1, 9:2, 10:2, 11:3,
12:2, 13:3, 14:3, 15:4
}
def count_bits(hex_string):
nof1s = 0
# print("----",hex_string)
for hex_ch in hex_string:
nof1s += bit_map[int(hex_ch, 16)]
return nof1s
# +
def square_fragment_bits(data):
fragment_bits = []
for i in range(0,128):
hash_str = data + "-" + str(i)
fragment_bits += [knot_hash(hash_str)]
return fragment_bits
def number_of_square_fragments(data):
nofsquares = 0
for hash_str in square_fragment_bits(data):
nofsquares += sum(bin(int(ch, 16))[2:].count('1') for ch in hash_str)
return nofsquares
# -
number_of_square_fragments(sample_data) == 8108
number_of_square_fragments(input_data)
# ### part 2
# +
def mark_regions(data, r, c, region):
if r > 127 or r < 0 or c > 127 or c < 0:
return
if data[r][c] == 1:
# mark
data[r][c] = region
# recurse near by squares
mark_regions(data, r+1, c, region)
mark_regions(data, r-1, c, region)
mark_regions(data, r, c-1, region)
mark_regions(data, r, c+1, region)
def count_regions(data):
fragment_bits = square_fragment_bits(data)
bit_strings = []
# lets make 128 x 128 bit strings
for hash_str in fragment_bits:
bit_string = bin(int(hash_str, 16))[2:]
bit_string = "".join(['0']*(128-len(bit_string))) + bit_string
bit_strings += [list(map(int,bit_string))]
# go thru each square
regions = 2
for r in range(0,128):
for c in range(0,128):
if bit_strings[r][c] == 1:
mark_regions(bit_strings, r, c, regions)
regions += 1
return regions - 2
# -
sys.getrecursionlimit()
count_regions(sample_data) == 1242
count_regions(input_data)
# ## Day 15: Dueling Generators
# input_data
#
# Generator A starts with 512
#
# Generator B starts with 191
#
# +
gen_A_input = 512
gen_B_input = 191
gen_A_sample = 65
gen_B_sample = 8921
# -
# ### part 1
# +
def generatorA(input_value):
current_value = input_value
while True:
current_value = (16807*current_value) % 2147483647
yield current_value
def generatorB(input_value):
current_value = input_value
while True:
current_value = (48271*current_value) % 2147483647
yield current_value
def num_matches(gena_input, genb_input, rounds=int(4e7)):
matches = 0
gena_output = generatorA(gena_input)
genb_output = generatorB(genb_input)
bits16 = 0xFFFF
for i in range(rounds):
if next(gena_output)&bits16 == next(genb_output)&bits16:
matches += 1
return matches
# -
num_matches(gen_A_sample, gen_B_sample, rounds=5) == 1
num_matches(gen_A_sample, gen_B_sample, rounds=int(4e7)) == 588
num_matches(gen_A_input, gen_B_input, rounds=int(4e7))
# ### part 2
# +
def generatorA(input_value):
current_value = input_value
while True:
current_value = (16807*current_value) % 2147483647
if current_value % 4 == 0:
yield current_value
def generatorB(input_value):
current_value = input_value
while True:
current_value = (48271*current_value) % 2147483647
if current_value % 8 == 0:
yield current_value
# -
num_matches(gen_A_sample, gen_B_sample, rounds=int(5e6)) == 309
num_matches(gen_A_input, gen_B_input, rounds=int(5e6))
# ## Day 16: Permutation Promenade
# ! cat day16_input.txt
input_data = None
with open('day16_input.txt') as f:
input_data = list(map(lambda s: (s[0],s[1:]) if s[0] == 's' else (s[0], tuple(s[1:].split('/'))), f.read().strip().split(",")))
# ### part 1
# +
def one_dance_move(programs, command, params):
if command == 's':
index = int(params)
programs = programs[-index:] + programs[:-index]
elif command == 'x':
i1,i2 = tuple(map(int, params))
c1, c2 = programs[i1], programs[i2]
programs = programs.replace(c1,'z')
programs = programs.replace(c2,c1)
programs = programs.replace('z',c2)
elif command == 'p':
c1,c2 = params
programs = programs.replace(c1,'z')
programs = programs.replace(c2,c1)
programs = programs.replace('z',c2)
return programs
def danceit1(data, programs=None, length=16):
programs = "".join([chr(ord('a')+i) for i in range(length)]) if programs is None else programs
for command, params in data:
programs = one_dance_move(programs, command, params)
return programs
# -
danceit1([('s','1'),('x',('3','4')),('p',('e','b'))], length=5) == 'baedc'
danceit1(input_data, length=16)
# ### part 2
def danceit2(data, length=16, rounds=int(1e9)):
programs = "".join([chr(ord('a')+i) for i in range(length)])
# lets find the cycle
hash_map = {}
prev_programs = programs
cycle_round = None
recycled_programs = None
for r in range(rounds):
programs = danceit1(input_data, prev_programs)
if prev_programs in hash_map:
cycle_round = r
recycled_programs = prev_programs
break
hash_map[prev_programs] = programs
prev_programs = programs
# thru inspection we know its the same starting pattern
# 'abcdefghijklmnop' occuring again, hence just harding coding this
# if we need a general program, we need to process hash table to
# figure out the starting and the ending cycle and form the transformation as necessary
programs = "".join([chr(ord('a')+i) for i in range(length)])
for r in range(rounds%cycle_round):
programs = danceit1(input_data, programs)
print("----aftr",programs)
return programs
danceit2(input_data)
# ## Day 17: Spinlock
#
| adventofcode/2017/.ipynb_checkpoints/codes-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pandas import read_csv
import numpy as np
df = read_csv('international-airline-passengers.csv', usecols=[1])
df.describe()
values = df.values.astype('float32')
values.shape
train_size = int((values.shape[0] * 0.67))
test_size = values.shape[0] - train_size
train = values[0:train_size]
test = values[train_size:]
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range =(0,1))
train = scaler.fit_transform(train)
test = scaler.transform(test)
def create_dataset(data, k):
dataX, dataY = [],[]
for i in range(data.shape[0] - k):
x = data[i:i + k, 0]
y = data[i + k, 0]
dataX.append(x)
dataY.append(y)
return np.array(dataX), np.array(dataY)
look_back = 12
trainX, trainY = create_dataset(train, look_back)
testX, testY = create_dataset(test, look_back)
trainX = np.reshape(trainX, (trainX.shape[0], trainX.shape[1], 1))
testX = np.reshape(testX, (testX.shape[0], testX.shape[1], 1))
trainX.shape
from keras.models import Sequential
from keras.layers import LSTM, Dense
model = Sequential()
model.add(SimpleRNN(4, input_shape=(look_back, 1)))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(trainX, trainY, epochs=10, batch_size = 1)
# +
testPredict = model.predict(testX)
testPredict = scaler.inverse_transform(testPredict).ravel()
trainPredict = model.predict(trainX)
trainPredict = scaler.inverse_transform(trainPredict).ravel()
testTrue = scaler.inverse_transform([testY]).ravel()
trainTrue = scaler.inverse_transform([trainY]).ravel()
testPredict.shape, testTrue.shape
# -
from matplotlib import pyplot as plt
plt.plot(trainTrue, c = 'g')
plt.plot(trainPredict, c = 'b')
plt.show()
combinedPredicted = np.concatenate((trainPredict, testPredict))
combinedTrue = np.concatenate((trainTrue, testTrue))
plt.plot(combinedTrue, c = 'g')
plt.plot(combinedPredicted, c = 'b')
plt.show()
| Lecture 29 RNN/Analysing the output/8. RNN-airlines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparse fascicle models
#
# As mentioned in the [DTI](DTI.ipynb) notebook, the tensor model has had large popularity and huge success in describing data measured in the white matter in many different situations. Nevertheless, already early on, in the work of <NAME> and colleagues, and continuing with the work of <NAME> (Frank, 2001, 2002), and others, there was an increasing understanding that though DTI may be a good *phenomenological* model of the diffusion data, it is not always a good description of the structure of the tissue. Thus, not necessarily a good *mechanistic* model of the physical causes of the signal. In particular, the principal diffusion direction (PDD) was initially thought to be a good description of the orientation of the nerve fiber fascicles within the voxel. However, in locations in which there is more than one fascicle, the PDD is oriented towards the average of these, rather than towards any of them in particular. Thus, the parameters of the model may be misleading, if one does not take care in interpreting them.
# +
import os.path as op
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
import nibabel as nib
import dipy.core.gradients as grad
import dipy.sims.voxel as sims
import dipy.reconst.dti as dti
import dipy.reconst.sfm as sfm
import dipy.direction.peaks as peaks
# -
gtab = grad.gradient_table(op.join('data', 'SUB1_b2000_1.bvals'), op.join('data', 'SUB1_b2000_1.bvecs'))
# Consider for example the following simulation of a signal from a simulated voxel with two different fascicles, occupying each an equal amount of partial volume of the voxel and oriented at an angle of 90 degrees from each other:
SNR = 100
S0 = 100
mevals = np.array(([0.0015, 0.0005, 0.0005],
[0.0015, 0.0005, 0.0005]))
angles = [(0, 0), (90, 0)]
sig, sticks = sims.multi_tensor(gtab, mevals, S0, angles=angles,
fractions=[50, 50], snr=SNR)
# In this case, the true fascicle directions are:
sticks
dti_model = dti.TensorModel(gtab)
dti_fit = dti_model.fit(sig)
# The principal diffusion direction of the model points in neither of these directions:
dti_fit.evecs[0]
# Even though the model provides an excellent fit to the data
plt.plot(sig, 'o-')
plt.plot(dti_fit.predict(gtab, S0=100))
# As a response to this challenge, several groups (Behrens et al. 2007, Tournier et al. 2007, Dell'Acqua et al. 2007) have proposed models and algorithms that describe the diffusion signal as a combination of signals due to different populations of fibers within the voxels. Since these models all restrict the number of fascicles in each voxel, we collectively refer to the as **Sparse Fascicle Models**.
#
# One way of formally describing these models is:
#
# $y = X\beta$
#
# Where $y$ is the signal and $\beta$ are weights on different points in the sphere. The columns of the design matrix, $X$ are the signals in each point in the measurement that would be predicted if there was a fascicle oriented in the direction of that column. Typically, this will be a prolate tensor with axial diffusivity 3-5 times higher than its radial diffusivity. The exact numbers can also be estimated from examining parts of the brain in which there is known to be only one fascicle (e.g. in corpus callosum).
#
# We have developed an algorithm that fits this model applying sparsity constraints on the fiber ODF ($\beta$) through the the Elastic Net algorihtm (Zou and Hastie, 2005)
#
# Elastic Net optimizes the following cost function:
#
#
# $\sum_{i=1}^{n}{(y_i - \hat{y}_i)^2} + \alpha (\lambda \sum_{j=1}^{m}{w_j} +
# (1-\lambda) \sum_{j=1}^{m}{w^2_j}$
#
# where $\hat{y}$ is the signal predicted for a particular setting of $\beta$, such that the left part of this expression is the squared loss function; $alpha$ is a parameter that sets the balance between the squared loss on the data, and the regularization constraints. The regularization parameter $\lambda$ sets the `l1_ratio`, which controls the balance between L1-sparsity (low sum of weights), and low L2-sparsity (low sum-of-squares of the weights).
#
# Below, we demonstrate the use of this algorithm to fit these data.
# For the simulated fasicle directions fitting such a model provides not only a good fit to the data, but also approximately correct fascicle directions
sf_model = sfm.SparseFascicleModel(gtab)
sf_fit = sf_model.fit(sig)
# Peak directions are derived from the orientation distribution function (ODF) of the model.
peak_dirs, _, _ = peaks.peak_directions(sf_fit.odf(sf_model.sphere), sf_model.sphere)
peak_dirs
# Let's consider a voxel in the actual data, in which the DTI model does not fit the data very well:
vox_idx = (53, 43, 47)
data1 = nib.load(op.join('data', 'SUB1_b2000_1.nii.gz')).get_fdata()[vox_idx]
data2 = nib.load(op.join('data', 'SUB1_b2000_2.nii.gz')).get_fdata()[vox_idx]
dti_fit = dti_model.fit(data1)
dti_predict = dti_fit.predict(gtab, S0=np.mean(data1[gtab.b0s_mask]))
fig, ax = plt.subplots(1)
ax.plot(data2,'o-')
ax.plot(dti_predict,'o-')
fig.set_size_inches([10,6])
# As before the DTI model can only represent a single principal diffusion direction:
dti_fit.evecs[0]
# while the SFM can find two distinct peaks in the ODF, or two different fascicle directions in this voxel
sf_fit = sf_model.fit(data1)
sf_predict = sf_fit.predict(gtab, S0=np.mean(data1[gtab.b0s_mask]))
peak_dirs, _, _ = peaks.peak_directions(sf_fit.odf(sf_model.sphere), sf_model.sphere)
print(peak_dirs)
# This also results in a more accurate fit of the model to the data, and a fit that is substantially more accurate than test-retest reliability
fig, ax = plt.subplots(1)
ax.plot(data2,'o-')
ax.plot(dti_predict,'o-')
ax.plot(sf_predict,'o-')
fig.set_size_inches([10,6])
plt.hist(np.abs(sf_predict - data2), histtype='step')
plt.hist(np.abs(dti_predict - data2), histtype='step')
plt.hist(np.abs(data1 - data2), histtype='step')
rmse_retest = np.sqrt(np.mean((data1 - data2)**2))
rmse_dti = np.sqrt(np.mean((dti_predict - data2)**2))
rmse_sfm = np.sqrt(np.mean((sf_predict - data2)**2))
print("Test-retest RMSE: %2.2f"%rmse_retest)
print("DTI RMSE: %2.2f"%rmse_dti)
print("SFM RMSE: %2.2f"%rmse_sfm)
| SFM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generative Adversarial Networks
#
# We’ve talked about how to make predictions. In some form or another, we used deep neural networks learned mappings from data points to labels. This kind of learning is called discriminative learning, as in, we’d like to be able to discriminate between photos cats and photos of dogs. Classifiers and regressors are both examples of discriminative learning. And neural networks trained by backpropagation have upended everything we thought we knew about discriminative learning on large complicated datasets. Classification accuracies on high-res images has gone from useless to human-level (withsome caveats) in just 5-6 years. We’ll spare you another spiel about all the other discriminative tasks where deep neural networks do astoundingly well.
#
# But there’s more to machine learning than just solving discriminative tasks. For example, given a largedataset, without any labels, we might want to learn a model that concisely captures the characteristics of this data. Given such a model, we could sample synthetic data points that resemble the distribution ofthe training data. For example, given a large corpus of photographs of faces, we might want to be able togenerate a new photorealistic image that looks like it might plausibly have come from the same dataset.This kind of learning is called `generative modeling`.
#
# Until recently, we had no method that could synthesize novel photorealistic images. But the success of deep neural networks for discriminative learning opened up new possibilities. One big trend over the last three years has been the application of discriminative deep nets to overcome challenges in problems that we don’t generally think of as supervised learning problems. The recurrent neural network language models are one example of using a discriminative network (trained to predict the next character) that once trained can actas a generative model.
#
# In 2014, a breakthrough paper introduced `Generative adversarial networks (GANs)`, a clever new way to leverage the power of discriminative models to get good generative models. At their heart, GANs rely on the idea that a data generator is good if we cannot tell fake data apart from real data. In statistics, this is calleda two-sample test - a test to answer the question whether datasets $X= {x_1;...;x_n}$ and $X'= {x'_1;...;x'_n}$ were drawn from the same distribution. The main difference between most statistics papers and GANs is that the latter use this idea in a constructive way. In other words, rather than just training a model to say “hey, these two datasets don’t look like they came from the same distribution”, they use the `two-sample test` to provide training signal to a generative model. This allows us to improve the data generator until it generates something that resembles the real data. At the very least, it needs to fool the classifier. And ifour classifier is a state of the art deep neural network.
#
# The GANs architecture is illustrated in `Figure 1`. As you can see, there are two pieces to GANs - first off, we need a device (say, a deep network but it really could be anything, such as a game rendering engine) that might potentially be able to generate data that looks just like the real thing. If we are dealing with images, this needs to generate images. If we’re dealing with speech, it needs to generate audio sequences, and soon. We call this the generator network. The second component is the discriminator network. It attempts to distinguish fake and real data from each other. Both networks are in competition with each other. The generator network attempts to fool the discriminator network. At that point, the discriminator network adapts to the new fake data. This information, in turn is used to improve the generator network, and so on.
#
# <p align="center">
# <img src="images/gan.svg"><br>
# Figure 1. Generative Adversarial Networks
# </p>
#
# The discriminator is a binary classifier to distinguish if the inputxis real (from real data) or fake (from the generator). Typically, the discriminator outputs a scalar prediction $ o \in R $ for input $\mathbf x$, such as using a dense layer with hidden size 1, and then applies sigmoid function to obtain the predicted probability $ D(x) = 1/(1 + e^{-x}) $. Assume the label y for true data is 1 and 0 for fake data. We train the discriminator to minimize the cross entropy loss, i.e $$ \min - y \log D(\mathbf x) - (1-y)\log(1-D(\mathbf x)) $$
#
# For the generator, it first draws some parameter $ z \in R^d $ from a source of randomness, e.g. a normal distribution $ \mathbf z \sim\mathcal(0;1) $. We often call z the `latent variable`. It then applies a function to generate $ \mathbf x′=G(\mathbf z) $. The goal of the generator is to fool the discriminator to classify $x′$ as true data. In other words, we update the parameters of the generator to maximize the cross entropy loss when y= 0, i.e. $$ \max - \log(1-D(\mathbf x'))$$
#
# If the discriminator does a perfect job, then $ D(\mathbf x')\approx 1 $ so the above loss near 0, which results the gradients are too small to make a good progress for the discriminator. So commonly we minimize the following loss $$ \max \log(D(\mathbf x')) $$
#
# which is just feed $\mathbf x' $ into the discriminator but giving label $y=1$.
#
# Many of the GANs applications are in the context of images. As a demonstration purpose, we're going to content ourselves with fitting a much simpler distribution first. We will illustrate what happens if we use GANs to build the world's most inefficient estimator of parameters for a Gaussian. Let's get started.
# ## 1. Import necessary dependencies
# -----------------------------------
# +
# %matplotlib inline
import d2l
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
# -
# ## 2. Generate some “real” data
# -------------------------------
#
# Since this is going to be the world’s lamest example, we simply generate data drawn from a Gaussian.
# +
X = np.random.normal(size=(1000, 2))
X = torch.from_numpy(X).float()
A = torch.tensor([[ 1. , 2. ],
[-0.1, 0.5]])
b = torch.tensor([1., 2.])
data = torch.matmul(X, A) + b
class GANDataset(Dataset):
def __init__(self, data, transforms=None):
self.dataset = data
def __len__(self):
return self.dataset.size(0)
def __getitem__(self, idx):
return self.dataset[idx]
# -
plt.scatter(data[:100, 0].numpy(), data[:100, 1].numpy())
plt.show()
print(f'The covariance matrix is {torch.matmul(A.T,A)}')
# ## 3. Generator
# ---------------
#
# Our generator network will be the simplest network possible - a single layer linear model. This is since we’llbe driving that linear network with a Gaussian data generator. Hence, it literally only needs to learn the parameters to fake things perfectly.
def get_generator(device=None):
net_G = nn.Sequential(
nn.Linear(2, 2)
)
if device is not None:
net_G.to(device)
return net_G
# ## 4. Discriminator
# -------------------
#
# For the discriminator we will be a bit more discriminating: we will use an MLP with 3 layers to make things a bit more interesting.
def get_discriminator(device=None):
net_D = nn.Sequential(
nn.Linear(2, 5),
nn.Tanh(),
nn.Linear(5, 3),
nn.Tanh(),
nn.Linear(3, 1)
)
if device is not None:
net_D.to(device)
return net_D
# ## 5. Training
# --------------
#
# First we define a function to update the discriminator.
def update_D(X, Z, net_D, net_G, loss, optimizer_D):
"""Updates discriminator."""
batch_size = X.size(0)
ones = X.new_ones(size=(batch_size, 1))
zeros = X.new_zeros(size=(batch_size, 1))
# zero the parameter gradients
optimizer_D.zero_grad()
real_Y = net_D(X)
fake_X = net_G(Z)
# Don't need to compute gradient for net_G, detach it from
# computing gradients.
fake_Y = net_D(fake_X.detach())
loss_D = (loss(real_Y, ones) + loss(fake_Y, zeros)) / 2
loss_D.backward()
optimizer_D.step()
return loss_D.sum().item()
# The generator is updated similarly. Here we reuse the cross-entropy loss but change the label of the fake data from $0$ to $1$.
def update_G(Z, net_D, net_G, loss, optimizer_G):
"""Updates generator."""
batch_size = Z.size(0)
ones = Z.new_ones(size=(batch_size, 1))
# zero the parameter gradients
optimizer_G.zero_grad()
# We could reuse fake_X from update_D to save computation.
fake_X = net_G(Z)
# Recomputing fake_Y is needed since net_D is changed.
fake_Y = net_D(fake_X)
loss_G = loss(fake_Y, ones)
loss_G.backward()
optimizer_G.step()
return loss_G.sum().item()
# Both the discriminator and the generator performs a binary logistic regression with the cross-entropy loss. We use Adam to smooth the training process. In each iteration, we first update the discriminator and then the generator. We visualize both losses and generated examples.
def train(net_D, net_G, data_iter, num_epochs, lr_D, lr_G, latent_dim, data):
loss = nn.BCEWithLogitsLoss()
optimizer_D = optim.Adam(net_D.parameters(), lr=lr_D)
optimizer_G = optim.Adam(net_G.parameters(), lr=lr_G)
animator = d2l.Animator(xlabel='epoch', ylabel='loss',
xlim=[1, num_epochs], nrows=2, figsize=(5,5),
legend=['generator', 'discriminator'])
animator.fig.subplots_adjust(hspace=0.3)
for epoch in range(num_epochs):
# Train one epoch
timer = d2l.Timer()
metric = d2l.Accumulator(3) # loss_D, loss_G, num_examples
for _, X in enumerate(data_iter):
batch_size = X.size(0)
X = X.view(-1, latent_dim)
X = X.to(device)
# z latent variables
Z = np.random.normal(0, 1, size=(batch_size, latent_dim))
Z = torch.from_numpy(Z).float()
Z = Z.view(-1, latent_dim)
Z = Z.to(device)
metric.add(update_D(X, Z, net_D, net_G, loss, optimizer_D),
update_G(Z, net_D, net_G, loss, optimizer_G),
batch_size)
# Visualize generated examples
Z = np.random.normal(0, 1, size=(100, latent_dim))
Z = torch.from_numpy(Z).float().to(device)
with torch.no_grad():
fake_X = net_G(Z).cpu()
fake_X = fake_X.numpy()
animator.axes[1].cla()
animator.axes[1].scatter(data[:,0], data[:,1])
animator.axes[1].scatter(fake_X[:,0], fake_X[:,1])
animator.axes[1].legend(['real', 'generated'])
# Show the losses
loss_D, loss_G = metric[0]/metric[2], metric[1]/metric[2]
animator.add(epoch, (loss_D, loss_G))
print(f'loss_D {loss_D:.3f}, loss_G {loss_G:.3f}, {metric[2]/timer.stop()} examples/sec')
# Now we specify the hyper-parameters to fit the Gaussian distribution.
# +
lr_D = 0.05
lr_G = 0.005
latent_dim = 2
num_epochs = 20
trainset = GANDataset(data)
trainloader = DataLoader(trainset, batch_size=2, shuffle=True, num_workers=4)
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
net_G = get_generator(device)
net_D = get_discriminator(device)
train(net_D, net_G, trainloader,
num_epochs, lr_D, lr_G,
latent_dim, data[:100].numpy())
# -
# ## Summary
# ----------
#
# - Generative adversarial networks (GANs) composes of two deep networks, the generator and the discriminator.
# - The generator generates the image as much closer to the true image as possible to fool the discriminator, via maximizing the cross-entropy loss, i.e., $\max \log(D(\mathbf{x'}))$.
# - The discriminator tries to distinguish the generated images from the true images, via minimizing the cross-entropy loss, i.e., $\min - y \log D(\mathbf{x}) - (1-y)\log(1-D(\mathbf{x}))$.
| gan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 5.7: 文字列の書式設定
# +
# リスト 5.7.1 テキストの描画
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
fig, ax = plt.subplots(1, 1)
# 軸範囲の設定
ticks = np.linspace(0, 10, 6)
ax.set_xticks(ticks)
ax.set_yticks(ticks)
# テキストの描画
ax.text(2, 4, "Jupyter")
# 目盛り線描画
ax.grid(linestyle="-")
# +
# リスト 5.7.2 フォントの設定
fig, ax = plt.subplots(1, 1)
string = "Jupyter"
# フォントの書式を個別設定をしてテキストを描画
ax.text(0.2, 0.6, string, alpha=0.5, color="blue", size=40)
# 辞書型でフォントの書式セットを設定してテキスト描画
fontdict = {
"family": "fantasy",
"style": "italic",
"weight": "heavy",
"size": 40,
"color": "red",
}
ax.text(0.2, 0.2, string, alpha=0.5, color="green", fontdict=fontdict)
ticks = np.linspace(0, 1, 6)
ax.set_xticks(ticks)
ax.set_yticks(ticks)
ax.grid(linestyle="-")
# +
# リスト 5.7.3 テキストボックスの塗りつぶし色を設定
fig, ax = plt.subplots(1, 1)
ax.text(0.1, 0.5, "塗りつぶし色の設定", size=25, color="white", backgroundcolor="blue")
# +
# リスト 5.7.4 書式辞書を用いたテキストボックスの書式設定
fig, ax = plt.subplots(1, 1)
# テキストボックスの書式辞書
boxprops = {
"facecolor": "pink",
"edgecolor": "red",
"alpha": 0.5,
"boxstyle": "roundtooth",
"linewidth": 2,
}
# テキストボックスの描画
ax.text(0.1, 0.4, "テキストボックスの\n書式辞書による設定", size=25, bbox=boxprops)
# +
# リスト 5.7.5 テキストボックスの水平方向の配置設定
fig, ax = plt.subplots(1, 1)
# テキストボックスの描画(水平方向の配置)
ax.text(0.5, 3, "left", horizontalalignment="left", size=25)
ax.text(0.5, 2, "center", horizontalalignment="center", size=25)
ax.text(0.5, 1, "right", horizontalalignment="right", size=25)
# 目盛りと罫線の設定
ax.set_xticks(np.linspace(0, 1, 3))
ax.set_yticks(np.linspace(0, 4, 5))
ax.minorticks_on()
ax.grid(which="major", axis="x", color="lightblue", linewidth=2)
ax.grid(which="minor", axis="x", color="grey", linewidth=0.5)
ax.grid(which="major", axis="y", color="grey", linewidth=0.5)
# +
# リスト 5.7.6 テキストボックスの垂直方向の配置設定
fig, ax = plt.subplots(figsize=(12, 2))
# テキストボックスの描画(垂直方向の配置)
ax.text(2, 0.5, "bottom", verticalalignment="bottom", size=20)
ax.text(8, 0.5, "baseline", verticalalignment="baseline", size=20)
ax.text(16, 0.5, "center", verticalalignment="center", size=20)
ax.text(22, 0.5, "center_baseline", verticalalignment="center_baseline", size=20)
ax.text(36, 0.5, "top", verticalalignment="top", size=20)
# 目盛りと罫線の設定
ax.set_xticks(np.linspace(0, 40, 5))
ax.set_yticks(np.linspace(0, 1, 3))
ax.minorticks_on()
ax.grid(which="major", axis="x", color="grey", linewidth=0.5)
ax.grid(which="minor", axis="x", color="grey", linewidth=0.5)
ax.grid(which="major", axis="y", color="lightblue", linewidth=2)
# +
# リスト 5.7.7 複数行文字列の水平方向の配置設定
fig, ax = plt.subplots(1, 1, figsize=(4, 7))
# テキストボックス内文字列の水平方向配置を設定して描画
ax.text(1, 10, "左揃え\nLeft\nalignment", multialignment="left", size=20)
ax.text(1, 6, "中央揃え\nCenter\nalignment", multialignment="center", size=20)
ax.text(1, 2, "右揃え\nRight\nalignment", multialignment="right", size=20)
# 目盛りと罫線の設定
ax.set_xticks(np.linspace(0, 5, 6))
ax.set_yticks(np.linspace(0, 14, 8))
ax.grid()
# +
# リスト 5.7.8 複数行文字列の行間を設定
fig, ax = plt.subplots(1, 1, figsize=(3.5, 5))
# 行間の設定
ax.text(1, 5, "行間 1\nspace 1", linespacing=1, size=20)
ax.text(1, 3, "行間 1.5\nspace 1.5", linespacing=1.5, size=20)
ax.text(1, 1, "行間 2\nspace 2", linespacing=2, size=20)
# 目盛りと罫線の設定
ax.set_xticks(np.linspace(0, 4, 5))
ax.set_yticks(np.linspace(0, 7, 8))
ax.grid()
# +
# リスト 5.7.9 テキストボックスの回転の設定
import itertools
fig, axes = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(8, 8))
# サブプロット1~3の描画
axes[0, 0].text(0.5, 0.5, "horizontal", rotation="horizontal", size=12)
axes[0, 1].text(0.5, 0.5, "vertical", rotation="vertical", size=12)
axes[1, 0].text(
0.5,
0.5,
"rotation=45\nwith anchor\nsetting",
rotation=45,
rotation_mode="anchor",
size=12,
)
# サブプロット4の描画
# 配置比較用テキストボックス描画
boxprops = {"facecolor": "Pink", "edgecolor": "Pink", "alpha": 0.5}
axes[1, 1].text(
0.5,
0.5,
"rotation=45\nwithout anchor\nsetting",
rotation=0,
bbox=boxprops,
color="white",
size=12,
)
# テキスト描画
axes[0, 0].text(0.02, 0.9, "1)", size=16)
axes[0, 1].text(0.02, 0.9, "2)", size=16)
axes[1, 0].text(0.02, 0.9, "3)", size=16)
axes[1, 1].text(0.02, 0.9, "4)", size=16)
# 回転したテキストボックス描画
boxprops = {"facecolor": "None", "edgecolor": "Pink"}
axes[1, 1].text(
0.5,
0.5,
"rotation=45\nwithout anchor\nsetting",
bbox=boxprops,
rotation=45,
size=12,
)
# 目盛りと罫線の設定
axes[0, 0].set_xticks(np.linspace(0, 1, 3))
axes[0, 0].set_yticks(np.linspace(0, 1, 3))
for l, c in itertools.product(range(2), range(2)):
axes[l, c].grid(axis="both", color="lightblue", linewidth=2)
| notebooks/5-07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alu0101046853/TFG_Chesen/blob/main/AutoKeras.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="e-9BoaAAd0lE"
# !pip install git+https://github.com/keras-team/keras-tuner.git
# !pip install autokeras
# + id="0FiF3UbTeiI-"
import numpy as np
import tensorflow as tf
from tensorflow.keras.datasets import mnist
import matplotlib.pyplot as plt
import autokeras as ak
# + id="EIBlgTAaevQ3"
(x_train, y_train), (x_test, y_test) = mnist.load_data()
num = 4
images = x_train[:num]
labels = y_train[:num]
num_row = 2
num_col = 2
fig, axes = plt.subplots(num_row, num_col, figsize=(1.5*num_col,2*num_row))
for i in range(num):
ax = axes[i//num_col, i%num_col]
ax.imshow(images[i], cmap='gray')
ax.set_title('Label: {}'.format(labels[i]))
plt.tight_layout()
plt.show()
# + id="blEOAowge2FL"
clf = ak.ImageClassifier(overwrite=True, max_trials=1)
clf.fit(x_train, y_train, epochs=10)
predicted_y = clf.predict(x_test)
print(predicted_y)
print(clf.evaluate(x_test, y_test))
modelo_final = clf.export_model()
modelo_final.summary()
tf.keras.utils.plot_model(modelo_final,show_shapes=True)
# + id="IE1yllAiMpWM"
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
import pandas as pd
data = load_iris()
X, y = data['data'], pd.get_dummies(data['target']).values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
model = ak.StructuredDataClassifier(overwrite=True, max_trials=50)
model.fit(X_train, y_train)
print("TEST")
loss, acc = model.evaluate(X_test, y_test)
print("Accuracy: ", acc)
y_predictions = model.predict(X_test)
modelo_final = model.export_model()
modelo_final.summary()
tf.keras.utils.plot_model(modelo_final,show_shapes=True)
# + id="HXuq0mBRF2zS"
from sklearn.metrics import confusion_matrix
from mlxtend.plotting import plot_confusion_matrix
matrix = confusion_matrix(y_test.argmax(axis=1), y_predictions.argmax(axis=1))
plot_confusion_matrix(conf_mat=matrix, figsize=(3,3), show_normed=False)
plt.tight_layout()
| AutoKeras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Application 1
import DSGRN
import Berry_2019_figures_results as Berry
from min_interval_posets import posets, poset_distance
from copy import deepcopy
from IPython import display
import matplotlib.pyplot as plt
from importlib import reload
from matplotlib import rc
rc('text', usetex=True)
fontsize=20
rc('axes', labelsize=fontsize) # fontsize of the x and y labels
rc('xtick', labelsize=fontsize) # fontsize of the tick labels
rc('ytick', labelsize=fontsize) # fontsize of the tick labels
rc('legend', fontsize=16) # legend fontsize
# %matplotlib inline
wavepool = DSGRN.Network("good_wavepool.txt")
swapped = DSGRN.Network("bad_wavepool.txt")
DSGRN.DrawGraph(wavepool)
DSGRN.DrawGraph(swapped)
wt1_file = "WT1_WT2_microarray_interpolated/wt1_microarray_coregenes_lifepoints_interpol_trim.csv"
wt2_file = "WT1_WT2_microarray_interpolated/wt2_microarray_coregenes_lifepoints_interpol.csv"
epsilons = [0.0, 0.01,0.04,0.05,0.06,0.08,0.09,0.1,0.14,0.15]
names = ["YOX1","SWI4","HCM1","NDD1"]
posets1 = Berry.getposets(wt1_file,"row",epsilons,names=names)
posets2 = Berry.getposets(wt2_file,"row",epsilons,names=names)
# graph data
def make_fig(fname,savename,start_time=None,end_time=None,names=None):
curves = Berry.row(fname)
subset_curves = deepcopy(curves)
if names is not None:
for name in curves:
if name not in names:
subset_curves.pop(name)
for name,curve in subset_curves.items():
n = curve.normalize()
if start_time is not None and end_time is not None:
n = curve.trim(start_time,end_time)
times,vals = zip(*n.items())
plt.plot(times,vals,label=r"${}$".format(name))
lgd = plt.legend(loc='upper left', bbox_to_anchor=(1, 1))
plt.ylabel(r"\textbf{normalized expression}")
plt.xlabel(r"\textbf{time points}")
plt.savefig(savename,bbox_extra_artists=(lgd,), bbox_inches='tight')
display.display(plt.show())
make_fig(wt1_file,"time_series_rep1.pdf",names=names)
make_fig(wt2_file,"time_series_rep2.pdf",names=names)
def make_posets(p,network):
events = list(p[1][0])
event_ordering = list(p[1][1])
poe = DSGRN.PosetOfExtrema(network, events, event_ordering)
return poe,len(events)
def create_pattern_graph(poset,network):
eps = poset[0]
poe,_ = make_posets(poset,network)
return DSGRN.PatternGraph(poe),eps
def create_search_graph(param):
domain_graph = DSGRN.DomainGraph(param)
return DSGRN.SearchGraph(domain_graph)
def get_matches(posets,network):
param_matches = {}
parameter_graph = DSGRN.ParameterGraph(network)
for poset in posets:
pattern_graph,eps = create_pattern_graph(poset,network)
param_matches[eps] = []
for pind in range(parameter_graph.size()):
parameter = parameter_graph.parameter(pind)
search_graph = create_search_graph(parameter)
matching_graph = DSGRN.MatchingGraph(search_graph, pattern_graph)
path_match = DSGRN.PathMatch(matching_graph)
if path_match:
param_matches[eps].append(pind)
return param_matches
# +
wavepool1_parameter_matches=get_matches(posets1,wavepool)
wavepool2_parameter_matches=get_matches(posets2,wavepool)
swapped11_parameter_matches=get_matches(posets1,swapped)
swapped12_parameter_matches=get_matches(posets2,swapped)
for eps in epsilons:
print("Epsilon = {}".format(eps))
print("Number of matches in rep 1, wavepool: {}".format(len(wavepool1_parameter_matches[eps])))
print("Number of matches in rep 2, wavepool: {}".format(len(wavepool2_parameter_matches[eps])))
print("Number of matches in rep 1, swapped: {}".format(len(swapped11_parameter_matches[eps])))
print("Number of matches in rep 2, swapped: {}".format(len(swapped12_parameter_matches[eps])))
# -
print(wavepool1_parameter_matches[0.04])
print(wavepool2_parameter_matches[0.04])
for (p,q) in zip(posets1,posets2):
print("Number of extrema is {},{} for replicates 1,2 at epsilon {}".format(len(p[1][0]),len(q[1][0]),p[0]))
# example poset between reps 1 and 2
# notice that even with the same number of extrema, the identity of the nodes vary
print("Replicate 1, eps 0.15")
poe, N = make_posets(posets1[-1],wavepool)
display.display(DSGRN.DrawGraph(poe))
with open("example_poset.dot","w") as f:
f.write(poe.graphviz())
print("Replicate 2, eps 0.15")
poe, N = make_posets(posets2[-1],wavepool)
display.display(DSGRN.DrawGraph(poe))
print("Replicate 1, eps 0.0")
poe, N = make_posets(posets1[0],wavepool)
display.display(DSGRN.DrawGraph(poe))
with open("example_poset0.dot","w") as f:
f.write(poe.graphviz())
print("Replicate 1, eps 0.01")
poe, N = make_posets(posets1[1],wavepool)
display.display(DSGRN.DrawGraph(poe))
print("Replicate 1, eps 0.04")
poe, N = make_posets(posets1[2],wavepool)
display.display(DSGRN.DrawGraph(poe))
# example pattern graph
pattern_graph,_ = create_pattern_graph(posets1[-1],wavepool)
with open("example_pattern_graph.dot","w") as f:
f.write(pattern_graph.graphviz())
DSGRN.DrawGraph(pattern_graph)
# example search graph
pind = wavepool1_parameter_matches[0.15][0]
parameter_graph = DSGRN.ParameterGraph(wavepool)
param = parameter_graph.parameter(pind)
search_graph = create_search_graph(param)
display.display(DSGRN.DrawGraph(search_graph))
with open("example_search_graph.dot","w") as f:
f.write(search_graph.graphviz())
# example matching graph
matching_graph = DSGRN.MatchingGraph(search_graph, pattern_graph)
path_match = DSGRN.PathMatch(matching_graph)
with open("example_matching_graph.dot","w") as f:
f.write(matching_graph.graphviz_with_highlighted_path(path_match))
DSGRN.DrawGraphWithHighlightedPath(matching_graph, path_match)
| scripts/Berry_2019_figures_results_Application1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nimitsolanki/Cricket-World-Cup-2019/blob/master/Cricket_World_Cup_2019.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="a4GuQITThGrn" colab_type="text"
# **CRICKET WORLD CUP 2019 - ICC CWC 2019**
#
# Venue : ENGLAND
#
# Total Teams : 10
#
#
# Dataset - ODI_DATASET.csv
#
# Starting from Jan-2013 to mid May-2019 - all the ODI results are included in this dataset
#
#
#
# + id="o0_Ll3Q4c-pF" colab_type="code" colab={}
import warnings
warnings.filterwarnings('ignore')
# linear algebra
import numpy as np
# data processing
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + id="m4HbipkidXfA" colab_type="code" colab={}
# ODI dataset from Jan-2013 to May-2019
# As I have used Google Colab to perform this notebook - hence added github raw URL to fetch the dataset
ODI_Data = pd.read_csv('https://raw.githubusercontent.com/nimitsolanki/Cricket-World-Cup-2019/master/data/ODI_DATASET.csv')
# + id="iqsp_GBfdp8V" colab_type="code" colab={}
# Scores_ID
ODI_Data["Scores_ID"] = ODI_Data["Unnamed: 0"]
ODI_Data.drop(columns="Unnamed: 0",inplace=True)
# + id="mOAMYy3mdtbn" colab_type="code" colab={}
# CWC-2019 pitches
WC_venue_pitches = ["The Oval, London","Trent Bridge, Nottingham","Sophia Gardens, Cardiff","County Ground, Bristol","Rose Bowl, Southampton","County Ground, Taunton","Old Trafford, Manchester","Edgbaston, Birmingham","Headingley, Leeds","Lord's, London","Riverside Ground, Chester-le-Street"]
#Total Grounds
WC_Ground_Stats = []
ODI_Grounds = ODI_Data.Ground
for i in ODI_Grounds:
for j in WC_venue_pitches:
if i in j:
WC_Ground_Stats.append((i,j))
# + id="WAc3I1HDd8W8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="823e1929-f02b-4747-9ba4-f8d12f59b3c4"
# Listing ground names
Ground_names = dict(set(WC_Ground_Stats))
def Full_Ground_names(value):
return Ground_names[value]
Ground_names
# + id="J0IkLYqDeA5l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="5718f266-40ae-422f-bcf6-79c6cde72587"
# Matching ODI's data with respect to the above listed grounds
WC_Grounds_History = ODI_Data[ODI_Data.Ground.isin([Ground[0] for Ground in WC_Ground_Stats])]
WC_Grounds_History["Ground"] = WC_Grounds_History.Ground.apply(Full_Ground_names)
WC_Grounds_History.head()
# + id="1tO53M8PeUB_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 576} outputId="f5e101ba-4eb7-4114-a46e-4b7c80acfb22"
# Finding the World cup team's played on these grounds
Team_Matches = WC_Grounds_History.Country.value_counts().reset_index()
plt.figure(figsize=(15,8))
sns.barplot(x = "index", y = "Country", data = Team_Matches).set_title("Total Matches Played by each Country")
plt.xlabel("Country")
plt.ylabel("Matches Played")
plt.xticks(rotation = 60)
# + id="RHDaJzGdfBv0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="74984497-f2bb-4af6-efa9-e9ca913d8d22"
# Team wise Winning Percentage in England Pitches after removing the currupt data result
WC_Grounds_History = WC_Grounds_History[~WC_Grounds_History.Result.isin(["-"])]
WC_Grounds_History.Result.value_counts()
# + id="rdCUT313fE1p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="19e89e71-fdf7-42b4-e5e5-154d7f4ebb2e"
# Country results in percentage
winnings = WC_Grounds_History[["Country","Result"]]
winnings["count"] = 1
Ground_Results_Per_Team = winnings.groupby(["Country","Result"]).aggregate(["sum"])
Ground_Results_Per_Team = Ground_Results_Per_Team.groupby(level=0).apply(lambda x:100 * x / float(x.sum())).reset_index()
Ground_Results_Per_Team.columns = ["Country","Result","Count"]
Ground_Results_Per_Team.head()
# + id="7xvKqHeNfHch" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 576} outputId="e4d202ba-017b-4160-83da-cebab10e268a"
# Plotting Results in percentage
plt.figure(figsize=(15,8))
sns.barplot(x = "Country", y = "Count", hue = "Result", data = Ground_Results_Per_Team)
plt.ylabel("Percentage")
plt.title("Country - Results")
plt.xticks(rotation = 60)
# + [markdown] id="VHm-Wc0pfbue" colab_type="text"
# It's clear that India and England have highest Green bars - highest winning percentage.
#
# Let's see what happens when the Top Two Teams face?
# + id="1WZsj6pyffBn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="d88785e4-d8b1-410e-c5da-54e2912d9ca4"
# It's WC final time - India Vs England
India_vs_England = WC_Grounds_History[WC_Grounds_History.Country == "India"]\
[WC_Grounds_History.Opposition.str.contains("England")]
India_vs_England = India_vs_England.Result.value_counts().reset_index()
sns.barplot(x = "index", y = "Result", data = India_vs_England).set_title("India against England")
plt.xlabel("India")
# + [markdown] id="jNfnoCDXkCnz" colab_type="text"
# **Prediction : India will win ICC Cricket World Cup 2019**
| Cricket_World_Cup_2019.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ## GeostatsPy: Basic Univariate Distribution Transformations Subsurface Data Analytics in Python
#
#
# ### <NAME>, Associate Professor, University of Texas at Austin
#
# #### [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
# ### PGE 383 Exercise: Basic Univariate Distribution Transformations Subsurface Data Analytics in Python with GeostatsPy
#
# Here's a simple workflow with some basic univariate distribution transformations for subsurface modeling workflows. This should help you get started data transformations.
#
# #### Data Distribution Transformations
#
# Why?:
# * variable has expected shape / correcting for too few data
# * a specific distribution assumption is required
# * correct for outliers
#
# How?:
#
# There are a variety of transformations. In general we are transforming the values from the cumulative distribution function (CDF), $F_{X}$, to a new CDF , $G_{Y}$. This can be generalized with the quantile - quantile transformation applied to all the sample data:
#
# * The forward transform:
#
# \begin{equation}
# Y = G_{Y}^{-1}(F_{X}(X))
# \end{equation}
#
# * The reverse transform:
#
# \begin{equation}
# X = F_{X}^{-1}(G_{Y}(Y))
# \end{equation}
#
# This may be applied to any data, nonparametric or samples from a parametric distribution. We just need to be able to map from one distribution to another through percentiles, so it is a:
#
# * Rank preserving transform
#
# We will cover three examples including:
#
# 1. Distribution rescaling
# 2. Normal score transform
#
# #### Objective
#
# In the PGE 383: Stochastic Subsurface Modeling class I want to provide hands-on experience with building subsurface modeling workflows. Python provides an excellent vehicle to accomplish this. I have coded a package called GeostatsPy with GSLIB: Geostatistical Library (Deutsch and Journel, 1998) functionality that provides basic building blocks for building subsurface modeling workflows.
#
# The objective is to remove the hurdles of subsurface modeling workflow construction by providing building blocks and sufficient examples. This is not a coding class per se, but we need the ability to 'script' workflows working with numerical methods.
#
# #### Getting Started
#
# Here's the steps to get setup in Python with the GeostatsPy package:
#
# 1. Install Anaconda 3 on your machine (https://www.anaconda.com/download/).
# 2. From Anaconda Navigator (within Anaconda3 group), go to the environment tab, click on base (root) green arrow and open a terminal.
# 3. In the terminal type: pip install geostatspy.
# 4. Open Jupyter and in the top block get started by copy and pasting the code block below from this Jupyter Notebook to start using the geostatspy functionality.
#
# You will need to copy the data file to your working directory. They are available here:
#
# * Tabular data - sample_data_biased.csv at https://git.io/fh0CW
#
# There are exampled below with these functions. You can go here to see a list of the available functions, https://git.io/fh4eX, other example workflows and source code.
import geostatspy.GSLIB as GSLIB # GSLIB utilies, visualization and wrapper
import geostatspy.geostats as geostats # GSLIB methods convert to Python
# We will also need some standard packages. These should have been installed with Anaconda 3.
import numpy as np # ndarrys for gridded data
import pandas as pd # DataFrames for tabular data
import os # set working directory, run executables
import matplotlib.pyplot as plt # for plotting
from scipy import stats # summary statistics
# #### Set the working directory
#
# I always like to do this so I don't lose files and to simplify subsequent read and writes (avoid including the full address each time).
os.chdir("c:/PGE383/Examples") # set the working directory
# #### Loading Tabular Data
#
# Here's the command to load our comma delimited data file in to a Pandas' DataFrame object.
df = pd.read_csv('sample_data_biased.csv') # load our data table (wrong name!)
# It worked, we loaded our file into our DataFrame called 'df'. But how do you really know that it worked? Visualizing the DataFrame would be useful and we already leard about these methods in this demo (https://git.io/fNgRW).
#
# We can preview the DataFrame by printing a slice or by utilizing the 'head' DataFrame member function (with a nice and clean format, see below). With the slice we could look at any subset of the data table and with the head command, add parameter 'n=13' to see the first 13 rows of the dataset.
print(df.iloc[0:5,:]) # display first 4 samples in the table as a preview
df.head(n=13) # we could also use this command for a table preview
# #### Summary Statistics for Tabular Data
#
# The table includes X and Y coordinates (meters), Facies 1 and 2 (1 is sandstone and 0 interbedded sand and mudstone), Porosity (fraction), and permeability as Perm (mDarcy).
#
# There are a lot of efficient methods to calculate summary statistics from tabular data in DataFrames. The describe command provides count, mean, minimum, maximum, and quartiles all in a nice data table. We use transpose just to flip the table so that features are on the rows and the statistics are on the columns.
df.describe().transpose()
# #### Visualizing Tabular Data with Location Maps
#
# It is natural to set the x and y coordinate and feature ranges manually. e.g. do you want your color bar to go from 0.05887 to 0.24230 exactly? Also, let's pick a color map for display. I heard that plasma is known to be friendly to the color blind as the color and intensity vary together (hope I got that right, it was an interesting Twitter conversation started by <NAME> from Agile if I recall correctly). We will assume a study area of 0 to 1,000m in x and y and omit any data outside this area.
xmin = 0.0; xmax = 1000.0 # range of x values
ymin = 0.0; ymax = 1000.0 # range of y values
pormin = 0.05; pormax = 0.25; # range of porosity values
cmap = plt.cm.plasma # color map
# Let's try out locmap. This is a reimplementation of GSLIB's locmap program that uses matplotlib. I hope you find it simpler than matplotlib, if you want to get more advanced and build custom plots lock at the source. If you improve it, send me the new code. Any help is appreciated. To see the parameters, just type the command name:
GSLIB.locmap
# Now we can populate the plotting parameters and visualize the porosity data.
GSLIB.locmap(df,'X','Y','Porosity',xmin,xmax,ymin,ymax,pormin,pormax,'Well Data - Porosity','X(m)','Y(m)','Porosity (fraction)',cmap,'locmap_Porosity')
# #### Decluster the Data
#
# Look carefully, and you'll notice the the spatial samples are more dense in the high porosity regions and less dense in the low porosity regions. There is preferential sampling. We cannot use the naive statistics to represent this region. We have to correct for the clustering of the samples in the high porosity regions.
#
# Let's try cell declustering. We can interpret that we will want to minimize the declustering mean and that a cell size of between 100 - 200m is likely a good cell size, this is 'an ocular' estimate of the largest average spacing in the sparsely sampled regions.
#
# Let's check out the declus program reimplimented from GSLIB.
geostats.declus
# We can now populate the parameters. We will run a very wide range of cell sizes, from 10m to 2,000m ('cmin' and 'cmax') and take the cell size that minimizes the declustered mean ('iminmax' = 1 minimize, and = 0 maximize). Multiple offsets (number of these is 'noff') uses multiple grid origins and averages the results to remove sensitivity to grid position. The ncell is the number of cell sizes.
#
# The output from this program is:
#
# * wts - an array with the weigths for each data (they sum to the number of data, 1 indicates nominal weight)
# * cell_sizes - an array with the considered cell sizes
# * dmeans - de an
wts, cell_sizes, dmeans = geostats.declus(df,'X','Y','Porosity',iminmax = 1, noff= 10, ncell=100,cmin=10,cmax=2000)
df['Wts'] = wts # add weights to the sample data DataFrame
df.head() # preview to check the sample data DataFrame
# Let's look at the location map of the weights.
GSLIB.locmap(df,'X','Y','Wts',xmin,xmax,ymin,ymax,0.5,2.5,'Well Data Weights','X(m)','Y(m)','Weights',cmap,'locmap_Weights')
# Does it look correct? See the weight varies with local sampling density?
#
# Now let's add the distribution of the weights and the naive and declustered porosity distributions. You should see the histogram bars adjusted by the weights. Also note the change in the mean due to the weights. There is a significant change.
# +
plt.subplot(121)
GSLIB.locmap_st(df,'X','Y','Porosity',xmin,xmax,ymin,ymax,pormin,pormax,'Well Data - Porosity','X(m)','Y(m)','Porosity (fraction)',cmap)
plt.subplot(122)
GSLIB.hist_st(df['Porosity'],0.05,0.25,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity",title="Declustered Porosity")
plt.ylim(0.0,40)
plt.subplots_adjust(left=0.0, bottom=0.0, right=3.0, top=1.5, wspace=0.2, hspace=0.2)
plt.show()
# -
# We are now ready to do some data transformations.
#
# #### Distribution Rescaling
#
# Distribution rescaling can be thought of as shifting, and stretching and squeezing a distribution. The common method is known as affine correction:
#
# \begin{equation}
# y = \frac{\sigma_y}{\sigma_x}(y - \overline{y}) + \overline{x}
# \end{equation}
#
# We can see that the metho first centers the distribution, the rescales the dispersion based on the ratio of the new standard deviation to the original standard deviation and then shifts the distribution to centered on the target mean.
#
# We have a function in GeostatsPy to do the affine correction of the distribution.
GSLIB.affine
# We just need to specify the new target mean and variance. Let's make 2 new rescaled distributions and then plot the results.
# +
por_original = df['Porosity'].values # extract the pororsity data as a ndarray
por1 = GSLIB.affine(por_original,0.0,1.0) # rescale the porosity to have a standard distribution
df['standPor'] = por1
plt.subplot(221)
GSLIB.hist_st(df['Porosity'],0.05,0.25,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity (fraction)",title="Declustered Porosity")
plt.ylim(0.0,40)
plt.subplot(222)
GSLIB.hist_st(df['standPor'],-3.0,3.0,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity Standardized",title="Declustered Porosity Standardized")
plt.ylim(0.0,40)
plt.subplot(223)
GSLIB.locmap_st(df,'X','Y','Porosity',xmin,xmax,ymin,ymax,pormin,pormax,'Well Data - Porosity','X(m)','Y(m)','Porosity (fraction)',cmap)
plt.subplot(224)
GSLIB.locmap_st(df,'X','Y','standPor',xmin,xmax,ymin,ymax,-3,3,'Well Data - Porosity - Standardized','X(m)','Y(m)','Standardized Porosity',cmap)
plt.subplots_adjust(left=0.0, bottom=0.0, right=3.0, top=2.5, wspace=0.2, hspace=0.2)
plt.show()
# -
# Notice that I did not say standard normal? A standard distribution has a mean of 0.0 and standard deviation of 1.0. The rescaling does not change the distribution shape; therefore, a non-normal (non-Gaussian) distribution cannot become normal just by rescaling. We'll cover that method in a bit.
#
# Also, notice that the shape is the same and the location maps look exactly the same? By adjusting the minimum and maximum values in the histogram x-axis and the location map color bar, we made them look unchanged! There are minor differences in bars due to the precise locations of the bin boundaries.
#
# Let's try a minor adjustment as in the case of correcting the porosity from well logs to a more reliable mean and standard deviation or in the case to produce multiple scenarios of the porosity distribution (more on these uncertainty methods later).
# +
por_original = df['Porosity'].values # extract the pororsity data as a ndarray
por2 = GSLIB.affine(por_original,0.11,0.02) # rescale the porosity to have a standard distribution
df['adjustedPor'] = por2
plt.subplot(221)
GSLIB.hist_st(df['Porosity'],0.05,0.25,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity (fraction)",title="Declustered Porosity")
plt.ylim(0.0,60)
plt.subplot(222)
GSLIB.hist_st(df['adjustedPor'],0.05,0.25,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity Corrected",title="Declustered Porosity Corrected")
plt.ylim(0.0,60)
plt.subplot(223)
GSLIB.locmap_st(df,'X','Y','Porosity',xmin,xmax,ymin,ymax,pormin,pormax,'Well Data - Porosity','X(m)','Y(m)','Porosity (fraction)',cmap)
plt.subplot(224)
GSLIB.locmap_st(df,'X','Y','adjustedPor',xmin,xmax,ymin,ymax,0.05,0.25,'Well Data - Porosity - Corrected','X(m)','Y(m)','Corrected Porosity',cmap)
plt.subplots_adjust(left=0.0, bottom=0.0, right=3.0, top=2.5, wspace=0.2, hspace=0.2)
plt.show()
# -
# #### Normal Score Transform / Gaussian Anamorphosis
#
# We showed that the correction of the mean to 0.0 and standard deviation to 1.0 with affine correction does not change the shape; therefore, does not make a Gaussian distributed property. For many statistic / geostatistical methods the assumption of Gaussian distributed is required. We need normal score transforms in many subsurface modeling workflows.
#
# Let's check out the GSLIB NSCORE program translated to Python in GeostatsPy.
geostats.nscore
# The inputs are primarily the DataFrame, the variable and the data weight columns ('df', 'vcol' and 'wcol'). The remainder of the variables are for the use of a reference distribution. When would you use a reference distribution? This would be the case when you have too few data to perform a reliable transformation and use analog information to inform a more complete distribution to support the transformation.
#
# As you can see the inputs from weights column ('wcol') have defaults of 0. You can run the function omitting these (e.g. just DataFrame and variable column etc.).
#
# The output form the program include the transformed data, and the trasformation table (discretized values in original and associated Gaussian space).
# +
ns_por,trans_vr,trans_ns = geostats.nscore(df,'Porosity','Wts')
df['NPor'] = ns_por
plt.subplot(221)
GSLIB.hist_st(df['Porosity'],0.05,0.25,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Porosity (fraction)",title="Declustered Porosity")
plt.ylim(0.0,40)
plt.subplot(222)
GSLIB.hist_st(df['NPor'],-3.0,3.0,log=False,cumul=False,bins=30,weights=df['Wts'],xlabel="Normal Scores Porosity",title="Declustered Normal Scores Porosity")
plt.ylim(0.0,40)
plt.subplot(223)
GSLIB.locmap_st(df,'X','Y','Porosity',xmin,xmax,ymin,ymax,pormin,pormax,'Well Data - Porosity','X(m)','Y(m)','Porosity (fraction)',cmap)
plt.subplot(224)
GSLIB.locmap_st(df,'X','Y','NPor',xmin,xmax,ymin,ymax,-3.0,3.0,'Well Data - Normal Scores Porosity','X(m)','Y(m)','Normal Scores Porosity',cmap)
plt.subplots_adjust(left=0.0, bottom=0.0, right=3.0, top=2.5, wspace=0.2, hspace=0.2)
plt.show()
# -
# That is interesting! Why is the new distribution not perfectly Gaussian in shape? Because it is the declustered distribution of the data transformed to Gaussian. It accounts for the spatial bias in the sampling.
#
# I'm not completely satified with the behavoir of the transformed data distribution at the tails. The NSCORE programs does not have any tail extrapolation model as found with simulation methods. The transform at the tails is hard to do just from the data alone. When we get into simulation methods we'll check that out.
#
# We should also visualize the transformation table.
# +
plt.subplot(111)
plt.scatter(trans_vr,trans_ns, c = "black", marker='o', alpha = 0.2, edgecolors = "none")
plt.xlabel('Porosity (%)')
plt.ylabel('Normal Score Transformed Porosity')
plt.title('Normal Score Transformed Porosity vs Untransformed Porosity p-p Plot')
plt.ylim(-4,4)
plt.xlim(0,.30)
plt.subplots_adjust(left=0.0, bottom=0.0, right=1.0, top=1.0, wspace=0.2, hspace=0.2)
plt.show()
# -
# This is a q-q plot that maps the transform from our original distribution to the Gaussian distribution. Notice how the declustering weights have shift up the lowe quantiles as they received more weight.
#
# As a final step we should check out the summary statistics of all the variants of porosity from our various data transformations.
df.describe().transpose()
# #### Comments
#
# This was a basic demonstration of data transformations. Much more could be done, I have other demonstrations on the basics of working with DataFrames, ndarrays, univariate statistics, plotting data, declustering and many other workflows available at https://github.com/GeostatsGuy/PythonNumericalDemos and https://github.com/GeostatsGuy/GeostatsPy.
#
# I hope this was helpful,
#
# *Michael*
#
# <NAME>, Ph.D., P.Eng. Associate Professor The Hildebrand Department of Petroleum and Geosystems Engineering, Bureau of Economic Geology, The Jackson School of Geosciences, The University of Texas at Austin
#
# #### More Resources Available at: [Twitter](https://twitter.com/geostatsguy) | [GitHub](https://github.com/GeostatsGuy) | [Website](http://michaelpyrcz.com) | [GoogleScholar](https://scholar.google.com/citations?user=QVZ20eQAAAAJ&hl=en&oi=ao) | [Book](https://www.amazon.com/Geostatistical-Reservoir-Modeling-Michael-Pyrcz/dp/0199731446) | [YouTube](https://www.youtube.com/channel/UCLqEr-xV-ceHdXXXrTId5ig) | [LinkedIn](https://www.linkedin.com/in/michael-pyrcz-61a648a1)
#
| GeostatsPy_transformations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../../../images/qiskit_header.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" align="middle">
# + [markdown] slideshow={"slide_type": "slide"}
# # Qiskit Aer: Building Noise Models
#
# The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorial.
# -
# ## Introduction
#
# This notebook introduces how to use the Qiskit Aer `noise` module to build custom noise models for noisy simulations.
# +
import numpy as np
from qiskit import execute, QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit.quantum_info.operators import Kraus, SuperOp
from qiskit.providers.aer import QasmSimulator
from qiskit.tools.visualization import plot_histogram
# Qiskit Aer noise module imports
from qiskit.providers.aer.noise import NoiseModel
from qiskit.providers.aer.noise.errors import QuantumError, ReadoutError
from qiskit.providers.aer.noise.errors import pauli_error
from qiskit.providers.aer.noise.errors import depolarizing_error
from qiskit.providers.aer.noise.errors import thermal_relaxation_error
# + [markdown] slideshow={"slide_type": "slide"}
# ## Qiskit Aer Noise Module
#
# The `aer.noise` module contains Python classes to build customized noise models for simulation. There are three key classes:
#
# 1. The `NoiseModel` class which stores a noise model used for noisy simulation
# 2. The `QuantumError` class which describes CPTP gate errors. These can be applied
# * After *gate* or *reset* instructions
# * Before *measure* instructions.
#
# 3. The `ReadoutError` class which describes classical readout errors.
# -
# ## Quantum Errors
#
# Rather than deal with the `QuantumError` object directly, many helper functions exist to automatically generate a specific type of parameterized quantum error. These are contained in the `noise.errors` submodule and include functions for many common errors types used in quantum computing research. The function names and the type of error they return are:
#
# | Standard error function | Details |
# | --- | --- |
# | `kraus_error` | a general n-qubit CPTP error channel given as a list of Kraus matrices $[K_0, ...]$. |
# | `mixed_unitary_error` | an n-qubit mixed unitary error given as a list of unitary matrices and probabilities $[(U_0, p_0),...]$. |
# | `coherent_unitary_error` | an n-qubit coherent unitary error given as a single unitary matrix $U$. |
# | `pauli_error` | an n-qubit Pauli error channel (mixed unitary) given as a list of Pauli's and probabilities $[(P_0, p_0),...]$ |
# | `depolarizing_error` | an n-qubit depolarizing error channel parameterized by a depolarization probability $p$. |
# | `reset_error` | a single-qubit reset error parameterized by a probabilities $p_0, p_1$ of reseting to the $|0\rangle$, $|1\rangle$ state.|
# | `thermal_relaxation_error` | a single qubit thermal relaxation channel parameterized by relaxation time constants $T_1$, $T_2$, gate time $t$, and excited state thermal population $p_1$. |
# | `phase_amplitude_damping_error` | A single-qubit generalized combined phase and amplitude damping error channel given by an amplitude damping parameter $\lambda$, a phase damping parameter $\gamma$, and an excited state thermal population $p_1$. |
# | `amplitude_damping_error` | A single-qubit generalized amplitude damping error channel given by an amplitude damping parameter $\lambda$, and an excited state thermal population $p_1$. |
# | `phase_damping_error` | A single-qubit phase damping error channel given by a phase damping parameter $\gamma$ |
#
# ### Combining quantum errors
#
# `QuantumError` instances can be combined by using composition, tensor product, and tensor expansion (reversed order tensor product) to produce new `QuantumErrors` as:
#
# * Composition: $\cal{E}(\rho)=\cal{E_2}(\cal{E_1}(\rho))$ as `error = error1.compose(error2)`
# * Tensor product: $\cal{E}(\rho) =(\cal{E_1}\otimes\cal{E_2})(\rho)$ as `error error1.tensor(error2)`
# * Expand product: $\cal{E}(\rho) =(\cal{E_2}\otimes\cal{E_1})(\rho)$ as `error error1.expand(error2)`
# ### Example
#
# For example to construct a 5% single-qubit Bit-flip error:
# Construct a 1-qubit bit-flip and phase-flip errors
p_error = 0.05
bit_flip = pauli_error([('X', p_error), ('I', 1 - p_error)])
phase_flip = pauli_error([('Z', p_error), ('I', 1 - p_error)])
print(bit_flip)
print(phase_flip)
# Compose two bit-flip and phase-flip errors
bitphase_flip = bit_flip.compose(phase_flip)
print(bitphase_flip)
# Tensor product two bit-flip and phase-flip errors with
# bit-flip on qubit-0, phase-flip on qubit-1
error2 = phase_flip.tensor(bit_flip)
print(error2)
# ### Converting to and from QuantumChannel operators
#
# We can also convert back and forth between `QuantumError` objects in Qiskit-Aer and `QuantumChannel` objects in Qiskit-Terra
# Convert to Kraus operator
bit_flip_kraus = Kraus(bit_flip)
print(bit_flip_kraus)
# Convert to Superoperator
phase_flip_sop = SuperOp(phase_flip)
print(phase_flip_sop)
# +
# Convert back to a quantum error
print(QuantumError(bit_flip_kraus))
# Check conversion is equivalent to original error
QuantumError(bit_flip_kraus) == bit_flip
# -
# ### Readout Error
#
# Classical readout errors are specified by a list of assignment probabilities vectors $P(A|B)$:
#
# * $A$ is the *recorded* classical bit value
# * $B$ is the *true* bit value returned from the measurement
#
# E.g. for 1 qubits: $ P(A|B) = [P(A|0), P(A|1)]$
# +
# Measurement miss-assignement probabilities
p0given1 = 0.1
p1given0 = 0.05
ReadoutError([[1 - p1given0, p1given0], [p0given1, 1 - p0given1]])
# -
# Readout errors may also be combined using `compose`, `tensor` and `expand` like with quantum errors
# ## Adding errors to a Noise Model
#
# When adding a quantum error to a noise model we must specify the type of *instruction* that it acts on, and what qubits to apply it to. There are three cases for Quantum Errors:
#
# 1. All-qubit quantum error
# 2. Specific qubit quantum error
# 3. Non-local quantum error
#
# ### All-qubit quantum error
#
# This applies the same error to any occurrence of an instruction, regardless of which qubits it acts on.
#
# It is added as `noise_model.add_all_qubit_quantum_error(error, instructions)`:
# +
# Create an empty noise model
noise_model = NoiseModel()
# Add depolarizing error to all single qubit u1, u2, u3 gates
error = depolarizing_error(0.05, 1)
noise_model.add_all_qubit_quantum_error(error, ['u1', 'u2', 'u3'])
# Print noise model info
print(noise_model)
# -
# ### Specific qubit quantum error
#
# This applies the error to any occurrence of an instruction acting on a specified list of qubits. Note that the order of the qubit matters: For a 2-qubit gate an error applied to qubits [0, 1] is different to one applied to qubits [1, 0] for example.
#
# It is added as `noise_model.add_quantum_error(error, instructions, qubits)`:
# +
# Create an empty noise model
noise_model = NoiseModel()
# Add depolarizing error to all single qubit u1, u2, u3 gates on qubit 0 only
error = depolarizing_error(0.05, 1)
noise_model.add_quantum_error(error, ['u1', 'u2', 'u3'], [0])
# Print noise model info
print(noise_model)
# -
# ### Non-local qubit quantum error
#
# This applies an error to a specific set of noise qubits after any occurrence of an instruction acting on a specific of gate qubits.
#
# It is added as `noise_model.add_quantum_error(error, instructions, instr_qubits, error_qubits)`:
# +
# Create an empty noise model
noise_model = NoiseModel()
# Add depolarizing error on qubit 2 forall single qubit u1, u2, u3 gates on qubit 0
error = depolarizing_error(0.05, 1)
noise_model.add_nonlocal_quantum_error(error, ['u1', 'u2', 'u3'], [0], [2])
# Print noise model info
print(noise_model)
# -
# ### Executing a noisy simulation with a noise model
#
# * To execute a noisy simulation we pass the noise model object to `QasmSimulator.run` or `execute` using the `noise_model` kwarg.
# * Eg: `qiskit.execute(circuits, QasmSimulator(), noise_model=noise)`
#
# **Important:** *When running a noisy simulation make sure you compile your qobj to the same basis gates as the noise model!*
#
# This can be done using `NoiseModel.basis_gates`
# + [markdown] slideshow={"slide_type": "subslide"}
# # Noise Model Examples
#
# We will now give some examples of noise models For our demonstrations we wil use a simple test circuit generating a n-qubit GHZ state:
# + slideshow={"slide_type": "fragment"}
# Simulator
simulator = QasmSimulator()
# System Specification
n_qubits = 4
qr = QuantumRegister(n_qubits)
cr = ClassicalRegister(n_qubits)
circ = QuantumCircuit(qr, cr)
# Test Circuit
circ.h(qr[0])
for qubit in range(n_qubits - 1):
circ.cx(qr[qubit], qr[qubit + 1])
circ.measure(qr, cr)
print(circ)
# -
# ### Ideal Simulation
# Ideal execution
job = execute(circ, simulator)
result_ideal = job.result()
plot_histogram(result_ideal.get_counts(0))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Noise Example 1: Basic bit-flip error noise model
#
# Lets consider a simple toy noise model example common in quantum information theory research:
#
# * When applying a single qubit gate, flip the state of the qubit with probability `p_gate1`.
# * When applying a 2-qubit gate apply single-qubit errors to each qubit.
# * When reseting a qubit reset to 1 instead of 0 with probability `p_reset`
# * When measuring a qubit, flip the state of the qubit before with probability `p_meas`.
# +
# Example error probabilities
p_reset = 0.03
p_meas = 0.1
p_gate1 = 0.05
# QuantumError objects
error_reset = pauli_error([('X', p_reset), ('I', 1 - p_reset)])
error_meas = pauli_error([('X',p_meas), ('I', 1 - p_meas)])
error_gate1 = pauli_error([('X',p_gate1), ('I', 1 - p_gate1)])
error_gate2 = error_gate1.tensor(error_gate1)
# Add errors to noise model
noise_bit_flip = NoiseModel()
noise_bit_flip.add_all_qubit_quantum_error(error_reset, "reset")
noise_bit_flip.add_all_qubit_quantum_error(error_meas, "measure")
noise_bit_flip.add_all_qubit_quantum_error(error_gate1, ["u1", "u2", "u3"])
noise_bit_flip.add_all_qubit_quantum_error(error_gate2, ["cx"])
print(noise_bit_flip)
# -
# ### Executing the noisy simulation
# + slideshow={"slide_type": "-"}
# Run the noisy simulation
job = execute(circ, simulator,
basis_gates=noise_bit_flip.basis_gates,
noise_model=noise_bit_flip)
result_bit_flip = job.result()
counts_bit_flip = result_bit_flip.get_counts(0)
# Plot noisy output
plot_histogram(counts_bit_flip)
# -
# ## Example 2: T1/T2 thermal relaxation
#
# * Now consider a more realistic error model based on thermal relaxation with the qubit environment
# * Each qubit parameterized by a thermal relaxation time constant $T_1$ and a dephasing time constant $T_2$.
# * Note that we must have $T_2 \le 2 T_1$
# * Error rates on instructions are determined by gate time and qubit $T_1$, $T_2$ values
# +
# T1 and T2 values for qubits 0-3
T1s = np.random.normal(50e3, 10e3, 4) # Sampled from normal distribution mean 50 microsec
T2s = np.random.normal(70e3, 10e3, 4) # Sampled from normal distribution mean 50 microsec
# Truncate random T2s <= T1s
T2s = np.array([min(T2s[j], 2 * T1s[j]) for j in range(4)])
# Instruction times (in nanoseconds)
time_u1 = 0 # virtual gate
time_u2 = 50 # (single X90 pulse)
time_u3 = 100 # (two X90 pulses)
time_cx = 300
time_reset = 1000 # 1 microsecond
time_measure = 1000 # 1 microsecond
# QuantumError objects
errors_reset = [thermal_relaxation_error(t1, t2, time_reset)
for t1, t2 in zip(T1s, T2s)]
errors_measure = [thermal_relaxation_error(t1, t2, time_measure)
for t1, t2 in zip(T1s, T2s)]
errors_u1 = [thermal_relaxation_error(t1, t2, time_u1)
for t1, t2 in zip(T1s, T2s)]
errors_u2 = [thermal_relaxation_error(t1, t2, time_u2)
for t1, t2 in zip(T1s, T2s)]
errors_u3 = [thermal_relaxation_error(t1, t2, time_u3)
for t1, t2 in zip(T1s, T2s)]
errors_cx = [[thermal_relaxation_error(t1a, t2a, time_cx).expand(
thermal_relaxation_error(t1b, t2b, time_cx))
for t1a, t2a in zip(T1s, T2s)]
for t1b, t2b in zip(T1s, T2s)]
# Add errors to noise model
noise_thermal = NoiseModel()
for j in range(4):
noise_thermal.add_quantum_error(errors_reset[j], "reset", [j])
noise_thermal.add_quantum_error(errors_measure[j], "measure", [j])
noise_thermal.add_quantum_error(errors_u1[j], "u1", [j])
noise_thermal.add_quantum_error(errors_u2[j], "u2", [j])
noise_thermal.add_quantum_error(errors_u3[j], "u3", [j])
for k in range(4):
noise_thermal.add_quantum_error(errors_cx[j][k], "cx", [j, k])
print(noise_thermal)
# -
# ### Executing the noisy simulation
# +
# Run the noisy simulation
job = execute(circ, simulator,
basis_gates=noise_thermal.basis_gates,
noise_model=noise_thermal)
result_thermal = job.result()
counts_thermal = result_thermal.get_counts(0)
# Plot noisy output
plot_histogram(counts_thermal)
| qiskit/advanced/aer/building_noise_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
plt.style.use('fivethirtyeight')
bureau=pd.read_csv('./data/bureau.csv')
bureau
previous_loan_counts = bureau.groupby('SK_ID_CURR', as_index=False)['SK_ID_BUREAU'].count().rename(columns = {'SK_ID_BUREAU': 'previous_loan_counts'})
previous_loan_counts.head()
df = pd.DataFrame({'A' : ['foo', 'bar', 'foo', 'bar',
...: 'foo', 'bar', 'foo', 'foo'],
...: 'B' : ['one', 'one', 'two', 'three',
...: 'two', 'two', 'one', 'three'],
...: 'C' : np.random.randn(8),
...: 'D' : np.random.randn(8)})
df
grouped = df.groupby(['A']).count()
grouped
train_df = pd.read_csv('./data/application_train.csv')
train_df=train_df.merge(previous_loan_counts, on='SK_ID_CURR', how='left')
train_df.head()
train_df['previous_loan_counts']
| HomeCredictDefaultRisk/kernel_try.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis
# This module will introduce and examine the stage of 'exploratory data analysis'.
#
# Rather than focusing on the statistical or technical techniques employed in modern data science though, we will approach this stage with a bias-aware perspective.
# However, we will make use of [Jupyter notebooks](https://jupyter.org)—a popular tool in data science—to aid our exploratory data analysis by visualising some data.
# You do not need to be familiar with either Python or Jupyter Notebooks if you just want to gain an understanding of how social, cognitive, and statistical biases interact and affect downstream stages in the research and innovation lifecycle.
# But the code is presented for those who wish to get more "hands-on".
#
# <!-- You can also edit this code in an interactive Jupyter notebook:
# [](https://mybinder.org/v2/gh/chrisdburr/turing-commons/master?labpath=https%3A%2F%2Fgithub.com%2Fchrisdburr%2Fturing-commons%2Fblob%2Fmaster%2Fguidebooks%2Frri%2Fchapter4%2Fproject_design%2Fdata_analysis.ipynb) -->
#
# ## What is Exploratory Data Analysis?
#
# Exploratory data analysis is a crucial stage in the project lifecycle. It is where a number of techniques are employed for the purpose of gaining a better understanding of the dataset and any relationships that exist between the relevant variables. Among other things, this could mean,
#
# - Describing the dataset and important variables
# - Cleaning the dataset
# - Identifying missing data and outliers, and deciding how to handle them
# - Provisional analysis of any relationships between variables
# - Uncovering possible limitations of the dataset (e.g. class imbalances) that could affect the project
#
# We will cover each of these sub-stages of EDA briefly, but to reiterate, our primary focus in this section is on the risks and challenges that stem from a variety of biases that can cause cascading issues that affect downstream tasks (e.g. model training).
#
# ## COVID-19 Hospital Data
#
# For the purpose of this section we have created a synthetic dataset that contains X records for fictional patients who were triaged (and possibly admitted) to a single hospital for treatment of COVID-19.
#
# The dataset has been designed with this pedagogical task in mind.
# Therefore, although we relied upon plausible assumptions when developing our generative model, the data are not intended to be fully representative of actual patients. Our methodology for generating this dataset can be [found here](https://github.com/chrisdburr/turing-commons/blob/master/guidebooks/rri/chapter4/project_design/synthetic_data_generation.ipynb).
#
# ### Importing Data
#
# First of all, we need to import our data and the software packages that we will use to describe, analyse, and visualise the data.
# The following lines of code achieve this by importing a series of software packages and then loading a csv file `covid_patients_syn_data.csv` into a DataFrame `df` using the `pd.read_csv` command from the Pandas package.
# +
# The following lines import necessary packages and renames them
import numpy as np
import pandas as pd
import matplotlib.pylab as plt
import seaborn as sns
# This line imports data from a csv file
df = pd.read_csv('covid_patients_syn_data.csv')
# -
# ### Describing the Data
#
# Once we have imported our data, we will then want to identify what variables there are, what their typical values are, and also assess a variety of other summary statistics.
# We can use several commands to help us describe our dataset and get a quick overview.
#
# First, we can use the `shape` attribute to list the number of rows and columns in our dataset.
# The output (27308, 12) means that there are 27308 rows and 12 columns.
#
df.shape
# Second, we can use the `head` attribute to return the first 5 rows of our dataset, which can be useful if you want to see a small sample of values for each variable.
df.head()
# Third, we can use the `columns` attribute to list all the names all of the columns in our dataset.
# This is helpful if you want to quickly see which variables you will have access to during your analysis.
df.columns
# Finally, if we want to see how many *unique* values there are for each of the variables, we can use the `nunique` attribute (i.e., number (n) of unique values).
# For example, in the ethnicity column there are 5 different values, which align with the formal list used by Public Health England in a report on the [Disparities in the risk and outcomes of COVID-19](https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/908434/Disparities_in_the_risk_and_outcomes_of_COVID_August_2020_update.pdf)—this report was used as the basis for generating our synthetic data.
df.nunique(axis=0)
# These commands can be helpful for describing some basic aspects of our dataset. But what about more useful statistical information?
# For that we have the `describe` attribute, which returns the numeric values for `count`, `mean`, `standard deviation`, `min`, and `max`.
# The code after the brackets (`apply(lambda s: s.apply(lambda x: format(x, 'f'))` helps improve readability.)
df.describe().apply(lambda s: s.apply(lambda x: format(x, 'f')))
# ### Cleaning the Data
#
# #### Removing Unnecessary Variables
#
# Our dataset has been created for this specific task, so there isn't much cleaning that is required.
# Datasets that are downloaded from public repositories may not be so well structured, and will likely require tidying up.
# For instance, there may be redundant columns that are not needed, such as the `site_id` column for our dataset, which is the same for all values due to the data being collected from a single hospital site (`UHJ_43643`).
# These can be easily removed with the `drop` function.
df_cleaned = df.drop(['site_id'], axis=1)
df_cleaned.head()
# #### Removing Outliers
#
# It is also possible that there may be some outliers that are probably the result of human mistakes in recording. For example, someone may enter a height in feet and inches, such that we end up with a value of greater than 5 in that column.
# It's probably a safe assumption to assume that this is a mistake!
df_cleaned.hist("height", log=True)
# We can see there is one entry with a height of between 5.5 and 6m! We can manually remove this row, and any others with a height greater than 2.5m:
df_cleaned = df_cleaned[df_cleaned.height< 2.5]
df_cleaned.hist("height")
# #### Handling Missing Data
#
# One of the biggest challenges with cleaning datasets is choosing how to handle missing data.
# As you can see from the following line of code, there are 1326 rows with missing data in the "sex" column, which are represented in our datasets using the `NaN` (not-a-number) value.
#
# <!-- This section needs to have something about how simply removing rows with null values or trying to impute the likelihood of a M or F based on the other variables may exacerbate representation bias. -->
#
len(df_cleaned[df_cleaned.sex.isnull()])
# The simplest thing that one can do is to remove rows containing missing data. However, this could introduce bias.
#
# The best case scenario is that the missingness occurs at random - in this case removing the rows would reduce our statistical precision (because we would have a smaller dataset), but would not bias the result.
#
# However, if the reason why the data is missing is correlated with some other variable, then simply removing them will mean that the remaining dataset is skewed in some way. The best thing to do in this case could be to _impute_ the missing values, based on other variables in the same row. However, this is often a complex process, and needs to be done with care!
# ### Analysing the Data
#
# <!-- This section needs to add some useful illustrations of how Python can help you identify and understand correlations between variables in your dataset, and help document your exploratory data analysis method by using Jupyter Notebooks. -->
# For convenience, we may want to calculate some new variables based on those already in the dataset, that we think may be relevant. For example, there have been reports that Body Mass Index (BMI) is a risk factor for Covid. We can calculate this easily given the height and weight:
df_cleaned["BMI"] = df_cleaned["weight"]/pow(df_cleaned["height"],2)
df_cleaned.head(20)
# A quick way to get an idea of what variables are correlated with one another is to plot a "heatmap". Higher numbers in this plot represent stronger correlations (note the diagonal line of red squares all containing "1" - every variable is 100% correlated with itself!)
# +
# The following lines return a correlation matrix, for the cleaned dataframe, using the seaborn package
corr = df_cleaned[["age","BMI","admitted","intrusive_ventilation","died"]].corr()# plot the heatmap
ax = sns.heatmap(corr, xticklabels=corr.columns, yticklabels=corr.columns, annot=True, cmap=sns.diverging_palette(220, 20, as_cmap=True))
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
# -
# Note that this heatmap only works for numeric (or boolean) variables. For categorical variables such as "ethnicity", we will use other methods to visualise the data.
# ### Visualising the Data
#
# <!-- This section should show how including people who died in care homes would affect the correlation between age and one or more of the outcomes. This will be used to help support the missing data narrative. -->
# As we saw from the heatmap above, the age of the patient is perhaps the most important variable with regard to outcomes such as hospital admission or death. We will therefore separate the data into different "bins" of age, to see if we can observe other dependencies.
#
# This is a bit tricky to do in pandas, we need to define the bin boundaries, and then "group by" the other variable that we're interested in, summing the numbers of patients and deaths over each category.
df_plot = df_cleaned.copy(deep=True)
# add a 'counter' variable, which is just 1 for each row, so that when we sum over categories, we know how many rows we
# originally had in that category
df_plot["counter"] = 1.0
# now we want to put the data in bins of age-range and ethnicity, and sum the numbers of
# patients (total, admitted, ventilated, died) over these bins
bins = np.arange(20,100,10)
df_plot = df_plot.groupby(["ethnicity",pd.cut(df_plot["age"], bins)]).sum()
# calculate probability that patient died by dividing num(died) by the total in each bin
df_plot["prob_died"] = df_plot["died"]/df_plot["counter"]
# we can also calculate the uncertainty on this probability, using binomial errors
df_plot["prob_died_std"] = np.sqrt(df_plot["prob_died"]*(1-df_plot["prob_died"])/df_plot["counter"])
# at this point, we have a redundant column "age" which is the sum of all the ages in the bin - drop this,
# so that we can then promote the index age-range to a column (which will also be called "age")
df_plot = df_plot.drop(["age"], axis=1)
# we can also drop the other columns that no longer make sense since we have summed over
# all the patients in the age range
df_plot = df_plot.drop(["Unnamed: 0","height","weight", "BMI"], axis=1)
df_plot = df_plot.reset_index()
# convert this new "age" column to a string
df_plot["age"] = df_plot["age"].astype(str)
df_plot.head(10)
fig, ax = plt.subplots(figsize=(9,5))
sns.pointplot(data=df_plot, x="age", y="prob_died", hue="ethnicity")
# Ideally we would plot error bars here so we could see the uncertainties on the probabilities, but it does seem to be a consistent pattern that "White" patients had a lower probability of death across a range of ages, while for older patients, the "Asian / Asian British" ethnicity had a higher probability of dying.
# ### Dependency on other factors
#
# It is generally very difficult to quantify correlations in situations like this where multiple factors might affect the probability of a death, especially when these factors might be correlated with one another.
# We can try though to see whether we can see a dependency on sex, and/or BMI. Since we know that the strongest dependency is on age, we will just choose a limited age range to make this plot - in order to have a reasonable number of patients we'll look at the rows with age between 70 and 80.
df_newplot = df_cleaned.copy(deep=True)
df_newplot = df_newplot[(df_newplot.age > 70) & (df_newplot.age < 80)]
df_newplot["counter"] = 1.0
# now we want to put the data in bins of BMI and sex, and sum the numbers of
# patients (total, admitted, ventilated, died) over these bins
bins = np.arange(10,45,6)
df_newplot = df_newplot.groupby(["sex",pd.cut(df_newplot["BMI"], bins)]).sum()
# calculate probability that patient died by dividing num(died) by the total in each bin
df_newplot["prob_died"] = df_newplot["died"]/df_newplot["counter"]
df_newplot["prob_died_std"] = np.sqrt(df_newplot["prob_died"]*(1-df_newplot["prob_died"])/df_newplot["counter"])
df_newplot = df_newplot.drop(["BMI","Unnamed: 0","age","height","weight"], axis=1)
df_newplot = df_newplot.reset_index()
# convert this new "BMI" column to a string
df_newplot["BMI"] = df_newplot["BMI"].astype(str)
df_newplot.head(8)
sns.pointplot(data=df_newplot, x="BMI", y="prob_died", hue="sex")
# Well, we can clearly see that there is a difference between men and women, with men having a higher probability of death right across the BMI range.
#
# But the dependence on BMI doesn't look like we might expect - why is this? Well, it would take a detailed analysis to find out for sure, but one plausible hypothesis is that BMI is negatively correlated with age. (In fact if we look at the [code used to generate this synthetic dataset](Synthetic_data_generation.ipynb) we can see that this is indeed the case - average BMI peaks at age 60 and decreases linearly with age after that). Even though we only took a limited age-range, the variation within this 10-year slice is still big enough that it is the dominant factor in determining the shape of this plot.
#
# This is just one example of the pitfalls of trying to ascertain relationships between outcomes and variables - it's not easy!
# ## Missing data
#
# As well as isolated rows with missing or incorrectly entered data, it is possible that there are entire categories that are under-represented in our dataset, as an artifact of the data collection procedure. For example, perhaps we are missing some of the older patients in our dataset, as they may have been too frail to even present at the hospital.
# We are in the fortunate situation here of using synthetic data, so we can see the effect of this by adding back in some rows that were artificially removed in the dataset we have used up to now:
df_all = pd.read_csv("covid_patients_syn_data_unbiased.csv")
df_all.shape
# To see how the data was affected, we can compare the histograms of ages of patients, and patients that died, between our dataset and the dataset with the missing rows restored.
