code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Justusm10moringaschool/100-Days-Of-ML-Code/blob/master/Copy_of_Copy_of_Moringa_Data_Science_Prep_W1_Independent_Project_2019_06_VIVIAN_NJAU_Python_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="KAT1A-SK_bCk" outputId="5861fead-26fb-4667-a604-29639604f731" colab={"base_uri": "https://localhost:8080/", "height": 86}
#converting pounds to kgs
x = float(input("Enter the first value in pounds "))
y = float(input("Enter the second value in pounds "))
#converting pounds to kgs
a = x*0.4535922
b = y*0.4535922
print (str(x) + "pounds in kgs is " + str(a) )
print (str(y) + "pounds in kgs is " + str(b) )
# + id="dusjOaTtBHcN" outputId="778f8a14-9705-4523-8c70-b2117104dc55" colab={"base_uri": "https://localhost:8080/", "height": 34}
#sum of the 2 values
z = x + y
print (" The sum is " + str(z))
# + id="moVRwu_AE2bg" outputId="44d754f4-c59c-43e4-a5ee-a2a91706cfc0" colab={"base_uri": "https://localhost:8080/", "height": 34}
#average of the 2 values
c = (x + y)/2
print (" The average is " + str(c))
# + id="G4iec9MgFB5i" outputId="4871c897-c886-48e6-de43-b58c78666c23" colab={"base_uri": "https://localhost:8080/", "height": 51}
#difference between the two values
d = x - y
print (" The difference between " + str(x) + " and " + str(y) + " is " + str(d))
g = y - x
print (" The difference between " + str(y) + " and " + str(x) + " is " + str(g))
# + id="Wsy90jSbFOUb" outputId="b655f652-85ac-4850-f7af-e7859176ea29" colab={"base_uri": "https://localhost:8080/", "height": 34}
#quotient of the 2 values
e = x / y
print (" The quotient is " + str(e))
# + id="ScCy6CsyFwrU" outputId="0bc7b037-e8f9-449e-e967-ff9b72819534" colab={"base_uri": "https://localhost:8080/", "height": 51}
#even or odd!
if x % 2 == 0:
print (" The number" + str(x) + " is even")
elif x % 2 != 0:
print (" The number " + str(x) + " is odd")
if y % 2 == 0:
print (" The number" + str(y) + " is even")
elif y % 2 != 0:
print (" The number " + str(y) + " is odd")
# + id="fkzlBKFeGbQe"
| Copy_of_Copy_of_Moringa_Data_Science_Prep_W1_Independent_Project_2019_06_VIVIAN_NJAU_Python_Notebook.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 📝 Exercise M1.04
#
# The goal of this exercise is to evaluate the impact of using an arbitrary
# integer encoding for categorical variables along with a linear
# classification model such as Logistic Regression.
#
# To do so, let's try to use `OrdinalEncoder` to preprocess the categorical
# variables. This preprocessor is assembled in a pipeline with
# `LogisticRegression`. The generalization performance of the pipeline can be
# evaluated by cross-validation and then compared to the score obtained when
# using `OneHotEncoder` or to some other baseline score.
#
# First, we load the dataset.
# +
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
# -
target_name = "class"
target = adult_census[target_name]
data = adult_census.drop(columns=[target_name, "education-num"])
# In the previous notebook, we used `sklearn.compose.make_column_selector` to
# automatically select columns with a specific data type (also called `dtype`).
# Here, we will use this selector to get only the columns containing strings
# (column with `object` dtype) that correspond to categorical features in our
# dataset.
# +
from sklearn.compose import make_column_selector as selector
categorical_columns_selector = selector(dtype_include=object)
categorical_columns = categorical_columns_selector(data)
data_categorical = data[categorical_columns]
# -
# Define a scikit-learn pipeline composed of an `OrdinalEncoder` and a
# `LogisticRegression` classifier.
#
# Because `OrdinalEncoder` can raise errors if it sees an unknown category at
# prediction time, you can set the `handle_unknown="use_encoded_value"` and
# `unknown_value` parameters. You can refer to the
# [scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OrdinalEncoder.html)
# for more details regarding these parameters.
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import OrdinalEncoder
from sklearn.linear_model import LogisticRegression
# Write your code here.
# -
# Your model is now defined. Evaluate it using a cross-validation using
# `sklearn.model_selection.cross_validate`.
# +
from sklearn.model_selection import cross_validate
# Write your code here.
# -
# Now, we would like to compare the generalization performance of our previous
# model with a new model where instead of using an `OrdinalEncoder`, we will
# use a `OneHotEncoder`. Repeat the model evaluation using cross-validation.
# Compare the score of both models and conclude on the impact of choosing a
# specific encoding strategy when using a linear model.
# +
from sklearn.preprocessing import OneHotEncoder
# Write your code here.
| notebooks/03_categorical_pipeline_ex_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func
engine = create_engine(f"sqlite:///Resources/hawaii.sqlite")
# reflect an existing database into a new model
Base=automap_base()
# reflect the tables
Base.prepare(engine,reflect=True)
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement=Base.classes.measurement
Station=Base.classes.station
# Create our session (link) from Python to the DB
session=Session(engine)
# Get all the columns of Measurement
from sqlalchemy import inspect
inspector=inspect(engine)
columns=inspector.get_columns("Measurement")
columns
# Get all the columns of Station
columns2=inspector.get_columns("Station")
columns2
# Get the last date
last_date=session.query(func.max(Measurement.date)).all()
last_date
# # Exploratory Climate Analysis
# Design a query to retrieve the last 12 months of precipitation data and plot the results
# Calculate the date 1 year ago from the last data point in the database
# Perform a query to retrieve the data and precipitation scores
precipitation=session.query(Measurement.date,Measurement.prcp).\
filter(Measurement.date>='2016-08-23').\
filter(Measurement.date<='2017-08-23').all()
precipitation[0:10]
# Save the query results as a Pandas DataFrame and set the index to the date column
precipitation_df=pd.DataFrame(precipitation,columns=["date","precipitation"])
precipitation_df.set_index("date",inplace=True)
precipitation_df.head(10)
# Sort the dataframe by date
precipitation_df=precipitation_df.sort_values(by="date")
precipitation_df.head(10)
# Use Pandas Plotting with Matplotlib to plot the data
fig1,ax1=plt.subplots(figsize=(20,12))
precipitation_df.plot(ax=ax1,grid=True,rot=90)
plt.xlabel("Date")
plt.ylabel("Inches")
plt.title("Precipitation of Past 12 Months")
plt.tight_layout()
plt.show()
# Use Pandas to calcualte the summary statistics for the precipitation data
precipitation_df["precipitation"].describe()
# Design a query to show how many stations are available in this dataset?
from sqlalchemy import distinct
number_of_stations=session.query(distinct(Station.id)).count()
number_of_stations
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
station_rows=session.query(Measurement.station,func.count(Measurement.station)).group_by(Measurement.station).order_by(func.count(Measurement.station).desc()).all()
station_rows
# ### USC00519281 has the highest number of rows.
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature of the most active station?
lowest_temp=session.query(func.min(Measurement.tobs)).filter(Measurement.station=="USC00519281").first()
highest_temp=session.query(func.max(Measurement.tobs)).filter(Measurement.station=="USC00519281").first()
average_temp=session.query(func.avg(Measurement.tobs)).filter(Measurement.station=="USC00519281").first()
print(f"The lowest temperature of station USC00519281 is {lowest_temp[0]}.")
print(f"The highest temperature of station USC00519281 is {highest_temp[0]}.")
print(f"The average temperature of station USC00519281 is {average_temp[0]}.")
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
temperatures=session.query(Measurement.date,Measurement.tobs).\
filter(Measurement.station=="USC00519281").\
filter(Measurement.date>='2016-08-23').\
filter(Measurement.date<='2017-08-23').all()
tobs_df=pd.DataFrame(temperatures,columns=["date","temperatures"])
tobs_df.head()
fig2,ax2=plt.subplots(figsize=(11,6))
tobs_df.plot.hist(ax=ax2,bins=12,grid=True)
plt.xlabel("Temperature")
plt.ylabel("Frequency")
plt.title("Temperature Frequency of USC00519281 Station")
plt.tight_layout()
plt.show()
# ## Bonus Challenge Assignment
# +
# Temperature Analysis I
# Filter temperatures of June
june_temp=session.query(Measurement.tobs).filter(func.strftime("%m",Measurement.date)=="06").all()
# Filter temperatures of December
december_temp=session.query(Measurement.tobs).filter(func.strftime("%m",Measurement.date)=="12").all()
# Run a t-test with the null hypothesis that temperatures of June and December are the same.
from scipy import stats
stats.ttest_ind(june_temp,december_temp)
# -
# ### Since the p-value of the t-test is less than 0.05, the null hypothesis will be rejected, which infers that the June temperatures of Hawaii are significantly different from December temperatures.
# +
# Temperature Analysis II
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# -
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
# I choose to start my trip on 08-01 and end my trip on 08-09.
trip_temp=calc_temps("2017-08-01","2017-08-09")
trip_temp
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
min_temp=trip_temp[0][0]
average_temp=trip_temp[0][1]
max_temp=trip_temp[0][2]
ptp=max_temp-min_temp
plt.figure(figsize=(2,5))
plt.bar(0.2,average_temp,yerr=ptp,width=0.3,color="red",alpha=0.5)
plt.title("Trip Avg Temp")
plt.ylabel("Temp(F)")
plt.ylim(0,max_temp+20)
plt.xticks([])
plt.show()
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
start_date="2017-08-01"
end_date="2017-08-09"
prcp_station=session.query(Station.station,func.sum(Measurement.prcp),Station.name,Station.latitude,Station.longitude,Station.elevation).\
filter(Measurement.date>=start_date).filter(Measurement.date<=end_date).filter(Measurement.station==Station.station).\
group_by(Station.station).order_by(func.sum(Measurement.prcp).desc()).all()
for row in prcp_station:
print(f"Station: {row[0]}")
print(f"Total amount of rainfall: {row[1]}")
print(f"Station name: {row[2]}")
print(f"Station latitude: {round(row[3],4)}")
print(f"Station longitude: {round(row[4],4)}")
print(f"Station elevation: {row[5]}")
print("\n")
# +
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# +
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
# Use the start and end date to create a range of dates
# Stip off the year and save a list of %m-%d strings
# Loop through the list of %m-%d strings and calculate the normals for each date
start_year=2017
start_month=8
start_day=1
duration=9
date_range=[]
date_list=[]
for i in range(duration):
date_range.append(dt.date(start_year,start_month,start_day+i))
date_list.append(dt.date(start_year,start_month,start_day+i).strftime("%m-%d"))
normals=[]
for date in date_list:
normals.append(daily_normals(date)[0])
normals
# -
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
trip_df=pd.DataFrame(normals,columns=["tmin","tavg","tmax"])
trip_df["date"]=date_range
trip_df.set_index("date",inplace=True)
trip_df
# Plot the daily normals as an area plot with `stacked=False`
x=trip_df.index
fig3,ax3=plt.subplots(figsize=(11,6))
trip_df.plot(ax=ax3,stacked=False,rot=45)
plt.ylabel("Temperature")
plt.title("Temperatures During the Trip")
plt.ylim(0,max(trip_df["tmax"]+5))
ax3.fill_between(x,0,trip_df["tmin"],alpha=0.2)
ax3.fill_between(x,trip_df["tmin"],trip_df["tavg"],alpha=0.2)
ax3.fill_between(x,trip_df["tavg"],trip_df["tmax"],alpha=0.2)
plt.tight_layout()
plt.show()
| climate_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" style="padding-right:10px; width:150px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/mindboggle_logo_200px_600dpi.jpg">
# This jupyter notebook provides a tutorial for the Mindboggle brain image analysis software.<br> Website: http://mindboggle.info<br> Reference: http://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1005350#sec007
# # Mindboggle tutorial
#
# 1. [Mindboggle processing steps](#processing)
# 2. [Run "mindboggle --help" on the command line](#help)
# 3. [Mindboggle on the command line](#command)
# 4. [Mindboggle Python library](#library)
# 5. [Run individual functions](#functions)
# <a id="processing"></a>
#
# # Mindboggle processing steps
#
# <img style="padding-right:10px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/mindboggle_nipype_flow.png">
#
# ## Output
# Mindboggle takes in (FreeSurfer- and optionally ANTs-) preprocessed T1-weighted MRI data, and outputs nifti volumes, vtk surfaces, and csv tables containing label, feature, and shape information for further analysis:
#
# - **labels/**: *integer-labeled vtk surfaces and nifti volumes*
# - **features/**: *integer-labeled sulci or fundi on vtk surfaces*
# - **shapes/**: *float shape value for every point on vtk surfaces*
# - **tables/**: *csv tables of shape values for every label/feature/vertex*
#
# <br><br>
# ## Processing steps
#
# ### 1. Combine FreeSurfer and ANTs gray/white segmentations:
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Figure2_FSvsANTs_segmentation_white_188mm_600dpi.png">
# <br>
# ### 2. Fill hybrid segmentation with (FreeSurfer- or ANTs-registered) labels.
# ### 3. Compute volume shape measures for each labeled region:
#
# <b>volume</b>
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/posters/roygbiv-volume.png">
# <br>
# ### 4. Compute surface shape measures for every cortical mesh vertex
#
# <b>surface area</b>
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Figure4_area_182mm_300dpi.png">
# <br><br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/HBM2015_ROYGBIV.png">
# <br>
# <br>
#
# <b>geodesic depth and travel depth</b>
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Figure6_geodesic_and_travel_depth_on_sulci_190mm_600dpi.png">
# <br>
# <br>
#
# <b>mean curvature</b>
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Mindboggle_bioRxiv2016_Fig5.png">
# <br>
# ### 5. Extract cortical surface features:
#
# <b>folds</b> (left and upper right, with manual labels)
# <br>
#
# <b>sulci</b> (lower right)
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Mindboggle_bioRxiv2016_Fig7.png">
#
#
# <b>fundi</b>
# <img style="padding-right:10px; width:300px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Figure8_fundus_views_78mm_600dpi_rotated.png">
# <br>
# ### 6. For each cortical surface label/sulcus, compute:
#
# <b>Zernike moments</b>
# <br>
#
# <b>Laplace-Beltrami spectrum</b> (2nd, 3rd, and 9th spectral components shown for two brains):
# <br>
# <br>
# <img style="padding-right:10px; width:600px;" src="https://raw.githubusercontent.com/nipy/mindboggle-assets/master/images/papers/Figure9_LaplaceBeltrami3rd6th9thSpectralComponents_190mm_600dpi.png">
# ### 7. Compute statistics for each shape measure across vertices for each label/feature:
# - median
# - median absolute deviation
# - mean
# - standard deviation
# - skew
# - kurtosis
# - lower and upper quartiles
# <br>
# <a id="help"></a>
#
# # Run "mindboggle --help" on the command line
#
# First, let's see what command-line options Mindboggle provides:
# ! mindboggle --help
# <a id="command"></a>
#
# # Mindboggle on the command line
#
# The following command computes shape measures for cortical surface labels and sulci from [example FreeSurfer and ANTs data](https://osf.io/3xfb8/?action=download&version=1). Without adding more restrictive arguments, this command takes an hour or two to run.
#
# mindboggle /home/jovyan/work/mindboggle_input_example/freesurfer/subjects/arno<br> --ants /home/jovyan/work/mindboggle_input_example/ants/subjects/arno/antsBrainSegmentation.nii.gz<br> --out /home/jovyan/work/mindboggled<br> --roygbiv<br><br><br>
# <a id="library"></a>
#
# # Mindboggle Python library
#
# Rather than call Mindboggle from the command line, we can also call individual Python functions within the Mindboggle library, which includes the following files in mindboggle/mindboggle/:
#
# - **mio**/ *-- input/output functions*
# - **[colors](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/colors.py)**.py *-- colormap-related functions*
# - **[convert_volumes](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/convert_volumes.py)**.py *-- read/write nifti volume files*
# - **[fetch_data](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/fetch_data.py)**.py *-- fetch data from a URL or from third party software*
# - **[labels](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/labels.py)**.py *-- information about labeling protocols*
# - **[plots](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/plots.py)**.py *-- plot functions*
# - **[tables](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/tables.py)**.py *-- read/write tables*
# - **[vtks](https://github.com/nipy/mindboggle/blob/master/mindboggle/mio/vtks.py)**.py *-- read/write VTK surface files*
#
# - **guts**/ *-- the "guts" underlying feature extraction and labeling code*
# - **[compute](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/compute.py)**.py *-- compute distances, etc.*
# - **[graph](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/graph.py)**.py *-- graph operations*
# - **[kernels](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/kernels.py)**.py *-- kernels for graph operations*
# - **[mesh](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/mesh.py)**.py *-- operate on surface mesh vertices*
# - **[paths](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/paths.py)**.py *-- connect surface mesh vertices*
# - **[rebound](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/rebound.py)**.py *-- adjust label borders on a surface mesh*
# - **[relabel](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/relabel.py)**.py *-- relabel surface or volume files*
# - **[segment](https://github.com/nipy/mindboggle/blob/master/mindboggle/guts/segment.py)**.py *-- segment a surface mesh*
#
# - **shapes**/ *-- shape measurement functions
# - **[surface_shapes](https://github.com/nipy/mindboggle/blob/master/mindboggle/shapes/surface_shapes.py)**.py *-- compute surface shapes (calls C++ library below)*
# - **[volume_shapes](https://github.com/nipy/mindboggle/blob/master/mindboggle/shapes/volume_shapes.py)**.py *-- compute volumes and thicknesses*
# - **[laplace_beltrami](https://github.com/nipy/mindboggle/blob/master/mindboggle/shapes/laplace_beltrami.py)**.py *-- compute a Laplace-Beltrami spectrum*
# - **[zernike/zernike](https://github.com/nipy/mindboggle/blob/master/mindboggle/shapes/zernike/zernike.py)**.py *-- compute Zernike moments of a collection of vertices*
# - **[likelihood](https://github.com/nipy/mindboggle/blob/master/mindboggle/shapes/likelihood.py)**.py *-- compute (fundus) likelihood values*
#
# - **features**/ *-- higher-level feature extraction (folds, fundi, sulci)*
# - **[folds](https://github.com/nipy/mindboggle/blob/master/mindboggle/features/folds.py)**.py *-- extract surface folds*
# - **[fundi](https://github.com/nipy/mindboggle/blob/master/mindboggle/features/fundi.py)**.py *-- extract fundus curves from folds*
# - **[sulci](https://github.com/nipy/mindboggle/blob/master/mindboggle/features/sulci.py)**.py *-- extract sulci from folds*
#
# <!--
# - **thirdparty/** *-- third-party code*
# - **[ants](https://github.com/nipy/mindboggle/blob/master/mindboggle/thirdparty/ants.py)**.py *-- call ANTs commands*
# - **[vtkviewer](https://github.com/nipy/mindboggle/blob/master/mindboggle/thirdparty/vtkviewer.py)**.py *-- VTK viewer (by <NAME>)*
# - mindboggle/vtk_cpp_tools *-- C++ tools for measuring shapes on VTK surface files*
# - **[area](https://github.com/nipy/mindboggle/blob/master/vtk_cpp_tools/area/PointAreaMain.cpp)**/
# - **[curvature](https://github.com/nipy/mindboggle/blob/master/vtk_cpp_tools/curvature/CurvatureMain.cpp)**/
# - **[geodesic_depth](https://github.com/nipy/mindboggle/blob/master/vtk_cpp_tools/geodesic_depth/GeodesicDepthMain.cpp)**/
# - **[travel_depth](https://github.com/nipy/mindboggle/tree/master/vtk_cpp_tools/travel_depth)**/
# - **[gradient](https://github.com/nipy/mindboggle/blob/master/vtk_cpp_tools/gradient/GradientMain.cpp)**/
# - **[surface_overlap](https://github.com/nipy/mindboggle/blob/master/vtk_cpp_tools/surface_overlap/SurfaceOverlapMain.cpp)**/
# -->
# <br>
# <br>
# <a id="functions"></a>
#
# # Run individual functions
#
# Let's run some functions within Mindboggle's Python library. The following code snippets are adapted from the above files' docstrings.
#
# ## Example: Compute statistics of depth measures in sulcus folds
# ### Measure travel depth for every vertex of a brain's left hemisphere
# Convert a FreeSurfer surface file to VTK format:
from mindboggle.mio.vtks import freesurfer_surface_to_vtk
subject_path = '/home/jovyan/work/mindboggle_input_example/freesurfer/subjects/arno/'
surface_file = freesurfer_surface_to_vtk(surface_file=subject_path + 'surf/lh.pial',
orig_file=subject_path + 'mri/orig.mgz',
output_vtk='lh.pial.vtk')
# Compute travel_depth for every vertex of the mesh in the VTK file:
import os
from mindboggle.shapes.surface_shapes import travel_depth
from mindboggle.mio.vtks import read_scalars
ccode_path = '/opt/vtk_cpp_tools'
command = os.path.join(ccode_path, 'travel_depth', 'TravelDepthMain')
depth_file = travel_depth(command=command,
surface_file=surface_file,
verbose=True)
depths, name = read_scalars(depth_file)
# Plot the depth values in 3-D: <b>UNDER CONSTRUCTION<b>
import numpy as np
import nilearn.plotting
depths_array = np.asarray(depths)
fs_surf_mesh = subject_path + 'surf/lh.pial'
nilearn.plotting.plot_surf(surf_mesh=fs_surf_mesh, surf_map=depths_array,
bg_map=None, hemi='left', view='lateral',
cmap=None, colorbar=True,
avg_method='mean', threshold=None,
alpha='auto', bg_on_data=False, darkness=1,
vmin=None, vmax=None, cbar_vmin=None, cbar_vmax=None,
title=None, output_file=None, axes=None, figure=None)
# ### Extract folds based on depth and curvature
# Plot a histogram of the depth values:
# %matplotlib inline
from mindboggle.mio.plots import histograms_of_lists
histograms_of_lists(columns=[depths],
column_name='Depth values',
ignore_columns=[],
nbins=100,
axis_limits=[],
titles='depth values')
# Find a depth threshold to extract folds from the surface:
from mindboggle.features.folds import find_depth_threshold
depth_threshold, bins, bin_edges = find_depth_threshold(depth_file=depth_file,
min_vertices=10000,
verbose=True)
depth_threshold
# Extract folds with the depth threshold:
from mindboggle.features.folds import extract_folds
folds, n_folds, folds_file = extract_folds(depth_file=depth_file,
depth_threshold=depth_threshold,
min_fold_size=50,
save_file=True,
output_file='folds.vtk',
background_value=-1,
verbose=True)
# Remove all vertices but the folds:
from mindboggle.mio.vtks import rewrite_scalars
rewrite_scalars(input_vtk=folds_file,
output_vtk='rewrite_scalars.vtk',
new_scalars=[folds],
new_scalar_names=['folds'],
filter_scalars=folds,
background_value=-1)
# Plot the folds in 3-D: <b>UNDER CONSTRUCTION<b>
folds_array = np.asarray(folds)
nilearn.plotting.plot_surf(surf_mesh=fs_surf_mesh, surf_map=folds_array,
bg_map=None, hemi='left', view='lateral',
cmap=None, colorbar=True,
avg_method='mean', threshold=None,
alpha='auto', bg_on_data=False, darkness=1,
vmin=None, vmax=None, cbar_vmin=None, cbar_vmax=None,
title=None, output_file=None, axes=None, figure=None)
# ### Extract sulci from folds
# Load a FreeSurfer .annot file and save as a VTK format file:
from mindboggle.mio.vtks import freesurfer_annot_to_vtk
labels, label_file = freesurfer_annot_to_vtk(annot_file=subject_path + 'label/lh.aparc.annot',
vtk_file=surface_file,
output_vtk='lh.aparc.annot.vtk',
background_value=-1)
# Relabel surface labels to match expected volume labels:
from mindboggle.guts.relabel import relabel_surface
from mindboggle.mio.labels import DKTprotocol
dkt = DKTprotocol()
relabel_file = relabel_surface(vtk_file=label_file,
hemi='lh',
old_labels=dkt.DKT31_numbers,
new_labels=[],
erase_remaining=True,
erase_labels=[0],
erase_value=-1,
output_file='relabeled.vtk')
# Extract sulci from folds using pairs of labels in the DKT labeling protocol (SLOW):
from mindboggle.features.sulci import extract_sulci
sulci, n_sulci, sulci_file = extract_sulci(labels_file=relabel_file,
folds_or_file=folds,
hemi='lh',
min_boundary=10,
sulcus_names=[],
save_file=True,
output_file='sulci.vtk',
background_value=-1,
verbose=True)
n_sulci
# Plot the sulci in 3-D: <b>UNDER CONSTRUCTION<b>
sulci_array = np.asarray(sulci)
nilearn.plotting.plot_surf(surf_mesh=fs_surf_mesh, surf_map=sulci_array,
bg_map=None, hemi='left', view='lateral',
cmap=None, colorbar=True,
avg_method='mean', threshold=None,
alpha='auto', bg_on_data=False, darkness=1,
vmin=None, vmax=None, cbar_vmin=None, cbar_vmax=None,
title=None, output_file=None, axes=None, figure=None)
# ### Compute statistics on sulcus depth values
from mindboggle.mio.tables import write_shape_stats
label_table, sulcus_table, fundus_table = write_shape_stats(labels_or_file=[],
sulci=sulci, fundi=[], affine_transform_files=[], inverse_booleans=[],
transform_format='itk', area_file='', normalize_by_area=False,
mean_curvature_file='',
travel_depth_file=depth_file, geodesic_depth_file='',
freesurfer_thickness_file='', freesurfer_curvature_file='',
freesurfer_sulc_file='',
labels_spectra=[], labels_spectra_IDs=[],
sulci_spectra=[], sulci_spectra_IDs=[],
labels_zernike=[], labels_zernike_IDs=[],
sulci_zernike=[], sulci_zernike_IDs=[],
exclude_labels=[-1], verbose=True)
# Show statistical summary table of sulcus depth values:
pd.read_csv(sulcus_table)
| docs/mindboggle_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Try to read a vtk file and a numpy array file of the same data and check that they are the same.
import numpy as np
import vtk
from vtk.util.numpy_support import vtk_to_numpy
import read_rectilinear_vtk as rrv
reload(rrv)
test_file_vtk = ("/home/jensv/rsx/jens_analysis/writing_to_vtk/"
"output/2017-01-16-18-15/Bdot_triple_probe_quantities0000.vtk")
test_file_npz = ("/home/jensv/rsx/jens_analysis/writing_to_vtk/"
"output/2017-01-20-21-03/Bdot_triple_probe_quantities0000.npz")
vtk_mesh, shape = rrv.get_mesh(vtk_data)
print shape
vtk_data = rrv.get_data(test_file_vtk)
vtk_mesh, shape = rrv.get_mesh(vtk_data)
(b_x_from_vtk,
b_y_from_vtk,
b_z_from_vtk) = rrv.get_vector(vtk_data, shape,
'B_x', 'B_y', 'B_z')
vtk_b_field = [b_x_from_vtk, b_y_from_vtk, b_z_from_vtk]
npz_file = np.load(test_file_npz)
npz_mesh = npz_file['mesh']
npz_b_field = npz_file['b_field']
np.allclose(npz_b_field[0], vtk_b_field[0])
# # Make test vtk file
# Below I made a test vtk file I plotted with seaborn and opened in Visit to make sure the axis swapping that I am doing is really that right thing.
# The visual inspection in Visit shows that the data is disaplyed with the correct axis ordering.
import matplotlib.pyplot as plt
import sys
sys.path.append('../../writing_to_vtk/source/')
import write_canonical_flux_tube_quantities as write
import seaborn as sns
x = 5
y = 6
z = 7
x_line = np.arange(x)
y_line = np.arange(y)
z_line = np.arange(z)
mesh = np.meshgrid(x_line, y_line, z_line)
test_field = np.arange(0, 5*6*7)
test_field_numpy = np.reshape(test_field, (y, x, z))
sns.heatmap(test_field_numpy[:,:,0], annot=True)
plt.show()
test_field_numpy.shape
test_field_numpy[0, :, 0]
x, y, z, variables = write.prepare_for_rectilinear_grid(mesh, (test_field_numpy,), ('test',))
write.write_fields_and_currents_to_structured_mesh('test', 'test', x, y, z, variables, 0)
# # Read grid
current_grid_file = '/home/jensv/rsx/jens_analysis/output/canonical_quantities/2017-02-22-11-01/canonical_quantities0000.vtk'
vtk_mesh, b_x = rrv.read_scalar(current_grid_file, 'B_x')
print vtk_mesh[0][0, 0 ,0], vtk_mesh[0][0, -1, 0]
print vtk_mesh[1][0, 0, 0], vtk_mesh[1][-1, 0, 0]
print vtk_mesh[2][0, 0, 0], vtk_mesh[2][0, 0, -1]
| vector_comparison/test_read_vtk_rectilinear_grids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## How do you evaluate goodness of embeddings across languages?
#
# If you have embeddings for language A and another set of embeddings for language B, how can you tell how good they are?
#
# We might want to evaluate each set on a task specific to that language. But a much more interesting proposition is aligning the embeddings in an unsupervised way and evaluating the performance on translation!
#
# For this, we need 3 components. [The embeddings](https://fasttext.cc/docs/en/pretrained-vectors.html), [a way to align embeddings](https://github.com/artetxem/vecmap) and [dictionaries](https://github.com/facebookresearch/MUSE) we can use to evaluate the results.
#
# Let's begin by grabbing embeddings for English and Polish.
# !wget -P data https://dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.en.vec
# !wget -P data https://dl.fbaipublicfiles.com/fasttext/vectors-wiki/wiki.pl.vec
# Time to align the embeddings. Let's translate from Polish to English. In order to do that, let's align our Polish embeddings with our English embeddings.
#
# Bear in mind that this is a challenging task - Polish and English come from distinct and very different language families (West Slavic languages of the Lechitic group and West Germanic language of the Indo-European language family respectively).
#
# Let us clone [vecmap](https://github.com/artetxem/vecmap) to a directory adjacent to this one and perform the alignment.
# %%time
# !python ../vecmap/map_embeddings.py --cuda --unsupervised data/wiki.pl.vec data/wiki.en.vec data/wiki.pl.aligned.vec data/wiki.en.aligned.vec
# Great! We now have the English and Polish embeddings aligned to live in the same embedding space!
#
# Let's grab a Polish to English dictionary from the MUSE repository.
# !wget -P data https://dl.fbaipublicfiles.com/arrival/dictionaries/pl-en.txt
# This dictionary is just a text file with a source and target word per line
# !head data/pl-en.txt
# Let's wrap it in a Python class to make usage of it easier.
# +
#export core
class Dictionary():
def __init__(self, path_to_dict):
self.read_words(path_to_dict)
self.create_dict()
def read_words(self, path_to_dict):
source_words, target_words = [], []
with open(path_to_dict) as f:
for line in f.readlines():
src, target = line.strip().split()
source_words.append(src)
target_words.append(target)
self.source_words = source_words
self.target_words = target_words
def create_dict(self):
self.dict = {}
for src, target in zip(self.source_words, self.target_words):
self.dict[src] = target
def __getitem__(self, source_word):
return self.dict[source_word]
def __len__(self):
return len(self.source_words)
# -
pl_en_dict = Dictionary('data/pl-en.txt')
pl_en_dict['królowa']
len(pl_en_dict)
# Thanks to [nbdev](https://github.com/fastai/nbdev), we can conveniently import the Embeddings class we defined in the earlier notebook and use it here.
from embedding_gym.core import Embeddings
# Let's limit ourselves to the most common 50 000 words from each set of embeddings.
# +
# %%time
embeddings_pl = Embeddings.from_txt_file('data/wiki.pl.aligned.vec', limit=50_000)
embeddings_en = Embeddings.from_txt_file('data/wiki.en.aligned.vec', limit=50_000)
# -
# Let's see if our mechanism for translation works!
embeddings_en.nn_words_to(embeddings_pl['jest'])
# That it does - 'jest' means 'is' in Polish!
# Our translation results will be adversaly affected by synonyms. What we can however do is limit the task to words where the source and the target word exist in source and target embeddings.
# +
# %%time
topk = 1
source_embeddings = embeddings_pl
target_embeddings = embeddings_en
correct, total = 0, 0
for source_word in source_embeddings.i2w:
if source_word in pl_en_dict.source_words and pl_en_dict[source_word] in target_embeddings.i2w:
total += 1
if pl_en_dict[source_word] in embeddings_en.nn_words_to(embeddings_pl[source_word], n=topk):
correct += 1
# -
correct / total, total
# Nearly 60% accuracy with topk@1 across 20 thousand words! That seems like a great result given that we are not making any accomodations for synonyms. Let's see what results we get with topk@5.
# +
# %%time
topk = 5
source_embeddings = embeddings_pl
target_embeddings = embeddings_en
correct, total = 0, 0
for source_word in source_embeddings.i2w:
if source_word in pl_en_dict.source_words and pl_en_dict[source_word] in target_embeddings.i2w:
total += 1
if pl_en_dict[source_word] in embeddings_en.nn_words_to(embeddings_pl[source_word], n=topk):
correct += 1
# -
correct / total, total
| 02_goodness_of_embeddings_across_languages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="a6stH3tqwKF2"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
# +
data = pd.read_csv("./data/Motor-Temperature-Dataset/measures_v2.csv")
data.head()
# +
profile_id = 7
time = np.arange(0,len(data.motor_speed[data.profile_id==profile_id]) * 0.5, 0.5)
fig, ax = plt.subplots()
ax.plot(time, data.stator_yoke[data.profile_id==profile_id], '-')
ax.set_xlabel('Time (secs)')
ax.set_ylabel('Stator Yoke Temperature (\u00b0C)')
plt.grid()
plt.tight_layout()
plt.show()
# +
from scipy import stats
m, c, r_value, p_value, std_err = stats.linregress(time[(time > 600) & (time < 6000)],data.stator_yoke[(data.profile_id==profile_id)].iloc[(time > 600) & (time < 6000)])
x = time[(time > 600) & (time < 6000)]
y = m*x + c
print("y = %fx + %f" % (m,c))
fig, ax = plt.subplots()
ax.plot(time, data.stator_yoke[data.profile_id==profile_id], '-', linewidth=0.5)
ax.plot(x, y, 'r-', linewidth=2)
ax.set_xlabel('Time (secs)')
ax.set_ylabel('Stator Yoke Temperature (\u00b0C)')
plt.grid()
plt.tight_layout()
plt.show()
# +
from matplotlib import cm
y_m = data.stator_yoke[(data.profile_id==profile_id)].iloc[(time > 600) & (time < 6000)]
x = time[(time > 600) & (time < 6000)]
error = list()
for m_ in np.arange(-0.0005, 0.001, 0.00001):
for c_ in np.arange(26, 27, 0.01):
y = m_ * x + c_
error.append(np.sum(np.square(np.subtract(y, y_m))))
error = np.array(error).reshape((len(np.arange(-0.0005, 0.001, 0.00001)), len(np.arange(26, 27, 0.01))))
fig, ax = plt.subplots(subplot_kw={"projection": "3d"})
X, Y = np.meshgrid(np.arange(-0.0005, 0.001, 0.00001), np.arange(26, 27, 0.01))
ax.plot_wireframe(X, Y, error.T, cmap=cm.coolwarm, linewidth=0.01, antialiased=False)
ax.plot(m, c, np.sum(np.square(np.subtract(m * x + c, y_m))), 'ro', markersize=5, label="Least square error")
ax.set_xlabel("Value of m")
ax.set_ylabel("Value of c")
ax.set_zlabel("Total square error")
plt.legend()
plt.grid()
plt.tight_layout()
plt.show()
# -
| Module-6/Motor dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recommender Systems 2020/21
#
# ### Practice - BPR for SLIM and MF
#
# ### State of the art machine learning algorithm
# ## A few info about gradient descent
# %matplotlib inline
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import random
from scipy import stats
from scipy.optimize import fmin
# ### Gradient Descent
# <b>Gradient descent</b>, also known as <b>steepest descent</b>, is an optimization algorithm for finding the local minimum of a function. To find a local minimum, the function "steps" in the direction of the negative of the gradient. <b>Gradient ascent</b> is the same as gradient descent, except that it steps in the direction of the positive of the gradient and therefore finds local maximums instead of minimums. The algorithm of gradient descent can be outlined as follows:
#
# 1: Choose initial guess $x_0$ <br>
# 2: <b>for</b> k = 0, 1, 2, ... <b>do</b> <br>
# 3: $s_k$ = -$\nabla f(x_k)$ <br>
# 4: choose $\alpha_k$ to minimize $f(x_k+\alpha_k s_k)$ <br>
# 5: $x_{k+1} = x_k + \alpha_k s_k$ <br>
# 6: <b>end for</b>
# As a simple example, let's find a local minimum for the function $f(x) = x^3-2x^2+2$
f = lambda x: x**3-2*x**2+2
x = np.linspace(-1,2.5,1000)
plt.plot(x,f(x))
plt.xlim([-1,2.5])
plt.ylim([0,3])
plt.show()
# We can see from plot above that our local minimum is gonna be near around 1.4 or 1.5 (on the x-axis), but let's pretend that we don't know that, so we set our starting point (arbitrarily, in this case) at $x_0 = 2$
# +
x_old = 0
x_new = 2 # The algorithm starts at x=2
n_k = 0.1 # step size
precision = 0.0001
x_list, y_list = [x_new], [f(x_new)]
# returns the value of the derivative of our function
def f_gradient(x):
return 3*x**2-4*x
while abs(x_new - x_old) > precision:
x_old = x_new
# Gradient descent step
s_k = -f_gradient(x_old)
x_new = x_old + n_k * s_k
x_list.append(x_new)
y_list.append(f(x_new))
print ("Local minimum occurs at: {:.2f}".format(x_new))
print ("Number of steps:", len(x_list))
# -
# The figures below show the route that was taken to find the local minimum.
plt.figure(figsize=[10,3])
plt.subplot(1,2,1)
plt.scatter(x_list,y_list,c="r")
plt.plot(x_list,y_list,c="r")
plt.plot(x,f(x), c="b")
plt.xlim([-1,2.5])
plt.ylim([0,3])
plt.title("Gradient descent")
plt.subplot(1,2,2)
plt.scatter(x_list,y_list,c="r")
plt.plot(x_list,y_list,c="r")
plt.plot(x,f(x), c="b")
plt.xlim([1.2,2.1])
plt.ylim([0,3])
plt.title("Gradient descent (zoomed in)")
plt.show()
# ## Recap on BPR
# <NAME> al. BPR: Bayesian Personalized Ranking from Implicit Feedback. UAI2009
#
# The usual approach for item recommenders is to predict a personalized score $\hat{x}_{ui}$ for an item that reflects the preference of the user for the item. Then the items are ranked by sorting them according to that score.
#
# Machine learning approaches are tipically fit by using observed items as a positive sample and missing ones for the negative class. A perfect model would thus be useless, as it would classify as negative (non-interesting) all the items that were non-observed at training time. The only reason why such methods work is regularization.
#
# BPR use a different approach. The training dataset is composed by triplets $(u,i,j)$ representing that user u is assumed to prefer i over j. For an implicit dataset this means that u observed i but not j:
# $$D_S := \{(u,i,j) \mid i \in I_u^+ \wedge j \in I \setminus I_u^+\}$$
#
# ### BPR-OPT
# A machine learning model can be represented by a parameter vector $\Theta$ which is found at fitting time. BPR wants to find the parameter vector that is most probable given the desired, but latent, preference structure $>_u$:
# $$p(\Theta \mid >_u) \propto p(>_u \mid \Theta)p(\Theta) $$
# $$\prod_{u\in U} p(>_u \mid \Theta) = \dots = \prod_{(u,i,j) \in D_S} p(i >_u j \mid \Theta) $$
#
# The probability that a user really prefers item $i$ to item $j$ is defined as:
# $$ p(i >_u j \mid \Theta) := \sigma(\hat{x}_{uij}(\Theta)) $$
# Where $\sigma$ represent the logistic sigmoid and $\hat{x}_{uij}(\Theta)$ is an arbitrary real-valued function of $\Theta$ (the output of your arbitrary model).
#
#
# To complete the Bayesian setting, we define a prior density for the parameters:
# $$p(\Theta) \sim N(0, \Sigma_\Theta)$$
# And we can now formulate the maximum posterior estimator:
# $$BPR-OPT := \log p(\Theta \mid >_u) $$
# $$ = \log p(>_u \mid \Theta) p(\Theta) $$
# $$ = \log \prod_{(u,i,j) \in D_S} \sigma(\hat{x}_{uij})p(\Theta) $$
# $$ = \sum_{(u,i,j) \in D_S} \log \sigma(\hat{x}_{uij}) + \log p(\Theta) $$
# $$ = \sum_{(u,i,j) \in D_S} \log \sigma(\hat{x}_{uij}) - \lambda_\Theta ||\Theta||^2 $$
#
# Where $\lambda_\Theta$ are model specific regularization parameters.
#
# ### BPR learning algorithm
# Once obtained the log-likelihood, we need to maximize it in order to find our obtimal $\Theta$. As the crierion is differentiable, gradient descent algorithms are an obvious choiche for maximization.
#
# Gradient descent comes in many fashions, you can find an overview on Cesare Bernardis thesis https://www.politesi.polimi.it/bitstream/10589/133864/3/tesi.pdf on pages 18-19-20. A nice post about momentum is available here https://distill.pub/2017/momentum/
#
# The basic version of gradient descent consists in evaluating the gradient using all the available samples and then perform a single update. The problem with this is, in our case, that our training dataset is very skewed. Suppose an item i is very popular. Then we habe many terms of the form $\hat{x}_{uij}$ in the loss because for many users u the item i is compared against all negative items j.
#
# The other popular approach is stochastic gradient descent, where for each training sample an update is performed. This is a better approach, but the order in which the samples are traversed is crucial. To solve this issue BPR uses a stochastic gradient descent algorithm that choses the triples randomly.
#
# The gradient of BPR-OPT with respect to the model parameters is:
# $$\frac{\partial BPR-OPT}{\partial \Theta} = \sum_{(u,i,j) \in D_S} \frac{\partial}{\partial \Theta} \log \sigma (\hat{x}_{uij}) - \lambda_\Theta \frac{\partial}{\partial\Theta} || \Theta ||^2$$
# $$ = \sum_{(u,i,j) \in D_S} \frac{-e^{-\hat{x}_{uij}}}{1+e^{-\hat{x}_{uij}}} \frac{\partial}{\partial \Theta}\hat{x}_{uij} - \lambda_\Theta \Theta $$
#
# ### BPR-MF
#
# In order to practically apply this learning schema to an existing algorithm, we first split the real valued preference term: $\hat{x}_{uij} := \hat{x}_{ui} − \hat{x}_{uj}$. And now we can apply any standard collaborative filtering model that predicts $\hat{x}_{ui}$.
