code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#容量制約付き施設配置問題
from gurobipy import *
def make_data():
I,d = multidict({1:80, 2:270, 3:250, 4:160, 5:180}) # demand
J,M,f = multidict({1:[500,1000], 2:[500,1000], 3:[500,1000]}) # capacity, fixed costs
c = {(1,1):4, (1,2):6, (1,3):9, # transportation costs
(2,1):5, (2,2):4, (2,3):7,
(3,1):6, (3,2):3, (3,3):4,
(4,1):8, (4,2):5, (4,3):3,
(5,1):10, (5,2):8, (5,3):4,
}
return I,J,d,M,f,c
#multidictを三つ一気にやると、keyの先のリストから一つずつ辞書の先にくっつく
make_data()
#変数の名前はつけないと結果の解釈ができない
def flp(I,J,d,M,f,c):
model = Model("flp")
x, y = {} ,{}
for j in J:
y[j] = model.addVar(vtype = "B", name = "facility(%s)" %j)
for i in I:
x[i,j] = model.addVar(vtype = "C", name = "transport(%s, %s)" %(i,j))
model.update()
for i in I:
model.addConstr(quicksum(x[i,j] for j in J) == d[i])
for j in J:
model.addConstr(quicksum(x[i,j] for i in I) <= M[j]* y[j])
for (i,j) in x:
model.addConstr(x[i,j] <= d[i] * y[j])
model.setObjective(quicksum(f[j]*y[j] for j in J) + quicksum(c[i,j]* x[i,j] for i in I for j in J))
model.__data = x,y
return model
if __name__ == "__main__":
I,J,d,c,f,M = make_data()
model = flp(I,J,d,c,f,M)
model.optimize()
for v in model.getVars():
print v.VarName ,v.X
#k-median problem
def kmedian(I, J, c, k):
model = Model("k-median")
x ,y = {} ,{}
for j in J:
y[j] = model.addVar(vtype= "B", name = "facility(%s)" %j)
for i in I:
x[i,j] = model.addVar(vtype="B" ,name = "satisfaction(%s, %s)" %(i,j))
model.update()
for i in I:
model.addConstr(quicksum(x[i,j] for j in J) == 1)
for j in J:
model.addConstr(x[i, j] <= y[j])
model.addConstr(quicksum(y[j] for j in J) == k)
model.setObjective(quicksum(c[i,j] *x[i,j] for i in I for j in J))
model.__data = x, y
return model
def make_data2():
I = [1,2,3,4,5]
J = [1,2,3]
c = c = {(1,1):4, (1,2):6, (1,3):9,
(2,1):5, (2,2):4, (2,3):7,
(3,1):6, (3,2):3, (3,3):4,
(4,1):8, (4,2):5, (4,3):3,
(5,1):10, (5,2):8, (5,3):4,}
k = 2
return I,J,c,k
if __name__ == "__main__":
I,J,c,k = make_data2()
model = kmedian( I,J,c,k)
model.optimize()
# +
for v in model.getVars():
print v.VarName, v.X
#1、2さんは施設1に
#3、4、5さんは施設3に
#施設2は開設されなかった
# -
#k-center problem
def kcenter(I, J, c, k):
model = Model("k-center")
z = model.addVar(vtype = "C", name = "max_distance" )
x, y = {}, {}
for j in J:
y[j] = model.addVar(vtype= "B", name = "facility(%s)" %j)
for i in I:
x[i,j] = model.addVar(vtype="B" ,name = "satisfaction(%s, %s)" %(i,j))
model.update()
for i in I:
model.addConstr(quicksum(x[i,j] for j in J) == 1)
for j in J:
model.addConstr(x[i, j] <= y[j])
model.addConstr(c[i,j] * x[i,j] <= z)
model.addConstr(quicksum(y[j] for j in J) == k)
model.setObjective(z)
model.__data = x, y
return model
if __name__ == "__main__":
I,J,c,k = make_data2()
model = kcenter( I,J,c,k)
model.optimize()
# +
for v in model.getVars():
print v.VarName, v.X
#この場合はk-median problemと解は同じ
# -
| chapter2_location.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="UOf5F00phAYJ" executionInfo={"status": "ok", "timestamp": 1627382069983, "user_tz": -480, "elapsed": 4436, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="10cdfcd2-2b46-40a7-8223-1bcb914fbf5b"
# !pip install ckip-segmenter
# + id="KykLDOAOgdr8" executionInfo={"status": "ok", "timestamp": 1627382069983, "user_tz": -480, "elapsed": 4, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
from ckip import CkipSegmenter
# + id="wFVL84Dkg165" executionInfo={"status": "ok", "timestamp": 1627382072201, "user_tz": -480, "elapsed": 2221, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
segmenter = CkipSegmenter()
text =( "這會是一件很耗時的工作,他決定先把後腿拖回去,然後再回來處理。這得花上一整晚,而且一定會很冷。"
"他花了將近二十分鐘才將腿拖回棚屋,回去時已經筋疲力盡了。他將肉沿著牆儲存好,然後回到母麋鹿那裡。"
"已經到了正午,他非常饑餓。於是花了十五分鐘收集木材,在麋鹿旁邊生火。當火燒得很旺盛時,他從抬走的後腿旁邊的臀部切下一條肉,懸在一根木棍上,整塊肉幾乎就在火燄裡。"
"烤肉的同時,他回頭去切割和剝皮。他切下右前肩,就跟切後腿一樣,將肩胛骨切開,接著切開腿,然後拖回營地。回來時,木棍上的肉已烤得非常完美:外頭有點焦,裡頭則熟透了。" )
result = segmenter.seg(text)
# + id="WxrJI2k2hYyp" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627382072202, "user_tz": -480, "elapsed": 5, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="48d014d7-9294-47be-9e73-c5103a6b7d11"
# result.res is a list of tuples contain a token and its pos-tag.
# print('result.res: {}\n'.format(result.res))
# result.tok and result.pos contains only tokens and pos-tags respectively.
print('result.tok: {}\n'.format(result.tok))
print('result.pos: {}\n'.format(result.pos))
| small_practice/WordSegmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Authors:
# * <NAME> (plotly figures)
# * <NAME> and <NAME> (original tutorial)
#
# [MNE-Python](http://martinos.org/mne/stable/mne-python.html) is a software package for processing [MEG](http://en.wikipedia.org/wiki/Magnetoencephalography)/[EEG](http://en.wikipedia.org/wiki/Electroencephalography) data.
#
# The first step to get started, ensure that mne-python is installed on your computer:
import mne # If this line returns an error, uncomment the following line
# # !easy_install mne --upgrade
# Let us make the plots inline and import numpy to access the array manipulation routines
# add plot inline in the page
# %matplotlib inline
import numpy as np
# We set the log-level to 'WARNING' so the output is less verbose
mne.set_log_level('WARNING')
# ## Access raw data
# Now we import the MNE sample dataset. If you don't already have it, it will be downloaded automatically (but be patient as it is approximately 2GB large)
# +
from mne.datasets import sample
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw.fif'
# -
# Read data from file:
raw = mne.io.Raw(raw_fname, preload=False)
print(raw)
# The data gets stored in the `Raw` object. If `preload` is `False`, only the header information is loaded into memory and the data is loaded on-demand, thus saving RAM.
#
# The `info` dictionary contains all measurement related information: the list of bad channels, channel locations, sampling frequency, subject information etc. The `info` dictionary is also available to the `Epochs` and `Evoked` objects.
print(raw.info)
# Look at the channels in raw:
print(raw.ch_names[:5])
# The raw object returns a numpy array when sliced
data, times = raw[:, :10]
print(data.shape)
# Read and plot a segment of raw data
start, stop = raw.time_as_index([100, 115]) # 100 s to 115 s data segment
data, times = raw[:306, start:stop]
print(data.shape)
print(times.shape)
print(times.min(), times.max())
# MNE-Python provides a set of helper functions to select the channels by type (see [here](http://imaging.mrc-cbu.cam.ac.uk/meg/VectorviewDescription#Magsgrads) for a brief overview of channel types in an MEG system). For example, to select only the magnetometer channels, we do this:
picks = mne.pick_types(raw.info, meg='mag', exclude=[])
print(picks)
# Similarly, `mne.mne.pick_channels_regexp` lets you pick channels using an arbitrary regular expression and `mne.pick_channels` allows you to pick channels by name. Bad channels are excluded from the selection by default.
#
# Now, we can use picks to select magnetometer data and plot it. The matplotlib graph can be converted into an interactive one using Plotly with just one line of code:
# +
picks = mne.pick_types(raw.info, meg='mag', exclude=[])
data, times = raw[picks[:10], start:stop]
import matplotlib.pyplot as plt
import plotly.plotly as py
plt.plot(times, data.T)
plt.xlabel('time (s)')
plt.ylabel('MEG data (T)')
update = dict(layout=dict(showlegend=True), data=[dict(name=raw.info['ch_names'][p]) for p in picks[:10]])
py.iplot_mpl(plt.gcf(), update=update)
# -
# But, we can also use MNE-Python's interactive data browser to get a better visualization:
raw.plot();
# Let us do the same using Plotly. First, we import the required classes
from plotly import tools
from plotly.graph_objs import Layout, YAxis, Scatter, Annotation, Annotations, Data, Figure, Marker, Font
# Now we get the data for the first 10 seconds in 20 gradiometer channels
# +
picks = mne.pick_types(raw.info, meg='grad', exclude=[])
start, stop = raw.time_as_index([0, 10])
n_channels = 20
data, times = raw[picks[:n_channels], start:stop]
ch_names = [raw.info['ch_names'][p] for p in picks[:n_channels]]
# -
# Finally, we create the plotly graph by creating a separate subplot for each channel
# +
step = 1. / n_channels
kwargs = dict(domain=[1 - step, 1], showticklabels=False, zeroline=False, showgrid=False)
# create objects for layout and traces
layout = Layout(yaxis=YAxis(kwargs), showlegend=False)
traces = [Scatter(x=times, y=data.T[:, 0])]
# loop over the channels
for ii in range(1, n_channels):
kwargs.update(domain=[1 - (ii + 1) * step, 1 - ii * step])
layout.update({'yaxis%d' % (ii + 1): YAxis(kwargs), 'showlegend': False})
traces.append(Scatter(x=times, y=data.T[:, ii], yaxis='y%d' % (ii + 1)))
# add channel names using Annotations
annotations = Annotations([Annotation(x=-0.06, y=0, xref='paper', yref='y%d' % (ii + 1),
text=ch_name, font=Font(size=9), showarrow=False)
for ii, ch_name in enumerate(ch_names)])
layout.update(annotations=annotations)
# set the size of the figure and plot it
layout.update(autosize=False, width=1000, height=600)
fig = Figure(data=Data(traces), layout=layout)
py.iplot(fig, filename='shared xaxis')
# -
# We can look at the list of bad channels from the ``info`` dictionary
raw.info['bads']
# Save a segment of 150s of raw data (MEG only):
picks = mne.pick_types(raw.info, meg=True, eeg=False, stim=True, exclude=[])
raw.save('sample_audvis_meg_raw.fif', tmin=0., tmax=150., picks=picks, overwrite=True)
# Filtering is as simple as providing the low and high cut-off frequencies. We can use the `n_jobs` parameter to filter the channels in parallel.
# +
raw_beta = mne.io.Raw(raw_fname, preload=True) # reload data with preload for filtering
# keep beta band
raw_beta.filter(13.0, 30.0, method='iir', n_jobs=-1)
# save the result
raw_beta.save('sample_audvis_beta_raw.fif', overwrite=True)
# check if the info dictionary got updated
print(raw_beta.info['highpass'], raw_beta.info['lowpass'])
# -
# ## Define and read epochs
# First extract events. Events are typically extracted from the trigger channel, which in our case is `STI 014`. In the sample dataset, there are [5 possible event-ids](http://martinos.org/mne/stable/manual/sampledata.html#babdhifj): 1, 2, 3, 4, 5, and 32.
events = mne.find_events(raw, stim_channel='STI 014')
print(events[:5]) # events is a 2d array
# Events is a 2d array where the first column contains the sample index when the event occurred. The second column contains the value of the trigger channel immediately before the event occurred. The third column contains the event-id.
#
# Therefore, there are around 73 occurences of the event with event-id 2.
len(events[events[:, 2] == 2])
# And the total number of events in the dataset is 319
len(events)
# We can index the channel name to find it's position among all the available channels
raw.ch_names.index('STI 014')
raw = mne.io.Raw(raw_fname, preload=True) # reload data with preload for filtering
raw.filter(1, 40, method='iir')
# Let us plot the trigger channel as an interactive plot:
d, t = raw[raw.ch_names.index('STI 014'), :]
plt.plot(d[0,:1000])
py.iplot_mpl(plt.gcf())
# We can also plot the events using the `plot_events` function.
# +
event_ids = ['aud_l', 'aud_r', 'vis_l', 'vis_r', 'smiley', 'button']
fig = mne.viz.plot_events(events, raw.info['sfreq'], raw.first_samp, show=False)
# convert plot to plotly
update = dict(layout=dict(showlegend=True), data=[dict(name=e) for e in event_ids])
py.iplot_mpl(plt.gcf(), update=update)
# -
# Define epochs parameters:
event_id = dict(aud_l=1, aud_r=2) # event trigger and conditions
tmin = -0.2 # start of each epoch (200ms before the trigger)
tmax = 0.5 # end of each epoch (500ms after the trigger)
event_id
# Mark two channels as bad:
raw.info['bads'] = ['MEG 2443', 'EEG 053']
print(raw.info['bads'])
# The variable raw.info[‘bads’] is just a python list.
#
# Pick the good channels:
picks = mne.pick_types(raw.info, meg=True, eeg=True, eog=True,
stim=False, exclude='bads')
# Alternatively one can restrict to magnetometers or gradiometers with:
mag_picks = mne.pick_types(raw.info, meg='mag', eog=True, exclude='bads')
grad_picks = mne.pick_types(raw.info, meg='grad', eog=True, exclude='bads')
# Define the baseline period for baseline correction:
baseline = (None, 0) # means from the first instant to t = 0
# Define peak-to-peak rejection parameters for gradiometers, magnetometers and EOG. If the data in any channel exceeds these thresholds, the corresponding epoch will be rejected:
reject = dict(grad=4000e-13, mag=4e-12, eog=150e-6)
# Now we create epochs from the `raw` object. The epochs object allows storing data of fixed length around the events which are supplied to the `Epochs` constructor.
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, proj=True,
picks=picks, baseline=baseline, reject=reject)
# Now let us compute what channels contribute to epochs rejection. The drop log stores the epochs dropped and the reason they were dropped. Refer to the MNE-Python documentation for further details:
# +
from mne.fixes import Counter
# drop bad epochs
epochs.drop_bad_epochs()
drop_log = epochs.drop_log
# calculate percentage of epochs dropped for each channel
perc = 100 * np.mean([len(d) > 0 for d in drop_log if not any(r in ['IGNORED'] for r in d)])
scores = Counter([ch for d in drop_log for ch in d if ch not in ['IGNORED']])
ch_names = np.array(list(scores.keys()))
counts = 100 * np.array(list(scores.values()), dtype=float) / len(drop_log)
order = np.flipud(np.argsort(counts))
# -
# And now we can use Plotly to show the statistics:
# +
from plotly.graph_objs import Data, Layout, Bar, YAxis, Figure
data = Data([
Bar(
x=ch_names[order],
y=counts[order]
)
])
layout = Layout(title='Drop log statistics', yaxis=YAxis(title='% of epochs rejected'))
fig = Figure(data=data, layout=layout)
py.iplot(fig)
# -
# And if you want to keep all the information about the data you can save your epochs in a fif file:
epochs.save('sample-epo.fif')
# ## Average the epochs to get [Event-related Potential](http://en.wikipedia.org/wiki/Event-related_potential)
evoked = epochs.average()
# Now let's visualize our event-related potential / field:
fig = evoked.plot(show=False) # butterfly plots
update = dict(layout=dict(showlegend=False), data=[dict(name=raw.info['ch_names'][p]) for p in picks[:10]])
py.iplot_mpl(fig, update=update)
# topography plots
evoked.plot_topomap(times=np.linspace(0.05, 0.15, 5), ch_type='mag');
evoked.plot_topomap(times=np.linspace(0.05, 0.15, 5), ch_type='grad');
evoked.plot_topomap(times=np.linspace(0.05, 0.15, 5), ch_type='eeg');
# ### Get single epochs for one condition:
#
# Syntax is `epochs[condition]`
epochs_data = epochs['aud_l'].get_data()
print(epochs_data.shape)
# epochs_data is a 3D array of dimension (55 epochs, 365 channels, 106 time instants).
evokeds = [epochs[k].average() for k in event_id]
from mne.viz import plot_topo
layout = mne.find_layout(epochs.info)
plot_topo(evokeds, layout=layout, color=['blue', 'orange']);
# ## Compute noise covariance
noise_cov = mne.compute_covariance(epochs, tmax=0.)
print(noise_cov.data.shape)
fig = mne.viz.plot_cov(noise_cov, raw.info)
# ## Inverse modeling: [dSPM](http://www.sciencedirect.com/science/article/pii/S0896627300811381) on evoked and raw data
# Inverse modeling can be used to estimate the source activations which explain the sensor-space data.
#
# First, Import the required functions:
from mne.forward import read_forward_solution
from mne.minimum_norm import (make_inverse_operator, apply_inverse,
write_inverse_operator)
# ## Read the forward solution and compute the inverse operator
# The forward solution describes how the currents inside the brain will manifest in sensor-space. This is required for computing the inverse operator which describes the transformation from sensor-space data to source space:
# +
fname_fwd = data_path + '/MEG/sample/sample_audvis-meg-oct-6-fwd.fif'
fwd = mne.read_forward_solution(fname_fwd, surf_ori=True)
# Restrict forward solution as necessary for MEG
fwd = mne.pick_types_forward(fwd, meg=True, eeg=False)
# make an M/EEG, MEG-only, and EEG-only inverse operators
info = evoked.info
inverse_operator = make_inverse_operator(info, fwd, noise_cov,
loose=0.2, depth=0.8)
write_inverse_operator('sample_audvis-meg-oct-6-inv.fif',
inverse_operator)
# -
# ## Compute inverse solution
# Now we can use the inverse operator and apply to MEG data to get the inverse solution
method = "dSPM"
snr = 3.
lambda2 = 1. / snr ** 2
stc = apply_inverse(evoked, inverse_operator, lambda2,
method=method, pick_ori=None)
print(stc)
stc.data.shape
# Show the result:
# +
import surfer
surfer.set_log_level('WARNING')
subjects_dir = data_path + '/subjects'
brain = stc.plot(surface='inflated', hemi='rh', subjects_dir=subjects_dir)
brain.set_data_time_index(45)
brain.scale_data_colormap(fmin=8, fmid=12, fmax=15, transparent=True)
brain.show_view('lateral')
# -
brain.save_image('dspm.jpg')
brain.close()
from IPython.display import Image
Image(filename='dspm.jpg', width=600)
# ## Time-frequency analysis
# +
from mne.time_frequency import tfr_morlet
freqs = np.arange(6, 30, 3) # define frequencies of interest
n_cycles = freqs / 4. # different number of cycle per frequency
power = tfr_morlet(epochs, freqs=freqs, n_cycles=n_cycles, use_fft=False,
return_itc=False, decim=3, n_jobs=1)
# -
# Now let''s look at the power plots
# +
# Inspect power
power.plot_topo(baseline=(-0.5, 0), tmin=0, tmax=0.4, mode='logratio', title='Average power');
power.plot([82], baseline=(-0.5, 0), tmin=0, tmax=0.4, mode='logratio');
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install publisher --upgrade
import publisher
publisher.publish(
'mne-tutorial.ipynb', 'ipython-notebooks/mne-tutorial/', 'Plotly visualizations for MNE-Python to process MEG/EEG data',
'Create interactive visualizations using MNE-Python and Plotly', name='Process MEG/EEG Data with Plotly',
redirect_from='ipython-notebooks/meeg-and-eeg-data-analysis/')
# -
| _posts/ipython-notebooks/mne-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Nouran-Khallaf/bert_score/blob/master/Ahmed_BERT.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] Collapsed="false" id="SEQK0la8C8GR"
# <strong><h1 align = center><font size = 6>Sentiment Analysis with Deep Learning using BERT</font></h1></strong>
# + [markdown] Collapsed="false" id="1pPP_1-mC8Ga"
# # __1. Introduction__
# + [markdown] id="SwrtUv0Eg1nr"
# ## __What is BERT ?__
# + [markdown] id="ttfSDF0ziSC9"
# __Bidirectional Encoder Representations from Transformers__
# + [markdown] Collapsed="true" id="PpkwzvnkC8Gb"
# - __BERT__ is basically the advancement of the __RNNs__, as its able to Parallelize the Processing and Training. For Example $\rightarrow$ In sentence we have to process each word sequentially, __BERT__ allow us to do the things in Parellel.
# - BERT is a large-scale transformer-based Language Model that can be finetuned for a variety of tasks.
#
#
#
# + [markdown] id="8xOmmKCz7lw1"
# > We will be using the __Hugging Face Transformer library__ that provides a __high-level API__ to state-of-the-art transformer-based models such as __BERT, GPT2, ALBERT, RoBERTa, and many more__. The Hugging Face team also happens to maintain another highly efficient and super fast library for text tokenization called Tokenizers.
# + [markdown] id="vBvCWSDlZzD8"
# - Bidirectional: Bert is naturally bi-directional
# - Generalizable: Pre-trained BERT model can be fine-tuned easily for downstream NLp task.
# - High Performace: Fine-tuned BERT models beats state-of-art results for many NLP tasks.
# - Universal: Trained on Wikipedia() + BookCorpus. No special Dataset needed,
# + [markdown] id="wPDY1mrog3fp"
# __Extension of Architecture:__
#
# - __RoBERTa__
# - __DistilBERT__
# - __AlBERT__
#
# __Other Languages:__
#
# - __CamemBERT(French)__
# - __AraBERT(Arabic)__
# - __mBERT(Multilingual)__
# + [markdown] id="quAl9tMiW5xF"
# Google Research recently __open-sourced__ implementation of __BERT__ and also released the following pre-trained models:
#
#
# ---
#
#
#
# - BERT-Base, Uncased: 12-layer, 768-hidden, 12-heads, 110M parameters
# - BERT-Large, Uncased: 24-layer, 1024-hidden, 16-heads, 340M parameters
#
#
#
# ---
#
#
# - BERT-Base, Cased: 12-layer, 768-hidden, 12-heads , 110M parameters
# - BERT-Large, Cased: 24-layer, 1024-hidden, 16-heads, 340M parameters
#
#
#
# ---
#
#
# - BERT-Base, Multilingual Cased (New, recommended): 104 languages, 12-layer, 768-hidden, 12-heads, 110M parameters
# - BERT-Base, Chinese: Chinese Simplified and Traditional, 12-layer, 768-hidden, 12-heads, 110M parameters
# + [markdown] id="ilalkPN5bcc6"
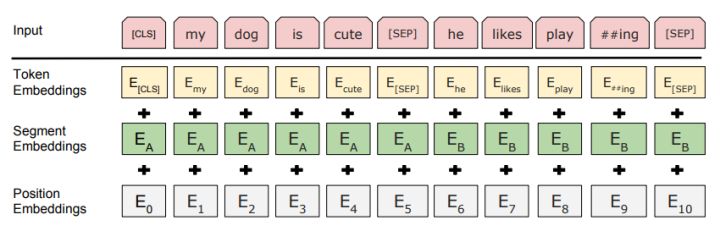
# ### __Embedding__
#
# In BERT, the embedding is the summation of three types of embeddings:
#
# 
#
#
# > __Token Embeddings__ is a word vector, with the first word as the __CLS flag__, which can be used for classification tasks.
#
#
# > __Segment Embeddings__ is used to distinguish between two sentences, since pre-training is not just a language modeling but also a classification task with two sentences as input
#
# > __Position Embedding__ is different from Transformer, __BERT__ learns a unique position embedding for the __input sequence__, and this __position-specific information__ can flow through the model to the __key__ and __query vectors__.
# + [markdown] id="ng2QG1GhdTIL"
# ### __Model Architecture__
#
# Here I use pre-trained BERT for binary sentiment analysis on Stanford Sentiment Treebank.
#
# - BertEmbeddings: Input embedding layer
# - BertEncoder: The 12 BERT attention layers
# - Classifier: Our multi-label classifier with out_features=2, each corresponding to our 2 labels
#
#
#
#
# ```
# - BertModel
# - embeddings: BertEmbeddings
# - word_embeddings: Embedding(28996, 768)
# - position_embeddings: Embedding(512, 768)
# - token_type_embeddings: Embedding(2, 768)
# - LayerNorm: FusedLayerNorm(torch.Size([768])
# - dropout: Dropout = 0.1
# - encoder: BertEncoder
# - BertLayer
# - attention: BertAttention
# - self: BertSelfAttention
# - query: Linear(in_features=768, out_features=768, bias=True)
# - key: Linear(in_features=768, out_features=768, bias=True)
# - value: Linear(in_features=768, out_features=768, bias=True)
# - dropout: Dropout = 0.1
# - output: BertSelfOutput(
# - dense: Linear(in_features=768, out_features=768, bias=True)
# - LayerNorm: FusedLayerNorm(torch.Size([768]),
# - dropout: Dropout =0.1
#
# - intermediate: BertIntermediate(
# - dense): Linear(in_features=768, out_features=3072, bias=True)
#
# - output: BertOutput
# - dense: Linear(in_features=3072, out_features=768, bias=True)
# - LayerNorm: FusedLayerNorm(torch.Size([768])
# - dropout: Dropout =0.1
# - pooler: BertPooler
# - dense: Linear(in_features=768, out_features=768, bias=True)
# - activation: Tanh()
# - dropout: Dropout =0.1
# - classifier: Linear(in_features=768, out_features = 2, bias=True)
# ```
#
#
# [Source: `mengxinji.github.io`](https://mengxinji.github.io/Blog/2019-03-27/pre-trained-bert/)
#
# + [markdown] id="y-I80ppLX4Gy"
# ### __Transformer model__
#
# The Transformer model was proposed in the paper: [Attention Is All You Need](https://arxiv.org/abs/1706.03762). In that paper they provide a new way of handling the sequence transduction problem (like the machine translation task) without complex recurrent or convolutional structure. Simply use a stack of attention mechanisms to get the latent structure in the input sentences and a special embedding (positional embedding) to get the locationality. The whole model architecture looks like this:
#
#
#
#
# 
# + [markdown] id="5W2V3JJQZjlZ"
# #### __Multi-Head Attention__
#
# Instead of using the __regular attention mechanism__, they split the __input vector__ to several pairs of __subvector__ and perform a __dot-product attention__ on each __subvector pairs__.
#
# 
#
# __Formula__:
#
# $
# Attention(Q, K, V) = softmax(\frac{QK^T}{\sqrt{d_k}})V
# $
#
# $
# MultiHead(Q, K, V) = Concat(head_1,..., head_h)W^O
# $
#
# $
# \text{where }head_i = Attention(QW^Q_i, KW^K_i, VW^V_i)
# $
# + [markdown] Collapsed="false" id="nBG8GiWjC8Ge"
# # __2. Exploratory Data Analysis and Preprocessing__
# + [markdown] Collapsed="false" id="1Gll1nBxC8Gf"
# __We will use the SMILE Twitter DATASET__.
#
# _<NAME>; <NAME>; <NAME>; <NAME>; <NAME>; <NAME> (2016): SMILE Twitter Emotion dataset. figshare. Dataset. https://doi.org/10.6084/m9.figshare.3187909.v2_
# + id="bmBPIEPGgVq1" outputId="29689ee1-61ba-459a-fe40-216afaf1b1a8" colab={"base_uri": "https://localhost:8080/"}
# ! pip install torch torchvision
# + id="NuCrFkvOhaHy" outputId="3e814311-9f73-4d6d-d4bd-d7be99e6be65" colab={"base_uri": "https://localhost:8080/"}
# ! pip install tqdm
# + [markdown] id="uyk1H_eIjMDX"
# [Python: Progress Bar with tqdm](https://youtu.be/qVHM3ly-Amg)
#
# > $Tqdm$ : Tqdm package is one of the more comprehensive packages for __Progress Bars__ with python and is handy for those instances you want to build scripts that keep the users informed on the status of your application.
#
# + id="DCneeeeig1Eq"
import torch
import pandas as pd
from tqdm.notebook import tqdm
# + id="Zqs_1FABicRn" outputId="f10a670c-4568-42d1-82b2-d0aaf3fc6d48" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 56}
from google.colab import files
uploaded = files.upload()
# + id="AqgLaF5GlbN5" outputId="e2700d93-7900-4ba1-da35-2e7f047d5d2b" colab={"base_uri": "https://localhost:8080/", "height": 34}
# ls
# + id="2r65PflHmUfM"
df = pd.read_csv('/content/ss.csv', names=[ 'text', 'category'])
# + [markdown] id="JpRANk8Yk5QO"
# > Pandas `set_index()` is a method to set a List, Series or Data frame as index of a Data Frame. Index column can be set while making a data frame too. But sometimes a data frame is made out of __two or more data frames__ and hence later index can be changed using this method.
#
#
# $Syntax$
# ```
# DataFrame.set_index(keys, drop=True, append=False, inplace=False, verify_integrity=False)
# ```
#
#
# + id="JZvENhJrAKNH" outputId="e578bfa9-38e6-44a1-dff3-bdaa7ed31e5b" colab={"base_uri": "https://localhost:8080/", "height": 423}
df
# + id="riMava0StvEM" outputId="e41431fc-7cea-4b06-fd38-286486fed520" colab={"base_uri": "https://localhost:8080/", "height": 122}
df.text.iloc[1]
# + id="c9tPCGUinTpD" outputId="c9ddb286-b1bf-4bc0-a4d5-97bc4b277aff" colab={"base_uri": "https://localhost:8080/", "height": 363}
df.head(10)
# + [markdown] id="Y8lMQxj6q2pD"
# $\color{red}{\textbf{NOTE:}}$ `id` is in bold because we set it as an __index__, So its no longer a data in the actual dataframe
# + id="3RSnIFOAnt9e" outputId="5159dc40-4d44-4f81-ab8f-e4b95c57b13f" colab={"base_uri": "https://localhost:8080/"}
df.category.value_counts() # it counts How many times each unique instance occur in your data
# + [markdown] id="9oXCQ-tPvyqc"
# - So we choose to ignore the _nodecode_ as it dose not contaion any emotions.
# - we also choose to ignore the multiple emotions as it makes our __BERT__ more Complicated.
# - So essentially we want. is $\rightarrow$ _one tweet to have one result._
#
# + id="TSPHN9HuoGdn"
# Removing the tweet with multiple category/nocode
df = df[~df.category.str.contains('\|')]
#str -> As we have to pull-it out of the string
#contain -> if the str contaion '|' -> Return True, Else False
# + id="oLmdt11eor9X"
df = df[df.category != 'nocode']
# + id="FbT4U8z5o3LA" outputId="eb4bc726-c61f-47ab-b951-543c29bbae3d" colab={"base_uri": "https://localhost:8080/"}
df.category.value_counts()
# + [markdown] id="CXX9UJyyzXE5"
# > This Shows that we have a __Class imbalance__ here, and we ned to take this into account.
#
# ```
# happy 1137
# not-relevant 214
# angry 57
# surprise 35
# sad 32
# disgust 6
# Name: category, dtype: int64
# ```
#
#
#
#
# + [markdown] id="fbCUTNiL0KX0"
# Building a _dictionary_ that can convert the emotions into the revelent number.
#
# _for example:_
#
# ```
# happy 1
# not-relevant 2
# angry 3
# surprise 4
# sad 5
# disgust 6
# ```
#
#
# + id="MKIPf5BopGoC"
possible_labels = df.category.unique() # Now we have the list that conatin all-of the labels
# + id="e3_Zwy24prMx"
label_dict = {} # Creating an empty Dict, & Looping over the possible labels
for index, possible_label in enumerate(possible_labels):
label_dict[possible_label] = index
# + id="VZK4jGlB4tvp" outputId="11a60fbd-152c-44fc-ca2e-e5de75fbb8db" colab={"base_uri": "https://localhost:8080/"}
label_dict
# + [markdown] id="RegoqlnB3BgN"
# _looping over the iterable and return the index_
#
# > `Enumerate()` in Python:
# A lot of times when dealing with iterators, we also get a need to keep a count of iterations. Python eases the programmers’ task by providing a built-in function `enumerate()` for this task.
#
# > `Enumerate()` method adds a counter to an iterable and returns it in a form of enumerate object. This enumerate object can then be used directly in for loops or be converted into a list of tuples using `list()` method.
# + [markdown] id="lZWdreQI4Mw7"
# $Synatx$
#
# ```
# enumerate(iterable, start=0)
#
# Parameters:
# Iterable: any object that supports iteration
# Start: the index value from which the counter is
# to be started, by default it is 0
# ```
#
#
# + id="Y_RuKe7FqSe9"
df['label'] = df.category.replace(label_dict)
# + [markdown] id="HJq2ojtqCjuT"
#
# + id="G6VNoepjqegz" outputId="ec9b7a0a-b9f1-41fe-e134-e393b04f89fa" colab={"base_uri": "https://localhost:8080/", "height": 676}
df.head(20)
# + [markdown] Collapsed="false" id="UqAcpMxIC8Hd"
# # __3. Training/Validation Split__
# + [markdown] id="cORWSHP6I9pF"
# [__train_test_split__ Vs __StratifiedShuffleSplit__](https://medium.com/@411.codebrain/train-test-split-vs-stratifiedshufflesplit-374c3dbdcc36)
# + id="wdIScfN7I9Q1"
from sklearn.model_selection import train_test_split
# + id="JT-HjRLRH8Nq"
x_train, x_val, y_train, y_val = train_test_split(df.index.values,
df.category.values,
test_size=0.15,
random_state=17,
)
# + [markdown] id="esQKksQ3mGK7"
# - the first thing we give in `train_test_split` is the _index value._ So as to uniquely identify each sample.
# - `df.label.values` it'll doing the random split based on index and label.
# - `test_size` is kept at `15%` so as to provide more data for training.
# - `random_state` ensures that the splits that you generate are __reproducible__. Scikit-learn uses random permutations to generate the splits. The random state that you provide is used as a __seed__ to the random number generator. This ensures that the random numbers are generated in the same order.
#
# > When the Random_state is not defined in the code for every run train data will change and accuracy might change for every run. When the `Random_state` = _"constant integer"_ is defined then train data will be constant For every run so that it will make easy to debug.
#
# - `stratify` to ensure that your training and validation datasets each contain the same percentage of classes
#
#
# + id="O7aTSWwvkknZ"
# Creating the New column in our dataframe --> 'data_type'
# data_type is Initally 'not_set' for all the samples
df['data_type'] = ['not_set']*df.shape[0]
# + id="QqxdWg_TqYBI" outputId="19ae3eeb-abf3-46ca-c625-dc895419a87c" colab={"base_uri": "https://localhost:8080/", "height": 206}
df.head()
# + id="UHpclaOTkkh-"
df.loc[x_train, 'data_type'] = 'train'
df.loc[x_val, 'data_type'] = 'val'
# + id="2n7DuorFleuq" outputId="b585c220-2607-46ff-f976-8219f8b7543a" colab={"base_uri": "https://localhost:8080/", "height": 238}
df.groupby(['category', 'data_type']).count()
# + [markdown] id="jozyup-8rIlF"
# Pandas `dataframe.groupby()` function is used to split the data into groups based on some criteria. pandas objects can be split on any of their axes. The abstract definition of grouping is to provide a mapping of labels to group names.
# + [markdown] Collapsed="false" id="aFCwQ08lC8Hz"
# # __4. Loading Tokenizer and Encoding our Data__
# + [markdown] id="7X9nLd1uGDAn"
# __BERT-Base__, uncased uses a vocabulary of __30,522__ words. The processes of __tokenization__ involves splitting the input text into list of tokens that are available in the vocabulary. In order to deal with the words not available in the vocabulary, BERT uses a technique called __BPE__ based WordPiece tokenization.
# + id="W1lNWlq54VoD" outputId="9f6269e7-ceb8-4a1e-ca9e-367d20e6ac91" colab={"base_uri": "https://localhost:8080/"}
# ! pip install transformers
# + id="6hB5gMZCH88P"
from transformers import BertTokenizer
from torch.utils.data import TensorDataset
# + [markdown] id="TryATa49ABGO"
# ### __Tokenizer__
#
# __Tokenizer__ takes the raw text as an input and splits it into the _Tokens_, Its a numerical number that represents a certain word.
#
# > __Tokenizer__ convert the text into the numerical data
#
# `TensorDataset`: It setup the data in the Pytorch enviorment. The Dataset wrapped into the tensors. Each sample will be retrieved by indexing tensors along the first dimension.
# + [markdown] id="viZIt_qA5S7E"
# > __BERT__ was trained using the WordPiece __tokenization__. It means that a word can be broken down into more than one __sub-words__. For example, _if I tokenize the sentence “Hi, my name is Dima”_ -- I'll get: tokenizer.tokenize('Hi my name is Dima')# OUTPUT. `['hi', 'my', 'name', 'is', 'dim', '##a']`
# + id="MpzN32Pg5SfO" outputId="faddb173-2208-43f7-f489-da5a8359ca20" colab={"base_uri": "https://localhost:8080/", "referenced_widgets": ["913a3a6c19d946208aa15ac27f006e30", "e1a0bafbcc9c4b4b9d94801ca2bd8e42", "01f5d2c9857446d1be7861522638dd69", "049f771f2fca40f88b990bbb34451137", "d2882040d7fe48b79ca9c0d16254470d", "5a2a41534868423e9e54cd7f913a131f", "080a7267bcf94e3e937ef6125d3dd7eb", "1f5fc387c7974b23ad6ec3dc030f79ce", "506e73ab21594ca3a1a979c2053e1539", "<KEY>", "5343e90f53104491bde623645148fe8a", "96ab64ec121d4df59d0175f11a534d58", "2d3762b90c68475e8572525462ef207a", "ee4e4de835ca423fb1ea3cea7cc8c14b", "<KEY>", "89a14afdc19e49609abe0ba7cb203e69", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "1fdd2e59e903428d8a3a474839b35618", "<KEY>", "5211ec83ee3d41d0b9423e29f8a2dbdd", "<KEY>", "61baf12d7b7b4ab3a4e6756571b3d6e4", "<KEY>", "c2319cd51d224104af27bed8160724c3", "<KEY>", "ac1e1fcc410e4d569a278bf47c7f0152", "<KEY>", "95c509ddb0e14dc19b91c41a8dbef623", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "07962dc1ebf047feba6dd9fbd279d735", "<KEY>", "68b2abe78e484278965c940ce50a3483", "<KEY>", "<KEY>", "<KEY>", "e520e59d38aa4f59ad181a191a90cf6d", "<KEY>", "5b54f605b290438c87df411f9e463f17"], "height": 145}
# The Tokenizer came from the Pre_trained BERT
# 'bert-base-uncased' means that we are using all lower case data
# `do_lower_case` Convert everything to lower-case.
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased',
do_lower_case=True)
# + [markdown] id="opFMB4NUG9sm"
# ### __Encoding__
#
#
# Convert all the Tweets into the encoded form.
#
#
# + id="NEiVBptX4oQy" outputId="bce6dc9d-5563-458e-be39-6f8e5e20b0f9" colab={"base_uri": "https://localhost:8080/"}
# Encoding the Training data
encoded_data_train = tokenizer.batch_encode_plus(
df[df.data_type=='train'].text.values,
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
# Encoding the Validation data
encoded_data_val = tokenizer.batch_encode_plus(
df[df.data_type=='val'].text.values,
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
# Spliting the data for the BERT training
'''
What the BERT needs for Training?
--> Inputs ids
--> Attention Masks
--> & Labels
'''
input_ids_train = encoded_data_train['input_ids']
attention_masks_train = encoded_data_train['attention_mask']
labels_train = torch.tensor(df[df.data_type=='train'].label.values)
input_ids_val = encoded_data_val['input_ids']
attention_masks_val = encoded_data_val['attention_mask']
labels_val = torch.tensor(df[df.data_type=='val'].label.values)
# + [markdown] id="AuUYGX2uIlG0"
# - `batch_encode_plus` is used to convert Multiple Strings into token as we need them. And this is perform seperately for both train and validation data.
#
# - `df[df.data_type=='train'].text.values`: we takes all the training data & takes the text values from it.
#
# - `add_special_tokens`: This is just the __BERT__ way of Knowing that when the sentence __ENDs__ and when the a __NEW__ one Begins.
#
# - `return_attention_mask`: Because we are using the _Fixed Input_. So, for an Instance we are having an sentence with $5$ words, and another sentence has $50$ $\rightarrow$ Everything has to be of same __Dimensionality__. So we set our `max_length` to a large value $256$, So as to contain all the Possible values. `attention_mask` tells where the actual values are, and where the blank[__Zeros__] are.
#
# - `max_length=256` as single Tweet dosen't have more than 256 words in it.
#
# - `return_tensors='pt'`: this represents how we wants to return these Tensors -- `pt` here represents __PyTorch__.
# + [markdown] id="VuX8URftU2QV"
# ### __We have to convert the input to the feature that is understood by BERT__
#
# - input_ids: list of numerical ids for the tokenized text
# - input_mask: will be set to 1 for real tokens and 0 for the padding tokens
# - segment_ids: for our case, this will be set to the list of ones
# - label_ids: one-hot encoded labels for the text
# + [markdown] id="kidV8EFmPVDp"
#
#
# ```python
# input_ids_dataset = encoded_data_dataset['input_ids']
# attention_masks_dataset = encoded_data_dataset['attention_mask']
# labels_dataset = torch.tensor(df[df.data_type=='dataset'].label.values)
# ```
#
# - `encoded_data_dataset` This will return the dictionary --> and we will pull out the `input_ids`, It represents each word as a number
#
# - similarly we will pull out the list of `attention_mask` as a PyTorch
# tensor.
#
# - Next we pulls the label, because thats the Numerical number we need.
#
# + id="Zgrqq3fK-9v5"
# Creating two different dataset
dataset_train = TensorDataset(input_ids_train, attention_masks_train, labels_train)
dataset_val = TensorDataset(input_ids_val, attention_masks_val, labels_val)
# + id="2T9xynJi_Z3U" outputId="3301eb2e-961e-46cd-9b47-f8e8add6d6af" colab={"base_uri": "https://localhost:8080/"}
len(dataset_train)
# + id="vtEXKIhs_aT-" outputId="236fa59c-1fee-4621-8098-c4e926ae0d35" colab={"base_uri": "https://localhost:8080/"}
len(dataset_val)
# + [markdown] Collapsed="false" id="gVyD956NC8IG"
# # __5. Setting up BERT Pretrained Model__
# + id="MwZiK26SH9va" outputId="c0990b94-f559-4d01-c835-5f92881a3d89" colab={"base_uri": "https://localhost:8080/", "referenced_widgets": ["<KEY>", "<KEY>", "<KEY>", "97a74df858514a66a123fea205dab0f1", "<KEY>", "dd9a652b33714ff7a54d04aea080b969", "0a769d0f6b8249ad9c6157fd6e23924f", "<KEY>", "<KEY>", "4b3856ad12984e0cbfe01a54edc97a3f", "3be7a4254b6b41cd863721d7900a0f80"], "height": 156}
from transformers import BertForSequenceClassification
model = BertForSequenceClassification.from_pretrained("bert-base-uncased",
num_labels=len(label_dict),
output_attentions=False,
output_hidden_states=False)
# + [markdown] id="_twV0ockrYLk"
# - Each tweet is treated as its own unique sequence.So one sequence will be classified into one of six classes
#
# - we are using the __BERT__ `bert-base` version as its Computationally efficent, & it's a smaller version.
#
# - `num_labels=len(label_dict)` which is how many output labels this final __BERT__ layout will have to be abel to classify.
#
# - `output_attentions=False` as we don't want any un-necessary inputs from the model.
#
# - we also don't care about the `output_hidden_states`, which is the state just before the prediction.
#
# + [markdown] id="FVj3PHarDJY4"
#
# + [markdown] Collapsed="false" id="DMJo2I-pC8IQ"
# # __6. Creating Data Loaders__
# + id="eoFfSpJ9xeim"
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# + [markdown] id="kvmcjn-yzNge"
# > __Dataloader__ Combines a `dataset` and a `sampler`, and provides single or multi-process __iterators__ over the dataset.
#
# Large datasets are _indispensable_ in the world of __Machine learning__ and __Deep learning__ these days. However, working with large datasets requires loading them into memory all at once.
#
# This leads to memory outage and slowing down of programs. PyTorch offers a solution for __parallelizing__ the data loading process with the support of automatic batching as well. This is the DataLoader class present within the `torch.utils.data package`
#
# <img src='https://cdn.journaldev.com/wp-content/uploads/2020/02/PyTorch-Data-Loader.png' width='400' height='450'>
#
# $\Rightarrow$ [How does data loader work PyTorch?](https://youtu.be/zN49HdDxHi8)
#
# $\Rightarrow$ [PyTorch-dataloader](https://www.journaldev.com/36576/pytorch-dataloader)
# + [markdown] id="HNQkqznC0niY"
# - `RandomSampler`, `SequentialSampler` - This is how to sample the data per batch. we use `RandomSampler` for traning, it randomize how our model is training & what data it's being Exposed to ans it also prevents the model from learning the sequence based differences while training.
#
# Where as the `SequentialSampler` return the samples sequentially contained in the dataset passed to the sampler, It takes in the dataset, not the set of indices.
#
# + id="uoHbpp61xeU3"
batch_size = 32
# We Need two different dataloder
dataloader_train = DataLoader(dataset_train,
sampler=RandomSampler(dataset_train),
batch_size=batch_size)
dataloader_validation = DataLoader(dataset_val,
sampler=RandomSampler(dataset_val),
batch_size=batch_size)
# + [markdown] Collapsed="false" id="kE4iakK_C8IY"
# # __7. Setting Up Optimiser and Scheduler__
# + id="19VqcD9yH-nk"
from transformers import AdamW, get_linear_schedule_with_warmup
# + [markdown] id="Tg89NhYY7aAv"
# __AdamW__
#
# > - Compute __weight decay__ before applying __gradient step__.
# - Multiply the weight decay by the learning rate.
#
# 
#
# The original Adam algorithm was proposed in Adam: 'A Method for Stochastic Optimization'. The AdamW variant was proposed in 'Decoupled Weight Decay Regularization'.
# + [markdown] id="e9bqfN95-VqL"
#
#
# ---
#
#
# `get_linear_schedule_with_warmup` Warm up steps is a parameter which is used to lower the __learning rate__ in order to reduce the impact of __deviating__ the model from learning on __sudden new data set exposure__.
#
# > _By default, number of warm up steps is 0._
#
# Then you make bigger steps, because you are probably not near the minima. But as you are approaching the minima, you make smaller steps to converge to it.
#
# Also, note that number of training steps is __number of batches * number of epochs__, but not just number of epochs. So, basically num_training_steps = N_EPOCHS+1 is not correct, unless your batch_size is equal to the training set size.
#
#
# __Source__:[Optimizer and scheduler for BERT fine-tuning](https://stackoverflow.com/questions/60120043/optimizer-and-scheduler-for-bert-fine-tuning)
# + id="Fmcvkx1fvX9n"
'''
Learning Rate as per the original paper: -- 2e-5 > 5e-5 --
'''
optimizer = AdamW(model.parameters(),
lr=1e-5,
eps=1e-8)
# + id="aaNqUERavXtn"
epochs = 10
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps=0,
num_training_steps=len(dataloader_train)*epochs)
# + [markdown] Collapsed="false" id="z8-HOaJdC8Ik"
# # __8. Defining our Performance Metrics__
# + [markdown] Collapsed="false" id="c8ZBTCCdC8Il"
# Accuracy metric approach originally used in accuracy function in [this tutorial](https://mccormickml.com/2019/07/22/BERT-fine-tuning/#41-bertforsequenceclassification).
# + id="o3ir6NIWoXv9"
import numpy as np
# + id="NRVzH3yiod0r"
from sklearn.metrics import f1_score
# + [markdown] id="zxqSsHwPsza9"
# There are total of Six labels to classify
#
# - preds-probability = [0.9, 0.05, 0.05, 0, 0, 0]
# - preds-binary-labels = [1, 0, 0, 0, 0, 0] --> These are Flat Values that we want
#
# __Flatten in contex of Keras__
#
# > __Flattening__ means. It breaks the spatial structure of the data and transforms your tridimensional $(W-(s-1), H - (s-1), N)$ tensor into a monodimensional tensor (a vector) of size $(W-(s-1))x(H - (s-1))xN$.
#
# 
#
# > Flatten make explicit how you serialize a __multidimensional tensor__ (tipically the input one). This allows the __Mapping__ between the (flattened) input tensor and the first hidden layer. If the first hidden layer is "dense" each element of the (serialized) input tensor will be connected with each element of the hidden array. If you do not use Flatten, the way the input tensor is mapped onto the first hidden layer would be ambiguous.
# + id="lPW6j1IZonLm"
def f1_score_func(preds, labels):
# Setting up the preds to axis=1
# Flatting it to a single iterable list of array
preds_flat = np.argmax(preds, axis=1).flatten()
# Flattening the labels
labels_flat = labels.flatten()
# Returning the f1_score as define by sklearn
return f1_score(labels_flat, preds_flat, average='weighted')
# + [markdown] id="H-RF2wABzkkD"
# [__sklearn.metrics.f1_score__](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html)
# + id="jHjqt1NxpkfW"
def accuracy_per_class(preds, labels):
label_dict_inverse = {v: k for k, v in label_dict.items()}
preds_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
# Iterating over all the unique labels
# label_flat are the --> True labels
for label in np.unique(labels_flat):
# Taking out all the pred_flat where the True alable is the lable we care about.
# e.g. for the label Happy -- we Takes all Prediction for true happy flag
y_preds = preds_flat[labels_flat==label]
y_true = labels_flat[labels_flat==label]
print(f'Class: {label_dict_inverse[label]}')
print(f'Accuracy: {len(y_preds[y_preds==label])}/{len(y_true)}\n')
# + [markdown] id="ztgTvJ-D0PLH"
# - ` label_dict_inverse` before we have [ __Happy__$\rightarrow$0 ] now we have [ 0$\rightarrow$__Happy__ ], So we have crated a _NEW inverse DICTIONARY_ , where insted of [ __Key__$\rightarrow$__Value__ ] we have [ __Value__$\rightarrow$__Key__ ]
#
#
#
#
#
# + [markdown] Collapsed="false" id="9ZEjEw63C8Ix"
# # __9. Create a training loop to control PyTorch finetuning of BERT using CPU or GPU acceleration__
# + [markdown] Collapsed="false" id="k2FQSUItC8Iy"
# Approach adapted from an older version of HuggingFace's `run_glue.py` script. Accessible [here](https://github.com/huggingface/transformers/blob/5bfcd0485ece086ebcbed2d008813037968a9e58/examples/run_glue.py#L128).
# + id="h4rcMNgJSFtv"
import random
seed_val = 17
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# + [markdown] id="9_t4pKXqSGyH"
# - A seed value specifies a particular stream from a set of possible random number streams. When you specify a seed, SAS generates the same set of pseudorandom numbers every time you run the program.
#
# - Seed function is used to save the state of a random function, so that it can generate same random numbers on multiple executions of the code on the same machine or on different machines (for a specific seed value). The seed value is the previous value number generated by the generator.
# + id="HEYBy1HxSFmJ" outputId="97552005-91ac-451b-8d8b-92009758b0ab" colab={"base_uri": "https://localhost:8080/"}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model.to(device)
print(device)
# + id="ieZu-O7VSFdv"
def evaluate(dataloader_val):
model.eval()
loss_val_total = 0
predictions, true_vals = [], []
for batch in tqdm(dataloader_val):
batch = tuple(b.to(device) for b in batch)
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
with torch.no_grad():
outputs = model(**inputs)
loss = outputs[0]
logits = outputs[1]
loss_val_total += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = inputs['labels'].cpu().numpy()
predictions.append(logits)
true_vals.append(label_ids)
loss_val_avg = loss_val_total/len(dataloader_val)
predictions = np.concatenate(predictions, axis=0)
true_vals = np.concatenate(true_vals, axis=0)
return loss_val_avg, predictions, true_vals
# + id="urde6RvHbbKH" outputId="c3c976e9-4a55-42b3-de53-41acfad95c8f" colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["c0e84400362c421a9aab65ab916cd205", "27ef963ce6ce4c04a6948f5ce8c8388b", "b13325c43ed641b2b1e8629ac7fd355e", "d1055e85af144ed19b3f5471d98534ce", "<KEY>", "<KEY>", "<KEY>", "e9017d618e6f4632bcd2d7ae43da1789", "be307fd8547a472bacd48071503b5809", "c689b5c17e3045ca8af9afe39ad311eb", "ee6dd4f0bdf447cb842e9afced8e0fd4", "1ede129058224a5490f56f6cc095a339", "<KEY>", "<KEY>", "<KEY>", "cf4e2345e3254edb94ba3b2766a8da7d", "a8289503bb6b4524bef6f429ad0609d7", "de87e2f2016c448c94e57a9a01f18fac", "5159598d7f5a4c82a93a126ba0cb0685", "<KEY>", "<KEY>", "33c883417b47472dadd11ff51d03ad95", "<KEY>", "653b56ab18574334ab709b89c2097857", "<KEY>", "0c708620e2024f05b7791a16fb734439", "41ee0d5ec5304c6c9c82e84538b53f39", "<KEY>", "18f95ac2af36411cb80513fe24881ed2", "d5e62f68d9ce434999b258d0208881c9", "7881fe9c5cc3477bb137cc85e3b4e464", "d3e80e34788d41ec9291d280930ecb9f", "162a071d96304c91a7ebba4b2e350a3f", "dde3868f9a1a4dfca8a05088080a5b59", "<KEY>", "410a58d16af242059ce8e585f3afef18", "<KEY>", "<KEY>", "<KEY>", "81a1d8b24c3a4cf887b089262940a225", "dee8801cbde34d74a38d7406cd719e43", "<KEY>", "<KEY>", "<KEY>", "6a7a2dbd9307462591829c9a4718255c", "<KEY>", "7577ed6300664f0a836a5018c4b57e97", "<KEY>", "f2a1e118199c4e6199203352254f77c1", "<KEY>", "4534a55d55da4047a1d14904db298952", "<KEY>", "<KEY>", "5890d3358d3e43f7a6ba0c754c1a4882", "<KEY>", "9139e3678c7246a797789ace6c8b41c7", "64461c27fa5745f68554e0ce20134f33", "a1db6bcdd45c40119ce5c399ef1ea5cc", "<KEY>", "<KEY>", "<KEY>", "1f0c7d62a15649829c61a2e372ec6135", "<KEY>", "<KEY>", "f74499e22a6849c19e014b74f9aaa383", "<KEY>", "<KEY>", "0d93d88f27064caeb31ee2d9d9a889a1", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e673489460dd42578305b75b171a3fa1", "48a7e9e0a4784d028d36c260414b7afe", "<KEY>", "91a49ea475524adeaf4e6c5d23fe4c09", "921f9a549ef7427e83bf76675ac3106a", "<KEY>", "5db739eef588470696df15d79998fe51", "<KEY>", "<KEY>", "<KEY>", "547f6f3715eb4182a3ef5e05e048b725", "<KEY>", "553e4f60f2494a5dae9a1fe71b8be32d", "<KEY>", "4edf8dc46e3741129f73ea3862fd9dac", "1e996adad4f34c9999fa497fc1c5a122", "<KEY>", "<KEY>", "4b6279ec1b9f43c8958e37c7c5e79c07", "b6f5ea15239d48f8a556e796f68a0fa9", "2af5aba8d93e4a26a7d1aa53ff95b3c3", "0ac08ae5ad064b3b9bceed130c8adbb5", "614c4b6b72524be38e8c6e1ba2d8d5cf", "<KEY>", "300276756a7d4d5bb3759b9e02a29dd9", "b4b4e7895e7a4a0c859acc5d7045edf6", "<KEY>", "87fa68ae5fa34d898fc344ea3b5332e2", "<KEY>", "e42bed205f6b4282a09b6739ffde1fcb", "<KEY>", "361f4678a1664695a03f7e2b7de314ff", "471ca7986dc940598368eb79e404c3f7", "<KEY>", "e208d95003884f7cbe7153d82328e444", "c88ed7d2a91c42f8a07656dc974091c5", "<KEY>", "7207d4b312af42e2a9c8de2170f48f29", "<KEY>", "<KEY>", "86682110ea384804b2a5ea64ba50580a", "7a16d412defb4ef4823d4123ed5309f6", "<KEY>", "538f8e8910024cb6bcee5e9e49af04db", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4f5a4a9b83e046feb7a2f878622958e4", "256b482f48a14425abed146d01c932e9", "39d0932230a34f6fbba0a595d9d7d162", "836665405fd64ee9b36b3501a39f5420", "<KEY>", "477667773f834ab2a7f6a758153ff3aa", "<KEY>", "8491b51d49f54ed6896d76147c6c9989", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "050e2a17ba764760a2c388983065ee50", "<KEY>", "<KEY>", "<KEY>", "875e574a07a540ab8eee85b6a1ad6737", "d1a88cd1fec244f08a55dd0d54fb0924", "02424474892f4fadaa4b12e2221aebff", "<KEY>", "e74c7c2530a24659b9fbedea59bf767c", "185cd6c878af44a09c235ce5891e61f3", "9ec6d098ac78443ea937bf899abae9a8", "7f45817ae9c9499b9ba6ce87820e1a22", "83c50b8695774121a3b6727de5f5e737", "<KEY>", "<KEY>", "35be77fa45e24901a1c2d605f7f1d4a1", "79640f5541484cd8bad6b39707b5ded2", "d1e1a1c4e06c42819b56176cd05e57ab", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "e351bc4d623746b480b34299e3be9df2", "<KEY>", "<KEY>", "10d8906227ed4efda2eeb3fb7807baf9", "966f943191a74bf184570e2da7663d62", "<KEY>", "be93491732a245da88bfec18a8f229ed", "1ae14d2048664274a63c5da923adedd8", "51a550d29ece48f0a5f22c60341dd1d7", "38badaec5dd343039269a7afa52e3c74", "0b3ad597080846ea9dabe115e9824a06", "<KEY>", "<KEY>", "<KEY>", "3e1f36326581486b8769b2adf2ac2f17", "<KEY>", "def7511564da4334a193a4df9aade18a", "<KEY>", "a237d4ad08ed4dc485a10effd038002b", "72f2638bce9d4f849adaf70acaf4a326", "348e019307914595aac584367f2c267b", "<KEY>", "<KEY>", "0228eeea8d78432e931d294b8261b9e7", "<KEY>", "<KEY>", "3eccd10383754ba8b9cdd9ad38c831e3", "<KEY>", "<KEY>", "<KEY>", "96fed45a09ab400894f8a1194680295f", "9ab303730d7c4ab2b127ee6b3ceb1e55", "a4467b3ed1f24eacb0ad07d9997a3dad", "<KEY>", "ecb37ef73fc24e4eb9da9f8718d1aaec", "b158a2d1a9b94c158245f3fed8ea41d5", "<KEY>", "f44614d7a4fe4af9af80700cb4273adb", "<KEY>", "650da115ba634669aa5d1ee1896660f4", "2cce602b8be742ce88c59aaa8e9c25f9", "<KEY>", "<KEY>", "<KEY>", "eca6540eb88f4905a3b06c5feece0ff1", "9dc2459ae69e4e759bdd5699e5cb9e8f", "67dbb1fa4c4c4fd790643a97d48e9ec6", "<KEY>", "<KEY>", "f45ad54204e1461e95089b157f9956ed", "ae9d6dda659249a08606253a5c76e92b", "<KEY>", "c32cc2a62b5244268a05ace1a481aef1", "<KEY>", "<KEY>", "160efa20088c4da1986c8b118964460b", "52a8b28b14824cec8e9fd8625ad02735", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "92f3d105371f4291be31ceb45a81ce2d", "3fba881cef054696b6ab6e2bc506f52c", "<KEY>", "8f37a1fdfd2c486a8817dc9e6ef87b27"]}
for epoch in tqdm(range(1, epochs+1)):
model.train() # Sending our model in Training mode
loss_train_total = 0 # Setting the training loss to zero initially
# Setting up the Progress bar to Moniter the progress of training
progress_bar = tqdm(dataloader_train, desc='Epoch {:1d}'.format(epoch), leave=False, disable=False)
for batch in progress_bar:
model.zero_grad() # As we not working with thew RNN's
# As our dataloader has '3' iteams so batches will be the Tuple of '3'
batch = tuple(b.to(device) for b in batch)
# INPUTS
# Pulling out the inputs in the form of dictionary
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
# OUTPUTS
outputs = model(**inputs) # '**' Unpacking the dictionary stright into the input
loss = outputs[0]
loss_train_total += loss.item()
loss.backward() # backpropagation
# Gradient Clipping -- Taking the Grad. & gives it a NORM value ~ 1
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
progress_bar.set_postfix({'training_loss': '{:.3f}'.format(loss.item()/len(batch))})
torch.save(model.state_dict(), f'finetuned_BERT_epoch_{epoch}.model')
tqdm.write(f'\nEpoch {epoch}')
loss_train_avg = loss_train_total/len(dataloader_train)
tqdm.write(f'Training loss: {loss_train_avg}')
val_loss, predictions, true_vals = evaluate(dataloader_validation)
val_f1 = f1_score_func(predictions, true_vals)
tqdm.write(f'Validation loss: {val_loss}')
tqdm.write(f'F1 Score (Weighted): {val_f1}')
# + [markdown] id="LjkW82FTCSU9"
# > __Gradient clipping__ is a technique to prevent __Exploding gradients__ in very deep networks, usually in recurrent neural networks -- This prevents any gradient to have norm greater than the threshold and thus the gradients are clipped.
# + [markdown] id="F3ju_W1nnebp"
# # __10. Loading finetuned BERT model and evaluate its performance__
# + id="foYSCdsJbeLl" outputId="2f3c4634-98aa-4531-b63c-4be817deef73" colab={"base_uri": "https://localhost:8080/"}
model = BertForSequenceClassification.from_pretrained("bert-base-uncased",
num_labels=len(label_dict),
output_attentions=False,
output_hidden_states=False)
model.to(device)
# + id="gzC3xAiMbgSM" outputId="bcda7f4e-2415-4d9e-9043-39517a615f5c" colab={"base_uri": "https://localhost:8080/", "height": 34}
model.load_state_dict(torch.load('/content/finetuned_BERT_epoch_10.model', map_location=torch.device('cpu')))
# + id="hSEaKqWybiSs" outputId="a95aa1fa-ce87-4b2c-8997-c376d95940fe" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["12a05e0e73384b4b808cf3c71e421492", "d1410b6189bc4b8ea0de298c4e05f467", "75787010208c41d4bd9f2030dd045943", "761d5de927404711b674baab03b138a1", "8700f708e5ad454c89e8592ee5119e07", "093671c6a19f4b60be5ffce45514b17b", "d2779d1dd58e4963b57450e966723a8e", "6489238102364e51b1351fe05ae2932d", "f4f64a98c7f342149bf6a51721bdf882", "ceefff46471340a18a9d6a05557b4579", "f325a98a05df41209327e6d47025d051"]}
_, predictions, true_vals = evaluate(dataloader_validation)
# + id="X0OBQg1ZbkKX" outputId="eda43adf-4120-4de9-fe29-f98f2a0dd6e4" colab={"base_uri": "https://localhost:8080/"}
accuracy_per_class(predictions, true_vals)
# + [markdown] id="fCeQOmNDbbEv"
# accuracy pred for finetuned_BERT_epoch_4.model
#
# ```
# Class: happy
# Accuracy: 168/171
#
# Class: not-relevant
# Accuracy: 16/32
#
# Class: angry
# Accuracy: 0/9
#
# Class: disgust
# Accuracy: 0/1
#
# Class: sad
# Accuracy: 0/5
#
# Class: surprise
# Accuracy: 0/5
# ```
#
#
# + [markdown] id="b6gwafXkb15g"
# # __11 Oth-Resources__
# + [markdown] id="8erP-JSCb9Bn"
#
#
# > 1. Paper: [Transformer](https://arxiv.org/abs/1706.03762)
#
# > 2. Paper: [BERT](https://arxiv.org/abs/1810.04805)
#
# 3. [Transformer Neural Networks - EXPLAINED!](https://youtu.be/TQQlZhbC5ps)
#
# 4. [BERT Neural Network - EXPLAINED!](https://youtu.be/xI0HHN5XKDo)
#
# 5. [HuggingFace documentation](https://huggingface.co/transformers/model_doc/bert.html)
#
# 6. [Hugging Face Write with Transformers](https://transformer.huggingface.co/)
#
# 7. [LSTM is dead. Long Live Transformers!](https://youtu.be/S27pHKBEp30)
#
# 8. [Hugging Face Releases New NLP ‘Tokenizers’ Library](https://www.analyticsvidhya.com/blog/2020/06/hugging-face-tokenizers-nlp-library/)
#
# 9. [Transfer Learning for NLP: Fine-Tuning BERT for Text Classification](https://www.analyticsvidhya.com/blog/2020/07/transfer-learning-for-nlp-fine-tuning-bert-for-text-classification/)
#
# 10. [Demystifying BERT: A Comprehensive Guide to the Groundbreaking NLP Framework](https://www.analyticsvidhya.com/blog/2019/09/demystifying-bert-groundbreaking-nlp-framework/)
#
# 11. [BERT Explained: State of the art language model for NLP](https://towardsdatascience.com/bert-explained-state-of-the-art-language-model-for-nlp-f8b21a9b6270)
#
# 12. [How do Transformers Work in NLP? A Guide to the Latest State-of-the-Art Models](https://www.analyticsvidhya.com/blog/2019/06/understanding-transformers-nlp-state-of-the-art-models/?utm_source=blog&utm_medium=demystifying-bert-groundbreaking-nlp-framework)
#
# 13. [BERT: Pre-Training of Transformers for Language Understanding](https://www.analyticsvidhya.com/blog/2019/09/demystifying-bert-groundbreaking-nlp-framework/)
#
# 14. [BERT Explained: A Complete Guide with Theory and Tutorial](https://towardsml.com/2019/09/17/bert-explained-a-complete-guide-with-theory-and-tutorial/)
#
# 15. [PyTorch_TDS](https://towardsdatascience.com/@theairbend3r)
#
| Ahmed_BERT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#using bs4 library for web scrapping and requests to extract data from URL
from bs4 import BeautifulSoup
import requests
r = requests.get("https://www.amazon.in/Redgear-Blaze-backlit-keyboard-aluminium/product-reviews/B073QQR2H2/ref=cm_cr_dp_d_show_all_btm?ie=UTF8&reviewerType=all_reviews")
print(r.url)
print(r.content)
# +
#By default we have to give HTML parser
soup = BeautifulSoup(r.text, 'html.parser')
#Use prettify to make the HTML code look better.
print(soup.prettify())
# +
Name = soup.findAll("span", {"class" : "a-profile-name"})
#from html code, there is span element with attribute class = a-profile-name
Reviewer = []
for i in range(2, len(Name)):
Reviewer.append(Name[i].get_text())
print(Reviewer)
# -
Title = soup.findAll("a", {"class" : "review-title-content"})
review_summary = []
for i in range(0, len(Title)):
review_summary.append(Title[i].get_text())
print(review_summary)
# +
#removing \n from all titles
#1. strip() :- This method is used to delete all the leading and trailing characters mentioned in its argument.
#2. lstrip() :- This method is used to delete all the leading characters mentioned in its argument.
review_summary[:] = [i.lstrip('\n').rstrip('\n') for i in review_summary]
print(review_summary)
# -
review_description = soup.findAll("span", {"class" : "review-text-content"})
Description = []
for i in range(0, len(review_description)):
Description.append(review_description[i].get_text())
print(Description)
# +
# We will remove the '\n' from before and after of every Review Desciption
Description[:] = [i.lstrip('\n').rstrip('\n') for i in Description]
print(Description)
# +
import pandas as pd
# -
#dataframes carry many additional useful functionalities at the cost of clarity and performance.
data = pd.DataFrame()
#adding info in dataframe
data["Title of Review"] = review_summary
data["Review by Customers"] = Description
data
| gamingkeyboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # INFO 3402 – Class 37: Analyzing and modelling autoregression
#
# [<NAME>, Ph.D.](http://brianckeegan.com/)
# [Assistant Professor, Department of Information Science](https://www.colorado.edu/cmci/people/information-science/brian-c-keegan)
# University of Colorado Boulder
#
# First things first, we need to install a new library: "fbprophet" to do some of the time series modeling later in the notebook. This takes a few minutes to install.
#
# **AT THE TERMINAL WINDOW**, run these two commands and agree to update when it requests:
#
# `conda update --all`
# `conda install -c conda-forge fbprophet`
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sb
import numpy as np
import pandas as pd
pd.options.display.max_columns = 200
import itertools
import statsmodels.formula.api as smf
import statsmodels.api as sm
from fbprophet import Prophet
# -
# We will return to the DIA passenger activity data we first explored back in Class 13 with data cleaning.
# +
dia_passengers = pd.read_csv('dia_passengers.csv',parse_dates=['date'])
dia_passengers.head()
# -
# Visualize the data.
# +
# Set up the plotting environment
f,ax = plt.subplots(1,1,figsize=(12,4))
# Put the "date" column as an index, access the remaining "passengers" column, and plot on the ax defined above
dia_passengers.set_index('date')['passengers'].plot(c='k',lw=3,ax=ax)
# Make a vertical red line on September 11, 2001
ax.axvline(pd.Timestamp('2001-09-11'),color='r',ls='--',lw=1)
# -
# Recall that statsmodels provides a variety of tools for "decomposing" a time-series into its seasonal, trend, and residual components.
# +
# This works best with a series having Timestamp/datetime objects as index
# So set the index to date and retrieve appropriate column to make a Series
decomposition = sm.tsa.seasonal_decompose(dia_passengers.set_index('date')['passengers'],model='additive')
plt.rcParams["figure.figsize"] = (8,8)
f = decomposition.plot()
f.axes[-1].axvline(pd.Timestamp('2001-09-11'),color='r',ls='--',lw=1)
# -
# Do the same feature engineering we did in Class 36.
# +
# Convert datetime/Timestamp objects into simpler floats
dia_passengers['months_since_opening'] = (dia_passengers['date'] - pd.Timestamp('1995-01-01'))/pd.Timedelta(1,'M')
# Get the rolling mean for trend modelling
dia_passengers['rolling_mean_passengers'] = dia_passengers['passengers'].rolling(12).mean()
# Extract month for fixed effects modelling
dia_passengers['month'] = dia_passengers['date'].apply(lambda x:x.month)
# Inspect
dia_passengers.tail()
# -
# Train the same linear regression models we used in Class 36.
# +
m0 = smf.ols('passengers ~ months_since_opening',data=dia_passengers).fit()
m1 = smf.ols('rolling_mean_passengers ~ months_since_opening',data=dia_passengers).fit()
m2 = smf.ols('passengers ~ months_since_opening + C(month)',data=dia_passengers).fit()
# -
# Create the DataFrame for predictions going forward in time.
# +
predict_passengers = pd.DataFrame({'date':pd.date_range('2017-01-01','2025-01-01',freq='M')},
index=range(263,263+96))
predict_passengers['months_since_opening'] = (predict_passengers['date'] - pd.Timestamp('1995-01-01'))/pd.Timedelta(1,'M')
predict_passengers['month'] = predict_passengers['date'].apply(lambda x:x.month)
# -
# Make the predictions.
# +
predict_passengers['m0'] = m0.predict({'months_since_opening':predict_passengers['months_since_opening']})
predict_passengers['m1'] = m1.predict({'months_since_opening':predict_passengers['months_since_opening']})
predict_passengers['m2'] = m2.predict({'months_since_opening':predict_passengers['months_since_opening'],
'month':predict_passengers['month']})
f,ax = plt.subplots(1,1,figsize=(8,6))
dia_passengers.plot(x='date',y='passengers',ax=ax,c='k',label='Observations',lw=3,alpha=.5)
predict_passengers.plot(x='date',y='m0',ax=ax,c='r',label='Simple',alpha=.5)
predict_passengers.plot(x='date',y='m1',ax=ax,c='g',label='Detrended',alpha=.5)
predict_passengers.plot(x='date',y='m2',ax=ax,c='b',label='Fixed effects',alpha=.5)
ax.set_xlim((pd.Timestamp('2012-01-01'),pd.Timestamp('2025-01-01')))
ax.set_ylim(3e6,7e6)
ax.legend(loc='center left',bbox_to_anchor=(1,.5))
ax.set_xlabel('Date')
ax.set_ylabel('Passengers');
# -
# ## Auto-correlation
#
# One of the key assumptions about regression we discussed in Classes 33 and 34 was the independence of observations. In many forms of data, this is a reasonable assumption: countries' behavior is independent from other countries, people's survey responses are independent from other people's, *etc*.
#
# This assumption breaks down with time series data where observations one day tend to be *strongly* correlated with observations for preceding and subsequent days: the weather yesterday is like the weather today, the price of a stock yesterday is like the price of a stock today, *etc*.
#
# Use pandas's `.shift()` method to create new columns that are shifted by a single or multiple rows so we can correlate passengers in one month with passenger values in adjacent months.
dia_passengers['passengers_shift_1'] = dia_passengers['passengers'].shift(1)
dia_passengers['passengers_shift_2'] = dia_passengers['passengers'].shift(2)
dia_passengers['passengers_shift_3'] = dia_passengers['passengers'].shift(3)
dia_passengers['passengers_shift_4'] = dia_passengers['passengers'].shift(4)
dia_passengers.head()
# Now correlate "passengers", "passengers_shift_1", "passengers_shift_2", *etc*. The correlations for "passengers" (first columns) at adjacent points in time is *extremely* strong: this is clearly a violation of the independence assumption because the values at different points in time are not independent but strongly correlated with the preceding and succeeding values.
# +
dia_passengers_corr = dia_passengers[['passengers','passengers_shift_1','passengers_shift_2','passengers_shift_3','passengers_shift_4']].corr()
# Using masking code from: https://seaborn.pydata.org/generated/seaborn.heatmap.html
dia_passengers_mask = np.zeros_like(dia_passengers_corr)
dia_passengers_mask[np.triu_indices_from(dia_passengers_mask)] = True
# Set up the plotting environment
f,ax = plt.subplots(1,1,figsize=(8,8))
# Make a heatmap
sb.heatmap(dia_passengers_corr,vmin=0,vmax=1,mask=dia_passengers_mask,annot=True,square=True,ax=ax,cmap='coolwarm_r')
# -
# This correlation in adjacent time points' values is called **[autocorrelation](https://en.wikipedia.org/wiki/Autocorrelation)**. Also note how even the shifted variables are also correlated with each other, we would need to control for the correlations in these other shifts/lags to recover the "true" correlation in the adjacent time series. **[partial autocorrelation](https://en.wikipedia.org/wiki/Partial_autocorrelation_function)** does exactly this.
#
# statsmodels has two functions in its `tsa` (time series analysis) class to plot out both the autocorrelation and partial autocorrelation in a time series. The x-axis is different lags (the shifting of different amounts we did above). The y-axis is the correlation values, identical to the "passengers" column in the heatmap above. These plots are called [correlograms](https://en.wikipedia.org/wiki/Correlogram).
#
# Passenger values are correlated at 0.87 at lag-1, 0.83 at lag-2, 0.77 as lag-3, and 0.72 at lag-4. In this case, we plotted out the correlations all the way to 50 lags with `plot_acf`. The blue curves are 95% confidence intervals, values *outside* this range are likely to be statistically significant correlations while values *within* this range cannot be distinguished from random noise.
#
# The *autocorrelation* is the simple correlation between values at different lags, but does not control for the fact that these correlations are correlated with other time lags: lag-1's correlation with lag-2 influences lag-1's correlation with lag-0, *etc*. The *partial autocorrelation* controls for these lagged values and recovers a more independent correlation value.
#
# The partial autocorrelations still show strong signals at 12 months, 24 months, *etc*. This captures the fact that monthly DIA activity in January one year is similar to January in the next year. In the figures below, I've marked lag-12, lag-24, *etc*. with dashed red lines.
# +
f,axs = plt.subplots(2,1,figsize=(8,8))
fig1 = sm.graphics.tsa.plot_acf(dia_passengers['passengers'],zero=False,lags=50,ax=axs[0],alpha=.05)
fig2 = sm.graphics.tsa.plot_pacf(dia_passengers['passengers'],zero=False,lags=50,ax=axs[1],alpha=.05)
for ax in axs:
ax.axvline(12,c='r',ls='--',lw=1)
ax.axvline(24,c='r',ls='--',lw=1)
ax.axvline(36,c='r',ls='--',lw=1)
ax.axvline(48,c='r',ls='--',lw=1)
# -
# pandas can also make an autocorrelation correlograms with `.plotting.autocorrelation_plot`. The dashed lines are the confidence interval where values above this are statistically significant correlation and unlikely to be noise (as above).
# +
f,ax = plt.subplots(1,1,figsize=(8,4))
ax = pd.plotting.autocorrelation_plot(dia_passengers['passengers'],ax=ax)
ax.set_xlim((1,50))
# -
# ## Prophet
#
# [Prophet](https://facebook.github.io/prophet/) is a time series forecasting tool developed (and obviously used by) data scientists at Facebook. There are a variety of more advanced statistical methods for modelling time series ([ARMA](https://en.wikipedia.org/wiki/Autoregressive%E2%80%93moving-average_model), [ARIMA](https://en.wikipedia.org/wiki/Autoregressive_integrated_moving_average), SARIMA, *etc*.) but these often involve doing data-drive grid search to find appropriate parameters or having deep understanding of the data or a theory to translate into parameters. Needless to say, these forecasting methods are outside the scope of the class.
#
# The reason we're looking at Prophet is because it does all of these (and more!), *mostly* automatically. Definitely check out the [documentation](https://facebook.github.io/prophet/) and examples in both R and Python.
#
# I've adapted the [Getting Started](https://facebook.github.io/prophet/docs/quick_start.html#python-api) documentation for our purposes here. Prophet wants a simple DataFrame containing only a column "ds" with timestamps and a column "y" with values for that date.
# +
# Copy the relevant columns to a new DataFrame
prophet_dia_passengers = dia_passengers[['date','passengers']].copy()
# Rename the columns how Prophet wants
prophet_dia_passengers.rename(columns={'date':'ds','passengers':'y'},inplace=True)
# Inspect
prophet_dia_passengers.head()
# -
# Fit the model. Since this data is at the monthly level, we do not need Prophet to estimate weekly or daily seasonality patterns. If you had data at the weekly or daily level, you probably would want it to estimate those. A few errors in red may be thrown and it should take a couple seconds for the estimation to finish.
m3 = Prophet(weekly_seasonality=False,daily_seasonality=False).fit(prophet_dia_passengers)
# We can use the `make_future_dataframe` method attached to the `m5` model we just estimated to make predictions into the future. Let's estimate a decade into the future, or 10 years \* 12 months/year = 120 months. Make sure to specify the "freq='M'" option, or it will default to daily estimates which doesn't make much sense for our monthly data.
# +
m3_future = m3.make_future_dataframe(periods=120,freq='M')
# Inspect the tail to make sure it's monthly
m3_future.tail()
# +
m3_forecast = m5.predict(m3_future)
# Lots of columns returned, just look at the choice ones
m3_forecast[['ds','yhat','yhat_lower','yhat_upper']].tail()
# -
# Inspect the predictions made by Prophet. This will include the historical data (black points), the line connecting these points, and the forecast into the future with errors.
# +
f,ax = plt.subplots(1,1,figsize=(10,5))
_ = m3.plot(m3_forecast,ax=ax)
# Always label your axes
ax.set_xlabel('Time')
ax.set_ylabel('Passengers')
ax.set_title('Observation and Forecast of DIA passengers')
# -
# You can also inspect the components of the Prophet model. It just estimates a trend and an annualy cyclical pattern.
f = m3.plot_components(m3_forecast)
# This annual cyclical variation should look similar to the pointplot we made in Class 34 for exploring the month-to-month differences that motivated our fixed effects model (`m2`).
f,ax = plt.subplots(1,1,figsize=(10,4))
sb.pointplot(x='month',y='passengers',data=dia_passengers,ax=ax)
# ## Make predictions
# +
# Make a range of dates, each month, going to June 1, 2028
future_date_range = pd.date_range('2017-01-01','2028-06-01',freq='M')
# Create an empty DataFrame for these dates, indexed from 263 going forward
predict_passengers = pd.DataFrame({'date':future_date_range},
index=range(263,263+len(future_date_range)))
# Feature engineer out the months since opening and month, like we did above
predict_passengers['months_since_opening'] = (predict_passengers['date'] - pd.Timestamp('1995-01-01'))/pd.Timedelta(1,'M')
predict_passengers['month'] = predict_passengers['date'].apply(lambda x:x.month)
predict_passengers.tail()
# -
# Use the `.predict` method on each of the statsmodels models and extract the "yhat" estimates from the Prophet model.
# +
# Store all the predictions in a dictionary, to be turned into a DataFrame shortly
predictions_d = {}
predictions_d['Observed'] = dia_passengers.loc[200:,'passengers']
predictions_d['Model 0'] = m0.predict({'months_since_opening':predict_passengers['months_since_opening']})
predictions_d['Model 1'] = m1.predict({'months_since_opening':predict_passengers['months_since_opening']})
predictions_d['Model 2'] = m2.predict({'months_since_opening':predict_passengers['months_since_opening'],
'month':predict_passengers['month']})
predictions_d['Model 3'] = m3_forecast['yhat']
# Turn dictionary into a DataFrame
predictions_df = pd.DataFrame(predictions_d)
predictions_df.head()
# -
# Visualize the predictions. Model 2 (fixed effects) and Model 3 (Prophet) look really similar and definitely capture the monthly cyclical pattern better than the simple regression models (Models 0 and 1).
# +
f,ax = plt.subplots(1,1,figsize=(8,6))
predictions_df.plot(lw=3,ax=ax)
ax.set_xlim((200,350))
ax.set_ylim((3e6,7e6))
ax.legend(loc='center left',bbox_to_anchor=(1,.5))
# Always label your axes
ax.set_xlabel('Months since opening')
ax.set_ylabel('Passengers')
# -
# Model 3 also made estimates going back in time. We can look at the performance of its predictions compared to the observed data by subtracting one from the other. The performance looks less good now: the Prophet model was off by more than 300,000 passengers (above *and* below) in many months.
#
# Over the course of a 30-day month, if an average of 10,000 more or fewer passengers were going through than you expected (about 50 planes worth of passengers each day), you might have either very angry passengers waiting in long lines or very angry merchants staffed for empty terminals.
# +
m3_performance = predictions_df['Model 3'] - predictions_df['Observed']
# Zoom in
ax = m3_performance.loc[180:300].plot(lw=3)
# Make a dashed line at 0, which would be perfect performance (no errors in prediction)
ax.axhline(0,c='k',ls='--',lw=1)
# Always label your axes
ax.set_xlabel("Months since opening")
ax.set_ylabel('Passengers difference')
ax.set_title('Model 3 performance compared to observations')
# -
# Models 2 and 3 are similar, but not identical. Model 3 has a clear trend of estimating more passengers than Model 2 going into the future. Over a decade, these differences between model predictions compound to a monthly difference of more than 300,000 passengers.
#
# Your manager or constituents would probably want to know whether they should plan for an extra quarter million passengers each month (or not) in the next decade to make appropriate investments.
# +
# Subtract Model 3 from Model 2
m2m3_prediction_diff = predictions_df['Model 2'] - predictions_df['Model 3']
# Make the plot, focus on future months by zooming in after 260
ax = m2m3_prediction_diff.loc[260:].plot(lw=3)
# Make a dashed line at 0, which would be perfect agreement between models
ax.axhline(0,c='k',ls='--',lw=1)
# Always label your axes
ax.set_xlabel("Months since opening")
ax.set_ylabel('Passengers difference')
ax.set_title('Difference in predictions, Models 2 and 3')
# -
| Module 07 - Extrapolating/Class 38 - Analyzing and modelling autoregression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Loading and using a trained model for Fashion
# Notebook demonstrating how to load a JointVAE model and use it for various things.
# +
from utils.load_model import load
path_to_model_folder = './trained_models/fashion/'
model = load(path_to_model_folder)
# -
# Print the latent distribution info
print(model.latent_spec)
# Print model architecture
print(model)
# +
from viz.visualize import Visualizer as Viz
# Create a Visualizer for the model
viz = Viz(model)
viz.save_images = False # Return tensors instead of saving images
# +
# %matplotlib inline
import matplotlib.pyplot as plt
samples = viz.samples()
plt.imshow(samples.numpy()[0, :, :], cmap='gray')
# -
traversals = viz.all_latent_traversals()
plt.imshow(traversals.numpy()[0, :, :], cmap='gray')
# Traverse 3rd continuous latent dimension across columns and first
# discrete latent dimension across rows
traversals = viz.latent_traversal_grid(cont_idx=2, cont_axis=1, disc_idx=0, disc_axis=0, size=(10, 10))
plt.imshow(traversals.numpy()[0, :, :], cmap='gray')
traversal = viz.latent_traversal_line(cont_idx=7, size=12)
plt.imshow(traversal.numpy()[0, :, :], cmap='gray')
# +
from utils.dataloaders import get_fashion_mnist_dataloaders
# Get MNIST test data
_, dataloader = get_fashion_mnist_dataloaders(batch_size=32)
# Extract a batch of data
for batch, labels in dataloader:
break
recon = viz.reconstructions(batch, size=(8, 8))
plt.imshow(recon.numpy()[0, :, :], cmap='gray')
# -
from torch.autograd import Variable
encodings = model.encode(Variable(batch))
# Continuous encodings for the first 5 examples
encodings['cont'][0][:5]
# Plot a grid of some traversals
from torchvision import datasets, transforms
from torch.utils.data import Dataset, DataLoader
import torch
path_to_data='../fashion_data'
all_transforms = transforms.Compose([
transforms.Resize(32),
transforms.ToTensor()
])
test_data = datasets.FashionMNIST(path_to_data, train=False,
transform=all_transforms)
test_loader = DataLoader(test_data, batch_size=10000, shuffle=True)
for data in test_loader:
pass
imges = data[0]
label = data[1]
latent_dist = model.encode(imges)
_, predict_label = torch.max(latent_dist['disc'][0], dim=1)
confusion = torch.zeros(10, 10)
for i in range(10000):
confusion[label[i].item(),predict_label[i].item()] += 1
for i in range(10):
confusion[i] = confusion[i] / confusion[i].sum()
# confusion = np.array([[0.9,0.1,0.0],[0.0,0.8,0.2],[0.1,0.7,0.2]])
# confusion = torch.tensor(confusion)
from matplotlib import cm
plt.imshow(confusion,interpolation='nearest',cmap=cm.Blues,aspect='auto',vmin=0,vmax=1.0)
value, predict_label = torch.max(confusion, dim=1)
list_price_positoin_address = []
seen = []
for i in predict_label:
if i in seen:
pass
else:
seen.append(i)
address_index = [x for x in range(len(predict_label)) if predict_label[x] == i]
list_price_positoin_address.append([i, address_index])
dict_address = dict(list_price_positoin_address)
print(dict_address)
for keys in dict_address.keys():
if(len(dict_address[keys])>1):
acc = confusion[dict_address[keys],keys.item()]
_, predict_label = torch.min(acc, dim=0)
confusion[dict_address[keys][predict_label.item()],keys.item()] = 0.0
value[dict_address[keys][predict_label.item()]], p = torch.max(confusion[dict_address[keys][predict_label.item()],:],dim=0)
value.mean()
| load_fasion.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// # Credit Card Transactions Fraud Detection Example:
//
// The notebook demonstrates how to develop a fraud detection application with the BigDL deep learning library on Apache Spark. We'll try to introduce some techniques that can be used for training a fraud detection model, but some advanced skills is not applicable since the dataset is highly simplified.
//
// **Dataset:**
// Credit Card Fraud Detection
// https://www.kaggle.com/dalpozz/creditcardfraud
//
// This dataset presents transactions that occurred in two days, where we got 492 frauds out of 284,807 transactions. The dataset is highly unbalanced, the positive class (frauds) account for 0.172% of all transactions.
//
// It contains only numerical input variables which are the result of a PCA transformation. Unfortunately, due to confidentiality issues, we cannot find the original features and more background information about the data. Features V1, V2, ... V28 are the principal components obtained with PCA, the only features which have not been transformed with PCA are 'Time' and 'Amount'. Feature 'Time' contains the seconds elapsed between each transaction and the first transaction in the dataset. The feature 'Amount' is the transaction Amount, this feature can be used for example-dependant cost-senstive learning. Feature 'Class' is the response variable and it takes value 1 in case of fraud and 0 otherwise.
//
// **Software stack:**
// Scala 2.11, Spark 2.0 or above, BigDL 0.3 or above
//
// **Contact:**
// <EMAIL>
//
// Loading data from csv files and output the schema:
// +
import org.apache.log4j.{Level, Logger}
import com.intel.analytics.bigdl.utils.Engine
import org.apache.spark.ml.Pipeline
import org.apache.spark.ml.linalg._
import org.apache.spark.ml.classification.{LogisticRegression, LogisticRegressionModel, MultilayerPerceptronClassifier, _}
import org.apache.spark.ml.evaluation.{BinaryClassificationEvaluator, MulticlassClassificationEvaluator}
import org.apache.spark.ml.feature.{MinMaxScaler, StandardScaler, VectorAssembler}
import org.apache.spark.ml.tuning.{ParamGridBuilder, TrainValidationSplit}
import org.apache.spark.mllib.evaluation.BinaryClassificationMetrics
import org.apache.spark.sql.{Row, SparkSession}
import org.apache.spark.sql.functions._
val conf = Engine.createSparkConf()
val spark = SparkSession.builder().master("local[1]").appName("Fraud Detection Example").config(conf).getOrCreate()
import spark.implicits._
Engine.init
val raw = spark.read.format("csv").option("header", "true").option("mode", "DROPMALFORMED").csv("data/creditcard.csv")
// cast all the column to Double type.
val df = raw.select(((1 to 28).map(i => "V" + i) ++ Array("Time", "Amount", "Class")).map(s => col(s).cast("Double")): _*)
println("num of records: " + df.count())
// select a few columns to show.
df.select("V1", "V2", "Time", "Amount", "Class").show()
println(" Class statistics: 1 represents fraud and 0 represents normal")
df.groupBy("Class").count().show()
// -
// **Feature analysis:**
//
// Normally it would improve the model if we could derive more features from the raw transaction records. E.g.
//
// days to last transaction,
// distance with last transaction,
// amount percentage over the last 1 month / 3months
// ...
//
// Yet with the public dataset, we can hardly derive any extention features from the PCA result. So here we only introduce several general practices:
//
// Usually there's a lot of categorical data in the raw dataset, E.g. post code, card type, merchandise id, seller id, etc.
// 1). For categorical feature with limited candidate values, like card type, channel id, just use OneHotEncoder.
// 2). For categorical feature with many candidate values, like merchandise id, post code or even phone number, suggest to use Weight of Evidence.
// 3). You can also use FeatureHasher from Spark MLlib which will be release with Spark 2.3.
//
// For this dataset, essentially it's a classification problem with highly unbalanced data set.
//
// ** Approach **
// 1. We will build a feature transform pipeline with Apache Spark and some of our transformers.
// 2. We will run some inital statistical analysis and split the dataset for training and validation.
// 3. We will build the model with BigDL.
// 4. We will compare different strategy to handle the unbalance.
//
// Details of each step is as follows:
//
// ***step 1. Build an inital pipeline for feature transform.***
//
// For each training records, we intend to aggregate all the features into one Spark Vector, which will then be sent to BigDL model for the training. First we'd like to introduce one handy transformer that we developed to help user build custom Transformers for Spark ML Pipeline.
//
//
//
// class FuncTransformer (
// override val uid: String,
// val func: UserDefinedFunction
// ) extends Transformer with HasInputCol with HasOutputCol with DefaultParamsWritable {
//
// `FuncTransformer` takes an udf as the constructor parameter and use the udf to perform the actual transform. The transformer can be saved/loaded as other transformer and can be integrated into a pipeline normally. It can be used widely in many use cases like conditional conversion(if...else...), , type conversion, to/from Array, to/from Vector and many string ops.
// Some examples:
//
// val labelConverter = new FuncTransformer(udf { i: Double => if (i >= 1) 1 else 0 })
//
// val shifter = new FuncTransformer(udf { i: Double => i + 1 })
//
// val toVector = new FuncTransformer(udf { i: Double => Vectors.dense(i) })
//
// We will use `VectorAssembler` to compose the all the Vx columns and append the Amount column. Then use `StandardScaler` to normlize the training records. Since in BigDL, the criterion generally only accepts 1, 2, 3... as the Label, so we will replace all the 0 with 2 in the training data.
//
//
//
// +
import com.intel.analytics.bigdl.nn._
import com.intel.analytics.bigdl.tensor.TensorNumericMath.TensorNumeric.NumericFloat
import org.apache.spark.ml.feature.{FuncTransformer, MinMaxScaler, StandardScaler, VectorAssembler}
import org.apache.spark.sql.functions._
// convert the label from {0, 1} to {1, 2}
val labelConverter = new FuncTransformer(udf {d: Double => if (d==0) 2 else d }).setInputCol("Class").setOutputCol("Class")
val assembler = new VectorAssembler().setInputCols((1 to 28).map(i => "V" + i).toArray ++ Array("Amount")).setOutputCol("assembled")
val scaler = new StandardScaler().setInputCol("assembled").setOutputCol("features")
val pipeline = new Pipeline().setStages(Array(assembler, scaler, labelConverter))
val pipelineModel = pipeline.fit(df)
val data = pipelineModel.transform(df)
println("Generate feature from raw data:")
data.select("features", "Class").show()
// -
// ***step 2. split the dataset into training and validation dataset.***
//
// Unlike some other training dataset, where the data does not have a time of occurance. For this case, we can know the sequence of the transactions from the Time column. Thus randomly splitting the data into training and validation does not make much sense, since in real world applications, we can only use the history transactions for training and use the latest transactions for validation. Thus we'll split the dataset according the time of occurance.
//
// get the time to split the data.
val splitTime = data.stat.approxQuantile("Time", Array(0.7), 0.001).head
val trainingData = data.filter(s"Time<$splitTime").cache()
val validData = data.filter(s"Time>=$splitTime").cache()
println("Split data into Training and Validation: ")
println("training records count: " + trainingData.count())
println("validation records count: " + validData.count())
// ***step 3. Build the model with BigDL***
//
// From the research community and industry feedback, a simple neural network turns out be the perfect candidate for the fraud detection training. We will quickly build a multiple layer Perceptron with linear layers.
// ```
// val bigDLModel = Sequential()
// .add(Linear(29, 10))
// .add(Linear(10, 2))
// .add(LogSoftMax())
// val criterion = ClassNLLCriterion()
// ```
// BigDL provides `DLEstimator` and `DLClassifier` for users with Apache Spark MLlib experience, which
// provides high level API for training a BigDL Model with the Apache Spark `Estimator`/`Transfomer`
// pattern, thus users can conveniently fit BigDL into a ML pipeline. The fitted model `DLModel` and
// `DLClassiferModel` contains the trained BigDL model and extends the Spark ML `Model` class.
// Alternatively users may also construct a `DLModel` with a pre-trained BigDL model to use it in
// Spark ML Pipeline for prediction.
//
// `DLClassifier` is a specialized `DLEstimator` that simplifies the data format for
// classification tasks. It only supports label column of DoubleType, and the fitted
// `DLClassifierModel` will have the prediction column of DoubleType.
//
// For this case we'll just use `DLClassifier` for the training. Note that users can set differet optimization mothod, batch size and epoch number.
//
//
// +
import com.intel.analytics.bigdl.nn._
import com.intel.analytics.bigdl.tensor.TensorNumericMath.TensorNumeric.NumericFloat
import com.intel.analytics.bigdl.utils.Engine
import org.apache.log4j.{Level, Logger}
import org.apache.spark.ml.ensemble.Bagging
import org.apache.spark.ml.evaluation.{BinaryClassificationEvaluator, MulticlassClassificationEvaluator}
import org.apache.spark.ml.feature.{FuncTransformer, MinMaxScaler, StandardScaler, VectorAssembler}
import org.apache.spark.ml.{DLClassifier, Pipeline}
import org.apache.spark.sql.functions._
import org.apache.spark.sql.{DataFrame, SparkSession}
val bigDLModel = Sequential().add(Linear(29, 10)).add(Linear(10, 2)).add(LogSoftMax())
val criterion = ClassNLLCriterion()
val dlClassifier = new DLClassifier(bigDLModel, criterion, Array(29)).setLabelCol("Class").setBatchSize(10000).setMaxEpoch(100)
val model = dlClassifier.fit(trainingData)
println("\ninitial model training finished.")
// -
// Now we have finished the training of our first model (which is certainly not the best, keep reading!).
//
// We'll need to think about how do evaluate the trained model:
//
// Given the class imbalance ratio, we recommend measuring the accuracy using the Area Under the Precision-Recall Curve (AUPRC). Confusion matrix accuracy is not meaningful for unbalanced classification. Since even if the model predicts all the records as normal transactions, it will still get an accuracy above 99%.
//
//
// +
val predictionDF = model.transform(validData)
// convert the prediction and label column back to {0, 1}
def evaluateModel(predictionDF: DataFrame): Unit = {
predictionDF.cache()
val labelConverter2 = new FuncTransformer(udf {d: Double => if (d==2) 0 else d }).setInputCol("Class").setOutputCol("Class")
val labelConverter3 = new FuncTransformer(udf {d: Double => if (d==2) 0 else d }).setInputCol("prediction").setOutputCol("prediction")
val finalData = labelConverter2.transform(labelConverter3.transform(predictionDF))
val metrics = new BinaryClassificationEvaluator().setRawPredictionCol("prediction").setLabelCol("Class")
val auPRC = metrics.evaluate(finalData)
println("\nArea under precision-recall curve: = " + auPRC)
val recall = new MulticlassClassificationEvaluator().setLabelCol("Class").setMetricName("weightedRecall").evaluate(finalData)
println("\nrecall = " + recall)
val precisoin = new MulticlassClassificationEvaluator().setLabelCol("Class").setMetricName("weightedPrecision").evaluate(finalData)
println("\nPrecision = " + precisoin)
predictionDF.unpersist()
}
evaluateModel(predictionDF)
// -
// To this point, we have finished the training and evaluation with a simple BigDL model. We can see that even though the recall and precision are high, the area under precision-recall curve is not optimistic. That's because we haven't really apply any technique to handle the imbalanced training data.
//
//
// Next we'll try to optimize the training process.
// ***step 4. handle the data imbalance***
//
// There are several ways to approach this classification problem taking into consideration this unbalance.
//
// 1. Collect more data? Nice strategy but not applicable in this case.
//
// 2. Resampling the dataset
// Essentially this is a method that will process the data to have an approximate 50-50 ratio.
// One way to achieve this is by OVER-sampling, which is adding copies of the under-represented class (better when there're little data)
// Another is UNDER-sampling, which deletes instances from the over-represented class (better when there are lots of data)
//
// 3. Apart from under and over sampling, there is a very popular approach called SMOTE (Synthetic Minority Over-Sampling Technique), which is a combination of oversampling and undersampling, but the oversampling approach is not by replicating minority class but constructing new minority class data instance via an algorithm.
//
// We'll start with Resampling.
//
// Since there're 492 frauds out of 284,807 transactions, to build a reasonable training dataset, we'll use UNDER-sampling for normal transactions and use OVER-sampling for fraud transactions. By using the sampling rate as
// fraud -> 10, normal -> 0.05, we can get a training dataset of (5K fraud + 14K normal) transactions. We can use the training data to fit a model.
//
// Yet we'll soon find that since there're only 5% of all the normal transactions are included in the training data, the model can only cover 5% of all the normal transactions, which is obviousely not optimistic. So how can we get a better converage for the normal transactions without breaking the ideal ratio in the training dataset?
//
// An immediate improvement would be to train multiple models. For each model, we will run the resampling from the original dataset and get a new training data set. After training, we can select best voting strategy for all the models to make the prediction.
//
// We'll use Ensembling of neural networks. That's where a Bagging classifier becomes handy. Bagging is an Estimator we developed for ensembling of multiple other Estimator.
//
// ```
// package org.apache.spark.ml.ensemble
//
// class Bagging[M <: Model[M]](override val uid: String)
// extends Estimator[BaggingModel[M]]
// with BaggingParams[M] {
// ```
//
// For usage, user need to set the specific Estimator to use and the number of models to be trained:
//
// ```
// val estimator = new Bagging()
// .setPredictor(dlClassifier)
// .setLabelCol("Class")
// .setIsClassifier(true)
// .setNumModels(10)
// ```
//
// Internally, Bagging will train $(numModels) models. Each model is trained with the resampled data from the original dataset.