# +
fig, axes = plt.subplots(1,2, figsize=(15,6))
ages_orig = list(df_cleaned["age"])
ages_all = list(df_all["age"])
ages_died_orig = list(df_cleaned[df_cleaned["died"]==True]["age"])
ages_died_all = list(df_all[df_all["died"]==True]["age"])
axes[0].hist(ages_all,bins=10,range=(0,100), alpha=0.5, label="unbiased dataset")
axes[0].hist(ages_orig,bins=10,range=(0,100), alpha=0.5, label="our dataset")
axes[0].set_xlabel("Age of patient")
axes[0].set_ylabel("Number of cases")
leg = axes[0].legend()
axes[1].hist(ages_died_all,bins=10,range=(0,100), alpha=0.5, label="unbiased dataset")
axes[1].hist(ages_died_orig,bins=10,range=(0,100), alpha=0.5, label="our dataset")
axes[1].set_xlabel("Age of patient")
axes[1].set_ylabel("Number of deaths")
leg = axes[1].legend()
# -
| guidebooks/_build/jupyter_execute/rri/chapter4/project_design/data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of Stack Over Flow 2020 Survey Results
#
# In this notebook, i will analyze the Stack Over Flow 2020 Survey Data.<br>
# I want to understand the factors effecting job satisfaction in different countries.<br>
# And as new starter on learning programing language, Python, i want to undestand the <br>
# popularity of the languages in different countries.<br>
# For this purpose i will try to asnwer below questions:
#
# 1. How is the distribution of the respondents according to countries, does the distribution represents<br>
# the countries well according to their population ?
# 2. What are the most popular (currently being worked and desired to work next year) programing languages<br>
# in mostly represented countries ?
# 3. How is the distribution of the job satisfaction in different countries?<br>
#
# 4. What are the most effected features on job satistaction ?
# ### Index
# [i. Importing Required Packages](#i)<br>
# [ii. Importing Data](#ii)<br>
#
# [Q1 : How is the distribution of the respondents according to countries, does the distribution represents the countries well according to their population ?](#Q1)<br>
# * [Q1.1 : What is the percentage of Respondents according to Countries?](#Q1.1)<br>
# * [Q1.2 : What is Respondent Density According to Population?](#Q1.2)<br>
#
# [Q2. What are the most popular (currently being worked and desired to work next year) programing languages in mostly represented countries ?](#Q2)<br>
# * [Q2.1: What is the most popular language among the respondents ?](#Q2.1)<br>
# * [Q2.2 What are the usage percents of languages in Countries, Which language is being used most in which Country?](#Q2.2)<br>
# * [Q2.3: What is the most desired language next year ?](#Q2.3)<br>
# * [Q2.4 What are the desired percents of languages in Countries, Which language is desired most in which country?](#Q2.4)<br>
#
# [Q3: How is the distribution of following features in those countries ? (Primary field of study, Education Level, Job satisfaction, Employment and Job seeking status, Education importance)](#Q3)<br>
# * [Q3.1 How is the distribution of the primary fields in top countries?](#Q3.1)<br>
# * [Q3.2 How is the distribution of the education level in top countries?](#Q3.2)<br>
# * [Q3.3 How is the distribution of the job satisfaction in top countries?](#Q3.3)<br>
#
# [Q4: 4.What are the most effected features on job satistaction ?](#Q4)<br>
#
#
# [Conclusion](#5)<br>
#
#
#
#
# <a id="i"></a>
# ### i. Importing Required Packages
# +
# Importing required packages
import numpy as np
import pandas as pd
from collections import defaultdict
import seaborn as sns
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# Ingore warnings
import warnings
warnings.filterwarnings("ignore")
# -
# <a id="ii"></a>
# ## ii. Importing Data
# I donwloaded the Stack Over Flow 2020 survey data using this [link].<br>
# For the Population 2020 data, i downloaded the data from [kaggle], thanks to <NAME>.
#
#
#
#
#
# [link]: http://insights.stackoverflow.com/survey/
# [kaggle]:https://www.kaggle.com/tanuprabhu/population-by-country-2020
#
# +
# Reading datas from csv files
# Read Stack Over Flow Survey Results
results_2020 = pd.read_csv(r"C:\Users\oozturk8\Desktop\data\survey_results_public_2020.csv")
# Read Stack Over Flow Survey Questions
schema_2020 = pd.read_csv(r"C:\Users\oozturk8\Desktop\data\survey_results_schema_2020.csv")
# Read World Population Data
population_2020 = pd.read_csv(r"C:\Users\oozturk8\Desktop\data\population_by_country_2020.csv")
# -
# <a id="Q1"></a>
# ## Q1 : How is the distribution of the respondents according to countries, does the distribution represents the countries well according to their population ?
# I will start looking at first rows of datas to understand how they are look like:
# Looking at first 3 rows:
results_2020.head(3)
# <a id="Q1.1"></a>
# ### Q1.1 : What is the percentage of Respondents according to Countries?
# #### Business Understanding:
# Knowing the individuals distribution in each country is a must to gain consistent insights with further analysis.<br>
# If there is not sufficient respondent for a country in survey, this means that the country is not well represented.<br>
# Thus, i need to understand percentage of respondents according to countries.
# #### Data Understanding
# I will use "Country" column in order to find the number of the respondent from each country.<br>
# Afterwards i will divide the sum of respondents by total number of respondents to get the mean.<br>
# First: I will check if there is any NaN row in Country column:
# How many respondents having NaN for Country?
country_nan_count = results_2020.Country.isnull().sum()
print("Survey respondents not having any country: ", country_nan_count)
# Check if the respondents who have NaN for "Country" have any NaN for the rest of the columns
not_na_percents = results_2020[results_2020.Country.isnull()==True].notnull().mean()
print("Not-NaN Percent > 0 :\n" ,not_na_percents[not_na_percents>0])
# Drop NaN rows and create new data frame
df = results_2020.dropna(subset = ["Country"], axis = 0)
# I will create a function to calculate percentage of respondents according to countries:
def create_count_table(df, col_name):
'''
INPUTS:
df - (dataframe)- The dataframe containing related column
col_name (string)- Column name of the column of which values to be counted
OUTPUT:
new dataframe containing columns:
- col_name:
- Counts : Counts of items in target column
- Percent : Counts of item / Total count of column
'''
counts = df[col_name].value_counts().values
items = df[col_name].value_counts().index
percent = np.round(counts / counts.sum()*100,2)
new_table = pd.DataFrame({col_name:items, "Counts":counts, "Percent":percent})
return new_table
# Creating a dataframe including countries, counts and percent.
country_count = create_count_table(df, "Country")
country_count.head()
# +
# Visualisation - Creating Bar Plot
# Filter the data as to have countries percent > 1
country_count_filtered = country_count[country_count.Percent>1]
# Creating ax object
ax = country_count_filtered.plot(kind = "bar", x = "Country", y = "Counts",legend = False, figsize = (15,6), edgecolor = "black")
# Showing percentages over the bar
i=0
for patch in ax.patches:
ax.text((patch.get_x()),
patch.get_height()+100,
country_count_filtered.Percent.values[i],
fontsize=9,
rotation=0,
color="red")
i = i + 1
# Plotting
plt.xlabel("Countries", fontsize = 12)
plt.ylabel("Respondent Count and Percent", fontsize = 12)
plt.title("Respondent Counts and Percents According to Countries (Countries having min %1 respondents are shown)",
fontsize=14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.xticks(rotation = 45, fontsize = 9)
plt.show()
# -
# From above graph: Most interested Country in the survey is United States with a 19.46 % .<br>
# And it is followed by India, United Kingdom and Germany.
# <a id="Q1.2"></a>
# ### Q1.2 : What is Respondent Density According to Population?
# #### Business Understanding:
# * We see that top 5 countries according to respondent percent are US, India, UK, Germany and Canada.<br>
# * I want to understand how is the distribution according to country population.<br>
# * Thus i will compare the Respondent Density of each country:
#
# Respondent Density = Respondent number in country / Population of Country <br>
# #### Data Understanding:
# * I will use Population data to get the population values corresponding Countries:
# +
# Creating new data frame with necessary columns from population_2020 dataframe
pop_df = population_2020[["Country (or dependency)","Population (2020)"]]
pop_df.rename(columns={"Country (or dependency)": "Country", "Population (2020)": "Population"}, inplace = True)
pop_df.info()
# +
# Checking the Country names if there are differences between pop_df and country_count df
# Existing countries in country_count but not existing pop_df
non_matches = sorted(set.difference(set(country_count.Country),set(pop_df.Country)))
non_matches
# +
# Manuel correction of above non-matches in pop_df
# correcting_list = ["country name in country_count", "country name in pop_df"]
correcting_list = [['Brunei Darussalam','Brunei'],['Cape Verde', 'Cabo Verde'],['Congo, Republic of the...', 'Congo'],
['Czech Republic', 'Czech Republic (Czechia)'],['Democratic Republic of the Congo', 'DR Congo'],
['Hong Kong (S.A.R.)', 'Hong Kong'],['Kosovo', "Kosovo"],["Lao People's Democratic Republic",'Laos'],
['Libyan Arab Jamahiriya','Libya'],['Micronesia, Federated States of...','Micronesia'],['Nomadic',"Nomadic"],
['Republic of Korea','South Korea'],['Republic of Moldova', 'Moldova'],['Russian Federation','Russia'],
['Saint Vincent and the Grenadines','St. Vincent & Grenadines'],['Swaziland',"Swaziland"],
['Syrian Arab Republic','Syria'],
['The former Yugoslav Republic of Macedonia', "The former Yugoslav Republic of Macedonia"],
['United Republic of Tanzania','Tanzania'],
['Venezuela, Bolivarian Republic of...','Venezuela'],['Viet Nam','Vietnam']]
for x,y in correcting_list:
pop_df.loc[pop_df.Country == y, "Country"] = x
# +
# Creating new column in country_count data frame for population info
country_count["Population"]= np.NaN # creating new column composed of NaN values
for k in country_count.Country: # look for each country name in country_count
for g in pop_df.Country: # look for each country name in pop_df
if k == g: # if country names are same
country_count.Population[country_count.Country == k] = pop_df.Population[pop_df.Country == g].values
# get the population value of country from pop_df and write in corresponding cell in country_count
# +
# Drop NaN
country_count.dropna(inplace = True)
# Change data type to int
country_count.Population = country_count.Population.astype(int)
# -
# Creating Respondent Density column in coutry_count data frame
country_count["Resp_Density*100k"] = country_count.Counts/country_count.Population*100000
# Sorting according to Respondent Density and stacking as new data frame
country_count_sorted = country_count.sort_values(by=['Resp_Density*100k'], ascending=False)
# Filtering according to Percentage and stacking as new data frame
country_count_filtered = country_count_sorted[country_count.Percent >1]
country_count_filtered
# Visualisation
country_count_filtered.plot.bar("Country",["Percent", "Resp_Density*100k"],figsize=(15,6), edgecolor = "black")
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Respondent Percent and Respondent Density (Respondent Count / Country Population) of Countries",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# * We see from above graph: Although the big portion (19.5 %)of the respondents from the US, <br>
# if we compare the respondent densities, it is in the 8th order.<br>
# * So, we can say that the people in the US, did not show big interest in survey according to respondent density.<br>
# * Most interested top five countries are Sweden, Netherlands, Israel, Canada and UK.<br>
# * Least intereste countries are Pakistan and India among the countries having percentage > 1%.<br>
# <a id="Q2"></a>
# ## Q2. What are the most popular (currently being worked and desired to work next year) programing languages in mostly represented countries ?
# #### Business Understanding
# * As a new starter on learning programing language, Python, i want to undestand the
# popularity of the languages in different countries.
# * I will define the countries which are mostly represented or mostly interested in Survey as the countries having <br>
# percentage > 1% (so they are the countries shown on above graphs).<br>
#
#
# #### Data Understanding
# * For uncovering most popular languages, i will use the columns "LanguageWorkedWith" and "LanguageDesireNextYear"<br>
#
# * I will also create a new data frame containing the columns in which i am interested for further analysis.
#
# Creating a list to store top country names
top_country_list = list(country_count[country_count.Percent >1].Country.values)
top_country_list
# Creating a feature list
feature_list = ["Age","Age1stCode","ConvertedComp","Country","EdLevel","Employment","Gender",
"Hobbyist", "UndergradMajor","JobSat","JobSeek","LanguageDesireNextYear",
"LanguageWorkedWith", "MainBranch","NEWEdImpt","OrgSize","WorkWeekHrs",
"YearsCode","YearsCodePro"]
# Create new data frame for top countries containing columns in feature list
top_country_df = df[df.Country.isin(top_country_list)][feature_list]
# <a id="Q2.1"></a>
# ### Q2.1: What is the most popular language among the respondents ?
# I will define a function:
def get_language_percent(df, col_country, col_lang, top_country_list ):
'''
INPUT:
df - Dataframe
col_country - Column name in df (as string) where the countries are stored
col_lang - Language column in df (as string) where the languages are stored
top_country_list - A list containig country names for which popularity to be calculated
OUTPUT:
sorted_lang_percent_table:
Overal Percentages of currently worked languages sorted as descending (as Series)
country_lang_percent_df:
Percentages of currently worked languages in Countries (as dataframe)
lang_list:
A list containing unique languages
'''
# create new df
new_df = df[[col_country,col_lang]]
# dropp na
new_df.dropna(inplace = True)
# create list for unique languages
lang_list = []
for each in list(new_df[col_lang].value_counts().index):
splited = each.split(";")
for each in splited:
lang_list.append(each)
lang_list = list(set(lang_list))
# Arranging LanguageWorkedWith by seperating into several columns
for lang in lang_list:
new_df[lang] = new_df[col_lang].str.split(";")
new_df[lang] = [lang in row for row in new_df[lang]]
new_df[lang] = new_df[lang].astype(int).replace({False: 0, True: 1})
# Percentages of languages
sorted_lang_percent_table = new_df.iloc[:,2:].mean().sort_values(ascending = False)
# Sorting languages descending and creating new list
sorted_lang_list = new_df.iloc[:,2:].mean().sort_values(ascending = False).index
# Creating series for language statistics of each country
# and creating df from those series
series_list = []
for country in top_country_list:
country_lang_percent = new_df[new_df.Country == country].iloc[:,2:].mean().reindex(sorted_lang_list)
series_list.append(country_lang_percent)
country_lang_percent_df = pd.concat(series_list, axis = 1)
country_lang_percent_df.columns = top_country_list
country_lang_percent_df = country_lang_percent_df.transpose()
return sorted_lang_percent_table, country_lang_percent_df, lang_list
# getting statistics for LanguageWorkedWith using above function
sorted_lang_percent_table, country_lang_percent_df, lang_list = get_language_percent(
top_country_df, "Country", "LanguageWorkedWith",top_country_list )
sorted_lang_percent_table
# Visualisation
sorted_lang_percent_table.plot.bar(figsize=(15,6), edgecolor = "black")
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Language", fontsize = 12)
plt.title("Working Language Percentage of Respondents", fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# From above graph :
# * Most popular language currently being worked is JavaScript with 67 % of usages among the respondents.
# * Followed by HTML/CSS, SQL, Python, Java...
# <a id="Q2.2"></a>
# ### Q2.2 What are the usage percents of languages in Countries, Which language is being used most in which Country?
# Using the other output of above function:
country_lang_percent_df
# +
# Visualisation
# Filtering as to get top 5 language for each country
country_lang_percent_df.iloc[:,:5].plot.bar(figsize = (20,4),
width = 0.8,
colormap= "Set3",
edgecolor = "black",
alpha = 0.8)
# Ploting
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Usage Percentage of Top 5 Languages in Top Countries", fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
# * Overal top 5 language are JavaScript, HTML, SQL, Python and Java from previous graph.
# * Still almost in all countries JavaScript at the first rank except Pakistan; in Pakistan HTML/CSS has greater usage than JavaScript.
# * Altough Python is at 4th rank, it is at 2nd rank in Israel and at 4th rank some other countries.
# * HTML/CSS is mostly used in Pakistan,<br>
# SQL is mostly used in Italy,<br>
# Python is mostly used in Israel and US,<br>
# Java is mostly used in Italy and Germany among the other countries.
# <a id="Q2.3"></a>
# ### Q2.3: What is the most desired language next year ?
# getting statistics for LanguageDesireNextYear using same function
sorted_desired_lang_percent_table, country_desired_lang_percent_df, lang_list = get_language_percent(
top_country_df, "Country", "LanguageDesireNextYear",top_country_list )
sorted_desired_lang_percent_table
# Visualisation
sorted_desired_lang_percent_table.plot.bar(figsize=(15,6), edgecolor = "black")
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Language", fontsize = 12)
plt.title("Desired Language Percentage of Respondents", fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# * Most desired language is Python with 49.5% desire among the respondents.
# * Followed by JavaScript, HTML/CSS, SQL and TypeScript.
# * So, Bash/Shell/PowerShell and Java seems to be losing popularity.
# * TypeScript seems to be getting more popular.
# <a id="Q2.4"></a>
# ### Q2.4 What are the desired percents of languages in Countries, Which language is desired most in which country?
country_desired_lang_percent_df
# +
# Visualisation
country_desired_lang_percent_df.iloc[:,:5].plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Set3",
edgecolor = "black",
alpha = 0.8)
# Plotting
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Desire Percentage of Top 5 Languages in Top Countries", fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
# * Python is mostly desired in Pakistan, India and Israel,
# * JavaScript is mostly desired Brazil and Pakistan,
# * HTML/CSS is mostly desired in Brazil,
# * SQL is mostly desired in Brazil, Italy and US,
# * TypeScript is mostly desired in Netherlands.
# <a id="Q3"></a>
# ## Q3: How is the distribution of following features in those countries ? (Primary field of study, Education Level, Job satisfaction, Employment and Job seeking status, Education importance)
#
# ### Business Understanding:
# More importantly, as a new programmer, i want to understand how well satisfied the programmer are.<br>
# And also in general, i wonder other features how effects the satisfaction.
#
# ### Data Understanding
# ### Exploratory Data Analysis
# +
# Arranging, cleaning columns
# EdLevel column, splitting unnecessary parts from column
top_country_df["EdLevel"] = top_country_df.EdLevel.str.split("\(", expand=True)[0]
# Employment column
top_country_df.Employment = top_country_df.Employment.str.split(", f", expand = True)[0]
# UndergradMajor column
top_country_df.UndergradMajor = top_country_df.UndergradMajor.str.split("\,|\(", expand = True)[0]
# Arranging OrgSize column
top_country_df.OrgSize = top_country_df.OrgSize.str.split("em|er", expand = True)[0]
# Dropping NaN in "LanguageDesireNextYear" and "LanguageWorkedWith"
top_country_df.dropna(subset = ["LanguageDesireNextYear","LanguageWorkedWith"], inplace = True)
# Dropping Gender column
top_country_df.drop("Gender", axis = 1, inplace = True)
# +
# Arranging Age1stCode column:
# Values "Younger than 5 years" in column to be replaced with 4.
# Data type to be changed from object to int
# Nan values to be filled with mean of the column
# Replace "Younger than 5 years" with 4.
top_country_df.Age1stCode = [4 if each == "Younger than 5 years" else each for each in top_country_df.Age1stCode]
# Fill na values with mean:
# Calculate the mean
Age1stCode_sum = sum([int(each) for each in top_country_df.Age1stCode.dropna(inplace = False)])
Age1stCode_len = len([int(each) for each in top_country_df.Age1stCode.dropna(inplace = False)])
Age1stCode_mean = Age1stCode_sum / Age1stCode_len
# Fill na with mean
top_country_df.Age1stCode = top_country_df.Age1stCode.fillna(Age1stCode_mean)
# Change data type to int
top_country_df.Age1stCode = top_country_df.Age1stCode.astype(int)
# +
# Arranging YearsCode column:
# "Less than 1 year" to be replaced with 0.5
# "More than 50 years" to be replaced with 51
# Data type to be changed from object to float
# Nan values to be filled with mean of the column
# Replacing "Less than 1 year" --> 0.5 and "More than 50 years" --> 51
top_country_df.YearsCode = [0.5 if each == "Less than 1 year" else
51 if each == "More than 50 years" else
each for each in top_country_df.YearsCode]
# Fill na values with mean:
# Calculate the mean
YearsCode_sum = sum([float(each) for each in top_country_df.YearsCode.dropna(inplace = False)])
YearsCode_len = len([float(each) for each in top_country_df.YearsCode.dropna(inplace = False)])
YearsCode_mean = YearsCode_sum / YearsCode_len
# Fill na with mean
top_country_df.YearsCode = top_country_df.YearsCode.fillna(YearsCode_mean)
# Change data type to float
top_country_df.YearsCode = top_country_df.YearsCode.astype(float)
# +
# Arranging YearsCodePro column:
# Less than 1 year --> 0.5
# More than 50 years --> 51
# Data type to be changed from object to float
# Nan values to be filled with mean of the column
# Replacing
top_country_df.YearsCodePro = [0.5 if each == "Less than 1 year" else
51 if each == "More than 50 years" else
each for each in top_country_df.YearsCodePro]
# Fill na values with mean:
# Calculate the mean
YearsCodePro_sum = sum([float(each) for each in top_country_df.YearsCodePro.dropna(inplace = False)])
YearsCodePro_len = len([float(each) for each in top_country_df.YearsCodePro.dropna(inplace = False)])
YearsCodePro_mean = YearsCodePro_sum / YearsCodePro_len
# Fill na with mean
top_country_df.YearsCodePro = top_country_df.YearsCodePro.fillna(YearsCodePro_mean)
# Change data type to float
top_country_df.YearsCodePro = top_country_df.YearsCodePro.astype(float)
# +
# Column WorkWeekHrs and Age
# Filling nan with mean
top_country_df.WorkWeekHrs = top_country_df.WorkWeekHrs.fillna(top_country_df.WorkWeekHrs.mean())
top_country_df.Age = top_country_df.Age.fillna(top_country_df.Age.mean())
# +
# Filling nan values with median in categorical features:
for each in ["MainBranch","Employment","EdLevel","JobSeek","UndergradMajor","NEWEdImpt","JobSat","OrgSize"]:
top_country_df[each].fillna(top_country_df[each].value_counts().index[0], inplace=True)
# +
# Checking the notnull percentage of columns:
top_country_df.notnull().mean().sort_values(ascending = False)
# -
# * So, i filled all the nan except ConvertedComp feature.
# * Almost 40 % of data in ConvertedComp is missing; i will drop the nans in this column later,
# after i complete the analysis in this section.
# * I do not want to loose data in other features by dropping it now.
# +
# Cleaning LanguageDesireNextYear:
# Creating new categorical columns for each Language
for lang in lang_list:
top_country_df[lang] = top_country_df["LanguageDesireNextYear"].str.split(";") # split each language
top_country_df[lang] = [lang in row for row in top_country_df[lang]] # True-False
top_country_df[lang] = top_country_df[lang].replace({False: 0, True: 1}).astype(int) # Change to 1 and 0
top_country_df.rename(columns={lang: "Desired_{}".format(lang)}, inplace = True) # Change type
# Deleting original column
top_country_df.drop(["LanguageDesireNextYear"], axis = 1, inplace = True)
# +
# Cleaning "LanguageWorkedWith":
# Creating new categorical columns for each Language
for lang in lang_list:
top_country_df[lang] = top_country_df["LanguageWorkedWith"].str.split(";")
top_country_df[lang] = [lang in row for row in top_country_df[lang]]
top_country_df[lang] = top_country_df[lang].replace({False: 0, True: 1}).astype(int)
top_country_df.rename(columns={lang: "Worked_{}".format(lang)}, inplace = True)
# Deleting original column
top_country_df.drop(["LanguageWorkedWith"], axis = 1, inplace = True)
# -
# Check the data
top_country_df.head()
# <a id="Q3.1"></a>
# ### Q3.1 How is the distribution of the primary fields in top countries?
# * I will define a function in order to calculate the percentages of feature pairs in data frame
def get_pair_statistics(top_country_df, in_col_pair1, col_pair2, top_country_list):
'''
INPUTS:
top_country_df - Dataframe in which the features to be investigated (as dataframe)
in_col_pair1 - Name of the column in which data to be grouped (as string)
col_pair2 - Name of the column which will be investigated in grouped data (as string)
top_country_list - country list in order the reindex the data (as list)
OUTPUTS:
percents_df - Percentage table of col_pair2 in in_col_pair1 (as dataframe)
ovrl_percents_df - Percentage table of col_pair2 in overall sum of col_pair2 for each in_col_pair1 (as dataframe)
'''
# Create gropued df
grouped_df = top_country_df.groupby([in_col_pair1])[col_pair2]
# Creating new df from groups in grouped df
pair1_list = []
series_list = []
for a,b in grouped_df:
pair1_list.append(a)
series_list.append(b.value_counts())
# Creating new df with series
percents_df = pd.concat(series_list, axis = 1)
# Updating column names
percents_df.columns = pair1_list
# Copying new_df for relative percentage
ovrl_percents_df = percents_df.copy()
# Getting percentages
for col in percents_df.columns:
percents_df[col] = percents_df[col].map(lambda x:x/(percents_df[col].sum())*100)
# Getting relative percentages
ovrl_percents_df = ovrl_percents_df.div(ovrl_percents_df.sum(axis = 1), axis = 0)*100
# Switching columns and rows in both dfs
percents_df = percents_df.transpose()
ovrl_percents_df = ovrl_percents_df.transpose()
# Reindex both dfs according to top_country_list
percents_df = percents_df.reindex(top_country_list)
ovrl_percents_df = ovrl_percents_df.reindex(top_country_list)
return percents_df, ovrl_percents_df
undergradMajor_percent_in_countries_df, undergradMajor_ovrl_percent_in_countries_df = get_pair_statistics(
top_country_df, "Country", "UndergradMajor", top_country_list)
undergradMajor_percent_in_countries_df
# +
# Visualisation of undergradMajor_percent_in_countries_df
undergradMajor_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title(
"Primary Field of Study Percentage of Respondents in Top Countries (Field count in country / Respondent Sum in Country)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# Visualisation of undergradMajor_ovrl_percent_in_countries_df
undergradMajor_ovrl_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title(
"Primary Field of Study Overall Percentage of Respondents in Top Countries (Field count in country / Field's Overall Count)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
# * Primary field of the respondents is Computer Science in all countries; roughly min 60% of respondent's primary field is computer science in each country.
# * From 2nd graph we can see that big portion of the computer science and information system employees, as well as fine arts and humanities dicipline, are in US.
# * In india, respondents are mostly from other engineering diciplines.
# <a id="Q3.2"></a>
# ### Q3.2 How is the distribution of the education level in top countries?
EdLevel_percent_in_countries_df, EdLevel_ovrl_percent_in_countries_df = get_pair_statistics(
top_country_df, "Country", "EdLevel", top_country_list)
# +
# Visualisation of EdLevel_percent_in_countries_df
EdLevel_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Education Level Percentage of Respondents in Top Countries (E.Level Count in country / E.Level Total in Country)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# Visualisation EdLevel_ovrl_percent_in_countries_df
EdLevel_ovrl_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Education Level Overall Percentage of Respondents in Top Countries (E.Level Count in country / E.Level Total)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
# * At least 70 % of the respondents in India and Pakistan have Bachelor's degree; which is the highest among the other countries.
# * However in overall, US has the big portion of Bachelor's Degrees than India, roughly 30 percent of Bachelor's Degree is in US and 23% is in India.
# * US has the big portion of respondents (50%) having master's degree,Canada at 2nd rank with 13%.
# * Spain has th big portion of repondents (18%) having professioal degree, Russia at 2nd rank with 13%.
# <a id="Q3.3"></a>
# ### Q3.3 How is the distribution of the job satisfaction in top countries?
JobSat_percent_in_countries_df, JobSat_ovrl_percent_in_countries_df = get_pair_statistics(
top_country_df, "Country", "JobSat", top_country_list)
# +
# Visualisation of JobSat_percent_in_countries_df
JobSat_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Job Satisfaction Percentage of Respondents in Top Countries (JobSat Count in country / JobSat Total in Country)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# Visualisation of JobSat_ovrl_percent_in_countries_df
JobSat_ovrl_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Job Satisfaction Overall Percentage of Respondents in Top Countries (JobSat Count in country / JobSat Total)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
Empl_percent_in_countries_df, Empl_ovrl_percent_in_countries_df = get_pair_statistics(
top_country_df, "Country", "Employment", top_country_list)
# +
# Visualisation of Empl_ovrl_percent_in_countries_df
Empl_ovrl_percent_in_countries_df.plot.bar(
figsize = (20,4),
width = 0.8,
colormap= "Paired",
edgecolor = "black",
alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("Employment Overall Percentage of Respondents in Top Countries (Employment Count in country / Employment Total",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
JobSeek_percent_in_countries_df, JobSeek_ovrl_percent_in_countries_df = get_pair_statistics(
top_country_df, "Country", "JobSeek", top_country_list)
# +
# Visualisation of JobSeek_ovrl_percent_in_countries_df
JobSeek_ovrl_percent_in_countries_df.plot.bar(
figsize = (20,4), width = 0.8, colormap= "Paired", edgecolor = "black", alpha = 0.9)
plt.legend(bbox_to_anchor=(1,1))
plt.xticks(rotation = 45, fontsize = 9)
plt.xlabel("Countries", fontsize = 12)
plt.title("JobSeek Overall Percentage of Respondents in Top Countries (JobSeek Count in country / JobSeek Total)",
fontsize = 14)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.show()
# -
# <a id="Q4"></a>
# ## Q4: What are the most effective features on job satisfaction?
# * I will use Sklearn library to create a Logistic Regression model.
# * Using model coefficients, features that has negative and positive effect on job satisfaction to be calculated.
# * Before creting model, i will prepare the data.
# Creating list for numerical and categorical variables
numericals = ["Age","Age1stCode","ConvertedComp","WorkWeekHrs","YearsCode","YearsCodePro"]
categoricals = ["Country","EdLevel","Employment","Hobbyist","UndergradMajor","JobSeek",
"MainBranch","NEWEdImpt","OrgSize"]
# +
# Checking JobSat column:
top_country_df.JobSat.value_counts()
# +
# Arranging JobSat column:
# I will accept "Very satisfied" and "Slightly satisfied" as "Satisfied"
# I will accept "Very dissatisfied" and "Slightly dissatisfied" as "Disatisfied"
# I will delete the rows having "Neither satisfied nor dissatisfied"
# Convert categorical variable "Satisfied" as 1 and "Dissatisfied" as 0.
# Delete rows "Neither satisfied nor dissatisfied" and create new dataframe
df = top_country_df.drop(top_country_df[top_country_df.JobSat == "Neither satisfied nor dissatisfied"].index)
# Replace "Very satisfied" and "Slightly satisfied" as "Satisfied"
# Replace "Very dissatisfied" and "Slightly dissatisfied" as "Disatisfied"
# Convert to categorical variable: "Satisfied" as 1 and "Dissatisfied" as 0.
df.JobSat = [1 if each == "Very satisfied" else
1 if each == "Slightly satisfied" else
0 if each == "Very dissatisfied"else
0 if each == "Slightly dissatisfied" else
each for each in df.JobSat]
# +
# Droppind nan in ConvertedComp:
df = df.dropna()
# -
# One hot encoding categorical variables
df = pd.get_dummies(df, columns = categoricals )
# Normalization of numerical variables
for each in numericals:
df[each] = (df[each] - df[each].min()) / (df[each].max() - df[each].min())
# +
# LOGISTIC REGRESSION MODEL
# Split data into X and y
X = df.drop("JobSat", axis = 1)
y = df.JobSat
# split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=7)
# Define the model
model = LogisticRegression()
# Fit the model
model.fit(X, y)
# get importance (effects of each feature or variable on JobSat)
importance = model.coef_[0]
# make predictions for test data and evaluate
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# +
# Create dataframe for feature importance results
results_df = pd.DataFrame()
results_df["Rates"] = importance.tolist()
results_df["Columns"] = X.columns
# Reindex dataframe and sort and filter
new_index = results_df.Rates.sort_values(ascending = False).index
sorted_results = results_df.reindex(new_index)
filtered_results = sorted_results[np.abs(sorted_results.Rates) > 0.15]
# Visualisation
plt.figure(figsize =(10,8))
plt.barh(filtered_results.Columns, filtered_results.Rates)
plt.grid(axis="both", color="gray", linewidth=0.1)
plt.title("Negative and Positive Effected Features on Job Satistaction",fontsize = 14)
plt.show()
# -
# * Features having effect rate > abs(0.15) are shown.
# * According to graph:
# * Top 3 features negatively effecting Job Satisfaction are age, actively looking for a job and age of startin to coding. So, in the elderly ages job satisfaction may decrease because of the personal expectation increases. As expected, respondents who are looking for a job would be dissatisfied. Respondents who started in early ages may be dissatisfied beacuse of they are well-qualified in their profession but the company they work may not be well-qualified comparing with their expectations. In the same way, as the professional coding years increase, satisfaction may decrease. Also increased working hours decreases satisfaction.
# * Among the countries; most dissatisfied countries are Israel, France, Poland, Spain, Brazil and Italy.
# * Primary-Elementary shcool graduations most dissatisfied, whereas having doctoral degrees mostly satisfied.
# * Most satisfied countries Pakistan, Sweden, US, Canada and Australia.
#
# <a id="5"></a>
# ## Conclusion
# Having the top respondent density and greater positive effect ratio on satisfaction, Sweden seems to have the most satisfied developers.<br>
#
# Although Pakistan has the greatest positive effect ratio on job satisfaction; because it has lowest respondent density, i can not conclude that it has the satisfied developers.<br>
#
# We can conclude that most dissatisfied developers are from Israel, due to it’s respondent density is higher and it has top negative effect ration among the countries.<br>
#
# As programing languages, Python and TypeScript have increasing popularity, however JavaScript is still mostly used language.
| Project1_Analysis-of-Stack-Overflow-2020-Survey-Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME> - praca domowa nr 2
# <h1 id="tocheading">Spis treści</h1>
# <div id="toc"></div>
# + language="javascript"
# $.getScript('https://kmahelona.github.io/ipython_notebook_goodies/ipython_notebook_toc.js')
# -
# # Wczytanie i ogląd
# Póki co wiem że przez te dwa tygodnie mamy się zabawić ze zbiorem dotyczącym *Allegro*. Czyli konkretnie... ?
#
# Na początek wczytam te obowiązkowe pakiety...
import pandas as pd
import numpy as np
# ... odtworzę moje dane...
data = pd.read_csv("allegro-api-transactions.csv")
data.sample(10)
data.shape
data.dtypes
# Wow - niby tylko 14 kolumn, ale ponad 400 tysięcy wierszy. Czy to już big data? Chyba jeszcze nie, ale więcej to niż pierwszy lepszy zbiór którym się wcześniej bawiłem.
#
# Zbiór chyba jasny - mamy informacje o produktach z serwisu aukcyjnego i ich parametry. Nawet bez znajomości treści naszego zadania instynktownie przychodzi co będzie naszą **zmienną celu** - i oczywiście myślę tu o kolumnie *price*. Do jej wyznaczania strzelam że najbardziej kluczowa będzie główna kategoria (*main category*), data \[wypuszczenia produktu?\] (*date*), a także *it_seller_rating* - punkty sprzedawcy \[zapewne więcej <-> bardziej doświadczony, bardziej zaufany\]. No ale zobaczymy.
# # Target encoding dla it_location
# Wykonajmy nasze pierwsze kodowanie dla kolumny *it_location* - ograniczona liczba miast do wyboru powinna dać nam satysfakcjonujący efekt prawdopodobnie kilkudziesięciu do kilkuset różnych lokacji.
# +
import category_encoders
te = category_encoders.target_encoder.TargetEncoder(data)
encoded = te.fit_transform(data['it_location'], data['price'])
data['it_location_encoded'] = encoded
# -
data[['it_location', 'price', 'it_location_encoded']].sample(10)
# Z oczywistych względów w wypadku naszej ramki *target jest zdecydowanie lepszy niż one-hot* - co najważniejsze mamy zchowane własności tego drugiego - każde miasto dostało swój unikalny numerek; ale co ważniejsze te liczby nie są przypadkowe i są ściśle powiązane ze średnią ceną dla tego miasta. Pojedyncze wiersze może nie dadzą nam tu multum informacji, ale biorąc pod uwagę, że dla takiej Woli Krzysztoporskiej dostajemy znacznie mniejszy identyfikator w porównaniu np. do takiej Warszawy co jest śliśle powiązane z naszą zmienną celu, jest już zdecydowanie satysfakcjonujące.
# # Encoding dla main_category
# W naszej ramce danych rzeczywiście większość kolumn nie wymaga category encoding - zwykle mamy do czynienia z wartościami prawda / fałsz \[np. *pay_option_transfer*, *pay_option_on_delivery*\] albo posiadających tak wiele wartości, że ciężko byłoby zrobić z nimi coś sensownego i nie miałoby to raczej sensu \[np. *seller*\].
#
# Mądze jest więc zakodować akurat *main_category* - jak sama nazwa wskazuje dotyczy ona tylko jednej kategorii. Przeciwnie także co do kolumn wielowartościowych takich jak identyfikator przedmiotu czy nazwa sprzedawcy na pewno będzie ona przydatna przy implementacji algorytmów uczenia maszynowego. Razem z chociażby datą, "punktami" sprzedawcy (*it_seller_rating*) czy parametrami prawda / fałsz produktu będziemy mogli otrzymać potencjalnie przybliżoną predykcję ceny.
# ## One-hot encoding
# +
values = np.array(data[["main_category"]])
from sklearn.preprocessing import LabelEncoder
# integer encode
le = LabelEncoder()
integer_encoded = le.fit_transform(values)
print(integer_encoded)
#invert
print(le.inverse_transform(integer_encoded))
# -
category_series = pd.concat([pd.DataFrame(integer_encoded),pd.DataFrame(le.inverse_transform(integer_encoded))], axis = 1)
category_series = category_series.drop_duplicates()
category_series.columns = ["Index", "Category"]
category_series = category_series.sort_values("Index")
category_series
# +
from sklearn.preprocessing import OneHotEncoder
# one hot encode
onehot_encoder = OneHotEncoder(sparse=False)
integer_encoded = integer_encoded.reshape(len(integer_encoded), 1)
onehot_encoded = onehot_encoder.fit_transform(integer_encoded)
print(onehot_encoded)
# invert
inverted = onehot_encoder.inverse_transform(onehot_encoded)
print(inverted.transpose())
# -
onehot_encoded = pd.DataFrame(onehot_encoded)
onehot_encoded.columns = category_series["Category"]
onehot_encoded
data_onehot = pd.concat([data, onehot_encoded], axis = 1)
data_onehot.sample(10)
# Tak oto otrzymaliśmy potężną 42-kolumnową data frame, gdzie ostatnie 27 odpowiadają za przyporządkowanie do głównej kategorii produktu.
# ## Binary encoding
# Brzmi wdzięcznie, pewnie coś z bitami.
#
# A zera i jedynki to sama przyjemność, czyż nie?
bin_e = category_encoders.BinaryEncoder(cols = ['main_category'])
bin_e.fit_transform(data)
# *Binary encoding* na chłopski rozum wydaje się być sensowniejszy niż *one hot* - zamiast 27 dodatkowych kolumn po jednej na każdą kategorię, tu dostajemy ich jedynie 5 - mimo, że wciąż zostajemy w obszarze bitów.
#
# Zalety? Mniejsza ramka danych, zachowanie unikalności oznaczeń kategorii, spójność i szybszy transfer.
#
# Wady? Słaba przejrzystość bez wiedzy o zastosowaniu metody \[zwłaszcza dla osób spoza branży\] i prawdopodobnie łatwe wpadnięcie w pułapkę wykrywania podobieństwa między kategoriami o większości bitów wspólnych z tych 5 kolumn przez źle napisane algorytmy uczenia maszynowego - np. *Antyki i sztuka* oraz *bilety* w czterech na pięć kolumn odpowiadających za bity kategorii mają tą samą wartość, co mogłoby mylić, że mają ze sobą coś wspólnego - a są to jednak dwa zupełnie odmienne byty.
# ## Polynomial encoding
# Kodowanie wielomianowe, a cóż to takiego?
pe = category_encoders.PolynomialEncoder(cols = ['main_category'])
pe.fit_transform(data)
# Jak udalo mi się zrozumieć owe kodowanie przypisuje w 27 dodatkowych kolumnach specjalne wartości wyliczone za pomocą mądrze dobranych wzorów i zależności matematycznych. Ale to chyba jeszcze nie ten poziom!
# # Uzupełnianie braków
# ## Przygotowanie pomocniczych ramek danych
# Ograniczmy nasz zbiór do wskazanych zmiennych numerycznych.
#
# W tym wypadku *price* będziemy traktować jako zmienną objaśniającą.
data_4 = data[["it_seller_rating", "it_quantity", "price"]]
data_4.head()
# Usuńmy losowe 10% wierszy.
np.random.seed(1234)
removed_ids = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids
removed_ids.sort()
removed_ids
to_remove = pd.DataFrame(data_4.iloc[removed_ids]).reset_index()
remove_ids = pd.DataFrame(removed_ids)
remove_ids.columns = ["index"]
to_remove = to_remove.merge(remove_ids, on = "index")
before_remove = to_remove.copy()
to_remove[["it_seller_rating"]] = np.nan
to_remove
removed = to_remove.copy()
removed[["it_seller_rating"]] = np.nan
removed
data_4_removed = data_4.copy().reset_index()
data_4_removed = data_4_removed.merge(to_remove, on = "index", how = "left")
data_4_removed = data_4_removed.drop(["index", "it_seller_rating_y", "price_y"], axis = 1)
data_4_removed["it_seller_rating"] = np.where(data_4_removed["it_quantity_x"] == data_4_removed["it_quantity_y"], np.nan, data_4_removed["it_seller_rating_x"])
data_4_removed = data_4_removed.drop(["it_seller_rating_x", "it_quantity_y"], axis = 1)
data_4_removed.columns = ["it_quantity", "price", "it_seller_rating"]
data_4_removed.sample(15)
# ## Multivariate feature imputation
# ### Pierwsze wypełnianie
# Użyjmy gotowych narzędzi do znalezienia spodziewanych wartości *it_seller_rating*.
# +
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
imp = IterativeImputer(max_iter=10, random_state=0)
imp.fit(data_4)
IterativeImputer(random_state=0)
filled = pd.DataFrame(imp.transform(to_remove.drop(["index"], axis = 1)))
filled.columns = ["it_seller_rating", "it_quantity", "price"]
# -
filled["index"] = removed_ids
filled
sample_10_to_check = np.random.choice(len(removed), 10, replace=False)
sample_10_to_check
compare = before_remove.iloc[sample_10_to_check].merge(filled.iloc[sample_10_to_check], on = 'price')
compare = compare.drop(["index_x", "it_quantity_x", "it_quantity_y", "price", "index_y"], axis = 1)
compare
# ### Wnioski z próby na pierwszej 10% usuniętych danych
# Na pierszy rzut oka *Multivariate feature imputation* zdaje się dać taki sobie efekt - przewidywana liczba punktów wystawcy zdaje się być bardzo zbliżona niezależnie od produktu.
# Do "mądrej" weryfikacji narzędzia skorzystamy z miary RMSE - jest to pierwiastek średniego kwadratu błędu prognoz.
# +
from sklearn.metrics import mean_squared_error
from math import sqrt
rms = sqrt(mean_squared_error(before_remove["it_seller_rating"], filled["it_seller_rating"]))
# -
rms
# Jak widać *Multivariate feature imputation* dla standardowych parametrów okazało się mało użyteczne - przewidziana ocena sprzedawcy nawet nie jest podobna do jego prawdziwej.
# ### Kolejne 9 prób i odchylenie standardowe MFI powtórzonego wielokrotnie
deviatons = [0] * 10
deviatons[0] = rms
# Powtórzmy nasz eksperyment 10 razy i obliczmy odchylenie standardowe wyniku.
#
# Wpierw wylosujmy seedy do kolejnych losowań.
np.random.seed(100)
random_seeds = np.random.choice(10000, 9)
random_seeds
for i in range(1, 10):
np.random.seed(random_seeds[i-1])
removed_ids = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids.sort()
to_remove = pd.DataFrame(data_4.iloc[removed_ids]).reset_index()
remove_ids = pd.DataFrame(removed_ids)
remove_ids.columns = ["index"]
to_remove = to_remove.merge(remove_ids, on = "index")
before_remove = to_remove.copy()
to_remove[["it_seller_rating"]] = np.nan
removed = to_remove.copy()
removed[["it_seller_rating"]] = np.nan
data_4_removed = data_4.copy().reset_index()
data_4_removed = data_4_removed.merge(to_remove, on = "index", how = "left")
data_4_removed = data_4_removed.drop(["index", "it_seller_rating_y", "price_y"], axis = 1)
data_4_removed["it_seller_rating"] = np.where(data_4_removed["it_quantity_x"] == data_4_removed["it_quantity_y"], np.nan, data_4_removed["it_seller_rating_x"])
data_4_removed = data_4_removed.drop(["it_seller_rating_x", "it_quantity_y"], axis = 1)
data_4_removed.columns = ["it_quantity", "price", "it_seller_rating"]
filled = pd.DataFrame(imp.transform(to_remove.drop(["index"], axis = 1)))
filled.columns = ["it_seller_rating", "it_quantity", "price"]
filled["index"] = removed_ids
rms = sqrt(mean_squared_error(before_remove["it_seller_rating"], filled["it_seller_rating"]))
deviatons[i] = rms
deviatons
import statistics
statistics.stdev(deviatons)
# <NAME>ie wartości miar RMSE, ich odchylenie standardowe jest satysfakcjonująco małe.
# ### Usunięcie wierszy z *it_quantity*
# Przygotuję ziarna do wylosowania 10 zestawów danych z usuniętymi po 10% kolejno: *it_seller_rating* i *it_quantity*.
np.random.seed(200)
random_seeds = np.random.choice(10000, 10)
random_seeds
deviatons2 = [0] * 10
# Wylosuję *id* wierszy do usunięcia - oczywiście mogą się one powtórzyć.
# +
np.random.seed(random_seeds[0])
removed_ids_1 = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids_2 = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids_1.sort()
removed_ids_2.sort()
print(removed_ids_1)
print(removed_ids_2)
# -
removed_ids = pd.concat((pd.DataFrame(removed_ids_1), pd.DataFrame(removed_ids_2)), axis = 0)
removed_ids = removed_ids.drop_duplicates().reset_index()
removed_ids = removed_ids.drop(["index"], axis = 1)
removed_ids.columns = ["index"]
removed_ids
# Przygotuję ramkę danych z wierszami do usunięcia - aby odróżnić gdzie usuwam *it_quantity*, gdzie *it_seller_rating*, a gdzie oba - dodam dwie pomocnicze binarne kolumny *no_rating* i *no_quantity* zawierające informacje o brakach w danym wierszu.
# +
to_remove_1 = pd.DataFrame(data_4.iloc[removed_ids_1]).reset_index()
to_remove_2 = pd.DataFrame(data_4.iloc[removed_ids_2]).reset_index()
to_remove = pd.concat((pd.DataFrame(to_remove_1), pd.DataFrame(to_remove_2)), axis = 0)
to_remove = to_remove.drop_duplicates()
before_remove = to_remove.copy()
to_remove
# +
to_remove = to_remove_1.merge(to_remove, on = "index", how = "right")
to_remove["no_rating"] = np.where(to_remove["price_x"] == to_remove["price_x"], 1, 0)
to_remove = to_remove.drop(["it_seller_rating_x", "it_quantity_x", "price_x"], axis = 1)
to_remove.columns = ["index", "it_seller_rating", "it_quantity", "price", "no_rating"]
to_remove = to_remove_2.merge(to_remove, on = "index", how = "right")
to_remove["no_quantity"] = np.where(to_remove["price_x"] == to_remove["price_x"], 1, 0)
to_remove = to_remove.drop(["it_seller_rating_x", "it_quantity_x", "price_x"], axis = 1)
to_remove.columns = ["index", "it_seller_rating", "it_quantity", "price", "no_rating", "no_quantity"]
to_remove.sample(15)
# -
# Jak widać zwykle jest jeden brak, ale raz na jakiś czas w wierszu będzie informacja tylko o cenie.
#
# No dobra, czas usunąć informacje!
removed = to_remove.copy()
removed["it_quantity"] = np.where(removed["no_quantity"] == 1, np.nan, removed["it_quantity"])
removed["it_seller_rating"] = np.where(removed["no_rating"] == 1, np.nan, removed["it_seller_rating"])
removed = removed.drop(["no_rating", "no_quantity"], axis = 1)
removed.sample(15)
# Znowu wszystko zgodnie z planem, wiwat!!
filled = pd.DataFrame(imp.transform(removed.drop(["index"], axis = 1)))
filled.columns = ["it_seller_rating", "it_quantity", "price"]
filled["index"] = removed_ids
filled
sample_10_to_check = np.random.choice(len(removed), 10, replace=False)
sample_10_to_check
compare = before_remove.iloc[sample_10_to_check].merge(filled.iloc[sample_10_to_check], on = 'index')
compare = compare.drop(["index", "price_x", "price_y"], axis = 1)
compare.columns = ["it_seller_rating_correct", "it_quantity_correct", "it_seller_rating_predicted", "it_quantity_predicted"]
compare
# Ciekawe wrażenie, ale nagle wartości przybliżone wydają się na pierwszy rzut oka być bardziej zbliżone.
#
# To zapewne iluzja spowodowana przedstawieniem ich w innej postaci.
#
# Co na to miara RMSE?
rms = sqrt(mean_squared_error(before_remove["it_seller_rating"], filled["it_seller_rating"]))
rms
# Jednak zgodnie ze zdrowym rozsądkiem większa!
# Więcej braków <-> większy błąd
# Dokojamy automatyzacji - takiej samej, jak wcześniej.
deviatons2[0] = rms
for i in range(1, 10):
np.random.seed(random_seeds[i-1])
removed_ids_1 = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids_2 = np.random.choice(len(data_4), len(data_4) // 10, replace=False)
removed_ids_1.sort()
removed_ids_2.sort()
removed_ids = pd.concat((pd.DataFrame(removed_ids_1), pd.DataFrame(removed_ids_2)), axis = 0)
removed_ids = removed_ids.drop_duplicates().reset_index()
removed_ids = removed_ids.drop(["index"], axis = 1)
removed_ids.columns = ["index"]
to_remove_1 = pd.DataFrame(data_4.iloc[removed_ids_1]).reset_index()
to_remove_2 = pd.DataFrame(data_4.iloc[removed_ids_2]).reset_index()
to_remove = pd.concat((pd.DataFrame(to_remove_1), pd.DataFrame(to_remove_2)), axis = 0)
to_remove = to_remove.drop_duplicates()
before_remove = to_remove.copy()
to_remove = to_remove_1.merge(to_remove, on = "index", how = "right")
to_remove["no_rating"] = np.where(to_remove["price_x"] == to_remove["price_x"], 1, 0)
to_remove = to_remove.drop(["it_seller_rating_x", "it_quantity_x", "price_x"], axis = 1)
to_remove.columns = ["index", "it_seller_rating", "it_quantity", "price", "no_rating"]
to_remove = to_remove_2.merge(to_remove, on = "index", how = "right")
to_remove["no_quantity"] = np.where(to_remove["price_x"] == to_remove["price_x"], 1, 0)
to_remove = to_remove.drop(["it_seller_rating_x", "it_quantity_x", "price_x"], axis = 1)
to_remove.columns = ["index", "it_seller_rating", "it_quantity", "price", "no_rating", "no_quantity"]
removed = to_remove.copy()
removed["it_quantity"] = np.where(removed["no_quantity"] == 1, np.nan, removed["it_quantity"])
removed["it_seller_rating"] = np.where(removed["no_rating"] == 1, np.nan, removed["it_seller_rating"])
removed = removed.drop(["no_rating", "no_quantity"], axis = 1)
filled = pd.DataFrame(imp.transform(removed.drop(["index"], axis = 1)))
filled.columns = ["it_seller_rating", "it_quantity", "price"]
filled["index"] = removed_ids
rms = sqrt(mean_squared_error(before_remove["it_seller_rating"], filled["it_seller_rating"]))
deviatons2[i] = rms
deviatons2
statistics.stdev(deviatons2)
# Odchylenie standardowe wyszło wyraźnie nieco większe.
# ### Wnioski i wykresy
# Jak można się było spodziewać, **przewidywanie wartości brakujących wartości z kolumny mając do dyspozycji jedynie trzy słabo skorelowane ze sobą kolumki ramki danych nie przyniosło należytych efektów**.
#
# Fajnie, że profesjonalne napisane już narzędzie informatyczne daje "jakieś" wyniki, ale jako porządni analitycy danych powinniśmy mocno zastanowić się nad warunkami i danymi wejściowymi przed ich użyciem.
#
# Prawdopodobnie chociażby **gdyby skorzystać z większej ilości kolumn niż tylko tych trzech, wynik byłby o wiele bardziej satysfakcjonujący** - niestety w tym wypadku przydatnych danych było o wiele za mało.
#
# Biorąc pod uwagę wartości miary RMSE i odchylenia standardowe dla przypadów gdzie: a) w jednej z trzech kolumn brakuje 10% wartości b) w dwóch trzech kolumn następuje owy szum; można więc strzelać, że im więcej braków danych, tym przybliżony wynik działania **Multivariate feature imputation jest gorszy**, a także i **wyniki są coraz mniej bliskie rzeczywistości** - jest to na szczęście jak najbardziej zgodne z intuicją.
#
# Podsumowawczo zobaczmy jeszcze wyniki miar RMSE dla obu problemów - wypełniania 10% braków z 1 z 3 kolumn z ramki danych zestawione z wynikami tej miary dla usunięcia po 10% danych dla 2 kolumn (przy losowaniu ze zwracaniem).
# +
import matplotlib.pyplot as plt
plt.plot(deviatons2)
plt.plot(deviatons)
plt.ylim(0, 50000)
plt.xlabel("Number of test")
plt.ylabel("Value of RMSE")
plt.title("RMSE deviations of imputations two data frames by multivariate feature imputation")
plt.show()
# -
# Jak będą wyglądać dane dla pierwotnego 3-kolumnowego zbioru, a jak dla uzupełnionymi brakami danych dla jednego z dziesięciu przypadków usunięcia 10% wartości *it_quantity* i 10% *it_seller_rating*?
# Przed usunięciem danych:
# +
plt.scatter(before_remove[["it_quantity"]], before_remove[["it_seller_rating"]], s = 1, alpha = 0.5)
plt.title("Allegro original data")
plt.xlabel("quantity")
plt.ylabel("seller rating")
plt.xlim(-100000, 100000)
plt.show()
# +
plt.scatter(filled[["it_quantity"]], filled[["it_seller_rating"]], s = 1, alpha = 0.5)
plt.title("Allegro filled data")
plt.xlabel("quantity")
plt.ylabel("seller rating")
plt.xlim(-100000, 100000)
plt.show()
# -
# Wizualizacje jeszcze konkretniej ukazują, że tym bardziej robotę zaimplementowanego *multivariate feature imputation* można wyrzucić do kosza - niedość, że dane są mocno skumulowane wokół dwóch prostych (jednej zbliżonej do x = 0, a drugiej do lekko pochylonej w prawo y = 0), to ponadto mamy styczność z pojedynczymi wartościami ujemnymi dla *it_quantity*... które docelowo są nieujemne. Tak czy inaczej trudno, żeby na podstawie tak małej ilości danych algorytm zrobił coś sensowniejszego - problem był po prostu nie do ogarnięcia korzystając z dostępnych narzędzi i danych.
# To by było na tyle. Dobra praca domowa, fajnie się bawiłem, dziękuję!
| Prace_domowe/Praca_domowa2/Grupa2/Kosterna_Jakub/pd2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #Cosmic integration
# ## 1. Introduction
#
# This section contains a series of tutorials on how to use and get the most out of the COMPAS Cosmic Integration tools. The COMPAS Cosmic Integration tools are based on work done in [Neijssel et al. 2019 ](https://arxiv.org/abs/1906.08136).
#
# The duration from the birth of the binary until the merger as a double compact object (DCO) can range from a few million years (lifetime of the stars) to more than 100 Gigayears depending on the evolution of the system.
#
# This means two things:
#
# 1. DCOs merge at different redshifts
#
# 2. Multiple DCOs merging at a specific redshift could have formed at different times
#
# We thus need to know the star formation that went into forming a single system. However, the star formation rate is non-constant over the lifetime of the universe. Furthermore, star formation is heavily dependent on the metallicity of the star forming gas, which also changes over the lifetime of the universe. Combined, we call this the metallicity-specific star formation rate (MSSFR).
#
# The cosmic-integration pipeline tries to predict the population of DCO mergers assuming a model for the MSSFR. Here we show how to use the combination of scripts: we show the purpose of each script in the pipeline, and how to combine them to derive our results. These notes show how to call the functions, and how to construct the pipeline: they do not offer any derivations. We assume that you have a COMPAS data set in `HDF5` file format. If not, see [COMPAS HDF5 files](../H5/0_H5Overview.ipynb).
#
# Note that although these pipelines are tailored for a COMPAS simulation, with some slight adjustments one could use these same pipelines for other (mock-)simulations, provided all the information is given (see section 7 of [Setting the Data](./1_ClassCOMPAS.ipynb)).
#
#
# If you make use of any these pipelines, we would appreciate you citing the following papers:
#
# For cosmic integration: [Neijssel et al. 2019 ](https://arxiv.org/abs/1906.08136)
#
# For selection effects: [Barrett et al. 2018](https://arxiv.org/pdf/1711.06287)
# ## Table of contents
#
# ### [Setting the data](./1_ClassCOMPAS.ipynb)
# Set the required columns and other information.
#
# ### [Setting the MSSFR](./2_MSSFR-or-SFRD-prescriptions.ipynb)
# Choose a model and plot it.
#
# ### [Setting the selection effects:](./3_SelectionEffects.ipynb)
# Choose the sensitivity and calculate probability.
#
# ### [Rate at a single redshift](./4_MergersAtSingleRedshift.ipynb)
# Combine data, MSSFR, and selection effects.
#
# ### [Rate as function of redshift](./5_MergersAsFunctionOfRedhisft.ipynb)
# Set the integral and combine results.
#
#
# ### [Fast cosmic integration](./6_FastCosmicIntegrator.ipynb)
# Do the whole routine in one go, using the phenomenological variation of the MSSFR and SFRD
| docs/online-docs/notebooks/cosmicIntegration/0_CosmicIntegrationOverview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.9 64-bit
# name: python3
# ---
# + [markdown] id="TA21Jo5d9SVq"
#
#
# 
#
# [](https://githubtocolab.com/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/streamlit_notebooks/NER_PT.ipynb)
#
#
#
# + [markdown] id="CzIdjHkAW8TB"
# # **Detect legal entities in Portuguese text**
# + [markdown] id="wIeCOiJNW-88"
# ## 1. Colab Setup
# + id="j2JGGTdMdovR"
import os
import json
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
sparknlp_version = license_keys["PUBLIC_VERSION"]
jsl_version = license_keys["JSL_VERSION"]
# + colab={"base_uri": "https://localhost:8080/"} id="vMaVR-rnldBM" outputId="3fa03fea-d6b7-45f6-b460-ac6fb3eec364"
print ('SparkNLP Version:', sparknlp_version)
print ('SparkNLP-JSL Version:', jsl_version)
# + [markdown] id="LAowjxBjd9Dt"
# Install dependencies
# + id="CGJktFHdHL1n"
# %%capture
for k,v in license_keys.items():
# %set_env $k=$v
# !wget https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/jsl_colab_setup.sh
# !bash jsl_colab_setup.sh
# Install Spark NLP Display for visualization
# !pip install --ignore-installed spark-nlp-display
# + [markdown] id="eCIT5VLxS3I1"
# ## 2. Start the Spark session
# + id="sw-t1zxlHTB7" colab={"base_uri": "https://localhost:8080/", "height": 214} outputId="2bcc653a-3b64-4ab9-8be9-62d2f764e437"
import json
import pandas as pd
import numpy as np
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
from sparknlp.annotator import *
from sparknlp.base import *
import sparknlp
from sparknlp.pretrained import PretrainedPipeline
import sparknlp_jsl
from sparknlp_jsl.annotator import *
spark = sparknlp_jsl.start(license_keys['SECRET'])
spark
# + [markdown] id="9RgiqfX5XDqb"
# ## 3. Select the DL model
# + id="LLuDz_t40be4"
# If you change the model, re-run all the cells below.
# Applicable models: lener_bert_base, lener_bert_large
MODEL_NAME = "lener_bert_base"
# + [markdown] id="2Y9GpdJhXIpD"
# ## 4. Some sample examples
# + id="vBOKkB2THdGI"
# Enter examples to be transformed as strings in this list
text_list = [
"""a primeira câmara desta corte , em acórdão constante da relação 31/2000 , ata 27/2000 , ministro marcos vinicios vilaça , julgou regulares com ressalva as contas de carlos aureliano motta de souza , ex-diretor-geral do stm , no ano de 1999 ( peça 1 , p. 49-51 ) .""",
"""com isso , a corte , por meio da decisão 877/2000 – plenário , manifestou-se nos seguintes termos : 8.1 - determinar à secex/rj que , com base nos artigos 41 e 43 , inciso ii , da lei nº 8.443/92 : 8.1.1 - promova a audiência do responsável acima identificado , para que , no prazo regimental , apresente razões de justificativa quanto a ocorrência de antecipações de pagamentos para fornecimento de esquadrias de alumínio , ar-condicionado e elevadores da obra de construção do edifício da 1ª cjm/rj , em afronta aos artigos 62 e 63 da lei nº 4.320/64 ; 38 do decreto nº 93.872/86 ; e 65 , inciso ii , letra `` c '' , da lei nº 8.666/93 , bem como quanto às alterações contratuais ."""
]
# + [markdown] id="XftYgju4XOw_"
# ## 5. Define Spark NLP pipeline
# + id="lBggF5P8J1gc" colab={"base_uri": "https://localhost:8080/"} outputId="10cf5a7c-06e8-4b81-fc14-34a42c04ad27"
document_assembler = DocumentAssembler() \
.setInputCol('text') \
.setOutputCol('document')
tokenizer = Tokenizer() \
.setInputCols(['document']) \
.setOutputCol('token')
# The model was trained with the Bert embeddings, we need to it.if MODEL_NAME == 'lener_bert_base': bert_model = 'bert_portuguese_base_cased'else: bert_model = 'bert_portuguese_large_cased'
embeddings = BertEmbeddings.pretrained(bert_model, 'pt')\
.setInputCols("document", "token") \
.setOutputCol("embeddings")
ner_model = MedicalNerModel.pretrained(MODEL_NAME, 'pt', 'clinical/models') \
.setInputCols(['document', 'token', 'embeddings']) \
.setOutputCol('ner')
ner_converter = NerConverter() \
.setInputCols(['document', 'token', 'ner']) \
.setOutputCol('ner_chunk')
nlp_pipeline = Pipeline(stages=[
document_assembler,
tokenizer,
embeddings,
ner_model,
ner_converter
])
# + [markdown] id="mv0abcwhXWC-"
# ## 6. Run the pipeline
# + id="EYf_9sXDXR4t"
empty_df = spark.createDataFrame([['']]).toDF('text')
pipeline_model = nlp_pipeline.fit(empty_df)
df = spark.createDataFrame(pd.DataFrame({'text': text_list}))
result = pipeline_model.transform(df)
# + [markdown] id="UQY8tAP6XZJL"
# ## 7. Visualize results
# + id="Ar32BZu7J79X" colab={"base_uri": "https://localhost:8080/", "height": 159} outputId="bc5b9b05-22c5-4c19-f7d7-d9e3b4c3d921"
from sparknlp_display import NerVisualizer
NerVisualizer().display(
result = result.collect()[0],
label_col = 'ner_chunk',
document_col = 'document'
)
# + id="-61DDZx9nEY0"
| tutorials/streamlit_notebooks/healthcare/NER_LEGAL_PT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from terminusdb_client import WOQLClient, WOQLQuery
db_id = "nlp_spacy"
client = WOQLClient(server_url = "http://localhost:6363")
client.connect(key="root", account="admin", user="admin")
client.db(db_id)
# +
import spacy
#import pprint as pp
def doc_id(doc, var_name):
return WOQLQuery().idgen("doc:SpaCyDoc",
[WOQLQuery().string(doc.text)], var_name)
def token_id(doc, token, var_name):
return WOQLQuery().idgen("doc:SpaCyToken",
[WOQLQuery().string(doc.text),
WOQLQuery().string(str(token.i))], var_name)
nlp = spacy.load("en_core_web_sm")
doc = nlp("Autonomous cars shift insurance liability toward manufacturers")
insert_doc = doc_id(doc, "v:doc_id") + WOQLQuery().insert("v:doc_id", "scm:SpaCyDoc", label=doc.text)
insert_doc.execute(client, "Add in the doc")
all_token = []
for token in doc:
print(token.text, token.lemma_, token.pos_, token.tag_, token.dep_,
token.shape_, token.is_alpha, token.is_stop)
insert_token = (WOQLQuery().insert(f"v:token_id_self{token.i}", "scm:SpaCyToken", label=token.text)
.property("lemma", token.lemma_)
.property("pos", token.pos_)
.property("tag", token.tag_)
.property("dep", token.dep_)
.property("shape", token.shape_)
.property("is_alpha", token.is_alpha)
.property("is_stop", token.is_stop)
.property("head", f"v:token_id_head{token.i}")
.property("doc", f"v:token_doc_id{token.i}")
)
all_token.append(
token_id(doc, token, f"v:token_id_self{token.i}")
+ token_id(doc, token.head, f"v:token_id_head{token.i}")
+ doc_id(token.doc, f"v:token_doc_id{token.i}")
+ insert_token
)
WOQLQuery().woql_and(*all_token).execute(client, "Add in all the tokens")
# -
for token in doc:
print(token.text, token.head.text)
# +
# Delete everything
# WOQLQuery().triple("v:A","v:B","v:C").delete_triple("v:A","v:B","v:C").execute(client, "Delete everything")
# +
import spacy
nlp = spacy.load("en_core_web_sm")
doc = nlp("Autonomous cars shift insurance liability toward manufacturers")
for token in doc:
print(token.text, token.dep_, token.head.text, token.head.pos_,
[child for child in token.children])
# -
| nlp-spacy/loading_spacy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import matplotlib
import matplotlib.pyplot as plt
from datasets import get_dataset
import numpy as np
np.random.seed(123)
import random
random.seed(123)
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.5
set_session(tf.Session(config=config))
from keras.utils import plot_model
from keras.models import Sequential, load_model
from keras.layers import Dense, Activation, Flatten, BatchNormalization, Dropout, Reshape
from keras.optimizers import Adadelta, SGD
from keras.callbacks import TensorBoard, EarlyStopping, ModelCheckpoint
from sklearn.preprocessing import label_binarize
import cv2
import pdb
import progressbar
import os
from scipy import stats
from sklearn.metrics import accuracy_score
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import accuracy_score
# -
n_views = 2
dataset = 'australian'
# 1. Partition L into (v_1, v_2, ..., v_n)
def get_dset(dataset):
ds = get_dataset(dataset, 0.7, 0.25)
[L_x, L_y], U, [test_x, test_y] = ds.get_data()
L_y = np.argmax(L_y, axis=1)
test_y = np.argmax(test_y, axis=1)
V = []
for ind in range(n_views):
left = int(ind * L_x.shape[0] / n_views)
right = int((ind+1) * L_x.shape[0] / n_views)
V.append([L_x[left:right], L_y[left:right]])
return ds, [L_x, L_y], U, [test_x, test_y], V
| co-learning.ipynb.LOCAL.8140.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#
# MultiLayerPerceptron Image classifier
#
# @author becxer
# @email <EMAIL>
# @reference https://github.com/sjchoi86/Tensorflow-101
#
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# %matplotlib inline
print ("packages are loaded")
# +
# Load npz data
npz_path = "images/MYIMG/my_img.npz"
load_data = np.load(npz_path)
print ("Load data : " + str(load_data.files))
train_img = load_data['train_img']
train_label = load_data['train_label']
valid_img = load_data['valid_img']
valid_label = load_data['valid_label']
test_img = load_data['test_img']
test_label = load_data['test_label']
print ("train_img shape : " + str(train_img.shape))
print ("valid_img shape : " + str(valid_img.shape))
print ("test_img shape : " + str(test_img.shape))
# +
# Plot image
rand_idx = np.arange(train_img.shape[0])
np.random.shuffle(rand_idx)
rand_idx = rand_idx[:5]
for idx in rand_idx:
label = np.argmax(train_label[idx])
img = np.reshape(train_img[idx], (64,64))
plt.matshow(img,cmap=plt.get_cmap('gray'))
plt.colorbar()
plt.title("Label : " + str(label))
plt.show()
# +
# Options for training
learning_rate = 0.001
training_epochs = 500
batch_size = 100
display_step = 100
# Options for Hidden Network Nodes
n_hidden = [1000, 500, 200] # hidden layer's node count
w_dev = 0.1
# Options for dropout
drop_out_ratio = 1.0
# +
# Build Graph of Dropout MLP
# Define placeholder & Variables
x = tf.placeholder("float", [None, train_img.shape[1]])
y = tf.placeholder("float", [None, train_label.shape[1]])
drop_out_prob = tf.placeholder("float")
def one_layer(_x, _W, _b, _dop):
return tf.nn.dropout(tf.nn.sigmoid(tf.add(tf.matmul(_x, _W),_b)),_dop)
WS = {}
BS = {}
last_input_layer = x
last_input_layer_size = train_img.shape[1]
for idx, hl_size in enumerate(n_hidden):
_W = tf.Variable(tf.random_normal([last_input_layer_size, hl_size], stddev=w_dev))
_b = tf.Variable(tf.random_normal([hl_size]))
last_input_layer = one_layer(last_input_layer, _W, _b, drop_out_prob)
last_input_layer_size = hl_size
WS['h_' + str(idx)] = _W
BS['b_' + str(idx)] = _b
WS['out'] = tf.Variable(tf.random_normal([last_input_layer_size, train_label.shape[1]], stddev=w_dev))
BS['out'] = tf.Variable(tf.random_normal([train_label.shape[1]], stddev=w_dev))
# Define operators
out = tf.add(tf.matmul(last_input_layer, WS['out']),BS['out'])
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(out, y))
optm = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
pred = tf.argmax(out, 1)
accr = tf.reduce_mean(tf.cast(tf.equal(pred, tf.argmax(y, 1)),"float"))
init = tf.initialize_all_variables()
print ("Graph build")
# +
# Training Graph
sess = tf.Session()
sess.run(init)
for epoch in range(training_epochs):
avg_cost = 0.
num_batch = int(train_img.shape[0]/batch_size)
for i in range(num_batch):
randidx = np.random.randint(train_img.shape[0], size=batch_size)
batch_xs = train_img[randidx, :]
batch_ys = train_label[randidx, :]
sess.run(optm, feed_dict={x: batch_xs, y: batch_ys, drop_out_prob : drop_out_ratio})
avg_cost += sess.run(cost, feed_dict={x: batch_xs, y: batch_ys, drop_out_prob : 1.})/num_batch
if epoch % display_step == 0:
train_img_acc = sess.run(accr , ({x: batch_xs, y: batch_ys, drop_out_prob : 1.}))
print ("epoch: %03d/%03d , cost: %.6f , train_img_acc: %.3f" \
% (epoch, training_epochs, avg_cost, train_img_acc))
test_batch_size = 10
avg_acc = 0.
num_batch_test = int(test_img.shape[0]/test_batch_size)
for i in range(num_batch_test):
batch_xs_test = test_img[i * test_batch_size : (i+1) * test_batch_size ]
batch_ys_test = test_label[i * test_batch_size : (i+1) * test_batch_size ]
avg_acc += sess.run(accr, feed_dict={x : batch_xs_test, y : batch_ys_test, drop_out_prob : 1.})
print ("Training complete, Accuracy : %.6f" \
% (avg_acc / num_batch_test,))
# -
| Jupyter/015_MLP_MYIMG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="2iYLQO5Evvqy"
# # Training a `Robust' Adapter with AdapterDrop
#
# This notebook extends our quickstart adapter training notebook to illustrate how we can use AdapterDrop
# to robustly train an adapter, i.e. adapters that allow us to dynmically dropp layers for faster multi-task inference.
# Please have a look at the original adapter training notebook for more details on the setup.
# + [markdown] id="a-XTIOLv0isn"
# ## Installation
#
# First, let's install the required libraries:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ju-alwbHmKYA" outputId="44bbe24a-0925-46c2-aaf8-d9b5cdd7d0e3"
# !pip install -U adapter-transformers
# !pip install datasets
# + colab={"base_uri": "https://localhost:8080/"} id="7Mx916lBCfoL" outputId="bec39206-a7d9-415a-a3fa-90b948d51489" pycharm={"name": "#%%\n"}
from datasets import load_dataset
from transformers import RobertaTokenizer
dataset = load_dataset("rotten_tomatoes")
tokenizer = RobertaTokenizer.from_pretrained("roberta-base")
def encode_batch(batch):
"""Encodes a batch of input data using the model tokenizer."""
return tokenizer(batch["text"], max_length=80, truncation=True, padding="max_length")
# Encode the input data
dataset = dataset.map(encode_batch, batched=True)
# The transformers model expects the target class column to be named "labels"
dataset.rename_column_("label", "labels")
# Transform to pytorch tensors and only output the required columns
dataset.set_format(type="torch", columns=["input_ids", "attention_mask", "labels"])
# + [markdown] id="S2-2CbfPGYvi"
# ## Training
# + colab={"base_uri": "https://localhost:8080/"} id="Tp9uG-pT-qgv" outputId="c6a19f25-26b9-44c7-887c-1d1a71e957fa"
from transformers import RobertaConfig, RobertaModelWithHeads
config = RobertaConfig.from_pretrained(
"roberta-base",
num_labels=2,
id2label={ 0: "👎", 1: "👍"},
)
model = RobertaModelWithHeads.from_pretrained(
"roberta-base",
config=config,
)
# Add a new adapter
model.add_adapter("rotten_tomatoes")
# Add a matching classification head
model.add_classification_head("rotten_tomatoes", num_labels=2)
# Activate the adapter
model.train_adapter("rotten_tomatoes")
# + [markdown] id="ev5t_8i8HzJB"
# To dynamically drop adapter layers during training, we make use of HuggingFace's `TrainerCallback'.
# + id="5FRft_5AAlQd"
import numpy as np
from transformers import TrainingArguments, Trainer, EvalPrediction, TrainerCallback
class AdapterDropTrainerCallback(TrainerCallback):
def on_step_begin(self, args, state, control, **kwargs):
skip_layers = list(range(np.random.randint(0, 11)))
kwargs['model'].set_active_adapters("rotten_tomatoes", skip_layers=skip_layers)
def on_evaluate(self, args, state, control, **kwargs):
# Deactivate skipping layers during evaluation (otherwise it would use the
# previous randomly chosen skip_layers and thus yield results not comparable
# across different epochs)
kwargs['model'].set_active_adapters("rotten_tomatoes", skip_layers=None)
training_args = TrainingArguments(
learning_rate=1e-4,
num_train_epochs=6,
per_device_train_batch_size=32,
per_device_eval_batch_size=32,
logging_steps=200,
output_dir="./training_output",
overwrite_output_dir=True,
remove_unused_columns=False
)
def compute_accuracy(p: EvalPrediction):
preds = np.argmax(p.predictions, axis=1)
return {"acc": (preds == p.label_ids).mean()}
trainer = Trainer(
model=model,
args=training_args,
train_dataset=dataset["train"],
eval_dataset=dataset["validation"],
compute_metrics=compute_accuracy,
)
trainer.add_callback(AdapterDropTrainerCallback())
# + [markdown] id="9iHhoYuLIdX3" pycharm={"name": "#%% md\n"}
# We can now train and evaluate our robustly trained adapter!
# + colab={"base_uri": "https://localhost:8080/", "height": 514} id="zZxaujENntNR" outputId="6700e4ac-1258-4bd8-ac30-a578c8d6c4ba" pycharm={"name": "#%%\n"}
trainer.train()
trainer.evaluate()
| notebooks/05_Adapter_Drop_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# __Analyzing Stock Prices__
#
# ---
# In this project, we'll look at stock market data from the yahoo_finance Python package.
#
# This data consists of the daily stock prices from 2007-1-1 to 2017-04-17 for several hundred stock symbols traded on the NASDAQ stock exchange, stored in the prices folder.
#
# The download_data.py script in the same folder as the Jupyter notebook was used to download all of the stock price data.
#
# Each file in the prices folder is named for a specific stock symbol, and contains the:
#
# - date -- date that the data is from.
# - close -- the closing price on that day, which is the price when the trading day ends.
# - open -- the opening price on that day, which is the price when the trading day starts.
# - high -- the highest price the stock reached during trading.
# - low -- the lowest price the stock reached during trading.
# - volume -- the number of shares that were traded during the day.
import os, pandas as pd
print("Files in directory: ", len(os.listdir('prices/')))
print('Rows across all directory files:' ,
(sum([ pd.read_csv("prices/" + i).shape[0] for i in os.listdir('prices/')])))
# Create dictionary of dataframes for directory files
directory = 'prices'
stocks = {}
for file in os.listdir(directory):
key = file.split('.')[0]
val = pd.read_csv(os.path.join(directory, file))
stocks[key] = val
stocks['aapl'].head()
# __Closing Prices__
# +
# Average closing price
closing_mean = {}
for df in stocks.keys():
close = stocks[df]['close'].mean()
closing_mean.update({df: close})
closing_mean = pd.DataFrame(data = closing_mean.values(),
index = closing_mean.keys(),
columns = ['close_mean'])
closing_mean = closing_mean.sort_values(by = 'close_mean', ascending = False)
# -
print('Top 5')
closing_mean.head().round(2)
print('Bottom 5')
closing_mean.tail().round(2)
# __Trades Per Day__
# +
day_trades = {}
for i in stocks.keys():
for ind, row in stocks[i].iterrows():
day = row['date']
vol = row['volume']
pair = (vol, i)
if day not in day_trades:
day_trades[day] = []
day_trades[day].append(pair)
# -
# __Most Traded Stock Per Day__
# +
top_daily = {}
for i in day_trades:
day_trades[i].sort()
top_daily[i] = day_trades[i][-1]
# -
# __High Volume Days__
# +
vols = []
for i in day_trades:
day_vol = sum([vol for vol, _ in day_trades[i]])
vols.append((round(day_vol,2), i))
vols.sort()
vols[-10:]
# -
# __Most Profitable Stocks__
# +
pct_change = []
for i in stocks:
data = stocks[i]
start = data.loc[0, 'close']
end = data.loc[len(data.index)-1, 'close']
pct = round(100 * (end - start) / start,2)
pct_change.append((pct, i))
pct_change.sort()
pct_change[-10:]
# -
# Within this dataset, the most profitable stock to have purchased wsa teh ADMP stock, growing by almost 8000% in value over the period.
#
# Potential further analysis:
# - What stocks would have been best to short at the start of the period?
# - Which stocks have the most after-hours trading, and show the biggest changes between the closing price and the next day open?
# - Can technical indicators like Bollinger Bands help us forecast the market?
# - What time periods have resulted in steady increases in prices, and what periods have resulted in steady declines?
# - Based on price, what was the optimal day to buy each stock if we wanted to hold them until now?
# - On days with high trading volume, do stocks move in one direction (up or down) more than the other one?
#
#
| i. Data Structures/AnalysingStocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting a full anp model from excel
# ## Importing needed libraries
import pandas as pd
import numpy as np
from pyanp.anp import ANPNetwork, anp_from_excel, get_matrix, is_pw_col_name, is_rating_col_name
# %pylab inline
# # Load in the data
exfile = "anp_data_ratings_manual_scales.xlsx"
anp = anp_from_excel(exfile)
# Let's verify our data actually has the manual scale defined
n = anp.node_prioritizer('n4-4', 'Cluster 4')
display(n.word_eval.values)
# # Let's calculate
# ## Global priorities for the group
anp.global_priority()
# ## Alternative scores for the group
anp.priority()
anp.priority().plot(kind="bar")
# ## Alternative scores for User 1
anp.priority(username="User 1")
anp.priority(username="User 1").plot(kind="bar")
anp.priority(username="User 2")
anp.priority(username="User 2").plot(kind="bar")
n = anp.node_obj("n4-4")
r= n.get_node_prioritizer("n4-1")
n.get_unscaled_column(username="User 1")
n.node_prioritizers
is_pw_col_name("a wrt b")
n.get_unscaled_column()
| examples/ANPModelFromExcelWithRatingManualScales.ipynb |
# # Interpreting predictions on MNIST
# +
# general imports
import warnings; warnings.filterwarnings("ignore", category=FutureWarning)
import tensorflow as tf; tf.logging.set_verbosity(tf.logging.ERROR) # suppress deprecation messages
import tempfile
import numpy as np
from matplotlib import pyplot as plt
from depiction.core import Task, DataType
# -
# plotting
_ = plt.gray()
plt.rcParams['figure.figsize'] = [20, 10]
# general variables
# NOTE: get a valid cache (but only once!)
CACHE_DIR = None
if CACHE_DIR is None:
CACHE_DIR = tempfile.mkdtemp()
# +
# general utils
def transform(x):
"""
Move to -0.5, 0.5 range and add channel dimension.
Args:
x (np.ndarray): a 2D-array representing an image.
Returns:
np.ndarray: a 3D-array representing the transformed image.
"""
return np.expand_dims(x.astype('float32') / 255 - 0.5, axis=-1)
def transform_sample(x):
"""
Add dimension representing the batch size.
Args:
x (np.ndarray): a 3D-array represnting an image.
Returns:
np.ndarray: a 4D-array representing a batch with a
single image.
"""
return np.expand_dims(transform(x), axis=0)
def inverse_transform(x):
"""
Apply an inverse transform on a batch with a single image.
Args:
x (np.ndarray): a 4D-array representing a batch with a
single image.
Returns:
np.ndarray: a 3D-array represnting an image.
"""
return (x.squeeze() + 0.5) * 255
def show_image(x, title=None):
"""
Show an image.
Args:
x (np.ndarray): a 4D-array representing a batch with a
single image.
title (str): optional title.
"""
axes_image = plt.imshow(x.squeeze())
axes_image.axes.set_xticks([], [])
axes_image.axes.set_yticks([], [])
if title is not None:
axes_image.axes.set_title(title)
return axes_image
# -
# ## Instantiate a model to intepret
# +
from depiction.models.keras import KerasModel
from tensorflow.keras.models import load_model
from depiction.models.base.utils import get_model_file
depiction_model = KerasModel(
load_model(
get_model_file(
filename='mninst_cnn.h5',
origin='https://ibm.box.com/shared/static/v3070m2y62qw4mpwl04pee75n0zg681g.h5',
cache_dir=CACHE_DIR
)
),
task=Task.CLASSIFICATION, data_type=DataType.IMAGE
)
depiction_model._model.summary()
# -
# ## Get data
from tensorflow.keras.datasets.mnist import load_data
(x_train, y_train), (x_test, y_test) = load_data()
print('x_train shape:', x_train.shape, 'y_train shape:', y_train.shape)
print('x_test shape:', x_test.shape, 'y_test shape:', y_test.shape)
index = 42
example = transform_sample(x_test[index])
label = y_test[index]
show_image(
example,
title=(
f'True={label} '
f'Predicted={np.argmax(depiction_model.predict(example))}'
)
)
# ## How is our model doing?
from tensorflow.keras.utils import to_categorical
score = depiction_model._model.evaluate(
transform(x_test), to_categorical(y_test), verbose=0
)
print(f'Test accuracy: {score[1]}')
# ## LIME
# +
from depiction.interpreters.u_wash import UWasher
lime_interpreter = UWasher('lime', depiction_model)
# -
lime_interpreter.interpret(example)
# ## CEM
# some utilities
def show_cem_explanation(explanation, mode):
"""
Show a CEM explanation for images.
Args:
explanation (dict): CEM explanation.
mode (str): CEM mode, PP or PN.
"""
prediction_key = f'{mode}_pred'
if prediction_key in explanation:
print(f'{mode} prediction: {explanation[prediction_key]}')
show_image(
explanation[mode],
title=(f'{mode} explanation for example provided.')
)
# setting some parameters
shape = example.shape
kappa = 0. # minimum difference needed between the prediction probability for the perturbed instance on the
# class predicted by the original instance and the max probability on the other classes
# in order for the first loss term to be minimized
beta = .1 # weight of the L1 loss term
gamma = 100 # weight of the optional auto-encoder loss term
c_init = 1. # initial weight c of the loss term encouraging predictions for the perturbed instance compared to the original instance to be explained
c_steps = 10 # updates for c
max_iterations = 10 # iterations per value of c
feature_range = (x_train.min(), x_train.max()) # feature range for the perturbed instance
clip = (-1000., 1000.) # gradient clipping
lr = 1e-2 # initial learning rate
no_info_val = -1. # picking value close to background
# NOTE: CEM supports the usage of an autoencoder to impose a variation on a latent manifold
ae = load_model(
get_model_file(
filename='mninst_ae.h5',
origin=
'https://ibm.box.com/shared/static/psogbwnx1cz0s8w6z2fdswj25yd7icpi.h5', # noqa
cache_dir=CACHE_DIR
)
)
ae.summary()
# +
from depiction.interpreters.alibi import CEM
cem_pn_interpreter = CEM(
depiction_model,
'PN', # pertinent negative
shape,
kappa=kappa,
beta=beta,
feature_range=feature_range,
gamma=gamma,
ae_model=ae,
max_iterations=max_iterations,
c_init=c_init,
c_steps=c_steps,
learning_rate_init=lr,
clip=clip,
no_info_val=no_info_val
)
# -
cem_pn_explanation = cem_pn_interpreter.interpret(example)
show_cem_explanation(cem_pn_explanation, 'PN')
# +
from depiction.interpreters.alibi import CEM
cem_pp_interpreter = CEM(
depiction_model,
'PP', # pertinent positive
shape,
kappa=kappa,
beta=beta,
feature_range=feature_range,
gamma=gamma,
ae_model=ae,
max_iterations=max_iterations,
c_init=c_init,
c_steps=c_steps,
learning_rate_init=lr,
clip=clip,
no_info_val=no_info_val
)
# -
cem_pp_explanation = cem_pp_interpreter.interpret(example)
show_cem_explanation(cem_pp_explanation, 'PP')
# ## Counterfactual explanations
def show_counterfactual_explanation(explanation):
"""
Show a counterfactual explanation for images.
Args:
explanation (dict): counterfactual explanation.