#
# The problem of predicting $\hat{x}_{ui}$ can be seen as the task of estimating a matrix $X:U×I$. With matrix factorization teh target matrix $X$ is approximated by the matrix product of two low-rank matrices $W:|U|\times k$ and $H:|I|\times k$:
# $$X := WH^t$$
# The prediction formula can also be written as:
# $$\hat{x}_{ui} = \langle w_u,h_i \rangle = \sum_{f=1}^k w_{uf} \cdot h_{if}$$
# Besides the dot product ⟨⋅,⋅⟩, in general any kernel can be used.
#
# We can now specify the derivatives:
# $$ \frac{\partial}{\partial \theta} \hat{x}_{uij} = \begin{cases}
# (h_{if} - h_{jf}) \text{ if } \theta=w_{uf}, \\
# w_{uf} \text{ if } \theta = h_{if}, \\
# -w_{uf} \text{ if } \theta = h_{jf}, \\
# 0 \text{ else }
# \end{cases} $$
#
# Which basically means: user $u$ prefer $i$ over $j$, let's do the following:
# - Increase the relevance (according to $u$) of features belonging to $i$ but not to $j$ and vice-versa
# - Increase the relevance of features assigned to $i$
# - Decrease the relevance of features assigned to $j$
#
# We're now ready to look at some code!
# # Let's implement SLIM BPR
import time
import numpy as np
# ### What do we need for a SLIM BPR?
#
# * Item-Item similarity matrix
# * Computing prediction
# * Update rule
# * Training loop and some patience
#
# +
from Notebooks_utils.data_splitter import train_test_holdout
from Data_manager.Movielens.Movielens10MReader import Movielens10MReader
data_reader = Movielens10MReader()
data_loaded = data_reader.load_data()
URM_all = data_loaded.get_URM_all()
URM_train, URM_test = train_test_holdout(URM_all, train_perc = 0.8)
# -
# ## Step 1: We create a dense similarity matrix, initialized as zero
n_users, n_items = URM_train.shape
item_item_S = np.zeros((n_items, n_items), dtype = np.float)
item_item_S
# ## Step 2: We sample a triplet
#
# #### Create a mask of positive interactions. How to build it depends on the data
# +
URM_mask = URM_train.copy()
URM_mask.data[URM_mask.data <= 3] = 0
URM_mask.eliminate_zeros()
URM_mask
# -
user_id = np.random.choice(n_users)
user_id
# ### Get user seen items and choose one
user_seen_items = URM_mask.indices[URM_mask.indptr[user_id]:URM_mask.indptr[user_id+1]]
user_seen_items
pos_item_id = np.random.choice(user_seen_items)
pos_item_id
# ### To select a negative item it's faster to just try again then to build a mapping of the non-seen items
# +
neg_item_selected = False
# It's faster to just try again then to build a mapping of the non-seen items
while (not neg_item_selected):
neg_item_id = np.random.randint(0, n_items)
if (neg_item_id not in user_seen_items):
neg_item_selected = True
neg_item_id
# -
# ## Step 2 - Computing prediction
#
# #### The prediction depends on the model: SLIM, Matrix Factorization...
# #### Note that here the data is implicit so we do not multiply for the user rating, because it is always 1, we just sum the similarities of the seen items.
# +
x_ui = item_item_S[pos_item_id, user_seen_items].sum()
x_uj = item_item_S[neg_item_id, user_seen_items].sum()
print("x_ui is {:.4f}, x_uj is {:.4f}".format(x_ui, x_uj))
# -
# ## Step 3 - Computing gradient
#
# #### The gradient depends on the objective function: RMSE, BPR...
x_uij = x_ui - x_uj
x_uij
# #### The original BPR paper uses the logarithm of the sigmoid of x_ij, whose derivative is the following
sigmoid_item = 1 / (1 + np.exp(x_uij))
sigmoid_item
# ## Step 4 - Update model
#
# #### How to update depends on the model itself, here we have just one paramether, the similarity matrix, so we perform just one update. In matrix factorization we have two.
#
# #### We need a learning rate, which influences how fast the model will change. Small ones lead to slower convergence but often higher results
# +
learning_rate = 1e-3
item_item_S[pos_item_id, user_seen_items] += learning_rate * sigmoid_item
item_item_S[pos_item_id, pos_item_id] = 0
item_item_S[neg_item_id, user_seen_items] -= learning_rate * sigmoid_item
item_item_S[neg_item_id, neg_item_id] = 0
# -
# #### Usually there is no relevant change in the scores over a single iteration
# +
x_i = item_item_S[pos_item_id, user_seen_items].sum()
x_j = item_item_S[neg_item_id, user_seen_items].sum()
print("x_i is {:.4f}, x_j is {:.4f}".format(x_i, x_j))
# -
# ## Now we put everything in a training loop
# +
def sample_triplet():
non_empty_user = False
while not non_empty_user:
user_id = np.random.choice(n_users)
user_seen_items = URM_mask.indices[URM_mask.indptr[user_id]:URM_mask.indptr[user_id+1]]
if len(user_seen_items)>0:
non_empty_user = True
pos_item_id = np.random.choice(user_seen_items)
neg_item_selected = False
# It's faster to just try again then to build a mapping of the non-seen items
while (not neg_item_selected):
neg_item_id = np.random.randint(0, n_items)
if (neg_item_id not in user_seen_items):
neg_item_selected = True
return user_id, pos_item_id, neg_item_id
# -
def train_one_epoch(item_item_S, learning_rate):
start_time = time.time()
for sample_num in range(n_users):
# Sample triplet
user_id, pos_item_id, neg_item_id = sample_triplet()
user_seen_items = URM_mask.indices[URM_mask.indptr[user_id]:URM_mask.indptr[user_id+1]]
# Prediction
x_ui = item_item_S[pos_item_id, user_seen_items].sum()
x_uj = item_item_S[neg_item_id, user_seen_items].sum()
# Gradient
x_uij = x_ui - x_uj
sigmoid_item = 1 / (1 + np.exp(x_uij))
# Update
item_item_S[pos_item_id, user_seen_items] += learning_rate * sigmoid_item
item_item_S[pos_item_id, pos_item_id] = 0
item_item_S[neg_item_id, user_seen_items] -= learning_rate * sigmoid_item
item_item_S[neg_item_id, neg_item_id] = 0
# Print some stats
if (sample_num +1)% 50000 == 0 or (sample_num +1) == n_users:
elapsed_time = time.time() - start_time
samples_per_second = (sample_num +1)/elapsed_time
print("Iteration {} in {:.2f} seconds. Samples per second {:.2f}".format(sample_num+1, elapsed_time, samples_per_second))
return item_item_S, samples_per_second
# +
learning_rate = 1e-6
item_item_S = np.zeros((n_items, n_items), dtype = np.float)
for n_epoch in range(5):
item_item_S, samples_per_second = train_one_epoch(item_item_S, learning_rate)
# -
estimated_seconds = 8e6 * 10 / samples_per_second
print("Estimated time with the previous training speed is {:.2f} seconds, or {:.2f} minutes".format(estimated_seconds, estimated_seconds/60))
# ### Common mistakes in using ML (based on last year's presentations)
#
# * Use default parameters and then give up when results are not good
# * Train for just 1 or 2 epochs
# * Use huge learning rate or regularization parameters: 1, 50, 100
# # BPR for MF
# ### What do we need for BPRMF?
#
# * User factor and Item factor matrices
# * Computing prediction
# * Update rule
# * Training loop and some patience
#
# ## Step 1: We create the dense latent factor matrices
# +
num_factors = 10
user_factors = np.random.random((n_users, num_factors))
item_factors = np.random.random((n_items, num_factors))
# -
user_factors
item_factors
# ## Step 2 - Computing prediction
user_id, pos_item_id, neg_item_id = sample_triplet()
(user_id, pos_item_id, neg_item_id)
# +
x_ui = np.dot(user_factors[user_id,:], item_factors[pos_item_id,:])
x_uj = np.dot(user_factors[user_id,:], item_factors[neg_item_id,:])
print("x_ui is {:.4f}, x_uj is {:.4f}".format(x_ui, x_uj))
# -
# ## Step 3 - Computing gradient
#
x_uij = x_ui - x_uj
x_uij
sigmoid_item = 1 / (1 + np.exp(x_uij))
sigmoid_item
# ## Step 4 - Update model
# +
regularization = 1e-4
learning_rate = 1e-2
H_i = item_factors[pos_item_id,:]
H_j = item_factors[neg_item_id,:]
W_u = user_factors[user_id,:]
user_factors[user_id,:] += learning_rate * (sigmoid_item * ( H_i - H_j ) - regularization * W_u)
item_factors[pos_item_id,:] += learning_rate * (sigmoid_item * ( W_u ) - regularization * H_i)
item_factors[neg_item_id,:] += learning_rate * (sigmoid_item * (-W_u ) - regularization * H_j)
# +
x_ui = np.dot(user_factors[user_id,:], item_factors[pos_item_id,:])
x_uj = np.dot(user_factors[user_id,:], item_factors[neg_item_id,:])
print("x_i is {:.4f}, x_j is {:.4f}".format(x_ui, x_uj))
# -
x_uij = x_ui - x_uj
x_uij
def train_one_epoch(user_factors, item_factors, learning_rate):
start_time = time.time()
for sample_num in range(n_users):
# Sample triplet
user_id, pos_item_id, neg_item_id = sample_triplet()
# Prediction
x_ui = np.dot(user_factors[user_id,:], item_factors[pos_item_id,:])
x_uj = np.dot(user_factors[user_id,:], item_factors[neg_item_id,:])
# Gradient
x_uij = x_ui - x_uj
sigmoid_item = 1 / (1 + np.exp(x_uij))
H_i = item_factors[pos_item_id,:]
H_j = item_factors[neg_item_id,:]
W_u = user_factors[user_id,:]
user_factors[user_id,:] += learning_rate * (sigmoid_item * ( H_i - H_j ) - regularization * W_u)
item_factors[pos_item_id,:] += learning_rate * (sigmoid_item * ( W_u ) - regularization * H_i)
item_factors[neg_item_id,:] += learning_rate * (sigmoid_item * (-W_u ) - regularization * H_j)
# Print some stats
if (sample_num +1)% 50000 == 0 or (sample_num +1) == n_users:
elapsed_time = time.time() - start_time
samples_per_second = (sample_num +1)/elapsed_time
print("Iteration {} in {:.2f} seconds. Samples per second {:.2f}".format(sample_num+1, elapsed_time, samples_per_second))
return user_factors, item_factors, samples_per_second
# +
learning_rate = 1e-6
num_factors = 10
user_factors = np.random.random((n_users, num_factors))
item_factors = np.random.random((n_items, num_factors))
for n_epoch in range(5):
user_factors, item_factors, samples_per_second = train_one_epoch(user_factors, item_factors, learning_rate)
# -
| Practice 09 - BPR for SLIM and MF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# default_exp models.maskrcnn
# -
# # MaskRCNN
#
# > MaskRCNN for instance segmentation.
#
# [MaskRCNN paper](https://arxiv.org/abs/1703.06870) - [Implementation](https://github.com/pytorch/vision/blob/master/torchvision/models/detection/mask_rcnn.py)
# +
#hide
# %load_ext autoreload
# %autoreload 2
from nbdev.showdoc import *
import warnings
warnings.filterwarnings("ignore")
# +
#export
from torchvision.models.detection.backbone_utils import resnet_fpn_backbone
from torchvision.models.utils import load_state_dict_from_url
from torchvision.models.detection.anchor_utils import AnchorGenerator
from torchvision.models.detection import MaskRCNN
from torchvision.ops.misc import FrozenBatchNorm2d
from functools import partial
from fastai.vision.all import delegates
# +
#export
#hide
_model_urls = {
'maskrcnn_resnet50_fpn_coco':
'https://download.pytorch.org/models/maskrcnn_resnet50_fpn_coco-bf2d0c1e.pth',
}
# -
# ## Function to create the model
# +
# export
@delegates(MaskRCNN)
def get_maskrcnn_model(arch_str, num_classes, pretrained=False, pretrained_backbone=True,
trainable_layers=5, **kwargs):
#if pretrained: pretrained_backbone = False
backbone = resnet_fpn_backbone(arch_str, pretrained=pretrained_backbone, trainable_layers=trainable_layers)
model = MaskRCNN(backbone,
num_classes,
image_mean = [0.0, 0.0, 0.0], # already normalized by fastai
image_std = [1.0, 1.0, 1.0],
**kwargs)
if pretrained:
try:
pretrained_dict = load_state_dict_from_url(_model_urls['maskrcnn_'+arch_str+'_fpn_coco'],
progress=True)
model_dict = model.state_dict()
pretrained_dict = {k: v for k, v in pretrained_dict.items() if
(k in model_dict) and (model_dict[k].shape == pretrained_dict[k].shape)}
model_dict.update(pretrained_dict)
model.load_state_dict(model_dict)
for module in model.modules():
if isinstance(module, FrozenBatchNorm2d):
module.eps = 0.0
except Exception as e:
#print(e)
print("No pretrained coco model found for maskrcnn_"+arch_str)
print("This does not affect the backbone.")
return model
# +
#export
#hide
maskrcnn_resnet18 = partial(get_maskrcnn_model, arch_str="resnet18")
maskrcnn_resnet34 = partial(get_maskrcnn_model, arch_str="resnet34")
maskrcnn_resnet50 = partial(get_maskrcnn_model, arch_str="resnet50")
maskrcnn_resnet101 = partial(get_maskrcnn_model, arch_str="resnet101")
maskrcnn_resnet152 = partial(get_maskrcnn_model, arch_str="resnet152")
# -
# ## Custom MaskRCNN
# To create a model, which you can pass to a `InstSegLearner` simply create a `partial` with the function `get_maskrcnn_model` (resnet backbone).
# +
from functools import partial
custom_maskrcnn = partial(get_maskrcnn_model, arch_str="resnet18",
pretrained=True, pretrained_backbone=True,
min_size=600, max_size=600)
# -
# When building the `Learner`, the number of classes are getting passed to this partial function and the model is ready for training.
custom_maskrcnn(num_classes=5)
# ## Prebuilt models
# There are some prebuilt model partials, which you can use instantly:
#
# * `maskrcnn_resnet18`
# * `maskrcnn_resnet34`
# * `maskrcnn_resnet50`
# * `maskrcnn_resnet101`
# * `maskrcnn_resnet152`
#
| notebooks/04b_models.maskrcnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import datetime
import numpy as np
import random
import time
import string
# # Functions to create random fields
# ## Random dates
# +
#Generate random date
def str_time_prop(start, end, format, prop):
"""Get a time at a proportion of a range of two formatted times.
start and end should be strings specifying times formated in the
given format (strftime-style), giving an interval [start, end].
prop specifies how a proportion of the interval to be taken after
start. The returned time will be in the specified format.
"""
stime = time.mktime(time.strptime(start, format))
etime = time.mktime(time.strptime(end, format))
ptime = stime + prop * (etime - stime)
return time.strftime(format, time.localtime(ptime))
def random_date(start, end, prop):
return str_time_prop(start, end, '%d/%m/%Y %I:%M %p', prop)
print(random_date("8/4/2020 12:00 PM", "31/05/2020 11:59 PM", np.random.power(1)))
# -
[random_date("8/4/2020 12:00 PM", "31/05/2020 11:59 PM", np.random.power(1)) for _ in range(3)]
# ## Random post codes
random_postcodes = ["SW1 3RN", "SW1V 4DG", "SE1 8XX", "B7 4NG", "EC1A 1BB", "W1A 0AX", "M1 1AE", "B33 8TH", "CR2 6XH", "DN55 1PT", "X", np.nan]
np.random.choice(random_postcodes, 5)
# ## First time agression
first_time = ["yes", "no"]
np.random.choice(first_time, 5)
# ## Agressor
agressor = ["partner", "father", "brother", "father-in-law"]
np.random.choice(agressor, 5)
# ## Display report or not
np.random.binomial(1, 0.1, 20)
# ## Random string
cupcake_paragraphs = pd.read_csv("https://raw.githubusercontent.com/rjcnrd/domestic_violence_covid-19/master/data/dummy-testimonials.csv").testimonials.tolist()
cupcake_paragraphs_big = (cupcake_paragraphs * int(np.ceil(1000/9)))
len((cupcake_paragraphs * int(np.ceil(1000/9)))[:1000])
# +
#Generate random string
def random_string_generator(size=6, chars=string.ascii_lowercase + string.digits):
return ''.join(random.choice(chars) for x in range(size))
random_string_generator()
# -
[random_string_generator(size = random.randint(0, 50)) for _ in range(3)]
# # Generate random data set
def generate_random_dataframe(i):
"""
i: number of rows
"""
df = pd.DataFrame({
"date_of_report" : [random_date("8/4/2020 12:00 PM", "31/05/2020 11:59 PM", np.random.power(2)) for _ in range(i)],
"postal_code": np.random.choice(random_postcodes, i),
"first_time_experience" : np.random.choice(first_time, i),
"aggressor" : np.random.choice(agressor, i),
"display_testimonial" : np.random.binomial(1, 0.1, i),
"written_report" : (cupcake_paragraphs * int(np.ceil(i/len(cupcake_paragraphs))))[:i]}
)
return df
data = generate_random_dataframe(1000)
data
data.to_csv("data/dummy_data_new.csv")
| generate_dummy_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.linear_model import LinearRegression
from sklearn.datasets import load_boston
# Load the data from the boston house-prices dataset
boston_data = load_boston()
x = boston_data['data']
y = boston_data['target']
# Make and fit the linear regression model
# TODO: Fit the model and assign it to the model variable
model = LinearRegression()
model.fit(x, y)
# Make a prediction using the model
sample_house = [[2.29690000e-01, 0.00000000e+00, 1.05900000e+01, 0.00000000e+00, 4.89000000e-01,
6.32600000e+00, 5.25000000e+01, 4.35490000e+00, 4.00000000e+00, 2.77000000e+02,
1.86000000e+01, 3.94870000e+02, 1.09700000e+01]]
# TODO: Predict housing price for the sample_house
prediction = model.predict(sample_house)
prediction
| Supervised_Learning/2. Linear_Regression/quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
>>> data = [[0, 0], [0, 0], [1, 1], [1, 1]]
>>> from sklearn.preprocessing import StandardScaler
>>> scaler = StandardScaler()
>>> print(scaler.fit(data))
>>> print(scaler.mean_)
>>> print(scaler.transform(data))
>>> print(scaler.transform([[2, 2]]))
| preprocessing_standardscaler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Which multiplication table do you want to learn?
#
# In this example notebook we demonstrate how Voilà can render different Jupyter widgets using [GridspecLayout](https://ipywidgets.readthedocs.io/en/latest/examples/Layout%20Templates.html#Grid-layout)
# +
from ipywidgets import GridspecLayout, Button, BoundedIntText, Valid, Layout, Dropdown
def create_expanded_button(description, button_style):
return Button(description=description, button_style=button_style, layout=Layout(height='auto', width='auto'))
rows = 11
columns = 6
gs = GridspecLayout(rows, columns)
def on_result_change(change):
row = int(change["owner"].layout.grid_row)
gs[row, 5].value = gs[0, 0].value * row == change["new"]
def on_multipler_change(change):
for i in range(1, rows):
gs[i, 0].description = str(change["new"])
gs[i, 4].max = change["new"] * 10
gs[i, 4].value = 1
gs[i, 4].step = change["new"]
gs[i, 5].value = False
gs[0, 0] = Dropdown(
options=[1, 2, 3, 4, 5, 6, 7, 8, 9, 10],
value=2,
)
gs[0, 0].observe(on_multipler_change, names="value")
multiplier = gs[0, 0].value
for i in range(1, rows):
gs[i, 0] = create_expanded_button(str(multiplier), "")
gs[i, 1] = create_expanded_button("*", "")
gs[i, 2] = create_expanded_button(str(i), "info")
gs[i, 3] = create_expanded_button("=", "")
gs[i, 4] = BoundedIntText(
min=0,
max=multiplier * 10,
layout=Layout(grid_row=str(i)),
value=1,
step=multiplier,
disabled=False
)
gs[i, 5] = Valid(
value=False,
description='Valid!',
)
gs[i, 4].observe(on_result_change, names='value')
gs
| notebooks/gridspecLayout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numerical Solution of the Schrödinger Equation Using a Finite Difference, Finite Element and Neural Network Approach
#
# Author: <NAME>
#
# Course: Modern Physics [FEMT08](https://nastava.fesb.unist.hr/nastava/predmeti/11624) taught by professor <NAME>
#
# Date: 2020, 26 Mar
# ## Content
#
# * [Content](#Content)
#
# * [Theory behind...](#Theory-behind...)
# * [1 Introduction to Quantum Mechanics](#1-Introduction-to-Quantum-Mechanics)
#
# * [2 Schrödinger Equation](#2-Schrödinger-Equation)
#
# * [3 The Use of Wave Equation](#3-The-Use-of-Wave-Equation)
#
# * [Numerical solution of the Schrödinger equation](#Numerical-solution-of-the-Schrödinger-equation)
# * [1 Finite difference method](#1-Finite-difference-method)
#
# * [2 Finite element method](#2-Finite-element-method)
#
# * [3 Artifical neural network method](#3-Artifical-neural-network-method)
# * [Results, discussion and conclusion](#Results,-discussion-and-conclussion)
# ## Theory behind...
# ### 1 Introduction to Quantum Mechanics
# One of the greatest modern-science successes surely has to be credited to sir **<NAME>** for his 1687. work titled *Philosophiæ Naturalis Principia Mathematica*. Immense knowledge of physics and mechanics accumulated over the centuries was beautifully summed up in three Newton laws. Those laws served as the foundation to many discoveries that will occur for few centuries to come.
# Also, great contribution in electricity and magnetism came from **Oersted**, **Faraday**, **Ampere**, **Henry** and **Ohm**. But not until **<NAME>** created theoretical synthesis of electricity and magnetism in his 1867. work titled *Treatise on Electricity and Magnetism*, the electromagnetism is understood as an indivisible phenomenon.
# The progress in 19th century was so huge that, to some, it seemed impossible there is anything fundamentally new to discover.
#
# However, the idea of (electromagnetic) waves propagating through vacuum was a mistery; also the black body radiation and the photoelectric effect were imposible to explain using the classical mechanics and Maxwell's classical electromagnetism.
#
# **<NAME>** had made the great first step into the creation of quantum hypothesis in 1900. Planck introduced the idea of discretized quantum oscilators that are able to radiate energy at certain discrete levels in order to explain black body radiation. Radiated energy was described using the frequency at which the source oscilates, multiplied by the extremely small constant, later known as the Planck constant $h$, mathematically formulated in the following expression:
#
# $$
# \begin{equation}
# B(\nu, T) = \frac{ 2 h \nu^3}{c^2} \frac{1}{e^\frac{h\nu}{k_B T} - 1}
# \end{equation}
# $$
#
# where $\nu$ is the frequency at which the source oscilates, T is the temperature, $k_B$ is the Boltzmann constant and $h$ is the Plank constant. $B(\nu, T)$ is the intensity of black body radiation, which can also be expressed using wavelength, $\lambda$, instead of frequency, $\nu$:
#
# $$
# \begin{equation}
# B(\lambda, T) =\frac{2hc^2}{\lambda^5}\frac{1}{ e^{\frac{hc}{\lambda k_B T}} - 1}
# \end{equation}
# $$
#
# Code for visual comparisson between the Planck's interpretation and the classical Rayleigh-Jeans interpretation of black body radiation is in the `black_body_radiation.py` script.
# %matplotlib inline
# %run src/scripts/black_body_radiation.py
# Young physicist **<NAME>** in 1905 using the Planck's hypothesis was able to explain the photoelectric effect.
# With 1913 **Bohr** atom model it was obvious that the quantum hypothesis reflects low level law of the nature and the modern physics was conceived.
#
# Even though the quantum hypothesis was able to explain some of the problems classical mechanics and physics in general had failed to explain, a lot of mysteries were yet to be unraveled. The wave-particle duality of nature by **<NAME>** captured through the expression:
#
# $$
# \begin{equation}
# \lambda = \frac{h}{p}
# \end{equation}
# $$
#
# where $\lambda$ is the wavelength, $h$ is the Planck constant and $p$ is the momentum of a particle, was the first step into a more general but also more complex way to explain the quantum hypothesis.
#
# The rather unusual De Broigle's hypothesis, presented in his 1924 PhD thesis, was soon after experimentally proved by **Davisson** and **Germer**. The conducted double-slit experiment is a demonstration that light and electrons can display characteristics of both classically defined waves and particles.
# use `inline` for inline plot or `notebook` for interactive plot w/ sliders
# %matplotlib inline
# %run src/scripts/double_slit_exp.py
# a - single slit width
# λ - wavelength
# L - screen distance
# d - distance between slits
# In 1926, assuming the wave-particle duality of nature, **<NAME>** formulates *the wave equation*. The wave equation describes the strenth of the field produced by the particle wave and has its solution as a function of both time and position. This particle wave is also known as the matter wave and is in the perfect accordance to the Davisson-Germer experiment.
# In the experiment, if the electrons were generated one by one, at first, the locations would seem to hit the sensor screen randomly. However, if the experiment is allowed to run for a very long time, a pattern shown in the previous figure will emerge. It is impossible to predict where each individual electron would strike the sensor screen, but using the Schrödinger equation, one can predict the probability of an electron striking a certain point of the sensor screen. This probability is proportional to the square of the magnitude of the wave function for the observed electron at that point, and is known as the Born rule, which will be described in more details in the following section. It was formulated by German physicist **<NAME>**, soon after Schrödinger formulated his equation.
#
# Davisson-Germer experiment also showed that if the observer wants to locate the position of an electron at every point, the sensor screen will not detect wave characteristics of a generated electron, rather the pattern of sensed electrons on the screen will be as if the elelctron has particle characteristics only. This occurance is called the wave function collapse and, more generally, it occurs when a wave function, naturally in a superposition of several eigenstates reduces to a single eigenstate due to the interaction with the external world. This interaction is called an *observation*. The problem of the wave function collapse is called the *measurement problem*. The inability to observe such a collapse directly has given rise to different interpretations of quantum mechanics and poses a key set of questions that each interpretation must answer. The oldest and most widely accepted interpretation is already mentioned Born rule, statistical interpretation which gave rise to the Copenhagen interpretation. Controversial Copenhagen philospohy, whose founding fathers are <NAME> and **<NAME>**, states that physical systems generally do not have definite properties prior to being measured, and quantum mechanics can only predict the probability distribution of a given measurement's possible results. Another opposing philosophy, supported by Einstein and Schrödinger, advocated the idea of hidden states and incomplete realization of quantum mechanics. From this, famous Schrödinger cat thought experiment emerged, illustrating the incompleteness of the Copenhagen interpretation. Great explanation of the Schrödinger cat thought experiment and intro to many-world interpretation by <NAME> is given [here](https://www.youtube.com/watch?v=kTXTPe3wahc).
#
# To take a few steps back, this study is based on, however flawed, still very powerful Copenhagen interpretation, without which most of the technology of the second half of the 20th century would not have been successfully realized. This probabilistic philosophy is based on the Heisenberg uncertainty principle. The uncertainty, in its most basic form, states that product of the position uncertainty and the momentum uncertainty of the observed electron at the same time is greater or equal to the reduced Planck constant $h$:
#
# $$
# \begin{equation}
# \Delta p \cdot \Delta x \geqslant \hbar
# \end{equation}
# $$
#
# where $\hbar=\frac{h}{2\pi}$.
#
# In order to determine precise position of the electron, one has to observe the electron using some electromagnetic-based detection mechanism. A photon with a low wavelength $\lambda_p$, emitted out of the detection mechanism, carries large momentum $p_p$ and the collision with an electron will deliver significant kinetic energy to an electron. Thus, an electron position at the certain time-point will be known with high probability but the initial momentum of an electron is changed and there is no way to know the exact value of the momentum.
# The principle is also applicable to all canonically conjugate variables (two variables, once multiplied, give the product of the physical dimension of the reduced Planck constant, e.q. time and energy):
#
# $$
# \begin{equation}
# \Delta E \cdot \Delta t \geqslant \hbar
# \end{equation}
# $$
#
# The uncertainty principle is an inherent property of the nature and comes from the fact that the matter acts both as waves and as particles. The uncertainty principle does not depend on the precision or tuning of a measuring device or a system.
# ### 2 Schrödinger Equation
# Assuming low radiation frequencies, where corpuscular nature of light is hardly detected, Maxwell's equations, in their differential form, are defined as follows:
#
# $$
# \begin{align}
# \nabla \times \mathbf{E} &= - \frac{\partial \mathbf{B}}{\partial t} \\
# \nabla \times \mathbf{B} &= \mu_0 ( \mathbf{J} + \epsilon_0\frac{\partial \mathbf{E}}{\partial t} ) \\
# \nabla \cdot \mathbf{E} &= \frac{\rho}{\epsilon_0} \\
# \nabla \cdot \mathbf{B} &= 0
# \end{align}
# $$
#
# where:
# * $\mathbf{E}$ is electric vector field;
# * $\mathbf{B}$ is magnetic pseudovector field;
# * $\mathbf{J}$ is electric current density;
# * $\epsilon_0$ is the permittivity of free space and
# * $\mu_0$ is the permeability of free space.
#
# The electromagnetic wave radiation velocity is defined with the following expression:
#
# $$
# \begin{equation}
# c = \lambda \cdot \nu
# \end{equation}
# $$
#
# where $\lambda$ is the wavelenght and $\nu$ is the frequency.
# Equation (10) can be rewritten as:
#
# $$
# \begin{equation}
# \frac{\nu^2}{c^2} - \frac{1}{\nu^2} = 0
# \end{equation}
# $$
#
# for later mathematical convenience.
#
# The value of the electric field of the plane electromagnetic wave expanding in $x$ direction is defined as follows:
#
# $$
# \begin{equation}
# E = E_0 \sin 2\pi \big(\nu t - \frac{x}{\lambda}\big)
# \end{equation}
# $$
#
# with the associated second partial derivatives:
#
# $$
# \begin{equation}
# \frac{\partial^2 E}{\partial t^2} = -4\pi^2 \nu^2E; \qquad \frac{\partial^2 E}{\partial x^2} = -\frac{4\pi^2}{\lambda^2} E
# \end{equation}
# $$
#
# Equation (11), once multiplied by $4\pi^2\mathbf{E}$ and rearanged considering the form of second partial derivatives defined in (13) generates the following expression:
#
# $$
# \begin{equation}
# \frac{\partial^2 \mathbf{E}}{\partial x^2} - \frac{1}{c^2}\frac{\partial^2 \mathbf{E}}{\partial t^2} = 0
# \end{equation}
# $$
#
# For a free-particle, i.e. an electron, moving rectilinearly, the matter wave takes the form of the plane wave. The relativistic relation between the energy and the momentum of a free-particle is defiend using the expression:
#
# $$
# \begin{equation}
# \frac{E^2}{c^2} = p^2 + m_0^2 c^2
# \end{equation}
# $$
#
# where $E$ is the relativistic energy of a free-particle, $p$ is the relativistic momentum and $m_o$ is the rest mass.
#
# Assuming the wave-particle duality for a free-particle where $E=h\nu$ and $p=\frac{h}{\nu}$, the relativistic relation between the energy and the momentum of a free-particle becomes:
#
# $$
# \begin{equation}
# \frac{h^2 \nu^2}{c^2} = \frac{h^2}{\lambda^2} + \frac{h^2 \nu_0^2}{c^2} \Rightarrow \frac{\nu^2}{c^2} = \frac{1}{\lambda^2} + \frac{\nu_0^2}{c^2}
# \end{equation}
# $$
#
# If we assume that the rest mass is 0, (16) becomes (11).
#
# The matter wave of an observed free particle is defined as:
#
# $$
# \begin{equation}
# \Phi = A \cdot e^{-2\pi i(\nu t - x/\lambda)}
# \end{equation}
# $$
#
# with the associated second partial derivatives:
#
# $$
# \begin{equation}
# \frac{\partial^2 \Phi}{\partial t^2} = -4\pi^2 \nu^2\Phi; \qquad \frac{\partial^2 \Phi}{\partial x^2} = -\frac{4\pi^2}{\lambda^2} \Phi
# \end{equation}
# $$
#
# Equation (16), once multiplied by $\Phi$ and rearanged considering the form of second partial derivatives defined in (18) generates the following expression:
#
# $$
# \begin{equation}
# \frac{\partial^2 \mathbf{\Phi}}{\partial x^2} - \frac{1}{c^2}\frac{\partial^2 \mathbf{\Phi}}{\partial t^2} = \frac{4\pi^2\mu_0^2}{c^2}\mathbf{\Phi}
# \end{equation}
# $$
#
# or, generalized in the 3-D space:
#
# $$
# \begin{equation}
# \frac{\partial^2 \mathbf{\Phi}}{\partial x^2} + \frac{\partial^2 \mathbf{\Phi}}{\partial y^2} + \frac{\partial^2 \mathbf{\Phi}}{\partial z^2} - \frac{1}{c^2}\frac{\partial^2 \mathbf{\Phi}}{\partial t^2} = \frac{4\pi^2\mu_0^2}{c^2}\mathbf{\Phi}
# \end{equation}
# $$
#
# $$\Downarrow$$
#
# $$
# \begin{equation}
# \Delta\mathbf{\Phi} - \frac{1}{c^2}\frac{\partial^2 \mathbf{\Phi}}{\partial t^2} = \frac{4\pi^2\mu_0^2}{c^2}\mathbf{\Phi}
# \end{equation}
# $$
#
# thus forming the relativistic version of the matter wave, in the literature often referenced as the **Klein-Gordon** equation.
#
# In practice, the difference between $\nu$ and $\nu_0$ is negligible and the non-relativistic version of the Klein-Gordon equation is of greater importance. From here, the famous time-dependant Schrödinger equation for a free-particle arises and is written as:
#
# $$
# \begin{equation}
# -\frac{\hbar^2}{2m_0} \Delta \mathbf{\Phi} = i \hbar \frac{\partial \mathbf{\Phi}}{\partial t}
# \end{equation}
# $$
#
# where $m_0$ is the rest mass of a free particle.
#
# Separating the spatial, $\psi(x, y, z)$, and temporal variables of the function $\Phi$, time-independent Schrödinger equation is written as:
#
# $$
# \begin{equation}
# -\frac{\hbar^2}{2m} \Delta \psi = E \psi
# \end{equation}
# $$
#
# where $E$ is the non-relativistic free-particle energy and the mass of a free particle is the same as the rest mass of a free particle. The non-relativistic energy is defined as:
#
# $$
# \begin{equation}
# E = \frac{p^2}{2m} + U
# \end{equation}
# $$
#
# where $U$ is the potential energy. Equation (24) must always be satisfied in order to obey the conservation of energy law.
#
# A solution of Schrödinger equation is a wave function, $\psi$, a quantity that described the displacement of the matter wave produced by particles, i.e. electrons. In Copenhagen interpretation, as mentioned in previous section, the square of the absolute value of the wave function results in the probability of finding the particle at given location.
# #### 1-D time-independent Schrödinger equation
# 1-D time-independent Schrödinger equation for a single non-relativistic particle is obtained from the time-dependant equation by the separation of variables and is written using the following expression:
#
# $$
# \begin{equation}
# \Big(-\frac{\hbar^2}{2m} \cdot \frac{d^2 \psi(x)}{dx^2} + U(x)\Big) \psi(x) = E \psi(x)
# \end{equation}
# $$
#
# where, in the language of linear algebra, this equation is the eigenvalue equation. The wave function is an eigenfunction, $\psi(x)$ of the Hamiltonian operator, $\hat H$, with corresponding eigenvalue $E$:
#
# $$
# \begin{equation}
# \hat H \psi(x) = \Big(-\frac{\hbar^2}{2m} \cdot \frac{d^2}{dx^2} + U(x)\Big) \psi(x)
# \end{equation}
# $$
#
# The form of the Hamiltonian operator comes from classical mechanics, where the Hamiltonian function is the sum of the kinetic and potential energies.
#
# The probability density function of a particle over the x-axis is written using the following expression:
#
# $$
# \begin{equation}
# \mbox{pdf} = \lvert \psi(x) \lvert^2
# \end{equation}
# $$
#
# and it can be shown, that the probability density function in space does not depend on time. From time-dependent Schrödinger equation (22), by letting the $\Phi(x,t) = \psi(x) \cdot \Phi(t)$ and applying the separation of variables, we obtain the following expression for temporal variable:
#
# $$
# \begin{equation}
# i\hbar \cdot \frac{1}{\Phi(t)} \cdot \frac{\partial \Phi(t)}{\partial t} = E
# \end{equation}
# $$
#
# The solution of (28) is obtained by integrating both sides:
#
# $$
# \begin{equation}
# \Phi(t) = e^{-\frac{iEt}{\hbar}}
# \end{equation}
# $$
#
# Finally the 1-D time-dependant wave function is written as:
#
# $$
# \begin{equation}
# \Phi(x,t) = \psi(x) e^{-\frac{iEt}{\hbar}}
# \end{equation}
# $$
#
# Following the Copenhagen interpretation, the square of the 1-D time-dependant wave function resolves the probability density of a single particle to be found at the certain location on the $x$-axis at a specific time $t$. This results in the following:
#
# $$
# \begin{equation}
# \lvert \Phi(x,t) \lvert^2 = \lvert \psi(x) \lvert^2 \cdot \lvert e^{-\frac{iEt}{\hbar}} \lvert^2 = \lvert \psi(x) \lvert^2
# \end{equation}
# $$
#
# From this, it is obvious that the probability density function in space does not depend on time, thus the rest of the seminar will be concentrated on the non-linear Schrödinger equation for non-relativitic particles.
#
# The probability of finding the observed particle anywhere in the domain $x \subseteq \Omega$ is 1 since the particle has to be somewhere. The actual position is not known, rather one can describe the uncertainty of finding the particle for position in $[x, x+\mbox{d}x]$. The sum of probabilities over the entire solution domain should be equal to one:
#
# $$
# \begin{equation}
# \int_{x \subseteq \Omega} \lvert \psi(x) \lvert^2 \mbox{d}x = 1
# \end{equation}
# $$
#
# or, more explicitly:
#
# $$
# \begin{equation}
# \int_{x \subseteq \Omega} \psi(x) \Psi^{*}(x) \mbox{d}x = 1
# \end{equation}
# $$
# #### Wave function constraints
# In order to represent a physically observable system in a measurable and meaningful fashion, the wave function must satisfy set of constraints:
#
# 1. the wave function must be a solution to Schrödinger equation;
# 2. the wave function and the first derivative of the wave function, $\frac{d \Psi(x)}{dx}$ must be continuous
# 3. the wave function must be normalizable - the 1-D wave function approaches to zero as $x$ approaches to infinity;
# ### 3 The Use of Wave Equation
# #### Electron in Infinite Potential Well
# Consider an electron of mass $m$ confined to 1-D rigid box of width L.
# The potential of the infinite potential well is defined as follows:
#
# $$
# \begin{align}
# U(x) &= 0, \qquad &0 < x < L\\
# U(x) &= +\infty, \qquad &x \leq 0, x \geq L
# \end{align}
# $$
#
# The electron can move along $x$-axis and the collisions with the walls are perfectly elastic. The described situation is known as an inifinitely deep square well potential.
# In order to determine the motion of the observed electron as it travels along the $x$-axis, we must determine the wave function using Schrödinger equation.
#
# Boundary conditions are pre-defined in (34) and (35). For $x=0$ and $x=L$, the potential energy is defined as $U(0) = U(L) = +\infty$, thus the wave function at boundaries is 0, $\psi(0) = \psi(L) = 0$, otherwise the product of $U(x) \cdot \psi(x)$ in Schrödinger equation would be phisically infeasible.
#
# Inside the potential well, the potential energy is equal to 0, which means that the Schrödinger equation becomes:
#
# $$
# \begin{equation}
# E \cdot \psi(x) = - \frac{\hbar^2}{2m} \cdot \frac{d^2 \psi(x)}{dx^2}
# \end{equation}
# $$
#
# and the wave function is written as:
#
# $$
# \begin{equation}
# \psi(x) = A \sin(kx) + B \cos(kx)
# \end{equation}
# $$
#
# In order to discover the unknown parameters $A$ and $B$, the boundary condtions have to be taken into consideration.
#
# If $x=0$,
#
# $\Psi(x=0) = 0 \qquad \Rightarrow \qquad \psi(0) = A\sin(0) + B\cos(0) = 0$,
#
# and
#
# $$
# \begin{equation}
# \boxed{B=0}
# \end{equation}
# $$
#
# ---
#
# Since $B=0$ the wave function takes the following form of $\psi(x) = A\sin(kx)$.
#
# ---
#
# If $x=L$,
#
# $\psi(x=L) = 0 \qquad \Rightarrow \qquad \psi(L) = A\sin(kL) = 0$.
#
# Since the electron is assumed to exists somewhere in the solution domain $ x \in (0, L)$, the term $\sin(kL)$ has to be equal to 0. This is true for $kL = 0, \pi, 2\pi...$ or, more generally, $kL=n\pi$ where $n=1,2,3,...$.
#
# From here, $k$ can be written as:
#
# $$
# \begin{equation}
# k = \frac{n\pi}{L}
# \end{equation}
# $$
#
# and, since the $k$ is known to be:
#
# $$
# \begin{equation}
# k = \sqrt{\frac{2mE}{\hbar^2}}
# \end{equation}
# $$
#
# Combining the expressions (39) and (40), the following expression occurs:
#
# $$
# \begin{equation}
# \frac{n\pi}{L} = \sqrt{\frac{2mE}{\hbar^2}}
# \end{equation}
# $$
#
# From (41), the non-relativistic energy of an electron is defined as:
#
# $$
# \begin{equation}
# E = \frac{n^{2}h^{2}}{8 m L^2}
# \end{equation}
# $$
#
# where, $n=1,2,3,...$ and $\hbar=h/2\pi$.
#
# Energy discretion is a direct consequence of boundary conditions and the initial assumption that the potential energy of an electron within the area is equal to zero.
#
# The positive integer $n$ is a principal quantum number. Quantum numbers describe values of conserved quantities in the dynamics of a quantum system, in this case the quantum system in an electron trapped inside of a infinite potential well. The quantum number $n$ represent principal quantum numbers, which give acceptable solutions to the Schrödinger equation. The standard model of a quantum system is described using four quantum numbers: principal quantum number $n$, azimuthal quantum number $l$, magentic quantum number $m_l$ and spin quantum number $m_s$. The first three quantum numbers are derived from Schrödinger equation, while the fourth comes from applying special relativity. The principal quantum number describes the electron shell - energy level of an electron, and it ranges from 1 to the shell containing the outermost electron of the specific atom, thus is of the greatest importance for the general solution of time-independent Schrödinger equation. Since the principal quantum number $n$ is natural number, the energy level of an electron is a discrete variable.
#
# The zero point energy represents the ground state of an electron or the least amount of energy that electron can contain, where $n=1$.
#
# A wave function has to obey [the set of constraints](#Wave-function-constraints) defined in the previous section:
#
# $$
# \begin{equation}
# \int_{-\infty}^{+\infty}\lvert \psi \lvert^2 dx = \int_{0}^{L} A^2 \sin^2 \big(\frac{n \pi}{L} \cdot x \big)dx = 1
# \end{equation}
# $$
#
# From (43):
#
# $$
# \begin{equation}
# \frac{A^2 L}{2} = 1 \qquad \Rightarrow \qquad \boxed{A=\sqrt{\frac{2}{L}}}
# \end{equation}
# $$
#
# ---
#
# Finally, the wave function describing the motion of the observed electron with potential energy $U(x)=0$ and with mass $m$ moving along the the $x$-axis trapped inside the infinite potential well, considering (38) and (44), is given using the following expression:
#
# $$
# \begin{equation}
# \psi_n(x) = \sqrt{\frac{2}{L}}\sin\big( \frac{n \pi}{L} \cdot x \big)
# \end{equation}
# $$
#
# where $n$ is the principal quantum number and $L$ is the width of the rigid box. The term $\sqrt{\frac{2}{L}}$ represents the amplitude of the wave function.
#
# In order to obtain the probability density of the electron in three different energy states for $n=1, 2 \mbox{ and } 3$, trapped inside the infinite potential well of width $L=1 \mbox{ nm}$, the square of the wave function has to be observed:
# %matplotlib inline
# %run src/scripts/infinite_potential_well.py
# The highest point of each probability density function plotted in previous figure represents the most likely placement of an electron at any given moment in time. Boundary conditions are defined *a priori* and are integrated in the analytical solution given in (45), thus the probability of an electron placement at locations $x=\{0, L\}$ is 0.