//
// +
// Train with Bagging model, takes 5-10 minutes
val bigDLModel = Sequential().add(Linear(29, 10)).add(Linear(10, 2)).add(LogSoftMax())
val criterion = ClassNLLCriterion()
val dlClassifier = new DLClassifier(bigDLModel, criterion, Array(29)).setLabelCol("Class").setBatchSize(10000).setMaxEpoch(100)
val estimator = new Bagging().setPredictor(dlClassifier).setLabelCol("Class").setIsClassifier(true).setNumModels(20)
val baggingModel = estimator.fit(trainingData)
// -
// After fitting, we can tune the voting strategy via `model.setThreshold(t)`. When using Threshold = 30, we can get an improved model.
evaluateModel(baggingModel.setThreshold(30).transform(validData))
// By using bagging model, we can increase the AUPRC from 0.78 to 0.93.
//
// The code used in this notebook can be found from https://github.com/intel-analytics/analytics-zoo/tree/master/pipeline/fraudDetection.
//
//
| apps/fraudDetection/Fraud Detction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ALENN - Demo Notebook
# ## Quickstart Guide
#
# <NAME>
# <br>
# Mathematical Institute, University of Oxford
# <br>
# Institute for New Economic Thinking at the Oxford Martin School
# <br>
# <br>
# Copyright (c) 2020, University of Oxford. All rights reserved.
# <br>
# Distributed under a BSD 3-Clause licence. See the accompanying LICENCE file for further details.
# # Overview
# This notebook provides, through the use of a simple illustrative example, a complete tutorial on the use of the ALENN package to perform Bayesian estimation for economic simulation models using the neural network-based approach introduced by Platt (2021) in the paper *[Bayesian Estimation of Economic Simulation Models Using Neural Networks](https://link.springer.com/article/10.1007/s10614-021-10095-9)*. In general, the workflow presented here should require minimal adjustment (changing the model function, empirical dataset, priors, and sampler settings) in order to be applied to new examples.
# # Step 1
# ## Importing of Packages
# As a natural starting point, we begin by importing any required Python packages. With the exception of ALENN, which we assume has already been installed as per the instructions provided in the accompanying README file, all other imported libraries are now fairly standard in most data science workflows.
# +
# Import the ALENN ABM Estimation Package
import alenn
# Import Plotting Libraries
import matplotlib.pyplot as plt
# Import Numerical Computation Libraries
import numpy as np
import pandas as pd
# Import General Mathematical Libraries
from scipy import stats
# Import Data Storage Libraries
import pickle as pkl
# Import System Libraries
import os
import logging
# +
# Disable Tensorflow Deprecation Warnings
logging.disable(logging.WARNING)
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
# Tensorflow 2.x deprecates many Tensorflow 1.x methods, causing Tensorflow 1.15.0 to output a large number
# of (harmless) deprecation warnings when performing the first likelihood calculation. This can be very
# distracting, leading us to disable them.
# -
# # Step 2
# ## Creating the Likelihood and Posterior Estimator Object
# The primary functionally of ALENN is implemented in the `MDNPosterior` class, which contains all the methods required to estimate the likelihood and posterior. It thus follows that the first step in the estimation pipeline is creating an `MDNPosterior` object by calling its constructor method, `alenn.mdn.MDNPosterior`.
# <br>
# <br>
# If no arguments are provided to the constructor, the default neural network architecture introduced in the paper is used. If an alternative is required, however, this can easily be specified through the use of keyword arguments. As an example, increasing the number of lags to 4 and decreasing the number of hidden layers to 2 could be achieved by calling `alenn.mdn.MDNPosterior(num_lags = 4, num_layers = 2)`. Further details can be obtained by consulting the class docstring:
# ```python
# # ?alenn.mdn.MDNPosterior
# ```
# Create an MDN Posterior Approximator Object (Uses Default Settings from the Paper)
posterior = alenn.mdn.MDNPosterior()
# # Step 3
# ## Specifying the Candidate Model
# At this stage, all we have done is defined a generic posterior estimator object. In order to actually apply the estimator to a given problem, we need to provide the object with additional information. We begin with the candidate model.
# <br>
# <br>
# From the perspective of ALENN, the model is a black box capable of producing simulated time series data. Therefore, the candidate model is provided to ALENN in the form of a function that takes in a 1-d numpy array or list of parameter values and returns a model output matrix as a 2-d numpy array. Ensuring that the model is correctly specified and matches ALENN's input, processing, and output requirements is perhaps the most critical part of this process and should therefore be approached with care.
# <br>
# <br>
# To elaborate, the model function should take, as input, a 1-d numpy array, $\mathbf{\theta}$, containing values for each of the model's free parameters (those that should be estimated). The function should then proceed to generate a corresponding set of $R$ model Monte Carlo replications. Each of these replications is a single time series of length $T_{sim}$ generated by the model for the same set of parameter values as the remaining replications, $\mathbf{\theta}$, but a different random seed, $i$. Once generated, each replication should be stored as a single column in a $T_{sim} \times R$ numpy array that is returned as the final output by the model function.
# <br>
# <br>
# It is important to note that, although the choice of seed for each replication is arbitrary, the same set of seeds must be used throughout the entire estimation experiment, i.e. the model function should always use the same set of seeds, regardless of the value of $\mathbf{\theta}$ at which the function is evaluated. Footnote 44 in the paper provides a more detailed discussion. Additionally, in most practical examples, the generation of simulated data using the candidate model is likely to be computationally-expensive and thus a bottleneck in the inference process. We therefore suggest that, if the model is costly to simulate, that the model function should generate the replications in parallel.
# <br>
# <br>
# Finally, as suggested by the model function output structure introduced above, this version of ALENN currently only supports univariate time series model outputs. Note, however, that the methodology itself is generally applicable to multivariate outputs and a multivariate extension to this library is likely to be released in the near future.
# +
# Specify the Simulated Data Characteristics
T_sim = 1000 # Length of each Monte Carlo replication
R = 100 # Number of Monte Carlo replications
seed_set = 7 # The set of seeds associated with the model replications
# In most cases, we suggest that either (T_sim = 1000 and R = 100) or (T_sim = 2000 and R = 50) be considered.
# The seed_set variable can be interpreted as defining an arbitrary set of 100 random seeds.
# Define the Candidate Model Function
def model(theta):
return np.diff(alenn.models.random_walk(700, 0.4, 0.5, theta[0], theta[1], T_sim, R, seed_set), axis = 0)
# Add the Model Function to the MDNPosterior Object
posterior.set_model(model)
# In the above, we have selected the random walk examined in the paper's comparative experiments. This model,
# along with the other models considered in the paper, are implemented as part of ALENN and can be accessed via
# alenn.models as above (see the corresponding file for more details).
#
# In this case, we are attempting to estimate the pre- and post-break volatility and have fixed all other parameters
# to their default values. Notice that we also consider the series of first differences to induce stationarity.
# While stationarity is not an assumption of the methodology, it may be advantageous to consider stationarity
# transformations if a given non-stationary model proves to be difficult to estimate.
# -
# # Step 4
# ## Specifying the Model Priors
# As in any Bayesian exercise, we must specify a prior over the model parameters. In ALENN, the prior is specified
# in the form of a special data structure. A prior function must be defined separately for each free parameter and each function of this type should take in a single value for that parameter and return a corresponding prior density value. These functions should be stored in a Python list.
# <br>
# <br>
# In all cases, the order of the density functions in the prior list must correspond to the order in which the parameters are passed to the model function. More concretely, if the model function takes in values for parameters $[\sigma_1, \sigma_2]$, the prior list must have form $[p(\sigma_1), p(\sigma_2)]$.
# +
# Define Parameter Priors
priors = [stats.uniform(loc = 0, scale = 10).pdf,
stats.uniform(loc = 0, scale = 10).pdf]
# Add the Model Priors to the MDNPosterior Object
posterior.set_prior(priors)
# In the above, we have defined uniform priors over [0, 10] for both the pre- and post-break volatility. In most
# applications, we recommend that users make use of SciPy's stats module to define the priors, as we have. This
# results in greater readability and can help avoid errors in the prior specification.
# -
# # Step 5
# ## Loading the Empirical Data
# To complete the problem specification, we are finally required to provide the `MDNPosterior` object with a set of empirical data. This process is rather straightforward and simply requires that the data be provided in the form of a 1-d numpy array.
# <br>
# <br>
# While longer empirical time series are always preferred if available, we typically consider $T_{emp} = 1000$ for problems involving $1-4$ free parameters and $T_{emp} = 2000$ for problems involving $5-10$ free parameters. In many cases, however, we suspect that a significant reduction in the number of data points would be viable, particularly when the data provides a reasonable level of information regarding the model parameters.
# +
# Load the Empirical Data
with open('data/Demo_Data', 'rb') as f:
empirical = pkl.load(f)
# Add the Empirical Data to the MDNPosterior Object
posterior.load_data(empirical)
# The empirical data loaded above is a synthetic series of 999 (first-differenced) observations generated by the
# random walk model when initialised using the parameter values associated with the first free parameter set
# introduced in the paper's comparative exercises. Our exercise here can thus be seen as a replication of the
# associated comparative experiment.
#
# In a true empirical application, this series would simply be replaced by a series measured from the actual
# real-world system being modelled.
# -
# # Step 6
# ## Sampling the Posterior
# With the `MDNPosterior` object now completely specified, we are able to evaluate the posterior for arbitrary values of $\mathbf{\theta}$ and hence sample it using MCMC. As discussed in detail in Appendix 2, we make use of the adaptive Metropolis-Hastings algorithm proposed by Griffin and Walker (2013).
# <br>
# <br>
# As in the case of the posterior, the sampler is also implemented as an object, in this case being an instantiation of the `AdaptiveMCMC` class. In order to perform the sampling procedure, a number of key components must be specified and passed to the object. These include:
# * Parameter ranges over which to conduct the initial sweep of the parameter space. This is specified in the form of two 1-d numpy arrays that contain, in the same order as is associated with the list of priors discussed in Step 4, the lower and upper bounds for each parameter respectively.
# * The desired number of samples per sample set. In general, we recommend that this is set to $K = 70$.
# * The desired number of sample sets to be generated. As a rule of thumb, we suggest generating $S = 5000$ sets for problems involving $1 - 4$ free parameters and $15000$ sets for problems involving $5 - 10$ free parameters. Of course, common convergence diagnostics, such as Galman and Ruben's R, could certainly be used to ensure that a sufficient number of samples has been generated.
# +
# Create an Adaptive MCMC Sampler Object
sampler = alenn.mcmc.AdaptiveMCMC(K = 70, S = 5000)
# Define the Parameter Bounds
theta_lower = np.array([0, 0])
theta_upper = np.array([10, 10])
# Add the Posterior Approximator and Parameter Ranges to the Newly-created Object
sampler.set_posterior(posterior)
sampler.set_initialisation_ranges(theta_lower, theta_upper)
# Please note that the set_posterior method must be called before the set_initialisation_ranges method.
# Initiate the Sampling Process
sampler.sample_posterior()
# -
# # Step 7
# ## Processing the Obtained Samples
# Once the sampling procedure has concluded, all that remains is the processing of the obtained samples into meaningful outputs, i.e. tables or marginal posterior plots.
# <br>
# <br>
# The aforementioned samples may be extracted from the `AdaptiveMCMC` object using the `process_samples` method, which requires the specification of a single integer argument, `burn_in`. This argument specifies the number of sample sets that should be discarded as part of an initial burning-in period, as is standard in all MCMC algorithms, and we typically recommend burning-in periods of $1500-2500$ sample sets for $S = 5000$ and $7500-10000$ sample sets for $S = 15000$. Of course, some problems may require alternative configurations depending on their associated convergence rates and we therefore recommend that multiple chains be generated by repeating Step 6 several times in order to diagnose convergence when applying the methodology.
# <br>
# <br>
# The `process_samples` method returns the obtained samples in the form of a 2-d numpy array, where each column represents the posterior samples obtained for a given parameter, with the columns following the same parameter order as the original model function. The method output also contains a final, extra column consisting of the the associated log-likelihood samples.
# +
# Result Table
# Note that we illustrate the construction of a result table for a single chain, whereas the corresponding result
# in Section 4.1 is associated with 5 chains.
# Process the Sampler Output
samples = sampler.process_samples(burn_in = 2500)
# Calculate the Posterior Mean
pos_mean = samples[:, :posterior.num_param].mean(axis = 0)
# Calculate the Posterior Standard Deviation
pos_std = samples[:, :posterior.num_param].std(axis = 0)
# Construct a Result Table
result_table = pd.DataFrame(np.array([pos_mean, pos_std]).transpose(), columns = ['Posterior Mean', 'Posterior Std. Dev.'])
result_table.index.name = 'Parameter'
result_table.index += 1
# Display the Result Table
print('Final Estimation Results:')
print('')
print(result_table)
# +
# Marginal Posterior Plots
# Note that we illustrate the construction of marginal posterior plots for a single chain, whereas the corresponding
# result in Section 4.1 is associated with 5 chains.
# Process the Sampler Output
samples = sampler.process_samples(burn_in = 2500)
# Set the Parameter Names
param_names = [r'$\sigma_1$', r'$\sigma_2$']
# Set-Up the Figure
fig = plt.figure(figsize = (5 * posterior.num_param, 5))
# Loop Over the Free Parameters
for i in range(posterior.num_param):
# Plot the Posterior Histogram
plt.subplot(1, posterior.num_param, i + 1)
plt.hist(samples[:, i], 25, density = True, color = 'b', alpha = 0.5)
# Plot the Prior Density
prior_range = np.linspace(samples[:, i].min() * 0.9, samples[:, i].max() * 1.1, 100)
plt.plot(prior_range, [priors[i](x) for x in prior_range], color = 'r', alpha = 0.75)
# Note that we are only plotting the prior for a limited range such that it extends only slightly
# beyond the posterior. This is done to improve the clarity of presentation. In reality, the prior is
# substantially wider than the posterior and would extend from 0 to 10 for this example.
# Plot the Posterior Mean
plt.axvline(x = samples[:, i].mean(), c = 'k', linestyle = 'dashed', alpha = 0.75)
# Label the Plot
plt.xlabel(param_names[i])
plt.ylabel(r'$p($' + param_names[i] + r'$)$')
plt.legend(['Prior Density', 'Posterior Mean', 'Posterior Density'], fontsize = 8)
# Set the Figure Layout
plt.tight_layout()
# Display the Figure
plt.show()
| demo/ALENN_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 自动求导
# 这次课程我们会了解 PyTorch 中的自动求导机制,自动求导是 PyTorch 中非常重要的特性,能够让我们避免手动去计算非常复杂的导数,这能够极大地减少了我们构建模型的时间,这也是其前身 Torch 这个框架所不具备的特性,下面我们通过例子看看 PyTorch 自动求导的独特魅力以及探究自动求导的更多用法。
import torch
from torch.autograd import Variable
# ## 简单情况的自动求导
# 下面我们显示一些简单情况的自动求导,"简单"体现在计算的结果都是标量,也就是一个数,我们对这个标量进行自动求导。
x = Variable(torch.Tensor([2]), requires_grad=True)
y = x + 2
z = y ** 2 + 3
print(z)
# 通过上面的一些列操作,我们从 x 得到了最后的结果out,我们可以将其表示为数学公式
#
# $$
# z = (x + 2)^2 + 3
# $$
#
# 那么我们从 z 对 x 求导的结果就是
#
# $$
# \frac{\partial z}{\partial x} = 2 (x + 2) = 2 (2 + 2) = 8
# $$
# 如果你对求导不熟悉,可以查看以下[网址进行复习](https://baike.baidu.com/item/%E5%AF%BC%E6%95%B0#1)
# 使用自动求导
z.backward()
print(x.grad)
# 对于上面这样一个简单的例子,我们验证了自动求导,同时可以发现发现使用自动求导非常方便。如果是一个更加复杂的例子,那么手动求导就会显得非常的麻烦,所以自动求导的机制能够帮助我们省去麻烦的数学计算,下面我们可以看一个更加复杂的例子。
# +
x = Variable(torch.randn(10, 20), requires_grad=True)
y = Variable(torch.randn(10, 5), requires_grad=True)
w = Variable(torch.randn(20, 5), requires_grad=True)
out = torch.mean(y - torch.matmul(x, w)) # torch.matmul 是做矩阵乘法
out.backward()
# -
# 如果你对矩阵乘法不熟悉,可以查看下面的[网址进行复习](https://baike.baidu.com/item/%E7%9F%A9%E9%98%B5%E4%B9%98%E6%B3%95/5446029?fr=aladdin)
# 得到 x 的梯度
print(x.grad)
# 得到 y 的的梯度
print(y.grad)
# 得到 w 的梯度
print(w.grad)
# 上面数学公式就更加复杂,矩阵乘法之后对两个矩阵对应元素相乘,然后所有元素求平均,有兴趣的同学可以手动去计算一下梯度,使用 PyTorch 的自动求导,我们能够非常容易得到 x, y 和 w 的导数,因为深度学习中充满大量的矩阵运算,所以我们没有办法手动去求这些导数,有了自动求导能够非常方便地解决网络更新的问题。
#
#
# ## 复杂情况的自动求导
# 上面我们展示了简单情况下的自动求导,都是对标量进行自动求导,可能你会有一个疑问,如何对一个向量或者矩阵自动求导了呢?感兴趣的同学可以自己先去尝试一下,下面我们会介绍对多维数组的自动求导机制。
m = Variable(torch.FloatTensor([[2, 3]]), requires_grad=True) # 构建一个 1 x 2 的矩阵
n = Variable(torch.zeros(1, 2)) # 构建一个相同大小的 0 矩阵
print(m)
print(n)
# 通过 m 中的值计算新的 n 中的值
n[0, 0] = m[0, 0] ** 2
n[0, 1] = m[0, 1] ** 3
print(n)
# 将上面的式子写成数学公式,可以得到
# $$
# n = (n_0,\ n_1) = (m_0^2,\ m_1^3) = (2^2,\ 3^3)
# $$
# 下面我们直接对 n 进行反向传播,也就是求 n 对 m 的导数。
#
# 这时我们需要明确这个导数的定义,即如何定义
#
# $$
# \frac{\partial n}{\partial m} = \frac{\partial (n_0,\ n_1)}{\partial (m_0,\ m_1)}
# $$
#
# 在 PyTorch 中,如果要调用自动求导,需要往`backward()`中传入一个参数,这个参数的形状和 n 一样大,比如是 $(w_0,\ w_1)$,那么自动求导的结果就是:
# $$
# \frac{\partial n}{\partial m_0} = w_0 \frac{\partial n_0}{\partial m_0} + w_1 \frac{\partial n_1}{\partial m_0}
# $$
# $$
# \frac{\partial n}{\partial m_1} = w_0 \frac{\partial n_0}{\partial m_1} + w_1 \frac{\partial n_1}{\partial m_1}
# $$
n.backward(torch.ones_like(n)) # 将 (w0, w1) 取成 (1, 1)
print(m.grad)
# 通过自动求导我们得到了梯度是 4 和 27,我们可以验算一下
# $$
# \frac{\partial n}{\partial m_0} = w_0 \frac{\partial n_0}{\partial m_0} + w_1 \frac{\partial n_1}{\partial m_0} = 2 m_0 + 0 = 2 \times 2 = 4
# $$
# $$
# \frac{\partial n}{\partial m_1} = w_0 \frac{\partial n_0}{\partial m_1} + w_1 \frac{\partial n_1}{\partial m_1} = 0 + 3 m_1^2 = 3 \times 3^2 = 27
# $$
# 通过验算我们可以得到相同的结果
#
#
# ## 多次自动求导
# 通过调用 backward 我们可以进行一次自动求导,如果我们再调用一次 backward,会发现程序报错,没有办法再做一次。这是因为 PyTorch 默认做完一次自动求导之后,计算图就被丢弃了,所以两次自动求导需要手动设置一个东西,我们通过下面的小例子来说明。
x = Variable(torch.FloatTensor([3]), requires_grad=True)
y = x * 2 + x ** 2 + 3
print(y)
y.backward(retain_graph=True) # 设置 retain_graph 为 True 来保留计算图
print(x.grad)
y.backward() # 再做一次自动求导,这次不保留计算图
print(x.grad)
# 可以发现 x 的梯度变成了 16,因为这里做了两次自动求导,所以讲第一次的梯度 8 和第二次的梯度 8 加起来得到了 16 的结果。
#
#
# **小练习**
#
# 定义
#
# $$
# x =
# \left[
# \begin{matrix}
# x_0 \\
# x_1
# \end{matrix}
# \right] =
# \left[
# \begin{matrix}
# 2 \\
# 3
# \end{matrix}
# \right]
# $$
#
# $$
# k = (k_0,\ k_1) = (x_0^2 + 3 x_1,\ 2 x_0 + x_1^2)
# $$
#
# 我们希望求得
#
# $$
# j = \left[
# \begin{matrix}
# \frac{\partial k_0}{\partial x_0} & \frac{\partial k_0}{\partial x_1} \\
# \frac{\partial k_1}{\partial x_0} & \frac{\partial k_1}{\partial x_1}
# \end{matrix}
# \right]
# $$
#
# 参考答案:
#
# $$
# \left[
# \begin{matrix}
# 4 & 3 \\
# 2 & 6 \\
# \end{matrix}
# \right]
# $$
# +
x = Variable(torch.FloatTensor([2, 3]), requires_grad=True)
k = Variable(torch.zeros(2))
k[0] = x[0] ** 2 + 3 * x[1]
k[1] = x[1] ** 2 + 2 * x[0]
# -
print(k)
# +
j = torch.zeros(2, 2)
k.backward(torch.FloatTensor([1, 0]), retain_graph=True)
j[0] = x.grad.data
x.grad.data.zero_() # 归零之前求得的梯度
k.backward(torch.FloatTensor([0, 1]))
j[1] = x.grad.data
# -
print(j)
# 下一次课我们会介绍两种神经网络的编程方式,动态图编程和静态图编程
| 03.PyTorch基础(入门)/autograd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
def intersect(list1, list2):
return [sample for sample in list1 if sample in list2]
# the true positives and negatives:
actual_positive = [2, 5, 6, 7, 8, 10, 18, 21, 24, 25, 29, 30, 32, 33, 38, 39, 42, 44, 45, 47]
actual_negative = [1, 3, 4, 9, 11, 12, 13, 14, 15, 16, 17, 19, 20, 22, 23, 26, 27, 28, 31, 34, 35, 36, 37, 40, 41, 43, 46, 48, 49]
# the positives and negatives we determine by running the experiment:
experimental_positive = [2, 4, 5, 7, 8, 9, 10, 11, 13, 15, 16, 17, 18, 19, 20, 21, 22, 24, 26, 27, 28, 32, 35, 36, 38, 39, 40, 45, 46, 49]
experimental_negative = [1, 3, 6, 12, 14, 23, 25, 29, 30, 31, 33, 34, 37, 41, 42, 43, 44, 47, 48]
#define type_i_errors and type_ii_errors here
type_i_errors = intersect(experimental_positive,actual_negative )
type_ii_errors = intersect(experimental_negative,actual_positive)
# -
# This is the error for the false positive
type_i_errors
# This is the error for the false negative
type_ii_errors
# +
def reject_null_hypothesis(p_value):
"""
Returns the truthiness of whether the null hypothesis can be rejected
Takes a p-value as its input and assumes p <= 0.05 is significant
"""
if p_value < 0.05:
res = True
else:
res = False
return res
hypothesis_tests = [0.1, 0.009, 0.051, 0.012, 0.37, 0.6, 0.11, 0.025, 0.0499, 0.0001]
for p_value in hypothesis_tests:
reject_null_hypothesis(p_value)
# -
| A-B Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: seaborn-refactor (py38)
# language: python
# name: seaborn-refactor
# ---
# Plot a univariate distribution along the x axis:
import seaborn as sns; sns.set()
tips = sns.load_dataset("tips")
sns.kdeplot(data=tips, x="total_bill")
# Flip the plot by assigning the data variable to the y axis:
sns.kdeplot(data=tips, y="total_bill")
# Plot distributions for each column of a wide-form dataset:
iris = sns.load_dataset("iris")
sns.kdeplot(data=iris)
# Use less smoothing:
sns.kdeplot(data=tips, x="total_bill", bw_adjust=.2)
# Use more smoothing, but don't smooth past the extreme data points:
ax= sns.kdeplot(data=tips, x="total_bill", bw_adjust=5, cut=0)
# Plot conditional distributions with hue mapping of a second variable:
sns.kdeplot(data=tips, x="total_bill", hue="time")
# "Stack" the conditional distributions:
sns.kdeplot(data=tips, x="total_bill", hue="time", multiple="stack")
# Normalize the stacked distribution at each value in the grid:
sns.kdeplot(data=tips, x="total_bill", hue="time", multiple="fill")
# Estimate the cumulative distribution function(s), normalizing each subset:
sns.kdeplot(
data=tips, x="total_bill", hue="time",
cumulative=True, common_norm=False, common_grid=True,
)
# Estimate distribution from aggregated data, using weights:
tips_agg = (tips
.groupby("size")
.agg(total_bill=("total_bill", "mean"), n=("total_bill", "count"))
)
sns.kdeplot(data=tips_agg, x="total_bill", weights="n")
# Map the data variable with log scaling:
diamonds = sns.load_dataset("diamonds")
sns.kdeplot(data=diamonds, x="price", log_scale=True)
# Use numeric hue mapping:
sns.kdeplot(data=tips, x="total_bill", hue="size")
# Modify the appearance of the plot:
sns.kdeplot(
data=tips, x="total_bill", hue="size",
fill=True, common_norm=False, palette="viridis",
alpha=.5, linewidth=0,
)
# Plot a bivariate distribution:
geyser = sns.load_dataset("geyser")
sns.kdeplot(data=geyser, x="waiting", y="duration")
# Map a third variable with a hue semantic to show conditional distributions:
sns.kdeplot(data=geyser, x="waiting", y="duration", hue="kind")
# Show filled contours:
sns.kdeplot(
data=geyser, x="waiting", y="duration", hue="kind", fill=True,
)
# Show fewer contour levels, covering less of the distribution:
sns.kdeplot(
data=geyser, x="waiting", y="duration", hue="kind",
levels=5, thresh=.2,
)
# Fill the axes extent with a smooth distribution, using a different colormap:
sns.kdeplot(
data=geyser, x="waiting", y="duration",
fill=True, thresh=0, levels=100, cmap="mako",
)
| doc/docstrings/kdeplot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from tf.app import use
from fixture import typeShow
# # Tisch
A = use("tisch:clone", checkout="clone", hoist=globals())
A.reuse()
typeShow(A, prettyTypes=False, baseTypes=set())
| zz_test/085-tisch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Measurements in stabilography
#
# > <NAME>
# > Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
# > Federal University of ABC, Brazil
# Posturography is a general term for all techniques concerned with quantifying postural sway of a standing person.
#
# Typically in posturography, instead of measuring the actual sway of each segment, measurements of the whole body sway have been used. The displacements of center of pressure (COP) and of the vertical projection (on the horizontal plane) of the center of gravity (COGv) are the most common measures of body sway (however, keep in mind that the COP displacement is not an actual measurement of postural sway of the body or of any of its segments). While the COP displacement can be easily measured with a force plate, the direct measurement of the COGv is more complicated and typically subject to a larger error. The measurement of the COGv is computed by measuring the position of each body segment combined with the mass of each segment. More commonly, the COGv displacement is determined indirectly from the COP displacement and different methods are available for such, which produce similar results (Lafond et al. 2004). See the notebook [The inverted pendulum model of the human standing posture](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/IP_Model.ipynb) for a description and code of one of these methods.
#
# It is possible to quantify a certain property of a phenomenon, like the postural control during upright standing, without any underlying scientific theory on what generated that property and what the implication of that measurement for the understanding of the phenomenon is. Sometimes, this is the only approach to start with. However, as our knowledge on the phenomenon advances, the need for a scientific theory or a hypothesis to interpret the results in a more meaningful way becomes evident. It is relatively easy to perform many measurements of the body sway during upright standing; far more difficult is to interpret those measurements to understand what they mean.
#
# The most widespread empirical interpretation of typical measurements of posture sway is that more sway means more instability which is seen as an indication of a deteriorated postural control system. This rationale is based on many experiments on aging and pathological conditions that, indeed, have observed increased sway in those conditions. However, bear in mind that it might not always be the case.
#
# Selecting which measurement will be used to quantify a certain characteristic of the postural control depends on which experiment we design and which task the individual under evaluation is performing. In this sense, posturography has been divided into two experimental paradigms: dynamic and static posturography. In static posturography, the individual is typically trying to stay as quiet as possible while his or her sway is measured, and no perturbation is applied during the task. In dynamic posturography, a momentary perturbation is applied and how the subject responded to that perturbation is measured.
#
# Let's show some of the most common measures of postural sway, more typically employed in static posturography.
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd # use Pandas to read data from a website
fileUrl = 'http://www.udel.edu/biology/rosewc/kaap686/reserve/cop/copdata.txt'
COP = pd.read_table(fileUrl, skipinitialspace=True, sep=None, engine='python') # Pandas dataframe
COP = COP.values / 10 # mm to cm
freq = 100
print('COP shape: ', COP.shape)
def cop_plot(freq, COP, units='cm'):
'''
Plot COP data from postural sway measurement.
'''
import matplotlib.gridspec as gridspec
t = np.linspace(0, COP.shape[0]/freq, COP.shape[0])
plt.rc('axes', labelsize=16, titlesize=16)
plt.rc('xtick', labelsize=12)
plt.rc('ytick', labelsize=12)
plt.figure(figsize=(10, 4))
gs = gridspec.GridSpec(1, 2, width_ratios=[2, 1])
ax1 = plt.subplot(gs[0])
ax1.plot(t, COP[:,0], lw=2, color=[0, 0, 1, 1], label='ap')
ax1.plot(t, COP[:,1], lw=2, color=[1, 0, 0, 1], label='ml')
ax1.set_xlim([t[0], t[-1]])
ax1.grid()
ax1.locator_params(axis='both', nbins=5)
ax1.set_xlabel('Time [s]')
ax1.set_ylabel('COP [%s]' %units)
ax1.set_title('Stabilogram')
ax1.legend(fontsize=12, loc='best', framealpha=.5)
ax2 = plt.subplot(gs[1])
ax2.plot(COP[:,1], COP[:,0], lw=2, color='g')
ax2.set_xlabel('COP ml [%s]' %units)
ax2.set_ylabel('COP ap [%s]' %units)
ax2.set_title('Statokinesigram')
if 0: # plot the axes with the same colors of the COP data
ax2.xaxis.label.set_color('red')
ax2.spines['bottom'].set_color('red')
ax2.tick_params(axis='x', colors='red')
ax2.yaxis.label.set_color('blue')
ax2.spines['left'].set_color('blue')
ax2.tick_params(axis='y', colors='blue')
ax2.grid()
ax2.locator_params(axis='both', nbins=5)
plt.tight_layout()
plt.show()
# plot data
cop_plot(freq, COP)
# ## Data detrend
#
# The mean value of the COP (or COGv) displacement is dependent where the individual stood on the force plate or in the space and usually has no particular interest to the understanding of postural sway. So, a typical procedure in the analysis of postural sway is to remove the mean value of the data. Related to that, the presence of a trend (a slow fluctuation) in the signal might also affect some of the measurements. Someone might argue that the trend itself could give valuable information about the signal (see Duarte and Zatsiorsky, 2000), but the problem is that most of the measurements one uses to describe a signal assumes that the signal is stationary. A signal is stationary if its statistical properties (such as mean and variance) do not change across time. So, to detrend the data might be necessary in certain cases. Another way to remove a trend in the data is to apply a high pass filter to the data with a cut-off frequency related to the period of data acquisition (Witt et al., 1998).
#
# Let's see a simple function for data detrend in Python and use it to remove the mean of the data.
from scipy.signal import detrend
COP = detrend(COP, axis=0, type='constant') # use 'linear' to remove a linear trend
# ## Measurements of spatial variability
#
# Standard deviation, RMS, range (or amplitude), and total path (or total displacement) are commonly employed to describe the spatial variability of some measurement of postural sway. For the data represented by $x$, these variables are defined as:
#
# $$ \bar{x} \; (mean) = \frac{1}{N}\sum_{i=1}^{N} x_i $$
#
# $$ SD = \sqrt{\frac{1}{N-1}\sum_{i=1}^{N} (x_i - \bar{x})^2} $$
#
# $$ RMS = \sqrt{\frac{1}{N}\sum_{i=1}^{N} x_i^2} $$
#
# $$ Range = max[x] - min[x] $$
#
# $$ Total \: path = \sum_{i=1}^{N-1} | x_{i+i}-x_i | $$
# +
m = np.mean(COP, axis=0) # mean
sd = np.std(COP, axis=0) # standard deviation
rms = np.sqrt(np.mean(COP ** 2, axis=0)) # root-mean square
rang = np.max(COP, axis=0) - np.min(COP, axis=0) # range (maximum - minimum)
tpath = np.sum(np.abs(np.diff(COP, axis=0)), axis=0) # total path (length of the COP displacement)
unit = 'cm'
print('Measurements of spatial variability')
print('{0:12} {1:^16}'.format('Variable', 'Direction'))
print('{0:12} {1:^8} {2:^5}'.format('', 'ap', 'ml'))
print('{0:12} {1:>6.2f} {2:>6.2f} {3:>3}'.format('Mean:', m[0], m[1], unit))
print('{0:12} {1:>6.2f} {2:>6.2f} {3:>3}'.format('SD:', sd[0], sd[1], unit))
print('{0:12} {1:>6.2f} {2:>6.2f} {3:>3}'.format('RMS:', rms[0], rms[1], unit))
print('{0:12} {1:>6.2f} {2:>6.2f} {3:>3}'.format('Range:', rang[0], rang[1], unit))
print('{0:12} {1:>6.2f} {2:>6.2f} {3:>3}'.format('Total path:', tpath[0], tpath[1], unit))
# -
# ## Mean velocity (or mean speed)
#
# The mean speed variable expresses the the average velocity of the COP displacement computed simply as the total path variable (the total displacement) divided by the total period. This variable is usually referred as velocity but as it a scalar, it should in fact be named speed.
# The mean resultant speed is the speed calculated in the vectorial form considering each direction (the square root of the sum of the squared speed in each direction). For the data represented by *x* and *y*, these variables are defined as:
# $$ Mean \: speed = \frac{1}{T}\sum_{i=1}^{N-1} | x_{i+i}-x_i | $$
#
# $$ Mean \: resultant \: speed = \frac{1}{T}\sum_{i=1}^{N-1} \sqrt{( x_{i+i}-x_i )^2 + ( y_{i+i}-y_i )^2} $$
mvel = np.sum(np.abs(np.diff(COP, axis=0)), axis=0) / 30
mvelr = np.sum(np.abs(np.sqrt(np.sum(np.diff(COP, axis=0) ** 2, axis=1))), axis=0) / 30
print('{0:15} {1:^16}'.format('Variable', 'Direction'))
print('{0:15} {1:^8} {2:^5}'.format('', 'ap', 'ml'))
print('{0:15} {1:>6.2f} {2:>6.2f} {3:>5}'.format('Mean velocity:', mvel[0], mvel[1], unit+'/s'))
print('')
print('{0:22} {1:>6.2f} {2:>5}'.format('Mean resultant velocity:', mvelr, unit+'/s'))
print('Squared sum of the mean speeds:')
print('{0:>6.2f} {1:>5}'.format(np.sqrt(np.sum(mvel ** 2)), unit+'/s'))
# ## Area
#
# Another measurement of postural sway is to compute the area that encompasses the COPap versus COPml data using a curve in a plane (e.g., a circle or ellipse) or a polygon (e.g., a rectangle). A common method for such is to compute a prediction ellipse, which is found by fitting an ellipse to the data using concepts from the statistical procedure known as principal component analysis. A 95% prediction ellipse is a prediction interval for the COP data (considered to be a bivariate random variable) such that there is 95% of probability that a new observation will lie inside the ellipse. For more details, see [Prediction ellipse and ellipsoid](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/PredictionEllipseEllipsoid.ipynb).
#
# The function `ellipseoid.py` (see [Prediction ellipse and ellipsoid](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/PredictionEllipseEllipsoid.ipynb)) calculates the ellipse area, some related parameters, and plots the results for a given multivariate random variable.
import sys
sys.path.insert(1, r'./../functions')
from hyperellipsoid import hyperellipsoid
area, axes, angles, center, R = hyperellipsoid(COP[:, 1], COP[:, 0], units='cm')
# ## Frequency analysis
#
# Frequency analysis refers to estimate the frequency content of a signal. Let's use standard Fourier analysis and the related power spectral analysis to estimate some frequency characteristics of the COP displacement. The function `psd.py` (see its code in the notebook [Fast Fourier Transform and Power Spectral Density](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/FFTandPSD.ipynb)) estimates power spectral density characteristics using Welch's method using the `scipy.signal.welch` function, estimates some frequency characteristics, and a plots the results.
#
# Let's use `psd.py` to estimate the frequency characteristics of the COP data.
from psd import psd
fp_ap, mf_ap, fmax_ap, Ptot_ap, F, P_ap = psd(COP[:, 0], fs=freq, scales='linear', xlim=[0, 2], units='cm')
fp_ml, mf_ml, fmax_ml, Ptot_ml, F, P_ml = psd(COP[:, 1], fs=freq, xlim=[0, 2], units='cm')
# ## Other analyses
#
#
# ## References
#
# - <NAME>, <NAME> (2000) [On the fractal properties of natural human standing](http://www.ncbi.nlm.nih.gov/pubmed/10754215). Neuroscience Letters, 283, 173-176.
# - <NAME>., <NAME>, et al. (2004). [Comparison of three methods to estimate the center of mass during balance assessment](http://www.ncbi.nlm.nih.gov/pubmed/15275850). J. Biomech. 37(9): 1421-1426.
# - <NAME>, <NAME>, <NAME> (1998) [Testing stationarity in time series](http://journals.aps.org/pre/abstract/10.1103/PhysRevE.58.1800). Physical Review E, 58, 1800-1810.
| notebooks/Stabilography.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS-109A Introduction to Data Science
#
#
# ## Lab 11: Neural Network Basics - Introduction to `tf.keras`
#
# **Harvard University**<br>
# **Fall 2019**<br>
# **Instructors:** <NAME>, <NAME>, <NAME><br>
# **Lab Instructors:** <NAME> and <NAME>. <br>
# **Authors:** <NAME>, <NAME>, and <NAME>.
## RUN THIS CELL TO PROPERLY HIGHLIGHT THE EXERCISES
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import pandas as pd
# %matplotlib inline
from PIL import Image
# +
from __future__ import absolute_import, division, print_function, unicode_literals
# TensorFlow and tf.keras
import tensorflow as tf
tf.keras.backend.clear_session() # For easy reset of notebook state.
print(tf.__version__) # You should see a 2.0.0 here!
# -
# #### Instructions for running `tf.keras` with Tensorflow 2.0:
#
# 1. Create a `conda` virtual environment by cloning an existing one that you know works
# ```
# conda create --name myclone --clone myenv
# ```
#
# 2. Go to [https://www.tensorflow.org/install/pip](https://www.tensorflow.org/install/pip) and follow instructions for your machine.
#
# 3. In a nutshell:
# ```
# pip install --upgrade pip
# pip install tensorflow==2.0.0
# ```
#
# All references to Keras should be written as `tf.keras`. For example:
#
# ```
# model = tf.keras.models.Sequential([
# tf.keras.layers.Flatten(input_shape=(28, 28)),
# tf.keras.layers.Dense(128, activation='relu'),
# tf.keras.layers.Dropout(0.2),
# tf.keras.layers.Dense(10, activation='softmax')
# ])
#
# model.compile(optimizer='adam',
# loss='sparse_categorical_crossentropy',
# metrics=['accuracy'])
#
# tf.keras.models.Sequential
# tf.keras.layers.Dense, tf.keras.layers.Activation,
# tf.keras.layers.Dropout, tf.keras.layers.Flatten, tf.keras.layers.Reshape
# tf.keras.optimizers.SGD
# tf.keras.preprocessing.image.ImageDataGenerator
# tf.keras.regularizers
# tf.keras.datasets.mnist
# ```
#
# You could avoid the long names by using
# ```
# from tensorflow import keras
# from tensorflow.keras import layers
# ```
# These imports do not work on some systems, however, because they pick up previous versions of `keras` and `tensorflow`. That is why I avoid them in this lab.
# ## Learning Goals
# In this lab we will understand the basics of neural networks and how to start using a deep learning library called `keras`. By the end of this lab, you should:
#
# - Understand how a simple neural network works and code some of its functionality from scratch.
# - Be able to think and do calculations in matrix notation. Also think of vectors and arrays as tensors.
# - Know how to install and run `tf.keras`.
# - Implement a simple real world example using a neural network.
# ## Part 1: Neural Networks 101
# Suppose we have an input vector $X=${$x_1, x_2, ... x_L$} to a $k$-layered network. <BR><BR>
# Each layer has its own number of nodes. For the first layer in our drawing that number is $J$. We can store the weights for each node in a vector $\mathbf{W} \in \mathbb{R}^{JxL+1}$ (accounting for bias). Similarly, we can store the biases from each node in a vector $\mathbf{b} \in \mathbb{R}^{I}$. The affine transformation is then written as $$\mathbf{a} = \mathbf{W^T}X + \mathbf{b}$$ <BR> What we then do is "absorb" $\mathbf{b}$ into $X$ by adding a column of ones to $X$. Our $X$ matrix than becomes $\mathbf{X} \in \mathbb{R}^{JxL+1}$ and our equation: <BR><BR>$$\mathbf{a} = \mathbf{W^T}_{plusones}X$$ <br>We have that $\mathbf{a} \in \mathbb{R}^{J}$ as well. Next we evaluate the output from each node. We write $$\mathbf{u} = \sigma\left(\mathbf{a}\right)$$ where $\mathbf{u}\in\mathbb{R}^{J}$. We can think of $\sigma$ operating on each individual element of $\mathbf{a}$ separately or in matrix notation. If we denote each component of $\mathbf{a}$ by $a_{j}$ then we can write $$u_{j} = \sigma\left(a_{j}\right), \quad j = 1, ... J.$$<br> In our code we will implement all these equations in matrix notation.
# `tf.keras` (Tensorflow) and `numpy` perform the calculations in matrix format.
# 
# <br><br>
# Image source: *"Modern Mathematical Methods for Computational Science and Engineering"* <NAME> and <NAME>.
# Let's assume that we have 3 input points (L = 3), two hidden layers ($k=2$), and 2 nodes in each layer ($J=2$)<br>
#
# ### Input Layer
#
# $𝑋$={$𝑥_1,𝑥_2,x_3$}
#
# ### First Hidden Layer
#
# \begin{equation}
# \begin{aligned}
# a^{(1)}_1 = w^{(1)}_{10} + w^{(1)}_{11}x_1 + w^{(1)}_{12}x_2 + w^{(1)}_{13}x_3 \\
# a^{(1)}_2 = w^{(1)}_{20} + w^{(1)}_{21}x_1 + w^{(1)}_{22}x_2 + w^{(1)}_{23}x_3 \\
# \end{aligned}
# \end{equation}
# <br> All this in matrix notation: $$\mathbf{a} = \mathbf{W^T}X$$
# <br> NOTE: in $X$ we have added a column of ones to account for the bias<BR><BR>
# **Then the sigmoid is applied**:
# \begin{equation}
# \begin{aligned}
# u^{(1)}_1 = \sigma(a^{(1)}_1) \\
# u^{(1)}_2 = \sigma(a^{(1)}_2) \\
# \end{aligned}
# \end{equation}
#
# or in matrix notation: $$\mathbf{u} = \sigma\left(\mathbf{a}\right)$$
#
# ### Second Hidden Layer
#
# \begin{equation}
# \begin{aligned}
# a^{(2)}_1 = w^{(2)}_{10} + w^{(2)}_{11}u^{(1)}_1 + w^{(2)}_{12}u^{(1)}_2 + w^{(2)}_{13}u^{(1)}_3 \\
# a^{(2)}_2 = w^{(2)}_{20} + w^{(2)}_{21}u^{(1)}_1 + w^{(2)}_{22}u^{(1)}_2 + w^{(2)}_{23}u^{(1)}_3 \\
# \end{aligned}
# \end{equation}
# <br>
#
# **Then the sigmoid is applied**:
#
# \begin{equation}
# \begin{aligned}
# u^{(2)}_1 = \sigma(a^{(2)}_1) \\
# u^{(2)}_2 = \sigma(a^{(2)}_2) \\
# \end{aligned}
# \end{equation}
#
# ### Output Layer
#
# #### If the output is categorical:
#
# For example with four classes ($M=4$): $Y$={$y_1, y_2, y_3, y_4$}, we have the affine and then the sigmoid is lastly applied:
#
# \begin{equation}
# \begin{aligned}
# a^{(3)}_1 = w^{(3)}_{10} + w^{(3)}_{11}u^{(2)}_1 + w^{(3)}_{12}u^{(2)}_2 \\
# a^{(3)}_2 = w^{(3)}_{20} + w^{(3)}_{21}u^{(2)}_1 + w^{(3)}_{22}u^{(2)}_2 \\
# a^{(3)}_3 = w^{(3)}_{30} + w^{(3)}_{31}u^{(2)}_1 + w^{(3)}_{32}u^{(2)}_2 \\
# a^{(3)}_4 = w^{(3)}_{40} + w^{(3)}_{41}u^{(2)}_1 + w^{(3)}_{42}u^{(2)}_2 \\
# \end{aligned}
# \end{equation}
# <br>
# \begin{equation}
# \begin{aligned}
# y_1 = \sigma(a^{(3)}_1) \\
# y_2 = \sigma(a^{(3)}_2) \\
# y_3 = \sigma(a^{(3)}_3) \\
# y_3 = \sigma(a^{(3)}_4) \\
# \end{aligned}
# \end{equation}
# $\sigma$ will be softmax in the case of multiple classes and sigmoid for binary.
# <BR>
#
# #### If the output is a number (regression):
#
# We have a single y as output:
#
# \begin{equation}
# \begin{aligned}
# y = w^{(3)}_{10}+ w^{(3)}_{11}u^{(2)}_1 + w^{(3)}_{12}u^{(2)}_2 + w^{(3)}_{13}u^{(2)}_3 \\
# \end{aligned}
# \end{equation}
#
# #### Matrix Multiplication and constant addition
a = np.array([[1, 0], [0, 1], [2, 3]])
b = np.array([[4, 1, 1], [2, 2, 1]])
print(np.matrix(a))
print('------')
print(np.matrix(b))
# both Tensorflow and numpy take care of transposing.
c = tf.matmul(a, b) # the tensorflow way
print(c)
d = np.dot(a, b) # the numpy way
print(d)
# how do we add the constant in the matrix
a = [[1, 0], [0, 1]]
ones = np.ones((len(a),1))
a = np.append(a, ones, axis=1)
a
# <div class="exercise"><b>1. In class exercise : Plot the sigmoid</b></div>
#
# Define the `sigmoid` and the `tanh`. For `tanh` you may use `np.tanh` and for the `sigmoid` use the general equation:
# \begin{align}
# \sigma = \dfrac{1}{1+e^{-2(x-c)/a}} \qquad\text{(1.1)}
# \textrm{}
# \end{align}
#
# Generate a list of 500 $x$ points from -5 to 5 and plot both functions. What do you observe? What do variables $c$ and $a$ do?
# +
# your code here
# +
# # %load solutions/sigmoid.py
# The smaller the `a`, the sharper the function is.
# Variable `c` moves the function along the x axis
def sigmoid(x,c,a):
z = ((x-c)/a)
return 1.0 / (1.0 + np.exp(-z))
x = np.linspace(-5.0, 5.0, 500) # input points
c = 1.
a = 0.5
plt.plot(x, sigmoid(x, c, a), label='sigmoid')
plt.plot(x, np.tanh(x), label='tanh')
plt.grid();
plt.legend();
# -
# <div class="exercise"><b>2. In class exercise: Approximate a Gaussian function using a node and manually adjusting the weights. Start with one layer with one node and move to two nodes.</b></div>
# The task is to approximate (learn) a function $f\left(x\right)$ given some input $x$. For demonstration purposes, the function we will try to learn is a Gaussian function:
# \begin{align}
# f\left(x\right) = e^{-x^{2}}
# \textrm{}
# \end{align}
#
# Even though we represent the input $x$ as a vector on the computer, you should think of it as a single input.
# #### 2.1 Start by plotting the above function using the $x$ dataset you created earlier
# +
x = np.linspace(-5.0, 5.0, 500) # input points
def gaussian(x):
return np.exp(-x*x)
f = gaussian(x)
plt.plot(x, f, label='gaussian')
plt.legend()
# -
f.shape
# #### 2.2 Now, let's code the single node as per the image above.
#
# Write a function named `affine` that does the transformation. The definition is provided below. Then create a simpler sigmoid with just one variable. We choose a **sigmoid** activation function and specifically the **logistic** function. Sigmoids are a family of functions and the logistic function is just one member in that family. $$\sigma\left(z\right) = \dfrac{1}{1 + e^{-z}}.$$ <br>
#
# Define both functions in code.
def affine(x, w, b):
"""Return affine transformation of x
INPUTS
======
x: A numpy array of points in x
w: An array representing the weight of the perceptron
b: An array representing the biases of the perceptron
RETURN
======
z: A numpy array of points after the affine transformation
z = wx + b
"""
# Code goes here
return z
# your code here
# +
# # %load solutions/affine-sigmoid.py
def affine(x, w, b):
return w * x + b
def sigmoid(z):
return 1.0 / (1.0 + np.exp(-z))
# -
# And now we plot the activation function and the true function. What do you think will happen if you change $w$ and $b$?
# +
w = [-5.0, 0.1, 5.0] # Create a list of weights
b = [0.0, -1.0, 1.0] # Create a list of biases
fig, ax = plt.subplots(1,1, figsize=(9,5))
SIZE = 16
# plot our true function, the gaussian
ax.plot(x, f, lw=4, ls='-.', label='True function')
# plot 3 "networks"
for wi, bi in zip(w, b):
h = sigmoid(affine(x, wi, bi))
ax.plot(x, h, lw=4, label=r'$w = {0}$, $b = {1}$'.format(wi,bi))
ax.set_title('Single neuron network', fontsize=SIZE)
# Create labels (very important!)
ax.set_xlabel('$x$', fontsize=SIZE) # Notice we make the labels big enough to read
ax.set_ylabel('$y$', fontsize=SIZE)
ax.tick_params(labelsize=SIZE) # Make the tick labels big enough to read
ax.legend(fontsize=SIZE, loc='best') # Create a legend and make it big enough to read
# -
# We didn't do an exhaustive search of the weights and biases, but it sure looks like this single perceptron is never going to match the actual function. Again, we shouldn't be suprised about this. The output layer of the network is simple the logistic function, which can only have so much flexibility.
#
# Let's try to make our network more flexible by using **more nodes**!
# ### Multiple Perceptrons in a Single Layer
# It appears that a single neuron is somewhat limited in what it can accomplish. What if we expand the number of nodes/neurons in our network? We have two obvious choices here. One option is to add depth to the network by putting layers next to each other. The other option is to stack neurons on top of each other in the same layer. Now the network has some width, but is still only one layer deep.
# +
x = np.linspace(-5.0, 5.0, 500) # input points
f = np.exp(-x*x) # data
w = np.array([3.5, -3.5])
b = np.array([3.5, 3.5])
# Affine transformations
z1 = w[0] * x + b[0]
z2 = w[1] * x + b[1]
# Node outputs
h1 = sigmoid(z1)
h2 = sigmoid(z2)
# -
# Now let's plot things and see what they look like.
# +
fig, ax = plt.subplots(1,1, figsize=(9,5))
ax.plot(x, f, lw=4, ls = '-.', label='True function')
ax.plot(x, h1, lw=4, label='First neuron')
ax.plot(x, h2, lw=4, label='Second neuron')
# Set title
ax.set_title('Comparison of Neuron Outputs', fontsize=SIZE)
# Create labels (very important!)
ax.set_xlabel('$x$', fontsize=SIZE) # Notice we make the labels big enough to read
ax.set_ylabel('$y$', fontsize=SIZE)
ax.tick_params(labelsize=SIZE) # Make the tick labels big enough to read
ax.legend(fontsize=SIZE, loc='best') # Create a legend and make it big enough to read
# -
# Just as we expected. Some sigmoids. Of course, to get the network prediction we must combine these two sigmoid curves somehow. First we'll just add $h_{1}$ and $h_{2}$ without any weights to see what happens.
#
# #### Note
# We are **not** doing classification here. We are trying to predict an actual function. The sigmoid activation is convenient when doing classification because you need to go from $0$ to $1$. However, when learning a function, we don't have as good of a reason to choose a sigmoid.
# Network output
wout = np.ones(2) # Set the output weights to unity to begin
bout = -1 # bias
yout = wout[0] * h1 + wout[1] * h2 + bout
# And plot.
# +
fig, ax = plt.subplots(1,1, figsize=(9,5))
ax.plot(x, f, ls='-.', lw=4, label=r'True function')
ax.plot(x, yout, lw=4, label=r'$y_{out} = h_{1} + h_{2}$')
# Create labels (very important!)
ax.set_xlabel('$x$', fontsize=SIZE) # Notice we make the labels big enough to read
ax.set_ylabel('$y$', fontsize=SIZE)
ax.tick_params(labelsize=SIZE) # Make the tick labels big enough to read
ax.legend(fontsize=SIZE, loc='best') # Create a legend and make it big enough to read
# -
# Very cool! The two nodes interact with each other to produce a pretty complicated-looking function. It still doesn't match the true function, but now we have some hope. In fact, it's starting to look a little bit like a Gaussian!
#
# We can do better. There are three obvious options at this point:
# 1. Change the number of nodes
# 2. Change the activation functions
# 3. Change the weights
#
# #### We will leave this simple example for some other time! Let's move on to fashion items!
# ## Part 2: Tensors, Fashion, and Reese Witherspoon
#
# We can think of tensors as multidimensional arrays of real numerical values; their job is to generalize matrices to multiple dimensions. While tensors first emerged in the 20th century, they have since been applied to numerous other disciplines, including machine learning. Tensor decomposition/factorization can solve, among other, problems in unsupervised learning settings, temporal and multirelational data. For those of you that will get to handle images for Convolutional Neural Networks, it's a good idea to have the understanding of tensors of rank 3.
#
# We will use the following naming conventions:
#
# - scalar = just a number = rank 0 tensor ($a$ ∈ $F$,)
# <BR><BR>
# - vector = 1D array = rank 1 tensor ( $x = (\;x_1,...,x_i\;)⊤$ ∈ $F^n$ )
# <BR><BR>
# - matrix = 2D array = rank 2 tensor ( $\textbf{X} = [a_{ij}] ∈ F^{m×n}$ )
# <BR><BR>
# - 3D array = rank 3 tensor ( $\mathscr{X} =[t_{i,j,k}]∈F^{m×n×l}$ )
# <BR><BR>
# - $\mathscr{N}$D array = rank $\mathscr{N}$ tensor ( $\mathscr{T} =[t_{i1},...,t_{i\mathscr{N}}]∈F^{n_1×...×n_\mathscr{N}}$ ) <-- Things start to get complicated here...
#
# #### Tensor indexing
# We can create subarrays by fixing some of the given tensor’s indices. We can create a vector by fixing all but one index. A 2D matrix is created when fixing all but two indices. For example, for a third order tensor the vectors are
# <br><BR>
# $\mathscr{X}[:,j,k]$ = $\mathscr{X}[j,k]$ (column), <br>
# $\mathscr{X}[i,:,k]$ = $\mathscr{X}[i,k]$ (row), and <BR>
# $\mathscr{X}[i,j,:]$ = $\mathscr{X}[i,j]$ (tube) <BR>
#
# #### Tensor multiplication
# We can multiply one matrix with another as long as the sizes are compatible ((n × m) × (m × p) = n × p), and also multiply an entire matrix by a constant. Numpy `numpy.dot` performs a matrix multiplication which is straightforward when we have 2D or 1D arrays. But what about > 3D arrays? The function will choose according to the matching dimentions but if we want to choose we should use `tensordot`, but, again, we **do not need tensordot** for this class.
# ### <NAME>
#
# This image is from the dataset [Labeled Faces in the Wild](http://vis-www.cs.umass.edu/lfw/person/Reese_Witherspoon.html) used for machine learning training. Images are 24-bit RGB images (height, width, channels) with 8 bits for each of R, G, B channel. Explore and print the array.
# load and show the image
FILE = '../fig/Reese_Witherspoon.jpg'
img = mpimg.imread(FILE)
imgplot = plt.imshow(img)
print(f'The image is a: {type(img)} of shape {img.shape}')
img[3:5, 3:5, :]
# #### Slicing tensors: slice along each axis
# we want to show each color channel
fig, axes = plt.subplots(1, 3, figsize=(10,10))
for i, subplot in zip(range(3), axes):
temp = np.zeros(img.shape, dtype='uint8')
temp[:,:,i] = img[:,:,i]
subplot.imshow(temp)
subplot.set_axis_off()
plt.show()
# #### Multiplying Images with a scalar (just for fun, does not really help us in any way)
temp = img
temp = temp * 2
plt.imshow(temp)
# For more on image manipulation by `matplotlib` see: [matplotlib-images](https://matplotlib.org/3.1.1/tutorials/introductory/images.html)
# ### Anatomy of an Artificial Neural Network
#
# In Part 1 we hand-made a neural network by writing some simple python functions. We focused on a regression problem where we tried to learn a function. We practiced using the logistic activation function in a network with multiple nodes, but a single or two hidden layers. Some of the key observations were:
# * Increasing the number of nodes allows us to represent more complicated functions
# * The weights and biases have a very big impact on the solution
# * Finding the "correct" weights and biases is really hard to do manually
# * There must be a better method for determining the weights and biases automatically
#
# We also didn't assess the effects of different activation functions or different network depths.
# ### 
# https://www.tensorflow.org/guide/keras
#
# `tf.keras` is TensorFlow's high-level API for building and training deep learning models. It's used for fast prototyping, state-of-the-art research, and production. `Keras` is a library created by <NAME>. After Google released Tensorflow 2.0, the creators of `keras` recommend that "Keras users who use multi-backend Keras with the TensorFlow backend switch to `tf.keras` in TensorFlow 2.0. `tf.keras` is better maintained and has better integration with TensorFlow features".
#
# #### IMPORTANT: In `Keras` everything starts with a Tensor of N samples as input and ends with a Tensor of N samples as output.
# ### The 3 parts of an ANN
#
# - **Part 1: the input layer** (our dataset)
# - **Part 2: the internal architecture or hidden layers** (the number of layers, the activation functions, the learnable parameters and other hyperparameters)
# - **Part 3: the output layer** (what we want from the network)
#
# In the rest of the lab we will practice with end-to-end neural network training
#
# 1. Load the data
# 2. Define the layers of the model.
# 3. Compile the model.
# 4. Fit the model to the train set (also using a validation set).
# 5. Evaluate the model on the test set.
# 6. Plot metrics such as accuracy.
# 7. Predict on random images from test set.
# 8. Predict on a random image from the web!
seed = 7
np.random.seed(seed)
# ### Fashion MNIST
#
# 
#
# MNIST, the set of handwritten digits is considered the Drosophila of Machine Learning. It has been overused, though, so we will try a slight modification to it.
#
# **Fashion-MNIST** is a dataset of clothing article images (created by [Zalando](https://github.com/zalandoresearch/fashion-mnist)), consisting of a training set of 60,000 examples and a test set of 10,000 examples. Each example is a **28 x 28** grayscale image, associated with a label from **10 classes**. The creators intend Fashion-MNIST to serve as a direct drop-in replacement for the original MNIST dataset for benchmarking machine learning algorithms. It shares the same image size and structure of training and testing splits. Each pixel is 8 bits so its value ranges from 0 to 255.
#
# Let's load and look at it!
# #### 1. Load the data
# +
# %%time
# get the data from keras
fashion_mnist = tf.keras.datasets.fashion_mnist
# load the data splitted in train and test! how nice!
(x_train, y_train),(x_test, y_test) = fashion_mnist.load_data()
# normalize the data by dividing with pixel intensity
# (each pixel is 8 bits so its value ranges from 0 to 255)
x_train, x_test = x_train / 255.0, x_test / 255.0
# classes are named 0-9 so define names for plotting clarity
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# plot
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(x_train[i], cmap=plt.cm.binary)
plt.xlabel(class_names[y_train[i]])
plt.show()
# -
plt.imshow(x_train[3], cmap=plt.cm.binary)
x_train.shape, x_test.shape
y_train.shape
# #### 2. Define the layers of the model.
# type together
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(154, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
#tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
# #### 3. Compile the model
# +
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
model.compile(optimizer=optimizer,
loss=loss_fn,
metrics=['accuracy'])
# -
model.summary()
tf.keras.utils.plot_model(
model,
#to_file='model.png', # if you want to save the image
show_shapes=True, # True for more details than you need
show_layer_names=True,
rankdir='TB',
expand_nested=False,
dpi=96
)
# [Everything you wanted to know about a Keras Model and were afraid to ask](https://www.tensorflow.org/api_docs/python/tf/keras/Model)
# #### 4. Fit the model to the train set (also using a validation set)
#
# This is the part that takes the longest.
#
# -----------------------------------------------------------
# **ep·och** <BR>
# noun: epoch; plural noun: epochs. A period of time in history or a person's life, typically one marked by notable events or particular characteristics. Examples: "the Victorian epoch", "my Neural Netwok's epochs". <BR>
#
# -----------------------------------------------------------
# +
# %%time
# the core of the network training
history = model.fit(x_train, y_train, validation_split=0.33, epochs=50,
verbose=2)
# -
# #### Save the model
#
# You can save the model so you do not have `.fit` everytime you reset the kernel in the notebook. Network training is expensive!
#
# For more details on this see [https://www.tensorflow.org/guide/keras/save_and_serialize](https://www.tensorflow.org/guide/keras/save_and_serialize)
# +
# save the model so you do not have to run the code everytime
model.save('fashion_model.h5')
# Recreate the exact same model purely from the file
#model = tf.keras.models.load_model('fashion_model.h5')
# -
# #### 5. Evaluate the model on the test set.
test_loss, test_accuracy = model.evaluate(x_test, y_test, verbose=0)
print(f'Test accuracy={test_accuracy}')
# #### 6. We learn a lot by studying History! Plot metrics such as accuracy.
#
# You can learn a lot about neural networks by observing how they perform while training. You can issue `kallbacks` in `keras`. The networks's performance is stored in a `keras` callback aptly named `history` which can be plotted.
print(history.history.keys())
# +
# plot accuracy and loss for the test set
fig, ax = plt.subplots(1,2, figsize=(20,6))
ax[0].plot(history.history['accuracy'])
ax[0].plot(history.history['val_accuracy'])
ax[0].set_title('Model accuracy')
ax[0].set_ylabel('accuracy')
ax[0].set_xlabel('epoch')
ax[0].legend(['train', 'val'], loc='best')
ax[1].plot(history.history['loss'])
ax[1].plot(history.history['val_loss'])
ax[1].set_title('Model loss')
ax[1].set_ylabel('loss')
ax[1].set_xlabel('epoch')
ax[1].legend(['train', 'val'], loc='best')
# -
# #### 7. Now let's use the Network for what it was meant to do: Predict!
predictions = model.predict(x_test)
predictions[0]
np.argmax(predictions[0]), class_names[np.argmax(predictions[0])]
# Let's see if our network predicted right! Is the first item what was predicted?
plt.figure()
plt.imshow(x_test[0], cmap=plt.cm.binary)
plt.xlabel(class_names[y_test[0]])
plt.colorbar()
# **Correct!!** Now let's see how confident our model is by plotting the probability values:
# +
# code source: https://www.tensorflow.org/tutorials/keras/classification
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array, true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array, true_label[i]
plt.grid(False)
plt.xticks(range(10))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# -
i = 406
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], y_test, x_test)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], y_test)
plt.show()
# #### 8. Predicting in the real world
#
# Let's see if our network can generalize beyond the MNIST fashion dataset. Let's give it an random googled image of a boot. Does it have to be a clothing item resembling the MNIST fashion dataset? Can it be a puppy?
#
# Download an image from the internet and resize it to 28x28.
# `tf.keras` models are optimized to make predictions on a batch, or collection, of examples at once. Accordingly, even though you're using a single image, you need to add it to a list:
| content/labs/lab11/notes/lab11_MLP_solutions_part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:fastai20] *
# language: python
# name: conda-env-fastai20-py
# ---
# ## Data Load Test 2
# env: fastai20 from ubuntu, local
# includes jupyter lab/notebook
# date: 7/24/2021 3:30 pm start
# date: 7/25/2021 11:10 pm end
# author: <NAME>
# desc: using fastai to load data, try PLANET_TINY fastai source data
# ### S0a. Setup for working locally
# Ubuntu WSL, conda env "fastai20"
# Python v 3.6, jupyter lab/nb, pytorch 1.7,
# nbdev 1.1, need nbdev for viewing docs
# +
# import data science libaries into each alias namespace
from PIL import Image
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# auto-reload all external modules
np.random.seed(42) # set seed for numpy.random
# -
# name.<shift+tab> # dot+shift+tab to view optioninfo.
# test versions, python, pytorch, nbdev
# !python --version # v3.6.10
import torch; print(torch.__version__) # v1.7.1
import nbdev; print(nbdev.__version__) # v1.1.11
# + jupyter={"outputs_hidden": true}
# dir() to see data science lib namespace
#dir()
# -
# Load fastai library, book contents
# install and import fastbook contents
# !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
# +
# import fastbook contents
from fastbook import *
# import all vision library namespace
from fastai.vision.all import *
# alternate vision library import statement
#from fastai.vision import *
# + jupyter={"outputs_hidden": true}
#dir() # prints all fastai vision imported namespace
# -
# ### S0b. Setup for working in Colab
#
# * mount gdrive
# * run fastbook install codes, import fastbook, call setup
# * from pathlib import Path (if "Path" object didn't get imported with fastbook import).
# * somehow upload data folder to gdrive or to Colab instance \contents\gdrive\ path
# * set Path object to point to uploaded data location, gdrive or root of Colab instance.
# * test Path by showing one image file
#
# #### Mount google drive with Google Colab
# ```
# from google.colab import drive
# drive.mount('/content/drive')
# #drive.mount("/content/drive", force_remount=True)
# ```
#
# #### Set data path
# ex1 remote google drive:
# ```Path('/content/gdrive/MyDrive/Colab Notebooks/data/planet_2k')```
# ex2 local:
# ```Path('c:/users/jyoon/repos/data-big/planet-jpg')```
#
# #### Explore path
# # !pwd # print working directory. linux bash command
# # !ls # list items, linux bash command
#
# #### Path object from Python pathlib.py
# path = Path('/contents/gdrive/Colab Notebooks/data/planet-2k/train/')
# path.ls()
# path.BASE_PATH = path
# path.BASE_PATH
#path = Path('/contents/gdrive/Colab Notebooks/data/planet_2k/train/')
#path.ls()
# +
#path.BASE_PATH = path
#path.BASE_PATH
# +
# Javascript keep alive widget, for working online with Google Colab
# #%%javascript
#function ClickConnect(){
# console.log(“Working”);
# document.querySelector(“colab-connect-button”).shadowRoot.getElementById(‘connect’).click();
#}
#setInterval(ClickConnect,60000)
# -
#
# ### S1a. untar_data, PLANET_TINY
#
# Try using fastai's "untar_data" function to download PLANET_TINY data from fastai's aws.s3 server.
# test untar_data namespace
path = untar_data(URLs.PETS)
path.ls()
# download data PLANET_TINY from fastai's aws.s3 storage server.
path2 = untar_data(URLs.PLANET_TINY)
path2.ls()
# shorten path
Path.BASE_PATH = path2
print(Path.BASE_PATH)
print(path2.ls())
path3 = path2/'labels.csv'
path3
filecsv = path2/'labels.csv'
print(filecsv)
df = pd.read_csv(filecsv)
df.head()
len(df)
# +
# load one or more images.
# do images show
# -
# #### S1b. Planet_2k data
# Working locally from Ubuntu fastai20 conda env.
# Data saved to "gdrive/Colab Notebooks/data/planet_2k" locally.
# +
# set path to planet-2k folder
# C:\Users\jyoon\gdrive\Colab Notebooks\data\planet_2k
# location: Path('/mnt/c/users/jyoon/gdrive/Colab Notebooks/data/planet_2k')
path2k = Path('/mnt/c/users/jyoon/gdrive/Colab Notebooks/data/planet_2k')
path2k.ls()
# -
Path.BASE_PATH = path2k
path2k.ls()
csv2k = path2k/'train_classes_2k.csv'
print(csv2k)
df2k = pd.read_csv(csv2k)
#df2k.tail()
df2k.head()
# +
# explore labels data frame
# df2k.describe()
# df2k.tags.unique()
# len(df2k)
# dk2k['tags'] == 'na'
# -
# #### S2b. Explore Planet_2k images
# explore files
files = get_image_files(path2k/"train")
len(files)
files[0], files[1], files[2], files[3], files[4]
from PIL import Image
# We can open an image with the Python Imaging Library's Image class
imlist = []
for i in [0, 1, 2]:
im = Image.open(files[i]).convert('RGB')
print("im")
imlist.append(im)
im
imlist
# +
# how to show 3 images at once? Use matplotlib.pyplot?
# After images loaded into dls, can use "dls.show_batch(3)"
# -
# #### S3a. DataBlock, DataLoaders, DLS
# Firt try loading PLANET_TINY into fastai dls.
# #### S3b. DataBlock, DataLoaders, DLS
# Next load planet_2k images into fastai dls.
# Need img.convert('RGB'), part of dataloader I think.
| rainforest/tests/nbs_data_load_tests/data_load_test2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text"
# This is a companion notebook for the book [Deep Learning with Python, Second Edition](https://www.manning.com/books/deep-learning-with-python-second-edition?a_aid=keras&a_bid=76564dff). For readability, it only contains runnable code blocks and section titles, and omits everything else in the book: text paragraphs, figures, and pseudocode.
#
# **If you want to be able to follow what's going on, I recommend reading the notebook side by side with your copy of the book.**
#
# This notebook was generated for TensorFlow 2.6.
# + [markdown] colab_type="text"
# ## Neural style transfer
# + [markdown] colab_type="text"
# ### The content loss
# + [markdown] colab_type="text"
# ### The style loss
# + [markdown] colab_type="text"
# ### Neural style transfer in Keras
# + [markdown] colab_type="text"
# **Getting the style and content images**
# + colab_type="code"
from tensorflow import keras
base_image_path = keras.utils.get_file(
"sf.jpg", origin="https://img-datasets.s3.amazonaws.com/sf.jpg")
style_reference_image_path = keras.utils.get_file(
"starry_night.jpg", origin="https://img-datasets.s3.amazonaws.com/starry_night.jpg")
original_width, original_height = keras.utils.load_img(base_image_path).size
img_height = 400
img_width = round(original_width * img_height / original_height)
# + [markdown] colab_type="text"
# **Auxiliary functions**
# + colab_type="code"
import numpy as np
def preprocess_image(image_path):
img = keras.utils.load_img(
image_path, target_size=(img_height, img_width))
img = keras.utils.img_to_array(img)
img = np.expand_dims(img, axis=0)
img = keras.applications.vgg19.preprocess_input(img)
return img
def deprocess_image(img):
img = img.reshape((img_height, img_width, 3))
img[:, :, 0] += 103.939
img[:, :, 1] += 116.779
img[:, :, 2] += 123.68
img = img[:, :, ::-1]
img = np.clip(img, 0, 255).astype("uint8")
return img
# + [markdown] colab_type="text"
# **Using a pretrained VGG19 model to create a feature extractor**
# + colab_type="code"
model = keras.applications.vgg19.VGG19(weights="imagenet", include_top=False)
outputs_dict = dict([(layer.name, layer.output) for layer in model.layers])
feature_extractor = keras.Model(inputs=model.inputs, outputs=outputs_dict)
# + [markdown] colab_type="text"
# **Content loss**
# + colab_type="code"
def content_loss(base_img, combination_img):
return tf.reduce_sum(tf.square(combination_img - base_img))
# + [markdown] colab_type="text"
# **Style loss**
# + colab_type="code"
def gram_matrix(x):
x = tf.transpose(x, (2, 0, 1))
features = tf.reshape(x, (tf.shape(x)[0], -1))
gram = tf.matmul(features, tf.transpose(features))
return gram
def style_loss(style_img, combination_img):
S = gram_matrix(style_img)
C = gram_matrix(combination_img)
channels = 3
size = img_height * img_width
return tf.reduce_sum(tf.square(S - C)) / (4.0 * (channels ** 2) * (size ** 2))
# + [markdown] colab_type="text"
# **Total variation loss**
# + colab_type="code"
def total_variation_loss(x):
a = tf.square(
x[:, : img_height - 1, : img_width - 1, :] - x[:, 1:, : img_width - 1, :]
)
b = tf.square(
x[:, : img_height - 1, : img_width - 1, :] - x[:, : img_height - 1, 1:, :]
)
return tf.reduce_sum(tf.pow(a + b, 1.25))
# + [markdown] colab_type="text"
# **Defining the final loss that you'll minimize**
# + colab_type="code"
style_layer_names = [
"block1_conv1",
"block2_conv1",
"block3_conv1",
"block4_conv1",
"block5_conv1",
]
content_layer_name = "block5_conv2"
total_variation_weight = 1e-6
style_weight = 1e-6
content_weight = 2.5e-8
def compute_loss(combination_image, base_image, style_reference_image):
input_tensor = tf.concat(
[base_image, style_reference_image, combination_image], axis=0
)
features = feature_extractor(input_tensor)
loss = tf.zeros(shape=())
layer_features = features[content_layer_name]
base_image_features = layer_features[0, :, :, :]
combination_features = layer_features[2, :, :, :]
loss = loss + content_weight * content_loss(
base_image_features, combination_features
)
for layer_name in style_layer_names:
layer_features = features[layer_name]
style_reference_features = layer_features[1, :, :, :]
combination_features = layer_features[2, :, :, :]
style_loss_value = style_loss(
style_reference_features, combination_features)
loss += (style_weight / len(style_layer_names)) * style_loss_value
loss += total_variation_weight * total_variation_loss(combination_image)
return loss
# + [markdown] colab_type="text"
# **Setting up the gradient-descent process**
# + colab_type="code"
import tensorflow as tf
@tf.function
def compute_loss_and_grads(combination_image, base_image, style_reference_image):
with tf.GradientTape() as tape:
loss = compute_loss(combination_image, base_image, style_reference_image)
grads = tape.gradient(loss, combination_image)
return loss, grads
optimizer = keras.optimizers.SGD(
keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=100.0, decay_steps=100, decay_rate=0.96
)
)
base_image = preprocess_image(base_image_path)
style_reference_image = preprocess_image(style_reference_image_path)
combination_image = tf.Variable(preprocess_image(base_image_path))
iterations = 4000
for i in range(1, iterations + 1):
loss, grads = compute_loss_and_grads(
combination_image, base_image, style_reference_image
)
optimizer.apply_gradients([(grads, combination_image)])
if i % 100 == 0:
print(f"Iteration {i}: loss={loss:.2f}")
img = deprocess_image(combination_image.numpy())
fname = f"combination_image_at_iteration_{i}.png"
keras.utils.save_img(fname, img)
# + [markdown] colab_type="text"
# ### Wrapping up
| chapter12_part03_neural-style-transfer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="LhtKaWCX4Uc7" colab_type="code" colab={} executionInfo={"status": "ok", "timestamp": 1596835103686, "user_tz": 420, "elapsed": 673, "user": {"displayName": "infinite sheets shees", "photoUrl": "", "userId": "02933683546179754709"}}
import requests
import json
import copy
class fetcher:
def fetch(c):
response = fetcher.fetch_from_musenet(c)
if fetcher.erred(response):
return fetcher.problems(response)
c.add_children(fetcher.process_raw_musenet(response))
return c
def fetch_from_musenet(c):
settings = c.settings
p = {"genre":settings.genre,
"instrument":settings.instrumentation,
"encoding":settings.enc,
"temperature":settings.temp,
"truncation":settings.truncation,
"generationLength": settings.length,
"audioFormat":settings.audioFormat}
h = {
"Content-Type": "application/json",
"path": "/sample"
}
r = requests.post("https://musenet.openai.com/sample",
data = json.dumps(p),
headers = h
)
return r.content.decode()
def process_raw_musenet(response):
return json.loads(response)["completions"]
def problems(response):
try:
out = fetcher.process_raw_musenet(response)
return None
except Exception as e:
return e
def erred(response):
return isinstance(fetcher.problems(response), Exception)
# + id="7Aheb-ijyfO3" colab_type="code" colab={}
| midimusic/api_interaction/fetcher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Atoms of Computation
#
# Programming a quantum computer is now something that anyone can do in the comfort of their own home.
#
# But what to create? What is a quantum program anyway? In fact, what is a quantum computer?
#
# These questions can be answered by making comparisons to standard digital computers. Unfortunately, most people don’t actually understand how digital computers work either. So in this article we’ll look at the basics principles behind these devices. To help us transition over to quantum computing later on, we’ll do it using the same tools as well use for quantum.
from qiskit import *
from qiskit.visualization import plot_histogram
# ### Splitting information into bits
# The first thing we need to know about is the idea of bits. These are designed to be the world’s simplest alphabet. With only two characters, 0 and 1, we can represent any piece of information.
#
# One example is numbers. In European languages, numbers are usually represented using a string of the ten digits 0, 1, 2, 3, 4, 5, 6, 7, 8 and 9. In this string of digits, each represents how many times the number contains a certain powers of ten. For example, when we write 9213, we mean
#
# $$ 9000 + 200 + 10 + 3 $$
#
# or, expressed in a way that emphasizes the powers of ten
#
# $$ (9\times10^3) + (2\times10^2) + (1\times10^1) + (3\times10^0) $$
#
# Though we usually use this system based on the number 10, we can just as easily use one based on any other number. The binary number system, for example, is based on the number two. This means using the two characters 0 and 1 to express numbers as multiples of powers of two. For example, 9213 becomes 10001111111101, since
#
# $$ 9213 = (1 \times 2^{13}) + (0 \times 2^{12}) + (0 \times 2^{11})+ (0 \times 2^{10}) +(1 \times 2^9) + (1 \times 2^8) + (1 \times 2^7) \\\\ \,\,\, + (1 \times 2^6) + (1 \times 2^5) + (1 \times 2^4) + (1 \times 2^3) + (1 \times 2^2) + (0 \times 2^1) + (1 \times 2^0) $$
#
# In this we are expressing numbers as multiples of 2, 4, 8, 16, 32, etc instead of 10, 100, 1000, etc.
#
# These strings of bits, known as binary strings, can be used to represent more than just numbers. For example, there is a way to represent any text using bits. For any letter, number or punctuation mark you want to use, you can find a corresponding string of at most eight bits using [this table.](https://www.ibm.com/support/knowledgecenter/en/ssw_aix_72/com.ibm.aix.networkcomm/conversion_table.htm) Though these are quite arbitrary, this is a widely agreed upon standard. In fact, it's what was used to transmit this article to you through the internet.
#
# This is how all information is represented in computers. Whether numbers, letters, images or sound, it all exists in the form of binary strings.
#
# Now let’s write a binary string using Qiskit. We’ll use the qubits as if they were just normal bits, and not do anything quantum with them.
# +
n = 8
qc_output = QuantumCircuit(n,n)
for j in range(n):
qc_output.measure(j,j)
qc_output.draw(output='mpl')
# -
# The picture above shows what is known (for historical reasons) as a circuit. It depicts the story of what happens to the qubits during the program, from left to right.
#
# In this particular example, not much is happening. Eight qubits have been set up and numbered from 0 to 7 (because that’s how programmers like to do things). The measure operation is then immediately applied to the qubits, which simply extracts an output of ```0``` or ```1``` from each.
# Qubits are always initialized to give the output ```0```. Since we don't do anything to our qubits in the circuit above, this is exactly the result we'll get when we measure them. We can see this by running the circuit many times and plotting the results in a histogram. From this we see that the result is always ```00000000```: a ```0``` from each qubit.
counts = execute(qc_output,Aer.get_backend('qasm_simulator')).result().get_counts()
plot_histogram(counts)
# The reason for running many times and showing the result as a histogram is because sometimes quantum computers have some randomness in their results. In this case, since we aren’t doing anything quantum, we get just the ```00000000``` result with certainty.
#
# Note that this result comes from a quantum simulator, which is a standard computer calculating what a quantum computer would do. Simulations are only possible for small numbers of qubits, but they are nevertheless a very useful tool when designing your first quantum circuits. To run on a real device you simply need to replace ```Aer.get_backend('qasm_simulator')``` with the backend object of the device you want to use.
#
# To encode a different binary string, we need what is known as a NOT gate. This is the most basic operation that you can do in a computer. It simply flips the bit value: ```0``` becomes ```1``` and ```1``` becomes ```0```. In Qiskit, this is done with an operation called ```x```.
# +
qc_encode = QuantumCircuit(n)
qc_encode.x(7)
qc_encode.draw(output='mpl')
# -
# We now need to add measurements to each qubit to extract the result. Since we already have these measurements set up in the circuit `qc_output`, we can just add that onto the end.
qc = qc_encode + qc_output
qc.draw(output='mpl',justify='none')
# Now we can run the combined circuit and look at the results.
counts = execute(qc,Aer.get_backend('qasm_simulator')).result().get_counts()
plot_histogram(counts)
# Now our computer outputs the string ```10000000``` instead.
#
# The bit we flipped, which comes from qubit 7, lives on the far left of the string. This is because Qiskit numbers the bits in a string from left to right. If this convention seems odd to you, don’t worry. It seems odd to lots of other people too, and some prefer to number their bits the other way around. But this system certainly has its advantages when we are using the bits to represent numbers. Specifically, it means that qubit 7 is telling us about how many $2^7$s we have in our number. So by flipping this bit, we’ve now written the number 128 in our simple 8-bit computer.
#
# Now try out writing another number for yourself. You could do your age, for example. Just use a search engine to find out what the number looks like in binary (if it includes a ‘0b’, just ignore it), and then add some 0s to the left side if you are younger than 64.
# +
qc_encode = QuantumCircuit(n)
qc_encode.x(1)
qc_encode.x(5)
qc_encode.draw(output='mpl')
# -
# Now we know how to encode information in a computer. The next step is to process it: To take an input that we have encoded, and turn it into an output that we need.
# ### Remembering how to add
# To look at turning inputs into outputs, we need a problem to solve. Let’s do some basic maths. In primary school you will have learned how to take large mathematical problems and break them down into manageable pieces. For example, how would you go about solving the following?
#
# ```
# 9213
# + 1854
# = ????
# ```
#
# One way is to do it digit by digit, from right to left. So we start with 3+4
# ```
# 9213
# + 1854
# = ???7
# ```
#
# And then 1+5
# ```
# 9213
# + 1854
# = ??67
# ```
#
# Then we have 2+8=10. Since this is a two digit answer, we need to carry the one over to the next column.
#
# ```
# 9213
# + 1854
# = ?067
# ¹
# ```
#
# Finally we have 9+1+1=11, and get our answer
#
# ```
# 9213
# + 1854
# = 11067
# ¹
# ```
#
# This may just be simple addition, but it demonstrates the principles behind all algorithms. Whether the algorithm is designed to solve mathematical problems or process text or images, we always break big tasks down into small and simple steps.
#
# To run on a computer, algorithms need to be compiled down to the smallest and simplest steps possible. To see what these look like, let’s do the above addition problem again, but in binary.
#
#
# ```
# 10001111111101
# + 00011100111110
#
# = ??????????????
# ```
#
# Note that the second number has a bunch of extra 0s on the left. This just serves to make the two strings the same length.
#
# Our first task is to do the 1+0 for the column on the right. In binary, as in any number system, the answer is 1. We get the same result for the 0+1 of the second column.
#
# ```
# 10001111111101
# + 00011100111110
#
# = ????????????11
# ```
#
# Next we have 1+1. As you’ll surely be aware, 1+1=2. In binary, the number 2 is written ```10```, and so requires two bits. This means that we need to carry the 1, just as we would for the number 10 in decimal.
#
# ```
# 10001111111101
# + 00011100111110
# = ???????????011
# ¹
# ```
#
# The next column now requires us to calculate ```1+1+1```. This means adding three numbers together, so things are getting complicated for our computer. But we can still compile it down to simpler operations, and do it in a way that only ever requires us to add two bits together. For this we can start with just the first two 1s.
#
# ```
# 1
# + 1
# = 10
# ```
#
# Now we need to add this ```10``` to the final ```1``` , which can be done using our usual method of going through the columns.
#
# ```
# 10
# + 01
# = 11
# ```
#
# The final answer is ```11``` (also known as 3).
#
# Now we can get back to the rest of the problem. With the answer of ```11```, we have another carry bit.
#
# ```
# 10001111111101
# + 00011100111110
# = ??????????1011
# ¹¹
# ```
#
# So now we have another 1+1+1 to do. But we already know how to do that, so it’s not a big deal.
#
# In fact, everything left so far is something we already know how to do. This is because, if you break everything down into adding just two bits, there’s only four possible things you’ll ever need to calculate. Here are the four basic sums (we’ll write all the answers with two bits to be consistent).
#
# ```
# 0+0 = 00 (in decimal, this is 0+0=0)
# 0+1 = 01 (in decimal, this is 0+1=1)
# 1+0 = 01 (in decimal, this is 1+0=1)
# 1+1 = 10 (in decimal, this is 1+1=2)
# ```
#
# This is called a *half adder*. If our computer can implement this, and if it can chain many of them together, it can add anything.
# ### Adding with Qiskit
# Let's make our own half adder using Qiskit. This will include a part of the circuit that encodes the input, a part that executes the algorithm, and a part that extracts the result. The first part will need to be changed whenever we want to use a new input, but the rest will always remain the same.
# <img src="https://s3.us-south.cloud-object-storage.appdomain.cloud/strapi/04498f84a69d4e859afd19b2760ced24atoms6.png" alt="" width="500" align="middle"/>
# The two bits we want to add are encoded in the qubits 0 and 1. The above example encodes a ```1``` in both these qubits, and so it seeks to find the solution of ```1+1```. The result will be a string of two bits, which we will read out from the qubits 2 and 3. All that remains is to fill in the actual program, which lives in the blank space in the middle.
#
# The dashed lines in the image are just to distinguish the different parts of the circuit (although they can have more interesting uses too). They are made by using the `barrier` command.
#
# The basic operations of computing are known as logic gates. We’ve already used the NOT gate, but this is not enough to make our half adder. We could only use it to manually write out the answers. But since we want the computer to do the actual computing for us, we’ll need some more powerful gates.
#
# To see what we need, let’s take another look at what our half adder needs to do.
#
# ```
# 0+0 = 00
# 0+1 = 01
# 1+0 = 01
# 1+1 = 10
# ```
#
# The rightmost bit in all four of these answers is completely determined by whether the two bits we are adding are the same or different. So for ```0+0``` and ```1+1```, where the two bits are equal, the rightmost bit of the answer comes out ```0```. For ```0+1``` and ```1+0```, where we are adding different bit values, the rightmost bit is ```1```.
#
# To get this part of our solution correct, we need something that can figure out whether two bits are different or not. Traditionally, in the study of digital computation, this is called an XOR gate.
#
# In quantum computers, the job of the XOR gate is done by the controlled-NOT gate. Since that's quite a long name, we usually just call it the CNOT. In Qiskit its name is ```cx```, which is even shorter. In circuit diagrams it is drawn as in the image below.
qc_cnot = QuantumCircuit(2)
qc_cnot.cx(0,1)
qc_cnot.draw(output='mpl')
# This is applied to a pair of qubits. One acts as the control qubit (this is the one with the little dot). The other acts as the *target qubit* (with the big circle).
#
# There are multiple ways to explain the effect of the CNOT. One is to say that it looks at its two input bits to see whether they are the same or different. Then it writes over the target qubit with the answer. The target becomes ```0``` if they are the same, and ```1``` if they are different.
#
# Another way of explaining the CNOT is to say that it does a NOT on the target if the control is ```1```, and does nothing otherwise. This explanation is just as valid as the previous one (in fact, it’s the one that gives the gate its name).
#
# Try the CNOT out for yourself by trying each of the possible inputs. For example, here's a circuit that tests the CNOT with the input ```01```.
qc = QuantumCircuit(2,2)
qc.x(0)
qc.cx(0,1)
qc.measure(0,0)
qc.measure(1,1)
qc.draw(output='mpl')
# If you execute this circuit, you’ll find that the output is ```11```. We can think of this happening because of either of the following reasons.
#
# - The CNOT calculates whether the input values are different and finds that they are, which means that it wants to output ```1```. It does this by writing over the state of qubit 1 (which, remember, is on the left of the bit string), turning ```01``` into ```11```.
#
# - The CNOT sees that qubit 0 is in state ```1```, and so applies a NOT to qubit 1. This flips the ```0``` of qubit 1 into a ```1```, and so turns ```01``` into ```11```.
#
# For our half adder, we don’t want to overwrite one of our inputs. Instead, we want to write the result on a different pair of qubits. For this we can use two CNOTs.
# +
qc_ha = QuantumCircuit(4,2)
# encode inputs in qubits 0 and 1
qc_ha.x(0) # For a=0, remove the this line. For a=1, leave it.
qc_ha.x(1) # For b=0, remove the this line. For b=1, leave it.
qc_ha.barrier()
# use cnots to write the XOR of the inputs on qubit 2
qc_ha.cx(0,2)
qc_ha.cx(1,2)
qc_ha.barrier()
# extract outputs
qc_ha.measure(2,0) # extract XOR value
qc_ha.measure(3,0)
qc_ha.draw(output='mpl')
# -
# We are now halfway to a fully working half adder. We just have the other bit of the output left to do: the one that will live on qubit 4.
#
# If you look again at the four possible sums, you’ll notice that there is only one case for which this is ```1``` instead of ```0```: ```1+1```=```10```. It happens only when both the bits we are adding are ```1```.
#
# To calculate this part of the output, we could just get our computer to look at whether both of the inputs are ```1```. If they are — and only if they are — we need to do a NOT gate on qubit 4. That will flip it to the required value of ```1``` for this case only, giving us the output we need.
#
# For this we need a new gate: like a CNOT, but controlled on two qubits instead of just one. This will perform a NOT on the target qubit only when both controls are in state ```1```. This new gate is called the *Toffoli*. For those of you who are familiar with Boolean logic gates, it is basically an AND gate.
#
# In Qiskit, the Toffoli is represented with the `ccx` command.
# +
qc_ha = QuantumCircuit(4,2)
# encode inputs in qubits 0 and 1
qc_ha.x(0) # For a=0, remove the this line. For a=1, leave it.
qc_ha.x(1) # For b=0, remove the this line. For b=1, leave it.
qc_ha.barrier()
# use cnots to write the XOR of the inputs on qubit 2
qc_ha.cx(0,2)
qc_ha.cx(1,2)
qc_ha.barrier()
# use ccx to write the AND of the inputs on qubit 3
qc_ha.ccx(0,1,3)
# extract outputs
qc_ha.measure(2,0) # extract XOR value
qc_ha.measure(3,1) # extract AND value
qc_ha.draw(output='mpl')
# -
# In this example we are calculating ```1+1```, because the two input bits are both ```1```. Let's see what we get.
counts = execute(qc_ha,Aer.get_backend('qasm_simulator')).result().get_counts()
plot_histogram(counts)
# The result is ```10```, which is the binary representation of the number 2. We have built a computer that can solve the famous mathematical problem of 1+1!
#
# Now you can try it out with the other three possible inputs, and show that our algorithm gives the right results for those too.
#
# The half adder contains everything you need for addition. With the NOT, CNOT and Toffoli gates, we can create programs that add any set of numbers of any size.
#
# These three gates are enough to do everything else in computing too. In fact, we can even do without the CNOT, and the NOT gate is only really needed to create bits with value ```1```. The Toffoli gate is essentially the atom of mathematics. It is simplest element into which every other problem-solving technique can be compiled.
#
# As we'll see, in quantum computing we split the atom.
| ch-states/atoms-computation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Instructions
#
# Data visualization is often a great way to start exploring your data and uncovering insights. In this notebook, you will initiate this process by creating an informative plot of the episode data provided to you. In doing so, you're going to work on several different variables, including the episode number, the viewership, the fan rating, and guest appearances. Here are the requirements needed to pass this project:
#
# 1. Create a `matplotlib` **scatter plot** of the data that contains the following attributes:
#
# 1. Each episode's **episode number plotted along the x-axis**.
# 2. Each episode's **viewership (in millions) plotted along the y-axis**.
# 3. A **color scheme** reflecting the **scaled ratings** (not the regular ratings) of each episode, such that:
# 1. Ratings < 0.25 are colored `"red"`.
# 2. Ratings >= 0.25 and < 0.50 are colored `"orange"`.
# 3. Ratings >= 0.50 and < 0.75 are colored `"lightgreen"`.
# 4. Ratings >= 0.75 are colored `"darkgreen"`.
# 4. A **sizing system**, such that episodes with guest appearances have a marker size of `250` and episodes without are sized `25`.
# 5. A **title**, reading `"Popularity, Quality, and Guest Appearances on the Office"`.
# 6. An **x-axis label** reading `"Episode Number"`.
# 7. A **y-axis label** reading `"Viewership (Millions)"`.
#
# 2. Provide the name of one of the guest stars (hint, there were multiple!) who was in the most watched Office episode. Save it as a string in the variable `top_star` (e.g. `top_star = "<NAME>"`).
#
# ## **Important!**
#
# To test your `matplotlib` plot, you will need to initalize a `matplotlib.pyplot` `fig` object, which you can do using the code `fig = plt.figure()` (provided you have imported `matplotlib.pyplot` as `plt`). In addition, in order to test it correctly, **please make sure to specify your plot (including the type, data, labels, etc) in the same cell as the one you initialize your figure** (`fig`)! _You are still free to use other cells to load data, experiment, and answer Question 2_.
#
# _In addition, if you want to be able to see a larger version of your plot, you can set the figure size parameters using this code (provided again you have imported `matplotlib.pyplot` as `plt`):_
#
# ```
# plt.rcParams['figure.figsize'] = [11, 7]
# ```
#
# ## **Bonus Step!**
#
# Although it was not taught in Intermediate Python, a useful skill for visualizing different data points is to use a different marker. You can learn more about them via the [Matplotlib documentation](https://matplotlib.org/api/markers_api.html) or via our course [Introduction to Data Visualization with Matplotlib](https://learn.datacamp.com/courses/introduction-to-data-visualization-with-matplotlib). Thus, as a bonus step, try to differentiate guest appearances not just with size, but also with a star!
#
# All other attributes still apply (data on the axes, color scheme, sizes for guest appearances, title, and axis labels).
| investigating_netflix_movies_and_guest_stars_in_the_office/project_instructions_unguided.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 5
#
# ## Video 24: Filtering by Product
# **Python for the Energy Industry**
#
# We can filter Cargo Movements by the carried product in much the same way as we filter by geography.
#
# ## Products Endpoint
#
# [Products endpoint documentation](https://vortechsa.github.io/python-sdk/endpoints/products/)
#
# We start by doing a search for 'crude':
# +
# Some initial imports / settings
from datetime import datetime
from dateutil.relativedelta import relativedelta
import vortexasdk as v
now = datetime.utcnow()
one_month_ago = now - relativedelta(months=1)
# -
crude_search = v.Products().search(term=['crude'])
len(crude_search)
# We get 19 results. Inspecting the first of these, we can see there is a product name and ID, plus information about parent products.
crude_search[0]
# Lets take a look at all the products returned by our search:
crude_search.to_df()
# There are different layers of product type, ranging from the broad group 'Crude/Condensates', down to specific grades. We can grab the ID of the Crude group/product to filter on.
# ## Filtering by Products
#
# An assert statement will check that we only get one result for our exact term match.
# +
# Grab the ID for Crude
crude_search = v.Products().search(term=['Crude'],exact_term_match=True)
assert len(crude_search) == 1
crude_ID = crude_search[0]['id']
cm_crude_query = v.CargoMovements().search(
filter_activity="loading_state",
filter_time_min=one_month_ago,
filter_time_max=now,
filter_products=crude_ID)
# -
# As before, we pick the colulmns of interest from our Cargo Movements search and make a DataFrame.
# +
required_columns = ["vessels.0.name","vessels.0.vessel_class","product.group.label","product.grade.label","quantity",
"status","events.cargo_port_load_event.0.location.port.label","events.cargo_port_load_event.0.end_timestamp",
"events.cargo_port_unload_event.0.location.port.label","events.cargo_port_unload_event.0.end_timestamp"]
new_labels = ["vessel_name","vessel_class","product_group","product_grade","quantity","status","loading_port","loading_finish","unloading_port","unloading_finish"]
relabel = dict(zip(required_columns,new_labels))
# -
cm_crude_query.to_df(columns=required_columns).rename(relabel,axis=1)
# ## Exploring the Products dataset
#
# Let's say you wanted to know which child products belonged to a particular group or category. One way to do this is to full the full products dataset:
search = v.Products().search()
len(search)
# We can convert this into a DataFrame. This allows us to, for example, find all products of 'group' type.
# +
search_df = search.to_df()
search_df[search_df['layer.0'] == 'group']
# -
# Let's say we then want to find the 'child' products of 'Crude/Condensates':
search_df[search_df['parent.0.name'] == 'Crude/Condensates']
# Going one step further:
search_df[search_df['parent.0.name'] == 'Crude']
# And finally, to grade level:
search_df[search_df['parent.0.name'] == 'Heavy-Sweet']
# This is a convenient way of exploring the different products that exist in the data.
#
# ### Exercise
#
# Assemble a DataFrame of Cargo Movements that were loading in the last 7 weeks, and that were carrying Diesel out of the United States.
#
# *Note: Diesel is a subset of wider group Diesel/Gasoil.*
| docs/examples/academy/24. Filtering by Product.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Short Jupyter Notebook Intro
#
# Also check this tutorial if you want more details:
#
# https://realpython.com/jupyter-notebook-introduction/
# +
# This is a code cell. It can contain any amount of python code.
print('Somebody spelled Jupiter wrong.')
# -
#you can have more than one line
for i in range(5):
print(f'I ate {i} kilograms of cheese.')
print('That was too much cheese.')