"""
predicted_class = explanation['cf']['class']
probability = explanation['cf']['proba'][0][predicted_class]
print(f'Counterfactual prediction: {predicted_class} with probability {probability}')
show_image(explanation['cf']['X'])
# setting some parameters
shape = example.shape
target_proba = 1.0
tol = 0.1 # tolerance for counterfactuals
max_iter = 10
lam_init = 1e-1
max_lam_steps = 10
learning_rate_init = 0.1
feature_range = (x_train.min(),x_train.max())
# +
from depiction.interpreters.alibi import Counterfactual
counterfactual_interpreter = Counterfactual(
depiction_model,
shape=shape, target_proba=target_proba, tol=tol,
target_class='other', # any other class
max_iter=max_iter, lam_init=lam_init,
max_lam_steps=max_lam_steps, learning_rate_init=learning_rate_init,
feature_range=feature_range
)
# -
counterfactual_explanation = counterfactual_interpreter.interpret(example)
show_counterfactual_explanation(counterfactual_explanation)
# +
from depiction.interpreters.alibi import Counterfactual
counterfactual_interpreter = Counterfactual(
depiction_model,
shape=shape, target_proba=target_proba, tol=tol,
target_class=1, # focusing on a specific class
max_iter=max_iter, lam_init=lam_init,
max_lam_steps=max_lam_steps, learning_rate_init=learning_rate_init,
feature_range=feature_range
)
# -
counterfactual_explanation = counterfactual_interpreter.interpret(example)
show_counterfactual_explanation(counterfactual_explanation)
| workshops/20191120_ODSC2019/notebooks/mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import warnings
origen_libs = '/home/scopatz/origen22/libs'
tape9 = 'pwru50.lib'
lib = os.path.join(origen_libs, tape9)
from pyne import utils
utils.toggle_warnings()
warnings.simplefilter('ignore')
from pyne.origen22 import parse_tape9
from pyne import nucname
from pyne import rxname
t9 = parse_tape9(lib)
nlb = tuple(sorted(t9.keys()))
nlb
set(t9[220].keys())
t9[219]['sigma_gamma'][30060], t9[221]['sigma_gamma'][30060]
XS_RXS = ['gamma', 'z_2n', 'z_3n', 'alpha', 'fission', 'proton', 'gamma_1', 'z_2n_1']
xs_2_o = {'gamma': 'gamma', 'z_2n': '2n', 'z_3n': '3n', 'alpha': 'alpha', 'fission': 'f', 'proton': 'p',
'gamma_1': 'gamma_x', 'z_2n_1': '2n_x'}
o_2_xs = {v: k for k, v in xs_2_o.items()}
sigma_maps = {}
nucs = set()
for n in nlb:
for rx in t9[n]:
if not rx.startswith('sigma_'):
continue
for nuc in t9[n][rx]:
if nuc not in sigma_maps:
sigma_maps[nuc] = {}
sigma_maps[nuc]['sigma_' + o_2_xs[rx[6:]]] = t9[n][rx][nuc]
nucs.add(nucname.name(int(nuc)))
# add absorption xs
for nuc in sigma_maps:
sigma_maps[nuc]['sigma_a'] = sum(sigma_maps[nuc].values())
nucs.add(nucname.name(int(nuc)))
nucname.name(nucname.zzaaam_to_id(10010))
sigma = {}
skip_rx = {'sigma_a', 'sigma_fission'}
for nuc in sigma_maps:
n = nucname.zzaaam_to_id(int(nuc))
name = nucname.name(n)
for srx in sigma_maps[nuc]:
if srx in skip_rx:
child = None
else:
_, _, rx = srx.partition('_')
print(name, rx, sigma_maps[nuc][srx])
child = nucname.name(rxname.child(n, rx))
sigma[srx + '_' + name] = [sigma_maps[nuc][srx], name, child]
nucs.add(name)
nucs.add(child)
child = rxname.child
sigma
import json
with open('sigma.json', 'w') as f:
json.dump(sigma, f, indent=' ', sort_keys=True)
with open('transmute_data.json', 'r') as f:
data = json.load(f)
nucs.discard(None)
nucs |= set(data['nucs'])
data['nucs'] = sorted(nucs, key=nucname.id)
with open('transmute_data.json', 'w') as f:
json.dump(data, f, indent=' ', sort_keys=True)
| tape9-to-sigma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Implementation of subspace alignment
#
# This is based on the following paper: <i>Unsupervised Visual Domain Adaptation Using Subspace Alignment</i>.
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA
import tensorflow_hub as hub
import warnings
warnings.filterwarnings("ignore")
# -
## feature extraction using pretrained feature extractor
universal_embed = hub.load("https://tfhub.dev/google/universal-sentence-encoder/4")
nnlm_embed = hub.load("https://tfhub.dev/google/nnlm-en-dim128/2")
# +
## getting the data from the different distributions
def get_data(file_name,data_dir="../data/sentiment/"):
""" returns the sentiment data for the given file
"""
all_text = []
all_labels = []
with open(data_dir+file_name) as data_file:
for line in data_file.readlines():
text,label = line.split("\t")
all_text.append(text.lower())
all_labels.append(int(label))
return np.asarray(all_text),np.asarray(all_labels)
amazon_x,amazon_y = get_data("amazon.txt")
imdb_x,imdb_y = get_data("imdb.txt")
print(amazon_x.shape,amazon_y.shape,imdb_x.shape,imdb_y.shape)
amazon_x = np.hstack([universal_embed(amazon_x),nnlm_embed(amazon_x)])
imdb_x = np.hstack([universal_embed(imdb_x),nnlm_embed(imdb_x)])
print(amazon_x.shape,imdb_x.shape)
# +
## distribution of data prior to the subspace alignment transformation
pca = PCA(n_components=200)
both_features = np.vstack([amazon_x,imdb_x])
combined_pca = pca.fit_transform(both_features)
print(sum(pca.explained_variance_ratio_))
tsne_embed = TSNE(n_components=2).fit_transform(combined_pca)
tsne_amazon = tsne_embed[:1000,:]
tsne_imdb = tsne_embed[1000:,:]
plt.scatter(tsne_amazon[:,0],tsne_amazon[:,1],color="cyan",label="amazon",)
plt.scatter(tsne_imdb[:,0],tsne_imdb[:,1],color="black",label="imdb",alpha=0.8)
plt.xlabel("TSNE 1")
plt.ylabel("TSNE 2")
plt.title("distribution of text data")
plt.legend()
plt.show()
# -
# ### Subspace alignment
#
# Treating the IMDB data as being the target, so the Amazon data will be transformed.
# +
pca = PCA(n_components=200)
amazon_pca = pca.fit_transform(amazon_x)
source = pca.components_
print(sum(pca.explained_variance_ratio_))
pca = PCA(n_components=200)
imdb_pca = pca.fit_transform(imdb_x)
target = pca.components_
print(sum(pca.explained_variance_ratio_))
print(source.shape,target.shape)
print(amazon_pca.shape,imdb_pca.shape)
# -
# Calculating the transformation matrix
M = source.dot(target.T)
M.shape
# Getting the transformed Amazon data
new_imdb_x = imdb_pca
new_amazon_x = amazon_pca.dot(M)
print(new_amazon_x.shape)
## text distributions more closely match than they did before
tsne_embed = TSNE(n_components=2).fit_transform(np.vstack([new_amazon_x,new_imdb_x]))
tsne_amazon = tsne_embed[:1000,:]
tsne_imdb = tsne_embed[1000:,:]
plt.scatter(tsne_amazon[:,0],tsne_amazon[:,1],color="cyan",label="amazon",)
plt.scatter(tsne_imdb[:,0],tsne_imdb[:,1],color="black",label="imdb",alpha=0.8)
plt.xlabel("TSNE 1")
plt.ylabel("TSNE 2")
plt.title("distribution of text data - post transformation")
plt.legend()
plt.show()
| domain_adaptation/Subspace_alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="V58rxea0HqSa" colab={"base_uri": "https://localhost:8080/"} outputId="bb1d9bb1-5d2a-42de-caa4-c0d8381494da"
import os
# Find the latest version of spark 3.0 from http://www.apache.org/dist/spark/ and enter as the spark version
# For example:
# spark_version = 'spark-3.1.2'
spark_version = 'spark-3.1.2'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-11-jdk-headless -qq > /dev/null
# !wget -q http://www.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + id="_xKwTpATHqSe" colab={"base_uri": "https://localhost:8080/"} outputId="71c0e9c5-a2fa-4360-e573-2213189ed634"
# Download the Postgres driver that will allow Spark to interact with Postgres.
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.16.jar
# + id="MMqDAjVS0KN9"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("M16-Amazon-Challenge").config("spark.driver.extraClassPath","/content/postgresql-42.2.16.jar").getOrCreate()
# + [markdown] id="cyBsySGuY-9V"
# ### Load Amazon Data into Spark DataFrame
# + id="CtCmBhQJY-9Z" colab={"base_uri": "https://localhost:8080/"} outputId="c198e2da-b419-415a-e758-22168c6e99ff"
from pyspark import SparkFiles
url = "https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Video_Games_v1_00.tsv.gz"
spark.sparkContext.addFile(url)
df = spark.read.option("encoding", "UTF-8").csv(SparkFiles.get(""), sep="\t", header=True, inferSchema=True)
df.show()
# + [markdown] id="2yUSe55VY-9t"
# ### Create DataFrames to match tables
# + id="C8REmY1aY-9u"
from pyspark.sql.functions import to_date
# Read in the Review dataset as a DataFrame
# + id="B0TESUDRY-90" colab={"base_uri": "https://localhost:8080/"} outputId="cb7dbda1-0298-42f7-b173-aa6e1fbda3d2"
# Create the customers_table DataFrame
customers_df = df.groupby("customer_id").agg({'customer_id': 'count'}).withColumnRenamed("count(customer_id)", "customer_count")
customers_df.show()
# + id="4FwXA6UvY-96" colab={"base_uri": "https://localhost:8080/"} outputId="1c9db91f-f895-4f2f-dee5-a220d27a2859"
# Create the products_table DataFrame and drop duplicates.
products_df = df.select(['product_id', 'product_title']).drop_duplicates()
products_df.show(truncate=False)
# + id="MkqyCuNQY-9-" colab={"base_uri": "https://localhost:8080/"} outputId="61e37406-ac54-45bd-aa5d-ff56a69eca6e"
# Create the review_id_table DataFrame.
# Convert the 'review_date' column to a date datatype with to_date("review_date", 'yyyy-MM-dd').alias("review_date")
review_id_df = df.select(['review_id', 'customer_id', 'product_id', 'product_parent', to_date("review_date", 'yyyy-MM-dd').alias("review_date")])
review_id_df.show()
# + id="lzMmkdKmY--D" colab={"base_uri": "https://localhost:8080/"} outputId="247febd3-3d4a-4777-c3c7-f09b3679c36a"
# Create the vine_table. DataFrame
vine_df = df.select(['review_id', 'star_rating', 'helpful_votes', 'total_votes', 'vine', 'verified_purchase'])
vine_df.show()
# + [markdown] id="jITZhLkmY--J"
# ### Connect to the AWS RDS instance and write each DataFrame to its table.
# + id="7jiUvs1aY--L"
# Configure settings for RDS
mode = "append"
jdbc_url="jdbc:postgresql://ajay-database.cbsz8urlbgic.us-east-2.rds.amazonaws.com:5432/postgres"
config = {"user":"root",
"password": password,
"driver":"org.postgresql.Driver"}
# + id="T2zgZ-aKY--Q"
# Write review_id_df to table in RDS
review_id_df.write.jdbc(url=jdbc_url, table='review_id_table', mode=mode, properties=config)
# + id="1m3yzn-LY--U"
# Write products_df to table in RDS
# about 3 min
products_df.write.jdbc(url=jdbc_url, table='products_table', mode=mode, properties=config)
# + id="KbXri15fY--Z"
# Write customers_df to table in RDS
# 5 min 14 s
customers_df.write.jdbc(url=jdbc_url, table='customers_table', mode=mode, properties=config)
# + id="XdQknSHLY--e"
# Write vine_df to table in RDS
# 11 minutes
vine_df.write.jdbc(url=jdbc_url, table='vine_table', mode=mode, properties=config)
# + id="Exuo6ebUsCqW"
| Amazon_Reviews_ETL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import datetime
import sys
import bisect
sys.path.insert(0, '..')
import keras.layers
from keras.layers import Input, Dense, LocallyConnected1D, Reshape
from keras.models import Model
from keras import regularizers
from sklearn import preprocessing
from IPython.display import SVG
from keras.utils import model_to_dot
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [15, 10]
# -
import umap
import umap.plot
raw_df = pd.read_hdf("../data/processed/summaries_protoss.hdf", "summaries")
raw_df = raw_df[raw_df.game_duration > 280]
raw_df.head().reset_index(drop=True)
# +
metadata_columns = """
self_won
game_start
game_duration
self_name self_race_is_protoss self_race_is_zerg self_race_is_terran
opponent_name
""".split()
weight_columns = [col for col in raw_df.columns if col.endswith("_weight")]
data_columns = [col for col in raw_df.columns if col not in metadata_columns and col not in weight_columns]
df = raw_df[data_columns]
min_max_scaler = preprocessing.MinMaxScaler()
df = pd.DataFrame(min_max_scaler.fit_transform(df), columns=df.columns, index=df.index)
encoding_dim = 10
# -
len(data_columns)
# +
input_data = Input(shape=(len(df.columns),))
encoded = Dense(encoding_dim, activation='relu')(input_data)
decoded = Dense(len(df.columns), activation='sigmoid')(encoded)
autoencoder = Model(input_data, decoded)
# -
encoder = Model(input_data, encoded)
autoencoder.compile(optimizer='adadelta', loss='mean_squared_error')
print("Model has {} parameters".format(autoencoder.count_params()))
SVG(model_to_dot(autoencoder).create(prog='dot', format='svg'))
x_all = df.sample(frac=1.0).values
num_samples = int(0.95 * x_all.shape[0])
x_train, x_test = x_all[:num_samples, :], x_all[num_samples:, :]
print("Training on {} samples. Testing on {}.".format(
num_samples, x_all.shape[0] - num_samples))
history = autoencoder.fit(x_train, x_train,
epochs=2000,
batch_size=1024,
shuffle=True,
verbose=0,
validation_data=(x_test, x_test))
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
cluster_games_df = raw_df[raw_df.game_start > datetime.datetime(2020, 1, 1)]
original_space = min_max_scaler.transform(cluster_games_df[data_columns].values)
encoded_space =autoencoder.predict(original_space)
# +
#encoded_space[0]
# +
#original_space[0]
# -
diffs = encoded_space - original_space
cluster_games_df['creativity'] = np.einsum("ij,ij->i", diffs, diffs)
creative_games_df= cluster_games_df.sort_values('creativity')
creative_games_won = creative_games_df[creative_games_df['self_won'] > 0]
creative_games_won.tail(20)[['game_start', 'self_name', 'opponent_name', 'creativity']].reset_index(drop=True)
sns.distplot(cluster_games_df.creativity)
# +
def summarize_player(*names):
names = [n.lower() for n in names]
series = cluster_games_df[cluster_games_df.self_name.str.lower().isin(names)].creativity
print(series.describe())
return sns.distplot(series, bins=15)
def summarize_player_contains(name):
all_names = set(creative_games_df[creative_games_df['self_name'].str.lower().str.contains(name.lower())].self_name)
print("Names to use: {}".format(sorted(all_names)))
return summarize_player(*all_names)
# -
summarize_player_contains("maxpax")
summarize_player_contains("dear")
summarize_player_contains("rotterdam")
summarize_player_contains("zest")
summarize_player_contains("skillous")
| notebooks/Autoencoder - selfdistance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.DataFrame(
{'A': [10, 20, 30, 40, 50], # 列Aとその値
'B': [0.8, 1.6, 2.4, 4.3, 7.6], # 列Bとその値
'C': [-1, -2.6, -3.5, -4.3, -5.1] }, # 列Cとその値
index = ['r1', 'r2', 'r3', 'r4', 'r5'] # 行名を設定
)
df['D'] = [1, 2, 3, 4, 5]
df
df_new = pd.DataFrame(
{'E': [10, 20, 30, 40, 50], # 列Eとその値
'F': [0.8, 1.6, 2.4, 4.3, 7.6]} # 列Fとその値
)
| notebooks/chap11/10_04/.ipynb_checkpoints/add_row-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import random
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm
from itertools import chain
from skimage.io import imread, imshow, imread_collection, concatenate_images
from skimage.transform import resize
from skimage.morphology import label
from tensorflow.keras import Model
#
from tensorflow.keras.layers import Input,Lambda,Conv2D,Conv2DTranspose,MaxPooling2D,concatenate
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.keras import backend as K
import tensorflow as tf
# Set some parameters
IMG_WIDTH = 256
IMG_HEIGHT = 256
IMG_CHANNELS = 3
TRAIN_PATH = 'input/stage1_train/'
TEST_PATH = 'input/stage1_test/'
warnings.filterwarnings('ignore', category=UserWarning, module='skimage')
random.seed = 40
np.random.seed = 40
# +
train_ids = next(os.walk(TRAIN_PATH))[1]
test_ids = next(os.walk(TEST_PATH))[1]
# +
# Get and resize train images and masks
X_train = np.zeros((len(train_ids), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), dtype=np.uint8)
Y_train = np.zeros((len(train_ids), IMG_HEIGHT, IMG_WIDTH, 1), dtype=np.bool)
print('Getting and resizing train images and masks ... ')
sys.stdout.flush()
for n, id_ in tqdm(enumerate(train_ids), total=len(train_ids)):
path = TRAIN_PATH + id_
img = imread(path + '/images/' + id_ + '.png')[:,:,:IMG_CHANNELS]
img = resize(img, (IMG_HEIGHT, IMG_WIDTH), mode='constant', preserve_range=True)
X_train[n] = img
mask = np.zeros((IMG_HEIGHT, IMG_WIDTH, 1), dtype=np.bool)
for mask_file in next(os.walk(path + '/masks/'))[2]:
mask_ = imread(path + '/masks/' + mask_file)
mask_ = np.expand_dims(resize(mask_, (IMG_HEIGHT, IMG_WIDTH), mode='constant',
preserve_range=True), axis=-1)
mask = np.maximum(mask, mask_)
Y_train[n] = mask
# Get and resize test images
X_test = np.zeros((len(test_ids), IMG_HEIGHT, IMG_WIDTH, IMG_CHANNELS), dtype=np.uint8)
sizes_test = []
print('Getting and resizing test images ... ')
sys.stdout.flush()
for n, id_ in tqdm(enumerate(test_ids), total=len(test_ids)):
path = TEST_PATH + id_
img = imread(path + '/images/' + id_ + '.png')[:,:,:IMG_CHANNELS]
sizes_test.append([img.shape[0], img.shape[1]])
img = resize(img, (IMG_HEIGHT, IMG_WIDTH), mode='constant', preserve_range=True)
X_test[n] = img
print('Done!')
# +
def iou_metric(y_true_in, y_pred_in, print_table=False):
labels = label(y_true_in > 0.5)
y_pred = label(y_pred_in > 0.5)
true_objects = len(np.unique(labels))
pred_objects = len(np.unique(y_pred))
intersection = np.histogram2d(labels.flatten(), y_pred.flatten(), bins=(true_objects, pred_objects))[0]
# Compute areas (needed for finding the union between all objects)
area_true = np.histogram(labels, bins = true_objects)[0]
area_pred = np.histogram(y_pred, bins = pred_objects)[0]
area_true = np.expand_dims(area_true, -1)
area_pred = np.expand_dims(area_pred, 0)
# Compute union
union = area_true + area_pred - intersection
# Exclude background from the analysis
intersection = intersection[1:,1:]
union = union[1:,1:]
union[union == 0] = 1e-9
# Compute the intersection over union
iou = intersection / union
# Precision helper function
def precision_at(threshold, iou):
matches = iou > threshold
true_positives = np.sum(matches, axis=1) == 1 # Correct objects
false_positives = np.sum(matches, axis=0) == 0 # Missed objects
false_negatives = np.sum(matches, axis=1) == 0 # Extra objects
tp, fp, fn = np.sum(true_positives), np.sum(false_positives), np.sum(false_negatives)
return tp, fp, fn
# Loop over IoU thresholds
prec = []
if print_table:
print("Thresh\tTP\tFP\tFN\tPrec.")
for t in np.arange(0.5, 1.0, 0.05):
tp, fp, fn = precision_at(t, iou)
if (tp + fp + fn) > 0:
p = tp / (tp + fp + fn)
else:
p = 0
if print_table:
print("{:1.3f}\t{}\t{}\t{}\t{:1.3f}".format(t, tp, fp, fn, p))
prec.append(p)
if print_table:
print("AP\t-\t-\t-\t{:1.3f}".format(np.mean(prec)))
return np.mean(prec)
def iou_metric_batch(y_true_in, y_pred_in):
batch_size = y_true_in.shape[0]
metric = []
for batch in range(batch_size):
value = iou_metric(y_true_in[batch], y_pred_in[batch])
metric.append(value)
return np.array(np.mean(metric), dtype=np.float32)
def my_iou_metric(label, pred):
metric_value = tf.py_function(iou_metric_batch, [label, pred], tf.float32)
return metric_value
# +
# Build U-Net model
inputs = Input((250, 250, 3))
s = Lambda(lambda x: x ) (inputs)
c1 = Conv2D(8, (3, 3), activation='relu', padding='same') (s)
c1 = Conv2D(8, (3, 3), activation='relu', padding='same') (c1)
p1 = MaxPooling2D((2, 2)) (c1)
c2 = Conv2D(16, (3, 3), activation='relu', padding='same') (p1)
c2 = Conv2D(16, (3, 3), activation='relu', padding='same') (c2)
p2 = MaxPooling2D((2, 2)) (c2)
c3 = Conv2D(32, (3, 3), activation='relu', padding='same') (p2)
c3 = Conv2D(32, (3, 3), activation='relu', padding='same') (c3)
p3 = MaxPooling2D((2, 2)) (c3)
c4 = Conv2D(64, (3, 3), activation='relu', padding='same') (p3)
c4 = Conv2D(64, (3, 3), activation='relu', padding='same') (c4)
p4 = MaxPooling2D(pool_size=(2, 2)) (c4)
c5 = Conv2D(128, (3, 3), activation='relu', padding='same') (p4)
c5 = Conv2D(128, (3, 3), activation='relu', padding='same') (c5)
u6 = Conv2DTranspose(64, (2, 2), strides=(2, 2), padding='same') (c5)
u6 = concatenate([u6, c4])
c6 = Conv2D(64, (3, 3), activation='relu', padding='same') (u6)
c6 = Conv2D(64, (3, 3), activation='relu', padding='same') (c6)
u7 = Conv2DTranspose(32, (2, 2), strides=(2, 2), padding='same') (c6)
u7 = concatenate([u7, c3])
c7 = Conv2D(32, (3, 3), activation='relu', padding='same') (u7)
c7 = Conv2D(32, (3, 3), activation='relu', padding='same') (c7)
u8 = Conv2DTranspose(16, (2, 2), strides=(2, 2), padding='same') (c7)
u8 = concatenate([u8, c2])
c8 = Conv2D(16, (3, 3), activation='relu', padding='same') (u8)
c8 = Conv2D(16, (3, 3), activation='relu', padding='same') (c8)
u9 = Conv2DTranspose(8, (2, 2), strides=(2, 2), padding='same') (c8)
u9 = concatenate([u9, c1], axis=3)
c9 = Conv2D(8, (3, 3), activation='relu', padding='same') (u9)
c9 = Conv2D(8, (3, 3), activation='relu', padding='same') (c9)
outputs = Conv2D(1, (1, 1), activation='sigmoid') (c9)
model = Model(inputs=[inputs], outputs=[outputs])
#model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=[my_iou_metric])
#model = tf.keras.utils.multi_gpu_model(mode,cpu_merge=True,cpu_relocation=False)
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
model.summary()
# -
def get_unet(n_ch,patch_height,patch_width):
concat_axis = 3
inputs = Input((patch_height, patch_width, n_ch))
conv1 = Conv2D(32, (3, 3), padding="same", name="conv1_1", activation="relu", data_format="channels_last")(inputs)
conv1 = Conv2D(32, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2), data_format="channels_last")(conv1)
conv2 = Conv2D(64, (3, 3), padding="same", activation="relu", data_format="channels_last")(pool1)
conv2 = Conv2D(64, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2), data_format="channels_last")(conv2)
conv3 = Conv2D(128, (3, 3), padding="same", activation="relu", data_format="channels_last")(pool2)
conv3 = Conv2D(128, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2), data_format="channels_last")(conv3)
conv4 = Conv2D(256, (3, 3), padding="same", activation="relu", data_format="channels_last")(pool3)
conv4 = Conv2D(256, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2), data_format="channels_last")(conv4)
conv5 = Conv2D(512, (3, 3), padding="same", activation="relu", data_format="channels_last")(pool4)
conv5 = Conv2D(512, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv5)
up_conv5 = UpSampling2D(size=(2, 2), data_format="channels_last")(conv5)
ch, cw = get_crop_shape(conv4, up_conv5)
crop_conv4 = Cropping2D(cropping=(ch,cw), data_format="channels_last")(conv4)
up6 = concatenate([up_conv5, crop_conv4], axis=concat_axis)
conv6 = Conv2D(256, (3, 3), padding="same", activation="relu", data_format="channels_last")(up6)
conv6 = Conv2D(256, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv6)
up_conv6 = UpSampling2D(size=(2, 2), data_format="channels_last")(conv6)
ch, cw = get_crop_shape(conv3, up_conv6)
crop_conv3 = Cropping2D(cropping=(ch,cw), data_format="channels_last")(conv3)
up7 = concatenate([up_conv6, crop_conv3], axis=concat_axis)
conv7 = Conv2D(128, (3, 3), padding="same", activation="relu", data_format="channels_last")(up7)
conv7 = Conv2D(128, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv7)
up_conv7 = UpSampling2D(size=(2, 2), data_format="channels_last")(conv7)
ch, cw = get_crop_shape(conv2, up_conv7)
crop_conv2 = Cropping2D(cropping=(ch,cw), data_format="channels_last")(conv2)
up8 = concatenate([up_conv7, crop_conv2], axis=concat_axis)
conv8 = Conv2D(64, (3, 3), padding="same", activation="relu", data_format="channels_last")(up8)
conv8 = Conv2D(64, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv8)
up_conv8 = UpSampling2D(size=(2, 2), data_format="channels_last")(conv8)
ch, cw = get_crop_shape(conv1, up_conv8)
crop_conv1 = Cropping2D(cropping=(ch,cw), data_format="channels_last")(conv1)
up9 = concatenate([up_conv8, crop_conv1], axis=concat_axis)
conv9 = Conv2D(32, (3, 3), padding="same", activation="relu", data_format="channels_last")(up9)
conv9 = Conv2D(32, (3, 3), padding="same", activation="relu", data_format="channels_last")(conv9)
#ch, cw = get_crop_shape(inputs, conv9)
#conv9 = ZeroPadding2D(padding=(ch[0],cw[0]), data_format="channels_last")(conv9)
#conv10 = Conv2D(1, (1, 1), data_format="channels_last", activation="sigmoid")(conv9)
flatten = Flatten()(conv9)
Dense1 = Dense(512, activation='relu')(flatten)
BN =BatchNormalization() (Dense1)
Dense2 = Dense(17, activation='sigmoid')(BN)
model = Model(input=inputs, output=Dense2)
return model.summary()
print(get_unet(3,256,256))
# +
results = model.fit(X_train, Y_train, validation_split=0.1, batch_size=8, epochs=5)
# +
# Predict on train, val and test
preds_train = model.predict(X_train[:int(X_train.shape[0]*0.9)], verbose=1)
preds_val = model.predict(X_train[int(X_train.shape[0]*0.9):], verbose=1)
preds_test = model.predict(X_test, verbose=1)
# Threshold predictions
preds_train_t = (preds_train > 0.5).astype(np.uint8)
preds_val_t = (preds_val > 0.5).astype(np.uint8)
preds_test_t = (preds_test > 0.5).astype(np.uint8)
# Create list of upsampled test masks
preds_test_upsampled = []
for i in range(len(preds_test)):
preds_test_upsampled.append(resize(np.squeeze(preds_test[i]),
(sizes_test[i][0], sizes_test[i][1]),
mode='constant', preserve_range=True))
# -
ix = 10
plt.figure(figsize=(20,20))
plt.subplot(131)
imshow(X_train[ix])
plt.title("Image")
plt.subplot(132)
imshow(np.squeeze(Y_train[ix]))
plt.title("Mask")
plt.subplot(133)
imshow(np.squeeze(preds_train_t[ix] > 0.5))
plt.title("Predictions")
plt.show()
iou_metric(np.squeeze(Y_train[ix]), np.squeeze(preds_train_t[ix]), print_table=True)
| kerasu test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def formingMagicSquare(square):
allSquares = [
[[4, 9, 2], [3, 5, 7], [8, 1, 6]],
[[2, 7, 6], [9, 5, 1], [4, 3, 8]],
[[6, 1, 8], [7, 5, 3], [2, 9, 4]],
[[8, 3, 4], [1, 5, 9], [6, 7, 2]],
[[2, 9, 4], [7, 5, 3], [6, 1, 8]],
[[6, 7, 2], [1, 5, 9], [8, 3, 4]],
[[8, 1, 6], [3, 5, 7], [4, 9, 2]],
[[4, 3, 8], [9, 5, 1], [2, 7, 6]],
]
minCost = 9 * 9
for magicSquare in allSquares:
cost = 0
for i in range(3):
for j in range(3):
cost += abs(magicSquare[i][j] - square[i][j])
if cost < minCost:
minCost = cost
return minCost
square = [
list(map(int, input().split())),
list(map(int, input().split())),
list(map(int, input().split()))
]
print(formingMagicSquare(square))
| Algorithm/21. forming a magic square.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # San Diego, California, US
# + active=""
# # install dependencies
# import sys
# !{sys.executable} -m pip install gdal shapely geopandas rasterio xarray \
# earthengine-api matplotlib contextily --upgrade >> /dev/null
# -
import xarray as xr
import numpy as np
import glob
import ee
from IPython.display import Image
from IPython.core.display import HTML
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Define helper functions
from geomed3dv4 import *
# +
def plot_fractality(ax, data):
from scipy.stats import linregress
import numpy as np
import matplotlib.ticker as ticker
ax.loglog(data.r, data, base=2, label='Calculated')
ax.set_xlabel('Wavelength, m', fontsize=18)
ax.axes.get_yaxis().set_visible(False)
ax.xaxis.set_major_formatter(ticker.FuncFormatter(lambda y, _: '{:g}'.format(y)))
res = linregress(np.log2(data.r), np.log2(data))
ax.plot(data.r, 2**(res.intercept + res.slope*np.log2(data.r)), 'r', label=f'Fitted R²={res.rvalue**2:.2f}', ls='--')
ax.legend(fontsize=18)
fractality = 1000*np.round((3 - (res.slope/2)),1)
return fractality
#def plot_preview(ax, GEEimage, filename):
# image = plt.imread(filename)
# ax.imshow(image, interpolation='bilinear', extent=gee_image2rect(GEEimage, True))
def plot_preview(ax, GEEimage, filename):
import matplotlib.ticker as ticker
image = plt.imread(filename)
ax.imshow(image, interpolation='bilinear', extent=gee_image2rect(GEEimage, True))
#ax.ticklabel_format(useOffset=False, style='plain')
ax.get_yaxis().set_major_formatter(ticker.FuncFormatter(lambda x, p: int(x/1000)))
ax.get_xaxis().set_major_formatter(ticker.FuncFormatter(lambda x, p: int(x/1000)))
ax.set_ylabel('Y, km', fontsize=18)
ax.set_xlabel('X, km', fontsize=18)
# -
# ## Connect to GEE
service_account = '<EMAIL>@gee-export-308512.iam.g<EMAIL>account.com'
service_key = '/Users/mbg/gee-export.json'
credentials = ee.ServiceAccountCredentials(service_account, service_key)
ee.Initialize(credentials)
# ## Define work area and scale
# +
# define center of work area, [m]
point = [575500, 3642500]
# radius, [m]
radius = 500
# scale, [m]
scale = 30
# EPSG code (coordinates system)
epsg = 26911
# redefine library function for projected coordinates
def gee_image2rect(GEEimage, reorder=False):
if not reorder:
return [point[0]-radius, point[1]-radius, point[0]+radius, point[1]+radius]
else:
return [point[0]-radius, point[0]+radius, point[1]-radius, point[1]+radius]
# define spatial components 1*scale .. N*scale [m]
gammas = np.arange(1, 6)
#(minx, miny, maxx, maxy)
GEEarea_proj = ee.Geometry.Rectangle(
coords=gee_image2rect(None),
proj=f'EPSG:{epsg}',
geodesic=False
)
GEEarea_proj.getInfo()
# -
# ## Prepare image bands
# https://developers.google.com/earth-engine/datasets/catalog/JAXA_ALOS_AW3D30_V3_2
GEEdsm = ee.ImageCollection("JAXA/ALOS/AW3D30/V3_2").mosaic().select('DSM')
# ## Build image mosaic
GEEimage = \
GEEdsm\
.reproject(crs=f'epsg:{epsg}',scale=scale)
print ('collected bands', GEEimage.bandNames().getInfo())
# ### Make image preview and save on local filesystem
# Note: we need to check the image preview quality before the image usage
# define visualization parameters
DEMvis = {'bands':['DSM'], 'min':230, 'max':330,
'palette': ['00A600','63C600','E6E600','E9BD3A','ECB176','EFC2B3','F2F2F2']}
print (gee_preview_tofile(GEEimage.clip(GEEarea_proj), DEMvis, (320, 320), 'dempreview.320x320.jpg')['url'])
Image('dempreview.320x320.jpg')
# ### Extract raw image bands and save on local filesystem
GEEurl = GEEimage.getDownloadURL({'filePerBand':True, 'scale': scale, 'region': GEEarea_proj})
print (f'downloading', GEEurl)
fname = f'image.{scale}m.zip'
geeurl_tofile(GEEurl, fname)
# ### Convert downloaded zipped images chunks to compressed NetCDF file
# %%time
ds = zipsbands2image(glob.glob(f'image.{scale}m.zip'))
# check data variables
for varname in ds.data_vars:
print (varname, float(ds[varname].min()), float(ds[varname].max()))
encoding = {var: dict(zlib=True, complevel=6) for var in ds.data_vars}
ds.to_netcdf(f'image.{scale}m.nc', encoding=encoding)
print (ds.dims)
# ## DEM Fractality
dem_power = xr.DataArray([raster_gamma_range(ds.DSM, g-1, g+1, backward=True).std() for g in gammas],
coords=[scale*gammas],
dims=['r'])
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 6))
plot_preview(ax1, GEEimage, 'dempreview.320x320.jpg')
dem_fractality = plot_fractality(ax2, dem_power)
plt.suptitle(f"ALOS AW3D30 DEM Fractality for San Diego, California, US\nFractality Density ρ={dem_fractality:.0f} kg/m³", fontsize=22)
fig.tight_layout(rect=[0.03, 0.03, .97, 0.97])
plt.savefig('ALOS AW3D30 DEM Fractality for San Diego, California, US.jpg', dpi=150)
plt.show()
| USGS_NED_one_meter/GEE ALOS AW3D30 DEM Fractality for San Diego, California, US.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview of key ideas
#
# + This notebook is part of the addition lecture *An overview of key ideas* in the OCW MIT course 18.06 by Prof <NAME> [1]
# + Created by me, Dr <NAME>
# + Head of Acute Care Surgery
# + Groote Schuur Hospital
# + University Cape Town
# + <a href="mailto:<EMAIL>">Email me with your thoughts, comments, suggestions and corrections</a>
# <a rel="license" href="http://creativecommons.org/licenses/by-nc/4.0/"><img alt="Creative Commons Licence" style="border-width:0" src="https://i.creativecommons.org/l/by-nc/4.0/88x31.png" /></a><br /><span xmlns:dct="http://purl.org/dc/terms/" href="http://purl.org/dc/dcmitype/InteractiveResource" property="dct:title" rel="dct:type">Linear Algebra OCW MIT18.06</span> <span xmlns:cc="http://creativecommons.org/ns#" property="cc:attributionName">IPython notebook [2] study notes by Dr <NAME></span> is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-nc/4.0/">Creative Commons Attribution-NonCommercial 4.0 International License</a>.
#
# + [1] <a href="http://ocw.mit.edu/courses/mathematics/18-06sc-linear-algebra-fall-2011/index.htm">OCW MIT 18.06</a>
# + [2] F<NAME>, <NAME>, IPython: A System for Interactive Scientific Computing, Computing in Science and Engineering, vol. 9, no. 3, pp. 21-29, May/June 2007, doi:10.1109/MCSE.2007.53. URL: http://ipython.org
from IPython.core.display import HTML, Image
css_file = 'style.css'
HTML(open(css_file, 'r').read())
from sympy import init_printing, Matrix, symbols, sqrt, Rational
from numpy import matrix, transpose, sqrt
from numpy.linalg import pinv, inv, det, svd, norm
from scipy.linalg import pinv2
from warnings import filterwarnings
init_printing(use_latex = 'mathjax')
filterwarnings('ignore')
# # An overview of key ideas
# ## Moving from vectors to matrices
# + Consider a position vector in three-dimensional space
# + It can be written as a column-vector
# $$ u=\begin{bmatrix}1\\-1\\0\end{bmatrix} \\ v=\begin{bmatrix}0\\1\\-1\end{bmatrix} $$
# + We can add constant scalar multiples of these vectors
# $$ {x}_{1}{u}+{x}_{2}{v}={b} $$
# + This is simple vector addition
# + Its easy to visualize that if we combine all possible combinations, that we start filling a plane through the origin
# + Adding a third vector that is not in this plane will extend all possible linear combinations to fill all of three-dimensional space
# $$ w=\begin{bmatrix}0\\0\\1\end{bmatrix} $$
# + We now have the following
# $$ {x}_{1}{u}+{x}_{2}{v}+{x}_{3}{w}={b} $$
# + Notice how this last equation can be written in matrix form A**x**=**b**
# $$ \begin{bmatrix} 1 & 0 & 0 \\ -1 & 1 & 0 \\ 0 & -1 & 1 \end{bmatrix}\begin{bmatrix} { x }_{ 1 } \\ { x }_{ 2 } \\ { x }_{ 3 } \end{bmatrix}=\begin{bmatrix} { x }_{ 1 } \\ { x }_{ 2 }-{ x }_{ 1 } \\ { x }_{ 3 }-{ x }_{ 2 } \end{bmatrix} $$
# + This is the column-view of matrix-vector multiplication as opposed to the row view
# + Matrices are seen a column, representing vectors
# + Each element of the column vector **x** is a scalar multiple of the corresponding column in the matrix A
# $$ { x }_{ 1 }\begin{bmatrix} 1 \\ -1 \\ 0 \end{bmatrix}+{ x }_{ 2 }\begin{bmatrix} 0 \\ 1 \\ -1 \end{bmatrix}+{ x }_{ 3 }\begin{bmatrix} 0 \\ 0 \\ 1 \end{bmatrix}=\begin{matrix} { x }_{ 1 } \\ -{ x }_{ 1 }+{ x }_{ 2 } \\ -{ x }_{ 2 }+{ x }_{ 3 } \end{matrix} = {x}_{1}{u}+{x}_{2}{v}+{x}_{3}{w}$$
# + Now consider the solution vector **b**
# $$ \begin{bmatrix} 1 & 0 & 0 \\ -1 & 1 & 0 \\ 0 & -1 & 1 \end{bmatrix}\begin{bmatrix} { x }_{ 1 } \\ { x }_{ 2 } \\ { x }_{ 3 } \end{bmatrix}=\begin{bmatrix} { x }_{ 1 } \\ { x }_{ 2 }-{ x }_{ 1 } \\ { x }_{ 3 }-{ x }_{ 2 } \end{bmatrix} = \begin{bmatrix}{b}_{1}\\{b}_{2}\\{b}_{3}\end{bmatrix} $$
# + By substitution we we now have the following
# $$ \begin{bmatrix} { x }_{ 1 } \\ { x }_{ 2 } \\ { x }_{ 3 } \end{bmatrix}=\begin{bmatrix} { b }_{ 1 } \\ { b }_{ 1 }+{ b }_{ 2 } \\ { b }_{ 1 }+{ b }_{ 2 }+{ b }_{ 2 } \end{bmatrix} $$
# + This, though, looks like a matrix times **b**
# $$ \begin{bmatrix}1&0&0\\1&1&0\\1&1&1\end{bmatrix}\begin{bmatrix}{b}_{1}\\{b}_{2}\\{b}_{3}\end{bmatrix} $$
# + This matrix is the inverse of A such that **x**=A<sup>-1</sup>**b**
# + The above matrix A is called a difference matrix as it took simple differences between the elements of vector **x**
# + It was lower triangular
# + Its inverse became a sum matrix
# + So it was a good matrix, able to transform between **x** and **b** (back-and-forth) and therefor invertible and for every **x** has a specific inverse
# + It transforms **x** into **b** (maps)
# + Let's look at the code for this matrix which replaces **w** above
x1, x2, x3, b1, b2, b3 = symbols('x1, x2, x3, b1, b2, b3') # Creating algebraic symbols
# This reserves these symbols so as not to see them as computer variable names
C = Matrix([[1, 0, -1], [-1, 1, 0], [0, -1, 1]]) # Creating a matrix and putting
# it into a computer variable called C
C # Displaying it to the screen
x_vect = Matrix([[x1], [x2], [x3]]) # Giving this columns vector a computer
# variable name
x_vect
C * x_vect
# + We now have three equations
# $$ { x }_{ 1 }-{ x }_{ 3 }={ b }_{ 1 }\\ { x }_{ 2 }-{ x }_{ 1 }={ b }_{ 2 }\\ { x }_{ 3 }-{ x }_{ 2 }={ b }_{ 3 } $$
# + Adding the left and right sides we get the following
# $$ 0={ b }_{ 1 }+{ b }_{ 2 }+{ b }_{ 3 } $$
# + We are now constrained for values of *b*<sub>i</sub>
# + The problem is clear to see geometrically as the new **w** is in the same plane as **u** and **v**
# + In essence **w** did not add anything
# + All combinations of **u**, **v**, and **w** will still be in the plane
# + The first matrix A above had three independent columns and their linear combinations could fill all of three-dimensional space
# + That made the first matrix A invertible as opposed to the second one (C), which is not invertible (i.e. it cannot take any vector in three-dimensional space back to **x**)
# + Let's look at the original column vectors in C
# + Remember the following dot product
# $$ {a}\cdot{b}=||a||||b||\cos{\theta} \\ \cos\left(\pi\right)=-1 $$
# + In linear algebra getting the dot product of two vectors is written as follows
# $$ a\cdot{b}={b}^{T}{a} $$
# + Which is the transpose of the second times the first
u = Matrix([[1], [-1], [0]])
v = Matrix([[0], [1], [-1]])
w = Matrix([[-1], [0], [1]])
u, v, w
v.transpose() * u
w.transpose() * u
w.transpose() * v
u.transpose() * v
u.transpose() * w
v.transpose() * w
# + The angle between all of them is π radians and therefor they must all lie in a plane
# ## Example problems
# ### Example problem 1
# + Suppose A is a matrix with the following solution
# $$ {A}{x}=\begin{bmatrix}1\\4\\1\\1\end{bmatrix} \\ {x}=\begin{bmatrix}0\\1\\1\end{bmatrix}+{c}\begin{bmatrix}0\\2\\1\end{bmatrix} $$
# + What can you say about the columns of A?
# #### Solution
c = symbols('c')
x_vect = Matrix([[0], [1 + 2 * c], [1 + c]])
b = Matrix([[1], [4], [1], [1]])
# + **x** is of size *m* × *n* is 3 × 1
# + **b** is of size 4 × 1
# + Therefor A must be of size 4 × 3 and each column vector in A is in ℝ<sup>4</sup>
# + Let's call these columns of A *C*<sub>1</sub>, *C*<sub>2</sub>, and *C*<sub>3</sub>
# $$ \begin{bmatrix} \vdots & \vdots & \vdots \\ { C }_{ 1 } & { C }_{ 2 } & { C }_{ 3 } \\ \vdots & \vdots & \vdots \\ \vdots & \vdots & \vdots \end{bmatrix} $$
# + With the particular way in which **x** was written we can say that we have a particular solution and a special solution
# $$ {A}\left({x}_{p}+{c}\cdot{x}_{s}\right)=b $$
# + For *c* = 0 we have:
# $$ {A}{x}_{p}=b $$
# + For *c* = 1 we have:
# $$ A{ x }_{ p }+A{ x }_{ s }=b\\ \because \quad A{ x }_{ p }=b\\ b+A{ x }_{ s }=b\\ \therefore \quad A{ x }_{ s }=0 $$
# + We also have that the following
# $$ { x }_{ p }=\begin{bmatrix} 0 \\ 1 \\ 1 \end{bmatrix},\quad { x }_{ s }=\begin{bmatrix} 0 \\ 2 \\ 1 \end{bmatrix} $$
# + For *x*<sub>p</sub> we have the following
# $$ \begin{bmatrix} \vdots & \vdots & \vdots \\ { C }_{ 1 } & { C }_{ 2 } & { C }_{ 3 } \\ \vdots & \vdots & \vdots \\ \vdots & \vdots & \vdots \end{bmatrix}\begin{bmatrix} 0 \\ 1 \\ 1 \end{bmatrix}=b\quad \Rightarrow \quad { C }_{ 2 }+{ C }_{ 3 }=b $$
# + For *x*<sub>s</sub> we have the following
# $$ \begin{bmatrix} \vdots & \vdots & \vdots \\ { C }_{ 1 } & { C }_{ 2 } & { C }_{ 3 } \\ \vdots & \vdots & \vdots \\ \vdots & \vdots & \vdots \end{bmatrix}\begin{bmatrix} 0 \\ 2 \\ 1 \end{bmatrix}=\underline { 0 } \quad \Rightarrow \quad 2{ C }_{ 2 }+{ C }_{ 3 }=0 $$
# + Solving for *C*<sub>2</sub> and *C*<sub>3</sub> we have the following
# $$ {C}_{3}=-2{C}_{2} \\ {C}_{2}-2{C}_{2}=b \\ {C}_{2}=-b \\ {C}_{3}=2b$$
# + As for the first column of A, we need to know more about ranks and subspaces
# + We see, though, that columns 2 and three are already constant multiples of each other
# + So, as long as column 1 is not a constant multiple of b, we are safe
# $$ A=\begin{bmatrix} \vdots & 1 & 2 \\ { C }_{ 1 } & 4 & 8 \\ \vdots & 1 & 2 \\ \vdots & 1 & 2 \end{bmatrix} $$
| Beginners_Guide_Math_LinAlg/Math/I_02_Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load data
# +
import pandas as pd, numpy as np, re
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.datasets import load_svmlight_file
from tqdm import tqdm
def parse_data(filename):
with open(filename, "rb") as f:
infoline = f.readline()
infoline = re.sub(r"^b'", "", str(infoline))
n_features = int(re.sub(r"^\d+\s(\d+)\s\d+.*$", r"\1", infoline))
features, labels = load_svmlight_file(f, n_features=n_features, multilabel=True)
mlb = MultiLabelBinarizer()
labels = mlb.fit_transform(labels)
features = np.array(features.todense())
features = np.ascontiguousarray(features)
return features, labels
X, y = parse_data("../data/Bibtex_data.txt")
print(X.shape)
print(y.shape)
# -
# # Own- AdaptiveGreedy
# - Adaptive greedy with threshold (as opposed to percentile) and beta prior (as opposed to smoothing).
# SUMMARY
#
# fit()
# 1) The fit method fits a model individually based upon the rewards we observed for each bandit in the batch (batch = 50 rows).
# - So if we decided to play Bandit 1, for say rounds 3, 40, 55, we would fit a model on the context we had for those three rounds
# with the rewards we observed (which can be 0 or 1).
# - SPECIAL SITUATION 1) There is a potential problem though if we observe only rewards 0 or only rewards 1, we really cannot fit a model on examples with
# only one label. In these situations we will predict a 0 or 1 automatically.
# - SPECIAL SITUATION 2) Furthermore, for cases where we have less than 2 positive or 2 negative for each bandit, we will use a beta prior and
# draw the label from a beta distribution as per the paper.
#
# 2) So in summary, at the start of every fit(), we first make a list containing unfit logistic regression models. We then overwrite these models with the beta prior above if the conditions hold. We then proceed to fit the models on the historical data (which increases over time). Over time, as we have more samples for each bandit, we will have less overwriting and end up using the logistic regression more and more for predicitons.
#
# Predict()
# 1) The predict method loops through all the models and predicts a probability. It then takes the argmax probability over all bandits and compares it to the threshold. If it is below the threshold, it takes a random bandit (hence exploring). If it is greater, it chooses the bandit with the highest probability as the action and the reward is revealed (0 or 1).
#
# 2) This threshold is then decayed after the end of every batch making it smaller which means the probability of choosing randomly becomes smaller (hence less exploration).
#
# Other things
# 1) In addition to beta priors, it is possible to do smoothing. r_smooth = (r * n + a) / (n + b) which is number of positives over total number per bandit.
#
# Extension
# 1) The extension adds weighting to the bandit fitting process. This means that examples further away in history will be weighted less (say 0.7 for a row vs 0.9 for a more recennt row). The intuition is that the policy is worse earlier (due to more exploraration) but as time progresses it becomes better so when we do a full-refit we may want to discount the early actions more.
# 2) New base learner. The original paper used logistic regression and we try several other ones such as random forest.
class BetaPredictor():
def __init__(self, a, b):
self.a = a
self.b = b
def fit(self, X=None, y=None, sample_weight=None):
pass
def predict_proba(self, X):
preds = np.random.beta(self.a, self.b, size = X.shape[0]).reshape((-1, 1))
return np.c_[1.0 - preds, preds]
def decision_function(self, X):
return np.random.beta(self.a, self.b, size = X.shape[0])
def predict(self, X):
return (np.random.beta(self.a, self.b, size = X.shape[0])).astype('uint8')
def discount_func(x, a = 0.1):
return x / (x + a)
class adaptive_greedy():
"""
Args
model: [Sklearn linear model] .Implements fit/predict
num_bandits: [int] Number of bandits/classes we choose from.
beta_prior: [tuple]. First element refers to alpha and beta for smoothing. Second element is number of positive/
negative labels we need to see for each bandit before we stop using the beta-prior
smoothing: [tuple]. First element is alpha in smoothing and second beta in smooting.
r_smooth = (r * n + a) / (n + b) which is number of positives over total number per bandit.
decay: [float]. We can decay the threshold after a certain number of rounds
"""
def __init__(self, model, num_bandits, beta_prior, smoothing = None, decay = 0.9998, discount_rate = None):
self.num_bandits = num_bandits
self.beta_prior = beta_prior #((3,7),2) used in paper
self.base_model = model
self.threshold = 1.0 / (np.sqrt(nchoices) * 2.0) # 1 / (2* sqrt(k)) used in paper
self.decay = decay
self.bandit_counter = np.zeros((3, num_bandits)) #[indicator whether to use Beta prior, num pos, num neg] per class
self.smoothing = smoothing
if self.smoothing is not None:
self.counters = np.zeros((1, n)) #where n counts the number of observations per class
else:
self.counters = None
if discount_rate is not None:
self.bandit_weights = [np.array([]) for _ in range(self.num_bandits)]
self.discount_rate = discount_rate
else:
self.discount_rate = None
self.num_rounds = 0
def fit(self, X, a, r):
self.algos = [deepcopy(self.base_model) for i in range(self.num_bandits)]
self.num_rounds += 1
for i in range(self.num_bandits):
self.fit_base_model(X, a, r, i)
def update_bandit_weights(self, X, bandit):
discount_rate = discount_func(self.num_rounds) #get discount rate dynamically.
prior_nob = len(self.bandit_weights[bandit])
if prior_nob < 1:
new_weights = np.ones((X.shape[0], 1))
self.bandit_weights[bandit] = new_weights
else:
new_weights = np.zeros((X.shape[0], 1))
new_weights[:prior_nob] = self.bandit_weights[bandit] * discount_rate
new_weights[prior_nob:] = 1
self.bandit_weights[bandit] = new_weights
def fit_base_model(self, X, a, r, bandit):
a_for_bandit = (a == bandit)
y_for_bandit = r[a_for_bandit]
num_pos = y_for_bandit.sum()
if self.smoothing is not None:
self.counters[0, i] += y_for_bandit.shape[0]
if (num_pos < self.beta_prior[1]) or ((y_for_bandit.shape[0] - num_pos) < self.beta_prior[1]):
self.algos[bandit] = BetaPredictor(self.beta_prior[0][0] + num_pos,self.beta_prior[0][1] + y_for_bandit.shape[0] - num_pos)
return None
xclass = X[a_for_bandit, :]
if self.discount_rate is not None:
self.update_bandit_weights(xclass, bandit)
self.algos[bandit].fit(xclass, y_for_bandit, self.bandit_weights[bandit].flatten())
else:
self.algos[bandit].fit(xclass, y_for_bandit)
if self.beta_prior[1] > 0:
self.increase_bandit_counter(y_for_bandit, bandit)
def increase_bandit_counter(self, y_for_bandit, bandit):
if (self.bandit_counter[0, bandit] == 0):
n_pos = y_for_bandit.sum()
self.bandit_counter[1, bandit] += n_pos
self.bandit_counter[2, bandit] += y_for_bandit.shape[0] - n_pos
if (self.bandit_counter[1, bandit] > beta_prior[1]) and (self.bandit_counter[2, bandit] > beta_prior[1]):
self.bandit_counter[0, bandit] = 1
def predict(self, X):
pred = self._predict(X)
self.threshold *= self.decay ** X.shape[0]
return pred
def _predict(self, X):
preds_proba = np.zeros((X.shape[0], self.num_bandits))
for bandit in range(self.num_bandits):
preds_proba[:, bandit] = self.algos[bandit].predict_proba(X)[:, 1]
if (self.smoothing is not None) and (self.counters is not None):
preds_proba[:, :] = (preds_proba * self.counters + smoothing[0]) / (self.counters + smoothing[1])
pred_max = preds_proba.max(axis = 1)
pred = np.argmax(preds_proba, axis = 1)
below_thr = pred_max <= self.threshold
if np.any(below_thr):
pred[below_thr] = np.random.randint(self.num_bandits, size = below_thr.sum())
return pred
# +
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from contextualbandits.online import AdaptiveGreedy
from copy import deepcopy
nchoices = y.shape[1]
base_algorithm = LogisticRegression(solver='lbfgs', warm_start=True)
base_algorithm2 = RandomForestClassifier(n_estimators=100, random_state=0)
beta_prior = ((3, 7), 2) # until there are at least 2 observations of each class, will use this prior
adaptive_greedy1 = adaptive_greedy(deepcopy(base_algorithm), nchoices , beta_prior)
adaptive_greedy_disc = adaptive_greedy(deepcopy(base_algorithm), nchoices , beta_prior, discount_rate = True)
adaptive_greedy_rf = adaptive_greedy(deepcopy(base_algorithm2), nchoices , beta_prior, discount_rate = True)
# -
# # Original Adaptive Greedy
# +
#adaptive_greedy_thr = AdaptiveGreedy(deepcopy(base_algorithm), nchoices=nchoices,
# decay_type='threshold',
# beta_prior = beta_prior)
# -
# # Comparison
models = [adaptive_greedy1, adaptive_greedy_disc, adaptive_greedy_rf]
# +
# These lists will keep track of the rewards obtained by each policy
rewards_agr, rewards_agr2, rewards_agr3 = [list() for i in range(len(models))]
lst_rewards = [rewards_agr, rewards_agr2, rewards_agr3]
# batch size - algorithms will be refit after N rounds
batch_size = 50
# initial seed - all policies start with the same small random selection of actions/rewards
first_batch = X[:batch_size, :]
np.random.seed(1)
action_chosen = np.random.randint(nchoices, size=batch_size)
rewards_received = y[np.arange(batch_size), action_chosen]
# fitting models for the first time
for model in models:
model.fit(X=first_batch, a=action_chosen, r=rewards_received)
# these lists will keep track of which actions does each policy choose
lst_a_agr, lst_a_agr2, lst_a_agr3 = [action_chosen.copy() for i in range(len(models))]
lst_actions = [lst_a_agr, lst_a_agr2, lst_a_agr3]
# rounds are simulated from the full dataset
def simulate_rounds(model, rewards, actions_hist, X_global, y_global, batch_st, batch_end):
np.random.seed(batch_st)
## choosing actions for this batch
actions_this_batch = model.predict(X_global[batch_st:batch_end, :]).astype('uint8')
# keeping track of the sum of rewards received
rewards.append(y_global[np.arange(batch_st, batch_end), actions_this_batch].sum())
# adding this batch to the history of selected actions
new_actions_hist = np.append(actions_hist, actions_this_batch)
# now refitting the algorithms after observing these new rewards
np.random.seed(batch_st)
model.fit(X_global[:batch_end, :], new_actions_hist, y_global[np.arange(batch_end), new_actions_hist])
return new_actions_hist
# now running all the simulation
for i in tqdm(range(int(np.floor(X.shape[0] / batch_size)))):
batch_st = (i + 1) * batch_size
batch_end = (i + 2) * batch_size
batch_end = np.min([batch_end, X.shape[0]])
for model in range(len(models)):
lst_actions[model] = simulate_rounds(models[model],
lst_rewards[model],
lst_actions[model],
X, y,
batch_st, batch_end)
# +
import matplotlib.pyplot as plt
from pylab import rcParams
# %matplotlib inline
def get_mean_reward(reward_lst, batch_size=batch_size):
mean_rew=list()
for r in range(len(reward_lst)):
mean_rew.append(sum(reward_lst[:r+1]) * 1.0 / ((r+1)*batch_size))
return mean_rew
rcParams['figure.figsize'] = 25, 15
lwd = 5
cmap = plt.get_cmap('tab20')
colors=plt.cm.tab20(np.linspace(0, 1, 20))
ax = plt.subplot(111)
plt.plot(get_mean_reward(rewards_agr), label="Adaptive Greedy (replicate)",linewidth=lwd,color=colors[10])
plt.plot(get_mean_reward(rewards_agr2), label="Adaptive Greedy (discount)",linewidth=lwd,color=colors[12])
plt.plot(get_mean_reward(rewards_agr3), label="Adaptive Greedy (rf)",linewidth=lwd,color=colors[14])
# import warnings
box = ax.get_position()
ax.set_position([box.x0, box.y0 + box.height * 0.1,
box.width, box.height * 1.25])
ax.legend(loc='upper center', bbox_to_anchor=(0.5, -0.05),
fancybox=True, ncol=3, prop={'size':20})
plt.tick_params(axis='both', which='major', labelsize=25)
plt.xticks([i*20 for i in range(8)], [i*1000 for i in range(8)])
plt.xlabel('Rounds (models were updated every 50 rounds)', size=30)
plt.ylabel('Cumulative Mean Reward', size=30)
plt.title('Comparison of Online Contextual Bandit Policies\n(Base Algorithm is Logistic Regression)\n\nBibtext Dataset\n(159 categories, 1836 attributes)',size=30)
plt.grid()
plt.show()
# -
| src/Extension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:semantique]
# language: python
# name: conda-env-semantique-py
# ---
# # Advanced usage
#
# In this notebook we will go a bit deeped into internal structures of the semantique package. We will also show some common operations for advanced users, that want more control over the behaviour of the query processor, and extend built-in structures of the package. It is assumed that you are already quite familiar with the structure and contents of the package, for example by reading the other provided notebooks.
#
# ## Content
#
# - [The query processor class](#The-query-processor-class)
# - [Internal data cube representations](#Internal-data-cube-representations)
# - [Applying verbs directly to xarray objects](#Applying-verbs-directly-to-xarray-objects)
# - [Converting cubes to other formats](#Converting-cubes-to-other-formats)
# - [Creating a spatio-temporal extent cube](#Creating-a-spatio-temporal-extent-cube)
# - [Aligning cubes to each other](#Aligning-cubes-to-each-other)
# - [Tracking value types](#Tracking-value-types)
# - [Adding custom operators](#Adding-custom-operators)
# - [Adding custom reducers](#Adding-custom-reducers)
# - [Tuning factbase configuration settings](#Tuning-factbase-configuration-settings)
# - [Creating custom factbase classes](#Creating-custom-factbase-classes)
# - [Creating custom ontology classes](#Creating-custom-ontology-classes)
# - [Logging progress](#Logging-progress)
#
# ## Prepare
#
# Import the semantique package:
import semantique as sq
# Import other packages we will use in this demo:
import geopandas as gpd
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
import copy
import json
import inspect
# Create the components for query processing. See the [Intro notebook](intro.ipynb) for details.
# +
# Query recipe.
with open("files/recipe.json", "r") as file:
recipe = sq.QueryRecipe(json.load(file))
# Ontology.
with open("files/ontology.json", "r") as file:
ontology = sq.ontology.Semantique(json.load(file))
# Factbase.
with open("files/factbase.json", "r") as file:
factbase = sq.factbase.GeotiffArchive(json.load(file), src = "files/resources.zip")
# Extent.
space = sq.SpatialExtent(gpd.read_file("files/footprint.geojson"))
time = sq.TemporalExtent("2019-01-01", "2020-12-31")
# Additional configuration.
config = {"crs": 3035, "tz": "UTC", "spatial_resolution": [-10, 10]}
# -
# ## The query processor class
#
# A semantic query is processed by a query processor. In semantique, this query processor is internally modelled as an object of class [QueryProcessor](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.html). An instance of this class is initialized whenever a query recipe is executed with respect to a given factbase, ontology and spatio-temporal extent. That is, whenever you call the [execute()](https://zgis.github.io/semantique/_generated/semantique.QueryRecipe.execute.html) method of a [QueryRecipe](https://zgis.github.io/semantique/_generated/semantique.QueryRecipe.html) instance, semantique internally creates a query processor object to take care of all processing tasks.
from semantique.processor.core import QueryProcessor
# The query processor processes a query in three three core phases: query parsing, query optimization and query execution. These align with the common phases in regular relational database querying (see for example [this paper](https://dsf.berkeley.edu/papers/fntdb07-architecture.pdf)). Each of these phases has their own, dedicated method. Below we will explain in more detail how each phase is implemented in semantique.
#
# ### Query parsing
#
# During query parsing, the required components for processing (i.e., the query recipe, the spatial and temporal extents, the ontology, the factbase and the additional configuration parameters) are read and converted all together into a single object which will be used internally for further processing of the query. Hence, query parsing takes care of initializing a [QueryProcessor](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.html) instance. It also rasterizes the given spatial extent and combines it with the temporal extent into a single spatio-temporal array. See the [Creating a spatio-temporal extent cube](#Creating-a-spatio-temporal-extent-cube) section for details.
#
# The query processor contains the [parse()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.parse.html) method (which is a [classmethod](https://www.programiz.com/python-programming/methods/built-in/classmethod)) to run the parsing phase. As arguments, it expects all required [components](intro.ipynb#Components) for semantic query processing. Additional configuration parameters can be provided as well.
#
# Ideally, parsing should also take care of validating the components and their interrelations. For example, it should check if referenced concepts in the provided query recipe are actually defined in the provided ontology. Such functionality is not implemented yet in the current version of semantique.
processor = QueryProcessor.parse(recipe, factbase, ontology, space, time, **config)
type(processor)
# ### Query optimization
#
# During query optimization, the query components are scanned and certain properties of the query processor are set. These properties influence some tweaks in how the query processor will behave when processing the query. For example, if the given spatial extent consists of multiple dispersed sub-areas, the query processor might instruct itself to load data separately for each sub-area, instead of loading data for the full extent and then subset it afterwards.
#
# The query processor contains the [optimize()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.optimize.html) method to run the optimization phase. It returns the same instance, but with updated properties. However, in the current version of semantique, the optimization phase only exists as a placeholder, and no properties are updated yet.
processor = processor.optimize()
type(processor)
# ### Query execution
#
# Finally, the execution phase is where the query recipe is executed. The query processor contains specific handlers for all kind of operations that may be specified in the recipe. Internally it will take care of the interaction with (and between) the given ontology and factbase, and forward required parameters to their translator and retriever accordingly.
#
# The query processor contains the [execute()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.execute.html) method to run the optimization phase. It returns the same instance, but with a [response](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.response.html) property containing the outputs of all formulated results in the query recipe.
# Before execution the response property is empty.
processor.response
processor = processor.execute()
type(processor)
# After execution the response property is filled.
processor.response
# When executing a query recipe, the query processor moves through the building blocks of the result instructions. For each building block, it has a specific **handler function** implemented as a method. Whenever the processor reaches a new block, it looks up its type and then calls its handler with the [call_handler()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.call_handler.html) method.
#
# For example, when the processor stumbles upon a reference to a semantic concept, it forwards it to the [call_handler()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.call_handler.html) method, which will notice this is a block of type "concept", and therefore call the handler [handle_concept()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.handle_concept.html). This handler knows exactly what to do with this type of block. In this specific example, it calls the translator function of the provided ontology to obtain the boolean data cube belonging to the referenced concept.
water = processor.handle_concept(sq.entity("water"))
values = [0, 1]
levels = [x - 0.5 for x in values + [max(values) + 1]]
colors = ["#d3d3d3", "#77abb7"]
legend = {"ticks": values, "label": "IsWater"}
water.unstack().plot(col = "time", levels = levels, colors = colors, cbar_kwargs = legend)
vegetation = processor.handle_concept(sq.entity("vegetation"))
legend = {"ticks": values, "label": "IsVegetation"}
vegetation.unstack().plot(col = "time", levels = levels, colors = colors, cbar_kwargs = legend)
# When moving through the building blocks of a processing chain, the object (i.e. data cube) under evaluation constantly changes. It starts as the evaluated reference in the *with* part of the chain. This object will be the input to the first action in the *do* part of the chain, which wrangles it into a different cube. That different cube will then be the input to the second action, which again wrangles it into a different cube, et cetera. We use the term **active evaluation object** to refer to the input object at each stage of the processing chain. Hence, at the first action, the active evaluation object is the evaluated reference. At the second action, the active evaluation object is the output of the first action, et cetera.
#
# In practice, this means that when the query processor stumbles upon a processing chain, it forwards it to the [call_handler()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.call_handler.html) method, which will notice this is a block of type "processing_chain", and therefore call the handler [handle_processing_chain()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.handle_processing_chain.html). This handler knows exactly what to do with this type of block. It first moves into the *with* part of the chain, and processes the reference (or a nested processing chain) it finds there by calling the corresponding handler. Then, it sets the obtained data cube as the new active evaluation object, and moves into the *do* part of the chain. For each verb, it calls the handler [handle_verb()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.handle_verb.html), which in turn calls the verb-specific handler (e.g. [handle_filter()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.handle_filter.html)) to apply the verb to the active evaluation object, and then replaces the active evaluation object with the output of the verb. Et cetera.
#
# > **NOTE** <br/> To get, set, and replace the active evaluation object, the query processor uses private methods such as `_get_eval_obj()`, `_set_eval_obj()` and `_replace_eval_obj()`.
# +
chain = sq.entity("water").\
filter_time("year", "equal", 2020).\
evaluate("or", sq.entity("vegetation"))
out = processor.handle_processing_chain(chain)
# -
values = [0, 1]
levels = [x - 0.5 for x in values + [max(values) + 1]]
legend = {"ticks": values, "label": "IsWaterOrVegetation"}
out.unstack().plot(col = "time", levels = levels, colors = colors, cbar_kwargs = legend)
# ### Query response
#
# To actually make the query processor return the response, you have to call the [respond()](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.respond.html) method of the query processor. Before returning the result arrays, this method will first post-process them.
#
# One of the post-processing steps is *trimming* the result arrays. This means that all coordinates for which all values are nodata, are dropped from the array. The spatial dimension (if present) is treated differently, by trimming it only at the edges, and thus maintaining the regularity of the spatial dimension.
#
# Trimming results can be disabled by adding the configuration parameter "trim_results" when initializing the query processor, and set its value to `False`.
#
# ```python
# config["trim_results"] = False
# ```
#
# Note the difference:
# +
new_recipe = sq.QueryRecipe()
new_config = copy.deepcopy(config)
new_recipe["foo"] = sq.entity("water").filter_time("year", "equal", 2020)
# +
out = new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"]
values = [0, 1]
levels = [x - 0.5 for x in values + [max(values) + 1]]
legend = {"ticks": values, "label": "IsWater"}
out.unstack().plot(col = "time", levels = levels, colors = colors, cbar_kwargs = legend)
# -
new_config["trim_results"] = False
# +
out = new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"]
values = [0, 1]
levels = [x - 0.5 for x in values + [max(values) + 1]]
legend = {"ticks": values, "label": "IsWater"}
out.unstack().plot(col = "time", levels = levels, colors = colors, cbar_kwargs = legend)
# -
# Another post-processing step is *unstacking* the result arrays. Internally the query processor always works with arrays that have a stacked spatial dimension, meaning the the x and y dimension are stacked together into a single multi-indexed space dimension. Before responding, the query processor unstacks the space dimension, such that the returned arrays have separate x and y dimensions again.
#
# Unstacking results can be disabled by adding the configuration parameter "unstack_results" when initializing the query processor, and set its value to `False`.
#
# ```python
# config["unstack_results"] = False
# ```
#
# Note the difference:
# +
new_recipe = sq.QueryRecipe()
new_config = copy.deepcopy(config)
new_recipe["foo"] = sq.entity("water").filter_time("year", "equal", 2020)
# -
out = new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"]
out
new_config["unstack_results"] = False
out = new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"]
out
# ## Internal data cube representations
#
# Internally the query processor uses [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) objects from the [xarray](https://docs.xarray.dev/en/stable/) package to represent data cubes. It extends these objects by adding a so-called *accessor* with the name **sq**. In practice, this means that all semantique-specific methods and properties for these objects can be called with a `.sq` prefix:
#
# ```python
# xarray_object.sq.method()
# xarray_object.sq.property
# ```
#
# Extending xarray in this way is recommended by its developers. Read more about it [here](https://docs.xarray.dev/en/stable/internals/extending-xarray.html).
#
# The semantique-specific properties for data arrays include for example the [value type](#Tracking-value-types) of the cube as well as spatio-temporal properties like the spatial resolution, the coordinate reference system in which the spatial coordinates are expressed and the time zone in which the temporal coordinates are expressed.
#
# The semantique-specific methods for data arrays include all the [verbs](#Applying-verbs-directly-to-xarray-objects), i.e. the actions that can be applied to a data cube. It also include other functionalities that are internally used by the query processor, such as [trimming](#Query-response), [unstacking](#Query-response) and [aligning](#Aligning-cubes-to-each-other). Besides that, there are methods for [conversions](#Converting-cubes-to-other-formats) to other objects or to files on disk.
#
# For all semantique specific methods and properties for data arrays, see the [API reference](https://zgis.github.io/semantique/_generated/semantique.processor.structures.Cube.html).
#
# ### Data cube collections
#
# For collections of multiple data cubes, semantique contains the [CubeCollection](https://zgis.github.io/semantique/_generated/semantique.processor.structures.CubeCollection.html) class. This is simply a list of [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) objects. Its methods include the [collection-specific verbs](verbs.ipynb#Verbs-for-data-cube-collections), as well as some single cube methods that are mapped over each member of the collection.
#
# For all specific methods and properties of these objects, see the [API reference](https://zgis.github.io/semantique/_generated/semantique.processor.structures.CubeCollection.html). To ensure compatible behaviour with single data cubes, you may call all these methods and properties as well with the prefix `.sq`. However, this is not an xarray accessor, and merely provided to simplify internal code. With cube collections, you can just as well call its methods and properties directly, without prefix.
# ## Applying verbs directly to xarray objects
#
# *To be added*
# ## Converting cubes to other formats
#
# *To be added*
# ## Creating a spatio-temporal extent cube
#
# During [query parsing](#Query-parsing), the query processor constructs a two-dimensional data cube that represents the spatio-temporal extent of the query. For that, it uses the [create_extent_cube()](https://zgis.github.io/semantique/_generated/semantique.processor.utils.create_extent_cube.html) function. The workflow is as follows:
#
# 1) The spatial extent is *reprojected* and *rasterized* given the specified coordinate reference system and spatial resolution in the configuration parameters. The resulting array is always rectangular and therefore covers the full *bounding box* of the area that was orginally given as spatial extent. Cells that don't overlap with the area boundaries itself are filled with nodata values. Others are filled with an integer. If the given area consisted of a single feature, they are all filled with a value of 1. If the given area consisted of multiple disconnected features, each feature gets a different index value.
#
# 2) The two spatial dimensions are *stacked* together into a single "space" dimension. This dimension has as many coordinates as the number of cells in the trimmed spatial array. Each coordinate value is a tuple of the x and y coordinate of the corresponding cell.
#
# 3) This stacked spatial array is *expanded* over a time dimension. The time dimension has two coordinates: the first coordinate value corresponds to the timestamp at the start of the given time interval, and the second coordinate value corresponds to the timestamp at the end of the given time interval. The time values are *converted* into the specified time zone in the configuration parameters.
#
# 4) The spatio-temporal array is *trimmed*, meaning that all coordinates for which all pixel values are nodata, are removed from the array. The spatial dimension (if present) is treated differently, by trimming it only at the edges, and thus maintaining the regularity of the spatial dimension.
#
# You can call the [create_extent_cube()](https://zgis.github.io/semantique/_generated/semantique.processor.utils.create_extent_cube.html) function directly to create a spatio-temporal extent cube from respectively a spatial and temporal extent. It may occur that due to the reprojection of the provided spatial extent, some pixels at the edges of the extent get a nodata value.
from semantique.processor.utils import create_extent_cube
crs = config["crs"]
tz = config["tz"]
res = config["spatial_resolution"]
extent = create_extent_cube(space, time, res, crs = crs, tz = tz)
extent
# Complying with common standards (see e.g. the discussion [here](https://github.com/opendatacube/datacube-core/issues/837)), the coordinate reference system of the spatial coordinates is stored as attribute of a specific non-dimension coordinate named "spatial_ref". Storing this inside a non-dimension coordinate instead of as direct attribute of the array guarantees that this information is preserved during any kind of operation. The coordinate itself serves merely as a placeholder.
print(extent["spatial_ref"].attrs)
# In the same style, semantique stores time zone information as attribute of a non-dimension coordinate named "temporal_ref". Note that time coordinates themselves are stored as [numpy.datetime64](https://numpy.org/doc/stable/reference/arrays.datetime.html) objects, which in itself don't support timezone information anymore. Therefore, the "temporal_ref" coordinate is an important piece of information in the spatio-temporal extent cube.
print(extent["temporal_ref"].attrs)
# The query processor obtains such spatio-temporal information by calling specific properties through the sq-accessor of the data cube, rather than scanning the above-mentioned non-dimension coordinates directly.
extent.sq.crs
type(extent.sq.crs)
extent.sq.tz
type(extent.sq.tz)
extent.sq.spatial_resolution
# ## Aligning cubes to each other
#
# When evaluating bivariate expressions with the [evaluate()](verbs.ipynb#Bivariate-expressions) verb, cube $Y$ does not necessarily have to be of the same shape as input cube $X$, but it should at least be possible to *align* it to that shape. This can be done in two ways.
#
# First consider the case where $Y$ has the same dimensions as $X$, but not all coordinate values of $X$ are present in $Y$. In that case, we can align $Y$ with $X$ such that pixel values at position $i$ in both cubes, i.e. $x_{i}$ and $y_{i}$ respectively, belong to pixels with the *same coordinates*. If $y_{i}$ was not originally part of $Y$, we assign it a nodata value. This also works vice-versa, with coordinate values in $Y$ that are not present in $X$.
#
# Secondly, consider a case where $Y$ has one or more dimensions with exactly the same coordinate values as $X$, but does not have *all* the dimensions that $X$ has. In that case, we can align $Y$ with $X$ by duplicating its values along those dimensions that are missing. This does *not* work vice versa. When cube $Y$ has more dimensions that cube $X$, there is no clear way to define how to subset the values in $Y$ to match the shape of $X$.
#
# See the [Verbs notebook](verbs.ipynb#Bivariate-expressions) for details.
#
# The bivariate operators of the query processor take care of aligning cube $Y$ to cube $X$ before applying the operation. To do so, they use the [align_with()](https://zgis.github.io/semantique/_generated/semantique.processor.structures.Cube.align_with.html) method of the [sq-accessor](#Internal-data-cube-representations) to data arrays.
#
# For example:
# Create a cube X.
x = water.isel(space = range(0, 9))
x
# Create a cube Y with only a subset of the coordinates of cube X.
y = vegetation.isel(space = range(3, 6))
y
# Create a cube z with only one dimension of cube x.
z = vegetation.isel(space = 0, drop = True)
z
# Aligning y to x makes sure that pixels with equal coordinates are at the same position.
y.sq.align_with(x)
# Aligning z to x duplicates for each timestamp the values of z along all spatial locations in x.
z.sq.align_with(x)
# ## Tracking value types
#
# During query processing references get translated into data cubes, and these data cubes are wrangled by specified actions. Along this road, the query processor keeps track of the *value type* of each data cube. Such a value type describes what kind of data the cube contains. It differs from the very technical, computer-oriented [numpy.dtype](https://numpy.org/doc/stable/reference/arrays.dtypes.html) categorization, which contains e.g. *int*, *float*, etc. Instead, the *value type* describes data on a more general, statistical level. Currently it makes a distinction between three main value types:
#
# - Numerical
# - Nominal
# - Ordinal
# - Binary
#
# Additional value types exist for spatio-temporal data:
#
# - datetime (for timestamps)
# - coords (for spatial coordinate tuples)
# - geometry (for spatial geometries stored [GeoDataFrame](https://geopandas.org/en/stable/docs/reference/api/geopandas.GeoDataFrame.html) objects)
#
# It is expected that whenever a data cube is retrieved from the factbase, the corresponding value type is stored as an semantique-specific array attribute [value_type](https://zgis.github.io/semantique/_generated/semantique.processor.structures.Cube.value_type.html). Even when the data are not qualitative, they are usually stored as numbers. In these cases, an additional attribute [value_labels](https://zgis.github.io/semantique/_generated/semantique.processor.structures.Cube.value_labels.html) may be used to define the mapping between character-encoded *labels* and integer-encoded *indices*.
colors = factbase.retrieve("appearance", "Color type", extent = extent)
colors.sq.value_type
colors.sq.value_labels
water = ontology.translate("entity", "water", extent = extent, factbase = factbase)
water.sq.value_type
# Whenever applying actions to a data cube, its value type might change. For example, when evaluating an expression (e.g. when evaluating an expression involving a comparison operator the resulting values are always binary) or applying a reducer (e.g. when counting the number of "true" values in a binary data cube the resulting values are numerical). This is what we also call **type promotion**. Each implemented operator and reducer function is able to promote the type of the output given the type(s) of the input(s) by using a type promotion manual.
#
# A common type promotion manual for a univariate operator and a reducer has a dictionary structure with as keys the supported input value types. The value of each key is the output value type. For example, the [count()](https://zgis.github.io/semantique/_generated/semantique.processor.reducers.count_.html) reducer should only be applied to data cubes containing binary data, since it counts the number of "true" values. The output of that operation, however, is a data cube with numerical data. Hence, the corresponding type promotion manual looks like this:
from semantique.processor.types import TYPE_PROMOTION_MANUALS
TYPE_PROMOTION_MANUALS["count"]
# The additional key "preserve_labels" defines if value labels should be preserved after the reducer is applied, with 0 being "no" and 1 being "yes".
#
# The reduction function will apply the type promotion whenever argument "track_types" is set to `True`, which is the default. When we apply the count reducer to a binary data cube, this means the value type of the output will be numerical:
from semantique.processor import reducers
new = water.sq.reduce("time", reducers.count_)
new.sq.value_type
# Now, when we call this reducer function on e.g. an ordinal data cube instead of a binary data cube, an [InvalidValueTypeError](https://zgis.github.io/semantique/_generated/semantique.exceptions.InvalidValueTypeError.html) will be thrown. Hence, the query processor uses type tracking also to *check* type promotions, and thus, to detect invalid operations.
from semantique.exceptions import InvalidValueTypeError
try:
colors.sq.reduce("time", reducers.count_)
except InvalidValueTypeError as e:
print(e)
# This will not happen when `track_types = False`. In that case, value types are not tracked and the output cube has no value type at all.
new = colors.sq.reduce("time", reducers.count_, track_types = False)
"value_type" in new.attrs
# For bivariate expressions, the type promotion manuals have an extra "layer" for the second operand. The first layer of keys refers to the value type of the first operand (i.e. x) and the second layer of keys to the value type of the second operand (i.e. y). For example, the boolean operator [and()](https://zgis.github.io/semantique/_generated/semantique.processor.operators.and_.html) accepts only data cubes with binary values, and returns binary values as well. Hence, the corresponding type promotion manual looks like this:
TYPE_PROMOTION_MANUALS["and"]
# In the case of these bivariate operators, the "preserve_labels" key may also have a value of 2, meaning that instead of the labels of the first operand, the labels of the second operand should be preserved (this is the case e.g. with the [assign()](https://zgis.github.io/semantique/_generated/semantique.processor.operators.assign_.html) operator).
#
# Just as with the reducers, the operators promote the value type of the output based on the manual, and throw an error if a combination of types is not supported.
from semantique.processor import operators
new = water.sq.evaluate(operators.and_, water)
new.sq.value_type
try:
colors.sq.evaluate(operators.and_, water)
except InvalidValueTypeError as e:
print(e)
# If the second operand in a bivariate expression is not a cube but a single value, its value type is determined based on its [numpy data type](https://numpy.org/doc/stable/reference/arrays.dtypes.html). In semantique, each of these numpy data types is mapped to one or more semantique value types. In the case of a mapping to multiple value types, all operations supporting at least one of these types will accept the value.
from semantique.processor.types import DTYPE_MAPPING
DTYPE_MAPPING
new = water.sq.evaluate(operators.and_, True)
new.sq.value_type
# Inside the reducer and operator function, an instance of the [TypePromoter](https://zgis.github.io/semantique/_generated/semantique.processor.types.TypePromoter.html) class is the worker that actually takes care of the type promotion. It can be initialized by providing it with the operand(s) of the process, and the name of the function. Then, it will find the type promotion manual belonging to that function. Alternatively (e.g. when you add your own reducers or operators, see the next sections) you can provide it your own custom type promotion manual.
#
# The promoter can determine the value types of the operands and check if they are supported by the operation. Subsequently, it can promote the value type of the operation output.
from semantique.processor.types import TypePromoter
promoter = TypePromoter(water, colors, function = "and")
try:
promoter.check()
except InvalidValueTypeError as e:
print(e)
promoter = TypePromoter(water, water, function = "and")
promoter.check()
promoter.promote(new)
# Now what if you don't want the query processor to track types at all? There are several reasons why you'd want to do so. For example, type tracking can be so strict that it limits your flexibility of using certain processes "out of the box". Some custom factbases you use may not set value types at all, and some custom operators or reducers that you use may not have any type promotion manual. Also, type tracking of course takes time and thus decreases query performance. No worries, it is easy to tell the query processor to *not* track types. Simply add the configuration parameter "track_types" when initializing the query processor, and set its value to `False`:
#
# ```python
# config["track_types"] = False
# ```
# ## Adding custom operators
#
# Operators are used when evaluating expressions on data cubes with the [evaluate()](https://zgis.github.io/semantique/_notebooks/verbs.html#Evaluate) verb. For example, you can compare each value in a data cube with a given constant (using e.g. the [greater()](https://zgis.github.io/semantique/_generated/semantique.processor.operators.greater_.html) operator), add two data cubes together (using the [add()](https://zgis.github.io/semantique/_generated/semantique.processor.operators.add_.html) operator), etcetera. When creating a query recipe, you can refer to these operators by their name, for example:
#
# ```python
# sq.result("water_count").evaluate("multiply", 2)
# ```
#
# The query processor instance has a [operators](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.operators.html) property in
# which it stores all supported operator functions. The keys of this dictionary are the operator names, such that the query processor can quickly retrieve the corresponding function of any referenced operator in the query recipe. For example:
processor.operators["multiply"]
# As can be seen, the operator function itself is taken from the [operators module](https://zgis.github.io/semantique/reference.html#operator-functions) of semantique. This is the module in which all built-in operator functions are defined. When initializing a query processor with default settings, these are the operators that get stored in the processors [operators](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.operators.html) property.
for x in processor.operators.keys():
print(x)
# That all operator functions are stored inside a property of the query processor means that you can easily overwrite them with your own custom operator functions (or add new ones to the list of existing ones) after initializing a query processor instance. However, you can also do this already at initialization itself, by setting specific configuration parameters.
#
# But let us first prepare a custom operator. As shown above this requires a name, an operator function and a type promotion manual. If you plan to disable type tracking either way, the latter is of course not needed. An operator function has to accept a [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) object as its first argument. This is the *active evaluation object* in the processing chain, to which the verb is applied. Bivariate operators also accept a second argument, which can either be a [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) object as well, or a single value. The return value of the operator must again be a [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) object. Usually it is a good idea to use the [xarray.apply_ufunc()](http://xarray.pydata.org/en/stable/generated/xarray.apply_ufunc.html) function inside operator functions, as shown below. Also, you should not forget to [align](#Aligning-cubes-to-each-other) the second operand to the input object before applying the core operation!
#
# > **NOTE** <br/> It is also possible to implement operator functions with more than two operands. All additional keyword arguments that the evaluate verb receives are simply forwarded to the operator function. You could for example build a query recipe like the following:
# > ```python
# sq.result("water_count").evaluate("custom", y = 1, foo = 10, bar = 100)
# > ```
# > You then need to implement an operator with four arguments:
# > ```python
# def custom(x, y, foo, bar)
# > ```
#
# If you want to [track value types](#Tracking-value-types), the custom operator should also have a binary "track_types" argument. Inside the function, the type promotion should be applied whenever the "track_types" argument is set to `True`. You can create your own type promotion manual.
#
# Lets create a custom operator that calculates the normalized difference between two values.
def normalized_difference(x, y, track_types = True, **kwargs):
if track_types:
manual = {
"numerical": {
"numerical": "numerical"
},
"__preserve_labels": 0
}
promoter = TypePromoter(x, y, manual = manual)
promoter.check()
f = lambda x, y: np.divide(np.subtract(x, y), np.add(x, y))
y = xr.DataArray(y).sq.align_with(x)
out = xr.apply_ufunc(f, x, y)
if track_types:
out = promoter.promote(out)
return out
# When executing a query recipe, we can now add this operator to the configuration parameters. There are two ways:
#
# 1) Setting the "operators" parameter will fully replace the default operators with the custom one(s).
# ```python
# config["operators"] = {"normalized_difference": normalized_difference}
# ```
#
# 2) Setting the "extra_operators" parameter will add the custom operator(s) to the default ones. If a custom operator has the same name as a default operator, the default operator will be replaced by the custom operator.
# ```python
# config["extra_operators"] = {"normalized_difference": normalized_difference}
# ```
# +
new_recipe = sq.QueryRecipe()
new_config = copy.deepcopy(config)
red = sq.reflectance("s2_band04")
nir = sq.reflectance("s2_band08")
new_recipe["ndvi"] = nir.evaluate("normalized_difference", red)
# -
from semantique.exceptions import UnknownOperatorError
try:
print(new_recipe.execute(factbase, ontology, space, time, **new_config)["ndvi"].round(1))
except UnknownOperatorError as e:
print(e)
new_config["extra_operators"] = {"normalized_difference": normalized_difference}
try:
print(new_recipe.execute(factbase, ontology, space, time, **new_config)["ndvi"].round(1))
except UnknownOperatorError as e:
print(e)
# ## Adding custom reducers
#
# Reducers are used inside the [reduce()](https://zgis.github.io/semantique/_notebooks/verbs.html#Reduce) verb as a function to aggregate values along a dimension of a data cube. For example, you can reduce the temporal dimension of a data cube by calculating the average value of each slice along the time dimension axis (using the [mean()](https://zgis.github.io/semantique/_generated/semantique.processor.reducers.mean_.html) reducer), etcetera. When creating a query recipe, you can refer to these reducers by their name, for example:
#
# ```python
# sq.result("water_count_space").reduce("time", "mean")
# ```
#
# The query processor instance has a [reducers](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.reducers.html) property in which it stores all supported reducer functions. The keys of this dictionary are the reducer names, such that the query processor can quickly retrieve the corresponding function of any referenced reducer in the query recipe. For example:
processor.reducers["mean"]
# As can be seen, the reducer function itself is taken from the [reducers module](https://zgis.github.io/semantique/reference.html#reducer-functions) of semantique. This is the module in which all built-in reducer functions are defined. When initializing a query processor with default settings, these are the reducers that get stored in the processors [reducers](https://zgis.github.io/semantique/_generated/semantique.processor.core.QueryProcessor.reducers.html) property.
for x in processor.reducers.keys():
print(x)
# That all reducer functions are stored inside a property of the query processor means that you can easily overwrite them with your own custom reducer functions (or add new ones to the list of existing ones) after initializing a query processor instance. However, you can also do this already at initialization itself, by setting specific configuration parameters.
#
# But let us first prepare a custom reducer. As shown above this requires a name, a reducer function and a type promotion manual. If you plan to disable type tracking either way, the latter is of course not needed. A reducer function has to accept a [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) object as its first argument. This is the *active evaluation object* in the processing chain, to which the verb is applied. The function should also accept a "dimension" argument, containing the name of the dimension to reduce over. The return value of the reducer must again be a [DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray) object. Usually it is a good idea to use the [xarray.reduce()](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.reduce.html) method inside reducer functions, as shown below. This function will take care e.g. of translating the dimension name into its corresponding axis number.
#
# > **NOTE** <br/> All additional keyword arguments that the reduce verb receives are simply forwarded to the reducers function. That allows you to include additional variables in your reducer functions (e.g. constants). You could for example build a query recipe like the following:
# >```python
# sq.result("water").reduce("time", "custom", alpha = 0.5)
# >```
# > You then need to implement a reducer with four arguments:
# > ```python
# def custom(x, dimension, alpha)
# > ```
#
# If you want to [track value types](#Tracking-value-types), the custom reducer should also have a binary "track_types" argument. Inside the function, the type promotion should be applied whenever the "track_types" argument is set to `True`. You can create your own type promotion manual.
#
# Lets create a custom operator that reduces a set of values to its sum of squares.
def sum_of_squares(x, dimension, track_types = False, **kwargs):
if track_types:
manual = {"numerical": "numerical", "__preserve_labels": 0}
promoter = TypePromoter(x, manual = manual)
promoter.check()
f = lambda x, axis: np.sum(np.square(x), axis)
out = x.reduce(f, dim = dimension)
if track_types:
promoter.promote(out)
return out
# When executing a query recipe, we can now add this reducer to the configuration parameters. There are two ways:
#
# 1) Setting the "reducers" parameter will fully replace the default reducers with the custom one(s).
# ```python
# config["reducers"] = {"sum_of_squares": sum_of_squares}
# ```
#
# 2) Setting the "extra_reducers" parameter will add the custom reducer(s) to the default ones. If a custom reducer has the same name as a default reducer, the default reducer will be replaced by the custom reducer.
# ```python
# config["extra_reducers"] = {"sum_of_squares": sum_of_squares}
# ```
# +
new_recipe = sq.QueryRecipe()
new_config = copy.deepcopy(config)
nir = sq.reflectance("s2_band08")
new_recipe["foo"] = nir.reduce("time", "sum_of_squares")
# -
from semantique.exceptions import UnknownReducerError
try:
print(new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"])
except UnknownReducerError as e:
print(e)
new_config["extra_reducers"] = {"sum_of_squares": sum_of_squares}
try:
print(new_recipe.execute(factbase, ontology, space, time, **new_config)["foo"])
except UnknownReducerError as e:
print(e)
# ## Tuning factbase configuration settings
#
# *To be added*
# ## Creating custom factbase classes
#
# *To be added*
# ## Creating custom ontology classes
#
# *To be added*
# ## Logging progress
#
# When processing a query, the query processor logs its progress. Most of these logs are of the "DEBUG" level. They are helpful to debug the query processing, since they show intermediate outputs when the query processor iterates through the building blocks in the query recipe.