# It is also important to notice that the area under each of the probability density curves shown above, is equal to 1. In order to be measurable and physically meaningful, a wave function must be normalized, mathematically formulated in (43). The total probability of finding the electron inside the $x \in \langle 0, L \rangle$ region is obtained by integrating the probability density function over $x$ and is always equal to 1. If the $L$ increases, the probability density peak will decrease and the probability density function curve will, consequently, be flatten.
# #### Electron in Finite Potential Well
#
# Consider an electron of mass $m$ confined to 1-D finite potential well of width L and potential barriers at energy level of $U_0$.
# The potential of the finite potential well is defined as follows:
#
# $$
# \begin{align}
# U(x) &= 0, &\qquad \lvert x \lvert > L/2\\
# U(x) &= U_0, &\qquad \lvert x \lvert \leq L
# \end{align}
# $$
#
# In order to determine the motion of the observed electron as it travels along the $x$-axis, we must determine the wave function using Schrödinger equation for three different regions.
#
# ##### Region II: $x \in \langle -L/2, L/2 \rangle$
# If particle is in region II, then the $U(x)=0$ due to (46). The Schrödinger equation is defined as follows:
#
# $$
# \begin{equation}
# E \cdot \psi (x) = -\frac{\hbar^2}{2m} \cdot \frac{d^2 \psi(x)}{dx^2}
# \end{equation}
# $$
#
# From (48) the wave function is defined as:
#
# $$
# \begin{equation}
# \psi(x) = A \sin(kx) + B \sin(kx)
# \end{equation}
# $$
#
# where $k = \frac{n\pi}{L}$.
#
# ##### Region I and III: $x \in \langle-\infty, -L/2] \cup [L/2, +\infty \rangle$
# If particle is in region I or III, then the $U(x) = U_0$ due to (47). The Schrödinger equation is defined as follows:
#
# $$
# \begin{equation}
# E \cdot \psi(x) = U_0 \cdot \psi(x) -\frac{\hbar^2}{2m} \cdot \frac{d^2 \psi(x)}{dx^2}
# \end{equation}
# $$
#
# After rearanging (50), the Schrödinger equation becomes:
#
# $$
# \begin{equation}
# \frac{d^2 \psi(x)}{dx^2} - \Big ( \frac{2m}{\hbar}(E - U_0) \Big ) \psi(x) = 0
# \end{equation}
# $$
#
# If the term $\frac{2m}{\hbar}(E - U_0)$ is written as $\alpha^2$, the equation takes simplified form of:
#
# $$
# \begin{equation}
# \frac{d^2 \psi(x)}{dx^2} - \alpha^2 \psi(x) = 0
# \end{equation}
# $$
#
# and the solution of the equation is:
#
# $$
# \begin{equation}
# \psi(x) = C \mbox{e}^{\alpha x} + D \mbox{e}^{-\alpha x}
# \end{equation}
# $$
#
# For the region I, if $x \rightarrow -\infty$, the term $\mbox{e}^{-\alpha x} \rightarrow \infty$ and $\psi(x) \rightarrow \infty$, which is physically infeasible. In order to avoid this, the term $D$ is set to be 0 for region I. The wave function for region I is defined as:
#
# $$
# \begin{equation}
# \psi(x) = C \mbox{e}^{\alpha x}
# \end{equation}
# $$
#
# the wave function of the region III is defined analogously as:
#
# $$
# \begin{equation}
# \psi(x) = D \mbox{e}^{-\alpha x}
# \end{equation}
# $$
#
# The wave function must obey [the set of constraints](#Wave-function-constraints) defined in the previous section, that is, the wave function must be continuous throughout $x$-axis in order to be physically feasible. The total wave function composed of wave functions for three different regions defined in (49), (54) and (55) must satisfy the following conditions on boundaries to achieve the continuity:
#
# $$
# \begin{align}
# \psi_1(x) = \psi_2(x), \quad \frac{\psi_1(x)}{dx}=\frac{\psi_2(x)}{dx} \qquad &\mbox{for } x=-\frac{L}{2};\\
# \psi_2(x) = \psi_3(x), \quad \frac{\psi_2(x)}{dx}=\frac{\psi_3(x)}{dx} \qquad &\mbox{for } x=\frac{L}{2}.
# \end{align}
# $$
#
# where $\psi_1(x)$, $\psi_2(x)$ and $\psi_3(x)$ are wave functions for regions I, II and III, respectively.
#
# Even though, the analytical solution for previously described situation exists, it is not going to be solved in this notebook, rather the high level intuition of the problem will be presented. According to classical mechanics, if $U_0 > E$, an electron cannot be found in the regions I and III. By solving the Schrödinger through the framework of quantum mechanics, it is shown that there is some probability of an electron to penetrate the potential barriers and to be found outside of the region II, even if the potential energy $U_0$ is greater than the total energy of an electron. The possibility of positioning an electron outside the potential well makes the energy conservation law momentarily invalid. This, however, can be explained via the Heisenberg uncertainty principle defined in (5). The uncertainty principle enables an electron to exist outside of the potential well for the amount of time $\Delta t$ during which a certain amount of energy uncertainty also co-exists.
# %matplotlib inline
# %run -i src/scripts/finite_potential_well.py
# ##### Quantum Tunneling
#
# As shown in the previous section, an electron is able to penetrate potential barrier and enter a region, which is not explainable through the lense of classical mechanics. According to quantum mechanics, there is some non-zero probability that the particle will end up passing through a potential barrier even though the potential energy level of the barrier is higher than the total non-relativistic energy of an electron. The wave function of an electron, once it enters the potential region, will start to decay exponentially. Before the amplitude of the wave function drops to zero, the wave function resurfaces and continues to propagate with a smaller amplitude on the other side of the barrier. Since there is existing wave function on the other side of the barrier, one can reason that electron is able to penetrate the barrier even though its kinetic energy is less than the potential energy of a barrier. This phenomenon is known as quantum tunneling. From here, two important coefficients can be calculated:
#
# **Transmission coefficient** ($T$) - the fraction of the transmitted wave function of an electron
#
# $$
# \begin{equation}
# T \approx \mbox{e}^{-2\alpha L}
# \end{equation}
# $$
#
# where L is the width of a potential barrier and $\alpha$ is formulated as follows:
#
# $$
# \begin{equation}
# \alpha = \frac{\sqrt{2m(U_0 - E}}{\hbar}
# \end{equation}
# $$
#
# where $m$ is the mass of an electron, $U_0$ is the poteential energy level of a barrier, $E$ is the kinetic energy of an electron and $\hbar$ is the reduced Planck constant.
#
# **Reflection coefficient** ($R$) - fraction of the reflected wave function of an electron:
#
# $$
# \begin{equation}
# R = 1 - T
# \end{equation}
# $$
#
# The total sum of the transmission and reflection coefficient is always 1.
#
# The following cell is the output of a Python solver for the 1-D Schrödinger equation beautfully explained through [this blog post](http://jakevdp.github.io/blog/2012/09/05/quantum-python/). The code is minimally modified to satisfy the current version of Python and some dependencies. All rights go to [<NAME>](http://vanderplas.com/).
# use `inline` for inline plot or `notebook` for interactive plot
# %matplotlib notebook
# %run -i src/scripts/quantum_tunneling.py
# ## Numerical solution of the Schrödinger equation
# ### 1 Finite difference method
# The wavefunction, $\psi(x)$, over $x$-axis is defined as eigenvector of the Hamiltonian operator as follows:
#
# $$
# \begin{equation}
# \hat{H} \psi = E \psi
# \end{equation}
# $$
#
# where $\hat{H}$ is the Hamiltonian operator, E is eigenvalues (energies) of an electron with the wave function $\psi$. This problem should be solved as eigenproblem, otherwise the solution to $\psi$ will be trivial.
#
# Hamiltonian operator is then defiend as $N \times N$ matrix, where $N$ is the chosen number of grid points.
# %matplotlib inline
# %run src/fdm_infinite_potential_well.py -n 50
# ### 2 Finite element method
# Applying the weighted residual approach to Schrödinger equation (25) with $U(x)=0$ the following term is obtained:
#
# $$
# \begin{equation}
# -\int_{0}^{L} \frac{\hbar^2}{2m}\frac{d^2\psi}{dx^2}W_{j} = \int_{0}^{L} E \psi W_{j} dx
# \end{equation}
# $$
#
# where $W_{j}$ are test functions for $j=1, 2, 3, ..., n$, where $n$ stands for the number of approximated solutions. From here, weak formulation is utilized as follows:
#
# $$
# \begin{equation}
# \int_{0}^{L} \frac{d\psi}{dx}\frac{dW_{j}}{dx} dx = \frac{2m}{\hbar^2}E \int_{0}^{L} \psi W_{j} dx + \frac{2m}{\hbar^2} \frac{d\psi}{dx} W_j \Big \lvert_{0}^L
# \end{equation}
# $$
#
# The approximate solution is formed as the linear combination of coefficients $\alpha_i$ and shape functions $N_i$. In the Galerkin-Bubnov framework test functions are the same as shape functions, $W_j = N_j$ and the vectorized Schrödinger equation is defined as follows:
#
# $$
# \begin{equation}
# [A]\{\alpha\} = [B]\{\alpha\}[E]
# \end{equation}
# $$
#
# where:
# * A is the left-hand-side matrix, $n \times n$ shaped, defined as:
#
# $$
# \begin{equation}
# A_{ji} = \int_{x_1}^{x_2} \frac{dN_i(x)}{dx}\frac{dN_{j}}{dx} dx
# \end{equation}
# $$
#
# * B is the right-hand-side vector, $n \times n$ shaped, defined as:
#
# $$
# \begin{equation}
# B_{ji} = \frac{2m}{\hbar^2}E \int_{x_1}^{x_2} N_i(x) N_{j}(x) dx
# \end{equation}
# $$
#
# * E is a diagonal matrix representing eigenvalues for different princple quantum numbers, $1$ to $N$.
# %matplotlib inline
# %run src/fem_infinite_potential_well.py -n 49
# ### 3 Artifical neural network method
# [The technique proposed by Lagaris et al.](https://pubmed.ncbi.nlm.nih.gov/18255782/) is extended and adjusted to modern machine learning practices in order to acquire the wave function in the particle-in-a-box quantum model. Firstly, the wave function is approximated using the following expression:
#
# $$
# \begin{equation}
# \hat \psi(x) = B(x) NN(x; \theta)
# \end{equation}
# $$
#
# where $B(x)$ is any function that satisfies boundary conditions, in this case $B(x) = x(1-x)$, and $NN(x; \theta)$ is the neural network that takes positional coordinate $x$ as the input argument and is parameterized using the weights and biases, $\theta$. The collocation method is applied in order to discretize the solution domain, $\Omega$, into the set of points for training, $x_i \in \Omega$ for $i \in [1, N]$, where $N$ is the total number of collocation points. The problem is transformed to the optimization problem with the following loss function:
#
# $$
# \begin{equation}
# J(x, \theta) = \frac{\sum_i [H \hat \psi(x_i; \theta) - E \hat \psi(x_i; \theta)]^2}{\int_\Omega \lvert \hat \psi \rvert^2 dx} + \lvert \psi(0) - \hat \psi(0) \rvert^2 + \lvert \psi(L) - \hat \psi(L) \rvert^2
# \end{equation}
# $$
#
# where
#
# $$
# \begin{equation}
# E = \frac{\int_\Omega \hat \psi^* H \hat \psi dx}{\int_\Omega \lvert \hat \psi \rvert^2 dx}
# \end{equation}
# $$
#
# Just like in the original work, the neural network is the single hidden layer multi-layer perceptron (MLP) with 40 hidden units total. Every unit is activated using hyperbolic tangent, *tanh*, activation function. Instead of manually applying symbolic derivatives, the automatic differentiation procedure is applied. Once the derivative of the loss function with respect to the network parameters has been acquired, the minimization can be performed. The Broyden-Fletcher-Goldfarb-Shanno (BFGS) algorithm, an iterative method for solving unconstrained nonlinear optimization problems, is applied.
# ## Results, discussion and conclussion
# %matplotlib inline
# %run src/compare.py -n 10
# %matplotlib inline
# %run -i src/compare.py -n 50
# %matplotlib inline
# %run -i src/compare.py -n 100
| schrodinger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Recurrent Neural networks
# =====
# ### RNN
# <img src ="imgs/rnn.png" width="20%">
# A recurrent neural network (RNN) is a class of artificial neural network where connections between units form a directed cycle. This creates an internal state of the network which allows it to exhibit dynamic temporal behavior.
keras.layers.recurrent.SimpleRNN(output_dim,
init='glorot_uniform', inner_init='orthogonal', activation='tanh',
W_regularizer=None, U_regularizer=None, b_regularizer=None,
dropout_W=0.0, dropout_U=0.0)
# #### Backprop Through time
# Contrary to feed-forward neural networks, the RNN is characterized by the ability of encoding longer past information, thus very suitable for sequential models. The BPTT extends the ordinary BP algorithm to suit the recurrent neural
# architecture.
# <img src ="imgs/rnn2.png" width="45%">
# %matplotlib inline
# +
import numpy as np
import pandas as pd
import theano
import theano.tensor as T
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.preprocessing import image
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from keras.datasets import imdb
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution2D, MaxPooling2D
from keras.utils import np_utils
from keras.preprocessing import sequence
from keras.layers.embeddings import Embedding
from keras.layers.recurrent import LSTM, GRU, SimpleRNN
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.cross_validation import train_test_split
from keras.layers.core import Activation, TimeDistributedDense, RepeatVector
from keras.callbacks import EarlyStopping, ModelCheckpoint
# -
# #### IMDB sentiment classification task
# This is a dataset for binary sentiment classification containing substantially more data than previous benchmark datasets.
#
# IMDB provided a set of 25,000 highly polar movie reviews for training, and 25,000 for testing.
#
# There is additional unlabeled data for use as well. Raw text and already processed bag of words formats are provided.
#
# http://ai.stanford.edu/~amaas/data/sentiment/
# #### Data Preparation - IMDB
# +
max_features = 20000
maxlen = 100 # cut texts after this number of words (among top max_features most common words)
batch_size = 32
print("Loading data...")
(X_train, y_train), (X_test, y_test) = imdb.load_data(nb_words=max_features, test_split=0.2)
print(len(X_train), 'train sequences')
print(len(X_test), 'test sequences')
print('Example:')
print(X_train[:1])
print("Pad sequences (samples x time)")
X_train = sequence.pad_sequences(X_train, maxlen=maxlen)
X_test = sequence.pad_sequences(X_test, maxlen=maxlen)
print('X_train shape:', X_train.shape)
print('X_test shape:', X_test.shape)
# -
# #### Model building
# +
print('Build model...')
model = Sequential()
model.add(Embedding(max_features, 128, input_length=maxlen))
model.add(SimpleRNN(128))
model.add(Dropout(0.5))
model.add(Dense(1))
model.add(Activation('sigmoid'))
# try using different optimizers and different optimizer configs
model.compile(loss='binary_crossentropy', optimizer='adam', class_mode="binary")
print("Train...")
model.fit(X_train, y_train, batch_size=batch_size, nb_epoch=1, validation_data=(X_test, y_test), show_accuracy=True)
# -
# ### LSTM
# A LSTM network is an artificial neural network that contains LSTM blocks instead of, or in addition to, regular network units. A LSTM block may be described as a "smart" network unit that can remember a value for an arbitrary length of time.
#
# Unlike traditional RNNs, an Long short-term memory network is well-suited to learn from experience to classify, process and predict time series when there are very long time lags of unknown size between important events.
# <img src ="imgs/gru.png" width="60%">
keras.layers.recurrent.LSTM(output_dim, init='glorot_uniform', inner_init='orthogonal',
forget_bias_init='one', activation='tanh',
inner_activation='hard_sigmoid',
W_regularizer=None, U_regularizer=None, b_regularizer=None,
dropout_W=0.0, dropout_U=0.0)
# ### GRU
# Gated recurrent units are a gating mechanism in recurrent neural networks.
#
# Much similar to the LSTMs, they have fewer parameters than LSTM, as they lack an output gate.
keras.layers.recurrent.GRU(output_dim, init='glorot_uniform', inner_init='orthogonal',
activation='tanh', inner_activation='hard_sigmoid',
W_regularizer=None, U_regularizer=None, b_regularizer=None,
dropout_W=0.0, dropout_U=0.0)
# ### Your Turn! - Hands on Rnn
# +
print('Build model...')
model = Sequential()
model.add(Embedding(max_features, 128, input_length=maxlen))
# Play with those! try and get better results!
#model.add(SimpleRNN(128))
#model.add(GRU(128))
#model.add(LSTM(128))
model.add(Dropout(0.5))
model.add(Dense(1))
model.add(Activation('sigmoid'))
# try using different optimizers and different optimizer configs
model.compile(loss='binary_crossentropy', optimizer='adam', class_mode="binary")
print("Train...")
model.fit(X_train, y_train, batch_size=batch_size,
nb_epoch=4, validation_data=(X_test, y_test), show_accuracy=True)
score, acc = model.evaluate(X_test, y_test, batch_size=batch_size, show_accuracy=True)
print('Test score:', score)
print('Test accuracy:', acc)
# -
# # Sentence Generation using RNN(LSTM)
# +
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout
from keras.layers import LSTM
from keras.optimizers import RMSprop
from keras.utils.data_utils import get_file
import numpy as np
import random
import sys
path = get_file('nietzsche.txt', origin="https://s3.amazonaws.com/text-datasets/nietzsche.txt")
text = open(path).read().lower()
print('corpus length:', len(text))
chars = sorted(list(set(text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
# cut the text in semi-redundant sequences of maxlen characters
maxlen = 40
step = 3
sentences = []
next_chars = []
for i in range(0, len(text) - maxlen, step):
sentences.append(text[i: i + maxlen])
next_chars.append(text[i + maxlen])
print('nb sequences:', len(sentences))
print('Vectorization...')
X = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
X[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
# build the model: a single LSTM
print('Build model...')
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(chars))))
model.add(Dense(len(chars)))
model.add(Activation('softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
# train the model, output generated text after each iteration
for iteration in range(1, 60):
print()
print('-' * 50)
print('Iteration', iteration)
model.fit(X, y, batch_size=128, nb_epoch=1)
start_index = random.randint(0, len(text) - maxlen - 1)
for diversity in [0.2, 0.5, 1.0, 1.2]:
print()
print('----- diversity:', diversity)
generated = ''
sentence = text[start_index: start_index + maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generated)
for i in range(400):
x = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(sentence):
x[0, t, char_indices[char]] = 1.
preds = model.predict(x, verbose=0)[0]
next_index = sample(preds, diversity)
next_char = indices_char[next_index]
generated += next_char
sentence = sentence[1:] + next_char
sys.stdout.write(next_char)
sys.stdout.flush()
print()
| 3.2 RNN and LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from itertools import permutations
import matplotlib.pyplot as plt
# **Purpose of this notebook**
#
# To optimize placement of magnets such that the cross-sectional area of the volume we are imaging is as homogeneous as possible.
# +
N = 8 #Input number of spaces in NMR Mandhala to optimize
numMagnets = 8 #Input number of magnets to choose from
R = 0.0145 #Input radius of NMR Mandhala to optimize in meters
#R measured from center of Mandhala to center of magnets
V = (0.25**2*1)*(0.025**3) #Volume of each magnet in meters^3
numPoints = 10 #Input number of test points around center to check homogeneity
frac = 0.10 #Input fraction of R to use as radius of circle over which test points will be spread evenly
# Input surface magnetic field values (in Telsa)
#for each of the N magnets to be placed in Mandhala
magnets = np.empty(numMagnets)
magnets[0] = 0.41
magnets[1] = 0.42
magnets[2] = 0.41
magnets[3] = 0.41
magnets[4] = 0.40
magnets[5] = 0.42
magnets[6] = 0.42
magnets[7] = 0.41
# -
# **Cell below sets up the particular MANDHALA geometry**
#
# `N` magnets arranged around circumference of circle of radius `R`
#
# `alpha` is the angle from the positive z-axis to the center of each magnet
#
# `beta` is the angle from the positive z-axis of the magnetic dipole orientation of each magnet
#
# `gamma` is the angle from the positive z-axis to each test point
#
# `n0` is a 2D array that gives the normalized position vector of the ith magnet
#
# `P` is a 2D array that gives the position vector of each test point
# +
# DEFINING THE ANGLE ARRAYS
#alpha is an array and alpha[n] gives the angle measured
#from center of first magnet to the center of nth magnet
#going clockwise from positive z-axis (which is in direction of generated magnetic field)
alpha = np.array([n*2*np.pi/N for n in range(N)])
#beta is an array that gives the magnetic field orientation of nth magnet
beta = 2*alpha
#gamma is an array that gives the angle from z axis to the jth test point
gamma = np.array([j*2*np.pi/numPoints for j in range(numPoints)])
#DEFINING POSITION VECTORS
#this creates a 2D array, n0
#such that n0[i] gives the normalized position vector of the ith magnet in Mandhala
x = np.zeros(N)
y = np.sin(alpha)
z = np.cos(alpha)
n0 = np.vstack([x, y, z]).T
#this creates a 2D array, P
#such that P[j] gives the position vector of the jth test point
x = frac*R*np.zeros(numPoints)
y = frac*R*np.sin(gamma)
z = frac*R*np.cos(gamma)
P = np.vstack([x, y, z]).T
# -
# **Cell below defines a function which will evaluate the magnetic field difference at a certain point relative to the magnetic field you would expect if all magnets had the same magnetic moment equal to the average of all the magnetic moments of current arrangement**
#
# >*Inputs:*
#
# >`m` 1D array of the magnetic moments for current arrangement of magnets
#
# >`beta` 1D array with directions of magnetic moments for each magnet
#
# >`rV` 2D array such that `rV[i]` gives the position vector from 'point' to the ith magnet
#
# >*Output:*
#
# >A vector that gives difference of magnetic field at that point relative to magnetic field if all magnets had the same magnetic moment.
#
# The function to calculate the magnetic field difference is given below:
#
# $$ \Delta \vec{B} = \frac{\mu_0}{4\pi} \sum_{i=0}^{N-1} \frac{3(\Delta \vec{m}_i \cdot \vec{n}_i) \vec{n}_i - \Delta \vec{m}_i}{\| \vec{r}_i \|^3} $$
#
# The cost function we are minimizing becomes the sum over all test point locations of the magnitude of this magnetic field difference.
#
# $$ \textrm{cost function} = \sum_\textrm{test points} \|\Delta \vec{B}\|_\textrm{point}$$
# +
def calcDeltaB(m, beta, rV):
N = len(m)
#calculating m0, the average magnetic moment
m0 = np.mean(m)
#mV gives 2D array that gives magnetic field vector for particular permutation of magnets
mV = np.array([m[i]*np.array([0, np.sin(beta[i]), np.cos(beta[i])]) for i in range(N)])
#delta gives an array of the difference in magnetic field values of nth magnet with mean
delta = m - m0
#delmV gives a 2D array of the magnetic field difference from mean and direction of the nth magnet as the nth column
#x,y,z positions (relative to center of circle) as rows
delmV = np.array([delta[i]*np.array([0, np.sin(beta[i]), np.cos(beta[i])]) for i in range(N)])
#rnorm finds the norm of rV
rnorm = np.array([np.linalg.norm(rV[i]) for i in range(N)])
#nV gives the n-vector to use for this point
nV = np.array([rV[i]/rnorm[i] for i in range(N)])
DeltaB = 10**(-7)*np.sum(np.array([(3*(np.dot(delmV[i], nV[i]) * nV[i]) - delmV[i])/(rnorm[i]**3) for i in range(N)]),0)
return DeltaB
def calcB(m, beta, rV):
N = len(m)
#mV gives 2D array that gives magnetic field vector for particular permutation of magnets
mV = np.array([m[i]*np.array([0, np.sin(beta[i]), np.cos(beta[i])]) for i in range(N)])
#rnorm finds the norm of rV
rnorm = np.array([np.linalg.norm(rV[i]) for i in range(N)])
#nV gives the n-vector to use for this point
nV = np.array([rV[i]/rnorm[i] for i in range(N)])
B = 10**(-7)*np.sum(np.array([(3*(np.dot(mV[i], nV[i]) * nV[i]) - mV[i])/(rnorm[i]**3) for i in range(N)]),0)
return B
# -
# **Cell below goes through different permutations of magnet arrangements and saves the one with the lowest cost function**
#
# Here we are treating each magnet as a magnetic dipole, and the equation for the magnetic field of a magnetic dipole requires the magnetic moment. The magnetic dipole is given by the following formulat in terms of the magnetization of the magnet $M$ and its volume $V$:
# $$ \vec{m} = \vec{M} V$$
#
# The magnetization can be found if we know the residual flux density of the magnet, $B_r$.
# $$\vec{M} = \frac{\vec{B}_r}{\mu_0}$$
#
# K&J magnetics gives the maximum residual flux density for the N-52 magnets as 1.48 Teslas and surface field of 0.5461 Teslas for the magnets we use ([Large magnets](https://www.kjmagnetics.com/proddetail.asp?prod=BY088-N52) and [smaller magnets](https://www.kjmagnetics.com/proddetail.asp?prod=BX044-N52)).
#
# Since we measure the surface magnetic fields of each magnet,$B_\textrm{surf}$, I propose using the following:
# $$B_r = \frac{B_\textrm{surf}}{0.5461} 1.48 = 2.71 B_\textrm{surf}$$
# +
import time
start = time.time()
#keeps track of current best permutation choice
p_min = np.zeros(N)
#keeps track of current best minimum value of optimization function
val_min = np.Infinity
for p in permutations(range(numMagnets),N):
#m gives an array of the magnetic moments for particular permutation of magnets being tested
m = 2.71*magnets.take(p)*V/(4*np.pi*10**(-7))
#FINDING COST FUNCTION CONTRIBUTION FROM CENTER OF MANDHALA
#costV gives a column vector that provides contribution to cost function
costV = calcDeltaB(m, beta, R*n0)
#val is the norm of the total cost function
val = np.linalg.norm(costV)
#FINDING COST FUNCTION CONTRIBUTIONS FROM OTHER POINTS IN YZ PLANE NEAR CENTER
for point in P:
#this creates a 2D array, rV
#such that rV[i] gives the position vector from 'point' to the ith magnet
rV = np.array([(R*n0[i] - point) for i in range(N)])
costV = calcDeltaB(m, beta, rV)
val += np.linalg.norm(costV)
if val < val_min:
val_min = val
p_min = p
end = time.time()
print("Execution Time:", end - start, " seconds")
print("Minimum Value:", val_min)
print("Permutation corresponding to the minimum value:", p_min)
# -
# **Cell below defines a function that will display information for inputted permutation of magnets**
def VisualizePermutation(p):
print(magnets.take(p))
m = 2.71*magnets.take(p)*V/(4*np.pi*10**(-7))
Bcenter = calcB(m, beta, R*n0)
print("Center Bz value: ", Bcenter[2])
k = 0
Bxpoint = np.zeros(numPoints)
Bypoint = np.zeros(numPoints)
Bzpoint = np.zeros(numPoints)
for testPoint in P:
rV = np.array([(R*n0[i] - testPoint) for i in range(N)])
Bpoint= calcB(m, beta, rV)
Bxpoint[k] = Bpoint[0]
Bypoint[k] = Bpoint[1]
Bzpoint[k] = Bpoint[2]
k += 1
deltaBz = max(Bzpoint) - min(Bzpoint)
print("Range of Bz values over test points: ", deltaBz)
# %matplotlib inline
plt.plot(gamma, Bzpoint, label = 'Bz')
plt.plot(gamma, Bypoint, label = 'By')
plt.plot(gamma, Bxpoint, label = 'Bx')
plt.legend()
plt.xlabel("Angle (Radians)")
plt.ylabel("Magnetic Field Strength (Telsa)")
plt.title("Magnetic Field Variation Around Center")
# **Cells below are visualizing different permutations**
VisualizePermutation(p_min)
p = [0, 1, 4, 5, 2, 3, 6, 7]
VisualizePermutation(p)
# **Possible future steps:**
# - This calculation over-estimates the magnetic field, because we are assuming the magnetic field contributed by each magnet is equal to a magnetic dipole with the same magnetic moment as the bar magnet centered at the center of each magnet.
#
# - It would be more accurate to break the bar magnet into cubic pieces of the same magnetization and treat each cube as a magnetic dipole. Since the magnetic moment of the bar magnet scales with the volume, breaking the 1-inch longer magnet into 4 1/4"-cubes would reduce each cubes magnetic dipole moment by 4 and each cube's contribution to the field would be smaller than assumed here due to slightly increased distance from the center plane.
#
# - Could see if making such an adjustment to our model would more accurately predict the magnetic field we measure with the actual magnets (and allow us to more accurately optimize our magnet design).
| EvenMoreOptimizedMandhala_2019.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# required libraries
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
# read dataset
dataset = pd.read_csv("wines.csv")
# split into components X and Y
X = dataset.iloc[:, 0:13].values
y = dataset.iloc[:, 13].values
# split into train and test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# +
# preprocessing
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# +
# pca
from sklearn.decomposition import PCA
pca = PCA(n_components = 2)
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
explained_variance = pca.explained_variance_ratio_
# -
explained_variance
pca = PCA().fit(X_train)
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
| select principal components/select_principal_components.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # 📃 Solution for Exercise M7.02
#
# We presented different classification metrics in the previous notebook.
# However, we did not use it with a cross-validation. This exercise aims at
# practicing and implementing cross-validation.
#
# We will reuse the blood transfusion dataset.
# + vscode={"languageId": "python"}
import pandas as pd
blood_transfusion = pd.read_csv("../datasets/blood_transfusion.csv")
data = blood_transfusion.drop(columns="Class")
target = blood_transfusion["Class"]
# -
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# First, create a decision tree classifier.
# + vscode={"languageId": "python"}
# solution
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier()
# -
# Create a `StratifiedKFold` cross-validation object. Then use it inside the
# `cross_val_score` function to evaluate the decision tree. We will first use
# the accuracy as a score function. Explicitly use the `scoring` parameter
# of `cross_val_score` to compute the accuracy (even if this is the default
# score). Check its documentation to learn how to do that.
# + vscode={"languageId": "python"}
# solution
from sklearn.model_selection import cross_val_score, StratifiedKFold
cv = StratifiedKFold(n_splits=10)
scores = cross_val_score(tree, data, target, cv=cv, scoring="accuracy")
print(f"Accuracy score: {scores.mean():.3f} +/- {scores.std():.3f}")
# -
# Repeat the experiment by computing the `balanced_accuracy`.
# + vscode={"languageId": "python"}
# solution
scores = cross_val_score(tree, data, target, cv=cv,
scoring="balanced_accuracy")
print(f"Balanced accuracy score: {scores.mean():.3f} +/- {scores.std():.3f}")
# -
# We will now add a bit of complexity. We would like to compute the precision
# of our model. However, during the course we saw that we need to mention the
# positive label which in our case we consider to be the class `donated`.
#
# We will show that computing the precision without providing the positive
# label will not be supported by scikit-learn because it is indeed ambiguous.
# + vscode={"languageId": "python"}
from sklearn.model_selection import cross_val_score
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier()
try:
scores = cross_val_score(tree, data, target, cv=10, scoring="precision")
except ValueError as exc:
print(exc)
# -
# <div class="admonition tip alert alert-warning">
# <p class="first admonition-title" style="font-weight: bold;">Tip</p>
# <p class="last">We catch the exception with a <tt class="docutils literal">try</tt>/<tt class="docutils literal">except</tt> pattern to be able to print it.</p>
# </div>
# We get an exception because the default scorer has its positive label set to
# one (`pos_label=1`), which is not our case (our positive label is "donated").
# In this case, we need to create a scorer using the scoring function and the
# helper function `make_scorer`.
#
# So, import `sklearn.metrics.make_scorer` and
# `sklearn.metrics.precision_score`. Check their documentations for more
# information.
# Finally, create a scorer by calling `make_scorer` using the score function
# `precision_score` and pass the extra parameter `pos_label="donated"`.
# + vscode={"languageId": "python"}
# solution
from sklearn.metrics import make_scorer, precision_score
precision = make_scorer(precision_score, pos_label="donated")
# -
# Now, instead of providing the string `"precision"` to the `scoring` parameter
# in the `cross_val_score` call, pass the scorer that you created above.
# + vscode={"languageId": "python"}
# solution
scores = cross_val_score(tree, data, target, cv=cv, scoring=precision)
print(f"Precision score: {scores.mean():.3f} +/- {scores.std():.3f}")
# -
# `cross_val_score` will only compute a single score provided to the `scoring`
# parameter. The function `cross_validate` allows the computation of multiple
# scores by passing a list of string or scorer to the parameter `scoring`,
# which could be handy.
#
# Import `sklearn.model_selection.cross_validate` and compute the accuracy and
# balanced accuracy through cross-validation. Plot the cross-validation score
# for both metrics using a box plot.
# + vscode={"languageId": "python"}
# solution
from sklearn.model_selection import cross_validate
scoring = ["accuracy", "balanced_accuracy"]
scores = cross_validate(tree, data, target, cv=cv, scoring=scoring)
scores
# + tags=["solution"] vscode={"languageId": "python"}
import pandas as pd
color = {"whiskers": "black", "medians": "black", "caps": "black"}
metrics = pd.DataFrame(
[scores["test_accuracy"], scores["test_balanced_accuracy"]],
index=["Accuracy", "Balanced accuracy"]
).T
# + tags=["solution"] vscode={"languageId": "python"}
import matplotlib.pyplot as plt
metrics.plot.box(vert=False, color=color)
_ = plt.title("Computation of multiple scores using cross_validate")
| notebooks/M7 Evaluating Model Performance - C4 Classification Metrics - S2 ex M7.02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Experiments for ER Graph
#
# ## Imports
# +
# %load_ext autoreload
# %autoreload 2
import os
import sys
from collections import OrderedDict
import logging
import math
from matplotlib import pyplot as plt
import networkx as nx
import numpy as np
import torch
from torchdiffeq import odeint, odeint_adjoint
sys.path.append('../../')
sys.path.append('../kuramoto_utilities')
# Baseline imports
from gd_controller import AdjointGD
from dynamics_driver import ForwardKuramotoDynamics, BackwardKuramotoDynamics
# Nodec imports
from neural_net import EluTimeControl, TrainingAlgorithm
# Various Utilities
from utilities import evaluate, calculate_critical_coupling_constant, comparison_plot, state_plot
from nnc.helpers.torch_utils.oscillators import order_parameter_cos
logging.getLogger().setLevel(logging.CRITICAL) # set to info to look at loss values etc.
# -
# ## Load graph parameters
# Basic setup for calculations, graph, number of nodes, etc.
# +
dtype = torch.float32
device = 'cpu'
graph_type = 'erdos_renyi'
adjacency_matrix = torch.load('../../data/'+graph_type+'_adjacency.pt')
parameters = torch.load('../../data/parameters.pt')
# driver vector is a column vector with 1 value for driver nodes
# and 0 for non drivers.
result_folder = '../../results/' + graph_type + os.path.sep
os.makedirs(result_folder, exist_ok=True)
# -
# ## Load dynamics parameters
# Load natural frequencies and initial states which are common for all graphs and also calculate the coupling constant which is different per graph. We use a coupling constant value that is $10%$ of the critical coupling constant value.
# +
total_time = parameters['total_time']
total_time = 5
natural_frequencies = parameters['natural_frequencies']
critical_coupling_constant = calculate_critical_coupling_constant(adjacency_matrix, natural_frequencies)
coupling_constant = 0.15*critical_coupling_constant
theta_0 = parameters['theta_0']
# -
# ## NODEC
# We now train NODEC with a shallow neural network. We initialize the parameters in a deterministic manner, and use stochastic gradient descent to train it. The learning rate, number of epochs and neural architecture may change per graph. We use different fractions of driver nodes.
# +
fractions = np.linspace(0,1,10)
order_parameter_mean = []
order_parameter_std = []
samples = 1000
for p in fractions:
sample_arr = []
for i in range(samples):
print(p,i)
driver_nodes = int(p*adjacency_matrix.shape[0])
driver_vector = torch.zeros([adjacency_matrix.shape[0],1])
idx = torch.randperm(len(driver_vector))[:driver_nodes]
driver_vector[idx] = 1
forward_dynamics = ForwardKuramotoDynamics(adjacency_matrix,
driver_vector,
coupling_constant,
natural_frequencies
)
backward_dynamics = BackwardKuramotoDynamics(adjacency_matrix,
driver_vector,
coupling_constant,
natural_frequencies
)
neural_net = EluTimeControl([2])
for parameter in neural_net.parameters():
parameter.data = torch.ones_like(parameter.data)/1000 # deterministic init!
train_algo = TrainingAlgorithm(neural_net, forward_dynamics)
best_model = train_algo.train(theta_0, total_time, epochs=3, lr=0.3)
control_trajectory, state_trajectory =\
evaluate(forward_dynamics, theta_0, best_model, total_time, 100)
nn_control = torch.cat(control_trajectory).squeeze().cpu().detach().numpy()
nn_states = torch.cat(state_trajectory).cpu().detach().numpy()
nn_e = (nn_control**2).cumsum(-1)
nn_r = order_parameter_cos(torch.tensor(nn_states)).cpu().numpy()
sample_arr.append(nn_r[-1])
order_parameter_mean.append(np.mean(sample_arr))
order_parameter_std.append(np.std(sample_arr,ddof=1))
# +
order_parameter_mean = np.array(order_parameter_mean)
order_parameter_std = np.array(order_parameter_std)
plt.figure()
plt.errorbar(fractions,order_parameter_mean,yerr=order_parameter_std/np.sqrt(samples),fmt="o")
plt.xlabel(r"fraction of controlled nodes")
plt.ylabel(r"$r(T)$")
plt.tight_layout()
plt.show()
# -
np.savetxt("ER_drivers_K015.csv",np.c_[order_parameter_mean,order_parameter_std],header="order parameter mean\t order parameter std")
| examples/kuramoto/ER_drivers_K015.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
df=pd.read_csv('D:\ExcelR\simple linear4\delivery_time.csv')
df.head()
df.info()
df.describe()
df.boxplot()
df.info()
import seaborn as sns
sns.pairplot(data=df)
# +
#there is positive assosiation between sorting time and delivery time
# -
df.corr()
# # simple linear regression MODEL 1
df.rename(columns = {'Delivery Time': 'Delivery_Time', 'Sorting Time': 'Sorting_Time'}, inplace = True)
import numpy as np
import statsmodels.formula.api as smf
# +
# Bulding model by ols method for prediction of delicery time.
# -
model1=smf.ols("Delivery_Time~Sorting_Time",data=df).fit()
model1.params
model1.summary()
pred=model1.predict(df.iloc[:,1]) #model1.predict(df["Sorting_Time"])
pred
rmse=np.sqrt(np.mean((np.array(df['Delivery_Time'])-np.array(pred))**2))
rmse
newdata=pd.Series([10,15,12,6,11,2,6,7])
data_pred=pd.DataFrame(newdata,columns=['Sorting_Time'])
model1.predict(data_pred)
model2=smf.ols('Delivery_Time~np.log(Sorting_Time)',data=df).fit()
model2.params
model2.summary()
pred2=model2.predict(df.iloc[:,1]) #model1.predict(df["Sorting_Time"])
pred2
rmse=np.sqrt(np.mean((np.array(df['Delivery_Time'])-np.array(pred2))**2))
rmse
#second exponential TRASFORMATION
model3=smf.ols('np.log(Delivery_Time)~Sorting_Time',data=df).fit()
model3.params
model3.summary()
model3.rsquared
pred3=model3.predict(df.iloc[:,1]) #model1.predict(df["Sorting_Time"])
pred3
rmse=np.sqrt(np.mean((np.array(df['Delivery_Time'])-np.array(pred3))**2))
rmse
newdata=pd.Series([10,15,12,6,11,2,6,7])
data_pred=pd.DataFrame(newdata,columns=['Sorting_Time'])
data_pred
model3.predict(data_pred)
# # model 3 is better than other models in this data set
| delivery time (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Vanilla LSTM for Sentiment Classification
# This LSTM performs sentiment analysis on the IMDB review dataset.
# +
import os
from tensorflow.keras.datasets import imdb
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding, Dense, Flatten, Dropout, Activation, SpatialDropout1D
from tensorflow.keras.layers import LSTM
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping, TensorBoard
from tensorflow.keras.models import load_model
import sklearn.metrics
from sklearn.metrics import roc_auc_score
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# #### Set Hyperparameters
# +
output_dir = 'model_output/vanillaLSTM'
e_param = 0.006
epochs = 5
batch_size = 64
patience = 3
val_split = .2
n_dim = 256
n_unique_words = 8000
max_review_length = 300
pad_type = trunc_type = 'pre'
n_lstm = 128
drop_lstm = 0.5
# -
# #### Load Data
(X_train, y_train), (X_valid, y_valid) = imdb.load_data(num_words=n_unique_words)
# #### Preprocess Data
X_train = pad_sequences(X_train, maxlen=max_review_length, padding=pad_type, truncating=trunc_type, value=0)
X_valid = pad_sequences(X_valid, maxlen=max_review_length, padding=pad_type, truncating=trunc_type, value=0)
# #### Design Deep Net Architecture
# +
model = Sequential()
model.add(Embedding(n_unique_words, n_dim, input_length=max_review_length))
model.add(LSTM(n_lstm, dropout=drop_lstm))
model.add(Dense(1, activation='sigmoid'))
# -
model.summary()
# #### Configure the Model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# +
modelCheckpoint = ModelCheckpoint(filepath=output_dir + '/imdb-vanilla-lstm.hdf5', save_best_only=True, mode='max', monitor='val_accuracy')
earlyStopping = EarlyStopping(mode='min', patience=patience)
if not os.path.exists(output_dir):
os.makedirs(output_dir)
# -
# #### TensorBoard
tensorboard = TensorBoard("../logs/imdb-vanilla-lstm")
# ### Train the Model
model.fit(X_train, y_train, batch_size=batch_size, epochs=epochs, verbose=1,
validation_split=val_split, callbacks=[modelCheckpoint, earlyStopping, tensorboard])
# #### Evaluate
model = load_model(output_dir + '/imdb-vanilla-lstm.hdf5')
y_hat = model.predict_proba(X_valid)
plt.hist(y_hat)
_ = plt.axvline(x=0.5, color='orange')
pct_auc = roc_auc_score(y_valid, y_hat) * 100
'{:0.2f}'.format(pct_auc)
# +
fpr, tpr, _ = sklearn.metrics.roc_curve(y_valid, y_hat)
roc_auc = sklearn.metrics.auc(fpr, tpr)
plt.figure()
lw = 2
plt.plot(fpr, tpr, color='darkorange',
lw=lw, label='ROC curve (area = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic')
plt.legend(loc="lower right")
plt.show()
# -
| notebooks/nlp/3_vanilla_lstm_in_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import torch
import gpytorch
import tqdm
import matplotlib.pyplot as plt
# %matplotlib inline
# # Modifying the Variational Strategy/Variational Distribution
#
# The predictive distribution for approximate GPs is given by
#
# $$
# p( \mathbf f(\mathbf x^*) ) = \int_{\mathbf u} p( f(\mathbf x^*) \mid \mathbf u) \: q(\mathbf u) \: d\mathbf u,
# \quad
# q(\mathbf u) = \mathcal N( \mathbf m, \mathbf S).
# $$
#
# $\mathbf u$ represents the function values at the $m$ inducing points.
# Here, $\mathbf m \in \mathbb R^m$ and $\mathbf S \in \mathbb R^{m \times m}$ are learnable parameters.
#
# If $m$ (the number of inducing points) is quite large, the number of learnable parameters in $\mathbf S$ can be quite unwieldy.