# you can define functions
def my_nice_function(a, b):
return (a - b) / (a + b)
# +
# and call it from another cell
my_nice_function(a=2, b=10)
# -
# The notebook will display the very last value thats being returned in the cell.
# No print needed
a = 1
b = 99
a + b
'Annecy has a lot of cheese'
print('A cell can be executed by pressing "Shift + Enter"')
# We can __plot__ some stuff into the notebook directly with matplotlib.
# First we need to import it.
# +
# same as with numpy. we import a module called 'matplotlib.pyplot' and use it under the name 'plt'
import matplotlib.pyplot as plt
# now use some notebook instructions so the plots will appear directly in the notebook instead of a new window.
# instructions for jupyter begin with % sign
# %matplotlib inline
# -
# now we plot a simple example.
plt.plot([1, 2, 3, 4], [1, 4, 9, 16])
plt.xlabel('Time in Hours')
plt.ylabel('Amount of cheese I eat in Annecy ')
# This is a markdown cell containing some text. It can be _styled_ in __different__ ways.
#
# You can even add some usefull images to explain some physics.
#
# 
# You can even add some equations here in latex-like syntax
#
# $$
# \rho =\sum _{i}w_{i}\left[\sum _{j}{\bar {c}}_{ij}(|\alpha _{ij}\rangle \otimes |\beta _{ij}\rangle )\right]\otimes \left[\sum _{k}c_{ik}(\langle \alpha _{ik}|\otimes \langle \beta _{ik}|)\right]
# $$
# # Short Numpy Intro
#
# Here we go really quickly over some numpy basics again
# the import always looks like this
import numpy as np # import a module called 'numpy' and make it accesible by a shorter name 'np'
# +
# the core data structure in numpy is called an array
x = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
x
# +
# arrays can have more dimensions
y = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
y
# -
y.shape
# +
# we can perform elementwise operations on numpy arrays
x / 10
# -
y + 10
# There a some nice predefined methods to create numpy arrays
# create a range of numbers from 0 to 100 in steps of 5
np.arange(0, 100, 5)
# create a range of 23 numbers from 0 to 1
np.linspace(0, 1, 23)
# create an array containing 3x3 uniformly distributed numbers
np.random.uniform(0, 1, size=(3, 3))
| scipy/basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optical recordings processing with pyCardiac
#
# This notebook demonstrates general pipeline of processing and analysis of Action Potential optical recordings.
#
# Please go through this notebook step by step.
# ## Content
# * 1. <a href="#section1" > Data loading </a>
#
# * 2. <a href="#section2" > Processing </a>
# * 2.1. <a href="#section2.1" > Fourier filtration</a>
# * 2.2. <a href="#section2.2" > Baseline removal </a>
# * 2.3. <a href="#section2.3" > Binning (spatial filtration) </a>
# * 2.4. <a href="#section2.4" > Rescaling (normalizing) </a>
# * 2.5. <a href="#section2.5" > Ensemble averaging </a>
# * 2.6. <a href="#section2.6" > Transform to phase </a>
#
# * 3.<a href="#section3" > Mapping </a>
# * 3.1. <a href="#section3.1" > APD and Alternance maps </a>
# * 3.2. <a href="#section3.2" > Activation time map </a>
# * 3.3. <a href="#section3.3" > Phase singularity points (PS) </a>
# +
import numpy as np
import matplotlib.pyplot as plt
import pyCardiac as pc
from pyCardiac.rhythm import *
from pyCardiac.signal.analysis import phase_singularity_detection
# -
# <a id='section1'></a>
# ## Data loading
data_raw_filename = "./source/sample_optical_recordings.npy"
data_raw = np.load(data_raw_filename)
# <a id='section2'></a>
# ## Processing
data = data_raw.copy()
mask = np.loadtxt("./source/sample_optical_recordings_mask.txt").astype(np.bool)
x, y, t = 80, 60, 15
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data[:, :, t])
plt.imshow(frame, cmap="jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
plt.imshow(mask, cmap = "Greys_r")
plt.plot(x, y, '*k', ms = 10)
plt.title("mask")
plt.show()
plt.figure(figsize = (10, 3.5))
plt.plot(data[y, x, :])
plt.title("signal: x = {x}, y = {y}".format(x=x, y=y))
plt.show()
# -
# <a id='section2.1'></a>
# ### Fourier filtration
kwargs = {'fs' : 1000, # sampling frequency (Hz)
'lp_freq' : 100, # lowpass frequency (Hz)
'hp_freq' : None, # highpass frequency (Hz)
'bs_freqs' : [60, ], # bandstop frequency (Hz)
'trans_width' : 2, # width of transition region between bands (Hz)
'band_width' : 2, # width of bandstop in (Hz)
}
# %time data_filtered = fourier_filter(data, **kwargs)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, data_filtered[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data filtered, t = {t}".format(t = t))
plt.show()
plt.figure(figsize = (10, 3.5))
plt.plot(data[y, x, :],
label = "data", color = "C0")
plt.plot(data_filtered[y, x, :],
label = "data filtered", color = "C1")
plt.title("Signal: x = {x}, y = {y}".format(x=x, y=y))
plt.legend()
plt.show()
# -
# <a id='section2.2'></a>
# ### Baseline removal
# +
#Linear detrending
# %time data_detrended = remove_baseline(data_filtered)
# Asymmetric-Least-Squares Method (slow but more powerful)
# uncomment to use
#niter = 2
# #%time data_detrended = remove_baseline(data_filtered, method_name="least_squares", niter = niter)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data_filtered[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data filtered, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, data_detrended[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data detrended, t = {t}".format(t = t))
plt.show()
fig, ax1 = plt.subplots(figsize = (10, 3.5))
ax1.plot(data_filtered[y, x, :], color = "C1")
ax1.set_title("Signal: x = {x}, y = {y}".format(x=x, y=y))
ax1.set_ylabel("data filtered", color='C1')
ax2 = ax1.twinx()
ax2.plot(data_detrended[y, x, :], color = "C2")
ax2.set_ylabel("data detrended", color='C2')
plt.show()
# -
# <a id='section2.3'></a>
# ### Binning (spatial filtration)
# %time data_binned = binning(data_detrended, 9, "gaussian", mask)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data_detrended[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data detrended, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, data_binned[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data binned, t = {t}".format(t = t))
plt.show()
plt.figure(figsize = (10, 3.5))
plt.plot(data_detrended[y, x, :],
label = "data detrended", color = "C2")
plt.plot(data_binned[y, x, :],
label = "data binned", color = "C3")
plt.title("Signal: x = {x}, y = {y}".format(x=x, y=y))
plt.legend()
plt.show()
# -
# <a id='section2.4'></a>
# ### Rescaling (normalizing)
# %time data_rescaled = rescale(data_binned)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data_binned[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data binned, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, data_rescaled[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data rescaled, t = {t}".format(t = t))
plt.show()
fig, ax1 = plt.subplots(figsize = (10, 3.5))
ax1.plot(data_binned[y, x, :], color = "C3")
ax1.set_title("Signal: x = {x}, y = {y}".format(x=x, y=y))
ax1.set_ylabel("data binned", color='C3')
ax2 = ax1.twinx()
ax2.plot(data_rescaled[y, x, :], '.', color = "C4")
ax2.set_ylabel("data rescaled", color='C4')
plt.show()
# -
# <a id='section2.5'></a>
# ### Ensemble averaging
# Now let's try to find *cycle length* of our signal and apply ensemble averaging.
#
# In case of known *cycle length* (pacing cycle length (PCL) for example) just use it.
# +
cycle_length = 75.
signal = data_rescaled[y, x]
plt.figure(figsize = (6, 2))
plt.plot(signal)
plt.plot(np.roll(signal, 75))
plt.show()
# -
# So it looks like *cycle_length = 75* is OK.
# %time data_averaged = ensemble_average(data_rescaled, cycle_length)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data_rescaled[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data rescaled, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, data_averaged[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data averaged, t = {t}".format(t = t))
plt.show()
fig, ax1 = plt.subplots(figsize = (10, 3.5))
ax1.plot(data_rescaled[y, x, :], color = "C4")
ax1.set_title("Signal: x = {x}, y = {y}".format(x=x, y=y))
ax1.set_ylabel("data rescaled", color='C4')
ax2 = ax1.twinx()
ax2.plot(data_averaged[y, x, :], color = "C5")
ax2.set_ylabel("data averaged", color='C5')
plt.show()
# -
# <a id='section2.6'></a>
# ### Transform to phase
# %time phase = transform_to_phase(data_rescaled)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, data_rescaled[:, :, t])
plt.imshow(frame, cmap = "jet")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("data rescaled, t = {t}".format(t = t))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, phase[:, :, t])
plt.imshow(frame, cmap = "hsv")
plt.plot(x, y, '*w', ms = 10)
plt.colorbar()
plt.title("phase, t = {t}".format(t = t))
plt.show()
fig, ax1 = plt.subplots(figsize = (10, 3.5))
ax1.plot(data_rescaled[y, x, :], color = "C4")
ax1.set_title("Signal: x = {x}, y = {y}".format(x=x, y=y))
ax1.set_ylabel("data rescaled", color='C4')
ax2 = ax1.twinx()
ax2.plot(phase[y, x, :], color = "C6")
ax2.set_ylabel("phase", color='C6')
plt.show()
# -
# <a id='section3'></a>
# ## Mapping
# <a id='section3.1'></a>
# ### APD and Alternance maps
t_start, t_end = 200, 300
percentage = 60
# %time apd_map = calculate_APD_map(data_rescaled, t_start, t_end, percentage).astype(float)
t_start, t_end = 200, 400
percentage = 60
# %time alt_map = calculate_alternance_map(data_rescaled, t_start, t_end, percentage).astype(float)
# +
plt.figure(figsize = (10, 3.5))
plt.subplot(1, 2, 1)
frame = np.ma.masked_where(~mask, apd_map)
plt.imshow(frame, cmap = "jet")
plt.colorbar(label = "AP duration")
plt.title("APD{percentage} map".format(percentage = percentage))
plt.subplot(1, 2, 2)
frame = np.ma.masked_where(~mask, alt_map)
alt_abs = np.nanmax(np.abs(alt_map))
plt.imshow(frame, cmap = "bwr",
vmin = -alt_abs, vmax = alt_abs)
plt.colorbar(label = "AP duration difference")
plt.title("Alternance map (for APD{percentage})".format(percentage = percentage))
plt.show()
# -
# <a id='section3.2'></a>
# ### Activation time map
t_start, t_end = 300, 310
# %time act_map = calculate_activation_map(data_rescaled, t_start, t_end, 90.)
# +
plt.figure(figsize = (1.1 * 3.5, 3.5))
frame = np.ma.masked_where(~mask, act_map)
frame = np.flipud(frame)
plt.contourf(frame, cmap = 'rainbow_r',
origin = "lower")
plt.colorbar(label = "Activation time", fraction=0.1, pad=0.04)
plt.title("Activation map")
plt.axis('equal')
plt.show()
# -
# <a id='section3.3'></a>
# ### Phase singularity points (PS)
ps = phase_singularity_detection(phase[:, :, t])
frame = np.ma.masked_where(~mask, phase[:, :, t])
plt.imshow(frame, cmap = 'hsv')
plt.colorbar()
plt.plot(ps[:, 1], ps[:, 0], 'k.', ms = 10)
plt.plot(ps[:, 1], ps[:, 0], 'w.', ms = 5)
plt.title("PS")
plt.show()
| examples/WelcomeGuide_Part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # CS446/546 - Class Session 13 - Similarity and Hierarchical Clustering
#
# In this class session we are going to hierachically cluster (based on Sorensen-Dice similarity) vertices in a directed graph from a landmark paper on human gene regulation (Neph et al., Cell, volume 150, pages 1274-1286, 2012; see PDF on Canvas)
# Let's start by having a look at the Neph et al. data file, `neph_gene_network.txt`. It is in edge-list format, with no header and no "interaction" column. Just two columns, first column contains the "regulator gene" and the second column contains the "target gene":
#
# head neph_gene_network.txt
# AHR BCL6
# AHR BHLHE41
# AHR BPTF
# AHR CEBPA
# AHR CNOT3
# AHR CREB1
# Now let's load the packages that we will need for this exercise
suppressPackageStartupMessages(library(igraph))
# Using `read.table`, read the file `shared/neph_gene_network.txt`; name the two columns of the resulting data frame, `regulator` and `target`. Since there is no header, we will use `header=FALSE`:
edge_list_neph <- read.table("shared/neph_gene_network.txt",
header=FALSE,
sep="\t",
stringsAsFactors=FALSE,
col.names=c("regulator","target"))
# Load the edge-list data into a Graph object in igraph, using `graph_from_data_frame`. Make the graph undirected
neph_graph <- graph_from_data_frame(edge_list_neph, directed=FALSE)
summary(neph_graph)
# Get the adjacency matrix for the graph, using `get.adjacency`, and assign to matrix `g_matrix`
g_matrix <- get.adjacency(neph_graph)
# Compute the Sorensen-Dice similarity of this graph, and assign to similarity matrix `S`, using the igraph `similarity` function and specifying `method="dice"`
# Compute a distance matrix by calling `as.dist` on the object 1-S; assign to object `D`
# Perform average-linkage hierarchical clustering on this distance object, using `hclust` with `method=average`
# Plot the dendrogram using `plot` and make the labels really small using `cex=0.01`, using `plot`
| class09_similarity_R_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AdityaLoth/Mask-Detection-using-Google-Colab/blob/master/mask_detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8gSinmvLiaJ4" colab_type="text"
# # **FACE MASK DETECTION**
#
# During this pandemic people are coming with various new and creative ideas to tackle Covid-19. One such idea is to detect whether a person is wearing mask or not at public places or at their workplaces. This has been made possible with the help of Deep Learning and Computer Vision.
# This notebook took inspiration from one such [PROJECT](https://github.com/chandrikadeb7/Face-Mask-Detection) by <NAME>.This is a Google colab implementation of Face Mask detection project.
#
#
# This notebook is divided into 4 parts:
#
# 1. Setting up the environment
# 2. Training the model
# 3. Detection on Images
# 4. Detection on Videos(using WebCam)
#
#
# + [markdown] id="VNsWU7VGo9s2" colab_type="text"
#
#
# ---
# # PART 1: Setting up the environment
#
# + [markdown] id="1tGhPsvBpmrQ" colab_type="text"
# **STEP 1. Connect the Colab notebook to Google Drive**
#
# We're gonna map your Google Drive folder. This first step is the only one that will require your manual interaction every time you run your notebook.
#
# * Execute the following cell _(Click on Play button or press CTRL + ENTER)_ and click on the link to authorize your notebook to access to your Google Drive.
# * Paste the code Google will give to you and push `enter`
# + id="PlyJlElKs4tW" colab_type="code" colab={}
# This cell imports the drive library and mounts your Google Drive as a VM local drive. You can access to your Drive files
# using this path "/content/drive/My Drive/"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="V3xT9RU3tg_d" colab_type="text"
# **STEP 2. Creating a new directory named MASKDETECTION and changing to that directory**
#
# Run the following cells to create a folder named MASKDETECTION on your drive.
#
# We then change the directory to this folder.
#
# **NOTE:** You need to create the directory only once. So skip creating directory on further runs.
# + id="zOqxY4eLvWWB" colab_type="code" colab={}
# This creates a new directory in your drive
# %mkdir drive/My\ Drive/MASKDETECTION
# + id="3OqXnABIKohZ" colab_type="code" colab={}
#This makes the newly created directory your working directory
# %cd drive/My\ Drive/MASKDETECTION
# + [markdown] id="i06osx-Lw3cW" colab_type="text"
# **STEP 3. Downloading dataset and required files**
#
# Now we download the image dataset of with and without mask images into our folder. We also need weights and configuration file.
#
# **NOTE:** Remember that you need to successfully download these files only once. You can skip this step on futher runs.
# + id="jB31VWwq3DTR" colab_type="code" colab={}
# !wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --quiet --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1P9YgcPTZNufjC45YhtIJoGMz5Y4KtJfo' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1P9YgcPTZNufjC45YhtIJoGMz5Y4KtJfo" -O FILE && rm -rf /tmp/cookies.txt
# + id="pJoAi336AZd2" colab_type="code" colab={}
# !unzip FILE
# + [markdown] id="wvcUGa3rA9i2" colab_type="text"
# **STEP 4. Importing required modules**
# + id="6rl4D9CEzdvh" colab_type="code" colab={}
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.applications import MobileNetV2
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.models import load_model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
import imutils
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import argparse
import os
import cv2
from google.colab.patches import cv2_imshow
from imutils.video import VideoStream
import time
import sys
# + [markdown] id="J9RCNIxJwfwo" colab_type="text"
# **Step 5. Importing the face detection model**
#
# Since our main focus here is to detect masks we will not go into creating a face detection model. We simply import a pretrained model res10_300x300_ssd_iter_140000.caffemodel.
# + id="-ZItzaMLpCc-" colab_type="code" colab={}
#Setting path to configuration file
prototxtPath = os.path.sep.join(["face_detector", "deploy.prototxt"])
#setting path to weights
weightsPath = os.path.sep.join(["face_detector",
"res10_300x300_ssd_iter_140000.caffemodel"])
#Creating the network to detect faces
net = cv2.dnn.readNet(prototxtPath, weightsPath)
# + [markdown] id="ep1qSlsHEqIY" colab_type="text"
#
#
#
#
#
#
# ---
#
#
#
#
#
# # PART 2: Training the model
#
#
# + [markdown] id="DfSP0XOxXCoJ" colab_type="text"
# **Step 1. Converting the images to array and storing their labels(with mask or withour mask)**
# + id="zfFk99I6MrIF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 74} outputId="04f14379-3e0f-481e-965e-08faf885716f"
imagePaths = list(paths.list_images("dataset"))
data = []
labels = []
# loop over the image paths
for imagePath in imagePaths:
# extract the class label from the filename
label = imagePath.split(os.path.sep)[-2]
# load the input image (224x224) and preprocess it
image = load_img(imagePath, target_size=(224, 224))
image = img_to_array(image)
image = preprocess_input(image)
# update the data and labels lists, respectively
data.append(image)
labels.append(label)
data = np.array(data, dtype="float32")
labels = np.array(labels)
# + id="LLjsItmfPKsx" colab_type="code" colab={}
# perform one-hot encoding on the labels
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
labels = to_categorical(labels)
# + [markdown] id="wocmrbz6ZjIP" colab_type="text"
# **Step 2. Dividing the dataset for training and testing purposes**
# + id="xj3pHKFUVxfo" colab_type="code" colab={}
(trainX, testX, trainY, testY) = train_test_split(data, labels,
test_size=0.20, stratify=labels, random_state=42)
# + [markdown] id="q7rji4J6Z0vz" colab_type="text"
# **Step 3. Creating the model**
#
# We will use MobileNetv2 as our base model and then apply further layers over it to create a final model.
# + id="LCmnVgzLaCf5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 93} outputId="87f610e5-7e8b-4cd2-bfd2-653e3e571233"
baseModel = MobileNetV2(weights="imagenet", include_top=False,
input_tensor=Input(shape=(224, 224, 3)))
# construct the head of the model that will be placed on top of the
# the base model
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(7, 7))(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(128, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(2, activation="softmax")(headModel)
# place the head FC model on top of the base model (this will become
# the actual model we will train)
model = Model(inputs=baseModel.input, outputs=headModel)
# loop over all layers in the base model and freeze them so they will
# *not* be updated during the first training process
for layer in baseModel.layers:
layer.trainable = False
# + [markdown] id="n18D8AfBaYLq" colab_type="text"
# Compiling our model
# + id="oRxQnai4YHXd" colab_type="code" colab={}
LR = 1e-4 #learning rate
EPOCHS = 20 #no. of epochs
BS = 32 #batch size
#We will use Adam Optimizer for our model
opt= Adam(lr=LR, decay=LR / EPOCHS)
#To compile the model we use 'binary crossentropy' as loss function and set the metrics to accuracy
model.compile(loss="binary_crossentropy", optimizer=opt,
metrics=["accuracy"])
# + [markdown] id="sKO9yvoHeCYE" colab_type="text"
# Construct the training image generator for data augmentation
#
# + id="RCb88i0-ew0q" colab_type="code" colab={}
aug = ImageDataGenerator(
rotation_range=20,
zoom_range=0.15,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.15,
horizontal_flip=True,
fill_mode="nearest")
# + [markdown] id="rziv9qM5fmfa" colab_type="text"
# Training our model
# + id="J8N4uMW6Ye-P" colab_type="code" colab={}
H = model.fit(
aug.flow(trainX, trainY, batch_size=BS),
steps_per_epoch=len(trainX) // BS,
validation_data=(testX, testY),
validation_steps=len(testX) // BS,
epochs=EPOCHS)
# + [markdown] id="ULcA2MVXfxPk" colab_type="text"
# Make predictions on the testing set
# + id="P0U5phmkowwM" colab_type="code" colab={}
preds = model.evaluate(testX, testY)
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
# + [markdown] id="5evDd92lf8hs" colab_type="text"
# Serialize the model to disk
# + id="iTlD5_aIxZhs" colab_type="code" colab={}
#Here "model" is the name of model. You can change it accordingly.
model.save("model", save_format="h5")
# + [markdown] id="5b7hRAAtgJE8" colab_type="text"
#
#
#
# ---
#
#
# # PART 3. Detection on Images
# + [markdown] id="SYPJy0HpgvGT" colab_type="text"
# Let us load the model which we created.
# + id="xhte2lIRxrho" colab_type="code" colab={}
#This will load the pretrained model named 'mask_detector'. You can replace it with your model name.
model = load_model("mask_detector.model")
# + [markdown] id="DGG-WILjiHnB" colab_type="text"
# Importing the image for mask detection
# + id="Ao3vBleYyN1e" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 93} outputId="29ebb185-6f3f-45f9-e383-6c7b398b234c"
from google.colab import files
uploaded = files.upload()
for name, data in uploaded.items():
with open(name, 'wb') as f:
f.write(data)
print ('saved file', name)
image = cv2.imread(name)
orig = image.copy()
(h, w) = image.shape[:2]
# + [markdown] id="MYD1I9WLibPT" colab_type="text"
# Construct a blob from the image and then pass it through the imported face detector network to obtain face detections
# + id="ne400mbLzLdK" colab_type="code" colab={}
blob = cv2.dnn.blobFromImage(image, 1.0, (300, 300),
(104.0, 177.0, 123.0))
net.setInput(blob)
detections = net.forward()
# + [markdown] id="Y641TS8mjQSO" colab_type="text"
# Looping over the detections and labelling them as 'Mask' or 'No mask'
# + id="n8Wmo4T20RJb" colab_type="code" colab={}
for i in range(0, detections.shape[2]):
# extract the confidence (i.e., probability) associated with
# the detection
confidence = detections[0, 0, i, 2]
# filter. out weak detections by ensuring the confidence is
# greater than the minimum confidence
if confidence > 0.5:
# compute the (x, y)-coordinates of the bounding box for
# the object
box = detections[0, 0, i, 3:7] * np.array([w, h, w, h])
(startX, startY, endX, endY) = box.astype("int")
# ensure the bounding boxes fall within the dimensions of
# the frame
(startX, startY) = (max(0, startX), max(0, startY))
(endX, endY) = (min(w - 1, endX), min(h - 1, endY))
# extract the face ROI, convert it from BGR to RGB channel
# ordering, resize it to 224x224, and preprocess it
face = image[startY:endY, startX:endX]
face = cv2.cvtColor(face, cv2.COLOR_BGR2RGB)
face = cv2.resize(face, (224, 224))
face = img_to_array(face)
face = preprocess_input(face)
face = np.expand_dims(face, axis=0)
# pass the face through the model to determine if the face
# has a mask or not
(mask, withoutMask) = model.predict(face)[0]
# determine the class label and color we'll use to draw
# the bounding box and text
label = "Mask" if mask > withoutMask else "No Mask"
color = (0, 255, 0) if label == "Mask" else (0, 0, 255)
# include the probability in the label
label = "{}: {:.2f}%".format(label, max(mask, withoutMask) * 100)
# display the label and bounding box rectangle on the output
# frame
cv2.putText(image, label, (startX, startY - 10),
cv2.FONT_HERSHEY_SIMPLEX, 0.45, color, 2)
cv2.rectangle(image, (startX, startY), (endX, endY), color, 2)
# + [markdown] id="PA8ZA5sjllXg" colab_type="text"
# Displaying the final image
# + id="2xnho_vz0waB" colab_type="code" colab={}
cv2_imshow(image)
# + [markdown] id="7rFgqWH9mHS6" colab_type="text"
#
#
# ---
#
# # Part 4. Detection on videos (using WebCam)
#
# + [markdown] id="mlKkTYjxm3Xr" colab_type="text"
# A major challange which I faced was the real time detection on objects(masks).
# Colab codes executes on a VM that doesn't have a webcam attached. So real time object detection is not possible here. But with the help of Javascript we can use the webcam to capture images and videos and use them for further actions. Though it is not real time but it works.
# + [markdown] id="sDO7KmVSrTRk" colab_type="text"
# First of all we need to set the webcam with the following code. Your camera will start if it is working.
# Clear the output afterwards to stop streaming.
# + id="j_VGKBWe7iBI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="1ccac090-8d47-4478-bb7e-1de917b3b73c"
# !pip install ffmpeg-python
from IPython.display import HTML, Javascript, display
from google.colab.output import eval_js
from base64 import b64decode
import numpy as np
import io
import ffmpeg
video_file= '/content/drive/My Drive/MASKDETECTION/train.mp4'
VIDEO_HTML = """
<script>
var my_div = document.createElement("DIV");
var my_p = document.createElement("P");
var my_btn = document.createElement("BUTTON");
var my_btn_txt = document.createTextNode("Press to start recording");
my_btn.appendChild(my_btn_txt);
my_div.appendChild(my_btn);
document.body.appendChild(my_div);
var base64data = 0;
var reader;
var recorder, videoStream;
var recordButton = my_btn;
var handleSuccess = function(stream) {
videoStream = stream;
var options = {
mimeType : 'video/webm;codecs=vp9'
};
recorder = new MediaRecorder(stream, options);
recorder.ondataavailable = function(e) {
var url = URL.createObjectURL(e.data);
var preview = document.createElement('video');
preview.controls = true;
preview.src = url;
document.body.appendChild(preview);
reader = new FileReader();
reader.readAsDataURL(e.data);
reader.onloadend = function() {
base64data = reader.result;
}
};
recorder.start();
};
recordButton.innerText = "Recording... press to stop";
navigator.mediaDevices.getUserMedia({video: true}).then(handleSuccess);
function toggleRecording() {
if (recorder && recorder.state == "recording") {
recorder.stop();
videoStream.getVideoTracks()[0].stop();
recordButton.innerText = "Original recording"
}
}
function sleep(ms) {
return new Promise(resolve => setTimeout(resolve, ms));
}
var data = new Promise(resolve=>{
recordButton.onclick = ()=>{
toggleRecording()
sleep(2000).then(() => {
// wait 2000ms for the data to be available
resolve(base64data.toString())
});
}
});
</script>
"""
def start_webcam():
js = Javascript('''
async function startWebcam() {
const div = document.createElement('div');
const video = document.createElement('video');
video.style.display = 'block';
const stream = await navigator.mediaDevices.getUserMedia({video: true});
document.body.appendChild(div);
div.appendChild(video);
video.srcObject = stream;
await video.play();
// Resize the output to fit the video element.
google.colab.output.setIframeHeight(document.documentElement.scrollHeight, true);
return;
}
''')
display(js)
data = eval_js('startWebcam()')
start_webcam()
def get_video():
display(HTML(VIDEO_HTML))
data = eval_js("data")
binary = b64decode(data.split(',')[1])
return binary
# + [markdown] id="b83Q4ZEtsKti" colab_type="text"
# Creating a function to detect and predict masks
# + id="nh3ljMsm9MkP" colab_type="code" colab={}
def detect_and_predict_mask(frame, faceNet, maskNet):
# grab the dimensions of the frame and then construct a blob
# from it
(h, w) = frame.shape[:2]
blob = cv2.dnn.blobFromImage(frame, 1.0, (300, 300),
(104.0, 177.0, 123.0))
# pass the blob through the network and obtain the face detections
faceNet.setInput(blob)
detections = faceNet.forward()
# initialize our list of faces, their corresponding locations,
# and the list of predictions from our face mask network
faces = []
locs = []
preds = []
# loop over the detections
for i in range(0, detections.shape[2]):
# extract the confidence (i.e., probability) associated with
# the detection
confidence = detections[0, 0, i, 2]
# filter out weak detections by ensuring the confidence is
# greater than the minimum confidence
if confidence > 0.5:
# compute the (x, y)-coordinates of the bounding box for
# the object
box = detections[0, 0, i, 3:7] * np.array([w, h, w, h])
(startX, startY, endX, endY) = box.astype("int")
# ensure the bounding boxes fall within the dimensions of
# the frame
(startX, startY) = (max(0, startX), max(0, startY))
(endX, endY) = (min(w - 1, endX), min(h - 1, endY))
# extract the face ROI, convert it from BGR to RGB channel
# ordering, resize it to 224x224, and preprocess it
face = frame[startY:endY, startX:endX]
face = cv2.cvtColor(face, cv2.COLOR_BGR2RGB)
face = cv2.resize(face, (224, 224))
face = img_to_array(face)
face = preprocess_input(face)
# add the face and bounding boxes to their respective
# lists
faces.append(face)
locs.append((startX, startY, endX, endY))
# only make a predictions if at least one face was detected
if len(faces) > 0:
# for faster inference we'll make batch predictions on *all*
# faces at the same time rather than one-by-one predictions
# in the above `for` loop
faces = np.array(faces, dtype="float32")
preds = maskNet.predict(faces, batch_size=32)
# return a 2-tuple of the face locations and their corresponding
# locations
return (locs, preds)
# + [markdown] id="TW38X-xCsVP7" colab_type="text"
# Load the model which we created.
# + id="FYBVZjzOAFeP" colab_type="code" colab={}
maskNet = load_model("mask_detector.model")
# + [markdown] id="xAQ-mig-tgWZ" colab_type="text"
# Capturing the video and detecting faces with or without mask
# + id="WXkLsgx4Awh7" colab_type="code" colab={}
vid = get_video()
with open(video_file, 'wb') as f:
f.write(vid)
print("Recorded the video")
video = cv2.VideoCapture('train.mp4')
out = cv2.VideoWriter('out.avi', cv2.VideoWriter_fourcc(*'XVID'), 20.0, (640,480))
# print(type(out))
while True:
# grab the frame from the threaded video stream and resize it
# to have a maximum width of 400 pixels
# frame = vs.read()
# frame = imutils.resize(frame, width=400)
boool, frame = video.read()
if(boool==False):
break
w, h, c = frame.shape
#syntax: cv2.resize(img, (width, height))
frame = cv2.resize(frame,(h, w))
# detect faces in the frame and determine if they are wearing a
# face mask or not
locs, preds = detect_and_predict_mask(frame, net, maskNet)
# loop over the detected face locations and their corresponding
# locations
for (box, pred) in zip(locs, preds):
# unpack # frame = vs.read()
# frame = imutils.resize(frame, width=400)the bounding box and predictions
(startX, startY, endX, endY) = box
(mask, withoutMask) = pred
# determine the class label and color we'll use to draw
# the bounding box and text
label = "Mask" if mask > withoutMask else "No Mask"
color = (0, 255, 0) if label == "Mask" else (0, 0, 255)
# include the probability in the label
label = "{}: {:.2f}%".format(label, max(mask, withoutMask) * 100)
# display the label and bounding box rectangle on the output
# frame
cv2.putText(frame, label, (startX, startY - 10),
cv2.FONT_HERSHEY_COMPLEX, 0.55, color, 2)
cv2.rectangle(frame, (startX, startY), (endX, endY), color, 2)
# show the output frame
out.write(frame)
# cv2_imshow(frame)
out.release()
# do a bit of cleanup
cv2.destroyAllWindows()
print("Detection complete.Now converting")
# !ffmpeg -i out.avi output.mp4 -loglevel quiet -y
from IPython.display import HTML
from base64 import b64encode
mp4 = open('output.mp4','rb').read()
data_url = "data:video/mp4;base64," + b64encode(mp4).decode()
HTML("""
<video controls>
<source src="%s" type="video/mp4">
</video>
""" % data_url)
# + [markdown] id="-glkNJBivaZ8" colab_type="text"
#
#
# ---
#
# # SOURCES
#
#
# 1. chandrikadeb7/Face-Mask-Detection repo [github](https://github.com/chandrikadeb7/Face-Mask-Detection)
# 2. [Tensorflow](https://www.tensorflow.org)
#
#
#
#
#
#
#
# ## **THANK YOU!**
| mask_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
# US data into dataframe
df = pd.read_csv('./output/US_cleaned_data_2018')
df.head()
# # Trending Popularity by Category
barp = df.groupby("category_id").count().sort_values("title").plot(kind='barh', y='title')
barp.set_title('Video Count by Category')
# # Trending Popularity by Channel
# 2207 channels made it to trending during 2017-2018 with ESPN as the most popular
df.get("channel_title").value_counts()
# # How Many Views Trending Videos Get
# Views range from 549 to 225,211,923
df['views'].describe()
# Very heavily skewed right
df['views'].skew()
# Very heavily leptokurtic
df['views'].kurtosis()
# Q1 and Q3 are roughly 242,329 and 1,823,157
# ## Views Boxplot
df.boxplot(['views'], vert = False).set_title('Boxplot of Views for All Trending Videos')
1823157 + (1.5 * (1823157 - 242329))
# Outliers begin at 4,194,399
df.boxplot(['views'], vert = False).set_title('Boxplot of Views Zoomed in to Zero to Five Million')
plt.xlim(0, 5000000)
plt.show()
# ## Views Histogram
df.hist(column='views').set_title('Video Count by Views')
mill = pd.DataFrame({})
mill['views'] = df['views']
indexNames = mill[mill['views'] >= 10000000].index
mill.drop(indexNames , inplace=True)
mill.hist(column='views')
hunthou = pd.DataFrame({})
hunthou['views'] = df['views']
indexNames = hunthou[hunthou['views'] >= 1000000 ].index
hunthou.drop(indexNames , inplace=True)
hunthou.hist(column='views')
# # Trending Like to Dislike Ratios
(df.dislikes == 0).sum()
df['dislikes'].describe()
(df.likes == 0).sum()
df['likes'].describe()
ratios = pd.DataFrame({})
ratios['likes'] = df['likes']
ratios['dislikes'] = df['dislikes']
indexNames = ratios[ratios['dislikes'] == 0].index
ratios.drop(indexNames , inplace=True)
ratios['ratio'] = ratios['likes'] / ratios['dislikes']
ratios['ratio'].describe()
df.boxplot(['likes'], vert = False)
df.boxplot(['dislikes'], vert = False)
ratios.boxplot(['ratio'], vert = False).set_title('Like to Dislike Ratios')
ratios.boxplot(['ratio'], vert = False).set_title('Like to Dislike Ratios Zoomed in to Zero to Two Hundred')
plt.xlim(0, 200)
plt.show()
# 576 trending videos have fewer likes than dislikes
(ratios.ratio < 1).sum()
# # Trending Comment Count
# 633 trending videos with comments disabled (from data set with 'comments_disabled' column)
(df.comment_count == 0).sum()
df['comment_count'].describe()
comments = pd.DataFrame({})
comments['comment_count'] = df['comment_count']
comments['comments_disabled'] = df['comments_disabled']
indexNames = comments[comments['comments_disabled']].index
comments.drop(indexNames , inplace=True)
comments['comment_count'].describe()
# # Days from Publish to Trending
| initial exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import cvxpy as cp
# %matplotlib inline
# 
# # Formulation
#
# $p \in \mathbb{R}^{n*n}$
#
# $\sum_i p(i,j) = M^{(1)}[j]$
#
# $\sum_j p(i,j) = M^{(0)}[i]$
#
# $E[R_1 R_2] = \rho \sigma_1 \sigma_2 + \mu_1 \mu_2$
#
# $\sum_{i,j} p(i,j) r[i]r[j] = \rho \sigma_1 \sigma_2 + \mu_1 \mu_2$
# # Problem data
# +
mu = [8, 20]
s = [6, 17.5]
rho = -0.25
n = 100
r = np.linspace(-30, 70, n)
marginal = [None, None]
for i in range(2):
mi = np.exp(-((r-mu[i])**2)/(2*(s[i]**2)))
mi /= mi.sum()
marginal[i] = mi
r_mul_ij = r.reshape(-1,1) @ r.reshape(1,-1)
r_sum_ij = r.reshape(-1,1) + r.reshape(1,-1)
# -
# # Solving
# +
p = cp.Variable((n,n), nonneg = True)
p_loss = cp.sum(p[r_sum_ij <= 0])
obj = cp.Maximize(p_loss)
C = [
cp.sum(p, axis = 0) == marginal[0],
cp.sum(p, axis = 1) == marginal[1],
# correlation
cp.sum(cp.multiply(r_mul_ij, p)) == rho*s[0]*s[1] + mu[0]*mu[1]
]
prob = cp.Problem(obj, C)
prob.solve()
assert prob.status == cp.OPTIMAL
print('max p_loss = ',prob.value)
p_ans = p.value
# +
# ((p_ans*r_mul_ij).sum() - mu[0]*mu[1])/(s[0]*s[1])
# -
fig, ax = plt.subplots(figsize = (10,10))
cs = ax.contourf(r, r, p_ans, np.linspace(1e-3, p_ans.max(), 10))
# ax.clabel(cs, fontsize=9, inline=1)
ax.grid()
ax.set_xlabel('R1')
ax.set_ylabel('R2')
cbar = fig.colorbar(cs)
plt.show()
| Statistical Estimation/Worst-case probability of loss.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # GHZ Game
#
# The **GHZ Game** quantum kata is a series of exercises designed
# to get you familiar with the GHZ game.
#
# In it three players (Alice, Bob and Charlie) try to win the following game:
#
# Each of them is given a bit (r, s and t respectively), and
# they have to return new bits (a, b and c respectively) so
# that r ∨ s ∨ t = a ⊕ b ⊕ c. The input bits will have
# zero or two bits set to true and three or one bits set to false.
# The trick is, the players can not communicate during the game.
#
# * You can read more about the GHZ game in the [lecture notes](https://cs.uwaterloo.ca/~watrous/CPSC519/LectureNotes/20.pdf) by <NAME>.
# * Another description can be found in the [lecture notes](https://staff.fnwi.uva.nl/m.walter/physics491/lecture1.pdf) by <NAME>.
#
# Each task is wrapped in one operation preceded by the description of the task.
# Your goal is to fill in the blank (marked with the `// ...` comments)
# with some Q# code that solves the task. To verify your answer, run the cell using Ctrl/⌘+Enter.
#
# To begin, first prepare this notebook for execution (if you skip this step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells):
%package Microsoft.Quantum.Katas::0.11.2003.3107
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
#
# ## Part I. Classical GHZ
#
# ### Task 1.1. Win Condition
# **Inputs:**
#
# 1. Alice, Bob and Charlie's input bits (r, s and t), stored as an array of length 3,
#
# 2. Alice, Bob and Charlie's output bits (a, b and c), stored as an array of length 3.
#
# The input bits will have zero or two bits set to true.
#
# **Output:**
# True if Alice, Bob and Charlie won the GHZ game, that is, if r ∨ s ∨ t = a ⊕ b ⊕ c, and false otherwise.
# +
%kata T11_WinCondition_Test
function WinCondition (rst : Bool[], abc : Bool[]) : Bool {
// ...
fail "Task 1.1 not implemented yet";
}
# -
# ### Task 1.2. Random classical strategy
#
# **Input:** The input bit for one of the players (r, s or t).
#
# **Output:** A random bit that this player will output (a, b or c).
#
# If all players use this strategy, they will win about 50% of the time.
# +
%kata T12_RandomClassical_Test
operation RandomClassicalStrategy (input : Bool) : Bool {
// ...
fail "Task 1.2 not implemented yet";
}
# -
# ### Task 1.3. Best classical strategy
#
# **Input:** The input bit for one of the players (r, s or t).
#
# **Output:** A bit that this player will output (a, b or c) to maximize their chance of winning.
#
# All players will use the same strategy.
# The best classical strategy should win about 75% of the time.
# +
%kata T13_BestClassical_Test
operation BestClassicalStrategy (input : Bool) : Bool {
// ...
fail "Task 1.3 not implemented yet";
}
# -
# ### Task 1.4. Referee classical GHZ game
#
# **Inputs:**
#
# 1. An operation which implements a classical strategy (i.e., takes an input bit and produces an output bit),
#
# 2. An array of 3 input bits that should be passed to the players.
#
# **Output:**
# An array of 3 bits that will be produced if each player uses this strategy.
# +
%kata T14_PlayClassicalGHZ_Test
operation PlayClassicalGHZ (strategy : (Bool => Bool), inputs : Bool[]) : Bool[] {
// ...
fail "Task 1.4 not implemented yet";
}
# -
# ## Part II. Quantum GHZ
#
# In the quantum version of the game, the players still can not
# communicate during the game, but they are allowed to share
# qubits from an entangled triple before the start of the game.
#
# ### Task 2.1. Entangled triple
#
# **Input:** An array of three qubits in the $|000\rangle$ state.
#
# **Goal:** Create the entangled state $|\Phi\rangle = \frac{1}{2} \big(|000\rangle - |011\rangle - |101\rangle - |110\rangle \big)$ on these qubits.
# +
%kata T21_CreateEntangledTriple_Test
operation CreateEntangledTriple (qs : Qubit[]) : Unit {
// ...
fail "Task 2.1 not implemented yet";
}
# -
# ### Task 2.2. Quantum strategy
#
# **Inputs:**
#
# 1. The input bit for one of the players (r, s or t),
#
# 2. That player's qubit of the entangled triple shared between the players.
#
# **Goal:** Measure the qubit in the Z basis if the bit is 0 (false), or the X basis if the bit is 1 (true), and return the result.
#
# The state of the qubit after the operation does not matter.
# +
%kata T22_QuantumStrategy_Test
operation QuantumStrategy (input : Bool, qubit : Qubit) : Bool {
// ...
fail "Task 2.2 not implemented yet";
}
# -
# ### Task 2.3. Play the GHZ game using the quantum strategy
#
# **Input:** Operations that return Alice, Bob and Charlie's output bits (a, b and c) based on
# their quantum strategies and given their respective qubits from the entangled triple.
# The players have already been told what their starting bits (r, s and t) are.
#
# **Goal:** Return an array of players' output bits (a, b and c).
# +
%kata T23_PlayQuantumGHZ_Test
operation PlayQuantumGHZ (strategies : (Qubit => Bool)[]) : Bool[] {
// ...
fail "Task 2.3 not implemented yet";
}
| GHZGame/GHZGame.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Register ONNX model and deploy as webservice
#
# Following this notebook, you will:
#
# - Learn how to register an ONNX in your Azure Machine Learning Workspace.
# - Deploy your model as a web service in an Azure Container Instance.
# ## Prerequisites
#
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the [configuration notebook](../../../configuration.ipynb) to install the Azure Machine Learning Python SDK and create a workspace.
# +
import azureml.core
# Check core SDK version number.
print('SDK version:', azureml.core.VERSION)
# -
# ## Initialize workspace
#
# Create a [Workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace%28class%29?view=azure-ml-py) object from your persisted configuration.
# + tags=["create workspace"]
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
# -
# ## Register model
#
# Register a file or folder as a model by calling [Model.register()](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py#register-workspace--model-path--model-name--tags-none--properties-none--description-none--datasets-none--model-framework-none--model-framework-version-none--child-paths-none-). For this example, we have provided a trained ONNX MNIST model(`mnist-model.onnx` in the notebook's directory).
#
# In addition to the content of the model file itself, your registered model will also store model metadata -- model description, tags, and framework information -- that will be useful when managing and deploying models in your workspace. Using tags, for instance, you can categorize your models and apply filters when listing models in your workspace. Also, marking this model with the scikit-learn framework will simplify deploying it as a web service, as we'll see later.
# + tags=["register model from file"]
from azureml.core import Model
model = Model.register(workspace=ws,
model_name='mnist-sample', # Name of the registered model in your workspace.
model_path='mnist-model.onnx', # Local ONNX model to upload and register as a model.
model_framework=Model.Framework.ONNX , # Framework used to create the model.
model_framework_version='1.3', # Version of ONNX used to create the model.
description='Onnx MNIST model')
print('Name:', model.name)
# -
# ## Deploy model
#
# Deploy your model as a web service using [Model.deploy()](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py#deploy-workspace--name--models--inference-config--deployment-config-none--deployment-target-none-). Web services take one or more models, load them in an environment, and run them on one of several supported deployment targets.
#
# For this example, we will deploy the ONNX model to an Azure Container Instance (ACI).
# ### Use a default environment (for supported models)
#
# The Azure Machine Learning service provides a default environment for supported model frameworks, including ONNX, based on the metadata you provided when registering your model. This is the easiest way to deploy your model.
#
# **Note**: This step can take several minutes.
# +
from azureml.core import Webservice
from azureml.exceptions import WebserviceException
service_name = 'onnx-mnist-service'
# Remove any existing service under the same name.
try:
Webservice(ws, service_name).delete()
except WebserviceException:
pass
service = Model.deploy(ws, service_name, [model])
service.wait_for_deployment(show_output=True)
# -
# After your model is deployed, perform a call to the web service.
# +
import requests
headers = {'Content-Type': 'application/json', 'Accept': 'application/json'}
if service.auth_enabled:
headers['Authorization'] = 'Bearer '+ service.get_keys()[0]
elif service.token_auth_enabled:
headers['Authorization'] = 'Bearer '+ service.get_token()[0]
scoring_uri = service.scoring_uri
print(scoring_uri)
with open('onnx-mnist-predict-input.json', 'rb') as data_file:
response = requests.post(
scoring_uri, data=data_file, headers=headers)
print(response.status_code)
print(response.elapsed)
print(response.json())
# -
# When you are finished testing your service, clean up the deployment.
service.delete()
| how-to-use-azureml/deployment/onnx/onnx-model-register-and-deploy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
#
# <table>
# <tr>
# <td width=15%><img src="./img/UGA.png"></img></td>
# <td><center><h1>Introduction to Python for Data Sciences</h1></center></td>
# <td width=15%><a href="http://www.iutzeler.org" style="font-size: 16px; font-weight: bold"><NAME></a><br/> Fall. 2018 </td>
# </tr>
# </table>
#
# <br/><br/><div id="top"></div>
#
# <center><a style="font-size: 40pt; font-weight: bold">Chap. 2 - Python for Scientific Computing </a></center>
#
# <br/>
#
# # ``2. Examples in Data Science``
#
# ---
# <a href="#style"><b>Package check and Styling</b></a><br/><br/><b>Outline</b><br/><br/>
# a) <a href="#sigEx"> Compressed Sensing </a><br/>
# ## <a id="sigEx"> a) Compressed Sensing </a>
#
# <p style="text-align: right; font-size: 10px;"><a href="#top">Go to top</a></p>
import numpy as np
import scipy.fftpack as spfft
import matplotlib.pyplot as plt
# %matplotlib inline
# In this example (borrowed from <NAME>. “Data-driven modeling and scientific computing: Methods for Integrating Dynamics of Complex Sys-tems and Big Data.” (2013)), we will create an artificial periodic signal, sample 5% of it, and try to reconstruct the original signal using the prior knowledge that the signal should be *simple* in the cosine (frequencial) domain.
#
# In practice:
# * The original temporal signal $x$ will be generated
# * Our observed subsampled signal $y$ will be obtained by randomly sampling 5% of the original signal, we will see that standard interpolation may be bad
# * The discrete cosine tranform (DCT) matrix $T$ will be obtained from <tt>scipy.fftpack</tt> (from the frequencial coefficients $Y$, the matrix $T$, s.t. $T_{i,j} \propto \cos(2\pi f_j t_i)$, enables to retreive the temporal signal $y = TY$)
# * We will recover an estimate $\hat{Y}$ of the DCT $X$ of $x$ by minimizing $\|y - TY\|$ over $Y$ under the contraint that $Y$ should be somehow sparse
# * From the coefficient $\hat{Y}$, we can oversample to get an estimate $\hat{x}$
#
# This is one dimensional **compressed sensing**.
# ### Problem Setup
#
# First, create a signal of two sinusoids.
# +
n = 200 # number of original discrete points
tMax = 2.0 # total signal time in sec
t = np.linspace(0, tMax , n)
f1 = 5.75; f2 = 15.0;
x = 2*np.sin(f1 * np.pi * t) + 1.5*np.sin(f2 * np.pi * t)
X = spfft.dct(x, norm='ortho')
f = (np.arange(n)+0.5)/tMax
# +
plt.subplot(121)
plt.plot(t,x)
plt.ylabel("Signal")
plt.subplot(122)
plt.plot(f,X)
plt.xlabel("frequency (Hz)")
# +
# extract small sample of signal
m = int(n*0.15) # 5% sample
iS = np.sort(np.random.choice(n, m, replace=False)) # random sample of indices
tS = t[iS]
y = x[iS]
# -
plt.plot(t,x,label="signal")
plt.scatter(tS,y , c = 'red', marker = 'o',label="observations")
plt.xlabel("time (s)")
plt.legend()
plt.xlim([0,tMax])
# ### Interpolation
#
#
# Scipy posseses an [interpolation module](https://docs.scipy.org/doc/scipy/reference/tutorial/interpolate.html) that enables direct interpolation using splines.
# +
import scipy.interpolate as spinter
tInt = np.clip(t, min(tS), max(tS)) # We can only interpolate between the sampled values
intL = spinter.interp1d(tS, y , kind = "linear")
xL = intL(tInt)
intC = spinter.interp1d(tS, y , kind = "cubic")
xC = intC(tInt)
# -
plt.plot(t,x ,c='0.7',label="true signal")
plt.scatter(tS,y , c = 'red', marker = 'o',label="observations")
plt.plot(tInt,xL ,c='m',label="linear interpolation")
plt.plot(tInt,xC ,c='g',label="cubic splines")
plt.xlabel("time (s)")
plt.xlim([0,tMax])
plt.ylim([-10,10])
plt.legend(loc="lower left")
# We see that interpolation is bad as we have *very few* points from a *complicated* signal in the temporal space. In Compressed sensing, we are going to use the signal *simplicity* in the cosine domain.
# ### Compressed Sensing
#
#
# The inverse discrete cosine tranform (IDCT) matrix $T$ will be obtained from <tt>scipy.fftpack</tt> (from the frequencial coefficients $Y$, the matrix $T$, s.t. $T_{i,j} \propto \cos(2\pi f_j t_i)$, enables to retreive the temporal signal $y = TY$).
# create idct matrix operator
T = spfft.idct(np.identity(n), norm='ortho', axis=0)
T = T[iS]
# We now want to recover an estimate $\hat{Y}$ of the DCT $X$ of $x$ by minimizing $\|y - TY\|$ over $Y$ under the contraint that $Y$ should be somehow sparse.
#
# To address this problem, two popular solutions are:
# * the **Dantzig Selector**, introduced in *<NAME> and <NAME> "The Dantzig selector: Statistical estimation when $p$ is much larger than $n$". The Annals of Statistics, 2007* can be used to compute $\hat{Y}$ in the case of an overparametrized problem, i.e. when the dimension $n$ of $\hat{Y}$ is well greater than the dimension $m$ of the observation $y$. <br/> In that case, the estimator $\widehat{Y}_{DS}$ is the solution of the optimization problem
# $$
# \widehat{Y}_{DS} \in \arg\min_{Y\in \mathbb{R}^n} \left\{\|Y\|_1,\;\mbox{with}\;\|T^T(TY-y)\|_\infty\leq \kappa\sigma\right\},
# $$
# where $\sigma>0$ is an *hyper-parameter*.
#
# * the **Least Absolute Shrinkage and Selection Operator** or **lasso**, introduced in *<NAME> "Regression shrinkage and selection via the lasso", Journal of the Royal Statistical Society, 1996* can also be used to estimate $Y$. <br/> The estimator $\widehat{Y}_{L}$ is the solution of the optimization problem
# $$
# \widehat{\theta}_{L} \in \arg\min_{\theta\in \mathbb{R}^n} \left\{ \|X\theta - y\|_2^2 + \kappa \sigma \|\theta\|_1 \right\},
# $$
# where $\sigma>0$ is an *hyper-parameter*.
# <div id="warn">
# <b>Warning:</b> Below is the code for both estimators using <tt>cvxopt</tt> for solving linear or quadratic reformulated problems. <b><i>you are strongly encouraged to take a deeper look at these functions.</i></b>
# </div>
# +
from cvxopt import matrix, solvers
from scipy.stats import norm
import numpy as np
def DantzigSelector(y,X,sigma):
# Extracting the sizes
m,n = X.shape
# Computing kappa
alpha = 0.05
nu = max(np.linalg.norm(X, axis=0))
kappa = nu*norm.ppf(1-alpha/(2.0*n))
# Computing LP
c = matrix(np.concatenate((np.zeros(n),np.ones(n))))
G1 = np.hstack((np.eye(n),-np.eye(n)))
G2 = np.hstack((-np.eye(n),-np.eye(n)))
G3 = np.hstack((np.dot(X.T,X),np.zeros((n,n))))
G4 = np.hstack((-np.dot(X.T,X),np.zeros((n,n))))
G = matrix(np.vstack((G1,G2,G3,G4)))
h = matrix(np.concatenate( ( np.zeros(n) , np.zeros(n) , kappa*sigma*np.ones(n) + np.dot(X.T,y) , kappa*sigma*np.ones(n) - np.dot(X.T,y) ) ))
# Solving LP
sol=solvers.lp(c,G,h)
theta = sol['x'][0:n]
return np.array(theta)
# +
sigmaDS = 1e-1
YDS = DantzigSelector(y,T,sigmaDS)
# -
plt.plot(f,X ,c='0.7',label="true signal")
plt.plot(f,YDS ,c='g',label="compressed sensing w/ Dantzig Sel.")
plt.xlabel("frequency (Hz)")
#plt.xlim([0,100])
plt.legend()
# From the *reconstructed frequential signal* $\hat{Y}$, we can retreive the full signal by inverse DCT.
xDS = spfft.idct(YDS, norm='ortho', axis=0)
plt.plot(t,x ,c='0.7',label="true signal")
plt.scatter(tS,y , c = 'red', marker = 'o',label="observations")
plt.plot(t,xDS ,c='g',label="compressed sensing w/ Dantzig")
plt.xlabel("time (s)")
plt.ylabel("Zoom")
plt.xlim([0,tMax])
plt.legend()
# <div class="exo">
# **Exercise 2-2.1:** Lasso
# <br/><br/>
#
# We can try the same thing for the lasso. Implement a Lasso function in the model of the Dantzig Selector.
#
# The reader is invited to play with the hyperparameter $\sigma$.
#
# </div>
#
#
def Lasso(y,X,sigma):
# Extracting the sizes
m,n = X.shape
return np.zeros((n,1)) #....................
# +
sigma = 0.1
YLa = Lasso(y,T,sigma)
# -
plt.plot(f,X ,c='0.7',label="true signal")
plt.plot(f,YLa ,c='g',label="compressed sensing w/ Lasso")
plt.xlabel("frequency (Hz)")
#plt.xlim([0,100])
plt.legend()
xLa = spfft.idct(YLa, norm='ortho', axis=0)
plt.plot(t,x ,c='0.7',label="true signal")
plt.scatter(tS,y , c = 'red', marker = 'o',label="observations")
plt.plot(t,xLa ,c='g',label="compressed sensing w/ Lasso" )
plt.xlabel("time (s)")
plt.ylabel("Zoom")
plt.xlim([0,tMax])
plt.legend()
# ---
# ---
# <div id="style"></div>
# ### Package Check and Styling
#
#
# <p style="text-align: right; font-size: 10px;"><a href="#top">Go to top</a></p>
#
# +
import lib.notebook_setting as nbs
packageList = ['IPython', 'numpy', 'scipy', 'matplotlib', 'cvxopt', 'pandas', 'seaborn', 'sklearn', 'tensorflow']
nbs.packageCheck(packageList)
nbs.cssStyling()
| 2-2_Examples_in_Data_Science.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (tensorflow)
# language: python
# name: tensorflow
# ---
# + [markdown] id="7bUMANd9dKsQ"
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_4_intro_transformers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nwue4RDMdKsS"
# # T81-558: Applications of Deep Neural Networks
# **Module 10: Time Series in Keras**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] id="iZEDDiENdKsS"
# # Module 10 Material
#
# * Part 10.1: Time Series Data Encoding for Deep Learning [[Video]](https://www.youtube.com/watch?v=dMUmHsktl04&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_1_timeseries.ipynb)
# * Part 10.2: Programming LSTM with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=wY0dyFgNCgY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_2_lstm.ipynb)
# * Part 10.3: Text Generation with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=6ORnRAz3gnA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_3_text_generation.ipynb)
# * **Part 10.4: Introduction to Transformers** [[Video]](https://www.youtube.com/watch?v=Z7FIdKVQ7kc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_4_intro_transformers.ipynb)
# * Part 10.5: Transformers for Timeseries [[Video]](https://www.youtube.com/watch?v=SX67Mni0Or4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_5_keras_transformers.ipynb)
# + [markdown] id="RrDcW1cUdKsT"
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# Running the following code will map your GDrive to ```/content/drive```.
# + id="E97PfPCGdKsT" outputId="882f0b5f-005b-4055-e708-0b62433f67ad"
try:
# %tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# + [markdown] id="NVW5XjandKsU"
# # Part 10.4: Introduction to Transformers
#
# Transformers are neural networks that provide state-of-the-art solutions for many of the problems previously assigned to recurrent neural networks. [[Cite:vaswani2017attention]](https://arxiv.org/abs/1706.03762) Sequences can form both the input and the output of a neural network, examples of such configurations include::
#
# * Vector to Sequence - Image captioning
# * Sequence to Vector - Sentiment analysis
# * Sequence to Sequence - Language translation
#
# Sequence-to-sequence allows an input sequence to produce an output sequence based on an input sequence. Transformers focus primarily upon this sequence-to-sequence configuration.
#
# ## High Level Overview of Transformers
#
# This course focuses primarily on the application of deep neural networks. The focus will be on presenting data to a transformer and a transformer's major components. As a result, we will not focus on implementing a transformer at the lowest level. The following section provides an overview of critical internal parts of a transformer, such as residual connections and attention. In the next chapter, we will use transformers from [Hugging Face](https://huggingface.co/) to perform natural language processing with transformers. If you are interested in implementing a transformer from scratch, Keras provides a comprehensive [example](https://www.tensorflow.org/text/tutorials/transformer).
#
# Figure 10.TRANS-1 presents a high-level view of a transformer for language translation.
#
# **Figure 10.TRANS-1: High Level View of a Translation Transformer**
# 
#
# We use a transformer that translates between English and Spanish for this example. We present the English sentence "the cat likes milk" and receive a Spanish translation of "al gato le gusta la leche."
#
# We begin by placing the English source sentence between the beginning and ending tokens. This input can be of any length, and we presented it to the neural network as a ragged Tensor. Because the Tensor is ragged, no padding is necessary. Such input is acceptable for the attention layer that will receive the source sentence. The encoder transforms this ragged input into a hidden state containing a series of key-value pairs representing the knowledge in the source sentence. The encoder understands to read English and convert to a hidden state. The decoder understands how to output Spanish from this hidden state.
#
# We initially present the decoder with the hidden state and the starting token. The decoder will predict the probabilities of all words in its vocabulary. The word with the highest probability is the first word of the sentence.
#
# The highest probability word is attached concatenated to the translated sentence, initially containing only the beginning token. This process continues, growing the translated sentence in each iteration until the decoder predicts the ending token.
#
# ## Transformer Hyperparameters
#
# Before we describe how these layers fit together, we must consider the following transformer hyperparameters, along with default settings from the Keras transformer example:
#
# * num_layers = 4
# * d_model = 128
# * dff = 512
# * num_heads = 8
# * dropout_rate = 0.1
#
# Multiple encoder and decoder layers can be present. The **num_layers** hyperparameter specifies how many encoder and decoder layers there are. The expected tensor shape for the input to the encoder layer is the same as the output produced; as a result, you can easily stack these layers.
#
# We will see embedding layers in the next chapter. However, you can think of an embedding layer as a dictionary for now. Each entry in the embedding corresponds to each word in a fixed-size vocabulary. Similar words should have similar vectors. The **d_model** hyperparameter specifies the size of the embedding vector. Though you will sometimes preload embeddings from a project such as [Word2vec](https://radimrehurek.com/gensim/models/word2vec.html) or [GloVe](https://nlp.stanford.edu/projects/glove/), the optimizer can train these embeddings with the rest of the transformer. Training your embeddings allows the **d_model** hyperparameter to set to any desired value. If you transfer the embeddings, you must set the **d_model** hyperparameter to the same value as the transferred embeddings.
#
# The **dff** hyperparameter specifies the size of the dense feedforward layers. The **num_heads** hyperparameter sets the number of attention layers heads. Finally, the dropout_rate specifies a dropout percentage to combat overfitting. We discussed dropout previously in this book.
#
# ## Inside a Transformer
#
# In this section, we will examine the internals of a transformer so that you become familiar with essential concepts such as:
#
# * Embeddings
# * Positional Encoding
# * Attention and Self-Attention
# * Residual Connection
#
# You can see a lower-level diagram of a transformer in Figure 10.TRANS-2.
#
# **Figure 10.TRANS-2: Architectural Diagram from the Paper**
# 
#
# While the original transformer paper is titled "Attention is All you Need," attention isn't the only layer type you need. The transformer also contains dense layers. However, the title "Attention and Dense Layers are All You Need" isn't as catchy.
#
# The transformer begins by tokenizing the input English sentence. Tokens may or may not be words. Generally, familiar parts of words are tokenized and become building blocks of longer words. This tokenization allows common suffixes and prefixes to be understood independently of their stem word. Each token becomes a numeric index that the transformer uses to look up the vector. There are several special tokens:
#
# * Index 0 = Pad
# * Index 1 = Unknow
# * Index 2 = Start token
# * Index 3 = End token
#
# The transformer uses index 0 when we must pad unused space at the end of a tensor. Index 1 is for unknown words. The starting and ending tokens are provided by indexes 2 and 3.
#
# The token vectors are simply the inputs to the attention layers; there is no implied order or position. The transformer adds the slopes of a sine and cosine wave to the token vectors to encode position.
#
# Attention layers have three inputs: key (k), value(v), and query (q). If query, key, value are the same, this layer is self-attention. The key and value pairs specify the information that the query operates upon. The attention layer learns what positions of data to focus upon.
#
# The transformer presents the position encoded embedding vectors to the first self-attention segment in the encoder layer. The output from the attention is normalized and ultimately becomes the hidden state after all encoder layers are processed.
#
# The hidden state is only calculated once per query. Once the input Spanish sentence becomes a hidden state, this value is presented repeatedly to the decoder until the final Spanish sentence is formed.
#
# This section presented a high-level introduction to transformers. In the next part, we will implement the encoder and apply it to time series. We will use [Hugging Face](https://huggingface.co/) transformers to perform natural language processing in the following chapter.
#
#
#
# + [markdown] id="pegE32yPJjzM"
#
# + id="3mOq6sardKsY"
| t81_558_class_10_4_intro_transformers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/allan-gon/DS-Unit-2-Kaggle-Challenge/blob/master/module1-decision-trees/LS_DS_221_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YqUUd8Uv6u-j" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 1*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Decision Trees
#
# ## Assignment
# - [ ] [Sign up for a Kaggle account](https://www.kaggle.com/), if you don’t already have one. Go to our Kaggle InClass competition website. You will be given the URL in Slack. Go to the Rules page. Accept the rules of the competition. Notice that the Rules page also has instructions for the Submission process. The Data page has feature definitions.
# - [ ] Do train/validate/test split with the Tanzania Waterpumps data.
# - [ ] Begin with baselines for classification.
# - [ ] Select features. Use a scikit-learn pipeline to encode categoricals, impute missing values, and fit a decision tree classifier.
# - [ ] Get your validation accuracy score.
# - [ ] Get and plot your feature importances.
# - [ ] Submit your predictions to our Kaggle competition. (Go to our Kaggle InClass competition webpage. Use the blue **Submit Predictions** button to upload your CSV file. Or you can use the Kaggle API to submit your predictions.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# ### Reading
#
# - A Visual Introduction to Machine Learning
# - [Part 1: A Decision Tree](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/)
# - [Part 2: Bias and Variance](http://www.r2d3.us/visual-intro-to-machine-learning-part-2/)
# - [Decision Trees: Advantages & Disadvantages](https://christophm.github.io/interpretable-ml-book/tree.html#advantages-2)
# - [How a Russian mathematician constructed a decision tree — by hand — to solve a medical problem](http://fastml.com/how-a-russian-mathematician-constructed-a-decision-tree-by-hand-to-solve-a-medical-problem/)
# - [How decision trees work](https://brohrer.github.io/how_decision_trees_work.html)
# - [Let’s Write a Decision Tree Classifier from Scratch](https://www.youtube.com/watch?v=LDRbO9a6XPU) — _Don’t worry about understanding the code, just get introduced to the concepts. This 10 minute video has excellent diagrams and explanations._
# - [Random Forests for Complete Beginners: The definitive guide to Random Forests and Decision Trees](https://victorzhou.com/blog/intro-to-random-forests/)
#
#
# ### Doing
# - [ ] Add your own stretch goal(s) !
# - [ ] Define a function to wrangle train, validate, and test sets in the same way. Clean outliers and engineer features. (For example, [what columns have zeros and shouldn't?](https://github.com/Quartz/bad-data-guide#zeros-replace-missing-values) What columns are duplicates, or nearly duplicates? Can you extract the year from date_recorded? Can you engineer new features, such as the number of years from waterpump construction to waterpump inspection?)
# - [ ] Try other [scikit-learn imputers](https://scikit-learn.org/stable/modules/impute.html).
# - [ ] Make exploratory visualizations and share on Slack.
#
#
# #### Exploratory visualizations
#
# Visualize the relationships between feature(s) and target. I recommend you do this with your training set, after splitting your data.
#
# For this problem, you may want to create a new column to represent the target as a number, 0 or 1. For example:
#
# ```python
# train['functional'] = (train['status_group']=='functional').astype(int)
# ```
#
#
#
# You can try [Seaborn "Categorical estimate" plots](https://seaborn.pydata.org/tutorial/categorical.html) for features with reasonably few unique values. (With too many unique values, the plot is unreadable.)
#
# - Categorical features. (If there are too many unique values, you can replace less frequent values with "OTHER.")
# - Numeric features. (If there are too many unique values, you can [bin with pandas cut / qcut functions](https://pandas.pydata.org/pandas-docs/stable/getting_started/basics.html?highlight=qcut#discretization-and-quantiling).)
#
# You can try [Seaborn linear model plots](https://seaborn.pydata.org/tutorial/regression.html) with numeric features. For this classification problem, you may want to use the parameter `logistic=True`, but it can be slow.
#
# You do _not_ need to use Seaborn, but it's nice because it includes confidence intervals to visualize uncertainty.
#
# #### High-cardinality categoricals
#
# This code from a previous assignment demonstrates how to replace less frequent values with 'OTHER'
#
# ```python
# # Reduce cardinality for NEIGHBORHOOD feature ...
#
# # Get a list of the top 10 neighborhoods
# top10 = train['NEIGHBORHOOD'].value_counts()[:10].index
#
# # At locations where the neighborhood is NOT in the top 10,
# # replace the neighborhood with 'OTHER'
# train.loc[~train['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# test.loc[~test['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# ```
#
# + colab_type="code" id="o9eSnDYhUGD7" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e9db7793-5893-4276-b2af-7db631409149"
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# !pip install pandas-profiling==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + colab_type="code" id="QJBD4ruICm1m" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b9f2cd79-d4bb-401b-80c4-6bd68939cb01"
import pandas as pd
from sklearn.model_selection import train_test_split
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
train.shape, test.shape
# + colab_type="code" id="2Amxyx3xphbb" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e410e418-6bd7-4bca-ee7a-5437ed48b13c"
from pandas_profiling import ProfileReport
#profile = ProfileReport(train, minimal=True).to_notebook_iframe()
#profile
# + [markdown] id="5eB4qoALhLro" colab_type="text"
# #Imports
# + id="2fG12Gf_hI7M" colab_type="code" colab={}
import numpy as np
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.tree import DecisionTreeClassifier
# + id="rZx69ELT6u-q" colab_type="code" colab={}
def wrangle(x):
df = x.copy()
#dropping
df.drop(['recorded_by','date_recorded','quantity_group'],axis=1,inplace=True)#dont drop id but i think dont include in model train
#replace latitude close to 0 values with 0
df.latitude.replace({-2.000000e-08:0},inplace=True)
# replace -s with anans here
cols_with_zeros = ['longitude', 'latitude']
for col in cols_with_zeros:
df[col] = df[col].replace(0, np.nan)
return df
# + id="LVJyififl1ad" colab_type="code" colab={}
train = wrangle(train)
test = wrangle(test)
# + [markdown] id="gz39VCpZtLe8" colab_type="text"
# # this removes the nans and puts other..., what its supposed to do is just lower each categorical columns cardinality
# + id="536CrwdtoVNF" colab_type="code" colab={}
for i in ['funder','installer','wpt_name','subvillage','lga','ward','scheme_name']:
top10 = train[i].value_counts()[:10].index
train.loc[~train[i].isin(top10), i] = 'OTHER'
test.loc[~test[i].isin(top10), i] = 'OTHER'
# + id="oQv5VKmOt_-9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 680} outputId="b74e76e8-787c-4999-e3b3-fdac393114db"
train.isnull().sum()
# + id="al_-vRGGz55M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="89402c2a-2b05-4255-f1f3-ced7dfbe9072"
train.shape
# + id="nyshYvpnz62j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9d2d8935-b1f3-4f78-d36a-3a6551aca9c7"
test.shape
# + id="HhqLBvLhz7wE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 663} outputId="61e61412-5875-42d4-93f2-68ac7efc490f"
test.isnull().sum()
# + id="2f5dUpyuz-cC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="203ecc33-3684-4a9b-9a53-6eeafad44f5a"
print(f"{test.dropna().shape}, {train.dropna().shape}")# i think its ok to drop here
#test.dropna(inplace=True)# i cant drop nans in test because i need an answer for each column
train.dropna(inplace=True)# i can here but i need to figure out what i wan tto do with the ananas in test
# + id="UxGqe4yx0OTh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 165} outputId="1f9004ed-4d9c-4416-9a9d-91ed4a3dde94"
test.head(2)
# + id="siKyfr5e1-oe" colab_type="code" colab={}
X_train,X_val,y_train,y_val = train_test_split(train.dropna().drop('status_group',axis=1),train.dropna().status_group,test_size=.2)
# + id="qI8RiNhK3Ouq" colab_type="code" colab={}
pipe_model = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
LogisticRegression(n_jobs=-1)
)
# Fit
pipe_model.fit(X_train, y_train);
# + id="lPxSCrfW3dKO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="70670872-cc16-4bad-878a-9384fc503e29"
print(f"Val Score: {pipe_model.score(X_val,y_val)}")
print(f"Train Score: {pipe_model.score(X_train,y_train)}")
print(f"Baseline: {max(train.status_group.value_counts(normalize=True))}")
# + id="n4sekNik4MIk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="7064ccf1-edf2-4f0e-f1df-ad27a23d7bdb"
pipe_model.predict(test)
# + id="OlHlOL3k4V9k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="645165bb-5cbb-42b2-9eba-2e07da32d757"
test.shape
# + id="8m_B6zSO4Z3s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 765} outputId="70d9d10c-7046-40a2-ce25-66b4c5638f1f"
test.info()
# + id="rZmF3vkf4az5" colab_type="code" colab={}
tree_pipe = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier()
)
# + id="oAyKwCjv40DA" colab_type="code" colab={}
tree_pipe.fit(X_train,y_train);
# + id="oeiOycuW45PA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="91d4be71-a402-4f27-b3c5-57c93b9ea2b3"
print(f"Val Score: {tree_pipe.score(X_val,y_val)}")
print(f"Train Score: {tree_pipe.score(X_train,y_train)}")
print(f"Baseline: {max(train.status_group.value_counts(normalize=True))}")
# + id="ZOWQDwth5BGv" colab_type="code" colab={}
y_pred = tree_pipe.predict(test)
# + id="OhqAMSzk7cei" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="6a4399cb-1b75-493b-bf4c-c56d51b1b585"
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
submission = sample_submission.copy()
submission['status_group'] = y_pred
submission.to_csv('second_try.csv', index=False)
from google.colab import files
files.download('second_try.csv')
# + id="D7o6zoOi5Lts" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 663} outputId="f234b395-659f-4a58-a4eb-80a7e261d76b"
test.isnull().sum()
# + id="mIXhGBig6f5U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="af091086-87bc-43a0-ee4c-51f374b41703"
import matplotlib.pyplot as plt
model = pipe_model.named_steps['logisticregression']
encoder = pipe_model.named_steps['onehotencoder']
encoded_columns = encoder.transform(X_val).columns
coefficients = pd.Series(model.coef_[0], encoded_columns)
plt.figure(figsize=(10,30))
coefficients.sort_values().plot.barh(color='grey');
# + id="JaBME6NO6qin" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6ad45f5b-36fa-4c07-f2b4-450d8e10a269"
y_pred.shape
# + id="iT3DkdXf840f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="93e85257-786e-4e37-8f2a-54945909a621"
test.shape[0]
# + id="Hzw7-ivk862-" colab_type="code" colab={}
from sklearn.ensemble import RandomForestClassifier
# + id="Mo8-g31u-A23" colab_type="code" colab={}
forest_pipe = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
RandomForestClassifier()
)
# + id="T9pmHwrD-JbA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="2d8513f7-1ae4-409a-a09f-d1e678b428dd"
forest_pipe.fit(X_train,y_train);
# + id="DoUsMbpP-Mfv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="19100a9c-045c-4f06-ebc9-300461f6ca9a"
print(f"Val Score: {forest_pipe.score(X_val,y_val)}")
print(f"Train Score: {forest_pipe.score(X_train,y_train)}")
print(f"Baseline: {max(train.status_group.value_counts(normalize=True))}")
# + id="1aTMlu_S-T-x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="91a19175-a8ac-4419-99b5-e31e40e2a30f"
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
submission = sample_submission.copy()
submission['status_group'] = y_pred
submission.to_csv('third_try.csv', index=False)
files.download('third_try.csv')
| module1-decision-trees/LS_DS_221_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/bro.png" width="100px"></div>
#
# # Bro to Parquet to Spark
# Apache Parquet is a columnar storage format focused on performance. Parquet data is often used within the Hadoop ecosystem and we will specifically be using it for loading data into both Pandas and Spark.
#
# <div style="float: right; margin: 30px -100px 0px 0px"><img src="images/parquet.png" width="300px"></div>
#
# ### Software
# - Bro Analysis Tools (BAT): https://github.com/Kitware/bat
# - Pandas: https://github.com/pandas-dev/pandas
# - Parquet: https://parquet.apache.org
# - Spark: https://spark.apache.org
#
# <div style="float: right; margin: 30px 0px 0px 0px"><img src="images/spark.png" width="200px"></div>
#
# ### Data
# - Sec Repo: http://www.secrepo.com (there's no Bro headers on these)
# - Kitware: https://data.kitware.com/#collection/58d564478d777f0aef5d893a (with headers)
#
# <div style="float: left; margin: 80px 20px 50px 20px"><img src="images/bleeding.jpg" width="250px"></div>
# ### Bleeding Edge Warning:
# You know you're on the bleeding edge when you link PRs that are still open/in-progess. There are **two open issues** with saving Parquet Files right now.
#
# - Timestamps in Spark: https://issues.apache.org/jira/browse/ARROW-1499
# - TimeDelta Support: https://issues.apache.org/jira/browse/ARROW-835
#
# For Spark timestamps, the BAT Parquet writer used below will output INT96 timestamps for now (we'll change over later when ARROW-1499 is complete).
#
# For the TimeDelta support we'll just have to wait until that gets pushed into the main branch and released.
# +
# Third Party Imports
import pyspark
from pyspark.sql import SparkSession
import pyarrow
# Local imports
import bat
from bat.log_to_parquet import log_to_parquet
# Good to print out versions of stuff
print('BAT: {:s}'.format(bat.__version__))
print('PySpark: {:s}'.format(pyspark.__version__))
print('PyArrow: {:s}'.format(pyarrow.__version__))
# -
# ## Bro log to Parquet File
# Here we're loading in a Bro HTTP log with ~2 million rows to demonstrate the functionality and do some simple spark processing on the data.
# - log_to_parquet is iterative so it can handle large files
# - 'row_group_size' defaults to 1 Million rows but can be set manually
# Create a Parquet file from a Bro Log with a super nice BAT method.
log_to_parquet('/Users/briford/data/bro/sec_repo/http.log', 'http.parquet')
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/compressed.jpeg" width="300px"></div>
#
# # Parquet files are compressed
# Here we see the first benefit of Parquet which stores data with compressed columnar format. There are several compression options available (including uncompressed).
#
# ## Original http.log = 1.3 GB
# ## http.parquet = 106 MB
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/spark.png" width="200px"></div>
#
# # Spark It!
# ### Spin up Spark with 4 Parallel Executors
# Here we're spinning up a local spark server with 4 parallel executors, although this might seem a bit silly since we're probably running this on a laptop, there are a couple of important observations:
#
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/spark_jobs.png" width="400px"></div>
#
# - If you have 4/8 cores use them!
# - It's the exact same code logic as if we were running on a distributed cluster.
# - We run the same code on **DataBricks** (www.databricks.com) which is awesome BTW.
#
#
# Spin up a local Spark Session (with 4 executors)
spark = SparkSession.builder.master('local[4]').appName('my_awesome').getOrCreate()
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/fast.jpg" width="350px"></div>
#
# # Parquet files are fast
# We see from the below timer output that the Parquet file only takes a few seconds to read into Spark.
# Have Spark read in the Parquet File
# %time spark_df = spark.read.parquet("http.parquet")
# <div style="float: right; margin: 0px 0px 0px -80px"><img src="images/spark_distributed.png" width="500px"></div>
#
# # Parquet files are Parallel
# We see that, in this case, the number of data partitions in our dataframe(rdd) equals the number of executors/workers. If we had 8 workers there would be 8 partitions (at least, often there are more partitions based on how big the data is and how the files were writen, etc).
#
#
# **Image Credit:** <NAME>, please see his excellent book - Mastering Apache Spark https://jaceklaskowski.gitbooks.io/mastering-apache-spark
spark_df.rdd.getNumPartitions()
# <div style="float: left; margin: 20px 20px 20px 20px"><img src="images/eyeball.jpeg" width="150px"></div>
# # Lets look at our data
# We should always inspect out data when it comes in. Look at both the data values and the data types to make sure you're getting exactly what you should be.
# Get information about the Spark DataFrame
num_rows = spark_df.count()
print("Number of Rows: {:d}".format(num_rows))
columns = spark_df.columns
print("Columns: {:s}".format(','.join(columns)))
spark_df.select(['`id.orig_h`', 'host', 'uri', 'status_code', 'user_agent']).show(5)
# <div style="float: right; margin: 20px 20px 20px 20px"><img src="images/fast.jpg" width="350px"></div>
#
# # Did we mention fast?
# The query below was executed on 4 workers. The data contains over 2 million HTTP requests/responses and the time to complete was **less than 1 second**. All this code is running on a 2016 Mac Laptop :)
# %time spark_df.groupby('method','status_code').count().sort('count', ascending=False).show()
# <div style="float: right; margin: 50px 0px 0px 20px"><img src="images/deep_dive.jpeg" width="350px"></div>
#
# # Data looks good, lets take a deeper dive
# Spark has a powerful SQL engine as well as a Machine Learning library. So now that we've got the data loaded into Parquet we're going to utilize the Spark SQL commands to do some investigation and clustering using the Spark MLLib. For this deeper dive we're going to go to another notebook :)
#
# ### Spark Clustering Notebook
# - [Bro Spark Clustering](https://github.com/Kitware/bat/blob/master/notebooks/Spark_Clustering.ipynb)
#
# <div style="float: left; margin: 0px 0px 0px 0px"><img src="images/spark_sql.jpg" width="150px"></div>
# <div style="float: left; margin: -20px 50px 0px 0px"><img src="images/mllib.png" width="150px"></div>
# <div style="float: right; margin: 50px 0px 0px -20px"><img src="https://www.kitware.com/img/small_logo_over.png" width="250px"></div>
# ## Wrap Up
# Well that's it for this notebook, we went from a Bro log to a high performance Parquet file and then did some digging with high speed, parallel SQL and groupby operations.
#
# If you liked this notebook please visit the [BAT](https://github.com/Kitware/bat) project for more notebooks and examples.
| notebooks/Bro_to_Parquet_to_Spark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import stats
import seaborn as sns
import warnings
import random
from datetime import datetime
random.seed(datetime.now())
warnings.filterwarnings('ignore')
from matplotlib import style
style.use("ggplot")
from sklearn import svm
from sklearn.model_selection import GridSearchCV
from sklearn import linear_model
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn import cross_validation # used to test classifier
from sklearn.cross_validation import KFold, cross_val_score, train_test_split
from sklearn import metrics
# %matplotlib inline
plt.rcParams['figure.figsize'] = (12,8)
# -
df = pd.read_csv("Replaced_small.csv",encoding="ISO-8859-1")
df.head()
#data formating Unnamed is a non-informative column
df = df.drop("Unnamed: 0", 1)
df.head()
# #### Drop the columns which are not required and not useful for predictions
# +
drop_cols = ['brand','categories','categories','dateAdded','dateUpdated','keys','manufacturer','name','reviewsdate','dateSeen','sourceURLs','text','title','userCity','upc','userProvince']
df = df.drop(drop_cols,axis=1)
df.head()
# -
# #### Fill the NaNs with suitable values
df['didPurchase'].fillna(True, inplace=True)
df['doRecommend'].fillna(True, inplace=True)
# #### Convert boolean values to binary values i.e. True to 1 and False to 0
df.didPurchase = (df.didPurchase)*1
df.doRecommend = (df.doRecommend)*1
df.fillna(0, inplace=True)
df.head()
# #### Convert string values to integer values by hashing the column values
# +
def get_hash(x):
return abs(hash(x)) % 10**9
df['username'] = df['username'].apply(get_hash)
df['id'] = df['id'].apply(get_hash)
df.head()
# -
df.groupby('doRecommend').count()
df.describe()
df.groupby('doRecommend').median()
df.groupby('doRecommend').mean()
# #### Scale the column values
def scaled_df(df):
scaled = pd.DataFrame()
for item in df:
if item in df.select_dtypes(include=[np.float]):
scaled[item] = ((df[item] - df[item].min()) /
(df[item].max() - df[item].min()))
else:
scaled[item] = df[item]
return scaled
df_scaled = scaled_df(df)
# +
f, ax = plt.subplots(figsize=(11, 15))
ax.set_axis_bgcolor('#FFFFFF')
plt.title("Box Plot Product Data Unscaled")
ax.set(xlim=(-.05, 1.05))
ax = sns.boxplot(data = df[:22],
orient = 'h',
palette = 'Set3')
# +
f, ax = plt.subplots(figsize=(11, 15))
ax.set_axis_bgcolor('#FFFFFF')
plt.title("Box Plot Product Data Scaled")
ax.set(xlim=(-.05, 1.05))
ax = sns.boxplot(data = df_scaled[:22],
orient = 'h',
palette = 'Set3')
# -
df.dtypes
df.head()
# #### Set predictor columns to determine the results
predictor_names=['id','didPurchase','username','rating']
predictor_names
# #### Find Rank for each of the predictor columns
def rank_predictors(dat,l,f='doRecommend'):
rank={}
max_vals=dat.max()
median_vals=dat.groupby(f).median() # We are using the median as the mean is sensitive to outliers
for p in l:
score=np.abs((median_vals[p][1]-median_vals[p][0])/max_vals[p])
rank[p]=score
return rank
cat_rank=rank_predictors(df,predictor_names)
cat_rank
# #### Sort the predictors by rank
cat_rank=sorted(cat_rank.items(), key=lambda x: x[1])
cat_rank
# #### Take the top predictors based on median difference
ranked_predictors=[]
for f in cat_rank[1:]:
ranked_predictors.append(f[0])
ranked_predictors
# #### Predicting if the product will be recommended or not using the predictor columns
X = df_scaled[predictor_names]
#setting target
y = df_scaled['doRecommend']
X_train, X_test, y_train, y_test = train_test_split(df, y,test_size=0.2)
# #### Find the accuracy score using SVM Classifier using Linear Kernel
# +
print("---------------------------------------------")
print("RBF Kernel")
svc = svm.SVC(kernel='rbf', C=1).fit(X, y)
print("KfoldCrossVal mean score using SVM is %s" %cross_val_score(svc,X,y,cv=10).mean())
#SVM metrics
sm = svc.fit(X_train, y_train)
y_pred = sm.predict(X_test)
print("Accuracy score using SVM is %s" %metrics.accuracy_score(y_test, y_pred))
print("---------------------------------------------")
print("RBF Kernel")
svc = svm.SVC(kernel='rbf', C=10).fit(X, y)
print("KfoldCrossVal mean score using SVM is %s" %cross_val_score(svc,X,y,cv=10).mean())
#SVM metrics
sm = svc.fit(X_train, y_train)
y_pred = sm.predict(X_test)
print("Accuracy score using SVM is %s" %metrics.accuracy_score(y_test, y_pred))
print("---------------------------------------------")
print("Poly Kernel")
svc = svm.SVC(kernel='poly', C=1).fit(X, y)
print("KfoldCrossVal mean score using SVM is %s" %cross_val_score(svc,X,y,cv=10).mean())
#SVM metrics
sm = svc.fit(X_train, y_train)
y_pred = sm.predict(X_test)
print("Accuracy score using SVM is %s" %metrics.accuracy_score(y_test, y_pred))
print("---------------------------------------------")
print("Sigmoid Kernel")
svc = svm.SVC(kernel='sigmoid', C=1, gamma=0.001).fit(X, y)
print("KfoldCrossVal mean score using SVM is %s" %cross_val_score(svc,X,y,cv=10).mean())
#SVM metrics
sm = svc.fit(X_train, y_train)
y_pred = sm.predict(X_test)
print("Accuracy score using SVM is %s" %metrics.accuracy_score(y_test, y_pred))
# -
# ### Changing hyper-parameter values does not change the accuracy score of predictions.
# +
#setting svm classifier
svc = svm.SVC(kernel='rbf', C=1).fit(X, y)
print("KfoldCrossVal mean score using SVM is %s" %cross_val_score(svc,X,y,cv=10).mean())
#SVM metrics
sm = svc.fit(X_train, y_train)
y_pred = sm.predict(X_test)
print("Accuracy score using SVM is %s" %metrics.accuracy_score(y_test, y_pred))
# -
# ### Accuracy score using SVM is 84.6%
# ### K-fold average accuracy score using SVM is ~75.35%
| Part A - SupportVectorMachine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd ../src
# # 1. `envGym`
#
# `envGym` is a wrapper around the OpenAI Gym environment. It has several methds that can be useful for using the environment. In this notebook, we shall explore this environment.
from lib.envs import envGym
from time import sleep
import numpy as np
import torch
# ## 1.1. The `Env()` context manager
#
# The `Env` class exposes a context manager that will allow this environment to generate an OpenAI environment within. The first time this context manager is entered, this creates a current state `self.state`. This can be reset with the `self.reset()` method.
name = 'Breakout-ramNoFrameskip-v4'
with envGym.Env(name, showEnv=False) as env:
print(f'Initial environment state:\n {env.state}')
env.reset()
print(f'Initial environment state after reset:\n {env.state}')
# ## 1.2. The `self.env` property
#
# The `self.env` property points to the OpenAI Gym environment. Use this for any of the OpenAI Gym methods. However, it is best not to use the internal environment directly. It would be much more preferabe to update this environment to create a new method for this specific class.
# +
name = 'Breakout-ramNoFrameskip-v4'
with envGym.Env(name, showEnv=False) as env:
print(f'Check environments action space: {env.env.action_space}')
env.env.render()
sleep(2)
env.env.close()
# -
# ## 1.3. Stepping and playing
#
# The `Env` class provides two main methods to interact with the environment - a `self.step(policy)` and an `selef.episode(policy, maxSteps)`. For making this environment compatible with the `envUnity` environment, which can simulate more than a single actor at a single time, this policy is a function that should return a number of actions, one for each actor. For this reason, the result of a policy should always be a list of actions. For the `envGym` environment, this will mean a list with a single action.
#
# ### 1.3.1. Let us take a couple of steps
#
# Note that the _Breakout_ takes a `Discrete(4)` pytorch tensor. Note allso that this returrns a result per actor (only one in this case). We shall specify a randoom action ...
# +
name = 'Breakout-ramNoFrameskip-v4'
policy = lambda m: [torch.randint(0, 4, (1,))]
with envGym.Env(name, showEnv=False) as env:
print('Taking the first step')
result = env.step(policy)
print(result)
print('\n\nTaking the second step')
result = env.step(policy)[0]
state, action, reward, nextState, done = result
print(f'''
state : \n{state}
nextState : \n{nextState}
action : {action}
done : {done}
''')
# -
# ### 1.3.1. Let us play an entire episode
#
# Note that the _Breakout_ takes a `Discrete(4)` pytorch tensor. Note allso that this returrns a result per actor (only one in this case). We shall specify a randoom action ...
# +
name = 'Breakout-ramNoFrameskip-v4'
policy = lambda m: [torch.randint(0, 4, (1,))]
with envGym.Env(name, showEnv=False) as env:
result = env.episode(policy, 10)[0]
for r in result:
state, action, reward, nextState, done = r
print(f'reward = {reward}, action = {action}, done = {done}')
print(f'final state:\n{state}')
# -
| notebooks/Environments - envGym.Env.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Augmentation Models Training
# +
"""
Supervised learning models are trained.