# +
import logging
import sys
logger = logging.getLogger("semantique.processor.core")
logger.setLevel(logging.DEBUG)
logger.addHandler(logging.StreamHandler(stream = sys.stdout))
# -
small_recipe = sq.QueryRecipe({"water_map": recipe["water_map"]})
small_recipe.execute(factbase, ontology, space, time, **config)
| demo/advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Module 4: lab practice - Subqueries and multiple tables
#
#
# ## Part A: Subqueries and nested selects
#
#
# ### Query A1: Introduce a fail (get an error) obtaining every employee which salary are bigger than avg salary.
#
# ```
# select *
# from employees
# where salary > avg(salary)
# ;
# ```
#
# ### Query A2: Introduce a query that works using a subquery to obtain every employee which salary are bigger than avg salary.
#
# ```
# select *
# from employees
# where salary > (select avg(salary) from employees)
# ;
# ```
#
# ### Query A3: Introduce a query that fails obtaining every row of employees table and the avg salary in every row.
#
# ```
# select *, avg(salary)
# from employees;
# ```
#
# ### Query A4: Introduce a column expression to get every employees and avg salary in the same row.
#
# ```
# select emp_id, salary, (select avg(salary) from employees) as "avg_salary"
# from employees;
# ```
#
# ### Query A5: Introduce a table expression to get only the columns with non confidencial data of every employee.
#
# ```
#
# select *
# from (select f_name, l_name, b_date from employees ) as EMNCF;
#
# ```
#
# ## Part B Multiple tables with sub-queries
#
#
# ### Query B1: Get only the rows of employees that are the same id of departments table.
#
# ```
# select * from
# employees where dep_id
# IN ( select dept_id_dep from departments );
# ```
#
# ### Query B2: Get the list of employees with place equals "L0002".
#
# ```
# select *
# from employees
# where dep_id in
# (select dept_id_dep from departments where loc_id = 'L0002');
# ```
#
# ### Query B3: Get the department ID and department name of the employee names that earn more than 70000.
#
# ```
# select dept_id_dep, dep_name
# from departments
# where dept_id_dep in
# (select dep_id from employees where salary > 70000);
# ```
#
# ### Query B4: Introduce a query 2 tables in the from clause.
#
# ```
# select *
# from employees, departments;
# ```
#
# ## Access to multiple tables with implicit joins
#
# ### Query B5: Get all the employees that are in departments table.
#
# ```
# select employees.*
# from employees, departments
# where employees.dep_id = departments.dept_id_dep;
#
# ```
#
# ### Query B6: Use aliases to shorten the query
#
# ```
# select em.*
# from employees em, departments dp
# where em.dep_id = dp.dept_id_dep;
# ```
#
# ### Query B7: Get only the employee ID and name department from the previous query
#
# ```
# select em.emp_id, dp.dep_name
# from employees em, departments dp
# where em.dep_id = dp.dept_id_dep;
# ```
#
# ### Query B8: From the previous query, specify the name the of columns with aliases
#
# ```
# select em.emp_id as "employee id", dp.dep_name as "dept name"
# from employees em, departments dp
# where em.dep_id = dp.dept_id_dep;
# ```
| module4/lab-practice-subqueries-multiple-tables/lab-practice-subquieries-multiple-tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow.keras.layers import Dense, Flatten, Conv2D, Input
from tensorflow.keras import Model, mixed_precision
import os
os.chdir('..')
# +
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# Add a channels dimension
x_train = x_train[..., tf.newaxis].astype("float32")
x_test = x_test[..., tf.newaxis].astype("float32")
# +
train_ds = tf.data.Dataset.from_tensor_slices((x_train, y_train)).shuffle(10000).batch(32)
test_ds = tf.data.Dataset.from_tensor_slices((x_test, y_test)).batch(32)
# +
from sam import sam_train_step
class MyModel(Model):
def __init__(self):
super(MyModel, self).__init__()
self.conv1 = Conv2D(32, 3, activation='relu')
self.flatten = Flatten()
self.d1 = Dense(128, activation='relu')
self.d2 = Dense(10, dtype='float32')
def call(self, x):
x = self.conv1(x)
x = self.flatten(x)
x = self.d1(x)
return self.d2(x)
def train_step(self, data):
return sam_train_step(self, data)
# Create an instance of the model
model = MyModel()
# -
model.compile(optimizer=tf.keras.optimizers.RMSprop(learning_rate=0.001, rho=0.9, momentum=0.0, epsilon=1e-07, centered=False),
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(train_ds, epochs=5,
validation_data=test_ds)
| examples/mnist_example_keras_fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
from torch import nn
from torchvision import models
from copy import deepcopy
import os
from nn_extrapolation import AcceleratedSGD
from nn_utils import *
# -
trainer = Trainer(
device="cuda:3",
loss_fn=nn.NLLLoss(reduction="mean"),
val_loss_fn=nn.NLLLoss(reduction="sum"),
)
dl = load_dataset(
dataset="CIFAR10",
root=os.path.join("/tmp", os.environ["USER"], "CIFAR"),
augmentation=transforms.RandomAffine(10, scale=(0.9, 1.1), translate=(0.2, 0.2)),
validation_split=0.2,
batch_size=128,
num_workers=10,
)
# +
model = models.vgg16(pretrained=False)
model.classifier[6] = nn.Linear(4096, 10)
model.classifier.add_module("7", nn.LogSoftmax(-1))
model.to(trainer.device)
# -
trainer.validation(model, dl["valid"])
# ## Momentum
optimizer = torch.optim.Adam(model.parameters(), 1e-4, weight_decay=1e-5)
logger = Logger("vgg_log_augmentation_adam.txt.no_resizing", overwrite=True)
# +
epochs = 70
for epoch in range(epochs):
train_loss = trainer.train_epoch(model, optimizer, dl["train"])
val_acc, val_loss = trainer.validation(model, dl["valid"])
logger.log("Epoch", epoch+1, "|",
f"Training loss: {train_loss:.4f}, validation accuracy: {val_acc:.4f}, validation loss: {val_loss:.4f}")
# -
train_score = trainer.validation(model, dl["train"])
valid_score = trainer.validation(model, dl["valid"])
logger.log("Train:", train_score)
logger.log("Valid:", valid_score)
| notebooks/nn_large/vgg-augmentation-adam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Sensitivity Analysis
# ## <NAME>
# ## Objective
# This notebook is dedicated to understand how social distancing will effect the spread of an epidemic.
# ## Setup
using DifferentialEquations, Plots, Plots.PlotMeasures
# ## Utility functions
# +
# change plot fonts
bplot(p, s) = plot!(p, titlefont=font(s, "Times"), xguidefontsize=s-4
, ytickfont=font(s-4, "Times"), margin=8mm, yguidefontsize=s-4
, xtickfont = font(s-4, "Times"), legendfont=font(s-8, "Times"))
# this function will generate a T period of social distancing at time t0
sinput(t, ts) = if t-ts<0 0 elseif t==ts 0.5 else 1 end
# -
# # $SIR$ Model
#
# ## Basics
# +
# model description
function SIR(dx,x,p,t)
b, db, g, ts = p
u = b - db.*sinput(t, ts)
dx[1] = -u*x[1]*x[2]
dx[2] = u*x[1]*x[2] - g*x[2]
dx[3] = g*x[2]
end
beta = 0.2
bdiff = 0.1
gamma = 0.05
# define initial values
u0 = [1-1e-3, 1e-3, 0.]
p = [beta, bdiff, gamma, 0.]
# ode solver and time span
alg = BS3()
tspan = (0.0, 5e2)
# resolution of numerical simulation
tsrng = 0:0.1:5e2
tsval = zeros(3,size(tsrng,1))
infpk = zeros(3,size(tsrng,1))
# show something
for i = 1:size(tsrng,1)
p[4] = tsrng[i]
prob = ODEProblem(SIR, u0, tspan, p)
sol = solve(prob, alg, reltol=1e-8, abstol=1e-8, saveat=0.01)
tsval[:,i] = sol(tsrng[i])
index = findmax(sol[2,:])
infpk[:,i] = sol[:,Int(index[2])]
end
# show something
p1 = plot(tsrng, tsval[1,:], linewidth=3, label="Ss")
p1 = plot!(p1,tsrng, tsval[2,:], linewidth=3, label="Is")
p1 = plot!(p1,tsrng, tsval[3,:], linewidth=3, label="Rs", title="Start Time",
xlabel="Start Time", ylabel="Normalized Population",legend=:topleft)
p1 = bplot(p1,20)
p2 = plot(tsrng, infpk[1,:], linewidth=3, label="Sp")
p2 = plot!(p2,tsrng, infpk[2,:], linewidth=3, label="Ip")
p2 = plot!(p2,tsrng, infpk[3,:], linewidth=3, label="Rp", title="Infected Peak",
xlabel="Start Time", ylabel="Normalized Population",legend=:topleft)
p2 = bplot(p2,20)
fig = plot(p1, p2, layout=(1,2), size=(960,470))
savefig(fig, "figures/sensitivity_peak.pdf")
fig
# -
# ## Differential Sensitivity
inx = findmax(tsval[2,:])
its = tsval[2,1:Int(inx[2])]
ipk = infpk[2,1:Int(inx[2])]
tsp = tsrng[1:Int(inx[2])]
p = plot(its, ipk, label=false, linewidth=4,
xlabel="Is", ylabel="Ip", title="Infection Population at Peak and Start Time")
bplot(p, 18)
p = plot(its[1:end-1], its[1:end-1]./ipk[1:end-1].*diff(ipk)./diff(its), label="Numerical",
linewidth=4, xlabel="I(ts)", ylabel="Sensitivity", title="Differential Sensitivity")
p = plot!(p, its, 0.5.*its./ipk./(1 .- its), label="Approximation",legend=:topleft, linewidth=4)
p = bplot(p, 20)
savefig(p, "figures/sensitivity.pdf")
p
# # Provenance
using Dates
Dates.format(now(), "Y/U/d HH:MM")
| notebooks/Sensitivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing the Parker Solar Probe flybys
#
# ## 1. Modulus of the exit velocity, some features of Orbit #2
#
# First, using the data available in the reports, we try to compute some of the properties of orbit #2. This is not enough to completely define the trajectory, but will give us information later on in the process.
from astropy import units as u
T_ref = 150 * u.day
T_ref
from poliastro.bodies import Earth, Sun, Venus
k = Sun.k
k
import numpy as np
# $$ T = 2 \pi \sqrt{\frac{a^3}{\mu}} \Rightarrow a = \sqrt[3]{\frac{\mu T^2}{4 \pi^2}}$$
a_ref = np.cbrt(k * T_ref**2 / (4 * np.pi**2)).to(u.km)
a_ref.to(u.au)
# $$ \varepsilon = -\frac{\mu}{r} + \frac{v^2}{2} = -\frac{\mu}{2a} \Rightarrow v = +\sqrt{\frac{2\mu}{r} - \frac{\mu}{a}}$$
energy_ref = (-k / (2 * a_ref)).to(u.J / u.kg)
energy_ref
# +
from poliastro.twobody import Orbit
from poliastro.util import norm
from astropy.time import Time
# -
flyby_1_time = Time("2018-09-28", scale="tdb")
flyby_1_time
r_mag_ref = norm(Orbit.from_body_ephem(Venus, epoch=flyby_1_time).r)
r_mag_ref.to(u.au)
v_mag_ref = np.sqrt(2 * k / r_mag_ref - k / a_ref)
v_mag_ref.to(u.km / u.s)
# ---
# ## 2. Lambert arc between #0 and #1
#
# To compute the arrival velocity to Venus at flyby #1, we have the necessary data to solve the boundary value problem.
d_launch = Time("2018-08-11", scale="tdb")
d_launch
ss0 = Orbit.from_body_ephem(Earth, d_launch)
ss1 = Orbit.from_body_ephem(Venus, epoch=flyby_1_time)
tof = flyby_1_time - d_launch
from poliastro import iod
(v0, v1_pre), = iod.lambert(Sun.k, ss0.r, ss1.r, tof.to(u.s))
v0
v1_pre
norm(v1_pre)
# ---
# ## 3. Flyby #1 around Venus
#
# We compute a flyby using poliastro with the default value of the entry angle, just to discover that the results do not match what we expected.
from poliastro.threebody.flybys import compute_flyby
V = Orbit.from_body_ephem(Venus, epoch=flyby_1_time).v
V
h = 2548 * u.km
d_flyby_1 = Venus.R + h
d_flyby_1.to(u.km)
V_2_v_, delta_ = compute_flyby(v1_pre, V, Venus.k, d_flyby_1)
norm(V_2_v_)
# ## 4. Optimization
#
# Now we will try to find the value of $\theta$ that satisfies our requirements.
def func(theta):
V_2_v, _ = compute_flyby(v1_pre, V, Venus.k, d_flyby_1, theta * u.rad)
ss_1 = Orbit.from_vectors(Sun, ss1.r, V_2_v, epoch=flyby_1_time)
return (ss_1.period - T_ref).to(u.day).value
# There are two solutions:
import matplotlib.pyplot as plt
theta_range = np.linspace(0, 2 * np.pi)
plt.plot(theta_range, [func(theta) for theta in theta_range])
plt.axhline(0, color='k', linestyle="dashed")
func(0)
func(1)
from scipy.optimize import brentq
theta_opt_a = brentq(func, 0, 1) * u.rad
theta_opt_a.to(u.deg)
theta_opt_b = brentq(func, 4, 5) * u.rad
theta_opt_b.to(u.deg)
V_2_v_a, delta_a = compute_flyby(v1_pre, V, Venus.k, d_flyby_1, theta_opt_a)
V_2_v_b, delta_b = compute_flyby(v1_pre, V, Venus.k, d_flyby_1, theta_opt_b)
norm(V_2_v_a)
norm(V_2_v_b)
# ## 5. Exit orbit
#
# And finally, we compute orbit #2 and check that the period is the expected one.
ss01 = Orbit.from_vectors(Sun, ss1.r, v1_pre, epoch=flyby_1_time)
ss01
# The two solutions have different inclinations, so we still have to find out which is the good one. We can do this by computing the inclination over the ecliptic - however, as the original data was in the International Celestial Reference Frame (ICRF), whose fundamental plane is parallel to the Earth equator of a reference epoch, we have change the plane to the Earth **ecliptic**, which is what the original reports use.
ss_1_a = Orbit.from_vectors(Sun, ss1.r, V_2_v_a, epoch=flyby_1_time)
ss_1_a
ss_1_b = Orbit.from_vectors(Sun, ss1.r, V_2_v_b, epoch=flyby_1_time)
ss_1_b
# Let's define a function to do that quickly for us, using the [`get_frame`](https://docs.poliastro.space/en/latest/safe.html#poliastro.frames.get_frame) function from poliastro.frames:
# +
from astropy.coordinates import CartesianRepresentation
from poliastro.frames import Planes, get_frame
def change_plane(ss_orig, plane):
"""Changes the plane of the Orbit.
"""
ss_orig_rv = ss_orig.frame.realize_frame(
ss_orig.represent_as(CartesianRepresentation)
)
dest_frame = get_frame(ss_orig.attractor, plane, obstime=ss_orig.epoch)
ss_dest_rv = ss_orig_rv.transform_to(dest_frame)
ss_dest_rv.representation_type = CartesianRepresentation
ss_dest = Orbit.from_vectors(
ss_orig.attractor,
r=ss_dest_rv.data.xyz,
v=ss_dest_rv.data.differentials['s'].d_xyz,
epoch=ss_orig.epoch,
plane=plane,
)
return ss_dest
# -
change_plane(ss_1_a, Planes.EARTH_ECLIPTIC)
change_plane(ss_1_b, Planes.EARTH_ECLIPTIC)
# Therefore, **the correct option is the first one**.
ss_1_a.period.to(u.day)
ss_1_a.a
# And, finally, we plot the solution:
from poliastro.plotting import OrbitPlotter
# +
frame = OrbitPlotter()
frame.plot(ss0, label=Earth)
frame.plot(ss1, label=Venus)
frame.plot(ss01, label="#0 to #1")
frame.plot(ss_1_a, label="#1 to #2");
| docs/source/examples/Analyzing the Parker Solar Probe flybys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''datasets'': conda)'
# language: python
# name: python37764bitdatasetscondae5d8ff60608e4c5c953d6bb643d8ebc5
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/huggingface/datasets/blob/master/notebooks/Overview.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zNp6kK7OvSUg" colab_type="text"
# # HuggingFace `🤗Datasets` library - Quick overview
#
# Models come and go (linear models, LSTM, Transformers, ...) but two core elements have consistently been the beating heart of Natural Language Processing: Datasets & Metrics
#
# `🤗Datasets` is a fast and efficient library to easily share and load dataset and evaluation metrics, already providing access to 150+ datasets and 12+ evaluation metrics.
#
# The library has several interesting features (beside easy access to datasets/metrics):
#
# - Build-in interoperability with PyTorch, Tensorflow 2, Pandas and Numpy
# - Lighweight and fast library with a transparent and pythonic API
# - Strive on large datasets: frees you from RAM memory limits, all datasets are memory-mapped on drive by default.
# - Smart caching with an intelligent `tf.data`-like cache: never wait for your data to process several times
#
# `🤗Datasets` originated from a fork of the awesome Tensorflow-Datasets and the HuggingFace team want to deeply thank the team behind this amazing library and user API. We have tried to keep a layer of compatibility with `tfds` and a conversion can provide conversion from one format to the other.
# + [markdown] id="dzk9aEtIvSUh" colab_type="text"
# # Main datasets API
#
# This notebook is a quick dive in the main user API for loading datasets in `datasets`
# + id="my95uHbLyjwR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="ef568bc2-33a8-4aa4-bf84-9a253df48b8d"
# install datasets
# !pip install datasets
# Make sure that we have a recent version of pyarrow in the session before we continue - otherwise reboot Colab to activate it
import pyarrow
if int(pyarrow.__version__.split('.')[1]) < 16 and int(pyarrow.__version__.split('.')[0]) == 0:
import os
os.kill(os.getpid(), 9)
# + id="PVjXLiYxvSUl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="88755377-1c1e-4ab5-83e2-b912f6d90da2"
# Let's import the library. We typically only need at most four methods:
from datasets import list_datasets, list_metrics, load_dataset, load_metric
from pprint import pprint
# + [markdown] id="TNloBBx-vSUo" colab_type="text"
# ## Listing the currently available datasets and metrics
# + id="d3RJisGLvSUp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 561} outputId="63017ebc-61a4-4e5f-bd52-09960ad03095"
# Currently available datasets and metrics
datasets = list_datasets()
metrics = list_metrics()
print(f"🤩 Currently {len(datasets)} datasets are available on the hub:")
pprint(datasets, compact=True)
print(f"🤩 Currently {len(metrics)} metrics are available on the hub:")
pprint(metrics, compact=True)
# + id="7T5AG3BxvSUr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="35665e60-2959-41b8-d511-9da3f64178b1"
# You can access various attributes of the datasets before downloading them
squad_dataset = list_datasets(with_details=True)[datasets.index('squad')]
pprint(squad_dataset.__dict__) # It's a simple python dataclass
# + [markdown] id="9uqSkkSovSUt" colab_type="text"
# ## An example with SQuAD
# + id="aOXl6afcvSUu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 241} outputId="4eda989b-73ab-4d37-acae-5cd7e1b6de42"
# Downloading and loading a dataset
dataset = load_dataset('squad', split='validation[:10%]')
# + [markdown] id="rQ0G-eK3vSUw" colab_type="text"
# This call to `datasets.load_dataset()` does the following steps under the hood:
#
# 1. Download and import in the library the **SQuAD python processing script** from HuggingFace AWS bucket if it's not already stored in the library. You can find the SQuAD processing script [here](https://github.com/huggingface/datasets/tree/master/datasets/squad/squad.py) for instance.
#
# Processing scripts are small python scripts which define the info (citation, description) and format of the dataset and contain the URL to the original SQuAD JSON files and the code to load examples from the original SQuAD JSON files.
#
#
# 2. Run the SQuAD python processing script which will:
# - **Download the SQuAD dataset** from the original URL (see the script) if it's not already downloaded and cached.
# - **Process and cache** all SQuAD in a structured Arrow table for each standard splits stored on the drive.
#
# Arrow table are arbitrarily long tables, typed with types that can be mapped to numpy/pandas/python standard types and can store nested objects. They can be directly access from drive, loaded in RAM or even streamed over the web.
#
#
# 3. Return a **dataset build from the splits** asked by the user (default: all), in the above example we create a dataset with the first 10% of the validation split.
# + id="fercoFwLvSUx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 734} outputId="b5fbf431-b40d-47b1-a609-58f308d81033"
# Informations on the dataset (description, citation, size, splits, format...)
# are provided in `dataset.info` (a simple python dataclass) and also as direct attributes in the dataset object
pprint(dataset.info.__dict__)
# + [markdown] id="GE0E87zsvSUz" colab_type="text"
# ## Inspecting and using the dataset: elements, slices and columns
# + [markdown] id="DKf4YFnevSU0" colab_type="text"
# The returned `Dataset` object is a memory mapped dataset that behave similarly to a normal map-style dataset. It is backed by an Apache Arrow table which allows many interesting features.
# + id="tP1xPqSyvSU0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="1f921b91-3deb-4dbd-b19e-dbddfd29e095"
print(dataset)
# + [markdown] id="aiO3rC8yvSU2" colab_type="text"
# You can query it's length and get items or slices like you would do normally with a python mapping.
# + id="xxLcdj2yvSU3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="5eb1eb3b-0f77-4935-c5f7-5f34747d0af7"
print(f"👉Dataset len(dataset): {len(dataset)}")
print("\n👉First item 'dataset[0]':")
pprint(dataset[0])
# + id="zk1WQ_cczP5w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="3fe3d8c7-c97f-4c71-a15a-42825f86dfa4"
# Or get slices with several examples:
print("\n👉Slice of the two items 'dataset[10:12]':")
pprint(dataset[10:12])
# + id="QXj2Qr5KvSU5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="9232ac73-1fe5-4bd2-de60-5716ef8f4201"
# You can get a full column of the dataset by indexing with its name as a string:
print(dataset['question'][:10])
# + [markdown] id="6Au7rqPMvSU7" colab_type="text"
# The `__getitem__` method will return different format depending on the type of query:
#
# - Items like `dataset[0]` are returned as dict of elements.
# - Slices like `dataset[10:20]` are returned as dict of lists of elements.
# - Columns like `dataset['question']` are returned as a list of elements.
#
# This may seems surprising at first but in our experiments it's actually a lot easier to use for data processing than returning the same format for each of these views on the dataset.
# + [markdown] id="6DB_y79cvSU8" colab_type="text"
# In particular, you can easily iterate along columns in slices, and also naturally permute consecutive indexings with identical results as showed here by permuting column indexing with elements and slices:
# + id="wjGocqArvSU9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="22389a1d-2e3a-4d32-e125-8aed384df07d"
print(dataset[0]['question'] == dataset['question'][0])
print(dataset[10:20]['context'] == dataset['context'][10:20])
# + [markdown] id="b1-Kj1xQvSU_" colab_type="text"
# ### Dataset are internally typed and structured
#
# The dataset is backed by one (or several) Apache Arrow tables which are typed and allows for fast retrieval and access as well as arbitrary-size memory mapping.
#
# This means respectively that the format for the dataset is clearly defined and that you can load datasets of arbitrary size without worrying about RAM memory limitation (basically the dataset take no space in RAM, it's directly read from drive when needed with fast IO access).
# + id="rAnp_RyPvSVA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="1ee0ace7-db77-4a9a-f6e4-c84cdb3f0bba"
# You can inspect the dataset column names and types
print("Column names:")
pprint(dataset.column_names)
print("Features:")
pprint(dataset.features)
# + [markdown] id="au4v3mOQvSVC" colab_type="text"
# ### Additional misc properties
# + id="efFhDWhlvSVC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="f5bd2739-e52f-4e50-b3fe-afbd3f1f2427"
# Datasets also have shapes informations
print("The number of rows", dataset.num_rows, "also available as len(dataset)", len(dataset))
print("The number of columns", dataset.num_columns)
print("The shape (rows, columns)", dataset.shape)
# + [markdown] id="1Ox7ppKDvSVN" colab_type="text"
# ## Modifying the dataset with `dataset.map`
#
# Now that we know how to inspect our dataset we also want to update it. For that there is a powerful method `.map()` which is inspired by `tf.data` map method and that you can use to apply a function to each examples, independently or in batch.
#
# `.map()` takes a callable accepting a dict as argument (same dict as the one returned by `dataset[i]`) and iterate over the dataset by calling the function on each example.
# + id="Yz2-27HevSVN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154, "referenced_widgets": ["08236188125a4c2e931feb58ebe648c0", "58f37f73168648a08edc0ae615260c24", "1da82f72f3fc46358fe3e4268e42d137", "b051786ab97145cbb88627588bbec7d2", "97948303212c4a1982a6dda9dfd4cc90", "f252b203b8d349edb87b0a81209746b2", "37b19294c7464eb4bd886d966e148219", "1aa28417f911424eb9ac87413e4572ee"]} outputId="183eafc7-fc4f-41ee-c342-7ff13ce09706"
# Let's print the length of each `context` string in our subset of the dataset
# (10% of the validation i.e. 1057 examples)
dataset.map(lambda example: print(len(example['context']), end=','))
# + [markdown] id="Ta3celHnvSVP" colab_type="text"
# This is basically the same as doing
#
# ```python
# for example in dataset:
# function(example)
# ```
# + [markdown] id="Z4Fjr0DJawuS" colab_type="text"
# The above examples was a bit verbose. We can control the logging level of `🤗Datasets` with it's logging module:
#
# + id="qAgptXFYaquI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="f4c780cf-3a15-4043-9c51-2af530bcc0c2"
from datasets import logging
logging.set_verbosity_warning()
dataset.map(lambda example: print(len(example['context']), end=','))
# + id="KfED6CEHa8J_" colab_type="code" colab={}
# Let's keep it verbose for our tutorial though
from datasets import logging
logging.set_verbosity_info()
# + [markdown] id="i_Ouw5gDvSVP" colab_type="text"
# The above example had no effect on the dataset because the method we supplied to `.map()` didn't return a `dict` or a `abc.Mapping` that could be used to update the examples in the dataset.
#
# In such a case, `.map()` will return the same dataset (`self`).
#
# Now let's see how we can use a method that actually modify the dataset.
# + [markdown] id="cEnCi9DFvSVQ" colab_type="text"
# ### Modifying the dataset example by example
# + [markdown] id="kA37VgZhvSVQ" colab_type="text"
# The main interest of `.map()` is to update and modify the content of the table and leverage smart caching and fast backend.
#
# To use `.map()` to update elements in the table you need to provide a function with the following signature: `function(example: dict) -> dict`.
# + id="vUr65K-4vSVQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="91d316b1-9fcc-44a7-b2f1-4af3b0c5c530"
# Let's add a prefix 'My cute title: ' to each of our titles
def add_prefix_to_title(example):
example['title'] = 'My cute title: ' + example['title']
return example
prefixed_dataset = dataset.map(add_prefix_to_title)
print(prefixed_dataset.unique('title')) # `.unique()` is a super fast way to print the unique elemnts in a column (see the doc for all the methods)
# + [markdown] id="FcZ_amDAvSVS" colab_type="text"
# This call to `.map()` compute and return the updated table. It will also store the updated table in a cache file indexed by the current state and the mapped function.
#
# A subsequent call to `.map()` (even in another python session) will reuse the cached file instead of recomputing the operation.
#
# You can test this by running again the previous cell, you will see that the result are directly loaded from the cache and not re-computed again.
#
# The updated dataset returned by `.map()` is (again) directly memory mapped from drive and not allocated in RAM.
# + [markdown] id="Skbf8LUEvSVT" colab_type="text"
# The function you provide to `.map()` should accept an input with the format of an item of the dataset: `function(dataset[0])` and return a python dict.
#
# The columns and type of the outputs can be different than the input dict. In this case the new keys will be added as additional columns in the dataset.
#
# Bascially each dataset example dict is updated with the dictionary returned by the function like this: `example.update(function(example))`.
# + id="d5De0CfTvSVT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="7ae32097-2117-4ebe-cd93-19eaad139579"
# Since the input example dict is updated with our function output dict,
# we can actually just return the updated 'title' field
titled_dataset = dataset.map(lambda example: {'title': 'My cutest title: ' + example['title']})
print(titled_dataset.unique('title'))
# + [markdown] id="Q5vny56-vSVV" colab_type="text"
# #### Removing columns
# You can also remove columns when running map with the `remove_columns=List[str]` argument.
# + id="-sPWnsz-vSVW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="3cac70f9-394f-4ef7-ffe3-e0d37d627d23"
# This will remove the 'title' column while doing the update (after having send it the the mapped function so you can use it in your function!)
less_columns_dataset = dataset.map(lambda example: {'new_title': 'Wouhahh: ' + example['title']}, remove_columns=['title'])
print(less_columns_dataset.column_names)
print(less_columns_dataset.unique('new_title'))
# + [markdown] id="G459HzD-vSVY" colab_type="text"
# #### Using examples indices
# With `with_indices=True`, dataset indices (from `0` to `len(dataset)`) will be supplied to the function which must thus have the following signature: `function(example: dict, indice: int) -> dict`
# + id="_kFL37R2vSVY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="d4f83cac-6e96-48c3-ba6b-6ffdd900712f"
# This will add the index in the dataset to the 'question' field
with_indices_dataset = dataset.map(lambda example, idx: {'question': f'{idx}: ' + example['question']},
with_indices=True)
pprint(with_indices_dataset['question'][:5])
# + [markdown] id="xckhVEWFvSVb" colab_type="text"
# ### Modifying the dataset with batched updates
# + [markdown] id="dzmicbSnvSVb" colab_type="text"
# `.map()` can also work with batch of examples (slices of the dataset).
#
# This is particularly interesting if you have a function that can handle batch of inputs like the tokenizers of HuggingFace `tokenizers`.
#
# To work on batched inputs set `batched=True` when calling `.map()` and supply a function with the following signature: `function(examples: Dict[List]) -> Dict[List]` or, if you use indices, `function(examples: Dict[List], indices: List[int]) -> Dict[List]`).
#
# Bascially, your function should accept an input with the format of a slice of the dataset: `function(dataset[:10])`.
# + id="pxHbgSTL0itj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="441ba953-8e69-4f74-d054-bb07804efecb"
# !pip install transformers
# + id="T7gpEg0yvSVc" colab_type="code" colab={}
# Let's import a fast tokenizer that can work on batched inputs
# (the 'Fast' tokenizers in HuggingFace)
from transformers import BertTokenizerFast, logging as transformers_logging
transformers_logging.set_verbosity_warning()
tokenizer = BertTokenizerFast.from_pretrained('bert-base-cased')
# + id="fAmLTPC9vSVe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 972} outputId="0182e1b5-aa19-4739-b6ba-dc60e151998e"
# Now let's batch tokenize our dataset 'context'
encoded_dataset = dataset.map(lambda example: tokenizer(example['context']), batched=True)
print("encoded_dataset[0]")
pprint(encoded_dataset[0], compact=True)
# + id="kNaJdKskvSVf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b7899e53-1bd9-4ea2-bf8d-60c127b3c6e2"
# we have added additional columns
pprint(dataset.column_names)
# + id="m3To8ztMvSVj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 154, "referenced_widgets": ["ae210ea515f94c6ab34c1113e823b92d", "86c8850654d54b47a2771cbb1804f5a1", "435d36683bc04e06a971b76129a88ed1", "78ae08f5803f4cb29f7f63c2843e9db0", "703443b26d7d40aea83417de59b64a79", "cf940cf9ea3043b5abeb4c851ad23b77", "e319f183228b4f0dba1140d9cc1573ec", "a3d89c58eda640a6a8289ba0c5d549e2"]} outputId="69eca4fb-d5ae-46c1-e586-29d75830f174"
# Let show a more complex processing with the full preparation of the SQuAD dataset
# for training a model from Transformers
def convert_to_features(batch):
# Tokenize contexts and questions (as pairs of inputs)
input_pairs = list(zip())
encodings = tokenizer(batch['context'], batch['question'], truncation=True)
# Compute start and end tokens for labels
start_positions, end_positions = [], []
for i, answer in enumerate(batch['answers']):
first_char = answer['answer_start'][0]
last_char = first_char + len(answer['text'][0]) - 1
start_positions.append(encodings.char_to_token(i, first_char))
end_positions.append(encodings.char_to_token(i, last_char))
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
return encodings
encoded_dataset = dataset.map(convert_to_features, batched=True)
# + id="KBnmSa46vSVl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="efca25d9-581e-46a2-f9e7-53aaf018ea7f"
# Now our dataset comprise the labels for the start and end position
# as well as the offsets for converting back tokens
# in span of the original string for evaluation
print("column_names", encoded_dataset.column_names)
print("start_positions", encoded_dataset[:5]['start_positions'])
# + [markdown] id="NzOXxNzQvSVo" colab_type="text"
# ## formatting outputs for PyTorch, Tensorflow, Numpy, Pandas
#
# Now that we have tokenized our inputs, we probably want to use this dataset in a `torch.Dataloader` or a `tf.data.Dataset`.
#
# To be able to do this we need to tweak two things:
#
# - format the indexing (`__getitem__`) to return numpy/pytorch/tensorflow tensors, instead of python objects, and probably
# - format the indexing (`__getitem__`) to return only the subset of the columns that we need for our model inputs.
#
# We don't want the columns `id` or `title` as inputs to train our model, but we could still want to keep them in the dataset, for instance for the evaluation of the model.
#
# This is handled by the `.set_format(type: Union[None, str], columns: Union[None, str, List[str]])` where:
#
# - `type` define the return type for our dataset `__getitem__` method and is one of `[None, 'numpy', 'pandas', 'torch', 'tensorflow']` (`None` means return python objects), and
# - `columns` define the columns returned by `__getitem__` and takes the name of a column in the dataset or a list of columns to return (`None` means return all columns).
# + id="aU2h_qQDvSVo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 666} outputId="15f14cbb-558b-4701-b6f8-0999d3570920"
columns_to_return = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
encoded_dataset.set_format(type='torch', columns=columns_to_return)
# Our dataset indexing output is now ready for being used in a pytorch dataloader
pprint(encoded_dataset[1], compact=True)
# + id="Wj1ukGIuvSVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aa7f4643-54c2-45ef-b33e-4881bc68a57d"
# Note that the columns are not removed from the dataset, just not returned when calling __getitem__
# Similarly the inner type of the dataset is not changed to torch.Tensor, the conversion and filtering is done on-the-fly when querying the dataset
print(encoded_dataset.column_names)
# + id="pWmmUdatasetsvSVs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 969} outputId="6e12d88f-8d39-414d-e340-fbecd65275b4"
# We can remove the formatting with `.reset_format()`
# or, identically, a call to `.set_format()` with no arguments
encoded_dataset.reset_format()
pprint(encoded_dataset[1], compact=True)
# + id="VyUOA07svSVu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="d3c2a6c9-a1ec-41c2-d438-66b68c5f7416"
# The current format can be checked with `.format`,
# which is a dict of the type and formatting
pprint(encoded_dataset.format)
# + [markdown] id="xyi2eMeSvSVv" colab_type="text"
# # Wrapping this all up (PyTorch)
#
# Let's wrap this all up with the full code to load and prepare SQuAD for training a PyTorch model from HuggingFace `transformers` library.
#
#
# + id="l0j8BPLi6Qlv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="de8ed5f7-069b-4c99-8b62-8fa32d755192"
# !pip install transformers
# + id="QvExTIZWvSVw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 526, "referenced_widgets": ["71d89e94d82f4335b7ca7aaf4dba83ed", "f38913964ddb4e3eb6fc0cf087ab0f52", "4f03c8400d014b3bbb5e8dd4f63c5441", "23f45949b7f949859ecabab44b591553", "<KEY>", "<KEY>", "d46f8f656ea34972a4dd9b7554d62315", "<KEY>", "<KEY>", "30e2b72ce2304332a202d92d606671f6", "810d81cd651d41f298ff4815fcf9f34a", "86fa8d2a2b204d8586cc8ab5ad2a1b7a", "6db8399610464d6e93ce82ad8bb7bfc4", "f684dcff9d4c4c10b6bb623c02465b14", "043e8d6ca6574bd4abcfdd7145162ceb", "ec53215a263e413eab01f74b66b9bff6"]} outputId="cb34dcd4-9ddc-4195-9b9d-5442659f9798"
import torch
from datasets import load_dataset
from transformers import BertTokenizerFast
# Load our training dataset and tokenizer
dataset = load_dataset('squad')
tokenizer = BertTokenizerFast.from_pretrained('bert-base-cased')
def get_correct_alignement(context, answer):
""" Some original examples in SQuAD have indices wrong by 1 or 2 character. We test and fix this here. """
gold_text = answer['text'][0]
start_idx = answer['answer_start'][0]
end_idx = start_idx + len(gold_text)
if context[start_idx:end_idx] == gold_text:
return start_idx, end_idx # When the gold label position is good
elif context[start_idx-1:end_idx-1] == gold_text:
return start_idx-1, end_idx-1 # When the gold label is off by one character
elif context[start_idx-2:end_idx-2] == gold_text:
return start_idx-2, end_idx-2 # When the gold label is off by two character
else:
raise ValueError()
# Tokenize our training dataset
def convert_to_features(example_batch):
# Tokenize contexts and questions (as pairs of inputs)
encodings = tokenizer(example_batch['context'], example_batch['question'], truncation=True)
# Compute start and end tokens for labels using Transformers's fast tokenizers alignement methods.
start_positions, end_positions = [], []
for i, (context, answer) in enumerate(zip(example_batch['context'], example_batch['answers'])):
start_idx, end_idx = get_correct_alignement(context, answer)
start_positions.append(encodings.char_to_token(i, start_idx))
end_positions.append(encodings.char_to_token(i, end_idx-1))
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
return encodings
encoded_dataset = dataset.map(convert_to_features, batched=True)
# Format our dataset to outputs torch.Tensor to train a pytorch model
columns = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
encoded_dataset.set_format(type='torch', columns=columns)
# Instantiate a PyTorch Dataloader around our dataset
# Let's do dynamic batching (pad on the fly with our own collate_fn)
def collate_fn(examples):
return tokenizer.pad(examples, return_tensors='pt')
dataloader = torch.utils.data.DataLoader(encoded_dataset['train'], collate_fn=collate_fn, batch_size=8)
# + id="4mHnwMx2vSVx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="1c0942f3-68df-490b-d37a-8f63d2aae717"
# Let's load a pretrained Bert model and a simple optimizer
from transformers import BertForQuestionAnswering
model = BertForQuestionAnswering.from_pretrained('distilbert-base-cased', return_dict=True)
optimizer = torch.optim.Adam(model.parameters(), lr=1e-5)
# + id="biqDH9vpvSVz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 426} outputId="1a55ede4-bde6-4bd7-e945-7e4b54352f68"
# Now let's train our model
device = 'cuda' if torch.cuda.is_available() else 'cpu'
model.train().to(device)
for i, batch in enumerate(dataloader):
batch.to(device)
outputs = model(**batch)
loss = outputs.loss
loss.backward()
optimizer.step()
model.zero_grad()
print(f'Step {i} - loss: {loss:.3}')
if i > 5:
break
# + [markdown] id="kxZQ9Ms_vSV1" colab_type="text"
# # Wrapping this all up (Tensorflow)
#
# Let's wrap this all up with the full code to load and prepare SQuAD for training a Tensorflow model (works only from the version 2.2.0)
# + id="ZE8VSTYovSV2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4f3c33f0-deb1-48d4-d778-c3d80172b22f"
import tensorflow as tf
import datasets
from transformers import BertTokenizerFast
# Load our training dataset and tokenizer
train_tf_dataset = datasets.load_dataset('squad', split="train")
tokenizer = BertTokenizerFast.from_pretrained('bert-base-cased', return_dict=True)
# Tokenize our training dataset
# The only one diff here is that start_positions and end_positions
# must be single dim list => [[23], [45] ...]
# instead of => [23, 45 ...]
def convert_to_tf_features(example_batch):
# Tokenize contexts and questions (as pairs of inputs)
encodings = tokenizer(example_batch['context'], example_batch['question'], truncation=True)
# Compute start and end tokens for labels using Transformers's fast tokenizers alignement methods.
start_positions, end_positions = [], []
for i, (context, answer) in enumerate(zip(example_batch['context'], example_batch['answers'])):
start_idx, end_idx = get_correct_alignement(context, answer)
start_positions.append([encodings.char_to_token(i, start_idx)])
end_positions.append([encodings.char_to_token(i, end_idx-1)])
if start_positions and end_positions:
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
return encodings
train_tf_dataset = train_tf_dataset.map(convert_to_tf_features, batched=True)
def remove_none_values(example):
return not None in example["start_positions"] or not None in example["end_positions"]
train_tf_dataset = train_tf_dataset.filter(remove_none_values, load_from_cache_file=False)
columns = ['input_ids', 'token_type_ids', 'attention_mask', 'start_positions', 'end_positions']
train_tf_dataset.set_format(type='tensorflow', columns=columns)
features = {x: train_tf_dataset[x].to_tensor(default_value=0, shape=[None, tokenizer.model_max_length]) for x in columns[:3]}
labels = {"output_1": train_tf_dataset["start_positions"].to_tensor(default_value=0, shape=[None, 1])}
labels["output_2"] = train_tf_dataset["end_positions"].to_tensor(default_value=0, shape=[None, 1])
tfdataset = tf.data.Dataset.from_tensor_slices((features, labels)).batch(8)
# + id="y0dfw8K8vSV4" colab_type="code" colab={}
# Let's load a pretrained TF2 Bert model and a simple optimizer
from transformers import TFBertForQuestionAnswering
model = TFBertForQuestionAnswering.from_pretrained("bert-base-cased")
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy(reduction=tf.keras.losses.Reduction.NONE, from_logits=True)
opt = tf.keras.optimizers.Adam(learning_rate=3e-5)
model.compile(optimizer=opt,
loss={'output_1': loss_fn, 'output_2': loss_fn},
loss_weights={'output_1': 1., 'output_2': 1.},
metrics=['accuracy'])
# + id="TcYtiykmvSV6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 207} outputId="4b755c1b-8e61-43ab-ffc6-d3e82ed4174d"
# Now let's train our model
model.fit(tfdataset, epochs=1, steps_per_epoch=3)
# + [markdown] id="eREDXWP6vSV8" colab_type="text"
# # Metrics API
#
# `datasets` also provides easy access and sharing of metrics.
#
# This aspect of the library is still experimental and the API may still evolve more than the datasets API.
#
# Like datasets, metrics are added as small scripts wrapping common metrics in a common API.
#
# There are several reason you may want to use metrics with `datasets` and in particular:
#
# - metrics for specific datasets like GLUE or SQuAD are provided out-of-the-box in a simple, convenient and consistant way integrated with the dataset,
# - metrics in `datasets` leverage the powerful backend to provide smart features out-of-the-box like support for distributed evaluation in PyTorch
# + [markdown] id="uUoGMMVKvSV8" colab_type="text"
# ## Using metrics
#
# Using metrics is pretty simple, they have two main methods: `.compute(predictions, references)` to directly compute the metric and `.add(prediction, reference)` or `.add_batch(predictions, references)` to only store some results if you want to do the evaluation in one go at the end.
#
# Here is a quick gist of a standard use of metrics (the simplest usage):
# ```python
# from datasets import load_metric
# sacrebleu_metric = load_metric('sacrebleu')
#
# # If you only have a single iteration, you can easily compute the score like this
# predictions = model(inputs)
# score = sacrebleu_metric.compute(predictions, references)
#
# # If you have a loop, you can "add" your predictions and references at each iteration instead of having to save them yourself (the metric object store them efficiently for you)
# for batch in dataloader:
# model_input, targets = batch
# predictions = model(model_inputs)
# sacrebleu_metric.add_batch(predictions, targets)
# score = sacrebleu_metric.compute() # Compute the score from all the stored predictions/references
# ```
#
# Here is a quick gist of a use in a distributed torch setup (should work for any python multi-process setup actually). It's pretty much identical to the second example above:
# ```python
# from datasets import load_metric
# # You need to give the total number of parallel python processes (num_process) and the id of each process (process_id)
# bleu_metric = datasets.load_metric('sacrebleu', process_id=torch.distributed.get_rank(),b num_process=torch.distributed.get_world_size())
#
# for batch in dataloader:
# model_input, targets = batch
# predictions = model(model_inputs)
# sacrebleu_metric.add_batch(predictions, targets)
# score = sacrebleu_metric.compute() # Compute the score on the first node by default (can be set to compute on each node as well)
# ```
# + [markdown] id="ySL-vDadvSV8" colab_type="text"
# Example with a NER metric: `seqeval`
# + id="f4uZym7MvSV9" colab_type="code" colab={}
ner_metric = load_metric('seqeval')
references = [['O', 'O', 'O', 'B-MISC', 'I-MISC', 'I-MISC', 'O'], ['B-PER', 'I-PER', 'O']]
predictions = [['O', 'O', 'B-MISC', 'I-MISC', 'I-MISC', 'I-MISC', 'O'], ['B-PER', 'I-PER', 'O']]
ner_metric.compute(predictions, references)
# + [markdown] id="ctY6AIAilLdH" colab_type="text"
# # Adding a new dataset or a new metric
#
# They are two ways to add new datasets and metrics in `datasets`:
#
# - datasets can be added with a Pull-Request adding a script in the `datasets` folder of the [`datasets` repository](https://github.com/huggingface/datasets)
#
# => once the PR is merged, the dataset can be instantiate by it's folder name e.g. `datasets.load_dataset('squad')`. If you want HuggingFace to host the data as well you will need to ask the HuggingFace team to upload the data.
#
# - datasets can also be added with a direct upload using `datasets` CLI as a user or organization (like for models in `transformers`). In this case the dataset will be accessible under the gien user/organization name, e.g. `datasets.load_dataset('thomwolf/squad')`. In this case you can upload the data yourself at the same time and in the same folder.
#
# We will add a full tutorial on how to add and upload datasets soon.
# + id="ypLjbtGrljk8" colab_type="code" colab={}
| notebooks/Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating simulated data from a mosaic image
# This notebook demonstrates how to use Mirage to create simulated data from a distortion-free mosaic image. In this case, we will use a mosaic of the GOODS-S region from the [CANDELS survey](https://archive.stsci.edu/prepds/candels/). Several stamp images containing additional objects will also be used. These images are cut-outs from a recent [HST/WFC3 SNAPSHOT program of some 3C sources](https://hz3c.stsci.edu/Observations.html).
#
# For each observation to be simulated, the appropriate area of the mosaic is extracted from the full mosaic, and is resampled in order to introduce the distortion associated with the JWST instrument to be used. This distorted image is then addded to the simulated data in one of two ways.
#
# If you wish to modify the image from the mosaic in any way, such as adding additional objects or scaling the brightness, then the mosaic image can be added to one of Mirage's "extended" source catalogs, along with additional sources.
#
# If you do not wish to modify the cropped mosaic image in any way (other than introducing the appropriate distortion), then the distorted image can be used directly as a seed image, and you only need to run the dark_prep and obs_generation steps of Mirage in order to create the final simulated data.
# ## Table of contents
# * [Imports](#imports)
# * [Download Data](#download)
# * [Using resampled image in an extended source catalog](#resample_into_catalog)
# * [Provide the PSF FWHM in the mosaic data](#provide_fwhm)
# * [Measure the FWHM by fitting a 2D Gaussian](#measure_fwhm)
# * [Run yaml_generator to create Mirage input yaml files](#yaml_generator_catalogs)
# * [Source Catalogs](#source_catalogs)
# * [Extract images from mosaic, resample, and add to catalog](#crop_and_blot_catalog)
# * [Create the simulated data](#create_data_catalog)
# * [Look at simulated data](#examine_data_catalog)
# * [Use resampled image as a seed image](#resample_seed)
# * [Run yaml_generator to create Mirage input yaml files](#yaml_generator_seed)
# * [Create the simulted data](#create_data_seed)
# * [Look at simulated data](#examine_data_seed)
# ---
# <a id='imports'></a>
# ## Imports
import os
import yaml
from astropy.io import fits
from astropy.modeling import models, fitting
import matplotlib.pyplot as plt
import numpy as np
from mirage.catalogs.catalog_generator import ExtendedCatalog
from mirage.catalogs.create_catalog import combine_catalogs
from mirage.dark.dark_prep import DarkPrep
from mirage.ramp_generator.obs_generator import Observation
from mirage.imaging_simulator import ImgSim
from mirage.reference_files.downloader import download_file
from mirage.seed_image.fits_seed_image import ImgSeed
from mirage.yaml import yaml_generator
# ---
# <a id='download'></a>
# ## Download Data
# Download FITS files containing the mosaic, as well as three small stamp images containing single objects.
base_url = ('https://data.science.stsci.edu/redirect/JWST/jwst-simulations/'
'mirage_reference_files/example_data_for_notebooks')
stamp_files = ['3C305.1_stamp.fits', '3C324_stamp.fits', '3C454.1_stamp.fits',
'hlsp_candels_hst_acs_gs-tot-sect23_f814w_v1.0_drz.fits']
for stamp_file in stamp_files:
stamp_url = '{}/{}'.format(base_url, stamp_file)
s_file = download_file(stamp_url, stamp_file, output_directory='./')
# ---
# <a id='resample_into_catalog'></a>
# ## Using resampled image in an extended source catalog
mosaicfile = 'hlsp_candels_hst_acs_gs-tot-sect23_f814w_v1.0_drz.fits'
xml_file = 'extended_object_test.xml'
pointing_file = xml_file.replace('.xml', '.pointing')
# ---
# <a id='provide_fwhm'></a>
# ### Provide the PSF FWHM in the mosaic data
# From the CANDELS documentation
mosaic_fwhm = 0.09 # arcseconds
# ---
# <a id='measure_fwhm'></a>
# ### Alternatively, measure the FWHM by fitting a 2D Gaussian
mosaic = fits.getdata(mosaicfile)
mosaic_header = fits.getheader(mosaicfile)
# Extract a subimage around a star
box = mosaic[3254: 3330, 7071: 7153]
yp, xp = box.shape
# Look at the extracted box. Make sure the PSF looks reasonable
plt.imshow(box)
# Generate grid of same size as box, to be used in fitting
y, x, = np.mgrid[:yp, :xp]
# Fit the model
p_init = models.Gaussian2D()
fit_p = fitting.LevMarLSQFitter()
fitted_psf = fit_p(p_init, x, y, box)
# Fit results. The FWHM is needed as an input to the
print('Amplitude: ', fitted_psf.amplitude.value)
print('X_mean: ', fitted_psf.x_mean.value)
print('Y_mean: ', fitted_psf.y_mean.value)
print('X_FWHM: ', fitted_psf.x_fwhm)
print('Y_FWHM: ', fitted_psf.y_fwhm)
print('X_stddev: ', fitted_psf.x_stddev.value)
print('Y_stddev: ', fitted_psf.y_stddev.value)
measured_mosaic_fwhm = fitted_psf.y_fwhm * (np.abs(mosaic_header['CD2_2']) * 3600.)
# Measured FWHM in arcseconds
measured_mosaic_fwhm
mosaic_fwhm = measured_mosaic_fwhm
# ---
# <a id='yaml_generator_catalogs'></a>
# ### Run yaml_generator to create Mirage input yaml files
# User-inputs to the yaml generator. Note that you can still use a catalogs input here and add
# point sources or galaxies. Extended source catalog names will be added later
cr = {'library': 'SUNMAX', 'scale': 1.0}
dates = '2019-5-25'
background = 'low'
pav3 = 0.0
#catalogs = {'NGC1234': {'nircam': {'point_source': 'ngc1234_ptsrc_nrc.cat',
# 'galaxy': 'ngc1234_galaxy_nrc.cat',
# }
# }
# }
# Run the yaml generator
yam = yaml_generator.SimInput(xml_file, pointing_file, verbose=True,
output_dir='yamls',
cosmic_rays=cr,
#catalogs=catalogs,
background=background, roll_angle=pav3, dates=dates,
simdata_output_dir='simdata',
datatype='raw')
yam.use_linearized_darks = True
yam.create_inputs()
# ---
# <a id='source_catalogs'></a>
# ### Source catalogs
# Get a list of all instruments, apertures, and filters used in the APT file
# +
instruments = yam.info['Instrument']
filter_keywords = ['FilterWheel', 'ShortFilter', 'LongFilter', 'Filter']
pupil_keywords = ['PupilWheel', 'ShortPupil', 'LongPupil']
yam.info
nrc_sw_optics = set([(f, p) for f, p in zip(yam.info['ShortFilter'], yam.info['ShortPupil'])])
nrc_lw_optics = set([(f, p) for f, p in zip(yam.info['LongFilter'], yam.info['LongPupil'])])
niriss_optics = set([(f, p) for f, p in zip(yam.info['FilterWheel'], yam.info['PupilWheel'])])
niriss_wfss_optics = set([(f, p) for f, p in zip(yam.info['Filter'], yam.info['PupilWheel'])])
print('NIRCam filters/pupils used in this proposal: ')
print(nrc_sw_optics)
print(nrc_lw_optics)
print('\nNIRISS filters/pupils used in this proposal: ')
print(niriss_optics)
print(niriss_wfss_optics)
print(('\nBe sure to add magnitude columns to the template catalog '
'for all filters you are going to simulate.\n'))
# -
# #### Create extended source catalog
# Create a template extended source catalog containing sources other than the mosaic image that you want to add to the seed image. The resampled mosaic will be added to this template later. Note that you must add magnitude values for these other sources in all filters that are used in the proposal.
# If you do not have any extended sources other than the mosaic, set the template_cat to None, so that later we know there is nothing to combine with the catalog containing the mosaic data.
template_cat = None
# If you do have extended sources in addition to the mosaic image, create template_cat here and add those sources.
filter1 = 'F150W'
filter2 = 'F444W'
# +
other_stamp_files = ['3C305.1_stamp.fits', '3C324_stamp.fits', '3C454.1_stamp.fits']
other_stamp_ra = [53.164375, 53.168375, 53.160375]
other_stamp_dec = [-27.815355, -27.811355, -27.819355]
other_stamp_pa = [0., 0., 0.]
other_stamp_f150w_mags = [18., 19., 19.5]
other_stamp_f444w_mags = [22.5, 23.5, 24.0]
# Magnitude values must be strings here because we will be combining them
# with values of 'None' for the resampled image magnitudes
f150w_mags_as_str = [str(element) for element in other_stamp_f150w_mags]
f444w_mags_as_str = [str(element) for element in other_stamp_f444w_mags]
template_extended_catalog_file = 'extended_sources_template.cat'
template_cat = ExtendedCatalog(filenames=other_stamp_files, ra=other_stamp_ra, dec=other_stamp_dec,
position_angle=other_stamp_pa)
template_cat.add_magnitude_column(f150w_mags_as_str, instrument='nircam', filter_name=filter1)
template_cat.add_magnitude_column(f444w_mags_as_str, instrument='nircam', filter_name=filter2)
template_cat.save(template_extended_catalog_file)
# -
# ---
# <a id='crop_and_blot_catalog'></a>
# ### Extract images from mosaic, resample, and add to catalog
# In this step, crop a roughly detector-sized subarray from the mosaic image at the location specified in the yaml file. Convolve the subarray with the proper kernel in order to adjust the PSF in the mosaic to match that of the specified JWST detector and filter. Note that this can only be done in cases where the mosaic PSF's FWHM is smaller than the JWST PSF's FWHM, otherwise we would be sharpening the image. If you attempt to run the code in a situation like that, an exception will be raised.
#
# After convolution, the subarray is resampled onto the JWST pixel grid. Resample is essentially the same as Astrodrizzle's blot functionality.
#
# The resampled image is then added to the previously created extended source catalog (or kept in its own catalog if template_cat is None). This leads to an extended source catalog that is specific to the input yaml file used to control the cropping and resampleing. This extended source catalog is added to the yaml file so that when the simulated data are created, it will be used.
yam.yaml_files
# In this case, with the mosaic image created from HST/ACS F814W data, with a FWHM of 0.09", we cannot run the code for the NIRCam shortwave detectors (B1 - B4) with F150W, because the corresponding FWHM for that is smaller than 0.09". However, we can run the code for the NIRCam longwave detector (B5) with the F444W filter, where the FWHM is larger than 0.09".
for yfile in [yam.yaml_files[-1]]:
# Read in the yaml file so that we know RA, Dec, PAV3
# of the exposure
with open(yfile) as file_obj:
params = yaml.safe_load(file_obj)
ra = params['Telescope']['ra']
dec = params['Telescope']['dec']
pav3 = params['Telescope']['rotation']
# Define the output files and directories
sim_data_dir = params['Output']['directory']
simulated_filename = params['Output']['file']
crop_file = simulated_filename.replace('.fits', '_cropped_from_mosaic.fits')
crop_file = os.path.join(sim_data_dir, crop_file)
blot_file = simulated_filename.replace('.fits', '_blotted_seed_image.fits')
# Crop from the mosaic and resample for the desired detector/aperture
seed = ImgSeed(paramfile=yfile, mosaic_file=mosaicfile, cropped_file=crop_file,
outdir=sim_data_dir, blotted_file=blot_file, mosaic_fwhm=mosaic_fwhm,
mosaic_fwhm_units='arcsec', gaussian_psf=False)
seed.crop_and_blot()
# Now add the resampled file to the extended source catalog template and
# save as a separate catalog file
# Need to add a magnitude entry for each filter/pupil
mosaic_f150w_mag = ['None']
mosaic_f444w_mag = ['None']
# Create the catalog containing only the resampled image
blotted_image_full_path = os.path.join(sim_data_dir, blot_file)
extended_catalog_file = simulated_filename.replace('.fits', '_extended_sources.cat')
ext_cat = ExtendedCatalog(filenames=[blotted_image_full_path], ra=[ra], dec=[dec], position_angle=[pav3])
ext_cat.add_magnitude_column(mosaic_f150w_mag, instrument='nircam', filter_name=filter1)
ext_cat.add_magnitude_column(mosaic_f444w_mag, instrument='nircam', filter_name=filter2)
# Combine the resampled image catalog and the template catalog
if template_cat is not None:
combined_cat = combine_catalogs(ext_cat, template_cat)
combined_cat.save(extended_catalog_file)
else:
ext_cat.save(extended_catalog_file)
# Now add this extended source catalog to the yaml file
params['simSignals']['extended'] = extended_catalog_file
# Save the updated yaml file
with open(yfile, 'w') as file_obj:
dump = yaml.dump(params, default_flow_style=False)
file_obj.write(dump)
# ---
# <a id='create_data_catalog'></a>
# ### Create the simulated data
# Run the imaging simulator using the yaml files. Again in this case we run only the case with the NIRCam longwave detector.
for yfile in [yam.yaml_files[-1]]:
sim = ImgSim(paramfile=yfile)
sim.create()
# ---
# <a id='examine_data_catalog'></a>
# ### Look at simulated data
def show(array,title,min=0,max=1000):
plt.figure(figsize=(12,12))
plt.imshow(array,clim=(min,max), origin='lower')
plt.title(title)
plt.colorbar().set_label('DN$^{-}$/s')
# #### Seed image
# Note that the sources are more difficult to see than you might expect. This is because background signal has been added to the data. This will be in addition to any background signal present in the original mosaic image. As with any Mirage simulation, the level of background signal can be controlled using the `bkgdrate` parameter in the yaml file.
#
# The three stamp images added on top of the mosaic are visible in a diagonal line in the center of the image, going from upper left to lower right.
# Look at the noiseless seed image
show(sim.seedimage,'Seed Image', max=0.4)
# Zoom in on the center, where the three added stamp images are. The sources are in the center, the upper left corner, and the lower right corner.
show(sim.seedimage[700: 1300, 700: 1300],'Seed Image', max=0.6)
# ---
# <a id='resample_seed'></a>
# ## Use resampled image as a seed image
# In this case, rather than adding the cropped and resampled images to extended source catalogs which are then used in the simulated data generation, we instead use the cropped and resampled images as Mirage's seed images. This means that no sources other than the mosaic image can be used. After cropping and resampling, Mirage's dark current prep and observation generator are run, creating the final simulated data directly.
mosaicfile = 'hlsp_candels_hst_acs_gs-tot-sect23_f814w_v1.0_drz.fits'
xml_file = 'extended_object_test.xml'
pointing_file = xml_file.replace('.xml', '.pointing')
# ---
# <a id='yaml_generator_seed'></a>
# ### Run yaml_generator to create Mirage input yaml files
# User-inputs to the yaml generator. Note that you cannot use a catalogs input here to add extra
# point sources or galaxies.
cr = {'library': 'SUNMAX', 'scale': 1.0}
dates = '2019-5-25'
background = 'low'
pav3 = 0.0
# Run the yaml generator
yam = yaml_generator.SimInput(xml_file, pointing_file, verbose=True,
output_dir='yamls',
cosmic_rays=cr,
background=background, roll_angle=pav3, dates=dates,
simdata_output_dir='simdata',
datatype='raw')
yam.use_linearized_darks = True
yam.create_inputs()
# ---
# <a id='create_data_seed'></a>
# ### Create the simulted data
# In this step, crop a roughly detector-sized subarray from the mosaic image at the location specified in the yaml file. Convolve the subarray with the proper kernel in order to adjust the PSF in the mosaic to match that of the specified JWST detector and filter. Note that this can only be done in cases where the mosaic PSF's FWHM is smaller than the JWST PSF's FWHM, otherwise we would be sharpening the image. If you attempt to run the code in a situation like that, an exception will be raised.
#
# After convolution, the subarray is resampled onto the JWST pixel grid. Resample is essentially the same as Astrodrizzle's blot functionality.
#
# The resampled image serves as Mirage's seed image. The dark_prep and observation generator steps are then run to complete the simulated data generation.
# As in the previous method, we run only the NIRCam longwave detector data here, as this is the only case where the NIRCam PSF FWHM is larger than the mosaic PSF FWHM.
for yfile in [yam.yaml_files[-1]]:
# Read in the yaml file so that we know RA, Dec, PAV3
with open(yfile) as file_obj:
params = yaml.safe_load(file_obj)
# Define output filenames and directories
sim_data_dir = params['Output']['directory']
simulated_filename = params['Output']['file']
crop_file = simulated_filename.replace('.fits', '_cropped_from_mosaic.fits')
crop_file = os.path.join(sim_data_dir, crop_file)
blot_file = simulated_filename.replace('.fits', '_blotted_seed_image.fits')
# Crop from the mosaic and then resample the image
seed = ImgSeed(paramfile=yfile, mosaic_file=mosaicfile, cropped_file=crop_file,
outdir=sim_data_dir, blotted_file=blot_file, mosaic_fwhm=mosaic_fwhm,
mosaic_fwhm_units='arcsec', gaussian_psf=False)
seed.crop_and_blot()
# Run dark_prep
dark = DarkPrep()
dark.paramfile = yfile
dark.prepare()
# Run the observation generator
obs = Observation()
obs.paramfile = yfile
obs.seed = seed.seed_image
obs.segmap = seed.seed_segmap
obs.seedheader = seed.seedinfo
obs.linDark = dark.prepDark
obs.create()
# ---
# <a id='examine_data_seed'></a>
# ### Look at simulated data
seed_file = 'simdata/jw00042001001_01101_00001_nrcb5_uncal_blotted_seed_image.fits'
seed_image = fits.getdata(seed_file)
# In this case, no background has been added to the mosaic image because the cutout from the mosaic is being used directly as the seed image. The only background signal present is that present in the original image.
# Look at the noiseless seed image
show(seed_image,'Seed Image',max=0.03)
| examples/Simulated_data_from_mosaic_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ntds_2018
# language: python
# name: ntds_2018
# ---
# +
import tensorflow as tf
import keras as ks
import numpy as np
import script_config as sc
# Calculated for 2 Hop
maximum_neighborhood_size = sc._config_maximum_neighborhood_size
data_folder = sc._config_data_folder
random_seed = sc._config_random_seed
availible_threads = sc._config_availible_threads
feature_permutations = sc._config_feature_permutations
# +
def create_undirected_adjacency(filename):
loaded_data = np.load(filename)
users = loaded_data['local_neighbors']
relations = loaded_data['local_relations']
adj = np.zeros((len(users),len(users)))
# Randomly Permute all nodes except the first one
users[1:] = np.random.permutation(users[1:])
for relation in relations:
idx_1 = np.where(users[:,0] is relations[2])
idx_2 = np.where(users[:,0] is relations[3])
adj[idx_1,idx_2] = 1
adj[idx_2,idx_1] = 1
return (tf.convert_to_tensor(adj),tf.constant(users[0,0]))
def create_spectral_decomposition(adjacency):
n_nodes = tf.shape(adjacency)[0]
degrees_inv_sqrt = tf.reshape(tf.math.pow(tf.math.sqrt(tf.math.reduce_sum(adjacency,axis=0)),
tf.constant([-1]*n_nodes)),[n_nodes,1])
normalized_laplacian = tf.subtract(tf.eye(n_nodes),
tf.multiply(adjacency,
tf.linalg.matmul(degrees_inv_sqrt,
tf.transpose(degrees_inv_sqrt)) ))
e_values, e_vectors = tf.linalg.eigh(normalized_laplacian)
features = tf.multiply(e_values,e_vectors[0,:])
return features
# -
# !mkdir /nvme/drive_1/NTDS_Final/spectral_decomp
# !rm -r /nvme/drive_1/NTDS_Final/spectral_decomp/filtered
# !mkdir /nvme/drive_1/NTDS_Final/spectral_decomp/filtered
# +
np.random.seed(seed=random_seed)
# Start with tensor of files containing adjacency lists
file_name_tensor = tf.data.Dataset.list_files("/nvme/drive_1/NTDS_Final/local_list/filtered/*.csv",
shuffle=True,seed=random_seed)
# Create adjacency matrices and interleave versions with permuted node indices
adjacency_tensor, ground_truth = file_name_tensor.interleave(lambda x:
tf.data.TextLineDataset(x)
.repeat(feature_permutations)
.map(create_undirected_adjacency,
num_parallel_calls=tf.constant(availible_threads)),
cycle_length=tf.constant(feature_permutations),
block_length=1,
num_parallel_calls=tf.constant(availible_threads))
#
feature_tensor = adjacency_tensor.shuffle(tf.constant(100),seed=random_seed)\
.map(create_spectral_decomposition,num_parallel_calls=tf.constant(availible_threads))
# +
model = ks.Sequential([
ks.layers.Dense(128, activation=tf.nn.relu),
ks.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
| raphael_temp/Spectral_Decomposition_(Pre-Filtered).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
filepath= './data/'
covid = pd.read_csv(filepath + '/covid19-US.csv')
flight_jan19 = pd.read_csv(filepath + '/flights-jan19.csv')
flight_jan19 = flight_jan19.drop(flight_jan19.columns[[1,2,3,5,6,8,10]] , axis = 1)
flight_jan19['DAY'] = flight_jan19['FL_DATE'].apply(lambda x: datetime.strptime(x, '%Y-%m-%d').day)
flight_jan20 = pd.read_csv(filepath + '/flights-jan20.csv')
flight_jan20 = flight_jan20.drop(flight_jan20.columns[[1,2,3,5,6,8,10]] , axis = 1)
flight_jan20['DAY'] = flight_jan20['FL_DATE'].apply(lambda x: datetime.strptime(x, '%Y-%m-%d').day)
flight_jan19.head()
flight_jan20.head()
flight_jan19_origin = flight_jan19.groupby(['ORIGIN_CITY_NAME','DAY']).sum() #['FL_DATE','ORIGIN_CITY_NAME']
flight_jan20_origin = flight_jan20.groupby(['ORIGIN_CITY_NAME','DAY']).sum()
flight_jan19_dest = flight_jan19.groupby(['DEST_CITY_NAME', 'DAY']).sum()
flight_jan20_dest = flight_jan20.groupby(['DEST_CITY_NAME', 'DAY']).sum()
flight_jan19_both = flight_jan19.groupby(['DAY','ORIGIN_CITY_NAME','DEST_CITY_NAME']).sum()
flight_jan20_both = flight_jan20.groupby(['DAY','ORIGIN_CITY_NAME','DEST_CITY_NAME']).sum()
flight_jan19_both.head()
diff_origin = flight_jan19_origin.merge(flight_jan20_origin, left_on = ['ORIGIN_CITY_NAME','DAY'], right_on = ['ORIGIN_CITY_NAME','DAY'])
diff_origin = diff_origin.rename(columns={'FLIGHTS_y':'FLIGHTS_2020', 'FLIGHTS_x':'FLIGHTS_2019'})
diff_origin['DIFFERENCE'] = diff_origin['FLIGHTS_2020'] - diff_origin['FLIGHTS_2019']
diff_dest = flight_jan19_dest.merge(flight_jan20_dest, left_on = ['DEST_CITY_NAME','DAY'], right_on = ['DEST_CITY_NAME','DAY'])
diff_dest = diff_dest.rename(columns={'FLIGHTS_y':'FLIGHTS_2020', 'FLIGHTS_x':'FLIGHTS_2019'})
diff_dest['DIFFERENCE'] = diff_dest['FLIGHTS_2020'] - diff_dest['FLIGHTS_2019']
diff_both = flight_jan19_both.merge(flight_jan20_both, left_on = ['DAY','ORIGIN_CITY_NAME','DEST_CITY_NAME'], right_on = ['DAY','ORIGIN_CITY_NAME','DEST_CITY_NAME'])
diff_both = diff_both.rename(columns={'FLIGHTS_y':'FLIGHTS_2020', 'FLIGHTS_x':'FLIGHTS_2019'})
diff_both['DIFFERENCE'] = diff_both['FLIGHTS_2020'] - diff_both['FLIGHTS_2019']
diff_both = diff_both.reset_index()
print(diff_origin['DIFFERENCE'].max())
print(diff_dest['DIFFERENCE'].max())
print(diff_both['DIFFERENCE'].max())
print(diff_both['DIFFERENCE'].min())
diff_daily = diff_both.groupby(['DAY']).sum().reset_index()
diff_daily.head()
import matplotlib.pyplot as plt
plt.figure(figsize=(10,7))
#plt.xticks(rotation=90)
plt.bar(diff_both.groupby(['DAY']).sum().reset_index()['DAY'], diff_both.groupby(['DAY']).sum().reset_index()['DIFFERENCE'])
# +
from mpl_toolkits import mplot3d
fig = plt.figure()
ax = plt.axes(projection='3d')
# data for three-dimensional scattered points
zdata = diff_both['DAY']
xdata = diff_both['ORIGIN_CITY_NAME']
ydata = diff_both['DEST_CITY_NAME']
c = diff_both['DIFFERENCE']
ax.scatter3D(xdata, ydata, zdata, c);
# -
diff_both['ORIGIN_CITY_NAME']
| Project/.ipynb_checkpoints/data_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### API로 날씨 정보 가져오기
# - darksky.net 서비스를 사용
# - 회원가입 및 로그인
import requests
KEY = "API KEY" # API KEY
lat, lng = 37.8267, -122.4233 # 위도, 경도
# url 설정
url = "https://api.darksky.net/forecast/{}/{},{}".format(KEY, lat, lng)
response = requests.get(url)
response
data = response.json()
data["timezone"], data["hourly"]["summary"]
def forecast(lat, lng):
url = "https://api.darksky.net/forecast/{}/{},{}".format(KEY, lat, lng)
response = requests.get(url)
data = response.json()
return data["timezone"], data["hourly"]["summary"]
# 서울 위도, 경도 : 37.5665, 126.9780
forecast(37.5665, 126.9780)
| Crawling/02_requests_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
__author__ = "<NAME>"
__email__ = "<EMAIL>"
import matplotlib.pyplot as plt
from skimage import io
import numpy as np
def threshold(image, th=None):
"""Returns a binarised version of given image, thresholded at given value.
Binarises the image using a global threshold `th`. Uses Otsu's method
to find optimal thrshold value if the threshold variable is None. The
returned image will be in the form of an 8-bit unsigned integer array
with 255 as white and 0 as black.
Parameters:
-----------
image : np.ndarray
Image to binarise. If this image is a colour image then the last
dimension will be the colour value (as RGB values).
th : numeric
Threshold value. Uses Otsu's method if this variable is None.
Returns:
--------
binarised : np.ndarray(dtype=np.uint8)
Image where all pixel values are either 0 or 255.
"""
# Setup
shape = np.shape(image)
binarised = np.zeros([shape[0], shape[1]], dtype=np.uint8)
if len(shape) == 3:
image = image.mean(axis=2)
elif len(shape) > 3:
raise ValueError('Must be at 2D image')
if th is None:
th = otsu_thval(image)
print(f'The threshold value is {th}')
binarised = image > th
return binarised
def histogram(image):
"""Returns the image histogram with 256 bins.
"""
# Setup
shape = np.shape(image)
histogram = np.zeros(256)
if len(shape) == 3:
image = image.mean(axis=2)
elif len(shape) > 3:
raise ValueError('Must be at 2D image')
# here [0] means we want to extract only pixel values
histogram = np.histogram(image.ravel(), np.array(range(0,257)))[0]
return histogram #we hust return pixel values which is used to find the otsu threshold value
def otsu_thval(image):
"""Finds the optimal thresholdvalue of given image using Otsu's method.
"""
hist = histogram(image)
th = 0
total_pixel = np.sum(hist) #finding total sum of the pixels of the image
levels = np.array(range(0,256))
temp = -1 #declaring temporary value
for i in range(1, len(levels)):
#finding weight of background and foreground
wb = (np.sum(hist[:i])) / total_pixel
wf = (np.sum(hist[i:])) / total_pixel
#finding mean of background and foreground
meanb = np.sum(levels[:i] * hist[:i]) / np.sum(hist[:i])
meanf = np.sum(levels[i:] * hist[i:]) / np.sum(hist[i:])
#finding between class variance(bcv)
bcv = wb * wf * (meanb - meanf) ** 2
#checking in such a way that biggest between class variance is obtained
if bcv > temp:
th = i
temp = bcv
return th
if __name__ == "__main__":
filename = "Bridge.jpg"
image = io.imread(filename)
binarised_image = threshold(image)
fig, axes = plt.subplots(ncols=2, figsize=(10,4))
ax = axes.ravel()
ax[0].imshow(image, cmap=plt.cm.gray)
ax[0].set_title("Original Image")
ax[1].imshow(binarised_image, cmap=plt.cm.gray)
ax[1].set_title("Binarised Image")
# -
| otsu_thresholding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Connector for Twitter
#
# In this example, we will be going over how to use Connector with Twitter.
# ## Prerequisites
#
# connector is a component in the dataprep library that aims to simplify the data access by providing a standard API set. The goal is to help the users skip the complex API configuration. In this tutorial, we demonstrate how to use connector library with Twitter.
#
# If you haven't installed dataprep, run command `pip install dataprep` or execute the following cell.
# +
# Run me if you'd like to install
# #!pip install dataprep
# -
# # Obtaining access token from Twitter
#
# In order for you to get the Twitter feed working, you need four keys - the **Consumer Key**, **Consumer Secret**, **Access Token** and **Access Token Secret**. These keys are unique identifiers of an application requesting access to the Twitter's API. Once an application creates the token, it will act as your credential when making an API request. These keys and tokens can be obtained with the following three simple steps.
#
# ##### 1. Create a Developer Account
# In order to access the data, you will need to create a server-side application on Twitter and to do so, you will need to create a developer account in order to create an application. You can sign up for the developer account on the [Twitter Developer Account](https://developer.twitter.com/en) by following the prompts. An email confirming your approval to the developer account will be sent to you shortly after the sign up.
#
# ##### 2. Create an Application
# Logging in to your developer account once you gain access, create an app in the [Apps section of your profile](https://developer.twitter.com/en/apps). While creating an application, fill in the name, description and URL of your website. You can leave the clalback URL and other URL sections if needed.
#
# ##### 3. Generate Auth Keys and tokens
#
# After you successfully create an application, you can find your access keys and token Keys in the **Keys and Tokens** section of your Application Details. Generate the Access token and Access Token Secret and store them in a secure location as it will be used to provide you access to the Twitter's data.
# # Initialize connector
#
# To initialize run the following code. Copy and paste the **Twitter Access Token Secret** into the **access_token** variable and ensure the connector path is correct. This returns an object establishing a connection with Twitter. Once you have that running you can use the built in functions available in connector.
# +
from dataprep.connector import Connector
client_id = '<Consumer API Key>'
client_secret = '<Consumer API Secret Key>'
dc = Connector("twitter", _auth={"client_id":client_id, "client_secret":client_secret})
dc
# -
# # Functionalities
#
# Connector has several functions you can perform to gain insight on the data downloaded from twitter.
# ### Connector.info
# The info method gives information and guidelines of using the connector. There are 3 sections in the response and they are table, parameters and examples.
# >1. Table - The table(s) being accessed.
# >2. Parameters - Identifies which parameters can be used to call the method. For Twitter, q is a required parameter that acts as a filter.
# >3. Examples - Shows how you can call the methods in the Connector class.
dc.info()
# ### Connector.show_schema
# The show_schema method returns the schema of the website data to be returned in a Dataframe. There are two columns in the response. The first column is the column name and the second is the datatype.
#
# As an example, lets see what is in the tweets table.
dc.show_schema("tweets")
# ### Connector.query
# The query method downloads the website data and displays it in a Dataframe. The parameters must meet the requirements as indicated in connector.info for the operation to run. You can use the **count** parameter to specify the number of tweets to be fetched. Each request can currently fetch a maximum of 100 requests.
#
# When the data is received from the server, it will either be in a JSON or XML format. The connector reformats the data in pandas Dataframe for the convenience of downstream operations.
#
# As an example, let's try to get 50 tweets related to covid-19 from the Twitter Website.
# #### Searching for tweets related to COVID-19
df = dc.query("tweets", q="covid-19", count=50)
df
# # That's all for now.
# If you are interested in writing your own configuration file or modify an existing one, refer to the [Configuration Files](https://github.com/sfu-db/DataConnectorConfigs>).
| examples/DataConnector_Twitter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.11 ('dp')
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from transformers import AutoModel, AutoTokenizer, DataCollatorForTokenClassification, AutoModelForTokenClassification, TrainingArguments, Trainer
import torch
import torch.nn as nn
import pickle
from helper import get_labeled_answers, get_predicted_answers
tokenizer = AutoTokenizer.from_pretrained('KB/bert-base-swedish-cased')
model = torch.load('../results/model_CA_3e_ISNS_weighted_loss.pkl')
with open(r'../data/CA/tokenized_CA_data_eval.pkl', "rb") as input_file:
test_data = pickle.load(input_file)
tokens = tokenizer.convert_ids_to_tokens(test_data[0]["input_ids"])
print(tokens)
dec = tokenizer.decode(test_data[0]["input_ids"])
print(dec)
# TODO: check why this function cannot be exported!!
def compare_labels_and_output(output, labels, tokens):
labels_stats = []
for idx, label in enumerate(labels):
if label == 1:
count = 1
next_label = labels[idx+1]
while idx+count < len(labels) and next_label in [2, -100]:
count += 1
next_label = labels[idx+count]
labels_stats.append((idx, count))
print(labels_stats)
num_matches = 0
for ans in labels_stats:
ans_start = ans[0]
is_match = True
answer_text = []
for i in range(ans[1]):
answer_text.append(tokens[ans_start+i])
if output[ans_start+i] != labels[ans_start+i]:
is_match = False
if is_match:
num_matches += 1
print('found match: ', ' '.join(answer_text))
return len(labels_stats), num_matches
# +
# Output class
# https://huggingface.co/docs/transformers/main_classes/output#transformers.modeling_tf_outputs.TFTokenClassifierOutput
model.eval()
test_input = []
test_labels = []
test_attn = []
token_type_ids = []
for i in range(len(test_data)):
test_input.append(test_data[i]['input_ids'])
test_labels.append(test_data[i]['labels'])
test_attn.append(test_data[i]['attention_mask'])
token_type_ids.append(test_data[i]['token_type_ids'])
print(len(test_input))
print(len(test_labels))
print(len(test_attn))
num_correct = 0
num_predicted = 0
num_pos_data = 0
total_num_answers = 0
total_exact_match = 0
for i in range(len(test_data)):
output = model(torch.tensor([test_data[i]['input_ids']]), attention_mask=torch.tensor([test_data[i]['attention_mask']]), token_type_ids=torch.tensor([test_data[i]['token_type_ids']]), labels=torch.tensor([test_data[i]['labels']]))
print('test idx: ', i)
print('instance loss: ', output.loss)
# print(output.logits)
m = nn.Softmax(dim=2)
max = m(output.logits)
out = torch.argmax(max, dim=2)
# print(max)
print('Output length: ', out[0])
# print('labels length: ', len(test_data[i]['labels']))
# print('Labels: ', test_data[i]['labels'])
tokens = tokenizer.convert_ids_to_tokens(test_data[i]["input_ids"])
true_labels = test_data[i]['labels']
# print(tokens)
get_labeled_answers(true_labels, tokens)
get_predicted_answers(out[0], tokens)
num_answers, num_exact_matches = compare_labels_and_output(out[0], true_labels, tokens)
total_num_answers += num_answers
total_exact_match += num_exact_matches
for idx, pred_label in enumerate(out[0]):
true_label = true_labels[idx]
if true_label > 0:
num_pos_data += 1
if pred_label > 0:
# print('label: ', pred_label)
# print('token: ', tokens[idx])
num_predicted += 1
if pred_label == true_label:
num_correct += 1
# calculate precision and recall
pr = num_correct/num_predicted
rec = num_correct/num_pos_data
print('precision: ', pr)
print('recall: ', rec)
print('percentage of exact matches: ', total_exact_match/total_num_answers)
# -
| model/evaluation/eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.random.seed(0)
# +
#computing manually reciprocal
def compute_reciprocal(value):
output = np.empty(len(value))
for i , val in enumerate(value):
output[i] = 1.0/val
return output
value = np.random.randint(1 , 10 , size=5)
compute_reciprocal(value)
# -
bing_arr = np.random.randint(1,100 , size=1000000)
# %timeit compute_reciprocal(bing_arr)
# # Functions
print(compute_reciprocal(value))
print(1.0/value)
# %timeit 1.0/bing_arr
#
#
# Vectorized operations in NumPy are implemented via ufuncs, whose main purpose is to quickly execute repeated operations on values in NumPy arrays. Ufuncs are extremely flexible – before we saw an operation between a scalar and an array, but we can also operate between two arrays:
#
np.arange(5) / np.arange(1,6)
x = np.arange(9).reshape(3 , 3)
2**x
# +
y = np.arange(4)
print(y)
print(y+1)
print(y-1)
print(y/2)
print(y//2)
print(y*2)
# -
print(-y)
print(y**3)
print(y %2)
-(0.5*y +1) **2
# # numpy function
# Operator Equivalent ufunc Description
# + (+) np.add Addition (e.g., 1 + 1 = 2)
# - (-) np.subtract Subtraction (e.g., 3 - 2 = 1)
# - (-) np.negative Unary negation (e.g., -2)
# * (*) np.multiply Multiplication (e.g., 2 * 3 = 6)
# * / np.divide Division (e.g., 3 / 2 = 1.5)
# * // np.floor_divide Floor division (e.g., 3 // 2 = 1)
# * ** np.power Exponentiation (e.g., 2 ** 3 = 8)
# * % np.mod Modulus/remainder (e.g., 9 % 4 = 1)
type(y)
x = np.array([-2, 4 , -6 , -8])
type(x)
abs(x)
np.absolute(x)
np.abs(x)
# This ufunc can also handle complex data, in which the absolute value returns the magnitude:
x = np.array([3 - 4j, 4 - 3j, 2 + 0j, 0 + 1j])
np.abs(x)
# # Trigonometric function
theta = np.linspace(0 , np.pi , 5)
theta
print(np.sin(theta) , np.cos(theta) , np.tan(theta) , )
x = [-1 , 0 , 1]
#inverse Trigonometric function
print(np.arccos(x) , np.arcsin(x) , np.arctan(x))
# # Exponent and logarithmics
# +
x = [2 , 3 , 4]
print("x = " , x)
print("e^x = " , np.exp(x))
print("2^x = " , np.exp2(x))
print("3^x= " , np.power(3 , x))
# -
x = [1, 2, 4, 10]
print("x =", x)
print("ln(x) =", np.log(x))
print("log2(x) =", np.log2(x))
print("log10(x) =", np.log10(x))
# There are also some specialized versions that are useful for maintaining precision with very small input:
# +
x = [0, 0.001, 0.01, 0.1]
print("exp(x) - 1 =", np.expm1(x))
print("log(1 + x) =", np.log1p(x))
# -
from scipy import special
# Gamma functions (generalized factorials) and related functions
x = [1, 5, 10]
print("gamma(x) =", special.gamma(x))
print("ln|gamma(x)| =", special.gammaln(x))
print("beta(x, 2) =", special.beta(x, 2))
# Error function (integral of Gaussian)
# its complement, and its inverse
x = np.array([0, 0.3, 0.7, 1.0])
print("erf(x) =", special.erf(x))
print("erfc(x) =", special.erfc(x))
print("erfinv(x) =", special.erfinv(x))
x = np.arange(5)
y = np.empty(5)
np.multiply(x , 10 , out=y)
y = np.zeros(10)
np.power( 2 , x , out=y[::2])
print(y)
# # Aggregates
x = np.arange(1, 5)
# A reduce repeatedly applies a given operation to the elements of an array until only a single result remains.
#
# For example, calling reduce on the add ufunc returns the sum of all elements in the array:
np.add.reduce(x)
np.multiply.reduce(x)
np.add.accumulate(x)
np.multiply.accumulate(x)
#
# Outer products
#
# Finally, any ufunc can compute the output of all pairs of two different inputs using the outer method. This allows you, in one line, to do things like create a multiplication table:
#
np.multiply.outer(x , x)
| NUMPY/Computation_numpy_array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Pittsburgh Neighborhood Parks Analysis**
#
# *CMPINF 0010 Spring 2022 Final Project* \
# *<NAME>*
# # Introduction
#
# We are performing a data-driven analysis to find the best neighborhood in Pittsburgh in terms of environmental quality. We are doing this by using datasets from the Western Pennsylvania Regional Data Center (WPRDC).
#
# # Parks
#
# My own submetric is parks in Pittsburgh. In my analysis, the number, density, and quality of parks is used to determinine the best neighborhood in Pittsburgh. [The Parks dataset](https://data.wprdc.org/dataset/parks/resource/fa329e3d-89ff-4708-8ddf-81bfedcad11d), [the neighborhood areas dataset](https://data.wprdc.org/dataset/neighborhoods2), and the neighborhood populations dataset were obtained from the Western Pennsylvania Regional Data Center.
import pandas as pd
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
parks = pd.read_csv("Parks.csv")
parks.head()
neighborhoods = parks["neighborhood"].unique()
print(neighborhoods)
print("There are " + str(parks["name"].size) + " parks listed in the dataset.")
print("There are " + str(len(neighborhoods)) + " neighborhoods listed in the Parks dataset.")
nbhdList = pd.read_csv("NeighborhoodList.csv")
print("There are a total of " + str(len(nbhdList["hood"])) + " neighborhoods in Pittsburgh.")