# Furthermore, a large $m$ might make some of the computations rather slow.
# Here we show a few ways to use different [variational distributions](https://gpytorch.readthedocs.io/en/latest/variational.html#variational-distributions) and
# [variational strategies](https://gpytorch.readthedocs.io/en/latest/variational.html#variational-strategies) to accomplish this.
# ### Experimental setup
#
# We're going to train an approximate GP on a medium-sized regression dataset, taken from the UCI repository.
# +
import urllib.request
import os
from scipy.io import loadmat
from math import floor
# this is for running the notebook in our testing framework
smoke_test = ('CI' in os.environ)
if not smoke_test and not os.path.isfile('../elevators.mat'):
print('Downloading \'elevators\' UCI dataset...')
urllib.request.urlretrieve('https://drive.google.com/uc?export=download&id=1jhWL3YUHvXIaftia4qeAyDwVxo6j1alk', '../elevators.mat')
if smoke_test: # this is for running the notebook in our testing framework
X, y = torch.randn(1000, 3), torch.randn(1000)
else:
data = torch.Tensor(loadmat('../elevators.mat')['data'])
X = data[:, :-1]
X = X - X.min(0)[0]
X = 2 * (X / X.max(0)[0]) - 1
y = data[:, -1]
train_n = int(floor(0.8 * len(X)))
train_x = X[:train_n, :].contiguous()
train_y = y[:train_n].contiguous()
test_x = X[train_n:, :].contiguous()
test_y = y[train_n:].contiguous()
if torch.cuda.is_available():
train_x, train_y, test_x, test_y = train_x.cuda(), train_y.cuda(), test_x.cuda(), test_y.cuda()
# +
from torch.utils.data import TensorDataset, DataLoader
train_dataset = TensorDataset(train_x, train_y)
train_loader = DataLoader(train_dataset, batch_size=500, shuffle=True)
test_dataset = TensorDataset(test_x, test_y)
test_loader = DataLoader(test_dataset, batch_size=500, shuffle=False)
# -
# ### Some quick training/testing code
#
# This will allow us to train/test different model classes.
# +
# this is for running the notebook in our testing framework
num_epochs = 1 if smoke_test else 10
# Our testing script takes in a GPyTorch MLL (objective function) class
# and then trains/tests an approximate GP with it on the supplied dataset
def train_and_test_approximate_gp(model_cls):
inducing_points = torch.randn(128, train_x.size(-1), dtype=train_x.dtype, device=train_x.device)
model = model_cls(inducing_points)
likelihood = gpytorch.likelihoods.GaussianLikelihood()
mll = gpytorch.mlls.VariationalELBO(likelihood, model, num_data=train_y.numel())
optimizer = torch.optim.Adam(list(model.parameters()) + list(likelihood.parameters()), lr=0.1)
if torch.cuda.is_available():
model = model.cuda()
likelihood = likelihood.cuda()
# Training
model.train()
likelihood.train()
epochs_iter = tqdm.notebook.tqdm(range(num_epochs), desc=f"Training {model_cls.__name__}")
for i in epochs_iter:
# Within each iteration, we will go over each minibatch of data
for x_batch, y_batch in train_loader:
optimizer.zero_grad()
output = model(x_batch)
loss = -mll(output, y_batch)
epochs_iter.set_postfix(loss=loss.item())
loss.backward()
optimizer.step()
# Testing
model.eval()
likelihood.eval()
means = torch.tensor([0.])
with torch.no_grad():
for x_batch, y_batch in test_loader:
preds = model(x_batch)
means = torch.cat([means, preds.mean.cpu()])
means = means[1:]
error = torch.mean(torch.abs(means - test_y.cpu()))
print(f"Test {model_cls.__name__} MAE: {error.item()}")
# -
# ## The Standard Approach
#
# As a default, we'll use the default [VariationalStrategy](https://gpytorch.readthedocs.io/en/latest/variational.html#id1) class with a [CholeskyVariationalDistribution](https://gpytorch.readthedocs.io/en/latest/variational.html#choleskyvariationaldistribution).
# The `CholeskyVariationalDistribution` class allows $\mathbf S$ to be on any positive semidefinite matrix. This is the most general/expressive option for approximate GPs.
class StandardApproximateGP(gpytorch.models.ApproximateGP):
def __init__(self, inducing_points):
variational_distribution = gpytorch.variational.CholeskyVariationalDistribution(inducing_points.size(-2))
variational_strategy = gpytorch.variational.VariationalStrategy(
self, inducing_points, variational_distribution, learn_inducing_locations=True
)
super().__init__(variational_strategy)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
train_and_test_approximate_gp(StandardApproximateGP)
# ## Reducing parameters
#
# ### MeanFieldVariationalDistribution: a diagonal $\mathbf S$ matrix
#
# One way to reduce the number of parameters is to restrict that $\mathbf S$ is only diagonal. This is less expressive, but the number of parameters is now linear in $m$ instead of quadratic.
#
# All we have to do is take the previous example, and change `CholeskyVariationalDistribution` (full $\mathbf S$ matrix) to [MeanFieldVariationalDistribution](https://gpytorch.readthedocs.io/en/latest/variational.html#meanfieldvariationaldistribution) (diagonal $\mathbf S$ matrix).
class MeanFieldApproximateGP(gpytorch.models.ApproximateGP):
def __init__(self, inducing_points):
variational_distribution = gpytorch.variational.MeanFieldVariationalDistribution(inducing_points.size(-2))
variational_strategy = gpytorch.variational.VariationalStrategy(
self, inducing_points, variational_distribution, learn_inducing_locations=True
)
super().__init__(variational_strategy)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
train_and_test_approximate_gp(MeanFieldApproximateGP)
# ### DeltaVariationalDistribution: no $\mathbf S$ matrix
#
# A more extreme method of reducing parameters is to get rid of $\mathbf S$ entirely. This corresponds to learning a delta distribution ($\mathbf u = \mathbf m$) rather than a multivariate Normal distribution for $\mathbf u$. In other words, this corresponds to performing MAP estimation rather than variational inference.
#
# In GPyTorch, getting rid of $\mathbf S$ can be accomplished by using a [DeltaVariationalDistribution](https://gpytorch.readthedocs.io/en/latest/variational.html#deltavariationaldistribution).
class MAPApproximateGP(gpytorch.models.ApproximateGP):
def __init__(self, inducing_points):
variational_distribution = gpytorch.variational.DeltaVariationalDistribution(inducing_points.size(-2))
variational_strategy = gpytorch.variational.VariationalStrategy(
self, inducing_points, variational_distribution, learn_inducing_locations=True
)
super().__init__(variational_strategy)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
train_and_test_approximate_gp(MAPApproximateGP)
# ## Reducing computation (through decoupled inducing points)
#
# One way to reduce the computational complexity is to use separate inducing points for the mean and covariance computations. The [Orthogonally Decoupled Variational Gaussian Processes](https://arxiv.org/abs/1809.08820) method of Salimbeni et al. (2018) uses more inducing points for the (computationally easy) mean computations and fewer inducing points for the (computationally intensive) covariance computations.
#
# In GPyTorch we implement this method in a modular way. The [OrthogonallyDecoupledVariationalStrategy](http://gpytorch.ai/variational.html#gpytorch.variational.OrthogonallyDecoupledVariationalStrategy) defines the variational strategy for the mean inducing points. It wraps an existing variational strategy/distribution that defines the covariance inducing points:
def make_orthogonal_vs(model, train_x):
mean_inducing_points = torch.randn(1000, train_x.size(-1), dtype=train_x.dtype, device=train_x.device)
covar_inducing_points = torch.randn(100, train_x.size(-1), dtype=train_x.dtype, device=train_x.device)
covar_variational_strategy = gpytorch.variational.VariationalStrategy(
model, covar_inducing_points,
gpytorch.variational.CholeskyVariationalDistribution(covar_inducing_points.size(-2)),
learn_inducing_locations=True
)
variational_strategy = gpytorch.variational.OrthogonallyDecoupledVariationalStrategy(
covar_variational_strategy, mean_inducing_points,
gpytorch.variational.DeltaVariationalDistribution(mean_inducing_points.size(-2)),
)
return variational_strategy
# Putting it all together we have:
class OrthDecoupledApproximateGP(gpytorch.models.ApproximateGP):
def __init__(self, inducing_points):
variational_distribution = gpytorch.variational.DeltaVariationalDistribution(inducing_points.size(-2))
variational_strategy = make_orthogonal_vs(self, train_x)
super().__init__(variational_strategy)
self.mean_module = gpytorch.means.ConstantMean()
self.covar_module = gpytorch.kernels.ScaleKernel(gpytorch.kernels.RBFKernel())
def forward(self, x):
mean_x = self.mean_module(x)
covar_x = self.covar_module(x)
return gpytorch.distributions.MultivariateNormal(mean_x, covar_x)
train_and_test_approximate_gp(OrthDecoupledApproximateGP)
| examples/04_Variational_and_Approximate_GPs/Modifying_the_variational_strategy_and_distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
from modsim import *
def make_system(*, gammatau=1.36*0.2, mu=1.36*0.001, beta=0.00027, rho=0.1, alpha=3.6*0.01, delta=0.33, sigma=2, pi=100, end=120):
init_state = State(R=200, L=0, E=0, V=0.0000004)
t0 = 0
return System(init=init_state, t0=t0, t_end=120, dt=0.1,
params={'gammatau':gammatau,
'mu':mu,
'beta':beta,
'rho':rho,
'alpha':alpha,
'delta':delta,
'sigma':sigma,
'pi': pi
}
)
sys = make_system()
def update_func(state: State, t: float, system: System) -> State:
dRdt = system.params['gammatau'] + -system.params['mu']*state.R + -system.params['beta']*state.R*state.V
#print(system.params['gammatau'],system.params['mu']*state.R,system.params['beta']*state.R*state.V)
dR = dRdt*system.dt
# print(dR)
dLdt = system.params['rho']*system.params['beta']*state.R*state.V + -system.params['mu']*state.L + -system.params['alpha']*state.L
dL = system.dt*dLdt
dEdt = (1-system.params['rho'])*system.params['beta']*state.R*state.V + system.params['alpha']*state.L + -system.params['delta']*state.E
dE = dEdt*system.dt
dVdt = system.params['pi']*state.E + -system.params['sigma']*state.V
dV = system.dt*dVdt
state.R += dR
state.L += dL
state.E += dE
state.V += dV
newState = State(R=state.R, L=state.L, E=state.E, V=state.V)
return newState
def run_sim(system, update_func):
frame = TimeFrame(columns=system.init.index)
frame.row[system.t0] = system.init
for index in linrange(0, 1200, 1):
frame.row[index+1] = update_func(frame.row[index], index, system)
return frame
sys = make_system()
result = run_sim(sys, update_func);
#plot(result.R, '-', label='r')
#plot(result.L, '-', label='l')
#plot(result.E, '-', label='e')
#plot(result.V, '-', label='v')
#decorate(xlabel='Time (idk)',
#ylabel='AAAAAA')
print(result)
plot(result.R, '-', label='r')
decorate(xlabel='Time (idk)',
ylabel='AAAAAA',
title='R')
plot(result.L, '-', label='l')
decorate(xlabel='Time (idk)',
ylabel='AAAAAA',
title='L')
plot(result.E, '-', label='e')
decorate(xlabel='Time (idk)',
ylabel='AAAAAA',
title='E')
plot(result.V, '-', label='v')
decorate(xlabel='Time (idk)',
ylabel='AAAAAA',
title='V')
index_array=linrange(0, 1200, 1)
len(index_array)
| code/Carlos_HIV_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="6MlHCEAdZPy1"
# **Initial step:** Please try to put the extracted CamVid folder in your Google Drive!
# So now you could mount your data to this ipynb!
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="UGAXY5HMYjzL" outputId="d6c3cebd-6453-4fe3-80e5-accd50c2730e"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="wTBYCMcpZoA8" outputId="c70bf925-0d59-41e0-db7f-6e85deae2c3b"
# if you mount Google drive correctly, the following commands should be able to executed correctly
# !ls /content/drive/
# %cd "/content/drive/My Drive"
# %cd "CamVid"
# !ls
# + colab={} colab_type="code" id="pfWmPZKyZ8gA"
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torch.utils.data import Dataset, DataLoader
from torchvision import utils
import torchvision
from torchvision import models
from torchvision.models.vgg import VGG
import random
from matplotlib import pyplot as plt
import numpy as np
import time
import sys
import os
from os import path
from PIL import Image
import pandas as pd
from torchvision.models.vgg import VGG
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="YMzL2KhcaI9u" outputId="86892b9b-9822-44e6-b09c-7f2c261fa7ee"
root_dir = "/content/drive/My Drive/CamVid/"
train_file = os.path.join(root_dir, "train.csv")
val_file = os.path.join(root_dir, "val.csv")
print("training csv exits:{}".format(path.exists(train_file)))
print("validation csv exits:{}".format(path.exists(val_file)))
# the folder to save results for comparison
folder_to_save_validation_result = '/content/drive/My Drive/CamVid/result_comparision/'
if os.path.isdir(folder_to_save_validation_result) == False:
os.mkdir(folder_to_save_validation_result)
# the number of segmentation classes
num_class = 11 # 32 for original CamVid # Q2-3:11->3
means = np.array([103.939, 116.779, 123.68]) / 255. # mean of three channels in the order of BGR
h, w = 256, 256
train_h = 256
train_w = 256
val_h = 256
val_w = 256
## parameters for Solver-Adam in this example
batch_size = 6 #
epochs = 20 # don't try to improve the performance by simply increasing the training epochs or iterations
lr = 1e-4 # achieved besty results
step_size = 100 # Won't work when epochs <=100
gamma = 0.5 #
#
## index for validation images
global_index = 0
# pixel accuracy and mIOU list
pixel_acc_list = []
mIOU_list = []
use_gpu = torch.cuda.is_available()
num_gpu = list(range(torch.cuda.device_count()))
class CamVidDataset(Dataset):
def __init__(self, csv_file, phase, n_class=num_class, crop=True, flip_rate=0.5):
self.data = pd.read_csv(csv_file)
self.means = means
self.n_class = n_class
self.flip_rate = flip_rate
self.resize_h = h
self.resize_w = w
if phase == 'train':
self.new_h = train_h
self.new_w = train_w
self.crop = crop
elif phase == 'val':
self.flip_rate = 0.
self.crop = False # False
self.new_h = val_h
self.new_w = val_w
def __len__(self):
return len(self.data)
def __getitem__(self, idx):
img_name = self.data.iloc[idx, 0]
img_name = root_dir + img_name
img = Image.open(img_name).convert('RGB')
label_name = self.data.iloc[idx, 1]
label_name = root_dir + label_name
label_image = Image.open(label_name)
label = np.asarray(label_image)
# In training mode, the crop strategy is random-shift crop.
# In validation model, it is center crop.
if self.crop:
w, h = img.size
A_x_offset = np.int32(np.random.randint(0, w - self.new_w + 1, 1))[0]
A_y_offset = np.int32(np.random.randint(0, h - self.new_h + 1, 1))[0]
img = img.crop((A_x_offset, A_y_offset, A_x_offset + self.new_w, A_y_offset + self.new_h)) # left, top, right, bottom
label_image = label_image.crop((A_x_offset, A_y_offset, A_x_offset + self.new_w, A_y_offset + self.new_h)) # left, top, right, bottom
else:
w, h = img.size
A_x_offset = int((w - self.new_w)/2)
A_y_offset = int((h - self.new_h)/2)
img = img.crop((A_x_offset, A_y_offset, A_x_offset + self.new_w, A_y_offset + self.new_h)) # left, top, right, bottom
label_image = label_image.crop((A_x_offset, A_y_offset, A_x_offset + self.new_w, A_y_offset + self.new_h)) # left, top, right, bottom
label_image_h, label_image_w = label_image.size
# we could try to revise the values in label for reducing the number of segmentation classes
label = np.array(label_image)
if random.random() < self.flip_rate:
img = np.fliplr(img)
label = np.fliplr(label)
# reduce mean in terms of BGR
img = np.transpose(img, (2, 0, 1)) / 255.
img[0] -= self.means[0]
img[1] -= self.means[1]
img[2] -= self.means[2]
# convert to tensor
img = torch.from_numpy(img.copy()).float()
label = torch.from_numpy(label.copy()).long()
# create one-hot encoding
h, w = label.size()
target = torch.zeros(self.n_class, h, w)
for c in range(self.n_class):
target[c][label == c] = 1
sample = {'X': img, 'Y': target, 'l': label}
return sample
train_data = CamVidDataset(csv_file=train_file, phase='train')
train_loader = DataLoader(train_data, batch_size=batch_size, shuffle=True, num_workers=8)
val_data = CamVidDataset(csv_file=val_file, phase='val', flip_rate=0)
val_loader = DataLoader(val_data, batch_size=1, num_workers=8)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="4h-a5moWhLhF" outputId="f1e637a8-0552-4185-f88e-ca3f1deccdad"
# cropped version from https://github.com/pytorch/vision/blob/master/torchvision/models/vgg.py
cfg = {
'vgg11': [64, 'M', 128, 'M', 256, 256, 'M', 512, 512, 'M', 512, 512, 'M'],
'vgg13': [64, 64, 'M', 128, 128, 'M', 256, 256, 'M', 512, 512, 'M', 512, 512, 'M'],
'vgg16': [64, 64, 'M', 128, 128, 'M', 256, 256, 256, 'M', 512, 512, 512, 'M', 512, 512, 512, 'M'],
'vgg19': [64, 64, 'M', 128, 128, 'M', 256, 256, 256, 256, 'M', 512, 512, 512, 512, 'M', 512, 512, 512, 512, 'M'],
}
ranges = {
'vgg11': ((0, 3), (3, 6), (6, 11), (11, 16), (16, 21)),
'vgg13': ((0, 5), (5, 10), (10, 15), (15, 20), (20, 25)),
'vgg16': ((0, 5), (5, 10), (10, 17), (17, 24), (24, 31)),
'vgg19': ((0, 5), (5, 10), (10, 19), (19, 28), (28, 37))
}
def make_layers(cfg, batch_norm=False):
layers = []
in_channels = 3
for v in cfg:
if v == 'M':
layers += [nn.MaxPool2d(kernel_size=2, stride=2)]
else:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3, padding=1)
if batch_norm:
layers += [conv2d, nn.BatchNorm2d(v), nn.ReLU(inplace=True)]
else:
layers += [conv2d, nn.ReLU(inplace=True)]
in_channels = v
return nn.Sequential(*layers)
class VGGNet(VGG):
def __init__(self, pretrained=True, model='vgg16', requires_grad=True, remove_fc=True, show_params=False):
super().__init__(make_layers(cfg[model]))
self.ranges = ranges[model]
if pretrained:
exec("self.load_state_dict(models.%s(pretrained=True).state_dict())" % model)
if not requires_grad:
for param in super().parameters():
param.requires_grad = False
if remove_fc: # delete redundant fully-connected layer params, can save memory
del self.classifier
if show_params:
for name, param in self.named_parameters():
print(name, param.size())
def forward(self, x):
output = {}
# get the output of each maxpooling layer (5 maxpool in VGG net)
for idx in range(len(self.ranges)):
for layer in range(self.ranges[idx][0], self.ranges[idx][1]):
x = self.features[layer](x)
output["x%d"%(idx+1)] = x
return output
class FCN8s(nn.Module):
#Ref: https://towardsdatascience.com/review-fcn-semantic-segmentation-eb8c9b50d2d1
#The layer description is accordance with the above fiture instead of the original paper. Alex 2019/12/03
def __init__(self, pretrained_net, n_class):
super().__init__()
self.n_class = n_class
self.pretrained_net = pretrained_net
self.relu = nn.ReLU(inplace=True)
self.deconv1 = nn.ConvTranspose2d(512, 512, kernel_size=3, stride=2, padding=1, dilation=1, output_padding=1)
#self.bn1 = nn.BatchNorm2d(512)
self.deconv2 = nn.ConvTranspose2d(512, 256, kernel_size=3, stride=2, padding=1, dilation=1, output_padding=1)
#self.bn2 = nn.BatchNorm2d(256)
self.deconv3 = nn.ConvTranspose2d(256, 128, kernel_size=3, stride=2, padding=1, dilation=1, output_padding=1)
#self.bn3 = nn.BatchNorm2d(128)
self.deconv4 = nn.ConvTranspose2d(128, 64, kernel_size=3, stride=2, padding=1, dilation=1, output_padding=1)
#self.bn4 = nn.BatchNorm2d(64)
self.deconv5 = nn.ConvTranspose2d(64, 32, kernel_size=3, stride=2, padding=1, dilation=1, output_padding=1)
#self.bn5 = nn.BatchNorm2d(32)
self.classifier = nn.Conv2d(32, n_class, kernel_size=1)
# eliminate the skip-connection so the output of convolution layers of FCN8s will be directly upsampled for 32x.
def forward(self, x):
output = self.pretrained_net(x)
x5 = output['x5'] # size=(N, 512, x.H/32, x.W/32)
#x4 = output['x4'] # size=(N, 512, x.H/16, x.W/16)
#x3 = output['x3'] # size=(N, 256, x.H/8, x.W/8)
score = self.relu(self.deconv1(x5)) # size=(N, 512, x.H/16, x.W/16)
#score = self.bn1(score + x4) # element-wise add, size=(N, 512, x.H/16, x.W/16)
score = self.relu(self.deconv2(score)) # size=(N, 256, x.H/8, x.W/8)
#score = self.bn2(score + x3) # element-wise add, size=(N, 256, x.H/8, x.W/8)
#score = self.bn3(self.relu(self.deconv3(score))) # size=(N, 128, x.H/4, x.W/4)
#score = self.bn4(self.relu(self.deconv4(score))) # size=(N, 64, x.H/2, x.W/2)
#score = self.bn5(self.relu(self.deconv5(score))) # size=(N, 32, x.H, x.W)
score = self.relu(self.deconv3(score))
score = self.relu(self.deconv4(score))
score = self.relu(self.deconv5(score))
score = self.classifier(score) # size=(N, n_class, x.H/1, x.W/1)
return score # size=(N, n_class, x.H/1, x.W/1)
# load pretrained weights and define FCN8s
vgg_model = VGGNet(requires_grad=True, remove_fc=True)
fcn_model = FCN8s(pretrained_net=vgg_model, n_class=num_class)
ts = time.time()
vgg_model = vgg_model.cuda()
fcn_model = fcn_model.cuda()
fcn_model = nn.DataParallel(fcn_model, device_ids=num_gpu)
print("Finish cuda loading, time elapsed {}".format(time.time() - ts))
criterion = nn.BCEWithLogitsLoss()
optimizer = optim.Adam(fcn_model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, step_size=step_size, gamma=gamma)
# + colab={} colab_type="code" id="lUjPeffKbm36"
def train():
for epoch in range(epochs):
scheduler.step()
ts = time.time()
for iter, batch in enumerate(train_loader):
optimizer.zero_grad()
if use_gpu:
inputs = Variable(batch['X'].cuda())
labels = Variable(batch['Y'].cuda())
else:
inputs, labels = Variable(batch['X']), Variable(batch['Y'])
outputs = fcn_model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
if iter % 10 == 0:
print("epoch{}, iter{}, loss: {}".format(epoch, iter, loss.data.item()))
print("Finish epoch {}, time elapsed {}".format(epoch, time.time() - ts))
val(epoch)
highest_pixel_acc = max(pixel_acc_list)
highest_mIOU = max(mIOU_list)
highest_pixel_acc_epoch = pixel_acc_list.index(highest_pixel_acc)
highest_mIOU_epoch = mIOU_list.index(highest_mIOU)
print("The highest mIOU is {} and is achieved at epoch-{}".format(highest_mIOU, highest_mIOU_epoch))
print("The highest pixel accuracy is {} and is achieved at epoch-{}".format(highest_pixel_acc, highest_pixel_acc_epoch))
def save_result_comparison(input_np, output_np):
means = np.array([103.939, 116.779, 123.68]) / 255.
global global_index
original_im_RGB = np.zeros((256,256,3))
original_im_RGB[:,:,0] = input_np[0,0,:,:]
original_im_RGB[:,:,1] = input_np[0,1,:,:]
original_im_RGB[:,:,2] = input_np[0,2,:,:]
original_im_RGB[:,:,0] = original_im_RGB[:,:,0] + means[0]
original_im_RGB[:,:,1] = original_im_RGB[:,:,1] + means[1]
original_im_RGB[:,:,2] = original_im_RGB[:,:,2] + means[2]
original_im_RGB[:,:,0] = original_im_RGB[:,:,0]*255.0
original_im_RGB[:,:,1] = original_im_RGB[:,:,1]*255.0
original_im_RGB[:,:,2] = original_im_RGB[:,:,2]*255.0
im_seg_RGB = np.zeros((256,256,3))
# the following version is designed for 11-class version and could still work if the number of classes is fewer.
for i in range(256):
for j in range(256):
if output_np[i,j] == 0:
im_seg_RGB[i,j,:] = [128, 128, 128]
elif output_np[i,j] == 1:
im_seg_RGB[i,j,:] = [128, 0, 0]
elif output_np[i,j] == 2:
im_seg_RGB[i,j,:] = [192, 192, 128]
elif output_np[i,j] == 3:
im_seg_RGB[i,j,:] = [128, 64, 128]
elif output_np[i,j] == 4:
im_seg_RGB[i,j,:] = [0, 0, 192]
elif output_np[i,j] == 5:
im_seg_RGB[i,j,:] = [128, 128, 0]
elif output_np[i,j] == 6:
im_seg_RGB[i,j,:] = [192, 128, 128]
elif output_np[i,j] == 7:
im_seg_RGB[i,j,:] = [64, 64, 128]
elif output_np[i,j] == 8:
im_seg_RGB[i,j,:] = [64, 0, 128]
elif output_np[i,j] == 9:
im_seg_RGB[i,j,:] = [64, 64, 0]
elif output_np[i,j] == 10:
im_seg_RGB[i,j,:] = [0, 128, 192]
# horizontally stack original image and its corresponding segmentation results
hstack_image = np.hstack((original_im_RGB, im_seg_RGB))
new_im = Image.fromarray(np.uint8(hstack_image))
file_name = folder_to_save_validation_result + str(global_index) + '.jpg'
global_index = global_index + 1
new_im.save(file_name)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="E7cK6h9vkbf7" outputId="886018b0-52da-4061-a151-29c2f8c54c8a"
def val(epoch):
fcn_model.eval()
total_ious = []
pixel_accs = []
for iter, batch in enumerate(val_loader): ## batch is 1 in this case
if use_gpu:
inputs = Variable(batch['X'].cuda())
else:
inputs = Variable(batch['X'])
output = fcn_model(inputs)
# only save the 1st image for comparison
if iter == 0:
print('---------iter={}'.format(iter))
# generate images
images = output.data.max(1)[1].cpu().numpy()[:,:,:]
image = images[0,:,:]
save_result_comparison(batch['X'], image)
output = output.data.cpu().numpy()
N, _, h, w = output.shape
pred = output.transpose(0, 2, 3, 1).reshape(-1, num_class).argmax(axis=1).reshape(N, h, w)
target = batch['l'].cpu().numpy().reshape(N, h, w)
for p, t in zip(pred, target):
total_ious.append(iou(p, t))
pixel_accs.append(pixel_acc(p, t))
# Calculate average IoU
total_ious = np.array(total_ious).T # n_class * val_len
ious = np.nanmean(total_ious, axis=1)
pixel_accs = np.array(pixel_accs).mean()
print("epoch{}, pix_acc: {}, meanIoU: {}, IoUs: {}".format(epoch, pixel_accs, np.nanmean(ious), ious))
global pixel_acc_list
global mIOU_list
pixel_acc_list.append(pixel_accs)
mIOU_list.append(np.nanmean(ious))
# borrow functions and modify it from https://github.com/Kaixhin/FCN-semantic-segmentation/blob/master/main.py
# Calculates class intersections over unions
def iou(pred, target):
ious = []
for cls in range(num_class):
pred_inds = pred == cls
target_inds = target == cls
intersection = pred_inds[target_inds].sum()
union = pred_inds.sum() + target_inds.sum() - intersection
if union == 0:
ious.append(float('nan')) # if there is no ground truth, do not include in evaluation
else:
ious.append(float(intersection) / max(union, 1))
# print("cls", cls, pred_inds.sum(), target_inds.sum(), intersection, float(intersection) / max(union, 1))
return ious
def pixel_acc(pred, target):
correct = (pred == target).sum()
total = (target == target).sum()
return correct / total
## perform training and validation
val(0) # show the accuracy before training
train()
# + [markdown] colab_type="text" id="Ej3AGy2a6X7A"
# Before eliminate the skip-connection, the highest mIOU is 0.42 achieved at epoch-19. The pixel accuracy is 0.84 achieved at epoch-19.
# After eliminate the skip-connection, the highest mIOU is 0.402 achieved at epoch-26. The pixel accuracy is 0.83 achieved at epoch-22.
# + colab={} colab_type="code" id="2KALOKFj67y0"
| Transfer Learning and Semantic Segmentation/Q2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Paraphrase generator
from parrot import Parrot
import torch
import warnings
warnings.filterwarnings("ignore")
# Initialize Parrot class with the pretrained model "prithivida/parrot_paraphraser_on_T5" which is available in huggingface. Loading of model might take sometime to complete so please patience. If there are errors during initializing of model then please rerun the below cell.
parrot = Parrot(model_tag="prithivida/parrot_paraphraser_on_T5")
# The below code snippet will iterate through the phrases and print parapharse sentences for each phrase
def print_paraphrase_sentences(phrases,phrase_diversity=False):
for phrase in phrases:
print("-"*100)
print("Input_phrase: ", phrase)
print("-"*100)
para_phrases = parrot.augment(input_phrase=phrase, use_gpu=False, do_diverse=phrase_diversity)
for para_phrase in para_phrases:
print(para_phrase)
# Provide list of phrases for which you would need paraphrase sentences
phrases = ["What are the famous places we should not miss in Russia?","Can you recommed some upscale restaurants in Newyork?"]
print_paraphrase_sentences(phrases)
# #### Getting phrasal diversity/variety in paraphrases
diversity_phrases = ["How are the new Macbook Pros with M1 chips?"]
# Without diversity flag set
print_paraphrase_sentences(diversity_phrases)
# With diversity flag set
# +
phrase_diversity = True
print_paraphrase_sentences(diversity_phrases,phrase_diversity)
# -
# #### Interactive shell simulation
while True:
usr_q = input("Enter a phrase (To quit enter exit): ")
if usr_q == "exit":
break
else:
if usr_q:
phrase_d = input("Do you want phrasal diversity (y or n): ")
phrases = [usr_q]
if phrase_d and phrase_d.lower() == "y":
print_paraphrase_sentences(phrases,phrase_diversity=True)
else:
print_paraphrase_sentences(phrases)
| paraphrase_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bUrwmz6asGBo"
# ### Libraries
# + id="Vd_OEKc-z0Mp" colab={"base_uri": "https://localhost:8080/"} outputId="80260108-9402-4fb4-ddef-41fa2f2f6649"
from google.colab import drive
drive.mount('/content/drive')
# + id="yzwXnz7RsFaB" colab={"base_uri": "https://localhost:8080/"} outputId="fa35db9f-bba8-4757-c480-5a7de396dfe0"
# !pip install opencv-contrib-python==3.4.2.17
# !pip install --upgrade imutils
# + id="9PlIoxBTsFW4" colab={"base_uri": "https://localhost:8080/"} outputId="38e7c425-6028-49a1-af10-26ae22dff2de"
import cv2
import numpy as np
import imutils
import matplotlib.pyplot as plt
from random import randrange
from google.colab.patches import cv2_imshow
import sys
import time
import os
import tqdm
from moviepy.editor import ImageSequenceClip
# For 24->8bit
from PIL import Image
# %matplotlib inline
# + [markdown] id="85nQ375HsBVn"
# #### For Stitcher
#
# + id="l9N5Wav3sLOj"
def cropping(img):
# create a 10 pixel border surrounding the stitched image
print("Cropping...")
stitched = cv2.copyMakeBorder(stitched, 10, 10, 10, 10,
cv2.BORDER_CONSTANT, (0, 0, 0))
# convert the stitched image to grayscale and threshold it
# such that all pixels greater than zero are set to 255
# (foreground) while all others remain 0 (background)
gray = cv2.cvtColor(stitched, cv2.COLOR_BGR2GRAY)
thresh = cv2.threshold(gray, 0, 255, cv2.THRESH_BINARY)[1]
# To get the intermediate results
plt.imshow(gray, cmap='gray')
plt.title("Gray Image")
plt.show()
plt.pause(0.2)
plt.imshow(thresh, cmap='gray')
plt.title("Threshold Image")
plt.show()
plt.pause(0.2)
# find all external contours in the threshold image then find
# the *largest* contour which will be the contour/outline of
# the stitched image
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
c = max(cnts, key=cv2.contourArea)
# allocate memory for the mask which will contain the
# rectangular bounding box of the stitched image region
mask = np.zeros(thresh.shape, dtype="uint8")
(x, y, w, h) = cv2.boundingRect(c)
h = int(0.95*h)
cv2.rectangle(mask, (x, y), (x + w, y + h), 255, -1)
plt.imshow(cv2.rectangle(mask, (x, y), (x + w, y + h), 255, -1), cmap='gray')
plt.title("Rectangular Mask")
plt.show()
plt.pause(0.2)
# create two copies of the mask: one to serve as our actual
# minimum rectangular region and another to serve as a counter
# for how many pixels need to be removed to form the minimum
# rectangular region
minRect = mask.copy()
sub = mask.copy()
# # keep looping until there are no non-zero pixels left in the
# # subtracted image
# while cv2.countNonZero(sub) > 2500:
# # erode the minimum rectangular mask and then subtract
# # the thresholded image from the minimum rectangular mask
# # so we can count if there are any non-zero pixels left
# minRect = cv2.erode(minRect, None)
# sub = cv2.subtract(minRect, thresh)
plt.imshow(minRect, cmap='gray')
plt.title('Minimized Image/Box')
plt.show()
plt.pause(0.2)
# find contours in the minimum rectangular mask and then
# extract the bounding box (x, y)-coordinates
cnts = cv2.findContours(minRect.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
c = max(cnts, key=cv2.contourArea)
(x, y, w, h) = cv2.boundingRect(c)
print(x, y, w, h)
# use the bounding box coordinates to extract the our final
# stitched image
stitched_cropped = stitched[100: 60 + 906, x:x + w]
# Display the output stitched image to our screen
return stitched_cropped
# + id="nBX7Sh3TrnlM"
def stitch_gt(img_arr, stitcher):
(status, stitched) = stitcher.stitch(img_arr)
if (status==cv2.STITCHER_OK):
# print('Panorama Generated')
stitched = cv2.cvtColor(stitched, cv2.COLOR_BGR2RGB)
# plt.imshow(stitched)
# plt.pause(0.2)
return stitched
else:
# print('Error Generating Panorama')
pass
# + id="k_ZPogxotaaS"
def run(left_video, mid_video, right_video, stitcher):
# Get information about the videos
n_frames = min( int(left_video.get(cv2.CAP_PROP_FRAME_COUNT)),
int(mid_video.get(cv2.CAP_PROP_FRAME_COUNT)),
int(right_video.get(cv2.CAP_PROP_FRAME_COUNT)) )
fps = int(left_video.get(cv2.CAP_PROP_FPS))
frames = []
count = -1
step_to_continue_from = 2499
fail_count = 0
for _ in tqdm.tqdm(np.arange(n_frames)):
count += 1
if (count>-1 and count<4000):
# Grab the frames from their respective video streams
ok, left = left_video.read()
_, mid = mid_video.read()
_, right = right_video.read()
if (count>step_to_continue_from) and ok:
left = cv2.cvtColor(left, cv2.COLOR_BGR2RGB)
mid = cv2.cvtColor(mid, cv2.COLOR_BGR2RGB)
right = cv2.cvtColor(right, cv2.COLOR_BGR2RGB)
# Stitch the frames together to form the panorama
stitched_frame = stitch_gt([left, mid, right], stitcher)
# No homography could not be computed
if stitched_frame is None:
print("[INFO]: Homography could not be computed!")
fail_count+=1
if fail_count>100:
print("Stopping the process!")
break
else:
continue
# Add frame to video
# stitched_frame = imutils.resize(stitched_frame, width=800)
stitched_frame = cv2.resize(stitched_frame, (1920, 720))
frames.append(stitched_frame)
elif (count==4000):
print("Finished Processing!")
cv2.destroyAllWindows()
return frames
# + id="v6j_xOmptx59"
# Defining the stitcher
stitcher = cv2.createStitcher(try_use_gpu=True)
# + id="RKn77u21t1iC"
# Defining the paths
left_video_in_path='/content/drive/My Drive/Colab Notebooks/Carla Dataset/City/Left.mp4'
mid_video_in_path='/content/drive/My Drive/Colab Notebooks/Carla Dataset/City/Front.mp4'
right_video_in_path='/content/drive/My Drive/Colab Notebooks/Carla Dataset/City/Right.mp4'
# + id="tDDHGldat6ju" colab={"base_uri": "https://localhost:8080/"} outputId="e8193147-6301-413a-de47-f01e20bf8a13"
# Set up video capture
left_video = cv2.VideoCapture(left_video_in_path)
mid_video = cv2.VideoCapture(mid_video_in_path)
right_video = cv2.VideoCapture(right_video_in_path)
print('[INFO]: {}, {} and {} loaded'.format( left_video_in_path.split('/')[-1],
mid_video_in_path.split('/')[-1],
right_video_in_path.split('/')[-1]) )
print('[INFO]: Video stitching starting....')
# The frames of the stitched images
frames = run(left_video, mid_video, right_video, stitcher)
# + id="BeDRM6Kd8esB" colab={"base_uri": "https://localhost:8080/"} outputId="38e95e0b-eb59-40c2-cf16-082cae78df8f"
print(len(frames))
# + id="_13jwN0Yubpa" colab={"base_uri": "https://localhost:8080/"} outputId="cd26d848-b135-4afc-c51f-d2c389f996ef"
print('[INFO]: Video stitching finished')
'''Writing via CV2'''
video_out_path='/content/drive/Shared drives/drive 4/Carla Dataset/Ground Truth'
for i in range(len(frames)):
cv2.imwrite(video_out_path+f"/{i+2462}"+".png", frames[i])
print("Creating : ",video_out_path,f"/{i+2462}",".png")
# + id="cxMHXCPnMoyX"
| Video_Stitching_Stitch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
from PIL import Image
import skvideo.io
import numpy as np
import torch
from torchvision import transforms
from tqdm import tqdm_notebook
from fast_neural_style.transformer_net import TransformerNet
from fast_neural_style.utils import recover_image, tensor_normalizer
# -
preprocess = transforms.Compose([
transforms.ToTensor(),
tensor_normalizer()
])
transformer = TransformerNet()
# ## Low Resolution GIF Animation
# Convert gif file to video file:
# ```
# ffmpeg -f gif -i cat.gif cat.mp4
# ```
skvideo.io.ffprobe("videos/cat.mp4")
transformer.load_state_dict(torch.load("../models/udine_10000.pth"))
frames = []
frames_orig = []
videogen = skvideo.io.vreader("videos/cat.mp4")
for frame in videogen:
frames_orig.append(Image.fromarray(frame))
frames.append(recover_image(transformer(
Variable(preprocess(frame).unsqueeze(0), volatile=True)).data.numpy())[0])
Image.fromarray(frames[3])
writer = skvideo.io.FFmpegWriter("cat.mp4")# tuple([len(frames)] + list(frames[0].shape)))
for frame in frames:
writer.writeFrame(frame)
writer.close()
# ## Higher Resolution Videos
skvideo.io.ffprobe("../videos/obama.mp4")
# Switch to GPU:
transformer.cuda()
BATCH_SIZE = 2
transformer.load_state_dict(torch.load("../models/mosaic_10000.pth"))
batch = []
videogen = skvideo.io.FFmpegReader("../videos/cod-2.mp4", {"-ss": "00:05:00", "-t": "00:01:00"})
writer = skvideo.io.FFmpegWriter("../videos/cod-clip-noise.mp4")
try:
with torch.no_grad():
for frame in tqdm_notebook(videogen.nextFrame()):
batch.append(preprocess(frame).unsqueeze(0))
if len(batch) == BATCH_SIZE:
for frame_out in recover_image(transformer(
torch.cat(batch, 0).cuda()).cpu().numpy()):
writer.writeFrame(frame_out)
batch = []
except RuntimeError as e:
print(e)
pass
writer.close()
transformer.load_state_dict(torch.load("../models/udine_10000_unstable.pth"))
batch = []
videogen = skvideo.io.FFmpegReader("../videos/cod-2.mp4", {"-ss": "00:05:00", "-t": "00:01:00"})
writer = skvideo.io.FFmpegWriter("../videos/cod-clip.mp4")
try:
with torch.no_grad():
for frame in tqdm_notebook(videogen.nextFrame()):
batch.append(preprocess(frame).unsqueeze(0))
if len(batch) == BATCH_SIZE:
for frame_out in recover_image(transformer(
torch.cat(batch, 0).cuda()).cpu().numpy()):
writer.writeFrame(frame_out)
batch = []
except RuntimeError as e:
pass
writer.close()
| vision/fast_neural_style_transfer/refs/03-Video-Creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preliminaries
#
# The `pandas` library allows the user several data structures for different data manipulation tasks:
# 1. Data storage through its `Series` and `DataFrame` data structures.
# 2. Data filtering using multiple methods from the package.
# 3. Reading data from many different file formats such as `csv`, `txt`, `xlsx`, ...
#
# Below we provide a brief overview of the `pandas` functionalities needed for these exercises. The complete documentation can be found on the [`pandas` website](https://pandas.pydata.org/).
#
# ## Pandas data structures
#
# ### Series
# The Pandas Series data structure is similar to a one-dimensional array. It can store any type of data. The values are mutable but the size not.
#
# To create `Series`, we call the `pd.Series()` method and pass an array. A `Series` may also be created from a numpy array.
# +
import pandas as pd
import numpy as np
first_series = pd.Series([1,10,100,1000])
print(first_series)
teams = np.array(['PSV','Ajax','Feyenoord','Twente'])
second_series = pd.Series(teams)
print('\n')
print(second_series)
# -
# ### DataFrame
# One can think of a `DataFrame` as a table with rows and columns (2D structure). The columns can be of a different type (as opposed to `numpy` arrays) and the size of the `DataFrame` is mutable.
#
# To create `DataFrame`, we call the `pd.DataFrame()` method and we can create it from scratch or we can convert a numpy array or a list into a `DataFrame`.
# +
# DataFrame from scratch
first_dataframe = pd.DataFrame({
"Position": [1, 2, 3, 4],
"Team": ['PSV','Ajax','Feyenoord','Twente'],
"GF": [80, 75, 75, 70],
"GA": [30, 25, 40, 60],
"Points": [79, 78, 70, 66]
})
print("From scratch: \n {} \n".format(first_dataframe))
# DataFrme from a list
data = [[1, 2, 3, 4], ['PSV','Ajax','Feyenoord','Twente'],
[80, 75, 75, 70], [30, 25, 40, 60], [79, 78, 70, 66]]
columns = ["Position", "Team", "GF", "GA", "Points"]
second_dataframe = pd.DataFrame(data, index=columns)
print("From list: \n {} \n".format(second_dataframe.T)) # the '.T' operator is explained later on
# DataFrame from numpy array
data = np.array([[1, 2, 3, 4], ['PSV','Ajax','Feyenoord','Twente'],
[80, 75, 75, 70], [30, 25, 40, 60], [79, 78, 70, 66]])
columns = ["Position", "Team", "GF", "GA", "Points"]
third_dataframe = pd.DataFrame(data.T, columns=columns)
print("From numpy array: \n {} \n".format(third_dataframe))
# -
# ### DataFrame attributes
# This section gives a quick overview of some of the `pandas.DataFrame` attributes such as `T`, `index`, `columns`, `iloc`, `loc`, `shape` and `values`.