@author: <NAME> - <EMAIL>
"""
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import Normalize
import pylab as pl
import seaborn as sns
plt.style.use(['ggplot','seaborn-paper'])
from time import process_time
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_predict
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import roc_auc_score
from sklearn.metrics import f1_score
from sklearn.metrics import recall_score
from sklearn.metrics import make_scorer
from sklearn.metrics import precision_recall_fscore_support
from sklearn import metrics
# -
"""
def log_reg(param_grid, X_train, y_train, X_test, y_test):
clf_log_reg = GridSearchCV(LogisticRegression(solver= 'liblinear'), param_grid=param_grid_log_reg,
verbose=0,return_train_score=True, n_jobs=-1, cv=5, iid = False)
clf_log_reg.fit(X_train, y_train)
y_pred_log_reg = clf_log_reg.predict(X_test)
score_micro = precision_recall_fscore_support(y_test, y_pred_log_reg, average='micro')
score_macro = precision_recall_fscore_support(y_test, y_pred_log_reg, average='macro')
score_weighted = precision_recall_fscore_support(y_test, y_pred_log_reg, average='weighted')
return score_micro[0], score_micro[1], score_micro[2], score_macro[0], score_macro[1], score_macro[2], score_weighted[0], score_weighted[1], score_weighted[2]
"""
def log_reg(X_train, y_train, X_test, y_test):
clf_log_reg = LogisticRegression(solver= 'liblinear', C=1)
clf_log_reg.fit(X_train, y_train)
y_pred_log_reg = clf_log_reg.predict(X_test)
score_micro = precision_recall_fscore_support(y_test, y_pred_log_reg, average='micro')
score_macro = precision_recall_fscore_support(y_test, y_pred_log_reg, average='macro')
score_weighted = precision_recall_fscore_support(y_test, y_pred_log_reg, average='weighted')
return score_micro[0], score_micro[1], score_micro[2], score_macro[0], score_macro[1], score_macro[2], score_weighted[0], score_weighted[1], score_weighted[2]
def SVM_rbf_clf(param_grid, X_train, y_train, X_test, y_test):
clf_svm_rbf = GridSearchCV(SVC(kernel='rbf'), param_grid=param_grid, verbose=0, iid = False,
return_train_score=True, cv=5)
clf_svm_rbf.fit(X_train, y_train)
y_pred_svm_rbf = clf_svm_rbf.predict(X_test)
score_micro = precision_recall_fscore_support(y_test, y_pred_svm_rbf, average='micro')
score_macro = precision_recall_fscore_support(y_test, y_pred_svm_rbf, average='macro')
score_weighted = precision_recall_fscore_support(y_test, y_pred_svm_rbf, average='weighted')
return score_micro[0], score_micro[1], score_micro[2], score_macro[0], score_macro[1], score_macro[2], score_weighted[0], score_weighted[1], score_weighted[2]
def random_forests_clf(param_grid, X_train, y_train, X_test, y_test):
clf_rf = GridSearchCV(RandomForestClassifier(random_state=42, n_jobs=-1), param_grid=param_grid_rf, verbose=0, iid = False,
return_train_score=True, n_jobs=-1, cv=5)
clf_rf.fit(X_train, y_train)
y_pred_rf = clf_rf.predict(X_test)
score_micro = precision_recall_fscore_support(y_test, y_pred_rf, average='micro')
score_macro = precision_recall_fscore_support(y_test, y_pred_rf, average='macro')
score_weighted = precision_recall_fscore_support(y_test, y_pred_rf, average='weighted')
return score_micro[0], score_micro[1], score_micro[2], score_macro[0], score_macro[1], score_macro[2], score_weighted[0], score_weighted[1], score_weighted[2]
def mlp_clf(param_grid, X_train, y_train, X_test, y_test):
clf_mlp = GridSearchCV(MLPClassifier(random_state=42), param_grid=param_grid_mlp, verbose=0, iid = False,
return_train_score=True, n_jobs=-1, cv=5)
clf_mlp.fit(X_train, y_train)
y_pred_mlp = clf_mlp.predict(X_test)
score_micro = precision_recall_fscore_support(y_test, y_pred_mlp, average='micro')
score_macro = precision_recall_fscore_support(y_test, y_pred_mlp, average='macro')
score_weighted = precision_recall_fscore_support(y_test, y_pred_mlp, average='weighted')
return score_micro[0], score_micro[1], score_micro[2], score_macro[0], score_macro[1], score_macro[2], score_weighted[0], score_weighted[1], score_weighted[2]
# +
X_train_low_set_1 = np.load('./data_augmentation/X_train_low_set_1.npy')
X_test_low_set_1 = np.load('./data_augmentation/X_test_low_set_1.npy')
y_train_low_set_1 = np.load('./data_augmentation/y_train_low_set_1.npy')
y_test_low_set_1 = np.load('./data_augmentation/y_test_low_set_1.npy')
X_train_med_set_1 = np.load('./data_augmentation/X_train_med_set_1.npy')
X_test_med_set_1 = np.load('./data_augmentation/X_test_med_set_1.npy')
y_train_med_set_1 = np.load('./data_augmentation/y_train_med_set_1.npy')
y_test_med_set_1 = np.load('./data_augmentation/y_test_med_set_1.npy')
X_train_high_set_1 = np.load('./data_augmentation/X_train_high_set_1.npy')
X_test_high_set_1 = np.load('./data_augmentation/X_test_high_set_1.npy')
y_train_high_set_1 = np.load('./data_augmentation/y_train_high_set_1.npy')
y_test_high_set_1 = np.load('./data_augmentation/y_test_high_set_1.npy')
# +
X_train_low_set_2 = np.load('./data_augmentation/X_train_low_set_2.npy')
X_test_low_set_2 = np.load('./data_augmentation/X_test_low_set_2.npy')
y_train_low_set_2 = np.load('./data_augmentation/y_train_low_set_2.npy')
y_test_low_set_2 = np.load('./data_augmentation/y_test_low_set_2.npy')
X_train_med_set_2 = np.load('./data_augmentation/X_train_med_set_2.npy')
X_test_med_set_2 = np.load('./data_augmentation/X_test_med_set_2.npy')
y_train_med_set_2 = np.load('./data_augmentation/y_train_med_set_2.npy')
y_test_med_set_2 = np.load('./data_augmentation/y_test_med_set_2.npy')
X_train_high_set_2 = np.load('./data_augmentation/X_train_high_set_2.npy')
X_test_high_set_2 = np.load('./data_augmentation/X_test_high_set_2.npy')
y_train_high_set_2 = np.load('./data_augmentation/y_train_high_set_2.npy')
y_test_high_set_2 = np.load('./data_augmentation/y_test_high_set_2.npy')
# +
X_train_low_set_3 = np.load('./data_augmentation/X_train_low_set_3.npy')
X_test_low_set_3 = np.load('./data_augmentation/X_test_low_set_3.npy')
y_train_low_set_3 = np.load('./data_augmentation/y_train_low_set_3.npy')
y_test_low_set_3 = np.load('./data_augmentation/y_test_low_set_3.npy')
X_train_med_set_3 = np.load('./data_augmentation/X_train_med_set_3.npy')
X_test_med_set_3 = np.load('./data_augmentation/X_test_med_set_3.npy')
y_train_med_set_3 = np.load('./data_augmentation/y_train_med_set_3.npy')
y_test_med_set_3 = np.load('./data_augmentation/y_test_med_set_3.npy')
X_train_high_set_3 = np.load('./data_augmentation/X_train_high_set_3.npy')
X_test_high_set_3 = np.load('./data_augmentation/X_test_high_set_3.npy')
y_train_high_set_3 = np.load('./data_augmentation/y_train_high_set_3.npy')
y_test_high_set_3 = np.load('./data_augmentation/y_test_high_set_3.npy')
# -
# +
X_train_low_cl = np.load('./data_augmentation/X_train_low.npy')
X_test_low_cl = np.load('./data_augmentation/X_test_low.npy')
y_train_low_cl = np.load('./data_augmentation/y_train_low.npy')
y_test_low_cl = np.load('./data_augmentation/y_test_low.npy')
X_train_med_cl = np.load('./data_augmentation/X_train_med.npy')
X_test_med_cl = np.load('./data_augmentation/X_test_med.npy')
y_train_med_cl = np.load('./data_augmentation/y_train_med.npy')
y_test_med_cl = np.load('./data_augmentation/y_test_med.npy')
X_train_high_cl = np.load('./data_augmentation/X_train_high.npy')
X_test_high_cl = np.load('./data_augmentation/X_test_high.npy')
y_train_high_cl = np.load('./data_augmentation/y_train_high.npy')
y_test_high_cl = np.load('./data_augmentation/y_test_high.npy')
# +
X_train_low = np.load('./results/X_train_low.npy')
X_test_low = np.load('./results/X_test_low.npy')
y_train_low = np.load('./results/y_train_low.npy')
y_test_low = np.load('./results/y_test_low.npy')
X_train_med = np.load('./results/X_train_med.npy')
X_test_med = np.load('./results/X_test_med.npy')
y_train_med = np.load('./results/y_train_med.npy')
y_test_med = np.load('./results/y_test_med.npy')
X_train_high = np.load('./results/X_train_high.npy')
X_test_high = np.load('./results/X_test_high.npy')
y_train_high = np.load('./results/y_train_high.npy')
y_test_high = np.load('./results/y_test_high.npy')
# +
#X_train_low_aug = np.concatenate((X_train_low_cl,X_train_low_set_1[:73],X_train_low_set_2[:73],X_train_low_set_3[:73]))
#y_train_low_aug = np.concatenate((y_train_low_cl,y_train_low_set_1[:73],y_train_low_set_2[:73],y_train_low_set_3[:73]))
# +
#X_train_med_aug = np.concatenate((X_train_med_cl,X_train_med_set_1[:73],X_train_med_set_2[:73],X_train_med_set_3[:73]))
#y_train_med_aug = np.concatenate((y_train_med_cl,y_train_med_set_1[:73],y_train_med_set_2[:73],y_train_med_set_3[:73]))
# +
#X_train_high_aug = np.concatenate((X_train_high_cl,X_train_high_set_1[:73],X_train_high_set_2[:73],X_train_high_set_3[:73]))
#y_train_high_aug = np.concatenate((y_train_high_cl,y_train_high_set_1[:73],y_train_high_set_2[:73],y_train_high_set_3[:73]))
# +
X_train_low_aug = np.concatenate((X_train_low_cl,X_train_low_set_1,X_train_low_set_2,X_train_low_set_3))
y_train_low_aug = np.concatenate((y_train_low_cl,y_train_low_set_1,y_train_low_set_2,y_train_low_set_3))
# +
X_train_med_aug = np.concatenate((X_train_med_cl,X_train_med_set_1,X_train_med_set_2,X_train_med_set_3))
y_train_med_aug = np.concatenate((y_train_med_cl,y_train_med_set_1,y_train_med_set_2,y_train_med_set_3))
# +
X_train_high_aug = np.concatenate((X_train_high_cl,X_train_high_set_1,X_train_high_set_2,X_train_high_set_3))
y_train_high_aug = np.concatenate((y_train_high_cl,y_train_high_set_1,y_train_high_set_2,y_train_high_set_3))
# -
len(y_train_high_set_1)
np.unique(y_train_high_aug)
y_train_low_set_1
unique, counts = np.unique(y_train_high_set_3, return_counts=True)
dict(zip(unique, counts))
y_train_low_set_1[:73]
step = 1
grid_log_reg = [i for i in range(1, 11, 1)]
param_grid_log_reg = {'C': [np.round((0.1*i), decimals =2) for i in grid_log_reg]}
print('Parameters:', param_grid_log_reg)
# +
Precision_micro_low_LR = np.array([])
Precision_macro_low_LR = np.array([])
Precision_weighted_low_LR = np.array([])
Recall_micro_low_LR = np.array([])
Recall_macro_low_LR = np.array([])
Recall_weighted_low_LR = np.array([])
F1_micro_low_LR = np.array([])
F1_macro_low_LR = np.array([])
F1_weighted_low_LR = np.array([])
Sample_Num_low_LR = np.array([])
for i in range(587, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = log_reg(X_train_low_aug[:i], y_train_low_aug[:i].astype(int), X_test_low, y_test_low)
Precision_micro_low_LR = np.append(Precision_micro_low_LR, pr_micro)
Precision_macro_low_LR = np.append(Precision_macro_low_LR, pr_macro)
Precision_weighted_low_LR = np.append(Precision_weighted_low_LR, pr_weighted)
Recall_micro_low_LR = np.append(Recall_micro_low_LR, re_micro)
Recall_macro_low_LR = np.append(Recall_macro_low_LR, re_macro)
Recall_weighted_low_LR = np.append(Recall_weighted_low_LR, re_weighted)
F1_micro_low_LR = np.append(F1_micro_low_LR, f1_micro)
F1_macro_low_LR = np.append(F1_macro_low_LR, f1_macro)
F1_weighted_low_LR = np.append(F1_weighted_low_LR, f1_weighted)
Sample_Num_low_LR = np.append(Sample_Num_low_LR, len(y_train_low_aug[:i]))
# +
df_plot_low_LR = pd.DataFrame()
df_plot_low_LR['Sample Number'] = Sample_Num_low_LR
df_plot_low_LR['Precision Micro'] = Precision_micro_low_LR
df_plot_low_LR['Precision Macro'] = Precision_macro_low_LR
df_plot_low_LR['Precision Weighted'] = Precision_weighted_low_LR
df_plot_low_LR['Recall Micro'] = Recall_micro_low_LR
df_plot_low_LR['Recall Macro'] = Recall_macro_low_LR
df_plot_low_LR['Recall Weighted'] = Recall_weighted_low_LR
df_plot_low_LR['F1 Micro'] = F1_micro_low_LR
df_plot_low_LR['F1 Macro'] = F1_macro_low_LR
df_plot_low_LR['F1 Weighted'] = F1_weighted_low_LR
df_plot_low_LR['Resolution'] = 'Low'
# +
Precision_micro_med_LR = np.array([])
Precision_macro_med_LR = np.array([])
Precision_weighted_med_LR = np.array([])
Recall_micro_med_LR = np.array([])
Recall_macro_med_LR = np.array([])
Recall_weighted_med_LR = np.array([])
F1_micro_med_LR = np.array([])
F1_macro_med_LR = np.array([])
F1_weighted_med_LR = np.array([])
Sample_Num_med_LR = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = log_reg(X_train_med_aug[:i], y_train_med_aug[:i].astype(int), X_test_med, y_test_med)
Precision_micro_med_LR = np.append(Precision_micro_med_LR, pr_micro)
Precision_macro_med_LR = np.append(Precision_macro_med_LR, pr_macro)
Precision_weighted_med_LR = np.append(Precision_weighted_med_LR, pr_weighted)
Recall_micro_med_LR = np.append(Recall_micro_med_LR, re_micro)
Recall_macro_med_LR = np.append(Recall_macro_med_LR, re_macro)
Recall_weighted_med_LR = np.append(Recall_weighted_med_LR, re_weighted)
F1_micro_med_LR = np.append(F1_micro_med_LR, f1_micro)
F1_macro_med_LR = np.append(F1_macro_med_LR, f1_macro)
F1_weighted_med_LR = np.append(F1_weighted_med_LR, f1_weighted)
Sample_Num_med_LR = np.append(Sample_Num_med_LR, len(y_train_med_aug[:i]))
# +
df_plot_med_LR = pd.DataFrame()
df_plot_med_LR['Sample Number'] = Sample_Num_med_LR
df_plot_med_LR['Precision Micro'] = Precision_micro_med_LR
df_plot_med_LR['Precision Macro'] = Precision_macro_med_LR
df_plot_med_LR['Precision Weighted'] = Precision_weighted_med_LR
df_plot_med_LR['Recall Micro'] = Recall_micro_med_LR
df_plot_med_LR['Recall Macro'] = Recall_macro_med_LR
df_plot_med_LR['Recall Weighted'] = Recall_weighted_med_LR
df_plot_med_LR['F1 Micro'] = F1_micro_med_LR
df_plot_med_LR['F1 Macro'] = F1_macro_med_LR
df_plot_med_LR['F1 Weighted'] = F1_weighted_med_LR
df_plot_med_LR['Resolution'] = 'Med'
# +
Precision_micro_high_LR = np.array([])
Precision_macro_high_LR = np.array([])
Precision_weighted_high_LR = np.array([])
Recall_micro_high_LR = np.array([])
Recall_macro_high_LR = np.array([])
Recall_weighted_high_LR = np.array([])
F1_micro_high_LR = np.array([])
F1_macro_high_LR = np.array([])
F1_weighted_high_LR = np.array([])
Sample_Num_high_LR = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = log_reg(X_train_high_aug[:i], y_train_high_aug[:i].astype(int), X_test_high, y_test_high)
Precision_micro_high_LR = np.append(Precision_micro_high_LR, pr_micro)
Precision_macro_high_LR = np.append(Precision_macro_high_LR, pr_macro)
Precision_weighted_high_LR = np.append(Precision_weighted_high_LR, pr_weighted)
Recall_micro_high_LR = np.append(Recall_micro_high_LR, re_micro)
Recall_macro_high_LR = np.append(Recall_macro_high_LR, re_macro)
Recall_weighted_high_LR = np.append(Recall_weighted_high_LR, re_weighted)
F1_micro_high_LR = np.append(F1_micro_high_LR, f1_micro)
F1_macro_high_LR = np.append(F1_macro_high_LR, f1_macro)
F1_weighted_high_LR = np.append(F1_weighted_high_LR, f1_weighted)
Sample_Num_high_LR = np.append(Sample_Num_high_LR, len(y_train_high_aug[:i]))
# +
df_plot_high_LR = pd.DataFrame()
df_plot_high_LR['Sample Number'] = Sample_Num_high_LR
df_plot_high_LR['Precision Micro'] = Precision_micro_high_LR
df_plot_high_LR['Precision Macro'] = Precision_macro_high_LR
df_plot_high_LR['Precision Weighted'] = Precision_weighted_high_LR
df_plot_high_LR['Recall Micro'] = Recall_micro_high_LR
df_plot_high_LR['Recall Macro'] = Recall_macro_high_LR
df_plot_high_LR['Recall Weighted'] = Recall_weighted_high_LR
df_plot_high_LR['F1 Micro'] = F1_micro_high_LR
df_plot_high_LR['F1 Macro'] = F1_macro_high_LR
df_plot_high_LR['F1 Weighted'] = F1_weighted_high_LR
df_plot_high_LR['Resolution'] = 'High'
# -
df_plot_LR = df_plot_low_LR.append(df_plot_med_LR, ignore_index=True)
df_plot_LR = df_plot_LR.append(df_plot_high_LR, ignore_index=True)
df_plot_LR.head()
size = (8, 7)
a4_dims = size
palette_3 = sns.color_palette("mako_r",3)
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
#plt.axvline(229, 0, 1, linestyle='--', color='black')
plt.axvline(587, 0, 1, linestyle='--', color='black')
plt.axvline(587, 0, 1, linestyle='--', color='black')
plt.axvline(587, 0, 1, linestyle='--', color='black')
plt.show()
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.axvline(587, 0, 1, linestyle='--', color='black')
plt.axvline(587+587, 0, 1, linestyle='--', color='black')
plt.axvline(587+587*2, 0, 1, linestyle='--', color='black')
plt.axvline(587+587*3, 0, 1, linestyle='--', color='black')
plt.show()
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.axvline(587, 0, 1, linestyle='--', color='black')
plt.axvline(587+73, 0, 1, linestyle='--', color='black')
plt.axvline(587+73, 0, 1, linestyle='--', color='black')
plt.axvline(587+73*2, 0, 1, linestyle='--', color='black')
plt.axvline(587+73*3, 0, 1, linestyle='--', color='black')
plt.show()
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
# +
#fig, ax = plt.subplots(figsize=a4_dims)
#ax = sns.lineplot(x="Thresholds", y="Score", hue="Resolution", style="Score Type",
# palette=palette_3, legend="full", data=df_plot)
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
#plt.show()
# -
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="Precision Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_LR)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
path_LR = './data_augmentation/results/df_LR.csv'
df_plot_LR.to_csv(path_LR, index=True)
grid_C = [i for i in range(-5, 7, 1)]
grid_gamma = [i for i in range(0, 11, 1)]
param_grid_svm_rbf = {'C': [2**i for i in grid_C],
'gamma': np.round([0.1*i for i in grid_gamma], decimals=2)}
print('Parameters:', param_grid_svm_rbf)
# +
Precision_micro_low_SVC = np.array([])
Precision_macro_low_SVC = np.array([])
Precision_weighted_low_SVC = np.array([])
Recall_micro_low_SVC = np.array([])
Recall_macro_low_SVC = np.array([])
Recall_weighted_low_SVC = np.array([])
F1_micro_low_SVC = np.array([])
F1_macro_low_SVC = np.array([])
F1_weighted_low_SVC = np.array([])
Sample_Num_low_SVC = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = SVM_rbf_clf(param_grid_svm_rbf, X_train_low_aug[:i], y_train_low_aug[:i].astype(int), X_test_low, y_test_low)
Precision_micro_low_SVC = np.append(Precision_micro_low_SVC, pr_micro)
Precision_macro_low_SVC = np.append(Precision_macro_low_SVC, pr_macro)
Precision_weighted_low_SVC = np.append(Precision_weighted_low_SVC, pr_weighted)
Recall_micro_low_SVC = np.append(Recall_micro_low_SVC, re_micro)
Recall_macro_low_SVC = np.append(Recall_macro_low_SVC, re_macro)
Recall_weighted_low_SVC = np.append(Recall_weighted_low_SVC, re_weighted)
F1_micro_low_SVC = np.append(F1_micro_low_SVC, f1_micro)
F1_macro_low_SVC = np.append(F1_macro_low_SVC, f1_macro)
F1_weighted_low_SVC = np.append(F1_weighted_low_SVC, f1_weighted)
Sample_Num_low_SVC = np.append(Sample_Num_low_SVC, len(y_train_low_aug[:i]))
print((i-590)/step)
# +
df_plot_low_SVC = pd.DataFrame()
df_plot_low_SVC['Sample Number'] = Sample_Num_low_SVC
df_plot_low_SVC['Precision Micro'] = Precision_micro_low_SVC
df_plot_low_SVC['Precision Macro'] = Precision_macro_low_SVC
df_plot_low_SVC['Precision Weighted'] = Precision_weighted_low_SVC
df_plot_low_SVC['Recall Micro'] = Recall_micro_low_SVC
df_plot_low_SVC['Recall Macro'] = Recall_macro_low_SVC
df_plot_low_SVC['Recall Weighted'] = Recall_weighted_low_SVC
df_plot_low_SVC['F1 Micro'] = F1_micro_low_SVC
df_plot_low_SVC['F1 Macro'] = F1_macro_low_SVC
df_plot_low_SVC['F1 Weighted'] = F1_weighted_low_SVC
df_plot_low_SVC['Resolution'] = 'Low'
# +
Precision_micro_med_SVC = np.array([])
Precision_macro_med_SVC = np.array([])
Precision_weighted_med_SVC = np.array([])
Recall_micro_med_SVC = np.array([])
Recall_macro_med_SVC = np.array([])
Recall_weighted_med_SVC = np.array([])
F1_micro_med_SVC = np.array([])
F1_macro_med_SVC = np.array([])
F1_weighted_med_SVC = np.array([])
Sample_Num_med_SVC = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = SVM_rbf_clf(param_grid_svm_rbf, X_train_med_aug[:i], y_train_med_aug[:i].astype(int), X_test_med, y_test_med)
Precision_micro_med_SVC = np.append(Precision_micro_med_SVC, pr_micro)
Precision_macro_med_SVC = np.append(Precision_macro_med_SVC, pr_macro)
Precision_weighted_med_SVC = np.append(Precision_weighted_med_SVC, pr_weighted)
Recall_micro_med_SVC = np.append(Recall_micro_med_SVC, re_micro)
Recall_macro_med_SVC = np.append(Recall_macro_med_SVC, re_macro)
Recall_weighted_med_SVC = np.append(Recall_weighted_med_SVC, re_weighted)
F1_micro_med_SVC = np.append(F1_micro_med_SVC, f1_micro)
F1_macro_med_SVC = np.append(F1_macro_med_SVC, f1_macro)
F1_weighted_med_SVC = np.append(F1_weighted_med_SVC, f1_weighted)
Sample_Num_med_SVC = np.append(Sample_Num_med_SVC, len(y_train_med_aug[:i]))
print((i-590)/step)
# +
df_plot_med_SVC = pd.DataFrame()
df_plot_med_SVC['Sample Number'] = Sample_Num_med_SVC
df_plot_med_SVC['Precision Micro'] = Precision_micro_med_SVC
df_plot_med_SVC['Precision Macro'] = Precision_macro_med_SVC
df_plot_med_SVC['Precision Weighted'] = Precision_weighted_med_SVC
df_plot_med_SVC['Recall Micro'] = Recall_micro_med_SVC
df_plot_med_SVC['Recall Macro'] = Recall_macro_med_SVC
df_plot_med_SVC['Recall Weighted'] = Recall_weighted_med_SVC
df_plot_med_SVC['F1 Micro'] = F1_micro_med_SVC
df_plot_med_SVC['F1 Macro'] = F1_macro_med_SVC
df_plot_med_SVC['F1 Weighted'] = F1_weighted_med_SVC
df_plot_med_SVC['Resolution'] = 'Med'
# +
Precision_micro_high_SVC = np.array([])
Precision_macro_high_SVC = np.array([])
Precision_weighted_high_SVC = np.array([])
Recall_micro_high_SVC = np.array([])
Recall_macro_high_SVC = np.array([])
Recall_weighted_high_SVC = np.array([])
F1_micro_high_SVC = np.array([])
F1_macro_high_SVC = np.array([])
F1_weighted_high_SVC = np.array([])
Sample_Num_high_SVC = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = SVM_rbf_clf(param_grid_svm_rbf, X_train_high_aug[:i], y_train_high_aug[:i].astype(int), X_test_high, y_test_high)
Precision_micro_high_SVC = np.append(Precision_micro_high_SVC, pr_micro)
Precision_macro_high_SVC = np.append(Precision_macro_high_SVC, pr_macro)
Precision_weighted_high_SVC = np.append(Precision_weighted_high_SVC, pr_weighted)
Recall_micro_high_SVC = np.append(Recall_micro_high_SVC, re_micro)
Recall_macro_high_SVC = np.append(Recall_macro_high_SVC, re_macro)
Recall_weighted_high_SVC = np.append(Recall_weighted_high_SVC, re_weighted)
F1_micro_high_SVC = np.append(F1_micro_high_SVC, f1_micro)
F1_macro_high_SVC = np.append(F1_macro_high_SVC, f1_macro)
F1_weighted_high_SVC = np.append(F1_weighted_high_SVC, f1_weighted)
Sample_Num_high_SVC = np.append(Sample_Num_high_SVC, len(y_train_high_aug[:i]))
print((i-590)/step)
# +
df_plot_high_SVC = pd.DataFrame()
df_plot_high_SVC['Sample Number'] = Sample_Num_high_SVC
df_plot_high_SVC['Precision Micro'] = Precision_micro_high_SVC
df_plot_high_SVC['Precision Macro'] = Precision_macro_high_SVC
df_plot_high_SVC['Precision Weighted'] = Precision_weighted_high_SVC
df_plot_high_SVC['Recall Micro'] = Recall_micro_high_SVC
df_plot_high_SVC['Recall Macro'] = Recall_macro_high_SVC
df_plot_high_SVC['Recall Weighted'] = Recall_weighted_high_SVC
df_plot_high_SVC['F1 Micro'] = F1_micro_high_SVC
df_plot_high_SVC['F1 Macro'] = F1_macro_high_SVC
df_plot_high_SVC['F1 Weighted'] = F1_weighted_high_SVC
df_plot_high_SVC['Resolution'] = 'High'
# -
df_plot_SVC = df_plot_low_SVC.append(df_plot_med_SVC, ignore_index=True)
df_plot_SVC = df_plot_SVC.append(df_plot_high_SVC, ignore_index=True)
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_SVC)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="Precision Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_SVC)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
path_SVC = './data_augmentation/results/df_SVC.csv'
df_plot_SVC.to_csv(path_SVC, index=True)
# +
ii = 0
for i in range(590, 2361, 15):
ii = ii +1
print(ii,' ', i)
# -
max_features_params_rf = [np.round(10**-1 * i, decimals=2) for i in range(1, 11, 1)]
param_grid_rf = {'n_estimators': [2**i for i in range(2, 12, 1)], 'max_features': max_features_params_rf}
print('Parameters:', param_grid_rf)
# +
Precision_micro_low_RF = np.array([])
Precision_macro_low_RF = np.array([])
Precision_weighted_low_RF = np.array([])
Recall_micro_low_RF = np.array([])
Recall_macro_low_RF = np.array([])
Recall_weighted_low_RF = np.array([])
F1_micro_low_RF = np.array([])
F1_macro_low_RF = np.array([])
F1_weighted_low_RF = np.array([])
Sample_Num_low_RF = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = random_forests_clf(param_grid_rf, X_train_low_aug[:i], y_train_low_aug[:i].astype(int), X_test_low, y_test_low)
Precision_micro_low_RF = np.append(Precision_micro_low_RF, pr_micro)
Precision_macro_low_RF = np.append(Precision_macro_low_RF, pr_macro)
Precision_weighted_low_RF = np.append(Precision_weighted_low_RF, pr_weighted)
Recall_micro_low_RF = np.append(Recall_micro_low_RF, re_micro)
Recall_macro_low_RF = np.append(Recall_macro_low_RF, re_macro)
Recall_weighted_low_RF = np.append(Recall_weighted_low_RF, re_weighted)
F1_micro_low_RF = np.append(F1_micro_low_RF, f1_micro)
F1_macro_low_RF = np.append(F1_macro_low_RF, f1_macro)
F1_weighted_low_RF = np.append(F1_weighted_low_RF, f1_weighted)
Sample_Num_low_RF = np.append(Sample_Num_low_RF, len(y_train_low_aug[:i]))
print((i-590)/step)
# +
df_plot_low_RF = pd.DataFrame()
df_plot_low_RF['Sample Number'] = Sample_Num_low_RF
df_plot_low_RF['Precision Micro'] = Precision_micro_low_RF
df_plot_low_RF['Precision Macro'] = Precision_macro_low_RF
df_plot_low_RF['Precision Weighted'] = Precision_weighted_low_RF
df_plot_low_RF['Recall Micro'] = Recall_micro_low_RF
df_plot_low_RF['Recall Macro'] = Recall_macro_low_RF
df_plot_low_RF['Recall Weighted'] = Recall_weighted_low_RF
df_plot_low_RF['F1 Micro'] = F1_micro_low_RF
df_plot_low_RF['F1 Macro'] = F1_macro_low_RF
df_plot_low_RF['F1 Weighted'] = F1_weighted_low_RF
df_plot_low_RF['Resolution'] = 'Low'
# +
Precision_micro_med_RF = np.array([])
Precision_macro_med_RF = np.array([])
Precision_weighted_med_RF = np.array([])
Recall_micro_med_RF = np.array([])
Recall_macro_med_RF = np.array([])
Recall_weighted_med_RF = np.array([])
F1_micro_med_RF = np.array([])
F1_macro_med_RF = np.array([])
F1_weighted_med_RF = np.array([])
Sample_Num_med_RF = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = random_forests_clf(param_grid_rf, X_train_med_aug[:i], y_train_med_aug[:i].astype(int), X_test_med, y_test_med)
Precision_micro_med_RF = np.append(Precision_micro_med_RF, pr_micro)
Precision_macro_med_RF = np.append(Precision_macro_med_RF, pr_macro)
Precision_weighted_med_RF = np.append(Precision_weighted_med_RF, pr_weighted)
Recall_micro_med_RF = np.append(Recall_micro_med_RF, re_micro)
Recall_macro_med_RF = np.append(Recall_macro_med_RF, re_macro)
Recall_weighted_med_RF = np.append(Recall_weighted_med_RF, re_weighted)
F1_micro_med_RF = np.append(F1_micro_med_RF, f1_micro)
F1_macro_med_RF = np.append(F1_macro_med_RF, f1_macro)
F1_weighted_med_RF = np.append(F1_weighted_med_RF, f1_weighted)
Sample_Num_med_RF = np.append(Sample_Num_med_RF, len(y_train_med_aug[:i]))
print((i-590)/step)
# +
df_plot_med_RF = pd.DataFrame()
df_plot_med_RF['Sample Number'] = Sample_Num_med_RF
df_plot_med_RF['Precision Micro'] = Precision_micro_med_RF
df_plot_med_RF['Precision Macro'] = Precision_macro_med_RF
df_plot_med_RF['Precision Weighted'] = Precision_weighted_med_RF
df_plot_med_RF['Recall Micro'] = Recall_micro_med_RF
df_plot_med_RF['Recall Macro'] = Recall_macro_med_RF
df_plot_med_RF['Recall Weighted'] = Recall_weighted_med_RF
df_plot_med_RF['F1 Micro'] = F1_micro_med_RF
df_plot_med_RF['F1 Macro'] = F1_macro_med_RF
df_plot_med_RF['F1 Weighted'] = F1_weighted_med_RF
df_plot_med_RF['Resolution'] = 'Med'
# +
Precision_micro_high_RF = np.array([])
Precision_macro_high_RF = np.array([])
Precision_weighted_high_RF = np.array([])
Recall_micro_high_RF = np.array([])
Recall_macro_high_RF = np.array([])
Recall_weighted_high_RF = np.array([])
F1_micro_high_RF = np.array([])
F1_macro_high_RF = np.array([])
F1_weighted_high_RF = np.array([])
Sample_Num_high_RF = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = random_forests_clf(param_grid_rf, X_train_high_aug[:i], y_train_high_aug[:i].astype(int), X_test_high, y_test_high)
Precision_micro_high_RF = np.append(Precision_micro_high_RF, pr_micro)
Precision_macro_high_RF = np.append(Precision_macro_high_RF, pr_macro)
Precision_weighted_high_RF = np.append(Precision_weighted_high_RF, pr_weighted)
Recall_micro_high_RF = np.append(Recall_micro_high_RF, re_micro)
Recall_macro_high_RF = np.append(Recall_macro_high_RF, re_macro)
Recall_weighted_high_RF = np.append(Recall_weighted_high_RF, re_weighted)
F1_micro_high_RF = np.append(F1_micro_high_RF, f1_micro)
F1_macro_high_RF = np.append(F1_macro_high_RF, f1_macro)
F1_weighted_high_RF = np.append(F1_weighted_high_RF, f1_weighted)
Sample_Num_high_RF = np.append(Sample_Num_high_RF, len(y_train_high_aug[:i]))
print((i-590)/step)
# +
df_plot_high_RF = pd.DataFrame()
df_plot_high_RF['Sample Number'] = Sample_Num_high_RF
df_plot_high_RF['Precision Micro'] = Precision_micro_high_RF
df_plot_high_RF['Precision Macro'] = Precision_macro_high_RF
df_plot_high_RF['Precision Weighted'] = Precision_weighted_high_RF
df_plot_high_RF['Recall Micro'] = Recall_micro_high_RF
df_plot_high_RF['Recall Macro'] = Recall_macro_high_RF
df_plot_high_RF['Recall Weighted'] = Recall_weighted_high_RF
df_plot_high_RF['F1 Micro'] = F1_micro_high_RF
df_plot_high_RF['F1 Macro'] = F1_macro_high_RF
df_plot_high_RF['F1 Weighted'] = F1_weighted_high_RF
df_plot_high_RF['Resolution'] = 'High'
# -
df_plot_RF = df_plot_low_RF.append(df_plot_med_RF, ignore_index=True)
df_plot_RF = df_plot_RF.append(df_plot_high_RF, ignore_index=True)
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_RF)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
path_RF = './data_augmentation/results/df_RF.csv'
df_plot_RF.to_csv(path_RF, index=True)
param_grid_mlp = {'alpha': [np.round(10**-1/4 * i, decimals=2) for i in range(1, 11, 1)],
'hidden_layer_sizes': [2**i for i in range(2, 12, 1)]}
print('Parameters:', param_grid_mlp)
# +
Precision_micro_low_MLP = np.array([])
Precision_macro_low_MLP = np.array([])
Precision_weighted_low_MLP = np.array([])
Recall_micro_low_MLP = np.array([])
Recall_macro_low_MLP = np.array([])
Recall_weighted_low_MLP = np.array([])
F1_micro_low_MLP = np.array([])
F1_macro_low_MLP = np.array([])
F1_weighted_low_MLP = np.array([])
Sample_Num_low_MLP = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = mlp_clf(param_grid_mlp, X_train_low_aug[:i], y_train_low_aug[:i].astype(int), X_test_low, y_test_low)
Precision_micro_low_MLP = np.append(Precision_micro_low_MLP, pr_micro)
Precision_macro_low_MLP = np.append(Precision_macro_low_MLP, pr_macro)
Precision_weighted_low_MLP = np.append(Precision_weighted_low_MLP, pr_weighted)
Recall_micro_low_MLP = np.append(Recall_micro_low_MLP, re_micro)
Recall_macro_low_MLP = np.append(Recall_macro_low_MLP, re_macro)
Recall_weighted_low_MLP = np.append(Recall_weighted_low_MLP, re_weighted)
F1_micro_low_MLP = np.append(F1_micro_low_MLP, f1_micro)
F1_macro_low_MLP = np.append(F1_macro_low_MLP, f1_macro)
F1_weighted_low_MLP = np.append(F1_weighted_low_MLP, f1_weighted)
Sample_Num_low_MLP = np.append(Sample_Num_low_MLP, len(y_train_low_aug[:i]))
print((i-590)/step)
# +
df_plot_low_MLP = pd.DataFrame()
df_plot_low_MLP['Sample Number'] = Sample_Num_low_MLP
df_plot_low_MLP['Precision Micro'] = Precision_micro_low_MLP
df_plot_low_MLP['Precision Macro'] = Precision_macro_low_MLP
df_plot_low_MLP['Precision Weighted'] = Precision_weighted_low_MLP
df_plot_low_MLP['Recall Micro'] = Recall_micro_low_MLP
df_plot_low_MLP['Recall Macro'] = Recall_macro_low_MLP
df_plot_low_MLP['Recall Weighted'] = Recall_weighted_low_MLP
df_plot_low_MLP['F1 Micro'] = F1_micro_low_MLP
df_plot_low_MLP['F1 Macro'] = F1_macro_low_MLP
df_plot_low_MLP['F1 Weighted'] = F1_weighted_low_MLP
df_plot_low_MLP['Resolution'] = 'Low'
# +
Precision_micro_med_MLP = np.array([])
Precision_macro_med_MLP = np.array([])
Precision_weighted_med_MLP = np.array([])
Recall_micro_med_MLP = np.array([])
Recall_macro_med_MLP = np.array([])
Recall_weighted_med_MLP = np.array([])
F1_micro_med_MLP = np.array([])
F1_macro_med_MLP = np.array([])
F1_weighted_med_MLP = np.array([])
Sample_Num_med_MLP = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = mlp_clf(param_grid_mlp, X_train_med_aug[:i], y_train_med_aug[:i].astype(int), X_test_med, y_test_med)
Precision_micro_med_MLP = np.append(Precision_micro_med_MLP, pr_micro)
Precision_macro_med_MLP = np.append(Precision_macro_med_MLP, pr_macro)
Precision_weighted_med_MLP = np.append(Precision_weighted_med_MLP, pr_weighted)
Recall_micro_med_MLP = np.append(Recall_micro_med_MLP, re_micro)
Recall_macro_med_MLP = np.append(Recall_macro_med_MLP, re_macro)
Recall_weighted_med_MLP = np.append(Recall_weighted_med_MLP, re_weighted)
F1_micro_med_MLP = np.append(F1_micro_med_MLP, f1_micro)
F1_macro_med_MLP = np.append(F1_macro_med_MLP, f1_macro)
F1_weighted_med_MLP = np.append(F1_weighted_med_MLP, f1_weighted)
Sample_Num_med_MLP = np.append(Sample_Num_med_MLP, len(y_train_med_aug[:i]))
print((i-590)/step)
# +
df_plot_Med_MLP = pd.DataFrame()
df_plot_Med_MLP['Sample Number'] = Sample_Num_Med_MLP
df_plot_Med_MLP['Precision Micro'] = Precision_micro_Med_MLP
df_plot_Med_MLP['Precision Macro'] = Precision_macro_Med_MLP
df_plot_Med_MLP['Precision Weighted'] = Precision_weighted_Med_MLP
df_plot_Med_MLP['Recall Micro'] = Recall_micro_Med_MLP
df_plot_Med_MLP['Recall Macro'] = Recall_macro_Med_MLP
df_plot_Med_MLP['Recall Weighted'] = Recall_weighted_Med_MLP
df_plot_Med_MLP['F1 Micro'] = F1_micro_Med_MLP
df_plot_Med_MLP['F1 Macro'] = F1_macro_Med_MLP
df_plot_Med_MLP['F1 Weighted'] = F1_weighted_Med_MLP
df_plot_Med_MLP['Resolution'] = 'Med'
# +
Precision_micro_high_MLP = np.array([])
Precision_macro_high_MLP = np.array([])
Precision_weighted_high_MLP = np.array([])
Recall_micro_high_MLP = np.array([])
Recall_macro_high_MLP = np.array([])
Recall_weighted_high_MLP = np.array([])
F1_micro_high_MLP = np.array([])
F1_macro_high_MLP = np.array([])
F1_weighted_high_MLP = np.array([])
Sample_Num_high_MLP = np.array([])
for i in range(590, 2361, step):
pr_micro, pr_macro, pr_weighted, re_micro, re_macro, re_weighted, f1_micro, f1_macro, f1_weighted = mlp_clf(param_grid_mlp, X_train_high_aug[:i], y_train_high_aug[:i].astype(int), X_test_high, y_test_high)
Precision_micro_high_MLP = np.append(Precision_micro_high_MLP, pr_micro)
Precision_macro_high_MLP = np.append(Precision_macro_high_MLP, pr_macro)
Precision_weighted_high_MLP = np.append(Precision_weighted_high_MLP, pr_weighted)
Recall_micro_high_MLP = np.append(Recall_micro_high_MLP, re_micro)
Recall_macro_high_MLP = np.append(Recall_macro_high_MLP, re_macro)
Recall_weighted_high_MLP = np.append(Recall_weighted_high_MLP, re_weighted)
F1_micro_high_MLP = np.append(F1_micro_high_MLP, f1_micro)
F1_macro_high_MLP = np.append(F1_macro_high_MLP, f1_macro)
F1_weighted_high_MLP = np.append(F1_weighted_high_MLP, f1_weighted)
Sample_Num_high_MLP = np.append(Sample_Num_high_MLP, len(y_train_high_aug[:i]))
print((i-590)/step)
# +
df_plot_high_MLP = pd.DataFrame()
df_plot_high_MLP['Sample Number'] = Sample_Num_high_MLP
df_plot_high_MLP['Precision Micro'] = Precision_micro_high_MLP
df_plot_high_MLP['Precision Macro'] = Precision_macro_high_MLP
df_plot_high_MLP['Precision Weighted'] = Precision_weighted_high_MLP
df_plot_high_MLP['Recall Micro'] = Recall_micro_high_MLP
df_plot_high_MLP['Recall Macro'] = Recall_macro_high_MLP
df_plot_high_MLP['Recall Weighted'] = Recall_weighted_high_MLP
df_plot_high_MLP['F1 Micro'] = F1_micro_high_MLP
df_plot_high_MLP['F1 Macro'] = F1_macro_high_MLP
df_plot_high_MLP['F1 Weighted'] = F1_weighted_high_MLP
df_plot_high_MLP['Resolution'] = 'High'
# -
df_plot_MLP = df_plot_low_MLP.append(df_plot_med_MLP, ignore_index=True)
df_plot_MLP = df_plot_MLP.append(df_plot_high_MLP, ignore_index=True)
fig, ax = plt.subplots(figsize=a4_dims)
ax = sns.lineplot(x="Sample Number", y="F1 Micro", legend="full", hue="Resolution", palette=palette_3,
data=df_plot_MLP)
#sns.scatterplot
#plt.legend(loc='center left', bbox_to_anchor=(1.0, 0.5))
#ax.set_xlim(-2.5,3.5)
#plt.savefig('./results/graphics/ST_LR.pdf', format='pdf', bbox_inches='tight')
plt.show()
path_MLP = './data_augmentation/results/df_MLP.csv'
df_plot_MLP.to_csv(path_MLP, index=True)
| 8_data_augmentation_models_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Creates text file with the considered criteria for the authors disambiguation
import pandas as pd
from unidecode import unidecode
def normalize_name(name):
normalized_name = unidecode(name)
normalized_name = str.lower(normalized_name)
return normalized_name
#Reads in all authors' names and associated dataset metadata
df = pd.read_csv("/DSKG_BETA_DISAMBIGUATION.csv")
number_id = 10000
author_id_names = []
author_id_coauthors = []
final_list = []
i = 0
while i < len(df):
authors_title = df['title'][i]
authors_years = df["issued"][i]
authors_years = str(authors_years).split("-")[0]
authors_dataset = df['dataset'][i]
authors_publisher = df['publisherName'][i]
authors_contributor = df['contributorName'][i]
authors_dataset_topic = df['theme'][i]
authors_string = df['creatorPersonName'][i]
authors = str(authors_string).split(", ")
authors_dataset_lda_topic_distribution = df['LDA_Topic_Distribution'][i]
for author in authors:
if str(author).startswith(" "):
max_len = len(author)
author = author[1:max_len]
author_id = str(number_id) + '\t' + author
coauthors = str(authors_string).replace(author, "")
coauthors = coauthors.replace(", ,", ", ")
coauthors_normalized = normalize_name(coauthors)
author_name_normalized = normalize_name(author)
authors_title_words = authors_title.split(" ")
authors_title_words_usefull = ""
for word in authors_title_words:
if len(word) > 4:
authors_title_words_usefull = authors_title_words_usefull + word + ", "
max_len = len(authors_title_words_usefull)
unwanted_cut_off = max_len - 2
authors_title_words_usefull = authors_title_words_usefull[0:unwanted_cut_off]
if coauthors_normalized.startswith(", "):
max_len = len(coauthors)
coauthors_normalized = coauthors_normalized[2:max_len]
if coauthors_normalized.endswith(", "):
max_len = len(coauthors)
unwanted_cut_off = max_len - 2
coauthors_normalized = coauthors_normalized[0:unwanted_cut_off]
if str(authors_dataset_topic) == "nan":
authors_dataset_topic = ""
if str(coauthors) == "nan":
coauthors = ""
if str(authors_publisher) == "nan":
authors_publisher = ""
if str(authors_contributor) == "nan":
authors_contributor = ""
if str(authors_years) == "nan":
authors_years = ""
if str(author) != "nan":
final_list.append(author_name_normalized + '\t' + str(author_id) + '\t' + str(authors_dataset_topic) + '\t' + str(authors_dataset) + '\t' + str(coauthors_normalized) + '\t' + str(authors_publisher) + '\t' + str(authors_contributor) + '\t' + str(authors_years) + '\t' + str(authors_title_words_usefull) + '\t' + str(authors_dataset_lda_topic_distribution) + '\n')
number_id += 1
i += 1
final_list.sort()
#Saves required criteria for author disambiguation in text file
with open("/Author_Disambiguation.txt", "w") as outp:
for line in final_list:
outp.write(line)
# +
#Developed Author Disambiguation
import pandas as pd
from pyjarowinkler import distance
import itertools
import math
import re
from scipy import spatial
def compare_first_names(author1, author2):
firstName1 = author1.split("\t")[0].strip().rsplit(' ', 1)[0]
firstName2 = author2.split("\t")[0].strip().rsplit(' ', 1)[0]
if len(str(firstName1)) >= 5:
if firstName1 == firstName2:
return 2
else:
return 0
else:
if firstName1 == firstName2:
return 1
else:
return 0
def compare_initials(author1, author2):
names1 = author1.split("\t")[0].strip().split(" ")
names2 = author2.split("\t")[0].strip().split(" ")
initials1 = ""
initials2 = ""
for name in names1:
initials1 += name[:1]
for name in names2:
initials2 += name[:1]
if len(initials1) == 1:
if initials1 == initials2:
return 1
else:
return 0
elif len(initials1) == 2:
if initials1 == initials2:
return 2
else:
return 0
else:
if initials1 == initials2:
return 3
else:
return 0
def compare_titles(author1, author2):
titles1 = author1.split("\t")[9].strip().split(", ")
titles2 = author2.split("\t")[9].strip().split(", ")
if len(titles1) == 0 or len(titles2) == 0:
return 0
else:
titles1_set = set(titles1)
titles2_set = set(titles2)
return len(titles1_set.intersection(titles2_set))
def compare_years(author1, author2):
if author1.split("\t")[8].strip() == "" or author2.split("\t")[8].strip() == "":
return False
else:
year1 = author1.split("\t")[8].strip()
year2 = author2.split("\t")[8].strip()
return abs(int(year1) - int(year2)) < 10
def compare_coauthors(author1, author2):
coauthors1 = set(author1.split("\t")[5].strip().split(", "))
coauthors2 = set(author2.split("\t")[5].strip().split(", "))
if len(coauthors1) == 0 or len(coauthors2) == 0:
return 0
else:
return len(coauthors1.intersection(coauthors2))
def compare_topic_datasets(author1, author2):
topics1 = set(author1.split("\t")[3].strip().split(", "))
topics2 = set(author2.split("\t")[3].strip().split(", "))
if len(topics1) == 0 or len(topics2) == 0:
return 0
else:
return len(topics1.intersection(topics2))
#Cosinus-Similarity LDA-Vectors
def compare_lda_topic_distribution(author1, author2):
author1 = author1.split("\t")[10].strip()
author2 = author2.split("\t")[10].strip()
author1 = re.sub(r'[0-9], ', '', author1)
author1 = author1.replace("[", "").replace("]", "").replace("(", "").replace(")", "")
lda_topic_distribution1 = [float(percent) for percent in author1.split(', ')]
author2 = re.sub(r'[0-9], ', '', author2)
author2 = author2.replace("[", "").replace("]", "").replace("(", "").replace(")", "")
lda_topic_distribution2 = [float(percent) for percent in author2.split(', ')]
cos_similarity = 1 - spatial.distance.cosine(lda_topic_distribution1, lda_topic_distribution2)
return cos_similarity
def compare_publisher(author1, author2):
publisher1 = set(author1.split("\t")[6].strip().split(", "))
publisher2 = set(author2.split("\t")[6].strip().split(", "))
if len(publisher1) == 0 or len(publisher2) == 0:
return 0
else:
return len(publisher1.intersection(publisher2))
def compare_contributer(author1, author2):
contributer1 = set(author1.split("\t")[7].strip().split(", "))
contributer2 = set(author2.split("\t")[7].strip().split(", "))
if len(contributer1) < 3 or len(contributer2) < 3:
return 0
else:
return len(contributer1.intersection(contributer2))
def compare_authors(author1, author2):
score = 0
if compare_initials(author1, author2) == 2:
score += 3
elif compare_initials(author1, author2) == 3:
score += 7
elif compare_initials(author1, author2) == 0:
score -= 5
if compare_first_names(author1, author2) == 1:
score += 2
elif compare_first_names(author1, author2) == 2:
score += 5
if compare_coauthors(author1, author2) == 1:
score += 4
elif compare_coauthors(author1, author2) == 2:
score += 7
elif compare_coauthors(author1, author2) > 2:
score += 10
if compare_titles(author1, author2) == 1:
score += 2
elif compare_titles(author1, author2) == 2:
score += 4
elif compare_titles(author1, author2) > 2:
score += 6
if compare_lda_topic_distribution(author1, author2) >= 0.99:
score += 4
elif compare_lda_topic_distribution(author1, author2) >= 0.95:
score += 3
elif compare_lda_topic_distribution(author1, author2) >= 0.75:
score += 2
if compare_years(author1, author2):
score += 1
if compare_topic_datasets(author1, author2) >= 1:
score += 1
if compare_publisher(author1, author2) >= 1:
score += 1
if compare_contributer(author1, author2) >= 1:
score += 3
return score
def get_id(author):
return author.split("\t")[1]
def add_to_mapping(dict_of_maps, entry1, entry2):
if entry2 not in dict_of_maps:
dict_of_maps[entry1] = entry2
return dict_of_maps
else:
return add_to_mapping(dict_of_maps, entry1, dict_of_maps[entry2])
def merge_authors(tuple_of_authors):
author1 = tuple_of_authors[0].strip("\n").split("\t")
author2 = tuple_of_authors[1].strip("\n").split("\t")
output = "\t".join(author1[0:11])
return output
def disambiguate(list_of_authors, result, positive, negative):
author_dictionary = {get_id(author): author.strip("\n") for author in list_of_authors}
author_list = [get_id(author) for author in list_of_authors]
mapping = {}
result = result.copy()
#Creates tuples of two authors each
comparisons = list(itertools.combinations(author_list, 2))
for item in comparisons:
try:
if compare_authors(author_dictionary[item[0]], author_dictionary[item[1]]) >= 11:
positive += 1
if item[0] not in mapping:
mapping = add_to_mapping(mapping, item[1], item[0])
result = add_to_mapping(result, item[1], item[0])
author_dictionary[item[0]] = merge_authors((author_dictionary[item[0]], author_dictionary[item[1]]))
del author_dictionary[item[1]]
else:
author_dictionary[mapping[item[0]]] = merge_authors((author_dictionary[mapping[item[0]]], author_dictionary[item[1]]))
mapping = add_to_mapping(mapping, item[1], item[0])
result = add_to_mapping(result, item[1], item[0])
del author_dictionary[item[1]]
else:
negative += 1
except KeyError:
pass
if not len(mapping) == 0:
return disambiguate([author_dictionary[author] for author in author_dictionary], result, positive, negative)
else:
return author_dictionary, result, positive, negative
# -
#Execute the author disambiguation
with open("/Author_Disambiguation.txt", "r") as inp:
with open("/results_summary.txt", "w") as outp:
with open("/all_positives.txt", "w") as outp2:
with open("/disambiguated_file.txt", "w") as outp3:
positive = 0
negative = 0
previous_name = ""
current_authors = []
line_count = 1
for line in inp:
name = line.split("\t")[0].strip()
if previous_name == "":
previous_name = name
current_authors.append(line)
elif distance.get_jaro_distance(str.lower(name), str.lower(previous_name), winkler=True, scaling=0.1) > 0.9:
previous_name = name
current_authors.append(line)
else:
result = {}
authors, result, positive, negative = disambiguate(current_authors, result, positive, negative)
previous_name = name
current_authors = [line]
for item in authors:
outp3.write(authors[item] + "\n")
for item in result:
outp2.write(item + "\t" + result[item] + "\n")
line_count += 1
result = {}
authors, result, positive, negative = disambiguate(current_authors, result, positive, negative)
for item in authors:
outp3.write(authors[item] + "\n")
for item in result:
outp2.write(item + "\t" + result[item] + "\n")
total_comparisons = positive + negative
outp.write("Total comparisons: " + str(total_comparisons) + "\n")
outp.write("Total positives: " + str(positive) + ": " + str(positive/total_comparisons) + "\n")
outp.write("Total negatives: " + str(negative) + ": " + str(negative/total_comparisons))
| author-disambiguation/author_disambiguation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import arcpy, os, datetime
from arcpy import env
import pandas as pd
env.workspace = r"T:\MPO\RTP\FY20 2045 Update\Data and Resources\Network_Analysis\Network_Analysis.gdb"
env.overwriteOutput = True
# need to restart the kernel after updating the function
import AccessibilitySpatialAnalysis as asa
asa.calculateAccessibility()
asa.AccessibilitySpatialJoin_HH()
asa.AccessibilitySpatialJoin_HH(AOI = "EFA")
AOIs = ["MPO", "EFA", "NEFA"]
travel_modes = ["Biking", "Walking"]
sas = ["Jobs", "Amenities"]
years = [2020, 2045]
outpath = r"T:\MPO\RTP\FY20 2045 Update\Data and Resources\Network_Analysis"
now = datetime.datetime.now()
for sa in sas:
byService = []
for travel_mode in travel_modes:
byTravelMode = []
colnms = []
for year in years:
byYear = []
colnm = []
for AOI in AOIs:
acc = asa.calculateAccessibility(service = sa,
travel_mode = travel_mode,
year = year,
AOI = AOI)
print("Got the accessibility number for {0} in {1} by {2} in {3}...".format(sa, AOI, travel_mode, year))
byYear.append(acc)
colnm.append(AOI+str(year))
byTravelMode += byYear
colnms += colnm
byService.append(byTravelMode)
print("Got the accessibility table for " + sa + ":")
df = pd.DataFrame(byService)
df.columns = colnms
df.index= travel_modes
print(df)
df.to_csv(os.path.join(outpath, sa+"AccessHH.csv"))
later = datetime.datetime.now()
elapsed = later - now
print("total time used: {0}".format(elapsed))
now = datetime.datetime.now()
for sa in sas:
byService = []
for travel_mode in travel_modes:
byTravelMode = []
colnms = []
for year in years:
out = asa.GetEFA_numbers_HH(service = sa,
travel_mode = travel_mode,
year = year)
byYear = out[0]
colnm = out[1]
byTravelMode += byYear
colnms += colnm
byService.append(byTravelMode)
print("Got the accessibility table for " + sa + ":")
df = pd.DataFrame(byService)
df.columns = colnms
df.index= travel_modes
print(df)
df.to_csv(os.path.join(outpath, sa + "AccessEFA_HH.csv"))
later = datetime.datetime.now()
elapsed = later - now
print("total time used: {0}".format(elapsed))
asa.AccessibilityEquityArea_HH()
asa.AccessibilityEquityArea_HH(service = "Amenities",
travel_mode = "Walking",
year = 2020)
asa.AccessibilityEquityArea_HH(service = "Amenities",
travel_mode = "Walking",
year = 2045)
for year in years:
for sa in sas:
for travel_mode in travel_modes:
asa.AccessibilityEquityArea_HH(service = sa,
travel_mode = travel_mode,
year = year)
for year in years:
for AOI in AOIs:
for sa in sas:
for travel_mode in travel_modes:
spatialJoin_out_name = sa + travel_mode + str(year) + "HH_SA"
out_name = AOI + spatialJoin_out_name
if arcpy.Exists(out_name):
arcpy.Delete_management(out_name)
for year in years:
for AOI in AOIs:
for sa in sas:
for travel_mode in travel_modes:
asa.AccessibilitySpatialJoin_HH(
AOI = AOI,
service = sa,
travel_mode = travel_mode,
year = year)
asa.AccessibilitySpatialJoin(keep = "only")
outpath = r"T:\MPO\RTP\FY20 2045 Update\Data and Resources\Network_Analysis"
# 2 years * 3 AOI * 2 travel modes * 2 services
layer_names = ["baseyearHH_FeatureToPoint", "forecastHH_FeatureToPoint"]
jobfields = ["ojobs", "jobs"]
AOIs = ["MPO", "EFA", "NEFA"]
travel_modes = ["Biking", "Walking"]
sas = ["Jobs", "Amenities"]
years = [2020, 2045]
for layer_name in layer_names:
for AOI in AOIs:
for sa in sas:
for travel_mode in travel_modes:
asa.AccessibilitySpatialJoin(layer_name = layer_name,
AOI = AOI,
jobfield = jobfields[layer_names.index(layer_name)],
service = sa,
travel_mode = travel_mode,
year = years[layer_names.index(layer_name)])
now = datetime.datetime.now()
for sa in sas:
byService = []
for travel_mode in travel_modes:
byTravelMode = []
colnms = []
for year in years:
out = asa.AccessibilityEquityArea(service = sa,
travel_mode = travel_mode,
year = year)
byYear = out[0]
colnm = out[1]
byTravelMode += byYear
colnms += colnm
byService.append(byTravelMode)
print("Got the accessibility table for " + sa + ":")
df = pd.DataFrame(byService)
df.columns = colnms
df.index= travel_modes
print(df)
df.to_csv(os.path.join(outpath, sa + "AccessEFA.csv"))
later = datetime.datetime.now()
elapsed = later - now
print("total time used: {0}".format(elapsed))
| analysis/spatial_analysis/asa_jobs_services.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''py_env'': conda)'
# language: python
# name: python37564bitpyenvconda30b1ae30d5b64d6794e2ee9f050e6b8a
# ---
table = {
'ATA':'I', 'ATC':'I', 'ATT':'I', 'ATG':'M',
'ACA':'T', 'ACC':'T', 'ACG':'T', 'ACT':'T',
'AAC':'N', 'AAT':'N', 'AAA':'K', 'AAG':'K',
'AGC':'S', 'AGT':'S', 'AGA':'R', 'AGG':'R',
'CTA':'L', 'CTC':'L', 'CTG':'L', 'CTT':'L',
'CCA':'P', 'CCC':'P', 'CCG':'P', 'CCT':'P',
'CAC':'H', 'CAT':'H', 'CAA':'Q', 'CAG':'Q',
'CGA':'R', 'CGC':'R', 'CGG':'R', 'CGT':'R',
'GTA':'V', 'GTC':'V', 'GTG':'V', 'GTT':'V',
'GCA':'A', 'GCC':'A', 'GCG':'A', 'GCT':'A',
'GAC':'D', 'GAT':'D', 'GAA':'E', 'GAG':'E',
'GGA':'G', 'GGC':'G', 'GGG':'G', 'GGT':'G',
'TCA':'S', 'TCC':'S', 'TCG':'S', 'TCT':'S',
'TTC':'F', 'TTT':'F', 'TTA':'L', 'TTG':'L',
'TAC':'Y', 'TAT':'Y', 'TAA':'_', 'TAG':'_',
'TGC':'C', 'TGT':'C', 'TGA':'_', 'TGG':'W',
}
| PhDtools/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Comparison of Charting Libraries Performance
# +
import json
import pandas as pd
from lets_plot import *
from lets_plot.mapping import as_discrete
LetsPlot.setup_html()
# -
# ### Preliminaries
# +
def get_json_data(file_path):
with open(file_path, "r") as f:
return json.load(f)
def get_data(file_path):
colors = {
'LightningChart': '#e41a1c',
'Plotly.js': '#4daf4a',
'Canvas.js': '#ff7f00',
'eCharts': '#ffff33',
'Lets-Plot groups': '#377eb8',
'Lets-Plot layers': '#984ea3',
}
data = {
'test': [],
'lib': [],
'channels_count': [],
'channel_data_points_per_second': [],
'time_domain_interval': [],
'loadup_speed_ms': [],
'fps_avg': [],
'fps_median': [],
'frame_time_avg': [],
'frame_time_median': [],
'color': [],
}
for test, test_data in get_json_data(file_path).items():
for lib, test_lib_data in test_data.items():
data['test'].append(test)
data['lib'].append(lib)
data['channels_count'].append(test_lib_data['config']['channelsCount'])
data['channel_data_points_per_second'].append(test_lib_data['config']['channelDataPointsPerSecond'])
data['time_domain_interval'].append(test_lib_data['config']['timeDomainInterval'])
data['loadup_speed_ms'].append(test_lib_data['loadupSpeedMs'])
data['fps_avg'].append(test_lib_data['fps']['avg'])
data['fps_median'].append(test_lib_data['fps']['median'])
data['frame_time_avg'].append(test_lib_data['frameTime']['avg'])
data['frame_time_median'].append(test_lib_data['frameTime']['median'])
data['color'].append(colors[lib])
return pd.DataFrame.from_dict(data)
# -
df = get_data("../../../data/lib_comparison.json")
df['load'] = df['channels_count'] * df['time_domain_interval']
df.head(6)
# Here "Lets-Plot groups" is a realization of the benchmark in which each data channel corresponds to a group in aesthetics. In "Lets-Plot layers" each data channel corresponds to a plot layer.
# +
def comparison1(col):
return ggplot(df) + \
geom_line(aes('load', col, color='color'), size=2, \
tooltips=layer_tooltips().line('@lib').format(col, '.2f').line('{0}|@{0}'.format(col))) + \
scale_x_log10() + scale_color_identity() + \
ggtitle('Comparison of {0} through load increasing'.format(col))
def comparison2(col):
return ggplot(df) + \
geom_bar(aes(as_discrete('lib', order_by=col, order=1), col, fill='color'), stat='identity', \
tooltips=layer_tooltips().line('@lib').format(col, '.2f').line('{0}|@{0}'.format(col))) + \
facet_grid(x='test') + \
scale_fill_identity() + \
coord_flip() + \
ggtitle('Comparison of {0} through tests'.format(col)) + \
theme_classic() + \
theme(axis_line_y='blank', axis_ticks_y='blank', axis_text_y='blank', \
strip_background='blank')
# -
# ### Comparison Charts
comparison1('loadup_speed_ms')
comparison1('fps_avg')
comparison1('frame_time_avg')
comparison2('loadup_speed_ms')
comparison2('fps_avg')
comparison2('frame_time_avg')
| source/examples/cookbook/lib_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''venv'': venv)'
# name: python3
# ---
# +
# !pip install pandas==1.3.1
import warnings
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# # %matplotlib inline
import tensorflow
from google.colab import drive
from matplotlib import rcParams
from pandas.tseries.offsets import DateOffset
from sklearn.preprocessing import MinMaxScaler
from tensorflow.keras.callbacks import EarlyStopping, ReduceLROnPlateau, ModelCheckpoint, TensorBoard
from tensorflow.keras.layers import Bidirectional
from tensorflow.keras.layers import Conv1D
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import LSTM
from tensorflow.keras.losses import Huber
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import SGD
from tensorflow.keras import Model
from tensorflow.keras.callbacks import History
rcParams.update({'figure.autolayout': True})
warnings.filterwarnings("ignore")
tensorflow.keras.backend.clear_session()
# Configuration
EPOCHS = 1
DROPOUT = 0.1
BATCH_SIZE = 128
LOOK_BACK = 100
UNITS = LOOK_BACK * 1
VALIDATION_SPLIT = .01
PREDICTION_RANGE = LOOK_BACK
DYNAMIC_RETRAIN = False
USE_SAVED_MODELS = False
SAVE_MODELS = False
def summary(for_model: Model) -> str:
summary_data = []
for_model.summary(print_fn=lambda line: summary_data.append(line))
return '\n'.join(summary_data)
def create_model_callbacks(es_patience: int = 40, lr_patience: int = 30) -> []:
es = EarlyStopping(monitor='loss', min_delta=1e-10, patience=es_patience, verbose=1)
rlr = ReduceLROnPlateau(monitor='loss', factor=0.2, patience=lr_patience, verbose=1)
mcp = ModelCheckpoint(filepath='weights.h5', monitor='loss', verbose=1, save_best_only=True,
save_weights_only=True)
tb = TensorBoard('logs')
return [es, rlr, mcp, tb]
def moving_average(array: [], w: int) -> []:
return np.concatenate((np.full(w - 1, array[w]), np.convolve(array, np.ones(w), 'valid') / w))
def df_info(name: str, data):
print(f'\n{name}.shape: {data.shape}')
print(f'{name}.describe(): {data.describe()}')
print(f'{name}.head(): {data.head()}')
print(f'{name}.tail(): {data.tail()}')
print('\n')
def build_model(n_output: int) -> Model:
new_model = Sequential()
new_model.add(Conv1D(filters=LOOK_BACK, kernel_size=5,
strides=1, padding="causal",
activation="relu",
input_shape=(x.shape[1], N_FEATURES)))
# new_model.add(
# Bidirectional(LSTM(units=UNITS, activation='relu', input_shape=(x.shape[1], N_FEATURES), return_sequences=True)))
# new_model.add(
# Bidirectional(LSTM(units=UNITS, activation='relu', input_shape=(x.shape[1], N_FEATURES))))
# new_model.add(Dropout(DROPOUT))
# new_model.add(LSTM(units=UNITS, return_sequences=True, input_shape=(x.shape[1], N_FEATURES)))
new_model.add(Bidirectional(LSTM(units=UNITS, activation='relu', return_sequences=True)))
# new_model.add(Dropout(DROPOUT))
new_model.add(Bidirectional(LSTM(units=UNITS, activation='tanh', return_sequences=True)))
# new_model.add(Dropout(DROPOUT))
# new_model.add(Bidirectional(LSTM(units=UNITS, activation='relu', return_sequences=True)))
# new_model.add(Dropout(DROPOUT))
new_model.add(Bidirectional(LSTM(units=UNITS, activation='linear')))
# new_model.add(Dropout(DROPOUT))
# new_model.add(LSTM(units=UNITS))
new_model.add(Dense(units=n_output))
# new_model.add(Dense(units=N_FEATURES))
new_model.compile(optimizer='adam', loss='mean_squared_error')
# optimizer = SGD(lr=1e-1, momentum=0.9)
# new_model.compile(loss=Huber(),
# optimizer=optimizer,
# metrics=["mae"])
return new_model
def fit_model(new_x: [], new_y: [], new_model: Model, epochs: int = EPOCHS, split: float = VALIDATION_SPLIT,
es_patience: int = 40, lr_patience: int = 30) -> [Model, History]:
new_model.fit(new_x, new_y, epochs=epochs, batch_size=BATCH_SIZE,
callbacks=create_model_callbacks(es_patience, lr_patience),
validation_split=split
)
new_model.load_weights(filepath="weights.h5")
return [new_model, new_model.history]
def get_updated_x(x_last: [], last_prediction: []) -> []:
# print(f'x_last input values: {x_last[-1]}')
x_last = np.append(x_last[1:], last_prediction)
x_last = x_last.reshape(LOOK_BACK, N_FEATURES)
# print(f'x_last new: {X_last}')
# print(f'x_last input values new: {x_last[-1]}')
return np.expand_dims(x_last, axis=0)
def get_stats() -> str:
return f'loss: {history.history.get("loss")[-1]} \n ' \
f'val_loss: {history_val.history.get("val_loss")[-1]} \n ' \
f'EPOCHS: {EPOCHS} DYNAMIC_RETRAIN: {DYNAMIC_RETRAIN} \n ' \
f'UNITS: {UNITS} \n ' \
f'BATCH_SIZE: {BATCH_SIZE} \n ' \
f'LOOK_BACK: {LOOK_BACK} \n ' \
f'VALIDATION_SPLIT: {VALIDATION_SPLIT} \n ' \
f'DROPOUT: {DROPOUT} \n ' \
f'N_FEATURES: {N_FEATURES} \n ' \
f'PREDICTION_RANGE: {PREDICTION_RANGE} '
def predict(model: Model, x: [], x_test: [], y:[], prediction_range: int = PREDICTION_RANGE) -> []:
y_predict = model.predict(x_test)
for prediction_steps in range(prediction_range):
x_predict = get_updated_x(x[-1], y_predict[-1])
y_predict_new = model.predict(x_predict)
# break at extreme values
if abs(y_predict_new[0, 1]) > 2:
break
print(y_predict_new[0, 1])
x = np.append(x, x_predict, axis=0)
y_predict = np.append(y_predict, y_predict_new, axis=0)
# y_predict = np.append(y_predict, [ct}')
# dynamic retrain
if DYNAMIC_RETRAIN:
y = np.append(y, y_predict_new, axis=0)
model, history = fit_model(x, y, model, epochs=5, es_patience=4, lr_patience=3)
# y = np.append(y, y_predict_new[, model, epochs=5, es_patience=4, lr_patience=3)
return y_predict
drive.mount('/drive')
# Download data
headers = {"User-Agent": "Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:89.0) Gecko/20100101 Firefox/89.0"}
df = pd.read_csv("https://www.coingecko.com/price_charts/export/1/usd.csv", parse_dates=['snapped_at'],
storage_options=headers)
df.to_csv('/Colab Notebooks/cryptofuture/data/btc_price.csv')
df = df.fillna(df.mean())
dates = df.iloc[:, [0]].values
df_info('df', df)
# https://docs.coinmetrics.io/info/metrics
df_coin = pd.read_csv('https://coinmetrics.io/newdata/btc.csv', parse_dates=['date'],
storage_options=headers)
df_coin.to_csv('/Colab Notebooks/cryptofuture/data/btc_metrics.csv')
df_coin = pd.read_csv('data/btc_metrics.csv', parse_dates=['date'])
df_coin = df_coin.drop(columns=['date'])
df_coin = df_coin.fillna(df_coin.mean())
df_coin = df_coin.drop(df_coin.index[:1577])
df_coin.rename(columns={'Unnamed: 0': 'index'}, inplace=True)
df_coin['index'] = df_coin['index'] - 1577
df_coin = df_coin.set_index('index')
df_info('df_coin', df_coin)
# Join dataframes
df = df.join(df_coin)
# Put the date column in the index.
df = df.set_index("snapped_at")
# add moving averages
open_values = df['price'].to_numpy()
print(f'open_values: {open_values}')
for m in range(10, 210, 10):
ma = moving_average(open_values, m).tolist()
print(f'ma_{m}: {ma}')
df[f'ma_{m}'] = ma
# Fill nan values
df = df.fillna(df.mean())
df_info('df', df)
input_feature = df.iloc[:, :].values
N_FEATURES = len(input_feature[0])
input_data = input_feature.copy()
scaler = MinMaxScaler(feature_range=(0, 1))
input_data[:, 0:N_FEATURES] = scaler.fit_transform(input_feature[:, :])
x = []
y = []
for i in range(len(df) - LOOK_BACK - 1):
t = []
for j in range(0, LOOK_BACK):
t.append(input_data[[(i + j)], :])
x.append(t)
y.append(input_data[i + LOOK_BACK, :])
x, y = np.array(x), np.array(y)
x_test = x[-2 * LOOK_BACK:]
x = x.reshape(x.shape[0], LOOK_BACK, N_FEATURES)
x_test = x_test.reshape(x_test.shape[0], LOOK_BACK, N_FEATURES)
print(f'x.shape: {x.shape}')
print(f'x_test.shape: {x_test.shape}')
if USE_SAVED_MODELS:
model_val = tensorflow.keras.models.load_model("models/model_val")
model = tensorflow.keras.models.load_model("models/model")
else:
model_val = build_model(N_FEATURES)
model = build_model(N_FEATURES)
model_val, history_val = fit_model(x, y, model_val)
tensorflow.keras.backend.clear_session()
model, history = fit_model(x, y, model, split=0)
if SAVE_MODELS:
model.save('models/model')
y_predict_val = predict(model_val, x, x_test, y, prediction_range=0)
y_predict = predict(model, x, x_test, y)
# Inverse scale value
y_predict = scaler.inverse_transform(y_predict)
y_predict = y_predict[:, 0]
y_predict_val = scaler.inverse_transform(y_predict_val)
y_predict_val = y_predict_val[:, 0]
plot_dates = dates[-2 * LOOK_BACK:]
add_dates = [dates[-1] + DateOffset(days=x) for x in range(0, PREDICTION_RANGE + 1)]
predict_dates = np.concatenate([plot_dates[:-1], add_dates])
# Plot graph
plt.figure(figsize=(20, 8))
plt.plot(dates[-2 * LOOK_BACK:, 0], input_feature[-2 * LOOK_BACK:, 0], color='green', label='Actual')
plt.plot(predict_dates[:-PREDICTION_RANGE], y_predict_val, color='orange', label='Validation')
plt.plot(predict_dates, y_predict, color='red', label='Prediction')
plt.axvline(dates[-1, 0], color='blue', label='Prediction split')
if VALIDATION_SPLIT > 0:
plt.axvline(dates[int(-1 - VALIDATION_SPLIT * len(x)), 0], color='purple', label='Validation split')
plt.title(f'BTC Price Prediction (NFA! No Warranties!) - USE_SAVED_MODELS: {USE_SAVED_MODELS}')
plt.legend(loc='best', fontsize='xx-large')
plt.xlabel("Time (latest-> oldest)")
plt.ylabel("Opening Price")
plt.figtext(0.7, 0.05, get_stats(), ha="center", fontsize=10, bbox={"facecolor": "orange", "alpha": 0.5, "pad": 5})
plt.annotate(summary(model), (0, 0), (0, -40), xycoords='axes fraction', textcoords='offset points', va='top')
plt.annotate(model.optimizer, (0, 0), (600, -40), xycoords='axes fraction', textcoords='offset points', va='top')
plt.savefig(
f'plots/BTC_price_{pd.to_datetime(df.index[-1]).date()}_{EPOCHS}_{BATCH_SIZE}_{LOOK_BACK}_{history.history.get("loss")[-1]}.png')
plt.show()
| cryptofuture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams['figure.figsize'] = (5,5)
matplotlib.rcParams['figure.dpi'] = 100
data = [1,1,2,3,5,8,13,21]
#plt.figure(figsize=(7,7), dpi=90)
plt.plot(data)
plt.figure()
# [xmin, xmax, ymin, ymax]
plt.axis([0, 5, 0, 5])
x = [1,2,3,4]
y = [4,3,2,1]
plt.plot(x, y)
# -
data = [1,2,4,8,16,32,64]
plt.figure()
plt.title('a Random Title', size='x-large')
plt.ylabel('Values', size='x-large')
plt.ylim([0,70])
plt.xlim([-0.5,6.5])
plt.plot(data, 'r*-', markersize=10, linewidth=2, label='Hello')
plt.tick_params(axis='both', which='major', labelsize=8)
plt.legend(loc=(0.25,0.75), scatterpoints=1)
# + [markdown] deletable=true editable=true
# # References
#
# * https://matplotlib.org/users/pyplot_tutorial.html
# * https://danieltakeshi.github.io/2016-01-16-ipython-jupyter-notebooks-and-matplotlib/
| Plot Size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Secara Visual
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn')
iris = pd.read_csv('data/iris.csv')
iris.shape
iris.head()
iris.plot(x='sepal_length', y='sepal_width')
iris.plot(x='sepal_length', y='sepal_width', kind='scatter')
plt.xlabel('sepal length (cm)')
plt.ylabel('sepal width (cm)')
iris.plot(y='sepal_length', kind='box')
plt.ylabel('sepal width (cm)')
iris.plot(y='sepal_length', kind='hist')
plt.xlabel('sepal length (cm)')
iris.plot(y='sepal_length', kind='hist', bins=30, range=(4,8), density=True)
plt.xlabel('sepal length (cm)')
plt.show()
iris.plot(y='sepal_length', kind='hist', bins=30, range=(4,8), cumulative=True, density=True)
plt.xlabel('sepal length (cm)')
plt.title('Cumulative distribution function (CDF)')
# # Secara Statistik
iris.describe()
iris.count()
iris['sepal_length'].count()
iris.mean()
iris.std()
iris.median()
q = 0.5
iris.quantile(q)
q = [0.25, 0.75]
iris.quantile(q)
iris.min()
iris.max()
iris.plot(kind= 'box')
plt.ylabel('[cm]')
# # Filtering
iris.head()
iris['species'].describe()
iris['species'].value_counts()
iris['species'].unique()
index_setosa = iris['species'] == 'setosa'
index_versicolor = iris['species'] == 'versicolor'
index_virginica = iris['species'] == 'virginica'
setosa = iris[index_setosa]
versicolor = iris[index_versicolor]
virginica = iris[index_virginica]
setosa['species'].unique()
versicolor['species'].unique()
virginica['species'].unique()
setosa.head(2)
versicolor.head(2)
virginica.head(2)
iris.plot(kind= 'hist', bins=50, range=(0,8), alpha=0.4)
plt.title('Entire iris data set')
plt.xlabel('[cm]')
setosa.plot(kind='hist', bins=50, range=(0,8), alpha=0.5)
plt.title('Setosa data set')
plt.xlabel('[cm]')
versicolor.plot(kind='hist', bins=50, range=(0,8), alpha=0.5)
plt.title('Versicolor data set')
plt.xlabel('[cm]')
virginica.plot(kind='hist', bins=50, range=(0,8), alpha=0.3)
plt.title('Virginica data set')
plt.xlabel('[cm]')
| Bagian 4 - Fundamental Pandas/2. Exploratory Data Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HighCWu/anime_biggan_toy/blob/main/colab/Anime_BigGAN_Demo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="BhN1AplL0Hpv"
# ##### Copyright 2019 The TensorFlow Hub Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + id="LMgeG2swVVi6"
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="HPIegH0UsrQ_"
# ##### Copyright 2020 <NAME>.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + id="GpdUkrqPsrRC"
# Copyright 2020 HighCWU. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="AqBuuwrIxlGs"
# # Generating Danbooru Anime Images 256x256 with BigGAN
#
# + [markdown] id="p5AWAusyySDA"
# To get started, connect to a runtime and follow these steps:
#
# 1. (Optional) Select a model in the second code cell below.
# 2. Click **Runtime > Run all** to run each cell in order.
# * Afterwards, the interactive visualizations should update automatically when you modify the settings using the sliders and dropdown menus.
#
# Note: if you run into any issues, youn can try restarting the runtime and rerunning all cells from scratch by clicking **Runtime > Restart and run all...**.
#
# + [markdown] id="_m5jsOM9kXWP"
# ## Setup
# + id="jry_aviZzX9Q"
from google.colab import drive
drive.mount('/content/drive')
# !cp drive/My\ Drive/anime-biggan-256px-run39-607250 ./ -r
# + cellView="both" id="NhlMa_tHs0_W"
# @title Imports and utility functions
import os
import IPython
from IPython.display import display
import numpy as np
import PIL.Image
import pandas as pd
import six
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
import tensorflow_hub as hub
def imgrid(imarray, cols=8, pad=1):
pad = int(pad)
assert pad >= 0
cols = int(cols)
assert cols >= 1
N, H, W, C = imarray.shape
rows = int(np.ceil(N / float(cols)))
batch_pad = rows * cols - N
assert batch_pad >= 0
post_pad = [batch_pad, pad, pad, 0]
pad_arg = [[0, p] for p in post_pad]
imarray = np.pad(imarray, pad_arg, 'constant')
H += pad
W += pad
grid = (imarray
.reshape(rows, cols, H, W, C)
.transpose(0, 2, 1, 3, 4)
.reshape(rows*H, cols*W, C))
return grid[:-pad, :-pad]
def imshow(a, format='png', jpeg_fallback=True):
a = np.asarray(a, dtype=np.uint8)
if six.PY3:
str_file = six.BytesIO()
else:
str_file = six.StringIO()
PIL.Image.fromarray(a).save(str_file, format)
png_data = str_file.getvalue()
try:
disp = display(IPython.display.Image(png_data))
except IOError:
if jpeg_fallback and format != 'jpeg':
print ('Warning: image was too large to display in format "{}"; '
'trying jpeg instead.').format(format)
return imshow(a, format='jpeg')
else:
raise
return disp
class Generator(object):
def __init__(self, module_spec):
self._module_spec = module_spec
self._sess = None
self._graph = tf.Graph()
self._load_model()
@property
def z_dim(self):
return self._z.shape[-1].value
@property
def conditional(self):
return self._labels is not None
def _load_model(self):
with self._graph.as_default():
self._generator = hub.Module(self._module_spec, name="gen_module",
tags={"gen", "bsNone"})
input_info = self._generator.get_input_info_dict()
inputs = {k: tf.placeholder(v.dtype, v.get_shape().as_list(), k)
for k, v in self._generator.get_input_info_dict().items()}
self._samples = self._generator(inputs=inputs, as_dict=True)["generated"]
print("Inputs:", inputs)
print("Outputs:", self._samples)
self._z = inputs["z"]
self._labels = inputs.get("labels", None)
def _init_session(self):
if self._sess is None:
self._sess = tf.Session(graph=self._graph)
self._sess.run(tf.global_variables_initializer())
def get_noise(self, num_samples, seed=None):
if np.isscalar(seed):
np.random.seed(seed)
return np.random.randn(num_samples, self.z_dim)
z = np.empty(shape=(len(seed), self.z_dim), dtype=np.float32)
for i, s in enumerate(seed):
np.random.seed(s)
z[i] = np.random.randn(self.z_dim)
return z
def get_samples(self, z, labels=None):
with self._graph.as_default():
self._init_session()
feed_dict = {self._z: z}
if self.conditional:
assert labels is not None
assert labels.shape[0] == z.shape[0]
feed_dict[self._labels] = labels
samples = self._sess.run(self._samples, feed_dict=feed_dict)
return np.uint8(np.clip(255 * samples, 0, 255))
class Discriminator(object):
def __init__(self, module_spec):
self._module_spec = module_spec
self._sess = None
self._graph = tf.Graph()
self._load_model()
@property
def conditional(self):
return "labels" in self._inputs
@property
def image_shape(self):
return self._inputs["images"].shape.as_list()[1:]
def _load_model(self):
with self._graph.as_default():
self._discriminator = hub.Module(self._module_spec, name="disc_module",
tags={"disc", "bsNone"})
input_info = self._discriminator.get_input_info_dict()
self._inputs = {k: tf.placeholder(v.dtype, v.get_shape().as_list(), k)
for k, v in input_info.items()}
self._outputs = self._discriminator(inputs=self._inputs, as_dict=True)
print("Inputs:", self._inputs)
print("Outputs:", self._outputs)
def _init_session(self):
if self._sess is None:
self._sess = tf.Session(graph=self._graph)
self._sess.run(tf.global_variables_initializer())
def predict(self, images, labels=None):
with self._graph.as_default():
self._init_session()
feed_dict = {self._inputs["images"]: images}
if "labels" in self._inputs:
assert labels is not None
assert labels.shape[0] == images.shape[0]
feed_dict[self._inputs["labels"]] = labels
return self._sess.run(self._outputs, feed_dict=feed_dict)
# + [markdown] id="msTFS1UPkugr"
# ## Select a model
# + cellView="both" id="-hBEi9IFdoI-"
# @title Select a model { run: "auto" }
model_name = "BigGAN 256x256 Danbooru Plus" # @param ["BigGAN 256x256 Danbooru Plus"]
models = {
"BigGAN 256x256 Danbooru Plus": "anime-biggan-256px-run39-607250/tfhub",
}
module_spec = models[model_name.split(" (")[0]]
print("Module spec:", module_spec)
tf.reset_default_graph()
print("Loading model...")
sampler = Generator(module_spec)
print("Model loaded.")
# + [markdown] id="ePQuAme_kxLj"
# ## Sample
# + cellView="both" id="kGgTXtFYq_FV"
# @title Sampling { run: "auto" }
num_rows = 2 # @param {type: "slider", min:1, max:16}
num_cols = 3 # @param {type: "slider", min:1, max:16}
noise_seed = 0 # @param {type:"slider", min:0, max:100, step:1}
label_str = "-1) Random" # @param ["-1) Random"]
num_classes = 1000
num_samples = num_rows * num_cols
z = sampler.get_noise(num_samples, seed=noise_seed)
label = int(label_str.split(')')[0])
if label == -1:
labels = np.random.randint(0, num_classes, size=(num_samples))
else:
labels = np.asarray([label] * num_samples)
samples = sampler.get_samples(z, labels)
imshow(imgrid(samples, cols=num_cols))
# + cellView="form" id="vCffdVZvTtxL"
# @title Interpolation { run: "auto" }
num_samples = 1 # @param {type: "slider", min: 1, max: 6, step: 1}
num_interps = 6 # @param {type: "slider", min: 2, max: 10, step: 1}
noise_seed_A = 0 # @param {type: "slider", min: 0, max: 100, step: 1}
noise_seed_B = 100 # @param {type: "slider", min: 0, max: 100, step: 1}
noise_seed_L = 1 # @param {type: "slider", min: 0, max: 5000, step: 1}
label_str = "-1) Random" # @param ["-1) Random"]
def interpolate(A, B, num_interps):
alphas = np.linspace(0, 1, num_interps)
if A.shape != B.shape:
raise ValueError('A and B must have the same shape to interpolate.')
return np.array([((1-a)*A + a*B)/np.sqrt(a**2 + (1-a)**2) for a in alphas])
def interpolate_and_shape(A, B, num_interps):
interps = interpolate(A, B, num_interps)
return (interps.transpose(1, 0, *range(2, len(interps.shape)))
.reshape(num_samples * num_interps, -1))
label = int(label_str.split(')')[0])
if label == -1:
label = np.random.RandomState(noise_seed_L).randint(0, num_classes)
print('Use label index:', label, '.')
print('Different labels basically cause only slight color changes.')
labels = np.asarray([label] * num_samples * num_interps)
z_A = sampler.get_noise(num_samples, seed=noise_seed_A)
z_B = sampler.get_noise(num_samples, seed=noise_seed_B)
z = interpolate_and_shape(z_A, z_B, num_interps)
samples = sampler.get_samples(z, labels)
imshow(imgrid(samples, cols=num_interps))
# + [markdown] id="esW0Up95Ob6U"
# ## Discriminator
# + id="ButxPSq0OzgL"
disc = Discriminator(module_spec)
batch_size = 4
num_classes = 1000
images = np.random.random(size=[batch_size] + disc.image_shape)
labels = np.random.randint(0, num_classes, size=(batch_size))
disc.predict(images, labels=labels)
| colab/Anime_BigGAN_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="VZHxjNNjn8m_" colab_type="text"
# # Imports and Installs
# + id="1_Wcb3Xjnx4Y" colab_type="code" outputId="084eb126-5144-405d-9d11-2bb913e0b1e8" executionInfo={"status": "ok", "timestamp": 1557126040305, "user_tz": -120, "elapsed": 13247, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 303}
# Downgrate numpy to fix a problem
# !pip install numpy==1.16.2
# + id="v_fMMMeun_fl" colab_type="code" colab={}
# import the required libraries
import matplotlib.pyplot as plt
import numpy as np
import time
import random
#import cPickle
import codecs
import collections
import os
import math
import json
import tensorflow as tf
from six.moves import xrange
# + id="wqzqLF23oCV_" colab_type="code" outputId="00908cdc-07c9-47ff-c6a0-92bec514535c" executionInfo={"status": "ok", "timestamp": 1557126069590, "user_tz": -120, "elapsed": 710, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
tf.logging.info("TensorFlow Version: %s", tf.__version__)
# + id="cuDoixZtok-P" colab_type="code" colab={}
# !pip install -q magenta
# + id="-ZmOq5gaolpE" colab_type="code" outputId="6e2a1fdf-8b0d-4f5c-f6c0-7d0e95a2cb4e" executionInfo={"status": "ok", "timestamp": 1557126084401, "user_tz": -120, "elapsed": 6501, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 141}
# import our command line tools
from magenta.models.sketch_rnn.sketch_rnn_train import *
from magenta.models.sketch_rnn.model import *
from magenta.models.sketch_rnn.utils import *
from magenta.models.sketch_rnn.rnn import *
# + [markdown] id="xaQtuIvUoOW3" colab_type="text"
# # Data Download
# + id="s-0PUBXIoQ8C" colab_type="code" outputId="e404d717-e1cd-400b-be6e-83b8901569bc" executionInfo={"status": "ok", "timestamp": 1557126095805, "user_tz": -120, "elapsed": 6506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 123}
# !git clone 'https://github.com/dbt-ethz/home-castle.git'
# + id="s2XYelzxozuZ" colab_type="code" colab={}
data_dir = './home-castle/datasets'
models_root_dir = './home-castle/models'
model_dir = './home-castle/models'
# + [markdown] id="i1Zej-oqpFkt" colab_type="text"
# # Load Model
# + id="fmg0TLX4pHwI" colab_type="code" colab={}
def load_env_compatible(data_dir, model_dir):
"""Loads environment for inference mode, used in jupyter notebook."""
# modified https://github.com/tensorflow/magenta/blob/master/magenta/models/sketch_rnn/sketch_rnn_train.py
# to work with depreciated tf.HParams functionality
model_params = sketch_rnn_model.get_default_hparams()
with tf.gfile.Open(os.path.join(model_dir, 'model_config.json'), 'r') as f:
data = json.load(f)
fix_list = ['conditional', 'is_training', 'use_input_dropout', 'use_output_dropout', 'use_recurrent_dropout']
for fix in fix_list:
data[fix] = (data[fix] == 1)
model_params.parse_json(json.dumps(data))
return load_dataset(data_dir, model_params, inference_mode=True)
def load_model_compatible(model_dir):
"""Loads model for inference mode, used in jupyter notebook."""
# modified https://github.com/tensorflow/magenta/blob/master/magenta/models/sketch_rnn/sketch_rnn_train.py
# to work with depreciated tf.HParams functionality
model_params = sketch_rnn_model.get_default_hparams()
with tf.gfile.Open(os.path.join(model_dir, 'model_config.json'), 'r') as f:
data = json.load(f)
fix_list = ['conditional', 'is_training', 'use_input_dropout', 'use_output_dropout', 'use_recurrent_dropout']
for fix in fix_list:
data[fix] = (data[fix] == 1)
model_params.parse_json(json.dumps(data))
model_params.batch_size = 1 # only sample one at a time
eval_model_params = sketch_rnn_model.copy_hparams(model_params)
eval_model_params.use_input_dropout = 0
eval_model_params.use_recurrent_dropout = 0
eval_model_params.use_output_dropout = 0
eval_model_params.is_training = 0
sample_model_params = sketch_rnn_model.copy_hparams(eval_model_params)
sample_model_params.max_seq_len = 1 # sample one point at a time
return [model_params, eval_model_params, sample_model_params]
# + id="uCsUZAAcpIyR" colab_type="code" outputId="23f6cf0f-a3ef-401f-e080-3779c0d9080e" executionInfo={"status": "ok", "timestamp": 1557126139615, "user_tz": -120, "elapsed": 11156, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 158}
[train_set, valid_set, test_set, hps_model, eval_hps_model, sample_hps_model] = load_env_compatible(data_dir, model_dir)
# + id="bsaX5cr4pPQb" colab_type="code" outputId="222894e8-80c8-45d2-e00b-358bd92ecd1b" executionInfo={"status": "ok", "timestamp": 1557126287880, "user_tz": -120, "elapsed": 4838, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 765}
# construct the sketch-rnn model here:
reset_graph()
model = Model(hps_model)
eval_model = Model(eval_hps_model, reuse=True)
sample_model = Model(sample_hps_model, reuse=True)
# + id="6ZQMsBuvpaES" colab_type="code" colab={}
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# + id="CwUxiDcNpddp" colab_type="code" outputId="4eedfa70-6a49-490b-c1b2-d2cfdc4e16df" executionInfo={"status": "ok", "timestamp": 1557126297272, "user_tz": -120, "elapsed": 630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 125}
# loads the weights from checkpoint into our model
load_checkpoint(sess, model_dir)
# + [markdown] id="paAYeiXlpegV" colab_type="text"
# # Helper Functions
# + id="PXz7SXvppgKg" colab_type="code" colab={}
def encode(input_strokes):
strokes = to_big_strokes(input_strokes, max_len=eval_model.hps.max_seq_len).tolist()
strokes.insert(0, [0, 0, 1, 0, 0])
seq_len = [len(input_strokes)]
return sess.run(eval_model.batch_z, feed_dict={eval_model.input_data: [strokes], eval_model.sequence_lengths: seq_len})[0]
# + id="1jYRrDd7pnAm" colab_type="code" colab={}
def decode(z_input=None, temperature=0.1, factor=0.2):
z = None
if z_input is not None:
z = [z_input]
sample_strokes, m = sample(sess, sample_model, seq_len=eval_model.hps.max_seq_len, temperature=temperature, z=z)
strokes = to_normal_strokes(sample_strokes)
return strokes
# + [markdown] id="G-Z1nij6px9D" colab_type="text"
# # Visualise
# + id="t_-Kg_eGpzMb" colab_type="code" colab={}
# get a sample drawing from the test set
stroke = test_set.random_sample()
# + id="3VCI_DUap2JV" colab_type="code" outputId="66a5490f-e6e0-4e06-eaa4-1e68f1367d2d" executionInfo={"status": "ok", "timestamp": 1557126324039, "user_tz": -120, "elapsed": 1328, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 264}
myabssk = strokes_to_lines(stroke)
plt.axis('equal')
plt.axis('off')
for s in myabssk:
xs = [v[0] for v in s]
ys = [-v[1] for v in s]
plt.plot(xs,ys,'#0088ff')
# + [markdown] id="AL4f-QQtqDq1" colab_type="text"
# # Encode and Decode
# + id="JtoF1SeKqFQm" colab_type="code" colab={}
z = encode(stroke)
d = decode(z, temperature=1)
# + id="usZ1Bre1qLVS" colab_type="code" outputId="4aa35416-0d92-458c-cc5d-32071cdf3ac2" executionInfo={"status": "ok", "timestamp": 1557126334277, "user_tz": -120, "elapsed": 573, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 264}
myabssk = strokes_to_lines(d)
plt.axis('equal')
plt.axis('off')
for s in myabssk:
xs = [v[0] for v in s]
ys = [-v[1] for v in s]
plt.plot(xs,ys,'#0088ff')
# + [markdown] id="gaZm8_xOqcnl" colab_type="text"
# # Random Samples
# + id="M73oNU28qd_B" colab_type="code" colab={}
# randomly unconditionally generate 10 examples
N = 10
reconstructions = []
for i in range(N):
reconstructions.append([decode(temperature=0.5), [0, i]])
# + id="RCwjbn6RqiIh" colab_type="code" outputId="a6432371-69d0-451b-c4a3-8c291f742979" executionInfo={"status": "ok", "timestamp": 1557126411504, "user_tz": -120, "elapsed": 1301, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 160}
plt.figure(figsize=(15,2))
for i in range(10):
plt.subplot(1,10,i+1)
myabssk = strokes_to_lines(reconstructions[i][0])
plt.axis('equal')
plt.axis('off')
for s in myabssk:
xs = [v[0] for v in s]
ys = [-v[1] for v in s]
plt.plot(xs,ys,'#0088ff')
# + [markdown] id="NXY9CO1EqrdB" colab_type="text"
# # Blend
# + id="VdiWhrT0qsx5" colab_type="code" colab={}
z_0 = np.random.randn(eval_model.hps.z_size)
z_1 = np.random.randn(eval_model.hps.z_size)
# + id="9Ls4ZnnYqw1p" colab_type="code" colab={}
z_list = [] # interpolate spherically between z_0 and z_1
N = 10
for t in np.linspace(0, 1, N):
z_list.append(slerp(z_0, z_1, t))
# for every latent vector in z_list, sample a vector image
reconstructions = []
for i in range(N):
reconstructions.append([decode(z_list[i], temperature=0.1), [0, i]])
# + id="DRXARbdm9_gt" colab_type="code" outputId="1ddc5f71-82d6-453e-e3ff-6ddc20def601" executionInfo={"status": "ok", "timestamp": 1557126449387, "user_tz": -120, "elapsed": 1333, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-i_bcow4kfsM/AAAAAAAAAAI/AAAAAAAAAA0/BKqcIMjm5-8/s64/photo.jpg", "userId": "09730771974529627842"}} colab={"base_uri": "https://localhost:8080/", "height": 160}
plt.figure(figsize=(15,2))
for i in range(10):
plt.subplot(1,10,i+1)
myabssk = strokes_to_lines(reconstructions[i][0])
plt.axis('equal')
plt.axis('off')
for s in myabssk:
xs = [v[0] for v in s]
ys = [-v[1] for v in s]
plt.plot(xs,ys,'#0088ff')
# + [markdown] id="bEExfiHVr5Xy" colab_type="text"
# # Exercise
# + [markdown] id="pE3ZJ54g3hOG" colab_type="text"
# ### Schrödinger’s Castle
#
# Explore the VAE model trained on houses and castles and find “sweet spots” in latent space. Generate new house/castle hybrids, either from random or synthetic input or with an existing sketch from the train-, test- or validation-set as a starting point. Use e.g. the dimensionality reduction / clustering presented in the last tutorial to find denser areas or generate stepwise blends. Upload the vector drawings (PDF) together with the notebook containing some comments what the idea was and how you approached it.
# + id="PzrjAWMS3nGV" colab_type="code" colab={}
| Exercises/Exercise04b_SchroedingersCastle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Segmenting and Clustering Neighborhoods in Toronto
# ## Section One
# Import required libraries
import pandas as pd
import requests
from IPython.display import display, HTML
# Fetch "List of postal codeds of Canada: M" then parse it into Pandas DataFrame
url = 'https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M'
r = requests.get(url)
wiki_table = pd.read_html(r.text, flavor='html5lib')
df = wiki_table[0]
df.columns = ['PostalCode', 'Borough', 'Neighborhood']
df
# Drop unassigned Borough
df.drop(df[df['Borough'] == 'Not assigned'].index, inplace=True)
df.reset_index(drop=True, inplace=True)
df
# Sort Postcode, Borough, and Neighbourhood then group by Postcode and Borough then aggregate the Neighbourhood columns by joining them into a string separated by "comma". Then check for "Not assigned" neighbourhood.
df.sort_values(['PostalCode', 'Borough', 'Neighborhood'], inplace=True)
df_grouped = df.groupby(['PostalCode', 'Borough'])['Neighborhood'].apply(', '.join).reset_index()
df_grouped[df_grouped['Neighborhood'] == 'Not assigned']
# Final DataFrame
df_grouped
df_grouped.shape
# ## Section Two
# Import required libraries
# +
# # !conda install -c conda-forge geopy --yes
# -
from geopy.geocoders import Nominatim
geolocator = Nominatim(user_agent="Toronto Geolocator")
df_location = df_grouped.copy()
# Because the geopy is unreliable I won't add new column manually
# df_location['Latitude'] = ''
# df_location['Longitude'] = ''
df_location
# __Note__: Unreliability proof; I limit the trial to about 10 times per postal code because each trial takes considerable time if you take into the account the time needed to get all the data for every postal code
lat_lon = []
for idx, row in df_location.iterrows():
print(idx)
try:
postcode = df_location.at[idx, 'PostalCode']
geo = None
for i in range(10):
geo = geolocator.geocode(f'{postcode}, Toronto, Ontario')
if geo: break
print(idx, postcode, geo)
# Save
if geo:
lat_lon.append(idx, geo.latitude, geo.longitude)
except:
continue
# As it said in the assignment page, the package is very unreliable. Fallback using provided data.
# +
# # !wget -q -O geo_data.csv https://cocl.us/Geospatial_data
# -
# Parse the geo data
df_geo = pd.read_csv('geo_data.csv')
df_geo.columns = ['PostalCode', 'Latitude', 'Longitude']
df_geo
df_toronto = df_location.merge(df_geo, left_on='PostalCode', right_on='PostalCode')
df_toronto
# ## Section Three
# Set Foursquare variables
# +
CLIENT_ID = 'EM0NULKILDUZUGSXYVR1TWWDQHMCB3CPMMB3CS0EWOSBDKML' # your Foursquare ID
CLIENT_SECRET = '<KEY>' # your Foursquare Secret
VERSION = '20180605' # Foursquare API version
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET: ' + CLIENT_SECRET)
# -
df_toronto['Borough'].value_counts()
def getNearbyVenues(names, latitudes, longitudes, radius=500, LIMIT=200):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
# Get 200 venues for each neighborhood.
toronto_venues = getNearbyVenues(names=df_toronto['Neighborhood'],
latitudes=df_toronto['Latitude'],
longitudes=df_toronto['Longitude'])
# Save to CSV
toronto_venues.to_csv('toronto_venues.csv')
toronto_venues.groupby('Neighborhood').count()
len(toronto_venues['Venue Category'].unique())
# In my case, `Venue Category` named `Neighborhood` must be get rid in order to avoid some error when transforming the DataFrame into one-hot form.
toronto_venues[toronto_venues['Venue Category'].str.contains('Nei')]
toronto_venues.drop(toronto_venues[toronto_venues['Venue Category'].str.contains('Nei')].index, inplace=True)
toronto_venues[toronto_venues['Venue Category'].str.contains('Nei')]
toronto_venues['Venue Category'].value_counts()[0:20]
# Transform to one-hot form to make it easier to cluster then.
toronto_onehot = pd.get_dummies(toronto_venues[['Venue Category']], prefix="", prefix_sep="")
list_columns = list(filter(lambda x: x != 'Neighborhood', list(toronto_onehot.columns)))
toronto_onehot['Neighborhood'] = toronto_venues['Neighborhood']
new_columns = ['Neighborhood'] + list_columns
toronto_onehot = toronto_onehot[new_columns]
toronto_onehot
# Grouping same neighborhood name, since initially it based on postal code and each neighborhood may have several postal code if it has big area.
toronto_grouped = toronto_onehot.groupby('Neighborhood').mean().reset_index()
toronto_grouped
# +
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
top_venues = 10
columns = ['1st', '2nd', '3rd', '4th', '5th', '6th', '7th', '8th', '9th', '10th']
columns = [i + ' most common' for i in columns]
columns = ['Neighborhood'] + columns
columns
toronto_venues_sorted = pd.DataFrame(columns=columns)
toronto_venues_sorted['Neighborhood'] = toronto_grouped['Neighborhood']
for idx, row in toronto_grouped.iterrows():
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
toronto_venues_sorted.loc[idx, 1:] = row_categories_sorted.index.values[:10]
toronto_venues_sorted
# -
from sklearn.cluster import KMeans
toronto_cluster = toronto_grouped.drop('Neighborhood', axis=1)
cluster_size = 5
kmeans = KMeans(n_clusters=cluster_size, random_state=42).fit(toronto_cluster)
kmeans.labels_[:10]
toronto_data1 = df_toronto[['Neighborhood', 'Latitude', 'Longitude']].groupby('Neighborhood').mean()
toronto_data1
toronto_data2 = toronto_venues_sorted
toronto_data2
toronto_final_data = toronto_data1.merge(toronto_data2, left_on='Neighborhood', right_on='Neighborhood')
toronto_final_data['Cluster'] = kmeans.labels_
toronto_final_data
# +
# # !conda install -c conda-forge folium --yes
# +
import folium
import numpy as np
import matplotlib.cm as cm
import matplotlib.colors as colors
latitude = 43.722365
longitude = -79.412422
# create map
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
# set color scheme for the clusters
x = np.arange(cluster_size)
ys = [i + x + (i*x)**2 for i in range(cluster_size)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
for idx, row in toronto_final_data.iterrows():
poi = row[0]
lat = row[1]
lon = row[2]
most_common = row[3]
cluster = row[-1]
label = folium.Popup(f'{poi} cluster {cluster} most common {most_common}', parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7
).add_to(map_clusters)
map_clusters
# -
map_clusters.save('toronto_cluster_map.html')
# In case the map is not showed, it can be seen in the [toronto_cluster_map.html](https://gpratama.github.io/toronto_cluster_map.html)
# Based on the cluster showed in rendered map it seems that the most dominant cluster, cluster 1, is centered at the city center and not so dense when it far from the city center. There also another dominant cluster, cluster 3, that seems to have no identifiable cluster center. The other cluster seems to not dominant compared to the first two. It can be said that there are two interesting cluster, cluster 1 and cluster 3.
toronto_final_data[toronto_final_data['Cluster'] == 0]
toronto_final_data[toronto_final_data['Cluster'] == 1]
# Cluster 1 seems to have most various kind of common venues apparently.
toronto_final_data[toronto_final_data['Cluster'] == 1]['1st most common'].value_counts()
# But when we see the count of most common venues it shows that it dominated by Coffee Shop
toronto_final_data[toronto_final_data['Cluster'] == 2]
toronto_final_data[toronto_final_data['Cluster'] == 3]
# Cluster 3 showed that most common venue there is Park
toronto_final_data[toronto_final_data['Cluster'] == 4]
# ### Cluster Summary
# | Cluster | Size | Most common |
# |---------|------|----------------------|
# | 0 | 1 | Playground |
# | 1 | 80 | Coffee Shop |
# | 2 | 1 | Fast Food Restaurant |
# | 3 | 11 | Park |
# | 4 | 1 | Food Service |
| Segmenting and Clustering Neighborhoods in Toronto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # L'Analyse Sémantique Latente
# Dans ce projet, nous allons voir comment « extraire » des concepts à partir d'un corpus (collection) de documents textuels.
#
# Je vais vous montrer comment extraire mathématiquement des "concepts" de ce corpus. La technique que nous allons utiliser s'appelle "l'analyse sémantique latente".
# -
from bs4 import BeautifulSoup
import nltk
from nltk.corpus import stopwords
from sklearn.decomposition import TruncatedSVD
from sklearn.feature_extraction.text import TfidfVectorizer
import requests as req
# J'ai scrapé un article sur le **Deep Learning** à travers le site **CNRS Le Journal**<br>
# [https://lejournal.cnrs.fr/billets/letonnante-acceptabilite-des-deep-fake](https://lejournal.cnrs.fr/billets/letonnante-acceptabilite-des-deep-fake)
# url_article = 'https://www.ledevoir.com/culture/medias/601874/medias-l-esprit-de-i-science-i-amp-i-vie-i-reinvente-dans-le-nouveau-magazine-i-epsiloon-i'
url_article = 'https://lejournal.cnrs.fr/billets/letonnante-acceptabilite-des-deep-fake'
reponse = req.get(url_article)
article = ''
if reponse.ok:
soup = BeautifulSoup(reponse.text)
articleText = soup.findAll('p')
# divArt = soup.findAll('div')
articleDocs = [paragraphe.text for paragraphe in articleText if len(paragraphe)>4]
article = [paragraphe.lower() for paragraphe in articleDocs]
# for i in divArt:
# if len(i)>5:
# article.append(i.text.lower())
else:
print('Pas de réponse')
import htmlentities as decodeEnt
def supprimer_nomber(texte):
import re
for i in re.findall('\d{1,}', texte):
texte = texte.replace(i, '')
return texte
def preproces(docs, entitie, separateur):
docs = [supprimer_nomber(art) for art in docs ]
return [art.replace(entitie, separateur) for art in docs ]
article = preproces(article, '\xa0', ' ')
article = preproces(article, '\t', ' ')
article = preproces(article, '\n', ' ')
# Les mots vides sont des mots que je ne veux pas convertir en fonctionnalités, car ils ne sont pas particulièrement utiles. Des mots comme **a**, **and** et **the** sont de bons mots vides en français. Je peux utiliser une liste intégrée de mots vides de nltk pour commencer. Ensuite, j'ajouterai des mots vides personnalisés qui sont **html indésirable** que je dois nettoyer de mes données
mot_inutils = set(stopwords.words('french'))
len(mot_inutils)
# ## TF-IDF Vectorizing
# Je vais utiliser le vectoriseur TF-IDF de scikit-learn pour prendre mon corpus et convertir chaque document en une matrice creuse de fonctionnalités TFIDF...
article[0]
vectorizer = TfidfVectorizer(
stop_words=mot_inutils, use_idf=True,
ngram_range=(1, 5)
)
X = vectorizer.fit_transform(article)
X[0]
# +
# print(X[1])
# -
# ## LSA
# * **Entrée : X**, une matrice où m est le nombre de documents que j'ai, et n est le nombre de termes
#
# * **Processus :** Je vais décomposer X en matrices appelées U, S et T. Lorsque nous effectuons la décomposition, nous devons choisir une valeur k, c'est le nombre de concepts que nous allons conserver.
# <div style="width:150px; margin: 10px auto;">X = ${USV}^T$</div>
# * **U :** sera une matrice mxk. Les lignes seront des documents et les colonnes seront des **concepts**
# * **S :** sera une matrice kxk. Les éléments seront la quantité de variation capturée à partir de chaque **concept**
# * **V :** sera une matrice mxk (attention à la transposition). la ligne sera les termes et les colonnes seront **concepts**
#
X.shape[0], X.shape[1]
lsa = TruncatedSVD(n_components=X.shape[0], n_iter=100)
model = lsa.fit(X)
V = model.components_[0]
V
# +
termes = vectorizer.get_feature_names()
for i, comp in enumerate(model.components_):
termeEtConc = zip(termes, comp)
sortedTermes = sorted(termeEtConc, key=lambda x: x[1], reverse=True)[:3]
print(f'Concept {i}')
for terme in sortedTermes:
print(f' - {terme[0]}')
print(' ')
# -
"""
Technologie,
les algorithmes contre images,
les application des transformations vocales,
analyses scientifiques
"""
t = "145 852 les application des transformations vocale"
supprimer_nomber(t)
| python/analyse_semantique.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Divyakathirvel26/Covid-19/blob/master/Covid_19_VGG_16_Model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="G9MPx0cacaE-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="77617985-8595-4aaf-adc4-bae7022611f6"
from google.colab import drive
drive.mount('/content/drive')
# + id="umKOqYwihxqX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f9a93596-d3b6-49db-f209-85502d84d7b5"
import pandas as pd
import numpy as np
import os
import tensorflow as tf
import keras
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D
from tensorflow.keras.applications.vgg16 import VGG16
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.vgg16 import preprocess_input
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
# + id="GON8PSTIiAwO" colab_type="code" colab={}
image_size = [224,224]
data_path = 'Data'
# + id="xFAb5Wm7iECH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="cf8e4649-c509-4bea-b99d-2ed3d40dbd97"
vgg = VGG16(input_shape= image_size+[3],weights='imagenet',include_top=False)
# + id="srm0dK_7iH5m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf554983-51d3-4626-c345-e188dee992e3"
vgg.output
# + id="NBfouL0jiPV-" colab_type="code" colab={}
x = vgg.output
x = GlobalAveragePooling2D()(x)
# + id="dLsAZl5PiSyy" colab_type="code" colab={}
x = Dense(1024, activation='relu')(x)
x = Dense(1024, activation='relu')(x)
x = Dense(512, activation='relu')(x)
# + id="3HU9SI9WiVPn" colab_type="code" colab={}
preds = Dense(2,activation='softmax')(x)
# + id="RmyVK6HliYzE" colab_type="code" colab={}
model = Model(inputs = vgg.input,outputs=preds)
# + id="WloYfVEkicfN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 969} outputId="220187d1-9386-4241-ebe6-8a216f1dd95c"
model.summary()
# + id="-BBmNBllie4a" colab_type="code" colab={}
for layer in vgg.layers:
layer.trainable = False
# + id="kw6sPslyilbP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="afdf8997-68e6-45e8-aa23-1f0d9c7f42c9"
train_datagen=ImageDataGenerator(preprocessing_function=preprocess_input)
train_generator=train_datagen.flow_from_directory('drive/My Drive/covid-data/' ,
target_size=(224,224),
color_mode='rgb' ,
batch_size=32,
class_mode='categorical' ,
shuffle = True)
# + id="TV_qjN6Vip7H" colab_type="code" colab={}
model.compile(optimizer='Adam',
loss='categorical_crossentropy' ,
metrics=['accuracy'])
# + id="1X_CFpzFjONi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="7fd8d2f4-c5bf-472b-ec06-1e9e10a73e20"
step_size_train=train_generator.n//train_generator.batch_size
r = model.fit_generator(generator=train_generator,
steps_per_epoch=step_size_train,
epochs=5)
# + id="kTcyL3RkkAjw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="0eafdca0-8761-42c8-f59d-1092c6a2bb02"
plt.plot(r.history['loss'], label='train loss')
plt.legend()
plt.show()
plt.savefig('LossVal_loss')
# + id="l00T0gJ6kItS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="b2e62085-525f-4c14-d17c-0d0c8dcd6158"
plt.plot(r.history['accuracy'])
plt.title('model accuracy')
plt.ylabel('acc')
plt.xlabel('epoch')
plt.show()
# + id="m-LjL3e3kPNr" colab_type="code" colab={}
from tensorflow.keras.models import load_model
# + id="ZpfFI1s5kqXa" colab_type="code" colab={}
model.save('covid.h5')
# + id="bNfIo2KDkuJd" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.preprocessing import image
import numpy as np
from tensorflow.keras.applications.vgg16 import preprocess_input
from tensorflow.keras.models import load_model
# + id="Wiz8yKVYkwuw" colab_type="code" colab={}
model = load_model('covid.h5')
# + id="ajf8DeiUk2Nz" colab_type="code" colab={}
img_path = 'drive/My Drive/covid-data/Test/Normal/2.jpeg'
# + id="NKyy-7I1lIZs" colab_type="code" colab={}
img = image.load_img(img_path,target_size=(224,224))
x= image.img_to_array(img)
x = np.expand_dims(x,axis=0)
img_data = preprocess_input(x)
# + id="ZZzCAF6MlL7b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="29849d8a-5321-4e81-8a2b-0ed6e6a5293b"
rslt = model.predict(img_data)
print(rslt)
# + id="azFxzXiylOgj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7b0d2037-e62f-4c12-8d31-0cbb967460e8"
if rslt[0][0] == 1:
prediction = 'Not a covid patient'
else:
prediction = 'Covid patient'
print(prediction)
# + id="y_eOcH-5lXL2" colab_type="code" colab={}
| Covid_19_VGG_16_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="XLXOhisptG1C" colab_type="text"
# # 第9回: RNN: Exercise1
# ## Echo State Networksで神経活動を学習する
# ### 概要
# 本演習ではpythonを用いてEcho State Networksを実装します。
#
# 学習データセットとして、運動イメージの脳波データを用い、与えられた脳波から2つの運動イメージをデコードします。
#
# まず、講師が概要を説明しますので、全体を掴んだところで演習に取り組んでください。
#
# ### 目標
# - numpyでESNを実装する
# - 神経活動を記録したデータ・セットを学習する
# - EEGから運動イメージを予測する
# + id="lL6MB4A9tQYZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="266cae87-a80e-4d02-ba5a-372edd502493" executionInfo={"status": "ok", "timestamp": 1520607323076, "user_tz": -540, "elapsed": 3240, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
# Install Requrements
# !pip install mne
# + id="X73DxGLQtBgm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Import required modules
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import mne
import pickle
from mne.decoding import CSP
# + [markdown] id="ALEmXKe3DvAU" colab_type="text"
# 使用するEEGデータセットをロードします.(稀にタイムアウトしますが,その場合は再度試してみてください)
# + id="XdZUhZ1stOSf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 656} outputId="0f2f5b6c-b19e-4ad1-964c-ea97f26bd2cc" executionInfo={"status": "ok", "timestamp": 1520607446392, "user_tz": -540, "elapsed": 1372, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
# Load EEG data
tmin, tmax = -1., 4.
event_id = dict(hands=2, feet=3)
subject = 1
# runs = [6, 10, 14] # Sessions to download
runs = [6, 10, 14] # Sessions to download
raw_fnames = mne.datasets.eegbci.load_data(subject, runs)
raw_files = [
mne.io.read_raw_edf(f, preload=True, stim_channel='auto')\
for f in raw_fnames
]
raw = mne.io.concatenate_raws(raw_files)
# strip channel names of "." characters
raw.rename_channels(lambda x: x.strip('.'))