# Some of the entries in the Parks dataset are not actually parks. Some are listed as a traffic island, median, or building grounds. Since we are focusing on the environment, we only want to look at parks because they are outdoor spaces.
parks = parks[parks.type == "Park"]
print("There are 209 entires in the dataset, but " + str(len(parks.index)) + " of them are just parks.")
parkCount = parks.groupby(by = "neighborhood").count()
parkCount = parkCount["id"].sort_values(ascending = False)
parkCount = parkCount.rename("parkCount")
parkCount
plt.style.use('ggplot')
parkCount.head(10).plot.bar(rot=50, color = "Green")
plt.title("Pittsburgh Neighborhood Parks")
plt.xlabel("Neighborhood")
plt.ylabel("Number of Parks")
# This is a graph of the top 10 Pittsburgh neighborhoods with the most parks. Now let's consider the density of the parks
# per total area of the neighborhood.
# # Park Density by Area
areas = pd.read_csv("NeighborhoodList.csv")
areas = areas[["hood","acres"]]
areas = areas.rename(columns={"hood":"Neighborhood","acres":"Acres"})
areas.head(5)
areas = areas.sort_values('Neighborhood')
areas.head(5)
parkCount = pd.DataFrame(parkCount)
parkCount = parkCount.sort_values(by="neighborhood")
parkCount.head(5)
parkCountArea = parkCount.merge(areas,how='inner',left_on="neighborhood",right_on="Neighborhood")
columns_titles = ["Neighborhood","parkCount","Acres"]
parkCountArea = parkCountArea.reindex(columns=columns_titles)
print(parkCountArea[parkCountArea["Neighborhood"]=="Perry North"].index.values)
parkCountArea.iat[48,2] = 700
parkCountArea.head(5)
# Perry North did not contain an acreage in the dataset. It looks roughly the same size as Brighton Heights, which is 718 acres. I will assign an estimate of 700 acres to Perry North. Since it will end up towards the bottom of our ranking list, the exact acreage is not needed.
parkCountArea["Parks per Acre"] = parkCountArea['parkCount']/parkCountArea['Acres']
parkCountArea.head(5)
parkCountArea = parkCountArea.sort_values("Parks per Acre",ascending = False)
parkCountArea['Rank'] = np.arange(len(parkCountArea))
parkCountArea.head(10)
plt.style.use('ggplot')
parkCountArea[["Neighborhood","Parks per Acre"]].head(10).plot.bar(rot=55, color = "Green",x='Neighborhood',y="Parks per Acre")
plt.title("Pittsburgh Neighborhood Park Density")
plt.xlabel("Neighborhood")
plt.ylabel("Number of Parks per Acre")
# Here are the top ten neighborhoods in parks per acre density. This means these neighborhoods have the most parks compared to their size, so it will take less distance to visit them. Having more parks per acre means there is more area reserved for outdoor spaces compared to roads and buildings. This contributes to the neighborhood having a higher quality environment.
# # Parks Density by Population
pops = pd.read_csv("NeighborhoodPopulations.csv")
pops = pops[["Neighborhood","2020_Total_Population"]]
pops.head(5)
pops = pops.sort_values('Neighborhood')
parkCount = pd.DataFrame(parkCount)
parkCount = parkCount.sort_values(by="neighborhood")
parkCountPop = parkCount.merge(pops,how='inner',left_on="neighborhood",right_on="Neighborhood")
columns_titles = ["Neighborhood","parkCount","2020_Total_Population"]
parkCountPop = parkCountPop.reindex(columns=columns_titles)
parkCountPop.head(5)
parkCountPop["Parks per Person"] = parkCountPop['parkCount']/parkCountPop['2020_Total_Population']
parkCountPop.head(5)
parkCountPop = parkCountPop.sort_values("Parks per Person",ascending = False)
parkCountPop['Rank'] = np.arange(len(parkCountPop))
parkCountPop
plt.style.use('ggplot')
parkCountPop[["Neighborhood","Parks per Person"]].head(10).plot.bar(rot=55, color = "Green",x='Neighborhood',y="Parks per Person")
plt.title("Pittsburgh Neighborhood Park Density")
plt.xlabel("Neighborhood")
plt.ylabel("Number of Parks per Person")
# Here are the top ten neighborhoods in parks per person. This means these neighborhoods have the most parks compared to their population, so there will be more parks per residence. Having more parks per person means there is more area reserved for outdoor spaces because of there is a higher density of parks when compared to the space that residences would take up. This contributes to the neighborhood having a higher quality environment.
# # Combining the two density submetrics
parkCountArea = parkCountArea.sort_values('Neighborhood')
parkCountPop = parkCountPop.sort_values('Neighborhood')
mergedRanks = parkCountArea.merge(parkCountPop,left_on="Neighborhood",right_on="Neighborhood")
mergedRanks = mergedRanks[["Neighborhood","Rank_x","Rank_y"]]
mergedRanks = mergedRanks.rename(columns={"Rank_x":"Area Rank","Rank_y":"Population Rank"})
mergedRanks
mergedRanks["Average Rank"] = (mergedRanks["Area Rank"] + mergedRanks["Population Rank"]/5)/2
mergedRanks
# When combining the ranks to an average rank, I weighted the Population Rank less by dividing it by 5. This is because density of parks per acre is a much better measurement of the quality of the environment compared to parks per person.
mergedRanks = mergedRanks.sort_values(by="Average Rank")
mergedRanks = mergedRanks[["Neighborhood","Average Rank"]]
print(mergedRanks[mergedRanks["Neighborhood"]=="Squirrel Hill South"].index.values)
plt.style.use('ggplot')
mergedRanks[["Neighborhood","Average Rank"]].head(10).plot.bar(rot=55, color = "Green",x='Neighborhood',y="Average Rank")
plt.title("Pittsburgh Neighborhood Parks Rank")
plt.xlabel("Neighborhood")
plt.ylabel("Average Ranking")
# This graph shows the average ranking of Pittsburgh neighborhoods between the metrics of parks per acre and parks per person. A lower ranking value is better because it means the neighborhood placed higher on the list.
# # Conclusion
#
# When considering the metric of both park density by area and by population, **East Liberty** has the highest parks ranking from my analysis.
#
# However, it is important to note that there many flaws in this data and this analysis:
# 1. The parks dataset only contains 73 out of 90 Pittsburgh neighborhoods.
# 2. The final ranking list contains less neighborhoods, at 68.
# 3. There are only 153 usable entries in this dataset, spread across 73 neighborhoods, which means many neighborhoods end up with 1 or 2 parks, and thus are easily skewed.
# 4. The size, quality, and construction of the parks are important to consider. For example, Schenley Park is clearly a much better and more expansive park than something like Able Long Park, which is merely a playground. In order to combat this flaw, it would be necessary to consider each of the 153 parks individually, rate them, and then use that data in this analysis. However, that is obviously unfeasible and unsuited for the nature of this project.
# 5. The area density of the parks would make more sense if the areas of the parks were included, not just the quantity.
#
# Personally, my favorite neighborhood in Pittsburgh would be Shadyside, but unfortunately it did not make the top ten in this metric.
| Parks/Parks Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # new_daily_price ratio
# ## Improvement to be made
# - add rate of return to the features **DONE**
# - add different transformations to the features
# - add benchmark(SVM is so) to the existing model sets
# - try some standardization methods
# - try some classifiers: add neural network, add adaboosting and gradient boosting, ensemble methods, etc **DONE**
# - conduct feature selection : Remove collineariry, RFE, tree-based methods, then rerun the models **DONE**
# - conduct dimension reduction, then rerun the models **DONE**
# - use RFC with logistic regression, and select top 5 features to build a logistic regression to study the effect of geo feature(s). **DONE**
# - add accuracy evaluation **DONE**
# # Feature Engeneering
# +
import numpy as np
import pandas as pd
import seaborn as sns
from datetime import datetime
import matplotlib.pylab as plt
from sklearn.model_selection import train_test_split #training and testing data split
# +
ua=pd.read_csv('UA.csv')
ua['time']=pd.DatetimeIndex(pd.to_datetime(ua['Date']))
ua=ua.set_index('time')
ua=ua.rename(columns={'Close':'ua_price'})
ua=ua.rename(columns={'Volume':'ua_stkv'})
ua.head()
# -
nike=pd.read_csv('NKE.csv')
nike['time']=pd.DatetimeIndex(pd.to_datetime(nike['Date']))
nike=nike.set_index('time')
nike=nike.rename(columns={'Close':'nike_price'})
nike=nike.rename(columns={'Volume':'nike_stkv'})
nike.head()
ua_price=ua.ua_price
nike_price=nike.nike_price
ua_stkv=ua.ua_stkv
nike_stkv=nike.nike_stkv
# load data
total=pd.read_csv('alldata.csv')
total=total.rename(columns={'Unnamed: 0':'time'})
total['time']=pd.DatetimeIndex(pd.to_datetime(total['time']))
total=total.set_index('time')
total.head(3)
# +
# transform the original 30-min data to daily data
count=total[['ua_count','nike_count']].resample('D').sum()
# price=total[['ua_price','nike_price']].resample('D').last()
# stkv=total[['ua_stkv','nike_stkv']].resample('D').sum()
tweet=total[['ua_tweet','nike_tweet']].resample('D').mean()
twtcount=total[['ua_twtcount','nike_twtcount']].resample('D').sum()
total = pd.concat([count,ua_price,nike_price,ua_stkv,nike_stkv,tweet,twtcount], axis=1)
# -
total.head()
# +
total['count_ratio']=total.ua_count/total.nike_count
total['price_ratio']=total.ua_price/total.nike_price
total['tweet_diff']=total.ua_tweet-total.nike_tweet
total['stkv_ratio']=total.ua_stkv/total.nike_stkv
total['twtcount_ratio']=total.ua_twtcount/total.nike_twtcount
# +
# total.tweet_diff.sort_values()
# -
total.head(3)
total.shape
# +
ssize = 8
msize = 10
bsize = 12
plt.rc('font', size=16) # controls default text sizes
plt.rc('axes', titlesize=12) # fontsize of the axes title
plt.rc('axes', labelsize=22, labelweight='bold') # fontsize of the x and y labels
plt.rc('xtick', labelsize=18) # fontsize of the tick labels
plt.rc('ytick', labelsize=22) # fontsize of the tick labels
plt.rc('legend', fontsize=18) # legend fontsize
plt.rc('figure', titlesize=12) # fontsize of the figure title
plt.rc('font',weight='bold')
# -
plt.rcParams['font.sans-serif'] = "Times New Roman"
# Then, "ALWAYS use sans-serif fonts"
plt.rcParams['font.family'] = "sans-serif"
# +
# plot the daily data
alldata1=total
figure=plt.figure(figsize=(13,15))
axes1=figure.add_subplot(2,1,1)
alldata1[['ua_count','nike_count']].plot(ax=axes1)
axes2=figure.add_subplot(2,1,2)
alldata1['ua_price'].plot(ax=axes2)
axes1.set_ylabel('Geo Location Customer Count',fontname="Times New Roman")
axes2.set_ylabel('UAA Stock Price',fontname="Times New Roman")
# -
alldata1=total
# +
figure=plt.figure(figsize=(13,15))
axes1=figure.add_subplot(2,1,1)
alldata1['count_ratio'].plot(ax=axes1)
axes2=figure.add_subplot(2,1,2)
alldata1['price_ratio'].plot(ax=axes2)
axes1.set_ylabel('Geo count ratio')
axes2.set_ylabel('Stock price ratio')
# -
# remove the missing values
total=total.dropna()
total.shape
# +
# create labels for classification
value_ua = total.ua_price.shift(-1)-total.ua_price
value_nike=total.nike_price.shift(-1)-total.nike_price
value_ratio=total.price_ratio.shift(-1)-total.price_ratio
value_ua[value_ua>=0]=1
value_ua[value_ua<0]=0
value_nike[value_nike>=0]=1
value_nike[value_nike<0]=0
value_ratio[value_ratio>=0]=1
value_ratio[value_ratio<0]=0
print(value_ua.shape)
print(value_nike.shape)
print(value_ratio.shape)
# -
# rename the geo-related features
total=total.rename(columns={'ua_count':'geo_ua_count'})
total=total.rename(columns={'nike_count':'geo_nike_count'})
total=total.rename(columns={'count_ratio':'geo_count_ratio'})
# add rate of returns
return_ua=total.ua_price/total.ua_price.shift(1)-1
return_nike=total.nike_price/total.nike_price.shift(1)-1
return_diff=return_ua-return_nike
# add labels and rate of returns to the dataframe
total['return_ua']=return_ua
total['return_nike']=return_nike
total['return_diff']=return_diff
total['value_ua']=value_ua
total['value_nike']=value_nike
total['value_ratio']=value_ratio
total[['ua_tweet','nike_tweet','tweet_diff']]=total[['ua_tweet','nike_tweet','tweet_diff']].fillna(method='bfill')
print(total.shape)
print(total.dropna().shape)
# +
total1=total.dropna()
total1.head()
# -
total1.columns
# +
#seperate the data into train and test sets
X=total1.drop(total1.columns[[18,19,20]],axis=1)
y=total1.value_ratio
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=0)
# -
print(X.shape)
print(y.shape)
# +
# X
# -
# # EDA
#correlation map
f,ax = plt.subplots(figsize=(10, 8))
sns.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax)
ax = sns.countplot(y,label="Count")
ups, downs = y.value_counts()
print('Number of ups: ',ups)
print('Number of downs : ',downs)
data_y = y
data_x = X
data_s = (data_x - data_x.mean()) / (data_x.std()) #standardize
X.shape
# # Model Fitting
#importing all the required ML packages
from sklearn.linear_model import LogisticRegression #logistic regression
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn import svm #support vector Machine
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.neighbors import KNeighborsClassifier #KNN
from sklearn.naive_bayes import GaussianNB #Naive bayes
from sklearn.tree import DecisionTreeClassifier #Decision Tree
from sklearn.neural_network import MLPClassifier #neural network
#import xgboost as xg #xgboost
#feature selection and evaluation
from sklearn import preprocessing
from sklearn.feature_selection import RFE
from sklearn.metrics import confusion_matrix,accuracy_score #for confusion matrix
from sklearn.model_selection import GridSearchCV, cross_val_score, StratifiedKFold, learning_curve,cross_val_predict,TimeSeriesSplit
# ## Baseline Model
# ### Hyper-Parameters Tuning
# ##### SVM
C=range(10,300,10)
gamma=[1.0,1.1,1.2,1.3,1.4,1.5,1.6,1.7,1.8,1.9,2.0]
# kernel=['rbf','linear']
hyper={'C':C,'gamma':gamma}
gd=GridSearchCV(estimator=svm.SVC(),param_grid=hyper,verbose=True)
gd.fit(X,y)
print(gd.best_score_)
print(gd.best_estimator_)
# #### Random Forest
n_estimators=range(10,100,10)
hyper={'n_estimators':n_estimators}
gd=GridSearchCV(estimator=RandomForestClassifier(random_state=0),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X,y)
print(gd.best_score_)
print(gd.best_estimator_)
# #### KNN
n_neighbors=[2,3,4,5,6,7,8,9,10]
weights=['uniform','distance']
hyper={'n_neighbors':n_neighbors,'weights':weights}
gd=GridSearchCV(estimator=KNeighborsClassifier(),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X,y)
print(gd.best_score_)
print(gd.best_estimator_)
# ### Cross Validation
# +
#two cross validation method
# kfold = TimeSeriesSplit(n_splits=3)
kfold = StratifiedKFold(n_splits=3)
# -
#standardization
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(X)
X_s= scaler.transform(X)
# X_s
X.shape
# +
# aggregate all the methods in one plot
random_state = 4
classifiers = []
classifiers.append(svm.SVC(random_state=random_state))
classifiers.append(DecisionTreeClassifier(random_state=random_state))
classifiers.append(RandomForestClassifier(n_estimators=10,random_state=random_state))
classifiers.append(KNeighborsClassifier(n_neighbors=2, p=2,weights='distance'))
classifiers.append(LogisticRegression(random_state = random_state))
classifiers.append(LDA())
classifiers.append(QDA())
classifiers.append(GaussianNB())
classifiers.append(MLPClassifier(random_state=random_state))
classifiers.append(AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1))
classifiers.append(GradientBoostingClassifier(random_state=random_state))
cv_results = []
for classifier in classifiers :
cv_results.append(cross_val_score(classifier, X, y, scoring = "accuracy", cv = kfold))
cv_means = []
cv_std = []
for cv_result in cv_results:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
cv_res = pd.DataFrame({"Method":["SVC","DecisionTree",
"RandomForest","KNeighboors","LogisticRegression","LinearDiscriminantAnalysis","QuadraticDiscriminantAnalysis","NaiveBayes","NeuralNetwork",
"AdaBoosting","GradientBoosting"],"CrossValMeans":cv_means,"CrossValerrors": cv_std})
cv_res = cv_res.sort_values(['CrossValMeans'],ascending=False).reset_index(drop=True)
cv_res
# +
g = sns.barplot("CrossValMeans","Method",data = cv_res, palette="Set2")
g.set_xlabel("Mean Accuracy")
g = g.set_title("Cross validation scores")
plt.show()
# -
# ### Confusion Matrix and Accuracy
# +
f,ax=plt.subplots(3,4,figsize=(12,10))
acc=[]
model=svm.SVC()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,0],annot=True,fmt='d')
ax[0,0].set_title('SVC is: ')
model=DecisionTreeClassifier()
model.fit(X_train,y_train)
y_pred =model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,1],annot=True,fmt='d')
ax[0,1].set_title('Decision Tree')
model=RandomForestClassifier(n_estimators=50)
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,2],annot=True,fmt='d')
ax[0,2].set_title('Random Forests')
model=KNeighborsClassifier(n_neighbors=2, p=2,weights='distance')
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,3],annot=True,fmt='d')
ax[0,3].set_title('KNN')
model=LogisticRegression()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,0],annot=True,fmt='d')
ax[1,0].set_title('Logistic Regression')
model=LDA()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,1],annot=True,fmt='d')
ax[1,1].set_title('LDA')
model=QDA()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,2],annot=True,fmt='d')
ax[1,2].set_title('QDA')
model=GaussianNB()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,3],annot=True,fmt='d')
ax[1,3].set_title('Naive Bayes')
model=pipeline
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,0],annot=True,fmt='d')
ax[2,0].set_title('Neural Network')
model=AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1)
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,1],annot=True,fmt='d')
ax[2,1].set_title('AdaBoosting')
model=GradientBoostingClassifier()
model.fit(X_train,y_train)
y_pred = model.predict(X_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,2],annot=True,fmt='d')
ax[2,2].set_title('Gradient Boosting')
plt.subplots_adjust(hspace=0.2,wspace=0.2)
plt.show()
# -
accdf = pd.DataFrame({"Accuracy":acc,"Algorithm":["SVC","DecisionTree",
"RandomForest","KNeighboors","LogisticRegression","LinearDiscriminantAnalysis","QuadraticDiscriminantAnalysis","NaiveBayes","NeuralNetwork",
"AdaBoosting","GradientBoosting"]})
accdf = accdf.sort_values(['Accuracy'],ascending=False).reset_index(drop=True)
accdf
# ### Feature selection and importance
# #### Correlation method
#correlation
X.columns
# ### take geolocation factors out
# +
# X2=X[['geo_ua_count','geo_nike_count','ua_price', 'nike_price', 'ua_stkv',
# 'nike_stkv', 'ua_tweet', 'nike_tweet', 'ua_twtcount', 'nike_twtcount',
# 'price_ratio', 'tweet_diff', 'stkv_ratio',
# 'twtcount_ratio', 'return_ua', 'return_nike', 'return_diff']]
# -
X1=X[['geo_count_ratio', 'price_ratio','tweet_diff','stkv_ratio','twtcount_ratio','return_diff']]
#correlation map
f,ax = plt.subplots(figsize=(10, 8))
sns.heatmap(X1.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax)
plt.show()
C=range(10,300,10)
gamma=[1.0,1.1,1.2,1.3,1.4,1.5,1.6,1.7,1.8,1.9,2.0]
# kernel=['rbf','linear']
hyper={'C':C,'gamma':gamma}
gd=GridSearchCV(estimator=svm.SVC(),param_grid=hyper,verbose=True)
gd.fit(X1,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_estimators=range(10,100,10)
hyper={'n_estimators':n_estimators}
gd=GridSearchCV(estimator=RandomForestClassifier(random_state=0),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X1,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_neighbors=[2,3,4,5,6,7,8,9,10]
weights=['uniform','distance']
hyper={'n_neighbors':n_neighbors,'weights':weights}
gd=GridSearchCV(estimator=KNeighborsClassifier(),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X1,y)
print(gd.best_score_)
print(gd.best_estimator_)
# +
# aggregate all the methods in one plot
random_state = 4
classifiers = []
classifiers.append(svm.SVC(random_state=random_state,C=10,gamma=1.0))
classifiers.append(DecisionTreeClassifier(random_state=random_state))
classifiers.append(RandomForestClassifier(n_estimators=30,random_state=random_state))
classifiers.append(KNeighborsClassifier(n_neighbors=2, p=2,weights='distance'))
classifiers.append(LogisticRegression(random_state = random_state))
classifiers.append(LDA())
classifiers.append(QDA())
classifiers.append(GaussianNB())
classifiers.append(MLPClassifier(random_state=random_state))
classifiers.append(AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1))
classifiers.append(GradientBoostingClassifier(random_state=random_state))
cv_results = []
for classifier in classifiers :
cv_results.append(cross_val_score(classifier, X1, y, scoring = "accuracy", cv = kfold))
cv_means = []
cv_std = []
for cv_result in cv_results:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
cv_res = pd.DataFrame({"Method":["SVM","DecisionTree",
"RandomForest","KNeighboors","LogisticRegression","LinearDiscriminantAnalysis","QuadraticDiscriminantAnalysis","NaiveBayes","NeuralNetwork",
"AdaBoosting","GradientBoosting"],"CrossValMeans":cv_means,"CrossValerrors": cv_std})
cv_res = cv_res.sort_values(['CrossValMeans'],ascending=False).reset_index(drop=True)
cv_res
# -
X1_train, X1_test, y_train, y_test = train_test_split(X1, y, test_size=0.4, random_state=0)
# +
f,ax=plt.subplots(3,4,figsize=(12,10))
acc=[]
model=svm.SVC()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,0],annot=True,fmt='d')
ax[0,0].set_title('SVC is: ')
model=DecisionTreeClassifier()
model.fit(X1_train,y_train)
y_pred =model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,1],annot=True,fmt='d')
ax[0,1].set_title('Decision Tree')
model=RandomForestClassifier(n_estimators=50)
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,2],annot=True,fmt='d')
ax[0,2].set_title('Random Forests')
model=KNeighborsClassifier(n_neighbors=2, p=2,weights='distance')
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[0,3],annot=True,fmt='d')
ax[0,3].set_title('KNN')
model=LogisticRegression()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,0],annot=True,fmt='d')
ax[1,0].set_title('Logistic Regression')
model=LDA()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,1],annot=True,fmt='d')
ax[1,1].set_title('LDA')
model=QDA()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,2],annot=True,fmt='d')
ax[1,2].set_title('QDA')
model=GaussianNB()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[1,3],annot=True,fmt='d')
ax[1,3].set_title('Naive Bayes')
model=pipeline
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,0],annot=True,fmt='d')
ax[2,0].set_title('Neural Network')
model=AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1)
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,1],annot=True,fmt='d')
ax[2,1].set_title('AdaBoosting')
model=GradientBoostingClassifier()
model.fit(X1_train,y_train)
y_pred = model.predict(X1_test)
acc.append(accuracy_score(y_test,y_pred))
sns.heatmap(confusion_matrix(y_test,y_pred),ax=ax[2,2],annot=True,fmt='d')
ax[2,2].set_title('Gradient Boosting')
plt.subplots_adjust(hspace=0.2,wspace=0.2)
plt.show()
# -
X11=X[['price_ratio','tweet_diff','stkv_ratio','twtcount_ratio','return_diff']]
C=range(10,300,10)
gamma=[1.0,1.1,1.2,1.3,1.4,1.5,1.6,1.7,1.8,1.9,2.0]
# kernel=['rbf','linear']
hyper={'C':C,'gamma':gamma}
gd=GridSearchCV(estimator=svm.SVC(),param_grid=hyper,verbose=True)
gd.fit(X11,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_estimators=range(10,100,10)
hyper={'n_estimators':n_estimators}
gd=GridSearchCV(estimator=RandomForestClassifier(random_state=0),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X11,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_neighbors=[2,3,4,5,6,7,8,9,10]
weights=['uniform','distance']
hyper={'n_neighbors':n_neighbors,'weights':weights}
gd=GridSearchCV(estimator=KNeighborsClassifier(),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X11,y)
print(gd.best_score_)
print(gd.best_estimator_)
# +
# aggregate all the methods in one plot
random_state = 4
classifiers = []
classifiers.append(svm.SVC(random_state=random_state,C=30,gamma=2.0))
classifiers.append(DecisionTreeClassifier(random_state=random_state))
classifiers.append(RandomForestClassifier(n_estimators=30,random_state=random_state))
classifiers.append(KNeighborsClassifier(n_neighbors=2, p=2,weights='uniform'))
classifiers.append(LogisticRegression(random_state = random_state))
classifiers.append(LDA())
classifiers.append(QDA())
classifiers.append(GaussianNB())
classifiers.append(MLPClassifier(random_state=random_state))
classifiers.append(AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1))
classifiers.append(GradientBoostingClassifier(random_state=random_state))
cv_results = []
for classifier in classifiers :
cv_results.append(cross_val_score(classifier, X11, y, scoring = "accuracy", cv = kfold))
cv_means = []
cv_std = []
for cv_result in cv_results:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
cv_res = pd.DataFrame({"Method":["SVC","DecisionTree",
"RandomForest","KNeighboors","LogisticRegression","LinearDiscriminantAnalysis","QuadraticDiscriminantAnalysis","NaiveBayes","NeuralNetwork",
"AdaBoosting","GradientBoosting"],"CrossValMeans":cv_means,"CrossValerrors": cv_std})
cv_res = cv_res.sort_values(['CrossValMeans'],ascending=False).reset_index(drop=True)
cv_res
# -
# ## Fix this plot
# +
# model=RandomForestClassifier(n_estimators=500,random_state=0)
# model.fit(X,y)
# model.feature_importances_
# X.columns
# imp = pd.DataFrame({"Features":["UA Customer Count","Nike Customer Count","UA Stock Price","Nike Stock Price","UA Stock Volume","Nike Stock Volume",
# "UA Tweet Attitude","Nike Tweet Attitude","UA Tweet Count","Nike Tweet Count","Geolocation Ratio","Stock Price Ratio","Tweet Attitude Difference",
# "Stock Volume Ratio","Tweet Count Ratio","UA Stock Return","Nike Stock Return","Stock Return Difference"],
# "Importance":model.feature_importances_})
# imp = imp.sort_values(['Importance'],ascending=False).reset_index(drop=True).head(5)
features=["UA Customer Count","Nike Customer Count","UA Stock Price","Nike Stock Price","UA Stock Volume","Nike Stock Volume",
"UA Tweet Attitude","Nike Tweet Attitude","UA Tweet Count","Nike Tweet Count","Geolocation Ratio","Stock Price Ratio","Tweet Attitude Difference",
"Stock Volume Ratio","Tweet Count Ratio","UA Stock Return","Nike Stock Return","Stock Return Difference"]
# pd.Series(model.feature_importances_,features).sort_values(ascending=True).tail(5)
# pd.Series(model.feature_importances_,features).sort_values(ascending=True).plot.barh(width=0.8,ax=ax[0,1],color='#ddff11')
# imp
# -
plt.text(weight='bold')
plt.rc('font', weight='bold')
# +
# tree-based feature selection
f,ax=plt.subplots(2,1,figsize=(10,12))
model=RandomForestClassifier(n_estimators=500,random_state=0)
model.fit(X,y)
pd.Series(model.feature_importances_,features).sort_values(ascending=True).tail(5).plot.barh(width=0.6,ax=ax[0],color='salmon')
ax[0].set_title('Feature Importance in Random Forests',fontsize=20)
#ax[0].set_label(fontsize=30)
ax[0].tick_params(axis='y', rotation=30)
model=AdaBoostClassifier(n_estimators=200,learning_rate=0.05,random_state=0)
model.fit(X,y)
pd.Series(model.feature_importances_,features).sort_values(ascending=True).tail(5).plot.barh(width=0.6,ax=ax[1],color='blue')
#pd.Series(model.feature_importances_,X.columns).sort_values(ascending=True).plot.barh(width=0.8,ax=ax[0,1],color='#ddff11')
ax[1].set_title('Feature Importance in AdaBoost',fontsize=20)
ax[1].tick_params(axis='y', rotation=30)
# model=GradientBoostingClassifier(n_estimators=500,learning_rate=0.1,random_state=0)
# model.fit(X,y)
# pd.Series(model.feature_importances_,features).sort_values(ascending=True).tail(5).plot.barh(width=0.6,ax=ax[1,0],cmap='RdYlGn_r')
# #pd.Series(model.feature_importances_,X.columns).sort_values(ascending=True).plot.barh(width=0.8,ax=ax[1,0],cmap='RdYlGn_r')
# ax[1,0].set_title('Feature Importance in Gradient Boosting')
# model=xg.XGBClassifier(n_estimators=900,learning_rate=0.1)
# model.fit(X,y)
# pd.Series(model.feature_importances_,features).sort_values(ascending=True).tail(5).plot.barh(width=0.6,ax=ax[1,1],color='#FD0F00')
# #pd.Series(model.feature_importances_,X.columns).sort_values(ascending=True).plot.barh(width=0.8,ax=ax[1,1],color='#FD0F00')
# ax[1,1].set_title('Feature Importance in XgBoost')
#plt.yticks(rotation=45)
#plt.show()
# -
X2=X[['nike_tweet','ua_stkv','geo_count_ratio','price_ratio','return_diff']]
X2=X[['return_ua','nike_tweet','geo_count_ratio','nike_price','geo_nike_count']]
C=range(10,300,10)
gamma=[1.0,1.1,1.2,1.3,1.4,1.5,1.6,1.7,1.8,1.9,2.0]
# kernel=['rbf','linear']
hyper={'C':C,'gamma':gamma}
gd=GridSearchCV(estimator=svm.SVC(),param_grid=hyper,verbose=True)
gd.fit(X2,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_estimators=range(10,100,10)
hyper={'n_estimators':n_estimators}
gd=GridSearchCV(estimator=RandomForestClassifier(random_state=0),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X2,y)
print(gd.best_score_)
print(gd.best_estimator_)
n_neighbors=[2,3,4,5,6,7,8,9,10]
weights=['uniform','distance']
hyper={'n_neighbors':n_neighbors,'weights':weights}
gd=GridSearchCV(estimator=KNeighborsClassifier(),param_grid=hyper,cv=3, scoring="accuracy",verbose=1)
gd.fit(X2,y)
print(gd.best_score_)
print(gd.best_estimator_)
# +
# aggregate all the methods in one plot
random_state = 4
classifiers = []
classifiers.append(svm.SVC(random_state=random_state))
classifiers.append(DecisionTreeClassifier(random_state=random_state))
classifiers.append(RandomForestClassifier(n_estimators=50,random_state=random_state))
classifiers.append(KNeighborsClassifier(n_neighbors=4, p=2,weights='uniform'))
classifiers.append(LogisticRegression(random_state = random_state))
classifiers.append(LDA())
classifiers.append(QDA())
classifiers.append(GaussianNB())
classifiers.append(MLPClassifier(random_state=random_state))
classifiers.append(AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1))
classifiers.append(GradientBoostingClassifier(random_state=random_state))
cv_results = []
for classifier in classifiers :
cv_results.append(cross_val_score(classifier, X2, y, scoring = "accuracy", cv = kfold))
cv_means = []
cv_std = []
for cv_result in cv_results:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
cv_res = pd.DataFrame({"Method":["SVC","DecisionTree",
"RandomForest","KNeighboors","LogisticRegression","LinearDiscriminantAnalysis","QuadraticDiscriminantAnalysis","NaiveBayes","NeuralNetwork",
"AdaBoosting","GradientBoosting"],"CrossValMeans":cv_means,"CrossValerrors": cv_std})
cv_res = cv_res.sort_values(['CrossValMeans'],ascending=False).reset_index(drop=True)
cv_res
# -
# #### RFE
#RFE with logistic regression
from sklearn.feature_selection import RFE
logreg = LogisticRegression()
rfe = RFE(estimator=logreg, n_features_to_select=5, step=1)
rfe = rfe.fit(X,y)
print('Chosen best 5 feature by rfe:',X_train.columns[rfe.support_])
X3=X[['geo_count_ratio', 'tweet_diff', 'stkv_ratio', 'twtcount_ratio',
'return_ua']]
# +
# X3=X[['geo_count_ratio']]
# -
score=cross_val_score(LogisticRegression(random_state = random_state), X3, y, scoring = "accuracy", cv = kfold)
score.mean()
from sklearn import preprocessing
scaler = preprocessing.StandardScaler().fit(X3)
X3 = scaler.transform(X3)
#####standardization
# ## Add name to this
import statsmodels.api as sm
logit_model=sm.Logit(y,X3)
result=logit_model.fit()
print(result.summary())
| Code_2019.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="OipjYZb__cCl"
# # ジンバルロック
# + [markdown] id="o0nn7eS7_cCn"
# 吉田勝俊(宇都宮大学)
# + [markdown] id="Dmk-Lck2_cCo"
# ## 参考情報
# - [ipywidgetsでインタラクティブなグラフを作る - Qiita](https://qiita.com/studio_haneya/items/adbaa01b637e7e699e75)
# - [Widget List — Jupyter Widgets 8.0.0a5 documentation](https://ipywidgets.readthedocs.io/en/latest/examples/Widget%20List.html)
# + id="jJCyzw0mAvRW"
import numpy as np #数値計算ライブラリ
from math import sin, cos, pi #低機能だが計算が速い数学関数
import matplotlib.pyplot as plt #描画ライブラリ
from mpl_toolkits.mplot3d import Axes3D #3次元座標用のモジュール
from mpl_toolkits.mplot3d.art3d import Poly3DCollection #3次元ポリゴン用のモジュール
import ipywidgets as ipw #対話的処理のモジュール
#↓Colab用の設定(グラフィックのインライン表示)
# %matplotlib inline
# + [markdown] id="NrJ6txkg9ArA"
# ## 3次元の回転行列
# + [markdown] id="ajmzKGDgctPX"
# ### 基本となる回転行列(回転軸$\boldsymbol{a}$,角度$\theta$)
# + id="wAmU2E10ctPY"
def Rotation(angle, axis):
a1, a2, a3 = axis #回転軸の成分
aa1 = a1*a1 #2乗を作っておく
aa2 = a2*a2
aa3 = a3*a3
C = cos(angle)
S = sin(angle)
R = np.array([
[ aa1 + (1-aa1)*C, a1*a2*(1-C) - a3*S, a1*a3*(1-C) + a2*S ],
[ a1*a2*(1-C) + a3*S, aa2 + (1-aa2)*C, a2*a3*(1-C) - a1*S ],
[ a1*a3*(1-C) - a2*S, a2*a3*(1-C) + a1*S, aa3 + (1-aa3)*C ],
])
return R
# + [markdown] id="f5wlC7rteGk-"
# ### オイラー角(XYZ型)
# + [markdown] id="ielCrW7HeMww"
# 各軸回転の行列を作っておく.
# + id="RGSJRlP98N6s"
def Rx(angle):
return Rotation(angle,[1,0,0])
def Ry(angle):
return Rotation(angle,[0,1,0])
def Rz(angle):
return Rotation(angle,[0,0,1])
Rx(0.5), Ry(0.5), Rz(0.5)
# + [markdown] id="KS_JUodEfSJQ"
# XYZ型オイラー角による回転行列を作る.
# + id="kn-tru6RffUv"
def Rxyz(angles):
th1, th2, th3 = angles
return Rz(th3).dot( Ry(th2) ).dot( Rx(th1) )
Rxyz( [0.5, 0.5, 0.5] )
# + [markdown] id="ZEvpQBm2AvRb"
# ### ポリゴンデータの操作
# + id="SxQqrEeAAvRb"
def Rotate_polys(R, polys):
'''
回転行列 R を,ポリゴンのリスト polys に掛ける
'''
POLYs = []
for poly in polys: #ポリゴン1枚の頂点セット in そのリスト
POLY = []
for point in poly: #各頂点 in あるポリゴンの頂点セット
newpoint = R.dot(point) #回転変換
POLY.append(newpoint) #POLYに格納
POLYs.append(POLY) #POLYSに格納
return POLYs
def plot_polys(ax, polys):
'''
ポリゴンのリスト polys をプロットする
'''
facecolors=['C1','C2','C6','C6','C3'] #角面の色
for i, p in enumerate(polys):
pc = Poly3DCollection([p])
pc.set_alpha(0.7) #alpha first
pc.set_facecolor(facecolors[i])
ax.add_collection3d(pc)
ax.set_xlim(-1,1)
ax.set_ylim(-1,1)
ax.set_zlim(-1,1)
# ax.set_xlabel('X', labelpad=5)
# ax.set_ylabel('Y', labelpad=5)
# ax.set_zlabel('Z', labelpad=0)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
# + [markdown] id="53QOHWLGAvRd"
# ### ポリゴンのデータサンプル
# + id="1M7gQWCgAvRd"
def create_triangle_polys(w, d, h):
'''
厚みのある二等辺三角形 (幅, 厚, 高)
'''
# 頂点
A = np.array([ w/2, -d/2, 0])
B = np.array([ w/2, d/2, 0])
C = np.array([-w/2, d/2, 0])
D = np.array([-w/2, -d/2, 0])
E = np.array([ 0, -d/2, h])
F = np.array([ 0, d/2, h])
# ポリゴンのリスト
polys =[
[F, B, C], #前面ポリゴン
[E, D, A], #後面ポリゴン
[A, B, F, E], #斜面ポリゴン
[C, D, E, F], #斜面ポリゴン
[A, B, C, D], #底面ポリゴン
]
return polys
# + [markdown] id="WJU5-TfgAvRe"
# ### お試しプロット
# + id="wIyoSh2PAvRf"
# 基準姿勢データのサンプル
body = create_triangle_polys(w=1, d=0.1, h=0.5)
# オイラー角による回転行列 R のサンプル
R = Rxyz( [pi/2, 0, 0] ) #X軸まわりにだけ回転
# 基準姿勢データを回転変換
newbody = Rotate_polys(R, body)
print(body)
print(newbody)
fig = plt.figure(figsize=(5,5)) #正方形のキャンバスに,
ax = fig.add_subplot(111, projection='3d') #3次元グラフ用紙を作る
plot_polys(ax, body)
plot_polys(ax, newbody)
fig.tight_layout()
plt.show()
# + [markdown] id="_UY79BuqEptw"
# ## 手動操作
# + [markdown] id="J3b_tHjysCjQ"
# 【注意】簡易的な実装のため,滑らかには動きません.
# + id="vQr9yjvGy5fZ"
def draw_body(th1deg, th2deg, th3deg):
'''
オイラー角の姿勢で物体を描く関数
'''
fig = plt.figure(figsize=(5,5)) #正方形のキャンバスに,
ax = fig.add_subplot(111, projection='3d') #3次元グラフ用紙を作る
ax.set_title('angles = (%d, %d, %d)'%(th1deg, th2deg, th3deg))
angles_rad = np.array([th1deg, th2deg, th3deg])/180*pi
# オイラー角による回転行列 R
R = Rxyz( angles_rad )
# 三角形のポリゴンデータ
body = create_triangle_polys(w=1, d=0.1, h=0.5)
# 三角形をオイラー角で回す
newbody = Rotate_polys( R, body )
# 描画する
plot_polys(ax, newbody)
fig.tight_layout()
plt.show()
### UI(ユーザーインターフェイス)を作る
#スライダの定義
slider_th1 = ipw.IntSlider(description='th1',
min=-180, max=180, step=10, value=0) #下限,上限,刻み,初期値
slider_th2 = ipw.IntSlider(description='th2',
min=-180, max=180, step=10, value=0)
slider_th3 = ipw.IntSlider(description='th3',
min=-180, max=180, step=10, value=0)
#スライダを縦に並べるUI
ui = ipw.VBox([slider_th1, slider_th2, slider_th3])
#スライダの値でグラフを更新するオブジェクト
out = ipw.interactive_output(
draw_body,
{
'th1deg': slider_th1,
'th2deg': slider_th2,
'th3deg': slider_th3,
}
)
out.layout.height = '350px' #サイズ固定で若干チラツキが緩和
out.layout.width = '350px'
display(ui, out) #UIとグラフを出力
# + [markdown] id="flxengt56HGt"
# - `th2deg = 90` に合わせると,ジンバルロックが発生します.
# - このとき,`th1deg` と `th3deg` は,効き方が同じになります.
# + id="1iDxOCzp0abS"
| m3d/Colab/Python_5.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from __future__ import print_function, absolute_import, division
import tensorflow as tf
print(tf.__version__)
tf.test.is_gpu_available()
from tensorflow.estimator import RunConfig
from training_functions import make_tfr_input_fn
# # Using a Tensorflow Estimator
# As you can see, the ```Estimator``` is the central working horse of ML Engineering. We'll have to provide it with
# - a model function: The model function creates appropriate versions of the hypothesis together with some parameters and the tools to evaluate and train the model.
# - The model function returns ```EstimatorSpec```s for the different phases of the ML lifecycle
# - ```EvalSpec``` and ```TrainSpec``` objects that determine the physical characteristics of the training and evaluation phases.
# - a ```RunConfig``` that essentially describes the execution environment.
#
# After that, the estimator performs all steps independently, creates logfiles, safe-points, performance metrics, and the entire model update life cycle. At the end we get to a model that we can use for prediction.
# ### Configuration
# +
import tempfile
temp_dir = tempfile.gettempdir()
import os
file_pattern = os.path.join(temp_dir, "training.tfr-*")
file_pattern
training_pattern = os.path.join(temp_dir, "training.tfr-*")
eval_pattern = os.path.join(temp_dir, "eval.tfr-*")
# remove this directory to start from scratch
model_dir = os.path.join(temp_dir, "models" )
# -
training_pattern = 'self_play_data/*-*.tfr'
eval_pattern = 'deleteme.tfr'
![ -d $model_dir ] && echo "Really delete $model_dir?"
# Uncomment if you really want to delete the model and start from scratch
# !rm -rf $model_dir
# +
training_options={
'num_epochs': None, # repeat infinitely
'shuffle_buffer_size': 1000,
'prefetch_buffer_size': 1000,
'reader_num_threads': 10,
'parser_num_threads': 10,
'sloppy_ordering': True,
'distribute': False}
eval_options={
'num_epochs': None, # repeat infinitely
'shuffle_buffer_size': 100,
'prefetch_buffer_size': 10,
'reader_num_threads': 10,
'parser_num_threads': 10,
'sloppy_ordering': True,
'distribute': False}
test_options={
'num_epochs': None, # repeat infinitely
'shuffle_buffer_size': 1000,
'prefetch_buffer_size': 1000,
'reader_num_threads': 10,
'parser_num_threads': 10,
'sloppy_ordering': True,
'distribute': False}
# -
# ### Input Functions
# +
train_input_fn = make_tfr_input_fn(
filename_pattern=training_pattern,
batch_size=256,
options = training_options)
eval_input_fn = make_tfr_input_fn(
filename_pattern=eval_pattern,
batch_size=80,
options = eval_options)
# -
# ### The model_function
# The model function provides ```EstimatorSpec```s, i.e. specifications how to build the model for each of the different cases: training, evaluation and test. Indeed, some models require the actual function to differ slightly between training and evaluation. The model function is the place to specify what exactly is to be calculated during each phase of the ML process. In our case, though, all specifications are essentially the same. Typically, you'd expect the *data scientist* to provide this function, so it's not so important that you fully understand the concept here.
class ResNet:
"""
After sufficient training, this instance of ResNet takes an array of dimensions 10x10
and returns 1 if the array contains the pattern you tought it to recognize.
"""
def __init__(self, size, n_blocks, n_filters, inps=None, lbls=None):
if inps == None:
inps = tf.placeholder(name="inp_resnet",
shape=[None, size+2, size+2, 2],
dtype=tf.float32)
if lbls == None:
lbls = tf.placeholder(name="lbl_resnet",
shape=[None, size+2, size+2, 1],
dtype=tf.float32)
self.inps = inps
self.lbls = lbls
out = self.inps
for i in range(n_blocks):
out = self._res_block(out, n_filters)
self.out = tf.layers.conv2d(kernel_size=1, filters=1, inputs=out,
padding='same', activation=None)
# TODO: Just wouldn't work, minimizes until a certain point then losses go up again
#lbls_1dim = tf.reshape(self.lbls, [-1, 21*21, 1])
#outs_1dim = tf.reshape(self.out, [-1, 21*21, 1])
#losses = tf.nn.softmax_cross_entropy_with_logits_v2(lbls_1dim, outs_1dim, axis=1)
#self.loss = tf.reduce_sum(losses)
self.loss = tf.losses.mean_squared_error(self.lbls, self.out)
self.errors = tf.losses.mean_squared_error(self.lbls, self.out)
self.accuracy=tf.reduce_sum(tf.cast(self.errors < .1, dtype=tf.int64))
self.optimizer = tf.train.AdamOptimizer(learning_rate=1e-3)
self.trainer = self.optimizer.minimize(self.loss)
def _res_block(self, inp, filters, activation='relu'):
out1_3 = tf.layers.conv2d(kernel_size=3, filters=filters, inputs=inp,
padding='same', activation=activation)
out1_5 = tf.layers.conv2d(kernel_size=5, filters=filters, inputs=inp,
padding='same', activation=activation)
out1 = tf.concat([out1_3, out1_5], axis=3)
out2_3 = tf.layers.conv2d(kernel_size=3, filters=filters, inputs=out1,
padding='same', activation=activation)
out2_5 = tf.layers.conv2d(kernel_size=5, filters=filters, inputs=out1,
padding='same', activation=activation)
out2 = tf.concat([out2_3, out2_5], axis=3)
out3_3 = tf.layers.conv2d(kernel_size=3, filters=filters, inputs=out2,
padding='same', activation=activation)
out3_5 = tf.layers.conv2d(kernel_size=5, filters=filters, inputs=out2,
padding='same', activation=activation)
out3 = tf.concat([out3_3, out3_5], axis=3)
bn = tf.layers.batch_normalization(inputs=out3)
skip = tf.layers.conv2d(kernel_size=1, filters=2, inputs=inp,
padding='same', activation=None)
return tf.concat([skip, bn], axis=3)
features, labels = train_input_fn()
features
resnet = ResNet(19, n_blocks=10, n_filters=32, inps=features['state'], lbls=labels)
# +
def model_function(features, labels, mode):
my_input_layer = features['state']
resnet = ResNet(19, n_blocks=2, n_filters=32, inps=my_input_layer, lbls=labels)
hypothesis = resnet.out
#
# For predictions, we just need the hypothesis.
#
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(
tf.estimator.ModeKeys.PREDICT,
predictions=hypothesis)
#
# For evaluation, we need to provide the loss function, too.
#
#loss = tf.losses.mean_squared_error(labels, hypothesis)
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(
tf.estimator.ModeKeys.EVAL,
loss = resnet.loss)
#
# And for training, we also need the optimizer
#
train_op = resnet.trainer
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
optimizer = tf.train.AdamOptimizer(1e-3)
#train_op = optimizer.minimize(loss)
global_step = tf.train.get_global_step()
update_global_step = tf.assign(global_step, global_step + 1,
name = 'update_global_step')
return tf.estimator.EstimatorSpec(
tf.estimator.ModeKeys.TRAIN,
loss = resnet.loss,
train_op = tf.group(train_op, update_global_step))
# -
# ### Serving Input Receiver
# This function returns a function that is going to be called by the estimator to create a ServingInputReciever. Sounds odd, but is pretty straight-forward. First, we provide a function that will return a tensor. We don't provide the tensor, because the tensor will have to be created in the context (graph and session) of the estimator methods. We use a function to create a function because we're passing a parameter that's necessary but not available to the estimator at runtime. Fine. But why are we doing that, anyway?
#
# Remember the scaling of the $\beta$s that we performed with our Beam pipeline. We saved the transform function as the last step of the pipeline. Here, we dig it out again and provide it to the estimator so it can attach it to the front of its computational graph such that the same scaling is applied to the *signature* data.
import tensorflow_transform as tft
def make_tft_serving_input_fn(metadata_dir):
def _input_fn():
# This is what signature data looks like: no feature cross yet
placeholders = {
'state': tf.placeholder(name='state', shape=[None, 21, 21, 2], dtype=tf.float32)
}
transform_output = tft.TFTransformOutput(transform_output_dir=metadata_dir)
features = placeholders #transform_output.transform_raw_features(placeholders)
return tf.estimator.export.ServingInputReceiver(features, placeholders)
return _input_fn
# ### The Estimator
config = RunConfig(
model_dir = model_dir,
save_summary_steps = 20000,
save_checkpoints_steps = 20000,
log_step_count_steps = 1000)
# !mkdir /tmp/metadata
metadata_dir = os.path.join(temp_dir, 'metadata')
# !ls $metadata_dir
serving_input_fn = make_tft_serving_input_fn(model_dir)
exporter = tf.estimator.LatestExporter('exporter', serving_input_fn)
estimator = tf.estimator.Estimator(
config=config,
model_fn=model_function)
# +
max_steps = (
5000 * # total number of records
5000 / # number of epochs I want for training
256 # batch size
)
print("Training up to %s steps now..." % max_steps)
train_spec = tf.estimator.TrainSpec(
input_fn=train_input_fn,
max_steps=max_steps)
eval_spec = tf.estimator.EvalSpec(
input_fn=eval_input_fn, exporters=exporter,
steps = 2, # 2 batches for evaluation
throttle_secs=1, # technical stuff - don't bother
start_delay_secs=0)
# -
tf.estimator.train_and_evaluate(
estimator,
train_spec=train_spec,
eval_spec=eval_spec)
import os
model_dir = os.path.join("/tmp/", "models/export/exporter")
# versions = !ls $model_dir
print( "Versions: %s" % versions)
latest_version = max(versions)
latest_model = os.path.join(model_dir, str(latest_version))
# !echo $latest_model
# !ls $latest_model
estimator = tf.contrib.predictor.from_saved_model(latest_model)
def _parse_function(example):
return tf.parse_single_example(example, feature_spec)
dataset = tf.data.TFRecordDataset("deleteme.tfr")
N_p=21
feature_spec = {
'state': tf.FixedLenFeature([N_p * N_p * 2], tf.float32),
'distr': tf.FixedLenFeature([N_p * N_p], tf.float32)
}
decoded = dataset.map(_parse_function).make_one_shot_iterator().get_next()
with tf.Session() as sess:
res2 = sess.run(decoded)
res2['state'].shape, res2['distr'].shape
from wgomoku import to_matrix12
sample=res2['state'].reshape(1,21,21,2)
m12 = to_matrix12(sample, 21)
m12
distr = estimator({'state': sample})
predi = distr['output'].reshape(21,21)[1:-1].T[1:-1].T
predi.shape
from wgomoku import (
GomokuBoard,
HeuristicGomokuPolicy, Heuristics, GomokuTools as gt)
size=19
p = HeuristicGomokuPolicy(style = 2, bias=.5, topn=5)
heuristics = Heuristics(kappa=3.0)
# +
import numpy as np
black = [gt.m2b(p, 19) for p, v in list(np.ndenumerate(m12)) if v == 1]
white = [gt.m2b(p, 19) for p, v in list(np.ndenumerate(m12)) if v == 2]
pairs = zip(black, white)
moves = []
for b,w in pairs:
moves.append(b)
moves.append(w)
board = GomokuBoard(N=19, stones=moves, heuristics=heuristics,
disp_width=10)
board.display(probas=lambda x: predi)
# -
from wgomoku import create_sample
smp = create_sample(board.stones, 19, 1-board.current_color)
smp.shape
sample = smp.reshape(1, 21, 21, 2)
distr = estimator({'state': sample})
def ai_policy(board):
smp = create_sample(board.stones, 19, 1-board.current_color)
sample = smp.reshape(1, 21, 21, 2)
pred = estimator({'state': sample})['output'][0].reshape(21,21)
return pred[1:-1].T[1:-1].T
ai_policy(board).shape
A,B,C,D,E,F,G,H,I,J,K,L,M,N,O,P,Q,R,S,T,U = \
1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21
BLACK=0
WHITE=1
EDGES=2
board.undo().undo().undo().undo().undo().undo().undo().undo(True);
board.display(probas = ai_policy)
from wgomoku import StochasticMaxSampler
sampler.choices[0][1]
sampler = StochasticMaxSampler(np.ndenumerate(ai_policy(board)), bias=8, topn=5)
rc = sampler.choices[0][1]
x, y = gt.m2b(rc, 19)
print(chr(64+x), y)
board.set(x, y);
board.display(probas = ai_policy)
board.set(H,8);
board.undo()
board.display(probas = ai_policy)
sampler = StochasticMaxSampler(np.ndenumerate(ai_policy(board)), bias=8, topn=5)
print(sampler.choices)
| Estimator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd #数据处理库
import numpy as np #数据处理库
import seaborn as sns #数据图像展示库
import matplotlib.pyplot as plt #数据图像展示库
# 导入第三方包,方便使用
# +
# 生成随机数展示
x=np.linspace(-10,10,50) #定义数据范围,起始值,终止值,样本个数
y=2*x+1 #定义y数据范围
y1=x**2
plt.figure(num=1,figsize=(8,5)) #定义一个图像窗口,编号为 1,大小为(8,5)
plt.plot(x,y,color='red',linewidth=4,linestyle='--') #plot()画出图像,颜色为红色,线宽度为4,线风格为 --
plt.plot(x,y1) #plot()画出曲线
plt.xlim(-9,10) #截取x坐标范围
plt.ylim(-18,80) #截取y坐标范围
plt.xlabel("I'm x") #设置x轴坐标
plt.ylabel("I'm y") #设置y轴坐标
plt.show() #显示图像
# -
# 设置边框属性,移动坐标轴
x=np.linspace(-10,10,50)
y1=2*x+1
y2=x**2
plt.figure(num=2,figsize=(8,5))
plt.plot(x,y1,color='red',linewidth=2,linestyle='--')
plt.plot(x,y2) #进行画图
new_ticks=np.linspace(-10,10,5) #小标从-10到10分为5个单位
plt.xticks(new_ticks) #进行替换新下标
plt.yticks([-0,20,60,80,],
[r'$really\ bad$','$bad$','$well$','$really\ well$'])
ax=plt.gca() #gca=get current axis
ax.spines['right'].set_color('none') #边框属性设置为none 不显示
ax.spines['top'].set_color('none')
#使用xaxis.set_ticks_position设置x坐标刻度数字或名称的位置 所有属性为top、bottom、both、default、none
ax.xaxis.set_ticks_position('bottom')
#使用.spines设置边框x轴;使用.set_position设置边框位置,y=0位置 位置所有属性有outward、axes、data
ax.spines['bottom'].set_position(('data', 0))
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data',0)) #坐标中心点在(0,0)位置
plt.show()
# 添加图例
x=np.linspace(-10,10,50)
y1=2*x+1
y2=x**2
plt.figure(num=2,figsize=(8,5))
plt.plot(x,y1,color='red',linewidth=2,linestyle='--')
plt.plot(x,y2) #进行画图
new_ticks=np.linspace(-10,10,5) #小标从-10到10分为5个单位
plt.xticks(new_ticks) #进行替换新下标
plt.yticks([-0,20,60,80,],
[r'$really\ bad$','$bad$','$well$','$really\ well$'])
l1,=plt.plot(x,y1,color='red',linewidth=2,linestyle='--',label='linear line')
l2,=plt.plot(x,y2,label='square line') #进行画图
#loc有很多参数 其中best自分配最佳位置
'''
'best' : 0,
'upper right' : 1,
'upper left' : 2,
'lower left' : 3,
'lower right' : 4,
'right' : 5,
'center left' : 6,
'center right' : 7,
'lower center' : 8,
'upper center' : 9,
'center' : 10,
'''
# plt.legend(loc='best') #显示在最好的位置
plt.legend(handles=[l1, l2], labels=['square', 'linear'], loc='best')
plt.show() #显示图
# +
x=np.linspace(-10,10,50)
y = 2*x + 1
plt.figure(num=1, figsize=(8, 5))
plt.plot(x, y,)
#移动坐标轴
ax = plt.gca()
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.spines['bottom'].set_position(('data', 0))
ax.yaxis.set_ticks_position('left')
ax.spines['left'].set_position(('data', 0))
#标注信息
x0=3
y0=2*x0+1
plt.scatter(x0,y0,s=50,color='b')
#连接(x0,y0)(x0,0) k表示黑色 lw=2.5表示线粗细
plt.plot([x0,x0],[y0,0],'k--',lw=1)
#xycoords='data'是基于数据的值来选位置,xytext=(+30,-30)和textcoords='offset points'
#对于标注位置描述和xy偏差值,arrowprops对图中箭头类型设置
plt.annotate(r'$2x0+1=%s$' % y0, xy=(x0, y0), xycoords='data', xytext=(+30, -30),
textcoords='offset points', fontsize=16,
arrowprops=dict(arrowstyle='->', connectionstyle="arc3,rad=.2"))
#添加注视text(-3.7,3)表示选取text位置 空格需要用\进行转译 fontdict设置文本字体
plt.text(-3.7, 3, r'$This\ is\ the\ some\ text. \mu\ \sigma_i\ \alpha_t$',
fontdict={'size': 18, 'color': 'r'})
plt.show()
# -
# Scatter散点图
n=100
x=np.random.normal(0,1,n) #每一个点的X值
y=np.random.normal(0,1,n) #每一个点的Y值
T=np.arctan2(y,x) #arctan2返回给定的X和Y值的反正切值
#scatter画散点图 size=75 颜色为T 透明度为50% 利用xticks函数来隐藏x坐标轴
plt.scatter(x,y,s=75,c=T,alpha=0.5)
plt.xlim(-1.5,1.5)
plt.xticks(()) #忽略xticks
plt.ylim(-1.5,1.5)
plt.yticks(()) #忽略yticks
plt.show()
# +
slices = [7,2,2,13]
activities = ['sleeping','eating','working','playing']
cols = ['c','m','r','b']
plt.pie(slices,
labels=activities,
colors=cols,
startangle=90,
shadow= True,
explode=(0,0.1,0,0),
autopct='%1.1f%%')
plt.title('Interesting Graph\nCheck it out')
plt.show()
# +
#条形图
n=8
X=np.arange(n)
Y1=(1-X/float(n))*np.random.uniform(0.5,1,n)
Y2=(1-X/float(n))*np.random.uniform(0.5,1,n)
plt.bar(X,+Y1,facecolor='#9999ff',edgecolor='white')
plt.bar(X,-Y2,facecolor='#ff9999',edgecolor='white')
#标记值
for x,y in zip(X,Y1): #zip表示可以传递两个值
plt.text(x+0.4,y+0.05,'%.2f'%y,ha='center',va='bottom') #ha表示横向对齐 bottom表示向下对齐
for x,y in zip(X,Y2):
plt.text(x+0.4,-y-0.05,'%.2f'%y,ha='center',va='top')
plt.xlim(-0.5,n)
plt.xticks(()) #忽略xticks
plt.ylim(-1.25,1.25)
plt.yticks(()) #忽略yticks
plt.show()
# -
# 等高线图
n=256
x=np.linspace(-3,3,n)
y=np.linspace(-3,3,n)
X,Y=np.meshgrid(x,y) #meshgrid从坐标向量返回坐标矩阵
#f函数用来计算高度值 利用contour函数把颜色加进去 位置参数依次为x,y,f(x,y),透明度为0.75,并将f(x,y)的值对应到camp之中
def f(x,y):
return (1 - x / 2 + x ** 5 + y ** 3) * np.exp(-x ** 2 - y ** 2)
plt.contourf(X,Y,f(X,Y),8,alpha=0.75,cmap=plt.cm.hot) #8表示等高线分成多少份 alpha表示透明度 cmap表示color map
#使用plt.contour函数进行等高线绘制 参数依次为x,y,f(x,y),颜色选择黑色,线条宽度为0.5
C=plt.contour(X,Y,f(X,Y),8,colors='black')
#使用plt.clabel添加高度数值 inline控制是否将label画在线里面,字体大小为10
plt.clabel(C,inline=True,fontsize=10)
# plt.xticks(()) #隐藏坐标轴
# plt.yticks(())
plt.show()
# +
# 多图合并显示
plt.figure()
plt.subplot(2,2,1) #表示整个图像分割成2行2列,当前位置为1
plt.plot([0,1],[0,1]) #横坐标变化为[0,1] 竖坐标变化为[0,2]
plt.subplot(2,3,4) #将整个窗口分割成2行3列,当前位置为4
plt.plot([0,1],[0,2])
plt.subplot(2,3,5)
plt.plot([0,1],[0,3])
plt.subplot(2,3,6)
plt.plot([0,1],[0,4])
plt.show()
# +
# 图中图
fig=plt.figure()
#创建数据
x=[1,2,3,4,5,6,7,8,9,10]
y=[1,3,4,2,6,8,5,7,10,9]
#绘制大图:假设大图的大小为10,那么大图被包含在由(1,1)开始,宽8高8的坐标系之中。
left, bottom, width, height = 0.1, 0.1, 0.8, 0.8
ax1 = fig.add_axes([left, bottom, width, height]) # main axes
ax1.plot(x, y, 'r') #绘制大图,颜色为red
ax1.set_xlabel('x') #横坐标名称为x
ax1.set_ylabel('y')
ax1.set_title('total')#图名称为 total
#绘制小图,注意坐标系位置和大小的改变
ax2 = fig.add_axes([0.2, 0.6, 0.25, 0.2])
ax2.plot(y, x, 'b') #颜色为buue
ax2.set_xlabel('x')
ax2.set_ylabel('y')
ax2.set_title('inside 1')
#绘制第二个小兔
plt.axes([0.6, 0.2, 0.25, 0.25])
plt.plot(y[::-1], x, 'g') #将y进行逆序
plt.xlabel('x')
plt.ylabel('y')
plt.title('inside 2')
plt.show()
# -
# 混合图形
fig, ax = plt.subplots()
ax.plot([1, 2, 3, 4], [10, 16, 26, 32], label='cd')
ax.plot([1, 2, 3, 4], [30, 24, 10,6], label='xa')
ax.scatter([1, 2, 3, 4], [18,6, 28, 16], label='Point')
ax.set(ylabel='Temperature(deg C)', xlabel='Time', title='The temperature of the two cities')
ax.legend()
plt.show()
# +
# 多条形图
np.random.seed(19680801)
n_bins = 6
x = np.random.randn(1000, 3)
fig, axes = plt.subplots(nrows=2, ncols=2)
ax0, ax1, ax2, ax3 = axes.flatten()
colors = ['red', 'tan', 'lime']
ax0.hist(x, n_bins, density=True, histtype='bar', color=colors, label=colors)
ax0.legend(prop={'size': 10})
ax0.set_title('bars with legend')
ax1.hist(x, n_bins, density=True, histtype='barstacked',rwidth=1.0)
ax1.set_title('stacked bar')
ax2.hist(x, histtype='barstacked', rwidth=0.8)
ax2.set_title('stacked bar')
ax3.hist(x[:, 0], rwidth=0.6)
ax3.set_title('different sample sizes')
fig.tight_layout()
plt.show()
# -
# 热力图
# Create a random dataset
data = pd.DataFrame(np.random.random((10,6)), columns=["SH","CD","XA","GY","GZ", "SZ"])
print(data)
# Plot the heatmap
heatmap_plot = sns.heatmap(data, center=0, cmap='gist_ncar')
plt.show()
| python-base/jupyter/numpy_matplotlib_learn/matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Vector Operations
import numpy as np
A = np.array([1+1j, 2+2j, 3+3j, 4+4j, 5+5j])
B = np.array([6-6j, 7-7j, 8-8j, 9-9j, 10-10j])
C = np.array([2,3,5,7,11])
Z = np.array([0,0,0,0,0])
A
B
C
Z
# ## Operations
# ### Equality
A == A
np.array_equal(A, A)
np.array_equal(A, B)
# ### Addition
A + B
A + (B + C) == (A + B) + C
A == A + 0
np.array_equal(
A + B,
[ A[n] + B[n] for n in range(0,len(A)) ]
)
np.array_equal(
A + 1,
[ A[n] + 1 for n in range(0,len(A)) ]
)
A + B == B + A
# ### Subtraction
A - B
A - B == B - A
A - (B + C) == A - B - C
np.array_equal(
A,
A - 0,
)
np.array_equal(
A - B,
[ A[n] - B[n] for n in range(0,len(A)) ]
)
np.array_equal(
A - 1,
[ A[n] - 1 for n in range(0,len(A)) ]
)
# ### Multiplicaton
A * B
A * 1 == A
-A == A * -1
np.array_equal(
A * 5,
[ A[n] * 5 for n in range(0,len(A)) ]
)
np.array_equal(
A * B,
[ A[n] * B[n] for n in range(0,len(A)) ]
)
# ### Division
A / B
np.array_equal(
A / 5,
[ A[n] / 5 for n in range(0,len(A)) ]
)
np.array_equal(
A / B,
[ A[n] / B[n] for n in range(0,len(A)) ]
)
# ### Prove (c1 + c2) · V == c1 · V + c2 · V
# ((c1 + c2)*V)[j]
# = (c1 + c2)*(V[j])
# = (c1 * V[j]) + (c2 * V[j])
# = (c1 * V)[j] + (c2 * V)[j]
# = ((c1 * V) + (c2 * V))[j]
c1 = 3+4j
c2 = 7-8j
np.array_equal(
(c1 + c2) * A,
(c1 * A) + (c2 * A)
)
np.array_equal(
[ (c1 + c2) * A[n] for n in range(0,len(A)) ],
[ (c1 * A[n]) + (c2 * A[n]) for n in range(0,len(A)) ]
)
| juypter/notebooks/linear-algebra/1_vector_operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python for Environmental Science Day 2
# ## Topics
# * Boolean Values
# * Flow Control
# * Import Stuff
# ## Getting started!
# Just like yesterday, MIT provides a [great video](https://youtu.be/0jljZRnHwOI?t=53) that sums up pretty much everything we will cover today. After a little recap on string concatenation (what we already learned yesterday), it covers the basics of flow control (here called branching, yet another name for the same thing is conditionals) and later the basics of loops.
# ## Boolean Values
# Up until now we only used strings and numbers, but missed out on another very important data type in Python: boolean values.
# Boolean values behave a bit different in comparison to other data types. They can only have two values:
# * **True**
#
#
# or
#
#
# * **False**
#
# For a short introduction see [this video](https://youtu.be/9OK32jb_TdI).
#
# As you can see, boolean values are mostly used to compare things and to determine if variables have the values you expect them to have. To connect them two types of operators exist:
# * **Comparison operators** (==, !=, <, >, <=, >=)
#
# and
#
# * **Boolean operators** (and, or, not)
#
# Comparison operators are mainly used to compare things and boolean operators to connect and negate other statements.
#
# ### Practice Questions
#
# To test your understanding here are a bunch of different boolean expressions. Let's go through them together.
1 > 0
not 5 == 4
a = 37
b = 50
c = 1
d = 1
not(a > b and c == d) or c
30 <= 30 and (31 - 15) == 16
# - What are the possible values for a boolean variable?
# - Name the boolean operators and what they are used for
# ## Flow Control
# ### Conditionals
# Conditionals allow your programs to take different routes. For example you want to have a program that either prints a string or the number of characters in this string. You could outcomment parts of your code to accomplish this.
name = "Anna"
# print(name)
print(len(name))
# But this is a rather unpractical way to do this. Here, the conditionals come into play. There are three types of conditionals in Python:
# * **if**
# * **elif**
# * **else**
#
# For an explanation of their uses watch [this video](https://youtu.be/2aA3VBdcl6A) and [here](https://youtu.be/apcI8MkJTOA) (if you did not grasp the concept from the first one) and if you are still confused [here](https://youtu.be/f4KOjWS_KZs) is yet another one. So, a more logical way to tackle our task could be the following:
import random
random_num = random.randint(0,10)
print(random_num)
if random_num < 3:
print("smaller than three")
elif random_num > 3 and random_num <6:
print("between 3 and 6")
else:
print("all other cases")
# As mentioned in the videos before, an important concept in Python becomes obvious:
# * Indentation
#
# Python is structured by using indentations. This just means that Python recognizes all lines in one indentation as one block. So in our example above both print statements are in different blocks.
#
# ### Practice Questions
# * What happens if multiple elif blocks are True?
# * Can I have an if statement within an if statement?
# * What is the difference between "if" and "elif"?
# * How does Python know what part of the code belongs to a certain if?
#
# ### Loops
# When writing code you often want to do something repeatedly. You could do this by simply writing the same code over and over again.
i = 1
print(i)
i = i + 1
print(i)
i = i + 1
print(i)
i = i + 1
print(i)
# As you can imagine, writing code this way sucks.
#
# 
#
#
# But we are not the first persons to recognize that using Python this way is not a good idea. To overcome such problems loops exist. Loops run the code in a predefined number of times (for-loop) or until a statement is not True anymore (while-loop). For a detailed explanation see [this video](https://youtu.be/6iF8Xb7Z3wQ) and [this video](https://youtu.be/FzpurxjwmsM).
# Here is a little reminder on how the [range function](https://youtu.be/0jljZRnHwOI?t=36m21s) works, which comes in handy when making for-loops.
#
# Here is the code above as a for-loop.
for i in range(1, 5):
print(i)
# Why did I have to write 1 as start explicitly and 5 as end? [Answer](https://www.johndcook.com/blog/2008/06/26/why-computer-scientists-count-from-zero/)
#
# You can also loop over words.
for character in "banana":
print(character)
#
# Now here is a use case for a while loop. We do not yet know how often the code will run.
import random
i = 1
while i != 5:
i = random.randint(0,10)
print(i)
# ### Practice Questions
# * How often does **for i in range(1, 3):** loop?
# * What is the difference between a for-loop and a while-loop?
# * Loops are super confusing for me. Will I ever understand them?
# * What happens if you write **while True:**?
# * My code often looks different than the one in the solutions, but I still get the same results. Is this normal?
#
# ## Break and Continue
# Often it happens that you have a loop where you calculate some stuff in the loop and this changes the way you want the program to run. One possible tool here are the continue and break keywords. In short they do the following:
# * **continue**: Ignore the code after the continue and jump back to the start of a loop
# * **break**: End the current loop, regardless of all other things and continue with the code below the loop
#
# A bit more detailed explanation can be found [here](https://youtu.be/G8ClpHu8DSE).
#
# ### Practice Questions
# * Give an example for a case where you might need break.
# * Can you break out of several loops at once?
# ## Import Stuff
# We now already know most parts of the Python language, so in theory we could write most programs now. However, this would be a lot of work. One of the benefits of Python is its large community and as Python has been around for a while, this community has written lots and lots of useful packages. Those packages contain the code to do all kinds of things. Ranging from math function to plotting or reading and writing files. As we use WinPython, we already have the most widely used packages. We just have to import them. And in Python this is pretty easy.
import math
# now we can use functions of this package
print("The square root of 9 is " + str(int(math.sqrt(9))))
# To access the functions and variables from the package simply type the name of the module, then add a dot and hit tab.
import math # Load the module
math. # Hit tab and Python will show you what is in the module
# Also, all common modules in Python have an extensive documentation on how they work. To access those simply google "Package Name Python Documentation".
# ## Plan your Code
# Beginners often read an exercise and start to code immediately. This often leads to massive confusion, as beginners still struggle with the basic syntax and the conceptual ideas behind programming at the same time. To entangle this mess, it is usually a good idea to do those things separately. First write down what you think your program should be doing. Only when you have a good feel that the ideas you have written down might work, you should start coding. Let's look at exercise 6 of day 1 to make this more clear:
#
# "Write a program that asks the user for two numbers. Then calculate first number divided by second number (normal division), first number divided by second number (integer division) and the remainder of dividing the first number by the second number. Print the results. What differences can you see?"
#
# And a possible way to write this program down would be:
# +
# Ask the user for the first number
# Ask the user for the second number
# Calculate first number divided (normal) by the second number and store it in a variable
# Calculate first number divided (integer) by the second number and store it in a variable
# Calculate the remainder of the division of the first number by the second number and store it in a variable
# Print the results of all three calculations
# -
# Now you only have to translate this comments to code, which is much easier than trying to find a working concept and the right way to code it in Python at the same time.
# ## Finding Errors
# Even if you have planned your code, errors will still occur. To make finding them easier, try the following:
# * Add additional print statements in your code, to check if variables have the values you think they have.
# * Use the type() function to check whether something has the type, you assume it has.
# * Test new functions, you want to use, in the interactive console. This way you can check, if they do what you assume.
# * Use a debugger. See [here](https://youtu.be/2Wnmocwz7u0) for a little tutorial (unfortunately the explanation is a bit all over the place. If the video left you still confused about the debugger, ask me). A written explanation can be found in this [stackoverflow question](https://stackoverflow.com/questions/28280308/how-do-i-debug-efficiently-with-spyder-in-python).
# * When you are using spyder, take a look at the variable explorer.
# ### Exercise 1
# Write a program that prints the numbers 1 to 10 using a for-loop
# ### Exercise 2
# The time module provides you with a function also named time. This function returns you the seconds that have passed since 1 January 1970. Write a program that saves the value that the time function gives you in a variable. Then use this value to calculate how many minutes have passed since 1 January 1970 and print it.
# ### Exercise 3
# Write a Python program to find the operating system name, platform name and platform release version.
#
# Hint: There are modules for that
# ### Exercise 4
# Long ago smart Greek people had three sticks and they wanted to know if they could form a triangle with those sticks. They found that if any of the sticks is longer than the sum of the other two, they could not form a triangle. Write a program that prompts the user for three stick lengths. Then determine if the triangle is valid. Print your results to the user.
# ### Exercise 5
# Write a program to recreate the following pattern, using loops.
#
# Hint: Doing this with a lot of math is the hard way. Be lazy!
1
22
333
4444
55555
666666
7777777
# ### Exercise 6
# Write a program that has a variable called month and prints the number of days in a month, depending on the value of month.
# ### Exercise 7
# Write a Python program that prints all the numbers from 0 to 6 except 3 and 6.
#
# Use the 'continue' statement.
#
# Expected output :
#
# 0
#
# 1
#
# 2
#
# 4
#
# 5
# ### Exercise 8
# Write a Python program to print the numbers which are divisible by 7 and multiple of 5, between 1500 and 2700 (both included).
# ### Exercise 9
# As a future athlete you just started your practice for an upcoming event. Given that on the first day you run x miles, and by the event you must be able to run y miles.
#
# Calculate the number of days required for you to finally reach the required distance for the event, if you increase your distance each day by 10% from the previous day.
#
# Print one integer representing the number of days to reach the required distance.
#
# Source: Snakify
# ### Exercise 10
# Let us assume that you have finished your master and started saving money for your first big holiday. You want to know how many months you have to save money to be able to afford your wonderful holiday. For this you decide to write a Python program, because this will be super helpful in this case!
# Consider the following:
# * The program should ask you how much your annual salary will be, how much of this salary you want to save (as a fraction) and how much your holiday will cost.
# * You need a variable to save current savings you have (starts with 0)
# * You will have to settle for an annual return **r**. Every month you will receive current_savings * (r / 12), e.g. 4 % return of investment each year.
# * At the end of each month your current_savings will improve by the returns and the percentage of your monthly salary you wanted to save
#
# Hint: A while loop might be quite useful here.
#
# Hint: First write down what you think the program should do, start coding only after you have done this!
#
# An example output looks like this:
# How expensive should the holiday be?
#
# 10000
#
# How much of your monthly salary do you want to save?
#
# 0.05
#
# How large is your annual salary?
#
# 25000
#
# You need to save for 84 months to afford your holiday.
# ### Exercise 11
# Program a guessing game which randomly picks a number between 1 and 20 (both numbers included) and asks you to guess which number the program chose. When you win, the program should congratulate to your great victory and state the number of guesses you took. To have a better chance, the program should tell you after every wrong guess whether your guess was too low or too high.
#
# Hint: Do you remember the import stuff? There is a good chance that there is a package which allows you to create random numbers.
#
# ### Exercise 12
# Write a Python program to check if a triangle is equilateral, isosceles or scalene.
#
# Hint:
# An equilateral triangle is a triangle in which all three sides are equal.
# A scalene triangle is a triangle that has three unequal sides.
# An isosceles triangle is a triangle with two equal sides.
#
# Source: w3resource.com
# ### Exercise 13
# Write a Python program to check the validity of a password input by a user. You should prompt the user until a valid password is entered.
#
# Validation :
# * At least 1 letter between [a-z] and 1 letter between [A-Z].
# * At least 1 number between [0-9].
# * Minimum length 6 characters.
# * Maximum length 16 characters.
#
# Hint: The 'string' module can be quite helpful here to get the characters to compare to.
#
# Hint: You can iterate over strings (e.g. for character in "hallo":).
#
# Hint: You can check if something is contained in something else using 'in' (e.g. "a" in "Hallo").
#
# Hint: Consider using variables that save if you have found a character that fulfills the conditions.
#
# Source: w3resource.com
| week_1/day_2_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ESRI Land Cover
#
# This notebook is a simple example of loading ESRI's global 10 m Land Cover dataset.
# For more information, see [the ESRI data page](https://www.arcgis.com/home/item.html?id=d6642f8a4f6d4685a24ae2dc0c73d4ac).
# +
import datacube
from matplotlib import colors as mcolours
import numpy as np
# %matplotlib inline
dc = datacube.Datacube()
# +
product = "esri_land_cover"
# This is a point in Hobart
lat, lon = -42.822771, 147.234277
buf = 0.25
lons = (lon - buf, lon + buf)
lats = (lat - buf, lat + buf)
# Load the data at 10 m resolution
ds = dc.load(
product=product,
longitude=lons,
latitude=lats,
resolution=(-10, 10),
output_crs="epsg:6933"
)
# -
# Colour it like the ESRI colour map
cmap = mcolours.ListedColormap([
np.array([0, 0, 0]) / 255,
np.array([65, 155, 223]) / 255,
np.array([57, 125, 73]) / 255,
np.array([136, 176, 83]) / 255,
np.array([122, 135, 198]) / 255,
np.array([228, 150, 53]) / 255,
np.array([223, 195, 90]) / 255,
np.array([196 ,40, 27]) / 255,
np.array([165, 155, 143]) / 255,
np.array([168, 235, 255]) / 255,
np.array([97, 97, 97]) / 255
])
bounds=range(0,12)
norm = mcolours.BoundaryNorm(np.array(bounds), cmap.N)
ds.isel(time=0).classification.plot.imshow(cmap=cmap, norm=norm, size=10)
# +
# This is the centre of Australia, approximately
lat, lon = -26.621795, 135.110935
buf = 18
lons = (lon - buf, lon + buf)
lats = (lat - buf, lat + buf)
# Load the data at 10 m resolution
ds_au = dc.load(
product=product,
longitude=lons,
latitude=lats,
resolution=(-1000, 1000),
output_crs="epsg:6933",
dask_chunks={} # Use dask to make it lazy load
)
# -
# %%time
ds_au.isel(time=0).classification.plot.imshow(cmap=cmap, norm=norm, size=10)
| notebooks/ESRI_Land_Cover.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# This exmaple will show you how to interpret anomalies in univariate *time-series* samples.
#
# Here we use a HDFS log anomaly detector, named *Deeplog* (CCS'17).
#
# # Prepare the Deeplog model
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
# Train Deeplog anomaly detection model
import numpy as np
import torch
from deeplog import train_deeplog, test_deeplog
train_data = np.load('./data/train_data.npz')
train_normal_seq = train_data['train_normal_seq']
train_normal_label = train_data['train_normal_label']
model = train_deeplog(train_normal_seq, train_normal_label)
torch.save(model, './save/LSTM_onehot.pth.tar')
# Validate the performance of trained model
test_normal_loader = np.load('./data/test_normal_loader.npy',allow_pickle=True)
test_abnormal_loader = np.load('./data/test_abnormal_loader.npy',allow_pickle=True)
test_deeplog(model, test_normal_loader, test_abnormal_loader)
# # Interpret your interested anomaly in four steps
# +
"""Step 1: Load your model"""
from deeplog import LSTM_onehot
import torch
model = torch.load('save/LSTM_onehot.pth.tar')
"""Step 2: Find an anomaly you are interested in"""
import sys
sys.path.append('../../deepaid/')
from utils import deeplogtools_seqformat
abnormal_data = np.load('data/abnormal_data.npy')
idx = 100
seq, label, anomaly_timeseries = deeplogtools_seqformat(model, abnormal_data, num_candidates=9, index=idx)
# print(seq.shape,label.shape)
"""Step 3: Create a DeepAID Interpreter"""
import sys
sys.path.append("../../deepaid/interpreters/")
from timeseries_onehot import UniTimeseriesAID
feature_desc = np.load('data/log_key_meanning.npy') # feature_description
my_interpreter = UniTimeseriesAID(model, feature_desc=feature_desc, class_num=28)
"""Step 4: Interpret your anomaly and show the result"""
interpretation = my_interpreter(seq, label)
my_interpreter.show_table(anomaly_timeseries, interpretation)
| demos/timeseries_uni_deeplog/timeseries_example_deeplog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Ale1167A/daa_2021_1/blob/master/30Septiembre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="uxBPIJccP-lW"
# #Palindromo
#
# ###plantemiento
# Encontrar todos los palindromos que existen en la franja horaria de un dia
# completo, tomando como horario inicial las 00:00 horas y como horario final
# las 23:59.
#
# El algoritmo debe mostrar todos los palindromos existen en es rengo y mostrarlo en pantalla y hacer el conteo final de todos los palindromos encontrados
# + id="QYn5tsj-Put1" outputId="07ff1a60-ffb6-4880-9787-c0fd938e3ff6" colab={"base_uri": "https://localhost:8080/", "height": 476}
#Solucion
def horas():
for m in range(0,24,1):
for h in range(0,60,1):
if m == h and h == m:
print(f'{m}:{h}')
print(f'{m+1} son palindromos')
def main():
hora = horas()
main()
| 30Septiembre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import matplotlib.pyplot as plt
import math
import scipy.integrate as integrate
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 10.0)
# ## a)
# $1 - p \; \Delta t $
# ## b)
# $P(n, t+ \Delta t, t_0) = P(n,t,t_0)(1 - p \Delta t) + P(n-1, t, t_0) p\Delta t$
# ## c)
# $P(n, t+ \Delta t, t_0) - P(n,t,t_0) = P(n-1, t, t_0) p\Delta t - P(n,t,t_0) p \Delta t \\$
#
# $\dfrac{P(n,t+\Delta t, t_0) - P(n,t,t_0)}{\Delta t} = P(n-1, t, t_0) p - P(n,t,t_0) p$
#
# now set $\Delta t$-> 0, and you get: $\dfrac{d P(n,t, t_0)}{d t} = P(n-1, t, t_0) p - P(n,t,t_0) p$
# ## d)
# $P(0, t+ \Delta t, t_0) = P(0,t,t_0)(1- p\Delta t) = P(0, t, t_0) - P(0,t,t_0) p \Delta t$
#
# $ \dfrac{P(0, t+\Delta t, t_0) - P(0,t,t_0)}{\Delta t} = - P(0,t,t_0) p$
#
# Same trick with $\Delta t$ as previously and get a partial:
#
# $ \dfrac{d P(0, t, t_0)}{d t} = - P(0,t,t_0) p$
# ## e)
# Let's introduce some notation: $A(t) = -p$, $B(t;n) = p\; x(n-1,t)$, $x(n,t)= P(n,t,t_0)$
# Now let's prove that eq. 2 solution is exactly the same as eq. 3 when n=0.
#
# ###1.
#
# $ P(0, t, t_0) = x(0,t) = C_0 e^{-p(t-t_0)} = e^{-p(t-t_0)}$ which we obtain by using the initial condition for $n=0$,$t=t_0$.
#
# ### 2.
# Now we prove that eq. 3 is the solution for eq. 1. First we note that we going to use linear non-autonomous O.D.E.s to prove with the previously introduced notation.
# Recall that linear non-autonomous O.D.E.s are O.D.Es of the form:
# $\frac{dx}{dt} = A(t)\;x + B(t) $, and the solution is $x= I^{-1} \int I \; B(t) dt $, where $I = e^{-\int A(t) dt}$
#
# $$ \frac {d P (n, t, t_0)}{dt} = \frac{d x(n, t)}{d t} = A(t) \; x(n,t) + B(t;n) \\
# I = e^{\int p dt} = e^{pt}$$
# Now we can obtain the solution: $ x(n,t) = e^{-pt} \int e^{pt}p \;x(n-1,t) dt$. Notice that it has a recursion, which we can unfold to obtain:
# $$
# x(n,t) = e^{-pt} \int e^{pt}p \;x(n-1,t) dt = e^{-pt} p^2 \int \int e^{pt} \; x(n-2,t) dt dt = \\
# = e^{-pt} p^n \int ... \int e^{pt} \; x(0,t) dt ... dt
# $$
# Now substitute our previously obtained solution for $x(0,t)$, and we get: $e^{-p(t-t_0)} p^n \int ... \int dt ... dt$
# Let's examine the integrals, we have $n$ of them:
# $\int ... \int dt ... dt = \int ... \int (t + C)dt ... dt$ (now we have $n-1$ integrals).
#
# To obtain the value of the constant, we use the initial condition:
#
# $e^{-p(t_0-t_0)} p^n \int ... \int (t_0 + C)dt ... dt = 0 => \int ... \int (t_0 + C)dt ... dt = 0$, so $C = -t_0$
#
# Using the same principle, we integrate once more: $e^{-p(t-t_0)} p^n \int ... \int (\frac{t^2}{2} - tt_0 + C)dt ... dt $, and again by using the initial condition we derive that $C = \frac{t_0^2}{2}$, therefore we obtain:
# $e^{-p(t-t_0)} p^n \int ... \int (\frac{t^2}{2} - tt_0 + C)dt ... dt = e^{-p(t-t_0)} p^n \frac{1}{2} \int ... \int (t^2 - 2tt_0 + t_0^2)dt ... dt = e^{-p(t-t_0)} p^n \frac{1}{2} \int ... \int (t - t_0)^2 dt ... dt $
#
# Notice that we have $n-2$ integrals left, the fraction is $2^{-1}$ and the power in the integrals is 2. Now if you integrate further the polynomial, you will get the solution inside of the integrals in the form : $\frac{1}{k+1}(t- t_0)^{k+1} + C $, and we notice that it's sufficient to set $t=t_0$ in order to satisfy the initial condition, and thus C = 0. Therefore, by performing the integration $n-2$ times we arrive to the eq. 3.