# transpose the index and columns
print(third_dataframe.T)
# index makes reference to the row labels
print(third_dataframe.index)
# columns makes reference to the column labels
print(third_dataframe.columns)
# iloc allows to access the index by integer-location (e.g. all team names, which are in the second columm)
print(third_dataframe.iloc[:,1])
# loc allows to access the index by label(s)-location (e.g. all team names, which are in the "Team" columm)
print(third_dataframe.loc[0, 'Team'])
# shape returns a tuple with the DataFrame dimension, similar to numpy
print(third_dataframe.shape)
# values return a Numpy representation of the DataFrame data
print(third_dataframe.values)
# ### DataFrame methods
# This section gives a quick overview of some of the `pandas.DataFrame` methods such as `head`, `describe`, `concat`, `groupby`,`rename`, `filter`, `drop` and `isna`. To import data from CSV or MS Excel files, we can make use of `read_csv` and `read_excel`, respectively.
# print the first few rows in your dataset with head()
print(third_dataframe.head()) # In this case, it is not very useful because we don't have thousands of rows
# get the summary statistics of the DataFrame with describe()
print(third_dataframe.describe())
# +
# concatenate (join) DataFrame objects using concat()
# first, we will split the above DataFrame in two different ones
df_a = third_dataframe.loc[[0,1],:]
df_b = third_dataframe.loc[[2,3],:]
print(df_a)
print('\n')
print(df_b)
print('\n')
# now, we concatenate both datasets
df = pd.concat([df_a, df_b])
print(df)
# +
# group the data by certain variable via groupby()
# here, we have grouped the data by goals for, which in this case is 75
group = df.groupby('GF')
print(group.get_group('75'))
# -
# rename() helps you change the column or index names
print(df.rename(columns={'Position':'Pos','Team':'Club'}))
# build a subset of rows or columns of your dataset according to labels via filter()
# here, items refer to the variable names: 'Team' and 'Points'; to select columns, we specify axis=1
print(df.filter(items=['Team', 'Points'], axis=1))
# dropping some labels
print(df.drop(columns=['GF', 'GA']))
# +
# search for NA (not available) entries in the DataFrame
print(df.isna()) # No NA values
print('\n')
# create a pandas Series with a NA value
# the Series as W (winnin matches)
tmp = pd.Series([np.NaN, 25, 24, 19], name="W")
# concatenate the Series with the DataFrame
df = pd.concat([df,tmp], axis = 1)
print(df)
print('\n')
# again, check for NA entries
print(df.isna())
# -
# ## Dataset
#
# For this week exercises we will use a dataset from the Genomics of Drug Sensitivity in Cancer (GDSC) project (https://www.cancerrxgene.org/). In this study (['Iorio et al., Cell, 2016']()), 265 compounds were tested on 1001 cancer cell lines for which different types of -omics data (RNA expression, DNA methylation, Copy Number Alteration, DNA sequencing) are available. This is a valuable resource to look for biomarkers of drugs sensitivity in order to try to understand why cancer patients responds very differently to cancer drugs and find ways to assign the optimal treatment to each patient.
#
# For this exercise we will use a subset of the data, focusing the response to the drug YM155 (Sepantronium bromide) on four cancer types, for a total of 148 cancer cell lines.
#
# | ID | Cancer type |
# |-------------|----------------------------------|
# | COAD/READ | Colorectal adenocarcinoma |
# | NB | Neuroblastoma |
# | KIRC | Kidney renal clear cell carcinoma|
# | BRCA | Breast carcinoma |
#
# We will use the RNA expression data (RMA normalised). Only genes with high variability across cell lines (variance > 5, resulting in 238 genes) have been kept.
#
# Drugs have been tested at different concentration, measuring each time the viability of the cells. Drug sensitivity is measured using the natural log of the fitted IC50 metric, which is defined as the half maximal inhibitory concentration. A lower IC50 corresponds to a more sensitive cell line because a lower amount of drug is sufficient to have a strong response, while a higher IC50 corresponds to a more resistant cell line because more drug is needed for killing the cells.
#
# Based on the IC50 metric, cells can be classified as sensitive or resistant. The classification is done by computing the $z$-score across all cell lines in the GDSC for each drug, and considering as sensitive the ones with $z$-score < 0 and resistant the ones with $z$-score > 0.
#
# The dataset is originally provided as 3 files ([original source](https://www.sciencedirect.com/science/article/pii/S0092867416307462?via%3Dihub)) :
#
# `GDSC_RNA_expression.csv`: gene expression matrix with the cell lines in the rows (148) and the genes in the columns (238).
#
# `GDSC_drug_response.csv`: vector with the cell lines response to the drug YM155 in terms of log(IC50) and as classification in sensitive or resistant.
#
# `GDSC_metadata.csv`: metadata for the 148 cell lines including name, COSMIC ID and tumor type (using the classification from ['The Cancer Genome Atlas TCGA'](https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga))
#
# For convenience, we provide the data already curated.
#
# `RNA_expression_curated.csv`: [148 cell lines , 238 genes]
#
# `drug_response_curated.csv`: [148 cell lines , YM155 drug]
#
# The curated data cam be read as `pandas` `DataFrame`s in the following way:
# +
import pandas as pd
gene_expression = pd.read_csv("./data/RNA_expression_curated.csv", sep=',', header=0, index_col=0)
drug_response = pd.read_csv("./data/drug_response_curated.csv", sep=',', header=0, index_col=0)
# -
# You can use the `DataFrame`s directly as inputs to the the `sklearn` models. The advantage over using `numpy` arrays is that the variable are annotated, i.e. each input and output has a name.
# ## Tools
# The `scikit-learn` library provides the required tools for linear regression/classification and shrinkage, as well as for logistic regression.
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import LogisticRegression
# Note that the notation used for the hyperparameters in the `scikit-learn` library is different from the one used in the lecture. More specifically, in the lecture $\alpha$ is the tunable parameter to select the compromise between Ridge and Lasso. Whereas, `scikit-learn` library refers to `alpha` as the tunable parameter $\lambda$. Please check the documentation for more details.
# # Exercises
#
# ## Selection of the hyperparameter
#
# Implement cross-validation (using `sklearn.grid_search.GridSearchCV`) to select the `alpha` hyperparameter of `sklearn.linear_model.Lasso`. To match scikit-learn nomenclature we explicitly define $\alpha$ as the regularisation hyperparameter instead of the commonly used $\lambda$ in literature.
# +
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 1
import matplotlib.pyplot as plt
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.metrics import mean_squared_error, make_scorer
from sklearn.linear_model import Lasso
from sklearn.model_selection import GridSearchCV, train_test_split
from sklearn.pipeline import Pipeline
import pandas as pd
import numpy as np
# Initialize (pseudo-) random number generator with a seed
# for reproducibility of results
np.random.seed(40)
def do_gridsearch_alpha(X_train, y_train, X_test, y_test, alphas, crossvals=5, plot=True):
"""
given a train and a test dataset, fit and evaluate polynomial regression models with given degrees
Parameters:
X_train: 2D (Datapoints x features) numpy array
the datapoints on which to train the models
y_train: 1D (Targets) numpy array
the targets on which to train the models
X_test: 2D (Datapoints x features) numpy array
the datapoints on which to test the models
y_test: 1D (Targets) numpy array
the targets on which to test the models
alphas: 1D numpy array
alpha candidates
crossvals: int
amount of cross-validation folds
plot: bool
True for plotting the MSE scores, False for surpressing the plots
returns:
a GridSearchCV object, containing the fitted models for every degree polynomial
"""
# Define the functions for polynomial features and linear regression
Lassoreg = Lasso(max_iter=10000, tol=0.001) # Increased tolerance to stop "ConvergenceWarning"
scaler = StandardScaler()
# Define the pipeline, containing the polynomial transformation and the subsequent linear regression
model = Pipeline([
("scaler", scaler),
("lasso", Lassoreg)
])
# Define the hyperparameters for the Grid Search
params = {'lasso__alpha': alphas}
# Define a custom scoring system to evaluate the mean squared error (lower is better)
mse = make_scorer(mean_squared_error,greater_is_better=False)
# Define the Grid Search method
gridsearch = GridSearchCV(model, params, scoring=mse, cv=crossvals)
# Run Grid Search
gridsearch.fit(X_train, y_train)
# Predict targets for test set
y_pred = gridsearch.predict(X_test)
# Calculate MSE of the predictions on the test set
MSE_test_set = mean_squared_error(y_test, y_pred)
print(f'Mean squared error (test set): {MSE_test_set:.4}')
if plot:
# Plot MSE scores
plt.figure()
plt.title('Scores for every degree (per fold different colors)')
colors = ['b.', 'r.', 'g.', 'y.', 'k.', 'c.']
for i in range(crossvals):
q = gridsearch.cv_results_[f'split{i}_test_score']
plt.plot(np.log(params['lasso__alpha']), np.log(-q), colors[i % len(colors)], markersize=12)
plt.xlabel('log alpha')
plt.ylabel('log(-MSE)')
plt.legend(list(range(1,crossvals+1)))
# Plot average MSE scores
plt.figure()
#plt.errorbar(np.log(params['lasso__alpha']), np.log(-gridsearch.cv_results_[f'mean_test_score']), np.log(gridsearch.cv_results_[f'std_test_score']), linestyle='None', marker='.',markersize=12)
plt.plot(np.log(params['lasso__alpha']), np.log(-gridsearch.cv_results_[f'mean_test_score']), 'k.', markersize=12)
plt.title('Average scores of all folds')
plt.xlabel('log alpha')
plt.ylabel('log(-MSE)');
return gridsearch
# Load data from CSV-files
gene_expression = pd.read_csv("./data/RNA_expression_curated.csv", sep=',', header=0, index_col=0)
drug_response = pd.read_csv("./data/drug_response_curated.csv", sep=',', header=0, index_col=0)
# Split data into training and test set
X_train, X_test, y_train, y_test = train_test_split(gene_expression, drug_response)
# Define value range for alpha (optimal value to be found using Grid Search)
alphaspace = np.logspace(-3, 1, num=15)
# Find optimal value for alpha using Grid Search
grids = do_gridsearch_alpha(X_train, y_train, X_test, y_test, alphaspace, 5)
alphamin = grids.best_params_['lasso__alpha']
print(f"Optimal value for alpha: {alphamin}")
feature_index = np.nonzero(grids.best_estimator_.named_steps['lasso'].coef_)
print(f"The selected features are: {gene_expression.columns.values[feature_index]}")
# -
# ## Feature selection
#
# Look at the features selected using the hyperparameter which corresponds to the minimum cross-validation error.
#
# <p><font color='#770a0a'>Is the partition in training and validation sets playing a role in the selection of the hyperparameter? How will this affect the selection of the relevant features?</font></p>
#
# <p><font color='#770a0a'>Should the value of the intercept also be shrunk to zero with Lasso and Ridge regression? Motivate your answer.</font></p>
# ***ANSWER:***
#
# Partitioning in training and validation do play a role in selection of alpha as we can see in the different cross validation runs above. For different partitions (colors), the least MSE is achieved with a different value for alpha. Without bootstrapping, one of these partitions would have been selected, changing the selected alpha. Following a change in alpha due to partitioning, the amount of features (and which specific features) that are selected will change as well, with a higher alpha leading to less selected features. Different train test splits can be shown to lead to different selected features as well by disabling the random seed in the above function, and running the code a few times. This can be seen to lead to different selected features.
#
# The intercept should be seperate, as the intercept is merely a correction of the mean in a complex feature space and thus not relate to whether a feature has high enough impact and robustness to be taken into account in the final model. Also:
#
# $$
# \hat{\beta}^{\text {lasso }}=\underset{\beta}{\operatorname{argmin}}\left\{\sum_{i=1}^{N}\left(y_{i}-\beta_{0}-\sum_{j=1}^{p} x_{i j} \beta_{j}\right)^{2}+\alpha \sum_{j=1}^{p}\left|\beta_{j}\right|\right\}
# $$
#
# and:
# $$
# \hat{\beta}^{\text {ridge }}=\underset{\beta}{\operatorname{argmin}}\left\{\sum_{i=1}^{N}\left(y_{i}-\beta_{0}-\sum_{j=1}^{p} x_{i j} \beta_{j}\right)^{2}+\alpha \sum_{j=1}^{p} \beta_{j}^{2}\right\}
# $$
#
# Intercept $\beta_0$ is, for both ridge and lasso, outside the penalty term. So, the intercept is never shrunk to zero.
# Find the resulting coefficients for the best estimator (best alpha) -->
# Effect of training and val sets: rerun and check if below func is different
grids.best_estimator_.steps[1][1].coef_
# As we can see Lasso automatically seperates the intercept from the Regularization --> It is and should not be included
grids.best_estimator_.steps[1][1].intercept_
# ## Bias-variance
#
# Show the effect of the regularization on the parameter estimates in terms of bias and variance. For this you can repeat the optimization 100 times using bootstrap and visualise the profile of the Lasso regression coefficient over a grid of the hyperparameter, optionally including the variability as error bars.
#
# <p><font color='#770a0a'>Based on the visual analysis of the plot, what are your observation on bias and variance in relation to model complexity? Motivate your answer.</font></p>
#
# <!--
# ***ANSWER:*** <font color='red'> Nog een keuze maken voor manier van bias berekenen (hieronder), ook nog tekst aanpassen van bias </font>
#
# <font color='red'>Below we implemented a procedure where the data is resampled for set amount `n_bootstrap_runs` times. The data is split into testing and training data every run. For every $\alpha$ in `alphaspace` a model was fitted on the training set and evaluated on the test set. For each run and $\alpha$ we keep track of the following quantities:
#
# <font color='red'>1. The mean squared error (MSE) of the test predictions
# <font color='red'>2. The lasso regression coefficients
# <font color='red'>3. The variance of the test predictions
#
# <font color='red'>Underneath the implementation we visualize the obtained quantities and discuss the effect of (L1-) regularisation on the parameter estimates.
#
# <font color='#FF0000'>Troy: Ik heb het nu zonder grid search gedaan. GridSearchCV heeft geen ondersteuning voor het retrieven van alle model coëfficiënten, alleen van de beste. Vorig jaar heb ik dit opgelost door een clone van GridSearchCV te maken en die zelf aan te passen (uiteraard in eigen code). Nu bedenk ik me wel dat GridSearch misschien helemaal niet nodig, aangezien er verder niet gevraagd wordt cross-validation toe te passen.</font>
#
# <font color='red'>**Vincent**: Waarom precies de opbouw van die "bootstrap_test_train_split"-functie? Daarnaast: de vraag is "Show the effect of the regularization on the parameter estimates in terms of bias and variance". Het enige wat we nu hebben is variance tov $\alpha$, maar we hebben niets qua bias (en wel MSE). Omdat $
# \operatorname{MSE}(\hat{\theta}) = \operatorname{Var_\theta}\left(\hat{\theta}\right) +\operatorname{Bias}_{\theta}\left(\hat{\theta}\right)^{2}
# $, kunnen we dan niet de bias op die manier uitrekenen?
#
# -->
# +
from sklearn.base import clone
from sklearn.model_selection import ParameterGrid
from sklearn.utils import resample
from mlxtend.evaluate import bias_variance_decomp
def bootstrap_test_train_split(X, y, n_runs, random_state = 0):
"""
A generator that samples X, y with replacement (used in bootstrapping) and splits
it into a test and train dataset.
Parameters:
X: 2D (Datapoints x features) array or dataframe
the datapoints on which to train the models
y: 1D (Targets) array or dataframe
the targets on which to train the models
n_runs: int
amount of times the data is resampled
random_state: int
random state of random number generators
yiels:
X and y resampled and split into a train and test set according:
X_train, X_test, y_train, y_test
"""
for i in range(n_runs):
# Bootstrapping is sampling WITH replacement
Xsampled, ysampled = resample(X, y, random_state=random_state+i, replace=True)
yield train_test_split(Xsampled, ysampled, shuffle=False)
# Load data from CSV files
gene_expression = pd.read_csv("./data/RNA_expression_curated.csv", sep=',', header=0, index_col=0)
drug_response = pd.read_csv("./data/drug_response_curated.csv", sep=',', header=0, index_col=0)
# Define values for alpha and set bootstrap options
alphaspace = np.logspace(-5, 2, num=20)
n_bootstrap_runs = 100
# Define empty arrays to store results in
MSEs = np.zeros((n_bootstrap_runs, alphaspace.shape[0]))
MSEs_train = np.zeros((n_bootstrap_runs, alphaspace.shape[0]))
coefs = np.zeros((n_bootstrap_runs, alphaspace.shape[0], gene_expression.shape[1]))
# Select training and test set using bootstrap function
for i, (X_train, X_test, y_train, y_test) in \
enumerate(bootstrap_test_train_split(gene_expression, drug_response, n_bootstrap_runs,40)):
# Define Lasso regression method and scaling method
Lassoreg = Lasso(max_iter=2000, tol=0.003)
scaler = StandardScaler()
# Define the pipeline
model = Pipeline([
("scaler", scaler),
("lasso", Lassoreg)
])
# Define parameters (values for alpha)
params = {'lasso__alpha': alphaspace}
# Loop over all values for alpha
for j, param in enumerate(ParameterGrid(params)):
# Clone model (to be able to use variable 'model' again, set value for alpha and fit model to training data
model_ = clone(model)
model_.set_params(**param)
model_.fit(X_train, y_train)
# Evaluate model, calculate MSE, standard deviation and bias
y_pred_train = model_.predict(X_train)
y_pred = model_.predict(X_test)
MSEs[i, j] = mean_squared_error(y_test, y_pred)
MSEs_train[i, j] = mean_squared_error(y_train, y_pred_train)
coefs[i, j, ...] = model_.named_steps['lasso'].coef_
# +
# Figure 1: Mean Squared Error (MSE) for different values of alpha
plt.figure()
plt.errorbar(np.log10(alphaspace), np.mean(MSEs, axis=0), yerr=np.std(MSEs, axis=0),
elinewidth=2, fmt='o-', markersize=10, capsize=14, color='orangered')
plt.xlabel('$\\mathrm{log}_{10}\\,\\alpha$')
plt.ylabel('$\\mathrm{MSE}$')
plt.title('(1) $\\mathrm{MSE}$ $\mathrm{vs.}$ $\\alpha$')
plt.show()
# Figure 2: Training Mean Squared Error (MSE) for different values of alpha
plt.figure()
plt.errorbar(np.log10(alphaspace), np.mean(MSEs_train, axis=0), yerr=np.std(MSEs_train, axis=0),
elinewidth=2, fmt='o-', markersize=10, capsize=14, color='orangered')
plt.xlabel('$\\mathrm{log}_{10}\\,\\alpha$')
plt.ylabel('train $\\mathrm{MSE}$')
plt.title('(2) train $\\mathrm{MSE}$ $\mathrm{vs.}$ $\\alpha$')
plt.show()
# Figure 3: Beta coefficient for different values of alpha
plt.figure()
for i in range(gene_expression.shape[1]):
plt.plot(np.log10(alphaspace), np.mean(coefs[:, :, i], axis=0))
plt.xlabel('$\\mathrm{log}_{10}\\,\\alpha$')
plt.ylabel('Lasso regression coefficient $\\beta$')
plt.title('(4) Lasso regression coefficient estimates (averaged over runs)')
plt.show()
# -
#
# ***ANSWER:*** In figure 1 (above) it can be seen that there is an optimal value for $\alpha$. This can be explained following the feature selection of the Lasso regression. An increase in $\alpha$ causes more feature weights to go to zero, thus reducing the model complexity. This leads to a higher bias, since the model can not fully describe the data anymore. On the contrary, an increase in $\alpha$ can be seen to correspond to a decrease in variance. The higher $\alpha$-coefficient pushes the feature weights closer to 0, decreasing the deviation. The model complexity at this optimal $\alpha$ is still high enough to accurately describe the data due to a lower bias, while it is also low enough to get accurate estimates on the features due to a lower variance.
#
# The increase in bias is also visualized in figure 2, which shows the train MSE. With an increasing $\alpha$, the train MSE goes up. This shows that the model can't accurately describe the training data, due to the decreased model complexity. Before reaching the optimum $\alpha$, the bias is increasing, while the test MSE is still decreasing. From $MSE = bias(\hat{\theta})^2 + Var(\hat{\theta})$, it can be concluded that the variance is also decreasing.
#
#
# <!--
# <font color='red'>In the cell below, the MSE on the test set is plotted against the including standard deviation (visualized by vertical errorbars) against $\alpha$ over a space of 20 points between $10^{-5}$ and $10^2$ (Figure 1). As the $\alpha$-coefficient increases, more regression coefficients are pushed towards zero. Effectively, as $\alpha$ increases the bias is increased and variance decreased. For reference we include a plot (Figure 2) of an estimate of the variance (or rather standard deviation) and we observe that from ~$10^{-2}$ the variance decreases (up to a point where it becomes zero). With this large increase comes just a modest increase in MSE. And as the MSE is effectively the sum of variance and the squared bias, we deduce that the bias must be increasing. Thus here we can see clearly that the theory of the bias-variance tradeoff holds true for a lasso regression model (on this particular dataset). </font>
#
# <font color='red'>If we dive into specifics as to why the bias increases as alpha increases, we take a glance at the regularisation component of the objective function for lasso regression:
#
# <font color='red'>$
# \begin{align*}
# \alpha \sum_{i=1}^p |\beta_i|
# \end{align*}
# $
#
# <font color='red'>Due to the $L_1$-norm, coefficients are pushed to zero if $\alpha$ increases. We can visualize this (Figure 3) by plotting the lasso regression coefficient estimates against $\alpha$ (averaged over the bootstrap runs).
#
# <font color='red'>It can be observed nicely from the figure that the lasso regression coefficients are pushed towards zero as $\alpha$ increases, up to a point where the coefficients all become zero. Naturally this will lead to zero variance of the model predictions, as the model will be of the form $\hat{y}=\beta_0$, where $\beta_0$ is the bias term. </font>
#
# <font color='red'>**Vincent:** Is dit stuk relevant?</font>
# -->
# ## Logistic regression
#
# <p><font color='#770a0a'>Write the expression of the objective function for the penalized logistic regression with $L_1$ and $L_2$ regularisation (as in Elastic net).</font></p>
# ***ANSWER:***
#
# The objective function $\mathcal{L}$ of logistic regression can be characterized by:
# \begin{align*}
# \mathcal{L}(\mathbf{\beta})=\sum_{i=1}^{N} [y_i(\beta_0 + \beta^T x_i) - \log(1+e^{(\beta_0+\beta^T x_i)}]
# \end{align*}
#
# The $L_1$ and $L_2$ regularisation terms can be added to the objective function:
#
# \begin{align*}
# \mathcal{L}(\mathbf{\beta})=\sum_{i=1}^{N} [y_i(\beta_0 + \beta^T x_i) - \log(1+e^{(\beta_0+\beta^T x_i)}] - \sum_{j=1}^{p} \left(\alpha_1\beta_j^2 + \alpha_2 |\beta_j|\right)
# \end{align*}
#
#
# Here $\alpha_1$, $\alpha_2$ are the regularisation parameters for $L_1$ and $L_2$ regularisation respectively. Alternatively, a trade-off parameter $V \in [0, 1]$ can be introduced:
#
# \begin{align*}
# \mathcal{L}(\mathbf{\beta})=\sum_{i=1}^{N} [y_i(\beta_0 + \beta^T x_i) - \log(1+e^{(\beta_0+\beta^T x_i)}] - \alpha \sum_{j=1}^{p} (V\beta_j^2 + (1-V) |\beta_j|)
# \end{align*}
#
# When $V=0$, Lasso regulatisation is used; when $V=1$, Ridge regularisation is used.
| practicals/week_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib import rcParams
# figure size in inches
rcParams['figure.figsize'] = 15,15
data = pd.read_excel('Data_Train.xlsx')
data_submit = pd.read_excel('Test_set.xlsx')
data.head(15)
data.info()
data_submit.info()
# One missing value, not that bad, we will drop it
data = data.dropna()
# # Data Transformation
# ## Date of The Journey
data.Date_of_Journey.unique().tolist()
# As you can see, there is only one year. I know that flight ticket are influenced by the period in the year, so I will keep only the month and the day. Plus as far as I'm concerned, flight tickets are more expensive during the week-end, so I will add a feature the numero of the day in week, and the name of the day
data['Date_of_Journey'] = pd.to_datetime(data['Date_of_Journey'])
data_submit['Date_of_Journey'] = pd.to_datetime(data_submit['Date_of_Journey'])
data['Month'] = data['Date_of_Journey'].dt.month
data['Day'] = data['Date_of_Journey'].dt.day
data['dayofweek_num'] = data['Date_of_Journey'].dt.dayofweek
data['dayofweek_name'] = data['Date_of_Journey'].dt.day_name()
data = data.drop(['Date_of_Journey'], axis=1)
data_submit['Month'] = data_submit['Date_of_Journey'].dt.month
data_submit['Day'] = data_submit['Date_of_Journey'].dt.day
data_submit['dayofweek_num'] = data_submit['Date_of_Journey'].dt.dayofweek
data_submit['dayofweek_name'] = data_submit['Date_of_Journey'].dt.day_name()
data_submit = data_submit.drop(['Date_of_Journey'], axis=1)
data.head(10)
# ## Route & Total Stops
data.Total_Stops.value_counts()
# As we can see, there is at max 4 stops
def check_and_return_stop(x,i):
x = x.split(' → ')
if len(x) < 2+i:
return float('nan')
else:
return x[i]
def transform_route_feature(df):
df['stopover_1'] = df['Route'].apply(check_and_return_stop, args=(1,))
df['stopover_2'] = df['Route'].apply(check_and_return_stop, args=(2,))
df['stopover_3'] = df['Route'].apply(check_and_return_stop, args=(3,))
df['stopover_4'] = df['Route'].apply(check_and_return_stop, args=(4,))
return df
data = transform_route_feature(data)
data_submit = transform_route_feature(data_submit)
data = data.drop(['Route'], axis=1)
data_submit = data_submit.drop(['Route'], axis=1)
def return_total_stop(x):
x = x.split()
if len(x) < 2:
return int(0)
else:
return int(x[0])
data['total_stop'] = data['Total_Stops'].apply(return_total_stop)
data_submit['total_stop'] = data_submit['Total_Stops'].apply(return_total_stop)
data = data.drop(['Total_Stops'], axis=1)
data_submit = data_submit.drop(['Total_Stops'], axis=1)
data.head()
# ## Dep Time & Arrival Time
def transform_dep_time(df):
df['Dep_Time'] = pd.to_datetime(df['Dep_Time'])
df['Hour_Dep'] = df['Dep_Time'].dt.hour
return df
data = transform_dep_time(data)
data_submit = transform_dep_time(data_submit)
data = data.drop(['Dep_Time'], axis=1)
data_submit = data_submit.drop(['Dep_Time'], axis=1)
data.head()
def transform_arr_time(df):
df['Arrival_Time'] = pd.to_datetime(df['Arrival_Time'])
df['Hour_Arr'] = df['Arrival_Time'].dt.hour
return df
data = transform_arr_time(data)
data_submit = transform_arr_time(data_submit)
data = data.drop(['Arrival_Time'], axis=1)
data_submit = data_submit.drop(['Arrival_Time'], axis=1)
data.head()
# I think that going at 12h and arriving at 18h is more expensive than going at 2h and arriving at 8h, that's why I decided to took the hour of the departure time and arrival time
# ## Duration
# This is one of the most feature since the price is highly dependent on it (the longer it is, the most expensive it is)
def extract_duration_hours(x):
x = x.split()
if len(x) < 2:
if 'm' in x[0]:
return float('0.' + x[0].split('m')[0])
return float(x[0].split('h')[0])
else:
return float(x[0].split('h')[0] + '.' + x[1].split('m')[0])
def duration(df):
df['Duration'] = df['Duration'].apply(extract_duration_hours)
return df
data = duration(data)
data_submit = duration(data_submit)
data.head()
## Stuff do the Additional_Info Feature
data = data.replace('No Info', 'No info')
data_submit = data_submit.replace('No Info', 'No info')
data.Additional_Info.unique().tolist()
import copy
data_train = copy.deepcopy(data)
data_submit_ = copy.deepcopy(data_submit)
# # OPTION 1 : One Hot Enconding
for col in ['Airline', 'Source', 'Destination', 'dayofweek_name']:
data = pd.concat([data, pd.get_dummies(data[col], prefix=col)], axis=1)
data = data.drop([col], axis=1)
data = pd.concat([data, pd.get_dummies(data['Additional_Info'], prefix='Info', drop_first=True)], axis=1)
data = data.drop(['Additional_Info'], axis=1)
data.head()
for col in ['Airline', 'Source', 'Destination', 'dayofweek_name']:
data_submit = pd.concat([data_submit, pd.get_dummies(data_submit[col], prefix=col)], axis=1)
data_submit = data_submit.drop([col], axis=1)
data_submit = pd.concat([data_submit, pd.get_dummies(data_submit['Additional_Info'], prefix='Info', drop_first=True)], axis=1)
data_submit = data_submit.drop(['Additional_Info'], axis=1)
data_submit.head()
[x for x in data.columns.tolist() if x not in data_submit.columns.tolist()]
data_submit[[ 'Airline_Trujet',
'Info_1 Short layover',
'Info_2 Long layover',
'Info_Red-eye flight']] = 0
data_submit.head()
# # Modelisation
# ## Baseline
from pycaret.regression import *
reg1 = setup(data, target = 'Price', session_id = 123,
ignore_low_variance = True, combine_rare_levels = True, silent=True)
compare_models(exclude = ['br', 'en', 'lar', 'llar', 'omp', 'ard', 'par', 'ransac', 'tr', 'mlp'])
# Take the 3 bests models
catb = create_model('catboost', verbose=False)
xgb = create_model('xgboost', verbose=False)
rf = create_model('rf', verbose=False)
# Blend all the 3 bests models
blend_all = blend_models(estimator_list = [catb, xgb, rf])
# Finalise models and make predictions
final_blender = finalize_model(blend_all)
predictions = predict_model(final_blender, data = data_submit)
predictions.head()
predictions['Price'] = predictions['Label']
predictions[['Price']].to_excel('submissions.xlsx', index=False)
# **We got a score of 0.9449 (rank : 85), I think we can do better than that**
# # OPTION 2 : Label Encode + HotEncode ?
# We will try the label encoding technique and hot encoding
all_ = pd.concat([data_train, data_submit_])
# plot
sns.set_style('ticks')
fig, ax = plt.subplots()
# the size of A4 paper
fig.set_size_inches(12, 12)
sns.boxplot(x="Additional_Info", y="Price", data=all_, ax=ax)
plt.xticks(rotation=90)
sns.despine()
## I will associate label according to the distribution of the price for each label in Additional Info
all_ = all_.replace('No info', float(0))
all_ = all_.replace('In-flight meal not included', -float(1)) # Less expensive
all_ = all_.replace('No check-in baggage included', -float(2)) # Mostly less expensive
all_ = all_.replace('1 Short layover', float(0.5)) # Since we only one data point
all_ = all_.replace('2 Long layover', float(0.5)) # Since we only have one data point
all_ = all_.replace('Change airports', float(1))
all_ = all_.replace('1 Long layover', float(1.5))
all_ = all_.replace('Red-eye flight', float(0)) # Since we only have one data point
all_ = all_.replace('Business class', float(3))
all_.Additional_Info.value_counts()
# ## Source / Destination
test = copy.deepcopy(data_train)
test['Comb_Source_Dest'] = test.apply(lambda x: x['Source'] + '/' + x['Destination'], axis=1)
# plot
sns.set_style('ticks')
fig, ax = plt.subplots()
# the size of A4 paper
fig.set_size_inches(12, 12)
sns.boxplot(x="Comb_Source_Dest", y="Price", data=test[test['total_stop']==0], ax=ax)
plt.xticks(rotation=90)
sns.despine()
# plot
sns.set_style('ticks')
fig, ax = plt.subplots()
# the size of A4 paper
fig.set_size_inches(12, 12)
sns.boxplot(x="Comb_Source_Dest", y="Price", data=test[test['total_stop']==1], ax=ax)
plt.xticks(rotation=90)
sns.despine()
# plot
sns.set_style('ticks')
fig, ax = plt.subplots()
# the size of A4 paper
fig.set_size_inches(12, 12)
sns.boxplot(x="Comb_Source_Dest", y="Price", data=test[test['total_stop']==3], ax=ax)
plt.xticks(rotation=90)
sns.despine()
# We can notice that the route (Source-Destination) can influence the price (taking into account the number total stop)
# I think I will hot encode this feature, and drop Source & Destination
all_['Comb_Source_Dest'] = all_.apply(lambda x: x['Source'] + '/' + x['Destination'], axis=1)
all_ = pd.concat([all_, pd.get_dummies(all_['Comb_Source_Dest'], prefix='CSD')], axis=1)
all_ = all_.drop(['Comb_Source_Dest', 'Source', 'Destination'], axis=1)
all_.head()
# ## AirLine
# plot
sns.set_style('ticks')
fig, ax = plt.subplots()
# the size of A4 paper
fig.set_size_inches(12, 12)
sns.boxplot(x="Airline", y="Price", data=all_, ax=ax)
plt.xticks(rotation=90)
sns.despine()
# I can give a rank to the airline company.
## I will associate label according to the distribution of the price for each label in Additional Info
all_ = all_.replace('Trujet', float(0))
all_ = all_.replace('SpiceJet', float(1))
all_ = all_.replace('IndiGo', float(2))
all_ = all_.replace('Air Asia', float(2.5))
all_ = all_.replace('GoAir', float(3))
all_ = all_.replace('Vistara', float(4))
all_ = all_.replace('Vistara Premium economy', float(4.5))
all_ = all_.replace('Air India', float(5))
all_ = all_.replace('Jet Airways', float(5.5))
all_ = all_.replace('Multiple carriers', float(6))
all_ = all_.replace('Multiple carriers Premium economy', float(6.5))
all_ = all_.replace('Jet Airways Business', float(8))
all_.Airline.value_counts()
all_.head()
# Im gonna hot encode month and dayweek_num
all_ = pd.concat([all_, pd.get_dummies(all_['Month'], prefix='Month')], axis=1)
all_ = all_.drop(['Month'], axis=1)
all_ = pd.concat([all_, pd.get_dummies(all_['dayofweek_name'], prefix='dname')], axis=1)
all_ = all_.drop(['dayofweek_name', 'dayofweek_num', 'Day'], axis=1)
all_.head()
# ## Pycaret With Advanced Settings
# Well, Pycaret handle missing value by computing the most common value, This is bad for the features such as stopover_1 etc ... I will fill NA with None
all_ = all_.fillna(value='None')
all_.head()
train = all_[:len(data_train)]
test = all_[len(data_train):].drop('Price',axis=1)
from pycaret.regression import *
exp_reg = setup(train, target='Price', session_id = 123, silent=True, normalize = True, normalize_method = 'zscore',
transformation = True, polynomial_features = True,
trigonometry_features = True, remove_outliers = True, outliers_threshold = 0.01, ignore_low_variance=True
)
compare_models(exclude = ['br', 'en', 'par', 'omp', 'llar', 'knn', 'lr', 'lasso', 'ridge', 'ada', 'lar'])
# Take the 4 bests models
catb = create_model('catboost', verbose=False)
xgb = create_model('xgboost', verbose=False)
rf = create_model('rf', verbose=False)
et = create_model('et', verbose=False)
# Blend all the 4 bests models
blend_all = blend_models(estimator_list = [catb, xgb, rf, et])
# Finalise models and make predictions
final_blender = finalize_model(blend_all)
predictions = predict_model(final_blender, data = test)
predictions.head()
predictions['Price'] = predictions['Label']
predictions[['Price']].to_excel('submission_2.xlsx', index=False)
# **The score is 0.9477 which is a (very) little better than the previous one (0.9449) , Nevertheless, My old rank was 85 and now it's 58 which is an improvment.**
| Flights_Price_Predictions/Flight_Price_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Network scanning
#
# Experiment setup:
#
# - select one node and setup AP on one interface
# - perform scanning from all nodes on all interfaces
# - repeat for all nodes and all interfaces
# +
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from IPython import display
import networkx as nx
from math import sqrt
from nxpd import draw
from nxpd import nxpdParams
nxpdParams['show'] = 'ipynb'
import bokeh.plotting as bkh
bkh.output_notebook()
# -
# Load scan data from `.csv`
# source = '../data/scan.csv'
source = '../data/scan_2017-08-04T13:00:46.185926.csv'
df = pd.DataFrame.from_csv(source)
df['signal'] = df['signal'].str.strip(' dBm').astype('float')
# df['label'] = df.apply(lambda x: '\n'.join([x['sta_dev'], str(x['signal'])]), axis=1)
df.head()
# Filter for own AP's (di
df_twist = df[df['ssid'] == 'twist-test']
df_twist = df_twist[df_twist['sta'].str.startswith('nuc')]
df_twist = df_twist[df_twist['ap'].str.startswith('nuc')]
df_twist
for freq in df_twist['freq'].unique():
df = df_twist[freq == df_twist['freq']]
G = nx.from_pandas_dataframe(
df,
'ap', 'sta',
'signal',
create_using=nx.DiGraph())
print('Frequency {}'.format(freq))
P = nx.nx_pydot.to_pydot(G)
display.display(display.Image(P.create_png()))
for freq in df_twist['freq'].unique():
plt.figure()
df = df_twist[freq == df_twist['freq']]
signal = df.pivot_table(
values='signal', index=['ap', 'ap_dev'], columns=['sta', 'sta_dev'])
sns.heatmap(signal);
plt.title('Frequency: {}'.format(freq))
| analysis/Scanning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 7.2 梯度下降和随机梯度下降
# %matplotlib inline
import numpy as np
import torch
import math
import sys
sys.path.append("..")
import d2lzh_pytorch as d2l
# ## 7.2.1 一维梯度下降
# +
def gd(eta):
x = 10
results = [x]
for i in range(10):
x -= eta * 2 * x # f(x) = x * x的导数为f'(x) = 2 * x
results.append(x)
print('epoch 10, x:', x)
return results
res = gd(0.2)
# +
def show_trace(res):
n = max(abs(min(res)), abs(max(res)), 10)
f_line = np.arange(-n, n, 0.1)
d2l.set_figsize((4,3))
d2l.plt.plot(f_line, [x**2 for x in f_line])
d2l.plt.plot(res, [x * x for x in res], '-o')
d2l.plt.xlabel('x')
d2l.plt.ylabel('f(x)')
show_trace(res)
# -
# ## 7.2.2 学习率
show_trace(gd(0.05))
show_trace(gd(0.49))
# ## 7.2.3 多维梯度下降
# +
def train_2d(trainer): # 本函数将保存在d2lzh_pytorch包中方便以后使用
x1, x2, s1, s2 = -5, -2, 0, 0 # s1和s2是自变量状态,本章后续几节会使用
results = [(x1, x2)]
for i in range(20):
x1, x2, s1, s2 = trainer(x1, x2, s1, s2)
results.append((x1, x2))
print('epoch %d, x1 %f, x2 %f' % (i + 1, x1, x2))
return results
def show_trace_2d(f, results): # 本函数将保存在d2lzh_pytorch包中方便以后使用
d2l.plt.plot(*zip(*results), '-o', color='#ff7f0e')
x1, x2 = np.meshgrid(np.arange(-5.5, 1.0, 0.1), np.arange(-3.0, 1.0, 0.1))
d2l.plt.contour(x1, x2, f(x1, x2), colors='#1f77b4')
print(x1)
d2l.plt.xlabel('x1')
d2l.plt.ylabel('x2')
# +
eta = 0.1
def f_2d(x1, x2): # 目标函数
return x1 ** 2 + 2 * x2 ** 2
def gd_2d(x1, x2, s1, s2):
return (x1 - eta * 2 * x1, x2 - eta * 4 * x2, 0, 0)
show_trace_2d(f_2d, train_2d(gd_2d))
# -
# ## 7.2.4 随机梯度下降
# +
def sgd_2d(x1, x2, s1, s2):
return (x1 - eta * (2 * x1 + np.random.normal(0.1)),
x2 - eta * (4 * x2 + np.random.normal(0.1)), 0, 0)
show_trace_2d(f_2d, train_2d(sgd_2d))
# +
from mpl_toolkits import mplot3d # 三维画图
x, y = np.mgrid[-1: 1: 31j, -1: 1: 31j]
z = x**2 - y**2
ax = d2l.plt.figure().add_subplot(111, projection='3d')
ax.plot_wireframe(x, y, z, **{'rstride': 2, 'cstride': 2})
ax.plot([0], [0], [0], 'rx')
ticks = [-1, 0, 1]
d2l.plt.xticks(ticks)
d2l.plt.yticks(ticks)
ax.set_zticks(ticks)
d2l.plt.xlabel('x')
d2l.plt.ylabel('y')
# -
| code/chapter07_optimization/7.2_gd-sgd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.preprocessing import StandardScaler, MinMaxScaler, LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
train_df = pd.read_csv(Path('Resources/2019loans.csv'))
test_df = pd.read_csv(Path('Resources/2020Q1loans.csv'))
# # Predictions:
# - I think the Random Forest Model will be a better model because its utilizing a bunch of random samples and averaging the results together. This allows it to better account for the 'noise' within the data. The Logistic Regression Model assumes there is a linear relationship which may not yeild accurate predictions.
# Convert categorical data to numeric and separate target feature for training data
X_train_df= train_df.drop(['loan_status', 'Unnamed: 0'], axis=1)
X_train_df.head()
X_train= pd.get_dummies(X_train_df)
X_train.head()
y_train= train_df['loan_status']
y_train.head()
# Convert categorical data to numeric and separate target feature for testing data
X_test_df= test_df.drop(['loan_status', 'Unnamed: 0'], axis=1)
X_test_df.head()
X_test= pd.get_dummies(X_test_df)
X_test.head()
y_test= test_df['loan_status']
y_test.head()
# add missing dummy variables to testing set
for column in X_train:
if column not in X_test:
X_test[column] = 0
print(column)
X_test.head()
# X_test['debt_settlement_flag_Y'] = 0
for index, value in enumerate(X_test['debt_settlement_flag_N']):
if value == 0:
X_test.at[index, 'debt_settlement_flag_Y'] = 1
else:
pass
X_test.head()
# Train the Logistic Regression model on the unscaled data and print the model score
clf = LogisticRegression()
clf.fit(X_train, y_train)
print(f"Training Data Score: {clf.score(X_train, y_train)}")
print(f"Testing Data Score: {clf.score(X_test, y_test)}")
# Train a Random Forest Classifier model and print the model score
rfc = RandomForestClassifier(random_state=1, n_estimators=500)
rfc.fit(X_train, y_train)
print(f' Random Forest Training Score: {rfc.score(X_train, y_train)}')
print(f' Random Forest Testing Score: {rfc.score(X_test, y_test)}')
# # Results 1:
# - The Random Forest model scored higher for both the train and test data
# Scale the data
scaler = StandardScaler().fit(X_train)
X_train_scaled=scaler.transform(X_train)
X_train_scaled
X_test_scaled=scaler.transform(X_test)
X_test_scaled
# Train the Logistic Regression model on the scaled data and print the model score
clf_scaled = LogisticRegression()
clf_scaled.fit(X_train_scaled, y_train)
print(f' Scaled Logistic Regression Score: {clf_scaled.score(X_train_scaled, y_train)}')
print(f' Scaled Logistic Regression Score: {clf_scaled.score(X_test_scaled, y_test)}')
# Train a Random Forest Classifier model on the scaled data and print the model score
rfc_scaled = RandomForestClassifier(random_state=1, n_estimators=500)
rfc_scaled.fit(X_train_scaled, y_train)
print(f' Scaled Random Forest Train Score: {rfc_scaled.score(X_train_scaled, y_train)}')
print(f' Scaled Random Forest Test Score: {rfc_scaled.score(X_test_scaled, y_test)}')
# # Results 2:
# - The Logistical regression model out performed the unscaled model and both Random Forest Models for the test data.
# - Scaling the model allowed the data to relate to itself in a way that works better in a Logistical Regression model.
# - Random Forest model results were unchanged after scaling the data. This is likely due to the random nature of the sampling/aggregating of data.
| Credit Risk Evaluator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementing Neural Network from Scratch.