# Apply band-pass filter
raw.filter(7., 30., fir_design='firwin', skip_by_annotation='edge')
# read events
events = mne.find_events(raw, stim_channel='STI 014')
# pick only EEG channels in datasets
# dropping bad channels
picks = mne.pick_types(
raw.info, meg=False, eeg=True, stim=False, eog=False, exclude='bads'
)
epochs = mne.Epochs(
raw, events, event_id, tmin, tmax,
proj=True, picks=picks, baseline=None, preload=True
)
# epochs_data = epochs.get_data()
# epochs_train = epochs.copy().crop(tmin=1., tmax=2.)
epochs_data_raw = epochs.get_data()
epochs_data = epochs.copy().crop(tmin=1., tmax=2.).get_data()
evoked = epochs.average()
# + [markdown] id="Rd8sHIiX6YPD" colab_type="text"
# 学習データセットを分割します.
# + id="llchxKJzxPav" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="db106399-f342-41b5-ceba-19891712e450" executionInfo={"status": "ok", "timestamp": 1520607448084, "user_tz": -540, "elapsed": 793, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//<KEY>", "userId": "104456185439409157314"}}
n_train = 40
data_train = epochs_data[:n_train]
data_test = epochs_data[n_train:]
labels = epochs.events[:, -1] - 2
train_labels = labels[:n_train]
test_labels = labels[n_train:]
print('Shape of the dataset. train:{}, test:{}'.format(data_train.shape, data_test.shape))
# + [markdown] id="V5gNGn6zDg3y" colab_type="text"
# EEGをそのまま処理することは難しいので,CSP (Common Spatial Pattern)を使って特徴抽出を行います.
# ※ CSPについては,[references](https://drive.google.com/file/d/10tTdNvn3qUq7aFMCjEdhtLLJ7XzYeGKD/view?usp=sharing)を参照.
# + id="ZMrc-ktPCYp6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}], "base_uri": "https://localhost:8080/", "height": 328} outputId="5562b898-fb62-46cb-f813-e58056cf2953" executionInfo={"status": "ok", "timestamp": 1520607450576, "user_tz": -540, "elapsed": 1521, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
# creates CSP feature extractor
csp = CSP(n_components=4, reg=None, log=True, norm_trace=False)
# plot CSP patterns estimated on full data for visualization
csp.fit_transform(epochs_data_raw, labels)
layout = mne.channels.read_layout('EEG1005')
csp.plot_patterns(epochs.info, layout=layout, ch_type='eeg',
units='Patterns (AU)', size=1.5)
# + id="FQNjlprckDgi" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
sfreq = raw.info['sfreq']
w_length = 20 # running classifier: window length
w_step = 10 # running classifier: window step size
w_start = np.arange(0, epochs_data.shape[2] - w_length, w_step)
# + id="qcWWee0dkP5S" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="c9fe5500-dae8-4227-cb6c-c3800d78c38a" executionInfo={"status": "ok", "timestamp": 1520607452592, "user_tz": -540, "elapsed": 603, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print(w_length, w_step, w_start)
# + id="M8X7IpJWjqqK" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="48619802-6795-4786-facd-06f14ed2d7cf" executionInfo={"status": "ok", "timestamp": 1520607453580, "user_tz": -540, "elapsed": 630, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
def feature_extractor(x_in):
X = []
for n in w_start:
# datas, channels, lengths
data = x_in[:, :, n:(n + w_length)]
X.append(csp.transform(data))
X = np.array(X).transpose((1, 2, 0))
return X
X_train = feature_extractor(data_train)
X_test = feature_extractor(data_test)
print('(datas, channels, length)', X_train.shape, X_test.shape)
# + id="HPnY0uyBj2Xy" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="2024d07d-e1d6-4dcd-8926-167ee4b6f81d" executionInfo={"status": "ok", "timestamp": 1520607454444, "user_tz": -540, "elapsed": 582, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
train_data_0 = []
train_data_1 = []
for i, c in enumerate(train_labels):
if c == 0:
train_data_0 += [X_train[i]]
else:
train_data_1 += [X_train[i]]
train_data_0 = np.array(train_data_0)
train_data_1 = np.array(train_data_1)
print(train_data_0.shape, train_data_1.shape)
# + id="xwSp-T1h_lME" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="8571a9e2-e1eb-4f45-fdc6-9e0e8c4cbb99" executionInfo={"status": "ok", "timestamp": 1520607455229, "user_tz": -540, "elapsed": 518, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
test_data_0 = []
test_data_1 = []
for i, c in enumerate(test_labels):
if c == 0:
test_data_0 += [X_test[i]]
else:
test_data_1 += [X_test[i]]
test_data_0 = np.array(test_data_0)
test_data_1 = np.array(test_data_1)
print(test_data_0.shape, test_data_1.shape)
# + [markdown] id="ZVlV0MaTyxem" colab_type="text"
# ## ESNの実装
# では,実際にESNの実装をしてみましょう.
# ### ハイパーパラメータの設定
# 以下ではハイパーパラメータの設定を行います。
# - 入力: `Nu`
# - 出力: `Ny`
# - 隠れ素子: `Nx`
# - leaking rate: `a`
# + id="NDF03_Gax7iW" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="4a5c0303-ae7f-4f0c-b60e-7d61eedd254c" executionInfo={"status": "ok", "timestamp": 1520607455978, "user_tz": -540, "elapsed": 524, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
# set hyper params
train_data = train_data_0
init_len = 5
Nu = train_data.shape[1] # Nu
Ny = train_data.shape[1] # Ny
Nx = 500 # Nx
Nt = train_data.shape[2] - init_len
Nb = train_data.shape[0]
a = 0.3 # leaking rate
print('Nu:{}, Nx:{}, Ny:{}, Nt:{}, Nb:{}'.format(Nu, Nx, Ny, Nt, Nb))
# + [markdown] id="EMVM00qwzQ5g" colab_type="text"
# ### Variableの設定
# ESNで使用するパラメータは,入力重み`Win`,reservoir重み`W`,出力重み`Wout`の3つです.
#
# この内学習するのは出力重みだけでしたね.
#
# 次元に気をつけて,パラメータの設定を行います。
# + id="YnAZfQqax9Wp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
np.random.seed(46)
# input weights
# Win: (Nx, 1 + Nu) bias included
Win = (np.random.rand("*** YOUR CODE HERE ***") - 0.5) * 1
# reservoir weights
# W: (Nx, Nx)
W = np.random.rand("*** YOUR CODE HERE ***") - 0.5
# rhoW = max(abs(np.linalg.eig(W)[0]))
# print(rhoW)
# rhoW = 13.1516
# W *= 18.25 / rhoW
W *= 0.8 # Magic!
# output weights (a placeholder)
# Wout: (Ny, 1 + Nx + Nu)
Wout = np.zeros("*** YOUR CODE HERE ***")
# allocated memory for the design (collected states) matrix
# X: (1 + Nu + Nx, Nb, Nt)
X = np.zeros("*** YOUR CODE HERE ***")
# set the corresponding target matrix directly
Yt = train_data.T[init_len:, :, :n_train]
x = np.zeros("*** YOUR CODE HERE ***")
ones = np.ones(Nb).reshape((-1, 1)).T
# + [markdown] id="jV_UGc2Azt_M" colab_type="text"
# 各パラメータの`shape`は以下のようになります
# + id="D9CTF160ztRe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="3d05244d-53ec-450f-b24c-6dfea72907a9" executionInfo={"status": "ok", "timestamp": 1520607457709, "user_tz": -540, "elapsed": 601, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print('Win:{}, W:{}, X:{}, Yt:{}, x:{}'.format(
Win.shape, W.shape, X.shape, Yt.shape, x.shape))
# + [markdown] id="fhq2PRiD11fp" colab_type="text"
# データを表現する際にはたとえ一次元であっても、列ベクトルで表記するべきです。
#
# すなわち、ベクトルのshapeは(25,)ではなく(25, 1)として計算することを心がけてください。
# + [markdown] id="1YSIStMkz67L" colab_type="text"
# ### Forwarding
# データをネットワークにfeedします.この時,ネットワークのreservoir層を活動を安定化させるため,初期化を行います.
# + id="Cbst8hnQ0ckC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="f7215e9d-21ad-4d2f-c460-90201a73a8ed" executionInfo={"status": "ok", "timestamp": 1520607463719, "user_tz": -540, "elapsed": 628, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
for t, un in enumerate(train_data.T):
# (Nx, Nb) = (Nx, Nu + 1)(Nu + 1, Nb) + (Nx, Nx)(Nx, Nb)
x_tld = "*** YOUR CODE HERE ***"
x = "*** YOUR CODE HERE ***"
if t >= init_len:
X[t - init_len, :, :] = np.vstack("*** YOUR CODE HERE ***")
print(X.shape)
# + [markdown] id="FaoUYI2e1WYI" colab_type="text"
# #### Hint
# ・行列同士の積は`np.dot()`を使用しましょう
# ```python
# # EXAMPLE
# >>> a = [[1, 0], [0, 1]]
# >>> b = [[4, 1], [2, 2]]
# >>> np.dot(a, b)
# array([[4, 1],
# [2, 2]])
# ```
#
# ・reservoir層の活性化関数には`np.tanh()`を使用しましょう
#
#
# ・行列を垂直方向に重ねるには`numpy.vstack()`が便利です
# ```python
# # EXAMPLE
# >>> a = np.array([1, 2, 3])
# >>> b = np.array([2, 3, 4])
# >>> np.vstack((a,b))
# array([[1, 2, 3],
# [2, 3, 4]])
#
# >>> np.vstack((0, [[1], [2], [3]]))
# array([[0],
# [1],
# [2],
# [3]])
# ```
# + id="pYKHkyCi-8yG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="027a76d2-8020-4412-a60a-113f1aae61db" executionInfo={"status": "ok", "timestamp": 1520607466466, "user_tz": -540, "elapsed": 598, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print(Yt.shape, X.shape)
# + id="AzThjW4c9xUE" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="87aa1802-11fd-42dd-8539-516bbd685ad4" executionInfo={"status": "ok", "timestamp": 1520607467778, "user_tz": -540, "elapsed": 640, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
X_mat = X.transpose((1, 0, 2)).reshape((X.shape[1], -1))
Yt_mat = Yt.transpose((1, 0, 2)).reshape((Yt.shape[1], -1))
print(X_mat.shape, Yt_mat.shape)
# + [markdown] id="dHUOWDYIFs99" colab_type="text"
# `np.reshape`関数の挙動には注意しましょう.変換後の形が一緒だからといって,内容が一致しているとは限りません.(see [docs](https://docs.scipy.org/doc/numpy/reference/generated/numpy.reshape.html))
# + id="v8i5xKbFDUX6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
_X_mat = X.reshape((X.shape[1], -1))
# + id="QiHuddpwD5di" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 52} outputId="4d7f864e-5383-4d04-a1f2-4d43686964ee" executionInfo={"status": "ok", "timestamp": 1520607473816, "user_tz": -540, "elapsed": 674, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print('{}\n{}'.format(list(_X_mat[5, :3]), list(X_mat[5, :3])))
# + [markdown] id="-MGjrau60j7W" colab_type="text"
# ### Training
# `X_mean`を用いて学習を行います.出力重み行列`Wout`を求めるには,擬似逆行列を用いるか,最小二乗法を用いる方法があります.
#
# 疑似逆行列の計算は`np.liniag.pinv()`を使いましょう.
# + id="HmVHkuTDyDHp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# using pseudo inverse matrix
# Wout: (Ny, 1 + Nu + Nx)
# Wout = np.dot(Yt_mat, np.linalg.pinv(X_mat))
# + [markdown] id="dL_2SE6Z2dN-" colab_type="text"
# 以下は,リッジ回帰の実装です.(両方試してみましょう)
# + id="wlBfJh-byEj0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# For regression
reg = 1e-8 # regularization coefficient
X_T = X_mat.T
Wout = np.dot(
np.dot(Yt_mat, X_mat.T),
np.linalg.inv(np.dot(X_mat, X_T) + reg * np.eye(1 + Nu + Nx))
)
# + [markdown] id="XY2OlkE-1G63" colab_type="text"
# ### Evaluation
# 以下では評価計算を行います。
# 実装はForwardingの部分と殆ど同じです。
# + id="X6v6-s1ayFnT" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# clear resv states
test_data = test_data_0
test_batch_size = test_data.shape[0]
x = np.zeros((Nx, 1))
X = np.zeros((Nt, 1 + Nu + Nx, test_batch_size))
Y = np.zeros((Nt, Ny, test_batch_size))
Yt = test_data.T[init_len: , :n_train, :]
# set the corresponding target matrix directly
ones = np.ones(test_batch_size).reshape((-1, 1)).T
# + id="xEEGpjqyyHLY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
generate = False
for t, un in enumerate(test_data.T):
# (Nx, Nb) = (Nx, Nu + 1)(Nu + 1, Nb) + (Nx, Nx)(Nx, Nb)
if t >= init_len + 1 and generate:
un = y_hat
x_tld = "*** YOUR CODE HERE ***"
x = "*** YOUR CODE HERE ***"
if t >= init_len:
Xk = np.vstack((ones, un, x))
X[t - init_len, :, :] = Xk
y_hat = np.dot(Wout, Xk)
Y[t - init_len, :, :] = y_hat
# + id="eDE4LdkByIdV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 52} outputId="1f2b2229-7f87-4152-dd3e-9df88247310b" executionInfo={"status": "ok", "timestamp": 1520607478703, "user_tz": -540, "elapsed": 842, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print('(Length, Channels, Samples)', Y.shape, Yt.shape)
mse = np.mean(np.sum((Y - Yt)**2, axis=2), axis=1)
print(np.mean(mse))
# + id="biI4weyr-JOq" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 225} outputId="2bda9332-c46f-467c-90ac-8200c74cca6b" executionInfo={"status": "ok", "timestamp": 1520607483678, "user_tz": -540, "elapsed": 4589, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
fig1 = plt.figure(figsize=(10, 3))
x = range(len(Y[:, 0, 0]))
ax = fig1.add_subplot(131)
ax.set_xlabel('time steps')
ax.set_ylabel('state')
ax.plot(x, Yt[:, 1, 0], label='teacher')
ax.plot(x, Y[:, 1, 0], label='predicted')
ax.set_title('Signal Prediction')
x = range(len(X[:, 0, 0]))
bx = fig1.add_subplot(132)
bx.set_xlabel('time steps')
bx.set_ylabel('activated')
rand_idx = np.random.choice(X.shape[1], 50, replace=False)
bx.plot(x, X[:, rand_idx, 1])
bx.set_title('Some reservoir activations $\mathbf{x}(n)$')
x = range(1 + Nu + Nx)
cx = fig1.add_subplot(133)
cx.set_xlabel('index')
cx.set_ylabel('value')
cx.bar(x, Wout[0].T, label='CP0')
cx.bar(x, Wout[1].T, label='CP1')
cx.bar(x, Wout[2].T, label='CP2')
cx.set_title('Output weights $\mathbf{W}^{out}$')
cx.set_ylim((0, 0.5))
plt.tight_layout()
plt.legend()
plt.show()
# + [markdown] id="V48TU7V_F5YP" colab_type="text"
# ## ESN Classの実装
# + [markdown] id="cL_SMg_URXTU" colab_type="text"
# ESNのクラスを実装します.
# + id="xYUk23CeFdlb" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class ESN():
def __init__(self, Nu, Ny, Nx=1000, a=0.3, seed=46):
self.Nu = Nu # size of input
self.Nx = Nx # size of reservoir
self.Ny = Ny # size of output
self.a = a # leaking rate
np.random.seed(seed)
self.Win = None
self.W = None
self.Wout = None
# reservoir placeholer
self.x = np.zeros((Nx, 1))
self._initialize_weights()
def __call__(self, un):
'''
ARGS:
un (array): (CHANNEL SIZE, BATCH SIZE)
RETURNS:
Yhatk (array): (Ny, BATCH SIZE)
Xk (array): (1 + Nu + Nx, BATCH SIZE)
'''
ones = np.ones(un.shape[-1]).reshape((-1, 1)).T
# (Nx, Nb) = (Nx, Nu + 1)(Nu + 1, Nb) + (Nx, Nx)(Nx, Nb)
x_tld = np.tanh(
"*** YOUR CODE HERE ***"
)
self.x = "*** YOUR CODE HERE ***" # update resv
Xk = np.vstack((ones, un, self.x))
Yhatk = np.dot(self.Wout, Xk)
return Yhatk, Xk
def train(self, X_mat, Yt_mat):
print('input shape: X_mat{}, Yt_mat{}\ntraining...'.format(
X_mat.shape, Yt_mat.shape
))
# train with inv or regression
self.Wout = np.dot(Yt_mat, np.linalg.pinv(X_mat))
def _initialize_weights(self):
# input weights (initialized random)
# Win: (Nx, 1 + Nu) bias included
self.Win = (np.random.rand("*** YOUR CODE HERE ***") - 0.5) * 1
# reservoir weights
# W: (Nx, Nx)
self.W = np.random.rand("*** YOUR CODE HERE ***") - 0.5
self.W *= 0.299 # Magic!
# output weights (a place holder)
self.Wout = np.zeros("*** YOUR CODE HERE ***")
def clearresv(self):
# clear reservoir status
self.x = np.zeros((self.Nx, 1))
# + [markdown] id="eOHzesXgRZ2z" colab_type="text"
# インスタンスを生成します.esn0がclass0の予測を,esn1がclass1の予測を担当します.
# + id="OGnfId2ZF8yZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
esn0 = ESN(Nu, Ny)
esn1 = ESN(Nu, Ny)
# + id="bhQM0tXlTg_1" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def compute_X(esn, inpts, init_len=50):
# inpts: (length, channel, batch)
esn.clearresv()
Nb = inpts.shape[-1]
Nt = inpts.shape[0]
X = np.zeros((Nt - init_len, 1 + esn.Nu + esn.Nx, Nb))
for t, un in enumerate(inpts):
_, Xk = esn(un)
if t >= init_len:
X[t - init_len, :, :] = Xk
return X
# + [markdown] id="llg2EtRYRhll" colab_type="text"
# ### Forwarding
# + id="TPbSKB5EWVLg" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="3731bc12-e525-47d0-b8b1-c32fa945f14e" executionInfo={"status": "ok", "timestamp": 1520601953507, "user_tz": -540, "elapsed": 541, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
X0 = compute_X(esn0, train_data_0.T, init_len)
X1 = compute_X(esn1, train_data_1.T, init_len)
X0_mat = X0.reshape((X0.shape[1], -1)) # reshape into (channel, length x batch)
X1_mat = X1.reshape((X1.shape[1], -1))
Yt0 = train_data_0.T[:X0.shape[0]] # Yt should have the same length as X
Yt1 = train_data_1.T[:X1.shape[0]]
Yt0_mat = Yt0.reshape((Yt0.shape[1], -1)) # reshape into (channel, length x batch)
Yt1_mat = Yt1.reshape((Yt1.shape[1], -1))
print(X0.shape ,X0_mat.shape, Yt0_mat.shape)
# + [markdown] id="FOky6dzyRp5p" colab_type="text"
# ### Training
# + id="160ipAy8f6nC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 86} outputId="d65d9a37-2203-49d5-fdde-64cc886252bf" executionInfo={"status": "ok", "timestamp": 1520597168178, "user_tz": -540, "elapsed": 566, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
# train networks
esn0.train(X0_mat, Yt0_mat)
esn1.train(X1_mat, Yt1_mat)
# + [markdown] id="pHP01j9aRtiy" colab_type="text"
# ### Evaluation
# + id="Gk7nEeLchZWN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def compute_Y(esn, inpts, init_len=50, generate=False):
# inpts: (length, channel, batch)
esn.clearresv()
Nb = inpts.shape[-1]
Nt = inpts.shape[0]
Y = np.zeros((Nt - init_len, esn.Ny, Nb))
for t, un in enumerate(inpts):
if t >= init_len + 1 and generate:
un = Yhatk
Yhatk, Xk = esn(un)
if t >= init_len:
Y[t - init_len, :, :] = Yhatk
return Y
# + id="Mzaf5hGBidpP" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
Y0 = compute_Y(esn0, X_test.T, init_len)
Y1 = compute_Y(esn1, X_test.T, init_len)
Yt = X_test.T[init_len:]
# + id="3wbMnIeFjD6G" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 34} outputId="3a10fa01-412c-4a90-ed2b-96bd58d858e3" executionInfo={"status": "ok", "timestamp": 1520597170844, "user_tz": -540, "elapsed": 624, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print(Y0.shape, Yt.shape)
# + id="xuweHXu4kvwo" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
mse0 = np.mean(np.sum((Y0 - Yt)**2, axis=1), axis=0)
mse1 = np.mean(np.sum((Y1 - Yt)**2, axis=1), axis=0)
# + id="kt3Wp2CslMKO" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
results = []
for v0, v1 in zip(mse0, mse1):
results += [0] if v0 > v1 else [1]
# + id="DnAUunJjn_Me" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 52} outputId="96fe66e6-8ccb-48ae-c711-18c380f845f0" executionInfo={"status": "ok", "timestamp": 1520597278162, "user_tz": -540, "elapsed": 505, "user": {"displayName": "\u677e\u68ee\u5320\u54c9", "photoUrl": "//lh5.googleusercontent.com/-H6w5JZ_91_U/AAAAAAAAAAI/AAAAAAAAAAs/_2w4RB2DUHQ/s50-c-k-no/photo.jpg", "userId": "104456185439409157314"}}
print('Predicted: {}\nGround Truth:{}'.format(results, list(test_labels)))
| lecture9/nico2ai_lecture9_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Citrus Leaves Classification Problem Using RMSProp Optimizer
# ## Team <NAME>
# ## Assignment 4-5
#
# **----------------------------------------------------------------------------------------------**
#
# ## Importing Libraries
# +
# Imports
import matplotlib.pyplot as plt
from matplotlib import gridspec
import numpy as np
import tensorflow as tf
from tensorflow.keras.preprocessing import image_dataset_from_directory
from tensorflow.keras.layers.experimental.preprocessing import Rescaling
# -
# ## Training-Testing-Validation Dataset Splitting
#
# Splitting Given Training Data into Training, Validation Set (3:1 |OR| 75:25)
# Selecting Whole Given Validation data as Training Dataset
#
# ie. Data split (Tr:Te:Va - 3:1:1)
# +
from keras.preprocessing.image import ImageDataGenerator as IDG
from sklearn.model_selection import train_test_split
idg_train = IDG(
rescale=1./ 255,
rotation_range=180,
zoom_range=0.3,
width_shift_range=0.3,
height_shift_range=0.3,
horizontal_flip=True,
vertical_flip=True,
validation_split=0.25)
idg_test = IDG(rescale=1./ 255)
ds_train=idg_train.flow_from_directory('../input/citrus-leaves-prepared/citrus_leaves_prepared/train',batch_size=32,shuffle=True,subset='training')
ds_valid=idg_train.flow_from_directory('../input/citrus-leaves-prepared/citrus_leaves_prepared/train',batch_size=8,shuffle=True,subset='validation')
ds_test=idg_test.flow_from_directory('../input/citrus-leaves-prepared/citrus_leaves_prepared/validation',batch_size=1,shuffle=True)
# -
# ## Model defining
# +
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import regularizers as rg
from tensorflow.keras import initializers as it
model = tf.keras.Sequential([
layers.Conv2D(16, (3,3), activation='relu', input_shape=(256, 256, 3),padding='same'),
layers.MaxPooling2D(2, 2),
layers.Conv2D(32, (3,3), activation='relu',padding='same'),
layers.MaxPooling2D(2,2),
layers.Conv2D(64, (3,3), activation='relu',padding='same'),
layers.MaxPooling2D(2,2),
layers.Conv2D(64, (3,3), activation='relu',padding='same'),
layers.MaxPooling2D(2,2),
layers.Conv2D(64, (3,3), activation='relu',padding='same'),
layers.MaxPooling2D(2,2),
layers.Flatten(),
layers.Dense(512, activation='relu'),
layers.Dense(4, activation='sigmoid')
])
model.summary()
# -
# ## Model Training and fitting
# Used RMSprop as optimizer,
# Categorical Cross Entropy as Loss
#
# Hyper Parameters
# * Learning Rate - 0.0004
# +
model.compile(
optimizer=tf.keras.optimizers.RMSprop(learning_rate=4*1e-4),
loss='categorical_crossentropy',
metrics=['accuracy','Precision','Recall']
)
history = model.fit(
ds_train,
validation_data=ds_valid,
epochs=35
)
# -
# ## Plotting the Graphs for Loss, Accuracy, Precision, Recall
# +
import pandas as pd
history_frame = pd.DataFrame(history.history)
history_frame.loc[:, ['loss', 'val_loss']].plot()
history_frame.loc[:, ['accuracy', 'val_accuracy']].plot();
history_frame.loc[:, ['precision', 'val_precision']].plot();
history_frame.loc[:, ['recall', 'val_recall']].plot();
# -
# ## Evaluating the Model using the Training Data
#
# **Output**
#
# * **loss:** 0.4764
# * **accuracy:** 0.8156
# * **precision:** 0.4140
# * **recall:** 0.9944
model.evaluate(ds_train)
| Assignment 4 & 5/CSP502_A4-5_The-Salvator-Brothers/code/cv-a-4-5-citrus-leaves-RMSprop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME> - Part 3: Parallel Parking
# In this section you will write a function that implements the correct sequence of steps required to parallel park a vehicle.
#
# **Note** for this segment the vehicle's maximum speed has been set to just over 4 mph. This should make parking a little easier.
#
# 
# %%HTML
<link rel="stylesheet" type="text/css" href="buttonStyle.css">
<button id="launcher">Launch Car Simulator</button>
<button id="restart">Restart Connection</button>
<script src="setupLauncher.js"></script>
<script src="kernelRestart.js"></script>
# +
# After running any code changes make sure to click the button "Restart Connection" above first.
# Also make sure to click Reset in the simulator to refresh the connection.
car_parameters = {"throttle": 0, "steer": 0, "brake": 0}
def control(pos_x, pos_y, time, velocity):
""" Controls the simulated car"""
global car_parameters
if(pos_y > 37.6):
car_parameters["throttle"] = -0.2
car_parameters["steer"] = 1
car_parameters["brake"] = 0
elif(pos_y > 35):
car_parameters["throttle"] = -0.01
car_parameters["steer"] = 0
car_parameters["brake"] = 0
elif(pos_y > 31.5):
car_parameters["throttle"] = -0.01
car_parameters["steer"] = -1
car_parameters["brake"] = 0
else:
car_parameters["throttle"] = 0
car_parameters["steer"] = 0
car_parameters["brake"] = 1
# TODO: Use WASD keys in simulator to gain an intuitive feel of parallel parking.
# Pay close attention to the time, position, and velocity in the simulator.
# TODO: Use this information to make decisions about how to set your car parameters
# In this example the car will drive forward for three seconds
# and then backs up until its y_pos is less than 32 then comes to a stop by braking
return car_parameters
import src.simulate as sim
sim.run(control)
# -
# # Submitting this Project!
# Your parallel park function is "correct" when:
#
# 1. Your car doesn't hit any other cars.
# 2. Your car stops completely inside of the right lane.
#
# Once you've got it working, it's time to submit. Submit by pressing the `SUBMIT` button at the lower right corner of this page.
| Project 0 - Joy Ride/ParallelParking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Managing Geometry Properties of Imported Networks
# The ``Imported`` geometry class is used to store the geometrical properties of imported networks. When importing an extracted network into OpenPNM using any of the ``io`` classes, all the geometrical and topological properties are lumped together on the *network* object. OpenPNM is generally designed such that geometrical properties are stored on a *geometry* object, so this class address this issue. The main function of the ``Imported`` class is to automatically strip the geometrical properties off of the network and transfer them onto itself.
# > **What problem does the Imported class solve?** Although OpenPNM can function with the geometrical properties on the network, a problem arises if the user wishes to add *more* pores to the network, such as boundary pores. In this case, they will probably wish to add pore-scale models to calculate size information, say 'pore.volume'. If they add this to the network, this model will overwrite the pre-existing 'pore.volume' values. The solution to this problem is an intrinsic part of OpenPNM: create a separate geometry object to manage it's own 'pore.volume' model and values. However, this **won't work**! OpenPNM will not allow an array called 'pore.volume' to exist on the network *and* a geometry object. The reason is that networks store values for *every* pore, so when adding new pores the network the 'pore.volume' array will increase to accommodate them. If you attempt to put 'pore.volume' values on the geometry object, you're are essentially putting *two* values in those locations. Therefore, the ``Imported`` class solves this problem by first transferring the 'pore.volume' array (and all other geometrical properties) from the network to itself.
import numpy as np
import openpnm as op
# %config InlineBackend.figure_formats = ['svg']
import matplotlib.pyplot as plt
ws = op.Workspace()
ws.settings['loglevel'] = 50 # Supress warnings, but see error messages
# Let's start by generating a random network using the Delaunay class. This will repreent an imported network:
np.random.seed(0)
pn = op.network.Delaunay(shape=[1, 1, 0], num_points=100)
# This network generator adds nicely defined boundary pores around the edges/faces of the network. Let's remove these for the sake of this example:
op.topotools.trim(network=pn, pores=pn.pores('boundary'))
# NBVAL_IGNORE_OUTPUT
fig, ax = plt.subplots(figsize=(5, 5))
op.topotools.plot_coordinates(network=pn, c='r', ax=ax)
op.topotools.plot_connections(network=pn, ax=ax)
# This network does not have any geometrical properties on it when generated. To mimic the situation of an imported network, let's manually enter some values for ``'pore.diameter'``. We'll just assign random numbers to illustrate the point:
pn['pore.diameter'] = np.random.rand(pn.Np)
# Now when we ``print`` the network we'll see all the topological data ('pore.coords' and 'throat.conns'), all the labels that were added by the generator (e.g. 'pore.left'), as well as the new geometry info we just added ('pore.diameter'):
print(pn)
# OpenPNM was designed to work by assigning geomtrical information to **Geometry** objects. The presence of 'pore.diameter' on the network can be a problem in some cases. For instance, let's add some boundary pores to the left edge:
Ps = pn['pore.surface']*(pn['pore.coords'][:, 0] < 0.1)
Ps = pn.toindices(Ps)
op.topotools.add_boundary_pores(network=pn, pores=Ps,
move_to=[0, None, None],
apply_label='left')
# Visualizing this networks shows the newl added pores where we intended:
# NBVAL_IGNORE_OUTPUT
fig, ax = plt.subplots(figsize=(7, 7))
op.topotools.plot_coordinates(network=pn, pores=pn.pores('left', mode='not'),
c='r', ax=ax)
op.topotools.plot_coordinates(network=pn, pores=pn.pores('left'), c='g', ax=ax)
op.topotools.plot_connections(network=pn, ax=ax)
# Now we have internal pores (red) and boundary pores (green). We would like to assign geometrical information to the boundary pores that we just created. This is typically done by creating a **Geometry** object, then either assigning numerical values or attaching a pore-scale model that calculates the values. The problem is that OpenPNM prevents you from having 'pore.diameter' on the network AND a geometry object at the same time.
Ps = pn.pores('left')
Ts = pn.find_neighbor_throats(pores=Ps)
geo_bndry = op.geometry.GenericGeometry(network=pn, pores=Ps, throats=Ts)
# Now we we try to assign ``'pore.diameter'``, we'll get the following exception (The "try-except" structure is used for the purpose of this notebook example, but is not needed in an actual script):
try:
geo_bndry['pore.diameter'] = 0
except Exception as e:
print(e)
# The solution is to remove the geometrical information from the network *before* adding the boundary pores, and place them on their own geometry. In this example it is easy to transfer the ``'pore.diameter'`` array, but in the case of a real extracted network there could be quite a few arrays to move. OpenPNM has a facility for doing this: the ``Imported`` geometry class.
# ## Using the Imported Geometry Class
# Let's create a network and add a geometric properties again, this time *before* adding boundary pores.
pn = op.network.Delaunay(shape=[1, 1, 0], num_points=100)
pn['pore.diameter'] = np.random.rand(pn.Np)
# Here we pass the network to the ``Imported`` geometry class. This class literally removes all numerical data from the network to itself. Everything is moved except topological info ('pore.coords' and 'throat.conns') and labels ('pore.left').
geo = op.geometry.Imported(network=pn)
# Printing ``geo`` reveals that the 'pore.diameter' array has been transferred from the network automatically:
print(geo)
# Now that the geometrical information is properly assigned to a geometry object, we can now use OpenPNM as intended. Let's extend this network by adding a single new pore.
op.topotools.extend(network=pn, pore_coords = [[1.2, 1.2, 0]], labels='new')
# The new pore can clearly be seen outside the top-right corner of the domain.
# NBVAL_IGNORE_OUTPUT
fig, ax = plt.subplots(figsize=(7, 7))
op.topotools.plot_coordinates(network=pn, pores=pn.pores('left', mode='not'),
c='r', ax=ax)
op.topotools.plot_coordinates(network=pn, pores=pn.pores('left'), c='g', ax=ax)
op.topotools.plot_connections(network=pn, ax=ax)
# We can now create a geometry just for this single pore and we will be free to add any properties we wish:
geo2 = op.geometry.GenericGeometry(network=pn, pores=pn.pores('new'))
geo2['pore.diameter'] = 2.0
print(geo2)
# Note that the network has the ability to fetch the 'pore.diameter' array from the geometry sub-domain object and create a single full array containing the values from all the locations. In the printout below we can see the value of 2.0 in the very last element, which is where new pores are added to the list.
print(pn['pore.diameter'])
| examples/notebooks/networks/extraction/managing_geometrical_properties_of_imported_networks.ipynb |
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Common Lisp (SBCL)
;; language: common-lisp
;; name: common-lisp_sbcl
;; ---
(#+quicklisp ql:quickload #-quicklisp asdf:load-system :ngl-clj)
(defparameter stage (ngl:make-stage :components (list (ngl:make-structure :value (alexandria:read-file-into-string "DPDP.pdb")
:ext "pdb"
:auto-view-duration 0
:representations (list (ngl:make-licorice))
:trajectories (list (ngl:make-trajectory :interpolate-type "spline"
:value (alexandria:read-file-into-byte-vector "DPDP.nc")
:ext "nc"))))
:layout (jw:make-layout :height "700px")))
(j:display stage)
(ngl:play stage)
(ngl:pause stage)
| examples/trajectory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # NumPy arrays: Creation and handling
#
# *Computer tools for Astronomers*
#
# This first part is an enhanced version of the slide about fundamental array handling in Numpy/SciPy.
#
# Basically, all I have done here is to add some plots that visualize the arrays with colors, because it makes it clearer what happens when we perform the different operations .For thispurpose, I am introducing the function `imshow()` from Matplotlib. We shall look more at plotting later; for now it is enough to know that `imshow()` can translate the value of each array element into a color, which can make it quite a bit easier to see whatis going on in the array than just looking at a bunch of numbers.
#
#
# > **Please note**: I am using the notebook interface to write this, since it is great for making
# > demos of all the feature. However, I recommend that you try the code below in the terminal or QtConsole
# > interface, as these have some features that this doesn't.
# > For example, the plots will show up in an interactive window that allow, panning zooming, etc., unlike this
# > interface that produces static raster images of the plots.
# -
import scipy as sp
import matplotlib.pyplot as plt
from scipy import random as ra # Purely laziness
# Only do this if you are working in the Notebook interface:
# %matplotlib inline
# + [markdown] slideshow={"slide_type": "slide"}
# ### Array construction
#
# The simplest way to define an array is by feeding it a list of numbers.
# -
a = sp.array([1., 2., 3.])
print a
# + [markdown] slideshow={"slide_type": "slide"}
# If you want a 2D array, feed it a list of lists of numbers:
# -
a = sp.array([[1., 2., 3.],
[4., 5., 6.],
[7., 8., 9.]])
print a
# NB! In the above, the line breaks are only there to look good. We could just as well have written:
# ```python
# a = sp.array([[1., 2., 3.], [4., 5., 6.], [7., 8., 9.]])
# ```
# Now I have been reassigning values to the variable name '`a`' a few times. Let's keep it the next time:
a = sp.zeros((8, 8)); print a
# What we wrote here was "create an array of $8 \times 8$ elements and set them all to zero".
#
# There's another convenience function called `ones()`, which can also be called with a tuple of numbers:
b = sp.ones((8, 8))
print b
# Besides, there are also the convenience functions `zeros_like()` and `ones_like()` that take another array as the input and returns an array of zeros resp. ones of the same size as the input array:
b = sp.ones_like(a)
print(b)
# An array can be reshaped to any shape you want, as long as it has the correct number of elements:
b = sp.ones_like(a).reshape((4, 16))
print b
print(b.transpose()) # Or b.T = easier to write but less explicit.
b = b.reshape((8, 8)) # Just to get the good ol' `b` array back.
# There are a number of different random number generating functions, the simplest of which is `scipy.random.random()`.
# Since in the imports section we have already abbreviated `scipy.random` to `ra`, we can write this as:
# ```python
# c = ra.random(b.shape)
# ```
# Remember, you can always get help for a function by writing e.g. `ra.random?` in your IPython console.
# Printing an array of random floating point numbers between zero and one doesn't show much, so we will visualize the array with a plotting function instead. Try running the below commands a few times to convince yourself that it is in fact a pretty well randomized set you get:
# +
# This cell should be ignored or deleted...
# plt.colormaps?
# -
plt.set_cmap('hot')
c = ra.random(b.shape) * .5
plt.matshow(c)
plt.colorbar()
# plt.show() # Remember to write this when calling it from the terminal
# Another convenience function gives you the Identity matrix, usually called $I$. Since `I` is a bad name for a function, they called it something very similar yet not the same:
d = sp.eye(b.shape[0]); print(d)
plt.matshow(d); plt.colorbar()
#b.shape # Arribute
#b.max() # Method
# ### Array vs. matrix
#
# Unlike e.g. Matlab$^®$, NumPy doesn't perform matrix operations by default, but only when you specifically ask about it. If we use the arrays `c` and `d` as examples, the difference is quite clear. Taking the sum of the two is the same in both languages, as the matrix sum is just defined as the element-wise sum like it is explained in wikipedia:
#
# 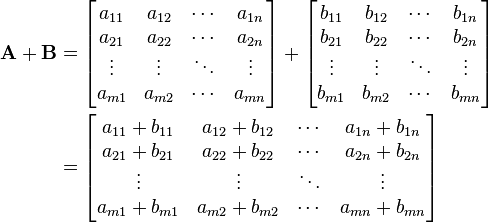
# In our case, the sum of `c` and `d`, hardly surprising, becomes:
plt.matshow(c + d, interpolation='nearest'); plt.colorbar()
# However, in in NumPy our array is ***not treated as a matrix unless we explicitly do this***.
#
# Here's the difference (by the way, compare the diagonals):
# +
array_product = c * d # Element-wise product
matrix_product = sp.dot(d, c) # Matrix product
# And now some plotting magic:
plt.subplot(121)
plt.imshow(array_product, interpolation='nearest')
plt.title('Element wise `Array` product \n default in NumPy')
plt.subplot(122)
plt.imshow(matrix_product, interpolation='nearest')
plt.title('`Matrix` product \n default in Matlab/Octave')
# plt.show()
# -
# Let's call `c` and `d` interpreted as matrices $C$ and $D$, respectively.
# Since $D$ is the identity matrix, it should come as no surprise that $D C = C$.
#
# ### Reshaping
#
# As mentioned, arrays are easily reshaped, as long as you don't mess up the total number of elements:
e = sp.arange(64).reshape(1, -1) # '-1' here means 'as many as it takes'.
plt.imshow(e, interpolation='nearest')
# plt.show()
e = sp.arange(64).reshape((8, 8)) # Same array, different shape!
plt.imshow(e, interpolation='nearest'); plt.colorbar()
f = e.reshape((4, 16))
plt.imshow(f, interpolation='nearest')
plt.colorbar(orientation='horizontal')
# ### On loops vs. arrays
#
# NumPy is optimized for performing element-wise operations on arrays in parallel.
# This means that you get both cleaner code and much faster computations if you utilize vectorization well.
# To give a completely unscientific idea about the kind of speed you gain, I have written my own function which raises an array`a` to the `x`th power, by looping through the array's indices $(i, j)$ and raise the elements **one by one**, where NumPy does it with all of them in parallel:
# +
def powerit2d(indata, expon): # Version for 2D array
for i in range(indata.shape[0]):
for j in range(indata.shape[1]):
indata[i, j] = indata[i, j] ** expon
return indata
def powerit3d(indata, expon): # Version for 3D array
for i in range(indata.shape[0]):
for j in range(indata.shape[1]):
for k in range(indata.shape[2]):
indata[i, j, k] = indata[i, j, k] ** expon
return indata
aa = ra.random((20, 20))
aaa = ra.random((20, 20, 20))
# -
# We now time the different methods operating on the 2D array. Note that different runs will give different results, this is *not* a strict benchmarking but only to give some rough feeling of the difference:
# %timeit aa ** 2000
# %timeit powerit2d(aa, 2000)
# A bit less than a factor of 50 in difference. Now for the 3D array:
# %timeit aaa ** 2000
# %timeit powerit3d(aaa, 2000)
# ...which gives a slightly larger difference.
#
# **We really want to utilize NumPy's array optimization when we can.**
# ## Slicing & Dicing
#
# *A short description of this section could be "How to elegantly select subsets of your dataset"*.
#
# Try to see what you get out of the following manipulations, and understand what they do. If you learn them, you can work very efficiently with $N$-simensional datasets:
print(e)
print(e[0, 0])
print(e[:, 2:3]) # In these intervals, the last index value is not included.
print(e[2:4, :])
print(e[2:4]) # Think a bit about this one.
print(e[2:-1, 0])
print(e[2:-1, 0:1]) # What's the difference here?
print e
print(e[::-1, 0:2])
print(e[1:6:2, :])
print(e[::2, :])
# You can of course always assign one of these subsets to a new variable:
f = e[1:6:2, :] # Etc.
# ### Logical indexing:
#
# This is a way to perform operations on elements in your array when you don't know exactly where they are, but you know that they live up to a certain logical criterion, e.g. below we say "please show me all elements in the array `g` for which we know that the element is an even number"
g = sp.arange(64).reshape((8, 8))
evens = sp.where(g % 2 == 0); print g[evens]
# The `where()` function can be used to pick elements by pretty complex criteria:
my_indices = sp.where(((g > 10) & (g < 20)) | (g > 45))
# To see exactly what that last one did, we'll try to "color" the elements that it selected and plot the array:
h = sp.ones_like(d); h[my_indices] = 0.
print(h)
plt.imshow(h, interpolation='nearest'); plt.colorbar()
# +
pp = sp.random.random((8, 8))
idx = sp.where(pp > .75)
qq = sp.zeros_like(pp)
qq[idx] = 1
plt.figure()
plt.gcf().set_size_inches(9,4)
plt.subplot(1, 2, 1)
plt.imshow(pp, interpolation='nearest')#; plt.colorbar()
plt.subplot(1, 2, 2)
plt.imshow(qq, interpolation='nearest')
# -
# ## Learning more:
#
# There's a NumPy tutorial at the [SciPy Wiki](http://wiki.scipy.org/Tentative_NumPy_Tutorial).
#
# For users of Matlab and Octave, the web site [NumPy for Matlab users](http://wiki.scipy.org/NumPy_for_Matlab_Users) on the same wiki could be very useful.
#
# There's also a [NumPy for IDL users](http://wiki.scipy.org/NumPy_for_Matlab_Users) page which could possibly be quite useful also if you never used IDL.
# ## A couple of more general videos about Python i Science
# A video about how to use IPython for several steps in your work flow: Computations, plotting, writing papers etc.
from IPython.display import YouTubeVideo
YouTubeVideo('iwVvqwLDsJo')
# A talk by an astronomer at Berkeley about how Python can be used for everything from running a remote telescope over auto-detection of interesting discoveries to the publication process.
YouTubeVideo('mLuIB8aW2KA')
| tools-package/NumpyArrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Classification Inference for High Resolution Images - ONNX Runtime
# In this example notebook, we describe how to use a pre-trained Classification model, using high resolution images, for inference.
# - The user can choose the model (see section titled *Choosing a Pre-Compiled Model*)
# - The models used in this example were trained on the ***ImageNet*** dataset because it is a widely used dataset developed for training and benchmarking image classification AI models.
# - We perform inference on a few sample images.
# - We also describe the input preprocessing and output postprocessing steps, demonstrate how to collect various benchmarking statistics and how to visualize the data.
#
# ## Choosing a Pre-Compiled Model
# We provide a set of precompiled artifacts to use with this notebook that will appear as a drop-down list once the first code cell is executed.
#
# <img src=docs/images/drop_down.PNG width="400">
#
# ## Image classification
# Image classification is a popular computer vision algorithm used in applications such as, object recongnition, traffic sign recongnition and traffic light recongnition. Image classification models are also used as feature extractors for other tasks such as object detection and semantic segmentation.
# - The image below shows classification results on few sample images.
# - Note: in this example, we used models trained with ***ImageNet*** because it is a widely used dataset developed for training and benchmarking image classifcation AI models
#
# <img src=docs/images/CLS.PNG width="500">
#
#
# ## ONNX Runtime based Work flow
# The diagram below describes the steps for ONNX Runtime based workflow.
#
# Note:
# - The user needs to compile models(sub-graph creation and quantization) on a PC to generate model artifacts.
# - For this notebook we use pre-compiled models artifacts
# - The generated artifacts can then be used to run inference on the target.
# - Users can run this notebook as-is, only action required is to select a model.
#
# <img src=docs/images/onnx_work_flow_2.png width="400">
# + tags=["parameters"]
import os
import cv2
import numpy as np
import ipywidgets as widgets
from scripts.utils import get_eval_configs
last_artifacts_id = selected_model_id.value if "selected_model_id" in locals() else None
prebuilt_configs, selected_model_id = get_eval_configs('classification','onnxrt', num_quant_bits = 8, last_artifacts_id = last_artifacts_id,high_resolution=True)
display(selected_model_id)
# -
print(f'Selected Model: {selected_model_id.label}')
config = prebuilt_configs[selected_model_id.value]
config['session'].set_param('model_id', selected_model_id.value)
config['session'].start()
# ## Define utility function to preprocess input images
#
# Below, we define a utility function to preprocess images for the model. This function takes a path as input, loads the image and preprocesses the images as required by the model. The steps below are shown as a reference (no user action required):
#
# 1. Load image
# 2. Convert BGR image to RGB
# 3. Scale image
# 4. Apply per-channel pixel scaling and mean subtraction
# 5. Convert RGB Image to BGR.
# 6. Convert the image to NCHW format
#
#
# - The input arguments of this utility function is selected automatically by this notebook based on the model selected in the drop-down
def preprocess(image_path, size, mean, scale, layout, reverse_channels):
# Step 1
img = cv2.imread(image_path)
# Step 2
img = img[:,:,::-1]
# Step 3
img = cv2.resize(img, (size, size), interpolation=cv2.INTER_CUBIC)
# Step 4
img = img.astype('float32')
for mean, scale, ch in zip(mean, scale, range(img.shape[2])):
img[:,:,ch] = ((img.astype('float32')[:,:,ch] - mean) * scale)
# Step 5
if reverse_channels:
img = img[:,:,::-1]
# Step 6
if layout == 'NCHW':
img = np.expand_dims(np.transpose(img, (2,0,1)),axis=0)
else:
img = np.expand_dims(img,axis=0)
return img
# ## Create the model using the stored artifacts
#
# <div class="alert alert-block alert-warning">
# <b>Warning:</b> It is recommended to use the ONNX Runtime APIs in the cells below without any modifications.
# </div>
# +
import onnxruntime as rt
onnx_model_path = config['session'].get_param('model_file')
delegate_options = {}
so = rt.SessionOptions()
delegate_options['artifacts_folder'] = config['session'].get_param('artifacts_folder')
EP_list = ['TIDLExecutionProvider','CPUExecutionProvider']
sess = rt.InferenceSession(onnx_model_path ,providers=EP_list, provider_options=[delegate_options, {}], sess_options=so)
input_details = sess.get_inputs()
output_details = sess.get_outputs()
# -
# ## Run the model for inference
#
# ### Preprocessing and Inference
#
# - You can use a portion of images provided in `/sample-images` directory to evaluate the classification inferences. In the cell below, we use a loop to preprocess the selected images, and provide them as the input to the network.
#
# ### Postprocessing and Visualization
#
# - Once the inference results are available, we postpocess the results and visualize the inferred classes for each of the input images.
# - Classification models return the results as a list of `numpy.ndarray`, containing one element which is an array with `shape` = `(1,1000)` and `dtype` = `'float32'`, where each element represents the activation for a particular ***ImageNet*** class. The results from the these inferences above are postprocessed using `argsort()` to get the `TOP-5` class IDs and the corresponding names using `imagenet_class_to_name()`.
# - Then, in this notebook, we use *matplotlib* to plot the original images and the corresponding results.
# +
from scripts.utils import get_preproc_props
# use results from the past inferences
images = [
('sample-images/elephant.bmp', 221),
('sample-images/laptop.bmp', 222),
('sample-images/bus.bmp', 223),
('sample-images/zebra.bmp', 224),
]
size, mean, scale, layout, reverse_channels = get_preproc_props(config)
print(f'Image size: {size}')
# +
import tqdm
import matplotlib.pyplot as plt
from scripts.utils import imagenet_class_to_name
plt.figure(figsize=(20,10))
for num in tqdm.trange(len(images)):
image_file, grid = images[num]
img = cv2.imread(image_file)[:,:,::-1]
ax = plt.subplot(grid)
img_in = preprocess(image_file , size, mean, scale, layout, reverse_channels)
if not input_details[0].type == 'tensor(float)':
img_in = np.uint8(img_in)
res = list(sess.run(None, {input_details[0].name: img_in}))[0]
# get the TOP-5 class IDs by argsort()
# and use utility function to get names
output = res.squeeze()
classes = output.argsort()[-5:][::-1]
names = [imagenet_class_to_name(x)[0] for x in classes]
# plot the TOP-5 class names
ax.text(20, 0 * img.shape[0] / 15, names[0], {'color': 'red', 'fontsize': 18, 'ha': 'left', 'va': 'top'})
ax.text(20, 1 * img.shape[0] / 15, names[1], {'color': 'blue', 'fontsize': 14, 'ha': 'left', 'va': 'top'})
ax.text(20, 2 * img.shape[0] / 15, names[2], {'color': 'blue', 'fontsize': 14, 'ha': 'left', 'va': 'top'})
ax.text(20, 3 * img.shape[0] / 15, names[3], {'color': 'blue', 'fontsize': 14, 'ha': 'left', 'va': 'top'})
ax.text(20, 4 * img.shape[0] / 15, names[4], {'color': 'blue', 'fontsize': 14, 'ha': 'left', 'va': 'top'})
# Show the original image
ax.imshow(img)
plt.show()
# -
# ## Plot Inference benchmarking statistics
#
# - During the model execution several benchmarking statistics such as timestamps at different checkpoints, DDR bandwidth are collected and stored. `get_TI_benchmark_data()` can be used to collect these statistics. This function returns a dictionary of `annotations` and the corresponding markers.
# - We provide the utility function plot_TI_benchmark_data to visualize these benchmark KPIs
#
# <div class="alert alert-block alert-info">
# <b>Note:</b> The values represented by <i>Inferences Per Second</i> and <i>Inference Time Per Image</i> uses the total time taken by the inference except the time taken for copying inputs and outputs. In a performance oriented system, these operations can be bypassed by writing the data directly into shared memory and performing on-the-fly input / output normalization.
# </div>
#
# +
from scripts.utils import plot_TI_performance_data, plot_TI_DDRBW_data, get_benchmark_output
stats = sess.get_TI_benchmark_data()
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10,5))
plot_TI_performance_data(stats, axis=ax)
plt.show()
tt, st, rb, wb = get_benchmark_output(stats)
print(f'SoC: J721E/DRA829/TDA4VM')
print(f' OPP:')
print(f' Cortex-A72 @2GHZ')
print(f' DSP C7x-MMA @1GHZ')
print(f' DDR @4266 MT/s\n')
print(f'{selected_model_id.label} :')
print(f' Inferences Per Second : {1000.0/tt :7.2f} fps')
print(f' Inference Time Per Image : {tt :7.2f} ms')
print(f' DDR usage Per Image : {rb+ wb : 7.2f} MB')
| examples/jupyter_notebooks/vcls-hr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Problem 2: Understanding Performance Tunning for Training High Accuracy Deep Models
# + id="iAve6DCL4JH4"
import tensorflow as tf
import time
import numpy as np
from PIL import Image
from tensorflow.keras import datasets, layers, models
import matplotlib.pyplot as plt
# + id="JWoEqyMuXFF4"
(train_images, train_labels), (test_images, test_labels) = datasets.cifar10.load_data()
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images / 255.0, test_images / 255.0
# + [markdown] id="7wArwCTJJlUa"
# ### Verify the data
#
# To verify that the dataset looks correct, let's plot the first 25 images from the training set and display the class name below each image:
#
# + id="K3PAELE2eSU9"
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i])
# The CIFAR labels happen to be arrays,
# which is why you need the extra index
plt.xlabel(class_names[train_labels[i][0]])
plt.show()
# -
# ### Utility functions for modular coding
def create_model():
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(32, 32, 3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.Flatten())
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(10))
return model
def train_model(model, model_name, train_data, train_label, test_data, test_label, lrate=0.01, moment=0.0):
opt = tf.keras.optimizers.SGD(
learning_rate=lrate, momentum=moment, nesterov=False, name='SGD')
model.compile(optimizer=opt,
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
t0 = time.time()
history = model.fit(train_data, train_label, epochs=20, validation_data=(test_data, test_label))
t1 = time.time()
print("Total training time to train ", model_name, t1-t0)
return history, model
def evaluate_model(fig_name, model, history, test_data, test_model):
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.0, 1])
plt.legend(loc='lower right')
plt.savefig(fig_name+'.png')
test_loss, test_acc = model.evaluate(test_data, test_model, verbose=2)
print('Validation accuracy ', test_acc)
# ### (1) Compile, train and calculate accuracy of model with default hyper parameters
model1 = create_model()
history1, model1 = train_model(model1, 'default', train_images, train_labels, test_images, test_labels)
evaluate_model('default', model1, history1, test_images, test_labels)
model1.summary()
# ### (2) Compile, train and calculate accuracy of model for lr = 0.001 and momentum = 0.1
model2 = create_model()
history2, model2 = train_model(model2, 'lr=0.001,m=0.1', train_images, train_labels, test_images, test_labels, 0.001, 0.1)
evaluate_model('lr=0.001,m=0.1', model2, history2, test_images, test_labels)
# ### (3) Compile, train and calculate accuracy of model for lr = 0.1 and momentum = 0.9
model3 = create_model()
history3, model3 = train_model(model3, 'lr=0.1,m=0.9', train_images, train_labels, test_images, test_labels, 0.1, 0.9)
evaluate_model('lr=0.1,m=0.9', model3, history3, test_images, test_labels)
# ### (4) Compile, train and calculate accuracy of model for lr = 0.1 and momentum = 0.1
model4 = create_model()
history4, model4 = train_model(model4, 'lr=0.1,m=0.1', train_images, train_labels, test_images, test_labels, 0.1, 0.1)
evaluate_model('lr=0.1,m=0.1', model4, history4, test_images, test_labels)
# ### (5) Compile, train and calculate accuracy of model for lr = 0.001 and momentum = 0.9
model5 = create_model()
history5, model5 = train_model(model5, 'lr=0.001,m=0.9', train_images, train_labels, test_images, test_labels, 0.001, 0.9)
evaluate_model('lr=0.001,m=0.9', model5, history5, test_images, test_labels)
# ### Confusion matrix analysis
def computeConfusion(model, model_name):
predictions = model.predict(test_images)
predictions = np.argmax(predictions,axis = 1)
confusion_matrix = tf.math.confusion_matrix(
test_labels, predictions, num_classes=None, weights=None, dtype=tf.dtypes.int32,name=None)
print("The confusion matrix is for ", model_name, confusion_matrix)
#Confusion matrix for model1
computeConfusion(model1, "model1")
computeConfusion(model2, "model2")
computeConfusion(model3, "model3")
computeConfusion(model4, "model4")
# ### Predicting images from classes using best model
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[60]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[60]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[2000]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[2000]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[1000]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[1000]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[30]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[30]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[9000]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[9000]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[3003]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[3003]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[3024]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[3024]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[3011]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[3011]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[3016]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[3016]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# +
plt.figure()
plt.imshow(Image.fromarray((test_images[14]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = test_images[14]
idx = np.argmax(model5.predict(image[np.newaxis,:]))
print(class_names[idx])
# -
# ### Model performance on outlier detection
# #### Preprocessing data to account for outlier class
# +
a = train_labels<7
arr_train_l7 = ["False"]*a.size
idx=0
for i in a:
arr_train_l7[idx] = i[0]
idx+=1
a = test_labels<7
arr_test_l7 = ["False"]*a.size
idx=0
for i in a:
arr_test_l7[idx] = i[0]
idx+=1
a = train_labels>=7
arr_train_g7 = ["False"]*a.size
idx=0
for i in a:
arr_train_g7[idx] = i[0]
idx+=1
a = test_labels>=7
arr_test_g7 = ["False"]*a.size
idx=0
for i in a:
arr_test_g7[idx] = i[0]
idx+=1
# -
train_images_modified = train_images[arr_train_l7]
train_labels_modified = train_labels[arr_train_l7]
test_images_modified = test_images[arr_test_l7]
test_labels_modified = test_labels[arr_test_l7]
outlier_images = test_images[arr_test_g7]
outlier_labels = test_labels[arr_test_g7]
# ## Training a new model on 7 classes
model6 = create_model()
history6, model6 = train_model(model6, 'outlier', train_images_modified, train_labels_modified, test_images_modified, test_labels_modified, 0.001, 0.9)
evaluate_model('outlier', model6, history6, test_images_modified, test_labels_modified)
# ### Using the remaining 3 classes as outliers to test the model
# #### Running each cell 3 times.
# +
plt.figure()
plt.imshow(Image.fromarray((outlier_images[14]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = outlier_images[14]
idx = np.argmax(model6.predict(image[np.newaxis,:]))
print(class_names[idx])
#Output on 3 runs: cat, cat, cat
# +
plt.figure()
plt.imshow(Image.fromarray((outlier_images[24]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = outlier_images[24]
idx = np.argmax(model6.predict(image[np.newaxis,:]))
print(class_names[idx])
#Output on 3 runs: airplane, airplane, airplane
# +
plt.figure()
plt.imshow(Image.fromarray((outlier_images[28]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = outlier_images[28]
idx = np.argmax(model6.predict(image[np.newaxis,:]))
print(class_names[idx])
#Output of 3 runs: airplane, airplane, airplane
# +
plt.figure()
plt.imshow(Image.fromarray((outlier_images[30]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = outlier_images[28]
idx = np.argmax(model6.predict(image[np.newaxis,:]))
print(class_names[idx])
#Output of 3 runs: airplane, airplane, airplane
# +
plt.figure()
plt.imshow(Image.fromarray((outlier_images[34]*255).astype(np.uint8)))
plt.colorbar()
plt.grid(False)
plt.show()
image = outlier_images[34]
idx = np.argmax(model6.predict(image[np.newaxis,:]))
print(class_names[idx])
#Output of 3 runs: cat, cat, cat
| Vidushi_Vashishth_Big_Data_HW2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import tensorflow as tf
from keras import backend as K
t1 = tf.constant(3.0, dtype=tf.float32)
t2 = tf.constant(4.0)
print t1, t2
f = K.function([t1], [t2])
s = tf.Session()
print s.run([t1, t2])
print f([3.0])
t3 = tf.add(t1, t2)
f1 = K.function([t1, t2], [t3])
print f1([3, 8])
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
t4 = tf.multiply(a, b)
f2 = K.function([a, b], [t4])
print f2([4.0, 5.0])
| examples/tensor-manipluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This code uses the freud package, alongwith PyLammps, to calculate the averaged RDF over a user-defined range of snapshots in a lammps trajectory file.
# +
import numpy as np
import time
from cycler import cycler
import matplotlib
from matplotlib import pyplot
from matplotlib.colors import colorConverter
myHexColors = ["#30A2DA", "#FC4F30", "#E5AE38", "#6D904F", "#9757DB", "#188487", "#FF7F00", "#9A2C66", "#626DDA", "#8B8B8B"]
myColorCycle = [colorConverter.to_rgb(i) for i in myHexColors]
from freud import parallel
parallel.setNumThreads(4)
from ipywidgets import IntProgress
from IPython.display import display
# matplotlib.rcParams.update({'font.sans-serif': 'Helvetica'})
# matplotlib.rcParams.update({'font.family': 'sans-serif'})
matplotlib.rcParams.update({'font.size': 24})
matplotlib.rcParams.update({'axes.labelsize': 24})
matplotlib.rcParams.update({'xtick.labelsize': 18})
matplotlib.rcParams.update({'ytick.labelsize': 18})
matplotlib.rcParams.update({'savefig.pad_inches': 0.025})
matplotlib.rcParams.update({"axes.prop_cycle": cycler(color=myColorCycle)})
matplotlib.rcParams.update({"lines.linewidth": 2})
# lammps
from lammps import lammps, PyLammps
lmp = lammps()
L = PyLammps(ptr=lmp) # Get PyLammps
# -
# All the user-defined stuff is in the next section :
# User-defined stuff
system = 'liq'
typeID = 1
frameNo = 500000 #500000
endFrame = 515000 #1000000
dumpFreq = 5000 #1000 #5000
nFrames = int((endFrame - frameNo)/dumpFreq) + 1
# Dumpfile name to read from
dumpfile = 'traj/dump-' + system + '.lammpstrj'
# Average the RDF over the frames specified by frameNo and endFrame (inclusive of both). If you specify a frame that is not present in the lammps trajectory file, PyLammps will throw an error.
#
# If you run the cell below more than once, make sure to uncomment the PyLammps command for reading in the data file, or else you will get a Lammps error.
# +
# import the freud object
# the rdf module is contained in the density module
from freud import box, density
# create the rdf object
# make rmax less than half the box?
rdf = density.RDF(rmax=4.0, dr=0.01) # 0.01
# compute the rdf for for all frames except the first
# Current frame
frame = frameNo
# compute the rdf for for all frames except the first (your syntax will vary based on your reader)
myProgressBar = IntProgress(min=1,max=nFrames)
display(myProgressBar)
# Lammps init
# load the data
# Read in the lammps input file
lmp.file('lammpsFiles/inp.'+ system) # LAMMPS input file for the system
# Get the number of atoms of one typeID
natoms = 0
L.read_dump(dumpfile, frameNo, "x y z box yes replace yes")
for i in range(L.atoms.natoms):
if L.atoms[i].type==typeID:
natoms += 1
# Define numPy coordinate array; here natoms is the no. of atoms of type 1
x = np.zeros((natoms,3), dtype=float)
start_time = time.time()
for i in range(nFrames):
# Progress bar
myProgressBar.value = int(frame/dumpFreq)
# --------------------------
# Lammps stuff
# read box, position data from the lammps dumpfile
L.read_dump(dumpfile, frame, "x y z box yes replace yes")
# Get coord
# Assigning values to numpy array x
for i in range(L.atoms.natoms):
if L.atoms[i].type==typeID:
for k in range(3):
x[i][k] = L.atoms[i].position[k]
box_limits = [L.system.xhi-L.system.xlo, L.system.yhi-L.system.ylo, L.system.zhi-L.system.zlo] # box from lammps
# --------------------------
# create the freud box object
fbox = box.Box(Lx=box_limits[0], Ly=box_limits[1], Lz=box_limits[2])
# accumulate
rdf.accumulate(fbox, x, x)
# ----------------------------
# Update frame number
print("Frame number ", frame)
frame = frame + dumpFreq
stop_time = time.time()
print("time to calc {} frames = {}".format(nFrames, stop_time-start_time))
print("speed of calc: {} (frames/sec)".format((nFrames)/(stop_time-start_time)))
# get the center of the histogram bins
r = rdf.getR()
# get the value of the histogram bins
y = rdf.getRDF()
pyplot.figure(figsize=(8, 5))
pyplot.title("RDF - Avg Frames")
pyplot.plot(r, y)
pyplot.xlabel(r"$r$")
pyplot.ylabel(r"$g\left(r\right)$")
pyplot.tight_layout()
pyplot.show()
pyplot.savefig('averaged_RDF.png')
# -
| RDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="YQCt0TA0uaNc"
# <!-- ---
# title: How to create Custom Events based on Forward or Backward Pass
# weight: 8
# downloads: true
# sidebar: true
# summary: Learn how to create custom events that depend on the loss calculated, backward pass, optimization step, etc.
# tags:
# - custom events
# --- -->
# # How to create Custom Events based on Forward or Backward Pass
# + [markdown] id="aMOY2iPOuaNk"
# This guide demonstrates how you can create [custom events](https://pytorch.org/ignite/concepts.html#custom-events) that depend on the loss calculated and backward pass.
#
# In this example, we will be using a ResNet18 model on the MNIST dataset. The base code is the same as used in the [Getting Started Guide](https://pytorch-ignite.ai/tutorials/getting-started/).
# + [markdown] id="THcUNAgpWMDF"
# ## Basic Setup
# + pycharm={"is_executing": false} id="Y0sJP9iFa1TB"
import pandas as pd
import torch
from torch import nn
from torch.utils.data import DataLoader
from torchvision.datasets import MNIST
from torchvision.models import resnet18
from torchvision.transforms import Compose, Normalize, ToTensor
from ignite.engine import Engine, EventEnum, Events, create_supervised_evaluator
from ignite.metrics import Accuracy, Loss
from ignite.handlers import Timer
from ignite.contrib.handlers import BasicTimeProfiler, HandlersTimeProfiler
# + id="iK_9cOP6a1TI"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.model = resnet18(num_classes=10)
self.model.conv1 = nn.Conv2d(1, 64, kernel_size=3, padding=1, bias=False)
def forward(self, x):
return self.model(x)
model = Net().to(device)
data_transform = Compose([ToTensor(), Normalize((0.1307,), (0.3081,))])
train_loader = DataLoader(
MNIST(download=True, root=".", transform=data_transform, train=True),
batch_size=128,
shuffle=True,
)
optimizer = torch.optim.RMSprop(model.parameters(), lr=0.005)
criterion = nn.CrossEntropyLoss()
# + [markdown] id="Q_u0IS8q9IY-"
# ## Create Custom Events
#
# First let's create a few custom events based on backpropogation. All user-defined custom events should inherit from the base class [`EventEnum`](https://pytorch.org/ignite/generated/ignite.engine.events.EventEnum.html#ignite.engine.events.EventEnum).
# + id="TbEoK_H8yIAj"
class BackpropEvents(EventEnum):
BACKWARD_STARTED = 'backward_started'
BACKWARD_COMPLETED = 'backward_completed'
OPTIM_STEP_COMPLETED = 'optim_step_completed'
# + [markdown] id="9lwr621Y9Lnx"
# ## Create `trainer`
#
# Then we define the `train_step` function to be applied on all batches. Within this, we use [`fire_event`](https://pytorch.org/ignite/generated/ignite.engine.engine.Engine.html#ignite.engine.engine.Engine.fire_event) to execute all handlers related to a specific event at that point.
# + id="8aqUFTEdxxvz"
def train_step(engine, batch):
model.train()
optimizer.zero_grad()
x, y = batch[0].to(device), batch[1].to(device)
y_pred = model(x)
loss = criterion(y_pred, y)
engine.fire_event(BackpropEvents.BACKWARD_STARTED)
loss.backward()
engine.fire_event(BackpropEvents.BACKWARD_COMPLETED)
optimizer.step()
engine.fire_event(BackpropEvents.OPTIM_STEP_COMPLETED)
return loss.item()
trainer = Engine(train_step)
# + [markdown] id="eiLRGHAK9Q12"
# ## Register Custom Events in `trainer`
#
# Finally, to make sure our events can be fired, we register them in `trainer` using [`register_events`](https://pytorch.org/ignite/generated/ignite.engine.engine.Engine.html#ignite.engine.engine.Engine.register_events).
# + id="4byi6J6N9d4K"
trainer.register_events(*BackpropEvents)
# + [markdown] id="WZbJwRUD9e-d"
# ## Attach handlers to Custom Events
#
# And now we can easily attach handlers to be executed when a particular event like `BACKWARD_COMPLETED` is fired.
# + id="9Dp6QBfQysOq"
@trainer.on(BackpropEvents.BACKWARD_COMPLETED)
def function_before_backprop(engine):
print(f"Iter[{engine.state.iteration}] Function fired after backward pass")
# + [markdown] id="XMKXagQk-VLl"
# And finally you can run the `trainer` for some epochs.
# + id="3G9DV6h767fj"
trainer.run(train_loader, max_epochs=3)
# + [markdown] id="x031SkP2-Lg9"
# ## Additional Links
#
# You can also checkout the source code of [TBPTT Trainer](https://pytorch.org/ignite/_modules/ignite/contrib/engines/tbptt.html#create_supervised_tbptt_trainer) for a detailed explanation.
| how-to-guides/08-custom-events.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Nl4ZgsC1wawx"
# The following additional libraries are needed to run this
# notebook. Note that running on Colab is experimental, please report a Github
# issue if you have any problem.
# + id="URk0QEy9waw2" colab={"base_uri": "https://localhost:8080/"} outputId="86d8b106-74dd-4ce1-cf53-43ef2c13e577"
# !pip install d2l==0.17.2
# + [markdown] origin_pos=0 id="cRQMGU8Owaw3"
# # Residual Networks (ResNet)
# :label:`sec_resnet`
#
# As we design increasingly deeper networks it becomes imperative to understand how adding layers can increase the complexity and expressiveness of the network.
# Even more important is the ability to design networks where adding layers makes networks strictly more expressive rather than just different.
# To make some progress we need a bit of mathematics.
#
#
# ## Function Classes
#
# Consider $\mathcal{F}$, the class of functions that a specific network architecture (together with learning rates and other hyperparameter settings) can reach.
# That is, for all $f \in \mathcal{F}$ there exists some set of parameters (e.g., weights and biases) that can be obtained through training on a suitable dataset.
# Let us assume that $f^*$ is the "truth" function that we really would like to find.
# If it is in $\mathcal{F}$, we are in good shape but typically we will not be quite so lucky.
# Instead, we will try to find some $f^*_\mathcal{F}$ which is our best bet within $\mathcal{F}$.
# For instance,
# given a dataset with features $\mathbf{X}$
# and labels $\mathbf{y}$,
# we might try finding it by solving the following optimization problem:
#
# $$f^*_\mathcal{F} \stackrel{\mathrm{def}}{=} \mathop{\mathrm{argmin}}_f L(\mathbf{X}, \mathbf{y}, f) \text{ subject to } f \in \mathcal{F}.$$
#
# It is only reasonable to assume that if we design a different and more powerful architecture $\mathcal{F}'$ we should arrive at a better outcome. In other words, we would expect that $f^*_{\mathcal{F}'}$ is "better" than $f^*_{\mathcal{F}}$. However, if $\mathcal{F} \not\subseteq \mathcal{F}'$ there is no guarantee that this should even happen. In fact, $f^*_{\mathcal{F}'}$ might well be worse.
# As illustrated by :numref:`fig_functionclasses`,
# for non-nested function classes, a larger function class does not always move closer to the "truth" function $f^*$. For instance,
# on the left of :numref:`fig_functionclasses`,
# though $\mathcal{F}_3$ is closer to $f^*$ than $\mathcal{F}_1$, $\mathcal{F}_6$ moves away and there is no guarantee that further increasing the complexity can reduce the distance from $f^*$.
# With nested function classes
# where $\mathcal{F}_1 \subseteq \ldots \subseteq \mathcal{F}_6$
# on the right of :numref:`fig_functionclasses`,
# we can avoid the aforementioned issue from the non-nested function classes.
#
#
# 
# :label:`fig_functionclasses`
#
# Thus,
# only if larger function classes contain the smaller ones are we guaranteed that increasing them strictly increases the expressive power of the network.
# For deep neural networks,
# if we can
# train the newly-added layer into an identity function $f(\mathbf{x}) = \mathbf{x}$, the new model will be as effective as the original model. As the new model may get a better solution to fit the training dataset, the added layer might make it easier to reduce training errors.
#
# This is the question that He et al. considered when working on very deep computer vision models :cite:`He.Zhang.Ren.ea.2016`.
# At the heart of their proposed *residual network* (*ResNet*) is the idea that every additional layer should
# more easily
# contain the identity function as one of its elements.
# These considerations are rather profound but they led to a surprisingly simple
# solution, a *residual block*.
# With it, ResNet won the ImageNet Large Scale Visual Recognition Challenge in 2015. The design had a profound influence on how to
# build deep neural networks.
#
#
#
# ## (**Residual Blocks**)
#
# Let us focus on a local part of a neural network, as depicted in :numref:`fig_residual_block`. Denote the input by $\mathbf{x}$.
# We assume that the desired underlying mapping we want to obtain by learning is $f(\mathbf{x})$, to be used as the input to the activation function on the top.
# On the left of :numref:`fig_residual_block`,
# the portion within the dotted-line box
# must directly learn the mapping $f(\mathbf{x})$.
# On the right,
# the portion within the dotted-line box
# needs to
# learn the *residual mapping* $f(\mathbf{x}) - \mathbf{x}$,
# which is how the residual block derives its name.
# If the identity mapping $f(\mathbf{x}) = \mathbf{x}$ is the desired underlying mapping,
# the residual mapping is easier to learn:
# we only need to push the weights and biases
# of the
# upper weight layer (e.g., fully-connected layer and convolutional layer)
# within the dotted-line box
# to zero.
# The right figure in :numref:`fig_residual_block` illustrates the *residual block* of ResNet,
# where the solid line carrying the layer input
# $\mathbf{x}$ to the addition operator
# is called a *residual connection* (or *shortcut connection*).
# With residual blocks, inputs can
# forward propagate faster through the residual connections across layers.
#
# 
# :label:`fig_residual_block`
#
#
# ResNet follows VGG's full $3\times 3$ convolutional layer design. The residual block has two $3\times 3$ convolutional layers with the same number of output channels. Each convolutional layer is followed by a batch normalization layer and a ReLU activation function. Then, we skip these two convolution operations and add the input directly before the final ReLU activation function.
# This kind of design requires that the output of the two convolutional layers has to be of the same shape as the input, so that they can be added together. If we want to change the number of channels, we need to introduce an additional $1\times 1$ convolutional layer to transform the input into the desired shape for the addition operation. Let us have a look at the code below.
#
# + origin_pos=3 tab=["tensorflow"] id="7mZjx3jMwaw7"
import tensorflow as tf
from d2l import tensorflow as d2l
class Residual(tf.keras.Model): #@save
"""The Residual block of ResNet."""
def __init__(self, num_channels, use_1x1conv=False, strides=1):
super().__init__()
self.conv1 = tf.keras.layers.Conv2D(
num_channels, padding='same', kernel_size=3, strides=strides)
self.conv2 = tf.keras.layers.Conv2D(
num_channels, kernel_size=3, padding='same')
self.conv3 = None
if use_1x1conv:
self.conv3 = tf.keras.layers.Conv2D(
num_channels, kernel_size=1, strides=strides)
self.bn1 = tf.keras.layers.BatchNormalization()
self.bn2 = tf.keras.layers.BatchNormalization()
def call(self, X):
Y = tf.keras.activations.relu(self.bn1(self.conv1(X)))
Y = self.bn2(self.conv2(Y))
if self.conv3 is not None:
X = self.conv3(X)
Y += X
return tf.keras.activations.relu(Y)
# + [markdown] origin_pos=4 id="huChYboHwaw7"
# This code generates two types of networks: one where we add the input to the output before applying the ReLU nonlinearity whenever `use_1x1conv=False`, and one where we adjust channels and resolution by means of a $1 \times 1$ convolution before adding. :numref:`fig_resnet_block` illustrates this:
#
# 
# :label:`fig_resnet_block`
#
# Now let us look at [**a situation where the input and output are of the same shape**].
#
# + origin_pos=7 tab=["tensorflow"] id="SiBCtkUKwaw8" outputId="008e7d4f-984f-4b8d-8951-84c57e5a9209" colab={"base_uri": "https://localhost:8080/"}
blk = Residual(3)
X = tf.random.uniform((4, 6, 6, 3))
Y = blk(X)
Y.shape
# + [markdown] origin_pos=8 id="yPOrfftRwaw9"
# We also have the option to [**halve the output height and width while increasing the number of output channels**].
#
# + origin_pos=11 tab=["tensorflow"] id="lvQxy5DFwaw-" outputId="acb8ae06-2cd0-4148-fa32-632c1be5e09c" colab={"base_uri": "https://localhost:8080/"}
blk = Residual(6, use_1x1conv=True, strides=2)
blk(X).shape
# + [markdown] origin_pos=12 id="kO155bjSwaw_"
# ## [**ResNet Model**]
#
# The first two layers of ResNet are the same as those of the GoogLeNet we described before: the $7\times 7$ convolutional layer with 64 output channels and a stride of 2 is followed by the $3\times 3$ maximum pooling layer with a stride of 2. The difference is the batch normalization layer added after each convolutional layer in ResNet.
#
# + origin_pos=15 tab=["tensorflow"] id="iRCJ9bALwaxA"
b1 = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(64, kernel_size=7, strides=2, padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('relu'),
tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same')])
# + [markdown] origin_pos=16 id="nOiOeRhiwaxA"
# GoogLeNet uses four modules made up of Inception blocks.
# However, ResNet uses four modules made up of residual blocks, each of which uses several residual blocks with the same number of output channels.
# The number of channels in the first module is the same as the number of input channels. Since a maximum pooling layer with a stride of 2 has already been used, it is not necessary to reduce the height and width. In the first residual block for each of the subsequent modules, the number of channels is doubled compared with that of the previous module, and the height and width are halved.
#
# Now, we implement this module. Note that special processing has been performed on the first module.
#
# + origin_pos=19 tab=["tensorflow"] id="GONXhKh4waxB"
class ResnetBlock(tf.keras.layers.Layer):
def __init__(self, num_channels, num_residuals, first_block=False,
**kwargs):
super(ResnetBlock, self).__init__(**kwargs)
self.residual_layers = []
for i in range(num_residuals):
if i == 0 and not first_block:
self.residual_layers.append(
Residual(num_channels, use_1x1conv=True, strides=2))
else:
self.residual_layers.append(Residual(num_channels))
def call(self, X):
for layer in self.residual_layers.layers:
X = layer(X)
return X
# + [markdown] origin_pos=20 id="7j9kyJFqwaxB"
# Then, we add all the modules to ResNet. Here, two residual blocks are used for each module.
#
# + origin_pos=23 tab=["tensorflow"] id="YbEfF5XDwaxC"
b2 = ResnetBlock(64, 2, first_block=True)
b3 = ResnetBlock(128, 2)
b4 = ResnetBlock(256, 2)
b5 = ResnetBlock(512, 2)
# + [markdown] origin_pos=24 id="Br4hEJU1waxC"
# Finally, just like GoogLeNet, we add a global average pooling layer, followed by the fully-connected layer output.
#
# + origin_pos=27 tab=["tensorflow"] id="cSo6BVE-waxC"
# Recall that we define this as a function so we can reuse later and run it
# within `tf.distribute.MirroredStrategy`'s scope to utilize various
# computational resources, e.g. GPUs. Also note that even though we have
# created b1, b2, b3, b4, b5 but we will recreate them inside this function's
# scope instead
def net():
return tf.keras.Sequential([
# The following layers are the same as b1 that we created earlier
tf.keras.layers.Conv2D(64, kernel_size=7, strides=2, padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Activation('relu'),
tf.keras.layers.MaxPool2D(pool_size=3, strides=2, padding='same'),
# The following layers are the same as b2, b3, b4, and b5 that we
# created earlier
ResnetBlock(64, 2, first_block=True),
ResnetBlock(128, 2),
ResnetBlock(256, 2),
ResnetBlock(512, 2),
tf.keras.layers.GlobalAvgPool2D(),
tf.keras.layers.Dense(units=10)])
# + [markdown] origin_pos=28 id="xJReL-2UwaxD"
# There are 4 convolutional layers in each module (excluding the $1\times 1$ convolutional layer). Together with the first $7\times 7$ convolutional layer and the final fully-connected layer, there are 18 layers in total. Therefore, this model is commonly known as ResNet-18.
# By configuring different numbers of channels and residual blocks in the module, we can create different ResNet models, such as the deeper 152-layer ResNet-152. Although the main architecture of ResNet is similar to that of GoogLeNet, ResNet's structure is simpler and easier to modify. All these factors have resulted in the rapid and widespread use of ResNet. :numref:`fig_resnet18` depicts the full ResNet-18.
#
# 
# :label:`fig_resnet18`
#
# Before training ResNet, let us [**observe how the input shape changes across different modules in ResNet**]. As in all the previous architectures, the resolution decreases while the number of channels increases up until the point where a global average pooling layer aggregates all features.
#
# + origin_pos=31 tab=["tensorflow"] id="YlEc_60_waxD" outputId="96fd8efa-2d81-41a5-e8f5-313f5cac3701" colab={"base_uri": "https://localhost:8080/"}
X = tf.random.uniform(shape=(1, 224, 224, 1))
for layer in net().layers:
X = layer(X)
print(layer.__class__.__name__,'output shape:\t', X.shape)
# + [markdown] origin_pos=32 id="IxAqylwjwaxE"
# ## [**Training**]
#
# We train ResNet on the Fashion-MNIST dataset, just like before.
#
# + origin_pos=33 tab=["tensorflow"] id="pVa-vRSBwaxE" outputId="2f4e5b55-ab0c-48b2-c7c8-140753272a70" colab={"base_uri": "https://localhost:8080/", "height": 316}
lr, num_epochs, batch_size = 0.05, 10, 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=96)
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
# + [markdown] origin_pos=34 id="MNDhv97dwaxF"
# ## Summary
#
# * Nested function classes are desirable. Learning an additional layer in deep neural networks as an identity function (though this is an extreme case) should be made easy.
# * The residual mapping can learn the identity function more easily, such as pushing parameters in the weight layer to zero.
# * We can train an effective deep neural network by having residual blocks. Inputs can forward propagate faster through the residual connections across layers.
# * ResNet had a major influence on the design of subsequent deep neural networks, both for convolutional and sequential nature.
#
#
# ## Exercises
#
# 1. What are the major differences between the Inception block in :numref:`fig_inception` and the residual block? After removing some paths in the Inception block, how are they related to each other?
# 1. Refer to Table 1 in the ResNet paper :cite:`He.Zhang.Ren.ea.2016` to
# implement different variants.
# 1. For deeper networks, ResNet introduces a "bottleneck" architecture to reduce
# model complexity. Try to implement it.
# 1. In subsequent versions of ResNet, the authors changed the "convolution, batch
# normalization, and activation" structure to the "batch normalization,
# activation, and convolution" structure. Make this improvement
# yourself. See Figure 1 in :cite:`He.Zhang.Ren.ea.2016*1`
# for details.
# 1. Why can't we just increase the complexity of functions without bound, even if the function classes are nested?
#
# + [markdown] origin_pos=37 tab=["tensorflow"] id="POaKWwxmwaxF"
# [Discussions](https://discuss.d2l.ai/t/333)
#
| week 12/resnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#xls2xlsx_converster
#created by <NAME>
#https://github.com/jinlee487/xls_to_xlsx_conversion_script
# -
import sys
import pandas as pd
import json
import os
import datetime
import glob
pd.set_option('display.width', 400)
pd.set_option('display.max_columns', None)
#TODO
#autoscan
# +
#https://www.geeksforgeeks.org/python-convert-an-html-table-into-excel/
#https://stackoverflow.com/questions/51898826/converting-object-column-in-pandas-dataframe-to-datetime
def convert_xls_to_xlsx(path,name):
try:
df = pd.read_html(path+"\\"+name+".xls")[0]
df[4] = df[4].str.split(' ').str[0]
# df[4] = pd.to_datetime(df[4].str.strip(), errors='ignore',format="%YYYY-%mm-%dd %HH:%MM:%SS")
#2021-03-02 13:30:22
print("\n")
print(df[[0, 2, 3, 4]])
print("\n")
choice = input("\n작업을 계속 진행하고 싶으시면 Y 아니면 y 를 입력해주세요.\n\n").strip()
if choice == "Y" or choice == "y":
df.to_excel(path+"\\"+name+".xlsx",header=False, index=False)
print("\n 다음 경로로"+path+"\n\n "+name+".xlsx\n\n 파일을 성공적으로 저장하였습니다.\n")
print("\nOperation successful.....Terminating the program.\n")
else:
print("\n.....Terminating the program.\n")
quit()
except Exception as e:
print("\n\nAn exception occurred!\n")
print(e)
print("\n\t.........Terminating the program.\n\n\n")
# -
def read_settings():
print("\n\n...Loading xls_to_xlsx_conversion_script\n\n...configuring settings\n")
with open(os.path.join(os.path.dirname(__file__),"config.json"), encoding='UTF8') as json_data:
f = json.load(json_data)
print(f"....xls file directory : {f['directory']}\n\n")
print("\n")
return f["directory"]
def get_subdirs(directory):
"Get a list of immediate subdirectories"
return next(os.walk(directory))[1]
# +
def choose_function():
directory = read_settings()
choice = input("\t변환하고 싶은 xls 파일의 path를 지정해주세요.\n \n\tEnter 1 : 직접 입력.\n \n\tEnter 2 : 자동 파일 찾기.\n \n\t기타: close program.\n\n")
if choice == "1":
print(f"\n\tYou entered {choice} : 직접 입력.\n")
path = input("\n\tEnter file path : ")
name = input("\n\tEnter file name (without extention): ")
if(len(path)>0):
print(f"\n\tYou entered \n{path}\\{name}.xls\n Initializing conversion ~~\n")
convert_xls_to_xlsx(path,name)
else:
print(f"\n\tAn exception occurred.\n You entered {path}. Path is unrecognizable ~~\n")
quit()
elif choice == "2":
print(f"\n\tYou entered {choice} : 자동 파일 찾기. \n")
today = datetime.datetime.now()
year = str(today.year)
month = str(today.month) +"월"
print("\tToday's date:",today,"\n")
year_dir = get_subdirs(directory)
if any(year in s for s in year_dir):
directory = directory + "\\" + year
month_dir = get_subdirs(directory)
if any(month in s for s in month_dir):
directory = directory + "\\" + month
file_dir=max(glob.glob(directory+"\\*"), key=os.path.getmtime)
file_name=os.path.basename(file_dir)
file_name_n_ext=os.path.splitext(file_name)[0]
print("\tfound file in the following path ... \n")
print(f"\tfull path : {file_dir}\n")
print("\tdirectory : " + directory + "\n")
print("\tfile_name_n_ext : " + file_name_n_ext+"\n Initializing conversion ~~\n")
convert_xls_to_xlsx(directory,file_name_n_ext)
else:
print("\n\t.........Terminating the program.\n\n\n")
quit()
# -
choose_function()
| xls2xlsx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Optimization for multiuser communications
# We consider a multiple-input multiple-output (MIMO) communication system
# with $n$ transmitters and $n$ receivers. Each transmitter transmits with power $p_j$ and the
# gain from transmitter $j$ to receiver $i$ is $G_{ij}$. The signal power from transmitter $i$ to receiver $i$ is then
#
# $$ S_i = G_{ii} p_i $$
#
# and the interference is
#
# $$ I_i = \sum_{j\neq i} G_{ij} p_j + \sigma_i $$
#
# where $\sigma_i$ is an additive noise component. In this notebook we consider different strategies for optimizing the signal-to-inference-plus-noise ratio (SINR)
#
# $$ s_i = \frac{G_{ii} p_i}{\sum_{j\neq i} G_{ij} p_j + \sigma_i} $$
#
# with a bound on the total transmitted power $ \sum_i p_i \leq P $.
# ## Minimizing total power for given SINRs
# Suppose we are given lower bounds $s_i \geq \gamma_i$. We can then minimize the required power
# $$
# \begin{array}{ll}
# \text{minimize} & \sum_i p_i \\
# \text{subject to} & s_i \geq \gamma_i \\
# & \sum_i p_i \leq P,
# \end{array}
# $$
# which is equivalent to a linear optimization problem
# $$
# \begin{array}{ll}
# \text{minimize} & \sum_i p_i \\
# \text{subject to} & G_{ii} p_i \geq \gamma_i\left ( \sum_{j\neq i} G_{ij} p_j + \sigma_i \right ) \\
# & \sum_i p_i \leq P.
# \end{array}
# $$
# ## Maximizing the worst SINR
# Alternatively we can maximize the smallest $s_i$,
#
# $$
# \begin{array}{ll}
# \text{maximize} & t \\
# \text{subject to} & s_i \geq t \\
# & \sum_i p_i \leq P.
# \end{array}
# $$
#
# Equivalently we can minimize the inverse,
#
# $$
# \begin{array}{ll}
# \text{minimize} & t^{-1} \\
# \text{subject to} & t \left ( \sum_{j\neq i} G_{ij} p_j + \sigma_i \right ) G_{ii}^{-1} p_i^{-1} \leq 1 \\
# & \sum_i p_i \leq P,
# \end{array}
# $$
#
# which can be rewritten as a geometric programming problem
#
# $$
# \begin{array}{ll}
# \text{minimize} & -z\\
# \text{subject to} &
# \log \left ( \sum_{j\neq i}e^{z + q_j - q_i + \log(G_{ij}/G_{ii})} + e^{z - q_i + \log(\sigma_i/G_{ii})} \right ) \leq 0\\
# & \log \left ( \sum_i e^{q_i-\log P}\right) \leq 0
# \end{array}
# $$
#
# with $p_i := e^{q_i}$ and $t := e^z$. To rewrite the geometric program into conic form, we note that
#
# $$
# \log \left( \sum_{i=1}^n e^{a_i^T x + b_i}\right) \leq 0 \qquad \Longleftrightarrow \qquad
# \sum_i u_i\leq 1, \quad (u_i, 1, a_i^Tx + b_i)\in K_\text{exp}, \: i=1,\dots n.
# $$
import sys
import numpy as np
from mosek.fusion import *
import matplotlib.pyplot as plt
def logsumexp(M, A, x, b):
u = M.variable(A.shape[0])
M.constraint( Expr.sum(u), Domain.lessThan(1.0))
M.constraint( Expr.hstack(u,
Expr.constTerm(A.shape[0], 1.0),
Expr.add(Expr.mul(A, x), b)), Domain.inPExpCone())
def max_worst_sinr(G, P, sigma):
n = G.shape[0]
with Model('MAX_WORST_SINR') as M:
qz = M.variable('qz', n+1) # concatenation of q and z
M.objective('Objective',ObjectiveSense.Minimize,Expr.neg(qz.index(n)))
for i in range(n):
A = np.zeros((n,n+1))
b = np.zeros(n)
for j in [k for k in range(n) if k!=i]:
A[j,[i,j,n]] = [-1, 1, 1]
b[j] = G[i,j]/G[i,i]
A[i, [i, n]] = [-1, 1]
b[i] = sigma[i]/G[i,i]
# If any Gij == 0, then we filter out row j
idx = np.nonzero(b)[0]
logsumexp(M, A[idx,:], qz, np.log(b[idx]))
logsumexp(M, np.eye(n), qz.slice(0,n), -np.log(P)*np.ones(n))
M.setLogHandler(sys.stdout)
M.solve()
pt = np.exp(qz.level())
return (pt[0:n], pt[n])
# +
P = 0.5
G = np.array([[1.0,0.1,0.2,0.1,0.0],
[0.1,1.0,0.1,0.1,0.0],
[0.2,0.1,2.0,0.2,0.2],
[0.1,0.1,0.2,1.0,0.1],
[0.0,0.0,0.2,0.1,1.0]])
sigma = 0.01*np.ones(G.shape[0])
# -
p1, t1 = max_worst_sinr(G, P, sigma)
p1, t1
SINR1 = (np.diagonal(G)*p1)/(np.dot(G,p1) - np.diagonal(G)*p1 + sigma)
SINR1
# ## Maximizing the best SINR
# The solution to
# $$
# \begin{array}{ll}
# \text{maximize} & t_i \\
# \text{subject to} & s_i \leq t_i \\
# & \sum_i p_i \leq P
# \end{array}
# $$
# is trivial; we choose the index $k$ maximizing $P_{ii}/\sigma_i$ and take $p_k=P$ and $p_j=0,\: j\neq k$.
def max_best_SINR(G,P,sigma):
GSD = [G[i][i]/sigma[i] for i in range(G.shape[0])]
P_max = max(GSD)
#Thus, maximum of the best SINR is equal to...
return(P_max*P)
max_best_SINR(G,P,sigma)
# ## Maximizing average SINR
# We can maximize the average SINR as
# $$
# \begin{array}{ll}
# \text{maximize} & \sum_i t_i \\
# \text{subject to} & s_i \geq t_i \\
# & 0 \leq p_i \leq P_i \\
# & \sum_i p_i \leq P,
# \end{array}
# $$
# which corresponds to an intractable non-convex bilinear optimization problem. However, in the low-SINR regime, we can approximate the above problem by maximizing $\sum_i \log t_i$, or equivalently minimizing $\prod_i t_i^{-1}$:
# $$
# \begin{array}{ll}
# \text{minimize} & \prod_i t_i^{-1} \\
# \text{subject to} & t_i \left ( \sum_{j\neq i} G_{ij} p_j + \sigma_i \right ) G_{ii}^{-1} p_i^{-1} \leq 1 \\
# & 0 \leq p_i \leq P_i \\
# & \sum_i p_i \leq P,
# \end{array}
# $$
# which again corresponds to a geometric programming problem.
def min_Geo_mean(G,P,sigma):
n = G.shape[0]
with Model('MIN_GEO_MEAN') as M:
t = M.variable('t',n)
x = M.variable('x',n)
q = M.variable('q',n)
logsumexp(M,np.eye(n),q,-np.log(P)*np.ones(n))
M.constraint(Expr.hstack(x,Expr.constTerm(n, 1.0),Expr.neg(t)),Domain.inPExpCone())
M.objective('Objective',ObjectiveSense.Minimize,Expr.sum(x))
for i in range(n):
A = np.zeros((n,n+1))
b = np.zeros(n)
for j in [k for k in range(n) if k!=i]:
A[j,[i,j,n]] = [-1,1,1]
b[j] = G[i,j]/G[i,i]
A[i,[i,n]] = [-1,1]
b[i] = sigma[i]/G[i,i]
idx = np.nonzero(b)[0]
logsumexp(M,A[idx,:],Expr.vstack(q,t.index(i)),np.log(b[idx]))
M.setLogHandler(sys.stdout)
M.solve()
T = t.level()
p = np.exp(q.level())
return(T,p)
t2,p2 = min_Geo_mean(G, P, sigma)
p2,t2
SINR2 = (np.diagonal(G)*p2)/(np.dot(G,p2) - np.diagonal(G)*p2 + sigma)
SINR2
# # Comparing the SINR for the cases above...
# +
fig,ax = plt.subplots(figsize = (13,10))
bar_width = 0.35
p_num = np.arange(1,G.shape[0]+1)
B1 = ax.bar(p_num,SINR1,bar_width,label = 'Max Worst SINR')
B2 = ax.bar(p_num+bar_width,SINR2,bar_width,label = 'Max Average SINR')
ax.set_ylabel('SINR')
ax.set_xticks(p_num + bar_width/2)
x_tiK = ['p{}'.format(i+1) for i in range(G.shape[0])]
ax.set_xticklabels(x_tiK)
ax.set_xlabel('Transmitter')
ax.legend()
plt.show()
# -
# <a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>. The **MOSEK** logo and name are trademarks of <a href="http://mosek.com">Mosek ApS</a>. The code is provided as-is. Compatibility with future release of **MOSEK** or the `Fusion API` are not guaranteed. For more information contact our [support](mailto:<EMAIL>).
| sinr-optimization/sinr-optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="B9Zrj8DMusOn"
# ## 1) Library & Data Import
# + executionInfo={"elapsed": 941, "status": "ok", "timestamp": 1632444254640, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="yzvddH4TuuwD"
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# + executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1632444254641, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="Q_0x25QSuxrp"
df = pd.read_csv("https://raw.githubusercontent.com/yoonkt200/FastCampusDataset/master/bourne_scenario.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 9, "status": "ok", "timestamp": 1632444254641, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="90IQykX7vZ3t" outputId="90949266-6a2c-4fcb-c32a-14f8e053e7cd"
df.head()
# + [markdown] id="F8mjnHsgvioB"
# #### Feature Description
# - page_no : 데이터가 위치한 pdf 페이지 정보
# - scene_title : 씬 제목
# - text : 씬에 해당하는 지문/대본 텍스트 정보
# + [markdown] id="i3P4CXUNvqtO"
# -----
# + [markdown] id="EmWaX52Rvuol"
# ## 2) 데이터셋 살펴보기
# + [markdown] id="Fjj86LoLv3Wl"
# ### 2-1) 기본 정보 탐색
# + [markdown] id="BZdMSr7Hyv3C"
# ##### 데이터셋 기본 정보 탐색
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 286, "status": "ok", "timestamp": 1632444916399, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="yTRz57Qpvgl4" outputId="63061583-09ea-41ed-bf83-4275dcff8ed1"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 305, "status": "ok", "timestamp": 1632444920530, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lJq9dP3qvlMg" outputId="c0fa336c-f3d5-48d3-a19f-e04e035901b7"
df.info()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 435, "status": "ok", "timestamp": 1632444941574, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="K7SiIjlDwFGG" outputId="d8d7eeca-e266-4a01-955f-8d8b0749bc2c"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"elapsed": 454, "status": "ok", "timestamp": 1632444931129, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="c51l1Bld6c15" outputId="5383991c-1378-4f28-9398-2230f45c0082"
df['text'][0]
# + [markdown] id="7Ob0pe0oxI6Y"
# -----
# + [markdown] id="vtLhtuYwxI86"
# ## 3) 텍스트 데이터 전처리
# + [markdown] id="ZAFsj3ppxI_J"
# ### 3-1) 정규 표현식 적용
# + executionInfo={"elapsed": 317, "status": "ok", "timestamp": 1632445318715, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="__acxhPiw8X2"
# 공백, 특수문자 거르고 대소문자를 소문자로 통일
import re
def apply_regular_expression(text):
text = text.lower()
english = re.compile('[^ a-z]') # 띄어쓰기를 포함한 알파벳 영어 추출
result = english.sub('',text) # 정규표현식을 text에 적용
result = re.sub(' +',' ',result) # 띄어쓰기 2개 이상인것 하나로 만들기
return result
# + colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"elapsed": 289, "status": "ok", "timestamp": 1632445336712, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="hE60OR0mxP0S" outputId="bab321a8-174e-4394-c15e-03a14f5ab813"
# 함수 테스트
apply_regular_expression(df['text'][0])
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 313, "status": "ok", "timestamp": 1632445380486, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="zBgiHxOW3KhY" outputId="a669a636-35db-455b-e0a4-2b8855743bd1"
df['processed_text'] = df['text'].apply(lambda x: apply_regular_expression(x))
df.head()
# + [markdown] id="4eAD9Zhs3ZjB"
# ### 3-2) Word Count
# + [markdown] id="IvmzcY4k7BBQ"
# ##### 말뭉치(코퍼스) 생성
# + id="WCtEGvDa3KkA"
# 말뭉치 생성
corpus = df['processed_text'].tolist()
corpus
# + [markdown] id="tsN3ydKi3cNB"
# ##### BoW 벡터 생성
# + executionInfo={"elapsed": 376, "status": "ok", "timestamp": 1632445589484, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="oMEmCNjQ3CE6"
from sklearn.feature_extraction.text import CountVectorizer
# filter stop words
vect = CountVectorizer(tokenizer=None, stop_words="english", analyzer='word').fit(corpus) # tokenizer : 문장단위를 어떻게 나눌건지, stop_words : 불용어 설정(실질적인 의미가 없는것 걸러줌), analyzer : 단어단위로
bow_vect = vect.fit_transform(corpus)
word_list = vect.get_feature_names()
count_list = bow_vect.toarray().sum(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 350, "status": "ok", "timestamp": 1632445618057, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="YOR-TYff3jzw" outputId="910c95ce-293c-4ba6-9db4-e4efe74f53f0"
word_list[:5]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 287, "status": "ok", "timestamp": 1632445613029, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="_PqufkPv7R6D" outputId="93452e4d-9b2f-41ae-d0f8-1ca21c543ecc"
count_list[:5] # word_list에 나온 저 단어들이 얼마나 나왔는지 볼 수 있다
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 340, "status": "ok", "timestamp": 1632445643451, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="nlY88bL27R9b" outputId="d58fd805-bbad-4960-aa9e-bb7e6980d6da"
bow_vect.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 288, "status": "ok", "timestamp": 1632445657324, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="f-rSHKzD7R_w" outputId="4b20a23a-232b-4f9f-cde9-a641d9990a43"
bow_vect.toarray()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 291, "status": "ok", "timestamp": 1632445671388, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="eqOGgz4B7SCd" outputId="a789be93-fcf7-4009-e9d6-4599675a7b19"
bow_vect.toarray().sum(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 313, "status": "ok", "timestamp": 1632445730189, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="xn3aU4l97V8K" outputId="ab323b1b-81b8-492c-98d1-8061262c5a75"
# 단어별 타입된 횟수 딕셔너리로
word_count_dict = dict(zip(word_list, count_list))
print(str(word_count_dict)[:100])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1632445804987, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="Lvqij9kP7Xgv" outputId="1a27fd02-bd31-4acd-8685-0990eb901dd6"
# 빈도수 순 정렬
import operator
sorted(word_count_dict.items(), key=operator.itemgetter(1), reverse=True)[:5]
# + id="Zu2v6oUn7aUx"
# + [markdown] id="huXcIK1D7eGI"
# ##### 단어 분포 탐색
# + colab={"base_uri": "https://localhost:8080/", "height": 265} executionInfo={"elapsed": 928, "status": "ok", "timestamp": 1632445874103, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="ePWpmhzL7f0A" outputId="5643bbcc-6d93-44ad-ba5d-28270ce5696e"
plt.hist(list(word_count_dict.values()), bins=150)
plt.show()
# + [markdown] id="QI0yqJ7PxS1q"
# -----
# + [markdown] id="3Nb-h1fM7nLc"
# ## 4) 텍스트 마이닝
# + [markdown] id="87E9Sz5g4YFE"
# ### 4-1) 단어별 빈도 분석
# + [markdown] id="MPC-EOxb4YIm"
# ##### 워드 클라우드 시각화
# + id="WXgfHMLP7004"
# # !pip install pytagcloud pygame simplejson
# + colab={"base_uri": "https://localhost:8080/", "height": 297} executionInfo={"elapsed": 693, "status": "ok", "timestamp": 1632446054145, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="PS27E0CT4h-7" outputId="a01c62a8-f507-4921-9d07-c84dea8fb594"
from collections import Counter
import random
import pytagcloud
import webbrowser
ranked_tags = Counter(word_count_dict).most_common(25) # 가장많이 나온 단어 상위 25개를 불러옴
taglist = pytagcloud.make_tags(sorted(word_count_dict.items(), key=operator.itemgetter(1), reverse=True)[:40], maxsize=60)
pytagcloud.create_tag_image(taglist, 'wordcloud_example.jpg', rectangular=False) # taglist : 이미지 생성, 뒤에 이름, 직사각형여부
from IPython.display import Image
Image(filename='wordcloud_example.jpg') # 이미지 출력
# + [markdown] id="_L_bAJUD-BYm"
# ##### 상위 빈도수 단어 출력
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 285, "status": "ok", "timestamp": 1632446088583, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lXvHnoTY96zZ" outputId="c78e3f82-b050-46c0-cb08-797b25dba5c6"
ranked_tags
# + [markdown] id="ocEMVqzM-MdE"
# -----
# + [markdown] id="dK64F8GZ-N8s"
#
# + [markdown] id="Pe7BOBrW-OAw"
# ### 4-2) 장면별 중요 단어 시각화
# + [markdown] id="EaalHFft-Sra"
# ##### TF-IDF 변환
# + executionInfo={"elapsed": 280, "status": "ok", "timestamp": 1632446161930, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="-Wxv-F3P-NEU"
# text의 의미 점수부여
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_vectorizer = TfidfTransformer()
tf_idf_vect = tfidf_vectorizer.fit_transform(bow_vect)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 377, "status": "ok", "timestamp": 1632446169072, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="rsDAFsCY-Unv" outputId="49a161f6-ce15-4d98-eb81-2509be7580cb"
print(tf_idf_vect.shape)
print(tf_idf_vect[0])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 284, "status": "ok", "timestamp": 1632446269030, "user": {"displayName": "\ud321\<PASSWORD>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="dWt8pyEQ-YeT" outputId="528996a1-638c-46ca-a675-a85620e2b18a"
print(tf_idf_vect[0].toarray().shape)
print(tf_idf_vect[0].toarray())
# + [markdown] id="LFT1BfiJ-Y8-"
# ##### 벡터 : 단어 맵핑
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 295, "status": "ok", "timestamp": 1632446332729, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="rzbmSQ8h-XGF" outputId="5cf40f95-108f-45ea-abf7-e8e080df4c1f"
invert_index_vectorizer = {v: k for k, v in vect.vocabulary_.items()}
print(str(invert_index_vectorizer)[:100]+'..')