#
#
#
#
# ## Here is an alternative solution:
# #### e) for the differential equations 1 and 2 that agree with the initial conditions.
#
# firstly calculate the FOC for the ansatz:
#
# $\frac{\partial P(n,t,t_0)}{\partial t} = \frac{\partial A_n(t)}{\partial t} e^{-p(t-t_0)} -p A_n(t) e^{-p(t-t_0)}$
#
# Now prove that this holds for the differential equation for 0.
#
# $\frac{\partial P(0;t,t_0)}{\partial t} = 0 - p A_n(t) e^{-p(t-t_0)} $
#
# $\frac{\partial P(0;t,t_0)}{\partial t} = - p P(0;t,t_0) $
#
# Prove that it holds for all other conditions.
#
# Take the derivative wrt $A_n$, where $A_n = \frac{p(t-t_0)^n}{n!}$
#
# $\frac{\partial A_n(t)}{\partial t} = \frac{p(pt)^{n-1}}{(n-1)!}$
#
# This simply states: $\frac{\partial A_n(t)}{\partial t} = p A_{n-1}$
#
# If we fill this in, we get the normal differential equation for the other cases:
#
# $\frac{\partial P(n,t,t_0)}{\partial t} = p A_{n-1} e^{-p(t-t_0)} -p A_n(t) e^{-p(t-t_0)}$
#
# #### prove the initial conditions are correct:
#
# $P(0,t_0,t_0)= \frac{p(0)^0}{0!} e^{-p(0)} = 1 $
#
# and,
#
# $P(n,t_0,t_0)= p \frac{p(0)^{n-1}}{(n-1)!} e^{-p(0)} - p \frac{p(0)^n}{n!} e^{-p(0)} = 0 $
#
# ## f)
# Since $n|t,t_0 \sim Pois(p(t-t_0))$ , so $E[n|t,t_0] = p(t-t_0)$
#
# ## And an alternative solution:
#
# From equation 3:
#
# $P(n;t,t_0) = \frac{p(t-t_0)^n e^{-p(t-t_0)}}{n!}$
#
# calculate the expectation:
#
# $E[P(n;t,t_0)] = \sum_n n \frac{p(t-t_0)^n e^{-p(t-t_0)}}{n!}$
#
# $E[P(n;t,t_0)] =e^{-p(t-t_0)}p(t-t_0) \sum_{n>0} \frac{p(t-t_0)^{n-1} }{(n-1)!}$
#
# Use the taylor series expansion of the exponential function:
#
# $E[P(n;t,t_0)] =e^{-p(t-t_0)}e^{p(t-t_0)} p(t-t_0)$
#
# $E[P(n;t,t_0)] =p(t-t_0)$
# ## g)
#
# Let's introduce some notation: $\lambda = p(t-t_0)$
# We want to find n, such that : 1) $\frac{P(n,t,t_0)}{P(n-1,t,t_0)} = \frac{\lambda}{n} > 1$
# and 2) $\frac{P(n,t,t_0)}{P(n+1, t,t_0)} = \frac{n+1}{\lambda} > 1$
# so we are looking for $n > \lambda -1 $ and $n< \lambda $, which is the integer part of lambda. e.g. if $\lambda = 1.5$, then $n>0.5$ and $n<1.5$, so it must be that $n=1$ (Recall that Poison is a discrete distribution, and also that e.g. 0.7 friends ariving to a party make no sense :))
# ## An alternative solution
#
# For P(n;t,t0) > P(n−1;t,t0), firstly fill in the equations:
#
# $\frac{(p(t-t_0))^n e^{-p(t-t_0)}}{n!} = \frac{(p(t-t_0))^{(n-1)} e^{-p(t-t_0)}}{(n-1)!}$
#
# $\frac{(p(t-t_0))^n (n-1)! }{n! (p(t-t_0))^{(n-1)}} = \frac{ e^{-p(t-t_0)}}{e^{-p(t-t_0)}}$
#
# $\frac{(p(t-t_0)) }{n} = 1$
#
# $(p(t-t_0)) = n$
#
# For P(n;t,t0) > P(n+1;t,t0), firstly fill in the equations:
#
# $\frac{(p(t-t_0))^n e^{-p(t-t_0)}}{n!} = \frac{(p(t-t_0))^(n+1) e^{-p(t-t_0)}}{(n+1)!}$
#
# $\frac{(p(t-t_0))^n (n+1)! }{n! (p(t-t_0))^{(n+1)}} = \frac{ e^{-p(t-t_0)}}{e^{-p(t-t_0)}}$
#
# $\frac{n + 1 }{(p(t-t_0))} = 1$
#
# $(p(t-t_0)) - 1 < n$
#
# Therefore,
#
# $(p(t-t_0)) - 1 < n < (p(t-t_0)) $
#
# ## h)
# +
def run_simulation_1(t_max, delta_t, p, N, t_0=0):
counts = []
for i in range(N):
count = 0
for j in range(t_0, t_max, delta_t):
count += np.random.binomial(1,p)
counts.append(count)
return counts
# poisson function, parameter lamb is the fit parameter
def poisson(x, lamb):
return (lamb**x/math.factorial(x)) * np.exp(-lamb)
# -
# setup
t_max = 60 # minutes
delta_t = 1
p = 0.5
N = 10000
counts = run_simulation_1(t_max, delta_t, p*delta_t, N)
fig, ax = plt.subplots()
x_line = np.linspace(0, 60, 61)
lamb = t_max * p
ax.hist(counts, 29, normed=1)
pois = [poisson(x, lamb) for x in x_line]
ax.plot(x_line, pois, 'r--', label='Poisson')
plt.xlabel("number of guests")
plt.ylabel("probability")
plt.title("the number of guests that arrived in an hour (with per minute simulation)")
plt.show()
# ## i)
# setup
t_max = 3600 # now in seconds
delta_t = 1
p = 0.5/60.0
N = 10000
counts = run_simulation_1(t_max, delta_t, p, N)
x_line = np.linspace(0, 60, 61)
fig, ax = plt.subplots()
lamb = t_max * p
ax.hist(counts, 40, normed=1)
pois = [poisson(x, lamb) for x in x_line]
ax.plot(x_line, pois, 'r--', label='Poisson')
plt.xlabel("number of guests")
plt.ylabel("probability")
plt.title("the number of guests that arrived in an hour (with per second simulation)")
plt.show()
# Simulations are different because in the first simulation we're assuming that a visitor can come every minute, and in the second one we're assuming that they can come every second, and the latter assumption is more realistic. In addition, the latter assumption is supported by the theory that binomial distribution ( we simulated it) can be approximated by Poison by setting probability of success close to 0, and the number of trials to infinity (limit).
# The second plot is more accurate, because it allows visitors to come on seconds basis instead of minutes.
# ### j)
# P($n_{th}$ visitor arrived at time $t$) = P(1 person arrives at time $t$, and $n-1$ people are present at time $t-\Delta t$) = P(1 person arrives at time $t$)P($n-1$ people are present at time $t-\Delta t$)
#
# Notice that those two events are independent, i.e. a person arrives to a party with equal probability regardless on the number of people already present, and vice-versa. Then, we notice that the second probability factorization component computes probability of Poisson r.v. taking value $n-1$, and since $\Delta t$ is infinitesmally small we can ignore it. Thus we obtain:
#
# $\Delta t \; p \dfrac{(p(t-t_0))^{n-1}}{(n-1)!} e^{-p(t-t_0)}$
# and by taking $\lim_{\Delta t ->0} \Delta t$, we get: $dt \; p \dfrac{p(p(t-t_0))^{n-1}}{(n-1)!} e^{-p(t-t_0)} = dt P(t;n,t_0)$
#
#
#
#
# ##k)
# Let $T \sim Expo(p)$, then we are interested in $E[T]$, and note that the support of T is $[0, + \inf]$. In addition, we note that here we mean that $T = \Delta t$ is the time between $t_0$ and $t$ as exponential distribution does not have any extra parameters, such as $t_0$.
# $E[T] = \int_{\Delta t=0}^\infty \Delta t \; p \; e^{- p \Delta t} d\Delta t = p \int_{t=0}^\infty \Delta t e^{- p \Delta t} d \Delta t
# = p (- \dfrac{e^{-p \Delta t + 1}}{p^2}) |_{t=0}^{+\infty} = \frac{1}{p}$
# ##l)
# Let $T_1 \sim Expo(p;t_0)$ be the time when the first person arrives to the party, and $T_2 \sim Expo(p;t_0)$ be the time for the second person to arrive(after the first one has arrived, so the second r.v. depends on the first one).
# $P(T_1 = t_1, T_2 = t_2| T_2 \geq t_1 ) = P(T_1 = t_1) P(T_2 = t_2 | T_2 \geq t_1)$
# $P(T_2 = t_2 | T_2 \geq t_1) = \dfrac{ P(T_2 = t_2 , T_2 \geq t_1)}{P(T_2 \geq t_1)}$ Here the density is zero unless $t_2 \geq t_1$, otherwise:
# $\dfrac{ P(T_2 = t_2 , T_2 \geq t_1)}{P(T_2 \geq t_1)} = \dfrac{p e^{-p(t_2 - t_0)} }{ 1 - (1 - e^{-p(t_1 - t_0)}) } = p e^{-p(t_2-t_1)}$
# $ P(T_1 = t_1) P(T_2 = t_2 | T_2 \geq t_1) = p^2 e^{-p(t_2 - t_0)}$
# $\int_{t_1 = t_0}^{t_2} P(T_1 = t_1, T_2 = t_2| T_2 \geq t_1 ) dt_1 = p^2 e^{-p(t_2 - t_0)}\int_{t_1 = t_0}^{t_2} dt_1 = p^2(t_2-t_0) e^{-p(t_2 - t_0)} $
# ## An alternative solution:
#
# Firstly we calculate independently the chances of 2 people arriving seperately in 2 time periods:
#
# $P(t_2;2,t_0) = P(t_1;1,t_0)P(t_2;1,t_1) = p^2e^{-p(t_1-t_0)} e^{-p(t_2-t_1)}= p^2 e^{-p((t_2-t_1) + (t_1-t_0))} $
#
# $P(t_2;2,t_0) = p^2 e^{-p(t_2-t_0)} $
#
# Now we integrate over this wrt $t_1$
#
#
# $P(t_2;2,t_0) = \int^{t_2}_{t_0}p^2 e^{-p(t_2-t_0)}\mathcal{d}t_1 = p^2 e^{-p(t_2-t_0)}t_2 -p^2 e^{-p(t_2-t_0)}t_0$
#
# $P(t_2;2,t_0) = p(p(t_2 - t_0))^{2-1} e^{-p(t_2-t_0)}$
#
# Which is equal to:
#
# $P(t_2;2,t_0) = \frac{p(p(t_2 - t_0))^{2-1} e^{-p(t_2-t_0)}}{(2-1)!}$
#
#
# ##m)
def simulate_expo(T, p = 0.5, N=10000):
visitors=[]
generations_count = 0
for _ in range(N):
samples=[]
while np.sum(samples) < T:
samples.append(np.random.exponential(1.0/p))
generations_count+=1
visitors.append(len(samples[:-1]))
return visitors, generations_count
t_max = 60.0
p = 0.5
fig, ax = plt.subplots()
visitors,_ = simulate_expo(t_max,p, N=10000)
x_line = np.linspace(0, 60, 61)
lamb = t_max * p
pois = [poisson(x, lamb) for x in x_line]
ax.plot(x_line, pois, 'r--', label='Poisson')
plt.xlabel("number of guests")
plt.ylabel("probability")
plt.title("the number of guests that arrived in an hour ( simulation via exponential)")
ax.hist(visitors, 40, normed=1)
plt.show()
# The shape is very similar to the one obtained in the previous sumulations. The previous distribution obtained via minutes-based simulation has higher mass around mean, but it's almost exactly the same as the one obtained via seconds-based simulation.
#
# Finally, the obtained normalized histogram is very similar to Poisson PDF, which can be explained by Poisson processes that encodes relationship between Exponential and Poison distributions.
# ## n)
_,generations_count = simulate_expo(t_max,p, N=10000)
print "the number of generations is %d " % generations_count
# We generate roughly 30 times for each simulation, where each generation can be interpreted as generation of the waiting time between a last visitor and a new one arriving to the party. We notice that previously we simulated Binomial process, which has mean of 30 visitors per simulation (which agrees with theoretical mean that is $np$, where n = 60 steps and $p =0.5$).
#
# The number of steps we previously simulated is $60 * 10k = 600k$, and here we have roughly 300k generations, which is twice less. It can be explained by the fact that when we generate a waiting time between new visitor's arrival we already assume that he is certainly going to come, while in Binomial simulation we are not, and that's why we "flip a coin" to determine if a visitor arrives(and thus we need to simulate all 600k flips). And as we expect on average 30 people to visit (Binomial mean), we analytically expect that we will have around $30 * 10k$ simulations in the last simulation.
# ## o)
# $P(t_n; n, t_0) P(0;t,t_n) = \dfrac{p(p(t_n - t_0))^{n-1}}{(n-1)!} e^{-pt_n +pt_0 - pt + pt_n} =\dfrac{p(p(t_n - t_0))^{n-1}}{(n-1)!} e^{-p(t - t_0)}$
#
# $ \int_{t_n=t_0}^{t} \dfrac{p(p(t_n - t_0))^{n-1}}{(n-1)!} e^{-p(t - t_0)} dt_n = \dfrac{(p(t-t_0))^n}{n!} e^{-p(t-t_0)}$
# ## Alternative solution
# We must prove the following:
#
# $P(n;t_0,t) =\int_{t_0}^{t} P(t_n;n,t_0) P(0;t,t_n) \mathcal{d}t_n$
#
# Let's begin:
#
# $P(n;t_0,t) = \int_{t_0}^{t} \frac{p(p(t_n-t_0))^{n-1}e^{-p(t_n-t_0)}}{(n-1)!} \frac{p(t-t_n)^0 e^{-p(t-t_n)}}{n!} \mathcal{d}t_n$
#
# $P(n;t_0,t) = \int_{t_0}^{t} \frac{p^n (t_n-t_0)^{n-1}e^{-p((t_n-t_0)+(t-t_n))}}{(n-1)!}\mathcal{d}t_n$
#
# $P(n;t_0,t) = \int_{t_0}^{t} \frac{p^n (t_n-t_0)^{n-1}e^{-p(t-t_0)}}{(n-1)!}\mathcal{d}t_n$
#
# $P(n;t_0,t) = \frac{p^n e^{-p(t-t_0)}}{(n-1)!} \int_{t_0}^{t} (t_n-t_0)^{n-1}\mathcal{d}t_n$
#
# $P(n;t_0,t) = \frac{p^n e^{-p(t-t_0)}}{(n-1)!} \frac{(t_n-t_0)^{n}}{n}$
#
# evaluate it at $t_0$ and $t$, this yields:
#
# $P(n;t_0,t) = \frac{p^n e^{-p(t-t_0)}}{(n-1)!} \frac{(t-t_0)^{n}}{n}$
#
# which simplifies to:
#
# $P(n;t_0,t) = \frac{(p(t-t_0))^n e^{-p(t-t_0)}}{n!} $
#
# This is correct!
# ## 0.2. Time dependent rate
# ## a)
# $P(n;t, t_0) = \dfrac{ (\int_{s=t_0}^t p(s) ds)^n \;}{n!} e^{-\int_{s=t_0}^t p(s) ds}$
# ##b)
# $\lim_{t-> \infty} P(0;t,t_0) = e^{- \lim_{t-> \infty} \int_{s=t_0}^{t} p(s) ds}$
# It's approaching zero if $p(t)$ has a large domain or infinite with $p(t)$ being non-zero (e.g. monotonically increasing).
# It's <b>not</b> approaching zero when $p(t) = 0$ on the domain $[\hat t,+\infty]$, and takes non-zero values on the domain $[t_0, \hat t]$, e.g. it's linear. Then the integral in the exponent can be broken down into 2 parts : $\int_{s=t_0}^{\infty} p(s) ds = \int_{s=t0}^{\hat t} p(s)ds + \int_{s=\hat t}^{+\infty} p(s) ds$, and using the initial assumption about the second component (it's zero) , we get : $\int_{s=t0}^{\hat t} p(s)ds$, which is a constant that we are free to choose to make the probability non-zero.
#
# Now let's take a limit of constant rate Poisson pmf. $\lim_{t->\infty} e^{-p(t-t_0)} = \lim_{t->\infty} e^{-pt} e^{pt_0} -> 0$, for all $p>0$, and when $p=0$, the probability is 1 for every $t$ and $t_0$.
# ## c)
# $ P(t,n,t_0) = p(t) \dfrac{(\int_{s=t_0}^{t} p(s) ds)^{n-1}} {(n-1)!} e^{- \int_{s=t_0}^{t} p(s) ds}$
#
#
# ## Alternative (more detailed) solution
# From question 1.j : $p(t;n,t_0) = \frac{p(p(t-t_0))^{n-1}e^{-p(t-t_0)}}{(n-1)!}$
#
# Apply pattern matching:
#
# $p = p(t)$, and $p(t-t_0)= \int_{t_0}^t p(s) \mathcal{d}s$
#
# $p(t;n,t_0) = \frac{p(t)(\int_{t_0}^t p(s) \mathcal{d}s)^{n-1}e^{-\int_{t_0}^t p(s) \mathcal{d}s}}{(n-1)!}$
# ## d)
# $ \int_{t_1 =t_0} ^{t_2} p(t_1) e^{-\int_{s=t_0}^{t_1} p(s) ds} p(t_2) e^{-\int_{s=t_1}^{t_2}p(s) ds} dt_1 = p(t_2) \int_{t_1=t_0}^{t_2} e^{-\int_{s=t_0}^{t_2} p(s) ds}p(t_1) dt_1 = \\
# =p(t_2) \int_{s=t_0} ^{t_2} p(s) ds \ e^{-\int_{s=t_0}^{t_2} p(s) ds}$
# ## e)
# setup
time_rates =[0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.3, 0.1]
time_rates = np.repeat(time_rates, 60)
# modified gamma pdf with non-constant rate
# t: upper_limit
# t_0: lower_limit
# n : number of sucesses
def gamma_mod(t_0, t, n, rates):
assert n>=0
if t > len(rates)-1:
return 0
rate = rates[t]
integr = np.sum(rates[t_0:t]) # computing the integral
fac_and_int_pow = 1.0 if n==0 else (integr**(n-1))/math.factorial(n-1)
return rate * np.exp(-integr) * fac_and_int_pow
# +
# it total we have 8 time slots according to table 1
hours_to_consider = 1.5
upper_limits_first = np.arange(0, hours_to_consider*60, 5)
theor_first_arrival_times = []
for upper_limit in upper_limits_first:
theor_first_arrival_times.append(gamma_mod(t_0=0, t=upper_limit, n=0, rates=time_rates))
# plotting
fig, ax = plt.subplots()
ax.plot(upper_limits_first, theor_first_arrival_times, 'r--', label='')
plt.xlabel("upper limit (minutes from 18.00)")
plt.ylabel("probability")
plt.title("probability of first visitor's arrival in different time")
plt.show()
# -
# ## f)
# +
# it total we have 8 time slots according to table 1
hours_to_consider = 3
upper_limits_tenth = np.arange(0, hours_to_consider*60, 5)
theor_10th_arrival_time = []
for upper_limit in upper_limits_tenth:
theor_10th_arrival_time.append(gamma_mod(t_0=0, t=upper_limit, n=10, rates=time_rates))
# print vals
# plotting
fig, ax = plt.subplots()
ax.plot(upper_limits_tenth, theor_10th_arrival_time, 'r--', label='')
plt.xlabel("upper limit (minutes from 18.00)")
plt.ylabel("probability")
plt.title("probability of 10th visitor's arrival in different time")
plt.show()
# -
# ## g)
# +
# for float comparison
# here we relax the relative tolerance because with the way we search over space of t.
def is_close(a, b, rel_tol=0.001, abs_tol=0.00):
return abs(a-b) <= max(rel_tol * max(abs(a), abs(b)), abs_tol)
def sample_from_expo(t_0, rates):
uni = np.random.uniform(0, 1)
# print "uni is %f" % uni
t_prev = t_0
t = t_0
lhs = rates[t_prev]
rhs = -np.log(1.0-uni) # right hand side
min_diff = float('infinity')
best_lhs = None
best_t = t_0
while True:
t += 1
if t >= len(rates):
return best_t + 1 # corner case
lhs += rates[t]
t_prev = t
if is_close(lhs, rhs):
return t
diff = abs(rhs - lhs)
# here the assumption about non-decreasing integral kicks-in
if diff < min_diff:
min_diff = diff
best_lhs = lhs
best_t = t
else:
# print "best lhs is %f" % best_lhs
# print "best t is %f" % best_t
return best_t
# -
# setup for g) and h)
# now we will look into seconds instead of minutes as we did previously
time_mult = 60.0 # 60 means that we will look at seconds instead of minutes, in this way we obtain more accurate results
time_rates = np.float32([0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.3, 0.1])/time_mult
time_rates = np.repeat(time_rates, 60*time_mult)
hours_to_consider = 8
# Sanity check of exponential
samples= []
for _ in range(5000):
samples.append(sample_from_expo(t_0 = 0, rates=time_rates))
samples = np.array(samples) / float(time_mult)
fig, ax = plt.subplots()
plt.hist(samples, normed=True, bins= 30)
ax.plot(upper_limits_first, theor_first_arrival_times, 'r--', label='theoretical pdf')
plt.xlabel("count")
plt.ylabel("density")
legend = ax.legend(loc='upper center', shadow=False)
plt.show()
print 'sampling mean is %d' % np.mean(samples)
# ## h)
def simulate_expo_2(t_0, rates, N=1000):
visitors=[]
generations_count = 0
first_visitor_arrivals = []
tenth_visitor_arrivals = []
# visitors_arrival_mean = []
T = len(rates)
# print "T is %d" % T
for i in range(N):
samples_count = 0
visitors_arrival_time = []
sample = 0
if i%100 == 0:
print i
while sample < T-1:
# prev_sample = sample
sample = sample_from_expo(t_0=sample, rates=rates)
# visitors_arrival_time.append(sample - prev_sample)
# print "sample %f" % sample
# elapsed_time+=sample
if sample < T :
samples_count+=1
if samples_count==1:
first_visitor_arrivals.append(sample)
if samples_count==10:
tenth_visitor_arrivals.append(sample)
generations_count+=samples_count
visitors.append(samples_count)
# visitors_arrival_mean.append(np.mean(visitors_arrival_time))
return visitors, generations_count, first_visitor_arrivals, tenth_visitor_arrivals, visitors_arrival_mean
visitors, generations_count, first_arrivals, tenth_arrivals, visitors_arrival_mean = simulate_expo_2(t_0 = 0, rates=time_rates, N=5000)
first_arrivals = np.array(first_arrivals)/time_mult
tenth_arrivals = np.array(tenth_arrivals)/time_mult
fig, ax = plt.subplots()
# pois = [poisson(x, lamb) for x in x_line]
# ax.plot(x_line, pois, 'r--', label='Poisson')
ax.hist(first_arrivals, 30, normed=1)
plt.xlabel("upper limit (minutes from 18.00)")
ax.plot(upper_limits_first, theor_first_arrival_times, 'r--', label='theoretical pdf')
plt.ylabel("density")
legend = ax.legend(loc='upper center', shadow=False)
plt.title("distribution of first person's arrival waiting time")
plt.show()
fig, ax = plt.subplots()
# pois = [poisson(x, lamb) for x in x_line]
# ax.plot(x_line, pois, 'r--', label='Poisson')
ax.hist(tenth_arrivals, 38, normed=1)
ax.plot(upper_limits_tenth, theor_10th_arrival_time, 'r--', label='theoretical pdf')
plt.xlabel("upper limit (minutes from 18.00)")
plt.ylabel("density")
legend = ax.legend(loc='upper center', shadow=False)
plt.title("distribution of tenth person's arrival waiting time")
plt.show()
fig, ax = plt.subplots()
ax.hist(visitors, 38, normed=1)
plt.xlabel("count")
plt.ylabel("density")
legend = ax.legend(loc='upper center', shadow=False)
plt.title("Distribution of the number of visitors")
plt.show()
print "the emperical number of visitors to the party is %f" % np.mean(visitors)
print "the expected number of visitors, according to Poisson is %f" % sum(time_rates)
# # 0.3) State dependent rate
# ## a)
# The probability/density we have now is different from the previous exponential density because now our rate is depends on the number of visitors already present. Thus, by using our rate, we can encode people preferences for visiting depending on the number of people present and the arrival time.
# ## b)
# $P(t_1; 0, t_0) P(t_2;1, t_1) = p(0,t_1) e^{-\int_{t_0}^{t_1} p(0,s) ds} p(1,t_2) e^{-\int_{t_1}^{t_2} p(1,s) ds} =
# p(0,t_1) p(1,t_0) e^{-\int_{t_0}^{t_1} p(0,s) ds -\int_{t_1}^{t_2} p(1,s) ds}
# $
# ## c)
# The reason why it does not work out nicely as previously is because now we have different rates(that depend on different n's) that can't be combined, see the power of the exponent.
# $P(t_1; 0, t_0) P(t_2;1, t_1) = \int_{t_1=t_0}^{t_2} (0,t_1) p(1,t_0) e^{-\int_{t_0}^{t_1} p(0,s) ds -\int_{t_1}^{t_2} p(1,s) ds} dt_1$
# ## d)
# +
import random
import numpy as np
import math as mt
l = list(np.array([0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.3, 0.1])/60.0)
hourly_rates = [x for x in l for _ in range(60*60)]
def cumulative_sum(rates, start):
return np.cumsum(rates[start:], axis = 0 )
def generate_times_n(hourly_rates):
times = []
x = - np.log(1 - np.random.rand())
t_index = 0
t = 0
while t_index <= 480*60:
n = len(times)
factor = np.exp(-0.05*(n - 100))/(1 + np.exp(-0.05*(n - 100)) )
#totalsum = np.cumsum([hourly_rates[t_index-t] for i in range(0,100)])
totalsum = factor * cumulative_sum(hourly_rates, t_index)
# catch if returns an empty list!
try:
t = np.argmax(totalsum[totalsum < x])
# calculate the differences and compare
diff_left = abs(totalsum[t] -x)
if t+1 < len(totalsum):
diff_right = abs(totalsum[t+1]-x)
if diff_right < diff_left:
t += 1
except:
# people can't arrive at the same time.
t = 1
# eventually the list is empty!
if 480*60 - t_index <= 3 and t ==0:
break
# add to index
t_index += t
if t_index > 480*60:
pass
times.append(mt.floor(t_index/60))
x = - np.log(1 - np.random.rand())
return times
y = generate_times_n(hourly_rates)
# -
# ## e)
# +
def run_simulation():
first= []
tenth = []
total = []
time = []
for x in range(0,1000):
guests = generate_times_n(hourly_rates)
first.append(guests[0])
tenth.append(guests[9])
total.append(len(guests))
time.append(np.mean(np.array(guests)))
return first, tenth, total, time
first, tenth, total, time = run_simulation()
# +
# show total number of guests
n, bins, patches = plt.hist(total, 20, normed=1)
plt.xlabel('length')
plt.ylabel('Relative counts')
plt.title('Total number of guests arriving.')
plt.show()
n, bins, patches = plt.hist(first, 20, normed=1)
plt.xlabel('first guest arrival time')
plt.ylabel('Relative counts')
plt.title('Arrival time of the first guest')
plt.show()
n, bins, patches = plt.hist(tenth, 20, normed=1)
plt.xlabel('tenth arrival time')
plt.ylabel('Relative counts')
plt.title('Arrival time of the tenth guest.')
plt.show()
print 'The average number of people arriving at our lame ass party is: %f'% np.mean(total)
# -
| hw2/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 10
from sympy import *
init_printing()
x = Symbol('x')
m = Symbol('m')
f0 = x**2 + 2 * x + m
f1 = x**2 + 3 * x + 2 *m
expand(f0)
expand(f1)
solve([f0, f1], [x, m])
# ### 手計算
# $$
# \begin{equation}
# x^2+ x + m = 0
# \label{eq:pythagoras}
# \tag{1}
# \end{equation}
# $$
# $$
# \begin{equation}
# 2x^2+ 2x + 2m = 0
# \label{eq:pythagoras}
# \tag{1'}
# \end{equation}
# $$
#
# $$
# \begin{equation}
# x^2+ 3x + 2m = 0
# \label{eq:pythagoras}
# \tag{2}
# \end{equation}
# $$
# (1') - (2)
# $$
# \begin{align*}
# x ^ 2 - x = 0 \\
# x(x - 1) = 0 \\
# x = 0, 1
# \end{align*}
# $$
# (2)の式
# $$
# \begin{align*}
# 0 ^ 2 + 3 * 0 + 2m = 0 (x=0)\\
# 1 ^ 2 + 3 * 1 + 2m = 0 (x=1)\\
# m = 0, -1
# \end{align*}
# $$
| markdown/python/anaconda/notebook/math/math.2018-09-11.10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
# $ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
# $ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
# $ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
# <h2> Basics of Python: Variables </h2>
#
# We review using variables in Python here.
#
# Please run each cell and check the results.
#
# <b> Indention of codes <u>matters</u> in Python!</b>
#
# In this notebook, each line of code should start from the left without any indention. Otherwise, you will get a syntax error.
#
# Comments can be indented.
#
# The codes belonging to a conditional or loop statement or a function/procedure are indented. We will see them later.
# +
# This is a comment
# A comment is used for explanations/descriptions/etc.
# Comments do not affect the programs
# +
# let's define an integer variable named a
a = 5
# let's print its value
print(a)
# +
# let's define three integer variables named a, b, and c
a = 2
b = 4
c = a + b # summation of a and b
# let's print their values together
print(a,b,c)
# a single space will automatically appear in between
# -
# let's print their values in reverse order
print(c,b,a)
# let's print their summation and multiplication
print(a+b+c,a*b*c)
# +
# let's define variables with string/text values
hw = "hello world" # we can use double quotes
hqw = 'hello quantum world' # we can use single quotes
# let's print them
print(hw)
print(hqw)
# -
# let's print them together by inserting another string in between
print(hw,"and",hqw)
# +
# let's concatenate a few strings
d = "Hello " + 'World' + " but " + 'Quantum ' + "World"
# let's print the result
print(d)
# -
# let's print numeric and string values together
print("a =",a,", b =",b,", a+b =",a+b)
# let's subtract two numbers
d = a-b
print(a,b,d)
# let's divide two numbers
d = a/b
print(a,b,d)
# let's divide integers over integers
# the result is always an integer (with possible integer remainder)
d = 33 // 6
print(d)
# +
# reminder/mod operator
r = 33 % 6
# 33 mod 6 = 3
# or when 33 is divided by 6 over integers, the reminder is 3
# 33 = 5 * 6 + 3
# let's print the result
print(r)
# +
# Booleen variables
t = True
f = False
# let's print their values
print(t,f)
# +
# print their negations
print(not t)
print("the negation of",t,"is",not t)
print(not f)
print("the negation of",f,"is",not f)
# +
# define a float variable
d = -3.4444
# let's print its value and its square
print(d, d * d)
# -
# Let's use parentheses in our expressions.
#
# $(23 * 13)-(11 * 15) $
#
# Here $*$ represents the multiplication operator
e = (23*13) - (11 * 15)
print(e)
# Let's consider a more complex expression.
#
# $ -3 * (123- 34 * 11 ) + 4 * (5+ (23 * 15) ) $
# +
# we can use more than one variable
# left is the variable for the left part of the expression
# we start with the multiplication inside the parentheses
left = 34*11
# we continue with the substruction inside the parentheses
# we reuse the variable named left
left = 123 - left
# we reuse left again for the multiplication with -3
left = -3 * left
# right is the variable for the right part of the expression
# we use the same idea here
right = 23 * 15
right = 5 + right
right = 4 * right
# at the end, we use left for the result
left = left + right
# let's print the result
print(left)
# -
# <h3> Task 1 </h3>
#
# Define three variables $n1$, $n2$, and $n3$, and set their values to $3$, $-4$, and $6$.
#
# Define a new variable $r1$, and set its value to $ (2 \cdot n1 + 3 \cdot n2) \cdot 2 - 5 \cdot n3 $, where $\cdot$ represents the multiplication operator.
#
# <i>The multiplication operator in python (and in many other programming languages) is *.</i>
#
# Then, print the value of $r1$.
#
# As you may verify it by yourself, the result should be $-42$.
#
# your solution is here
#
# <a href="Python08_Basics_Variables_Solutions.ipynb#task1">click for our solution</a>
# <h3> Task 2 </h3>
#
# By using the same variables (you may not need to define them again), please print the following value
# $$
# \dfrac{(n1-n2)\cdot(n2-n3)}{(n3-n1)\cdot(n3+1)}
# $$
#
# You should see $ -3.3333333333333335 $ as the outcome.
#
# your solution is here
#
# <a href="Python08_Basics_Variables_Solutions.ipynb#task2">click for our solution</a>
# <h3> Task 3 </h3>
#
# Define variables N and S, and set their values to your name and surname.
#
# Then, print the values of N and S with a prefix phrase "hello from the quantum world to".
#
# your solution is here
#
# <a href="Python08_Basics_Variables_Solutions.ipynb#task3">click for our solution</a>
| python/Python08_Basics_Variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Mission to Mars Scrapping
# **Step 1 - Scraping
# !pip install --user splinter
# !pip install selenium
# +
#import required dependencies and modules.
#import requests
#from requests_html import AsyncHTMLSession
#asession = AsyncHTMLSession()
from bs4 import BeautifulSoup
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.action_chains import ActionChains
import time
import pymongo
import requests
from bs4 import BeautifulSoup
import pandas as pd
# +
#from splinter import Browser
#browser = Browser()
# -
# ## 1.NASA Mars News
# Scrape the [NASA Mars News Site](https://mars.nasa.gov/news/) and collect the latest News Title and Paragraph Text. Assign the text to variables that you can reference later.
# +
# declaring element names to grab
article_header = 'slide'
content_title = 'content_title'
content_body = 'article_teaser_body'
wait_element = 'news'
mars_news_url = "https://mars.nasa.gov/news/"
driver = webdriver.Chrome(r'C:\Users\sonal\Documents\USCBootCamp\Drivers\chrome\chromedriver.exe')
driver.get(mars_news_url)
time.sleep(10)
try:
element=WebDriverWait(driver, 10).until(
EC.presence_of_element_located((By.ID,wait_element))
)
except:
pass
slide = driver.find_elements_by_class_name(article_header)[0]
# +
# Scrape Latest article title and teaser on NASA's Mars website
first_article_headline = slide.find_element_by_class_name(content_title).text
first_article_paragraph = slide.find_element_by_class_name(content_body).text
print(f"Title: {first_article_headline}")
print(f"Para: {first_article_paragraph}")
# -
# ## 2.JPL Mars Space Images - Featured Image
# +
featured_image_url = "https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars"
featured_image_response = requests.get(featured_image_url)
if featured_image_response.status_code == 200:
featured_image_html = featured_image_response.text
# +
featured_image_soup = BeautifulSoup(featured_image_html, "html.parser")
image_element = featured_image_soup.find("a", {"class": "button fancybox"}).get('data-fancybox-href')
featured_image_url = "https://www.jpl.nasa.gov" + image_element
# -
featured_image_url
# ## 3.Mars Weather
# Connect to url for Mars Weather Twitter page
weather_tweets_url = "https://twitter.com/marswxreport?lang=en"
driver = webdriver.Chrome(r'C:\Users\sonal\Documents\USCBootCamp\Drivers\chrome\chromedriver.exe')
driver.get(weather_tweets_url)
time.sleep(10)
#Get using Xpath
mars_weather = driver.find_element_by_xpath("/html/body/div/div/div/div[2]/main/div/div/div/div/div/div/div/div/div[2]/section/div/div/div[2]/div/div/div/article/div/div[2]/div[2]/div[2]/div[1]/div/span")
weather_tweet = mars_weather.text
# +
print(weather_tweet)
# -
# ## 4.Mars Facts
# URL of page to be scraped
mars_fact_url = "https://space-facts.com/mars/"
all_mars_fact_tables = pd.read_html(mars_fact_url)
df_mars_facts = all_mars_fact_tables[0]
df_mars_facts.columns = ["Measure", "Number"]
df_mars_facts
#Use Pandas to convert the data to a HTML table string.
facts_df_html = df_mars_facts.to_html()
facts_df_html_table=facts_df_html.replace("\n","")
facts_df_html_table
# ## 5.Mars Hemispheres
# +
mars_hemisphere_url = "https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars"
mars_hemisphere_response = requests.get(mars_hemisphere_url)
# +
if mars_hemisphere_response.status_code == 200:
mars_hemisphere_html = mars_hemisphere_response.text
# -
def get_full_image_link(link):
response = requests.get(link)
if response.status_code == 200:
html = response.text
soup = BeautifulSoup(html, "html.parser")
img_link = soup.find("a", text="Original").get("href")
title = soup.find("h2", class_="title").text.split(" Enhanced")[0]
return [title, img_link]
mars_hemisphere_soup = BeautifulSoup(mars_hemisphere_html, "html.parser")
link_elements = mars_hemisphere_soup.findAll("a", {"class": "itemLink product-item"})
full_page_links = [dict(title=get_full_image_link("https://astrogeology.usgs.gov" + le.get('href'))[0], img_url=get_full_image_link("https://astrogeology.usgs.gov" + le.get('href'))[1]) for le in link_elements]
full_page_links
| mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp linkcheck
# -
# # fastlinkcheck API
# > API for fast local and online link checking
#export
from fastcore.all import *
from html.parser import HTMLParser
from urllib.parse import urlparse,urlunparse
from fastcore.script import SCRIPT_INFO
import sys
# ## Find links in an HTML file
#export
class _HTMLParseAttrs(HTMLParser):
def reset(self):
super().reset()
self.found = set()
def handle_starttag(self, tag, attrs):
a = first(v for k,v in attrs if k in ("src","href"))
if a: self.found.add(a)
handle_startendtag = handle_starttag
#export
def get_links(fn):
"List of all links in file `fn`"
h = _HTMLParseAttrs()
h.feed(Path(fn).read_text())
return L(h.found)
# We can use `get_links` to parse an HTML file for different types of links. For example, this is the contents of `./example/broken_links/test.html`:
example = Path('_example/broken_links/test.html')
print(example.read_text())
# Calling `get_links` with the above file path will return a list of links:
links = get_links(example)
test_eq(set(links), {'test.js',
'//somecdn.com/doesntexist.html',
'http://www.bing.com','http://fastlinkcheck.com/test.html',
'/test'})
#export
def _local_url(u, root, host, fname):
"Change url `u` to local path if it is a local link"
fpath = Path(fname).parent
islocal=False
# remove `host` prefix
for o in 'http://','https://','http://www.','https://www.':
if host and u.startswith(o+host): u,islocal = remove_prefix(u, o+host),True
# remove params, querystring, and fragment
p = list(urlparse(u))[:5]+['']
# local prefix, or no protocol or host
if islocal or (not p[0] and not p[1]):
u = p[2]
if u and u[0]=='/': return (root/u[1:]).resolve()
else: return (fpath/u).resolve()
# URLs without a protocol are "protocol relative"
if not p[0]: p[0]='http'
# mailto etc are not checked
if p[0] not in ('http','https'): return ''
return urlunparse(p)
#export
class _LinkMap(dict):
"""A dict that pretty prints Links and their associated locations."""
def _repr_locs(self, k): return '\n'.join(f' - `{p}`' for p in self[k])
def __repr__(self):
rstr = L(f'\n- {k!r} was found in the following pages:\n{self._repr_locs(k)}' for k in self).concat()
return '\n'.join(rstr)
_repr_markdown_ = __repr__
#export
def local_urls(path:Path, host:str):
"returns a `dict` mapping all HTML files in `path` to a list of locally-resolved links in that file"
path=Path(path)
fns = L(path.glob('**/*.html'))+L(path.glob('**/*.htm'))
found = [(fn.resolve(),_local_url(link, root=path, host=host, fname=fn))
for fn in fns for link in get_links(fn)]
return _LinkMap(groupby(found, 1, 0))
# The keys of the `dict` returned by `local_urls` are links found in HTML files, and the values of this `dict` are a list of paths that those links are found in.
#
# Furthermore, local links are returned as `Path` objects, whereas external URLs are strings. For example, notice how the link:
#
#
# `http://fastlinkcheck.com/test.html`
#
#
# is resolved to a local path, because the `host` parameter supplied to `local_urls`, `fastlinkcheck.com` matches the url in the link:
path = Path('./_example/broken_links/')
links = local_urls(path, host='fastlinkcheck.com')
links
# ## Finding broken links
#export
def html_exists(o):
"If a path without a suffix is proivded, see if the same path with a .html suffix exists"
return (o.parent / (o.name + '.html') if not o.suffix else o).exists() or o.exists()
# the path `_example/broken_links/test` doesn't exist, but `_example/broken_links/test.html` does:
# +
p = Path("_example/broken_links/test")
assert not p.exists()
assert html_exists(p)
# -
# the path `_example/broken_links/really_doesnt_exist` and neither does `_example/broken_links/really_doesnt_exist.html`:
p = Path("_example/broken_links/really_doesnt_exist")
assert not p.exists()
assert not html_exists(p)
# Since `test.js` does not exist in the `example/` directory, `broken_local` returns this path:
#export
def broken_local(links, ignore_paths=None):
"List of items in keys of `links` that are `Path`s that do not exist"
ignore_paths = setify(ignore_paths)
return L(o for o in links if isinstance(o,Path) and o not in ignore_paths and not html_exists(o))
broken_local(links)
assert not all([x.exists() for x in broken_local(links)])
#export
def broken_urls(links, ignore_urls=None):
"List of items in keys of `links` that are URLs that return a failure status code"
ignore_urls = setify(ignore_urls)
its = L(o for o in links if isinstance(o, str) and o not in ignore_urls)
working_urls = parallel(urlcheck, its, n_workers=32, threadpool=True)
return L(o for o,p in zip(its,working_urls) if not p)
# Similarly the url `http://somecdn.com/doesntexist.html` doesn't exist, which is why it is returned by `broken_urls`
assert broken_urls(links) == ['http://somecdn.com/doesntexist.html']
#export
@call_parse
def link_check(path:Param("Root directory searched recursively for HTML files", str),
host:Param("Host and path (without protocol) of web server", str)=None,
config_file:Param("Location of file with urls to ignore",str)=None):
"Check for broken links recursively in `path`."
path = Path(path)
assert path.exists(), f"{path.absolute()} does not exist."
is_cli = (SCRIPT_INFO.func == 'link_check')
if config_file: assert Path(config_file).is_file(), f"{config_file} is either not a file or doesn't exist."
ignore = L(x.strip() for x in (Path(config_file).readlines() if config_file else ''))
links = local_urls(path, host=host)
ignore_paths = set((path/o).resolve() for o in ignore if not urlvalid(o))
ignore_urls = set(ignore.filter(urlvalid))
lm = _LinkMap({k:links[k] for k in (broken_urls(links, ignore_urls) + broken_local(links, ignore_paths))})
if is_cli and lm:
sys.stderr.write(f'\nERROR: The Following Broken Links or Paths were found:\n{lm}')
sys.exit(1)
if is_cli and not lm: print('No broken links found!')
return lm
link_check(path='_example/broken_links/', host='fastlinkcheck.com')
# Similarly if there are no broken links, `link_check` will not return any data. In this case, there are no broken links in the directory `_example/no_broken_links/`:
assert not link_check(path='_example/no_broken_links/')
# ### Ignore links with a configuration file
# You can choose to ignore files with a a plain-text file containing a list of urls to ignore. For example, the file `linkcheck.rc` contains a list of urls I want to ignore:
print((path/'linkcheck.rc').read_text())
# In this case `example/test.js` will be filtered out from the list:
link_check(path='_example/broken_links/', host='fastlinkcheck.com', config_file='_example/broken_links/linkcheck.rc')
# `link_check` can also be called use from the command line like this:
#
# > Note: the `!` command in Jupyter allows you [run shell commands](https://stackoverflow.com/questions/38694081/executing-terminal-commands-in-jupyter-notebook/48529220)
#
# The `-h` or `--help` flag will allow you to see the command line docs:
# !link_check -h
#hide
from nbdev.export import *
notebook2script()
| linkcheck.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#[Tutorial-Title]¶" data-toc-modified-id="[Tutorial-Title]¶-1"><span class="toc-item-num">1 </span>[Tutorial Title]¶</a></span><ul class="toc-item"><li><span><a href="#Introduction" data-toc-modified-id="Introduction-1.1"><span class="toc-item-num">1.1 </span>Introduction</a></span></li><li><span><a href="#Prerequisites" data-toc-modified-id="Prerequisites-1.2"><span class="toc-item-num">1.2 </span>Prerequisites</a></span></li><li><span><a href="#Initialization" data-toc-modified-id="Initialization-1.3"><span class="toc-item-num">1.3 </span>Initialization</a></span><ul class="toc-item"><li><span><a href="#Ensure-database-is-running" data-toc-modified-id="Ensure-database-is-running-1.3.1"><span class="toc-item-num">1.3.1 </span>Ensure database is running</a></span></li><li><span><a href="#Download-and-install-additional-components." data-toc-modified-id="Download-and-install-additional-components.-1.3.2"><span class="toc-item-num">1.3.2 </span>Download and install additional components.</a></span></li><li><span><a href="#Connect-to-database." data-toc-modified-id="Connect-to-database.-1.3.3"><span class="toc-item-num">1.3.3 </span>Connect to database.</a></span></li><li><span><a href="#Populate-database-with-test-data." data-toc-modified-id="Populate-database-with-test-data.-1.3.4"><span class="toc-item-num">1.3.4 </span>Populate database with test data.</a></span></li><li><span><a href="#Create-secondary-indexes-if-necessary" data-toc-modified-id="Create-secondary-indexes-if-necessary-1.3.5"><span class="toc-item-num">1.3.5 </span>Create secondary indexes if necessary</a></span></li></ul></li></ul></li><li><span><a href="#[Tutorial-Section]" data-toc-modified-id="[Tutorial-Section]-2"><span class="toc-item-num">2 </span>[Tutorial Section]</a></span><ul class="toc-item"><li><span><a href="#[Subsection-1]" data-toc-modified-id="[Subsection-1]-2.1"><span class="toc-item-num">2.1 </span>[Subsection 1]</a></span></li><li><span><a href="#[Subsection-2]" data-toc-modified-id="[Subsection-2]-2.2"><span class="toc-item-num">2.2 </span>[Subsection 2]</a></span></li></ul></li><li><span><a href="#[Next-Section]" data-toc-modified-id="[Next-Section]-3"><span class="toc-item-num">3 </span>[Next Section]</a></span><ul class="toc-item"><li><span><a href="#[Subsection-1]" data-toc-modified-id="[Subsection-1]-3.1"><span class="toc-item-num">3.1 </span>[Subsection 1]</a></span></li><li><span><a href="#[Subsection-2]" data-toc-modified-id="[Subsection-2]-3.2"><span class="toc-item-num">3.2 </span>[Subsection 2]</a></span></li></ul></li><li><span><a href="#Takeaways-and-Conclusion" data-toc-modified-id="Takeaways-and-Conclusion-4"><span class="toc-item-num">4 </span>Takeaways and Conclusion</a></span></li><li><span><a href="#Clean-up" data-toc-modified-id="Clean-up-5"><span class="toc-item-num">5 </span>Clean up</a></span></li><li><span><a href="#Further-Exploration-and-Resources" data-toc-modified-id="Further-Exploration-and-Resources-6"><span class="toc-item-num">6 </span>Further Exploration and Resources</a></span><ul class="toc-item"><li><span><a href="#Next-steps" data-toc-modified-id="Next-steps-6.1"><span class="toc-item-num">6.1 </span>Next steps</a></span></li></ul></li></ul></div>
# -
# # Implementing Read-Write Transactions with R-M-W Pattern
# This tutorial explains how to use the Read-Modify-Write pattern in order to ensure atomicity and isolation for read-write single-record transactions.
#
# This notebook requires Aerospike datbase running on localhost and that python and the Aerospike python client have been installed (`pip install aerospike`). Visit [Aerospike notebooks repo](https://github.com/aerospike-examples/interactive-notebooks) for additional details and the docker container.
# ## Introduction
# In Aerospike, the transactional boundaries are "single request, single record". While multiple operations may be specified in a single request on a single record, each such operation can involve a single bin and only certain write operations are allowed. Therefore, neither updates involving multiple bins (e.g, "a=a+b") nor general logic (e.g., "concatenate alternate letters and append") are possible as server-side operations. Of course, UDFs allow complex logic in a transactional update of a single record, however they are not suitable for all situations for various reasons such as performance and ease. Therefore most updates entail the R-M-W pattern or Reading the record, Modifying bins on the client side, and then Writing the record updates back to the server.
#
# The tutorial first demonstrates how read-write operations can result in lost writes in a concurrent multi-client environment.
#
# Then we show how to specify conditional writes with version check to address the problem by disallowing intereaved read-write and thus protecting against lost writes.
# ## Prerequisites
# This tutorial assumes familiarity with the following topics:
#
# [Provide topics and links. For example:]
# - [Hello World](hello_world.ipynb)
# - [Aerospike Basic Operations](basic_operations.ipynb)
# ## Initialization
# ### Ensure database is running
# This notebook requires that Aerospike datbase is running.
# [Include the right code cell for Java or Python from the two cells below.]
# !asd >& /dev/null
# !pgrep -x asd >/dev/null && echo "Aerospike database is running!" || echo "**Aerospike database is not running!**"
# ### Connect to database.
# +
# import the modules
import sys
from __future__ import print_function
import aerospike
# Configure the client
config = {
'hosts': [ ('127.0.0.1', 3000) ],
'policy' : {'key': aerospike.POLICY_KEY_SEND}
}
# Create a client and connect it to the cluster
try:
client = aerospike.client(config).connect()
except:
print("failed to connect to the cluster with", config['hosts'])
sys.exit(1)
print('Client successfully connected to the database.')
# -
# ### Populate database with test data.
# We create one record with an integer bin "gen-times-2" (the names will become clear below), initialized to 1.
# +
namespace = 'test'
tutorial_set = 'rmw-tutorial-set'
user_key = 'id-1'
# Records are addressable via a tuple of (namespace, set, user_key)
rec_key = (namespace, tutorial_set, user_key)
rmw_bin = 'gen-times-2'
try:
# Create the record
client.put(rec_key, {rmw_bin: 1})
except Exception as e:
print("error: {0}".format(e), file=sys.stderr)
print('Test data populated.')
# -
# # The Problem of Lost Writes
# In a concurrent setting, multiple clients may be performaing Read-Modify-Write on the same record in a way that get in each other's way. Since various R-M-W transactions can interleave, a transaction can be lost, if another client updates the record without reading the transaction's update.
#
# To demonstrate this, we make use of a record's "generation" or version, that is available as the record metadata, and is automatically incremented on each successful update of the record.
#
# The integer bin “gen-times-2” holds the value that is 2 times the value of the current generation of the record. A client first reads the current generation of the record, and then updates the bin value 2 times that value.
#
# In the case of a single client, there are no issues in maintaining the semantics of the bin. However when there are multiple clients, the interleaving of reads and writes of different transactions can violate the semantics. By updating the bin using an older generation value, it may not be 2 times the current generation, which is the constraint that we want to preserve.
#
# First, we will show how transaction writes are lost in a simple concurrent case by observing whether the relationship between record's current generation and the bin value is maintained. Then we will show how the problem is solved using a conditional write with version check.
#
# ## Test Framework
# We spawn multiple (num_threads) threads to simulate concurrent access. Each thread repeatedly (num_txns) does the following:
# - waits for a random duration (with average of txn_wait_ms)
# - executes a passed-in R-M-W function that returns the failure type (string, null if success).
#
# At the end the thread prints out the aggregate counts for each error type. In aggregate, they signify the likelihood of a read-write transaction failing.
# +
import threading
import time
import random
num_txns = 10
txn_wait_ms = 500
def thread_fn(thread_id, rmw_fn):
random.seed(thread_id)
lost_writes_count = 0
failures = {}
for i in range(num_txns):
failure = rmw_fn()
if failure:
if not failure in failures:
failures[failure] = 1
else:
failures[failure] += 1
print('\tThead {0} failures: {1}'.format(thread_id, failures))
return
def run_test(num_threads, rmw_fn):
threads = list()
print('{0} threads, {1} transcations per thread:'.format(num_threads, num_txns))
for thread_index in range(num_threads):
thread = threading.Thread(target=thread_fn, args=(thread_index, rmw_fn))
threads.append(thread)
thread.start()
for thread in threads:
thread.join()
return
# -
# ## Simple RMW Function
# Next we implement a simple RMW function simple_rmw_fn to pass into the above framework. The function:
# - Reads the record.
# - Computes new value of gen_times_2 (= 2 * read generation). Then waits for a random duration, with average of write_wait_ms average to simulate the application computation time between read and write.
# - Writes the new bin value. In the same (multi-op) request, reads back the record for the record's new generation value.
# - Returns "lost writes" if the updated value of gen_times_2/2 is smaller than the new gen. If they are the same, it returns None.
# +
write_wait_ms = 50
def rmw_simple():
#read
_, meta, bins = client.get(rec_key)
# wait before write to simulate computation time
time.sleep(random.uniform(0,2*write_wait_ms/1000.0))
# modify
read_gen = meta['gen']
new_rmw_bin_value = 2*(read_gen+1)
# write and read back bin_inc to compare
ops = [op_helpers.write(rmw_bin, new_rmw_bin_value),
op_helpers.read(rmw_bin)]
try:
_, meta, bins = client.operate(rec_key, ops)
except Exception as e:
print("error: {0}".format(e), file=sys.stderr)
exit(-1)
# compare new_rmw_bin_value//2 and new gen; if different return 'lost writes'
new_gen = meta['gen']
if new_rmw_bin_value//2 != new_gen:
#print('gen: {0}, bin: {1}, lost: {2}'.format(new_gen, new_rmw_bin_value//2, new_gen-new_rmw_bin_value//2))
return 'lost writes'
return None
# -
# ## Test Results
# For various values of concurrency (num_threads), we can see that with greater concurrent updates, a larger percentage of read-write transactions are lost, meaning greater likelihood of the semantics of the gen_times_2 bin not being preserved.
run_test(num_threads=1, rmw_fn=rmw_simple)
run_test(num_threads=2, rmw_fn=rmw_simple)
run_test(num_threads=3, rmw_fn=rmw_simple)
run_test(num_threads=4, rmw_fn=rmw_simple)
# # Using Generation Check
# To solve the problem of lost writes, the simple R-M-W is modified with how the Write is done: by making it conditional on the record not having been modified since the Read. It is a "check-and-set (CAS)" like operation that succeeds if the record generation (version) is still the same as at the time of Read. Otherwise it fails, and the client must retry the whole R-M-W pattern. The syntax and usage is shown in the code below.
# ## RMW Function with Version Check and Retries
# In the rmw_with_gen_check function below, a failed read-write due to generation mismatch is retried for max_retries attempts or until the write is successful. Each retry is attempted after a exponential backoff wait of (retry_number ** 2) * retry_wait_ms.
#
# A write can still fail after max_retries attempts, and the client can suitably handle it. However no writes are overwritten or lost, and the intended semantics of the gen-times-2 bin is always preserved.
#
# We perform the same concurrent test with the version check at Write. We expect no interleaved_writes reported in any thread.
# +
from aerospike_helpers.operations import operations as op_helpers
from aerospike import exception as ex
max_retries = 3
retry_wait_ms = 20
def rmw_with_gen_check():
retryRMWCount = 0
done = False
while (not done):
#read
_, meta, bins = client.get(rec_key)
# wait before write to simulate computation time
time.sleep(random.uniform(0,2*write_wait_ms/1000.0))
# modify
read_gen = meta['gen']
new_rmw_bin_value = 2*(read_gen+1)
# write and read back bin_inc to compare
ops = [op_helpers.write(rmw_bin, new_rmw_bin_value),
op_helpers.read(rmw_bin)]
write_policy = { 'gen': aerospike.POLICY_GEN_EQ }
try:
_, meta, bins = client.operate(rec_key, ops, meta={'gen': read_gen}, policy=write_policy)
except ex.RecordGenerationError as e:
if retryRMWCount < max_retries:
retryRMWCount += 1
time.sleep((2**retryRMWCount)*retry_wait_ms/1000.0)
else:
return 'max retries exceeded'
except Exception as e:
print("error: {0}".format(e), file=sys.stderr)
exit(-1)
else:
done = True
# compare new_rmw_bin_value//2 and new gen; if different
new_gen = meta['gen']
if new_rmw_bin_value//2 != new_gen:
return 'lost writes'
return None
# -
# ## Test Results
# Let's execute for various levels of concurrency and see the results. We expect to see no lost writes. Even when max-retries are exceeded, transaction and database integrity is preserved.
run_test(num_threads=2, rmw_fn=rmw_with_gen_check)
run_test(num_threads=3, rmw_fn=rmw_with_gen_check)
run_test(num_threads=4, rmw_fn=rmw_with_gen_check)
# # Takeaways
# In the tutorial we showed:
# - the need for read-write transactions in Aerospike to use the R-M-W pattern
# - how writes can be overwritten and lost in a concurrent environment if performed simply
# - how the developer can ensure atomicity and isolation of a read-write transaction by using version check logic and syntax.
# # Clean up
# Remove data and close connection.
client.truncate(namespace, tutorial_set, 0)
# Close the connection to the Aerospike cluster
client.close()
print('Removed tutorial data. Connection closed.')
# # Further Exploration and Resources
# For further exploration of transactions support in Aerospike, check out the following resources:
#
# - Blog posts
# - [Developers: Understanding Aerospike Transactions](https://www.aerospike.com/blog/developers-understanding-aerospike-transactions/)
# - [Twelve Do's of Consistency in Aerospike](https://www.aerospike.com/blog/twelve-dos-of-consistency-in-aerospike/)
# - Video
# - [Strong Consistency in Databases. What does it actually guarantee?](https://www.aerospike.com/resources/videos/strong-consistency-in-databases-what-does-it-actually-guarantee/)
# ## Next steps
#
# Visit [Aerospike notebooks repo](https://github.com/aerospike-examples/interactive-notebooks) to run additional Aerospike notebooks. To run a different notebook, download the notebook from the repo to your local machine, and then click on File->Open, and select Upload.
| notebooks/transactions_rmw_pattern.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3c97b0a2-00f9-46fa-8e3d-661a6a5ef59d", "showTitle": false, "title": ""}
# ##Exercise Overview
# In this exercise we will play with Spark [Datasets & Dataframes](https://spark.apache.org/docs/latest/sql-programming-guide.html#datasets-and-dataframes), some [Spark SQL](https://spark.apache.org/docs/latest/sql-programming-guide.html#sql), and build a couple of binary classifiaction models using [Spark ML](https://spark.apache.org/docs/latest/ml-guide.html) (with some [MLlib](https://spark.apache.org/mllib/) too).
# <br><br>
# The set up and approach will not be too dissimilar to the standard type of approach you might do in [Sklearn](http://scikit-learn.org/stable/index.html). Spark has matured to the stage now where for 90% of what you need to do (when analysing tabular data) should be possible with Spark dataframes, SQL, and ML libraries. This is where this exercise is mainly trying to focus.
# <br>
# Feel free to adapt this exercise to play with other datasets readily availabe in the Databricks enviornment (they are listed in a cell below).
# #####Getting Started
# To get started you will need to create and attach a databricks spark cluster to this notebook. This notebook was developed on a cluster created with:
# - Databricks Runtime Version 4.0 (includes Apache Spark 2.3.0, Scala 2.11)
# - Python Version 3
#
# #####Links & References
#
# Some useful links and references of sources used in creating this exercise:
#
# **Note**: Right click and open as new tab!
# <br>
# 1. [Latest Spark Docs](https://spark.apache.org/docs/latest/index.html)
# 1. [Databricks Homepage](https://databricks.com/)
# 1. [Databricks Community Edition FAQ](https://databricks.com/product/faq/community-edition)
# 1. [Databricks Self Paced Training](https://databricks.com/training-overview/training-self-paced)
# 1. [Databricks Notebook Guide](https://docs.databricks.com/user-guide/notebooks/index.html)
# 1. [Databricks Binary Classification Tutorial](https://docs.databricks.com/spark/latest/mllib/binary-classification-mllib-pipelines.html#binary-classification)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "695a1fed-7ad1-4a91-bd70-46e0275a793e", "showTitle": false, "title": ""}
# ####Get Data
#
# Here we will pull in some sample data that is already pre-loaded onto all databricks clusters.
#
# Feel free to adapt this notebook later to play around with a different dataset if you like (all available are listed in a cell below).
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "22e9b324-2b85-41ea-b2a9-cc7a9aa97fa3", "showTitle": false, "title": ""}
# Help for completing this assignment came from my mentor
# display datasets already in databricks
display(dbutils.fs.ls("/databricks-datasets")) #dbutils is not defined on the local host. It works fine with Databricks!
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c7388cb6-bf4f-44d6-a7cd-50d35e20a22d", "showTitle": false, "title": ""}
# Lets take a look at the '**adult**' dataset on the filesystem. This is the typical US Census data you often see online in tutorials. [Here](https://archive.ics.uci.edu/ml/datasets/adult) is the same data in the UCI repository.
#
# _As an aside: [here](https://github.com/GoogleCloudPlatform/cloudml-samples/tree/master/census) this same dataset is used as a quickstart example for Google CLoud ML & Tensorflow Estimator API (in case youd be interested in playing with tensorflow on the same dataset as here)._
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "db143421-89ed-4e9b-9272-0ddf8f2a4fe9", "showTitle": false, "title": ""}
# %fs ls databricks-datasets/adult/adult.data
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6df40add-37cc-47ff-adc1-3eb0852a5afc", "showTitle": false, "title": ""}
# **Note**: Above %fs is just some file system cell magic that is specific to databricks. More info [here](https://docs.databricks.com/user-guide/notebooks/index.html#mix-languages).
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "84d3180c-0f4b-4375-8f97-7975ca99ef77", "showTitle": false, "title": ""}
# ####Spark SQL
# Below we will use Spark SQL to load in the data and then register it as a Dataframe aswell. So the end result will be a Spark SQL table called _adult_ and a Spark Dataframe called _df_adult_.
# <br><br>
# This is an example of the flexibility in Spark in that you could do lots of you ETL and data wrangling using either Spark SQL or Dataframes and pyspark. Most of the time it's a case of using whatever you are most comfortable with.
# <br><br>
# When you get more advanced then you might looking the pro's and con's of each and when you might favour one or the other (or operating direclty on RDD's), [here](https://databricks.com/blog/2016/07/14/a-tale-of-three-apache-spark-apis-rdds-dataframes-and-datasets.html) is a good article on the issues. For now, no need to overthink it!
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c1ab6a7a-748b-49a7-a1dc-afdccd2695c8", "showTitle": false, "title": ""}
# %sql
-- drop the table if it already exists
DROP TABLE IF EXISTS adult
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f22f5e37-a6e4-4d5c-863e-61e9fda375f7", "showTitle": false, "title": ""}
# %sql
-- create a new table in Spark SQL from the datasets already loaded in the underlying filesystem.
-- In the real world you might be pointing at a file on HDFS or a hive table etc.
CREATE TABLE adult (
age DOUBLE,
workclass STRING,
fnlwgt DOUBLE,
education STRING,
education_num DOUBLE,
marital_status STRING,
occupation STRING,
relationship STRING,
race STRING,
sex STRING,
capital_gain DOUBLE,
capital_loss DOUBLE,
hours_per_week DOUBLE,
native_country STRING,
income STRING)
USING com.databricks.spark.csv
OPTIONS (path "/databricks-datasets/adult/adult.data", header "true")
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b73f000c-7068-4d9c-a559-18dc7bda95f0", "showTitle": false, "title": ""}
# look at the data
#spark.sql("SELECT * FROM adult LIMIT 5").show()
# this will look prettier in Databricks if you use display() instead
display(spark.sql("SELECT * FROM adult LIMIT 5"))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7b41c5e9-3101-41ba-a7e9-9e84399f3150", "showTitle": false, "title": ""}
# If you are more comfortable with SQL then as you can see below, its very easy to just get going with writing standard SQL type code to analyse your data, do data wrangling and create new dataframes.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1219e3d2-a3ff-4d81-b688-c6751f950822", "showTitle": false, "title": ""}
# Lets get some summary marital status rates by occupation
result = spark.sql(
"""
SELECT
occupation,
SUM(1) as n,
ROUND(AVG(if(LTRIM(marital_status) LIKE 'Married-%',1,0)),2) as married_rate,
ROUND(AVG(if(lower(marital_status) LIKE '%widow%',1,0)),2) as widow_rate,
ROUND(AVG(if(LTRIM(marital_status) = 'Divorced',1,0)),2) as divorce_rate,
ROUND(AVG(if(LTRIM(marital_status) = 'Separated',1,0)),2) as separated_rate,
ROUND(AVG(if(LTRIM(marital_status) = 'Never-married',1,0)),2) as bachelor_rate
FROM
adult
GROUP BY 1
ORDER BY n DESC
""")
display(result)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ce745db8-1b57-47cd-976b-4ce3ee440604", "showTitle": false, "title": ""}
# You can easily register dataframes as a table for Spark SQL too. So this way you can easily move between Dataframes and Spark SQL for whatever reason.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9007db35-22d8-4432-bcea-9bbb90017fa3", "showTitle": false, "title": ""}
# register the df we just made as a table for spark sql
sqlContext.registerDataFrameAsTable(result, "result")
spark.sql("SELECT * FROM result").show(5)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "24bbcd7e-55ec-4929-a898-09a27acf9269", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 1</span>
#
# 1. Write some spark sql to get the top 'bachelor_rate' by 'education' group?
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "964b6472-2d89-4158-b517-0955f1e3c587", "showTitle": false, "title": ""}
### Question 1.1 Answer ###
# register the df we just made as a table for spark sql
def run_sql(statement):
try:
result = sqlContext.sql(statement)
except Exception as e:
print(e.desc, '\n', e.stackTrace)
return
return result
sqlContext.registerDataFrameAsTable(result, "result")
tbls = run_sql('show tables')
tbls.toPandas()
result = run_sql(
"""
SELECT
education,
SUM(1) as n,
ROUND(AVG(if(LTRIM(marital_status) = 'Never-married',1,0)),2) as bachelor_rate
FROM
adult
GROUP BY 1
ORDER BY n DESC
""")
result.limit(5).toPandas()
#result = # fill in here
result.show()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "82050620-e0ac-45a1-96ac-11ddce045cb9", "showTitle": false, "title": ""}
# ####Spark DataFrames
# Below we will create our DataFrame from the SQL table and do some similar analysis as we did with Spark SQL but using the DataFrames API.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cbad390a-677d-4b3b-8e01-9dd7bb211ab2", "showTitle": false, "title": ""}
# register a df from the sql df
df_adult = spark.table("adult")
cols = df_adult.columns # this will be used much later in the notebook, ignore for now
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "da26c930-94a3-4289-ab00-61f5b283f9c5", "showTitle": false, "title": ""}
# look at df schema
df_adult.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0443d5cc-85c8-462c-8998-0334ac36d6c1", "showTitle": false, "title": ""}
# look at the df
display(df_adult.show(5))
#df_adult.show(5)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c6f3c980-fda5-4a86-8a83-75846e041b74", "showTitle": false, "title": ""}
# Below we will do a similar calculation to what we did above but using the DataFrames API
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "460c47b6-0405-4ce6-8015-39625fbc514a", "showTitle": false, "title": ""}
# import what we will need
from pyspark.sql.functions import when, col, mean, desc, round
# wrangle the data a bit
df_result = df_adult.select(
df_adult['occupation'],
# create a 1/0 type col on the fly
when( col('marital_status') == ' Divorced' , 1 ).otherwise(0).alias('is_divorced')
)
# do grouping (and a round)
df_result = df_result.groupBy('occupation').agg(round(mean('is_divorced'),2).alias('divorced_rate'))
# do ordering
df_result = df_result.orderBy(desc('divorced_rate'))
# show results
df_result.show(5)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2e688a7f-a03c-4c74-94ae-42d999aca3bc", "showTitle": false, "title": ""}
# As you can see the dataframes api is a bit more verbose then just expressing what you want to do in standard SQL.<br><br>But some prefer it and might be more used to it, and there could be cases where expressing what you need to do might just be better using the DataFrame API if it is too complicated for a simple SQL expression for example of maybe involves recursion of some type.
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "252f073d-4272-4703-886a-4b37c9cd47f6", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 2</span>
# 1. Write some pyspark to get the top 'bachelor_rate' by 'education' group using DataFrame operations?
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7d5c2957-a783-47ec-bcfd-a769ba0cf294", "showTitle": false, "title": ""}
# wrangle the data
df_result = df_adult.select(
df_adult['education'],
# create a yes/no type col on the fly
when( col('marital_status') == ' Never-married' , 1 ).otherwise(0).alias('is_bachelor')
)
# do grouping
df_result = df_result.groupBy('education').agg(round(mean('is_bachelor'),2).alias('bachelor_rate'))
# do ordering
df_result = df_result.orderBy(desc('bachelor_rate'))
# show result
df_result.show(1)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "99f244c2-4b2e-4727-8c65-66cbd75a69f8", "showTitle": false, "title": ""}
# ####Explore & Visualize Data
# It's very easy to [collect()](https://spark.apache.org/docs/latest/rdd-programming-guide.html#printing-elements-of-an-rdd) your Spark DataFrame data into a Pandas df and then continue to analyse or plot as you might normally.
# <br><br>
# Obviously if you try to collect() a huge DataFrame then you will run into issues, so usually you would only collect aggregated or sampled data into a Pandas df.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7640afab-8c43-46e2-91c6-43b5834e1a23", "showTitle": false, "title": ""}
import pandas as pd
# do some analysis
result = spark.sql(
"""
SELECT
occupation,
AVG(IF(income = ' >50K',1,0)) as plus_50k
FROM
adult
GROUP BY 1
ORDER BY 2 DESC
""")
# collect results into a pandas df
df_pandas = pd.DataFrame(
result.collect(),
columns=result.schema.names
)
# look at df
print(df_pandas.head())
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5e7e5b12-632a-4dd9-b66b-6fb03ea902d4", "showTitle": false, "title": ""}
print(df_pandas.describe())
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a71f0594-35e9-480b-b3c9-7793765e3af2", "showTitle": false, "title": ""}
print(df_pandas.info())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b1090b4c-4789-4b83-84c7-6150b8158026", "showTitle": false, "title": ""}
# Here we will just do some very basic plotting to show how you might collect what you are interested in into a Pandas DF and then just plot any way you normally would.
#
# For simplicity we are going to use the plotting functionality built into pandas (you could make this a pretty as you want).
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "946501d8-b915-4fbd-b9cb-a0a31a802078", "showTitle": false, "title": ""}
import matplotlib.pyplot as plt
# i like ggplot style
plt.style.use('ggplot')
# get simple plot on the pandas data
myplot = df_pandas.plot(kind='barh', x='occupation', y='plus_50k')
# display the plot (note - display() is a databricks function -
# more info on plotting in Databricks is here: https://docs.databricks.com/user-guide/visualizations/matplotlib-and-ggplot.html)
display(myplot.figure)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1c9aafee-4f84-4f93-96ac-70a73bc2a588", "showTitle": false, "title": ""}
# You can also easily get summary stats on a Spark DataFrame like below. [Here](https://databricks.com/blog/2015/06/02/statistical-and-mathematical-functions-with-dataframes-in-spark.html) is a nice blog post that has more examples.<br><br>So this is an example of why you might want to move from Spark SQL into DataFrames API as being able to just call describe() on the Spark DF is easier then trying to do the equivilant in Spark SQL.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b2fa9c46-ff91-4424-ad28-9dd4f74f078e", "showTitle": false, "title": ""}
# describe df
df_adult.select(df_adult['age'],df_adult['education_num']).describe().show()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3e7bfbde-68e3-47a3-9f3e-278bce83808b", "showTitle": false, "title": ""}
# ### ML Pipeline - Logistic Regression vs Random Forest
#
# Below we will create two [Spark ML Pipelines](https://spark.apache.org/docs/latest/ml-pipeline.html) - one that fits a logistic regression and one that fits a random forest. We will then compare the performance of each.
#
# **Note**: A lot of the code below is adapted from [this example](https://docs.databricks.com/spark/latest/mllib/binary-classification-mllib-pipelines.html).
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "73a32ba3-ff2c-4ca0-91ca-8e6187f24363", "showTitle": false, "title": ""}
from pyspark.ml import Pipeline
from pyspark.ml.feature import OneHotEncoder, StringIndexer, VectorAssembler
categoricalColumns = ["workclass", "education", "marital_status", "occupation", "relationship", "race", "sex", "native_country"]
stages = [] # stages in our Pipeline
for categoricalCol in categoricalColumns:
# Category Indexing with StringIndexer
stringIndexer = StringIndexer(inputCol=categoricalCol, outputCol=categoricalCol + "Index")
# Use OneHotEncoder to convert categorical variables into binary SparseVectors
# encoder = OneHotEncoderEstimator(inputCol=categoricalCol + "Index", outputCol=categoricalCol + "classVec")
encoder = OneHotEncoder(inputCols=[stringIndexer.getOutputCol()], outputCols=[categoricalCol + "classVec"])
# Add stages. These are not run here, but will run all at once later on.
stages += [stringIndexer, encoder]
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f2657356-694c-4a30-80bd-02ede201a55c", "showTitle": false, "title": ""}
# Convert label into label indices using the StringIndexer
label_stringIdx = StringIndexer(inputCol="income", outputCol="label")
stages += [label_stringIdx]
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5a2fa539-5109-4b69-bc89-e7a7439a1469", "showTitle": false, "title": ""}
# Transform all features into a vector using VectorAssembler
numericCols = ["age", "fnlwgt", "education_num", "capital_gain", "capital_loss", "hours_per_week"]
assemblerInputs = [c + "classVec" for c in categoricalColumns] + numericCols
assembler = VectorAssembler(inputCols=assemblerInputs, outputCol="features")
stages += [assembler]
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c96a56cc-57f4-4c4b-856e-f2e5701e95fe", "showTitle": false, "title": ""}
# Create a Pipeline.
pipeline = Pipeline(stages=stages)
# Run the feature transformations.
# - fit() computes feature statistics as needed.
# - transform() actually transforms the features.
pipelineModel = pipeline.fit(df_adult)
dataset = pipelineModel.transform(df_adult)
# Keep relevant columns
selectedcols = ["label", "features"] + cols
dataset = dataset.select(selectedcols)
display(dataset.show(5))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0a5e74a5-dff1-45a3-9155-b9944d95e9aa", "showTitle": false, "title": ""}
### Randomly split data into training and test sets. set seed for reproducibility
(trainingData, testData) = dataset.randomSplit([0.7, 0.3], seed=100)
print(trainingData.count())
print(testData.count())
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c8de9b08-6ff6-4b6a-b67f-a78f2db52ccf", "showTitle": false, "title": ""}
from pyspark.sql.functions import avg
# get the rate of the positive outcome from the training data to use as a threshold in the model
training_data_positive_rate = trainingData.select(avg(trainingData['label'])).collect()[0][0]
print("Positive rate in the training data is {}".format(training_data_positive_rate))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5fb85c94-76bc-42aa-82b4-d00e6369e6bc", "showTitle": false, "title": ""}
# ####Logistic Regression - Train
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "06e5691c-1522-4b7e-a914-57b1bfa7a3a5", "showTitle": false, "title": ""}
from pyspark.ml.classification import LogisticRegression
# Create initial LogisticRegression model
lr = LogisticRegression(labelCol="label", featuresCol="features", maxIter=10)
# set threshold for the probability above which to predict a 1
lr.setThreshold(training_data_positive_rate)
# lr.setThreshold(0.5) # could use this if knew you had balanced data
# Train model with Training Data
lrModel = lr.fit(trainingData)
# get training summary used for eval metrics and other params
lrTrainingSummary = lrModel.summary
# Find the best model threshold if you would like to use that instead of the empirical positve rate
fMeasure = lrTrainingSummary.fMeasureByThreshold
maxFMeasure = fMeasure.groupBy().max('F-Measure').select('max(F-Measure)').head()
lrBestThreshold = fMeasure.where(fMeasure['F-Measure'] == maxFMeasure['max(F-Measure)']) \
.select('threshold').head()['threshold']
print("Best threshold based on model performance on training data is {}".format(lrBestThreshold))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "50f512e9-7d60-427d-8f17-530e08639605", "showTitle": false, "title": ""}
# ####GBM - Train
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e4b1128b-d223-4d3d-b202-5ad887e9cda3", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 3</span>
# 1. Train a GBTClassifier on the training data, call the trained model 'gbModel'
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "998d8fa4-1162-4f46-bd6a-40d05beadf31", "showTitle": false, "title": ""}
### Question 3.1 Answer ###
from pyspark.ml.classification import GBTClassifier
# Create initial LogisticRegression model
gb = GBTClassifier(labelCol="label", featuresCol="features", maxIter=10)
# Train model with Training Data
gbModel = gb.fit(trainingData)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f52bee49-0815-4252-a35f-d6a9b8b52be2", "showTitle": false, "title": ""}
# ####Logistic Regression - Predict
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6cb54547-1466-46ac-81bc-a9ae5d32dc5c", "showTitle": false, "title": ""}
# make predictions on test data
lrPredictions = lrModel.transform(testData)
# display predictions
display(lrPredictions.select("label", "prediction", "probability").show(5))
#display(lrPredictions)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c2b53bb9-d298-4af9-bdc8-fd94a5f0b388", "showTitle": false, "title": ""}
# ###GBM - Predict
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5f2f8c8c-070a-466c-a315-d26562880c66", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 4</span>
# 1. Get predictions on the test data for your GBTClassifier. Call the predictions df 'gbPredictions'.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "88af5d4c-6bc4-45c0-a4d5-39656e5aea3a", "showTitle": false, "title": ""}
### Question 4.1 Answer ###
# make predictions on test data
gbPredictions = gbModel.transform(testData)
display(gbPredictions.show(5))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "16eabc78-ad9f-4a58-a66d-b8bdc6669417", "showTitle": false, "title": ""}
# ####Logistic Regression - Evaluate
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f3194131-7989-4854-b39a-622c6ef6f2cd", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 5</span>
# 1. Complete the print_performance_metrics() function below to also include measures of F1, Precision, Recall, False Positive Rate and True Positive Rate.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e20e5588-1c65-428e-b125-9f2cb6ed8a0d", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.mllib.evaluation import BinaryClassificationMetrics, MulticlassMetrics
def print_performance_metrics(predictions):
# Evaluate model
evaluator = BinaryClassificationEvaluator(rawPredictionCol="rawPrediction")
auc = evaluator.evaluate(predictions, {evaluator.metricName: "areaUnderROC"})
aupr = evaluator.evaluate(predictions, {evaluator.metricName: "areaUnderPR"})
print("auc = {}".format(auc))
print("aupr = {}".format(aupr))
# get rdd of predictions and labels for mllib eval metrics
predictionAndLabels = predictions.select("prediction","label").rdd
# Instantiate metrics objects
binary_metrics = BinaryClassificationMetrics(predictionAndLabels)
multi_metrics = MulticlassMetrics(predictionAndLabels)
# Area under precision-recall curve
print("Area under PR = {}".format(binary_metrics.areaUnderPR))
# Area under ROC curve
print("Area under ROC = {}".format(binary_metrics.areaUnderROC))
# Accuracy
print("Accuracy = {}".format(multi_metrics.accuracy))
# Confusion Matrix
print(multi_metrics.confusionMatrix())
### Question 5.1 Answer ###
# F1
print("F1 = {}".format(multi_metrics.weightedFMeasure(1.0)))
# Precision
print("Precision = {}".format(multi_metrics.weightedPrecision))
# Recall
print("Recall = {}".format(multi_metrics.weightedRecall))
# FPR
print("FPR = {}".format(multi_metrics.weightedFalsePositiveRate))
# TPR
print("TPR = {}".format(multi_metrics.weightedTruePositiveRate))
print_performance_metrics(lrPredictions)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "21aa0bfe-9510-470d-9692-7c252c9bc58f", "showTitle": false, "title": ""}
# ####GBM - Evaluate
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "111e82b8-8470-42d0-9485-5fdb59912e56", "showTitle": false, "title": ""}
print_performance_metrics(gbPredictions)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c9d7ac8b-69d9-47d4-9412-9342faad7c05", "showTitle": false, "title": ""}
# ## Cross Validation
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "766d216e-da4c-4d99-8693-44c74ef75370", "showTitle": false, "title": ""}
# For each model you can run the below comand to see its params and a brief explanation of each.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "dcfef6fc-f469-4556-bebc-9ed17b605b8f", "showTitle": false, "title": ""}
print(lr.explainParams())
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "077280d3-39ad-4b29-b97e-ca45993c7527", "showTitle": false, "title": ""}
print(gb.explainParams())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5daa3f89-6335-47e0-96e6-b1e0c43a3aea", "showTitle": false, "title": ""}
# ####Logisitic Regression - Param Grid
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0d0ad441-c23a-484b-81e4-14b7fbf0b27e", "showTitle": false, "title": ""}
from pyspark.ml.tuning import ParamGridBuilder, CrossValidator
# Create ParamGrid for Cross Validation
lrParamGrid = (ParamGridBuilder()
.addGrid(lr.regParam, [0.01, 0.5, 2.0])
.addGrid(lr.elasticNetParam, [0.0, 0.5, 1.0])
.addGrid(lr.maxIter, [2, 5])
.build())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b72cc844-4efe-498a-aff0-18403c629de6", "showTitle": false, "title": ""}
# ####GBM - Param Grid
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "47696aef-d15e-4885-94aa-e263c7a56121", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 6</span>
# 1. Build out a param grid for the gb model, call it 'gbParamGrid'.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "34b95f49-81f1-4edc-949d-e453c6214d22", "showTitle": false, "title": ""}
### Question 6.1 Answer ###
# Create ParamGrid for Cross Validation
gbParamGrid = (ParamGridBuilder()
.addGrid(gb.maxDepth, [5, 10])
.addGrid(gb.maxIter, [2, 5])
.build())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cb7eecc2-5510-4563-8fc1-ec0f8efa7ecd", "showTitle": false, "title": ""}
# ####Logistic Regression - Perform Cross Validation
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9d2856c2-cec9-43e5-8b42-61f93429b97f", "showTitle": false, "title": ""}
# set up an evaluator
evaluator = BinaryClassificationEvaluator(rawPredictionCol="rawPrediction")
# Create CrossValidator
lrCv = CrossValidator(estimator=lr, estimatorParamMaps=lrParamGrid, evaluator=evaluator, numFolds=2)
# Run cross validations
lrCvModel = lrCv.fit(trainingData)
# this will likely take a fair amount of time because of the amount of models that we're creating and testing
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "919f51ca-6b9d-43ac-a32b-16bf811f3a0d", "showTitle": false, "title": ""}
# below approach to getting at the best params from the best cv model taken from:
# https://stackoverflow.com/a/46353730/1919374
# look at best params from the CV
print(lrCvModel.bestModel._java_obj.getRegParam())
print(lrCvModel.bestModel._java_obj.getElasticNetParam())
print(lrCvModel.bestModel._java_obj.getMaxIter())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ea6bd5d5-df8c-4eaa-aa24-7dcc2e7627cc", "showTitle": false, "title": ""}
# ####GBM - Perform Cross Validation
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fb2030b9-f96b-4402-b6cf-ab05effc3944", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 7</span>
# 1. Perform cross validation of params on your 'gb' model.
# 1. Print out the best params you found.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "98a9e5e6-449d-4f6e-8fee-3fc3e1e56c2f", "showTitle": false, "title": ""}
### Question 7.1 Answer ###
# Create CrossValidator
gbCv = CrossValidator(estimator=gb, estimatorParamMaps=gbParamGrid, evaluator=evaluator, numFolds=2)
# Run cross validations
gbCvModel = gbCv.fit(trainingData)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "60e1eff8-e84e-4a8b-bab1-3b5ce2e5e9f4", "showTitle": false, "title": ""}
### Question 7.2 Answer ###
# look at best params from the CV
print(gbCvModel.bestModel._java_obj.getMaxDepth())
print(gbCvModel.bestModel._java_obj.getMaxIter())
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "efd7e902-4c5c-48ca-8dfc-93a8ca1192e0", "showTitle": false, "title": ""}
# ####Logistic Regression - CV Model Predict
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "402209de-a5da-4195-ac8a-fe20fb1c9b87", "showTitle": false, "title": ""}
# Use test set to measure the accuracy of our model on new data
lrCvPredictions = lrCvModel.transform(testData)
display(lrCvPredictions.show(5))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cff1a654-95f5-4e08-9705-76e133141de7", "showTitle": false, "title": ""}
# ####GBM - CV Model Predict
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b7e0cddb-99ed-4fe9-9c46-ed9dc12f5044", "showTitle": false, "title": ""}
gbCvPredictions = gbCvModel.transform(testData)
display(gbCvPredictions.show(5))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f9db16de-2dcd-45ee-9e00-50009530a361", "showTitle": false, "title": ""}
# ####Logistic Regression - CV Model Evaluate
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "30e58e24-4d40-4c50-bbb0-73e059994495", "showTitle": false, "title": ""}
print_performance_metrics(lrCvPredictions)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f959c275-36a7-42a3-b1b7-551e28e651c9", "showTitle": false, "title": ""}
# ####GBM - CV Model Evaluate
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "62cfc105-53b6-4690-b732-e6446b965087", "showTitle": false, "title": ""}
print_performance_metrics(gbCvPredictions)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "39417456-a688-48c7-9aa4-58801fbf147e", "showTitle": false, "title": ""}
# ####Logistic Regression - Model Explore
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f567014e-5897-4b4f-b8a5-0aa1b536fe9c", "showTitle": false, "title": ""}
print('Model Intercept: ', lrCvModel.bestModel.intercept)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c5f8f9be-7672-4cee-83e1-3f72e632a07f", "showTitle": false, "title": ""}
lrWeights = lrCvModel.bestModel.coefficients
lrWeights = [(float(w),) for w in lrWeights] # convert numpy type to float, and to tuple
lrWeightsDF = sqlContext.createDataFrame(lrWeights, ["Feature Weight"])
display(lrWeightsDF)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "857e8f3b-45fb-4af6-b8ce-dd30d3190095", "showTitle": false, "title": ""}
# ### Feature Importance
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "03af6259-aa64-4309-a2a8-6207c8675e26", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 8</span>
# 1. Print out a table of feature_name and feature_coefficient from the Logistic Regression model.
# <br><br>
# Hint: Adapt the code from here: https://stackoverflow.com/questions/42935914/how-to-map-features-from-the-output-of-a-vectorassembler-back-to-the-column-name
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "eb525282-a308-446e-a14a-37fae0a9019d", "showTitle": false, "title": ""}
### Question 8.1 Answer ###
# fill in here
# from: https://stackoverflow.com/questions/42935914/how-to-map-features-from-the-output-of-a-vectorassembler-back-to-the-column-name
from itertools import chain
attrs = sorted(
(attr["idx"], attr["name"]) for attr in (chain(*lrCvPredictions
.schema[lrCvModel.bestModel.summary.featuresCol]
.metadata["ml_attr"]["attrs"].values())))
gbCvFeatureCoefficient = pd.DataFrame([(name, gbCvModel.bestModel.featureImportances[idx]) for idx, name in attrs],columns=['feature_name','feature_coefficient'])
print(gbCvFeatureCoefficient.sort_values(by=['feature_coefficient'],ascending =False))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "432159f9-6383-42eb-8ced-489f722d68e0", "showTitle": false, "title": ""}
gbCvFeatureCoefficient = pd.DataFrame([(name, gbCvModel.bestModel.featureImportances[idx]) for idx, name in attrs],columns=['feature_name','feature_coefficient'])
print(gbCvFeatureCoefficient.sort_values(by=['feature_coefficient'],ascending =False))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "bd47490f-2880-4b55-9b80-e74faa0a7fef", "showTitle": false, "title": ""}
# ####<span style="color:darkblue">Question 9</span>
# 1. Build and train a RandomForestClassifier and print out a table of feature importances from it.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9d63f4af-4566-4657-8f7e-a6b848f03356", "showTitle": false, "title": ""}
### Question 9.1 Answer ###
from pyspark.ml.classification import RandomForestClassifier
rf = RandomForestClassifier(labelCol="label", featuresCol="features")
rfModel = rf.fit(trainingData)
rfFeatureImportance = pd.DataFrame([(name, rfModel.featureImportances[idx]) for idx, name in attrs],columns=['feature_name','feature_importance'])
print(rfFeatureImportance.sort_values(by=['feature_importance'],ascending=False))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "467705fd-5bd3-4d1e-a57b-95a0b8156235", "showTitle": false, "title": ""}
| Spark_DF_SQL_ML_Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from numpy import array, median
from hw import Jamshidian as jamsh
from hw import Henrard as henr
from hw import calibration as hw_calib
from hw.const import *
from fox_toolbox.utils import rates
from fox_toolbox.utils import xml_parser
from fox_toolbox.utils.rates import RateCurve
from tsr import csv_parser, tsr
from random import choice
import matplotlib.pyplot as plt
from tsr import linear
from copy import deepcopy
import seaborn as sns
sns.set_context("poster")
sns.set(rc={'figure.figsize': (16, 9.)})
sns.set_style("whitegrid")
import pandas as pd
pd.set_option("display.max_rows", 120)
pd.set_option("display.max_columns", 120)
# %load_ext autoreload
# %autoreload 2
# -
# ### Read IRSM FORM
# +
cms_xml = xml_parser.get_files('irsmform_mono.xml', folder = '../linear TSR logs/monocurve')
cms_csv = xml_parser.get_files('CMS_Replication', folder = '../linear TSR logs/monocurve')
main_curve, estim_curves = xml_parser.get_rate_curves(cms_xml)
dsc_curve = main_curve
try:
estim_curve = estim_curves[0]
except TypeError:
estim_curve = main_curve
cms_replic_basket = csv_parser.parse_csv(cms_csv)
cal_basket = list(xml_parser.get_calib_basket(cms_xml))
# -
for (caplet, floorlet) in cms_replic_basket:
print('model a: ', caplet.model.a, ' model b: ', caplet.model.b)
caplet1 = cms_replic_basket[0][0]
swo1 = cal_basket[0]
mr = 0.06
linear.get_a(caplet1.pmnt_date, dsc_curve, swo1, mr, estim_curve)
diff_a, diff_b = [], []
for (caplet, floorlet), swo in zip(cms_replic_basket, cal_basket):
diff_a.append(caplet.model.a - linear.get_a(caplet.pmnt_date, dsc_curve, swo, mr, estim_curve)[0])
diff_b.append(caplet.model.b - linear.get_a(caplet.pmnt_date, dsc_curve, swo, mr, estim_curve)[1])
plt.plot(array(diff_a), label='a mistmatch')
plt.plot(array(diff_b), label='b mistmatch')
plt.title('mistmatch between csv logs and python replication')
plt.ylabel('mistmatch')
plt.xlabel('caplet expiry')
plt.legend()
| notebooks/unittest/linear_tsr_param_monocurve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: flaresinclustersii
# language: python
# name: flaresinclustersii
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
df = pd.read_csv("../flare_tables/20200227_vetted_flares_lacking_some.csv")
df = df[df.complex==0]
df1 = df[(df.real==1) &
(df.st>6) &
(df.ampl_rec > 0.05) &
(df.ampl_rec < .5)]
df2 = df[(df.real==1) &
(df.st==6) &
(df.ampl_rec > 0.05) &
(df.ampl_rec < .5)]
df1.ampl_rec.hist(bins=np.linspace(.05,.5,20), histtype="step", linewidth=2, label="later")
df2.ampl_rec.hist(bins=np.linspace(.05,.5,20), histtype="step", linewidth=2, label="equal")
plt.legend()
(df1.cstop-df1.cstart).hist(bins=np.linspace(3,60,50), histtype="step", linewidth=2, label="later")
(df2.cstop-df2.cstart).hist(bins=np.linspace(3,60,50), histtype="step", linewidth=2, label="equal")
plt.legend()
for i in range(0,df2.shape[0]):
l = df2[["TIC","sector","cstart"]].iloc[i]
s =f"{str(l.TIC).zfill(16)}_{l.sector}_{l.cstart}.csv"
f = pd.read_csv(f"../flare_snippets/{s}")
plt.figure()
plt.plot(f.c, f.f, label=f"TIC {l.TIC} sector {l.sector}, c={l.cstart}")
plt.legend()
plt.savefig(f"../plots/{i}.png")
plt.close()
| notebook/Compare_M6_and_later_M_types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import ast
from collections import Counter
import csv
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer, TfidfTransformer
from factor_analyzer.factor_analyzer import calculate_kmo
import numpy as np
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
from advanced_pca import CustomPCA
import gensim
import scipy
import seaborn as sns; sns.set()
from sklearn.cluster import MiniBatchKMeans, KMeans
from sklearn.manifold import TSNE
import matplotlib.cm as cm
from sklearn.cluster import DBSCAN
# The data is used from the 'Mediacloud_Analysis.ipynb' (<a href='Mediacloud_Analysis.ipynb'>link</a>). It already contains preprocessed and tokenized text for each article. Also it has a column with corona terms specifically and their frequency.