# Neural Networks are really powerful algorithms used for classification. <br>
# Dataset = Iris_Dataset <br>
# Link = http://scikit-learn.org/stable/auto_examples/datasets/plot_iris_dataset.html <br>
#
# ### Import required libraries
from sklearn import datasets #for dataset
import numpy as np #for maths
import matplotlib.pyplot as plt #for plotting
# ### Get Dataset
# +
iris = datasets.load_iris() #load the dataset
data = iris.data #get features
target = iris.target #get labels
shape = data.shape #shape of data
#convert into numpy array
data = np.array(data).reshape(shape[0],shape[1])
target = np.array(target).reshape(shape[0],1)
#print shape
print("Data Shape = {}".format(data.shape))
print("Target Shape = {}".format(target.shape))
print('Classes : {}'.format(np.unique(target)))
print('Sample data : {} , Target = {}'.format(data[70],target[70]))
# -
# ### Define Parameters and Hyperparameters
#
#
#
#
#
# One hidden layer Neural Network.
# 
# Input Units = 4 <br>
# Hidden Units = 8 <br>
# Output Units = 3 <br>
# +
#HYPERPARAMETERS
#num of target labels
num_classes = len(np.unique(target))
#define layer_neurons
input_units = 4 #neurons in input layer
hidden_units = 8 #neurons in hidden layer
output_units = 3 #neurons in output layer
#define hyper-parameters
learning_rate = 0.03
#regularization parameter
beta = 0.00001
#num of iterations
iters = 4001
# -
# ### Dimesions of Parameters
# Shape of layer1_weights (Wxh) = (4,8) <br>
# Shape of layer1_biasess (Bh) = (8,1) <br>
# Shape of layer2_weights (Why) = (8,3) <br>
# Shape of layer2_biasess (By) = (3,1) <br>
# +
#PARAMETERS
#initialize parameters i.e weights
def initialize_parameters():
#initial values should have zero mean and 0.1 standard deviation
mean = 0 #mean of parameters
std = 0.03 #standard deviation
layer1_weights = np.random.normal(mean,std,(input_units,hidden_units))
layer1_biases = np.ones((hidden_units,1))
layer2_weights = np.random.normal(mean,std,(hidden_units,output_units))
layer2_biases = np.ones((output_units,1))
parameters = dict()
parameters['layer1_weights'] = layer1_weights
parameters['layer1_biases'] = layer1_biases
parameters['layer2_weights'] = layer2_weights
parameters['layer2_biases'] = layer2_biases
return parameters
# -
# ### Activation Function
#
# **Sigmoid**
#
# 
#
# +
#activation function
def sigmoid(X):
return 1/(1+np.exp((-1)*X))
#softmax function for output
def softmax(X):
exp_X = np.exp(X)
exp_X_sum = np.sum(exp_X,axis=1).reshape(-1,1)
exp_X = (exp_X/exp_X_sum)
return exp_X
# -
# ### Define Utility Functions
# #### 1. Forward Propagation
# ---- Logits = matmul(X,Wxh) + Bh <br>
# ---- A = sigmoid(logits) <br>
# ---- logits = matmul(A,Why) + By <br>
# ---- output = softmax(logits) <br>
#
# Store output and A in cache to use it in backward propagation <br>
#
# #### 2. Backward Propagation
# ---- Error_output = output - train_labels <br>
# ---- Error_activation = (matmul(error_output,Why.T))(A)(1-A) <br>
# ---- dWhy = (matmul(A.T,error_output))/m <br>
# ---- dWxh = (matmul(train_dataset.T,error_activation))/m <br>
#
# m = len(train_dataset) <br>
# Store derivatives in derivatives dict
#
# #### 3. Update Parameters
# ---- Wxh = Wxh - learning_rate(dWxh + beta*Wxh) <br>
# ---- Why = Why - learning_rate(dWhy + beta*Why) <br>
#
# #### 4. Calculate Loss and Accuracy
# ---- Loss = (-1(Y log(prediction)) + (1-Y) (log(1-predictions))) + beta * (sum(Wxh^2) + sum(Why^2)))/m <br>
# ---- Accuracy = sum(Y==predictions)/m
# +
#forward propagation
def forward_propagation(train_dataset,parameters):
cache = dict() #to store the intermediate values for backward propagation
m = len(train_dataset) #number of training examples
#get the parameters
layer1_weights = parameters['layer1_weights']
layer1_biases = parameters['layer1_biases']
layer2_weights = parameters['layer2_weights']
layer2_biases = parameters['layer2_biases']
#forward prop
logits = np.matmul(train_dataset,layer1_weights) + layer1_biases.T
activation1 = np.array(sigmoid(logits)).reshape(m,hidden_units)
activation2 = np.array(np.matmul(activation1,layer2_weights) + layer2_biases.T).reshape(m,output_units)
output = np.array(softmax(activation2)).reshape(m,num_classes)
#fill in the cache
cache['output'] = output
cache['activation1'] = activation1
return cache,output
#backward propagation
def backward_propagation(train_dataset,train_labels,parameters,cache):
derivatives = dict() #to store the derivatives
#get stuff from cache
output = cache['output']
activation1 = cache['activation1']
#get parameters
layer1_weights = parameters['layer1_weights']
layer2_weights = parameters['layer2_weights']
#calculate errors
error_output = output - train_labels
error_activation1 = np.matmul(error_output,layer2_weights.T)
error_activation1 = np.multiply(error_activation1,activation1)
error_activation1 = np.multiply(error_activation1,1-activation1)
#calculate partial derivatives
partial_derivatives2 = np.matmul(activation1.T,error_output)/len(train_dataset)
partial_derivatives1 = np.matmul(train_dataset.T,error_activation1)/len(train_dataset)
#store the derivatives
derivatives['partial_derivatives1'] = partial_derivatives1
derivatives['partial_derivatives2'] = partial_derivatives2
return derivatives
#update the parameters
def update_parameters(derivatives,parameters):
#get the parameters
layer1_weights = parameters['layer1_weights']
layer2_weights = parameters['layer2_weights']
#get the derivatives
partial_derivatives1 = derivatives['partial_derivatives1']
partial_derivatives2 = derivatives['partial_derivatives2']
#update the derivatives
layer1_weights -= (learning_rate*(partial_derivatives1 + beta*layer1_weights))
layer2_weights -= (learning_rate*(partial_derivatives2 + beta*layer2_weights))
#update the dict
parameters['layer1_weights'] = layer1_weights
parameters['layer2_weights'] = layer2_weights
return parameters
#calculate the loss and accuracy
def cal_loss_accuray(train_labels,predictions,parameters):
#get the parameters
layer1_weights = parameters['layer1_weights']
layer2_weights = parameters['layer2_weights']
#cal loss and accuracy
loss = -1*np.sum(np.multiply(np.log(predictions),train_labels) + np.multiply(np.log(1-predictions),(1-train_labels)))/len(train_labels) + np.sum(layer1_weights**2)*beta/len(train_labels) + np.sum(layer2_weights**2)*beta/len(train_labels)
accuracy = np.sum(np.argmax(train_labels,axis=1)==np.argmax(predictions,axis=1))
accuracy /= len(train_dataset)
return loss,accuracy
# -
# ### Train Function
#
# 1. Initialize Parameters
# 2. Forward Propagation
# 3. Backward Propagation
# 4. Calculate Loss and Accuracy
# 5. Update the parameters
#
# Repeat the steps 2-5 for the given number of iterations
# +
#Implementation of 3 layer Neural Network
#training function
def train(train_dataset,train_labels,iters=2):
#To store loss after every iteration.
J = []
#WEIGHTS
global layer1_weights
global layer1_biases
global layer2_weights
global layer2_biases
#initialize the parameters
parameters = initialize_parameters()
layer1_weights = parameters['layer1_weights']
layer1_biases = parameters['layer1_biases']
layer2_weights = parameters['layer2_weights']
layer2_biases = parameters['layer2_biases']
#to store final predictons after training
final_output = []
for j in range(iters):
#forward propagation
cache,output = forward_propagation(train_dataset,parameters)
#backward propagation
derivatives = backward_propagation(train_dataset,train_labels,parameters,cache)
#calculate the loss and accuracy
loss,accuracy = cal_loss_accuray(train_labels,output,parameters)
#update the parameters
parameters = update_parameters(derivatives,parameters)
#append loss
J.append(loss)
#update final output
final_output = output
#print accuracy and loss
if(j%500==0):
print("Step %d"%j)
print("Loss %f"%loss)
print("Accuracy %f%%"%(accuracy*100))
return J,final_output
# +
#shuffle the dataset
z = list(zip(data,target))
np.random.shuffle(z)
data,target = zip(*z)
#make train_dataset and train_labels
train_dataset = np.array(data).reshape(-1,4)
train_labels = np.zeros([train_dataset.shape[0],num_classes])
#one-hot encoding
for i,label in enumerate(target):
train_labels[i,label] = 1
#normalizations
for i in range(input_units):
mean = train_dataset[:,i].mean()
std = train_dataset[:,i].std()
train_dataset[:,i] = (train_dataset[:,i]-mean)/std
# -
#train data
J,final_output = train(train_dataset,train_labels,iters=4001)
# #### Reached an Accuracy of 97%
# ### Plot the loss vs iteration graph
#plot loss graph
plt.plot(list(range(1,len(J))),J[1:])
plt.xlabel('Iterations')
plt.ylabel('Loss')
plt.title('Iterations VS Loss')
plt.show()
| Neural Network/NeuralNetwork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Workflows
#
# Although it would be possible to write analysis scripts using just Nipype [Interfaces](basic_interfaces.ipynb), and this may provide some advantages over directly making command-line calls, the main benefits of Nipype are the workflows.
#
# A workflow controls the setup and the execution of individual interfaces. Let's assume you want to run multiple interfaces in a specific order, where some have to wait for others to finish while others can be executed in parallel. The nice thing about a nipype workflow is, that the workflow will take care of input and output of each interface and arrange the execution of each interface in the most efficient way.
#
# A workflow therefore consists of multiple [Nodes](basic_nodes.ipynb), each representing a specific [Interface](basic_interfaces.ipynb) and directed connection between those nodes. Those connections specify which output of which node should be used as an input for another node. To better understand why this is so great, let's look at an example.
# ## Interfaces vs. Workflows
#
# Interfaces are the building blocks that solve well-defined tasks. We solve more complex tasks by combining interfaces with workflows:
#
# <table style="width: 100%; font-size: 14px;">
# <thead>
# <th style="text-align:left">Interfaces</th>
# <th style="text-align:left">Workflows</th>
# </thead>
# <tbody>
# <tr>
# <td style="text-align:left">Wrap *unitary* tasks</td>
# <td style="text-align:left">Wrap *meta*-tasks
# <li style="text-align:left">implemented with nipype interfaces wrapped inside ``Node`` objects</li>
# <li style="text-align:left">subworkflows can also be added to a workflow without any wrapping</li>
# </td>
# </tr>
# <tr>
# <td style="text-align:left">Keep track of the inputs and outputs, and check their expected types</td>
# <td style="text-align:left">Do not have inputs/outputs, but expose them from the interfaces wrapped inside</td>
# </tr>
# <tr>
# <td style="text-align:left">Do not cache results (unless you use [interface caching](advanced_interfaces_caching.ipynb))</td>
# <td style="text-align:left">Cache results</td>
# </tr>
# <tr>
# <td style="text-align:left">Run by a nipype plugin</td>
# <td style="text-align:left">Run by a nipype plugin</td>
# </tr>
# </tbody>
# </table>
# ## Preparation
#
# Before we can start, let's first load some helper functions:
# +
import numpy as np
import nibabel as nb
import matplotlib.pyplot as plt
# Let's create a short helper function to plot 3D NIfTI images
def plot_slice(fname):
# Load the image
img = nb.load(fname)
data = img.get_data()
# Cut in the middle of the brain
cut = int(data.shape[-1]/2) + 10
# Plot the data
plt.imshow(np.rot90(data[..., cut]), cmap="gray")
plt.gca().set_axis_off()
# -
# # Example 1 - ``Command-line`` execution
#
# Let's take a look at a small preprocessing analysis where we would like to perform the following steps of processing:
#
# - Skullstrip an image to obtain a mask
# - Smooth the original image
# - Mask the smoothed image
#
# This could all very well be done with the following shell script:
# + language="bash"
# ANAT_NAME=sub-01_ses-test_T1w
# ANAT=/data/ds000114/sub-01/ses-test/anat/${ANAT_NAME}
# bet ${ANAT} /output/${ANAT_NAME}_brain -m -f 0.3
# fslmaths ${ANAT} -s 2 /output/${ANAT_NAME}_smooth
# fslmaths /output/${ANAT_NAME}_smooth -mas /output/${ANAT_NAME}_brain_mask /output/${ANAT_NAME}_smooth_mask
# -
# This is simple and straightforward. We can see that this does exactly what we wanted by plotting the four steps of processing.
f = plt.figure(figsize=(12, 4))
for i, img in enumerate(["T1w", "T1w_smooth",
"T1w_brain_mask", "T1w_smooth_mask"]):
f.add_subplot(1, 4, i + 1)
if i == 0:
plot_slice("/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_%s.nii.gz" % img)
else:
plot_slice("/output/sub-01_ses-test_%s.nii.gz" % img)
plt.title(img)
# # Example 2 - ``Interface`` execution
# Now let's see what this would look like if we used Nipype, but only the Interfaces functionality. It's simple enough to write a basic procedural script, this time in Python, to do the same thing as above:
# +
from nipype.interfaces import fsl
# Skullstrip process
skullstrip = fsl.BET(
in_file="/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz",
out_file="/output/sub-01_T1w_brain.nii.gz",
mask=True)
skullstrip.run()
# Smoothing process
smooth = fsl.IsotropicSmooth(
in_file="/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz",
out_file="/output/sub-01_T1w_smooth.nii.gz",
fwhm=4)
smooth.run()
# Masking process
mask = fsl.ApplyMask(
in_file="/output/sub-01_T1w_smooth.nii.gz",
out_file="/output/sub-01_T1w_smooth_mask.nii.gz",
mask_file="/output/sub-01_T1w_brain_mask.nii.gz")
mask.run()
f = plt.figure(figsize=(12, 4))
for i, img in enumerate(["T1w", "T1w_smooth",
"T1w_brain_mask", "T1w_smooth_mask"]):
f.add_subplot(1, 4, i + 1)
if i == 0:
plot_slice("/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_%s.nii.gz" % img)
else:
plot_slice("/output/sub-01_%s.nii.gz" % img)
plt.title(img)
# -
# This is more verbose, although it does have its advantages. There's the automated input validation we saw previously, some of the options are named more meaningfully, and you don't need to remember, for example, that fslmaths' smoothing kernel is set in sigma instead of FWHM -- Nipype does that conversion behind the scenes.
#
# ### Can't we optimize that a bit?
#
# As we can see above, the inputs for the **``mask``** routine ``in_file`` and ``mask_file`` are actually the output of **``skullstrip``** and **``smooth``**. We therefore somehow want to connect them. This can be accomplisehd by saving the executed routines under a given object and than using the output of those objects as input for other routines.
# +
from nipype.interfaces import fsl
# Skullstrip process
skullstrip = fsl.BET(
in_file="/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz", mask=True)
bet_result = skullstrip.run() # skullstrip object
# Smooth process
smooth = fsl.IsotropicSmooth(
in_file="/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz", fwhm=4)
smooth_result = smooth.run() # smooth object
# Mask process
mask = fsl.ApplyMask(in_file=smooth_result.outputs.out_file,
mask_file=bet_result.outputs.mask_file)
mask_result = mask.run()
f = plt.figure(figsize=(12, 4))
for i, img in enumerate([skullstrip.inputs.in_file, smooth_result.outputs.out_file,
bet_result.outputs.mask_file, mask_result.outputs.out_file]):
f.add_subplot(1, 4, i + 1)
plot_slice(img)
plt.title(img.split('/')[-1].split('.')[0].split('test_')[-1])
# -
# Here we didn't need to name the intermediate files; Nipype did that behind the scenes, and then we passed the result object (which knows those names) onto the next step in the processing stream. This is somewhat more concise than the example above, but it's still a procedural script. And the dependency relationship between the stages of processing is not particularly obvious. To address these issues, and to provide solutions to problems we might not know we have yet, Nipype offers **Workflows.**
# # Example 3 - ``Workflow`` execution
#
# What we've implicitly done above is to encode our processing stream as a directed acyclic graphs: each stage of processing is a node in this graph, and some nodes are unidirectionally dependent on others. In this case there is one input file and several output files, but there are no cycles -- there's a clear line of directionality to the processing. What the Node and Workflow classes do is make these relationships more explicit.
#
# The basic architecture is that the Node provides a light wrapper around an Interface. It exposes the inputs and outputs of the Interface as its own, but it adds some additional functionality that allows you to connect Nodes into a Workflow.
#
# Let's rewrite the above script with these tools:
# +
# Import Node and Workflow object and FSL interface
from nipype import Node, Workflow
from nipype.interfaces import fsl
# For reasons that will later become clear, it's important to
# pass filenames to Nodes as absolute paths
from os.path import abspath
in_file = abspath("/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz")
# Skullstrip process
skullstrip = Node(fsl.BET(in_file=in_file, mask=True), name="skullstrip")
# Smooth process
smooth = Node(fsl.IsotropicSmooth(in_file=in_file, fwhm=4), name="smooth")
# Mask process
mask = Node(fsl.ApplyMask(), name="mask")
# -
# This looks mostly similar to what we did above, but we've left out the two crucial inputs to the ApplyMask step. We'll set those up by defining a Workflow object and then making *connections* among the Nodes.
# Initiation of a workflow
wf = Workflow(name="smoothflow", base_dir="/output/working_dir")
# The Workflow object has a method called ``connect`` that is going to do most of the work here. This routine also checks if inputs and outputs are actually provided by the nodes that are being connected.
#
# There are two different ways to call ``connect``:
#
# connect(source, "source_output", dest, "dest_input")
#
# connect([(source, dest, [("source_output1", "dest_input1"),
# ("source_output2", "dest_input2")
# ])
# ])
#
# With the first approach you can establish one connection at a time. With the second you can establish multiple connects between two nodes at once. In either case, you're providing it with four pieces of information to define the connection:
#
# - The source node object
# - The name of the output field from the source node
# - The destination node object
# - The name of the input field from the destination node
#
# We'll illustrate each method in the following cell:
# +
# First the "simple", but more restricted method
wf.connect(skullstrip, "mask_file", mask, "mask_file")
# Now the more complicated method
wf.connect([(smooth, mask, [("out_file", "in_file")])])
# -
# Now the workflow is complete!
#
# Above, we mentioned that the workflow can be thought of as a directed acyclic graph. In fact, that's literally how it's represented behind the scenes, and we can use that to explore the workflow visually:
wf.write_graph("workflow_graph.dot")
from IPython.display import Image
Image(filename="/output/working_dir/smoothflow/workflow_graph.png")
# This representation makes the dependency structure of the workflow obvious. (By the way, the names of the nodes in this graph are the names we gave our Node objects above, so pick something meaningful for those!)
#
# Certain graph types also allow you to further inspect the individual connections between the nodes. For example:
wf.write_graph(graph2use='flat')
from IPython.display import Image
Image(filename="/output/working_dir/smoothflow/graph_detailed.png")
# Here you see very clearly, that the output ``mask_file`` of the ``skullstrip`` node is used as the input ``mask_file`` of the ``mask`` node. For more information on graph visualization, see the [Graph Visualization](./basic_graph_visualization.ipynb) section.
# But let's come back to our example. At this point, all we've done is define the workflow. We haven't executed any code yet. Much like Interface objects, the Workflow object has a ``run`` method that we can call so that it executes. Let's do that and then examine the results.
# +
# Specify the base directory for the working directory
wf.base_dir = "/output/working_dir"
# Execute the workflow
wf.run()
# -
# **The specification of ``base_dir`` is very important (and is why we needed to use absolute paths above), because otherwise all the outputs would be saved somewhere in the temporary files.** Unlike interfaces, which by default spit out results to the local directry, the Workflow engine executes things off in its own directory hierarchy.
#
# Let's take a look at the resulting images to convince ourselves we've done the same thing as before:
f = plt.figure(figsize=(12, 4))
for i, img in enumerate(["/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz",
"/output/working_dir/smoothflow/smooth/sub-01_ses-test_T1w_smooth.nii.gz",
"/output/working_dir/smoothflow/skullstrip/sub-01_ses-test_T1w_brain_mask.nii.gz",
"/output/working_dir/smoothflow/mask/sub-01_ses-test_T1w_smooth_masked.nii.gz"]):
f.add_subplot(1, 4, i + 1)
plot_slice(img)
# Perfet!
#
# Let's also have a closer look at the working directory:
# !tree /output/working_dir/smoothflow/ -I '*js|*json|*html|*pklz|_report'
# As you can see, the name of the working directory is the name we gave the workflow ``base_dir``. And the name of the folder within is the name of the workflow object ``smoothflow``. Each node of the workflow has its' own subfolder in the ``smoothflow`` folder. And each of those subfolders contains the output of the node as well as some additional files.
# # The #1 gotcha of nipype Workflows
#
# Nipype workflows are just DAGs (Directed Acyclic Graphs) that the runner ``Plugin`` takes in and uses to compose an ordered list of nodes for execution. As a matter of fact, running a workflow will return a graph object. That's why you often see something like `<networkx.classes.digraph.DiGraph at 0x7f83542f1550>` at the end of exectuion stream when running a workflow.
#
# The principal implication is that ``Workflow``s *don't have inputs and outputs*, you can just access them through the ``Node`` decoration.
#
# In practical terms, this has one clear consequence: from the resulting object of the workflow execution you don't generally have access to the value of the outputs of the interfaces. This is particularly true for Plugins with asynchronous execution.
# # A workflow inside a workflow
# When you start writing full-fledged analysis workflows, things can get quite complicated. Some aspects of neuroimaging analysis can be thought of as a coherent step at a level more abstract than the execution of a single command line binary. For instance, in the standard FEAT script in FSL, several calls are made in the process of using `susan` to perform nonlinear smoothing on an image. In Nipype, you can write **nested workflows**, where a sub-workflow can take the place of a Node in a given script.
#
# Let's use the prepackaged `susan` workflow that ships with Nipype to replace our Gaussian filtering node and demonstrate how this works.
from nipype.workflows.fmri.fsl import create_susan_smooth
# Calling this function will return a pre-written `Workflow` object:
susan = create_susan_smooth(separate_masks=False)
# Let's display the graph to see what happens here.
susan.write_graph("susan_workflow.dot")
from IPython.display import Image
Image(filename="susan_workflow.png")
# We see that the workflow has an `inputnode` and an `outputnode`. While not strictly necessary, this is standard practice for workflows (especially those that are intended to be used as nested workflows in the context of a longer analysis graph) and makes it more clear how to connect inputs and outputs from this workflow.
#
# Let's take a look at what those inputs and outputs are. Like Nodes, Workflows have `inputs` and `outputs` attributes that take a second sub-attribute corresponding to the specific node we want to make connections to.
print("Inputs:\n", susan.inputs.inputnode)
print("Outputs:\n", susan.outputs.outputnode)
# Note that `inputnode` and `outputnode` are just conventions, and the Workflow object exposes connections to all of its component nodes:
susan.inputs
# Let's see how we would write a new workflow that uses this nested smoothing step.
#
# The susan workflow actually expects to receive and output a list of files (it's intended to be executed on each of several runs of fMRI data). We'll cover exactly how that works in later tutorials, but for the moment we need to add an additional ``Function`` node to deal with the fact that ``susan`` is outputting a list. We can use a simple `lambda` function to do this:
from nipype import Function
extract_func = lambda list_out: list_out[0]
list_extract = Node(Function(input_names=["list_out"],
output_names=["out_file"],
function=extract_func),
name="list_extract")
# Now let's create a new workflow ``susanflow`` that contains the ``susan`` workflow as a sub-node. To be sure, let's also recreate the ``skullstrip`` and the ``mask`` node from the examples above.
# +
# Initiate workflow with name and base directory
wf2 = Workflow(name="susanflow", base_dir="/output/working_dir")
# Create new skullstrip and mask nodes
skullstrip2 = Node(fsl.BET(in_file=in_file, mask=True), name="skullstrip")
mask2 = Node(fsl.ApplyMask(), name="mask")
# Connect the nodes to each other and to the susan workflow
wf2.connect([(skullstrip2, mask2, [("mask_file", "mask_file")]),
(skullstrip2, susan, [("mask_file", "inputnode.mask_file")]),
(susan, list_extract, [("outputnode.smoothed_files",
"list_out")]),
(list_extract, mask2, [("out_file", "in_file")])
])
# Specify the remaining input variables for the susan workflow
susan.inputs.inputnode.in_files = abspath(
"/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz")
susan.inputs.inputnode.fwhm = 4
# -
# First, let's see what this new processing graph looks like.
wf2.write_graph(dotfilename='/output/working_dir/full_susanflow.dot', graph2use='colored')
from IPython.display import Image
Image(filename="/output/working_dir/full_susanflow.png")
# We can see how there is a nested smoothing workflow (blue) in the place of our previous `smooth` node. This provides a very detailed view, but what if you just wanted to give a higher-level summary of the processing steps? After all, that is the purpose of encapsulating smaller streams in a nested workflow. That, fortunately, is an option when writing out the graph:
wf2.write_graph(dotfilename='/output/working_dir/full_susanflow_toplevel.dot', graph2use='orig')
from IPython.display import Image
Image(filename="/output/working_dir/full_susanflow_toplevel.png")
# That's much more managable. Now let's execute the workflow
wf2.run()
# As a final step, let's look at the input and the output. It's exactly what we wanted.
f = plt.figure(figsize=(12, 4))
for i, e in enumerate([["/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz", 'input'],
["/output/working_dir//susanflow/mask/sub-01_ses-test_T1w_smooth_masked.nii.gz",
'output']]):
f.add_subplot(1, 2, i + 1)
plot_slice(e[0])
plt.title(e[1])
# # So, why are workflows so great?
#
# So far, we've seen that you can build up rather complex analysis workflows. But at the moment, it's not been made clear why this is worth the extra trouble from writing a simple procedural script. To demonstrate the first added benefit of the Nipype, let's just rerun the ``susanflow`` workflow from above and measure the execution times.
# %time wf2.run()
# That happened quickly! **Workflows (actually this is handled by the Node code) are smart, and know if their inputs have changed from the last time they are run. If they have not, they don't recompute; they just turn around and pass out the resulting files from the previous run.** This is done on a node-by-node basis, also.
#
# Let's go back to the first workflow example. What happened if we just tweak one thing:
wf.inputs.smooth.fwhm = 1
wf.run()
# By changing an input value of the ``smooth`` node, this node will be re-executed. This triggers a cascade such that any file depending on the ``smooth`` node (in this case, the ``mask`` node, also recompute). However, the ``skullstrip`` node hasn't changed since the first time it ran, so it just coughed up its original files.
#
# That's one of the main benefit of using Workflows: **efficient recomputing**.
#
# Another benefits of Workflows is parallel execution, which is covered under [Plugins and Distributed Computing](./basic_plugins.ipynb). With Nipype it is very easy to up a workflow to an extremely parallel cluster computing environment.
#
# In this case, that just means that the `skullstrip` and `smooth` Nodes execute together, but when you scale up to Workflows with many subjects and many runs per subject, each can run together, such that (in the case of unlimited computing resources), you could process 50 subjects with 10 runs of functional data in essentially the time it would take to process a single run.
#
# To emphasize the contribution of Nipype here, you can write and test your workflow on one subject computing on your local CPU, where it is easier to debug. Then, with the change of a single function parameter, you can scale your processing up to a 1000+ node SGE cluster.
# ### Exercise 1
#
# Create a workflow that connects three nodes for:
# - skipping the first 3 dummy scans using ``fsl.ExtractROI``
# - applying motion correction using ``fsl.MCFLIRT`` (register to the mean volume, use NIFTI as output type)
# - correcting for slice wise acquisition using ``fsl.SliceTimer`` (assumed that slices were acquired with interleaved order and time repetition was 2.5, use NIFTI as output type)
# + solution2="hidden" solution2_first=true
# write your solution here
# + solution2="hidden"
# importing Node and Workflow
from nipype import Workflow, Node
# importing all interfaces
from nipype.interfaces.fsl import ExtractROI, MCFLIRT, SliceTimer
# + [markdown] solution2="hidden"
# Defining all nodes
# + solution2="hidden"
# extracting all time levels but not the first four
extract = Node(ExtractROI(t_min=4, t_size=-1, output_type='NIFTI'),
name="extract")
# using MCFLIRT for motion correction to the mean volume
mcflirt = Node(MCFLIRT(mean_vol=True,
output_type='NIFTI'),
name="mcflirt")
# correcting for slice wise acquisition (acquired with interleaved order and time repetition was 2.5)
slicetimer = Node(SliceTimer(interleaved=True,
output_type='NIFTI',
time_repetition=2.5),
name="slicetimer")
# + [markdown] solution2="hidden"
# Creating a workflow
# + solution2="hidden"
# Initiation of a workflow
wf_ex1 = Workflow(name="exercise1", base_dir="/output/working_dir")
# connect nodes with each other
wf_ex1.connect([(extract, mcflirt, [('roi_file', 'in_file')]),
(mcflirt, slicetimer, [('out_file', 'in_file')])])
# providing a input file for the first extract node
extract.inputs.in_file = "/data/ds000114/sub-01/ses-test/func/sub-01_ses-test_task-fingerfootlips_bold.nii.gz"
# -
# ### Exercise 2
# Visualize and run the workflow
# + solution2="hidden" solution2_first=true
# write your solution here
# + [markdown] solution2="hidden"
# We learnt 2 methods of plotting graphs:
# + solution2="hidden"
wf_ex1.write_graph("workflow_graph.dot")
from IPython.display import Image
Image(filename="/output/working_dir/exercise1/workflow_graph.png")
# + [markdown] solution2="hidden"
# And more detailed graph:
# + solution2="hidden"
wf_ex1.write_graph(graph2use='flat')
from IPython.display import Image
Image(filename="/output/working_dir/exercise1/graph_detailed.png")
# + [markdown] solution2="hidden"
# if everything works good, we're ready to run the workflow:
# + solution2="hidden"
wf_ex1.run()
# + [markdown] solution2="hidden"
# we can now check the output:
# + solution2="hidden"
# ! ls -lh /output/working_dir/exercise1
| notebooks/basic_workflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 상미분 방정식의 엄밀해<br>An Exact Solution of an Ordinary Differential Equation
#
#
# 미분 방정식의 한 간단한 예는 다음과 같다.<br>A simple example of a differential equation is as follows.
#
#
# $$
# \begin{cases}
# \begin{align}
# a_0 \frac{d}{dt}x(t)+a_1 x(t)&=0 \\
# x(0)&=x_0 \\
# \end{align}
# \end{cases}
# $$
#
#
# 위 미분방정식의 해는 함수 $x(t)$ 이다.<br>
# A solution for the differential equation above is the function $x(t)$.
#
#
# 어떤 미분방정식을 푼다는 것은, 해당 미분방정식을 만족하는 함수 $x(t)$를 찾는 것을 뜻한다.<br>
# Solving for a differential equation means searching for a function $x(t)$ satisfying the differential equation.
#
#
# 처음에는 어떤 형태의 함수가 해당 미분 방정식을 만족시킬 것인지 잘 모를 수도 있겠지만 몇 가지 함수를 생각해 보자.<br>
# At first, we may not be certain about which type of function may satisfy the equation above, but let's consider a few possible candiates.
#
#
# ## 다항식?<br>A polynomial?
#
#
# 다음과 같은 다항식이 만족시킬 수 있을까?<br>Is a polynomial as follows suitable?
#
#
# $$
# x(t) = A_0 t^2 + A_1 t + A_2
# $$
#
#
# 미분해 보자<br>Let's differentiate.
#
#
# $$
# \frac{d}{dt}x(t) = 2 A_0 t + A_1
# $$
#
#
# 각각 계수로 곱해보자.<br>Let's multiply with respective coefficients.
#
#
# $$
# \begin{cases}
# \begin{align}
# a_0 \frac{d}{dt}x(t) &= a_0 \left(2 A_0 t + A_1 \right)\\
# a_1 x(t)&=a_1 \left( A_0 t^2 + A_1 t + A_2 \right) \\
# \end{align}
# \end{cases}
# $$
#
#
# 등호의 오른쪽은 오른쪽 끼리, 등호의 왼쪽은 왼쪽 끼리 더해 보자.<br>Let's add each side respectively.
#
#
# $$
# \begin{align}
# a_1 x(t) + a_0\frac{d}{dt}x(t) &= a_1 \left( A_0 t^2 + A_1 t + A_2 \right) + a_0 \left(2 A_0 t + A_1 \right) \\
# & = a_1 A_0 t^2 + \left(2 a_0 A_0 + a_1 A_1\right) t + \left(a_0 A_1 + a_1 A_2\right)
# \end{align}
# $$
#
#
# 위에서<br>Remember
#
#
# $$
# a_0 \frac{d}{dt}x(t)+a_1 x(t)=0
# $$
#
#
# 이었으므로 $t$가 어떤 값을 가지든 <br> thus for all $t$
#
#
# $$
# a_1 A_0 t^2 + \left(2 a_0 A_0 + a_1 A_1\right) t + \left(a_0 A_1 + a_1 A_2\right) = 0
# $$
#
#
# 이어야 한다. <br> should be satisfied.
#
#
# 이러한 조건이 전혀 만족될 수 없는 것은 아닐 것이나, 일단은 좀 더 일반적으로 유용할 수 있어 보이는 경우를 살펴보도록 하자.<br>
# Although this could be satisfied in some way or the other, however, for now let's take a look at a different case that seems more generally useful.
#
#
# ## 지수 함수?<br>Exponential Function?
#
#
# 아래와 같은 지수함수를 생각해 보자.<br>
# Let's think about an exponential function as follows.
#
#
# $$
# x(t) = A_0 e^{\lambda t}
# $$
#
#
# 전과 마찬가지로, 미분하고 계수를 곱해 더해보자.<br>
# As before, let's differentiate and add after multiplying coefficients.
#
#
# $$
# \frac{d}{dt}x(t) = A_0 \lambda e^{\lambda t}
# $$
#
#
# $$
# \begin{cases}
# \begin{align}
# a_0 \frac{d}{dt}x(t) &= a_0 \left(A_0 \lambda e^{\lambda t} \right)\\
# a_1 x(t)&=a_1 \left( A_0 e^{\lambda t} \right) \\
# \end{align}
# \end{cases}
# $$
#
#
# $$
# a_0 \frac{d}{dt}x(t) + a_1 x(t) = a_0 \left(A_0 \lambda e^{\lambda t} \right) + a_1 \left( A_0 e^{\lambda t} \right) =0
# $$
#
#
# $t$가 어떤 값을 가지든 상관 없이 위 관계는 만족되어야한다.<br>For all $t$, the relationship above must be satisfied.
#
#
# $A_0 e^{\lambda t}$ 에 관해 묶어 보자.<br>Let's factor out $A_0 e^{\lambda t}$.
#
#
# $$
# \begin{align}
# a_0 \left(A_0 \lambda e^{\lambda t} \right) + a_1 \left( A_0 e^{\lambda t} \right) &= A_0 e^{\lambda t} a_0 \lambda + A_0 e^{\lambda t} a_1 \\
# &= A_0 e^{\lambda t} \left(a_0 \lambda + a_1 \right)=0
# \end{align}
# $$
#
#
# 모든 $t$ 에 대해 위 식이 $0$이 되려면, $A_0=0$ 이거나 $a_0 \lambda + a_1 = 0$ 이어야 한다.<br>
# To make the equation above to be $0$ for all $t$, either $A_0=0$ or $a_0 \lambda + a_1 = 0$.
#
#
# $A_0=0$ 인 경우:<br>If $A_0=0$:
#
#
# $$
# x(t)=0 \cdot e^{\lambda t} = 0
# $$
#
#
# $a_0 \lambda + a_1 = 0$ 인 경우:<br>If $a_0 \lambda + a_1 = 0$:
#
#
# $$
# \begin{align}
# a_0 \lambda + a_1 &= 0 \\
# \lambda &= -\frac{a_1}{a_0} \\
# x(t)&=A_0 e^{\lambda t}=A_0 e^{-\frac{a_1}{a_0} t}=A_0 exp\left(-\frac{a_1}{a_0} t\right)
# \end{align}
# $$
#
#
# 다항식의 경우와 비교해 보면 지수함수쪽이 조금 더 흥미있어 보일 수 있다.<br>Comparing with the polynomial, the exponential functions may seem somewhat more interesting.
#
#
# ## 초기 조건<br>Initial condition
#
#
# 그런데 $A_0$는 어떻게 정하는 것이 좋을까? 처음의 식으로 돌아가 보자.<br>However, what could be a good way to decide $A_0$? Let's go back to the first equation.
#
#
# $$
# \begin{cases}
# \begin{align}
# a_0 \frac{d}{dt}x(t)+a_1 x(t)&=0 \\
# x(0)&=x_0 \\
# \end{align}
# \end{cases}
# $$
#
#
# 따라서 $x(0)=x_0$ 였음을 이용해 보자. 여기서 $x_0$는 어떤 알려진 상수이다.<br>Thus let's use $x(0)=x_0$. Here, $x_0$ is a known constant.
#
#
# $$
# \begin{align}
# x(t)&=A_0 e^{-\frac{a_1}{a_0} t} \\
# x(0)&=A_0 e^{-\frac{a_1}{a_0}\cdot 0}=A_0 e^0=A_0 \cdot 1 = A_0=x_0 \\
# x(t)&=x_0 e^{-\frac{a_1}{a_0} t}
# \end{align}
# $$
#
#
# 이렇게 어떤 미분방정식의 엄밀해를 구할 수 있었다.<br>This way, we could find an exact solution of a differential equation.
#
#
# ## Final Bell<br>마지막 종
#
#
# +
# stackoverfow.com/a/24634221
import os
os.system("printf '\a'");
# -
| 00_Ordinary_Differential_Equation_Exact_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import os
import numpy as np
from IPython.display import HTML
#source: in class examples
# Study data files
pwd = os.path.abspath('.')
weather_path = os.path.join(pwd, 'cities.csv')
# Read the mouse data and the study results
weather_data = pd.read_csv(weather_path)
weather_data.head()
# +
weather_data.to_html('weather_data.html', index=False)
html_table = weather_data.to_html
print(html_table)
# -
| weather_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iVibudh/Hugging-Face-Course/blob/main/notebooks/01_labeling_VCs_with_zero_shot_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eisQjUpD-eLf"
# # Labeling VCs with Zero-Shot Learning using Huggingface Transformers
#
# This notebook has the goal of taking the text from ESA tables and trying to classify the VCs for a specific table. We use an approach called Zero-Shot Learning to do this. I won't go over the nitty-gritty details of Zero-Shot, but you can go to the following link if you are interested: https://towardsdatascience.com/zero-shot-text-classification-evaluation-c7ba0f56688e.
#
# Since we have to classify *a lot* of tables, it will be better to use a GPU to speed up inference/prediction speed.
#
# # Installations
# + colab={"base_uri": "https://localhost:8080/"} id="KGzdoHOghfJH" outputId="b806ccf3-471f-40ce-89c5-a1efa0f5dd92"
# !pip install transformers --quiet
# !pip install tqdm --quiet
# + [markdown] id="_8rHj6q5-tC5"
# # Imports
# + id="aEvJ1LmnO9Hh" colab={"base_uri": "https://localhost:8080/"} outputId="cb0377b0-8f4b-40e7-a08b-0d0d9887805a"
import os
import requests
import pickle
import pandas as pd
from transformers import pipeline
from bs4 import BeautifulSoup
import nltk
from nltk.stem.porter import *
from tqdm import tqdm
nltk.download('stopwords', quiet=True)
# + id="yAZmqv-XctM6"
pickled_dataset = 'zero_shot_vcs_train'
# + [markdown] id="tBRuBx98-wWg"
# # Loading the Data
# + id="AYzuCqZUPjCo"
csv_url = 'https://raw.githubusercontent.com/JayThibs/huggingface-course-cer-workshop/main/data/zero_shot_esa_index_train_128_max_tokens.csv'
df_joined = pd.read_csv(csv_url, index_col=0)
# + colab={"base_uri": "https://localhost:8080/"} id="y_VYcHfZJKTk" outputId="af93ad8a-5a06-40a6-de2f-954852f04f8a"
len(df_joined)
# + colab={"base_uri": "https://localhost:8080/", "height": 608} id="1N82Q4fFi4S2" outputId="c58039b5-2b24-459a-d063-517256ed5abc"
df_joined.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="Ip3S0PYHPmPJ" outputId="9be38864-160e-4340-e251-b19bf42dd0e5"
df_joined.iloc[100]['text']
# + id="7hM_whVXPpDW"
sequences = []
for x in range(len(df_joined)):
sequences.append(df_joined.iloc[x]['text'][0:250])
# + colab={"base_uri": "https://localhost:8080/"} id="y2i8SmQVVF0J" outputId="e54e952f-e0d5-4399-8b29-2cd36ba7b36a"
len(sequences)
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="dwT0g0J56L2q" outputId="91c03c4f-96e6-4210-b471-5c5010ffa41e"
sequences[0]
# + [markdown] id="XQV6qG0kX77D"
# # Loading the Model with `pipeline`
# + id="FDuUc81sPkjT" colab={"base_uri": "https://localhost:8080/", "height": 226, "referenced_widgets": ["6acc0c12ede74f56ae2d2f6abceda90f", "2e2a8761f188447eaa13f2516908b657", "848fc13b7f2d43f1b3bf205ae668c87d", "8875737dea3c44ad9ef85451307095af", "3f770c5724ae4b3a908ed5cff6b6e8bc", "260952bf2da942919cb61b0ce1876b9a", "e568a84f45d94c03b937b488a705b72b", "69ece20357c743bd8ebc866a372081a9", "2ba1064fcbcf4a7f9bc715363504dbd5", "168fe836e28e42099ff08a4474cf84cf", "0c2b4745ee6f4806a4ca21447e0fbdb4", "2d3b8a758b984a68a863567b16a082fe", "<KEY>", "d06b09ce9c7945c38b14508b3e8d1a8b", "<KEY>", "<KEY>", "fc7fa832810c4e92ba2369cf6ec4cdb2", "<KEY>", "c18e7835630d44aaa9497fd8795fee5f", "3e95c435c29a48c2a6d39b5a1ea72c2a", "0fc3e431ad444a2c9dedc0b041cb8a35", "<KEY>", "52c1f261511c412ca80f2e1beea9d578", "<KEY>", "2cf696340013425a966e17d98edd45ba", "<KEY>", "<KEY>", "<KEY>", "af227674ada641f8ac3a0378805ef275", "8fb1e7a2312344ca8f8be8fa3350c166", "<KEY>", "<KEY>", "f6ca533c94ab4a66aa79b9dabd587dd2", "<KEY>", "<KEY>", "<KEY>", "fcf9ad0e1fbf4d7294841d480e2efe5a", "9ac53ac2b10d4ef9bf33b8e3923df392", "<KEY>", "b0812840381145df995f5395737984f5", "f27d7b5ad833480c8054bd78ec07afc3", "be9e6895a54048b7880dd46b5b63d043", "<KEY>", "<KEY>", "ce5576885d7947f1806dff4eccdfb5b1", "<KEY>", "c958182800e942589650e16283bc7403", "45b202a9fd2948e3b932ea9e936702a4", "47827342be544cf49f2061fcc8e5e7e2", "582274340f3a4523a6418553d90916c1", "fd7ac13075634c45869e0095d90608b6", "39343b83e90c4216a41b541e08fab077", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8e0c0a8798dc4cab9a06b1f232d35d07", "4c96bae9df954e57a61502579b4e3b75", "<KEY>", "7728b5c6b98e4983879ef98f433f5483", "ed28d76d76ea4d0db7d07f66015fec28", "<KEY>", "<KEY>", "<KEY>"]} outputId="7545d441-0763-4fe0-d017-2b9a1cefaf60"
# Explicitly ask for tensor allocation on CUDA device :0
classifier = pipeline("zero-shot-classification", device=0)
# + colab={"base_uri": "https://localhost:8080/"} id="loDVpdHvP2XQ" outputId="a1e6fd65-5f8a-448e-9442-89e0a2df34a9"
candidate_labels = """Physical and meteorological environment,Soil and soil productivity,Vegetation,Water quality and quantity,Fish and fish habitat,Wetlands,Wildlife and wildlife habitat,Species at Risk or Species of Special Status and related habitat,Greenhouse gas (GHG) emissions and climate change,GHG Emissions and Climate Change – Assessment of Upstream GHG Emissions,Air emissions,Acoustic environment,Electromagnetism and Corona Discharge,Human occupancy and resource use,Heritage resources,Navigation and navigation safety,Traditional land and resource use,Social and cultural well-being,Human health and aesthetics,Infrastructure and services,Employment and economy,Environmental Obligations,Rights of Indigenous Peoples""".split(',')
hypothesis_template = "The data from this table is about {}."
candidate_labels
# + [markdown] id="N9TRFVGaaAvE"
# The following code cell sometimes helps when Colab's GPU is full. That said, I don't find it helps very much even though people mention it quite often in StackOverflow threads.