# + id="ttxpR5npYeD0"
vect.vocabulary_
# + [markdown] id="XSmFxYD6-tec"
# ##### 중요 단어 추출 - Top 3 TF-IDF
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 288, "status": "ok", "timestamp": 1632446350818, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="RPmzms_d-dzM" outputId="7a84444b-816a-45b2-d6b7-312e2458a888"
# 첫번째 문장의 tf_idf 값이 높은 순으로 정렬(3개)
np.argsort(tf_idf_vect[0].toarray())[0][-3:]
# + id="JBnbCA0d-ztq"
np.argsort(tf_idf_vect.toarray())[:, -3:]
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 432, "status": "ok", "timestamp": 1632446466627, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="sKpKmuM7-0ul" outputId="23797821-73c8-49ae-8d45-c690ee9e2418"
top_3_word = np.argsort(tf_idf_vect.toarray())[:, -3:]
df['important_word_indexes'] = pd.Series(top_3_word.tolist())
df.head()
# + executionInfo={"elapsed": 285, "status": "ok", "timestamp": 1632446522225, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="v4C0B2xO-191"
def convert_to_word(x):
word_list = []
for word in x:
word_list.append(invert_index_vectorizer[word])
return word_list
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 309, "status": "ok", "timestamp": 1632446529465, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="WWCeMCOM-5KF" outputId="b3e7c2fa-9e15-4cd9-954c-92de66904131"
df['important_words'] = df['important_word_indexes'].apply(lambda x: convert_to_word(x))
df.head()
# -
#
#
#
#
#
#
# + [markdown] id="B9Zrj8DMusOn"
# ## 1) Library & Data Import
# + executionInfo={"elapsed": 873, "status": "ok", "timestamp": 1632447782112, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="yzvddH4TuuwD"
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# + executionInfo={"elapsed": 315, "status": "ok", "timestamp": 1632447782426, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="Q_0x25QSuxrp"
df = pd.read_csv("https://raw.githubusercontent.com/yoonkt200/FastCampusDataset/master/tripadviser_review.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 273, "status": "ok", "timestamp": 1632447782695, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="90IQykX7vZ3t" outputId="5973e55a-fb49-4f4e-cf4e-9db5ba1c39bd"
df.head()
# + [markdown] id="F8mjnHsgvioB"
# #### Feature Description
# - rating : 이용자 리뷰의 평가 점수
# - text : 이용자 리뷰 평가 내용
# + [markdown] id="i3P4CXUNvqtO"
# -----
# + [markdown] id="EmWaX52Rvuol"
# ## 2) 데이터셋 살펴보기
# + [markdown] id="Fjj86LoLv3Wl"
# ### 2-1) 기본 정보 탐색
# + [markdown] id="BZdMSr7Hyv3C"
# ##### 데이터셋 기본 정보 탐색
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 282, "status": "ok", "timestamp": 1632447997300, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="yTRz57Qpvgl4" outputId="a95de910-6498-4cc9-9a65-9217881e335c"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1632447998515, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lJq9dP3qvlMg" outputId="736f11d4-92e0-4afa-f18b-8cf40c8f5e1e"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 444, "status": "ok", "timestamp": 1632448000623, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="K7SiIjlDwFGG" outputId="30eaec34-ab18-4206-e7b1-e6efb87a0c42"
df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 271, "status": "ok", "timestamp": 1632448003134, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="c51l1Bld6c15" outputId="2f347a68-056f-4793-88e7-da1602ce966d"
df['text'][0]
# + colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"elapsed": 268, "status": "ok", "timestamp": 1632448027695, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="u9MFm3kOfAJ_" outputId="79ab452b-fd91-4d98-b77a-87b0203a6065"
df['text'][100]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1, "status": "ok", "timestamp": 1632448005883, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="JjX8EUde6f-g" outputId="7e1e9f63-4996-4bdf-c6b1-7d92d80cec08"
len(df['text'].values.sum())
# + [markdown] id="7Ob0pe0oxI6Y"
# -----
# + [markdown] id="vtLhtuYwxI86"
# ## 3) 한국어 텍스트 데이터 전처리
# + [markdown] id="ZAFsj3ppxI_J"
# ### 3-0) konlpy 설치
# + id="__acxhPiw8X2"
# konlpy 0.5.2의 JVM 버그로 인해, 0.5.1 버전으로 install
# !pip install konlpy==0.5.1 jpype1 Jpype1-py3
# + [markdown] id="yzf3UKBOAgQn"
# ### 3-1) 정규표현식 적용
# + executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1632448084827, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="hE60OR0mxP0S"
import re
# 띄어쓰기 포함한 한글 받아서 넣기 / 띄어쓰기를 두번이상 하는 경우가 거의 없다고 판단
def apply_regular_expression(text):
hangul = re.compile('[^ ㄱ-ㅣ가-힣]') # 한글의 정규표현식
result = hangul.sub('', text)
return result
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 331, "status": "ok", "timestamp": 1632448112378, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="zBgiHxOW3KhY" outputId="73508b5c-bc8e-4ffc-b086-df9552441da1"
apply_regular_expression(df['text'][0])
# + colab={"base_uri": "https://localhost:8080/", "height": 52} executionInfo={"elapsed": 438, "status": "ok", "timestamp": 1632448179796, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="ySDH-32mfl5v" outputId="4a4acb45-becb-4767-bf0b-d3eae66af93d"
apply_regular_expression(df['text'][100])
# + [markdown] id="pghj3X8ZBaD5"
# -----
# + [markdown] id="4eAD9Zhs3ZjB"
# ### 3-2) 한국어 형태소분석 - 명사 단위
# + [markdown] id="huXcIK1D7eGI"
# ##### 명사 형태소 추출
# + executionInfo={"elapsed": 7853, "status": "ok", "timestamp": 1632448202074, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="ePWpmhzL7f0A"
from konlpy.tag import Okt
from collections import Counter
nouns_tagger = Okt()
nouns = nouns_tagger.nouns(apply_regular_expression(df['text'][0]))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 271, "status": "ok", "timestamp": 1632448248305, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="XMdNZNRZAvmh" outputId="ba789f9a-c371-4050-8949-8d26e71cadd6"
nouns
# + executionInfo={"elapsed": 11349, "status": "ok", "timestamp": 1632448288719, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="w4KTE9U5AvrJ"
# 전체 말뭉치(corpus)에서 명사 형태소 추출
nouns = nouns_tagger.nouns(apply_regular_expression("".join(df['text'].tolist())))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 19, "status": "ok", "timestamp": 1632448288720, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="Y5yywiItAvuK" outputId="33c2e2d5-a1b5-4218-a6b2-012d3f5161cb"
# 빈도 탐색
counter = Counter(nouns)
counter.most_common(10)
# + [markdown] id="wpr7mEG5A2At"
# ##### 한글자 명사 제거
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 266, "status": "ok", "timestamp": 1632448334335, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="fDZfri14Avw1" outputId="a135defb-541e-40c5-df06-3adca6c54202"
available_counter = Counter({x : counter[x] for x in counter if len(x) > 1}) # 한글자 이상의 단어만 출력
available_counter.most_common(10)
# + [markdown] id="QI0yqJ7PxS1q"
# -----
# + [markdown] id="Jbr6rURvBPJ6"
# ### 3-3) 불용어 사전
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 391, "status": "ok", "timestamp": 1632448376893, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="7ZCmuyS8BUU9" outputId="620b8f2c-3782-450c-975c-ca80241aee41"
# source - https://www.ranks.nl/stopwords/korean
stopwords = pd.read_csv("https://raw.githubusercontent.com/yoonkt200/FastCampusDataset/master/korean_stopwords.txt").values.tolist()
print(stopwords[:10])
# + executionInfo={"elapsed": 283, "status": "ok", "timestamp": 1632448390366, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="IJTLaAlwBUYE"
# 데이터 특징을 반영한 불용어 추가
jeju_hotel_stopwords = ['제주', '제주도', '호텔', '리뷰', '숙소', '여행', '트립']
for word in jeju_hotel_stopwords:
stopwords.append(word)
# + [markdown] id="qb9hCD7GBWOJ"
# -----
# + [markdown] id="6-XoxM4KBWRB"
# ### 3-4) Word Count
# + [markdown] id="0hNlz_-kCq3v"
# ##### BoW 벡터 생성
# + executionInfo={"elapsed": 9033, "status": "ok", "timestamp": 1632448534781, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="ae-mZ9HOCcX1"
from sklearn.feature_extraction.text import CountVectorizer
def text_cleaning(text):
hangul = re.compile('[^ ㄱ-ㅣ가-힣]')
result = hangul.sub('', text)
tagger = Okt()
nouns = nouns_tagger.nouns(result)
nouns = [x for x in nouns if len(x) > 1]
nouns = [x for x in nouns if x not in stopwords]
return nouns
vect = CountVectorizer(tokenizer = lambda x: text_cleaning(x))
bow_vect = vect.fit_transform(df['text'].tolist())
word_list = vect.get_feature_names()
count_list = bow_vect.toarray().sum(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 365, "status": "ok", "timestamp": 1632449173631, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="I7WlEPkKCca9" outputId="406cf8a1-e307-4a34-975a-ab9da20f653d"
word_list
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 259, "status": "ok", "timestamp": 1632448541882, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="6-FIjghaCcdb" outputId="d42b3916-16d7-44a8-fd30-c9cea5fd6915"
count_list
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 364, "status": "ok", "timestamp": 1632448545516, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="3od1nB2ECcf_" outputId="696a9fa9-3d6d-4133-a8f8-089566e28a0c"
bow_vect.shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1632448547995, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="H-jKjWBVCyAZ" outputId="36ccb75c-fa86-4724-83e3-350b5d277c7d"
bow_vect.toarray()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 258, "status": "ok", "timestamp": 1632448550027, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="YHf8jMXgCyDN" outputId="8d4d22fd-eaed-4378-b565-0a3aab433c5d"
bow_vect.toarray().sum(axis=0)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 255, "status": "ok", "timestamp": 1632448557211, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="7Oeolrt9CyFZ" outputId="c35c8494-c456-4efa-e656-64527bafb9da"
bow_vect.toarray().sum(axis=0).shape
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 290, "status": "ok", "timestamp": 1632449180552, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="M5oVFqzSCyIK" outputId="7f034551-fa03-49ed-e309-54177935f866"
word_count_dict = dict(zip(word_list, count_list))
word_count_dict
# + [markdown] id="tv8-qE6MC79d"
# -----
# + [markdown] id="WMwtBHzKC85L"
# ### 3-5) TF-IDF 적용
# + [markdown] id="d6kLDjZYDHs7"
# ##### TF-IDF 변환
# + executionInfo={"elapsed": 260, "status": "ok", "timestamp": 1632448586481, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="eE5AR1IzDCAD"
from sklearn.feature_extraction.text import TfidfTransformer
tfidf_vectorizer = TfidfTransformer()
tf_idf_vect = tfidf_vectorizer.fit_transform(bow_vect)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 261, "status": "ok", "timestamp": 1632448588372, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="bCq6b-3TDCC-" outputId="71ea89f1-f16c-432d-bfe1-4e80f36e53f0"
print(tf_idf_vect.shape)
print(tf_idf_vect[0])
# + [markdown] id="LAXZrfU8DRPx"
# ##### 벡터 : 단어 맵핑
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 262, "status": "ok", "timestamp": 1632448590959, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="jpOhHaKqDCFm" outputId="6358cb09-5034-4834-981c-bfe82301be5f"
invert_index_vectorizer = {v: k for k, v in vect.vocabulary_.items()}
print(str(invert_index_vectorizer)[:100]+'..')
# + [markdown] id="nvwPAdU2DA_v"
# -----
# + [markdown] id="3Nb-h1fM7nLc"
# ## 4) Logistic Regression 분류
# + [markdown] id="87E9Sz5g4YFE"
# ### 4-1) 데이터셋 생성
# + [markdown] id="dcVGBXHVDeIu"
# ##### Rating 데이터 이진으로 변환
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 262, "status": "ok", "timestamp": 1632448647379, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="PS27E0CT4h-7" outputId="9f6440d7-632b-470a-ace4-4f96b0f83562"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} executionInfo={"elapsed": 758, "status": "ok", "timestamp": 1632448650075, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lXvHnoTY96zZ" outputId="8dd49e86-fe19-4d50-806e-aac3b7c87286"
df.rating.hist()
# + executionInfo={"elapsed": 260, "status": "ok", "timestamp": 1632448667143, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lhjSTR4HDlXS"
def rating_to_label(rating):
if rating > 3:
return 1
else:
return 0
df['y'] = df['rating'].apply(lambda x: rating_to_label(x))
# + colab={"base_uri": "https://localhost:8080/", "height": 202} executionInfo={"elapsed": 294, "status": "ok", "timestamp": 1632448669220, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="fADwGYKLDlaU" outputId="6091e997-154c-46bf-89b3-dddf90800907"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1632448672286, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="FZi8MUaxDp2U" outputId="7490033a-e20e-4773-a8e7-2212495e6108"
df.y.value_counts()
# + [markdown] id="ocEMVqzM-MdE"
# -----
# + [markdown] id="dK64F8GZ-N8s"
#
# + [markdown] id="Pe7BOBrW-OAw"
# ### 4-2) 데이터셋 분리
# + executionInfo={"elapsed": 419, "status": "ok", "timestamp": 1632448716799, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="-Wxv-F3P-NEU"
from sklearn.model_selection import train_test_split
y = df['y']
x_train, x_test, y_train, y_test = train_test_split(tf_idf_vect, y, test_size=0.30)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1632448717221, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="rsDAFsCY-Unv" outputId="667c5278-39f9-4ed6-e768-aa0c2d06cce4"
print(x_train.shape)
print(x_test.shape)
# + [markdown] id="pMkSIg7JD7rj"
# -----
# + [markdown] id="lHGviAneD7vN"
# ### 4-3) 모델 학습
# + [markdown] id="kbez8JYXD7x-"
# ##### Logistic Regression 학습
# + executionInfo={"elapsed": 285, "status": "ok", "timestamp": 1632448732818, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="rx7Ojv4TEBq3"
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
# Train LR model
lr = LogisticRegression(random_state=0)
lr.fit(x_train, y_train)
# classifiacation predict
y_pred = lr.predict(x_test)
# + [markdown] id="eYOegz_7EKg8"
# ##### 분류 결과 평가
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1, "status": "ok", "timestamp": 1632448733188, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="lyLhjDwFEB8h" outputId="dff84761-753a-4243-cec6-4131a8ea1af7"
# classification result for test dataset
print("accuracy: %.2f" % accuracy_score(y_test, y_pred))
print("Precision : %.3f" % precision_score(y_test, y_pred))
print("Recall : %.3f" % recall_score(y_test, y_pred))
print("F1 : %.3f" % f1_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1, "status": "ok", "timestamp": 1632448733486, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="6gd3tmYXEJ32" outputId="77190ddc-a200-4d56-c5c2-4eab6e66f9a4"
from sklearn.metrics import confusion_matrix
# print confusion matrix
confmat = confusion_matrix(y_true=y_test, y_pred=y_pred)
print(confmat)
# + [markdown] id="UHt-LenyEOnp"
# -----
# + [markdown] id="N70AyTZDEOuz"
# ### 4-4) 샘플링 재조정
# + [markdown] id="3u1UonfVEWnR"
# ##### 1:1 Sampling
# + executionInfo={"elapsed": 268, "status": "ok", "timestamp": 1632448822545, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="3BYUcp08ER7-"
positive_random_idx = df[df['y']==1].sample(275, random_state=33).index.tolist()
negative_random_idx = df[df['y']==0].sample(275, random_state=33).index.tolist()
# + executionInfo={"elapsed": 1, "status": "ok", "timestamp": 1632448823611, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="HUTZegD7EVkc"
# dataset split to train/test
random_idx = positive_random_idx + negative_random_idx
X = tf_idf_vect[random_idx]
y = df['y'][random_idx]
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=33)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1632448823987, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="YHvy8WzGEaBy" outputId="f9472d9a-27e9-446e-a960-597e3005229b"
print(x_train.shape)
print(x_test.shape)
# + [markdown] id="TDPgGZJeEfSd"
# ##### 모델 재학습
# + executionInfo={"elapsed": 384, "status": "ok", "timestamp": 1632448840094, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="uFO_UsnCEbGB"
lr = LogisticRegression(random_state=0)
lr.fit(x_train, y_train)
y_pred = lr.predict(x_test)
# + [markdown] id="y3PqFXYAEjzQ"
# ##### 분류 결과 평가
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1632448840394, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="QKiAhKirEeOQ" outputId="3caf1d57-0953-45aa-a01b-2d39c7e3b008"
print("accuracy: %.2f" % accuracy_score(y_test, y_pred))
print("Precision : %.3f" % precision_score(y_test, y_pred))
print("Recall : %.3f" % recall_score(y_test, y_pred))
print("F1 : %.3f" % f1_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2, "status": "ok", "timestamp": 1632448841232, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="uMRrUbU0Ei-B" outputId="94f53c35-3ba6-45d9-a8d5-aad9a64e0672"
confmat = confusion_matrix(y_true=y_test, y_pred=y_pred)
print(confmat)
# + [markdown] id="GeQGRMLaFFLk"
# -----
# + [markdown] id="3Ob_KEwxFGvr"
# ## 5) 긍정/부정 키워드 분석
# + [markdown] id="HVYQEDe0FNG0"
# ##### Logistic Regression 모델의 coef 분석
# + colab={"base_uri": "https://localhost:8080/", "height": 500} executionInfo={"elapsed": 9256, "status": "ok", "timestamp": 1632448918462, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="D5rVvWCQElIa" outputId="e2f3159b-751e-4421-cda3-7d58d6b437a2"
# print logistic regression's coef
# 단어의 분포 확인
plt.rcParams['figure.figsize'] = [10, 8]
plt.bar(range(len(lr.coef_[0])), lr.coef_[0])
# + [markdown] id="MvLq3hSpFUO7"
# ##### 긍정/부정 키워드 출력
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 286, "status": "ok", "timestamp": 1632448934580, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="me2xwEybFTnD" outputId="0684233f-1067-42fd-ec18-e2a38c4eedbf"
# 긍정/부정키워드 5개씩
print(sorted(((value, index) for index, value in enumerate(lr.coef_[0])), reverse=True)[:5])
print(sorted(((value, index) for index, value in enumerate(lr.coef_[0])), reverse=True)[-5:])
# + id="yGxlW37CFbd7"
# 상위 5개 인덱스
coef_pos_index = sorted(((value, index) for index, value in enumerate(lr.coef_[0])), reverse=True)
coef_neg_index = sorted(((value, index) for index, value in enumerate(lr.coef_[0])), reverse=False)
coef_pos_index
# + executionInfo={"elapsed": 277, "status": "ok", "timestamp": 1632448982821, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="bQuDlc-GFbwJ"
invert_index_vectorizer = {v: k for k, v in vect.vocabulary_.items()}
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 285, "status": "ok", "timestamp": 1632449014941, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="eHkWYQM0FdNU" outputId="ad7fbc2b-8830-471f-cf82-81edec3ab9d3"
for coef in coef_pos_index[:15]:
print(invert_index_vectorizer[coef[1]], coef[0])
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1, "status": "ok", "timestamp": 1632449015829, "user": {"displayName": "\ud321\ud321", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "08351897761321616592"}, "user_tz": -540} id="O0gRzUtXFeLX" outputId="3718ebf5-5b31-4775-a68f-16b00077e5c2"
for coef in coef_neg_index[:15]:
print(invert_index_vectorizer[coef[1]], coef[0])
# + id="fF8BtpB_FfkB"
# + id="TDYsTQ6HZTyw"
| study/python/210924-Python-text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Human numbers
from fastai.text import *
bs=64
# ## Data
path = untar_data(URLs.HUMAN_NUMBERS)
path.ls()
def readnums(d): return [', '.join(o.strip() for o in open(path/d).readlines())]
train_txt = readnums('train.txt'); train_txt[0][:80]
valid_txt = readnums('valid.txt'); valid_txt[0][-80:]
# +
train = TextList(train_txt, path=path)
valid = TextList(valid_txt, path=path)
src = ItemLists(path=path, train=train, valid=valid).label_for_lm()
data = src.databunch(bs=bs)
# -
train[0].text[:80]
len(data.valid_ds[0][0].data)
data.bptt, len(data.valid_dl)
13017/70/bs
it = iter(data.valid_dl)
x1,y1 = next(it)
x2,y2 = next(it)
x3,y3 = next(it)
it.close()
x1.numel()+x2.numel()+x3.numel()
x1.shape,y1.shape
x2.shape,y2.shape
x1[:,0]
y1[:,0]
v = data.valid_ds.vocab
v.textify(x1[:,0])
v.textify(y1[:,0])
v.textify(x2[:,0])
v.textify(x3[:,0])
v.textify(x1[:,1])
v.textify(x2[:,1])
v.textify(x3[:,1])
v.textify(x3[:,-1])
data.show_batch(ds_type=DatasetType.Valid)
# ## Single fully connected model
data = src.databunch(bs=bs, bptt=3, max_len=0, p_bptt=1.)
x,y = data.one_batch()
x.shape,y.shape
nv = len(v.itos); nv
nh=64
def loss4(input,target): return F.cross_entropy(input, target[-1])
def acc4 (input,target): return accuracy(input, target[-1])
class Model0(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh) # green arrow
self.h_h = nn.Linear(nh,nh) # brown arrow
self.h_o = nn.Linear(nh,nv) # blue arrow
self.bn = nn.BatchNorm1d(nh)
def forward(self, x):
h = self.bn(F.relu(self.i_h(x[0])))
if x.shape[0]>1:
h = h + self.i_h(x[1])
h = self.bn(F.relu(self.h_h(h)))
if x.shape[0]>2:
h = h + self.i_h(x[2])
h = self.bn(F.relu(self.h_h(h)))
return self.h_o(h)
learn = Learner(data, Model0(), loss_func=loss4, metrics=acc4)
learn.fit_one_cycle(6, 1e-4)
# ## Same thing with a loop
class Model1(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh) # green arrow
self.h_h = nn.Linear(nh,nh) # brown arrow
self.h_o = nn.Linear(nh,nv) # blue arrow
self.bn = nn.BatchNorm1d(nh)
def forward(self, x):
h = torch.zeros(x.shape[1], nh).to(device=x.device)
for xi in x:
h = h + self.i_h(xi)
h = self.bn(F.relu(self.h_h(h)))
return self.h_o(h)
learn = Learner(data, Model1(), loss_func=loss4, metrics=acc4)
learn.fit_one_cycle(6, 1e-4)
# ## Multi fully connected model
data = src.databunch(bs=bs, bptt=20)
x,y = data.one_batch()
x.shape,y.shape
class Model2(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh)
self.h_h = nn.Linear(nh,nh)
self.h_o = nn.Linear(nh,nv)
self.bn = nn.BatchNorm1d(nh)
def forward(self, x):
h = torch.zeros(x.shape[1], nh).to(device=x.device)
res = []
for xi in x:
h = h + self.i_h(xi)
h = self.bn(F.relu(self.h_h(h)))
res.append(self.h_o(h))
return torch.stack(res)
learn = Learner(data, Model2(), metrics=accuracy)
learn.fit_one_cycle(10, 1e-4, pct_start=0.1)
# ## Maintain state
class Model3(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh)
self.h_h = nn.Linear(nh,nh)
self.h_o = nn.Linear(nh,nv)
self.bn = nn.BatchNorm1d(nh)
self.h = torch.zeros(x.shape[1], nh).cuda()
def forward(self, x):
res = []
h = self.h
for xi in x:
h = h + self.i_h(xi)
h = F.relu(self.h_h(h))
res.append(h)
self.h = h.detach()
res = torch.stack(res)
res = self.h_o(self.bn(res))
return res
learn = Learner(data, Model3(), metrics=accuracy)
learn.fit_one_cycle(20, 3e-3)
# ## nn.RNN
class Model4(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh)
self.rnn = nn.RNN(nh,nh)
self.h_o = nn.Linear(nh,nv)
self.bn = nn.BatchNorm1d(nh)
self.h = torch.zeros(1, x.shape[1], nh).cuda()
def forward(self, x):
res,h = self.rnn(self.i_h(x), self.h)
self.h = h.detach()
return self.h_o(self.bn(res))
learn = Learner(data, Model4(), metrics=accuracy)
learn.fit_one_cycle(20, 3e-3)
# ## 2-layer GRU
class Model5(nn.Module):
def __init__(self):
super().__init__()
self.i_h = nn.Embedding(nv,nh)
self.rnn = nn.GRU(nh,nh,2)
self.h_o = nn.Linear(nh,nv)
self.bn = nn.BatchNorm1d(nh)
self.h = torch.zeros(2, bs, nh).cuda()
def forward(self, x):
res,h = self.rnn(self.i_h(x), self.h)
self.h = h.detach()
return self.h_o(self.bn(res))
learn = Learner(data, Model5(), metrics=accuracy)
learn.fit_one_cycle(10, 1e-2)
# ## fin
| nbs/dl1/lesson7-human-numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Harmonic oscillator visualization
#
# version 02.0: based on pendulum code; v2 adds driving force curve
#
# * Created 12-Jan-2019 by <NAME> (<EMAIL>)
# * Last revised 16-Jan-2019 by <NAME> (<EMAIL>).
# %matplotlib inline
# +
import numpy as np
from scipy.integrate import ode, odeint
import matplotlib.pyplot as plt
# -
# ## Harmonic oscillator code
class Harmonic_oscillator():
"""
Harmonic oscillator class implements the parameters and differential
equation for a damped, driven, simple harmonic oscillator.
Parameters
----------
omega0 : float
natural frequency of the oscillator (e.g., \sqrt{k/m} if a spring)
beta : float
coefficient of damping term (with a factor of 2)
f_ext : float
amplitude of external force (this is f_0 in Taylor)
omega_ext : float
frequency of external force
phi_ext : float
phase angle for external force (taken to be zero in Taylor)
Methods
-------
dy_dt(y, t)
Returns the right side of the differential equation in vector y,
given time t and the corresponding value of y.
"""
def __init__(self,
omega0=1.,
beta=0.2,
f_ext=0.2,
omega_ext=0.689,
phi_ext=0.
):
self.omega0 = omega0
self.beta = beta
self.f_ext = f_ext
self.omega_ext = omega_ext
self.phi_ext = phi_ext
def dy_dt(self, y, t):
"""
This function returns the right-hand side of the diffeq as a vector:
[d theta/dt
d^2theta/dt^2]
Parameters
----------
y : float
A 2-component vector with y[0] = theta(t) and y[1] = dtheta/dt
t : float
time
Returns
-------
"""
F_ext = self.driving_force(t)
return [y[1], -self.omega0**2*y[0] - 2.*self.beta*y[1] + F_ext]
def driving_force(self, t):
"""Returns the driving force as a function of time t."""
return self.f_ext * np.cos(self.omega_ext*t + self.phi_ext)
def plot_y_vs_x(x, y, axis_labels=None, label=None, title=None, ax=None):
"""
Generic plotting function: return a figure axis with a plot of y vs. x.
"""
if ax is None: # if the axis object doesn't exist, make one
ax = plt.gca()
ax.plot(x, y, label=label)
if label is not None: # if a label if passed, show the legend
ax.legend()
if title is not None: # set a title if one if passed
ax.set_title(title)
if axis_labels is not None: # set x-axis and y-axis labels if passed
ax.set_xlabel(axis_labels[0])
ax.set_ylabel(axis_labels[1])
return ax
# ## Interface using ipywidgets with interactive_output
#
# We'll make a more elaborate interface so we can adjust all of the parameters.
# +
# Import the widgets we will use (add more if needed!)
import ipywidgets as widgets
from ipywidgets import HBox, VBox, Layout, Tab, Label, Checkbox
from ipywidgets import FloatSlider, IntSlider, Play, Dropdown, HTMLMath
from IPython.display import display
from time import sleep
# +
# This function generates the main output here, which is a grid of plots
def ho_plots(theta_vs_time_plot=True, theta_dot_vs_time_plot=True,
phase_space_plot=True, omega0=10.*np.pi, beta=np.pi/2.,
f_ext=1000., omega_ext=2.*np.pi, phi_ext=0.,
theta0=0.0, theta_dot0=0.0,
t_start=0, t_end=10, delta_t=0.01, plot_start=0,
font_size=18):
"""
Create plots for interactive_output according to the inputs.
Based on generating a Harmonic_oscillator instance and associated graphs.
Notes
-----
1. We generate a new Harmonic_oscillator instance every time *and*
solve the ODE every time, even if the only change is to parameters
like t_start and t_end. Should we care or is this just so
cheap to recalculate that it doesn't matter?
How could we structure this differently?
2. Should we delete ho1 at some point? E.g., is there a memory issue?
"""
# add delta_t it goes at least to t_end (probably should use linspace!)
t_pts = np.arange(t_start, t_end+delta_t, delta_t)
# Instantiate an oscillator with the passed (or default) values of the
# natural frequency omega0, damping beta, driving amplitude, frequency,
# and phase (f_ext, omega_ext, phi_ext).
ho1 = Harmonic_oscillator(omega0=omega0, beta=beta, f_ext=f_ext,
omega_ext=omega_ext, phi_ext=phi_ext)
y0 = [theta0, theta_dot0] # initial conditions for the oscillator ODE
# ODE solver parameters
abserr = 1.0e-8
relerr = 1.0e-6
# For now we solve with odeint; give more options in the future.
# The .T is for transpose, so that the matrix from odeint can changed
# to the correct form for reading off theta and theta_dot.
theta, theta_dot = odeint(ho1.dy_dt, y0, t_pts,
atol=abserr, rtol=relerr).T
# Update the common font size
plt.rcParams.update({'font.size': font_size})
# Labels for individual plot axes
theta_vs_time_labels = (r'$t$', r'$\theta$')
theta_dot_vs_time_labels = (r'$t$', r'$d\theta/dt$')
phase_space_labels = (r'$\theta$', r'$d\theta/dt$')
# Figure out how many rows and columns [one row for now]
plot_flags = [theta_vs_time_plot, theta_dot_vs_time_plot, phase_space_plot]
plot_num = plot_flags.count(True)
plot_rows = 1
figsize_rows = plot_rows*6
plot_cols = plot_num
figsize_cols = min(plot_cols*8, 16) # at most 16
# Make the plot!
fig = plt.figure(figsize=(figsize_cols,figsize_rows))
, axes = plt.subplots(plot_rows, plot_cols,
# finds nearest index to plot_start in t_pts array
start_index = (np.fabs(t_pts-plot_start)).argmin()
next_axis = 1 # keep track of the axis number
if theta_vs_time_plot:
ax_theta = fig.add_subplot(plot_rows, plot_cols, next_axis)
plot_y_vs_x(t_pts, theta, axis_labels=theta_vs_time_labels,
label='oscillator', title=r'$\theta$ vs. time',
ax=ax_theta)
# add a line where the phase space plot starts
ax_theta.axvline(t_pts[start_index], lw=3, color='red')
next_axis += 1
if theta_dot_vs_time_plot:
ax_theta_dot = fig.add_subplot(plot_rows, plot_cols, next_axis)
plot_y_vs_x(t_pts, theta_dot, axis_labels=theta_dot_vs_time_labels,
label='oscillator', title=r'$d\theta/dt$ vs. time',
ax=ax_theta_dot)
# add a line where the phase space plot starts
ax_theta_dot.axvline(t_pts[start_index], lw=3, color='red')
next_axis += 1
if phase_space_plot:
ax_phase_space = fig.add_subplot(plot_rows, plot_cols, next_axis)
plot_y_vs_x(theta[start_index:-1], theta_dot[start_index:-1],
axis_labels=phase_space_labels, title='Phase space',
ax=ax_phase_space)
next_axis += 1
fig.tight_layout()
return fig
# +
# Widgets for the various inputs.
# For any widget, we can set continuous_update=False if we don't want the
# plots to shift until the selection is finished (particularly relevant for
# sliders).
# Widgets for the plot choice (plus a label out front)
plot_choice_w = Label(value='Which plots: ',layout=Layout(width='100px'))
def plot_choice_widget(on=True, plot_description=None):
"""Makes a Checkbox to select whether to show a plot."""
return Checkbox(value=on, description=plot_description,
disabled=False, indent=False, layout=Layout(width='150px'))
theta_vs_time_plot_w = plot_choice_widget(True, r'$\theta$ vs. time')
theta_dot_vs_time_plot_w = plot_choice_widget(False, r'$d\theta/dt$ vs. time')
phase_space_plot_w = plot_choice_widget(True, 'phase space')
# Widgets for the oscillator parameters (all use FloatSlider, so we made
# it a function)
def float_widget(value, min, max, step, description, format):
"""Makes a FloatSlider with the passed parameters and continuous_update
set to False."""
slider_border = Layout(border='solid 1.0px')
return FloatSlider(value=value,min=min,max=max,step=step,disabled=False,
description=description,continuous_update=False,
orientation='horizontal',layout=slider_border,
readout=True,readout_format=format)
omega0_w = float_widget(value=10.*np.pi, min=0.0, max=20.*np.pi, step=0.1,
description=r'natural $\omega_0$:', format='.2f')
beta_w = float_widget(value=np.pi/2., min=0.0, max=2.*np.pi, step=0.1,
description=r'damping $\beta$:', format='.2f')
f_ext_w = float_widget(value=1000., min=0.0, max=2000., step=1.,
description=r'drive $f_{\rm ext}$:', format='.0f')
omega_ext_w = float_widget(value=2.*np.pi, min=0.0, max=6.*np.pi, step=0.1,
description=r'freq. $\omega_{\rm ext}$:', format='.2f')
phi_ext_w = float_widget(value=0.0, min=0, max=2.*np.pi, step=0.1,
description=r'phase $\phi_{\rm ext}$:', format='.1f')
# Widgets for the initial conditions
theta0_w = float_widget(value=0.0, min=0., max=2.*np.pi, step=0.1,
description=r'$\theta_0$:', format='.1f')
theta_dot0_w = float_widget(value=0.0, min=-100., max=100., step=1.,
description=r'$(d\theta/dt)_0$:', format='.1f')
# Widgets for the plotting parameters
t_start_w = float_widget(value=0., min=0., max=10., step=1.,
description='t start:', format='.1f')
t_end_w = float_widget(value=5., min=0., max=20., step=1.,
description='t end:', format='.1f')
delta_t_w = float_widget(value=0.001, min=0.001, max=0.1, step=0.001,
description='delta t:', format='.3f')
plot_start_w = float_widget(value=0., min=0., max=20., step=1.,
description='start plotting:', format='.1f')
# Widgets for the styling parameters
font_size_w = Dropdown(options=['12', '16', '18', '20', '24'], value='18',
description='Font size:',disabled=False,
continuous_update=False,layout=Layout(width='140px'))
############## Begin: Explicit callback functions #######################
# Make sure that t_end is at least t_start + 10
def update_t_end(*args):
if t_end_w.value < t_start_w.value:
t_end_w.value = t_start_w.value + 10
t_end_w.observe(update_t_end, 'value')
t_start_w.observe(update_t_end, 'value')
# Make sure that plot_start is at least t_start and less than t_end
def update_plot_start(*args):
if plot_start_w.value < t_start_w.value:
plot_start_w.value = t_start_w.value
if plot_start_w.value > t_end_w.value:
plot_start_w.value = t_end_w.value
plot_start_w.observe(update_plot_start, 'value')
t_start_w.observe(update_plot_start, 'value')
t_end_w.observe(update_plot_start, 'value')
############## End: Explicit callback functions #######################
# Set up the interactive_output widget
plot_out = widgets.interactive_output(ho_plots,
dict(
theta_vs_time_plot=theta_vs_time_plot_w,
theta_dot_vs_time_plot=theta_dot_vs_time_plot_w,
phase_space_plot=phase_space_plot_w,
omega0=omega0_w,
beta=beta_w,
f_ext=f_ext_w,
omega_ext=omega_ext_w,
phi_ext=phi_ext_w,
theta0=theta0_w,
theta_dot0=theta_dot0_w,
t_start=t_start_w,
t_end=t_end_w,
delta_t=delta_t_w,
plot_start=plot_start_w,
font_size=font_size_w)
)
# Now do some manual layout, where we can put the plot anywhere using plot_out
hbox1 = HBox([plot_choice_w, theta_vs_time_plot_w, theta_dot_vs_time_plot_w,
phase_space_plot_w]) # choice of what plots to show
hbox2 = HBox([omega0_w, f_ext_w, omega_ext_w, phi_ext_w]) # external driving parameters
hbox3 = HBox([theta0_w, theta_dot0_w, beta_w]) # initial conditions and damping
hbox4 = HBox([t_start_w, t_end_w, delta_t_w, plot_start_w]) # time and plot ranges
hbox5 = HBox([font_size_w]) # font size
# We'll set up Tabs to organize the controls. The Tab contents are declared
# as tab0, tab1, ... (probably should make this a list?) and the overall Tab
# is called tab (so its children are tab0, tab1, ...).
tab_height = '70px' # Fixed minimum height for all tabs. Specify another way?
tab0 = VBox([hbox2, hbox3], layout=Layout(min_height=tab_height))
tab1 = VBox([hbox1, hbox4], layout=Layout(min_height=tab_height))
tab2 = VBox([hbox5], layout=Layout(min_height=tab_height))
tab = Tab(children=[tab0, tab1, tab2])
tab.set_title(0, 'Physics')
tab.set_title(1, 'Plotting')
tab.set_title(2, 'Styling')
# Release the Kraken!
vbox2 = VBox([tab, plot_out])
display(vbox2)
# -
| 2020_week_2/Harmonic_oscillator_visualization_v01.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tse]
# language: python
# name: conda-env-tse-py
# ---
# +
# default_exp tokenizers
# -
# ### Explore Tokenizers
#
# >Exploring tokenizers offered and used by transformers models
#
# +
#export
import tokenizers; print(f"tokenizers: {tokenizers.__version__}")
import fastai; print(f"fastai: {fastai.__version__}")
from fastai.text import *
from tse.preprocessing import *
# -
# Preprocess
train_df = pd.read_csv("../data/train.csv").dropna().reset_index(drop=True)
test_df = pd.read_csv("../data/test.csv")
strip_text(train_df, "text")
strip_text(train_df, "selected_text")
strip_text(test_df, "text")
replace_whitespace(train_df, "text")
replace_whitespace(train_df, "selected_text")
replace_whitespace(test_df, "text")
replace_URLs(train_df, "text")
replace_URLs(train_df, "selected_text")
replace_URLs(test_df, "text")
replace_user(train_df, "text")
replace_user(train_df, "selected_text")
replace_user(test_df, "text")
is_wrong = train_df.apply(lambda o: is_wrong_selection(o['text'], o['selected_text']), 1)
train_df = train_df[~is_wrong].reset_index(drop=True)
list(train_df['text'])
# ### Tokenizers
#
# `tokenizers==0.7.0` package. https://twitter.com/moi_anthony/status/1251193880302759938
#
# There are many models offered and each of these pretrained models use a specific tokenizer.
#
# Python binding [docs](https://github.com/huggingface/tokenizers/blob/71b7830d1b4b633e05cfc2b5271f08a215db2a04/bindings/python/tokenizers/__init__.pyi#L330-L337)
# 
# +
#export
from tokenizers import Tokenizer, AddedToken, pre_tokenizers, decoders, processors
from tokenizers.models import BPE
from tokenizers.normalizers import BertNormalizer, Lowercase
def init_roberta_tokenizer(vocab_file, merges_file, max_length=192, do_lower_case=True):
roberta = Tokenizer(BPE(vocab_file, merges_file))
if do_lower_case: roberta.normalizer = Lowercase()
roberta.pre_tokenizer = pre_tokenizers.ByteLevel()
roberta.decoder = decoders.ByteLevel()
roberta.enable_padding(pad_id=roberta.token_to_id("<pad>"),
pad_token="<pad>",
max_length=max_length)
roberta.enable_truncation(max_length=max_length, strategy="only_second")
roberta.add_special_tokens([
AddedToken("<mask>", lstrip=True),
"<s>",
"</s>"
])
roberta.post_processor = processors.RobertaProcessing(
("</s>", roberta.token_to_id("</s>")),
("<s>", roberta.token_to_id("<s>")),
)
roberta.pad_token_id = roberta.token_to_id("<pad>")
roberta.eos_token_id = roberta.token_to_id("</s>")
roberta.bos_token_id = roberta.token_to_id("<s>")
roberta.unk_token_id = roberta.token_to_id("<unk>")
roberta.mask_token_id = roberta.token_to_id("<mask>")
return roberta
# -
tokenizer = init_roberta_tokenizer("../roberta-base/vocab.json", "../roberta-base/merges.txt")
tokenizer.bos_token_id, tokenizer.pad_token_id, tokenizer.eos_token_id, tokenizer.unk_token_id, tokenizer.mask_token_id
train_inp = list(tuple(zip(train_df.sentiment, train_df.text)))
test_inp = list(tuple(zip(train_df.sentiment, test_df.text)))
train_inp[:3]
# %time
train_outputs = tokenizer.encode_batch(train_inp)
test_outputs = tokenizer.encode_batch(test_inp)
# +
# trn_npads = [sum(array(o.tokens) == '<pad>') for o in train_outputs]
# test_npads = [sum(array(o.tokens) == '<pad>') for o in test_outputs]
# min(trn_npads), min(test_npads) == (195, 227)
# -
output = train_outputs[0]
input_ids = output.ids
attention_mask = output.attention_mask
token_type_ids = output.type_ids
offsets = output.offsets
offsets[0]
# ### export
from nbdev.export import notebook2script
notebook2script()
| nbs/02-tokenizers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D1_RealNeurons/student/W3D1_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="GWwMJ-Wpdz_q"
# # Neuromatch Academy: Week 3, Day 1, Tutorial 2
# # Real Neurons: Effects of Input Correlation
# __Content creators:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
# + [markdown] colab_type="text" id="foBnMV3PS9tN"
# ---
# # Tutorial Objectives
# In this tutorial, we will use the leaky integrate-and-fire (LIF) neuron model (see Tutorial 1) to study how they transform input correlations to output properties (transfer of correlations). In particular, we are going to write a few lines of code to:
#
# - inject correlated GWN in a pair of neurons
#
# - measure correlations between the spiking activity of the two neurons
#
# - study how the transfer of correlation depends on the statistics of the input, i.e. mean and standard deviation.
# + [markdown] colab_type="text" id="aeLO8mtpeGQw"
# ---
# # Setup
# + cellView="both" colab={} colab_type="code" id="bQ1Ue4PZd6ur"
# Import libraries
import matplotlib.pyplot as plt
import numpy as np
import time
# + cellView="form" colab={} colab_type="code" id="_tDBYa7uTn0c"
# @title Figure Settings
import ipywidgets as widgets # interactive display
# %config InlineBackend.figure_format = 'retina'
# use NMA plot style
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
my_layout = widgets.Layout()
# + cellView="form" colab={} colab_type="code" id="nktgc1MmQocS"
# @title Helper functions
def default_pars(**kwargs):
pars = {}
### typical neuron parameters###
pars['V_th'] = -55. # spike threshold [mV]
pars['V_reset'] = -75. # reset potential [mV]
pars['tau_m'] = 10. # membrane time constant [ms]
pars['g_L'] = 10. # leak conductance [nS]
pars['V_init'] = -75. # initial potential [mV]
pars['V_L'] = -75. # leak reversal potential [mV]
pars['tref'] = 2. # refractory time (ms)
### simulation parameters ###
pars['T'] = 400. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
### external parameters if any ###
for k in kwargs:
pars[k] = kwargs[k]
pars['range_t'] = np.arange(0, pars['T'], pars['dt']) # Vector of discretized
# time points [ms]
return pars
def run_LIF(pars, Iinj):
"""
Simulate the LIF dynamics with external input current
Args:
pars : parameter dictionary
Iinj : input current [pA]. The injected current here can be a value or an array
Returns:
rec_spikes : spike times
rec_v : mebrane potential
"""
# Set parameters
V_th, V_reset = pars['V_th'], pars['V_reset']
tau_m, g_L = pars['tau_m'], pars['g_L']
V_init, V_L = pars['V_init'], pars['V_L']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tref = pars['tref']
# Initialize voltage and current
v = np.zeros(Lt)
v[0] = V_init
Iinj = Iinj * np.ones(Lt)
tr = 0.
# simulate the LIF dynamics
rec_spikes = [] # record spike times
for it in range(Lt - 1):
if tr > 0:
v[it] = V_reset
tr = tr - 1
elif v[it] >= V_th: # reset voltage and record spike event
rec_spikes.append(it)
v[it] = V_reset
tr = tref / dt
# calculate the increment of the membrane potential
dv = (-(v[it] - V_L) + Iinj[it] / g_L) * (dt / tau_m)
# update the membrane potential
v[it + 1] = v[it] + dv
rec_spikes = np.array(rec_spikes) * dt
return v, rec_spikes
def my_GWN(pars, sig, myseed=False):
"""
Function that calculates Gaussian white noise inputs
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
myseed : random seed. int or boolean
the same seed will give the same random number sequence
Returns:
I : Gaussian white noise input
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Set random seed. You can fix the seed of the random number generator so
# that the results are reliable however, when you want to generate multiple
# realization make sure that you change the seed for each new realization
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# generate GWN
# we divide here by 1000 to convert units to sec.
I_GWN = sig * np.random.randn(Lt) * np.sqrt(pars['tau_m'] / dt)
return I_GWN
def Poisson_generator(pars, rate, n, myseed=False):
"""
Generates poisson trains
Args:
pars : parameter dictionary
rate : noise amplitute [Hz]
n : number of Poisson trains
myseed : random seed. int or boolean
Returns:
pre_spike_train : spike train matrix, ith row represents whether
there is a spike in ith spike train over time
(1 if spike, 0 otherwise)
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# generate uniformly distributed random variables
u_rand = np.random.rand(n, Lt)
# generate Poisson train
poisson_train = 1. * (u_rand < rate * (dt / 1000.))
return poisson_train
def example_plot_myCC():
pars = default_pars(T=50000, dt=.1)
c = np.arange(10) * 0.1
r12 = np.zeros(10)
for i in range(10):
I1gL, I2gL = correlate_input(pars, mu=20.0, sig=7.5, c=c[i])
r12[i] = my_CC(I1gL, I2gL)
plt.figure()
plt.plot(c, r12, 'bo', alpha=0.7, label='Simulation', zorder=2)
plt.plot([-0.05, 0.95], [-0.05, 0.95], 'k--', label='y=x',
dashes=(2, 2), zorder=1)
plt.xlabel('True CC')
plt.ylabel('Sample CC')
plt.legend(loc='best')
def LIF_output_cc(pars, mu, sig, c, bin_size, n_trials=20):
""" Simulates two LIF neurons with correlated input and computes output correlation
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
c : correlation coefficient ~[0, 1]
bin_size : bin size used for time series
n_trials : total simulation trials
Returns:
r : output corr. coe.
sp_rate : spike rate
sp1 : spike times of neuron 1 in the last trial
sp2 : spike times of neuron 2 in the last trial
"""
r12 = np.zeros(n_trials)
sp_rate = np.zeros(n_trials)
for i_trial in range(n_trials):
I1gL, I2gL = correlate_input(pars, mu, sig, c)
_, sp1 = run_LIF(pars, pars['g_L'] * I1gL)
_, sp2 = run_LIF(pars, pars['g_L'] * I2gL)
my_bin = np.arange(0, pars['T'], bin_size)
sp1_count, _ = np.histogram(sp1, bins=my_bin)
sp2_count, _ = np.histogram(sp2, bins=my_bin)
r12[i_trial] = my_CC(sp1_count[::20], sp2_count[::20])
sp_rate[i_trial] = len(sp1) / pars['T'] * 1000.
return r12.mean(), sp_rate.mean(), sp1, sp2
def plot_c_r_LIF(c, r, mycolor, mylabel):
z = np.polyfit(c, r, deg=1)
c_range = np.array([c.min() - 0.05, c.max() + 0.05])
plt.plot(c, r, 'o', color=mycolor, alpha=0.7, label=mylabel, zorder=2)
plt.plot(c_range, z[0] * c_range + z[1], color=mycolor, zorder=1)
# + [markdown] colab_type="text" id="ZjA3lyb3REcK"
# The helper function contains the:
#
# - Parameter dictionary: `default_pars( **kwargs)`
# - LIF simulator: `run_LIF`
# - Gaussian white noise generator: `my_GWN(pars, sig, myseed=False)`
# - Poisson type spike train generator: `Poisson_generator(pars, rate, n, myseed=False)`
# - Two LIF neurons with correlated inputs simulator: `LIF_output_cc(pars, mu, sig, c, bin_size, n_trials=20)`
# - Some additional plotting utilities
# + [markdown] colab_type="text" id="L66i6gE6Tdll"
# ---
# # Section 1: Correlations (Synchrony)
# Correlation or synchrony in neuronal activity can be described for any readout of brain activity. Here, we are concerned with the spiking activity of neurons.
#
# In the simplest way, correlation/synchrony refers to coincident spiking of neurons, i.e., when two neurons spike together, they are firing in **synchrony** or are **correlated**. Neurons can be synchronous in their instantaneous activity, i.e., they spike together with some probability. However, it is also possible that spiking of a neuron at time $t$ is correlated with the spikes of another neuron with a delay (time-delayed synchrony).
#
# ## Origin of synchronous neuronal activity:
# - Common inputs, i.e., two neurons are receiving input from the same sources. The degree of correlation of the shared inputs is proportional to their output correlation.
# - Pooling from the same sources. Neurons do not share the same input neurons but are receiving inputs from neurons which themselves are correlated.
# - Neurons are connected to each other (uni- or bi-directionally): This will only give rise to time-delayed synchrony. Neurons could also be connected via gap-junctions.
# - Neurons have similar parameters and initial conditions.
#
# ## Implications of synchrony
# When neurons spike together, they can have a stronger impact on downstream neurons. Synapses in the brain are sensitive to the temporal correlations (i.e., delay) between pre- and postsynaptic activity, and this, in turn, can lead to the formation of functional neuronal networks - the basis of unsupervised learning (we will study some of these concepts in a forthcoming tutorial).
#
# Synchrony implies a reduction in the dimensionality of the system. In addition, correlations, in many cases, can impair the decoding of neuronal activity.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="T-2sPjeZTTmY" outputId="c595e911-f042-415b-c895-46fb6d1ab7b6"
# @title Video 1: Input & output correlations
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="nsAYFBcAkes", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
# + [markdown] colab_type="text" id="rekiswDRoOR6"
# ## How to study the emergence of correlations
#
# + [markdown] colab_type="text" id="Is8piETq2_J0"
# A simple model to study the emergence of correlations is to inject common inputs to a pair of neurons and measure the output correlation as a function of the fraction of common inputs.
#
# Here, we are going to investigate the transfer of correlations by computing the correlation coefficient of spike trains recorded from two unconnected LIF neurons, which received correlated inputs.
#
#
# The input current to LIF neuron $i$ $(i=1,2)$ is:
#
# \begin{equation}
# \frac{I_i}{g_L} =\mu_i + \sigma_i (\sqrt{1-c}\xi_i + \sqrt{c}\xi_c) \quad (1)
# \end{equation}
#
# where $\mu_i$ is the temporal average of the current. The Gaussian white noise $\xi_i$ is independent for each neuron, while $\xi_c$ is common to all neurons. The variable $c$ ($0\le c\le1$) controls the fraction of common and independent inputs. $\sigma_i$ shows the variance of the total input.
#
# So, first, we will generate correlated inputs.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 247} colab_type="code" id="ZLLRFa2YJOQD" outputId="dd69178b-4d57-433f-b754-3285809bfed6"
# @title
#@markdown Execute this cell to get a function for generating correlated GWN inputs
def correlate_input(pars, mu=20., sig=7.5, c=0.3):
"""
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
c. : correlation coefficient ~[0, 1]
Returns:
I1gL, I2gL : two correlated inputs with corr. coe. c
"""
# generate Gaussian whute noise xi_1, xi_2, xi_c
xi_1 = my_GWN(pars, sig)
xi_2 = my_GWN(pars, sig)
xi_c = my_GWN(pars, sig)
# Generate two correlated inputs by Equation. (1)
I1gL = mu + np.sqrt(1. - c) * xi_1 + np.sqrt(c) * xi_c
I2gL = mu + np.sqrt(1. - c) * xi_2 + np.sqrt(c) * xi_c
return I1gL, I2gL
print(help(correlate_input))
# + [markdown] colab_type="text" id="q8CfqV7g4XAG"
# ### Exercise 1: Compute the correlation
#
# The _sample correlation coefficient_ between two input currents $I_i$ and $I_j$ is defined as the sample covariance of $I_i$ and $I_j$ divided by the square root of the sample variance of $I_i$ multiplied with the square root of the sample variance of $I_j$. In equation form:
#
# \begin{align}
# r_{ij} &= \frac{cov(I_i, I_j)}{\sqrt{var(I_i)} \sqrt{var(I_j)}}\\
# cov(I_i, I_j) &= \sum_{k=1}^L (I_i^k -\bar{I}_i)(I_j^k -\bar{I}_j) \\
# var(I_i) &= \sum_{k=1}^L (I_i^k -\bar{I}_i)^2
# \end{align}
#
# where $\bar{I}_i$ is the sample mean, k is the time bin, and L is the length of $I$. This means that $I_i^k$ is current i at time $k\cdot dt$. Note that the equations above are not accurate for sample covariances and variances as they should be additionally divided by L-1 - we have dropped this term because it cancels out in the sample correlation coefficient formula.
#
# The _sample correlation coefficient_ may also be referred to as the _sample Pearson correlation coefficient_. Here, is a beautiful paper that explains multiple ways to calculate and understand correlations [Rodgers and Nicewander 1988](https://www.stat.berkeley.edu/~rabbee/correlation.pdf).
#
# In this exercise, we will create a function, `my_CC` to compute the sample correlation coefficient between two time series. Note that while we introduced this computation here in the context of input currents, the sample correlation coefficient is used to compute the correlation between any two time series - we will use it later on binned spike trains.
# + colab={} colab_type="code" id="cNRakP99Hlqr"
def my_CC(i, j):
"""
Args:
i, j : two time series with the same length
Returns:
rij : correlation coefficient
"""
########################################################################
## TODO for students: compute rxy, then remove the NotImplementedError #
# Tip1: array([a1, a2, a3])*array([b1, b2, b3]) = array([a1*b1, a2*b2, a3*b3])
# Tip2: np.sum(array([a1, a2, a3])) = a1+a2+a3
# Tip3: square root, np.sqrt()
# Fill out function and remove
raise NotImplementedError("Student exercise: compute the sample correlation coefficient")
########################################################################
# Calculate the covariance of i and j
cov = ...
# Calculate the variance of i
var_i = ...
# Calculate the variance of j
var_j = ...
# Calculate the correlation coefficient
rij = ...
return rij
# Uncomment the line after completing the my_CC function
# example_plot_myCC()
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 431} colab_type="text" id="sbvh6eyCzwgl" outputId="20776a5e-d19e-4157-9261-e40a319017f5"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_RealNeurons/solutions/W3D1_Tutorial2_Solution_03e44bdc.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D1_RealNeurons/static/W3D1_Tutorial2_Solution_03e44bdc_0.png>
#
#
# + [markdown] colab_type="text" id="SgKLPRqwM5_z"
# ### Exercise 2: Measure the correlation between spike trains
#
# After recording the spike times of the two neurons, how can we estimate their correlation coefficient?
#
# In order to find this, we need to bin the spike times and obtain two time series. Each data point in the time series is the number of spikes in the corresponding time bin. You can use `np.histogram()` to bin the spike times.
#
# Complete the code below to bin the spike times and calculate the correlation coefficient for two Poisson spike trains. Note that `c` here is the ground-truth correlation coefficient that we define.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 247} colab_type="code" id="U9MU6wgtPpuU" outputId="5c73ae5e-d946-4112-cff2-fa8123f4c366"
# @title
# @markdown Execute this cell to get a function for generating correlated Poisson inputs (generate_corr_Poisson)
def generate_corr_Poisson(pars, poi_rate, c, myseed=False):
"""
function to generate correlated Poisson type spike trains
Args:
pars : parameter dictionary
poi_rate : rate of the Poisson train
c. : correlation coefficient ~[0, 1]
Returns:
sp1, sp2 : two correlated spike time trains with corr. coe. c
"""
range_t = pars['range_t']
mother_rate = poi_rate / c
mother_spike_train = Poisson_generator(pars, rate=mother_rate,
n=1, myseed=myseed)[0]
sp_mother = range_t[mother_spike_train > 0]
L_sp_mother = len(sp_mother)
sp_mother_id = np.arange(L_sp_mother)
L_sp_corr = int(L_sp_mother * c)
np.random.shuffle(sp_mother_id)
sp1 = np.sort(sp_mother[sp_mother_id[:L_sp_corr]])
np.random.shuffle(sp_mother_id)
sp2 = np.sort(sp_mother[sp_mother_id[:L_sp_corr]])
return sp1, sp2
print(help(generate_corr_Poisson))
# + colab={} colab_type="code" id="oEPDC5YGSXPc"
def corr_coeff_pairs(pars, rate, c, trials, bins):
"""
Calculate the correlation coefficient of two spike trains, for different
realizations
Args:
pars : parameter dictionary
rate : rate of poisson inputs
c : correlation coefficient ~ [0, 1]
trials : number of realizations
bins : vector with bins for time discretization
Returns:
r12 : correlation coefficient of a pair of inputs
"""
r12 = np.zeros(n_trials)
for i in range(n_trials):
##############################################################
## TODO for students: Use np.histogram to bin the spike time #
## e.g., sp1_count, _= np.histogram(...)
# Use my_CC() compute corr coe, compare with c
# Note that you can run multiple realizations and compute their r_12(diff_trials)
# with the defined function above. The average r_12 over trials can get close to c.
# Note: change seed to generate different input per trial
# Fill out function and remove
raise NotImplementedError("Student exercise: compute the correlation coefficient")
##############################################################
# Generate correlated Poisson inputs
sp1, sp2 = generate_corr_Poisson(pars, ..., ..., myseed=2020+i)
# Bin the spike times of the first input
sp1_count, _ = np.histogram(..., bins=...)
# Bin the spike times of the second input
sp2_count, _ = np.histogram(..., bins=...)
# Calculate the correlation coefficient
r12[i] = my_CC(..., ...)
return r12
poi_rate = 20.
c = 0.2 # set true correlation
pars = default_pars(T=10000)
# bin the spike time
bin_size = 20 # [ms]
my_bin = np.arange(0, pars['T'], bin_size)
n_trials = 100 # 100 realizations
# Uncomment to test your function
# r12 = corr_coeff_pairs(pars, rate=poi_rate, c=c, trials=n_trials, bins=my_bin)
# print(f'True corr coe = {c:.3f}')
# print(f'Simu corr coe = {r12.mean():.3f}')
# + [markdown] colab_type="text" id="1puI4yGdwktJ"
# Sample output
#
# ```
# True corr coe = 0.200
# Simu corr coe = 0.197
# ```
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="text" id="Mim54TJNSk9x" outputId="4bba72fd-a0b1-4ca6-f87f-57be663c929a"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_RealNeurons/solutions/W3D1_Tutorial2_Solution_38477225.py)
#
#
# + [markdown] colab_type="text" id="6_PutaGqPC9_"
# ---
# # Section 2: Investigate the effect of input correlation on the output correlation
#
# Now let's combine the aforementioned two procedures. We first generate the correlated inputs by Equation (1). Then we inject the correlated inputs $I_1, I_2$ into a pair of neurons and record their output spike times. We continue measuring the correlation between the output and
# investigate the relationship between the input correlation and the output correlation.
# + [markdown] colab_type="text" id="VpDGQSuQmUR0"
# ## Drive a neuron with correlated inputs and visualize its output
# In the following, you will inject correlated GWN in two neurons. You need to define the mean (`gwn_mean`), standard deviation (`gwn_std`), and input correlations (`c_in`).
#
# We will simulate $10$ trials to get a better estimate of the output correlation. Change the values in the following cell for the above variables (and then run the next cell) to explore how they impact the output correlation.
# + colab={} colab_type="code" id="3EPOvX4jFfzp"
# Play around with these parameters
pars = default_pars(T=80000, dt=1.) # get the parameters
c_in = 0.3 # set input correlation value
gwn_mean = 10.
gwn_std = 10.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 484} colab_type="code" id="TzWrUz1tmUmc" outputId="1b96ffca-d19d-4f0e-b370-44efb426fe9c"
# @title
# @markdown Do not forget to execute this cell to simulate the LIF
bin_size = 10. # ms
starttime = time.perf_counter() # time clock
r12_ss, sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=gwn_mean, sig=gwn_std, c=c_in,
bin_size=bin_size, n_trials=10)
# just the time counter
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
print(f"Input correlation = {c_in}")
print(f"Output correlation = {r12_ss}")
plt.figure(figsize=(12, 6))
plt.plot(sp1, np.ones(len(sp1)) * 1, '|', ms=20, label='neuron 1')
plt.plot(sp2, np.ones(len(sp2)) * 1.1, '|', ms=20, label='neuron 2')
plt.xlabel('time (ms)')
plt.ylabel('neuron id.')
plt.xlim(1000, 8000)
plt.ylim(0.9, 1.2)
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="nKQVWR60qqdi"
# ## Think!
# - Is the output correlation always smaller than the input correlation? If yes, why?
# - Should there be a systematic relationship between input and output correlations?
#
# You will explore these questions in the next figure but try to develop your own intuitions first!
# + [markdown] colab_type="text" id="3zpGLfwgqXgq"
# Lets vary `c_in` and plot the relationship between the `c_in` and output correlation. This might take some time depending on the number of trials.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 447} colab_type="code" id="-27REsp8rGy5" outputId="56ca7a39-23ee-4927-e89d-5a99afa0de7d"
#@title
#@markdown Don't forget to execute this cell!
pars = default_pars(T=80000, dt=1.) # get the parameters
bin_size = 10.
c_in = np.arange(0, 1.0, 0.1) # set the range for input CC
r12_ss = np.zeros(len(c_in)) # small mu, small sigma
starttime = time.perf_counter() # time clock
for ic in range(len(c_in)):
r12_ss[ic], sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=10)
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
plt.figure(figsize=(7, 6))
plot_c_r_LIF(c_in, r12_ss, mycolor='b', mylabel='Output CC')
plt.plot([c_in.min() - 0.05, c_in.max() + 0.05],
[c_in.min() - 0.05, c_in.max() + 0.05],
'k--', dashes=(2, 2), label='y=x')
plt.xlabel('Input CC')
plt.ylabel('Output CC')
plt.legend(loc='best', fontsize=16)
plt.show()
# + [markdown] colab={} colab_type="text" id="WHlfpa6qGxL3"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_RealNeurons/solutions/W3D1_Tutorial2_Solution_ecc2167a.py)
#
#
# + [markdown] colab_type="text" id="i4CqOWkIMAJ-"
# ---
# # Section 3: Correlation transfer function
# The above plot of input correlation vs. output correlation is called the __correlation transfer function__ of the neurons.
# + [markdown] colab_type="text" id="YEB9LWD0smps"
# ## Section 3.1: How do the mean and standard deviation of the GWN affect the correlation transfer function?
#
# The correlations transfer function appears to be linear. The above can be taken as the input/output transfer function of LIF neurons for correlations, instead of the transfer function for input/output firing rates as we had discussed in the previous tutorial (i.e., F-I curve).
#
# What would you expect to happen to the slope of the correlation transfer function if you vary the mean and/or the standard deviation of the GWN?
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 448} colab_type="code" id="kH9WSUy9dvUf" outputId="1abc589d-afeb-4c53-db46-33a309f7528a"
#@markdown Execute this cell to visualize correlation transfer functions
pars = default_pars(T=80000, dt=1.) # get the parameters
no_trial = 10
bin_size = 10.
c_in = np.arange(0., 1., 0.2) # set the range for input CC
r12_ss = np.zeros(len(c_in)) # small mu, small sigma
r12_ls = np.zeros(len(c_in)) # large mu, small sigma
r12_sl = np.zeros(len(c_in)) # small mu, large sigma
starttime = time.perf_counter() # time clock
for ic in range(len(c_in)):
r12_ss[ic], sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
r12_ls[ic], sp_ls, sp1, sp2 = LIF_output_cc(pars, mu=18.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
r12_sl[ic], sp_sl, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=20.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
plt.figure(figsize=(7, 6))
plot_c_r_LIF(c_in, r12_ss, mycolor='b', mylabel=r'Small $\mu$, small $\sigma$')
plot_c_r_LIF(c_in, r12_ls, mycolor='y', mylabel=r'Large $\mu$, small $\sigma$')
plot_c_r_LIF(c_in, r12_sl, mycolor='r', mylabel=r'Small $\mu$, large $\sigma$')
plt.plot([c_in.min() - 0.05, c_in.max() + 0.05],
[c_in.min() - 0.05, c_in.max() + 0.05],
'k--', dashes=(2, 2), label='y=x')
plt.xlabel('Input CC')
plt.ylabel('Output CC')
plt.legend(loc='best', fontsize=14)
plt.show()
# + [markdown] colab_type="text" id="jrzxU3RULDDI"
# ### Think!
# Why do both the mean and the standard deviation of the GWN affect the slope of the correlation transfer function?
# + [markdown] colab={} colab_type="text" id="i9XTRTC_HpWM"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_RealNeurons/solutions/W3D1_Tutorial2_Solution_bb194a8f.py)
#
#
# + [markdown] colab_type="text" id="67gsO37w-8Qj"
# ## Section 3.2: What is the rationale behind varying $\mu$ and $\sigma$?
# The mean and the variance of the synaptic current depends on the spike rate of a Poisson process. We can use [Campbell's theorem](https://en.wikipedia.org/wiki/Campbell%27s_theorem_(probability)) to estimate the mean and the variance of the synaptic current:
#
# \begin{align}
# \mu_{\rm syn} = \lambda J \int P(t) \\
# \sigma_{\rm syn} = \lambda J \int P(t)^2 dt\\
# \end{align}
#
# where $\lambda$ is the firing rate of the Poisson input, $J$ the amplitude of the postsynaptic current and $P(t)$ is the shape of the postsynaptic current as a function of time.
#
# Therefore, when we varied $\mu$ and/or $\sigma$ of the GWN, we mimicked a change in the input firing rate. Note that, if we change the firing rate, both $\mu$ and $\sigma$ will change simultaneously, not independently.
#
# Here, since we observe an effect of $\mu$ and $\sigma$ on correlation transfer, this implies that the input rate has an impact on the correlation transfer function.
#
# + [markdown] colab_type="text" id="3_dyceUAEzh8"
# ### Think!
#
# - What are the factors that would make output correlations smaller than input correlations? (Notice that the colored lines are below the black dashed line)
# - What does it mean for the correlation in the network?
# - Here we have studied the transfer of correlations by injecting GWN. But in the previous tutorial, we mentioned that GWN is unphysiological. Indeed, neurons receive colored noise (i.e., Shot noise or OU process). How do these results obtained from injection of GWN apply to the case where correlated spiking inputs are injected in the two LIFs? Will the results be the same or different?
#
# Reference
# - <NAME>, Jaime, et al. "Correlation between neural spike trains increases with firing rate." Nature (2007) (https://www.nature.com/articles/nature06028/)
#
# - <NAME>, <NAME>, <NAME>. Role of input correlations in shaping the variability and noise correlations of evoked activity in the neocortex. Journal of Neuroscience. 2015 Jun 3;35(22):8611-25. (https://www.jneurosci.org/content/35/22/8611)
# + [markdown] colab={} colab_type="text" id="bHPafQAfI2eT"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D1_RealNeurons/solutions/W3D1_Tutorial2_Solution_4575a736.py)
#
#
# + [markdown] colab_type="text" id="LJfdR6Xu7Vyi"
# ---
# # Summary
#
# In this tutorial, we studied how the input correlation of two LIF neurons is mapped to their output correlation. Specifically, we:
#
# - injected correlated GWN in a pair of neurons,
#
# - measured correlations between the spiking activity of the two neurons, and
#
# - studied how the transfer of correlation depends on the statistics of the input, i.e., mean and standard deviation.
#
# Here, we were concerned with zero time lag correlation. For this reason, we restricted estimation of correlation to instantaneous correlations. If you are interested in time-lagged correlation, then we should estimate the cross-correlogram of the spike trains and find out the dominant peak and area under the peak to get an estimate of output correlations.
#
# We leave this as a future to-do for you if you are interested.
# + [markdown] colab_type="text" id="6IdP4bt2NpB2"
# ---
# # Bonus 1: Example of a conductance-based LIF model
# Above, we have written code to generate correlated Poisson spike trains. You can write code to stimulate the LIF neuron with such correlated spike trains and study the correlation transfer function for spiking input and compare it to the correlation transfer function obtained by injecting correlated GWNs.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 354} colab_type="code" id="z7-8LaIWnAKY" outputId="99b6e02d-d979-441c-ea24-090c191d70df"
# @title Function to simulate conductance-based LIF
def run_LIF_cond(pars, I_inj, pre_spike_train_ex, pre_spike_train_in):
"""
conductance-based LIF dynamics
Args:
pars : parameter dictionary
I_inj : injected current [pA]. The injected current here can
be a value or an array
pre_spike_train_ex : spike train input from presynaptic excitatory neuron
pre_spike_train_in : spike train input from presynaptic inhibitory neuron
Returns:
rec_spikes : spike times
rec_v : mebrane potential
gE : postsynaptic excitatory conductance
gI : postsynaptic inhibitory conductance
"""
# Retrieve parameters
V_th, V_reset = pars['V_th'], pars['V_reset']
tau_m, g_L = pars['tau_m'], pars['g_L']
V_init, E_L = pars['V_init'], pars['E_L']
gE_bar, gI_bar = pars['gE_bar'], pars['gI_bar']
VE, VI = pars['VE'], pars['VI']
tau_syn_E, tau_syn_I = pars['tau_syn_E'], pars['tau_syn_I']
tref = pars['tref']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize
tr = 0.
v = np.zeros(Lt)
v[0] = V_init
gE = np.zeros(Lt)
gI = np.zeros(Lt)
Iinj = I_inj * np.ones(Lt) # ensure I has length Lt
if pre_spike_train_ex.max() == 0:
pre_spike_train_ex_total = np.zeros(Lt)
else:
pre_spike_train_ex_total = pre_spike_train_ex * np.ones(Lt)
if pre_spike_train_in.max() == 0:
pre_spike_train_in_total = np.zeros(Lt)
else:
pre_spike_train_in_total = pre_spike_train_in * np.ones(Lt)
# simulation
rec_spikes = [] # recording spike times
for it in range(Lt - 1):
if tr > 0:
v[it] = V_reset
tr = tr - 1
elif v[it] >= V_th: # reset voltage and record spike event
rec_spikes.append(it)
v[it] = V_reset
tr = tref / dt
# update the synaptic conductance
gE[it+1] = gE[it] - (dt / tau_syn_E) * gE[it] + gE_bar * pre_spike_train_ex_total[it + 1]
gI[it+1] = gI[it] - (dt / tau_syn_I) * gI[it] + gI_bar * pre_spike_train_in_total[it + 1]
# calculate the increment of the membrane potential
dv = (-(v[it] - E_L) - (gE[it + 1] / g_L) * (v[it] - VE) - \
(gI[it + 1] / g_L) * (v[it] - VI) + Iinj[it] / g_L) * (dt / tau_m)
# update membrane potential
v[it + 1] = v[it] + dv
rec_spikes = np.array(rec_spikes) * dt
return v, rec_spikes, gE, gI
print(help(run_LIF_cond))
# + [markdown] colab_type="text" id="DHs35UCoRpG4"
# ## Interactive Demo: Correlated spike input to an LIF neuron
#
# In the following you can explore what happens when the neurons receive correlated spiking input.
#
# You can vary the correlation between excitatory input spike trains. For simplicity, the correlation between inhibitory spike trains is set to 0.01.
#
# Vary both excitatory rate and correlation and see how the output correlation changes. Check if the results are qualitatively similar to what you observed previously when you varied the $\mu$ and $\sigma$.
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 634, "referenced_widgets": ["39a85ae1d21e45ca950d285232b751c9", "<KEY>", "ecdafef7540f49aca41eeee1d395f3d7", "ec0d46c3a8d040ae994d557ad1d24b8d", "e30c189bb30742548b8e3a52975c0292", "1c5e4c9c99274292b9c90e79fc686cad", "4e7d29683c494c09b2e74f090a640e66", "c94d14fd962143cb9111ebf4d80890a5", "e147deaa9c2a4907874822756522f42e", "3e49d2d1532b44388c0b1227146ede4f", "55d2e7c2a9d84ec6b3b22db6390e44ec"]} colab_type="code" id="bksg7PSWnGWV" outputId="0e48fd2c-63ae-4200-a8fc-3b93f5e5d39c"
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
pwc_ee=widgets.FloatSlider(0.3, min=0.05, max=0.99, step=0.01,
layout=my_layout),
exc_rate=widgets.FloatSlider(1e3, min=500., max=5e3, step=50.,
layout=my_layout),
inh_rate=widgets.FloatSlider(500., min=300., max=5e3, step=5.,
layout=my_layout),
)
def EI_isi_regularity(pwc_ee, exc_rate, inh_rate):
pars = default_pars(T=1000.)
# Add parameters
pars['V_th'] = -55. # spike threshold [mV]
pars['V_reset'] = -75. # reset potential [mV]
pars['tau_m'] = 10. # membrane time constant [ms]
pars['g_L'] = 10. # leak conductance [nS]
pars['V_init'] = -65. # initial potential [mV]
pars['E_L'] = -75. # leak reversal potential [mV]
pars['tref'] = 2. # refractory time (ms)
pars['gE_bar'] = 4.0 # [nS]
pars['VE'] = 0. # [mV] excitatory reversal potential
pars['tau_syn_E'] = 2. # [ms]
pars['gI_bar'] = 2.4 # [nS]
pars['VI'] = -80. # [mV] inhibitory reversal potential
pars['tau_syn_I'] = 5. # [ms]
my_bin = np.arange(0, pars['T']+pars['dt'], .1) # 20 [ms] bin-size
# exc_rate = 1e3
# inh_rate = 0.4e3
# pwc_ee = 0.3
pwc_ii = 0.01
# generate two correlated spike trains for excitatory input
sp1e, sp2e = generate_corr_Poisson(pars, exc_rate, pwc_ee)
sp1_spike_train_ex, _ = np.histogram(sp1e, bins=my_bin)
sp2_spike_train_ex, _ = np.histogram(sp2e, bins=my_bin)
# generate two uncorrelated spike trains for inhibitory input
sp1i, sp2i = generate_corr_Poisson(pars, inh_rate, pwc_ii)
sp1_spike_train_in, _ = np.histogram(sp1e, bins=my_bin)
sp2_spike_train_in, _ = np.histogram(sp2e, bins=my_bin)
v1, rec_spikes1, gE, gI = run_LIF_cond(pars, 0, sp1_spike_train_ex, sp1_spike_train_in)
v2, rec_spikes2, gE, gI = run_LIF_cond(pars, 0, sp2_spike_train_ex, sp2_spike_train_in)
# bin the spike time
bin_size = 20 # [ms]
my_bin = np.arange(0, pars['T'], bin_size)
spk_1, _ = np.histogram(rec_spikes1, bins=my_bin)
spk_2, _ = np.histogram(rec_spikes2, bins=my_bin)
r12 = my_CC(spk_1, spk_2)
print(f"Input correlation = {pwc_ee}")
print(f"Output correlation = {r12}")
plt.figure(figsize=(14, 7))
plt.subplot(211)
plt.plot(sp1e, np.ones(len(sp1e)) * 1, '|', ms=20,
label='Exc. input 1')
plt.plot(sp2e, np.ones(len(sp2e)) * 1.1, '|', ms=20,
label='Exc. input 2')
plt.plot(sp1e, np.ones(len(sp1e)) * 1.3, '|k', ms=20,
label='Inh. input 1')
plt.plot(sp2e, np.ones(len(sp2e)) * 1.4, '|k', ms=20,
label='Inh. input 2')
plt.ylim(0.9, 1.5)
plt.legend()
plt.ylabel('neuron id.')
plt.subplot(212)
plt.plot(pars['range_t'], v1, label='neuron 1')
plt.plot(pars['range_t'], v2, label='neuron 2')
plt.xlabel('time (ms)')
plt.ylabel('membrane voltage $V_{m}$')
plt.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="z7TXjCQa7CR7"
#
# Above, we are estimating the output correlation for one trial. You can modify the code to get a trial average of output correlations.
#
#
#
# + [markdown] colab_type="text" id="tbhWzUDm5hiY"
# ---
# # Bonus 2: Ensemble Response
#
# Finally, there is a short BONUS lecture video on the firing response of an ensemble of neurons to time-varying input. There are no associated coding exercises - just enjoy.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 537} colab_type="code" id="xkrEq6Jzn-lc" outputId="c6dd925f-eb98-45d5-a5f4-beb55b198202"
#@title Video 2 (Bonus): Response of ensemble of neurons to time-varying input
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="78_dWa4VOIo", width=854, height=480, fs=1)
print("Video available at https://youtube.com/watch?v=" + video.id)
video
| tutorials/W3D1_RealNeurons/student/W3D1_Tutorial2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow2_latest_p37]
# language: python
# name: conda-env-tensorflow2_latest_p37-py
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 184, "status": "ok", "timestamp": 1622586402138, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="v7eQSqoQuYgg" outputId="efd9f857-657f-4851-e1aa-fbe208c9538f"
# TODO: Enter the foldername in your Drive where you have saved the unzipped
# assignment folder, e.g. 'cs231n/assignments/assignment1/'
FOLDERNAME = 'home/ubuntu/Vision-Classifiers/Microsoft-Vision-Classifier/'
assert FOLDERNAME is not None, "[!] Enter the foldername."
# Now that we've mounted your Drive, this ensures that
# the Python interpreter of the Colab VM can load
# python files from within it.
import sys
sys.path.append('/home/ubuntu/Vision-Classifiers/Microsoft-Vision-Classifier')
# This downloads the CIFAR-10 dataset to your Drive
# if it doesn't already exist.
# %cd /$FOLDERNAME/flowers/
# !bash get_datasets.sh
# %cd /$FOLDERNAME
# + _uuid="4902ee7b7f4d66a42d59b971180bba213d0133c9" executionInfo={"elapsed": 816, "status": "ok", "timestamp": 1622586579831, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="z19bVm7o9zeu"
import warnings
warnings.filterwarnings('always')
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import style
import seaborn as sns
# %matplotlib inline
style.use('fivethirtyeight')
sns.set(style='whitegrid',color_codes=True)
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score,precision_score,recall_score,confusion_matrix,roc_curve,roc_auc_score
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import LabelEncoder
import tensorflow
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras import backend as K
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.optimizers import Adam,SGD,Adagrad,Adadelta,RMSprop
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.callbacks import ReduceLROnPlateau
from tensorflow.keras.layers import Dropout, Flatten,Activation
from tensorflow.keras.layers import Conv2D, MaxPooling2D, BatchNormalization
import tensorflow as tf
import random as rn
import cv2
import numpy as np
from tqdm import tqdm
import os
from random import shuffle
from zipfile import ZipFile
from PIL import Image
from tensorflow.keras.applications.vgg16 import VGG16
# + _uuid="346f2fdc1b809eb19158d411af4288d83c79f389" colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 142, "status": "ok", "timestamp": 1622586583796, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="0TvTNjLftcgt" outputId="e10e80a6-eeaa-4186-ef2e-f4b9400fd802"
import os
path = '/home/ubuntu/Vision-Classifiers/Microsoft-Vision-Classifier'
os.listdir('/home/ubuntu/Vision-Classifiers/Microsoft-Vision-Classifier/flowers')
# + _uuid="7b0c13e69deaf6449739ba2104bb6238be376f05" executionInfo={"elapsed": 104, "status": "ok", "timestamp": 1622586596499, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="abZS8dPk9ze1"
X=[]
Z=[]
IMG_SIZE=150
FLOWER_DAISY_DIR=f'{path}/flowers/daisy'
FLOWER_SUNFLOWER_DIR=f'{path}/flowers/sunflower'
FLOWER_TULIP_DIR=f'{path}/flowers/tulip'
FLOWER_DANDI_DIR=f'{path}/flowers/dandelion'
FLOWER_ROSE_DIR=f'{path}/flowers/rose'
weights_path= f'{path}/trans-learn-weights/vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5'
# + _uuid="1c467392d43ee29671c5498bd1feea5db5ef862d" executionInfo={"elapsed": 117, "status": "ok", "timestamp": 1622586599890, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="2qPgwo1d9ze4"
def assign_label(img,flower_type):
return flower_type
# + _uuid="861b4e251d97f7601a3bc2c3077183de4122e3d9" executionInfo={"elapsed": 162, "status": "ok", "timestamp": 1622586608535, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjaCV5m2H_od77fWW_j8T9E5_Nd_t82pjGdydJF=s64", "userId": "14896880010903859463"}, "user_tz": 420} id="vlY8PywM9ze7"
def make_train_data(flower_type,DIR):
for img in tqdm(os.listdir(DIR)):
label=assign_label(img,flower_type)
path = os.path.join(DIR,img)
img = cv2.imread(path,cv2.IMREAD_COLOR)
img = cv2.resize(img, (IMG_SIZE,IMG_SIZE))
X.append(np.array(img))
Z.append(str(label))
# + _uuid="04ee1723256e1836f56600bf041e67d1d8370314" colab={"base_uri": "https://localhost:8080/"} id="9hn_RjL29ze_" outputId="a158fa49-13f2-4e6f-9064-4f7ba1280fb0"
make_train_data('Daisy',FLOWER_DAISY_DIR)
print(len(X))
# + _uuid="d4cd8f4d88756086d747d72cad465433c7e1e0ff" id="6XZpRkLK9zfC"
make_train_data('Sunflower',FLOWER_SUNFLOWER_DIR)
print(len(X))
# + _uuid="9c86247c9d3651de45e3f6883df1453cf594b65b" id="YvJMmnNx9zfH"
make_train_data('Tulip',FLOWER_TULIP_DIR)
print(len(X))
# + _uuid="0ff3f7e5d2aae2b06fe87a49cf89007833d5c83d" id="KgYj8x-H9zfL"
make_train_data('Dandelion',FLOWER_DANDI_DIR)
print(len(X))
# + _uuid="8e6a148321afbfaa53dfc05a5d700c5a24fed336" id="3mtQH6Vg9zfQ"
make_train_data('Rose',FLOWER_ROSE_DIR)
print(len(X))
# + _uuid="1f006ba66f46d8c3355ecdd3b27b37bdea635944" id="_Gug0CHU9zfe"
le=LabelEncoder()
Y=le.fit_transform(Z)
Y=to_categorical(Y,5)
X=np.array(X)
X=X/255
# + _uuid="e9b04ed0a732e99941d6a347a14d66eb2cb4727b" id="4xogXfvm9zfg"
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.25,random_state=42)
# + _uuid="f0e2ae22e3bca8e3143d5b4d312460283b833878" id="S_nM3vLf9zfj"
np.random.seed(42)
rn.seed(42)
#tf.set_random_seed(42)
tf.random.set_seed(42)
# + _uuid="33b31b3df570b37c968f4c4e81ea24b48c864caa" id="SERVVhIgkxXV"
base_model=VGG16(include_top=False, weights=None,input_shape=(150,150,3), pooling='avg')
# + _uuid="9155aceff9ad2d42cbd5b9151389d2f15e8c7abd" id="iap7xkPgtcgz"
base_model.load_weights(weights_path)
# + _uuid="3bca9ce0ba74b4bec90b228d7805ce6d64e00062" id="xHOoktp-k4tw"
base_model.summary()
# + _uuid="61461000564043bcdd7dcafc2f99bf931eaecfa0" id="gGgMscM_eIYS"
model=Sequential()
model.add(base_model)
model.add(Dense(256,activation='relu'))
model.add(Dense(5,activation='softmax'))
# + _uuid="5859d726c293f373e7ee955bb931bd28f39db8d5" id="lH038cfsgkvZ"
datagen = ImageDataGenerator(
featurewise_center=False,
samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,
rotation_range=10,
zoom_range = 0.1,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True,
vertical_flip=False)
datagen.fit(x_train)
# + _uuid="0953cdcd6425381a3551aeb28af6c4b61654849d" id="B_6-fsX6gky4"
epochs=50
batch_size=128
red_lr=ReduceLROnPlateau(monitor='val_acc', factor=0.1, epsilon=0.0001, patience=2, verbose=1)
# + _uuid="8ad3563c162ea79510b967f9c2a53c0cd6fbc2d4" id="gPolpADLYv9p"
fig,ax=plt.subplots(5,2)
fig.set_size_inches(15,15)
for i in range(5):
for j in range (2):
l=rn.randint(0,len(Z))
ax[i,j].imshow(X[l])
ax[i,j].set_title('Flower: '+Z[l])
plt.tight_layout()
# + _uuid="c18c97375cbdde67228b22780bff5a0230598c71" id="IVxD9F-TeIdi"
model.summary()
# + _uuid="d2bc03ec5faf3e57a7504680d065a8ba30e403e3" id="CqU7SQXzTNof"
base_model.trainable=False
# + _uuid="b5fc67aea4c22dec0cf54adc0d475a9aa49e3dc7" id="Hx2arlHIeIhS"
model.compile(optimizer=Adam(lr=1e-4),loss='categorical_crossentropy',metrics=['accuracy'])
# + _uuid="2dc473930156c22f817d91349cd3aa233e623d65" id="cS9WpEjOeIjz"
gpus= tf.config.experimental.list_physical_devices('GPU')
History = model.fit_generator(datagen.flow(x_train,y_train, batch_size=batch_size),
epochs = 50, validation_data = (x_test,y_test),
verbose = 1, steps_per_epoch=x_train.shape[0] // batch_size)
# + [markdown] _uuid="7957e2a9e323de85d99f72cf1af44f078ccc5e11" id="8MRNUXM1tcg2"
# #### 3.5.2 ) FINE TUNING BY UNFREEZING THE LAST BLOCK OF VGG16
# + [markdown] _uuid="0deafdf0a87be6b94cf8e34e4e69e0c525afd1b0" id="ILVy9eWMtcg3"
# In this section I have done fine tuning. To see the effect of the fine tuning I have first unfreezed the last block of the VGG16 model and have set it to trainable.
# + _uuid="1efcc1bd4b8c809998052b4edf9bfd72a7467ad9" id="ZcwBRzYYPVx8"
for i in range (len(base_model.layers)):
print (i,base_model.layers[i])
for layer in base_model.layers[15:]:
layer.trainable=True
for layer in base_model.layers[0:15]:
layer.trainable=False
# + _uuid="67f268c689a28c44745f42658b1937d54ea2504c" id="U6-PRjikXAoK"
model.compile(optimizer=Adam(lr=1e-4),loss='categorical_crossentropy',metrics=['accuracy'])
# + _uuid="21fbf26cc0c2aab15e7c5903c2f92ef6a736c213" id="8HOQuHdyTfj9"
History = model.fit_generator(datagen.flow(x_train,y_train, batch_size=batch_size),
epochs = 50, validation_data = (x_test,y_test),
verbose = 1, steps_per_epoch=x_train.shape[0] // batch_size)
# + _uuid="9244d358ff5774c52799bd2009a9b27cce6ce00c" id="mPW6EdQ6zsVG"
for i in range (len(base_model.layers)):
print (i,base_model.layers[i])
for layer in base_model.layers[11:]:
layer.trainable=True
for layer in base_model.layers[0:11]:
layer.trainable=False
# + _uuid="82a58b8e1b1a8bf539377e53d3e9129f9dfa838c" id="8r2vkhzxzri4"
model.compile(optimizer=Adam(lr=1e-4),loss='categorical_crossentropy',metrics=['accuracy'])
# + _uuid="af35be1c41a5b83459a9ea7f1b544ec62fcee54f" id="Mu4-v7Tl0JnF"
History = model.fit_generator(datagen.flow(x_train,y_train, batch_size=batch_size),
epochs = 50, validation_data = (x_test,y_test),
verbose = 1, steps_per_epoch=x_train.shape[0] // batch_size)
# + _uuid="e662853bfaf2f12c0bba1a311a5498d3eea618bc" id="uVG9lwNqahjL"
plt.plot(History.history['loss'])
plt.plot(History.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epochs')
plt.legend(['train', 'test'])
plt.show()
# + _uuid="3a6beed19c34da7cbd224bc733b23bc81bc232ce" id="7_R7EV33ahsp"
plt.plot(History.history['accuracy'])
plt.plot(History.history['val_accuracy'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epochs')
plt.legend(['train', 'test'])
plt.show()
# + _uuid="adba788f8da8644c9e268bfe4063ecf8623e5e3b" id="pFbqDROC0Zqv"
History = model.fit_generator(datagen.flow(x_train,y_train, batch_size=batch_size),
epochs = 50, validation_data = (x_test,y_test),
verbose = 1, steps_per_epoch=x_train.shape[0] // batch_size)
# + _uuid="9f55b5b6a11cb49117b92960ff7925a8a03b836e" id="w44nLg9rZ7oY"
plt.plot(History.history['accuracy'])
plt.plot(History.history['val_accuracy'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epochs')
plt.legend(['train', 'test'])
plt.show()
# + _uuid="52eb4c22f88ffa062c479d1114ebb9c856eb14b6" id="HXtrIOjvtcg7"
| Flower-Classification-Transfer-Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:TwitterAPI] *
# language: python
# name: conda-env-TwitterAPI-py
# ---
import sqlite3 as sql
import pandas as pd
# connection
con = sql.connect('database.sqlite')
cursor = con.cursor()
for row in cursor.execute('''SELECT name FROM sqlite_master WHERE type='table';'''):
print(row)
country = pd.read_sql_query('SELECT * FROM Country', con)
league = pd.read_sql_query('SELECT * FROM League', con)
player = pd.read_sql_query('SELECT * FROM Player', con)
type(country)
player.info
height_over_150 = pd.read_sql_query('''SELECT * FROM player WHERE height > 150''', con)
height_over_150
| module2-sql-for-analysis/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tuples Data Structure
# ## Python Tuples (a,b)
# * Immutable - can NOT be changed
# * Use - passing data that does not need changing
# * Faster than list
# * "safer" than list
# * Can be key in dict unlike list
# * For Heterogeneous data - meaning mixing different data types(int,str,list et al) inside
# * https://docs.python.org/3/tutorial/datastructures.html#tuples-and-sequences
# * https://realpython.com/python-lists-tuples/
my_tuple = ("Valdis", "programmer", 45, 200, [1, 2, 3])
print(my_tuple)
my_values = 5, 10 # i do not even need ( )
type(my_values)
my_values[1] = 20 # cant mutate tuples with primitive values!
print(my_tuple[0])
# however we do have a list inside a tuple and we can modify that one!
print(my_tuple[-1])
my_tuple
my_list = my_tuple[-1] # so i create a shortcut/alias/reference/pointer to the last item in my tuple
# this item is a list
my_list
my_list.append(9000)
my_list
my_tuple # so tuple contained shortcuts to actual items, so list inside could be mutated IN PLACE only!
my_tuple.count(45)
my_tuple.index(45)
name, job, age, top_speed, favorite_list = my_tuple # tuple unpacking into individual variables
name, top_speed
favorite_list.append(5000)
my_tuple
print(job)
# small problem how to exchange values in two variables?
a = 2001
b = 2021
# how to exchange?
print(a,b)
# in older languages you would use temp variable
print(a,b)
t = a
a = b
b = t
print(a,b)
print(a,b)
a, b = b, a # in fact we could do a, b, c = c, b, a etc etc
print(a,b)
my_list = list(my_tuple)
my_tuple, my_list
my_list[1] = "teacher"
my_list
my_tuple = tuple(my_list)
my_tuple
# so if you need to change some value in a tuple you convert it list and then change and go back to tuple if need be
# +
# tuples also are used to return multiple values from functions which we will see in another lecture
# -
for item in my_tuple:
print(item) # each item has differrent type so not much we can do on each item without checking the type
for item in my_tuple:
print(item)
for item in my_tuple:
if type(item) == str: # if our iterables(lists,tuples) have multiple types inside we can check them
print(item.upper())
else:
print(f"Not a string {item}")
# +
# type(my_tuple)
# len(my_tuple)
# my_tuple[-1]
# my_tuple[-1][1]
# mini_3 = my_tuple[:3]
# print(mini_3, type(mini_3))
# my_tuple[::2]
# my_tuple[::-1]
# reverse_inner = tuple(el * 2 for el in my_tuple)
# my_list = []
# for el in my_tuple:
# if type(el) == int:
# my_list.append(1/el)
# else:
# my_list.append(el[::-1])
# my_rev_tuple = tuple(my_list)
# my_rev_tuple
# print(my_tuple)
# # my_tuple[1] = "scientist"
# my_list = list(my_tuple)
# print(my_list)
# my_list[1] = "scientist"
# new_tuple = tuple(my_list)
# print(new_tuple)
# t = () # empty tuple only question where would you use it?
# print(t, type(t))
# t = (1, 2, 55) # 2 or more elements
# print(t, type(t))
# t = (5,) # if you really need a tuple of one element
# my_tuple.count("programmer")
# new_tuple.count("programmer")
# new_tuple.index("scientist")
# my_tuple.index(45)
# a = 10
# b = 20
# print(a, b)
# # how to change them
# temp = a
# a = b
# b = temp
# print(a, b)
# # in Python the above is simpler!
# print(a, b)
# a, b = b, a # we can even change a,b,c,d = d,c,b,a and more
# print(a, b)
# (name, job, age, top_speed, favorite_list) = my_tuple # tuple unpacking
# print(name, job, age, top_speed, favorite_list)
# name is my_tuple[0]
# # tuple unpacking and using _ for values that we do need
# (name, job, _, top_speed, _) = my_tuple
# print(name, _) # so _ will have value of last unpacking
| core/Tuples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from Testdata.Builder import simulation_test, plotter, plot_cut
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Merton Jump Diffusion Model
#
#
# Create a random time series with marked jumps.
# Where **(v)** is the standard deviation of the jumps, **(l)** the intensity of the jumps and **(sigma)** the standard deviation of the Wiener process.