# +
#reading the dataframe with pre-processed tokens
df = pd.read_csv("preprocessed_results/mediacloud_parsed_corona_df_feb.csv")
temp = pd.read_csv("preprocessed_results/mediacloud_parsed_corona_df_may.csv")
temp_2 = pd.read_csv("preprocessed_results/mediacloud_parsed_corona_df_sep.csv")
df = pd.concat([df,temp])
df = pd.concat([df,temp_2])
df = df[~df.Text.isnull()] #removing rows withh no text
df['tokens'] = df['tokens'].apply(ast.literal_eval) #transforming string of tokens to list
df.head()
# +
# sample = df.sample(n=1)
temp = pd.read_csv('ncov-or-cov-19-or-covid-or-all-story-urls-20201012133126.csv')
sample = temp.sample(n=1)
print(sample['url'].values[0])
print(sample['title'].values[0])
print(sample['publish_date'])
# -
# For further procedures we use 500 most frequent tokens, that are later manually reviewed. All names, countries, dates as well as words that do not carry any strong meaning are excluded. They are saved to the 'most_frequent_tokens.csv' file
#finding 500 most frequent tokens
flatten_tokens = [token for sublist in df['tokens'].tolist() for token in sublist]
counter_tokens = Counter(flatten_tokens)
most_frequent = counter_tokens.most_common(500)
#saving them to csv file
with open('most_frequent_tokens.csv', "w") as the_file:
csv.register_dialect("custom", delimiter=",", skipinitialspace=True)
writer = csv.writer(the_file, dialect="custom")
for tup in most_frequent:
writer.writerow(tup)
# +
#finding 500 most frequent tokens for SEPTEMBER
flatten_tokens = [token for sublist in df['bigrams'][85298:].tolist() for token in sublist]
counter_tokens = Counter(flatten_tokens)
most_frequent = counter_tokens.most_common(500)
#saving them to csv file
with open('most_frequent_bigrams_SEP.csv', "w") as the_file:
csv.register_dialect("custom", delimiter=",", skipinitialspace=True)
writer = csv.writer(the_file, dialect="custom")
for tup in most_frequent:
writer.writerow(tup)
# -
# ## Unigrams
# Reading file with reviewed tokens (<a href="most_frequent_tokens_cleaned_v2.csv">file link</a>)
tokens = pd.read_csv('most_frequent_tokens_cleaned_v2.csv', header=None, names=['token', 'frequency'])
#tokens['tfidf'] = 0
# Firstly the original tokenized texts are converted to the tfidf scores. The result is sparse tfidf matrix. After that for each row only tfidf scores of frequent tokens are kept (for each sparse vector we match id of the tfidf value with dictionary token and check if this token is in the clean list). As a result for each row in the dataframe there is a vector of length n (nuber of cleaned frequent tokens) with tfidf values.
# +
def dummy_fun(doc):
return doc
cv = CountVectorizer(analyzer='word',
tokenizer=dummy_fun,
preprocessor=dummy_fun,
token_pattern=None)
data = cv.fit_transform(df['tokens'])
tfidf_transformer = TfidfTransformer()
tfidf_matrix = tfidf_transformer.fit_transform(data)
# +
tfidf_dict = cv.get_feature_names() #all tokens there are in the original texts
df['transformed_tokens'] = np.empty((len(df), 0)).tolist()
for i in range(tfidf_matrix.shape[0]):
print(i)
df.at[i, 'transformed_tokens'] = [tfidf_matrix[i].toarray()[0][j] for j in range(len(tfidf_dict)) if tfidf_dict[j] in tokens['token'].tolist()]
# +
temp = df['transformed_tokens'].tolist()
temp = [np.array(x) for x in temp]
tfidf_frequent = np.array(temp)
tfidf_frequent.shape #= [np.array(token_list) for token_list in tokens_transformed]
# -
with open("tfidf_transformed_tokens.csv", "w", newline="") as f:
writer = csv.writer(f)
writer.writerows(temp)
# KMO score is calculated (according to the <a href="https://www.tandfonline.com/doi/full/10.1080/1369183X.2017.1282813">paper</a>). KMO is a measure for sampling adequacy applied in factor analysis. It informs about the general strength of the relationship among items and thus indicates whether an item (i.e. a word) should be included in a factor analysis or not. Following Backhaus et al. (2006), terms with a KMO value below .50 were subsequently excluded.
kmo_all,kmo_model=calculate_kmo(tfidf_frequent)
kmo_model
# +
features_pca = np.zeros((tfidf_frequent.shape[0], len(kmo_all)))
for i in range(len(kmo_all)):
if kmo_all[i] > 0.5: #keeping only those that have kmo over 0.5
features_pca[i] = tfidf_frequent[i]
print(len(features_pca), tfidf_frequent.shape)
# -
# Running PCA on the filtered tokens. PCA is applied using <a href="https://pypi.org/project/advanced-pca/"> advanced PCA package</a>. For each number of components factor loadings are calculated (for each term) based on the <a href="https://www.r-bloggers.com/p-is-for-principal-components-analysis-pca/">tutorial here</a>. Only significant terms are taken (with a threshold of 0.1)
# +
scaler = StandardScaler()
features_pca_scaled = scaler.fit_transform(features_pca)
pca_results = {'Num_of_components': [],
'Explained_variance': [],
'Sum_Explained_variance': [],
'Terms':[]
}
for n in range (3, 21):
pca_model = (CustomPCA(n_components=n)
.fit(features_pca_scaled))
pca_results['Num_of_components'].append(n)
pca_results['Explained_variance'].append(pca_model.explained_variance_ratio_)
pca_results['Sum_Explained_variance'].append(sum(pca_model.explained_variance_ratio_))
all_terms = []
for i in range(n):
scores = [score for score in pca_model.components_[i].round(1) if score>0.1 or score<-0.1]
# tokens_sign = (pca_model.components_[i].round(1)>0.1) or (pca_model.components_[i].round(1)<-0.1)
terms = tokens.token[(pca_model.components_[i].round(1)>0.1) | (pca_model.components_[i].round(1)<-0.1)]
all_terms.append(list(zip(terms, scores)))
pca_results['Terms'].append(all_terms)
pca_results_df = pd.DataFrame(pca_results)
# -
# Example with a custom PCA with 3 components, printing variance ratio for each component and factor loadings:
pca_model = (CustomPCA(n_components=5)
.fit(features_pca_scaled))
print(pca_model.explained_variance_ratio_)
pca_model.components_[1]
pca_results_df['Terms'][0]
# Saving results of the PCA to the csv file 'results/mediacloud_pca_results_shortlist.csv'. Plot the sum of explained variance based on the number of components:
pca_results_df.to_csv('results/mediacloud_pca_results_shortlist.csv')
pca_results_df.plot.line(x='Num_of_components', y='Sum_Explained_variance')
# Save the 'significant' terms for all components (each n of components) with corresponding factor loadings to csv file 'results/pca_terms.csv':
pca_results_df['Terms'].to_csv('results/pca_terms.csv')
print(pca_results_df['Terms'][5])
# A plot that shows cumulative explained variance and explained variance of each component (with max 20):
# +
cummulative_pca = PCA(n_components=20).fit(features_pca_scaled)
fig, ax = plt.subplots(figsize=(8,6))
x_values = range(1, cummulative_pca.n_components_+1)
ax.plot(x_values, cummulative_pca.explained_variance_ratio_, lw=2, label='explained variance')
ax.plot(x_values, np.cumsum(cummulative_pca.explained_variance_ratio_), lw=2, label='cumulative explained variance')
ax.set_title('PCA on filtered tokens : explained variance of components')
ax.set_xlabel('principal component')
ax.set_ylabel('explained variance')
plt.show()
# -
# ## Bigrams
# Creating bigrams from the original texts. The bigrams are then saved to file 'most_frequent_tokens_bigrams.csv' and reviewed the same way as the unigrams in the file 'most_frequent_tokens_bigrams.csv' (<a href='most_frequent_tokens_bigrams.csv'>link</a>). The final list contains 87 terms
bigram = gensim.models.Phrases(df['tokens'], min_count=3, threshold=50) # higher threshold fewer phrases.
bigram_mod = gensim.models.phrases.Phraser(bigram)
# +
def make_bigrams(texts):
return [bigram_mod[doc] for doc in texts]
df['bigrams'] = make_bigrams(df['tokens'])
df['bigrams']
# -
flatten_bigrams = [token for sublist in df['bigrams'].tolist() for token in sublist]
counter_bigrams = Counter(flatten_bigrams)
most_frequent = counter_bigrams.most_common(500)
#saving them to csv file
with open('most_frequent_tokens_bigrams.csv', "w") as the_file:
csv.register_dialect("custom", delimiter=",", skipinitialspace=True)
writer = csv.writer(the_file, dialect="custom")
for tup in most_frequent:
writer.writerow(tup)
# +
# tokens_bigrams = pd.read_csv('most_frequent_tokens_bigrams.csv', header=None, names=['token', 'frequency'])
cv = CountVectorizer(analyzer='word',
tokenizer=dummy_fun,
preprocessor=dummy_fun,
token_pattern=None)
data = cv.fit_transform(df['bigrams'])
tfidf_transformer = TfidfTransformer()
tfidf_matrix = tfidf_transformer.fit_transform(data)
# -
tfidf_matrix
# +
tfidf_dict_bigrams = cv.get_feature_names() #all tokens there are in the original texts
df['transformed_tokens_bigrams'] = np.empty((len(df), 0)).tolist()
for i in range(tfidf_matrix.shape[0]):
print(i)
df.at[i, 'transformed_tokens_bigrams'] = [tfidf_matrix[i].toarray()[0][j] for j in range(len(tfidf_dict_bigrams)) if tfidf_dict_bigrams[j] in tokens_bigrams['token'].tolist()]
# -
with open("tfidf_transformed_bigrams.csv", "w", newline="") as f:
writer = csv.writer(f)
writer.writerows(df['transformed_tokens_bigrams'].tolist())
# +
temp = df['transformed_tokens_bigrams'].tolist()
temp = [np.array(x) for x in temp]
tfidf_frequent_bigrams = np.array(temp)
tfidf_frequent_bigrams.shape #= [np.array(token_list) for token_list in tokens_transformed]
# -
kmo_all_bi,kmo_model_bi=calculate_kmo(np.array(tfidf_frequent_bigrams))
kmo_model_bi
# +
features_bigrams = np.zeros((tfidf_frequent_bigrams.shape[0], len(kmo_all_bi)))
for i in range(len(kmo_all_bi)):
if kmo_all_bi[i] > 0.5: #keeping only those that have kmo over 0.5
features_bigrams[i] = tfidf_frequent_bigrams[i]
print(len(features_bigrams), tfidf_frequent_bigrams.shape)
# +
scaler = StandardScaler()
features_bi_scaled = scaler.fit_transform(features_bigrams)
pca_results_bi = {'Num_of_components': [],
'Explained_variance': [],
'Terms':[]
}
for n in range (3, 21):
pca_model = (CustomPCA(n_components=n)
.fit(features_bi_scaled))
pca_results_bi['Num_of_components'].append(n)
pca_results_bi['Explained_variance'].append(sum(pca_model.explained_variance_ratio_))
all_terms = []
for i in range(n):
scores = [score for score in pca_model.components_[i].round(1) if score>0.1]
# tokens_sign = (pca_model.components_[i].round(1)>0.1) or (pca_model.components_[i].round(1)<-0.1)
terms = tokens_bigrams.token[pca_model.components_[i].round(1)>0.1]
all_terms.append(list(zip(terms, scores)))
pca_results_bi['Terms'].append(all_terms)
pca_results_bi_df = pd.DataFrame(pca_results_bi)
# -
pca_model = (CustomPCA(n_components=3)
.fit(features_bi_scaled))
print(pca_model.explained_variance_ratio_)
pca_model.components_[1]
pca_results_bi_df['Terms'][0]
# +
temp = tokens_bigrams['token'].tolist()
pca_dict = {}
for token in temp:
pca_dict[token] = []
for topic in pca_results_bi_df['Terms'][17]:
if token in [term[0] for term in topic]:
pca_dict[token].append([term[1] for term in topic if term[0]==token][0])
else:
pca_dict[token].append(0)
pca_df = pd.DataFrame(pca_dict).transpose()
# -
pca_df[pca_df[5]!=0]
pca_results_bi_df.to_csv('results/mediacloud_pca_bigrams_results_shortlist.csv')
pca_results_bi_df.plot.line(x='Num_of_components', y='Explained_variance')
# +
cummulative_pca = PCA(n_components=20).fit(features_bi_scaled)
fig, ax = plt.subplots(figsize=(8,6))
x_values = range(1, cummulative_pca.n_components_+1)
ax.plot(x_values, cummulative_pca.explained_variance_ratio_, lw=2, label='explained variance')
ax.plot(x_values, np.cumsum(cummulative_pca.explained_variance_ratio_), lw=2, label='cumulative explained variance')
ax.set_title('PCA on filtered tokens : explained variance of components')
ax.set_xlabel('principal component')
ax.set_ylabel('explained variance')
plt.show()
# -
# ## Toy example
#
# The perfect curated list is created, that contains 39 words for 4 frames: economic, medical, travel and restrictions/prevention. The list is available <a href="most_frequent_tokens_toy.csv">here</a>
tokens_toy = pd.read_csv('most_frequent_tokens_toy.csv', header=None, names=['token', 'frequency'])
toy = tokens_toy['token'].sort_values().tolist()
bigrams_sep = pd.read_csv('most_frequent_bigrams_SEP.csv', header=None, names=['token', 'frequency'])
tokens_sep = bigrams_sep['token'].sort_values().tolist()
# +
# tokens_bigrams = pd.read_csv('most_frequent_tokens_bigrams.csv', header=None, names=['token', 'frequency'])
def dummy_fun(doc):
return doc
cv = CountVectorizer(analyzer='word',
tokenizer=dummy_fun,
preprocessor=dummy_fun,
token_pattern=None)
data = cv.fit_transform(df['bigrams'][85298:])
tfidf_transformer = TfidfTransformer()
tfidf_matrix = tfidf_transformer.fit_transform(data)
# -
df.reset_index(inplace=True)
df.drop(['Unnamed: 0','Unnamed: 0.1'],axis=1,inplace=True)
tfidf_matrix.shape[0]
# +
# tfidf_dict = cv.get_feature_names() #all tokens there are in the original texts
tfidf_dict_bigrams = cv.get_feature_names()
transformed_tokens_sep = np.empty((tfidf_matrix.shape[0], 86))
for i in range(0, tfidf_matrix.shape[0]):
print(i)
# print([tfidf_matrix[i].toarray()[0][j] for j in range(len(tfidf_dict_bigrams)) if tfidf_dict_bigrams[j] in tokens_sep])
transformed_tokens_sep[i] = [tfidf_matrix[i].toarray()[0][j] for j in range(len(tfidf_dict_bigrams)) if tfidf_dict_bigrams[j] in tokens_sep]
# -
with open("tfidf_transformed_toy_sep.csv", "w", newline="") as f:
writer = csv.writer(f)
writer.writerows(df['transformed_tokens_toy2'][1136:].tolist())
with open("tfidf_transformed_toy_sep.csv", newline='') as csvfile:
data = list(csv.reader(csvfile))
# +
temp = data
temp = [np.array(x) for x in temp]
tfidf_frequent_toy = np.array(temp)
tfidf_frequent_toy.shape #= [np.array(token_list) for token_list in tokens_transformed]
# -
tfidf_frequent_sep = transformed_tokens_sep[:1136].astype(np.float)
kmo_all_toy,kmo_model_toy=calculate_kmo(tfidf_frequent_sep)
kmo_model_toy
# +
features_sep = np.zeros((tfidf_frequent_sep.shape[0], len(kmo_all_toy)))
for i in range(len(kmo_all_toy)):
if kmo_all_toy[i] > 0.5: #keeping only those that have kmo over 0.5
features_sep[i] = tfidf_frequent_sep[i]
print(len(features_sep), tfidf_frequent_sep.shape)
# -
# Kmeans clustering. For each number of k model is created and fitted on above features (consisting of 36 manually chosen words). Number of texts assigned to each cluster is printed below. Then top words are presented and a tsne graph of them in 2d
# +
random_state = 20
k = 3
model = KMeans(n_clusters=k, random_state=random_state)
clusters = model.fit_predict(features_sep)
# tsne = TSNE().fit_transform(features_sep)
Counter(clusters)
# max_items = np.random.choice(range(features_toy.shape[0]), size=10000, replace=False)
# +
#FEBRUARY
def get_top_keywords(data, clusters, labels, n_terms):
df = pd.DataFrame(data).groupby(clusters).mean()
for i,r in df.iterrows():
print('\nCluster {}'.format(i))
print(','.join([labels[t] for t in np.argsort(r)[-n_terms:]]))
get_top_keywords(features_toy, clusters, tokens_toy, 5)
# -
#SEPTEMBER
get_top_keywords(features_sep, clusters, tokens_sep, 5)
# +
#kmeans and dbscan, 3 to 5 k
def plot_tsne_pca(tsne, labels):
max_label = max(labels)
label_subset = [cm.hsv(i/max_label) for i in labels]
plt.scatter(tsne[:, 0], tsne[:, 1], c=label_subset)
plt.title('TSNE Cluster Plot')
plot_tsne_pca(tsne[clusters!=0], clusters[clusters!=0])
# plot_tsne_pca(tsne, clusters)
# -
# DBSCAN.
# +
eps = 3
min_samples = 3
dbscan = {
'eps':[],
'min_samples':[],
'labels':[]
}
for eps in np.arange(0.01,0.05, 0.01):
for min_samples in range (3, 10, 1):
db1 = DBSCAN(eps=eps, min_samples=min_samples).fit(features_toy)
labels1 = db1.labels_
print(f"eps: {eps}, min samples: {min_samples}")
print(Counter(labels1))
dbscan['eps'].append(eps)
dbscan['min_samples'].append(min_samples)
dbscan['labels'].append(labels1)
# -
# PCA. Number of components ranging from 3 to 5, printing explained variance ratio, factor loading matrix and significant terms for each component.
scaler = StandardScaler()
features_toy_scaled = scaler.fit_transform(features_sep)
pca_model_toy = (CustomPCA(n_components=4)
.fit(features_toy_scaled))
print(pca_model_toy.explained_variance_ratio_)
pca_model_toy.components_[1]
# +
all_terms = []
for i in range(4):
scores = [score for score in pca_model_toy.components_[i].round(2) if score>=0.2]
print(scores)
terms = bigrams_sep.token[pca_model_toy.components_[i].round(2)>=0.2]
all_terms.append(list(zip(terms, scores)))
all_terms
# -
pca_model_toy.components_[3]
| Mediacloud_PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# some (artificial) data
X = np.array([[0,0,1],
[0,1,1],
[1,0,1],
[0,0,1]])
y = np.array([[0],
[1],
[1],
[0]])
# define how to compute the nonlinearity and it's derivative
def nonlin(x, deriv=False):
pass
# +
# seed the NRG for reproducible results
np.random.seed(1)
# set up the parameters for a 2-layer fully connected network
layer0 = ???
layer1 = ???
learning_rate = 1
# iterate...
for j in range(0, 60000):
# compute forward pass through the network; retain values
# compute error: how much did we miss the target value?
# compute delta at output level
# recursively compute deltas at lower levels
# print final params
print(layer0, layer1)
# -
# test model
| jupyter/06-neural-nets-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
lista = ['manzana', 'platano', 'melocoton', 'pera', 'manzana', 'limon', 'platano', 'naranaja']
conjunto = set(lista)
conjunto
# +
final_list = []
for i in conjunto:
final_list.append(i)
# -
final_list
| Machine Learning/exercise_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # School Board Minutes
#
# Scrape all of the school board minutes from http://www.mineral.k12.nv.us/pages/School_Board_Minutes
#
# Save a CSV called `minutes.csv` with the date and the URL to the file. The date should be formatted as YYYY-MM-DD.
#
# **Bonus:** Download the PDF files
#
# **Bonus 2:** Use [PDF OCR X](https://solutions.weblite.ca/pdfocrx/index.php) on one of the PDF files and see if it can be converted into text successfully.
#
# * **Hint:** If you're just looking for links, there are a lot of other links on that page! Can you look at the link to know whether it links or minutes or not? You'll want to use an "if" statement.
# * **Hint:** You could also filter out bad links later on using pandas instead of when scraping
# * **Hint:** If you get a weird error that you can't really figure out, you can always tell Python to just ignore it using `try` and `except`, like below. Python will try to do the stuff inside of 'try', but if it hits an error it will skip right out.
# * **Hint:** Remember the codes at http://strftime.org
# * **Hint:** If you have a date that you've parsed, you can use `.dt.strftime` to turn it into a specially-formatted string. You use the same codes (like %B etc) that you use for converting strings into dates.
#
# ```python
# try:
# blah blah your code
# your code
# your code
# except:
# pass
# ```
| 08-school-board-minutes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
# %matplotlib inline
import matplotlib.pyplot as plt
from matplotlib import style
from sklearn.datasets import make_regression
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_squared_error
from sklearn.datasets.samples_generator import make_blobs
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report
from sklearn.cluster import KMeans
# # Read the CSV and Perform Basic Data Cleaning
df = pd.read_csv("cumulative.csv")
df = df.drop(columns=["rowid", "kepid", "kepoi_name", "kepler_name", "koi_pdisposition", "koi_score", "koi_tce_delivname"])
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df.head()
# # Create a Train Test Split
#
# Use `koi_disposition` for the y values
target = df["koi_disposition"]
target_names = ["CONFIRMED", "FALSE POSITIVE", "CANDIDATE"]
target.head()
data = df.drop("koi_disposition", axis=1)
feature_names = data.columns
data.head()
# Use train_test_split to create training and testing data
X_train, X_test, y_train, y_test = train_test_split(data, target, random_state=42)
X_train.head()
# # Pre-processing
#
# Scale the data using the MinMaxScaler
# +
from sklearn.preprocessing import MinMaxScaler
# Create a StandardScater model and fit it to the training data
X_scaler = MinMaxScaler().fit(X_train)
# Transform the training and testing data using the X_scaler and y_scaler models
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
# # Train the Kmeans
from sklearn.cluster import KMeans
import numpy as np
kmeans = KMeans(n_clusters=3)
kmeans.fit(X_train_scaled)
# Predict the clusters
predicted_clusters = kmeans.predict(X_test_scaled)
predicted_clusters
np.unique(predicted_clusters)
np.unique(y_test)
# Plot the predicted clusters to see if the model predicted the correct clusters
# This is visual validation that the model was trained correctly.
plt.scatter(X_train_scaled, X_test_scaled, c=predicted_clusters, s=50, cmap='viridis')
| starter_code/Model_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Processor temperature
#
# We have a temperature sensor in the processor of our company's server. We want to analyze the data provided to determinate whether we should change the cooling system for a better one. It is expensive and as a data analyst we cannot make decisions without a basis.
#
# We provide the temperatures measured throughout the 24 hours of a day in a list-type data structure composed of 24 integers:
# ```
# temperatures_C = [33,66,65,0,59,60,62,64,70,76,80,69,80,83,68,79,61,53,50,49,53,48,45,39]
# ```
#
# ## Goals
#
# 1. Treatment of lists
# 2. Use of loop or list comprenhention
# 3. Calculation of the mean, minimum and maximum.
# 4. Filtering of lists.
# 5. Interpolate an outlier.
# 6. Logical operators.
# 7. Print
# ## Temperature graph
# To facilitate understanding, the temperature graph is shown below. You do not have to do anything in this section. The test starts in **Problem**.
# +
# import
import matplotlib.pyplot as plt
# %matplotlib inline
# axis x, axis y
y = [33,66,65,0,59,60,62,64,70,76,80,81,80,83,90,79,61,53,50,49,53,48,45,39]
x = list(range(len(y)))
# plot
plt.plot(x, y)
plt.axhline(y=70, linewidth=1, color='r')
plt.xlabel('hours')
plt.ylabel('Temperature ºC')
plt.title('Temperatures of our server throughout the day')
# -
# ## Problem
#
# If the sensor detects more than 4 hours with temperatures greater than or equal to 70ºC or any temperature above 80ºC or the average exceeds 65ºC throughout the day, we must give the order to change the cooling system to avoid damaging the processor.
#
# We will guide you step by step so you can make the decision by calculating some intermediate steps:
#
# 1. Minimum temperature
# 2. Maximum temperature
# 3. Temperatures equal to or greater than 70ºC
# 4. Average temperatures throughout the day.
# 5. If there was a sensor failure at 03:00 and we did not capture the data, how would you estimate the value that we lack? Correct that value in the list of temperatures.
# 6. Bonus: Our maintenance staff is from the United States and does not understand the international metric system. Pass temperatures to Degrees Fahrenheit.
#
# Formula: F = 1.8 * C + 32
#
# web: https://en.wikipedia.org/wiki/Conversion_of_units_of_temperature
#
# +
# assign a variable to the list of temperatures
temperatures_C = [33,66,65,0,59,60,62,64,70,76,80,69,80,83,68,79,61,53,50,49,53,48,45,39]
# 1. Calculate the minimum of the list and print the value using print()
print(min(temperatures_C))
# 2. Calculate the maximum of the list and print the value using print()
print(max(temperatures_C))
# 3. Items in the list that are greater than 70ºC and print the result
greater70=[]
for i in range(len(temperatures_C)):
if temperatures_C[i]>70:
greater70.append(temperatures_C[i])
print(greater70)
# 4. Calculate the mean temperature throughout the day and print the result
print(sum(temperatures_C)/len(temperatures_C))
# 5.1 Solve the fault in the sensor by estimating a value
missing=(temperatures_C[3-1]+temperatures_C[3+1])/2
# 5.2 Update of the estimated value at 03:00 on the list
temperatures_C[3]=missing
# Bonus: convert the list of ºC to ºFarenheit
temperatures_F=[]
for i in range(len(temperatures_C)):
temperatures_F.append(round((1.8*temperatures_C[i])+32,2))
# -
# ## Take the decision
# Remember that if the sensor detects more than 4 hours with temperatures greater than or equal to 70ºC or any temperature higher than 80ºC or the average was higher than 65ºC throughout the day, we must give the order to change the cooling system to avoid the danger of damaging the equipment:
# * more than 4 hours with temperatures greater than or equal to 70ºC
# * some temperature higher than 80ºC
# * average was higher than 65ºC throughout the day
# If any of these three is met, the cooling system must be changed.
#
# +
# Print True or False depending on whether you would change the cooling system or not
decision=bool
if (len(greater70)>4) | (max(temperatures_C)>80) | (sum(temperatures_C)/len(temperatures_C) > 65):
decision = True
else:
decision = False
print(decision)
# -
# ## Future improvements
# 1. We want the hours (not the temperatures) whose temperature exceeds 70ºC
# 2. Condition that those hours are more than 4 consecutive and consecutive, not simply the sum of the whole set. Is this condition met?
# 3. Average of each of the lists (ºC and ºF). How they relate?
# 4. Standard deviation of each of the lists. How they relate?
#
# 1. We want the hours (not the temperatures) whose temperature exceeds 70ºC
for i in range(len(temperatures_C)):
if (temperatures_C[i]>70):
print('hour',i)
# 2. Condition that those hours are more than 4 consecutive and consecutive, not simply the sum of the whole set. Is this condition met?
hours70=[]
for i in range(len(temperatures_C)):
if (temperatures_C[i]>70):
hours70.append(i)
fourinarow=bool
for i in range(len(hours70)-3):
if(hours70[i+1]==hours70[i]+1)&(hours70[i+2]==hours70[i]+2)&(hours70[i+3]==hours70[i]+3):
fourinarow=True
break #to not overwrite the boolean, if we find, it met
else:
fourinarow=False
print(fourinarow)
# 3. Average of each of the lists (ºC and ºF). How they relate?
ave_C=sum(temperatures_C)/len(temperatures_C)
ave_F=sum(temperatures_F)/len(temperatures_F)
print(ave_C, ave_F)
print("They relate in the same way they relate piecewise. Proof: ave_C*1.8+32 = ", ave_C*1.8+32 ,", which is exactly the average of\n the Fahrenheit list (rounded to two decimal places).")
# +
# 4. Standard deviation of each of the lists. How they relate?
import statistics
stdev_C=statistics.stdev(temperatures_C)
stdev_F=statistics.stdev(temperatures_F)
print(stdev_C, stdev_F)
print("The relation between them is, that the standard deviation of Celsius times 1.8 is the standard deviation of\n Fahrenheit: stdev_C*1.8=", stdev_C*1.8, ". It means, that the standard deviation doesn't express the summand, \n but the multiplicative summand.")
# -
| temperature/temperature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:gpt]
# language: python
# name: conda-env-gpt-py
# ---
import runpy
file_globals = runpy.run_path("../run_generation.py")
sample_sequence = file_globals['sample_sequence']
yt_encoder = runpy.run_path("../yt_encoder.py")
YTEncoder = yt_encoder['YTEncoder']
# +
from transformers import GPT2LMHeadModel
import threading
import regex as re
import logging
logging.basicConfig(filename="rest.log", level=logging.INFO)
logger = logging.getLogger(__name__)
from os import environ
device = environ.get('DEVICE', 'cuda:0')
model_path = '../gpt2/medium'
tokenizer = YTEncoder.from_pretrained(model_path)
poetry_model = GPT2LMHeadModel.from_pretrained(model_path + '/poetry')
poetry_model.to(device)
poetry_model.eval()
def get_sample(model, prompt, length:int, num_samples:int, allow_linebreak:bool):
logger.info("*" * 200)
logger.info(prompt)
filter_n = tokenizer.encode('\n')[-1:]
filter_single = [1]
filter_single += [] if allow_linebreak else filter_n
context_tokens = tokenizer.encode(prompt)
out = sample_sequence(
model=model,
context=context_tokens,
length=length,
temperature=1,
top_k=0,
top_p=0.9,
device=device,
filter_single=filter_single,
filter_double=filter_n,
num_samples=num_samples,
).to('cpu')
prompt = tokenizer.decode(context_tokens)
len_prompt = len(prompt)
replies = [out[item, :].tolist() for item in range(len(out))]
text = [tokenizer.decode(item)[len_prompt:] for item in replies]
reg_text = [re.match(r'[\w\W]*[\.!?]\n', item) for item in text]
result = [reg_item[0] if reg_item else item for reg_item, item in zip(reg_text,text)]
logger.info("=" * 200)
logger.info(result)
return result
# -
from rupo.api import Engine
from rupo.rhymes.rhymes import Rhymes
engine = Engine(language="ru")
engine.is_rhyme("корова", "здорова")
engine.is_rhyme('привет', 'ответ')
engine.is_rhyme('пизда', 'джигурда')
from transitions import Machine
def get_sample(model, prompt, length:int, num_samples:int, allow_linebreak:bool):
logger.info("*" * 200)
logger.info(prompt)
filter_n = tokenizer.encode('\n')[-1:]
filter_single = [1]
filter_single += [] if allow_linebreak else filter_n
context_tokens = tokenizer.encode(prompt)
out = sample_sequence(
model=model,
context=context_tokens,
length=length,
temperature=1,
top_k=0,
top_p=0.9,
device=device,
filter_single=filter_single,
filter_double=filter_n,
num_samples=num_samples,
).to('cpu')
prompt = tokenizer.decode(context_tokens)
len_prompt = len(prompt)
replies = [out[item, :].tolist() for item in range(len(out))]
text = [tokenizer.decode(item)[len_prompt:] for item in replies]
reg_text = [re.match(r'[\w\W]*[\.!?]\n', item) for item in text]
result = [reg_item[0] if reg_item else item for reg_item, item in zip(reg_text,text)]
logger.info("=" * 200)
logger.info(result)
return result
# +
import torch
import torch.nn.functional as F
import numpy as np
from tqdm import trange
from transformers import (GPT2Config, OpenAIGPTConfig, XLNetConfig, TransfoXLConfig,
GPT2LMHeadModel, GPT2Tokenizer,
OpenAIGPTLMHeadModel, OpenAIGPTTokenizer,
XLNetLMHeadModel, XLNetTokenizer,
TransfoXLLMHeadModel, TransfoXLTokenizer, )
logging.basicConfig(format = '%(asctime)s - %(levelname)s - %(name)s - %(message)s',
datefmt = '%m/%d/%Y %H:%M:%S',
level = logging.INFO)
logger = logging.getLogger(__name__)
MAX_LENGTH = int(10000) # Hardcoded max length to avoid infinite loop
ALL_MODELS = sum((tuple(conf.pretrained_config_archive_map.keys()) for conf in (GPT2Config, OpenAIGPTConfig, XLNetConfig, TransfoXLConfig)), ())
MODEL_CLASSES = {
'gpt2': (GPT2LMHeadModel, GPT2Tokenizer),
'openai-gpt': (OpenAIGPTLMHeadModel, OpenAIGPTTokenizer),
'xlnet': (XLNetLMHeadModel, XLNetTokenizer),
'transfo-xl': (TransfoXLLMHeadModel, TransfoXLTokenizer),
}
FILTER_VALUE=-float('Inf')
# -
model = poetry_model
length = 150
prompt = 'мы с иваном ильичем ехали на дизеле\nон мудак и я мудак\n'
context = tokenizer.encode(prompt)
num_samples = 1
temperature=1
top_k=0
top_p=0.0
is_xlnet=False
device='cuda'
max_input=1023
filter_single=[]
filter_double=[]
def top_k_top_p_filtering(logits, top_k=0, top_p=0.0, filter_value=-float('Inf')):
""" Filter a distribution of logits using top-k and/or nucleus (top-p) filtering
Args:
logits: logits distribution shape (vocabulary size)
top_k > 0: keep only top k tokens with highest probability (top-k filtering).
top_p > 0.0: keep the top tokens with cumulative probability >= top_p (nucleus filtering).
Nucleus filtering is described in Holtzman et al. (http://arxiv.org/abs/1904.09751)
From: https://gist.github.com/thomwolf/1a5a29f6962089e871b94cbd09daf317
"""
assert logits.dim() == 1 # batch size 1 for now - could be updated for more but the code would be less clear
top_k = min(top_k, logits.size(-1)) # Safety check
if top_k > 0:
# Remove all tokens with a probability less than the last token of the top-k
indices_to_remove = logits < torch.topk(logits, top_k)[0][..., -1, None]
logits[indices_to_remove] = filter_value
if top_p > 0.0:
sorted_logits, sorted_indices = torch.sort(logits, descending=True)
cumulative_probs = torch.cumsum(F.softmax(sorted_logits, dim=-1), dim=-1)
# Remove tokens with cumulative probability above the threshold
sorted_indices_to_remove = cumulative_probs > top_p
# Shift the indices to the right to keep also the first token above the threshold
sorted_indices_to_remove[..., 1:] = sorted_indices_to_remove[..., :-1].clone()
sorted_indices_to_remove[..., 0] = 0
indices_to_remove = sorted_indices[sorted_indices_to_remove]
logits[indices_to_remove] = filter_value
return logits
if '\n' not in prompt:
initial = 'firstline'
else:
initial = 'midline'
# +
states = ['firstline', 'midline', 'rhymed', 'del']
transitions = [
{'trigger': 'newline', 'source': 'firstline', 'dest': 'midline'},
{'trigger': 'newline', 'source': 'midline', 'dest': 'del'},
{'trigger': 'newline', 'source': 'rhymed', 'dest': 'midline'},
{'trigger': 'rhyme', 'source': 'firstline', 'dest': 'del'},
{'trigger': 'rhyme', 'source': 'midline', 'dest': 'rhymed'},
{'trigger': 'rhyme', 'source': 'rhymed', 'dest': 'del'},
{'trigger': 'word', 'source': 'firstline', 'dest': 'firstline'},
{'trigger': 'word', 'source': 'rhymed', 'dest': 'del'},
{'trigger': 'word', 'source': 'midline', 'dest': 'midline'},
{'trigger': 'newline', 'source': 'del', 'dest': 'del'},
{'trigger': 'rhyme', 'source': 'del', 'dest': 'del'},
{'trigger': 'word', 'source': 'del', 'dest': 'del'},
]
class Stih(object):
def __init__(self, text=''):
self.tokens = tokenizer.encode(text)
self.text = text
self.p = 1
if '\n' not in text:
initial = 'firstline'
else:
initial = 'midline'
self.machine = Machine(model=self, states=states, transitions=transitions, initial=initial)
def add(self, token, p):
token = int(token)
self.tokens += [token]
self.last = tokenizer.decode([token])
self.text = tokenizer.decode(self.tokens)
self.p *= p
if '\n' in self.last:
self.newline()
if re.search(r'(\w)', self.last):
self.word()
return self.last
def __repr__(self):
return self.text
# -
stih = Stih(prompt)
import copy
def filter_rhyme(stih, filtered_logits):
def check_rhyme(token):
last = tokenizer.decode([int(token)])
if not re.search(r'([а-яА-Я])', last):
return False
total_stih = tokenizer.decode(stih.tokens + [token])
last_word = re.findall(r'\b\w+\b', total_stih)[-1]
lines = total_stih.splitlines()
rhyme_line = lines[max(-3, -len(lines))]
rhyme_word = re.findall(r'\b\w+\b', rhyme_line)[-1]
#print(last_word, rhyme_word)
return engine.is_rhyme(last_word, rhyme_word)
arr = np.array(np.argsort(filtered_logits.cpu()))[::-1][:6400]
rhymes = Pool(64).map(check_rhyme, arr)
for itoken in arr:
if filtered_logits[itoken] > FILTER_VALUE:
if rhymes[itoken]:
return itoken
# +
from mezmorize import Cache
cache = Cache(CACHE_TYPE='filesystem', CACHE_DIR='cache')
@cache.memoize()
def is_rhyme(word1, word2):
#return engine.is_rhyme(word1, word2)
markup_word1 = engine.get_markup(word1).lines[0].words[0]
markup_word1.set_stresses(engine.get_stresses(word1))
markup_word2 = engine.get_markup(word2).lines[0].words[0]
markup_word2.set_stresses(engine.get_stresses(word2))
return Rhymes.is_rhyme(markup_word1, markup_word2, 5)
# -
def check_rhyme(stih, token):
last = tokenizer.decode([int(token)])
if not re.search(r'([а-яА-Я])', last):
return False
total_stih = tokenizer.decode(stih.tokens + [token])
last_word = re.findall(r'\b\w+\b', total_stih)[-1]
lines = total_stih.splitlines()
rhyme_line = lines[max(-3, -len(lines))]
rhyme_word = re.findall(r'\b\w+\b', rhyme_line)[-1]
if rhyme_word == last_word: return False
is_rhymed = is_rhyme(last_word, rhyme_word)
if is_rhymed: print(last_word, rhyme_word)
return is_rhymed
from profilehooks import profile
#@profile(immediate=True)
def filter_rhyme(stih, filtered_logits):
for itoken in np.array(np.argsort(filtered_logits.cpu()))[::-1][:64000]:
if filtered_logits[itoken] > 0:
if check_rhyme(stih, itoken):
return itoken
def filter_after_rhyme(stih, filtered_logits):
for itoken in np.array(np.argsort(filtered_logits.cpu()))[::-1][:64000]:
if filtered_logits[itoken] > 0:
if not re.search(r'(\w)', tokenizer.decode([int(itoken)])):
return itoken
stih.text
stihi = [stih]
with torch.no_grad():
for _ in trange(length):
context = torch.tensor([stih.tokens for stih in stihi], dtype=torch.long, device=device)
inputs = {'input_ids': context[:,-max_input:]}
outputs = model(**inputs) # Note: we could also use 'past' with GPT-2/Transfo-XL/XLNet (cached hidden-states)
next_tokens = torch.zeros(num_samples, dtype=torch.long).to(device)
n_stihi = []
for isample in range(len(stihi)):
next_token_logits = outputs[0][isample, -1, :] / temperature
next_token_logits[filter_single] = FILTER_VALUE
# filter blank line = double \n
if context[isample, -1] in filter_double:
next_token_logits[generated[isample, -1]] = FILTER_VALUE
filtered_logits = top_k_top_p_filtering(next_token_logits, top_k=top_k, top_p=top_p)
stih1 = copy.deepcopy(stihi[isample])
if stih1.state != 'rhymed':
i_logit = torch.multinomial(F.softmax(filtered_logits, dim=-1), num_samples=1)
else:
i_logit = filter_after_rhyme(stih, filtered_logits)
i_logit = int(i_logit)
stih1.add(i_logit, filtered_logits[i_logit])
n_stihi += [stih1]
stih2 = copy.deepcopy(stihi[isample])
stih2.rhyme()
if stih2.state != 'del':
rhymed_logit = filter_rhyme(stih2, filtered_logits)
print(rhymed_logit)
if rhymed_logit:
stih2.add(rhymed_logit, filtered_logits[rhymed_logit])
stih2.machine.set_state('rhymed')
n_stihi += [stih2]
stihi = [stih for stih in n_stihi if stih.state != 'del']
stihi = sorted(stihi, key=lambda stih: stih.p, reverse=True)[:64]
print(stihi)
n_stihi[0].text
print(stih.text)
print(tokenizer.decode(generated.to('cpu')))
get_sample(poetry_model, 'тест', 50, 1, True)
| poetry/rhyme.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import datetime as dt
def epoch_to_millis(epoch):
datetime_time = dt.datetime.fromtimestamp(epoch)
s = (datetime_time - dt.datetime(1970, 1, 1)).total_seconds()-3600
return s
# +
#import e parsing file di log
import pandas as pd
def start(path):
file = open(path, 'r')
Lines = file.readlines()
return Lines
def rmse_distance(Lines):
coppie = []
for line in Lines:
if 'COUPLE(N_ITER,DISTANCE RMSE)' in line:
split = line.split(':')
s = split[3].replace('[','')
s = s.replace('(','')
s = s.replace(')','')
s = s.replace(']','')
s = s.split(',')
a = int(s[0])
b = float(s[1])
coppie.append((a,b))
return coppie
def execution_time_gurobi(Lines):
for line in Lines:
if 'TEMPO_ESECUZIONE GUROBI' in line:
split = line.split(':')
return epoch_to_millis(float(split[3]))
def execution_time_tensorflow(Lines):
for line in Lines:
if 'TEMPO_ESECUZIONE TENSORFLOW' in line:
split = line.split(':')
return epoch_to_millis(float(split[3]))
def time_per_iter(Lines):
coppie = []
for line in Lines:
if 'TEMPO PER ITERAZIONI (N_ITER,TEMPO) TENSORFLOW' in line:
split = line.split(':')
s = split[3].replace('[','')
s = s.replace('(','')
s = s.replace(')','')
s = s.replace(']','')
s = s.split(',')
a = int(s[0])
b = epoch_to_millis(float(s[1]))
coppie.append((a,b))
return coppie
# +
#funzione per graficare
import matplotlib.pyplot as plt
def graph_rmse_distance(coppie):
x_val = [x[0] for x in coppie]
y_val = [x[1] for x in coppie]
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(x_val,y_val)
ax.plot(x_val,y_val,'or')
ax.set_title('Rmse Distance per number of iterations')
ax.set_ylabel('RMSE Distance (Gurobi -TensorFlow)')
ax.set_xlabel('Number of Iterations')
return plt.show()
def graph_time_per_iter(coppie):
x_val = [x[0] for x in coppie]
y_val = [x[1] for x in coppie]
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(x_val,y_val)
ax.plot(x_val,y_val,'or')
ax.set_title('Time Tensorflow per Number of Iterations')
ax.set_ylabel('Total Time Tensorflow)')
ax.set_xlabel('Number of Iteration')
return plt.show()
def graph_rmse_distance_and_execution_time(couples_rmse, couples_time):
x1 = [x[0] for x in couples_rmse]
x2 = [x[0] for x in couples_time]
y1 = [x[1] for x in couples_rmse]
y2 = [x[1] for x in couples_time]
plt.subplot(2, 1, 1)
plt.plot(x1, y1, 'ko-')
plt.title('Rmse Distance per Number of Iterations')
plt.xlabel('Number of Iterations')
plt.ylabel('RMSE Distance (Gurobi -TensorFlow)')
plt.subplot(2, 1, 2)
plt.plot(x2, y2, 'r.-')
plt.title('Time Tensorflow per Number of Iterations')
plt.xlabel('Number of Iterations')
plt.ylabel('Total Time Tensorflow)')
return plt.show()
# +
#funzione per tabella
def tab_rmse_distance(coppie):
print ("N_ITER RMSE_DISTANCE")
for i in coppie:
print ("{:<14}{:<11}".format(*i))
# +
path = "../../../log/Prove-4/Setosa/c1_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma01_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma025_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma05_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c1_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma01_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma025_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma05_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c75_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma01_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma01_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma025_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma025_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
#graph_time_per_iter(coppie_time)
#graph_rmse_distance_and_execution_time(coppie_rmse, coppie_time)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma05_penalization05_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# +
path = "../../../log/Prove-4/Setosa/c200_sigma05_penalization1_1e-05.log"
lines = start(path)
coppie_rmse = rmse_distance(lines)
coppie_time = time_per_iter(lines)
graph_rmse_distance(coppie_rmse)
tab_rmse_distance(coppie_rmse)
# -
| notebook/Grafici/Prove-4/Setosa-Rmse-Corretto-Tempi-Esecuzione.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import math
import statsmodels
from pandas_datareader import data
import datetime
import quandl
start= datetime.datetime(2015, 1, 1) # The start of the year for example
end= datetime.datetime(2020, 7, 30)
ticker_1 = "mts.ax"
ticker_2 = "wow.ax"
df_tickr1 = data.DataReader(name= ticker_1, data_source= "yahoo", start= start, end= end)
df_tickr2 = data.DataReader(name= ticker_2, data_source= "yahoo", start= start, end= end)
df_tickr1.tail()
df_tickr2.tail()
df_tickr1['Adj Close'].plot(label='{}'.format(ticker_1),figsize=(12,8))
df_tickr2['Adj Close'].plot(label='{}'.format(ticker_2))
plt.legend();
spread_ratio = df_tickr1['Adj Close']/df_tickr2['Adj Close']
spread_ratio.plot(label='Spread ratio',figsize=(12,8))
plt.axhline(spread_ratio.mean(),c='r')
plt.legend();
def zscore(stocks):
return (stocks - stocks.mean()) / np.std(stocks)
# +
#1 day moving average of the price spread
spreadratio_mavg1 = spread_ratio.rolling(1).mean()
# 30 day moving average of the price spread
spreadratio_mavg30 = spread_ratio.rolling(60).mean()
# Take a rolling 30 day standard deviation
std_30ratio = spread_ratio.rolling(60).std()
# Compute the z score for each day
zscore_30_1_ratio = (spreadratio_mavg1 - spreadratio_mavg30)/std_30ratio
zscore_30_1_ratio.plot(figsize=(12,8),label='Rolling 30 day Z score for spread ratio')
plt.axhline(0, color='black')
plt.axhline(2.5, color='red', linestyle='--')
plt.axhline(3.5, color='red', linestyle='-')
plt.axhline(-3.5, color='red', linestyle='-')
plt.axhline(-2.5, color='red',linestyle='--')
plt.legend(['Rolling 30 day z-score spread ratio', 'Mean', '+2.5 Sd', '-2.5 Sd']);
# -
from statsmodels.tsa.stattools import coint
y0 = df_tickr1['Adj Close']
y1 = df_tickr2['Adj Close']
t_stat_summary = statsmodels.tsa.stattools.coint(y0, y1, trend='c', method='aeg', maxlag=None, autolag='aic', return_results=True)
print('----------------------------------------')
print('T-stat for the pair {} VS {}:'.format(ticker_1,ticker_2), t_stat_summary[0])
print('Alpha value for the T-stat: {}'.format(t_stat_summary[1]))
print('--------------------------------------')
print('T-stat 99%: {}'.format(t_stat_summary[2][0]))
print('T-stat 95%: {}'.format(t_stat_summary[2][1]))
print('T-stat 90%: {}'.format(t_stat_summary[2][2]))
np.corrcoef(df_tickr1['Adj Close'],df_tickr2['Adj Close'])
np.cov(df_tickr1['Adj Close'],df_tickr2['Adj Close'])
# # Differencing method of spread; GARCH volatility method of spread and final use log returns minus average log returns of spread over GARCH - to be done next
zscore_30_1_ratio[-20:]
statsmodels.tsa.stattools.adfuller(spread_ratio, maxlag=None, regression='c', autolag='t-stat', store=False, regresults=False)
| Quant trading test models - pair co-integration test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building and Training Deep Neural Networks (DNNs) with Pytorch
#
#
# This lecture includes:
#
# 1. Build DNNs
# 3. Train MNIST with DNNs
# ## Class in Python
# Classes in Python provide a means of data and functionality together
# +
# Define a class
class Math497():
def __init__(self):
self.num_students = 40
num = 100
def add_students(self,x):
self.num_students += x
summer_course = Math497() # create a object
print('The number of students is:', summer_course.num_students)
summer_course.add_students(5)
print('The number of students is:', summer_course.num_students)
summer_course.add_students(-5)
print('The number of students is:', summer_course.num_students)
# -
# Define a child class of Math497()
class undergraduate(Math497):
def __init__(self):
super().__init__() # super().__init__() inherit parent's __init__() function.
self.num_undergraduate_students = 34
def add_undergraduate_students(self,x):
self.num_students += x # self.add_students(x) also works
self.num_undergraduate_students += x
# +
ug =undergraduate()
print('The number of undergraduate students in Math497 is:', ug.num_undergraduate_students)
print('The number of students in Math497 is:', ug.num_students)
ug.add_undergraduate_students(2)
print('The number of undergraduate students in Math497 is:', ug.num_undergraduate_students)
print('The number of students in Math497 is:', ug.num_students)
# -
# ## Module in Python
# Consider a module to be the same as a code library. To create a module just save the code you want in a file with the file extension .py
#
# See Course.py as example
# +
import Course
# If our module changed, we would have to reload it with the following commands
import imp
imp.reload(Course)
my_class = Course.Math497()
print('The number of students in my class is:', my_class.num_students)
# -
# ## 1. Build DNNs
# DNN includes:
#
# * input layer: given images $x$
#
# * $l$-hidden layers: denote $x^{0}=x$
# $$
# \begin{eqnarray}
# &x^{1} = \sigma (x^0{W^0}^{T}+b^0), &&\text{first hidden layer}\\
# &x^{2} = \sigma (x^1{W^1}^{T}+b^1), &&\text{second hidden layer}\\
# &\vdots &&\\
# &x^{l} = \sigma (x^{l-1}{W^{l-1}}^{T}+b^{l-1}), &&l\text{-th hidden layer}\\
# \end{eqnarray}
# $$
#
# * output layer: outputs$=(x^{l}{W^{l}}^{T}+b^{l})$
# +
import torch.nn as nn
import torch.nn.functional as F
# Note that:
# (1)torch.nn.Module is a Class
# (2)torch.nn is a Module
# You can not import torch.nn.Modules
class model(nn.Module): #
def __init__(self,input_size,num_classes):
super().__init__()
self.fc1 = nn.Linear(input_size, 500)
self.fc2 = nn.Linear(500, 250)
self.fc3 = nn.Linear(250, num_classes)
def forward(self, x): #Defines the computation performed at every call.
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
input_size = 784
num_classes = 10
hidden_size = 500
my_model =model(input_size, num_classes)
print(my_model.fc1.weight.size())
print(my_model.fc1.bias.size())
print(my_model.fc2.weight.size())
print(my_model.fc2.bias.size())
print(my_model.fc3.weight.size())
print(my_model.fc3.bias.size())
# Question: When we call model(images), the forward(self,x) will run automatically. Why? check __call__
# -
# ## 2. Train a DNN model on MNIST
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
import torch.nn.functional as F
# Define a 1-hidden layer neural network.
class model(nn.Module):
def __init__(self,input_size,hidden_size,num_classes):
super().__init__()
self.fc1 = nn.Linear(input_size, hidden_size)
self.fc2 = nn.Linear(hidden_size, num_classes)
def forward(self, x):
x = x.reshape(x.size(0), input_size) # you can reshape the iamges here.
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
input_size = 784
hidden_size = 500
num_classes = 10
minibatch_size = 128
num_epochs = 2
lr = 0.1
# Step 1: Define a model
my_model =model(input_size,hidden_size, num_classes)
# Step 2: Define a loss function and training algorithm
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(my_model.parameters(), lr=lr)
# Step 3: load dataset
MNIST_transform = torchvision.transforms.Compose([torchvision.transforms.ToTensor()])
trainset = torchvision.datasets.MNIST(root='./data', train= True, download=True, transform=MNIST_transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=minibatch_size)
testset = torchvision.datasets.MNIST(root='./data', train= False, download=True, transform=MNIST_transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=1)
#Step 4: Train the NNs
# One epoch is when an entire dataset is passed through the neural network only once.
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(trainloader):
#images = images.reshape(images.size(0), 28*28) # move this reshape to model class
# Forward pass to get the loss
outputs = my_model(images)
loss = criterion(outputs, labels)
# Backward and compute the gradient
optimizer.zero_grad()
loss.backward() #backpropragation
optimizer.step() #update the weights/parameters
# Training accuracy
correct = 0
total = 0
for i, (images, labels) in enumerate(trainloader):
#images = images.reshape(images.size(0), 28*28) # move this reshape to model class
outputs = my_model(images)
p_max, predicted = torch.max(outputs, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
training_accuracy = float(correct)/total
# Test accuracy
correct = 0
total = 0
for i, (images, labels) in enumerate(testloader):
#images = images.reshape(images.size(0), 28*28) # move this reshape to model class
outputs = my_model(images)
p_max, predicted = torch.max(outputs, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
test_accuracy = float(correct)/total
print('Epoch: {}, the training accuracy: {}, the test accuracy: {}' .format(epoch+1,training_accuracy,test_accuracy))
# -
# ## Reading material
#
# 1. Details of torch.nn https://pytorch.org/docs/stable/nn.html
#
# 2. Details of torch package https://pytorch.org/docs/stable/torch.html
#
| _build/jupyter_execute/Module3/C08_DNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 2*
#
# ---
#
#
# # Permutation & Boosting
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] If you haven't completed assignment #1, please do so first.
# - [ ] Continue to clean and explore your data. Make exploratory visualizations.
# - [ ] Fit a model. Does it beat your baseline?
# - [ ] Try xgboost.
# - [ ] Get your model's permutation importances.
#
# You should try to complete an initial model today, because the rest of the week, we're making model interpretation visualizations.
#
# But, if you aren't ready to try xgboost and permutation importances with your dataset today, that's okay. You can practice with another dataset instead. You may choose any dataset you've worked with previously.
#
# The data subdirectory includes the Titanic dataset for classification and the NYC apartments dataset for regression. You may want to choose one of these datasets, because example solutions will be available for each.
#
#
# ## Reading
#
# Top recommendations in _**bold italic:**_
#
# #### Permutation Importances
# - _**[Kaggle / <NAME>: Machine Learning Explainability](https://www.kaggle.com/dansbecker/permutation-importance)**_
# - [<NAME>: Interpretable Machine Learning](https://christophm.github.io/interpretable-ml-book/feature-importance.html)
#
# #### (Default) Feature Importances
# - [<NAME>: Selecting good features, Part 3, Random Forests](https://blog.datadive.net/selecting-good-features-part-iii-random-forests/)
# - [<NAME>, et al: Beware Default Random Forest Importances](https://explained.ai/rf-importance/index.html)
#
# #### Gradient Boosting
# - [A Gentle Introduction to the Gradient Boosting Algorithm for Machine Learning](https://machinelearningmastery.com/gentle-introduction-gradient-boosting-algorithm-machine-learning/)
# - _**[A Kaggle Master Explains Gradient Boosting](http://blog.kaggle.com/2017/01/23/a-kaggle-master-explains-gradient-boosting/)**_
# - [_An Introduction to Statistical Learning_](http://www-bcf.usc.edu/~gareth/ISL/ISLR%20Seventh%20Printing.pdf) Chapter 8
# - [Gradient Boosting Explained](http://arogozhnikov.github.io/2016/06/24/gradient_boosting_explained.html)
# - _**[Boosting](https://www.youtube.com/watch?v=GM3CDQfQ4sw) (2.5 minute video)**_
# -
# ### Continue to clean and explore your data. Make exploratory visualizations.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
train = pd.read_csv('train-01.csv')
# +
# Copying from the previous assignment.
df = pd.read_csv('assists-data-01.csv')
df2 = df.dropna(subset=['Target'])
test = df2[df2['Year'] == 2018]
temp = df2[df2['Year'] < 2018]
val = temp[temp['Year'] >= 2015]
test = test.reset_index(drop=True)
val = val.reset_index(drop=True)
# -
val.isnull().sum()
# Only missing values are the shooting percentages. I'll set those to 0 like I did for the train set.
val = val.fillna(value=0)
val.isnull().sum()
# +
# Need to temporarily fill in the missing values in Train before doing visualizations.
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean')
)
train_imputed = pd.DataFrame(pipeline.fit_transform(train), columns=train.columns)
# +
# %matplotlib inline
for i in range(len(train.columns)):
plt.scatter(train_imputed.iloc[:,i], train_imputed['Target'])
plt.xlabel(train_imputed.columns[i])
plt.ylabel('Target')
plt.show()
# -
# ### Fit a model. Does it beat your baseline?
# +
# Repeating from yesterday, here's my baseline:
from sklearn.metrics import mean_absolute_error
mean_absolute_error(val['AST'], val['Target'])
# +
train.describe(exclude=np.number)
# I'm going to exclude "Player." "League" can be one-hot encoded. As for "Pos"...
# -
train['Pos'].value_counts()
# +
# The positions have a natural order: PG = 1; SG = 2; SF = 3; PF = 4; C = 5.
# Numbers for the mixed positions will be obtained by averaging the respective numbers.
# 'G' will be interpreted as a PG/SG mix, and 'F' as a SF/PF mix.
# Often there are mixes in both orders (e.g. both PG-SG and SG-PG). While the intent here
# may have been to signal a small difference--e.g., a PG-SG is perhaps more of a "point guard"
# while a SG-PG is more of a "shooting guard"--for the purposes of this analysis, I'll just
# ignore such possible subtleties.
pos_encode = {'Pos': {
'PF': 4, 'C': 5, 'SG': 2, 'SF': 3, 'PG': 1,
'G': 1.5, 'G-F': 2.5, 'F': 3.5, 'F-C': 4.25, 'F-G': 2.5,
'C-F': 4.25, 'C-PF': 4.5, 'PG-SG': 1.5, 'SF-SG': 2.5, 'SG-SF': 2.5,
'PF-C': 4.5, 'PF-SF': 3.5, 'SG-PG': 1.5, 'SF-PF': 3.5, 'SG-PF': 3,
'PG-SF': 2, 'C-SF': 4, 'SF-PG': 2
}}
train2 = train.replace(to_replace=pos_encode)
train2['Pos'].value_counts()
# +
val2 = val.replace(to_replace=pos_encode)
val2['Pos'].value_counts()
# -
val2['Pos'] = val2['Pos'].astype(float)
val2.dtypes
# +
# The last decision I have to make regarding categorical encoding is for "Team."
pd.set_option('display.max_rows', 500)
train2['Tm'].value_counts(ascending=False)
# +
# It would probably be best to just drop this column, and I may do that eventually.
# But for a first pass, I'll reduce the cardinality and do one-hot encoding.
top50 = train2['Tm'].value_counts()[:50].index
train2.loc[~train2['Tm'].isin(top50), 'Tm'] = 'OTH'
train2['Tm'].value_counts()
# -
X_train = train2.drop(['Player', 'Target'], axis=1)
y_train = train2['Target']
X_val = val2.drop(['Player', 'Target'], axis=1)
y_val = val2['Target']
# +
from sklearn.linear_model import LinearRegression
from sklearn.feature_selection import f_regression, SelectKBest
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import f_regression, SelectKBest
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import RandomizedSearchCV
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
SelectKBest(f_regression),
LinearRegression()
)
param_distributions = {
'simpleimputer__strategy': ['mean', 'median', 'most_frequent'],
'selectkbest__k': range(1, len(X_train.columns)+1),
}
search = RandomizedSearchCV(
pipeline,
param_distributions=param_distributions,
n_iter=100,
cv=5,
scoring='neg_mean_absolute_error',
verbose=10,
return_train_score=True,
n_jobs=-1
)
search.fit(X_train, y_train);
# -
print('Best hyperparameters', search.best_params_)
print('Cross-validation mae', -search.best_score_)
# +
pipeline = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(strategy='mean'),
StandardScaler(),
SelectKBest(f_regression, k=30),
LinearRegression()
)
pipeline.fit(X_train, y_train)
# -
y_pred = pipeline.predict(X_val)
mean_absolute_error(y_val, y_pred)
baseline_mae = mean_absolute_error(val['AST'], val['Target'])
model1_mae = mean_absolute_error(y_val, y_pred)
print('The linear regression model improves on the baseline by', baseline_mae - model1_mae)
print(f'Relative to the target mean, the improvment is by {(baseline_mae - model1_mae) / np.mean(y_val) * 100}%')
# An improvement, but a really small one--less than 1%. Might just be statistical noise.
# +
# Let's try a random forest!
from sklearn.ensemble import RandomForestRegressor
from scipy.stats import randint, uniform
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestRegressor(random_state=110)
)
param_distributions = {
'simpleimputer__strategy': ['mean', 'median', 'most_frequent'],
'randomforestregressor__n_estimators': randint(50, 500),
'randomforestregressor__max_depth': [5, 10, 15, 20, None],
'randomforestregressor__max_features': uniform(0, 1),
}
search = RandomizedSearchCV(
pipeline,
param_distributions=param_distributions,
n_iter=10,
cv=3,
scoring='neg_mean_absolute_error',
verbose=10,
return_train_score=True,
n_jobs=-1
)
search.fit(X_train, y_train);
# -
print('Best hyperparameters', search.best_params_)
print('Cross-validation MAE', search.best_score_)
# +
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
RandomForestRegressor(max_depth=15, max_features=0.77, n_estimators=499, random_state=50)
)
pipeline.fit(X_train, y_train)
y_pred = pipeline.predict(X_val)
# +
model2_mae = mean_absolute_error(y_val, y_pred)
print('Random Forest mean absolute error:', model2_mae)
print('The radnom forest model improves on the baseline by', baseline_mae - model2_mae)
print(f'Relative to the target mean, the improvment is by {(baseline_mae - model2_mae) / np.mean(y_val) * 100}%')
# -
# A one percent improvement. Better!
# ### Try xgboost.
# +
from xgboost import XGBRegressor
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='median'),
XGBRegressor(n_estimators=100, random_state=30, n_jobs=-1)
)
pipeline.fit(X_train, y_train)
# +
y_pred = pipeline.predict(X_val)
model3_mae = mean_absolute_error(y_val, y_pred)
print('XGBoost mean absolute error:', model3_mae)
print('The XGBoost model improves on the baseline by', baseline_mae - model3_mae)
print(f'Relative to the target mean, the improvement is by {(baseline_mae - model3_mae) / np.mean(y_val) * 100}%')
# -
# Still not a very high improvement, but it's getting better!
# +
Encoder = ce.OrdinalEncoder()
Imputer = SimpleImputer(strategy='median')
X_train_encoded = Encoder.fit_transform(X_train)
X_val_encoded = Encoder.transform(X_val)
X_train_imputed = Imputer.fit_transform(X_train_encoded)
X_val_imputed = Imputer.fit_transform(X_val_encoded)
eval_set = [(X_train_imputed, y_train),
(X_val_imputed, y_val)]
model = XGBRegressor(
n_estimators=1000,
max_depth=4,
learning_rate=0.1, # Tried various combinations of max_depth and learning_rate and this seems to be the best.
n_jobs=-1
)
model.fit(X_train_imputed, y_train, eval_set=eval_set, eval_metric='mae', early_stopping_rounds=50)
# -
results = model.evals_result()
results
# +
train_error = results['validation_0']['mae']
val_error = results['validation_1']['mae']
epoch = range(1, len(train_error)+1)
plt.plot(epoch, train_error, label='Train')
plt.plot(epoch, val_error, label='Validation')
plt.ylabel('Mean Absolute Error')
plt.xlabel('Model Complexity (n_estimators)')
plt.legend();
# Is it a problem that these are smooth curves?
# +
y_pred = model.predict(X_val_imputed)
model4_mae = mean_absolute_error(y_val, y_pred)
print('Tuned XGBoost mean absolute error:', model4_mae)
print('The tuned XGBoost model improves on the baseline by', baseline_mae - model4_mae)
print(f'Relative to the target mean, the improvement is by {(baseline_mae - model4_mae) / np.mean(y_val) * 100}%')
# -
# Best improvement yet...slightly over 1.5%!
# ### Get your model's permutation importances.
# +
import eli5
from eli5.sklearn import PermutationImportance
permuter = PermutationImportance(
model,
scoring='neg_mean_absolute_error',
n_iter=5,
random_state=77
)
permuter.fit(X_val_imputed, y_val)
# +
feature_names = X_val.columns.tolist()
eli5.show_weights(
permuter,
top=None,
feature_names=feature_names
)
# -
# Unsurprisingly, previous year's assists is by far the most important feature. But a few of the others have some impact. In the future, I'll try removing the extraneous features and see if that helps the models.
| module2/assignment_applied_modeling_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"elapsed": 896, "status": "ok", "timestamp": 1594609251474, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="Omj1sv1rK1IG" outputId="35be439f-3ce9-49eb-e501-09c14c71eec1"
import nltk
from nltk.corpus.reader import ConllCorpusReader
# -
emergingE = ConllCorpusReader('emerging_entities_17-master/','.conll', ('words', 'pos', 'chunk'))
# + [markdown] id="Qh6qToBNRro4"
# ### Data Preparation
# + colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"elapsed": 2865, "status": "ok", "timestamp": 1594609255272, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="Gw331TN8SViw" outputId="3c7aad6b-1744-4997-851f-cb6c50353b89"
## Training and testing
train_sents = list(emergingE.tagged_sents('wnut17train.conll'))
valid_sents = list(emergingE.tagged_sents('emerging.dev.conll'))
test_sents = list(emergingE.tagged_sents('emerging.test.conll'))
print(train_sents[0])
#each tuple contains token, syntactic tag, ner label
# + id="JWIshaAS1mTk"
# functions of sentence representations for sequence labelling
def word2features(sent, i):
word = sent[i][0]
postag = sent[i][1]
features = {
'bias': 1.0,
'word.lower()': word.lower(),
'word[-3:]': word[-3:],
'word[-2:]': word[-2:],
'word.isupper()': word.isupper(),
'word.istitle()': word.istitle(),
'word.isdigit()': word.isdigit(),
'postag': postag,
'postag[:2]': postag[:2],
}
if i > 0:
word1 = sent[i-1][0]
postag1 = sent[i-1][1]
features.update({
'-1:word.lower()': word1.lower(),
'-1:word.istitle()': word1.istitle(),
'-1:word.isupper()': word1.isupper(),
'-1:postag': postag1,
'-1:postag[:2]': postag1[:2],
})
else:
# Indicate that it is the 'beginning of a document'
features['BOS'] = True
if i < len(sent)-1:
word1 = sent[i+1][0]
postag1 = sent[i+1][1]
features.update({
'+1:word.lower()': word1.lower(),
'+1:word.istitle()': word1.istitle(),
'+1:word.isupper()': word1.isupper(),
'+1:postag': postag1,
'+1:postag[:2]': postag1[:2],
})
else:
# Features for words that are not at the end of a document
features['EOS'] = True
return features
def sent2features(sent):
return [word2features(sent, i) for i in range(len(sent))]
def sent2labels(sent):
return [label for token, label in sent]
def sent2tokens(sent):
return [token for token, label in sent]
# + id="AmjwjGB817dp"
# sentence representations for sequence labelling
X_train = [sent2features(s) for s in train_sents]
y_train = [sent2labels(s) for s in train_sents]
X_valid = [sent2features(s) for s in valid_sents]
y_valid = [sent2labels(s) for s in valid_sents]
X_test = [sent2features(s) for s in test_sents]
y_test = [sent2labels(s) for s in test_sents]
# + colab={"base_uri": "https://localhost:8080/", "height": 225} executionInfo={"elapsed": 994, "status": "ok", "timestamp": 1594609267345, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="dIuFaNU82l-6" outputId="c7764679-e72e-46ab-ddb7-b36ba6011ecd"
train_sents[0], y_train[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 629, "status": "ok", "timestamp": 1594609268351, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="TVDPXw9FRrpm" outputId="aae004cd-89fe-4376-82f5-949679b871ea"
X_train[0], y_train[0]
# + [markdown] id="OkDc2ijbRrpq"
# ### Training
# Here we are using L-BFGS training algorithm (it is default) with Elastic Net (L1 + L2) regularization.
# + colab={"base_uri": "https://localhost:8080/", "height": 104} executionInfo={"elapsed": 4431, "status": "ok", "timestamp": 1594609275010, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="jJyWWkbx2I7p" outputId="0d9a56bb-6927-43e2-fb99-2ed0f65a1f08"
# train CRF model
import sklearn_crfsuite
crf = sklearn_crfsuite.CRF(
algorithm='lbfgs',
c1=0.1,
c2=0.1,
max_iterations=100,
all_possible_transitions=True
)
# + colab={"base_uri": "https://localhost:8080/", "height": 193} executionInfo={"elapsed": 968, "status": "ok", "timestamp": 1594609275987, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="Ja9VIf7SRrpt" outputId="5513aa73-40b8-469a-af96-e1cb2b36a64f"
crf
# + colab={"base_uri": "https://localhost:8080/", "height": 193} executionInfo={"elapsed": 45571, "status": "ok", "timestamp": 1594609320602, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="gCX9S-VbRrpw" outputId="697226db-d931-4787-f9ca-b44f790519af"
crf.fit(X_train, y_train, X_dev= X_valid, y_dev= y_valid)
# training model parameters
# + [markdown] id="u1rTPZHVRrpz"
# ### Evaluation
# There is much more O entities in data set, but we’re more interested in other entities. To account for this we’ll use averaged F1 score computed for all labels except for O. sklearn-crfsuite.metrics package provides some useful metrics for sequence classification task, including this one.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 45104, "status": "ok", "timestamp": 1594609320606, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="hN-RvWGZ25xJ" outputId="10efdf56-8a8c-49b4-e4be-0ef207be5fc7"
# get label set
labels = list(crf.classes_)
labels.remove('O')
print(labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 44592, "status": "ok", "timestamp": 1594609321103, "user": {"displayName": "AI3 member", "photoUrl": "", "userId": "02800686538475887838"}, "user_tz": -480} id="PdTxgk0Z3EX4" outputId="41425b81-b038-4f6d-b8d6-5a930e890359"
# evaluate CRF model
from sklearn_crfsuite import metrics
y_pred = crf.predict(X_test)
metrics.flat_f1_score(y_test, y_pred, average='weighted', labels=labels)
| A1/CRF_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
from library_task_3 import read_json, words
from wordcloud import WordCloud
battles = read_json('battles.json')
print(words(battles))
# Generate a word cloud image
wordcloud = WordCloud().generate(' '.join(words(battles)))
# Display the generated image:
# the matplotlib way:
plt.imshow(wordcloud, interpolation='bilinear', aspect='auto')
plt.axis("off")
plt.show()
# -
| 03/Cloud of Words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits import mplot3d
# %matplotlib inline
# -
# # Creating $x_i$, $y_i$ by drawing the data such that $x_i\epsilon\mathscr{N}(0, \sigma^2)$
#
# ### $$d = 1000, n = 500, \sigma^2 = 1, k = 100$$
#
# ### $$y_i = w^T . x_i + \epsilon_i$$
n = 500
d = 1000
k = 100 # k here denotes the number of relevant features
sigma = 1
mu = 0
# +
# Create the feature matrix using the numpy random standard normal with nxd dimension
X = mu + sigma**2 * np.random.standard_normal(size=(n,d))
# gaussian error
error = mu + sigma**2 * np.random.standard_normal(n)
# +
# creating the vector of weights w over the length of features
w_i = []
for j in range(d):
if j in range(k):
w_j = j/k
else:
w_j = 0
w_i.append(w_j)
w_i = np.array(w_i)
# +
y = []
for i in range(n):
y_i = np.inner(w_i, X[i,]) + error[i]
y.append(y_i)
y = (np.array(y)).reshape(-1,1)
# -
# # Solving the Lasso coordinate descent algorithm
# Define a function for soft thresholding
def soft_thresh(c_k, lamda):
"""
The soft threshold function based on a_k, c_k and lamda
"""
if c_k < - lamda:
return (c_k + lamda)
elif c_k > lamda:
return (c_k - lamda)
else:
return 0
# function to solve inner loop for the features till d
def inner_loop(weight, X, y, lamda, n_it=100):
n,d = X.shape
for it in range(n_it):
for k_it in range(k):
X_of_k = X[:,k_it].reshape(-1,1)
b = X @ weight
c_k = X_of_k.T @ (y - b + weight[k_it]*X_of_k)
weight[k_it] = soft_thresh(c_k, lamda)
return weight.flatten()
# +
int_weights = np.ones((X.shape[1],1))
weight_list = []
lamda = np.logspace(3.0, -3, num=100)
# the Outer loop for lamda
for lam in lamda:
weight = inner_loop(int_weights, X, y, lamda=lam)
weight_list.append(weight)
#Stack into numpy array
weights_lasso = np.stack(weight_list).T
#Plot results
n,_ = weights_lasso.shape
plt.figure(figsize = (14,8))
for i in range(n):
plt.plot(lamda[:len(weight_list)], weights_lasso[i])
plt.xscale('log')
plt.xlabel('Log($\\lambda$)')
plt.ylabel('Weights')
plt.title('Regularization Path')
plt.axis('tight')
# -
c = np.array([[ 1, 2, 3],
[-1, 1, 4]])
c
from tqdm import tqdm
import time
for i in tqdm(range(1000)):
a = i+5
| src/HW1/HW1_lasso_Coordinate_descent_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Как выложить бота на HEROKU
#
# *Подготовил <NAME>*
# Сразу оговоримся, что мы на heroku выкладываем
#
# **echo-Бота в телеграме, написанного с помощью библиотеки [pyTelegramBotAPI](https://github.com/eternnoir/pyTelegramBotAPI)**.
#
# А взаимодействие его с сервером мы сделаем с использованием [flask](http://flask.pocoo.org/)
#
# То есть вы боту пишете что-то, а он вам отвечает то же самое.
# ## Регистрация
# Идем к **@BotFather** в Telegram и по его инструкции создаем нового бота командой **/newbot**.
#
# Это должно закончиться выдачей вам токена вашего бота. Например последовательность команд, введенных мной:
#
# * **/newbot**
# * **my_echo_bot** (имя бота)
# * **ian_echo_bot** (ник бота в телеграме)
#
# Завершилась выдачей мне токена **1403467808:AAEaaLPkIqrhrQ62p7ToJclLtNNINdOopYk**
#
# И ссылки t.me/ian_echo_bot
#
# <img src="botfather.png">
# ## Регистрация на HEROKU
#
# Идем сюда: https://signup.heroku.com/login
#
# Создаем пользователя (это бесплатно)
#
# Попадаем на https://dashboard.heroku.com/apps и там создаем новое приложение:
# <img src="newapp1.png">
# Вводим название и регион (Я выбрал Европу), создаем.
# <img src="newapp2.png">
#
# После того, как приложение создано, нажмите, "Open App" и скопируйте адрес оттуда.
#
# <img src="newapp3.png">
#
# У меня это https://ian-echo-bot.herokuapp.com
# ## Установить интерфейсы heroku и git для командной строки
# Теперь надо установить Интерфейсы командной строки heroku и git по ссылкам:
#
# Сначала проверяем установлен ли git с помощью команды
#
# git --version
#
# Если терминал выдает ошибку, значит он не установлен
#
# Идем по ссылке, читаем инструкцию
#
# * https://git-scm.com/book/en/v2/Getting-Started-Installing-Git
#
# переходим на https://git-scm.com/download/mac
#
# и далее ставим GIT по указанной инструкции
#
# После чего переходим по ссылке и ставим интерфейс heroku
#
# * https://devcenter.heroku.com/articles/heroku-cli
#
# ## Установить библиотеки
# Теперь в вашем редакторе (например PyCharm) надо установить библиотеку для Телеграма и flask:
#
# * pip install pyTelegramBotAPI
# * pip install flask
# ## Код нашего echo-бота
#
# Вот этот код я уложил в файл main.py
# +
import os
import telebot
from flask import Flask, request
TOKEN = '<KEY>' # это мой токен
bot = telebot.TeleBot(token=TOKEN)
server = Flask(__name__)
# Если строка на входе непустая, то бот повторит ее
@bot.message_handler(func=lambda msg: msg.text is not None)
def reply_to_message(message):
bot.send_message(message.chat.id, message.text)
@server.route('/' + TOKEN, methods=['POST'])
def getMessage():
bot.process_new_updates([telebot.types.Update.de_json(request.stream.read().decode("utf-8"))])
return "!", 200
@server.route("/")
def webhook():
bot.remove_webhook()
bot.set_webhook(url='https://ian-echo-bot.herokuapp.com/' + TOKEN) #
return "!", 200
if __name__ == "__main__":
server.run(host="0.0.0.0", port=int(os.environ.get('PORT', 5000)))
# -
# ## Теперь создаем еще два файла для запуска
# **Procfile**(файл без расширения). Его надо открыть текстовым редактором и вписать туда строку:
#
# web: python main.py
# **requirements.txt** - файл со списком версий необходимых библиотек.
#
# Зайдите в PyCharm, где вы делаете проект и введите в терминале команду:
#
# pip freeze > requirements.txt
#
# В файле записи должны иметь вид:
#
# Название библиотеки==Версия библиотеки
#
# Если вдруг вы выдите что-то такое:
# <img src="versions.png">
#
# Удалите этот кусок текста, чтоб остался только номер версии и сохраните файл.
#
# Теперь надо все эти файлы уложить на гит, привязанный к Heroku и запустить приложение.
#
#
# ## Последний шаг
# Надо залогиниться в heroku через командную строку.
#
# Введите:
#
# heroku login
#
# Вас перебросит в браузер на вот такую страницу:
# <img src="login.png">
#
# После того, как вы залогинились, удостоверьтесь, что вы находитесь в папке, где лежат фаши файлы:
#
# main.py
# Procfile
# requirements.txt
# **Вводите команды:**
# git init
# git add .
# git commit -m "first commit"
# heroku git:remote -a ian-echo-bot
# git push heroku master
# По ходу выкатки вы увидите что-то такое:
# <img src="process.png">
#
# Готово, вы выложили вашего бота.
# Материалы, которыми можно воспользоваться по ходу выкладки бота на сервер:
#
# https://towardsdatascience.com/how-to-deploy-a-telegram-bot-using-heroku-for-free-9436f89575d2
#
# https://mattrighetti.medium.com/build-your-first-telegram-bot-using-python-and-heroku-79d48950d4b0
| lect13/heroku.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="9YOJF4IfmVMR" outputId="d6dab74e-a866-426a-c314-fb4521f16613"
_parsed_rows = []
def parse_csv():
import csv
_file_path = "../nyc_weather.csv"
global _parsed_rows
with open(_file_path, "r") as f:
reader = csv.reader(f, delimiter=',')
reader.next()
for row in reader:
_parsed_rows.append({
'date': row[0],
'temperature': row[1],
'DewPoint': row[2],
'Humidity': row[3],
'Sea_Level_PressureIn': row[4],
'VisibilityMiles': row[5],
'WindSpeedMPH': row[6],
'PrecipitationIn': row[7],
'CloudCover': row[8],
'Events': row[9],
'WindDirDegrees': row[10]
})
def get_max_temperature():
max_temp = 0
for row in _parsed_rows:
if int(row['temperature']) > max_temp:
max_temp = int(row['temperature'])
return max_temp
def get_days_of_rain(event):
days = []
for row in _parsed_rows:
if row['Events'] == event:
days.append(row['date'])
return days
def get_avg_wind_speed():
total = 0
count = 0
for row in _parsed_rows:
speed = 0 if row['WindSpeedMPH']=='' else int(row['WindSpeedMPH'])
total += speed
count += 1
return float(total/count)
if __name__ == '__main__':
parse_csv()
print "Max Temperature is : " + str(get_max_temperature())
print "Days of rain : " + str(get_days_of_rain('Rain'))
print "Average wind speed is : " + str(get_avg_wind_speed())
# + colab={} colab_type="code" id="9-9zh0ldmVMb"
| Module 1 - Fundamentals of programming/pandas/pandas introduction/pandas_intro.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ENV/ATM 415: Climate Laboratory
#
# [<NAME>](http://www.atmos.albany.edu/facstaff/brose/index.html), University at Albany
#
# # Lecture 17: Ice-albedo feedback and Snowball Earth in the EBM
# ## Contents
#
# 1. [Review of the 1D EBM](#section1)
# 2. [Interactive snow/ice line in the EBM](#section2)
# 3. [Solving the EBM with variable snow/ice line in CLIMLAB](#section3)
# 4. [Polar-amplified warming in the EBM](#section4)
# 5. [A different kind of climate forcing: changing the solar constant](#section5)
# 6. [The large ice cap instability](#section6)
# 7. [The Neoproterozoic Snowball Earth](#section7)
# 8. [Computing the complete hysteresis curve for the 1D diffusive EBM](#section8)
# ____________
# <a id='section1'></a>
#
# ## 1. Review of the 1D EBM
# ____________
# Last time we derived the equation for the one-dimensional EBM with diffusive heat transport:
# $$ C \frac{\partial T}{\partial t} = (1-\alpha) ~ Q - \left( A + B~T \right) + \frac{D}{\cos\phi } \frac{\partial }{\partial \phi} \left( \cos\phi ~ \frac{\partial T}{\partial \phi} \right) $$
#
# We have chosen the following parameter values, which seems to give a reasonable fit to the observed **annual mean temperature and energy budget**:
#
# - $ A = 210 ~ \text{W m}^{-2}$ (emission at 0$^\circ$C)
# - $ B = 2 ~ \text{W m}^{-2}~^\circ\text{C}^{-1} $ (increase in emission per degree, related to net longwave climate feedback)
# - $ D = 0.6 ~ \text{W m}^{-2}~^\circ\text{C}^{-1} $ (thermal diffusivity of the climate system)
# We looked at the adjustment of this model to equilibrium, with annual mean insolation $\overline{Q(\phi)}$ and specified albedo $\alpha(\phi)$ (giving a reasonable fit to observations).
#
# We also tuned the diffuvisity $D$ so that our annual mean solution has a reasonable pole-to-equator temperature gradient and peak poleward heat transport.
#
# Actually for the new version of this model with interactive ice line, we are going to reduce the diffusivity down to $ D = 0.55 ~ \text{W m}^{-2}~^\circ\text{C}^{-1} $.
# ____________
# <a id='section2'></a>
#
# ## 2. Interactive snow/ice line in the EBM
# ____________
# What we want to do today is introduce another process into our model: an **interactive snow and ice line**.
#
# The idea is simply that, as the climate gets warmer, the snow and ice will retreat poleward, and the planetary albedo will decrease (or vice-versa).
# We modeled this in the zero-dimensional model by using a kind of ramp function for the global mean albedo as a function of global mean temperature.
# Here, since our model resolves temperature at each latitude, we want to do something more physical: *suppose that the surface is covered in ice and snow wherever the temperature is below some threshold $T_f$.*
# ### Temperature-dependent ice line
#
# Let the surface albedo be larger wherever the temperature is below some threshold $T_f$:
#
# $$ \alpha\left(\phi, T(\phi) \right) = \left\{\begin{array}{ccc}
# \alpha_0 + \alpha_2 P_2(\sin\phi) & ~ & T(\phi) > T_f & \text{(no ice)} \\
# a_i & ~ & T(\phi) \le T_f & \text{(ice-covered)} \\
# \end{array} \right. $$
#
# where $P_2(\sin\phi) = \frac{1}{2}\left( 3\left(\sin\phi\right)^2 - 1 \right) $ is called the *second Legendre Polynomial* (just a mathematically convenient description of a smooth variation between the equator and pole).
# Empirically, we follow classic work by Budyko and set the threshold temperature
#
# $$ T_f = -10^\circ\text{C} $$
# This is known as a "step function" formula, because the value of $\alpha$ steps or jumps up to a higher value as we cross the ice line.
# ____________
# <a id='section3'></a>
#
# ## 3. Solving the EBM with variable snow/ice line in CLIMLAB
# ____________
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import climlab
from climlab import constants as const
# for convenience, set up a dictionary with our reference parameters
param = {'D':0.55, 'A':210, 'B':2, 'a0':0.3, 'a2':0.078, 'ai':0.62, 'Tf':-10.}
model1 = climlab.EBM_annual( num_lat=180, D=0.55, A=210., B=2., Tf=-10., a0=0.3, a2=0.078, ai=0.62)
print(model1)
# Because we provided a parameter `ai` for the icy albedo, our model now contains several sub-processes contained within the process called `albedo`. Together these implement the step-function formula above.
#
# The process called `iceline` simply looks for grid cells with temperature below $T_f$.
print(model1.param)
# +
# A python shortcut... we can use the dictionary to pass lots of input arguments simultaneously:
# same thing as before, but written differently:
model1 = climlab.EBM_annual( num_lat=180, **param)
print(model1)
# -
def ebm_plot(e, return_fig=False):
templimits = -60,32
radlimits = -340, 340
htlimits = -6,6
latlimits = -90,90
lat_ticks = np.arange(-90,90,30)
fig = plt.figure(figsize=(8,12))
ax1 = fig.add_subplot(3,1,1)
ax1.plot(e.lat, e.Ts)
ax1.set_ylim(templimits)
ax1.set_ylabel('Temperature (deg C)')
ax2 = fig.add_subplot(3,1,2)
ax2.plot(e.lat, e.ASR, 'k--', label='SW' )
ax2.plot(e.lat, -e.OLR, 'r--', label='LW' )
ax2.plot(e.lat, e.net_radiation, 'c-', label='net rad' )
ax2.plot(e.lat, e.heat_transport_convergence, 'g--', label='dyn' )
ax2.plot(e.lat, e.net_radiation.squeeze() + e.heat_transport_convergence, 'b-', label='total' )
ax2.set_ylim(radlimits)
ax2.set_ylabel('Energy budget (W m$^{-2}$)')
ax2.legend()
ax3 = fig.add_subplot(3,1,3)
ax3.plot(e.lat_bounds, e.heat_transport )
ax3.set_ylim(htlimits)
ax3.set_ylabel('Heat transport (PW)')
for ax in [ax1, ax2, ax3]:
ax.set_xlabel('Latitude')
ax.set_xlim(latlimits)
ax.set_xticks(lat_ticks)
ax.grid()
if return_fig:
return fig
model1.integrate_years(5)
# Diagnostics in this model are mostly on the latitude axis, e.g.:
model1.ASR
# There is a built-in utility to take properly area-weighted global averages!
# This gives us the net global energy imbalance:
climlab.global_mean(model1.ASR - model1.OLR)
# There is also a built-in diagnostic `net_radiation` that has the same information (i.e. ASR - OLR):
climlab.global_mean(model1.net_radiation)
# Since it's not fully equilibrated yet, we can run it out a little longer:
# Integrate out to equilibrium.
model1.integrate_years(5)
# Check for energy balance
print(climlab.global_mean(model1.net_radiation))
f = ebm_plot(model1)
# There is a diagnostic that tells us the current location of the ice edge:
model1.icelat
# This model is tuned up to reasonable "present-day" conditions.
# ____________
# <a id='section4'></a>
#
# ## 4. Polar-amplified warming in the EBM
# ____________
# ### Add a small radiative forcing
#
# The equivalent of doubling CO2 in this model is something like
#
# $$ A \rightarrow A - \delta A $$
#
# where $\delta A = 4$ W m$^{-2}$.
#
#
model1.subprocess['LW'].A
# +
deltaA = 4.