#
# Generally, if you get a CUDA error during inference or training, it will likely be because you've loaded too much data onto the GPU and it is out of space. To resolve this, you will need to do a Factory Reset of the notebook instance, which you can find in the Runtime tab. After this, you will need to rerun all of your code.
# + id="p5AbPnG318bj"
import gc
import torch
gc.collect()
torch.cuda.empty_cache()
# + colab={"base_uri": "https://localhost:8080/", "height": 88} id="Ne-OdCs32Fyb" outputId="380daed5-e99e-49b8-b519-4e6f181871c4"
torch.cuda.memory_summary(device=None, abbreviated=False)
# + [markdown] id="J9fNbAH3avev"
# Below is where we do inference on all of the table texts. Since there can be a lot of text in tables, we needed to cut down the length of words when creating the dataset, otherwise you will use up all the memory and inference will crash.
#
# However, this is not the only thing you need to worry about. When you are doing inference, you can actually to inference in "batches". This means that you are loading multiple examples of text at the same time onto the GPU. The more batches of data you load on the GPU at the same time, the faster your code will run. However, you also have the issue of running out of memory if you load too much data at once! So, if you get a "CUDA out of memory" type error, your first instinct should be to reduce the batch size. As you can see below in the following line:
#
# ```
# for sequence_batch in tqdm(grouper(sequences, 2, "")):
# ```
#
# I am only loading to examples at once! This is because more than two was too much for the Colab GPUs. Each GPU has different amounts of memory available, so they won't all have the same limit. In practice, this is just something you will need to play around with until you can find a "batch size" that doesn't overload the GPU, but still runs fast.
# + id="9F2Z8U3XYDSa" colab={"base_uri": "https://localhost:8080/"} outputId="94f0d65a-0213-426d-dc7c-510d0635be18"
from itertools import zip_longest
def grouper(iterable_obj, count, fillvalue=None):
args = [iter(iterable_obj)] * count
return zip_longest(*args, fillvalue=fillvalue)
output = []
for sequence_batch in tqdm(grouper(sequences, 2, "")):
# print(sequence_batch)
output.append(classifier(sequence_batch, candidate_labels, hypothesis_template=hypothesis_template, multi_label=True))
# + [markdown] id="zrKeTC7JcTVM"
# Phew! That took just a little over 3 hours.
#
# Now, let's check the output:
# + colab={"base_uri": "https://localhost:8080/"} id="0RS5OvrhS-Km" outputId="ffa09ccd-f104-4862-ea54-2385b9f65246"
output[0]
# + [markdown] id="MsN-rvbvcdQt"
# As you can see, we have a list of the VC labels along with a corresponding list of confidence scores. The closer the confidence score is to 1, the more likely the table should be labeled as that VC. Closer to 0 means it shouldn't be labeled that VC.
# + id="YbaOlI23wPEE"
# Since we have a list of lists for the outputs (because inference was done in batches), we need to flatten the lists together into one.
flatOutput = [item for elem in output for item in elem]
# + [markdown] id="KEBK8eH_cyln"
# # Saving the Predictions in a Pickle File
# + id="-_X0rMIzw4Jl"
with open(f'{pickled_dataset}.pkl', 'wb') as f:
pickle.dump(flatOutput, f)
# + [markdown] id="Nlbu10V3c5mq"
# We don't want to have to rerun inference every time if we don't have to, so let's make sure to save the predictions for future use and experimentation.
# + [markdown] id="JB9cCF8cc2Gz"
# # Loading the Predictions from the Pickle File
# + id="8qe26VkfvX7O"
infile = open(f'{pickled_dataset}.pkl', 'rb')
zs_vcs_dict_list = pickle.load(infile)
infile.close()
# + colab={"base_uri": "https://localhost:8080/"} id="2l7lupmVwiAl" outputId="3df6ad31-135a-4ea9-ab1f-1b4e715c0fa0"
zs_vcs_dict_list[0]
# + colab={"base_uri": "https://localhost:8080/"} id="qSKSYIm5wr5Z" outputId="0c90c327-e908-422e-89a2-1d9367446ace"
len(zs_vcs_dict_list)
# + [markdown] id="KxLfcxAXhOVE"
# Now, let's prepare our data so that is can be placed in a Pandas DataFrame.
#
# Note: I excluded the last element from the loop since it's an empty string.
# + id="cR4YTbLuvvtJ"
vcs = []
scores = []
texts = []
for i in range(0, len(zs_vcs_dict_list) - 1):
vcs.append(zs_vcs_dict_list[i]['labels'])
scores.append(zs_vcs_dict_list[i]['scores'])
texts.append(zs_vcs_dict_list[i]['sequence'])
# + colab={"base_uri": "https://localhost:8080/", "height": 423} id="rO4n3M2EQBNj" outputId="10295aea-5807-4786-d04a-d1ca60e7f13c"
d = {'vcs': vcs, 'scores': scores, 'texts': texts}
df_output = pd.DataFrame(data=d)
df_output
# + [markdown] id="49rB9yWUhZcB"
# Now, we could set a scoring threshold to decide whether we want to label the table with a VC or not. For example, we could say that all scores under 0.6 should not label the text.
# + id="66K6Dgv9h6WI"
| notebooks/01_labeling_VCs_with_zero_shot_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] tags=[]
# TSG091 - Get the azdata CLI logs
# ================================
#
# Steps
# -----
#
# ### Get the azdata logs from the local machine
#
# Gets the contents of the most recent log. There may be old logs in
# azdata.log.1, azdata.log.2 etc.
# + tags=[]
import os
from pathlib import Path
home = str(Path.home())
with open(os.path.join(home, ".azdata", "logs", "azdata.log"), "r") as file:
line = file.readline()
while line:
print(line.replace("\n", ""))
line = file.readline()
# + tags=[]
print("Notebook execution is complete.")
| Big-Data-Clusters/CU14/public/content/log-files/tsg091-get-azdata-logs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import dependencies
import tweepy
import csv
import pandas as pd
# Consumer key and Access Token
consumer_key = "YOUR KEY"
consumer_secret = "YOUR KEY"
access_token = "YOUR KEY"
access_secret = "YOUR KEY"
# Create OAuthHandler object
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
# Set access token and secret token
auth.set_access_token(access_token, access_secret)
# Create Tweepy API Object to Fetch Tweets
api = tweepy.API(auth)
# Open/Create a file to append data
csvFile = open('yang.csv', 'a')
#Use csv Writer
csvWriter = csv.writer(csvFile)
# Append header to csv
header = ["Date","User Name", "Retweeted", "Favorite Count", "Retweet Count", "Tweet Text"]
with open('yang.csv',newline='') as f:
r = csv.reader(f)
data = [line for line in r]
with open('yang.csv','w',newline='') as f:
w = csv.writer(f)
w.writerow(header)
w.writerows(data)
# Fetch Tweets (& filter retweets)
for tweet in tweepy.Cursor(api.search,q="<NAME> -filter:retweets",
lang="en",
count=12000).items():
print (tweet.created_at)
print (tweet.user.name)
print (tweet.text)
print (tweet.truncated)
print ("~~~~~~~~~~~~~~~~~~~~~~~~~~")
# Append individual tweets & other relevant information to csv
csvWriter.writerow([tweet.created_at, tweet.user.name, tweet.retweeted, tweet.favorite_count, tweet.retweet_count, tweet.text.encode('utf-8')])
| tweet_sentiment_analysis/Tweets -- yang.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5.3. Accelerating array computations with Numexpr
import numpy as np
import numexpr as ne
x, y, z = np.random.rand(3, 1000000)
# %timeit x + (y**2 + (z*x + 1)*3)
# %timeit ne.evaluate('x + (y**2 + (z*x + 1)*3)')
ne.ncores
for i in range(1, 5):
ne.set_num_threads(i)
# %timeit ne.evaluate('x + (y**2 + (z*x + 1)*3)',)
| 001-Jupyter/001-Tutorials/002-IPython-Cookbook/chapter05_hpc/03_numexpr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from aide_design.play import*
from aide_design import floc_model as floc
from pytexit import py2tex
#Below are the items that were imported by the code above so that you know what abbreviations to use in your code.
# Third-party imports
#import numpy as np
#import pandas as pd
#import matplotlib.pyplot as plt
#import matplotlib
# AIDE imports
#import aide_design
#import aide_design.pipedatabase as pipe
#from aide_design.units import unit_registry as u
#from aide_design import physchem as pc
#import aide_design.expert_inputs as exp
#import aide_design.materials_database as mat
#import aide_design.utility as ut
#import aide_design.k_value_of_reductions_utility as k
#import aide_design.pipeline_utility as pipeline
#import warnings
# -
# # DC Stacked Rapid Sand Filtration
#
# The stacked rapid sand filter at Tamara, Hondruas is treating 12 L/s of water with six 20 cm deep layers of sand. The sand has an effective size of 0.5 mm and a uniformity coefficient of 1.6. The backwash velocity of the filter is 11 mm/s. Defined below are many of the necessary inputs for the filtration analysis.
# +
flow_plant = 12 * u.L / u.s
density_sand = 2650 * u.kg / u.m**3
k_Kozeny = 5
H_filter_layer = 20 * u.cm
N_filter_layer = 6
#We will use 20 deg C as the typical operating temperature. Honduras is quite warm.
T_design = 20*u.degC
# Notes on Uniformity Coefficient and Effective size from http://www.clean-water-for-laymen.com/sand-sieve.html
#Effective Size (ES) – This is defined as the size of screen opening that will pass 10% of the sand sample (see more explanation below)
#Uniformity Coefficient (UC) – This is defined as a ratio and is calculated as the size opening that will just pass 60% of the sand (d60 value) divided by the size opening that will just pass 10% of the sand sample (d10 value).
#ES is basically a value describing the average size of sand grains in a sand sample.
#UC is basically a value describing the range of grain sizes present in a sample
D_filter_sand_ES = 0.5 * u.mm
UC_filter_sand = 1.6
porosity_filter_sand = 0.4
V_filter_backwash = 11 * u.mm / u.s
g = pc.gravity
# -
# <div class="alert alert-block alert-info">
# ## Remember: don't break continuity!
#
# Ensure that you use the variables defined above in your code, do not hard code any numbers if you do not have to.
# ### 1)
# Calculate the total sand depth of all 6 sand layers.
# +
H_filter_sand_total = (H_filter_layer * N_filter_layer).to(u.m)
print('The total depth of the filter sand is', H_filter_sand_total)
# -
# ### 2)
# Calculate the diameter that is larger than 60% of the sand (D60 of the filter sand).
# +
D60_filter_sand = D_filter_sand_ES * UC_filter_sand
print('The D60 for the sand grain size is', D60_filter_sand)
# -
# ### 3)
# What is the total filter bed plan view area for both filters in Tamara?
A_filter_total = (flow_plant / V_filter_backwash).to(u.m**2)
print('The filter bed plan view area is ',A_filter_total)
# ### 4)
# What is the velocity of water through a filter during filtration? Recall that the flow through the filter is the same in filter and backwash modes.
# +
V_filter = V_filter_backwash / N_filter_layer
print('The filtration velocity is', V_filter)
# -
# ### 5)
# **Create a function** to calculate the head loss through the filter at the beginning of filtration with a clean filter bed. Then use that function to find the head loss through the clean bed of the Tamara filter. Assume that each flow path receives the same flow.
#
# Recall:
# - If you have flow paths in parallel, the headloss is NOT the sum of the head loss in each path.
# - Instead, the head loss in each path is the same as the total head loss.
#
# +
def headloss_kozeny(PorosityFilterSand, D60_filter_sand, VelocityFilter, DepthFilter):
return (36*k_Kozeny*((1 - PorosityFilterSand)**2 / PorosityFilterSand**3) * (
pc.viscosity_kinematic(T_design) * VelocityFilter)/(g * D_filter_sand_ES**2)* DepthFilter).to(u.cm)
hl_filter_clean = headloss_kozeny(porosity_filter_sand, D60_filter_sand, V_filter, H_filter_layer)
print('The headloss through the clean filter sand is', ut.sig(hl_filter_clean, 4))
# -
# ### 6)
# **Create a function** to estimate the minimum fluidization velocity for this filter bed. Then use that function to find the minimum fluidization velocity of the Tamara filter. Fluidization occurs at the beginning of backwash as all of the water flows through the bottom inlet. Note that this is not the actual velocity used for backwashing the sand.
# +
def velocity_backwash_min(porosity_sand, D60_filter_sand,T):
return ((porosity_sand**3 * g * D60_filter_sand**2) / (
36 * k_Kozeny * pc.viscosity_kinematic(T) * (1 - porosity_sand)) * (
density_sand / pc.density_water(T)-1)).to(u.mm/u.s)
print('The minimum fluidization velocity for this filter bed is', ut.sig(velocity_backwash_min(porosity_filter_sand, D60_filter_sand,T_design),2))
# -
# ### 7)
# First, plot the minimum backwash velocity as a function of water temperature from 0°C to 30°C. Then use your plot to answer the following question: if you have a water treatment plant with a single filter and there is a drought that is reducing flow to the plant, when should you backwash the filter? Double click this cell and bold the correct answer. Recall that bolding is done by placing \** around the words you wish to bold.
#
# 1. **At the time of the day when the water is coolest**
# 1. At the time of the day when the water is warmest
# +
T_graph=np.linspace(0,30,50)*u.degC
ypoints = velocity_backwash_min(porosity_filter_sand, D60_filter_sand,T_graph)
plt.plot(T_graph,ypoints,'-')
plt.xlabel(r'Temperature (°C)')
plt.ylabel('Minimum backwash velocity')
plt.title('Minimum backwash velocity vs water temperature')
plt.show()
# -
# ### 8)
# What is the residence time of water in the filter during backwash, when the bed is fluidized? You may assume the sand bed expansion ratio is 1.3.
# +
Pi_filter_backwash_expansion = 1.3
t_res_filter_backwash = ((porosity_filter_sand * H_filter_sand_total + (
Pi_filter_backwash_expansion - 1) * H_filter_sand_total
) / V_filter_backwash).to(u.s)
print('The residence time in the fluidized bed during backwash is', t_res_filter_backwash)
# -
# Our next overall goal is to determine the ratio of water wasted in a Stacked Rapid Sand Filter (SRSF) to water treated in a SRSF. Given that the backwash water that ends up above the filter bed never returns to the filter it isn't necessary to completely clear the water above the filter bed during a backwash cycle. Therefore we anticipate that backwash can be ended after approximately 3 expanded bed residence times. In addition it takes about 1 minute to initiate backwash by lowering the water level above the filter bed.
#
# ### 9)
#
# To start, estimate the time between beginning backwash and finishing the cleaning of the bed.
# +
t_filter_backwash_drain = 1 * u.min
t_filter_backwash = (t_res_filter_backwash * 3 + t_filter_backwash_drain).to(u.s)
print('The time to backwash the filter is', t_filter_backwash)
# -
# ### 10)
# Estimate the total **depth** of water that is wasted while backwash is occuring.
# +
H_filter_backwash_water = (V_filter_backwash * t_filter_backwash).to(u.m)
print('The total depth of water that is wasted is', H_filter_backwash_water)
# -
# ### 11)
# Estimate the total depth of water that is lost due to refilling the filter box at the end of backwash plus the slow refilling to the maximum dirty bed height. You may ignore the influence of plumbing head loss and you may assume that the dirty bed head loss is about 40 cm. The water level in the filter during backwash is lower than the water level at the end of filtration by both the head loss during backwash AND the headloss at the end of filtration. There is also an additional 20 cm of lost water that is required for the hydraulic controls.
#
# To reiterate, the three components that contribute to the depth of water lost in refilling the filter box after backwash are as follows:
#
# 1. Head loss during clean-bed filtration.
# 1. Difference in head loss between clean-bed filtration and dirty-bed filtration, just before backwash.
# 1. Height of the pipe that initiates backwash, also called the hydraulic control. This is actuall the pipe's diameter, since it is laying sideways in the fitler.
# +
hl_filter_final = 40 * u.cm
H_hydraulic_controls = 20*u.cm
H_filter_backwash_water_refill = H_filter_sand_total + H_hydraulic_controls + hl_filter_final
print('The total depth of water that is lost due to refilling the filter box is', H_filter_backwash_water_refill)
# -
# ### 12)
# Now calculate the total length (or depth) of water that is wasted due to backwash by adding the two previous lengths. The lengthh found in problem 10 represents water wasted while backwash is occuring, while the length in problem 11 represents the water lost in the transition to and from backwash.
H_filter_backwash_water_wasted = H_filter_backwash_water + H_filter_backwash_water_refill
print('The depth of the water that is wasted due to backwash is', H_filter_backwash_water_wasted)
# ### 13)
# Assume that the filter is backwashed every 12 hours. This means that the filter is producing clean water for 12 hours before it need to be backwashed. What is the total height (or length) of water that would be treated by the filter during this time? This length when multiplied by the area of the filter would give the total volume of water processed by a filter.
t_filter_cycle = 12 * u.hr
H_water_filtered_per_cycle = (t_filter_cycle * V_filter_backwash).to(u.m)
print('The height of water that would enter the filter in 12 hours is', H_water_filtered_per_cycle)
# ### 14)
# Finally, what is the ratio of water lost due to backwash and related water level changes in the filter box to water treated?
Pi_water_wasted = (H_filter_backwash_water_wasted / H_water_filtered_per_cycle)
print('The fraction of the total water that is lost due to backwash is', Pi_water_wasted)
# # Extra analysis that * is already completed*
# ### 15:
# Now we will evaluate the very first data set from a full scale SRSF. The performance data given below is the settled water turbidity and then the filtered water turbidity during one filter run. The time step is 5 minutes. Plot pC\* for the filter as well as effuent turbidity as a function of time on two separate graphs.
filter_influent = np.array([7.06201,7.14465,7.00537,6.33032,5.4502,4.98511,4.64221,4.23853,3.93707,3.72717,3.67126,3.55866,3.55292,3.45593,3.48163,3.50226,3.45093,3.50232,3.41095,3.55341,3.46643,3.50732,3.49146,3.51208,3.487,3.58893,3.54315,3.61469,3.58429,3.55835,3.72723,3.7829,3.74201,3.82398,3.74725,3.83423,3.72717,3.72705,3.87079,3.77338,3.70148,3.74762,3.76227,3.58875,3.63477,3.67566,3.52789,3.59296,3.66577,3.51709,3.63965,3.57843,3.47546,3.66016,3.58386,3.53259,3.57886,3.51392,3.63983,3.64972,3.64606,3.49121,3.51691,3.5119,3.61414,3.5835,3.46558,3.63965,3.60962,3.50147,3.51678,3.46039,3.49133,3.48566,3.50104,3.5943,3.47778,3.53766,3.55811,3.50635,3.42535,3.48077,3.54242,3.55274,3.59369,3.48596,3.53296,3.51746,3.45056,3.59387,3.5426,4.26868,3.99201,3.45569,3.86975,3.54407,3.49097,3.9823,3.58325,3.77789,3.70111,3.59839,4.09424,3.56769,3.83325,3.76019,3.49121,4.52917,3.63037,4.21228,3.60858,4.72827,4.00757,3.71674,3.87647,3.8288,3.44067,3.74219,3.64941,3.87439,3.79327,4.10486,4.16638,3.84418,4.11792,3.46082,3.71661,3.5061,3.48596,4.42175,3.57806,4.01294,3.63965,3.80408,3.60431,3.67572,3.61981,3.66022,3.67554,3.46076,3.72669,3.56287,3.66003,3.60004,3.4812,3.55823,6.19641,3.74146,3.88025,4.18713,4.27881,3.6496,3.45111,3.8656,3.90106,3.68597,3.66101,3.88513,3.74652,3.70123,3.79828,3.84369,3.59418,3.60968,3.49127,3.45081,3.40424,3.35852,3.32788,3.29211,3.21008,3.79279,])*u.NTU
filter_effluent = np.array([0.926376,0.645981,0.69725,0.625855,0.600449,0.472694,0.381546,0.340439,0.309883,0.289848,0.27813,0.254097,0.249432,0.253769,0.203117,0.253891,0.34063,0.223385,0.365952,0.264389,0.259193,0.340576,0.279671,0.309776,0.304878,0.279404,0.299896,0.340294,0.25399,0.350731,0.34053,0.487877,0.391518,0.309715,0.360901,0.442352,0.30967,0.391251,0.457253,0.447685,0.386322,0.549149,0.33036,0.426437,0.513458,0.279022,0.462692,0.589768,0.381073,0.532669,0.538529,0.350472,0.559036,0.51828,0.462578,0.594772,0.47242,0.538681,0.605087,0.402168,0.538818,0.66658,0.447243,0.574585,0.691544,0.513382,0.655701,0.655884,0.605408,0.666153,0.599976,0.651047,0.702515,0.92572,0.90535,0.732117,0.605026,0.849335,0.691925,0.584976,0.803314,0.783569,0.706787,0.783066,0.640701,0.732269,0.803497,0.625458,0.625687,1.00089,1.03265,1.00687,0.834213,0.732224,0.859665,0.559174,1.08862,0.93634,0.722717,0.966141,1.03278,0.890366,0.874756,0.813675,0.864746,0.874863,0.625504,0.874847,0.889893,0.931473,0.880112,1.30231,0.813965,0.961502,0.966324,0.839432,0.889969,0.930618,1.20074,0.93631,0.951584,0.78331,0.961609,0.996796,0.940994,0.966675,1.04788,0.951645,0.966888,1.01251,0.96167,1.0076,0.900436,0.996628,1.12949,0.910263,1.03256,1.04788,0.997604,1.05817,1.05801,1.06311,1.05805,1.00711,1.05811,1.11896,1.06299,1.07318,0.966049,1.07343,1.09918,1.15497,1.07303,1.13937,1.12402,1.155,1.15436,1.2056,1.14954,1.17526,1.15457,1.20569,1.17517,1.0376,0.915253,0.935471,0.884979,0.89035,0.940781,0.885071,0.874817,0.828796])*u.NTU
# +
t_delta = 5 * u.min
t_end = np.shape(filter_effluent)[0]
t_data = np.arange(0,t_end)*t_delta
c_dim = np.divide(filter_effluent,filter_influent)
filter_pc = -np.log10(c_dim)
plt.plot(t_data.to(u.hour),filter_pc)
plt.xlabel('Filter run time (hrs)')
plt.ylabel('pC*')
plt.title('Filter run time vs removal efficiency')
plt.grid(True)
plt.show()
plt.plot(t_data.to(u.hour),filter_effluent)
plt.xlabel('Filter run time (hrs)')
plt.ylabel('Effluent turbidity (NTU)')
plt.title('Filter run time vs effluent turbidity')
plt.grid(True)
plt.show()
# -
# The filter performance deteriorated over the length of the filter run. This does not match the expectations that we have based on laboratory experiments with filters. AguaClara has limited data of filter performance as a function of time. However, the [recent data from Tamara](http://aguaclara.github.io/index.html) (select Tamara from the drop down menu of plants) suggests that filtered water turbidity is consistently lower than in this first run of the filter that you plotted above.
# ### 16)
# How many kg of suspended solids per square meter of filter were removed during this filter run. Use the plan view area for the filter (don't multiply by the number of layers)
# +
M_filter_solids = (np.sum((filter_influent - filter_effluent) * (
flow_plant * t_delta)/ A_filter_total)
).to(u.kg/u.m**2)
print('The mass of the suspended solids removed is', ut.sig(M_filter_solids,3))
# -
# ### 17)
# Another useful way to express the solids capacity of the filter is to calculate the turbidty removed the run time and then express the results with units of NTU hrs. What was the capacity of the filter in NTU hrs?
# +
solids_capacity_filter = (np.sum((filter_influent-filter_effluent) * t_delta)).to(u.NTU * u.hr)
print('The filter capacity is',solids_capacity_filter,)
# -
# ### 18)
# How long was the filter run?
t_filter_cycle = t_data[np.size(t_data)-1]
print('The filter was run for', t_filter_cycle.to(u.hour))
# ### 19)
# What is the total volume of pores per square meter (plan view area) of StarS filter bed (includes all 6 layers) (in L/m^2)?
volume_filter_pores = (H_filter_sand_total * porosity_filter_sand).to(u.L/u.m **2)
print('The total volume of pores is', volume_filter_pores)
# ### 20)
# The next step is to estimate the volume of flocs per plan view area of the filter. Assume the density of the flocs being captured by the filter are approximated by the density of flocs that have a sedimentation velocity of 0.10 mm/s (slightly less than the capture velocity of the plate settlers). (see slides in flocculation notes for size of the floc and then density of that floc. I've provided this value below to simplify the analysis
density_floc = pc.density_water(T_design) + 100 * u.kg/u.m**3
density_clay = 2650 * u.kg/u.m**3
# Given the floc density, calculate fraction of floc volume that is clay.
#
# Given that floc mass is the sum of clay mass and water mass and given that floc volume is the sum of clay volume and water volume, derive an equation for the volume of flocs per plan view area of a stacked rapid sand filter (includes all 6 layers) given the floc, clay, and water densities and the mass of the clay. Show the equations that you derive using Latex
# Mass conservation gives
# $$ Vol_{Floc} \cdot \rho_{Floc} = M_{Clay} + M_{Water} $$
#
# $M_{Water}$ is an unknown.
# $$ M_{Water} = Vol_{Floc} \cdot \rho{Floc} - M_{Clay} $$
#
# Volume conservation gives
# $$ Vol_{Floc} = Vol_{Clay} + Vol_{Water} $$
#
# $$ Vol_{Floc} = \frac{M_{Clay}}{\rho_{Clay}} + \frac{M_{Water}}{\rho_{Water}} $$
#
# Substitute to eliminate $M_{Water}$
# $$ Vol_{Floc} = \frac{M_{Clay}}{\rho_{Clay}} + \frac{Vol_{Floc} \cdot \rho_{Floc}}{\rho_{Water}} -\frac{M_{Clay}}{\rho_{Water}} $$
#
# Solve for $Vol_{Floc}$
# $$ Vol_{Floc} - \frac{Vol_{Floc} \cdot \rho_{Floc}}{\rho_{Water}} = \frac{M_{Clay}}{\rho_{Clay}} - \frac{M_{Clay}}{\rho_{Water}} $$
#
# $$ Vol_{Floc}\left ( 1-\frac{\rho_{Floc}}{\rho_{Water}} \right ) = M_{Clay}\left ( \frac{1}{\rho_{Clay}} -\frac{1}{\rho_{Water}}\right ) $$
#
# $$ Vol_{Floc} = M_{Clay}\left ( \frac{\frac{1}{\rho_{Clay}}-\frac{1}{\rho_{Water}}}{ 1-\frac{\rho_{Floc}}{\rho_{Water}}} \right ) $$
#
# $$ Vol_{Floc} = { \frac{M_{Clay}\rho_{Water}}{\rho_{Floc}-\rho_{Water}}}\left ( \frac{1}{\rho_{Water}}-\frac{1}{\rho_{Clay}} \right )$$
vol_floc = ((M_filter_solids *pc.density_water(T_design)/(density_floc-pc.density_water(T_design))) *
((1/pc.density_water(T_design))-1/density_clay)).to(u.l / u.m **2)
print('The volume of the flocs per plan view area is', vol_floc)
# ### 21)
# What percent of the filter pore volume is occupied by the flocs? This fraction of pore space occupied is quite small and suggests that much of the filter bed has a very low particle concentration at the end of a filter run.
# +
Pi_flocvolume_porevolume = (vol_floc / volume_filter_pores)
print('The fraction of filter pore volume that is occupied by flocs is', ut.sig(Pi_flocvolume_porevolume,3))
# -
# # Filter constriction hypothesis
# The following analysis is completed for you and is intended to illustrate the hypothesis that flocs that are removed by the filter form a small diameter flow constriction at each place where the sand grains form a flow constriction.
# Final head loss for the filter was 50cm. Assume that this is caused by minor losses due to creation of a floc orifice (constriction) in each pore. Find the minor loss contribution by subtracting the clean bed head loss to find the head loss created by the flow constrictions that were created by the flocs.
# +
hl_filter_final = 50 * u.cm
hl_constriction = hl_filter_final-hl_filter_clean
print('The minor loss contribution is', hl_constriction)
# -
# If we assume that at the end of the filter run every pore in the filter had a flow constricting orifice from the deposition of flocs in the pore, then what was the diameter of each of the flow constrictions? We will calculate this in several steps. To begin, estimate how many flow constrictions are created by the sand grains before any flocs are added with the assumption that there is one flow constriction per sand grain. How many sand grains are there per cubic meter of filter bed? Use D60_filter_sand to estimate the number of sand grains. We will assume there is a one to one correspondence between sand grains and flow constrictions.
# +
vol_filter_sand_grain = D60_filter_sand**3 * np.pi/6
vol_filter_sand_grain_and_pore = vol_filter_sand_grain / (1 - porosity_filter_sand)
N_sand_grains = 1 /(vol_filter_sand_grain_and_pore)
print('There are this many sand grains in a cubic millimeter', N_sand_grains.to(1/u.mm**3))
# -
# Estimate the average vertical distance between flow constriction based on the cube root of the volume occupied by a sand grain
# +
L_grain_separation = (vol_filter_sand_grain_and_pore **(1/3)).to(u.mm)
print('The distance between flow constriction is', L_grain_separation)
# -
# On average, how many sand grain flow constriction does a water molecule flow through on its way through the filter?
# +
N_constriction = (H_filter_layer/L_grain_separation).to(u.dimensionless)
print('A water molecule flows through', N_constriction,'constriction through the filter')
# -
# What is the head loss per flow constriction?
hl_per_constriction = (hl_constriction / N_constriction).to(u.mm)
print('The head loss per constriction is', hl_per_constriction)
# If each constriction was partially clogged with flocs at the end of the filter run, estimate the velocity in the constriction using the expansion head loss equation. You can use the average pore water velocity as a good estimate of the expanded flow velocity.
# $$ h_{e} = \frac{(V_{in}-V_{out})^2}{2g} $$
# +
V_pore = V_filter / porosity_filter_sand
V_constriction = ((2 * g * hl_per_constriction)**(1/2) + V_pore).to(u.mm /u.s)
print('The velocity in the constriction is', V_constriction)
# -
# What is flow rate of water through each pore in μL/s? You can estimate this from the number of pores per square meter given the average separation distance.
#
# +
N_pore_per_area = 1 / L_grain_separation**2
flow_per_pore = (V_filter/ N_pore_per_area).to(u.microliter/u.s)
print('The flow rate through each pore is', flow_per_pore)
# -
# What is the inner diameter of the flow constriction created by the flocs if the vena contracta is 0.62?
Pi_vena_contracta = 0.62
A_constriction = flow_per_pore / V_constriction/Pi_vena_contracta
D_constriction = pc.diam_circle(A_constriction)
print('The inner diameter of the flow constriction created by the flocs is', D_constriction.to(u.micrometer))
# Plot the fractional removal per constriction as a function of particle size.
# +
D_clay = 7 * u.micrometer
#create an array of floc sizes from clay diameter up to the diameter of the constriction
def D_floc(D_constriction):
return (np.linspace(D_clay.to(u.micrometer), D_constriction.to(u.micrometer)))*u.micrometer
#below is an estimate of the floc removal efficiency as a function of the floc size
def c_star_constriction(D_constriction):
return ((D_constriction - D_floc(D_constriction))
/ D_constriction)**2
c_star = c_star_constriction(D_constriction)
plt.xlabel('Particle diameter(micrometer)')
plt.ylabel('Fractional remaining')
plt.title('Diameter vs fractional remaining')
plt.grid(True)
plt.plot(D_floc(D_constriction), c_star_constriction(D_constriction),'black' )
plt.show()
# -
# There are many constrictions in series and the filter fraction remaining is the pore fraction remaining raised to the power of the number of pores in series.
| DC Solutions 2017/Jupyter Notebooks/DC_Filtration_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from scipy import linalg
import scipy as sp
from sklearn import decomposition
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import train_test_split
import tensorflow as tf
import keras
from keras.layers import Dense, Embedding, LSTM, SpatialDropout1D, Dropout
from keras.models import Sequential
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.wrappers.scikit_learn import KerasClassifier
from nltk.corpus import stopwords
import pickle
import json
import gc
import re
rw = open("News_Category_Dataset_v2.json", "rb")
t = rw.readlines()
data = []
for i in range(len(t)):
data.append(json.loads(t[i].decode("utf-8").replace("\r\n", "")))
data = pd.io.json.json_normalize(data)
data.to_csv("news_dataset.csv", index=False)
def cleaner(s):
rem = re.compile('[^a-zA-Z0-9\'#-]')
s = rem.sub(" ", s).lower()
return s
data = data[data.headline.notnull()&data.short_description.notnull()]
print(len(data))
data['descrp'] = data['headline'] + " " + data['short_description']
data['descrp'] = data['descrp'].apply(lambda s: cleaner(s))
data.drop(['authors', 'link', 'headline', 'short_description', 'date'], axis=1, inplace=True)
data['descrp'].apply(lambda s: len(s.split())).value_counts(bins=10)
#Closing out similar category listings: pair[0] is old categories being closed to pair[1] new category
newcats = [(['WELLNESS', 'HEALTHY LIVING'], "WELLNESS"), (['STYLE & BEAUTY', 'HOME & LIVING', 'STYLE'], "LIVING"),
(['PARENTS', 'PARENTING', 'WEDDINGS', 'DIVORCE', 'FIFTY'], "FAMILY"), (['CRIME'], "CRIME"),
(['WORLD NEWS', 'THE WORLDPOST', 'WORLDPOST', 'POLITICS', 'RELIGION'], "POLITICS"),
(['TECH', 'SCIENCE'], "TECHNOLOGY"), (['TRAVEL', 'FOOD & DRINK', 'TASTE'], "FOOD & TRAVEL"),
(['ARTS & CULTURE', 'CULTURE & ARTS', 'ARTS'], "CULTURE"), (['COLLEGE', 'EDUCATION'], "EDUCATION"),
(['ENTERTAINMENT', 'COMEDY', 'SPORTS'], "ENTERTAINMENT"), (['ENVIRONMENT', 'GREEN'], "ENVIRONMENT"),
(['WOMEN', 'QUEER VOICES', 'BLACK VOICES', 'LATINO VOICES'], "REPRESENTATIVE VOICES"),
(['BUSINESS', 'MEDIA', 'IMPACT', 'MONEY'], "BUSINESS")]
for pair in newcats:
data.loc[data.category.isin(pair[0]), "category"] = pair[1]
data.drop(data[data.category.isin(['GOOD NEWS', 'WEIRD NEWS'])].index, inplace=True)
#Let us set our maximum sequence length to 60 since that covers about 96.7% of our data
data['descrp'] = data['descrp'].apply(lambda s: s.split())
data = data[data.descrp.apply(len) < 60]
data['descrp'] = data['descrp'].apply(lambda s: s + [" "] * (60-len(s)))
#Label-Encode the Categories
tp = data.category.value_counts().index.values
data['response'] = data.category.apply(lambda s: np.where(tp==s)[0][0])
tokenizer = keras.preprocessing.text.Tokenizer(num_words=50000, filters='!"#$%&()*+,-./:;<=>?@[\]^_`{|}~', lower=True)
tokenizer.fit_on_texts(data['descrp'].values)
wordindex = tokenizer.word_index
X = tokenizer.texts_to_sequences(data['descrp'].values)
X = np.array([i + [wordindex[' ']] * (60 - len(i)) for i in X])
Y = data.response.values.reshape(-1, 1)
Y = keras.utils.to_categorical(Y)
trainX, testX, trainY, testY = train_test_split(np.array(X), np.array(Y), random_state=1, test_size=0.3)
trainX.shape, testX.shape, trainY.shape, testY.shape
def build_mdr():
model = Sequential()
model.add(Embedding(50000, 100, input_length=X.shape[1]))
model.add(SpatialDropout1D(0.1))
model.add(LSTM(75, dropout=0.2, recurrent_dropout=0.2, return_sequences=True))
model.add(LSTM(75, dropout=0.2, recurrent_dropout=0.2, return_sequences=True))
model.add(LSTM(75, dropout=0.2, recurrent_dropout=0.2, return_sequences=False))
model.add(Dense(25, activation='relu'))
model.add(Dense(Y.shape[1], activation="softmax"))
model.compile(loss="categorical_crossentropy", optimizer="adam", metrics=["accuracy"])
return model
chk = ModelCheckpoint('./djiarnn2.h5', monitor='loss', save_best_only=True, period=10)
callbacklist = [chk]
mdl = KerasClassifier(build_fn=build_mdr, epochs=10, batch_size=500, verbose=True, callbacks=callbacklist,
validation_data=(testX, testY))
mdl.fit(trainX, trainY)
mdl.model.save("news.h5")
res = mdl.predict(testX)
sum(res==np.array([np.argmax(i) for i in testY]))
| economic-data/Google-News-Classification-LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Gabrieleas10/TELCO-Customer-Churn/blob/main/churn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="QcFoHtRs8yS8"
# importing libs
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn
from sklearn.preprocessing import LabelEncoder , StandardScaler
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split , RandomizedSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import *
from sklearn.ensemble import RandomForestClassifier
# + id="oV0uCRTN-GEx"
df = pd.read_csv('/content/WA_Fn-UseC_-Telco-Customer-Churn.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="QPiK-b7UF84r" outputId="4eac2269-fa9c-481e-a6dd-768da4783423"
print('Observations: ',df.shape[0])
print('Features: ',df.shape[1])
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="vXyO28Yw-N9y" outputId="f98a4c78-8611-42b7-9ea6-7105f1ed4778"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="2wYrFl9R-OAC" outputId="1e61b5d3-31e7-42f4-9613-f5fa7aba45be"
df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="G-Tre0XP-ODY" outputId="b5dcc05b-b4a6-48e5-9c0b-6272e27fbef7"
df.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="WTDy2XVi-f8n" outputId="bdc1e74d-1e29-4433-f4ed-d5afe432900a"
df.isnull().sum()
# + [markdown] id="zXOSXOpO-8ju"
# ## Data Exploration
# + id="MovS-rxT5aNW" colab={"base_uri": "https://localhost:8080/", "height": 567} outputId="cc46a92a-f5ff-4ac4-a316-7a288f7b81f5"
sns.pairplot(df , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="Oq9FTVe2-f-9" outputId="5e7ccf4b-2d2b-405a-94f4-a944b0a3de31"
# plot count of Senior Citizen in dataset
# more representative churn in senior citizen
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'SeniorCitizen' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="fvJJs_Z8-gCT" outputId="1fc44ce6-00fe-402b-dea1-001c60e0d7eb"
# plot count of dependents in dataset
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'Dependents' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="2J3SheJ4J_lf" outputId="c524207b-d398-45e4-d944-26f06c233a1f"
# histplot of tenure in dataset
# there is a tendency to decrease churn with increased recurrence
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.histplot(data = df , x = 'tenure' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="Vrdy5HqoSSFa" outputId="2af78609-19e1-4382-889e-e8f6b70593b5"
# histplot of tenure in dataset
# there is a tendency to decrease churn with increased recurrence
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.violinplot(data = df , y = 'tenure' , x = 'Churn', ax=ax )
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="zwZqSVAKJ_o6" outputId="7797eabc-4962-408f-dc99-b998b6ca59a8"
# plot count of phone service in dataset
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'PhoneService' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="hcKUdiAOK3dc" outputId="0fb7b701-d75a-4cb7-b6cd-fcb0a1066e61"
# plot count of multiple lines service in dataset
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'MultipleLines' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="6i68bdTHK3gr" outputId="6904e683-2212-4410-d384-a5a78e5e9973"
# plot count of contract service in dataset
# the most representative churn in month-to-month class and the lower churn in two year contract
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'Contract' , ax=ax , hue = 'Churn')
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="o7TL1ErcK3qI" outputId="1e267204-b319-4cda-9450-2a72b7f5ea59"
# plot count of payment method in dataset
# eletronic check is the class with the most churn
sns.set_theme()
fig, ax = plt.subplots(figsize= (7,5))
sns.countplot(data = df , x = 'PaymentMethod' , ax=ax , hue = 'Churn')
plt.xticks(rotation=25)
# + [markdown] id="Ck5apgX5VITP"
# ## Feature Engineering
# + id="XKwsy1vVXnAt"
# label encoding the features
le = LabelEncoder()
for col in list(df.columns):
le.fit(df[col])
df[col] = le.transform(df[col])
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="it2bp_IjjVcv" outputId="2de37cc8-c303-49c4-bb48-c8a37d321868"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="rxrmHZ-wTiy2" outputId="ff1c2f45-b421-479a-e350-f3c5c69296d5"
df.describe()
# + [markdown] id="wd5_g2eCF9uC"
# ## Principal Component Analysis (PCA)
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="t8q_Uh_k75er" outputId="5fb62240-f077-4e6a-93a3-673a487de9dc"
# features to apply PCA
X = df.drop(['customerID','Churn'] , axis =1)
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="U0ch0UQfj8CV" outputId="0dc3d015-6158-41a9-e006-79a6681aa312"
# pca
pca = PCA(n_components=2)
pca.fit(X)
components = np.round(pca.explained_variance_ratio_ , 2)
components
# + [markdown] id="bPU8ALwGGNVW"
# ## Feature Selection
# + id="CGQInCJSj8Fo"
# applying the 2 components in dataset
pca_1 = pca.transform(X)[:,0]
pca_2 = pca.transform(X)[:,1]
df['PCA1'] = pca_1
df['PCA2'] = pca_2
# + colab={"base_uri": "https://localhost:8080/", "height": 466} id="ZfTSYmcVACSy" outputId="5186dec8-6ae1-47af-a631-4c5048d264a3"
# correlation matrix
fig, ax = plt.subplots(figsize= (8,6))
cm = df.corr(method='spearman')
sns.heatmap(cm , ax= ax)
plt.show()
# + [markdown] id="TW_jgzS9GKsX"
#
# + colab={"base_uri": "https://localhost:8080/"} id="m0sUyTIhACWD" outputId="33a34063-24dd-4d2a-a171-aa289c25d775"
# spearman correlation factor
cm['Churn'].sort_values(ascending = False)
# + [markdown] id="LUiEH4h9Ek4y"
# ## Features Scale and Split
# + id="xnNSXWapACmx"
# feature selection
features = df[['Contract','tenure','OnlineSecurity','TechSupport','Dependents',
'PaperlessBilling','MonthlyCharges','DeviceProtection','SeniorCitizen']]
label = df['Churn']
# + id="7D-SRl2kBWCz"
# scalling and train/test split
scaler = sklearn.preprocessing.StandardScaler()
features = scaler.fit_transform(features)
train_features, test_features, train_labels, test_labels = train_test_split(features , label,
test_size = 0.25,
random_state = 0)
# + [markdown] id="tXXWON0tHCuM"
# ## Model and Hyperparams Selection
# + [markdown] id="sYEXz0OtOQFB"
# ### **Logistic Regression**
# + id="jBy37k5yHIFa"
# build the params grid of logistic regression
params_grid = {'penalty':['l1', 'l2', 'elasticnet'],
'solver':['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'],
'max_iter':[25,50,100,200,500]}
# + id="SVclQGXcKoV_"
# build classifier
clf = LogisticRegression()
# + id="NVlcm9XdHIIy"
# run randomized search
n_search = 100
random_search = RandomizedSearchCV(clf, param_distributions = params_grid,
n_iter = n_search)
# + colab={"base_uri": "https://localhost:8080/"} id="xHVPf486LLEy" outputId="d5001fbc-1490-4dd2-9ba6-84b686ed3d29"
# search for best hyperparams
random_search.fit(train_features, train_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="8hoUE9yOLLGa" outputId="75f1b657-fba4-4a8c-bfa5-ecc260f79b03"
random_search.best_params_
# + id="t56tJE7NLLJ8"
clf = LogisticRegression(max_iter = random_search.best_params_['max_iter'],
penalty = random_search.best_params_['penalty'],
solver = random_search.best_params_['solver'])
clf.fit(train_features, train_labels)
predictions = clf.predict(test_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="DhhZYKwJOsTm" outputId="2bd28807-92f0-4935-aeec-e2c46c84dfd9"
# confusion matrix
cfm = confusion_matrix(test_labels , predictions)
sns.heatmap(cfm, annot=True , cmap ='cividis')
plt.title('Confusion Matrix' , size = 15 , weight = 'bold')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="AJkl2yGlMzja" outputId="63555d8c-75f7-485d-d1dc-2e7ad76d1445"
# evaluate the classifier model
acc = accuracy_score(test_labels , predictions)
f1 = f1_score(test_labels , predictions)
rcall = recall_score(test_labels , predictions)
print('Logistic Regression Accuracy: ', str(round(acc,3)*100) , ' %')
print('Logistic Regression F1 Score: ', str(round(f1,3)))
print('Logistic Regression Recall: ', str(round(rcall,3)))
# + [markdown] id="fdQT7e92dVRU"
# ### **Random Forest Classifier**
# + id="E9Hvlv6QdUrY"
# build the params grid of random forest classifier
params_grid = {'n_estimators':[25,50,75,150,200,250,300],
'criterion':['gini', 'entropy'],
'max_depth':[5,10,50,100,150,200]}
# + id="1wpgWrAjdeT_"
# build classifier
clf = RandomForestClassifier()
# + id="3BQ4K2U9deXg"
# run randomized search
n_search = 100
random_search = RandomizedSearchCV(clf, param_distributions = params_grid,
n_iter = n_search)
# + colab={"base_uri": "https://localhost:8080/"} id="L2VCK35Sh0ss" outputId="001536f5-8b64-4f71-e727-b3e286efba99"
# search for best hyperparams
random_search.fit(train_features, train_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="nw_pzj1vh656" outputId="13d5ba7e-68c9-4ee3-c5ca-0ee9294bd1e2"
random_search.best_params_
# + id="_qXr81yxh68w"
clf = RandomForestClassifier(n_estimators = random_search.best_params_['n_estimators'],
criterion = random_search.best_params_['criterion'],
max_depth = random_search.best_params_['max_depth'])
clf.fit(train_features, train_labels)
predictions = clf.predict(test_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="NaLjHFQuh7AO" outputId="d61abde2-0018-49c1-9fc5-253b4dce8451"
# confusion matrix
cfm = confusion_matrix(test_labels , predictions)
sns.heatmap(cfm, annot=True , cmap ='cividis')
plt.title('Confusion Matrix' , size = 15 , weight = 'bold')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="8dq4MZvDlIU5" outputId="813967f8-9b57-4005-b38c-6e29b5519c91"
# evaluate the classifier model
acc = accuracy_score(test_labels , predictions)
f1 = f1_score(test_labels , predictions)
rcall = recall_score(test_labels , predictions)
print('Logistic Regression Accuracy: ', str(round(acc,3)*100) , ' %')
print('Logistic Regression F1 Score: ', str(round(f1,3)*100) , ' %')
print('Logistic Regression Recall: ', str(round(f1,3)*100) , ' %')
| churn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # YouTube on Android
#
# The goal of this experiment is to run Youtube videos on a Pixel device running Android and collect results.
from conf import LisaLogging
LisaLogging.setup()
# +
# %pylab inline
import json
import os
# Support to access the remote target
import devlib
from env import TestEnv
# Import support for Android devices
from android import System, Screen, Workload
# Support for trace events analysis
from trace import Trace
# Suport for FTrace events parsing and visualization
import trappy
import pandas as pd
import sqlite3
# -
# ## Support Functions
# This function helps us run our experiments:
def experiment():
# Configure governor
target.cpufreq.set_all_governors('sched')
# Get workload
wload = Workload(te).getInstance(te, 'YouTube')
# Run Youtube workload
wload.run(te.res_dir, 'https://youtu.be/XSGBVzeBUbk?t=45s',
video_duration_s=60, collect='ftrace')
# Dump platform descriptor
te.platform_dump(te.res_dir)