# + pycharm={"name": "#%%\n"}
data = simulation_test(v=0.035, l=9, step=1000, sigma=0.25,N=1)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Output of the Time Series
#
# Representation of the time series with marked jumps and detected anomalies with the Isolation Forest and features.
#
#
# + pycharm={"name": "#%%\n"}
plotter(data)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Output CutOff
# + pycharm={"name": "#%%\n"}
plot_cut(data,'Return log')
| Testdata/Notebooks/PlotPath.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python388jvsc74a57bd097ae724bfa85b9b34df7982b8bb8c7216f435b92902d749e4263f71162bea840
# ---
# This notebook was written by <NAME> for a workshop held 18-19 May 2021 as part of a contract from the [National Oceanography Centre](https://noc.ac.uk) to [Bolding & Bruggeman ApS](https://bolding-bruggeman.com/). It is licensed under a [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
import numpy
import scipy.stats
# %matplotlib widget
from matplotlib import pyplot
# ## The Sheldon-Sutcliffe size spectrum [(Sheldon et al. 1972)](https://doi.org/10.4319/lo.1972.17.3.0327)
# +
# Generate a Sheldon-Sutcliffe spectrum (equal biomass in log-spaced bins)
# with some random noise superimposed
noise_sd = 0.2 # coefficient of variation of log biomass
binbounds = numpy.arange(-3, 7) # log10 of individual mass
bincentres = 0.5 * (binbounds[1:] + binbounds[:-1])
binwidth = 10.**binbounds[1:] - 10.**binbounds[:-1]
biomass = numpy.ones_like(bincentres)
biomass = 10.**(1. + noise_sd * numpy.random.normal(size=bincentres.shape))
# -
# Plot size spectrum (biomass per bin)
fig, ax = pyplot.subplots()
ax.bar(10.**binbounds[:-1], biomass, width=.9*binwidth, align='edge')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylabel('biomass (g)')
ax.set_xlabel('individual wet mass (g)')
ax.set_ylim(1, 100)
ax.grid()
# ## Biomass density
# Convert to biomass density by dividing by bin width
fig, ax = pyplot.subplots()
ax.plot(10.**bincentres, biomass / binwidth, 'o')
x = bincentres
y = numpy.log10(biomass / binwidth)
regr = scipy.stats.linregress(x, y)
ax.plot([10.**x[0], 10.**x[-1]], [10.**(regr.intercept + regr.slope * x[0]), 10.**(regr.intercept + regr.slope * x[-1])], '-r')
ax.text(0.55, 0.5, 'slope = %.3f' % regr.slope, color='r', transform=ax.transAxes)
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylabel('biomass density (g/g)')
ax.set_xlabel('individual wet mass (g)')
ax.grid()
# ## Abundance density
# Convert to abundance density by dividing by biomass at bin centre
fig, ax = pyplot.subplots()
x = bincentres
y = numpy.log10(biomass / binwidth) - bincentres
regr = scipy.stats.linregress(x, y)
ax.plot(10.**bincentres, biomass / binwidth / 10.**bincentres, 'o')
ax.plot([10.**x[0], 10.**x[-1]], [10.**(regr.intercept + regr.slope * x[0]), 10.**(regr.intercept + regr.slope * x[-1])], '-r')
ax.text(0.55, 0.5, 'slope = %.3f' % regr.slope, color='r', transform=ax.transAxes)
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_ylabel('abundance density (#/g)')
ax.set_xlabel('individual wet mass (g)')
ax.grid()
| notebooks/1 - size spectra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="temjRZWJjOfH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="91d1317e-a96f-496a-f25e-14c22aac12bd"
import numpy as np
import pandas as pd
import nltk
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
import seaborn as sns
import re
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
# + id="EpjjhQpujcRs" colab_type="code" colab={}
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="1B7C55zDjgDD" colab_type="code" colab={}
#https://drive.google.com/file/d/1LxF2TdIqrZ71l8IQIbnVUejcpyoVVgKH/view?usp=sharing
downloaded = drive.CreateFile({'id':'1LxF2TdIqrZ71l8IQIbnVUejcpyoVVgKH'})
downloaded.GetContentFile('training.1600000.processed.noemoticon')
data = pd.read_csv('training.1600000.processed.noemoticon',encoding='latin-1',header=None)
# + id="TbBHVys2jxt7" colab_type="code" colab={}
stemmer = PorterStemmer()
# + id="1_zFKtvgj6e1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="ebf05bcb-9adf-4eec-e832-5287dafa45f3"
nltk.download('stopwords')
# + id="yOcBWy8kj96Y" colab_type="code" colab={}
text1=[]
# + id="pd7LJyskkAm1" colab_type="code" colab={}
for i in range(750000,850000):
sentence = re.sub('[^a-zA-Z123456789]', ' ', data[5][i])
sentence = sentence.lower()
sentence = sentence.split()
sentence = [stemmer.stem(word) for word in sentence if not word in stopwords.words('english')]
sentence = ' '.join(sentence)
text1.append(sentence)
# + id="n1x6m-OqkhCu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="51a46643-1a9e-4ca2-a53d-f744fb8b95fe"
len(text1)
# + id="WnlCDY3Lkk31" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="49144096-921e-4d8a-9c26-7fe7982e0d1a"
text1[-1]
# + id="THrnEgwjklKL" colab_type="code" colab={}
y = data[0][750000:850000]
# + id="k35UJlAekleK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="92552382-5112-4868-e690-5deb750a076c"
data[5][849999]
# + id="i8ob9xULklx-" colab_type="code" colab={}
cv = CountVectorizer(max_features=5000,ngram_range=(1,3))
X = cv.fit_transform(text1[0]).toarray()
# + id="apolHkHkk29O" colab_type="code" colab={}
xtrain,xtest,ytrain,ytest = train_test_split(X,y,test_size=0.33,random_state=42)
# + id="lbGBQCqck3LH" colab_type="code" colab={}
import matplotlib.pyplot as plt
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
See full source and example:
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + id="LDE_rIyck3Xy" colab_type="code" colab={}
from sklearn.naive_bayes import MultinomialNB
classifier=MultinomialNB()
# + id="UzD0lh1EnUtw" colab_type="code" colab={}
from sklearn import metrics
# + id="fZzB-uEZmshw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="78bedbcf-e856-4cf7-b6de-203736326f31"
classifier.fit(xtrain, ytrain)
pred = classifier.predict(xtest)
score = metrics.accuracy_score(ytest, pred)
print("accuracy: %0.3f" % score)
#cm = metrics.confusion_matrix(ytest, pred)
#plot_confusion_matrix(cm, classes=['FAKE', 'REAL'])
# + id="1xZgmySRnOOR" colab_type="code" colab={}
# https://drive.google.com/file/d/1c6f6uGi9YGjsv2Oernkr7cf9bVTgzpRj/view?usp=sharing
downloaded = drive.CreateFile({'id':'1c6f6uGi9YGjsv2Oernkr7cf9bVTgzpRj'})
downloaded.GetContentFile('test_middle.csv')
data = pd.read_csv('test_middle.csv')
# + id="stxfACXfo04O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="019067d9-7bca-44eb-b42e-8d66ff0949f5"
data.head()
# + id="W8AjukvQHgEv" colab_type="code" colab={}
# https://drive.google.com/file/d/1c6f6uGi9YGjsv2Oernkr7cf9bVTgzpRj/view?usp=sharing
downloaded = drive.CreateFile({'id':'1_VWF8om2pO4Bn77WygW-ggXKHFffY8k6'})
downloaded.GetContentFile('ocr_final_data.csv')
data1 = pd.read_csv('ocr_final_data.csv')
# + id="IgP_ICVoqZQj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="2bca3b42-3968-4b0b-9de2-cc3fbf1c2ba7"
data1.head()
# + id="M-D55HW0qlvE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="19f3ff95-60e7-47f5-cc83-c188896218dc"
type(data['Text'][0])
# + id="4sL6w1mEqOiC" colab_type="code" colab={}
predtext=[]
# + id="osjRudilrCXc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="42bc489d-131e-4fdb-90b6-f0cc5f727fef"
sentence = re.sub('[^a-zA-Z123456789]', ' ', data['Text'][0])
sentence.lower()
# + id="8z666d7-qXQd" colab_type="code" colab={}
for i in range(239):
sentence = re.sub('[^a-zA-Z123456789\n]', ' ', str(data1['Text'][i]))
sentence = sentence.lower()
sentence = sentence.split()
sentence = [stemmer.stem(word) for word in sentence if not word in stopwords.words('english')]
sentence = ' '.join(sentence)
predtext.append(sentence)
# + id="LatJV9yLqzB0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3b2b6685-729f-447b-ba29-16a2c2fb6946"
len(predtext)
# + id="wloh4uzErnBB" colab_type="code" colab={}
cv1 = CountVectorizer(max_features=5000,ngram_range=(1,3))
Xpredict = cv1.fit_transform(predtext).toarray()
# + id="NEl36i_Osxle" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f88d1d3f-0737-4fd6-8a20-a74e9674a07a"
Xpredict.shape
# + id="NolHqZrqtNzz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="025cd718-a2f9-47e2-c77b-c7a2f2ba85a1"
type(Xpredict)
# + id="dOWL6DXouNN5" colab_type="code" colab={}
x_predicted = np.zeros((239,5000))
# + id="47fTo-P1uOY3" colab_type="code" colab={}
x_predicted[:,0:3466] = Xpredict
# + id="YrC_27TCuNCu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f8197dfe-a645-43b9-d673-dc0424687064"
x_predicted.shape
# + id="S00x_hGjsQYn" colab_type="code" colab={}
finalpred = classifier.predict(Xpredict)
# + id="ujH9bIHdsaw5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="f8ad01d7-c301-41aa-e53d-2ec4f8603fff"
finalpred
# + id="qaxAdum7uveR" colab_type="code" colab={}
finalprediction = []
for i in range(239):
if(finalpred[i]==4):
finalprediction.append('Positive')
else:
finalprediction.append('Negative')
# + id="36w5GTXnvSVg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5a297ab9-7289-42b0-d8a7-8d08d71a2f17"
finalprediction[0]
# + id="Y9OBTvkMvgUT" colab_type="code" colab={}
#csv_input = pd.read_csv('input.csv')
data1['finalprediction'] = finalprediction
data.to_csv('test_middle.csv', index=False)
# + id="qegt4xhOxYdz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="76d68e8e-e96e-47bd-dcba-65de75094140"
# Write the DataFrame to CSV file.
with open('/gdrive/My Drive/test_middle.csv', 'w') as f:
data.to_csv(f)
# + id="6OMmHrP9ym-b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="2ea186df-7c81-4707-87ab-71559c702190"
from google.colab import files
data.to_csv('df.csv')
files.download('df.csv')
# + id="xhKkdz0S6wqJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d10afe6e-d4d8-46f6-80e6-8c9b0b2fabe8"
data['Text'].isnull().sum()
# + id="3UPeqsL6y7UK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5d9bdf70-42c7-4959-ae66-e58a8e78e4ef"
type(data['Text'][0])
# + id="Cy3bQfEK00V4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="417b38fb-c122-40f1-cf29-0ae975db1d65"
j=0
for i in range(239):
if type(data1['Text'][i])==float:
data1['finalprediction'][i]="Random"
j=j+1
print(j)
# + id="koqxOtHi2miS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1aefe8bd-f264-4df9-c72c-be550c203f68"
i
# + id="iAOOO9rH6Ng1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="84100c8e-60f4-4ffc-b6e1-939c9a0d4bd2"
sns.countplot(data1['finalprediction'])
# + id="6zdE_uYm7U1a" colab_type="code" colab={}
mydata = data1['Filename']
# + id="bQcjQhba7jxE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="45f3dea6-75d8-4b71-a449-801800a83239"
type(mydata)
# + id="dvQI8i457mAL" colab_type="code" colab={}
mydata = data['Filename']
# + id="B7XTSFv672fU" colab_type="code" colab={}
finalcsv = pd.DataFrame(mydata)
# + id="CSwLX5ty7-0i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="0633cd95-78b1-439c-a2ef-dac5c76658b3"
finalcsv.head()
# + id="UbIfps4l8fBZ" colab_type="code" colab={}
finalcsv['Category'] = data1['finalprediction']
# + id="czWuFDqm8lr1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="112ae30a-8e9d-4126-8fbf-f851b222f380"
finalcsv.head()
# + id="IWT3Z0Nl8n3a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="fede0856-b074-4e7a-af17-f9fc68b87844"
from google.colab import files
finalcsv.to_csv('submission1.csv')
files.download('submission1.csv')
# + id="P8a--RZz82b5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="caf6ae3b-b507-429f-83b0-422b8520621c"
sns.countplot(finalcsv['Category'])
# + id="Pj1Scs5D9EMb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0f8308f6-2911-43fd-b33c-c9556a50772f"
finalcsv.shape
# + id="0ovBOZBx9PaH" colab_type="code" colab={}
textcsv = pd.DataFrame(text1)
# + id="MxbixT-UScU0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="19a7ee69-968d-4cdc-a159-8b7d1c06bd6f"
textcsv.shape
# + id="nGxOK-u2SgH2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="426c5e22-14d5-4f55-a8c2-bad11f961df3"
textcsv.head()
# + [markdown] id="T5H0-gC_Sn8Y" colab_type="text"
# As cleaning the data takes alot time, So i save the text of index 750000 to 850000 in textcsv file
# + id="lcQOZ837SirB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="7beb5b93-e97d-4e2f-fb83-cd901a299e4e"
from google.colab import files
textcsv.to_csv('textcsv75-85.csv')
files.download('textcsv75-85.csv')
# + id="1_3dDzBVTClM" colab_type="code" colab={}
| HackerEarth_train_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kartikay-99k/Cough-detector-app/blob/master/Cough.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qCeXU-tfhY5_" colab_type="text"
# ## 1. Importing Lybraries
#
# + id="_vq8NclUhgf8" colab_type="code" outputId="afdf7697-4812-4409-dd9f-03b965d347d5" colab={"base_uri": "https://localhost:8080/", "height": 71}
import glob
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import seaborn as sns
from tensorflow.keras.preprocessing import image
from tensorflow.keras.datasets import mnist
from tensorflow.keras.layers import Conv2D, MaxPool2D, Dense, Flatten, Dropout, BatchNormalization
from tensorflow.keras.models import Sequential, load_model
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping
from tensorflow.keras.applications.vgg16 import preprocess_input
from sklearn.model_selection import train_test_split
from pathlib import Path
# + [markdown] id="AT5eT3cXpPr2" colab_type="text"
# ## 3. Data Preprocessing
# + id="EBpi-pROpKwY" colab_type="code" colab={}
# loading training data
def train_data():
# Loading Important Files
Cough_Path = Path("/content/drive/My Drive/Colab Notebooks/Data/Cough/")
NonCough_Path = Path("/content/drive/My Drive/Colab Notebooks/Data/NonCough/")
# list to hold Test data
Labels = []
Images = []
# for loading spectrogram Images
for img in Cough_Path.glob("*.png"):
# Load the image from disk
img = image.load_img(img)
# Convert the image to a numpy array
image_array = image.img_to_array(img)
# Add the image to the list of images
Images.append(image_array)
# For each 'not dog' image, the expected value should be 0
Labels.append(1)
print("***** C_ Done *****")
for img in NonCough_Path.glob("*.png"):
# Load the image from disk
img = image.load_img(img)
# Convert the image to a numpy array
image_array = image.img_to_array(img)
# Add the image to the list of images
Images.append(image_array)
# For each 'not dog' image, the expected value should be 0
Labels.append(0)
print("***** NC_ Done *****")
x_train = np.array(Images)
y_train = np.array(Labels)
x_train = x_train/255
x_train, val_x, y_train, val_y = train_test_split(x_train, y_train, test_size=0.2)
return x_train, val_x, y_train, val_y
# + id="_4XEw8QMsLkf" colab_type="code" outputId="0f8b5554-3981-4018-e423-59193c47c0d7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
x_train, val_x, y_train, val_y = train_data()
# + id="ztngGQ3Ku8OZ" colab_type="code" outputId="16900ffe-8116-4564-c253-b83c9fdb1246" colab={"base_uri": "https://localhost:8080/", "height": 34}
x_train.shape
# + [markdown] id="5mDzFwi_tzxg" colab_type="text"
# ## 3. Defining Importan Parameters
# + id="MJGkabPgtCyF" colab_type="code" colab={}
es = EarlyStopping(monitor='loss', patience=3)
filepath="/content/drive/My Drive/Colab Notebooks/Saved_M/bestmodel.h5"
md = ModelCheckpoint(filepath, monitor='loss', verbose=1, save_best_only=True, save_weights_only=False, mode='min')
# + id="YtilSgj_uDpa" colab_type="code" colab={}
##### Important Variables
epochs = 20
num_classes = 2
batch_size = 64
input_shape = (181, 279, 3)
adam = tf.keras.optimizers.Adam(learning_rate=0.0001, beta_1=0.9, beta_2=0.999, amsgrad=False)
# + [markdown] id="ZaWwESsauUMS" colab_type="text"
# ## 4. 4. Building CNN
# + id="k-Lwwtw8uOVp" colab_type="code" outputId="f909aa2e-6bee-484f-973c-fe67f39401e3" colab={"base_uri": "https://localhost:8080/", "height": 561}
model = Sequential()
# Filter 1
model.add(Conv2D(32, (3, 3), padding='same', input_shape=input_shape, activation= 'relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
#model.add(BatchNormalization())
# Filter 2
model.add(Conv2D(16, (3, 3), padding='same', activation= 'relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
#model.add(BatchNormalization())
# Filter 3
#model.add(Conv2D(16, (3, 3), padding='same', activation= 'relu'))
#model.add(Conv2D(16, (3, 3), padding='same', activation= 'relu'))
#model.add(MaxPool2D(pool_size=(2, 2)))
#model.add(Dropout(0.25))
#model.add(BatchNormalization())
# 1st Dense Layer
model.add(Flatten())
#model.add(Dense(1024, activation='relu'))
#model.add(Dropout(0.25))
#model.add(BatchNormalization())
# 2nd Dense Layer
#model.add(Dense(512, activation='relu'))
#model.add(Dropout(0.25))
# 3rd Dense Layer
#model.add(Dense(256, activation='relu'))
#model.add(Dropout(0.3))
# 4th Dense Layer
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.3))
# 5th Dense Layer
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
# 6th Dense Layer
#model.add(Dense(32, activation='relu'))
#model.add(Dropout(0.5))
# Output Layer
model.add(Dense(1, activation= 'sigmoid'))
# Model Compile
model.compile(optimizer= adam, loss= tf.keras.losses.binary_crossentropy, metrics=["accuracy"])
# Model Summery
model.summary()
# + [markdown] id="UqabsRY12IAe" colab_type="text"
# ## 5. Training Model
# + id="Gb5euMVy2osy" colab_type="code" outputId="12cddb3c-5716-4647-9e6a-115cb1bbda79" colab={"base_uri": "https://localhost:8080/", "height": 1000}
History = model.fit(x_train,
y_train,
batch_size=batch_size,
#steps_per_epoch=2048,
epochs = epochs,
verbose=2,
validation_data = (val_x, val_y),
callbacks = [es,md],
shuffle= True
)
# + [markdown] id="X1ZS46gt-o4R" colab_type="text"
# ## 6. Saving Model
# + id="2oYHLR6s-vql" colab_type="code" outputId="3e9659c3-80a3-4e27-bb08-65593f73981c" colab={"base_uri": "https://localhost:8080/", "height": 561}
cnn = load_model("/content/drive/My Drive/Colab Notebooks/Saved_M/bestmodel.h5")
cnn.summary()
# + id="baRLvm2e-4cl" colab_type="code" colab={}
model_structure = cnn.to_json()
saving_m = Path("/content/drive/My Drive/Colab Notebooks/Saved_M/N_model_structure_17_256_0.1293_0.9722_0.1457_0.9812.jason")
saving_m.write_text(model_structure)
# Saving Model
model.save("/content/drive/My Drive/Colab Notebooks/Saved_M/N_20_64_0.1089_0.9487_0.1305_0.9517.h5")
# Saving weights only
model.save_weights("/content/drive/My Drive/Colab Notebooks/Saved_M/N_W_20_64_0.1089_0.9487_0.1305_0.9517.h5")
| Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow import keras
import numpy as np
print(tf.__version__)
# +
imdb = keras.datasets.imdb
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=10000)
# -
print("Training entries: {}, labels: {}".format(len(train_data), len(train_labels)))
print(train_data[0])
len(train_data[0]), len(train_data[1])
# +
# A dictionary mapping words to an integer index
word_index = imdb.get_word_index()
# The first indices are reserved
word_index = {k:(v+3) for k,v in word_index.items()}
word_index["<PAD>"] = 0
word_index["<START>"] = 1
word_index["<UNK>"] = 2 # unknown
word_index["<UNUSED>"] = 3
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
# -
decode_review(train_data[0])
# +
train_data = keras.preprocessing.sequence.pad_sequences(train_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
test_data = keras.preprocessing.sequence.pad_sequences(test_data,
value=word_index["<PAD>"],
padding='post',
maxlen=256)
# -
len(train_data[0]), len(train_data[1])
print(train_data[0])
# +
# input shape is the vocabulary count used for the movie reviews (10,000 words)
vocab_size = 10000
model = keras.Sequential()
model.add(keras.layers.Embedding(vocab_size, 16))
model.add(keras.layers.GlobalAveragePooling1D())
model.add(keras.layers.Dense(16, activation=tf.nn.relu))
model.add(keras.layers.Dense(1, activation=tf.nn.sigmoid))
model.summary()
# -
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='binary_crossentropy',
metrics=['accuracy'])
# +
x_val = train_data[:10000]
partial_x_train = train_data[10000:]
y_val = train_labels[:10000]
partial_y_train = train_labels[10000:]
# -
history = model.fit(partial_x_train,
partial_y_train,
epochs=40,
batch_size=512,
validation_data=(x_val, y_val),
verbose=1)
# +
results = model.evaluate(test_data, test_labels)
print(results)
# -
history_dict = history.history
history_dict.keys()
# +
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# +
plt.clf() # clear figure
acc_values = history_dict['acc']
val_acc_values = history_dict['val_acc']
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# -
| Imdb_datatest.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Cvičení 2 - Pravděpodobnost
# ## <NAME>, <NAME>, <NAME>
# V tomto cvičení projdeme úvod do pravděpodobnosti. Předpokládáme znalosti z přednášky, především pojmy: **definice pravděpodobnosti, podmíněná pravděpodobnost, věta o úplné pravděpodobnosti, Bayesova věta**.
#
# # Pomocné funkce
#
# ## Úplná pravděpodobnost
# $P(A)=\sum_{i=1}^{n}P(B_i)P(A|B_i)$
# spočítání pravděpodobnosti P(A) - věta o úplné pravděpodobnosti
uplna_pravdepodobnost = function(P_B, P_AB)
{ # uvažujeme P_B jako vektor hodnot P(B_i) a P_BA jako vektor hodnot P(A|B_i)
P_A = 0
for (i in 1:length(P_B))
{
P_A = P_A + P_B[i]*P_AB[i]
}
return(P_A)
}
# ## Bayesova věta
# $P(B_k|A)=\frac{P(B_k)P(A|B_k)}{\sum_{i=1}^{n}P(B_i)P(A|B_i)}$
# spočítání podmíněné pravděpodobnosti P(B_k|A) - Bayesova věta
bayes = function(P_B, P_AB, k)
{ # uvažujeme P_B jako vektor hodnot P(B_i), P_BA jako vektor hodnot P(A|B_i)
P_A = uplna_pravdepodobnost(P_B, P_AB)
P_BkA = P_B[k]*P_AB[k]/P_A
return(P_BkA)
}
# **Přidáme funkce z munulého cvičení pro počítání kombinatorických výběrů, jsou v skriptu kombinatorika.R**
source('kombinatorika.R')
# # Příklady
# ## Příklad 1.
# Určete pravděpodobnost, že při hodu 20stěnnou spravedlivou (férovou) kostkou padne číslo větší než 14.
omega = 1:20
A = c(15,16,17,18,19,20)
# pravděpodobnost jako podíl příznivých ku všem
length(A)/length(omega)
# ## Příklad 2.
# Určete pravděpodobnost, že při hodu 20stěnnou kostkou padne číslo větší než 14, víte-li, že sudá čísla padají 2x častěji než lichá.
p_liche = 1/(20+10)
p_sude = 2*p_liche
pravd = c(p_liche, p_sude, p_liche, p_sude, p_liche, p_sude, p_liche, p_sude,
p_liche, p_sude, p_liche, p_sude, p_liche, p_sude, p_liche, p_sude,
p_liche, p_sude, p_liche, p_sude)
pravd
#pravdepodobnost je
sum(pravd[15:20])
# ## Příklad 3.
# Určete pravděpodobnost, že ve sportce uhodnete 4 čísla. (Ve sportce se losuje 6 čísel ze 49.)
(kombinace(6,4)*kombinace(43,2))/kombinace(49,6)
# ## Příklad 4.
# Z abecedního seznamu studentů zapsaných na dané cvičení vybere učitel prvních 12 a nabídne jim sázku: „Pokud se každý z Vás narodil v jiném znamení zvěrokruhu, dám každému z Vás 100 Kč. Pokud jsou však mezi Vámi alespoň dva studenti, kteří se narodili ve stejném znamení, dá mi každý z Vás 100 Kč.“ Vyplatí se studentům sázku přijmout? S jakou pravděpodobností studenti vyhrají?
permutace(12)/variace_opak(12,12)
# ## Příklad 5.
# Spočtěte pravděpodobnost toho, že z bodu 1 do bodu 2 bude protékat elektrický proud, je-li část el. obvodu včetně pravděpodobnosti poruch jednotlivých součástek vyznačen na následujícím obrázku. (Poruchy jednotlivých součástek jsou na sobě nezávislé.)
# 
# rozdělíme na bloky I=(A,B) a II=(C,D,E)
PI = 1 - (1 - 0.1)*(1 - 0.3)
PI
PII = 0.2*0.3*0.2
PII
# výsledek
(1 - PI)*(1-PII)
# ## Příklad 6.
# Ohrada má obdélníkový tvar, východní a západní stěna mají délku 40 m, jižní a severní pak 100 m. V této ohradě běhá kůň. Jaká je pravděpodobnost, že je k jižní stěně blíž než ke zbývajícím třem?
# geometrická pravděpodobnost
ohrada = 40*100
#blize k jihu
blize_J = 20*60 + 20*20
#pravdepodobnosti
blize_J/ohrada
# ## Příklad 7.
# U pacienta je podezření na jednu ze čtyř vzájemně se vylučujících nemocí - N1, N2, N3, N4 s pravděpodobností výskytu P(N1)=0,1; P(N2)=0,2; P(N3)=0,4; P(N4)=0,3. Laboratorní zkouška A je pozitivní v případě první nemoci v 50 % případů, u druhé nemoci v 75 % případů, u třetí nemoci v 15 % případů a u čtvrté v 20 % případů. Jaká je pravděpodobnost, že výsledek laboratorní zkoušky bude pozitivní?
# věta o úplné pravděpodobnosti
P_N = c(0.1,0.2,0.4,0.3) # P(N1), P(N2), ...
P_PN = c(0.5,0.75,0.15,0.2) # P(P|N1), P(P|N2), ...
P_P = uplna_pravdepodobnost(P_B = P_N, P_AB = P_PN) # P(P)
P_P
# ## Příklad 8.
# Telegrafické znaky se skládají ze signálů „tečka“, „čárka“. Je statisticky zjištěno, že se zkomolí 25 % sdělení „tečka“ a 20 % signálů „čárka“. Dále je známo, že signály se používají v poměru 3:2. Určete pravděpodobnost, že byl přijat správně signál, jestliže byl přijat signál „tečka“.
# Bayesova věta
P_O = c(0.6, 0.4) # P(O.), P(O-)
P_PO = c(0.75, 0.2) # P(P.|O.), P(P.|O-)
bayes(P_B = P_O, P_AB = P_PO, k = 1) # k = 1 protože správně = O.
# ## Příklad 9.
# V jednom městě jezdí 85 % zelených taxíků a 15 % modrých. Svědek dopravní nehody vypověděl, že nehodu zavinil řidič modrého taxíku, který pak ujel. Testy provedené za obdobných světelných podmínek ukázaly, že svědek dobře identifikuje barvu taxíku v 80 % případů a ve 20 % případů se mýlí.
# - Jaká je pravděpodobnost, že viník nehody skutečně řídil modrý taxík?
# - Následně byl nalezen další nezávislý svědek, který rovněž tvrdí, že taxík byl modrý. Jaká je nyní pravděpodobnost, že viník nehody skutečně řídil modrý taxík?
# - Ovlivní pravděpodobnost, že viník nehody skutečně řídil modrý taxík to, zda dva výše
# zmínění svědci vypovídali postupně nebo najednou?
# a) opět Bayesova věta
P_B = c(0.85, 0.15) # P(Z), P(M)
P_SB = c(0.20, 0.80) # P(SM|Z), P(SM|M)
bayes(P_B = P_B, P_AB = P_SB, k = 2) # modrý je druhý
# b) první možnost - druhý průchod Bayesem
P_M = bayes(P_B = P_B, P_AB = P_SB, k = 2)
P_B = c(1 - P_M, P_M) # P(Z), P(M)
P_SB = c(0.20, 0.80) # P(S2M|Z), P(S2M|M)
bayes(P_B = P_B, P_AB = P_SB, k = 2)
# c) nebo odpověděli najednou
P_B = c(0.85, 0.15) # P(Z), P(M)
P_SB = c(0.20^2, 0.80^2) # P(S1M&S2M|Z), P(S1M&S2M|M)
bayes(P_B = P_B, P_AB = P_SB, k = 2)
# ## Příklad 10.
# Potřebujeme zjistit odpověď na určitou citlivou otázku. Jak odhadnout, kolik procent dotazovaných na otázku odpoví ANO a přitom všem respondentům zaručit naprostou anonymitu? Jedním z řešení je tzv. dvojitě anonymní anketa:<br>
# Necháme respondenty hodit korunou a dvojkorunou a ti, kterým padl na koruně líc napíšou na lísteček odpověď (ANO/NE) na citlivou otázku. Ostatní respondenti napíší, zda jim padl na dvojkoruně líc (ANO/NE). Jakým způsobem určíme podíl studentů, kteří na citlivou otázku odpověděli ANO?<br>
# Předpokládejme, že respondenti byli dotazování, zda podváděli u zkoušky. Z anketních lístků se zjistilo, že „ANO“ odpovědělo 120 respondentů a „NE“ odpovědělo 200 respondentů. Kolik procent studentů podvádělo u zkoušky?
# věta o úplné pravděpodobnosti
# P(A) = P(K_lic)*P(A|K_lic)+P(K_rub)*P(D_lic|K_rub)
# rovnice 120/320=0.5*x+0.5*0.5
(120/320-0.5^2)/0.5
# ## Bonus - Monty Hall Problem
#
# Začneme s vygenerováním n instancí soutěže - cena bude náhodný index dveří (1,2,3) za kterými se může nacházet cena
n = 10000 # počet pokusů
cena = sample.int(n = 3, size = n, replace = TRUE) # náhdný výběr dveří
head(cena) # head vykresli prvních 6 prvků/řádků
# Totéž pro naši původní volbu - náhodný index dveří.
volba_orig = sample.int(n = 3, size = n, replace = TRUE) # původní volba
head(volba_orig)
# V prvním kole moderátor jedny prázdné dveře otevře, takto se to dá nasimulovat:
otevrene_dvere = rep(0, n) # inicializace vektoru
dvere_c = 1:3 # pomocná proměnná - identifikátory dveří
for (i in 1:n){
dvere_k_otevereni = c(TRUE, TRUE, TRUE) # inicializace
dvere_k_otevereni[cena[i]] = FALSE # nesmíme otevřít dveře s cenou
dvere_k_otevereni[volba_orig[i]] = FALSE # ani naše vybrané dveře
# ve zbytku jsou buď 2 (pokud jsme se trefili) nebo 1 dveře (pokud ne)
idx_dvere = dvere_c[dvere_k_otevereni]
if (length(idx_dvere) == 1){
otevrene_dvere[i] = idx_dvere # pokud jedny otevřeme je
} else { # pokud 2 tak jedny náhodně vybereme a otevřeme je
otevrene_dvere[i] = sample(x = idx_dvere, size = 1)
}
}
head(otevrene_dvere)
# Naše nová volba pokud se tak rozhodneme - součet indexů je 1+2+3=6 takže pokud my máme vybraný nějaký index, dále nějaký index se otevře, tak do zbytku 6 jsou ty třetí = naše nová volba.
nova_volba = 6 - (volba_orig + otevrene_dvere)
head(nova_volba)
# Úspěšnost při originální volbě:
p_orig = sum(cena == volba_orig)/n
p_orig
# Úspěšnost při výměně:
p_zmena = sum(cena == nova_volba)/n
p_zmena
p_orig + p_zmena
| CV2/cv2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
################################################################################
## Imports
import sys
import math
import numpy
numpy.set_printoptions(threshold=sys.maxsize)
import random
from pyeasyga import pyeasyga
import matplotlib as mpl
import matplotlib.ticker as ticker
import matplotlib.pyplot as plt
plt.style.use('ggplot')
################################################################################
## Common code
# Error
ERR = 1e-10
# Data
data = [0] * 5
# Define fitness function
def fitness_function(individual, data=None):
sum = 0
for u in individual:
sum += (u ** 2)
return sum
# Define the key field for sorting
def get_key(obj):
return obj.fitness
def round_up(n, decimals=10):
multiplier = 10 ** decimals
return math.floor(n * multiplier) / multiplier
################################################################################
## Simple Genetic Algorithm (sGA)
# Initialize genetic algorithm
sga = pyeasyga.GeneticAlgorithm(data, maximise_fitness=False)
# Set fitness function
sga.fitness_function = fitness_function
# Covariance matrices
sga_cm = {}
# Create a new individual
def create_individual(data):
# Set mutation range
mrange = len(data)
# Generate a random individual
individual = []
for d in data:
individual.append(random.uniform(d - mrange, d + mrange))
# Return a new individual
return individual
sga.create_individual = create_individual
# Fix mutate function
def mutate_function(individual):
# Set mutation range
mrange = max(individual)
mutate_index = random.randrange(len(individual))
d = individual[mutate_index]
individual[mutate_index] = random.uniform(d - mrange, d + mrange)
sga.mutate_function = mutate_function
# Fix population fitness calculation
def calculate_population_fitness(self):
for individual in self.current_generation:
individual.fitness = self.fitness_function(individual.genes, self.seed_data)
individual.fitness = round_up(individual.fitness)
individual.fitness = 0 if ERR >= individual.fitness else individual.fitness
sga.calculate_population_fitness = calculate_population_fitness
# Set initial population generation function (fix rank population call)
def create_first_generation(self):
self.create_initial_population()
self.calculate_population_fitness(self)
self.rank_population()
sga.create_first_generation = create_first_generation
# Set next population generation function (fix rank population call)
def create_next_generation(self):
self.create_new_population()
self.calculate_population_fitness(self)
self.rank_population()
sga.create_next_generation = create_next_generation
# Set evolution function
def run(self):
# Initialize seed data with random values
model_size = len(self.seed_data)
for i in range(model_size):
self.seed_data[i] = random.uniform(0, model_size)
# Run evolution
self.create_first_generation(self)
# Initial Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in self.current_generation]
sga_cm['icm'] = numpy.cov(arrs)
for i in range(1, self.generations):
self.create_next_generation(self)
# Intermediary
if i == int(self.generations / 2):
# Intermediary Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in self.current_generation]
sga_cm['tcm'] = numpy.cov(arrs)
# Final Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in self.current_generation]
sga_cm['fcm'] = numpy.cov(arrs)
sga.run = run
# Run sGA
sga.run(sga)
# Get best individual
result = sga.best_individual()
# Print result
print('The sGA best solution is: {}'.format(result))
################################################################################
## Compact Genetic Algorithm (cGA)
# Initialize genetic algorithm
cga = pyeasyga.GeneticAlgorithm(data, maximise_fitness=False)
# Set fitness function
cga.fitness_function = fitness_function
# Covariance matrices
cga_cm = {}
# Generate probability vector
def generate_prob(model_size):
prob = []
std_stdev = 1
for i in range(model_size):
bound = model_size + std_stdev
mean = random.uniform(-bound, bound)
pair = (mean, std_stdev)
prob.append(pair)
return prob
# Update probability vector
def update_prob(elite, prob):
# Update probability vector with the best results
for i in range(len(prob)):
# Mean and standard deviation of the ith element
aux = []
for item in elite:
mean = item.genes[i]
aux.append(mean)
# Update mean and stdev
prob[i] = numpy.mean(aux), numpy.std(aux, ddof=1)
# Create a new individual
def create_individual(prob):
individual = []
for p in prob:
mean, stdev = p
value = random.uniform(mean - stdev, mean + stdev)
individual.append(value)
mean = numpy.mean(individual)
stdev = numpy.std(individual, ddof=1)
individual = numpy.random.normal(mean, stdev, len(individual))
# Return a new individual
return pyeasyga.Chromosome(individual)
cga.create_individual = create_individual
icm_pop = None
tcm_pop = None
fcm_pop = None
# Set evolution function
def run(self):
# Initialize the max number of individuals in a offspring
offspring_max = self.population_size
# Initialize best solution
best = None
# Initialize best individuals population
k = int(self.population_size / 2)
population = []
arrs = []
# Initialize probability vector
prob = generate_prob(len(self.seed_data))
# Run `i` generations
for i in range(self.generations):
# Create individuals
for _ in range(offspring_max):
downward = self.create_individual(prob)
downward.fitness = round_up(self.fitness_function(downward.genes))
downward.fitness = 0 if ERR >= downward.fitness else downward.fitness
population.append(downward)
# Update best individuals population
population.sort(key=get_key)
population = population[:self.population_size]
elite = population[:k]
best = population[0]
# Initial population
if i == 0:
# Initial Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in population]
cga_cm['icm'] = numpy.cov(arrs)
# Intermediary
if i == int(self.generations / 2):
# Intermediary Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in population]
cga_cm['tcm'] = numpy.cov(arrs)
# Update the probability vector based on the success of each bit
update_prob(elite, prob)
# Add final solution
self.current_generation.append(best)
# Update best individuals population
population.sort(key=get_key)
population = population[:self.population_size]
# Final Covariance Matrix
arrs = [numpy.transpose(i.genes) for i in population]
cga_cm['fcm'] = numpy.cov(arrs)
cga.run = run
# Run evolution
cga.run(cga)
# Get best individual
result = cga.best_individual()
# Print result
print('The cGA best solution is: {}'.format(result))
################################################################################
## Heat map
def fmt(x, pos):
return r'{:.2f}'.format(x)
# sGA
hmmin, hmmax = -10, 10
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('sGA\nCovariance Matrix of Initial Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(sga_cm['icm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Blues)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('sGA\nCovariance Matrix of Intermediary Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(sga_cm['tcm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Blues)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('sGA\nCovariance Matrix of Final Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(sga_cm['fcm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Blues)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
# cGA
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('cGA\nCovariance Matrix of Initial Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(cga_cm['icm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Reds)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('cGA\nCovariance Matrix of Intermediary Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(cga_cm['tcm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Reds)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
fig, axis = plt.subplots(figsize=(13, 10))
fig.suptitle('cGA\nCovariance Matrix of Final Population',
fontsize=14, fontweight='bold', y=0.94)
heatmap = axis.pcolor(cga_cm['fcm'], vmin=hmmin, vmax=hmmax, cmap=plt.cm.Reds)
plt.colorbar(heatmap, format=ticker.FuncFormatter(fmt))
plt.show()
################################################################################
| ies8-hm3-sga-vs-cga-quadratic-function-rn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
# %matplotlib inline
data = pd.read_csv('train.csv')
data.describe()
# +
# Cabin and Ticket columns have a lot of missing values, thus they aren't useful features
# We drop those two columns
data = data.drop(['Ticket','Cabin'], axis=1)
# We also remove all rows with NaN in any column
data.fillna(data.mean(), inplace=True)
data.head(n=10)
# -
# #### More men died as compared to women
sns.barplot(x="Sex", y="Survived", data=data)
fig, ax = plt.subplots(figsize=(12, 8))
ax.hist([data[data['Survived'] == 1]['Age'], data[data['Survived'] == 0]['Age']], stacked=True, color=['g','b'], bins=30,
label=['Survived','Dead'])
ax.set_xlabel('Age')
ax.set_ylabel('Number of passengers')
ax.legend()
# +
survived_sex = data[data['Survived'] == 1]['Sex'].value_counts().sort_index()
dead_sex = data[data['Survived'] == 0]['Sex'].value_counts().sort_index()
survived_sex = survived_sex / float(survived_sex.sum())
dead_sex = dead_sex / float(dead_sex.sum())
df_survived = pd.DataFrame([survived_sex, dead_sex])
df_survived.index = ['Survived','Dead']
df_survived.plot(kind='barh', figsize = (12, 8), title="Survival with respect to gender (Proportionally adjusted)")
# -
fig, ax = plt.subplots(figsize=(12, 8))
ax.hist([data[data['Survived'] == 1]['Fare'], data[data['Survived'] == 0]['Fare']], stacked=True, color=['g','b'], bins=30,
label=['Survived','Dead'])
ax.set_xlabel('Fare')
ax.set_ylabel('Number of passengers')
ax.set_title("Survival as per the amount of fare paid")
ax.legend()
# +
survived = data[data['Survived'] == 1]['Embarked'].value_counts()
dead = data[data['Survived'] == 0]['Embarked'].value_counts()
df_embarked = pd.DataFrame([survived, dead])
df_embarked.index = ['Survived', 'Dead']
df_embarked.plot(kind='barh', stacked=True, figsize = (12, 8), title="Who survived with respect to port of embarkation?")
# +
survived = data[data['Survived'] == 1]['Pclass'].value_counts()
dead = data[data['Survived'] == 0]['Pclass'].value_counts()
df_embarked = pd.DataFrame([survived, dead])
df_embarked.index = ['Survived', 'Dead']
df_embarked.plot(kind='barh', stacked=True, figsize = (12, 8), title="Who survived with respect to ticket class?")
# +
port_s = data[data['Embarked'] == 'S']['Pclass'].value_counts()
port_c = data[data['Embarked'] == 'C']['Pclass'].value_counts()
port_q = data[data['Embarked'] == 'Q']['Pclass'].value_counts()
# Adjust proportion
df_embark = pd.DataFrame([port_s / float(port_s.sum()), port_c / float(port_c.sum()), port_q / float(port_q.sum())])
df_embark.index = ["S", "C", 'Q']
df_embark.plot(kind='barh', stacked=True, figsize = (12, 8), title="Cabin class with respect to port embarked")
# -
from sklearn import preprocessing
from sklearn.model_selection import GridSearchCV, train_test_split, learning_curve, cross_val_score
from sklearn.svm import SVC
from sklearn.metrics import make_scorer, accuracy_score
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None,
n_jobs=1, train_sizes=np.linspace(.1, 1.0, 5)):
"""
Generate a simple plot of the test and traning learning curve.
Parameters
----------
estimator : object type that implements the "fit" and "predict" methods
An object of that type which is cloned for each validation.
title : string
Title for the chart.
X : array-like, shape (n_samples, n_features)
Training vector, where n_samples is the number of samples and
n_features is the number of features.
y : array-like, shape (n_samples) or (n_samples, n_features), optional
Target relative to X for classification or regression;
None for unsupervised learning.
ylim : tuple, shape (ymin, ymax), optional
Defines minimum and maximum yvalues plotted.
cv : integer, cross-validation generator, optional
If an integer is passed, it is the number of folds (defaults to 3).
Specific cross-validation objects can be passed, see
sklearn.cross_validation module for the list of possible objects
n_jobs : integer, optional
Number of jobs to run in parallel (default 1).
"""
plt.figure(figsize=(12, 8))
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
return plt
# +
# Feature Engineering
def extract_titles(data):
# we extract the title from each name
data['Title'] = data['Name'].map(lambda name:name.split(',')[1].split('.')[0].strip())
# a map of more aggregated titles
Title_Dictionary = {
"Capt": "Officer",
"Col": "Officer",
"Major": "Officer",
"Jonkheer": "Royalty",
"Don": "Royalty",
"Sir" : "Royalty",
"Dr": "Officer",
"Rev": "Officer",
"the Countess":"Royalty",
"Dona": "Royalty",
"Mme": "Mrs",
"Mlle": "Miss",
"Ms": "Mrs",
"Mr" : "Mr",
"Mrs" : "Mrs",
"Miss" : "Miss",
"Master" : "Master",
"Lady" : "Royalty"
}
# we map each title
data['Title'] = data.Title.map(Title_Dictionary)
label_encoder = preprocessing.LabelEncoder()
data['FamilySize'] = data['SibSp'] + data['Parch']
data['Age*Class'] = data ['Age'] * data['Pclass']
data['Fare_Per_Person'] = data['Fare'] / (data['FamilySize']+1)
extract_titles(data)
data['Sex'] = label_encoder.fit_transform(data['Sex'].astype('str'))
data['Embarked'] = label_encoder.fit_transform(data['Embarked'].astype('str'))
data['Title'] = label_encoder.fit_transform(data['Title'].astype('str'))
# +
y = data['Survived']
X = data[['Sex', 'Age*Class', 'Fare', 'Fare_Per_Person', 'Title']]
# Testing / Training split
X_train, X_test, y_train, y_test = train_test_split(X, y)
# Using the Support Vector Machine (hyperparameters chosen using gridsearch)
clf = SVC(C=10, gamma=0.001, kernel='poly')
clf.fit(X_train, y_train)
clf.score(X_test, y_test)
# -
cv_scores = cross_val_score(clf, X, y, cv=5)
cv_scores.sum() / cv_scores.shape[0]
X = data[['Sex', 'Age*Class', 'Fare', 'Fare_Per_Person', 'Title']]
plot_learning_curve(clf, "Learning Curve", X.values, y.values)
# +
test_df = pd.read_csv("test.csv")
test_df = test_df.drop(['Ticket','Cabin'], axis=1)
test_df.head(n=10)
# -
# Fill NA age with median age
test_df['Age'].fillna(test_df['Age'].median(), inplace=True)
test_df.fillna(test_df.mean(), inplace=True)
test_df.head()
# +
test_df['FamilySize'] = test_df['SibSp'] + test_df['Parch']
test_df['Age*Class'] = test_df['Age'] * test_df['Pclass']
test_df['Fare_Per_Person'] = test_df['Fare'] / (test_df['FamilySize'] + 1)
extract_titles(test_df)
test_df['Sex'] = label_encoder.fit_transform(test_df['Sex'].astype('str'))
test_df['Embarked'] = label_encoder.fit_transform(test_df['Embarked'].astype('str'))
test_df['Title'] = label_encoder.fit_transform(test_df['Title'].astype('str'))
# +
X = test_df[['Sex', 'Age*Class', 'Fare', 'Fare_Per_Person', 'Title']]
pred = clf.predict(X=X)
submission = pd.DataFrame({"PassengerId": test_df["PassengerId"], "Survived": pred})
submission.to_csv('titanic1.csv', index=False)
| Titanic Machine Learning From Disaster/Titanic Machine Learning From Disaster .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming with Python
#
# ## Session 3
# ## Functions
#
# Functions are reusable blocks of code that you can name and execute any number of times from different parts of your script(s). This reuse is known as "calling" the function. Functions are important building blocks of a software.
#
# There are several built-in functions of Python, which can be called anywhere (and any number of times) in your current program. You have been using built-in functions already, for example, `len()`, `range()`, `sorted()`, `max()`, `min()`, `sum()` etc.
# #### Structure of writing a function:
#
# - `def` (keyword) + function name (you choose) + `()`.
# - newline with 4 spaces or a tab + block of code # Note: Codes at the 0 position are always read
# - Call your function using its name
# +
# Examples of the functions that you already know, i.e. print(), len(), max(), min()
myvar = [12,23,67,45,58,19]
# -
# Non parametric function
# Define a function that prints a sum of number1 and number2 defined inside the function
# Parametric function
# Example: a function to convert temperatures in Fahrenheit to Celcius
# tC = (tF - 32) * (5/9)
# Returning values
# change function above so that user can save the result, rather than printing it to screen
# +
# Local Vs. global variable
# variables defined inside a function are *not* visible to the "higher" levels of the program
# however, variables defined outside the function can be accessed inside the function
# be careful with this!
pi = 3.1415
def calculate_circumference(radius):
circ = radius * 2 * pi
return circ
# -
# ## Exercise 1 - Composing functions
#
# Write a function, similar to the one above, to convert a temperature in Celcius to the equivalent value in Kelvin. The equation is `tK = tC + 273.15`
# ## Exercise 2 - Combining functions
#
# Using the two temperature converting functions that we've already defined, write a new function that will convert a temperature in Fahrenheit to the equivalent temperature in __Kelvin__. Hint: remember that the point of encapsulating our code in functions is to avoid repeating ourselves.
# ## Exercise 3 - Combining strings
#
# “Adding” two strings produces their concatenation: `'a' + 'b'` is `'ab'`. Write a function called `fence` that takes two parameters called `original` and `wrapper` and returns a new string that has the wrapper character at the beginning and end of the original. A call to your function should look like this:
#
# ```Python
# print(fence('name', '*'))
# ```
# ```
# *name*
# ```
# ## Documenting Functions
# remember that you can use help() get help on how to use a function
help(range)
# +
# what happens if we call help() on one of the functions that we've defined ourselves?
help(calculate_circumference)
# how can we improve this? *why* should we?
| python/3-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
from SimPEG import np, sp, EM, Mesh
from SimPEG.EM.Static import DC
# %pylab inline
# Modified Pole-Pole array
def getDCdata(fname, offset=0.):
fid = open(fname, "rb")
lines = fid.readlines()
n = int(np.array(map(float, lines[5].split())) [0])
nelecl = int(np.array(map(float, lines[5].split())) [1])
nelec = nelecl+3
i_n = 0
data = []
print n, nelec
for iline, line in enumerate(lines[6:]):
temp = np.array(map(float, line.split()))
data.append(temp)
if temp.size is not 10:
i_n += 1
if i_n == n:
# aspacing = float(lines[6+iline+1].split()[1])
break
data = np.hstack(data)
fid.close()
ndata = data.size
B = []
M = []
for i_n in range(n):
tempB = np.array(np.arange(nelecl-i_n)+1)
tempM = np.array(np.arange(nelecl-i_n)+2+i_n)
B.append(tempB)
M.append(tempM)
B = np.hstack(B)
M = np.hstack(M)
A = np.ones(ndata, dtype=int)
N = np.ones(ndata, dtype=int) * (nelec-1)
aspacing = 5.
uniqB = np.unique(B)
srcList = []
dobsres = []
for isrc in range (nelecl):
locA = np.r_[0.+offset, 0., 0.]
locB = np.r_[uniqB[isrc]*aspacing+offset, 0., 0.]
rxinds = B == uniqB[isrc]
locsM = np.c_[M[rxinds]*aspacing+offset, np.zeros(rxinds.sum()), np.zeros(rxinds.sum())]
locsN = np.c_[N[rxinds]*aspacing+offset, np.zeros(rxinds.sum()), np.zeros(rxinds.sum())]
rx = DC.Rx.Dipole(locsM, locsN)
src = DC.Src.Dipole([rx], locA, locB)
srcList.append(src)
dobsres.append(data[rxinds])
dobsres = np.hstack(dobsres)
survey = DC.Survey(srcList)
survey.dobs = dobsres
return survey
fname2007 = "../data/resis2007/lakeside_1st.apr"
fname2015 = "../data/resis2015/IW1-1(20m).APR"
survey2007 = getDCdata(fname2007)
survey2015 = getDCdata(fname2015, offset=-10.)
fig, ax = plt.subplots(1,1, figsize=(10,1.5))
out = EM.Static.Utils.plot_pseudoSection(survey2007, ax, sameratio=False, dtype="volt", clim=(0.7, 80))
geom = np.hstack(out[3])
dobs2007 = survey2007.dobs*geom
ax.set_xlim(80, 280)
fig, ax = plt.subplots(1,1, figsize=(10,1.5))
out = EM.Static.Utils.plot_pseudoSection(survey2015, ax, sameratio=False, dtype="volt", clim=(0.7, 80))
geom = np.hstack(out[3])
temp = survey2015.dobs.copy()
temp [temp <= 0.] = np.nan
dobs2015 = temp*geom
ax.set_xlim(80, 280)
fig, ax = plt.subplots(1,1, figsize=(7,3))
out = hist(np.log10(temp[~np.isnan(temp)]), bins=100, color='b', alpha=0.5)
out = hist(np.log10(survey2007.dobs), bins=100, color='r', alpha=0.3)
ax.grid(True)
ax.legend(("Mar 01", "Apr 01"))
# +
from SimPEG.Survey import Data
def removeRxsfromDC(survey, inds, DClow=-np.inf, DChigh=np.inf, surveyType="2D"):
srcList = survey.srcList
srcListNew = []
dobs = survey.dobs
dobs[inds] = np.nan
data = Data(survey, survey.dobs)
rxData = []
for iSrc, src in enumerate(srcList):
rx = src.rxList[0]
data_temp = data[src, rx]
rxinds = np.isnan(data_temp) | (np.logical_or(DClow>data_temp, DChigh<data_temp))
nrxact_temp = rxinds.sum()
nrx_temp = len(rxinds)
rxlocM = rx.locs[0]
rxlocN = rx.locs[1]
srcloc = src.loc
rxData.append(data_temp[~rxinds])
# All Rxs are active
if nrxact_temp == 0:
if surveyType == "2D":
rxNew = DC.Rx.Dipole_ky(rxlocM, rxlocN)
else:
rxNew = DC.Rx.Dipole(rxlocM, rxlocN)
srcNew = DC.Src.Dipole([rxNew], srcloc[0], srcloc[1])
srcListNew.append(srcNew)
# All Rxs are nan then remove src
elif nrx_temp == nrxact_temp:
print ("Remove %i-th Src") % (iSrc)
# Some Rxs are not active
else:
if surveyType == "2D":
rxNew = DC.Rx.Dipole_ky(rxlocM[~rxinds,:], rxlocN[~rxinds,:])
else:
rxNew = DC.Rx.Dipole(rxlocM[~rxinds,:], rxlocN[~rxinds,:])
srcNew = DC.Src.Dipole([rxNew], srcloc[0], srcloc[1])
srcListNew.append(srcNew)
if surveyType == "2D":
surveyNew = DC.Survey_ky(srcListNew)
else:
surveyNew = DC.Survey(srcListNew)
surveyNew.dobs = np.hstack(rxData)
return surveyNew
def from3Dto2Dsurvey(survey):
srcLists2D = []
nSrc = len(survey.srcList)
for iSrc in range (nSrc):
src = survey.srcList[iSrc]
locsM = np.c_[src.rxList[0].locs[0][:,0], np.ones_like(src.rxList[0].locs[0][:,0])*-0.75]
locsN = np.c_[src.rxList[0].locs[1][:,0], np.ones_like(src.rxList[0].locs[1][:,0])*-0.75]
rx = DC.Rx.Dipole_ky(locsM, locsN)
locA = np.r_[src.loc[0][0], -0.75]
locB = np.r_[src.loc[1][0], -0.75]
src = DC.Src.Dipole([rx], locA, locB)
srcLists2D.append(src)
survey2D = DC.Survey_ky(srcLists2D)
survey2D.dobs = survey.dobs.copy()
return survey2D
# -
survey2007.dobs = dobs2007
fig, ax = plt.subplots(1,1, figsize=(10,1.5))
out = EM.Static.Utils.plot_pseudoSection(survey2007, ax, sameratio=False, dtype="appr", scale="log", clim=(-0.2, 2))
ax.set_xlim(80, 280)
survey2015.dobs = dobs2015
survey2015New = removeRxsfromDC(survey2015, [])
fig, ax = plt.subplots(1,1, figsize=(10,1.5))
out = EM.Static.Utils.plot_pseudoSection(survey2015New, ax, sameratio=False, dtype="appr", scale="log", clim=(-0.2, 2))
ax.set_xlim(80, 280)
cs = 2.5
npad = 6
hx = [(cs,npad, -1.3),(cs,150),(cs,npad, 1.3)]
hy = [(cs,npad, -1.3),(cs,20),(cs/2, 5)]
mesh = Mesh.TensorMesh([hx, hy])
mesh = Mesh.TensorMesh([hx, hy],x0=[-mesh.hx[:6].sum()-0.25-10, -mesh.hy.sum()])
from SimPEG import (Mesh, Maps, Utils, DataMisfit, Regularization,
Optimization, Inversion, InvProblem, Directives)
from pymatsolver import PardisoSolver
mapping = Maps.ExpMap(mesh)
survey2D = from3Dto2Dsurvey(survey2015New)
problem = DC.Problem2D_N(mesh, mapping=mapping)
problem.pair(survey2D)
problem.Solver = PardisoSolver
m0 = np.ones(mesh.nC)*np.log(1e-1)
mref = np.ones(mesh.nC)*np.log(1e-2)
# +
# from ipywidgets import interact
# nSrc = len(survey2D.srcList)
# def foo(isrc):
# figsize(10, 5)
# mesh.plotImage(np.ones(mesh.nC)*np.nan, gridOpts={"color":"k", "alpha":0.5}, grid=True)
# # isrc=0
# src = survey2D.srcList[isrc]
# plt.plot(src.loc[0][0], src.loc[0][1], 'bo')
# plt.plot(src.loc[1][0], src.loc[1][1], 'ro')
# locsM = src.rxList[0].locs[0]
# locsN = src.rxList[0].locs[1]
# plt.plot(locsM[:,0], locsM[:,1], 'ko')
# plt.plot(locsN[:,0], locsN[:,1], 'go')
# plt.gca().set_aspect('equal', adjustable='box')
# interact(foo, isrc=(0, nSrc-1, 1))
# -
weight = 1./abs(mesh.gridCC[:,1])**1.5
survey2D.eps = 10**(-2.5)
survey2D.std = 0.02
dmisfit = DataMisfit.l2_DataMisfit(survey2D)
regmap = Maps.IdentityMap(nP=int(mesh.nC))
reg = Regularization.Simple(mesh,mapping=regmap)
# reg = Regularization.Simple(mesh,mapping=regmap,cell_weights=weight)
opt = Optimization.ProjectedGNCG(maxIter=10)
opt.upper = np.log(1e0)
opt.lower = np.log(1e-3)
invProb = InvProblem.BaseInvProblem(dmisfit, reg, opt)
# Create an inversion object
beta = Directives.BetaSchedule(coolingFactor=5, coolingRate=2)
betaest = Directives.BetaEstimate_ByEig(beta0_ratio=1e0)
inv = Inversion.BaseInversion(invProb, directiveList=[beta, betaest])
problem.counter = opt.counter = Utils.Counter()
opt.LSshorten = 0.5
opt.remember('xc')
reg.mref = mref
mopt = inv.run(m0)
# sigma = np.ones(mesh.nC)
sigma = mapping*mopt
modelname = "sigma2015_lakeside.npy"
np.save(modelname, sigma)
fig, ax = plt.subplots(1,1, figsize = (20, 1.5))
sigma = mapping*mopt
dat = mesh.plotImage(np.log10(1./sigma), grid=False, ax=ax, pcolorOpts={"cmap":"jet"}, clim=(0, 3))
ax.set_ylim(-15, 0)
ax.set_xlim(-10, 380)
plt.colorbar(dat[0], ax=ax)
plt.plot(np.r_[1, 1]*342, np.r_[-20, 20], lw=3)
| notebook/ReadDCdata2015.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 13 - Text Similarity
# by [<NAME>](http://www.albahnsen.com/)
#
# version 1.0, June 2020
#
# ## Part of the class [AdvancedMethodsDataAnalysisClass](https://github.com/albahnsen/AdvancedMethodsDataAnalysisClass/tree/master/notebooks)
#
# This notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://creativecommons.org/licenses/by-sa/3.0/deed.en_US). Special thanks goes to [
# <NAME>](https://medium.com/@adriensieg/text-similarities-da019229c894)
# ## Text Similarities : Estimate the degree of similarity between two texts
#
# We always need to compute the similarity in meaning between texts.
# * Search engines need to model the relevance of a document to a query, beyond the overlap in words between the two. For instance, question-and-answer sites such as Quora or Stackoverflow need to determine whether a question has already been asked before.
# * In legal matters, text similarity task allow to mitigate risks on a new contract, based on the assumption that if a new contract is similar to a existent one that has been proved to be resilient, the risk of this new contract being the cause of financial loss is minimised. Here is the principle of Case Law principle. Automatic linking of related documents ensures that identical situations are treated similarly in every case. Text similarity foster fairness and equality. Precedence retrieval of legal documents is an information retrieval task to retrieve prior case documents that are related to a given case document.
# * In customer services, AI system should be able to understand semantically similar queries from users and provide a uniform response. The emphasis on semantic similarity aims to create a system that recognizes language and word patterns to craft responses that are similar to how a human conversation works. For example, if the user asks “What has happened to my delivery?” or “What is wrong with my shipping?”, the user will expect the same response.
#
# ### What is text similarity?
# Text similarity has to determine how ‘close’ two pieces of text are both in surface closeness [lexical similarity] and meaning [semantic similarity].
# # 1. Jaccard Similarity
#
# Jaccard similarity or intersection over union is defined as size of intersection divided by size of union of two sets. Let’s take example of two sentences:
# - Sentence 1: AI is our friend and it has been friendly
# - Sentence 2: AI and humans have always been friendly
#
# <img src="ts1.png"
# style="margin-right: 10px;" />
#
def jaccard_similarity(query, document):
intersection = set(query).intersection(set(document))
union = set(query).union(set(document))
return len(intersection)/len(union)
s1 = "AI is our friend and it has been friendly"
s2 = "AI and humans have always been friendly"
jaccard_similarity(s1, s2)
# # 2. Cosine Similarity
#
# Cosine similarity calculates similarity by measuring the cosine of angle between two vectors.
#
#
# <img src="ts2.png"
# style="margin-right: 10px;" />
#
#
# Mathematically speaking, Cosine similarity is a measure of similarity between two non-zero vectors of an inner product space that measures the cosine of the angle between them. The cosine of 0° is 1, and it is less than 1 for any angle in the interval (0,π] radians. It is thus a judgment of orientation and not magnitude: two vectors with the same orientation have a cosine similarity of 1, two vectors oriented at 90° relative to each other have a similarity of 0, and two vectors diametrically opposed have a similarity of -1, independent of their magnitude.
#
# The cosine similarity is advantageous because even if the two similar documents are far apart by the Euclidean distance (due to the size of the document), chances are they may still be oriented closer together. The smaller the angle, higher the cosine similarity.
#
# +
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import numpy as np
def cosine_distance_countVectorizer(s1, s2):
vect = CountVectorizer()
X_dtm = vect.fit_transform([s1, s2]).todense()
return cosine_similarity(X_dtm)
# -
cosine_distance_countVectorizer(s1, s2)
# # 3. Sentence Encoding + Cosine Similarity
# Word embedding is one of the most popular representation of document vocabulary. It is capable of capturing context of a word in a document, semantic and syntactic similarity, relation with other words, etc.
#
# It is common to find in many sources that the first step to cluster text data is to transform text units to vectors. This is not 100% true. But this step depends mostly on the similarity measure and the clustering algorithm. Some of the best performing text similarity measures don’t use vectors at all. This is the case of the winner system in SemEval2014 sentence similarity task which uses lexical word alignment. However, vectors are more efficient to process and allow to benefit from existing ML/DL algorithms.
#
#
# <img src="ts3.png"
# style="margin-right: 10px;" />
#
# import tensorflow as tf
import tensorflow.compat.v1 as tf
#To make tf 2.0 compatible with tf1.0 code, we disable the tf2.0 functionalities
tf.disable_eager_execution()
import tensorflow_hub as hub
# +
module_url = "https://tfhub.dev/google/universal-sentence-encoder/2" #@param ["https://tfhub.dev/google/universal-sentence-encoder/2", "https://tfhub.dev/google/universal-sentence-encoder-large/3"]
# Import the Universal Sentence Encoder's TF Hub module
embed = hub.Module(module_url)
# +
s1 = "AI is our friend and it has been friendly"
s2 = "AI and humans have always been friendly"
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
sentences_embeddings = session.run(embed([s1, s2]))
cosine_similarity(sentences_embeddings)
# +
s1 = "it our been and is has AI friendly friend"
s2 = "AI and humans have always been friendly"
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
sentences_embeddings = session.run(embed([s1, s2]))
cosine_similarity(sentences_embeddings)
# -
sentences_embeddings
sentences_embeddings.shape
# +
s1 = 'i like indian food because it is very spicy'
s2 = 'i like spicy food. For example, indian food'
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
sentences_embeddings = session.run(embed([s1, s2]))
# -
cosine_similarity(sentences_embeddings)
# +
s1 = 'i like indian food because it is very spicy'
s2 = 'soccer is a very fast sport'
s3 = 'bogotá is a very cold city'
s4 = 'the temperatures in my city are very low'
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
sentences_embeddings = session.run(embed([s1, s2, s3, s4]))
# -
cosine_similarity(sentences_embeddings)
| notebooks/.ipynb_checkpoints/13-TextSimilarity-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright © 2021, SAS Institute Inc., Cary, NC, USA. All Rights Reserved.
# SPDX-License-Identifier: Apache-2.0
# # Imports
import swat
import os
import sys
import re
import pandas as pd
sys.path.append(os.path.join(os.path.dirname(os.getcwd()),r"../../../common/python"))
import visualization as viz
import cas_connection as cas
# # CAS Connection
s=cas.reconnect()
s.loadActionSet('network')
s.loadActionSet('fcmpact')
# # Generate Word Embeddings
# Assign weights to word pairs (1 = perfect synonym, 0 = unrelated word).
# In practice, pre-trained word embeddings could be used instead of learning them here.
# These would typically be trained based on co-occurence frequency in a large corpus.
synonym_links = [['music', 'dvd', 0.1],
['music', 'mp3', 0.9],
['music', 'video', 0.3],
['book', 'mp3', 0.1],
['dvd', 'video', 0.95],
['mp3', 'video', 0.1]]
df_synonym_links = pd.DataFrame(synonym_links, columns = ['from', 'to', 'weight'])
s.upload(df_synonym_links, casout = {'name':'synonymLinks', 'replace':True})
s.CASTable('synonymLinks').head(6)
# Train word embeddings using vector node similarity, first order proximity
n_dim=10
convergence_threshold=0.00001
n_samples=1000000
result = s.network.nodeSimilarity(
links = {"name":"synonymLinks"},
vector = True,
jaccard = False,
convergenceThreshold = convergence_threshold,
nSamples = n_samples,
nDimensions = n_dim,
proximityOrder = "first",
outSimilarity = {"name":"outSim", "replace":True},
outNodes = {"name":"wordEmbeddings", "replace":True}
)
# Show the word similarity computed based on trained embeddings
# +
grouped_df = s.CASTable('outSim').to_frame().groupby('source')
for key, item in grouped_df:
display(grouped_df.get_group(key).head(100))
# -
# # Approximate Pattern Matching Based on Word Embeddings
# ## Main graph: purchase history for 6 people for 5 categories of products
nodes_purchase = [['1', 'person', 'Amy'],
['2', 'person', 'Blaine'],
['3', 'person', 'Catherine'],
['4', 'person', 'Dexter'],
['5', 'person', 'Edwin'],
['6', 'person', 'Faye'],
['MUSIC1','music', 'Dark_Side_of_the_Moon'],
['MUSIC2','music', 'Led_Zeppelin'],
['MP3A', 'mp3', 'Back_in_Black'],
['MP3B', 'mp3', 'From_This_Moment_On'],
['VIDEO1','video', 'Star_Wars'],
['DVD1', 'dvd', 'The_West_Wing'],
['DVD2', 'dvd', 'King_Kong'],
['BOOK1', 'book', 'Catcher_in_the_Rye'],
['BOOK2', 'book', 'Little_Women'],
['BOOK3', 'book', 'The_Bell_Jar']]
links_purchase = [['1', 'MUSIC1'],
['1', 'MP3A'],
['1', 'VIDEO1'],
['2', 'MUSIC1'],
['2', 'MP3A'],
['3', 'VIDEO1'],
['3', 'MUSIC1'],
['3', 'DVD1'],
['3', 'DVD2'],
['3', 'BOOK1'],
['4', 'MP3A'],
['4', 'BOOK1'],
['4', 'BOOK2'],
['4', 'BOOK3'],
['5', 'MP3B'],
['5', 'MUSIC2'],
['5', 'DVD1'],
['5', 'DVD2'],
['5', 'VIDEO1'],
['6', 'DVD2'],
['6', 'VIDEO1'],
['6', 'BOOK3'],
['6', 'MP3A']]
df_nodes_purchase = pd.DataFrame(nodes_purchase, columns = ['node', 'type', 'longName'])
df_links_purchase = pd.DataFrame(links_purchase, columns = ['from', 'to'])
s.upload(df_nodes_purchase, casout = {'name':'nodesPurchase', 'replace':True})
s.upload(df_links_purchase, casout = {'name':'linksPurchase', 'replace':True})
# ### Add color and label columns
s.datastep.runCode(
code="""
data nodesPurchase;
set nodesPurchase;
length color $8 label $52;
if type EQ 'music' then color='1';
else if type EQ 'mp3' then color='2';
else if type EQ 'video' then color='3';
else if type EQ 'dvd' then color='4';
else if type EQ 'book' then color='5';
else color='white';
label = CATS(longName,'\n','(',type,')');
"""
)
s.CASTable('nodesPurchase').head(16)
# ## Query graph: find pair of persons who purchased the same 2 video items
nodes_query = [['Person1', 'person'],
['Person2', 'person'],
['Video1', 'video' ],
['Video2', 'video' ]]
links_query = [['Person1', 'Video1'],
['Person1', 'Video2'],
['Person2', 'Video1'],
['Person2', 'Video2']]
df_nodes_query = pd.DataFrame(nodes_query, columns = ['node', 'type'])
df_links_query = pd.DataFrame(links_query, columns = ['from', 'to'])
s.upload(df_nodes_query, casout = {'name':'nodesQuery', 'replace':True})
s.upload(df_links_query, casout = {'name':'linksQuery', 'replace':True})
# ### Merge nodes with word embeddings
s.dataStep.runCode(
code = ''' data casuser.nodesQueryEmbed;
merge casuser.nodesQuery(in = nodeIn) casuser.wordEmbeddings(rename=(node=type));
by type;
if nodeIn;
run;'''
)
s.dataStep.runCode(
code = ''' data casuser.nodesPurchaseEmbed;
merge casuser.nodesPurchase(in = nodeIn) casuser.wordEmbeddings(rename=(node=type));
by type;
if nodeIn;
run;'''
)
# ## Define FCMP function for fuzzy match
# +
# Generate expressions for fcmp variable list
vars_comma_n = ', '.join([f'n.vec_{i}, nQ.vec_{i}' for i in range(n_dim)])
vars_dotproduct_n = '+ '.join([f'n.vec_{i}*nQ.vec_{i}' for i in range(n_dim)])
# Consider two types to be equivalent if the vector dot product value exceeds this threshold
fuzzy_match_threshold=0.7
# -
s.addRoutines(
routineCode = f'''
/** Node filter: we require exact match for type=person, approximate match otherwise **/
function nodeFilter(n.type $, nQ.type $, {vars_comma_n});
if (nQ.type EQ 'person') then return (n.type EQ nQ.type);
if ({vars_dotproduct_n} > {fuzzy_match_threshold}) then return (1);
return (0);
endsub;
/** Node pair filter: don't enumerate redundant (symmetric) permutations **/
function nodePairFilter(n.node[*] $, nQ.node[*] $);
if(nQ.node[1] EQ 'Person1' AND nQ.node[2] EQ 'Person2') then return (n.node[1] LT n.node[2]);
if(nQ.node[1] EQ 'Video1' AND nQ.node[2] EQ 'Video2') then return (n.node[1] LT n.node[2]);
return (1);
endsub;
''',
package = "myPackage",
saveTable = True,
funcTable = {"name":"myRoutines", "caslib":"casuser","replace":True}
)
s.sessionProp.setSessOpt(cmplib = "casuser.myRoutines")
# ## Approximate PatternMatch
nodes_var = ["type", "longName"]
nodes_var.extend([f"vec_{i}" for i in range(n_dim)])
nodes_query_var = ["type"]
nodes_query_var.extend([f"vec_{i}" for i in range(n_dim)])
result = s.network.patternMatch(
direction = "directed",
nodes = {"name":"nodesPurchaseEmbed"},
links = {"name":"linksPurchase"},
nodesQuery = {"name":"nodesQueryEmbed"},
linksQuery = {"name":"linksQuery"},
nodesVar = {"vars": nodes_var},
nodesQueryVar = {"vars": nodes_query_var, "varsMatch" : []},
nodePairFilter = "nodePairFilter(n.node, nQ.node)",
nodeFilter = f"nodeFilter(n.type, nQ.type, {vars_comma_n})",
outMatchNodes = {"name":"outMatchNodes", "replace":True},
outMatchLinks = {"name":"outMatchLinks", "replace":True}
)
num_matches = result.numMatches
# ## Input Graph Visualization
viz.graph2dot(nodesDf = s.CASTable('nodesPurchase'),
linksDf = s.CASTable('linksPurchase'),
directed = 1,
nodesLabel = "label",
nodesAttrs = {"colorscheme":"paired8", "style":"filled", "shape":"oval", "color":"black"},
nodeAttrs = {"fillcolor":"color", "label":"label"},
graphAttrs = {"layout":"dot", "rankdir":"TB"},
stdout=False
)
# ## Query Graph Visualization
viz.graph2dot(
linksDf = s.CASTable('linksQuery'),
directed = 1,
nodesAttrs = {"shape":"oval"},
graphAttrs = {"layout":"dot", "rankdir":"TB"},
size = 5,
stdout = False
)
# ## Matches Found Visualization
# +
highlightColor = 'blue'
highlightThickness = 3
def find_match(selectedMatch):
s.dataStep.runCode(
code = f'''
data casuser.linksPurchaseHighlighted;
merge casuser.linksPurchase
casuser.outMatchLinks(in=inMatch where=(match={selectedMatch}));
by from to;
if inMatch then do;
color= "{highlightColor}";
thickness={highlightThickness};
end;
run;''')
s.dataStep.runCode(
code = f'''
data casuser.nodesPurchaseHighlighted;
merge casuser.nodesPurchase
casuser.outMatchNodes(in=inMatch where=(match={selectedMatch}));
by node;
if inMatch then do;
pencolor="{highlightColor}";
thickness={highlightThickness};
end;
run;
'''
)
return viz.graph2dot(
nodesDf = s.CASTable('nodesPurchaseHighlighted'),
linksDf = s.CASTable('linksPurchaseHighlighted'),
nodesAttrs = {"colorscheme":"paired8", "style":"filled", "shape":"oval", "color":"black"},
nodeAttrs = {"fillcolor":"color", "label":"label", "color":"pencolor", "penwidth":"thickness"},
linkAttrs = {"color":"color", "penwidth":"thickness"},
graphAttrs = {"layout":"dot", "rankdir":"TB"},
directed = 1,
stdout = False
)
# -
for selected_match in range(num_matches):
display(find_match(selected_match))
| applications/natural-language-processing/word-embeddings-for-approximate-pattern-matching/python/patternmatch_with_embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import sys
sys.path.append('..')
# +
import pdb
#import dill as pickle
import torch
from transformers import BertTokenizer, BertModel, BertForMaskedLM
#optional
# import logging
# logging.basicConfig(level=logging.INFO)
# -
torch.cuda.set_device(1)
print(f'Using GPU #{torch.cuda.current_device()}')
# ## Utility methods
class Config(dict):
def __init__(self, **kwargs):
super().__init__(**kwargs)
for k, v in kwargs.items():
setattr(self, k, v)
def set(self, key, val):
self[key] = val
setattr(self, key, val)
def convert_to_snakecase(name):
s1 = re.sub('(.)([A-Z][a-z]+)', r'\1_\2', name)
return re.sub('([a-z0-9])([A-Z])', r'\1_\2', s1).lower().replace('__', '_')
# ## Quickstart
# ### Bert
# Use `BertTokenizer` to tokenize/numericalize input text
# load pre-trained model tokenizer (vocab)
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
# +
# tokenize input
text = '[CLS] What is George Lucas famous for ? [SEP] George Lucas created Star Wars [SEP]'
tokenized_text = tokenizer.tokenize(text)
print(tokenized_text)
# +
# mask a token to predict with `BertForMaskedLM`
masked_idx = 9
tokenized_text[masked_idx] = '[MASK]'
assert tokenized_text == ['[CLS]', 'what', 'is', 'george', 'lucas', 'famous', 'for', '?', '[SEP]', '[MASK]', 'lucas', 'created', 'star', 'wars', '[SEP]']
# +
# convert tokens to vocab idxs
tokenized_idxs = tokenizer.convert_tokens_to_ids(tokenized_text)
# define segment_ids
segment_idxs = [0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1]
print(tokenized_idxs)
# -
# convert inputs to pytorch tensors
T_tokenized_idxs = torch.tensor([tokenized_idxs])
T_segment_idxs = torch.tensor([segment_idxs])
# Use `BertModel` to **encode our inputs in hidden-states**:
# load pre-trained model (weights)
model = BertModel.from_pretrained('bert-base-uncased')
# set model to eval (deactivates Dropout and BatchNorm)
model = model.eval()
# (optional) put everything on GPU
T_tokenized_idxs = T_tokenized_idxs.to('cuda')
T_segment_idxs = T_segment_idxs.to('cuda')
model = model.to('cuda')
# +
# predict hidden states features for each layer
with torch.no_grad():
outputs = model(T_tokenized_idxs, token_type_ids=T_segment_idxs)
# outputs are tuples; here first element = the hidden state of the last layer of Bert model
encoded_layers = outputs[0]
# we have encoded our input sequence in a FloatTensor (bsz, seq_len, model_hidden)
print(encoded_layers.shape)
assert tuple(encoded_layers.shape) == (1, len(tokenized_idxs), model.config.hidden_size)
# -
# And how to use `BertForMaskedLM` to **predict a masked token**:
# +
# load pre-trained model (weights)
model = BertForMaskedLM.from_pretrained('bert-base-uncased')
model.eval()
# (optional) put everything on GPU
T_tokenized_idxs = T_tokenized_idxs.to('cuda')
T_segment_idxs = T_segment_idxs.to('cuda')
model = model.to('cuda')
# predict all tokens
with torch.no_grad():
outputs = model(T_tokenized_idxs, token_type_ids=T_segment_idxs)
predictions = outputs[0]
# +
predicted_idx = torch.argmax(predictions[0, masked_idx]).item()
predicted_token = tokenizer.convert_ids_to_tokens([predicted_idx])
assert predicted_token[0] == 'george'
# -
predicted_idx, predicted_token[0]
# ### OpenAI GPT-2
# Use `GPT2Tokenizer` to tokenize text
import torch
from transformers import GPT2Tokenizer, GPT2LMHeadModel
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
# +
# tokenize input
text = ' What is <NAME> famous for ? <NAME> created Star'
indexed_tokens = tokenizer.encode(text)
print(indexed_tokens)
# -
# convert tokens to idxs
tokens_tensor = torch.tensor([indexed_tokens])
# Use `GPT2LMHeadModel` to **predict the next token from a text prompt**
# +
model = GPT2LMHeadModel.from_pretrained('gpt2')
model = model.eval()
tokens_tensor = tokens_tensor.to('cuda')
model = model.to('cuda')
with torch.no_grad():
outputs = model(tokens_tensor)
preds = outputs[0]
# -
preds.shape
# get the predicted next sub-word
predicted_index = torch.argmax(preds[0, -1, :]).item()
# predicted_text = tokenizer.decode(indexed_tokens + [predicted_index])
tokenizer.decode(indexed_tokens + [predicted_index])
| nbs/_transformers-quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Racket
# language: racket
# name: racket
# ---
(require circuitous redex/reduction-semantics)
# Specialize to kn+2 = 2 for now
(define start
(circuit
#:inputs (RES GO KILL SUSP)
#:outputs (SEL K0 K1 K2)
;; exit 2
(lem = (and SEL (and RES (not lsel))))
(l0 = false)
(l1 = false)
(l2 = GO)
(lsel = false)
;; pause
(rem = (and SEL (and RES (not rsel))))
(r0 = (and reg-out RES))
(r1 = GO)
(r2 = false)
(rsel = reg-out)
(reg reg-in reg-out = (and (not do-kill) reg-set))
(reg-set = (or GO is-susp))
(is-susp = (and SUSP rsel))
;; K interface
(K0 = I0)
(K1 = I1)
(K2 = I2)
(SEL = (or rsel lsel))
;; synchronizer
(do-kill = (or KILL I2))
(left0 = (or l0 lem))
(both0 = (or l0 r0))
(right0 = (or r0 rem))
(I0 = (and left0 (and both0 right0)))
(left1 = (or left0 l1))
(both1 = (or l1 r1))
(right1 = (or right0 r1))
(I1 = (and left1 (and both1 right1)))
(left2 = (or left1 l2))
(both2 = (or l2 r2))
(right2 = (or right1 r2))
(I2 = (and left2 (and both2 right2)))))
(define end
(circuit
#:inputs (GO)
#:outputs (SEL K0 K1 K2)
(K0 = false)
(K1 = false)
(K2 = GO)
(SEL = false)))
# Okay first we show that in the first instant the two circuits are quivalent
(assert-same start end)
| circuits/[par-1exit].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NYC High Schools Analysis
#
# Date March 3, 2018
#
# New York City has a significant immigrant population and is very diverse, so comparing demographic data such as race, income, and gender with SAT (Scholastic Aptitude Test) scores is a good way to determine whether the SAT is a fair test.
#
# This notebook is an exercise analysis that looks for any relationships or correlations, if any, among those factors.
#
# The dataset is via NYC Open Data circa 2012 https://data.cityofnewyork.us/Education/2012-SAT-Results/f9bf-2cp4
#
# +
import pandas as pd
import numpy
import re
data_files = [
"ap_2010.csv",
"class_size.csv",
"demographics.csv",
"graduation.csv",
"hs_directory.csv",
"sat_results.csv"
]
data = {}
for f in data_files:
d = pd.read_csv("schools/{0}".format(f))
data[f.replace(".csv", "")] = d
# -
data
# # Read in the surveys
#
# Read in the surveys, which are in delimited text files, and do some housecleaning.
# +
all_survey = pd.read_csv("schools/survey_all.txt", delimiter="\t", encoding='windows-1252')
d75_survey = pd.read_csv("schools/survey_d75.txt", delimiter="\t", encoding='windows-1252')
survey = pd.concat([all_survey, d75_survey], axis=0)
survey["DBN"] = survey["dbn"]
survey_fields = [
"DBN",
"rr_s",
"rr_t",
"rr_p",
"N_s",
"N_t",
"N_p",
"saf_p_11",
"com_p_11",
"eng_p_11",
"aca_p_11",
"saf_t_11",
"com_t_11",
"eng_t_11",
"aca_t_11",
"saf_s_11",
"com_s_11",
"eng_s_11",
"aca_s_11",
"saf_tot_11",
"com_tot_11",
"eng_tot_11",
"aca_tot_11",
]
survey = survey.loc[:,survey_fields]
data["survey"] = survey
# -
data
# # Add DBN feature
#
# Construct an additional DBN feature
# +
data["hs_directory"]["DBN"] = data["hs_directory"]["dbn"]
def pad_csd(num):
string_representation = str(num)
if len(string_representation) > 1:
return string_representation
else:
return "0" + string_representation
data["class_size"]["padded_csd"] = data["class_size"]["CSD"].apply(pad_csd)
data["class_size"]["DBN"] = data["class_size"]["padded_csd"] + data["class_size"]["SCHOOL CODE"]
# -
# # Convert SAT features to numeric
# +
cols = ['SAT Math Avg. Score', 'SAT Critical Reading Avg. Score', 'SAT Writing Avg. Score']
for c in cols:
data["sat_results"][c] = pd.to_numeric(data["sat_results"][c], errors="coerce")
data['sat_results']['sat_score'] = data['sat_results'][cols[0]] + data['sat_results'][cols[1]] + data['sat_results'][cols[2]]
def find_lat(loc):
coords = re.findall("\(.+, .+\)", loc)
lat = coords[0].split(",")[0].replace("(", "")
return lat
def find_lon(loc):
coords = re.findall("\(.+, .+\)", loc)
lon = coords[0].split(",")[1].replace(")", "").strip()
return lon
data["hs_directory"]["lat"] = data["hs_directory"]["Location 1"].apply(find_lat)
data["hs_directory"]["lon"] = data["hs_directory"]["Location 1"].apply(find_lon)
data["hs_directory"]["lat"] = pd.to_numeric(data["hs_directory"]["lat"], errors="coerce")
data["hs_directory"]["lon"] = pd.to_numeric(data["hs_directory"]["lon"], errors="coerce")
# -
# # Condense datasets
# +
class_size = data["class_size"]
class_size = class_size[class_size["GRADE "] == "09-12"]
class_size = class_size[class_size["PROGRAM TYPE"] == "GEN ED"]
class_size = class_size.groupby("DBN").agg(numpy.mean)
class_size.reset_index(inplace=True)
data["class_size"] = class_size
data["demographics"] = data["demographics"][data["demographics"]["schoolyear"] == 20112012]
data["graduation"] = data["graduation"][data["graduation"]["Cohort"] == "2006"]
data["graduation"] = data["graduation"][data["graduation"]["Demographic"] == "Total Cohort"]
# -
# # Convert AP scores to numeric
# +
cols = ['AP Test Takers ', 'Total Exams Taken', 'Number of Exams with scores 3 4 or 5']
for col in cols:
data["ap_2010"][col] = pd.to_numeric(data["ap_2010"][col], errors="coerce")
# -
# # Combine the datasets
# +
combined = data["sat_results"]
combined = combined.merge(data["ap_2010"], on="DBN", how="left")
combined = combined.merge(data["graduation"], on="DBN", how="left")
to_merge = ["class_size", "demographics", "survey", "hs_directory"]
for m in to_merge:
combined = combined.merge(data[m], on="DBN", how="inner")
combined = combined.fillna(combined.mean())
combined = combined.fillna(0)
# -
# # Add a school district feature for mapping
# +
def get_first_two_chars(dbn):
return dbn[0:2]
combined["school_dist"] = combined["DBN"].apply(get_first_two_chars)
# -
# # Find correlations
correlations = combined.corr()
correlations = correlations["sat_score"]
print(correlations)
# # Plotting survey correlations
# Remove DBN since it's a unique identifier, not a useful numerical value for correlation.
survey_fields.remove("DBN")
correlations.dropna()
len(correlations.dropna().values)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from numpy import arange
import numpy as np
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
bar_heights = correlations.dropna().values
bar_positions = np.arange(len(correlations.dropna().values)) + 0.75
ax.bar(bar_positions, bar_heights, 0.5)
plt.show()
# -
dir(ax)
# # Correlation between Safety Scores and SAT
#scatter plot exercise 2
# %matplotlib inline
fig2 = plt.figure()
ax2 = fig2.add_subplot(1,1,1)
x=combined['saf_s_11'].values
y=combined['sat_score'].values
ax2.scatter(x,y,alpha=0.5)
ax2.set_xlabel('saf_s_11')
ax2.set_ylabel('sat_score')
plt.show()
y
x
combined[["saf_s_11","school_dist"]]
import pandas as pd
safety_scores = pd.pivot_table(combined,values='saf_s_11',index='school_dist',aggfunc=np.average)
safety_scores
import numpy
districts = combined.groupby("school_dist").agg(numpy.mean)
districts.reset_index(inplace=True)
print(districts.head())
# # Correlate demographics and SAT
# %matplotlib inline
fig3 = plt.figure()
ax3 = fig3.add_subplot(1,1,1)
x=combined['white_per'].values
y=combined['sat_score'].values
ax3.scatter(x,y,alpha=0.5)
ax3.set_xlabel('white_per')
ax3.set_ylabel('sat_score')
plt.show()
# %matplotlib inline
fig4 = plt.figure()
ax4 = fig4.add_subplot(1,1,1)
x=combined['asian_per'].values
y=combined['sat_score'].values
ax4.scatter(x,y,alpha=0.5)
ax4.set_xlabel('asian_per')
ax4.set_ylabel('sat_score')
plt.show()
# %matplotlib inline
fig5 = plt.figure()
ax5 = fig5.add_subplot(1,1,1)
x=combined['black_per'].values
y=combined['sat_score'].values
ax5.scatter(x,y,alpha=0.5)
ax5.set_xlabel('black_per')
ax5.set_ylabel('sat_score')
plt.show()
# %matplotlib inline
fig6 = plt.figure()
ax6 = fig6.add_subplot(1,1,1)
x=combined['hispanic_per'].values
y=combined['sat_score'].values
ax6.bar(x,y,alpha=0.5)
ax6.set_xlabel('hispanic_per')
ax6.set_ylabel('sat_score')
plt.show()
# %matplotlib inline
fig7 = plt.figure()
ax7 = fig7.add_subplot(1,1,1)
x=combined['hispanic_per'].values
y=combined['sat_score'].values
ax7.scatter(x,y,alpha=0.5)
ax7.set_xlabel('hispanic_per')
ax7.set_ylabel('sat_score')
plt.show()
# # Correlate Gender and SAT
# %matplotlib inline
fig8 = plt.figure()
ax8 = fig8.add_subplot(1,1,1)
x=combined['male_per'].values
y=combined['sat_score'].values
ax8.scatter(x,y,alpha=0.5)
ax8.set_xlabel('male_per')
ax8.set_ylabel('sat_score')
plt.show()
# %matplotlib inline
fig9 = plt.figure()
ax9 = fig9.add_subplot(1,1,1)
x=combined['female_per'].values
y=combined['sat_score'].values
ax9.scatter(x,y,alpha=0.5)
ax9.set_xlabel('female_per')
ax9.set_ylabel('sat_score')
plt.show()
combined
combined['ap_per']=combined['AP Test Takers ']/combined['total_enrollment']
# %matplotlib inline
fig10 = plt.figure()
ax10 = fig10.add_subplot(1,1,1)
x=combined['ap_per'].values
y=combined['sat_score'].values
ax10.scatter(x,y,alpha=0.5)
ax10.set_xlabel('ap_per')
ax10.set_ylabel('sat_score')
plt.show()
| Exercises/NYC_High_Schools/.ipynb_checkpoints/Schools-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Optimal lockdown start time
using DifferentialEquations, Plots, Plots.PlotMeasures
pyplot()
# ## Utility functions
# +
# change plot fonts
bplot(p, s) = plot!(p, titlefont=font(s, "Times"), xguidefontsize=s-4
, ytickfont=font(s-4, "Times"), margin=8mm, yguidefontsize=s-4
, xtickfont = font(s-4, "Times"), legendfont = font(s-4, "Times"))
# this function will generate a T period of social distancing at time t0
sinput(t, ts, td) = if (t-ts<0 || t-ts-td>0) 0 else 1 end
# -
# # Numerical investigating functions
function numericalsearch(model, u0, pr, icomp)
if(size(icomp,1)==1) pointer=2 else pointer=1 end
# setup ode problem setup based on no lockdown mandate
β, βl, γ = pr[1:3]
tspn = (0., 1000.)
prob = ODEProblem(model, u0, tspn, pr)
nlck = solve(prob, BS3(), reltol=1e-8,abstol=1e-8, saveat=1e-2)
inf = sum(nlck[icomp,:],dims=pointer)
if (pointer==1) inf=inf' end
v = maximum(inf)
vind = argmax(inf)
if (pointer==1) vind=vind[1] end
# max duration of lockdown
maxduration = floor(Int,nlck.t[vind])
A = v*ones(maxduration,1)
O = v*ones(maxduration,1)
# for each lockdown durations
for td = 1:maxduration
# simulation with start time formula (based on β=0)
Is = v/(2-exp(-γ*td))
ts = nlck.t[argmin(abs.(inf[1:vind].-Is))]
pr[end-1:end] = [ts, td]
prob = ODEProblem(model, u0, tspn, pr)
appx = solve(prob,BS3(),reltol=1e-8,
abstol=1e-8,saveat=1e-2)
infa = sum(appx[icomp,:],dims=pointer)
if(pointer==1) infa=infa' end
A[td]= maximum(infa)
# computation search for numerical optimal start time
tsr = ts-maxduration:2:ts+maxduration
peak = v
# for each duration start time
for i=1:size(tsr,1)
pr[end-1] = tsr[i]
prob = ODEProblem(model, u0, tspn, pr)
sol = solve(prob, BS3(), reltol=1e-8,
abstol=1e-8, saveat=1e-2)
info = sum(sol[icomp,:],dims=pointer)
if(pointer==1) info=info' end
peak = min(peak,maximum(info))
end
O[td]= peak
end
[A, O]
end
function lockdown(model, u0, pr, icomp)
A, O = numericalsearch(model, u0, pr,icomp)
β, βl, γ, ts, td = pr
td = 1:size(A,1)
p1 = plot(td, A, label="approximation",
ylabel="infected peak", linewidth=3,
title=string(model))
p1 = plot!(p1, td, O, label="optimal lockdown",
linewidth=3, linestyle=:dash, xlabel="Lockdown Duration")
p1 = bplot(p1, 12)
savefig(p1, "figures/"*string(model)*"_app1_"*string(βl)*".png")
savefig(p1, "figures/"*string(model)*"_app1_"*string(βl)*".svg")
p2 = plot(td, 100*abs.((O-A)./O), label=:false,
ylabel="% difference", linewidth=3,
title="βl/β="*string(round(βl/β,digits=3)))
p2 = bplot(p2, 12)
savefig(p2, "figures/"*string(model)*"_app2_"*string(βl)*".png")
savefig(p2, "figures/"*string(model)*"_app2_"*string(βl)*".svg")
p = plot(p1, p2, layout=(2,1))
savefig(p, "figures/"*string(model)*"_app_"*string(βl)*".png")
savefig(p, "figures/"*string(model)*"_app_"*string(βl)*".svg")
end
# # SIR
# +
# model description ts and td are pr[end-1:end]
function SIR(dx,x,p,t)
β, βl, γ, ts, td = p
u = β - (β-βl).*sinput(t, ts, td)
dx[1] = -u*x[1]*x[2]
dx[2] = u*x[1]*x[2] - γ*x[2]
dx[3] = γ*x[2]
end
pr = [0.2,0.02,0.05,0,0]
u0 = [1-1e-3, 1e-3, 0.0]
icomp = 2
lockdown(SIR, u0, pr, icomp)
# -
pr = [0.2,0.01,0.05,0,0]
lockdown(SIR, u0, pr, icomp)
# # SAIR (I)
# +
# model description
function SAIR(dx,x,p,t)
βᵢ, βlᵢ, γᵢ, βₐ, βlₐ, γₐ, δ, ts, td = p
u1 = βₐ - (βₐ-βlₐ).*sinput(t, ts, td)
u2 = βᵢ - (βᵢ-βlᵢ).*sinput(t, ts, td)
dx[1] = -u1*x[1]*x[2] - u2*x[1]*x[3]
dx[2] = u1*x[1]*x[2] + u2*x[1]*x[3] - (δ+γₐ)*x[2]
dx[3] = δ*x[2] - γᵢ*x[3]
dx[4] = γₐ*x[2] + γᵢ*x[3]
end
# define initial values
u0 = [1-11e-4, 1e-3, 1e-4, 0.]
pr = [0.2, 0.02, 0.05, 0.3, 0.03, 0.09, 0.15, 0, 0]
icomp = 3
lockdown(SAIR, u0, pr, icomp)
# -
# # SAIR (A)
icomp = 2
lockdown(SAIR, u0, pr, icomp)
# # SAIR (I+A)
icomp = 2:3
lockdown(SAIR, u0, pr, icomp)
# # SIDARTHE (I+D+A+R+T)
# +
# SIDARHE model description
function SIDARTHE(du,u,p,t)
β, βl = p[1:2]
s = p[3:18]
N, ts, td = p[19:21]
beta = β - (β-βl).*sinput(t, ts, td)
du[1] = -β*u[1]*(s[1]*u[2]+s[2]*u[3]+s[3]*u[4]+s[4]*u[5])/N
du[2] = β*u[1]*(s[1]*u[2]+s[2]*u[3]+s[3]*u[4]+s[4]*u[5])/N
- (s[5]+s[6]+s[7])*u[2]
du[3] = s[5]*u[2] - (s[8]+s[9])*u[3]
du[4] = s[6]*u[2] - (s[10]+s[11]+s[12])*u[4]
du[5] = s[8]*u[3] + s[10]*u[4] - (s[13]+s[14])*u[5]
du[6] = s[11]*u[4] + s[13]*u[5] - (s[15]+s[16])*u[6]
du[7] = s[7]*u[2] + s[9]*u[3] + s[12]*u[4] + s[14]*u[5]
+ s[15]*u[6]
du[8] = s[16]*u[6]
end
# parameters and initial conditions
pr = [1, 0.1, 0.570, 0.011, 0.456, 0.011, 0.171, 0.371, 0.125,
0.125, 0.012, 0.027, 0.003, 0.034, 0.034, 0.017, 0.017,
0.017, 1e7, 0, 0]
u0 = [1e7-83.333, 83.333, 0, 0, 0, 0, 0, 0]
icomp = 2:6
lockdown(SIDARTHE, u0, pr, icomp)
# -
# # Provenance
using Dates
println("mahdiar")
Dates.format(now(), "Y/U/d HH:MM")
| notebooks/.ipynb_checkpoints/Approximation-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# # Imports
from pathlib import Path
import os
import pandas as pd
import numpy as np
from typing import Set, Dict, Callable
# # Defining Constants
# +
# Define constants
CWD = os.path.abspath('')
CWD = Path(CWD)
CRISPR_DATA = CWD/'CRISPR_Data'
DEPMAP = CWD / 'DepMap'
SOD_LBL = 'PSOD_LT_FDR.05'
SOD_CUTOFFS = (.05, .1)
IPE_CUTOFFS = (.25, -.1)
IPE_LBL = 'PIPE_GT_.25FDR_-.1LFC'
COMM_ESS = 'Common_Essential'
MAIN_OUT = 'PEL_All_Data.csv'
SODS_OUT = 'PSODs.csv'
IPES_OUT = 'PIPEs.csv'
LRT_OUT = 'NormLRT.csv'
EXPORT_SETTINGS = {
'sep': ',',
'header': True,
'index': True
}
# -
# # Defining Functions
# +
def import_and_merge(dir: Path = CRISPR_DATA, suffix: str = '.csv',
sep: str = ',') -> pd.DataFrame:
"""
Imports MAGeCK data from multiple screens into one dataframe
Parameters
----------
dir : Path, optional
Directory containing MAGeCK data, by default CRISPR_DATA
file names should be meaningful (i.e. cell line/condition names)
suffix : str, optional
The file suffix of target MAGeCK Output Files, by default '.csv'
sep : str, optional
Field separator for MAGeCK data, by default ','
Returns
-------
pd.DataFrame
A dataframe with rows of genes and columns of MAGeCK depletion
metrics (NegFDR/LFC)
"""
df_dict ={}
for f in list(dir.glob(f'*{suffix}')):
cell_line = f.name.strip(suffix)
df_dict[cell_line] = pd.read_csv(f, sep=',', header=0, index_col=0)
for cell_line, df in df_dict.items():
new_cols = {
'neg|fdr': f'{cell_line}|NegFDR',
'neg|lfc': f'{cell_line}|NegLFC'
}
df.rename(columns=new_cols, inplace=True)
df_dict[cell_line] = df[[f'{cell_line}|NegFDR', f'{cell_line}|NegLFC']]
dfs_list = [v for v in df_dict.values()]
merged_df = pd.concat(dfs_list,axis=1,join="inner")
return merged_df
# Should be run on MAGeCK data before anything else
# Typically twice (Achilles+SCORE combined vs Achilles-only genes)
def check_comm_ess(df: pd.DataFrame, prefix: str, common_essentials: Set[str],
included_guides: Set[str]):
"""
Checks whether each gene is included in a subset of DepMap.
Modifies in-place, does not return.
Parameters
----------
df : pd.DataFrame
The input dataframe, containing MAGeCK screen data
prefix : str
A label to signify this DepMap subset (i.e. Achilles-only)
common_essentials : Set[str]
A set of guides designated as common essentials in a subset of DepMap.
included_guides : Set[str]
The set of guides included in this subset of DepMap.
"""
comm_ess = f'{prefix}_Comm_Ess'
df[comm_ess] = np.nan
df[f'{prefix}_hasGuide'] = df.index.isin(included_guides)
has_guide = df[f'{prefix}_hasGuide']
df.loc[has_guide, comm_ess] = df[has_guide].index.isin(common_essentials)
# Run after labeling common essentials from one or more datasets
def if_not_na_else(df: pd.DataFrame, new_col: str, if_not_na_col: str,
else_col: str):
"""
Creates a new column C in a "if A != NA, B" manner.
Modifies in-place, does not return.