# This is a very handy way to "clone" an existing model:
model2 = climlab.process_like(model1)
# Now change the longwave parameter:
model2.subprocess['LW'].A = param['A'] - deltaA
# and integrate out to equilibrium again
model2.integrate_years(5, verbose=False)
plt.plot(model1.lat, model1.Ts, label='model1')
plt.plot(model2.lat, model2.Ts, label='model2')
plt.legend(); plt.grid()
# -
# **The warming is polar-amplified**: more warming at the poles than elsewhere.
#
# Why?
#
# Also, the current ice line is now:
model2.icelat
# There is no ice left!
# Let's do some more greenhouse warming:
# +
model3 = climlab.process_like(model1)
model3.subprocess['LW'].A = param['A'] - 2*deltaA
model3.integrate_years(5, verbose=False)
plt.plot(model1.lat, model1.Ts, label='model1')
plt.plot(model2.lat, model2.Ts, label='model2')
plt.plot(model3.lat, model3.Ts, label='model3')
plt.xlim(-90, 90)
plt.grid()
plt.legend()
# -
# In the ice-free regime, there is no polar-amplified warming. A uniform radiative forcing produces a uniform warming.
# ____________
# <a id='section5'></a>
#
# ## 5. A different kind of climate forcing: changing the solar constant
# ____________
# Historically EBMs have been used to study the climatic response to a change in the energy output from the Sun.
#
# We can do that easily with `climlab`:
m = climlab.EBM_annual( num_lat=180, **param )
# The current (default) solar constant, corresponding to present-day conditions:
m.subprocess.insolation.S0
# ### What happens if we decrease $S_0$?
# First, get to equilibrium
m.integrate_years(5.)
# Check for energy balance
climlab.global_mean(m.net_radiation)
m.icelat
# Now make the solar constant smaller:
m.subprocess.insolation.S0 = 1300.
# Integrate to new equilibrium
m.integrate_years(10.)
# Check for energy balance
climlab.global_mean(m.net_radiation)
m.icelat
ebm_plot(m)
# A much colder climate! The ice line is sitting at 54º. The heat transport shows that the atmosphere is moving lots of energy across the ice line, trying hard to compensate for the strong radiative cooling everywhere poleward of the ice line.
# ____________
# <a id='section6'></a>
#
# ## 6. The large ice cap instability
# ____________
# ### What happens if we decrease $S_0$ even more?
#
# Now make the solar constant smaller:
m.subprocess.insolation.S0 = 1200.
# First, get to equilibrium
m.integrate_years(5.)
# Check for energy balance
climlab.global_mean(m.net_radiation)
m.integrate_years(10.)
# Check for energy balance
climlab.global_mean(m.net_radiation)
ebm_plot(m)
# Something very different happened! Where is the ice line now?
m.icelat
# ### Now what happens if we set $S_0$ back to its present-day value?
# Now make the solar constant smaller:
m.subprocess.insolation.S0 = 1365.2
# First, get to equilibrium
m.integrate_years(5.)
# Check for energy balance
climlab.global_mean(m.net_radiation)
ebm_plot(m)
# Is this the same climate we started with?
# This is an example of a **hysteresis** in the climate system: the state of the climate depends on its history!
#
# - A global cooling caused snow and ice to expand to the equator
# - External conditions (i.e. the solar constant) returned back to its present-day value
# - The climate stayed cold and completely ice-covered.
#
# If the oceans froze over and the Earth were covered in ice and snow today, it would remain that way!
# ____________
# <a id='section7'></a>
#
# ## 7. The Neoproterozoic Snowball Earth
# ____________
# ### The Geologic Time Scale
#
# First, some information on the nomenclature for Earth history:
# 
# > <NAME>. and <NAME>. (2009). Geologic time scale. Technical report, Geological Society of America.
# The long view of glacial epochs on Earth:
# <img src="http://www.atmos.albany.edu/facstaff/brose/classes/ENV415_Spring2018/images/Hoffman_etal_SciAdv_Fig2.pdf" width="800">
# > <NAME> al. (2017), Science Advances 3:e1600983, doi:10.1126/sciadv.1600983
# ### Extensive evidence for large glaciers at sea level in the tropics
#
# Evidently the climate was **very cold** at these times (635 Ma and 715 Ma)
# 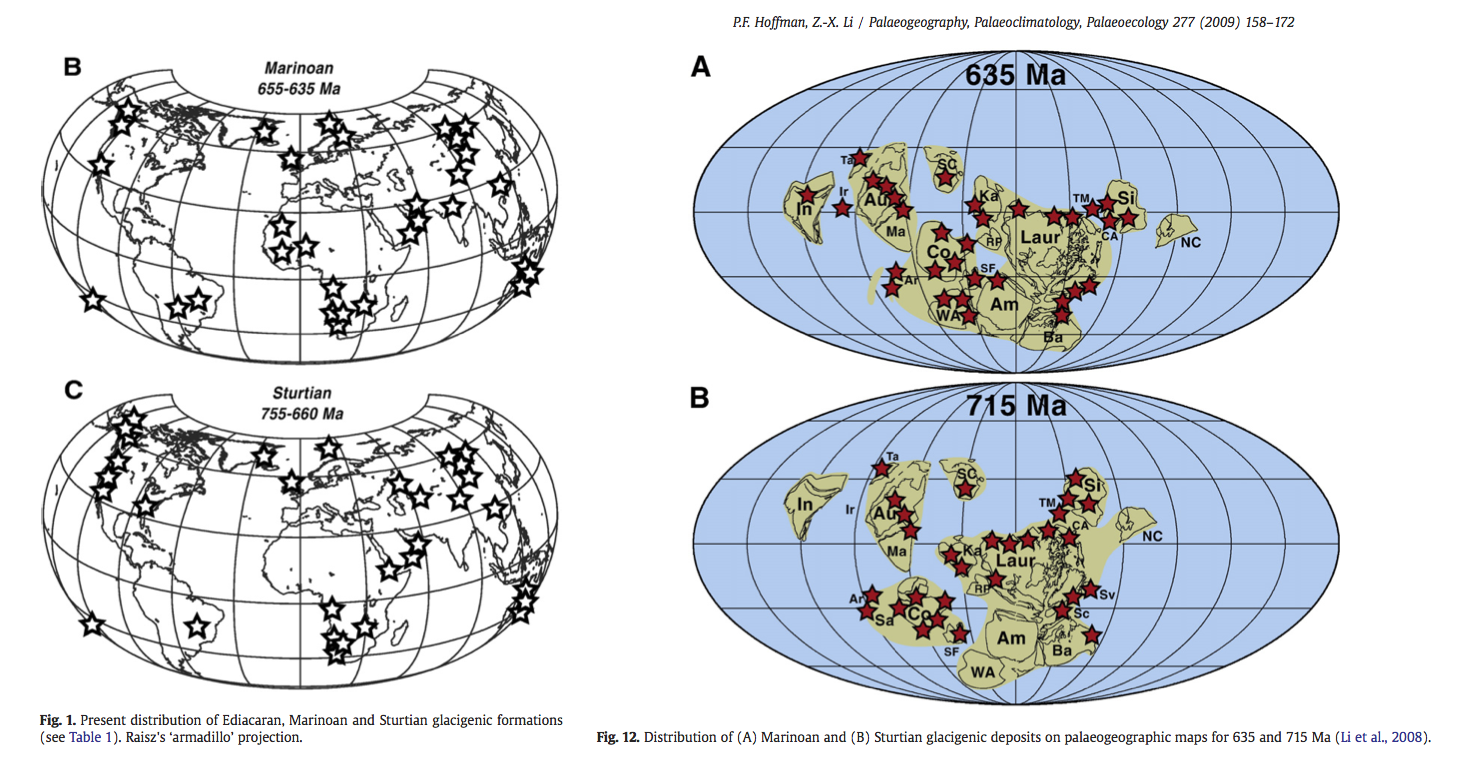
# > <NAME>. and <NAME>. (2009). A palaeogeographic context for Neoproterozoic glaciation. Palaeogeogr. Palaeoclimatol. Palaeoecol., 277:158–172.
# ### The Snowball Earth hypothesis
#
# Various bizarre features in the geological record from 635 and 715 Ma ago indicate that the Earth underwent some very extreme environmental changes… at least twice. The **Snowball Earth hypothesis** postulates that:
#
# - The Earth was completely ice-covered (including the oceans)
# - The total glaciation endured for millions of years
# - CO$_2$ slowly accumulated in the atmosphere from volcanoes
# - Weathering of rocks (normally acting to reduce CO$_2$) extremely slow due to cold, dry climate
# - Eventually the extreme greenhouse effect is enough to melt back the ice
# - The Earth then enters a period of extremely hot climate.
# The hypothesis rests on a phenomenon first discovered by climate modelers in the Budyko-Sellers EBM: **runaway ice-albedo feedback** or **large ice cap instability**.
# ____________
# <a id='section8'></a>
#
# ## 8. Computing the complete hysteresis curve for the 1D diffusive EBM
# ____________
# <div class="alert alert-warning">
# The calculations in this section may take a long time to complete, depending on the speed of your computer.
# </div>
# The ice edge in our model is always where the temperature crosses $T_f = -10^\circ$C. The system is at **equilibrium** when the temperature is such that there is a balance between ASR, OLR, and heat transport convergence everywhere.
#
# Suppose that sun was hotter or cooler at different times (in fact it was significantly cooler during early Earth history). That would mean that the solar constant $S_0 = 4Q$ was larger or smaller. We should expect that the temperature (and thus the ice edge) should increase and decrease as we change $S_0$.
# $S_0$ during the Neoproterozoic Snowball Earth events is believed to be about 93% of its present-day value, or about 1270 W m$^{-2}$.
#
# We are going to look at how the **equilibrium** ice edge depends on $S_0$, by integrating the model out to equilibrium for lots of different values of $S_0$. We will start by slowly decreasing $S_0$, and then slowly increasing $S_0$.
model2 = climlab.EBM_annual(num_lat = 360, **param)
S0array = np.linspace(1400., 1200., 200)
model2.integrate_years(5)
icelat_cooling = np.empty_like(S0array)
icelat_warming = np.empty_like(S0array)
# First cool....
for n in range(S0array.size):
model2.subprocess['insolation'].S0 = S0array[n]
model2.integrate_years(10, verbose=False)
icelat_cooling[n] = np.max(model2.icelat)
# Then warm...
for n in range(S0array.size):
model2.subprocess['insolation'].S0 = np.flipud(S0array)[n]
model2.integrate_years(10, verbose=False)
icelat_warming[n] = np.max(model2.icelat)
# For completeness -- also start from present-day conditions and warm up.
model3 = climlab.EBM_annual(num_lat=360, **param)
S0array3 = np.linspace(1350., 1400., 50)
icelat3 = np.empty_like(S0array3)
for n in range(S0array3.size):
model3.subprocess['insolation'].S0 = S0array3[n]
model3.integrate_years(10, verbose=False)
icelat3[n] = np.max(model3.icelat)
fig = plt.figure( figsize=(10,6) )
ax = fig.add_subplot(111)
ax.plot(S0array, icelat_cooling, 'r-', label='cooling' )
ax.plot(S0array, icelat_warming, 'b-', label='warming' )
ax.plot(S0array3, icelat3, 'g-', label='warming' )
ax.set_ylim(-10,100)
ax.set_yticks((0,15,30,45,60,75,90))
ax.grid()
ax.set_ylabel('Ice edge latitude', fontsize=16)
ax.set_xlabel('Solar constant (W m$^{-2}$)', fontsize=16)
ax.plot( [const.S0, const.S0], [-10, 100], 'k--', label='present-day' )
ax.legend(loc='upper left')
ax.set_title('Solar constant versus ice edge latitude in the EBM with albedo feedback', fontsize=16);
# There are actually up to 3 different climates possible for a given value of $S_0$!
# ### How to un-freeze the Snowball
# The graph indicates that if the Earth were completely frozen over, it would be perfectly happy to stay that way even if the sun were brighter and hotter than it is today.
#
# Our EBM predicts that (with present-day parameters) the equilibrium temperature at the equator in the Snowball state is about -33ºC, which is much colder than the threshold temperature $T_f = -10^\circ$C. How can we melt the Snowball?
# We need to increase the avaible energy sufficiently to get the equatorial temperatures above this threshold! That is going to require a much larger increase in $S_0$ (could also increase the greenhouse gases, which would have a similar effect)!
#
# Let's crank up the sun to 1830 W m$^{-2}$ (about a 35% increase from present-day).
# +
my_ticks = [-90,-60,-30,0,30,60,90]
model4 = climlab.process_like(model2) # initialize with cold Snowball temperature
model4.subprocess['insolation'].S0 = 1830.
model4.integrate_years(40)
plt.plot(model4.lat, model4.Ts)
plt.xlim(-90,90); plt.ylabel('Temperature'); plt.xlabel('Latitude')
plt.grid(); plt.xticks(my_ticks)
print('The ice edge is at ' + str(model4.icelat) + ' degrees latitude.' )
# -
# Still a Snowball... but just barely! The temperature at the equator is just below the threshold.
#
# Try to imagine what might happen once it starts to melt. The solar constant is huge, and if it weren't for the highly reflective ice and snow, the climate would be really really hot!
#
# We're going to increase $S_0$ one more time...
model4.subprocess['insolation'].S0 = 1840.
model4.integrate_years(10)
plt.plot(model4.lat, model4.Ts)
plt.xlim(-90,90); plt.ylabel('Temperature'); plt.xlabel('Latitude')
plt.grid(); plt.xticks(my_ticks);
# Suddenly the climate looks very very different again! The global mean temperature is
model4.global_mean_temperature()
# A roasty 58ºC, and the poles are above 20ºC. A tiny increase in $S_0$ has led to a very drastic change in the climate.
# Now we will complete the plot of ice edge versus solar constant.
# +
S0array_snowballmelt = np.linspace(1400., 1900., 50)
icelat_snowballmelt = np.empty_like(S0array_snowballmelt)
icelat_snowballmelt_cooling = np.empty_like(S0array_snowballmelt)
for n in range(S0array_snowballmelt.size):
model2.subprocess['insolation'].S0 = S0array_snowballmelt[n]
model2.integrate_years(10, verbose=False)
icelat_snowballmelt[n] = np.max(model2.icelat)
for n in range(S0array_snowballmelt.size):
model2.subprocess['insolation'].S0 = np.flipud(S0array_snowballmelt)[n]
model2.integrate_years(10, verbose=False)
icelat_snowballmelt_cooling[n] = np.max(model2.icelat)
# -
fig = plt.figure( figsize=(18,6) )
ax = fig.add_subplot(111)
ax.plot(S0array, icelat_cooling, 'r-', label='cooling' )
ax.plot(S0array, icelat_warming, 'b-', label='warming' )
ax.plot(S0array3, icelat3, 'g-', label='warming' )
ax.plot(S0array_snowballmelt, icelat_snowballmelt, 'b-' )
ax.plot(S0array_snowballmelt, icelat_snowballmelt_cooling, 'r-' )
ax.set_ylim(-10,100)
ax.set_yticks((0,15,30,45,60,75,90))
ax.grid()
ax.set_ylabel('Ice edge latitude', fontsize=16)
ax.set_xlabel('Solar constant (W m$^{-2}$)', fontsize=16)
ax.plot( [const.S0, const.S0], [-10, 100], 'k--', label='present-day' )
ax.legend(loc='upper left')
ax.set_title('Solar constant versus ice edge latitude in the EBM with albedo feedback', fontsize=16);
# The upshot:
#
# - For extremely large $S_0$, the only possible climate is a hot Earth with no ice.
# - For extremely small $S_0$, the only possible climate is a cold Earth completely covered in ice.
# - For a large range of $S_0$ including the present-day value, more than one climate is possible!
# - Once we get into a Snowball Earth state, getting out again is rather difficult!
| notes/L17_albedo_feedback.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dealing with Messy Data
# > This chapter introduces you to the reality of messy and incomplete data. You will learn how to find where your data has missing values and explore multiple approaches on how to deal with them. You will also use string manipulation techniques to deal with unwanted characters in your dataset. This is the Summary of lecture "Feature Engineering for Machine Learning in Python", via datacamp.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Datacamp, Machine_Learning]
# - image:
import pandas as pd
import numpy as np
# ## Why do missing values exist?
# - How gaps in data occur
# - Data not being collected properly
# - Collection and management errors
# - Data intentionally being omitted
# - Could be created due to transformations of the data
# - Why we care?
# - Some models cannot work with missing data (Nulls/NaN)
# - Missing data may be a sign a wider data issue
# - Missing data can be a useful feature
# ### How sparse is my data?
# Most data sets contain missing values, often represented as NaN (Not a Number). If you are working with Pandas you can easily check how many missing values exist in each column.
#
# Let's find out how many of the developers taking the survey chose to enter their age (found in the `Age` column of `so_survey_df`) and their gender (`Gender` column of `so_survey_df`).
so_survey_df = pd.read_csv('./dataset/Combined_DS_v10.csv')
so_survey_df.head()
# +
# Subset the DataFrame
sub_df = so_survey_df[['Age', 'Gender']]
# Print the number of non-missing values
print(sub_df.notnull().sum())
# -
# ### Finding the missing values
# While having a summary of how much of your data is missing can be useful, often you will need to find the exact locations of these missing values. Using the same subset of the StackOverflow data from the last exercise (`sub_df`), you will show how a value can be flagged as missing.
# Print the top 10 entries of the DataFrame
sub_df.head(10)
# Print the locations of the missing values
sub_df.head(10).isnull()
# Print the locations of the missing values
sub_df.head(10).notnull()
# ## Dealing with missing values (I)
# - Issues with deletion
# - It deletes vaild data points
# - Relies on randomness
# - Reduces information
# ### Listwise deletion
# The simplest way to deal with missing values in your dataset when they are occurring entirely at random is to remove those rows, also called **'listwise deletion'**.
#
# Depending on the use case, you will sometimes want to remove all missing values in your data while other times you may want to only remove a particular column if too many values are missing in that column.
# Print the number of rows and columns
print(so_survey_df.shape)
# +
# Create a new DataFrame dropping all incomplete rows
no_missing_values_rows = so_survey_df.dropna()
# Print the shape of the new DataFrame
print(no_missing_values_rows.shape)
# +
# Create a new DataFrame dropping all columns with incomplete rows
no_missing_values_cols = so_survey_df.dropna(axis=1)
# Print the shape fo the new DataFrame
print(no_missing_values_cols.shape)
# +
# Drop all rows where Gender is missing
no_gender = so_survey_df.dropna(subset=['Gender'])
# Print the shape of the new DataFrame
print(no_gender.shape)
# -
# ### Replacing missing values with constants
# While removing missing data entirely maybe a correct approach in many situations, this may result in a lot of information being omitted from your models.
#
# You may find categorical columns where the missing value is a valid piece of information in itself, such as someone refusing to answer a question in a survey. In these cases, you can fill all missing values with a new category entirely, for example 'No response given'.
# Print the count of occurrence
print(so_survey_df['Gender'].value_counts())
# +
# Replace missing values
so_survey_df['Gender'].fillna('Not Given', inplace=True)
# Print the count of each value
print(so_survey_df['Gender'].value_counts())
# -
# ## Dealing with missing values (II)
# - Deleting missing values
# - Can't delete rows with missing values in the test set
# - What else can you do?
# - Categorical columns: Replace missing values with the most common occurring value or with a string that flags missing values such as 'None'
# - Numerical columns: Replace missing values with a suitable value
# ### Filling continuous missing values
# In the last lesson, you dealt with different methods of removing data missing values and filling in missing values with a fixed string. These approaches are valid in many cases, particularly when dealing with categorical columns but have limited use when working with continuous values. In these cases, it may be most valid to fill the missing values in the column with a value calculated from the entries present in the column.
#
#
# Print the first five rows of StackOverflowJobsRecommend column
so_survey_df['StackOverflowJobsRecommend'].head()
# +
# Fill missing values with the mean
so_survey_df['StackOverflowJobsRecommend'].fillna(so_survey_df['StackOverflowJobsRecommend'].mean(),
inplace=True)
# Round the StackOverflowJobsRecommend values
so_survey_df['StackOverflowJobsRecommend'] = round(so_survey_df['StackOverflowJobsRecommend'])
# Print the first five rows of StackOverflowJobsRecommend column
so_survey_df['StackOverflowJobsRecommend'].head()
# -
# ## Dealing with other data issues
#
# ### Dealing with stray characters (I)
# In this exercise, you will work with the `RawSalary` column of `so_survey_df` which contains the wages of the respondents along with the currency symbols and commas, such as `$42,000`. When importing data from Microsoft Excel, more often that not you will come across data in this form.
#
#
# +
# Remove the commas in the column
so_survey_df['RawSalary'] = so_survey_df['RawSalary'].str.replace(',', '')
# Remove the dollar signs in the column
so_survey_df['RawSalary'] = so_survey_df['RawSalary'].str.replace('$', '')
# -
# ### Dealing with stray characters (II)
# In the last exercise, you could tell quickly based off of the `df.head()` call which characters were causing an issue. In many cases this will not be so apparent. There will often be values deep within a column that are preventing you from casting a column as a numeric type so that it can be used in a model or further feature engineering.
#
# One approach to finding these values is to force the column to the data type desired using `pd.to_numeric()`, coercing any values causing issues to NaN, Then filtering the DataFrame by just the rows containing the NaN values.
#
# Try to cast the `RawSalary` column as a float and it will fail as an additional character can now be found in it. Find the character and remove it so the column can be cast as a float.
# +
# Attempt to convert the column to numeric values
numeric_vals = pd.to_numeric(so_survey_df['RawSalary'], errors='coerce')
# find the indexes of missing values
idx = so_survey_df['RawSalary'].isna()
# Print the relevant raws
print(so_survey_df['RawSalary'][idx])
# +
# Replace the offending characters
so_survey_df['RawSalary'] = so_survey_df['RawSalary'].str.replace('£', '')
# Convert the column to float
so_survey_df['RawSalary'] = so_survey_df['RawSalary'].astype(float)
# Print the column
so_survey_df['RawSalary']
# -
# ### Method chaining
# When applying multiple operations on the same column (like in the previous exercises), you made the changes in several steps, assigning the results back in each step. However, when applying multiple successive operations on the same column, you can "chain" these operations together for clarity and ease of management. This can be achieved by calling multiple methods sequentially:
#
# ```python
# # Method chaining
# df['column'] = df['column'].method1().method2().method3()
#
# # Same as
# df['column'] = df['column'].method1()
# df['column'] = df['column'].method2()
# df['column'] = df['column'].method3()
# ```
# In this exercise you will repeat the steps you performed in the last two exercises, but do so using method chaining.
# +
so_survey_df = pd.read_csv('./dataset/Combined_DS_v10.csv')
# Use method chaining
so_survey_df['RawSalary'] = so_survey_df['RawSalary']\
.str.replace(',', '')\
.str.replace('$', '')\
.str.replace('£', '')\
.astype(float)
# Print the RawSalary column
print(so_survey_df['RawSalary'])
| _notebooks/2020-07-12-02-Dealing-with-Messy-Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
fidx = 'ens'
midx = '66'
import numpy as np
import pickle
# class-specific ensemble weights
weights = pickle.load(open('ens_weights66.pkl','rb'))
print(len(weights))
glist = ['air_u67',
'preresnet_u67','preresnet_u101',
'resnet_w65'
]
glist += ['iv3_q11','irv2_q10','resnet_q24','resnet_q25','resnet_q40',
'gap_q12','nas_q11','nas_q12','se_resnext_q26','se_resnext_q28',
'bni_q0','bni_q1']
glist += ['iv3_q12','iv3_u11','iv3_u12','irv2_u12',
'resnet_q26','resnet_q27',
'resnet_u25','resnet_u31','resnet_u56',
'resnet_u60','resnet_u61','resnet_u63',
'resnet_u65','resnet_u101',
'se_resnet_u50','se_resnext_u29',
'gap_u14','bni_u1','bni_u2']
# -
# list of unique model names
allm = [weights[i]['mod'].values for i in range(len(weights))]
mnames = list(set().union(*allm))
print(mnames)
print(len(mnames))
# +
nfold = 4
threshold = 0.52
sub_dir = './sub/'
print(fidx,midx,nfold,len(mnames))
# +
blist = ['blend_0459']
tlist = ['iv3_a4','se_resnext_d10','se_resnext_d11']
klist = ['resnet34_46','resnet50','resnet34']
print(glist)
print(blist)
print(tlist)
print(klist)
gpct = 97.5
gscale = 10.
# +
import numpy as np
import pandas as pd
import pickle
import os
from os import path
from random import randint
import matplotlib.pyplot as plt
plt.style.use('seaborn-white')
import seaborn as sns
sns.set_style("white")
from sklearn.model_selection import train_test_split
from keras.preprocessing.image import load_img
import pydensecrf.densecrf as dcrf
from pydensecrf.utils import unary_from_labels, create_pairwise_bilateral
from skimage.color import gray2rgb
from skimage.color import rgb2gray
from skimage.transform import resize
from scipy.special import logit, expit
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import jaccard_similarity_score, f1_score
import scipy.optimize as opt
import gc
import cv2
from tqdm import tqdm_notebook
from numpy import inf
# -
name_label_dict = {
0: "Nucleoplasm",
1: "Nuclear membrane",
2: "Nucleoli",
3: "Nucleoli fibrillar center",
4: "Nuclear speckles",
5: "Nuclear bodies",
6: "Endoplasmic reticulum",
7: "Golgi apparatus",
8: "Peroxisomes",
9: "Endosomes",
10: "Lysosomes",
11: "Intermediate filaments",
12: "Actin filaments",
13: "Focal adhesion sites",
14: "Microtubules",
15: "Microtubule ends",
16: "Cytokinetic bridge",
17: "Mitotic spindle",
18: "Microtubule organizing center",
19: "Centrosome",
20: "Lipid droplets",
21: "Plasma membrane",
22: "Cell junctions",
23: "Mitochondria",
24: "Aggresome",
25: "Cytosol",
26: "Cytoplasmic bodies",
27: "Rods & rings"
}
LABEL_MAP = name_label_dict
np.set_printoptions(precision=3, suppress=True, linewidth=100)
test_df = pd.read_csv("sample_submission.csv", index_col="Id")
print(test_df.head())
print(test_df.shape)
# + _uuid="b18c1f50cefd7504eae7e7b9605be3814c7cad6d"
# test_df["images"] = [np.array(load_img("images/{}.png".format(idx), \
# color_mode = "grayscale")) / 255. \
# # grayscale=True)) / 255. \
# for idx in tqdm_notebook(test_df.index)]
# print(test_df.shape)
# -
test_df.columns
# save a base copy for reuse below
test_df0 = test_df.copy()
# +
# build submission ensemble as weighted average on logit scale
test_df = test_df0.copy()
idx = test_df.index.values
firstm = True
sumw = 0.
for m in mnames:
firstf = True
navg = 0
if m in blist: suffix = '.npy'
elif m in klist: suffix = '.pkl'
else: suffix = '_mm.pkl'
for fold in range(nfold):
# print('')
# print(m+'_'+str(fold))
first = True
nr = 0
for rep in ['','a','b','c','d','e','f','g','h','i']:
if m in blist:
fname = sub_dir + m + rep + suffix
if fold > 0: fname = fname + '_' + str(fold)
else:
fname = sub_dir + m + rep + '_' + str(fold) + suffix
if os.path.exists(fname):
print()
print(fname)
if m in blist:
ptestb = np.load(fname)
idb = idb0
elif m in klist:
idb, ptestb = pickle.load(open(fname,'rb'))
# ptestb = logit(ptestb)
elif m in tlist:
idb, ptestba, ptestb, ptestbb = pickle.load(open(fname,'rb'))
ptestb = expit(ptestb)
elif m in glist:
idb, ptestb, ptestba = pickle.load(open(fname,'rb'))
ptestb = np.percentile(ptestb, gpct, axis=(2,3))
ptestb = 1./(1. + np.exp(-ptestb/gscale))
else:
idb, ptestb, ptestba = pickle.load(open(fname,'rb'))
ptestb = expit(ptestb)
# ptestb = np.clip(ptestb,-20.0,20.0)
# ptestb[ptestb==-inf] = -6.0
# ptestb[ptestb==inf] = 6.0
print(ptestb.min(),ptestb.mean(),ptestb.max())
if first:
ptestf = ptestb.copy()
idb0 = idb
first = False
else:
checki = [i0 != i1 for i0,i1 in zip(idb0,idb)]
si = np.array(checki).sum()
# print(si)
assert si == 0
print(np.corrcoef(np.array(ptestf).flatten(),
np.array(ptestb).flatten()))
ptestf += ptestb
nr += 1
if nr > 0:
ptestf /= nr
print(ptestf.shape,ptestf.min(),ptestf.mean(),ptestf.max())
if firstf:
id = idb
ptest = ptestf.copy()
firstf = False
else:
print(np.corrcoef(np.array(ptest).flatten(),
np.array(ptestf).flatten()))
ptest += ptestf
navg += 1
if navg != nfold: raise ValueError('Found %d folds for %s instead of %d' % (navg,m,nfold))
ptest /= navg
pmask = (ptest > threshold).astype(int)
print(m,navg,ptest.shape,ptest.min(),ptest.mean(),ptest.max(),pmask.mean())
td = pd.DataFrame({'id':id})
td[m] = [a for a in ptest]
td.set_index('id',inplace=True)
test_df = test_df.join(td)
# -
test_df.columns
print(idx[:5])
f = []
clist = list(test_df.columns[1:])
for c in clist:
ff = np.array(list(test_df[c])).flatten()
print(ff.shape,c)
f.append(ff)
# print(clist)
r = np.corrcoef(f)
print(r)
# +
d = pd.DataFrame(r, columns=mnames)
d['id'] = mnames
d.set_index('id',inplace=True)
del d.index.name
sns.clustermap(d)
# import scipy.cluster.hierarchy as hc
# from matplotlib import pyplot
# link = hc.linkage(d.values, method='centroid')
# o1 = hc.leaves_list(link)
# mat = d.iloc[o1,:]
# mat = mat.iloc[:, o1[::-1]]
# pyplot.imshow(mat)
# -
flat = np.transpose(np.array(f))
print(flat.shape)
fname = 'flat.csv'
fdf = pd.DataFrame(flat)
fdf.columns = clist
fdf.to_csv(fname)
print(fname)
# +
ens = np.zeros((test_df.shape[0],28))
# apply class-specific ensemble weights
for i in range(28):
f = []
clist = weights[i]['mod'].values
for c in clist:
f.append(np.array(list(test_df[c])))
m = np.array(f)
print(m.shape)
w = np.array(weights[i]['weight'].values)
print(i, clist, w)
ens[:,i] = np.average(m,axis=0,weights=w)[:,i]
print(ens.shape)
# -
print(ens[1])
# +
# fname = 'mthresh.pkl'
# if os.path.isfile(fname): mthresh = pickle.load(open(fname,'rb'))
# mthresh[mname]
# +
# desired class proportions from lb probing
desired = {
0 : 0.36239782,
1 : 0.043841336,
2 : 0.075268817,
3 : 0.059322034,
4 : 0.075268817,
5 : 0.075268817,
6 : 0.043841336,
7 : 0.075268817,
8 : 0.0018,
9 : 0.0014,
10 : 0.0009,
11 : 0.043841336,
12 : 0.043841336,
13 : 0.014198783,
14 : 0.043841336,
15 : 0.0007,
16 : 0.028806584,
17 : 0.014198783,
18 : 0.028806584,
19 : 0.059322034,
20 : 0.0056,
21 : 0.126126126,
22 : 0.028806584,
23 : 0.075268817,
24 : 0.0104,
25 : 0.222493888,
26 : 0.028806584,
27 : 0.0004
}
print(desired)
# -
# fixed global threshold
pred = (ens > threshold).astype(int)
print('fixed threshold',threshold)
for j in range(pred.shape[1]):
prop = np.mean(pred[:,j])
print(j,'%6.4f' % desired[j],'%6.4f' % prop,name_label_dict[j],)
# +
lb_prob = [
0.362397820,0.043841336,0.075268817,0.059322034,0.075268817,
0.075268817,0.043841336,0.075268817,0.010000000,0.010000000,
0.010000000,0.043841336,0.043841336,0.014198783,0.043841336,
0.010000000,0.028806584,0.014198783,0.028806584,0.059322034,
0.010000000,0.126126126,0.028806584,0.075268817,0.010000000,
0.222493880,0.028806584,0.010000000]
from scipy.special import expit
def sigmoid_np(x):
return 1.0/(1.0 + np.exp(-x))
def Count_soft(preds,th=0.5,d=50.0):
preds = sigmoid_np(d*(preds - th))
return preds.mean(axis=0)
def fit_test(x,y):
params = 0.5*np.ones(len(name_label_dict))
wd = 1e-5
error = lambda p: np.concatenate((Count_soft(x,p) - y,
wd*(p - 0.5)), axis=None)
p, success = opt.leastsq(error, params)
return p
pred_t = expit(ens)
th_t = fit_test(pred_t,lb_prob)
th_t[th_t<0.1] = 0.1
np.set_printoptions(precision=3, suppress=True, linewidth=100)
print('Thresholds: ',th_t)
print('Fractions: ',(pred_t > th_t).mean(axis=0))
print('Fractions (th = 0.5): ',(pred_t > 0.5).mean(axis=0))
pred = (pred_t > th_t).astype(int)
# -
ens.shape
p = pd.DataFrame(ens)
p.columns = ['p'+str(i) for i in range(ens.shape[1])]
subp = pd.DataFrame({'id':ids,'data':'sub'})
subp = pd.concat((subp,p),axis=1)
print(subp.shape)
fname = 'sub/sub' + midx + '_probs.csv'
subp.to_csv(fname, index=False)
print(fname)
# custom thresholds to match lb proportions
thresholds = np.linspace(0.9, 0.1, 100001)
pred = ens.copy()
th = []
for j in range(pred.shape[1]):
for t in thresholds:
pred[:,j] = (ens[:,j] > t).astype(int)
prop = np.mean(pred[:,j])
if prop >= desired[j]: break
th.append(t)
print(j,'%6.4f' % t,'%6.4f' % desired[j],'%6.4f' % prop,name_label_dict[j])
# +
# # linear adjustment of thresholds estimated in ens_oof.ipynb
# for j in range(pred.shape[1]):
# t = 0.094 + 0.827*th[j]
# pred[:,j] = (ens[:,j] > t).astype(int)
# prop = np.mean(pred[:,j])
# print(j,'%6.4f' % t,'%6.4f' % desired[j],'%6.4f' % prop,name_label_dict[j])
# -
np.set_printoptions(linewidth=100)
print(pred[:5].astype(int))
xps = np.sum(pred,axis=1)
print(xps.shape)
print(xps.min(),xps.mean(),xps.max())
print(sum(xps==0)/pred.shape[0])
subs = []
ids = []
for i,fid in enumerate(test_df.index.values):
subrow = ' '.join(list([str(i) for i in np.nonzero(pred[i])[0]]))
subs.append(subrow)
ids.append(fid)
if i % 1000 == 0: print(i,fid,subrow)
print({'ids':ids[:10], 'subs':subs[:10]})
print(len(ids), len(subs))
subm = pd.DataFrame.from_dict({'Id':ids, 'Predicted':subs}, orient='index').T
fname = 'sub/' + fidx + midx + '.csv'
subm.to_csv(fname, index=False)
print(fname)
from itertools import chain
from collections import Counter
max_idx = 27
subm['target_list'] = subm['Predicted'].map(lambda x: \
[int(a) if a != '' else -1 for a in str(x).split(' ')])
subm['target_vec'] = subm['target_list'].map(lambda ck: \
[i in ck for i in range(max_idx+1)])
all_labels = list(chain.from_iterable(subm['target_list'].values))
c_val = Counter(all_labels)
n_keys = c_val.keys()
max_idx = max(n_keys)
for k,v in name_label_dict.items():
print(k,v, 'count', c_val[k] if k in c_val else 0,
'prop', '%6.4f' % (c_val[k]/len(ids) if k in c_val else 0))
train_sum_vec = np.sum(np.stack(subm['target_vec'].values, 0), 0)
_ = plt.bar(n_keys, [train_sum_vec[k] for k in n_keys])
# +
from sklearn.metrics import f1_score
from sklearn.metrics import confusion_matrix
# computute confusion matrices between two submission files
def f1_confusion(csv0, csv1, num_classes=28):
c0 = pd.read_csv(csv0)
c1 = pd.read_csv(csv1)
assert c0.shape == c1.shape
s0 = [s if isinstance(s,str) else '' for s in c0.Predicted]
s1 = [s if isinstance(s,str) else '' for s in c1.Predicted]
p0 = [s.split() for s in s0]
p1 = [s.split() for s in s1]
y0 = np.zeros((c0.shape[0],num_classes)).astype(int)
y1 = np.zeros((c0.shape[0],num_classes)).astype(int)
# print(p0[:5])
for i in range(c0.shape[0]):
for j in p0[i]: y0[i,int(j)] = 1
for j in p1[i]: y1[i,int(j)] = 1
# print(y0[:5])
y0avg = np.average(y0,axis=0)
y1avg = np.average(y1,axis=0)
cm = [confusion_matrix(y0[:,i], y1[:,i]) for i in range(y0.shape[1])]
fm = [f1_score(y0[:,i], y1[:,i]) for i in range(y0.shape[1])]
for i in range(y0.shape[1]):
print(LABEL_MAP[i])
print(cm[i],' %4.2f' % fm[i],' %6.4f' % y0avg[i],' %6.4f' % y1avg[i],
' %6.4f' % (y0avg[i] - y1avg[i]))
print()
# print('y0avg')
# print(y0avg)
# print('y1avg')
# print(y1avg)
# print('y0avg - y1avg')
# print(y0avg-y1avg)
print('f1 macro')
print(np.mean(fm))
return f1_score(y0, y1, average='macro')
# compute f1 score between two submission files
def f1_sub(csv0, csv1, num_classes=28):
c0 = pd.read_csv(csv0)
c1 = pd.read_csv(csv1)
assert c0.shape == c1.shape
s0 = [s if isinstance(s,str) else '' for s in c0.Predicted]
s1 = [s if isinstance(s,str) else '' for s in c1.Predicted]
p0 = [s.split() for s in s0]
p1 = [s.split() for s in s1]
y0 = np.zeros((c0.shape[0],num_classes)).astype(int)
y1 = np.zeros((c0.shape[0],num_classes)).astype(int)
# print(p0[:5])
for i in range(c0.shape[0]):
for j in p0[i]: y0[i,int(j)] = 1
for j in p1[i]: y1[i,int(j)] = 1
# print(y0[:5])
return f1_score(y0, y1, average='macro')
# -
f1_sub(fname,'sub/ens45.csv')
f1_sub(fname,'sub/ens46.csv')
f1_sub(fname,'sub/ens47.csv')
f1_sub(fname,'sub/ens48.csv')
f1_sub(fname,'sub/ens49.csv')
f1_sub(fname,'sub/ens53.csv')
f1_sub(fname,'sub/ens53c.csv')
f1_sub(fname,'sub/ens53d.csv')
f1_sub(fname,'sub/ens55.csv')
f1_sub(fname,'sub/ens55d.csv')
f1_sub(fname,'sub/ens56.csv')
f1_sub(fname,'sub/ens56d.csv')
f1_sub(fname,'sub/ens58.csv')
f1_sub(fname,'sub/ens58d.csv')
f1_sub(fname,'sub/ens59.csv')
f1_sub(fname,'sub/ens59d.csv')
f1_sub(fname,'sub/ens60d.csv')
f1_sub(fname,'sub/ens61.csv')
f1_sub(fname,'sub/ens61d.csv')
f1_sub(fname,'sub/ens62.csv')
f1_sub(fname,'sub/preresnet0.csv')
f1_sub(fname,'sub/preresnet0d.csv')
f1_sub(fname,'sub/resnet11.csv')
f1_sub(fname,'sub/resnet12.csv')
f1_sub(fname,'sub/resnet13.csv')
f1_sub(fname,'sub/resnet15.csv')
f1_sub(fname,'sub/resnet15c.csv')
f1_sub(fname,'sub/resnet16.csv')
f1_sub(fname,'sub/resnet17.csv')
f1_sub(fname,'sub/resnet17.csv')
f1_sub(fname,'sub/se_resnext11.csv')
f1_sub(fname,'sub/se_resnext11d.csv')
print(fname)
# +
# f1_confusion(fname, 'sub/ens56.csv')
| wienerschnitzelgemeinschaft/src/Russ/ens_sub66.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Random Diffusion in 2D**
#
# **Authors:** <NAME>, <NAME> and <NAME>
#
# <i class="fa fa-home fa-2x"></i><a href="../index.ipynb" style="font-size: 20px"> Go back to index</a>
#
# **Source code:** https://github.com/osscar-org/quantum-mechanics/blob/master/notebook/statistical-mechanics/diffusion_2d.ipynb
#
# Diffusion is a common physical phenomenon. We simulated the diffusion of the
# two-dimensional systems in this notebook.
#
# <hr style="height:1px;border:none;color:#cccccc;background-color:#cccccc;" />
# ## **Goals**
#
# * Understand random diffusion process in two dimension
# * Learn how to calculate diffusion coefficient from standard radius.
# * Understand the connection between microscopic kinetics and macro observables.
# ## **Background theory**
#
# [More on the background theory.](./theory/theory_diffusion_2d.ipynb)
# ## **Tasks and exercises**
#
# 1. Click run button and play the random walk simulation. Compare the global behavior and the path of an individual point, what do you observe? (The individual path will be shown when the "show trajectory" box is checked.)
#
# <details>
# <summary style="color: red">Solution</summary>
# The individual path is random, which is physically a Brownian motion since
# all particles have constant kinetic energy. The global behavior, however,
# shows a uniform spherical pattern, where the diffusion radius is
# characterized by $r_{std}$.
# <br>
# </details>
#
# 2. What statistical distribution should the diffusion pattern obey? See if the red analytical curve drawn in the plot agrees with you.
#
# <details>
# <summary style="color: red">Solution</summary>
# Brownian motion will obey normal distribution. Note that in plot 2,
# $G(r)r$ is shown where G(r) is the normal probability density function
# and r denotes the distance to center which comes from the integration of
# probability density in sphereical coordinates i.e.
# $\int_0^{2\pi} d\theta \int_0^r rdr G(r) =1$
# </details>
#
#
# 3. How can the diffusion coefficient D be obtained from the global pattern observed?
#
# <details>
# <summary style="color: red">Solution</summary>
# Combining equation (1) and (5), we can see how the microscopic step
# size of random walk $l$ is controlling the macroscopic diffusion radius,
# $r_{std}$ and the speed of diffusion is characterized by diffusion
# coefficient D. Through the statistical study of the system (i.e.
# calculating mean and standard radius of the coordinates), we can
# obtain the standard radius or MSD without the knowledge on the
# microscopic kinetics (i.e. stepsize l). Thus, by doing a linear fitting
# on $r_{std}^2$ and time as shown in plot 3, D is given by:
# $$D = \frac{\text{slope}}{2 n D dt}$$ where dt is the time interval
# of one timestep.
# </details>
# <hr style="height:1px;border:none;color:#cccccc;background-color:#cccccc;" />
#
# ## Interactive visualization
# (be patient, it might take a few seconds to load)
# +
# %matplotlib widget
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as ipw
from scipy.stats import linregress
# -
box_xrange = (-10, 10)
box_yrange = (-10, 10)
starting_radius = 0.1
r = np.linspace(0,10,100)
# +
layout = ipw.Layout(width='auto', height='30px')
ndots_slider = ipw.IntSlider(value=1000, min=1, max=5000, step=100, description='Number of points', style= {'description_width': 'initial'}, layout=layout) # number of points
stepsize_slider = ipw.FloatSlider(value=0.05, min=0.01, max=0.1, step=0.01, description='Step size', continuous_update=False, readout=True, readout_format='.2f', style= {'description_width': 'initial'}, layout=layout) # max step size
frame_slider = ipw.IntSlider(value=0, min=0, max=ndots_slider.value, step=100, description='Time step', continuous_update=False, readout=True, disabled=True, style= {'description_width': 'initial'}, layout=layout) # step index indicator and slider
nsteps_slider = ipw.IntSlider(value=5000, min=100, max=10000, step=100, description='Number of steps', continuous_update=False, disabled=False, style= {'description_width': 'initial'}, layout=layout)
traj_chkbox = ipw.Checkbox(value=False,description='Show trajectory', disabled=False, indent=False)
map_chkbox = ipw.Checkbox(value=False,description='Show density map', disabled=False, indent=False)
run_btn = ipw.Button(description='Simulate')
run_btn.style.button_color = 'green'
play = ipw.Play(value=0, min=0, max=nsteps_slider.value, step=100, disabled=True, interval=500) # iterate frame with 500ms interval
# +
trajectory = [] # trajectory of all dots
r_std_sq = np.array([]) # square standard radius
slope = 0. # slope of linear fit in plot 3
intercept = 0. # intercept of the fit
def plot_dots_circle(ax):
show_traj = traj_chkbox.value
show_map = map_chkbox.value
frame_idx = frame_slider.value
r_l = np.sqrt(frame_idx) * stepsize_slider.value * np.sqrt(2) # analytical radius = sqrt(N) * stepsize * sqrt(2), a factor of sqrt(2) since we need mean stepsize along radius direction
r_std = np.sqrt(r_std_sq[frame_idx, 1]) # standard radius from simulation
frame_coords = trajectory[frame_idx]
ax.clear()
ax.set_xlim(box_xrange)
ax.set_ylim(box_yrange)
ticks_ax1 = [-10., -5., 0., 5., 10]
ax.xaxis.set_ticks(ticks_ax1)
ax.yaxis.set_ticks(ticks_ax1)
ax.set_aspect(1.)
ax.set_xlabel('x')
ax.set_ylabel('y')
# draw dots
ax.plot(frame_coords[:,0], frame_coords[:,1], '.', alpha=0.1, zorder=11)
# draw circle
circle_std = plt.Circle((0, 0), r_std, color='green', linewidth=2, fill=False,zorder=12, label='$r_{std}$')
circle_l = plt.Circle((0, 0), r_l, color='red', fill=False, linestyle='dashed',zorder=12, label='$r_{l}$')
ax.add_patch(circle_std)
ax.add_patch(circle_l)
# draw trajectory of first dots
if show_traj:
ax.plot(trajectory[:frame_idx:100,0,0], trajectory[:frame_idx:100,0,1], linewidth=2, color='purple', zorder=13, label='trajectory')
# analytical Gaussian density map for the diffusion plot as a comparison for the actual simulation pattern
if show_map:
x = np.linspace(-10, 10, 30)
y = np.linspace(-10, 10, 30)
N = frame_idx
l = stepsize_slider.value
gx = gaussian_1d(x, N, l)
gy = gaussian_1d(y, N, l)
H = np.ma.outerproduct(gx, gy).data
ax.imshow(H, origin='lower', interpolation='none', extent=[box_xrange[0], box_xrange[1], box_yrange[0], box_yrange[1]],aspect='equal', alpha=1, cmap='Reds')
ax.legend(loc='lower right', bbox_to_anchor=(1, 1.05))
def gaussian_1d(x, N, l):
"""A helper function for plot 2.
x: range
N: number of steps
l: stepsize
Return Gaussian/ Normal distributino on 1D
"""
if N == 0:
return np.zeros(len(x)) # for simplicity of visualization, zeros is returned instead of a Dirac distribution
var = N * l**2
return (1 / np.sqrt(2 * np.pi * var)) * np.exp(-x**2/ (2 * var))
def plot_1d_hist(ax):
""" draw plot 2
Histogram is obtained consider only x direction, which should fits under
1D Gaussian distribution. Note that histogram may deviates from Gaussian
after prolonged time due to PBC.
"""
frame_idx = frame_slider.value
N = ndots_slider.value
stepsize = stepsize_slider.value
x_coords = trajectory[frame_idx][:,0]
nbins = 30
bin_width = (box_xrange[1] - box_xrange[0]) / nbins
hist, bins= np.histogram(x_coords, bins=30, range=box_xrange, density=True)
# hist = hist / (bin_width * N) # normalized count by count/ (N * width) to get f(r)
h_offset = 0.5 * bin_width # horizontal offset for histogram plot so the first column starts at 0
r = np.linspace(box_xrange[0], box_xrange[1], 100)
gr = gaussian_1d(r, frame_idx, stepsize)
ax.clear()
ax.set_xlim(-10, 10)
ax.set_ylim(0, 0.6)
ax.set_xlabel("x")
ax.set_ylabel("frequency")
ax.bar(bins[:-1]+h_offset, hist, ec='k', width=bin_width)
ax.plot(r, gr, 'r--',label='Gaussian distribution')
ax.legend(loc='lower right', bbox_to_anchor=(1, 1.05))
def plot_radii(ax):
"""draw Plot 3
"""
frame_idx = frame_slider.value
nsteps = nsteps_slider.value
ax.clear()
# plot r_std^2 (MSD) vs t
interval = 500
ax.plot(r_std_sq[::interval,0], r_std_sq[::interval,1], 'o') # plot every 100 steps
ax.plot(frame_idx, r_std_sq[frame_idx, 1], 'o', color='green', label='current step')
# plot linear fitting line
lx = np.linspace(0,nsteps,10)
ly = lx * slope + intercept
ax.plot(lx, ly, 'r--', lw=2, label='y = {:.2e} x + {:.2f}'.format(slope, intercept))
ax.set_xlabel('time step')
ax.set_ylabel('$r_{std}^2$')
ax.legend(loc='lower right', bbox_to_anchor=(1, 1.05))
def plot_frame(change):
''' plot current frame for all axis'''
# check if trajectory is already stored
if len(trajectory) == 0:
return
# plot 1
plot_dots_circle(ax1)
# plot 2
plot_1d_hist(ax2) # in x direction
# plot_circle_hist(ax2) # in spherical coords, along radius
# plot 3
plot_radii(ax3)
def run(change):
'''Main function for simulation
- generate initial particle coords
- run diffusion simulation and store trajectory of all dots in trajectory
- do linear fitting on r_std and t for plot 3
'''
global trajectory, r_std_sq, slope, intercept
run_btn.style.button_color = 'red'
N = ndots_slider.value
# Initial coords with a random radial distribution generated by creating normal
# random coords and take first N points in the initial circle. Arguably, we can
# start with all particles at origin but that is less realistic. A demo
# is attached as commented out code at the end of the notebook.
stepsize = stepsize_slider.value # mean stepsize
coords = (np.random.random((10*N, 2)) - 0.5)*2 * stepsize
coords = coords[(coords**2).sum(axis=1) < starting_radius**2][:N]
assert len(coords) == N # check if enough points are in the circle
# run simulation and store trajectory
trajectory = [coords]
num_steps = nsteps_slider.value
for i in range(num_steps):
# two different ways of displacement with same distribution
# random_displacement = (np.random.random((N, 2)) - 0.5) * 2 * stepsize # continuous
random_displacement = (np.random.choice([-1,1],(N, 2))) * stepsize # discrete
new_positions = trajectory[-1] + random_displacement
# Some points might have gone beyond the box.
# I could either reflect them back as a hard wall, or just use PBC. For simplicity, I use PBC
new_positions[:,0] = (new_positions[:,0] - box_xrange[0]) % (box_xrange[1] - box_xrange[0]) + box_xrange[0]
new_positions[:,1] = (new_positions[:,1] - box_yrange[0]) % (box_yrange[1] - box_yrange[0]) + box_yrange[0]
trajectory.append(new_positions)
trajectory = np.array(trajectory)
# calculate r_std by sqrt(mean**2 + std**2) and do the fitting
radii = np.sqrt((trajectory**2).sum(axis=2))
r_std_sq = radii.mean(axis=1)**2 + radii.std(axis=1)**2
r_std_sq = np.c_[np.arange(len(r_std_sq)), r_std_sq]
res = linregress(r_std_sq)
slope = res.slope
intercept = res.intercept
# enable play and frame slider after the simulation run
play.disabled = False
frame_slider.disabled = False
plot_frame('init')
run_btn.style.button_color = 'green'
def stop(change):
''' disable play widget and reset frame slider'''
global dots_art, traj_art, circle
play.disabled = True
frame_slider.value = 0
# reset all the axes
for ax in [ax1, ax2, ax3]:
ax.clear()
initialize_plot()
def initialize_plot():
"""Initialized plot to specify ranges, ticks or labels on x, y axis
Called when first run the notebook or the simulation parameters change."""
global ax1, ax2, ax3
ax = ax1
ax.set_xlim(box_xrange)
ax.set_ylim(box_yrange)
ticks_ax1 = [-10., -5., 0., 5., 10]
ax.xaxis.set_ticks(ticks_ax1)
ax.yaxis.set_ticks(ticks_ax1)
ax.set_aspect(1.)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax = ax2
ax.set_xlim(-10, 10)
ax.set_ylim(0, 0.6)
ax.set_xlabel("x")
ax.set_ylabel("frequency")
ax = ax3
ax.set_xlabel('time step')
ax.set_ylabel('$r_{std}^2$')
# link widgets
ipw.jslink((play, 'value'), (frame_slider, 'value'))
ipw.jslink((nsteps_slider, 'value'), (frame_slider,'max'))
frame_slider.observe(plot_frame, names='value', type='change')
# click run for simmulation and collect trajectory
run_btn.on_click(run)
# change simulation parameters will disable play and frame slider until finish run
ndots_slider.observe(stop, names='value', type='change')
stepsize_slider.observe(stop, names='value', type='change')
nsteps_slider.observe(stop, names='value', type='change')
# group widgets
play_wdgt = ipw.HBox([run_btn, play])
ctrl_widgets = ipw.VBox([ndots_slider, stepsize_slider, nsteps_slider, play_wdgt, traj_chkbox, map_chkbox, frame_slider])
# frame_idx = 0
# use Output to wrap the plot for better layout
plotup_out = ipw.Output()
with plotup_out:
fig_up, (ax1,ax2) = plt.subplots(1,2,constrained_layout=True, figsize=(6,3))
plt.show()
plotdwn_out = ipw.Output()
with plotdwn_out:
fig_dwn, ax3 = plt.subplots(constrained_layout=True, figsize=(3,2))
plt.show()
initialize_plot()
display(ipw.VBox([ipw.HBox([plotup_out]), ipw.HBox([ctrl_widgets, plotdwn_out])]))
# -
# <hr style="height:1px;border:none;color:#cccccc;background-color:#cccccc;" />
#
# ## **Legend**
#
# ### Interactive Figures
#
# Plots are named plot 1, 2 and 3 from top to bottom and left to right.
# - Plot 1 is the main plot showing directly the Brownian diffusion of free particles in 2D. The green circle has radius of standard radius, $r_{std}$ which is given by $\sqrt{\sigma^2 + \mu^2}$ and the radius of red circle is $r_l$ which is given by $\sqrt{(N)} l$. Optionally, trajectory and density map can be displayed. The former corresponds to one arbitrary particles for every 100 steps and the latter is the analytical Gaussian density in 2D given by the outer product of two 1D distribution vectors.
# - Plot 2 shows the 1D histogram of the particles along x direction. The analytical Gaussian distribution is obtained by substitute equation (3), (4) into (2).
# - Plot 3 shows the liner relationship of $r_{std}^2$ and time step, t. The data points for every 500 steps are shown and the current step is labeled in green. The red dashed line is the result of linear regression on the data.
#
#
# ### Controls
#
# The panel at lower left controls all 3 plots. Top three sliders set respectively the number of points, step size and the number of points for the diffusion simulation. Choose the value as you wish and then click "Run" to run the simulation in the background. To visualize the result, use the play widget next to it. Note all 3 plots are synced with the time step.
# The bottom three widgets control the visualization. Click the checkbox to show corresponding elements in plot 1. The time step slider indicates and also controls the current step corresponding to plots.
| notebook/statistical-mechanics/diffusion_2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nbase] *
# language: python
# name: conda-env-nbase-py
# ---
import numpy as np
import pandas as pd
import os
import pickle
raw_data_dir = "../data/ft_data/"
new_dir = './BeFree/n_out/'
# !cp ../data/ft_data/labels_n.txt ./BeFree/n_out/labels.txt
FT_MODE = True
# +
# raw_data_dir = "../data/abs_data/2nd_ann/"
# new_dir = './BeFree/abs_out/'
# # !cp ../data/abs_data/2nd_ann/labels.txt ./BeFree/abs_out/labels.txt
# FT_MODE = False
# -
# +
# generate gene/disease annotation for BeFree
# +
sentence_path = os.path.join(raw_data_dir, "sentences.txt")
ner_path = os.path.join(raw_data_dir, "anns.txt")
text_path = os.path.join(raw_data_dir, "docs.txt")
new_gene_file = open(new_dir+"genes_FINAL.befree", 'w')
new_disease_file = open(new_dir+"diseases_FINAL.befree", 'w')
text_file = open(text_path, "r")
sentence_file = open(sentence_path, "r")
ner_file = open(ner_path, "r")
tar_pmid_lst = None
def get_sent_offset(sentences, text):
sent_offset = []
for i, sentence in enumerate(sentences):
if i == 0:
offset = 0
else:
offset = last_offset + len(last_sentence)
if sentence != text[offset: offset+len(sentence)]:
offset = offset + text[offset:].find(sentence)
sent_offset.append(offset)
last_offset = offset
last_sentence = sentence
return sent_offset
def Sent_no(sent_offset, offset):
return np.digitize(offset, sent_offset).tolist() - 1
def offset_in_sent(sentences_offset, sent_no, begin, end):
begin_offset = begin - sentences_offset[sent_no]
end_offset = end - sentences_offset[sent_no]
return str(begin_offset) + "#" + str(end_offset)
def get_sentence(sentences, row):
return sentences[sent_no]
_idx = 1
while (1):
line = text_file.readline()
if line == '':
break
pmid = line.strip()
title = text_file.readline()
title = title[:-1] if len(title) > 0 else title
abstract = text_file.readline()
abstract= abstract[:-1] if len(abstract) > 0 else abstract
para = text_file.readline()
para = para[:-1] if len(para) > 0 else para
#fix space between title, abstract, para
text = title + " " + abstract + " " + para
text_file.readline()
#text = abstract title + text
sentences = []
line = sentence_file.readline()
if line == "\n":
sentence_file.readline()
while (1):
line = sentence_file.readline()
if line == "\n":
break
sentence = line.strip()
sentences.append(sentence)
sent_offset = get_sent_offset(sentences, text)
anns = []
while (1):
line = ner_file.readline()
if line == "\n":
break
ann = line.strip().split("\t")
if ';' in ann[5]:
ann[5] = ann[5].split(";")[1]
# for ft only
if FT_MODE:
ann = ann[:-1]
# print(ann)
ann[1] = int(ann[1])
ann[2] = int(ann[2])
ann[-1], ann[-2] = ann[-2], ann[-1]
if ann[4] == 'None' or len(ann[4]) < 1:
continue
anns.append(ann[:])
# print(ann)
begin, end, mention, _id, _type = ann[1], ann[2], ann[3], ann[4], ann[5]
if tar_pmid_lst and (pmid not in tar_pmid_lst):
continue
# print(_type)
sent_no = Sent_no(sent_offset, begin)
off_tar = offset_in_sent(sent_offset, sent_no, begin, end)
# print(sent_no, off_tar)
_pmid = pmid
_year, _journal_name, _journal_ISSN = '#', '#', '#'
_section, _section_nb, _sent_no = 'TITLE', 0, 00.8
if sent_no > 0:
_section, _section_nb, _sent_no = 'ALL_TEXT', 1, sent_no
_entity_id = _id
_entity_type = 'LONGTERM|DICTIONARY'
_entity_norm = mention.lower()
_mention = mention
_offset = off_tar
_entity_parent = 'nan'
_sentence = sentences[sent_no]
# 'pubmed', 'year','journal_name',
# 'journal_ISSN', 'section', 'section_nb', 'sent_no', 'entity_id', 'entity_type', 'entity_norm', 'mention', 'offset',
# 'entity_parent', 'sentence'
# print('%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t' % (\
# _pmid, _year, _journal_name, _journal_ISSN, \
# _section, _section_nb, _sent_no, \
# _entity_id, _entity_type, _entity_norm, \
# _mention, _offset, _entity_parent,\
# _sentence))
# print(_type)
if _type == 'Disease':
new_disease_file.write('%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\n' % (\
_pmid, _year, _journal_name, _journal_ISSN, \
_section, _section_nb, _sent_no, \
_entity_id, _entity_type, _entity_norm, \
_mention, _offset, _entity_parent,\
_sentence))
else:
new_gene_file.write('%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\t%s\n' % (\
_pmid, _year, _journal_name, _journal_ISSN, \
_section, _section_nb, _sent_no, \
_entity_id, _entity_type, _entity_norm, \
_mention, _offset, _entity_parent,\
_sentence))
if tar_pmid_lst and (pmid not in tar_pmid_lst):
# if pmid in tar_pmid_lst:
print(_idx, pmid)
_idx += 1
# print(abstract)
# print(para)
# print(sentences)
# print(sent_offset)
# print('\n'.join([str(i) for i in anns]))
# break
print('end')
new_gene_file.close()
new_disease_file.close()
text_file.close()
sentence_file.close()
ner_file.close()
# -
# +
# run BeFree
# -
| benchmark/Generate_BeFree_Input.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
print(tf.__version__)
# # Data Pipeline
# ## Coding tutorials
# #### [1. Keras datasets](#coding_tutorial_1)
# #### [2. Dataset generators](#coding_tutorial_2)
# #### [3. Keras image data augmentation](#coding_tutorial_3)
# #### [4. The Dataset class](#coding_tutorial_4)
# #### [5. Training with Datasets](#coding_tutorial_5)
# ***
# <a id="coding_tutorial_1"></a>
# ## Keras datasets
#
# For a list of Keras datasets and documentation on recommended usage, see [this link](https://keras.io/datasets/).
import numpy as np
import matplotlib.pyplot as plt
# #### Load the CIFAR-100 Dataset
from tensorflow.keras.datasets import cifar100
# +
# Load the CIFAR-100 dataset
(x_train, y_train), (x_test, y_test) = cifar100.load_data(label_mode='fine')
# +
# Confirm that reloading the dataset does not require a download
(x_train, y_train), (x_test, y_test) = cifar100.load_data(label_mode='fine')
# -
# #### Examine the Dataset
# +
# Examine the shape of the data.
print(x_train.shape)
print(y_train.shape)
# +
# Examine one of the images and its corresponding label
plt.imshow(x_train[500])
print(y_train[500])
# +
# Load the list of labels from a JSON file
import json
with open('data/cifar100_fine_labels.json', 'r') as fine_labels:
cifar100_fine_labels = json.load(fine_labels)
# -
# The list of labels for the CIFAR-100 dataset are available [here](https://www.cs.toronto.edu/~kriz/cifar.html).
# +
# Print a few of the labels
cifar100_fine_labels[0:10]
# +
# Print the corresponding label for the example above
cifar100_fine_labels[41]
# -
# #### Load the data using different label modes
# +
# Display a few examples from category 87 (index 86) and the list of labels
examples = x_train[(y_train.T == 86)[0]][:3]
fig, ax = plt.subplots(1,3)
ax[0].imshow(examples[0])
ax[1].imshow(examples[1])
ax[2].imshow(examples[2])
# +
# Reload the data using the 'coarse' label mode
(x_train, y_train), (x_test, y_test) = cifar100.load_data(label_mode='coarse')
# +
# Display three images from the dataset with the label 6 (index 5)
examples = x_train[(y_train.T == 5)[0]][:3]
fig, ax = plt.subplots(1,3)
ax[0].imshow(examples[0])
ax[1].imshow(examples[1])
ax[2].imshow(examples[2])
# +
# Load the list of coarse labels from a JSON file
with open('data/cifar100_coarse_labels.json', 'r') as coarse_labels:
cifar100_coarse_labels = json.load(coarse_labels)
# +
# Print a few of the labels
cifar100_coarse_labels[0:10]
# +
# Print the corresponding label for the example above
print(cifar100_fine_labels[86])
print(cifar100_coarse_labels[5])
# -
# #### Load the IMDB Dataset
from tensorflow.keras.datasets import imdb
# +
# Load the IMDB dataset
(x_train, y_train), (x_test, y_test) = imdb.load_data()
# +
# Print an example from the training dataset, along with its corresponding label
print(x_train[0])
print(y_train[0])
# +
# Get the lengths of the input sequences
sequence_lengths = [len(seq) for seq in x_train]
# +
# Determine the maximum and minimum sequence length
print(np.max(sequence_lengths))
print(np.min(sequence_lengths))
# -
# #### Using Keyword Arguments
# +
# Load the data ignoring the 50 most frequent words, use oov_char=2 (this is the default)
(x_train, y_train), (x_test, y_test) = imdb.load_data(skip_top=50, oov_char=2)
# +
# Get the lengths of the input sequences
sequence_lengths = [len(seq) for seq in x_train]
# +
# Determine the maximum and minimum sequence length
print(np.max(sequence_lengths))
print(np.min(sequence_lengths))
# +
# Define functions for filtering the sequences
def remove_oov_char(element):
''' Filter function for removing the oov_char. '''
return [word for word in element if word!=2]
def filter_list(lst):
''' Run remove_oov_char on elements in a list. '''
return [remove_oov_char(element) for element in lst]
# +
# Remove the oov_char from the sequences using the filter_list function
x_train = filter_list(x_train)
# +
# Get the lengths of the input sequences
sequence_lengths = [len(seq) for seq in x_train]
# +
# Determine the maximum and minimum sequence length
print(np.max(sequence_lengths))
print(np.min(sequence_lengths))
# -
# ***
# <a id="coding_tutorial_2"></a>
# ## Dataset generators
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# #### Load the UCI Fertility Dataset
#
# We will be using a dataset available at https://archive.ics.uci.edu/ml/datasets/Fertility from UC Irvine.
# +
# Load the fertility dataset
headers = ['Season', 'Age', 'Diseases', 'Trauma', 'Surgery', 'Fever', 'Alcohol', 'Smoking', 'Sitting', 'Output']
fertility = pd.read_csv('data/fertility_diagnosis.txt', delimiter=',', header=None, names=headers)
# +
# Print the shape of the DataFrame
print(fertility.shape)
# +
# Show the head of the DataFrame
fertility.head()
# -
# #### Process the data
# +
# Map the 'Output' feature from 'N' to 0 and from 'O' to 1
fertility['Output'] = fertility['Output'].map(lambda x : 0.0 if x=='N' else 1.0)
# +
# Show the head of the DataFrame
fertility.head()
# +
# Convert the DataFrame so that the features are mapped to floats
fertility = fertility.astype('float32')
# +
# Shuffle the DataFrame
fertility = fertility.sample(frac=1).reset_index(drop=True)
# +
# Show the head of the DataFrame
fertility.head()
# +
# Convert the field Season to a one-hot encoded vector
fertility = pd.get_dummies(fertility, prefix='Season', columns=['Season'])
# +
# Show the head of the DataFrame
fertility.head()
# +
# Move the Output column such that it is the last column in the DataFrame
fertility.columns = [col for col in fertility.columns if col != 'Output'] + ['Output']
# +
# Show the head of the DataFrame
fertility.head()
# +
# Convert the DataFrame to a numpy array.
fertility = fertility.to_numpy()
# -
# #### Split the Data
# +
# Split the dataset into training and validation set
training = fertility[0:70]
validation = fertility[70:100]
# +
# Verify the shape of the training data
print(training.shape)
# +
# Separate the features and labels for the validation and training data
training_features = training[:,0:-1]
training_labels = training[:,-1]
validation_features = validation[:,0:-1]
validation_labels = validation[:,-1]
# -
# #### Create the Generator
# +
# Create a function that returns a generator producing inputs and labels
def get_generator(features, labels, batch_size=1):
for n in range(int(len(features)/batch_size)):
yield (features[n*batch_size: (n+1)*batch_size], labels[n*batch_size: (n+1)*batch_size])
# +
# Apply the function to our training features and labels with a batch size of 10
train_generator = get_generator(training_features, training_labels, batch_size=10)
# +
# Test the generator using the next() function
next(train_generator)
# -
# #### Build the model
# +
# Create a model using Keras with 3 layers
from tensorflow.keras import Model
from tensorflow.keras.layers import Dense, Input, BatchNormalization
input_shape = (12,)
output_shape = (1,)
model_input = Input(input_shape)
batch_1 = BatchNormalization(momentum=0.8)(model_input)
dense_1 = Dense(100, activation='relu')(batch_1)
batch_2 = BatchNormalization(momentum=0.8)(dense_1)
output = Dense(1, activation='sigmoid')(batch_2)
model = Model([model_input], output)
# +
# Display the model summary to show the resultant structure
model.summary()
# -
# #### Compile the model
# +
# Create the optimizer object
optimizer = tf.keras.optimizers.Adam(learning_rate=1e-2)
# +
# Compile the model with loss function and metric
model.compile(optimizer=optimizer, loss='binary_crossentropy', metrics=['accuracy'])
# -
# #### Train and evaluate the model using the generator
# +
# Calculate the number of training steps per epoch for the given batch size.
batch_size = 5
train_steps = len(training) // batch_size
# +
# Set the epochs to 3
epochs = 3
# +
# Train the model
for epoch in range(epochs):
train_generator = get_generator(training_features, training_labels, batch_size=batch_size)
validation_generator = get_generator(validation_features, validation_labels, batch_size=30)
model.fit_generator(train_generator, steps_per_epoch=train_steps, validation_data=validation_generator,validation_steps=1)
# +
# Try to run the fit_generator function once more; observe what happens
model.fit_generator(train_generator, steps_per_epoch=train_steps)
# -
# #### Make an infinitely looping generator
# +
# Create a function that returns an infinitely looping generator
def get_generator_cyclic(features, labels, batch_size=1):
while True:
for n in range(int(len(features)/batch_size)):
yield (features[n*batch_size: (n+1)*batch_size], labels[n*batch_size: (n+1)*batch_size])
permuted = np.random.permutation(len(features))
features = features[permuted]
labels = labels[permuted]
# +
# Create a generator using this function.
train_generator_cyclic = get_generator_cyclic(training_features, training_labels, batch_size=batch_size)
# +
# Assert that the new cyclic generator does not raise a StopIteration
for i in range(2*train_steps):
next(train_generator_cyclic)
# +
# Generate a cyclic validation generator
validation_generator_cyclic = get_generator_cyclic(validation_features, validation_labels, batch_size=batch_size)
# +
# Train the model
model.fit_generator(train_generator_cyclic, steps_per_epoch=train_steps,
validation_data=validation_generator_cyclic, validation_steps=1, epochs=3)
# -
# #### Evaluate the model and get predictions
# +
# Let's obtain a validation data generator.
validation_generator = get_generator(validation_features, validation_labels, batch_size=30)
# +
# Get predictions on the validation data
predictions = model.predict_generator(validation_generator, steps=1)
print(np.round(predictions.T[0]))
# +
# Print the corresponding validation labels
print(validation_labels)
# +
# Obtain a validation data generator
validation_generator = get_generator(validation_features, validation_labels, batch_size=30)
# +
# Evaluate the model
print(model.evaluate(validation_generator))
# -
# ***
# <a id="coding_tutorial_3"></a>
# ## Keras image data augmentation
import matplotlib.pyplot as plt
import numpy as np
# #### Load the CIFAR-10 Dataset
from tensorflow.keras.datasets import cifar10
# +
# Load the CIFAR-10 dataset
(training_features, training_labels), (test_features, test_labels) = cifar10.load_data()
# +
# Convert the labels to a one-hot encoding
num_classes = 10
training_labels = tf.keras.utils.to_categorical(training_labels, num_classes)
test_labels = tf.keras.utils.to_categorical(test_labels, num_classes)
# -
# #### Create a generator function
# +
# Create a function that returns a data generator
def get_generator(features, labels, batch_size=1):
for n in range(int(len(features)/batch_size)):
yield (features[n*batch_size:(n+1)*batch_size], labels[n*batch_size:(n+1)*batch_size])
# +
# Use the function we created to get a training data generator with a batch size of 1
training_generator = get_generator(training_features, training_labels)
# +
# Assess the shape of the items generated by training_generator using the `next` function to yield an item.
image, label = next(training_generator)
print(image.shape)
print(label.shape)
# +
# Test the training generator by obtaining an image using the `next` generator function, and then using imshow to plot it.
# Print the corresponding label
from matplotlib.pyplot import imshow
image, label = next(training_generator)
image_unbatched = image[0,:,:,:]
imshow(image_unbatched)
print(label)
# +
# Reset the generator by re-running the `get_generator` function.
train_generator = get_generator(training_features, training_labels)
# -
# #### Create a data augmention generator
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# +
# Create a function to convert an image to monochrome
def monochrome(x):
def func_bw(a):
average_colour = np.mean(a)
return [average_colour, average_colour, average_colour]
x = np.apply_along_axis(func_bw, -1, x)
return x
# +
# Create an ImageDataGenerator object
image_generator = ImageDataGenerator(preprocessing_function=monochrome,
rotation_range=100,
rescale=(1/255.))
image_generator.fit(training_features)
# -
# Check [the documentation](https://keras.io/preprocessing/image/) for the full list of image data augmentation options.
# +
# Create an iterable generator using the `flow` function
image_generator_iterable = image_generator.flow(training_features, training_labels, batch_size=1, shuffle=True)
# +
# Show a sample from the generator and compare with the original
image, label = next(image_generator_iterable)
image_orig, label_orig = next(train_generator)
figs, axes = plt.subplots(1,2)
axes[0].imshow(image[0,:,:,:])
axes[0].set_title('Transformed')
axes[1].imshow(image_orig[0,:,:,:])
axes[1].set_title('Original')
plt.show()
# -
# #### Flow from directory
# +
# Inspect the directory structure
train_path = 'data/flowers-recognition-split/train'
val_path = 'data/flowers-recognition-split/val'
# +
# Create an ImageDataGenerator object
datagenerator = ImageDataGenerator(rescale=(1/255.0))
# -
classes = ['daisy', 'dandelion', 'rose', 'sunflower', 'tulip']
# +
# Create a training data generator
train_generator = datagenerator.flow_from_directory(train_path, batch_size=64, classes=classes, target_size=(16, 16))
# +
# Create a validation data generator
val_generator = datagenerator.flow_from_directory(val_path, batch_size=64, classes=classes, target_size=(16, 16))
# +
# Get and display an image and label from the training generator
x = next(train_generator)
imshow(x[0][4])
print(x[1][4])
# +
# Reset the training generator
train_generator = datagenerator.flow_from_directory(train_path, batch_size=64, classes=classes, target_size=(16, 16))
# -
# #### Create a model to train
# +
# Build a CNN model
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Input, Flatten, Dense
model = tf.keras.Sequential()
model.add(Input((16,16,3)))
model.add(Conv2D(8, (8, 8), padding='same', activation='relu'))
model.add(MaxPooling2D((4,4)))
model.add(Conv2D(8, (8, 8), padding='same', activation='relu'))
model.add(MaxPooling2D((2,2)))
model.add(Conv2D(4, (4, 4), padding='same', activation='relu'))
model.add(Flatten())
model.add(Dense(16, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(5, activation='softmax'))
# +
# Create an optimizer object
optimizer = tf.keras.optimizers.Adam(1e-3)
# +
# Compile the model
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['accuracy'])
# +
# Print the model summary
model.summary()
# -
# #### Train the model
# +
# Calculate the training generator and test generator steps per epoch
train_steps_per_epoch = train_generator.n // train_generator.batch_size
val_steps = val_generator.n // val_generator.batch_size
print(train_steps_per_epoch, val_steps)
# +
# Fit the model
model.fit_generator(train_generator, steps_per_epoch=train_steps_per_epoch, epochs=5)
# -
# #### Evaluate the model
# +
# Evaluate the model
model.evaluate_generator(val_generator, steps=val_steps)
# -
# #### Predict using the generator
# +
# Predict labels with the model
predictloss = model.predictloss
# -
# ***
# <a id="coding_tutorial_4"></a>
# ## The Dataset Class
import matplotlib.pyplot as plt
import numpy as np
import os
# #### Create a simple dataset
x = np.zeros((100,10,2,2))
# +
# Create a dataset from the tensor x
dataset1 = tf.data.Dataset.from_tensor_slices(x)
# +
# Inspect the Dataset object
print(dataset1)
print(dataset1.element_spec)
# -
x2 = [np.zeros((10,2,2)), np.zeros((5,2,2))]
# +
# Try creating a dataset from the tensor x2
dataset2 = tf.data.Dataset.from_tensor_slices(x2)
# -
x2 = [np.zeros((10,1)), np.zeros((10,1)), np.zeros((10,1))]
# +
# Create another dataset from the new x2 and inspect the Dataset object
dataset2 = tf.data.Dataset.from_tensor_slices(x2)
# +
# Print the element_spec
print(dataset2.element_spec)
# -
# #### Create a zipped dataset
# +
# Combine the two datasets into one larger dataset
dataset_zipped = tf.data.Dataset.zip((dataset1, dataset2))
# +
# Print the element_spec
print(dataset_zipped.element_spec)
# +
# Define a function to find the number of batches in a dataset
def get_batches(dataset):
iter_dataset = iter(dataset)
i = 0
try:
while next(iter_dataset):
i = i+1
except:
return i
# +
# Find the number of batches in the zipped Dataset
get_batches(dataset_zipped)
# -
# #### Create a dataset from numpy arrays
# +
# Load the MNIST dataset
(train_features, train_labels), (test_features, test_labels) = tf.keras.datasets.mnist.load_data()
print(type(train_features), type(train_labels))
# +
# Create a Dataset from the MNIST data
mnist_dataset = tf.data.Dataset.from_tensor_slices((train_features, train_labels))
# +
# Inspect the Dataset object
print(mnist_dataset.element_spec)
# +
# Inspect the length of an element using the take method
element = next(iter(mnist_dataset.take(1)))
print(len(element))
# +
# Examine the shapes of the data
print(element[0].shape)
print(element[1].shape)
# -
# #### Create a dataset from text data
# +
# Print the list of text files
text_files = sorted([f.path for f in os.scandir('data/shakespeare')])
print(text_files)
# +
# Load the first file using python and print the first 5 lines.
with open(text_files[0], 'r') as fil:
contents = [fil.readline() for i in range(5)]
for line in contents:
print(line)
# +
# Load the lines from the files into a dataset using TextLineDataset
shakespeare_dataset = tf.data.TextLineDataset(text_files)
# +
# Use the take method to get and print the first 5 lines of the dataset
first_5_lines_dataset = iter(shakespeare_dataset.take(5))
lines = [line for line in first_5_lines_dataset]
for line in lines:
print(line)
# +
# Compute the number of lines in the first file
lines = []
with open(text_files[0], 'r') as fil:
line = fil.readline()
while line:
lines.append(line)
line = fil.readline()
print(len(lines))
# +
# Compute the number of lines in the shakespeare dataset we created
shakespeare_dataset_iterator = iter(shakespeare_dataset)
lines = [line for line in shakespeare_dataset_iterator]
print(len(lines))
# -
# #### Interleave lines from the text data files
# +
# Create a dataset of the text file strings
text_files_dataset = tf.data.Dataset.from_tensor_slices(text_files)
files = [file for file in text_files_dataset]
for file in files:
print(file)
# +
# Interleave the lines from the text files
interleaved_shakespeare_dataset = text_files_dataset.interleave(tf.data.TextLineDataset, cycle_length=9)
print(interleaved_shakespeare_dataset.element_spec)
# +
# Print the first 10 elements of the interleaved dataset
lines = [line for line in iter(interleaved_shakespeare_dataset.take(10))]
for line in lines:
print(line)
# -
# ***
# <a id="coding_tutorial_5"></a>
# ## Training with Datasets
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
# #### Load the UCI Bank Marketing Dataset
# +
# Load the CSV file into a pandas DataFrame
bank_dataframe = pd.read_csv('data/bank/bank-full.csv', delimiter=';')
# +
# Show the head of the DataFrame
bank_dataframe.head()
# +
# Print the shape of the DataFrame
print(bank_dataframe.shape)
# +
# Select features from the DataFrame
features = ['age', 'job', 'marital', 'education', 'default', 'balance', 'housing',
'loan', 'contact', 'campaign', 'pdays', 'poutcome']
labels = ['y']
bank_dataframe = bank_dataframe.filter(features + labels)
# +
# Show the head of the DataFrame
bank_dataframe.head()
# -
# #### Preprocess the data
# +
# Convert the categorical features in the DataFrame to one-hot encodings
from sklearn.preprocessing import LabelBinarizer
encoder = LabelBinarizer()
categorical_features = ['default', 'housing', 'job', 'loan', 'education', 'contact', 'poutcome']
for feature in categorical_features:
bank_dataframe[feature] = tuple(encoder.fit_transform(bank_dataframe[feature]))
# +
# Show the head of the DataFrame
bank_dataframe.head()
# +
# Shuffle the DataFrame
bank_dataframe = bank_dataframe.sample(frac=1).reset_index(drop=True)
# -
# #### Create the Dataset object
# +
# Convert the DataFrame to a Dataset
# +
# Inspect the Dataset object
# -
# #### Filter the Dataset
# +
# First check that there are records in the dataset for non-married individuals
def check_divorced():
bank_dataset_iterable = iter(bank_dataset)
for x in bank_dataset_iterable:
if x['marital'] != 'divorced':
print('Found a person with marital status: {}'.format(x['marital']))
return
print('No non-divorced people were found!')
check_divorced()
# +
# Filter the Dataset to retain only entries with a 'divorced' marital status
bank_dataset = bank_dataset.filter(lambda x : tf.equal(x['marital'], tf.constant([b'divorced']))[0] )
# +
# Check the records in the dataset again
check_divorced()
# -
# #### Map a function over the dataset
# +
# Convert the label ('y') to an integer instead of 'yes' or 'no'
# +
# Inspect the Dataset object
bank_dataset.element_spec
# +
# Remove the 'marital' column
# +
# Inspect the Dataset object
bank_dataset.element_spec
# -
# #### Create input and output data tuples
# +
# Create an input and output tuple for the dataset
def map_feature_label(x):
features = [[x['age']], [x['balance']], [x['campaign']], x['contact'], x['default'],
x['education'], x['housing'], x['job'], x['loan'], [x['pdays']], x['poutcome']]
return (tf.concat(features, axis=0), x['y'])
# +
# Map this function over the dataset
# +
# Inspect the Dataset object
# -
# #### Split into a training and a validation set
# +
# Determine the length of the Dataset
dataset_length = 0
for _ in bank_dataset:
dataset_length += 1
print(dataset_length)
# +
# Make training and validation sets from the dataset
# -
# #### Build a classification model
#
# Now let's build a model to classify the features.
# +
# Build a classifier model
from tensorflow.keras.layers import Dense, Input, Concatenate, BatchNormalization
from tensorflow.keras import Sequential
model = Sequential()
model.add(Input(shape=(30,)))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(400, activation='relu'))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(400, activation='relu'))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(1, activation='sigmoid'))
# +
# Compile the model
optimizer = tf.keras.optimizers.Adam(1e-4)
model.compile(optimizer=optimizer, loss='binary_crossentropy', metrics=['accuracy'])
# +
# Show the model summary
model.summary()
# -
# #### Train the model
# +
# Create batched training and validation datasets
# +
# Shuffle the training data
# +
# Fit the model
# +
# Plot the training and validation accuracy
| Customising your models with TensorFlow 2/Week 2/utf-8''Coding Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Basic Usage
#
# `hypml` is utility for loading valid Hypothesis Markup Language (.hypml) documents.
#
# You can import the module into your Python script:
import hypml
# ## Features
#
# ### `load`
#
# Loads a .hypml document. By default, only valid documents will be returned.
#
# #### Parameters
#
# - path : string
#
# a path to a .hypml document
#
# - ignore_validation : bool
#
# False (default) - returns nothing if validation fails
#
# True - returns the document contents even if invalid
#
# #### Returns
#
# - object : an object of the .hypml contents for downstream usage
#
# #### Example
#
# Load an example .hypml representing an Iris classifier. The file looks like this:
print(open('../examples/iris.hypml','r').read())
# Using the `load` function, we can load and validate the file and represent it as Python object:
hyp = hypml.load('../examples/iris.hypml')
hyp
# Which means you can do things like this:
hyp['task']
# And this:
for feature, properties in hyp['features'].items():
print(feature, 'is of type', properties['type'])
| python/Basic Usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
EA, l, F1, F2 = var("EA, l, F1, F2")
sub_list = [
( EA, 2 *Pa*m**2 ),
( l, 1 *m ),
( F1, 1 *Newton /2 ), # due to symmetry
( F2, 2 *Newton /2 ), # due to symmetry
]
def k(phi):
""" element stiffness matrix """
# phi is angle between:
# 1. vector along global x axis
# 2. vector along 1-2-axis of truss
# phi is counted positively about z.
# pprint("phi / deg:")
# pprint(N(deg(phi),3))
(c, s) = ( cos(phi), sin(phi) )
(cc, ss, sc) = ( c*c, s*s, s*c)
return Matrix(
[
[ cc, sc, -cc, -sc],
[ sc, ss, -sc, -ss],
[-cc, -sc, cc, sc],
[-sc, -ss, sc, ss],
])
(p1, p2, p3) = (315*pi/180, 0 *pi/180, 45 *pi/180)
# k2 uses only 1/2 A due to symmetry:
(k1, k2, k3) = (EA/l*k(p1), EA/2/l*k(p2), EA/l*k(p3))
pprint("\nk1 / (EA / l): ")
pprint(k1 / (EA/l) )
pprint("\nk2 / (EA / l): ")
pprint(k2 / (EA/l) )
pprint("\nk3 / (EA / l): ")
pprint(k3 / (EA/l) )
K = EA/l*Matrix([
[ 1 , -S(1)/2 ],
[ -S(1)/2, 1 ]
])
u2x, u3x = var("u2x, u3x")
u = Matrix([u2x , u3x ])
f = Matrix([F1 , F2 ])
u2x, u3x = var("u2x, u3x")
eq = Eq(K*u , f)
sol = solve(eq, [u2x, u3x])
pprint("\nSolution:")
pprint(sol)
u2x, u3x = sol[u2x], sol[u3x]
pprint("\nu2x / m:")
tmp = u2x.subs(sub_list)
tmp /= m
pprint(tmp)
pprint("\nu3x / m:")
tmp = u3x.subs(sub_list)
tmp /= m
pprint(tmp)
pprint("\nF1x / N:")
tmp = - EA/l * u2x/2
tmp = tmp.subs(sub_list)
tmp /= Newton
pprint(tmp)
# k1 / (EA / l):
# ⎡1/2 -1/2 -1/2 1/2 ⎤
# ⎢ ⎥
# ⎢-1/2 1/2 1/2 -1/2⎥
# ⎢ ⎥
# ⎢-1/2 1/2 1/2 -1/2⎥
# ⎢ ⎥
# ⎣1/2 -1/2 -1/2 1/2 ⎦
#
# k2 / (EA / l):
# ⎡1/2 0 -1/2 0⎤
# ⎢ ⎥
# ⎢ 0 0 0 0⎥
# ⎢ ⎥
# ⎢-1/2 0 1/2 0⎥
# ⎢ ⎥
# ⎣ 0 0 0 0⎦
#
# k3 / (EA / l):
# ⎡1/2 1/2 -1/2 -1/2⎤
# ⎢ ⎥
# ⎢1/2 1/2 -1/2 -1/2⎥
# ⎢ ⎥
# ⎢-1/2 -1/2 1/2 1/2 ⎥
# ⎢ ⎥
# ⎣-1/2 -1/2 1/2 1/2 ⎦
#
# Solution:
# ⎧ 2⋅l⋅(2⋅F₁ + F₂) 2⋅l⋅(F₁ + 2⋅F₂)⎫
# ⎨u2x: ───────────────, u3x: ───────────────⎬
# ⎩ 3⋅EA 3⋅EA ⎭
#
# u2x / m:
# 2/3
#
# u3x / m:
# 5/6
#
# F1x / N:
# -2/3
| ipynb/WB-Klein/5/5.4.ipynb |