Parameters
----------
df : pd.DataFrame
The input dataframe, containing MAGeCK screen data
new_col : str
Name of the new column (i.e. col C)
if_not_na_col : str
Name of the default column to use (if not NA) (i.e. col A)
else_col : str
Name of the backup column to use (i.e. col B)
False if there is no backup (propagates NAs from col A)
"""
non_null = df[if_not_na_col].isnull()
if else_col:
vals = np.where(non_null, df[else_col], df[if_not_na_col])
else:
df[new_col] = df[if_not_na_col]
df[new_col] = vals
# Must before labeling SODs
def lbl_highly_selective(df: pd.DataFrame,
highly_selectives: Set[str]):
"""
Labels genes as highly selective or not.
Modifies in-place, does not return.
Parameters
----------
df : pd.DataFrame
The input dataframe, containing MAGeCK screen data
highly_selectives : Set[str]
A set or other container to check for membership.
"""
df['Highly_Selective'] = False
df.loc[df.index.isin(highly_selectives),'Highly_Selective'] = True
# Must run lbl_highly_selective first
def lbl_cohort_sods(df: pd.DataFrame, output_col: str, fdr_thresh: float = .05,
lfc_thresh: float = 0, metric: str = 'median'):
"""
Labels genes as a cohort-specific oncogenic dependency (SOD) in the
non-DepMap cohort, based on thresholded filtering.
Modifies in-place, does not return.
Parameters
----------
df : pd.DataFrame
The input dataframe, containing MAGeCK screen data
output_col : str
Name for output column
fdr_thresh : float, optional
Maximum FDR to allow, by default .05
lfc_thresh : float, optional
Maximum LFC to allow, by default 0
metric : str, optional
How to aggregate cohort data (mean or median), by default 'median'
"""
fdrs = [i for i in df.columns if 'NegFDR' in i]
lfcs = [i for i in df.columns if 'NegLFC' in i]
if metric == 'median':
df['Median|NegFDR'] = np.median(df[fdrs], axis=1)
df['Median|NegLFC'] = np.median(df[lfcs], axis=1)
fdr_col = 'Median|NegFDR'
lfc_col = 'Median|NegLFC'
elif metric == 'mean':
df['Mean|NegFDR'] = np.mean(df[fdrs], axis=1)
df['Mean|NegLFC'] = np.mean(df[lfcs], axis=1)
fdr_col = 'Mean|NegFDR'
lfc_col = 'Mean|NegLFC'
not_comm_ess = df['Common_Essential'] == False
highly_specific = df['Highly_Selective'] == True
uncommon = np.logical_or(not_comm_ess, highly_specific)
meets_thresh = (df[fdr_col] <= fdr_thresh) & (df[lfc_col] <= lfc_thresh)
df[output_col] = np.where(uncommon & meets_thresh, True, False)
def lbl_cohort_ipes(df: pd.DataFrame, output_col: str, fdr_thresh: float = .25,
lfc_thresh: float = 0, metric: str = 'median') -> None:
"""
Labels genes as a cohort-insensitive (SOD) in the non-DepMap
cohort, based on filtering.
Parameters
----------
df : pd.DataFrame
The input dataframe, containing MAGeCK screen data
output_col : str
Name for output column
fdr_thresh : float, optional
Minimum FDR to allow, by default .25
lfc_thresh : float, optional
Minimum LFC to allow, by default 0
metric : str, optional
How to aggregate cohort data (mean or median), by default 'median'
"""
if metric == 'median':
fdr_col = 'Median|NegFDR'
lfc_col = 'Median|NegLFC'
elif metric == 'mean':
fdr_col = 'Mean|NegFDR'
lfc_col = 'Mean|NegLFC'
comm_ess = df['Common_Essential'] == True
meets_thresh = (df[fdr_col] >= fdr_thresh) & (df[lfc_col] >= lfc_thresh)
df[output_col] = np.where(comm_ess & meets_thresh, True, False)
# Used to return a df of NormLRT values; optional.
def run_r_func(func: Callable[[pd.DataFrame, dict], pd.DataFrame],
ge_tables: Dict[str, pd.DataFrame]) -> pd.DataFrame:
"""
Calculates NormLRT values from CHRONOS data (assumes no overlap in genes)
Parameters
----------
func : Callable[[pd.DataFrame, dict], pd.DataFrame]
An R function (intended to be LRT_test from DepMap consortium)
Must be defined properly with RPy2 or equivalent. See norm_lrt.py
ge_tables : Dict[str, pd.DataFrame]
A dictionary of CHRONOS dataframes (rows screens/columns genes).
Dictionary key is used for progress bar.
It is assumed that dataframes have already been filtered to avoid
overlap between genes.
Returns
-------
pd.DataFrame
A dataframe, with genes as rows and a single 'NormLRT' when used with
the LRT_test() function from DepMap consortium.
"""
lrts = []
for k,v in ge_tables.items():
lrts.append(func(v, k))
lrt_df = pd.concat(lrts, axis=0, join='inner', verify_integrity=True)
return lrt_df
# -
# # Loading and examining data
# +
#### Load in DepMap data
print("Loading and processing DepMap data")
# CRISPR (Achilles + Score)
crispr_comm_essential = pd.read_csv(DEPMAP/'CRISPR_common_essentials.csv')
crispr_comm_essential = crispr_comm_essential['gene'].tolist()
crispr_comm_essential = [g.split(' ')[0] for g in crispr_comm_essential]
crispr_ge = pd.read_csv(DEPMAP/'CRISPR_gene_effect.csv', index_col=0)
renames = {c:c.split(' ')[0] for c in crispr_ge.columns}
crispr_ge.rename(columns=renames,inplace=True)
crispr_guides = set(crispr_ge.columns)
# Achilles(Achilles)
ach_comm_essential = pd.read_csv(DEPMAP/'Achilles_common_essentials.csv')
ach_comm_essential = ach_comm_essential['gene'].tolist()
ach_comm_essential = [g.split(' ')[0] for g in ach_comm_essential]
ach_ge = pd.read_csv(DEPMAP/'Achilles_gene_effect.csv', index_col=0)
renames = {c:c.split(' ')[0] for c in ach_ge.columns}
ach_ge.rename(columns=renames,inplace=True)
ach_guides = set(ach_ge.columns)
depmap_guides = crispr_guides.union(ach_guides)
#Subset Achille's GE matrix to exclude CRISPR (Achilles+Score) genes.
ach_ge = ach_ge[[c for c in ach_ge.columns if c not in crispr_guides]]
#Load DepMap list of highly-selective genes
hs_genes = pd.read_csv(DEPMAP/'DepMap_Selective_Genes.csv')
hs_genes = set(hs_genes['gene'].values)
# +
ach_only_comm_essential = set([g for g in ach_comm_essential if g not in crispr_guides])
all_comm_essentials = ach_only_comm_essential.union(crispr_comm_essential)
hs_comm_essentials = all_comm_essentials.intersection(hs_genes)
print(f'There are {len(all_comm_essentials)} common essential genes in this DepMap release.')
print(f'There are {len(hs_genes)} highly-selective genes.')
print(f'There are {len(hs_comm_essentials)} highly-selective, common essential genes.')
# -
print("Loading and processing MAGeCK data")
merged_df = import_and_merge()
merged_df.head()
# +
cohort_genes = set(merged_df.index)
overlap_comm = all_comm_essentials.intersection(cohort_genes)
overlap_hs = hs_genes.intersection(cohort_genes)
overlap_hs_comm = hs_comm_essentials.intersection(cohort_genes)
print(f'{len(overlap_comm)} of {len(all_comm_essentials)} DepMap common essentials are in the cohort library.')
print(f'{len(overlap_hs)} of {len(hs_genes)} DepMap highly-selective genes are in the cohort library')
print(f'{len(overlap_hs_comm)} of {len(hs_comm_essentials)} DepMap highly-selective, common essentials genes are in the cohort library')
# +
# Check if highly LRTs have been calculated
LRT_FILE = CWD / LRT_OUT
if LRT_FILE .is_file():
print('Loading pre-calculated LRT values')
lrt_initialized = True
lrt_df = pd.read_csv(LRT_FILE, sep=',', index_col=0)
else:
from norm_lrt import lrt_test
# -
# # Analysis/Filtering
print("Beginning analysis")
check_comm_ess(merged_df, 'Ach+SCORE', crispr_comm_essential, crispr_guides)
merged_df.head()
check_comm_ess(merged_df, 'Ach', ach_comm_essential, ach_guides)
merged_df.head()
if_not_na_else(merged_df, COMM_ESS, 'Ach+SCORE_Comm_Ess', 'Ach_Comm_Ess')
merged_df.head()
# +
if not(lrt_initialized):
lrt_df = run_r_func(_lrt_test, {'Achilles+Score': crispr_ge, 'Achilles-only': ach_ge})
lrt_df = lrt_df[lrt_df.index.isin(merged_df.index)]
lrt_df.head()
# -
merged_df = pd.concat([merged_df, lrt_df], axis=1, join='outer')
merged_df.head()
lbl_highly_selective(merged_df, hs_genes)
merged_df.head()
lbl_cohort_sods(merged_df, SOD_LBL, *SOD_CUTOFFS)
merged_df.head()
lbl_cohort_ipes(merged_df, IPE_LBL, *IPE_CUTOFFS)
merged_df.head()
# # Export results and summarize
# +
print("Exporting reports")
merged_df.to_csv(CWD/MAIN_OUT, **EXPORT_SETTINGS)
norm_lrt_df = merged_df['Skew_LRT']
norm_lrt_df.to_csv(CWD / LRT_FILE, sep=',', index=True)
# -
sods = merged_df.loc[merged_df[SOD_LBL] == True]
sods = sods.drop(columns=[IPE_LBL])
has_guides = sods.index.isin(depmap_guides)
high_conf = sods.loc[has_guides]
high_conf = high_conf.loc[sods[COMM_ESS] == False]
high_conf.head()
comm_ess = sods[COMM_ESS] == True
print(f'{len(high_conf)} high-confidence SODs')
is_hs = high_conf['Highly_Selective'] == True
print(f'{len(high_conf[is_hs])} high-confidence, highly-selective SODs')
high_conf.head()
high_conf.groupby(['Highly_Selective'])['Highly_Selective'].agg('count')
# +
putative = sods[np.logical_or(comm_ess, ~has_guides)].copy()
putative['Reason'] = ''
putative.loc[comm_ess, 'Reason'] = 'Pan-Essential'
putative.loc[~comm_ess, 'Reason'] = 'No Data'
# -
print(f'{len(putative)} putative SODs.')
putative.head()
putative.groupby(['Reason'])['Reason'].agg('count')
putative.groupby(['Highly_Selective']).agg('count')
high_conf_out = 'HighConf_'+SODS_OUT
putative_out = 'Putative_'+SODS_OUT
high_conf.to_csv(CWD/high_conf_out, **EXPORT_SETTINGS)
putative.to_csv(CWD/putative_out, **EXPORT_SETTINGS)
ipes = merged_df[merged_df[IPE_LBL] == True]
ipes = ipes.drop(columns=[SOD_LBL])
ipes.to_csv(CWD/IPES_OUT, **EXPORT_SETTINGS)
print(f'{len(ipes)} IPEs')
is_hs = ipes['Highly_Selective'] == True
print(f'{len(ipes[is_hs])} highly-selective IPEs')
ipes.head()
print("Complete!")
| MAGeCK_DepMap_Comparisons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# name: ir
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/hongqin/Use-R-in-CoLab/blob/master/simple_stat_election20.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 786} id="BJV8y8TApBHE" outputId="5cbc88ab-dec3-4d54-aafe-3cf233380740"
library(tidyverse)
Election20df = read_csv("https://raw.githubusercontent.com/tonmcg/US_County_Level_Election_Results_08-20/master/2020_US_County_Level_Presidential_Results.csv")
head(Election20df)
# + id="7C5gJ-T1pZF1"
# Question: What is the total number of votes?
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="LztQJtzGpMOy" outputId="29f00109-f6fc-410e-9adb-c99e13402d53"
Election20df %>% select ( total_votes ) %>% sum()
sum( Election20df$total_votes ) /1E6
# + [markdown] id="CM6zcB9FpcsN"
# # Question: What are the total number votes for GOP or DEM?
#
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="lk0NvppnpkUg" outputId="c8255a94-8384-45ba-ca63-fa867a82f7d0"
Election20df %>% select ( votes_gop ) %>% sum()
Election20df %>% select ( votes_dem ) %>% sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="PMkFa4BDpsp9" outputId="806f0088-c1c8-4183-f637-a9ef3dcba698"
mystate = "California"
Californiadf <-
Election20df %>% filter( state_name == mystate) %>% arrange( per_point_diff)
names( Californiadf )[8] = "percentage_for_GOP"
ggplot(Californiadf, aes(percentage_for_GOP)) + geom_histogram()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="qaHlEZn9ptl8" outputId="acabf880-5b3d-4bbf-9620-f637d5c574d5"
Statedf <-
Election20df %>% select( state_name, votes_gop, votes_dem, total_votes ) %>% group_by( state_name ) %>% summarise_if( is.numeric, sum)
Statedf$percentage_for_GOP = Statedf$votes_gop / Statedf$total_votes
ggplot(Statedf, aes(percentage_for_GOP)) + geom_histogram()
# + colab={"base_uri": "https://localhost:8080/", "height": 129} id="QX0CnXGFpz8r" outputId="2039e71f-7b8e-470c-96e0-309fab90ca1e"
# Question: Find out which state has the nearly 95% DEM voting percentage?
# There are many ways to do this.
Statedf %>% filter( percentage_for_GOP < 0.1 )
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="jtmK34yTp38V" outputId="ede4bf6e-26df-4450-bc7c-19646f522d42"
mean( Statedf$percentage_for_GOP) #average
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="1NB2kah2p-wX" outputId="accf68ea-271d-439e-9a3a-edb8f82c5484"
quantile( Statedf$percentage_for_GOP )
# + [markdown] id="Mir7OPJRqHI9"
# # Label the states to deep red, red, swing, blue, deep blue
#
# Reference:
# https://stackoverflow.com/questions/21050021/create-category-based-on-range-in-r
#
# + id="bwtxjr2aqAcL"
groups = cut( Statedf$percentage_for_GOP, c(0, 0.4, 0.47, 0.53, 0.6, 1) )
levels(groups) = c("deepblue", "blue", "swing", "red", "deepred")
Statedf$groups = groups
# + colab={"base_uri": "https://localhost:8080/", "height": 579} id="9pEz7KyfqKKn" outputId="03afa6dd-f438-459c-e02b-b51b372594fc"
Censusdf = read_csv("https://raw.githubusercontent.com/hongqin/USA-census-county-level/main/USA-County-level-census-2010-2019.csv")
head(Censusdf)
# + colab={"base_uri": "https://localhost:8080/", "height": 276} id="qlwB-n60qNMf" outputId="17e66947-a862-4351-c6f6-60ca00181304"
Election20df$Location = paste( Election20df$county_name, Election20df$state_name, sep=", " )
Election20df$Location %in% Censusdf$Location
# + id="gMMWHz0JqQlf"
EleCen.df = merge( Election20df, Censusdf, by="Location")
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="Yf_-NYlPqUGI" outputId="7e058f41-c8a5-4f64-dd80-fca10de875ed"
Statedf2 <- EleCen.df %>% select( state_name, votes_gop, votes_dem, total_votes, '2019' ) %>% group_by( state_name ) %>% summarise_if( is.numeric, sum)
head(Statedf2)
names( Statedf2)[5] = "population"
# + id="lXKtC_nFqWdf"
Statedf$population = Statedf2$population[match( Statedf$state_name , Statedf2$state_name ) ]
# + colab={"base_uri": "https://localhost:8080/", "height": 345} id="BXJn3C2IqfpA" outputId="b09a37d1-813d-406a-d1d4-611cd2d80468"
model1 = lm( Statedf$percentage_for_GOP ~ Statedf$population)
summary(model1)
# + colab={"base_uri": "https://localhost:8080/", "height": 852} id="o9fdrwAUqhq0" outputId="99e520f4-56f5-43ed-d676-77264d8c303e"
model2 = lm( Statedf$population ~ Statedf$groups)
summary(model2)
ggplot( Statedf, aes(x=groups, y=population)) + geom_point()
# + colab={"base_uri": "https://localhost:8080/"} id="DQvkFzhbqkkW" outputId="53870142-0d4f-4861-c4cc-8accd0225723"
StateArea = read_csv("https://raw.githubusercontent.com/hongqin/data-USstates/master/state-areas.csv")
names( StateArea) = c("state_name", "area")
# + colab={"base_uri": "https://localhost:8080/", "height": 345} id="Euukp1giqoYM" outputId="9f3d73a4-e2ea-4fc6-f255-c09d1fdd038a"
Statedf$area = StateArea$area[ match( Statedf$state_name , StateArea$state_name ) ]
Statedf$pop_density = Statedf$population / Statedf$area
model = lm( Statedf$percentage_for_GOP ~ Statedf$pop_density)
summary(model)
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="YUGCpGx-qq4Z" outputId="88b37be7-fe6b-4822-afbc-c63af1a64043"
ggplot( Statedf, aes(x=pop_density, y=percentage_for_GOP)) + geom_point()
# + colab={"base_uri": "https://localhost:8080/", "height": 541} id="O-n397roqtOk" outputId="3a9b35a1-44c1-496d-89f6-ee8df85c2ce1"
Statedf3 <- Statedf %>% filter( percentage_for_GOP > 0.1)
ggplot( Statedf3, aes(x=pop_density, y=percentage_for_GOP)) +
geom_point() +
geom_smooth(method='lm')
# + colab={"base_uri": "https://localhost:8080/", "height": 345} id="JGCJTB_-qwGW" outputId="0146eb98-f502-4b36-b20a-90c4129c8288"
summary(lm(Statedf3$percentage_for_GOP~ Statedf3$pop_density))
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="XmdqY57WqzbW" outputId="4e7a7ec2-202f-4b9d-86c2-59b34b282015"
ggplot( Statedf3, aes(x=groups, y=pop_density)) + geom_point()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="w8DrnMa6q2NG" outputId="000cb22b-ad26-4b16-b164-efe4a1422048"
ggplot( Statedf3, aes(x=groups, y=pop_density)) + geom_boxplot()
# + colab={"base_uri": "https://localhost:8080/", "height": 207} id="gARTlZbaq5qZ" outputId="c84a29ad-8210-4bf0-c113-8b3e039484c4"
deepred_pop_densities <-
Statedf3 %>% filter( groups == "deepred") %>% select( pop_density)
deepblue_pop_densities <-
Statedf3 %>% filter( groups == "deepblue") %>% select( pop_density)
t.test( deepblue_pop_densities, deepred_pop_densities, alternative = "greater")
# + colab={"base_uri": "https://localhost:8080/", "height": 437} id="aaTYvrauq943" outputId="b73b8f11-e4b3-401f-e2e6-040aaef3b614"
ggplot(Statedf, aes(x=state_name, y=percentage_for_GOP)) + geom_bar(stat='identity', width=.5)+ theme(axis.text.x = element_text(angle = 90, vjust = 0.5, hjust=1))
# + colab={"base_uri": "https://localhost:8080/", "height": 437} id="R9yRQ93XrA0U" outputId="02f8a1fe-b706-4991-c889-005a5c0f3d69"
ggplot(Statedf, aes(x=reorder(state_name, percentage_for_GOP), y=percentage_for_GOP)) + geom_bar(stat='identity', width=.5)+ theme(axis.text.x = element_text(angle = 90, vjust = 0.5, hjust=1))
# + id="dKZekocpvp7F"
| simple_stat_election20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 33} colab_type="code" id="gEsmboX0vW9X" outputId="e96e2a3c-ebc7-4923-90aa-ab9329fff93e"
import cv2
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.models import Model
from tensorflow.keras.regularizers import l2
from tensorflow.compat.v1 import ConfigProto
from sklearn.preprocessing import LabelBinarizer
from tensorflow.compat.v1 import InteractiveSession
from sklearn.model_selection import train_test_split
from tensorflow.keras.layers import Dense, BatchNormalization
from tensorflow.keras.applications.densenet import DenseNet121
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
# + colab={"base_uri": "https://localhost:8080/", "height": 304} colab_type="code" id="muuwkPL069DC" outputId="a9a7e223-cb81-48bf-fe37-7a89d854a384"
model = DenseNet121(include_top = False, weights = 'imagenet', input_shape=(128,128,3), pooling = 'max')
model.trainable = False
set_trainable = False
for layer in model.layers:
if layer.name.startswith("conv5"):
set_trainable = True
if set_trainable:
layer.trainable = True
else:
layer.trainable = False
x = model.output
bin_classifier = Dense(64, kernel_regularizer = l2(0.05), activation = "relu")(x)
bin_classifier = BatchNormalization(axis = -1)(bin_classifier)
bin_classifier = Dense(1, kernel_regularizer = l2(0.05), activation = "sigmoid")(bin_classifier)
reg_head = Dense(64, kernel_regularizer = l2(0.05), activation = "relu")(x)
reg_head = BatchNormalization(axis = -1)(reg_head)
reg_head = Dense(1, name = "reg_head", kernel_regularizer = l2(0.05))(reg_head)
base_model = Model(model.input, [bin_classifier, reg_head])
base_model.summary()
# + colab={} colab_type="code" id="DW9gHpKgDcVb"
images = np.load('../imfdb-dataset/imfdb_images.npy')
gender = np.load('../imfdb-dataset/imfdb_gender_labels.npy')
age = np.load('../imfdb-dataset/imfdb_age_labels.npy')
# + colab={} colab_type="code" id="i6F1xc6fEBiJ"
images = images.astype("float") / 255.0
# + colab={} colab_type="code" id="BEt2AgjyEEtx"
(x_train, x_test, gender_train, gender_test, age_train, age_test) = \
train_test_split(images, gender, age, test_size = 0.2, random_state = 42)
# + colab={} colab_type="code" id="cCzqTlKsEEyZ"
lb = LabelBinarizer()
gender_train = lb.fit_transform(gender_train)
gender_test = lb.fit_transform(gender_test)
# + colab={} colab_type="code" id="cVku12hrEE25"
loss_weights = {'reg_head': 1., 'bin_classifier': 6.}
losses = {'reg_head': 'mse', 'bin_classifier': 'binary_crossentropy'}
# + colab={"base_uri": "https://localhost:8080/", "height": 120} colab_type="code" id="Ext7oTz3EE7I" outputId="d483c93a-796d-4d51-c986-4e356236d7f3"
base_model.compile(optimizer = "nadam", loss = losses, loss_weights = loss_weights, metrics = ["acc"])
# + colab={} colab_type="code" id="IqkY63SgFKTw"
chkpt = ModelCheckpoint(filepath = "model_initial.h5", monitor = 'val_bin_classifier_acc',\
mode = 'max', save_best_only = True, verbose = 1)
early = EarlyStopping(monitor = "val_acc", mode = "max", patience = 5, verbose = 1)
redonplat = ReduceLROnPlateau(monitor = "val_acc", mode = "max", patience = 3, verbose = 2)
callbacks_list = [chkpt, early, redonplat]
# + colab={} colab_type="code" id="EkJf0qpYFKWw"
epochs = 25
batch_size = 16
# + colab={"base_uri": "https://localhost:8080/", "height": 406} colab_type="code" id="jcKL7mSMFKZg" outputId="892a476d-1b4d-4d72-99d0-81dce8aa13d3"
base_model.fit(x_train, {'reg_head': age_train, 'bin_classifier': gender_train}, batch_size = batch_size, epochs = epochs, callbacks = callbacks_list, validation_data = (x_test, {'reg_head': age_test, 'bin_classifier': gender_test}))
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="zOwwzW2lP8R6" outputId="06824a06-188f-4f8e-e424-9676ddcc8bf1"
np.unique(age_train)
| Trained/Age_Gender_Module_continous.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(1, 'C:/Users/peter/Desktop/volatility-forecasting/midas')
from volatility import Panel_GARCH, Panel_GARCH_SLSQP, GARCH
from stats import panel_DM, dm_test
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import t
import math
# -
# # First let's check what can we observe in the standard DM test:
np.random.seed(14)
params = [0.0, 0.1, 0.05, 0.85]
model = GARCH()
r, sigma2 = model.simulate(params = params)
model.fit(['', '01', '01', '01'], r)
pred1 = model.model_filter(model.optimized_params, r)
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.2, size = pred1.shape)
pred2 = pred1 * eps
plt.figure(figsize =(15,4))
plt.plot(sigma2, label = 'Actual')
plt.plot(pred1, label = 'Estimated')
plt.plot(pred2, label = 'Estimated with noise')
plt.legend(loc = 'best')
plt.grid()
plt.tight_layout()
plt.show()
# Negative number means that the first prediction is better then the second:
dm_test(sigma2, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, r)
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.05, size = pred1.shape)
pred2 = pred1 * eps
plt.figure(figsize =(15,4))
plt.plot(sigma2, label = 'Actual')
plt.plot(pred1, label = 'Estimated')
plt.plot(pred2, label = 'Estimated with noise')
plt.legend(loc = 'best')
plt.grid()
plt.tight_layout()
plt.show()
# With that much noise the first prediction is much better:
dm_test(sigma2, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, r)
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.01, size = pred1.shape)
pred2 = pred1 * eps
plt.figure(figsize =(15,4))
plt.plot(sigma2, label = 'Actual')
plt.plot(pred1, label = 'Estimated')
plt.plot(pred2, label = 'Estimated with noise')
plt.legend(loc = 'best')
plt.grid()
plt.tight_layout()
plt.show()
# The null hypothesis can be accepted if the added noise is that much as you can see above
dm_test(sigma2, pred1, pred2)
# # Now, let's turn into the panel version:
# +
def family_of_loss_func(actual, predicted, degree):
"""
Implemented from:
<NAME>., 2011. Volatility forecasting comparison using imperfect
volatility proxies, Journal of Econometrics 160, 246-256.
"""
if degree == -2:
# QLIKE
loss = actual / predicted - np.log(actual / predicted) - 1
elif degree == -1:
loss = predicted - actual + actual * np.log(actual / predicted)
else:
# MSE if degree = 0
loss = (np.sqrt(actual) ** (2 * degree + 4) - predicted ** (degree + 2)) / ((degree + 1) * (degree + 2))
loss -= (1 / (degree + 1)) * (predicted ** (degree + 1)) * (actual - predicted)
return loss
def panel_DM_V1(act, pred1, pred2, degree = 0):
"""
Implemented from:
<NAME>., <NAME>., 2019. Comparing Forecasting Performance with Panel Data
"""
l1 = family_of_loss_func(act, pred1, degree)
l2 = family_of_loss_func(act, pred2, degree)
l1_mean = np.nanmean(l1)
l2_mean = np.nanmean(l2)
delta_l = l1 - l2
n_T_sqrt = np.sqrt(np.sum(~np.isnan(delta_l)))
sum_delta_l = np.nansum(delta_l)
sigma_delta_l = np.nanstd(delta_l)
j_dm = sum_delta_l / (sigma_delta_l * n_T_sqrt)
p_value = 2 * t.cdf(-np.abs(j_dm), df = np.sum(~np.isnan(delta_l)) - 1)
return j_dm, p_value
def panel_DM_V2(act, pred1, pred2, degree = 0):
l1 = family_of_loss_func(act, pred1, degree)
l2 = family_of_loss_func(act, pred2, degree)
d12 = l1 - l2
nT = np.sum(~np.isnan(d12))
d12_m = np.nanmean(d12, axis = 0)
# Itt azért osztunk csak nT-vel és nem np.sqrt(nT)-vel, mert
# amikor az nT-t kihozzuk a Var-ból akkor annak a négyzetével számolunk
var = np.nansum((d12 - np.nanmean(d12, axis = 0)) * (d12 - np.nanmean(d12, axis = 0))) / nT
std = math.sqrt(var)
m = np.nansum(d12) / math.sqrt(nT)
DM = m / std
p_value = 2 * t.cdf(-np.abs(DM), df = nT-1)
return DM, p_value
def panel_DM_V3(act, pred1, pred2, degree = 0):
l1 = family_of_loss_func(act, pred1, degree)
l2 = family_of_loss_func(act, pred2, degree)
d12 = l1 - l2
nT = np.sum(~np.isnan(d12))
n = np.sum(~np.isnan(d12), axis = 1)
T = d12.shape[0]
Rt = np.sqrt(n) * np.nansum(d12, axis = 1) / n
Rt_hat = Rt - np.nansum(Rt) / T
gamma0 = np.nansum(Rt_hat * Rt_hat) / T
Rt1 = Rt[1:] - np.nansum(Rt[1:]) / (T - 1)
Rt11 = Rt[:-1] - np.nansum(Rt[:-1]) / (T - 1)
gamma1 = np.nansum(Rt1 * Rt11) / (T - 1)
std = math.sqrt(gamma0 + 2 * gamma1)
m = np.nansum(d12) / np.sqrt(nT)
DM = m / std
p_value = 2 * t.cdf(-np.abs(DM), df = nT-1)
return DM, p_value
# -
np.random.seed(14)
params = [0.05, 0.9]
model = Panel_GARCH()
act, r = model.simulate(params = params)
model.fit(['01', '01'], pd.DataFrame(r))
pred1 = model.model_filter(model.optimized_params, pd.DataFrame(r))
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.2, size = pred1.shape)
pred2 = pred1 * eps
panel_DM(act, pred1, pred2)
panel_DM_V1(act, pred1, pred2)
panel_DM_V2(act, pred1, pred2)
panel_DM_V3(act, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, pd.DataFrame(r))
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.05, size = pred1.shape)
pred2 = pred1 * eps
panel_DM(act, pred1, pred2)
panel_DM_V1(act, pred1, pred2)
panel_DM_V2(act, pred1, pred2)
panel_DM_V3(act, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, pd.DataFrame(r))
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.01, size = pred1.shape)
pred2 = pred1 * eps
panel_DM(act, pred1, pred2)
panel_DM_V1(act, pred1, pred2)
panel_DM_V2(act, pred1, pred2)
panel_DM_V3(act, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, pd.DataFrame(r))
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.005, size = pred1.shape)
pred2 = pred1 * eps
panel_DM(act, pred1, pred2)
panel_DM_V1(act, pred1, pred2)
panel_DM_V2(act, pred1, pred2)
panel_DM_V3(act, pred1, pred2)
pred1 = model.model_filter(model.optimized_params, pd.DataFrame(r))
np.random.seed(2)
eps = np.ones(pred1.shape) + np.random.normal(scale = 0.001, size = pred1.shape)
pred2 = pred1 * eps
panel_DM(act, pred1, pred2)
panel_DM_V1(act, pred1, pred2)
panel_DM_V2(act, pred1, pred2)
panel_DM_V3(act, pred1, pred2)
| Examples/Panel_DM_backtest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Задание 2.2 - Введение в PyTorch
#
# Для этого задания потребуется установить версию PyTorch 1.0
#
# https://pytorch.org/get-started/locally/
#
# В этом задании мы познакомимся с основными компонентами PyTorch и натренируем несколько небольших моделей.<br>
# GPU нам пока не понадобится.
#
# Основные ссылки:
# https://pytorch.org/tutorials/beginner/deep_learning_60min_blitz.html
# https://pytorch.org/docs/stable/nn.html
# https://pytorch.org/docs/stable/torchvision/index.html
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision.datasets as dset
from torch.utils.data.sampler import SubsetRandomSampler, Sampler
from torchvision import transforms
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# -
# ## Как всегда, начинаем с загрузки данных
#
# PyTorch поддерживает загрузку SVHN из коробки.
# First, lets load the dataset
data_train = dset.SVHN('./data/', split='train',
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
])
)
data_test = dset.SVHN('./data/', split='test',
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean=[0.43,0.44,0.47],
std=[0.20,0.20,0.20])
]))
# Теперь мы разделим данные на training и validation с использованием классов `SubsetRandomSampler` и `DataLoader`.
#
# `DataLoader` подгружает данные, предоставляемые классом `Dataset`, во время тренировки и группирует их в батчи.
# Он дает возможность указать `Sampler`, который выбирает, какие примеры из датасета использовать для тренировки. Мы используем это, чтобы разделить данные на training и validation.
#
# Подробнее: https://pytorch.org/tutorials/beginner/data_loading_tutorial.html
# +
batch_size = 64
data_size = data_train.data.shape[0]
validation_split = .2
split = int(np.floor(validation_split * data_size))
indices = list(range(data_size))
np.random.shuffle(indices)
train_indices, val_indices = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_indices)
val_sampler = SubsetRandomSampler(val_indices)
train_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=train_sampler)
val_loader = torch.utils.data.DataLoader(data_train, batch_size=batch_size,
sampler=val_sampler)
# -
# В нашей задаче мы получаем на вход изображения, но работаем с ними как с одномерными массивами. Чтобы превратить многомерный массив в одномерный, мы воспользуемся очень простым вспомогательным модулем `Flattener`.
# +
sample, label = data_train[0]
print("SVHN data sample shape: ", sample.shape)
# As you can see, the data is shaped like an image
# We'll use a special helper module to shape it into a tensor
class Flattener(nn.Module):
def forward(self, x):
batch_size, *_ = x.shape
return x.view(batch_size, -1)
# -
# И наконец, мы создаем основные объекты PyTorch:
# - `nn_model` - собственно, модель с нейросетью
# - `loss` - функцию ошибки, в нашем случае `CrossEntropyLoss`
# - `optimizer` - алгоритм оптимизации, в нашем случае просто `SGD`
# +
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 10),
)
nn_model.type(torch.FloatTensor)
# We will minimize cross-entropy between the ground truth and
# network predictions using an SGD optimizer
loss = nn.CrossEntropyLoss().type(torch.FloatTensor)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-2, weight_decay=1e-1)
# -
# ## Тренируем!
#
# Ниже приведена функция `train_model`, реализующая основной цикл тренировки PyTorch.
#
# Каждую эпоху эта функция вызывает функцию `compute_accuracy`, которая вычисляет точность на validation, эту последнюю функцию предлагается реализовать вам.
# +
# This is how to implement the same main train loop in PyTorch. Pretty easy, right?
def train_model(model, train_loader, val_loader, loss, optimizer, num_epochs, scheduler=False):
loss_history = []
train_history = []
val_history = []
for epoch in range(num_epochs):
model.train() # Enter train mode
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y) in enumerate(train_loader):
prediction = model(x)
loss_value = loss(prediction, y)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
_, indices = torch.max(prediction, 1)
correct_samples += torch.sum(indices == y)
total_samples += y.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / (i_step + 1)
train_accuracy = float(correct_samples) / total_samples
val_accuracy = compute_accuracy(model, val_loader)
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
val_history.append(val_accuracy)
if scheduler:
scheduler.step()
print("Average loss: %f, Train accuracy: %f, Val accuracy: %f" % (ave_loss, train_accuracy, val_accuracy))
return loss_history, train_history, val_history
def compute_accuracy(model, loader):
"""
Computes accuracy on the dataset wrapped in a loader
Returns: accuracy as a float value between 0 and 1
"""
model.eval() # Evaluation mode
# TODO: Implement the inference of the model on all of the batches from loader,
# and compute the overall accuracy.
# Hint: PyTorch has the argmax function!
accuracies = []
for i, (x, y) in enumerate(loader):
probabilities = model(x)
prediction = torch.argmax(probabilities,dim=1)
accuracies.append(len(y[prediction==y])/len(y))
return np.mean(accuracies)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 3)
# -
# ## После основного цикла
#
# Посмотрим на другие возможности и оптимизации, которые предоставляет PyTorch.
#
# Добавьте еще один скрытый слой размера 100 нейронов к модели
# +
# Since it's so easy to add layers, let's add some!
# TODO: Implement a model with 2 hidden layers of the size 100
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.ReLU(inplace=True),
nn.Linear(100,10),
)
nn_model.type(torch.FloatTensor)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-2, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5)
# -
# Добавьте слой с Batch Normalization
# +
# We heard batch normalization is powerful, let's use it!
# TODO: Add batch normalization after each of the hidden layers of the network, before or after non-linearity
# Hint: check out torch.nn.BatchNorm1d
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100,10),
)
#nn_model.type(torch.FloatTensor)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-3, weight_decay=1e-1)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5)
# -
# Добавьте уменьшение скорости обучения по ходу тренировки.
# +
# Learning rate annealing
# Reduce your learning rate 2x every 2 epochs
# Hint: look up learning rate schedulers in PyTorch. You might need to extend train_model function a little bit too!
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100, 100),
nn.BatchNorm1d(100),
nn.ReLU(inplace=True),
nn.Linear(100,10),
)
optimizer = optim.SGD(nn_model.parameters(), lr=1e-3, weight_decay=1e-1)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=2, gamma=0.5)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5, scheduler)
# -
# # Визуализируем ошибки модели
#
# Попробуем посмотреть, на каких изображениях наша модель ошибается.
# Для этого мы получим все предсказания модели на validation set и сравним их с истинными метками (ground truth).
#
# Первая часть - реализовать код на PyTorch, который вычисляет все предсказания модели на validation set.
# Чтобы это сделать мы приводим код `SubsetSampler`, который просто проходит по всем заданным индексам последовательно и составляет из них батчи.
#
# Реализуйте функцию `evaluate_model`, которая прогоняет модель через все сэмплы validation set и запоминает предсказания модели и истинные метки.
# +
class SubsetSampler(Sampler):
r"""Samples elements with given indices sequentially
Arguments:
indices (ndarray): indices of the samples to take
"""
def __init__(self, indices):
self.indices = indices
def __iter__(self):
return (self.indices[i] for i in range(len(self.indices)))
def __len__(self):
return len(self.indices)
def evaluate_model(model, dataset, indices):
"""
Computes predictions and ground truth labels for the indices of the dataset
Returns:
predictions: np array of ints - model predictions
grount_truth: np array of ints - actual labels of the dataset
"""
model.eval() # Evaluation mode
# TODO: Evaluate model on the list of indices and capture predictions
# and ground truth labels
# Hint: SubsetSampler above could be useful!
predictions = []
ground_truth = []
#for i in indices:
probabilities = model(torch.FloatTensor(dataset.data[indices]))
predictions = np.array(torch.argmax(probabilities,dim=1).double())
ground_truth = np.array(dataset.labels[indices])
return predictions, ground_truth
# Evaluate model on validation
predictions, gt = evaluate_model(nn_model, data_train, val_indices)
assert len(predictions) == len(val_indices)
assert len(gt) == len(val_indices)
assert gt[100] == data_train[val_indices[100]][1]
assert np.any(np.not_equal(gt, predictions))
# -
# ## Confusion matrix
# Первая часть визуализации - вывести confusion matrix (https://en.wikipedia.org/wiki/Confusion_matrix ).
#
# Confusion matrix - это матрица, где каждой строке соответствуют классы предсказанный, а столбцу - классы истинных меток (ground truth). Число с координатами `i,j` - это количество сэмплов класса `j`, которые модель считает классом `i`.
#
# 
#
# Для того, чтобы облегчить вам задачу, ниже реализована функция `visualize_confusion_matrix` которая визуализирует такую матрицу.
# Вам осталось реализовать функцию `build_confusion_matrix`, которая ее вычислит.
#
# Результатом должна быть матрица 10x10.
# +
def visualize_confusion_matrix(confusion_matrix):
"""
Visualizes confusion matrix
confusion_matrix: np array of ints, x axis - predicted class, y axis - actual class
[i][j] should have the count of samples that were predicted to be class i,
but have j in the ground truth
"""
# Adapted from
# https://stackoverflow.com/questions/2897826/confusion-matrix-with-number-of-classified-misclassified-instances-on-it-python
assert confusion_matrix.shape[0] == confusion_matrix.shape[1]
size = confusion_matrix.shape[0]
fig = plt.figure(figsize=(10,10))
plt.title("Confusion matrix")
plt.ylabel("predicted")
plt.xlabel("ground truth")
res = plt.imshow(confusion_matrix, cmap='GnBu', interpolation='nearest')
cb = fig.colorbar(res)
plt.xticks(np.arange(size))
plt.yticks(np.arange(size))
for i, row in enumerate(confusion_matrix):
for j, count in enumerate(row):
plt.text(j, i, count, fontsize=14, horizontalalignment='center', verticalalignment='center')
def build_confusion_matrix(predictions, ground_truth):
"""
Builds confusion matrix from predictions and ground truth
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
Returns:
np array of ints, (10,10), counts of samples for predicted/ground_truth classes
"""
confusion_matrix = np.zeros((10,10), np.int)
for i in range(confusion_matrix.shape[0]):
for j in range(confusion_matrix.shape[1]):
confusion_matrix[i,j] = np.sum(predictions[ground_truth == j] == i)
# TODO: Implement filling the prediction matrix
return confusion_matrix
confusion_matrix = build_confusion_matrix(predictions, gt)
visualize_confusion_matrix(confusion_matrix)
# -
# Наконец, посмотрим на изображения, соответствующие некоторым элементам этой матрицы.
#
# Как и раньше, вам дана функция `visualize_images`, которой нужно воспрользоваться при реализации функции `visualize_predicted_actual`. Эта функция должна вывести несколько примеров, соответствующих заданному элементу матрицы.
#
# Визуализируйте наиболее частые ошибки и попробуйте понять, почему модель их совершает.
# +
data_train_images = dset.SVHN('./data/', split='train')
def visualize_images(indices, data, title='', max_num=10):
"""
Visualizes several images from the dataset
indices: array of indices to visualize
data: torch Dataset with the images
title: string, title of the plot
max_num: int, max number of images to display
"""
to_show = min(len(indices), max_num)
fig = plt.figure(figsize=(10,1.5))
fig.suptitle(title)
for i, index in enumerate(indices[:to_show]):
plt.subplot(1,to_show, i+1)
plt.axis('off')
sample = data[index][0]
plt.imshow(sample)
def visualize_predicted_actual(predicted_class, gt_class, predictions, groud_truth, val_indices, data):
"""
Visualizes images of a ground truth class which were predicted as the other class
predicted: int 0-9, index of the predicted class
gt_class: int 0-9, index of the ground truth class
predictions: np array of ints, model predictions for all validation samples
ground_truth: np array of ints, ground truth for all validation samples
val_indices: np array of ints, indices of validation samples
"""
# TODO: Implement visualization using visualize_images above
# predictions and ground_truth are provided for validation set only, defined by val_indices
# Hint: numpy index arrays might be helpful
# https://docs.scipy.org/doc/numpy/user/basics.indexing.html#index-arrays
# Please make the title meaningful!
images = val_indices[(predictions == predicted_class) & (groud_truth == gt_class)]
title = 'predict: ' + str(predicted_class) + ' true: ' + str(gt_class)
visualize_images(images, data,title=title)
return None
visualize_predicted_actual(6, 8, predictions, gt, np.array(val_indices), data_train_images)
visualize_predicted_actual(1, 7, predictions, gt, np.array(val_indices), data_train_images)
# -
# # Переходим к свободным упражнениям!
#
# Натренируйте модель как можно лучше - экспериментируйте сами!
# Что следует обязательно попробовать:
# - перебор гиперпараметров с помощью валидационной выборки
# - другие оптимизаторы вместо SGD
# - изменение количества слоев и их размеров
# - наличие Batch Normalization
#
# Но ограничиваться этим не стоит!
#
# Точность на тестовой выборке должна быть доведена до **80%**
# +
# Experiment here!
#firstlayersize = 128
#secondlayersize = 128
#lr = 1e-3
#weight_decay = 1e-2
#num_epoch = 20
#
#nn_model = nn.Sequential(
# Flattener(),
# nn.Linear(3*32*32, firstlayersize),
# nn.BatchNorm1d(firstlayersize),
# nn.ReLU(inplace=True),
# nn.Linear(firstlayersize, secondlayersize),
# nn.BatchNorm1d(secondlayersize),
# nn.ReLU(inplace=True),
# nn.Linear(secondlayersize,10),
# )
#
#optimizer = optim.Adam(nn_model.parameters(), lr=lr, weight_decay=weight_decay)
#scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=2, gamma=0.5)
#loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, num_epoch, scheduler)
#
firstlayersizes= [32,64,128,256]
secondlayersizes= [64,128,256]
lrs = [1e-2,1e-3,1e-4]
weight_decays = [1e-1,1e-2,1e-3]
num_epochs = [10,20,30]
for firstlayersize in firstlayersizes:
for lr in lrs:
for weight_decay in weight_decays:
print('firstlayersize',firstlayersize,'lr',lr,'weight_decay',weight_decay)
nn_model = nn.Sequential(
Flattener(),
nn.Linear(3*32*32, firstlayersize),
nn.BatchNorm1d(firstlayersize),
nn.ReLU(inplace=True),
nn.Linear(firstlayersize, firstlayersize),
nn.BatchNorm1d(firstlayersize),
nn.ReLU(inplace=True),
nn.Linear(firstlayersize, firstlayersize),
nn.BatchNorm1d(firstlayersize),
nn.ReLU(inplace=True),
nn.Linear(firstlayersize,10),
)
optimizer = optim.Adam(nn_model.parameters(), lr=lr, weight_decay=weight_decay)
scheduler = torch.optim.lr_scheduler.StepLR(optimizer, step_size=2, gamma=0.5)
loss_history, train_history, val_history = train_model(nn_model, train_loader, val_loader, loss, optimizer, 5, scheduler)
# -
# Как всегда, в конце проверяем на test set
test_loader = torch.utils.data.DataLoader(data_test, batch_size=batch_size)
test_accuracy = compute_accuracy(nn_model, test_loader)
print("Test accuracy: %2.4f" % test_accuracy)
'firstlayersize 64 secondlayersize 64 lr 0.001 weight_decay 0.01'
| assignments/assignment2/.ipynb_checkpoints/PyTorch-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''venv'': venv)'
# language: python
# name: python38264bitvenvvenv3181ffefc85747bfb9f84936641d758d
# ---
from crossing import data, experiment, gpt2
import datetime
# ## GPT-2 Experiments
# + tags=["outputPrepend"]
# Load model
model_fn = gpt2.get_model_fn()
# Run all experiments.
n_forms = 7
for i in range(7):
print('[Start]\t Experiment {}\t {}'.format(i+1, datetime.datetime.now().time()))
inputs = data.read_sents('data/sents%d.tsv' % (i+1))
experiment.run_experiment(inputs, model_fn, outfile='data/results_GPT2-%d.tsv' % (i+1))
print('[End]\t Experiment {}\t {}'.format(i+1, datetime.datetime.now().time()))
| run_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# # Load modules
# +
import sys
sys.path.append('..')
from datefeatures import MonthCircle
import numpy as np
import pandas as pd
from randdate import randdate
from datetime import datetime
from sklearn.pipeline import Pipeline, FeatureUnion
from mlxtend.feature_selection import ColumnSelector
# -
# # Example 1
# +
# generate fake dates
X = np.c_[np.array(randdate(10)), np.array(randdate(10))]
# transform date variable to fetures
cmp = MonthCircle()
cmp.fit(X)
Z = cmp.transform(X)
Z.head()
# -
cmp.feature_names_
# # Example 2
# +
# generate fake dates
X = np.c_[np.array(randdate(10)), np.array(randdate(10))]
# emulate missing value
X[1,0] = np.nan
# transform date variable to fetures
cmp = MonthCircle()
cmp.fit(X)
Z = cmp.transform(X)
Z.head()
# -
# # Example 3
n_samples = 100000
X = np.c_[np.array(randdate(n_samples)), np.array(randdate(n_samples)), np.array(randdate(n_samples))]
cmp = MonthCircle()
# %time Z = cmp.fit_transform(X)
# # Example 4
# +
# generate fake dates
n_samples = 5
X = np.c_[np.array(randdate(n_samples))]
X[1,0] = np.nan
# make pipeline
pipe = Pipeline(steps=[
('pre', MonthCircle(out=['sin', 'cos', 'frac']))
])
Z = pipe.fit_transform(X)
Z
# -
# # Example 5
# generate fake dates
n_samples = 5
X = pd.DataFrame(data=randdate(n_samples), columns=['this_date'])
X['some_numbers'] = np.random.randn(n_samples)
X
# +
# make pipeline
pipe = Pipeline(steps=[
# process column by column
('col_by_col', FeatureUnion(transformer_list=[
('dates', Pipeline(steps=[
('sel1', ColumnSelector(cols=('this_date'))),
('pre1', MonthCircle())
])),
('numbers', ColumnSelector(cols=('some_numbers')))
]))
# do some other stuff ..
])
Z = pipe.fit_transform(X)
Z
# -
colnam = list(pipe.steps[0][1].transformer_list[0][1].steps[1][1].feature_names_)
colnam += ['some_numbers']
colnam
pd.DataFrame(Z, columns=colnam)
| examples/MonthCircle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Notebook exploring the geodata extracted from adam4adam profiles based in MA
# ### by <NAME> under supervision of Professor <NAME>
#
# 1. Using geopy, I can use town data I scraped and convert it to lat/long values
# !pip install geopy
from geopy.geocoders import Nominatim
geolocator = Nominatim()
location = geolocator.geocode("Chicago Illinois")
print(location.raw)
help(location)
import pandas as pd
import plotly
# csv file ma_plus_race.csv stores data (last updated to include a section on race/ethnicity for each profile)
ma = pd.DataFrame.from_csv("ma_plus_race.csv")
one_city_list = list(set([i + " Massachusetts" for i in ma["town"].values.tolist()\
if len(str(i).split(", ")) == 1 and str(i) != "nan"]))
lats = []
lons = []
# print(len(one_city_list))
for i in one_city_list:
location = geolocator.geocode(i, timeout=10)
lats.append(location.latitude)
lons.append(location.longitude)
two_city_list = list(set([i + " Massachusetts" for i in ma["town"].values.tolist()\
if len(str(i).split(", ")) == 2 and str(i) != "nan"]))
for i in two_city_list:
if "Other" in i:
two_city_list.remove(i)
two_city_list = [i.split(", ")[1] for i in two_city_list]
all_cities = one_city_list + two_city_list
len(all_cities)
# **Edge cases:**
for i,v in enumerate(all_cities):
if "Leather" in v:
all_cities[i] = "Leather District, Boston"
elif "Bay Village" in v:
all_cities[i] = "Bay Village"
elif "Brickell" in v:
all_cities.remove(v)
elif "Boston Airport" in v:
all_cities[i] = "Boston Logan International Airport, Boston"
# Store conversion data in dictionary and pandas frame
geo_dict = {}
for i in all_cities:
location = geolocator.geocode(i, timeout=10)
try:
lat = location.latitude
lon = location.longitude
except AttributeError:
print(i)
geo_dict[i] = {"lat": lat, "lon": lon}
geo_dict["Boston Logan Airport"] = {"lat": 42.366828, "lon": -71.027330}
geo_dict["Leather District"] = {"lat": 42.350807, "lon": -71.057969}
geo_frame = pd.DataFrame(geo_dict).T
geo_frame.index
# use conversion dictionary to add a lat and long value to each profile in pandas dataframe
clean_towns = []
lats_list = []
lons_list = []
ma_city_values = [str(i) for i in ma["town"].values.tolist()]
for ind,i in enumerate(ma_city_values):
if str(i) == "nan" or ":" in i:
clean_towns.append("nan")
lats_list.append("nan")
lons_list.append("nan")
else:
if "," in i:
val = i.split(", ")[1]
else:
val = i
try:
lats_list.append(geo_dict[val + " Massachusetts"]["lat"])
lons_list.append(geo_dict[val + " Massachusetts"]["lon"])
clean_towns.append(val + " Massachusetts")
except KeyError:
if "Bay Village" in val:
lats_list.append(42.349176)
lons_list.append(-71.069591)
clean_towns.append("Bay Village")
elif "Leather" in val:
lats_list.append(42.350807)
lons_list.append(-71.057969)
clean_towns.append("Leather District")
elif "Airport" in val:
lats_list.append(42.366828)
lons_list.append(-71.027330)
clean_towns.append("Boston Logan Airport")
else:
clean_towns.append("nan")
lats_list.append("nan")
lons_list.append("nan")
ma["towns"] = clean_towns
ma["lat"] = lats_list
ma["lon"] = lons_list
len(ma["towns"])
from itertools import groupby
lengtsh = [len(list(group)) for key, group in groupby(clean_towns)]
# **Check out the frequency breakdown for users around the state**
import collections
counter=collections.Counter(clean_towns)
# print(counter)
# print(counter.values())
# print(counter.keys())
frequencies = (counter.most_common(134))
geo_dict[u'Westborough Massachusetts']['lat']
freq = []
for i,v in enumerate(frequencies):
try:
freq.append([geo_dict[v[0]]["lat"], geo_dict[v[0]]["lon"], v[1]])
except KeyError:
print v
prep_towns = ma["towns"][ma["hiv_status"] == "HIV Negative, on PrEP"].values.tolist()
counter=collections.Counter(prep_towns)
frequencies = (counter.most_common(58))
freq = []
for i,v in enumerate(frequencies):
try:
freq.append([geo_dict[v[0]]["lat"], geo_dict[v[0]]["lon"], v[1], frequencies[i][0]])
except KeyError:
print v
freq
# Use plotly's geoscatter plot to try to visualize that frequency
import plotly.tools as tls
tls.set_credentials_file(username='mrngos', api_key='<KEY>')
# ## Biggest issue:
# The map data in plotly doesn't scale to city/state scope, stays only at state/country level in the States. must find alternative
# +
import plotly.plotly as py
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2011_february_us_airport_traffic.csv')
df.head()
# ma['text'] = df['town'] + '' + df['city'] + ', ' + df['state'] + '' + 'Arrivals: ' + df['cnt'].astype(str)
scl = [ [0,"rgb(5, 10, 172)"],[0.35,"rgb(40, 60, 190)"],[0.5,"rgb(70, 100, 245)"],\
[0.6,"rgb(90, 120, 245)"],[0.7,"rgb(106, 137, 247)"],[1,"rgb(220, 220, 220)"] ]
data = [ dict(
type = 'scattergeo',
locationmode = 'MA-cities',
lon = ma['lon'],
lat = ma['lat'],
mode = 'markers',
marker = dict(
size = 8,
opacity = 0.8,
reversescale = True,
autocolorscale = False,
symbol = 'circle',
line = dict(
width=1,
color='rgba(102, 102, 102)'
),
colorscale = scl,
cmin = 0,
color = df['cnt'],
cmax = df['cnt'].max(),
colorbar=dict(
title="Incoming flightsFebruary 2011"
)
))]
layout = dict(
title = 'Most trafficked US airports<br>(Hover for airport names)',
colorbar = True,
geo = dict(
scope='usa',
projection=dict( type='albers usa' ),
showland = True,
landcolor = "rgb(250, 250, 250)",
subunitcolor = "rgb(217, 217, 217)",
countrycolor = "rgb(217, 217, 217)",
countrywidth = 0.5,
subunitwidth = 0.5
),
)
fig = dict( data=data, layout=layout )
py.iplot( fig, validate=False, filename='d3-airports' )
# -
| .ipynb_checkpoints/geoscatter_data_collection-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generalized Canonical Correlation Analysis (GCCA)
# +
from mvlearn.datasets import load_UCImultifeature
from mvlearn.embed import GCCA
from mvlearn.plotting import crossviews_plot
from graspy.plot import pairplot
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
# ## Load Data
# We load three views from the UCI handwritten digits multi-view data set. Specificallym the Profile correlations, Karhunen-Love coefficients, and pixel averages from 2x3 windows.
# Load full dataset, labels not needed
Xs, y = load_UCImultifeature()
Xs = [Xs[1], Xs[2], Xs[3]]
# Check data
print(f'There are {len(Xs)} views.')
print(f'There are {Xs[0].shape[0]} observations')
print(f'The feature sizes are: {[X.shape[1] for X in Xs]}')
# ### Embed Views
# Create GCCA object and embed the
gcca = GCCA()
Xs_latents = gcca.fit_transform(Xs)
print(f'The feature sizes are: {[X.shape[1] for X in Xs_latents]}')
# ## Plot the first two views against each other
# The top three dimensions from the latents spaces of the profile correlation and pixel average views are plotted against each other. However, their latent spaces are influenced the the Karhunen-Love coefficients, not plotted.
crossviews_plot(Xs_latents[[0,2]], dimensions=[0,1,2], labels=y, cmap='Set1', title=f'Profile correlations vs Pixel Averages', scatter_kwargs={'alpha':0.4, 's':2.0})
| docs/tutorials/embed/gcca_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
A = 2.414*10**-5
B = 247.8
C = 140
celsious = np.linspace(0,100,1000)
T = np.linspace(273,373,1000)
mu_T = A*10**(B/(T-C))
plt.figure(figsize=(10,5),dpi=250)
plt.plot(celsious,mu_T)
plt.xlabel('Temperature in C')
plt.ylabel('')
| SimPy/.ipynb_checkpoints/water_viscosity-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Introduction to Geospatial Data
# ### Part 5 of 5
# # Storing geography in the computer
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## Reminder
# <a href="#/slide-2-0" class="navigate-right" style="background-color:blue;color:white;padding:8px;margin:2px;font-weight:bold;">Continue with the lesson</a>
#
# <font size="+1">
#
# By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
#
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
# If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# For the full description please navigate to this website: <a href="../../gateway-lesson/gateway/gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
#
# </font>
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# ## The world is infinitely complex
# <table>
# <tr style="background: #fff">
# <td width=50%> <img src='supplementary/queenstown.jpg' alt='Picture of Queenstown, New Zealand'></td>
# <td align=left valign=top>This is a photo looking down towards the SE on the mountain resort town of Queenstown, New Zealand (at 45.03 N lat, 168.66 E long).<br><br>
# How many different kinds of things do you see here?
# <br><br>
# How can we decide what to measure and record?
# <br><br>
# And how can we structure data about this complex world into tables to represent this?????</td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# ## A famous GIScientist once said
# <i>"People cultivate fields (but manipulate objects)"</i> <small>**</small></p>
# This phrase summarizes the most important distinction we make when capturing geospatial data --
# <table>
# <tr style="background: #fff">
# <td width=40%> <img src='supplementary/raster_vector.png' alt='Raster or vector?'></td>
# <td align=left valign=top><b><u>Is the world made up of <i>fields</i> or <i>objects</i>?</u></b></td>
# </tr>
# </table>
# <small>** by <NAME>, 1992, <a href="https://www.researchgate.net/publication/221589734_People_Manipulate_Objects_but_Cultivate_Fields_Beyond_the_Raster-Vector_Debate_in_GIS">"People Manipulate Objects (but Cultivate Fields): Beyond the Raster-Vector Debate in GIS"</a> from the book *Theories and Methods of Spatio-Temporal Reasoning in Geographic Space: International Conference GIS — From Space to Territory: Theories and Methods of Spatio-Temporal Reasoning* Pisa, Italy, September 21–23, 1992 (pp.65-77)</small>
# + [markdown] slideshow={"slide_type": "slide"}
# Think about the picture of Queenstown we looked at earlier.
#
# The rolling surface of the landscape is continuous. There's land or water, at various elevations, everywhere. That's a *field*. Elevation is the classic field. There is a value of elevation everywhere.
#
# Then consider all the manmade structures in the picture. There are buildings, lightposts, roads. These are *objects*. The object world view is mostly empty, with objects scattered around.
# + [markdown] slideshow={"slide_type": "slide"} tags=["6A", "init"] variables={"IFrame(\"supplementary/sort-field-object.html\", width=970, height=530)": "\n <iframe\n width=\"970\"\n height=\"530\"\n src=\"supplementary/sort-field-object.html\"\n frameborder=\"0\"\n allowfullscreen\n \n ></iframe>\n "}
# So, let's see if you can separate these two perspectives.
#
# {{IFrame("supplementary/sort-field-object.html", width=970, height=530)}}
# + [markdown] slideshow={"slide_type": "slide"}
# <center><large>Now let's look at some geospatial data that are coded as either objects or fields.
# + [markdown] slideshow={"slide_type": "slide"}
# Starting with field data, here is a file of elevation measurements in the area to the south of Queenstown which is located near the center top of the image. You can see the lakes in the pale grey color.
# + tags=["6B"]
import rasterio
from matplotlib import pyplot
filepath = 'https://dds.cr.usgs.gov/srtm/version1/Islands/S46E168.hgt.zip'
raster = rasterio.open(filepath, 'r')
pyplot.imshow(raster.read(1), cmap='terrain')
pyplot.show()
# + [markdown] slideshow={"slide_type": "slide"}
# Now we can look at how the field data is actually stored.
# + tags=["6C"]
raster.read()
# -
# What we're seeing here is the beginning and end of the first three and last three lines of the file. What's all this???
# + [markdown] slideshow={"slide_type": "slide"}
# Field data is usually stored as *rasters*.
#
# To store the world into a raster, the surface of the earth is divided into a grid of equal sized cells that covers a specific chunk of the earth, say a square that is 10 m by 10 m.
#
# <table>
# <tr style="background: #fff">
# <td width=40%> <img src='supplementary/world_to_raster_sm.png' alt='World to raster'></td>
# <td align=left valign=top>Each cell is given a value that represents the data that has been measured on the earth in that cell.<br><br>
# In the raster in this graphic, the building has been coded with the value green and the road has been coded with the value red.
# </td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# So, let's look again at that field data. Run both of these code chunks.
# + tags=["6D"]
print("The Raster is", raster.width, "cells wide and", raster.height, "cells high")
# + tags=["6E"]
raster.bounds
# + [markdown] slideshow={"slide_type": "-"}
# These show us that the NW (top left) corner of the area covered is 45 S latitude and 168 E longitude and the area covered is 1 degree of latitude high and 1 degree of longitude wide. Since 1 degree is 3600 seconds and we have ~1200 cells, this means the cell dimensions are 3600/1200 = ~3 arc seconds of a degree (that's approx 64m wide and 90m high at this latitude).
# + [markdown] slideshow={"slide_type": "slide"}
# Each row in the file shows us the average elevation value (in meters) in each cell across a row of the grid. Run this code to see the file again.
# + tags=["6F"]
raster.read()
# -
# Note that the elevations are much higher in the NE corner (as evidenced by the high values at the end of the first few rows) and lower along the southern edge (shown in the final rows).
# + [markdown] slideshow={"slide_type": "slide"}
# Now let's look at how object data is stored - hint, it's completely different! And WAY more complex.
#
# We'll start simple. When you ask Google to show you all the nearby restaurants on a map, you get a map with a bunch of pins, some with labels. You can click on them and find out information about those places. Those dots represent restaurant objects.
#
# For example...
# + [markdown] slideshow={"slide_type": "slide"}
# Here's a map of Queenstown showing some points of interest. Now we're looking north and the camera point for the photo used earlier is the cleared area at the top of the hill on the left.
#
# <a href="https://www.google.com/maps/place/Queenstown,+New+Zealand/@-45.0514839,168.6648181,1609a,35y,345.22h,54.63t/data=!3m1!1e3!4m5!3m4!1s0xa9d51df1d7a8de5f:0x500ef868479a600!8m2!3d-45.0301511!4d168.6616206">This link will take you to this map live in Google Maps.</a>
# 
#
# + [markdown] slideshow={"slide_type": "slide"}
# Now, let's see how that data is stored in a file. In Try-it Exercise #1 you looked at a point dataset. Remember this? (click the arrow to the left of the code)
# -
# !wget -O ne_50m_populated_places_simple.zip https://www.naturalearthdata.com/http//www.naturalearthdata.com/download/50m/cultural/ne_50m_populated_places_simple.zip
# !unzip -n ne_50m_populated_places_simple.zip
# + tags=["6G"]
import geopandas
cities = geopandas.read_file("ne_50m_populated_places_simple.shp")
cities.head()
# + [markdown] slideshow={"slide_type": "slide"}
# In the table we just generated, each row has
# - an object ID
# - some data about various attributes for that object
# - then a column with an entry that is the point location
#
# Click back one slide to check this out.
# + [markdown] slideshow={"slide_type": "slide"}
# Now let's see again how that table can generate the dots on a map...
# + tags=["6H"]
from ipyleaflet import Map, GeoData
cities_layer= GeoData(geo_dataframe = cities)
mymap = Map(center=(-43,168), zoom = 5)
mymap.add_layer(cities_layer)
mymap
# + [markdown] slideshow={"slide_type": "slide"}
# OK, let's get back to fields and objects and how we put them into the computer. Remember this?
#
# <table>
# <tr style="background: #fff">
# <td width=40%> <img src='supplementary/raster_vector.png' alt='Raster or vector'></td>
# <td align=left valign=top>These two graphics show the two most common <i><u>data models</u></i> for geospatial data.<br><br>
# Fields are stored as grids called <i>rasters</i> and there is a value everywhere. <br><br>
# Objects, which are scattered around mostly empty space, are stored as <i>vectors</i>.<br><br><br>
# So, tell me more about vectors, you say...</td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# Vectors usually come in three varieties - points, lines and polygons.
# <table>
# <tr style="background: #fff">
# <td width=30%> <img src='supplementary/vectors_sm.png' alt='Raster or vector'></td>
# <td align=left valign=top>Points are good for things like cities on a world map, or lightpoles and signposts on a neighborhood map.<br><br>
# Lines are for rivers, roads, railways, boundaries - that sort of thing.<br><br>
# Polygons are areas. So they're used for lakes, building footprints, parks.</td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# Vector data has two components.
# <table>
# <tr style="background: #fff">
# <td width=40%> <img src='supplementary/vector_structure.png' alt='Vector structure'></td>
# <td align=left valign=top>These components can be stored together in a table by including one or more columns that provide the direct georeference (e.g. lat and long).<br><br>
# <i>OR</i>, these components can be stored separately. Attributes with an object ID in one table and the geometry labelled with the same IDs in a separate file. </td>
# </tr>
# </table>
# + [markdown] slideshow={"slide_type": "slide"}
# By the way, it's important to know that you can't mix up points, lines and polygons in a single geospatial data file. If you want a map that shows points, lines and polygons, then you'll need at least three different datasets, one for each type of vector object.
#
# Remember the rivers data in our Try-it Exercise #1? Let's add it to the map.
#
# First, we'll get it again, just in case it's not currently loaded. (click the arrow to the left)
# -
# !wget -O ne_10m_rivers_lake_centerlines.zip https://www.naturalearthdata.com/http//www.naturalearthdata.com/download/10m/physical/ne_10m_rivers_lake_centerlines.zip
# !unzip -n ne_10m_rivers_lake_centerlines.zip
# + tags=["6I"]
rivers = geopandas.read_file("ne_10m_rivers_lake_centerlines.shp")
rivers_layer = GeoData(geo_dataframe = rivers, style={'color':'blue'})
# -
# (wait for the asterisk to turn into a number...) then go to the next slide and we'll add it to the cities data...
# + slideshow={"slide_type": "slide"} tags=["6J"]
mymap2 = Map(center=(-43,168), zoom = 5)
mymap2.add_layer(cities_layer)
mymap2.add_layer(rivers_layer)
mymap2
# + [markdown] slideshow={"slide_type": "slide"} tags=["6K"] variables={"IFrame(\"supplementary/sort-raster-vector.html\", width=970, height=430)": "\n <iframe\n width=\"970\"\n height=\"430\"\n src=\"supplementary/sort-raster-vector.html\"\n frameborder=\"0\"\n allowfullscreen\n \n ></iframe>\n "}
# OK, now let's practice these concepts. For each of the following kinds of geospatial data, choose the data model (raster or vector) that it's most likely to be stored in.<p>
#
# {{IFrame("supplementary/sort-raster-vector.html", width=970, height=430)}}
# + [markdown] slideshow={"slide_type": "slide"}
# Well done! Now you know a little bit about geospatial data.
#
# If you have worked through this lesson carefully, you should now be able to:
# 1. Explain what is special about geospatial data.
# 2. Describe how location can be measured and recorded in geospatial data.
# 3. Explain the difference between raster and vector data.
# 4. Identify several different types of geospatial data.
# 5. Load and view different kinds of geospatial data in Python Notebooks.
# + [markdown] slideshow={"slide_type": "slide"}
# If you still have time, feel free to go back to the two Try-It exercises and try out downloading some different datasets from the sources. Make maps of different parts of the earth or of different days from the Johns Hopkins data server.
#
# If you want to learn more about geospatial data, you can go on to the intermediate Geospatial Data lesson.
#
# Or you can go back and complete some of the other introductory lessons as they all touch on the use of geospatial data.
#
#
# -
#
# # Congratulations!
#
#
# **You have finished an Hour of CI!**
#
#
# But, before you go ...
#
# 1. Please fill out a very brief questionnaire to provide feedback and help us improve the Hour of CI lessons. It is fast and your feedback is very important to let us know what you learned and how we can improve the lessons in the future.
# 2. If you would like a certificate, then please type your name below and click "Create Certificate" and you will be presented with a PDF certificate.
#
# <font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="https://forms.gle/JUUBm76rLB8iYppN7">Take the questionnaire and provide feedback</a></font>
#
#
#
# + hide_input=true slideshow={"slide_type": "-"} tags=["Hide", "Init"]
# This code cell loads the Interact Textbox that will ask users for their name
# Once they click "Create Certificate" then it will add their name to the certificate template
# And present them a PDF certificate
from PIL import Image
from PIL import ImageFont
from PIL import ImageDraw
from ipywidgets import interact
def make_cert(learner_name, lesson_name):
cert_filename = 'hourofci_certificate.pdf'
img = Image.open("../../supplementary/hci-certificate-template.jpg")
draw = ImageDraw.Draw(img)
cert_font = ImageFont.load_default()
cert_font = ImageFont.truetype('../../supplementary/times.ttf', 150)
cert_font2 = ImageFont.truetype('../../supplementary/times.ttf', 100)
w,h = cert_font.getsize(learner_name)
draw.text( xy = (1650-w/2,1100-h/2), text = learner_name, fill=(0,0,0),font=cert_font)
draw.text( xy = (1650-w/2 - 12*int(len(lesson_name)),1100-h/2 + 750), text = lesson_name, fill=(0,0,0),font=cert_font2)
img.save(cert_filename, "PDF", resolution=100.0)
return cert_filename
interact_cert=interact.options(manual=True, manual_name="Create Certificate")
@interact_cert(name="Your Name")
def f(name):
print("Congratulations",name)
filename = make_cert(name, 'The Geospatial Data Beginner Lesson')
print("Download your certificate by clicking the link below.")
# -
# <font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="hourofci_certificate.pdf?download=1" download="hourofci_certificate.pdf">Download your certificate</a></font>
# + hide_input=true slideshow={"slide_type": "-"} tags=["Hide", "Init"]
IFrame(src = '../../supplementary/confetti.html', width="700", height="430")
| beginner-lessons/geospatial-data/gd-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cifrum as lib
import pandas as pd
import numpy as np
# # Inflation
# To compute the inflation, the library provides `inflation` method. The library supports kinds of inflation as follows:
#
# - `cumulative` - cumulative inflation
# - `a_mean` - arithmetic mean of inflation
# - `g_mean` - geometric mean of inflation
# - `yoy` - Year on Year inflation
# - `values` - raw values of inflation
#
# Another parameter is `currency` to compute the inflation against.
#
# And the last ones are periods:
#
# - `end_period` - the period till which to compute. The default value is the latest date for the existing inflation values
# - `start_period` - the period till which to compute. The default value is the earliest date for the existing inflation values
# - `years_ago` that can be only specified if `start_period` doesn't. Corresponding `start_period` is then computed as `end_period - 12 * years_ago`
lib.inflation(currency='usd', start_period='2016-1', end_period='2019-4', kind='values')
lib.inflation(currency='usd', start_period='2016-1', end_period='2019-4', kind='yoy')
lib.inflation(currency='usd', end_period='2019-4', years_ago=10, kind='yoy')
| examples/inflation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + cell_id="00001-80603ce7-5e43-4aaa-9c2e-fc333c20a663" execution_millis=4 execution_start=1603914492064 output_cleared=false source_hash="bbacf3a0" tags=[] id="09g62Azr5U1R" executionInfo={"status": "ok", "timestamp": 1603916886579, "user_tz": 420, "elapsed": 1675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}}
import nltk
import numpy as np
# + cell_id="00002-c68972ca-99d0-4fb8-8b2c-1c296e224a4a" execution_millis=15 execution_start=1603914493054 output_cleared=false source_hash="2f335b27" tags=[] id="kOWus-a95U1Z" executionInfo={"status": "ok", "timestamp": 1603916888361, "user_tz": 420, "elapsed": 849, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}}
import requests
url = "http://www.gutenberg.org/files/2554/2554.txt"
response = requests.get(url)
raw_html = response.content
text = raw_html.decode("utf-8-sig")
# + [markdown] cell_id="00005-0e3e3085-9f12-4073-b9bf-a699a216c404" tags=[] id="OXQvFzyj5U1f"
# ### Bag of Words
#
# Bag of Words based encoding or TF-IDF vector is a frequentist based approach to NLP applications.
# + cell_id="00006-381941ee-0649-4a76-9841-1923426edffe" execution_millis=248 execution_start=1603914496124 output_cleared=false source_hash="abe7455d" tags=[] id="iSDR5KNt5U1g" executionInfo={"status": "ok", "timestamp": 1603917274958, "user_tz": 420, "elapsed": 2459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}}
import urllib.request
url = "https://www.gutenberg.org/files/829/829-0.txt" # gulliver's travels
#Alt
#url = 'https://www.gutenberg.org/files/2701/2701-0.txt' # Moby Dick
file = urllib.request.urlopen(url)
text = [line.decode('utf-8') for line in file]
text = ''.join(text)
# + [markdown] cell_id="00007-e8474323-77fb-4af2-b76b-cf21b17e2b07" tags=[] id="S3hK5Ldw5U1j"
# #### Tokenize
# + cell_id="00008-c92e3a1e-acf6-4466-8344-0ca3510a6b75" execution_millis=2119 execution_start=1603914497851 output_cleared=false source_hash="dd04af38" tags=[] id="43g6RdUf5U1k" executionInfo={"status": "ok", "timestamp": 1603917279960, "user_tz": 420, "elapsed": 1306, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="243880a7-7b85-4f86-98b9-379827ce3afb" colab={"base_uri": "https://localhost:8080/"}
nltk.download('punkt')
from nltk import word_tokenize
tokens = word_tokenize(text)
# + cell_id="00009-e513d93c-17df-4f64-ba5e-748f1628283d" execution_millis=218 execution_start=1603914500757 output_cleared=false source_hash="204a6f56" tags=[] id="oc9TB80i5U1p" executionInfo={"status": "ok", "timestamp": 1603917288906, "user_tz": 420, "elapsed": 647, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}}
import string
tokens = [word for word in tokens if word.isalpha()]
table = str.maketrans('', '', string.punctuation)
tokens = [w.translate(table) for w in tokens]
tokens = [word.lower() for word in tokens]
# + [markdown] cell_id="00010-128bff7f-6c9c-41c7-af8b-34e14d7c8cda" tags=[] id="d-kUPb7k5U1s"
# Removing **stop-words** and **stemming**
# + cell_id="00011-cdf4882f-23c8-4056-852c-7aa7f2e398d9" execution_millis=2889 execution_start=1603914502637 output_cleared=false source_hash="203b141e" tags=[] id="Syfo7BOb5U1t" executionInfo={"status": "ok", "timestamp": 1603917291637, "user_tz": 420, "elapsed": 1571, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="842060a1-2a34-4b07-c1d2-e90caa8c4d71" colab={"base_uri": "https://localhost:8080/"}
from nltk.corpus import stopwords
nltk.download('stopwords')
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
from nltk.stem.porter import PorterStemmer
porter = PorterStemmer()
tokens = [porter.stem(word) for word in tokens]
tokens[200:202]
# + [markdown] cell_id="00012-044ce2ce-6815-490b-9f18-0645470e543f" tags=[] id="NjzIKn_85U1w"
# **Understanding the vocabulary**
#
# * A vocabulary of a document represents all the words in that document and the frequency they appear.
# * `FreqDist` class
#
# + cell_id="00013-fdd3703a-21ab-4f5c-bef8-ecccb46953e3" execution_millis=40 execution_start=1603914505591 output_cleared=false source_hash="60a04f4f" tags=[] id="2Yv0397E5U1x" executionInfo={"status": "ok", "timestamp": 1603917300119, "user_tz": 420, "elapsed": 683, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="9ceba789-32ca-425d-af6c-26b40c38e744" colab={"base_uri": "https://localhost:8080/"}
from nltk.probability import FreqDist
word_counts = FreqDist(tokens)
word_counts
# + [markdown] cell_id="00014-72d4693b-04ed-488c-ae66-8f0ce7199aed" tags=[] id="RhDlzRqE5U1z"
# **Scoring words with frequency**
# + cell_id="00015-02387729-4804-4ce6-9161-419c503be606" execution_millis=9 execution_start=1603914506105 output_cleared=false source_hash="d999d5d7" tags=[] id="eapZUCqh5U10" executionInfo={"status": "ok", "timestamp": 1603917302419, "user_tz": 420, "elapsed": 513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="54d8f9b6-ffa1-4158-dcd9-aa00c84730f5" colab={"base_uri": "https://localhost:8080/"}
top = 100
vocabulary = word_counts.most_common(top)
vocabulary[:10]
# + cell_id="00016-a40281d3-efa8-460f-bbe0-e4090eee870a" execution_millis=4 execution_start=1603914507044 output_cleared=false source_hash="a2651e8b" tags=[] id="SO1eID8a5U14" executionInfo={"status": "ok", "timestamp": 1603917303987, "user_tz": 420, "elapsed": 599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="70e04656-fbc9-4c47-a1f0-36bf5282b984" colab={"base_uri": "https://localhost:8080/"}
voc_size = len(vocabulary)
doc_vector = np.zeros(voc_size)
word_vector = [(idx,word_counts[word[0]]) for idx, word in enumerate(vocabulary) if word[0] in word_counts.keys()]
word_vector[10]
# + cell_id="00017-36e9c930-b190-4fa9-9b2d-dbda227ca607" execution_millis=415 execution_start=1603914507882 output_cleared=false source_hash="ae320f3a" tags=[] id="NStzHzfH5U17" executionInfo={"status": "ok", "timestamp": 1603917335032, "user_tz": 420, "elapsed": 3213, "user": {"displayName": "Y<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggu3bTBcysUxxtmnLI9n_OEGc92nfpZXGRAhDZ5vQ=s64", "userId": "12984101785953050215"}} outputId="48efa19f-9fe3-400a-81c5-8f1a91f05975" colab={"base_uri": "https://localhost:8080/"}
# Generating a model of Bag of Words
from nltk import sent_tokenize
docs = sent_tokenize(text)[703:706]
docs
from sklearn.feature_extraction.text import CountVectorizer
count_vectorizer=CountVectorizer(stop_words='english')
word_count_vector=count_vectorizer.fit_transform(docs)
word_count_vector.shape
word_count_vector.toarray()
count_vectorizer.get_feature_names()
# + id="DqneBlSy5exp"
| part1_labs/01_NLP basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Solución Titanic</h1>
# <h2>Introducción</h2>
# <p>En este notebook se resuelve el problema del <a href="https://www.kaggle.com/c/titanic">Titanic</a>. En específicio, se desarrollarán las siguientes etapas:</p>
#
# <ol>
# <li><strong>Análisis de datos</strong></li>
# <ol>
# <li>Preguntas</li>
# <li>Data wrangling</li>
# <li>Exploratory data analysis</li>
# <li>Conclusiones</li>
# </ol>
# <li><strong>Clasificación con Machine Learning</strong></li>
# <ol>
# <li>Pre-procesamiento de variables</li>
# <li>Train, validation y test sets</li>
# <li>3 modelos para clasificación</li>
# <li>Entrenamiento</li>
# <li>Diagnóstico</li>
# <li>Resultados</li>
# </ol>
# </ol>
# Manipulacion de datos y tensores.
import numpy as np
import pandas as pd
# Aprendizaje automatico.
import sklearn
from sklearn.model_selection import train_test_split
# Visualizaciones.
import matplotlib.pyplot as plt
from matplotlib import style
style.use("ggplot")
import seaborn as sns
# Retina command that makes things look good
# %config InlineBackend.figure_format = 'retina'
# <h2>1. Análisis de datos</h2>
# <p style="text-justify;">Antes de aplicar <strong>Inteligencia Artificial</strong> analizaremos los datos siguiendo el proceso descrito previamente. En la primera fase, leeremos los datos y los cargaremos a una estructura de datos. Seguidamente, limpiaremos los datos utilizando estadística descriptiva para encontrar outliers y missing values. Despues, aplicaremos análisis de datos uni-variables y multi-variables con el fin de responder preguntas sobre el dataset. Finalmente, presentaremos conclusiones sobre nuestras preguntas.</p>
# <h3>A. Preguntas</h3>
# <p>En esta sección:</p>
# <ol>
# <li>Propondemos preguntas sobre el dataset del Titanic.</li>
# </ol>
# <h4></h4>
# <ol>
# <li>Cual es el sexo que tuvo mayor opcion de sobrevivir?.</li>
# <li>Cual es el rango de edad que tuvo mayor opcion de sobrevivir?</li>
# <li>Que clase social tenia mas oportunidad de sobrevivir?</li>
# </ol>
# <h3>B. Data wrangling</h3>
# <p>En esta sección:</p>
# <ol>
# <li>Cargaremos los datos a una estructura de datos</li>
# <li>Exploraremos cada variable del dataset</li>
# <li>Limpiaremos las variables del dataset</li>
# </ol>
# Cargar datos
df_original = pd.read_csv("data/train.csv")
# Leer datos
df_original.head(3)
# <p>Las features PassengerId, Name, Cabin, Embarked, Parch no tienen relacion con la variable de salida. Es decir, el id de identificacion de cada pasajero no decidira la muerte de un pasajero. El nombre de un pasajero tampoco interferira con su posibilidad de supervivencia. Asimismo, el numero de cabina o el puerto de abordo al barco tampoco afectan la supervivencia del pasajero. Por tanto eliminamos estas variables.</p>
df_dropped_features = df_original.drop(["PassengerId", "Name", "Cabin", "Embarked", "Parch", "Ticket"], axis = 1)
df_dropped_features.head()
# <p>Antes de seguir con el analisis de datos, debemos transformar las features con valores no numericos. Estas features son Sex y Ticket. Veamos las siguientes celdas para entender la situacion.</p>
#
# <h4>Feature Sex</h4>
# Veamos las clases unicas de la variable Sex.
print("Clases unicas: ", df_dropped_features["Sex"].unique())
print("Cantidad de variables unicas: ", len(df_dropped_features["Sex"].unique()))
# <p>Como se puede observar, la variable de sexo esta compuesta por dos categorias o clases: male y female. Asimismo, la variable ticket esta compuesta por 681 categorias (el tipo de ticket nos sirve porque tiene correlacion con la la clase social a la cual el pasajero pertenece).</p>
# <h4>Categorizacion de feature sex</h4>
# Categorizamos las features con el metodo get_dummies
df_categorized = pd.get_dummies(df_dropped_features)
df_categorized.head()
# Cuantas features tenemos?
print("Features:", [i for i in df_categorized.columns], "\n")
print("Cantidad total de features: ", len(df_categorized.columns))
# <p>Utilizemos variables descriptivas para analizar los datos:</p>
df_categorized.describe()
# <h3>Conclusiones sobre la tabla</h3>
# <o>
# <li>Con respecto a la cantidad de datos por feature podemos concluir que la feature Age tiene una menor cantidad de datos. Por tanto, probablemente hay valores nan.</li>
# </o>
# <h2>Limpieza de datos</h2>
# <h3>Ejercicio</h3>
# <p>Debemos limpiar los valores nan en la feature edad de nuestros datos. Tenemos varias opciones:</p>
# <ol>
# <li><strong>OPCION 1</strong> Rellenar los valores nan con la <strong>media</strong> de las edades.</li>
# <li><strong>OPCION 2</strong> Rellenar los valores nan con la <strong>mediana</strong> de las edades.</li>
# <li><strong>OPCION 3</strong> Eliminar las filas con los valores nan.</li>
# </ol>
# <p>Por el momento tu trabajo es escoger una de las estrategias. Mas adelante evaluaras tus resultados al aplicar un algoritmo de machine learning para predecir el test set.</p>
# +
# ESCOJE UNA OPCION.
opcion = # LLena la variable con uno de los siguientes numeros: 1, 2, 3
# Assertions.
assert (type(opcion) == int), "La variable opcion debe ser un entero."
assert ((opcion in [1,2,3]) == True), "opcion debe tener un valor entre [1, 2, 3]"
# Local variables
df_cleaned = df_categorized.copy()
# Opciones de limpieza para valores nan.
if (opcion == 1):
media_edad = df_categorized["Age"].mean()
df_cleaned["Age"] = df_categorized["Age"].fillna(media_edad)
elif (opcion == 2):
mediana_edad = df_categorized["Age"].median()
df_cleaned["Age"] = df_categorized["Age"].fillna(mediana_edad)
elif (opcion == 3):
df_cleaned["Age"] = df_categorized.dropna(axis = 1)
# -
# <h3>C. Exploratory data analysis</h3>
# <p>En esta sección:</p>
# <ol>
# <li>Responderemos a las preguntas formuladas previamente</li>
# </ol>
# <h4>Cual es el sexo que tuvo mayor opcion de sobrevivir?</h4>
# <p>La siguiente celda tiene codigo que te ayudara a graficar mas sencillamente.</p>
def plot_with_bar(values = None, title = None, xticks = None):
# Assertions
if (values == None):
raise ValueError("Values no puede estar vacio.")
if (type(values) == list) or (type(values) == tuple):
pass
else:
raise TypeError("Values debe ser una lista o tupla.")
ind = [i for i in range(len(values))]
plt.bar(ind, values)
plt.title(title)
plt.xticks(ind, xticks)
plt.show()
# Cuantos hombres y mujeres habian entre pasajeros.
male = df_cleaned.query("Sex_male == 1")
female = df_cleaned.query("Sex_female == 1")
print("Cantidad de hombres: ", male.shape[0])
print("Cantidad de mujeres: ", female.shape[0])
print("Cantidad total de pasajeros: ", df_original.shape[0])
# Plot a bar
plot_with_bar(values = [male.shape[0], female.shape[0]],
title = "Cantidad de pasajeros en el barco por sexo",
xticks = ["Hombres", "Mujeres"])
# Del numero total de mujeres, cuantas sobrevivieron y cuantas no.
females_survived = df_cleaned.query("Survived == 1 and Sex_female == 1")
females_not_survived = df_cleaned.query("Survived == 0 and Sex_female == 1")
# Del numero total de hombres, cuantos sobrevivieron y cuantos no.
males_survived = df_cleaned.query("Survived == 1 and Sex_male == 1")
males_not_survived = df_cleaned.query("Survived == 0 and Sex_male == 1")
# Print
print("Mujeres")
print("Mujeres sobrevivieron: ", females_survived.shape[0])
print("Mujeres no sobrevivieron: ", females_not_survived.shape[0])
plot_with_bar(values = [females_survived.shape[0], females_not_survived.shape[0]],
title = "Supervivencia de mujeres",
xticks = ["Sobrevivieron", "No sobrevivieron"])
print("\nHombres")
print("Hombres sobrevivieron: ", males_survived.shape[0])
print("Hombres no sobrevivieron: ", males_not_survived.shape[0])
plot_with_bar(values = [males_survived.shape[0], males_not_survived.shape[0]],
title = "Supervivencia de hombres",
xticks = ["Sobrevivieron", "No sobrevivieron"])
# <h4>Respuesta</h4>
# <ol>
# <li>Determinar la cantidad de pasajeros nos permitio concluir que hubo una mayor cantidad de hombres con respecto a mujeres en el barco.</li>
# <li>Seguidamente obtuvimos una cantidad de 233 de 314 mujeres que sobrevivieron. Por tanto, si eras mujer tenias un 74% de probabilidad de sobrevivir.</li>
# <li>Despues obtuvimos una cantidad de 109 de 577 hombres que sobrevivieron. Por tanto, si eras hombre tenias un 18% de probabilidad de sobrevivir.</li>
# <li>Ademas, podemos agregar estos datos y determinar que 341 de 891 pasajeros sobrevivieron al accidente. Esto representa 38% del total de personas en el barco, menos de la mitad.</li>
# </ol>
# <h4>Cual es el rango de edad que tuvo mayor opcion de sobrevivir?</h4>
# Primero veamos el rango de edades.
df_cleaned["Age"].describe()
# Podemos observar que la edad minima es 0.42 y la edad maxima es 80.
# Crearemos tres categorias para segmentar los pasajeros.
children = df_cleaned.query("Age < 18")
adults = df_cleaned.query("Age >= 18 and Age <= 55")
elder = df_cleaned.query("Age > 55 and Age <= 80")
# Grafiquemos los datos.
plot_with_bar(values = [children.shape[0], adults.shape[0], elder.shape[0]],
title = "Edades por categorias",
xticks = ["Niños", "Adultos", "3ra edad"])
# Ahora veamos la cantidad de ninos, adultos y 3ra edad que sobrevivieron.
children_survived = children.query("Survived == 1")
children_not_survived = children.query("Survived == 0")
adults_survived = adults.query("Survived == 1")
adults_not_survived = adults.query("Survived == 0")
elder_survived = elder.query("Survived == 1")
elder_not_survived = elder.query("Survived == 0")
per_ch_surv = (children_survived.shape[0] / (children_survived.shape[0] + children_not_survived.shape[0]))*100
per_ad_surv = (adults_survived.shape[0] / (adults_survived.shape[0] + adults_not_survived.shape[0]))*100
per_el_surv = (elder_survived.shape[0] / (elder_survived.shape[0] + elder_not_survived.shape[0]))*100
# Grafiquemos los datos
plot_with_bar(values = [children_survived.shape[0], children_not_survived.shape[0]],
title = "Supervivencia de niños",
xticks = ["Sobrevivieron", "No sobrevivieron"])
plot_with_bar(values = [adults_survived.shape[0], adults_not_survived.shape[0]],
title = "Supervivencia de adultos",
xticks = ["Sobrevivieron", "No sobrevivieron"])
plot_with_bar(values = [elder_survived.shape[0], elder_not_survived.shape[0]],
title = "Supervivencia de 3ra edad",
xticks = ["Sobrevivieron", "No sobrevivieron"])
print("Supervivencia niños: ", str(per_ch_surv)[:6])
print("Supervivencia adultos: ", str(per_ad_surv)[:6])
print("Supervivencia 3ra edad: ", per_el_surv)
# <h4>Respuesta</h4>
# <ol>
# <li>Los niños tenian un 53% porcentaje de supervivencia.</li>
# <li>Los adultos tenian un 36% porcentaje de supervivencia.</li>
# <li>Los 3ra eda tenian un 30% porcentaje de supervivencia.</li>
# <li>En conclusion, los niños tenian una mayor ventaja en supervivencia.</li>
# </ol>
# <h4>Que clase social tenia una mayor posibilidad de supervivencia?</h4>
# Veamos lost tipos de clases sociales.
print("Tres tipos de clases: ", df_cleaned["Pclass"].unique())
# Veamos la distribucion de clases en el barco.
first_class = df_cleaned.query("Pclass == 1")
second_class = df_cleaned.query("Pclass == 2")
third_class = df_cleaned.query("Pclass == 3")
# Plots.
plot_with_bar(values = [first_class.shape[0], second_class.shape[0], third_class.shape[0]],
title = "Distribucion clases sociales en los pasajeros",
xticks = ["1ra", "2da", "3ra"])
# Ahora veamos el porcentaje de supervivencia de cada clase.
first_class_survived = first_class.query("Survived == 1")
first_class_not_survived = first_class.query("Survived == 0")
second_class_survived = second_class.query("Survived == 1")
second_class_not_survived = second_class.query("Survived == 0")
third_class_survived = third_class.query("Survived == 1")
third_class_not_survived = third_class.query("Survived == 0")
print("Supervivencia 1ra clase: ", (first_class_survived.shape[0] / (first_class_survived.shape[0] + first_class_not_survived.shape[0])))
print("Supervivencia 2da clase: ", (second_class_survived.shape[0] / (second_class_survived.shape[0] + second_class_not_survived.shape[0])))
print("Supervivencia 3ra clase: ", (third_class_survived.shape[0] / (third_class_survived.shape[0] + third_class_not_survived.shape[0])))
# Grafiquemos.
survived = (first_class_survived.shape[0], second_class_survived.shape[0], third_class_survived.shape[0])
not_survived = (first_class_not_survived.shape[0], second_class_not_survived.shape[0], third_class_not_survived.shape[0])
ind = np.arange(3) # the x locations for the groups
width = 0.35 # the width of the bars: can also be len(x) sequence
p1 = plt.bar(ind, survived, width)
p2 = plt.bar(ind, not_survived, width, bottom=survived)
plt.ylabel("Cantidad")
plt.title("Supervivencia por clase social")
plt.xticks(ind, ("1ra clase", "2da clase", "3ra clase"))
# plt.yticks(np.arange(0, 81, 10))
plt.legend((p1[0], p2[0]), ("Sobrevivieron", "No sobrevivieron"))
plt.show()
# <h4>Respuesta</h4>
# <ol>
# <li>La tercera clase doblaba a la 2da y 1ra clase en cantidad de pasajeros.</li>
# <li>Si eras de 1ra clase tenias un 62% de supervivencia.</li>
# <li>Si eras de 2da clase tenias un 47% de supervivencia.</li>
# <li>Si eras de 3ra clase tenias un 24% de supervivencia.</li>
# </ol>
# <h3>D. Conclusiones</h3>
# <p>En esta sección:</p>
# <ol>
# <li>Presentaremos las conclusiones de nuestro estudio</li>
# </ol>
# <p>Las conclusiones son las respuestas a las preguntas previas.</p>
# <h5>Puedes agregar mas conclusiones?</h5>
# <ol>
# <li>Agrega aqui tus conclusiones.</li>
# </ol>
# <h2>2. Clasificación con Machine Learning</h2>
# <h3>A. Pre-procesamiento de variables</h3>
# Dividimos la matriz en X y Y.
Y = df_cleaned["Survived"]
X = df_cleaned.drop("Survived", axis = 1)
print(X.shape, Y.shape)
# <h3>B. Train, validation y test sets</h3>
# <h3>EJERCICIO</h3>
# <p>Debes escoger el porcentaje de datos que ocupara el training set y lo mismo para el validation set. Recuerda los valores que vimos en clase.</p>
# <p><strong>Nota: </strong>En esta ocacion solo necesitamos un train set y un validation set. El test set lo provee Kaggle.</p>
# TRAIN SIZE Y VALIDATION SIZE deben sumar 1.0.
# Por ejemplo: TRAIN_SIZE = 0.5 y VALIDATION_SIZE = 0.5
TRAIN_SIZE = # ESCOJE EL VALOR DEL TRAINING SET
VALIDATION_SIZE = # ESCOJE EL VALOR DEL VALIDATION SET
assert TRAIN_SIZE+VALIDATION_SIZE==1.0, "TRAIN SIZE y VALIDATION SIZE deben sumar 1.00"
# Training y validation set.
x_train, x_validation, y_train, y_validation = train_test_split(X, Y, test_size=TEST_SIZE, random_state=42)
print("Tamaños train set: ", x_train.shape, y_train.shape)
print("Tamaños validation set: ", x_validation.shape, y_validation.shape)
# <h3>C. Modelos para clasificación</h3>
# <h3>Ejercicios</h3>
# <p>Tu trabajo es entrenar los modelos de aprendizaje automatico evitando el overfitting y underfitting. Para lo cual analizaras la precision de tu modelo utilizando el train y el validation set. <strong>Recuerda que ambas precisiones deben estar lo mas cerca posible entre ellas para obtener un buen resultado.</strong></p>
# <h3>Random forest</h3>
# <p>Utiliza la documentacion en el siguiente link para tunear los parametros de un random forest: <a href="http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html">Random forest</a></p>
# Random forest
from sklearn.ensemble import RandomForestClassifier
randForClf = RandomForestClassifier(n_estimators = , # CAMBIA EL VALOR DE ESTE PARAMETRO
max_features = , # CAMBIA EL VALOR DE ESTE PARAMETRO
max_depth = , # CAMBIA EL VALOR DE ESTE PARAMETRO
min_samples_leaf = , # CAMBIA EL VALOR DE ESTE PARAMETRO
min_samples_split = ) # CAMBIA EL VALOR DE ESTE PARAMETRO
randForClf.fit(x_train, y_train)
# <p>Utiliza la siguiente celda para debugear tu entrenamiento. Recuerda que el score del training y el validation deben estar lo mas cerca posible.</p>
print("Score en training set: ", randForClf.score(x_train, y_train))
print("Score en validation set: ", randForClf.score(x_validation, y_validation))
# <h3>SVM</h3>
# <p>Utiliza la documentacion en el siguiente link para tunear los parametros de un SVM: <a href="http://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html">SVM</a></p>
# <br>
# <p>Puedes escoger entre los siguientes tipos de kernel: </p>
# <ol>
# <li>"linear"</li>
# <li>"rbf"</li>
# <li>"sigmoid"</li>
# </ol>
# SVM
from sklearn.svm import SVC
svcclf = SVC(kernel = "sigmoid") # ESCOJE EL TIPO DE KERNEL DE EN ESTE PARAMETRO
svcclf.fit(x_train, y_train)
print("Score en training set: ", svcclf.score(x_train, y_train))
print("Score en validation set: ", svcclf.score(x_validation, y_validation))
# <h3>Adaboost</h3>
# <p>Utiliza la documentacion en el siguiente link para tunear los parametros de un Adaboost: <a href="http://scikit-learn.org/stable/auto_examples/ensemble/plot_adaboost_hastie_10_2.html#sphx-glr-auto-examples-ensemble-plot-adaboost-hastie-10-2-py">Adaboost</a></p>
# Adaboost
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import zero_one_loss
from sklearn.ensemble import AdaBoostClassifier
# Clasifier
treeclf = DecisionTreeClassifier(max_depth= , # ESCOJE EL MAX DEPTH
min_samples_leaf= ) # ESCOJE EL MIN SAMPLES LEAF
treeclf.fit(x_train, y_train)
adareal = AdaBoostClassifier(
base_estimator=treeclf,
learning_rate = , # ESCOJE UNA TASA DE APRENDIZAJE
n_estimators = , # ESCOJE UN NUMERO DE ESTIMADORES
algorithm="SAMME.R")
adareal.fit(x_train, y_train)
print("Score en training set: ", adareal.score(x_train, y_train))
print("Score en validation set: ", adareal.score(x_validation, y_validation))
# <h3>F. Resultados</h3>
# <p>Cuando creas que tus resultados son correctos. Prueba con el test set provisto por Kaggle. Para esto corre el codigo de las siguientes celdas.</p>
# <h3>EJERCICIO</h3>
# <p>Escribe tu nombre y la version de tu archivo en las siguientes variables.</p>
your_name = "Rodrigo" # ESCRIBE TU NOMBRE AQUI
your_version = "1" # ESCRIBE LA VERSION DE TU ARHIVO
# Leer datos de entrada.
df_test_set = pd.read_csv("data/test.csv")
df_passenger_id = df_test_set["PassengerId"]
# Eliminar features que no usamos.
df_test_set = df_test_set.drop(["PassengerId", "Name", "Cabin", "Embarked", "Parch", "Ticket"], axis = 1)
# Limpiar datos.
if (opcion == 1):
df_test_set["Age"] = df_test_set["Age"].fillna(df_test_set["Age"].mean())
df_test_set["Fare"] = df_test_set["Fare"].fillna(df_test_set["Fare"].mean())
elif (opcion == 2):
df_test_set["Age"] = df_test_set["Age"].fillna(df_test_set["Age"].median())
df_test_set["Fare"] = df_test_set["Fare"].fillna(df_test_set["Fare"].median())
elif (opcion == 3):
df_test_set = df_test_set.dropna(axis = 1)
# Categorizar variables.
df_test_set = pd.get_dummies(df_test_set)
# Preprocesar datos.
x_test = df_test_set
# <h3>ESCRIBE EL NOMBRE DEL MODELO QUE ESCOJES PARA EL TEST SET EN LA SIGUIENTE CELDA.</h3>
# <p>Opciones:</p>
# <ol>
# <li>"Random forests"</li>
# <li>"SVM"</li>
# <li>"Adaboost"</li>
# </ol>
# Escribe aqui el nombre de tu modelo.
MODELO_ESCOGIDO = # ESCRIBE EL MODELO EN UN STRING
# Procesar resultados del modelo
if (MODELO_ESCOGIDO == "Random forests"):
resultados = randForClf.predict(x_test)
elif (MODELO_ESCOGIDO == "SVM"):
resultados = svcclf.predict(x_test)
elif (MODELO_ESCOGIDO == "Adaboost"):
resultados = adareal.predict(x_test)
else:
print("El nombre no es valido")
# Escribir archivo de salida.
file = open("data/enviar_archivo_" + your_name + "_" + your_version + "_kaggle.csv", "w")
file.write("PassengerId,Survived\n")
for i, j in zip(df_passenger_id, resultados):
file.write(str(i) + "," + str(j) +"\n")
file.close()
# <p>El archivo con tus resultados se encuentra en la carpeta data y se llama: <strong>enviar_archivo_YOURNAME_YOURVERSION_kaggle.csv</strong></p>
# <p>La siguiente imagen te guiara donde debes enviar tu archivo en kaggle.</p>
# 
| Proyecto de curso Titanic/Notebook_titanic.ipynb |