# ## Test environment setup
# For more details on this please check out **examples/utils/testenv_example.ipynb**.
# **devlib** requires the ANDROID_HOME environment variable configured to point to your local installation of the Android SDK. If you have not this variable configured in the shell used to start the notebook server, you need to run a cell to define where your Android SDK is installed or specify the ANDROID_HOME in your target configuration.
# In case more than one Android device are conencted to the host, you must specify the ID of the device you want to target in **my_target_conf**. Run **adb devices** on your host to get the ID.
# Setup target configuration
my_conf = {
# Target platform and board
"platform" : 'android',
"board" : 'pixel',
# Device
#"device" : "FA6A10306347",
# Android home
"ANDROID_HOME" : "/usr/local/google/home/kevindubois/Android/Sdk",
# Folder where all the results will be collected
"results_dir" : "Youtube_example",
# Define devlib modules to load
"modules" : [
'cpufreq' # enable CPUFreq support
],
# FTrace events to collect for all the tests configuration which have
# the "ftrace" flag enabled
"ftrace" : {
"events" : [
"sched_switch",
"sched_wakeup",
"sched_wakeup_new",
"sched_overutilized",
"sched_load_avg_cpu",
"sched_load_avg_task",
"cpu_capacity",
"cpu_frequency",
"clock_enable",
"clock_disable",
"clock_set_rate"
],
"buffsize" : 100 * 1024,
},
# Tools required by the experiments
"tools" : [ 'trace-cmd', 'taskset'],
}
# Initialize a test environment using:
te = TestEnv(my_conf, wipe=False)
target = te.target
# ## Workloads execution
#
# This is done using the **experiment** helper function defined above which is configured to run a **Youtube** experiment.
# Intialize Workloads for this test environment
results = experiment()
# ## Benchmarks results
# +
# Benchmark statistics
db_file = os.path.join(te.res_dir, "framestats.txt")
# !sed '/Stats since/,/99th/!d;/99th/q' {db_file}
# For all results:
# # !cat {results['db_file']}
# -
# ## Traces visualisation
#
# For more information on this please check **examples/trace_analysis/TraceAnalysis_TasksLatencies.ipynb**.
# +
# Parse all traces
platform_file = os.path.join(te.res_dir, 'platform.json')
with open(platform_file, 'r') as fh:
platform = json.load(fh)
trace_file = os.path.join(te.res_dir, 'trace.dat')
trace = Trace(trace_file, my_conf['ftrace']['events'], platform)
trappy.plotter.plot_trace(trace.ftrace)
# -
try:
trace.analysis.frequency.plotClusterFrequencies();
logging.info('Plotting cluster frequencies for [sched]...')
except: pass
trace.analysis.frequency.plotClusterFrequencies()
trace.analysis.frequency.plotPeripheralClock(title="Bus Clock", clk="bimc_clk")
| ipynb/deprecated/examples/android/workloads/Android_YouTube.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# One Machine against Infinite Bus (OMIB) simulation with [PowerSimulationsDynamics.jl](https://github.com/NREL-SIIP/PowerSimulationsDynamics.jl)
# **Originally Contributed by**: <NAME> and <NAME>
# # Introduction
# This tutorial will introduce you to the functionality of `PowerSimulationsDynamics`
# for running power system dynamic simulations.
# This tutorial presents a simulation of a two-bus system with an infinite bus
# (represented as a voltage source behind an impedance) at bus 1, and a classic
# machine on bus 2. The perturbation will be the trip of one of the two circuits
# (doubling its resistance and impedance) of the line that connects both buses.
# ## Dependencies
using SIIPExamples #hide
using PowerSimulationsDynamics
using PowerSystems
using Sundials
using Plots
gr()
# `PowerSystems` (abbreviated with `PSY`) is used to properly define the data structure and establish an equilibrium
# point initial condition with a power flow routine, while `Sundials` is
# used to solve the problem defined in `PowerSimulationsDynamics`.
# ## Load the system
# _The following command requires that you have executed the
# [dynamic systems data example](../../notebook/2_PowerSystems_examples/loading_dynamic_systems_data.jl.ipynb)
# previously to generate the json file._
file_dir = joinpath(
dirname(dirname(pathof(SIIPExamples))),
"script",
"4_PowerSimulationsDynamics_examples",
"Data",
)
omib_sys = System(joinpath(file_dir, "omib_sys.json"))
# ## Build the simulation and initialize the problem
# The next step is to create the simulation structure. This will create the indexing
# of our system that will be used to formulate the differential-algebraic system of
# equations. To do so, it is required to specify the perturbation that will occur in
# the system. `PowerSimulationsDynamics` supports three types of perturbations:
# - Network Switch: Change in the Y-bus values.
# - Branch Trip: Disconnects a line from the system.
# - Change in Reference Parameter
# Here, we will use a Branch Trip perturbation, that is modeled by modifying the
# specifying which line we want to trip. In this case we disconnect one of the lines
# that connects BUS 1 and BUS 2, named "BUS 1-BUS 2-i_1".
# With this, we are ready to create our simulation structure:
time_span = (0.0, 30.0)
perturbation_trip = BranchTrip(1.0, "BUS 1-BUS 2-i_1")
sim = Simulation(pwd(), omib_sys, time_span, perturbation_trip)
# This will automatically initialize the system by running a power flow
# and update `V_ref`, `P_ref` and hence `eq_p` (the internal voltage) to match the
# solution of the power flow. It will also initialize the states in the equilibrium,
# which can be printed with:
print_device_states(sim)
# To examine the calculated initial conditions, we can export them into a dictionary:
x0_init = get_initial_conditions(sim)
# ## Run the Simulation
# Finally, to run the simulation we simply use:
execute!(
sim, #simulation structure
IDA(), #Sundials DAE Solver
dtmax = 0.02,
); #Arguments: Maximum timestep allowed
# In some cases, the dynamic time step used for the simulation may fail. In such case, the
# keyword argument `dtmax` can be used to limit the maximum time step allowed for the simulation.
# ## Exploring the solution
# After execution, `sim` will contain the solution.
# - `sim.solution.t` contains the vector of time
# - `sim.solution.u` contains the array of states
sim.solution
# In addition, `PowerSimulationsDynamics` has two functions to obtain different
# states of the solution:
# - `get_state_series(sim, ("generator-102-1", :δ))`: can be used to obtain the solution as
# a tuple of time and the required state. In this case, we are obtaining the rotor angle `:δ`
# of the generator named `"generator-102-1"`.
angle = get_state_series(sim, ("generator-102-1", :δ));
Plots.plot(angle, xlabel = "time", ylabel = "rotor angle [rad]", label = "rotor angle")
# - `get_voltagemag_series(sim, 102)`: can be used to obtain the voltage magnitude as a
# tuple of time and voltage. In this case, we are obtaining the voltage magnitude at bus 102
# (where the generator is located).
volt = get_voltagemag_series(sim, 102);
Plots.plot(volt, xlabel = "time", ylabel = "Voltage [pu]", label = "V_2")
# ## Optional: Small Signal Analysis
# `PowerSimulationsDynamics` uses automatic differentiation to compute the reduced Jacobian
# of the system for the differential states. This can be used to analyze the local stability
# of the linearized system. We need to re-initialize our simulation:
# +
sim2 = Simulation(pwd(), omib_sys, time_span, perturbation_trip)
small_sig = small_signal_analysis(sim2)
# -
# The `small_sig` result can report the reduced jacobian for ``\delta`` and ``\omega``,
small_sig.reduced_jacobian
# and can also be used to report the eigenvalues of the reduced linearized system:
small_sig.eigenvalues
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| notebook/4_PowerSimulationsDynamics_examples/01_omib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to remove motion noise from K2 data using the `SFFCorrector`?
# You can use `lightkurve` to remove the spacecraft motion noise from K2 data. Targets in K2 data move over multiple pixels during the exposure due to thruster firings. This can be corrected using the Self Flat Fielding method (SFF), which you can read more about [here](https://docs.lightkurve.org/tutorials/04-replicate-vanderburg-2014-k2sff).
#
# This tutorial demonstrates how you can apply the method on your light curves Using `lightkurve`.
#
# Let's start by downloading a K2 light curve of an exoplanet host star.
# +
from lightkurve import search_lightcurvefile
lcf = search_lightcurvefile('EPIC 247887989').download() # returns a KeplerLightCurveFile
lc = lcf.PDCSAP_FLUX # returns a KeplerLightCurve
# Remove nans and outliers
lc = lc.remove_nans().remove_outliers()
# Remove long term trends
lc = lc.flatten(window_length=401)
# -
lc.scatter();
# This light curve of the object [K2-133](https://exoplanetarchive.ipac.caltech.edu/cgi-bin/DisplayOverview/nph-DisplayOverview?objname=K2-133+b&type=CONFIRMED_PLANET), which is known to host an exoplanet with a period of 3.0712 days. The light curve shows a lot of motion noise on K2's typical 6-hour motion timescale.
#
# Let's plot the folded version of it to see what the signal of the known transiting exoplanet looks like.
lc.fold(period=3.0712).scatter();
# We can see the hint of an exoplanet transit close to the center, but the motion of the spacecraft has made it difficult to make out above the noise.
#
# We can use the `SFFCorrector` class to remove this motion. An in-depth look into how the algorithm works can be found [here](https://docs.lightkurve.org/tutorials/04-replicate-vanderburg-2014-k2sff.html). You can tune the algorithm using a number of optional keywords, including:
#
# * `degree` : *int*
# The degree of polynomials in the splines in time and arclength.
# * `niters` : *int*
# Number of iterations
# * `bins` : *int*
# Number of bins to be used to create the piece-wise interpolation of arclength vs flux correction.
# * `windows` : *int*
# Number of windows to subdivide the data. The SFF algorithm is run independently in each window.
#
# This [tutorial](https://docs.lightkurve.org/tutorials/04-replicate-vanderburg-2014-lightkurve.html) will teach you more about how to tune these parameters. For this problem, we will use the defaults, but increase the number of windows to 20.
corr_lc = lc.to_corrector("sff").correct(windows=20)
# Note: this is identical to the following command:
from lightkurve import SFFCorrector
corr_lc = SFFCorrector(lc).correct(windows=20)
# Now when we compare the two light curves we can see the clear signal from the exoplanet.
ax = lc.fold(period=3.0712).scatter(color='red', alpha=0.5, label='Original light curve')
ax = corr_lc.fold(period=3.0712).scatter(ax=ax, color='blue', alpha=0.5, label='Motion noise removed using SFF');
# Voilà! Correcting motion systematics can vastly improve the signal-to-noise ratio in your lightcurve.
| docs/source/tutorials/04-how-to-detrend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tabular data handling
# This module defines the main class to handle tabular data in the fastai library: [`TabularDataBunch`](/tabular.data.html#TabularDataBunch). As always, there is also a helper function to quickly get your data.
#
# To allow you to easily create a [`Learner`](/basic_train.html#Learner) for your data, it provides [`tabular_learner`](/tabular.learner.html#tabular_learner).
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.tabular import *
# + hide_input=true
show_doc(TabularDataBunch)
# -
# The best way to quickly get your data in a [`DataBunch`](/basic_data.html#DataBunch) suitable for tabular data is to organize it in two (or three) dataframes. One for training, one for validation, and if you have it, one for testing. Here we are interested in a subsample of the [adult dataset](https://archive.ics.uci.edu/ml/datasets/adult).
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
valid_idx = range(len(df)-2000, len(df))
df.head()
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race', 'sex', 'native-country']
dep_var = 'salary'
# The initialization of [`TabularDataBunch`](/tabular.data.html#TabularDataBunch) is the same as [`DataBunch`](/basic_data.html#DataBunch) so you really want to use the factory method instead.
# + hide_input=true
show_doc(TabularDataBunch.from_df)
# -
# Optionally, use `test_df` for the test set. The dependent variable is `dep_var`, while the categorical and continuous variables are in the `cat_names` columns and `cont_names` columns respectively. If `cont_names` is None then we assume all variables that aren't dependent or categorical are continuous. The [`TabularProcessor`](/tabular.data.html#TabularProcessor) in `procs` are applied to the dataframes as preprocessing, then the categories are replaced by their codes+1 (leaving 0 for `nan`) and the continuous variables are normalized.
#
# Note that the [`TabularProcessor`](/tabular.data.html#TabularProcessor) should be passed as `Callable`: the actual initialization with `cat_names` and `cont_names` is done during the preprocessing.
procs = [FillMissing, Categorify, Normalize]
data = TabularDataBunch.from_df(path, df, dep_var, valid_idx=valid_idx, procs=procs, cat_names=cat_names)
# You can then easily create a [`Learner`](/basic_train.html#Learner) for this data with [`tabular_learner`](/tabular.learner.html#tabular_learner).
# + hide_input=true
show_doc(tabular_learner)
# -
# `emb_szs` is a `dict` mapping categorical column names to embedding sizes; you only need to pass sizes for columns where you want to override the default behaviour of the model.
# + hide_input=true
show_doc(TabularList)
# -
# Basic class to create a list of inputs in `items` for tabular data. `cat_names` and `cont_names` are the names of the categorical and the continuous variables respectively. `processor` will be applied to the inputs or one will be created from the transforms in `procs`.
# + hide_input=true
show_doc(TabularList.from_df)
# + hide_input=true
show_doc(TabularList.get_emb_szs)
# + hide_input=true
show_doc(TabularList.show_xys)
# + hide_input=true
show_doc(TabularList.show_xyzs)
# + hide_input=true
show_doc(TabularLine, doc_string=False)
# -
# An object that will contain the encoded `cats`, the continuous variables `conts`, the `classes` and the `names` of the columns. This is the basic input for a dataset dealing with tabular data.
# + hide_input=true
show_doc(TabularProcessor)
# -
# Create a [`PreProcessor`](/data_block.html#PreProcessor) from `procs`.
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
show_doc(TabularProcessor.process_one)
show_doc(TabularList.new)
show_doc(TabularList.get)
show_doc(TabularProcessor.process)
show_doc(TabularList.reconstruct)
# ## New Methods - Please document or move to the undocumented section
| docs_src/tabular.data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
1lI
o0O
あ井上お氏い
aabbccddeess
氏
zz
xx
a = int(input())
tmp = input().split(' ')
b, c = int(tmp[0]), int(tmp[1])
s = input()
print('{} {}'.format(a+b+c, s))
# ABC 086 A - Product
a, b = map(int, input().split())
ab = a * b
if ab % 2:
print('Odd')
else:
print('Even')
# ABC 081 A - Placing Marbles
print(input().count('1'))
# ABC 081 B - Shift Only
# +
def are_even(data):
return all([i % 2 == 0 for i in data])
N = input()
data = list(map(int, input().split()))
count = 0
while are_even(data):
data = [i/2 for i in data]
count += 1
print(count)
# -
# ABC 087 B - Coins
# +
# %%time
A = int(input())
B = int(input())
C = int(input())
X = int(input())
count = 0
for a in range(A+1):
for b in range(B+1):
for c in range(C+1):
x = (a*500) + (b*100) + (c*50)
if x == X:
count += 1
print(count)
# +
N, A, B = map(int, input().split())
x = 0
for n in range(1, N+1):
n_sum = 0
for i in str(n):
n_sum += int(i)
if A <= n_sum <= B:
x += n
print(x)
# -
# ABC 088 B - Card Game for Two
# +
N = int(input())
A = list(map(int, input().split()))
A.sort(reverse=True)
Alice = sum(A[::2])
Bob = sum(A[1::2])
print(Alice - Bob)
# -
pip list
| python/practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.io)
# # Gestión de *URLs* y funciones de vista.
# **ADVERTENCIA:** Es necesario haber creado previamente el proyecto definido en la notebook [```01_introduccion_a_django.ipynb```](01_introduccion_a_django.ipynb) localizado en el directorio [```tutorial```](tutorial).
# ## Los scripts ```urls.py``` y ```views.py```.
#
# .
#
# ### Procesamiento de una petición en *Django*.
#
# Cuando un cliente envía una petición a una *URL* apuntando a un servidor de *Django* ocurre lo siguiente:
#
# * El servidor de *Django* creará un objeto de tipo ```request```, el cual contiene todos los datos de la petición del cliente.
# * La ruta de la petición debe de coincidir con patrón de ruta definido en algún script ```urls.py```.
# * Cada patrón de ruta está ligado a una función de vista, la cual será ejecutada cuando una petición coincida con el patrón de ruta correspondiente.
# * Las funciones de vista son definidas generalmente en los scripts ```views.py```, los cuales son importados como módulos por los scripts ```urls.py```.
# * *Django* ejecutará una función de vista ingresando al objeto ```request``` como primer argumento de dicha función.
# * La función de vista deberá de regresar un objeto ```response```, el cual será utilizado por el servidor de *Django* para transmitir una respuesta al cliente.
#
# **Nota:** Los objetos ```request``` y ```response``` serán estudiados en capítulos posteriores.
# ## Configuración del script ```urls.py``` raíz de un proyecto.
#
# Un proyecto de *Django* debe de tener al menos un script ```urls.py``` a partir del cual es posible mapear todas las *URLs* de dicho poyecto y de sus aplicaciones.
#
# La localización de este script es definida con la variable ```ROOT_URLCONF``` del script ```settings.py``` del proyecto.
# **Ejemplo:**
# * La siguiente celda importará el módulo ```settings``` del paquete ```tutorial.tutorial```.
from tutorial.tutorial import settings
# * Las siguiente celda regresará el valor de ```ROOT_URLCONF``` del proyecto ```tutorial```, el cual corresponde al módulo ```'tutorial.urls'```.
settings.ROOT_URLCONF
# * A continación se muestra el script ```tutorial/tutorial/urls.py```, el cual fue generado cuando se creó el proyecto ```tutorial```.
# * Este script define al patrón de ruta ```'admin/'```, el cual es ligado a la función ```django.contrib.admin.site.urls```.
#
# ``` python
# from django.contrib import admin
# from django.urls import path
#
# urlpatterns = [
# path('admin/', admin.site.urls),
# ]
# ```
#
# **Nota:** *Django* ofrece la funcionalidad de desplegar un panel de administración mediante el paquete ```django.contrib.admin```. Dicho panel de administración se estudiará en capítulos posteriores.
# * La siguiente celda desplegará el contenido del script ```tutorial/tutorial/urls.py```.
# %pycat tutorial/tutorial/urls.py
# ## El objeto ```urlpatterns``` de los scripts ```urls.py```.
#
# El objeto ```urlpatterns``` define una lista de reglas para patrones de ruta dentro de una *URL*.
#
# ```
# urlpattern = [<regla 1>, <regla 2>... <regla n>]
#
# ```
#
# Donde:
#
# * ```<regla i>``` es una regla que relaciona a un patrón de ruta con una función de vista .
# ## El paquete ```django.urls```.
#
# Este paquete contiene las herramientas que permitirán identificar patrones de ruta y ligarlos con funciones de vista.
#
# La siguiente liga apunta a la documentación de referencian del paquete.
#
# https://docs.djangoproject.com/en/3.1/ref/urls/
# ### La función ```django.urls.path()```.
#
# Esta función permite relacionar a un patrón de ruta dentro de un *URL* con una función.
#
# ```
# django.urls.path('<patrón>', <función>, <kwargs>, name='<nombre>')
# ```
# Donde:
#
# * ```<patrón>``` corresponde a un patrón de ruta.
# * ```<función>``` corresponde a una función de vista que será ejecutado.
# * ```<kwargs>``` corresponde a argumentos ```clave=valor```.
# * ```<nombre>``` es el nombre que se le dará a esta relación (opcional).
#
# ### La función ```django.urls.include()```.
#
# En los lenguajes de programación como en el caso de *C*, se utliza la palabra ```include``` cuando se desea importar algún módulo de forma similar a la palabra reservada de Python ```import```.
#
# En este caso, la función ```django.urls.include()``` permite referenciar a un script de tipo ```urls.py``` localizado en otro sitio del proyecto, permitiendo acceder al objeto ```urlpatterns``` de dicho script, extendiendo de ese modo los patrones de ruta.
#
# ``` python
# django.urls.include('<patrón>', include('<script url>')
# ```
#
# Donde:
#
# * ```<patrón>``` corresponde a un patrón de *URL*.
# * ```<script url>``` corresponde la localización de un un script ```urls.py``` en otro sitio del proyecto.
# ## Ejemplo de definiciones de patrones de ruta.
# A continuación se crearán las reglas de patrones de *URLs* del proyecto ```tutorial``` y de la aplicación ```tutorial/main```.
# ### El script ```tutorial/tutorial/urls.py```.
#
# El script ```src/05/tutorial_urls.py``` contiene el siguiente código:
#
# ``` python
# from django.contrib import admin
# from django.urls import include, path
#
# urlpatterns = [path('admin/', admin.site.urls),
# path('main/', include('main.urls')),]
# ```
#
# * El primer elemento de ```urlpatterns``` relaciona al patron de ruta ```admin/``` con la función ```django.contrib.admin.site.urls()```.
# * El segundo elemento de ```urlpatterns``` indica que los patrones de ruta a partir del patrón ```main/``` serán definidos por el módulo ```main.urls```, el cual corresponde al script ```tutorial/main/urls.py``` .
# * La siguiente celda sustituirá al script ```tutorial/tutorial/urls.py``` con el script ```src/tutorial_urls.py```.
# * En entornos *GNU/Linux* y *MacOS X*.
# !cp src/05/tutorial_urls.py tutorial/tutorial/urls.py
# * En entornos *Windows*.
# !copy src\05\tutorial_urls.py tutorial\tutorial\urls.py
# * La siguiente celda desplegará los cambios en el script ```tutorial\tutorial\urls.py```.
# %pycat tutorial/tutorial/urls.py
# ### El script ```tutorial/main/urls.py```.
#
# En vista de que ```tutorial/tutorial/urls.py``` cargará como módulo el contenido de
# ```tutorial/main/urls.py``` es necesario que el script de referencia exista.
#
# El script ```src/05/main_urls.py``` contiene el siguiente código:
#
# ``` python
# from django.urls import path
# from . import views
#
# urlpatterns = [path('', views.index, name="inicio"),
# path('vista', views.vista)]
# ```
#
# Este código indica lo siguiente:
#
# * Importar la función ```django.urls.path()```.
# * Importar el módulo ```views``` localizado en el mismo directorio donde se encuentra el script ```tutorial/main/urls.py``` (denotado por el punto ```.```).
# * La lista ```urlpatterns``` definen dos patrones de rutas:
# * La ruta de ```main/```, la cual se denota por una cadena de caracteres vacía ```''```, ligándola a la función ```views.index()```.
# * La ruta de ```main/vista```, la cual se denota por una cadena de caracteres ```'vista'```, ligándola a la función ```views.vista()```.
# * La siguiente celda copiará el script ```src/05/main_urls.py``` a ```tutorial/main/urls.py```.
# * En entornos *GNU/Linux* y *MacOS X*.
# !cp src/05/main_urls.py tutorial/main/urls.py
# * En entornos *Windows*.
# !copy src\05\main_urls.py tutorial\main\urls.py
# * La siguiente celda desplegará los cambios en el script ```tutorial\main\urls.py```.
# %pycat tutorial/main/urls.py
# ### El script ```tutorial/main/views.py```.
#
# El script ```tutorial/main/views.py``` es referido como un módulo en el script ```tutorial/main/urls.py```.
#
# Hasta el momento, este script no contiene ninguna función y su código se vería similar a lo siguiente:
#
# ```
# from django.shortcuts import render
#
# # Create your views here.
# ```
# * La siguiente celda desplegará el contenido del script ```tutorial/main/views.py```.
# %pycat tutorial/main/views.py
# ## Ejemplo ilustrativo sobre los scripts ```view.py```.
# ### El script ```tutorial/main/views.py```.
#
# Este script contiene las funciones de vista habilitadas para la aplicación ```main``` del proyecto ```tutorial```.
# El script ```src/05/views.py``` contiene lo siguiente:
#
# ``` python
# from django.http import HttpResponse
#
# # Create your views here.
# def index(request):
# return HttpResponse("<h1>Hola, mundo.</h1>")
#
# def vista(request):
# pass
# ```
#
# Este código realiza lo siguiente:
#
# * Importa la clase ```HttpResponse``` desde el módulo ```django.http```, la cual es la clase base de los objetos ```response```.
# * Define la función de vista ```index()```, la cual regresa un objeto ```response``` conteniendo código en *HTML*.
# * Define la función de vista ```vista()```, la cual regresa ```None```.
#
# * La siguiente celda sustituirá al script ```tutorial/main/views.py``` con el script ```src/05/views.py```.
# * En entornos *GNU/Linux* y *MacOS X*.
# !cp src/05/views.py tutorial/main/views.py
# * En entornos *Windows*.
# !copy src\05\views.py tutorial\main\views.py
# * La siguiente celda desplegará los cambios en el script ```tutorial/main/views.py```.
# %pycat tutorial/main/views.py
# ### El paquete ```django.contrib.admin```.
#
# Este paquete contiene un conjunto de herramientas capaces de desplegar una aplicación de administración de un proyecto de *Django*.
#
# La documentación de este paquete puede ser consultada en:
#
# https://docs.djangoproject.com/en/3.1/ref/contrib/admin/
#
# **Notas:**
# * Aún cuando esta la aplicación de administración es habilitada automáticamente al momento en el que se crea un proyecto, puede ser un recurso opcional.
# * La aplicación de administración requiere que se definan usuarios con los roles y permisos correspondientes. Ese tema se verá en capítulos posteriores.
# ## Arranque del servicio web.
# ### Arranque desde la *notebook*.
# %cd tutorial
# !python manage.py runserver 0.0.0.0:8000
# **Nota:** Es necesario interrumpir el *kernel* de la *notebook* para terminar la ejecución del servidor.
# ### Arranque desde una terminal.
#
# #### Carga del entorno.
#
# * En caso de que se acceda a la terminal de la máquina virtual proporcionada por *Pythonista<sup>®</sup>*, es necesario cargar el entormno con el siguiente comando.
#
# ```
# source ~/pythonista/bin/activate
# ```
# #### Inicio del servidor.
#
# * Desde la terminal ubicar el shell en el directorio ```tutorial```, en el cual se encuentra el script ```manage.py```.
#
# ```
# python manage.py runserver 0.0.0.0:8000
# ```
#
# **Nota:**
# Es necesario que el firewall de su equipo esté configurado para transmitir desde el puerto ```8000```.
# ## Resultados.
# ### La ruta ```/main/```.
#
# Al acceder a http://localhost:8000/main/ aparecerá un mensaje de encabezado desplegando ```Hola, mundo.```.
#
# El navegador desplegará algo similar a la siguiente imagen:
#
# <img src="imagenes/05/hola.png" width="400px">
# ### La ruta ```/main/vista```.
#
# Al acceder a http://localhost:8000/main/vista se mostrará un mensaje de error en vista de que el objeto que regresa la función ```tutorial.main.views.vista()``` no regresa un objeto ```response```. Debido a que *Django* está configurado en modo de depuración, desplegará los detalles del error de ejecución.
#
# El navegador desplegará algo similar a la siguiente imagen:
#
#
# <img src="imagenes/05/error.png" width="600px">
# ### Rutas no definidas.
#
# * Al acceder a cualquier otra *URL* distinta a las indicadas, se mostrará un mensaje de estado ```404```. Debido a que *Django* está configurado en modo de depuracion, desplegará los detalles y los patrones de ```URL``` válidos.
#
# En este caso, el navegador desplegará algo similar a la siguiente imagen cuando se trate de acceder a la *URL* http://localhost:8000/main/incorrecto:
#
# <img src="imagenes/05/404.png" width="600px">
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2020.</p>
| 05_urls_y_funciones_de_vista.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Much data produced is unlabeled data, data where the target vale or class is unknown. Unsupervised learning gives us the tools to find hidden structure in unlabeled data. There are many techniques, one of which is clustering of the data. [Clustering](https://en.wikipedia.org/wiki/Cluster_analysis) is a method that places objects that are more similar into the same group. One of the steps in clustering is to determine the "correct" number of cluster. There are several diagnostics for this step, three of which will be shown in this post. In doing this exercise, we will demonstrate the use of [kmeans clustering](https://en.wikipedia.org/wiki/K-means_clustering) using H2O's Python API and how to retrieve results of the modeling work from H2O.
# Our first step is to import the H2O Python library start the H2O engine. H2O.ai provides detailed [documentation](http://docs.h2o.ai/) about the Python API and the rest of H2O. When H2O is started it provides summary information about the amount of memory and number of cores being used by the H2O engine.
import h2o
import imp
from h2o.estimators.kmeans import H2OKMeansEstimator
# Start a local instance of the H2O engine.
h2o.init();
# The next step of using H2O is to parse and load data into H2O's in-memory columnar compressed storage. Today we will be using the [Iris flower data set](https://en.wikipedia.org/wiki/Iris_flower_data_set).
iris = h2o.import_file(path="https://github.com/h2oai/h2o-3/raw/master/h2o-r/h2o-package/inst/extdata/iris_wheader.csv")
# H2O provides convenient commands to understand the H2OFrame object, the data structure for data that will be used by H2O's machine learning algorithms. Because H2O is often used for very large datasets and in a cluster computing configuration information about how much the data is compressed in memory and the distribution of the data across the H2O nodes, along with standard summary statics on the data in the H2OFrame, is provided.
iris.describe()
# The iris data set is labeled into three classes; there are four measurements that were taken for each iris. While we will not be using the labeled data for clustering, it does provide us a convenient comparison and visualization of the data as it was provided. In this example I use [Seaborn](http://stanford.edu/~mwaskom/software/seaborn/) for the visualization of the data.
#
# <sub><sup>(As an aside, the approach taken here of using all the data for visualization does not scale to large datasets. One approach to dealing with large data sets is to sample the data in H2O and then transfer the sample of data to the Python environment for plotting).</sup></sub>
# +
try:
imp.find_module('pandas')
can_pandas = True
import pandas as pd
except:
can_pandas = False
try:
imp.find_module('seaborn')
can_seaborn = True
import seaborn as sns
except:
can_seaborn = False
# %matplotlib inline
if can_seaborn:
sns.set()
# -
if can_seaborn:
sns.set_context("notebook")
sns.pairplot(iris.as_data_frame(True), vars=["sepal_len", "sepal_wid", "petal_len", "petal_wid"], hue="class");
# The next step is to model the data using H2O's kmeans algorithm. We will do this across a range of cluster options and collect each H2O model object as an element in an array. In this example the initial position of the cluster centers is selected at random and the random number seed is set for reproducibility. Because H2O is designed for high performance it is quick and easy to explore many different hyper-parameter settings during modeling to find the model that best suits your needs.
results = [H2OKMeansEstimator(k=clusters, init="Random", seed=2, standardize=True) for clusters in range(2,13)]
for estimator in results:
estimator.train(x=iris.col_names[0:-1], training_frame = iris)
# There are three diagnostics that will be demonstrated to help with determining the number of clusters: total within cluster sum of squares, AIC, and BIC.
#
# Total within cluster sum of squares measures sums the distance from each point in a cluster to that point's assigned cluster center. This is the minimization criteria of kmeans. The standard guideline for picking the number of clusters is to look for a 'knee' in the plot, showing where the total within sum of squares stops decreasing rapidly. Total within cluster sum of squares can be difficult to intepret, with the criteria being to look for an arbitrary knee in the plot.
#
# With this challenge from total within cluster sum of squares, we will also use two merit statistics for determining the number of clusters. [AIC](https://en.wikipedia.org/wiki/Akaike_information_criterion) and [BIC](https://en.wikipedia.org/wiki/Bayesian_information_criterion) are both measures of the relative quality of a statistical model. AIC and BIC introduce penality terms for the number of parameters in the model to counter the problem of overfitting; BIC has a larger penality term than AIC. With these merit statistics one is to look for the number of clusters that minimize the statistic.
#
# Here we build a method for extracting the inputs for each diagnostics and calculating the AIC and BIC values on a model. Each model is then inspected by the method and the results plotted for quick analysis.
# +
import math as math
def diagnostics_from_clusteringmodel(model):
total_within_sumofsquares = model.tot_withinss()
number_of_clusters = len(model.centers()[0])
number_of_dimensions = len(model.centers())
number_of_rows = sum(model.size())
aic = total_within_sumofsquares + 2 * number_of_dimensions * number_of_clusters
bic = total_within_sumofsquares + math.log(number_of_rows) * number_of_dimensions * number_of_clusters
return {'Clusters':number_of_clusters,
'Total Within SS':total_within_sumofsquares,
'AIC':aic,
'BIC':bic}
# -
if can_pandas:
diagnostics = pd.DataFrame( [diagnostics_from_clusteringmodel(model) for model in results])
diagnostics.set_index('Clusters', inplace=True)
# From the plot below, to me, it is difficult to find a 'knee' in the rate of decrease of the total within cluster sum of square. It might be at 4 clusters, it might be 7. AIC is minimized at 7 clusters, and BIC is minimized at 4 clusters.
if can_pandas:
diagnostics.plot(kind='line');
# For demonstration purposes, I will selected the number of clusters to be 4. I will use the H2O Model for 4 clusters previously created, and use that to assign the membership in each of the original data points. This predicted cluster assignment is then added to the original iris data frames as a new vector (mostly to make plotting easy).
clusters = 4
predicted = results[clusters-2].predict(iris)
iris["Predicted"] = predicted["predict"].asfactor()
# Finally, I will plot the predicted cluster membership using the same layout as on the original data earlier in the notebook.
if can_seaborn:
sns.pairplot(iris.as_data_frame(True), vars=["sepal_len", "sepal_wid", "petal_len", "petal_wid"], hue="Predicted");
# This iPython notebook is available for [download](https://github.com/h2oai/h2o-3/blob/master/h2o-py/demos/kmeans_aic_bic_diagnostics.ipynb). Grab the [latest version of H2O](http://h2o.ai/download/) to try it out and be sure to check out other [H2O Python demos](https://github.com/h2oai/h2o-3/tree/master/h2o-py/demos).
#
# -- [Hank](http://twitter.com/hankroark)
| implementation/h2o-fakegame/h2o-py/demos/kmeans_aic_bic_diagnostics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob
fileNames = glob.glob("*.obj")
print(len(fileNames), fileNames)
fileNameWithoutExt = [fileName[:-4] for fileName in fileNames]
print(len(fileNameWithoutExt), fileNameWithoutExt)
| static/models/factoryobj/get file names.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dheerajjoshim/machinelearningcourse/blob/master/DBSCAN_ForestType_EDP_Practice.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vHsoq1bWc9UH" colab_type="text"
# # # DBSCAN - Density-Based Spatial Clustering of Applications with Noise
# + id="EamnuGxrc9UI" colab_type="code" colab={}
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
# + id="R3TSQ4ogc9UL" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 72} outputId="1bbde9b2-37a4-40b6-a192-b4e36f4d65eb"
# Upload to Google Colab
from google.colab import files
uploaded = files.upload()
# + id="8xCK5pJNc9UO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="e209c051-292e-47a1-a402-20386b6abfe8"
# Read the dataset
df = pd.read_csv("ForestType.csv")
df.head()
# + id="6P1CKp4Rc9UR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="83e399de-6e39-4d75-95d5-e9529dddf003"
df.shape
# + id="OV4iexw7c9UT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="813ee362-bb36-444d-c169-fd9b48e98063"
df.info()
# + id="-BFv4RdJc9UW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="775b64bc-cfc7-48b7-d6ac-c636a25e033e"
# Check for Null values
df.isnull().sum()
# + id="mYbyEXu1c9UY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="625380c4-4f80-41e7-d19a-ece890abf358"
# have a look on the dataset details
df.describe()
# + id="RRJ9vNXVc9Ua" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="f5b175cc-84fe-4a1e-82eb-f2c572b8b718"
# Outlier check
df[ df['b3'] > 63.676768 + 3*17.314545 ]
# + [markdown] id="CrmrLsVXc9Ud" colab_type="text"
# ### Scatter plot data
# + id="EAnCcx9Nc9Ud" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="d0602d96-fe98-4d3a-9cbd-79087662c3d2"
# plt.scatter(df.b1,df.b2,df.b3)
plt.scatter(df.b1,df.b2)
# + [markdown] id="mzi2DjkBc9Uf" colab_type="text"
# ### Call DBSCAN method passing epsilon and min sample as argument to form a cluster
# + id="gKF5KTUTc9Ug" colab_type="code" colab={}
from sklearn.cluster import DBSCAN
model = DBSCAN(eps = 6.5 , min_samples=5).fit(df)
# model = DBSCAN(eps = 6.7).fit(df)
# + id="xTBJAvD9c9Ui" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7f38d44d-43a3-434b-f714-294a6a77a6c4"
labels=model.labels_
print(type(labels))
# + [markdown] id="hAdLr4Vpc9Uk" colab_type="text"
# ### # DBSCAN forms cluster automattically, check how many clusters are formed
# + id="uuosez9Uc9Ul" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6c7ee565-d5d0-4782-f574-a079c5e25e1a"
print(len(set(labels)))
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
print(n_clusters_)
# + [markdown] id="aODkvO0Qc9Un" colab_type="text"
# ### # Check clusters lables
# + id="B8P4cWoTc9Un" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="9c1d0c79-60fb-4c4e-9b15-d54ef45d5137"
labels=model.labels_
print(labels)
# + [markdown] id="V0CTg-oWc9Up" colab_type="text"
# ## Create dataframe for each clusters and outlier
# + id="TTxgjUOtc9Up" colab_type="code" colab={}
df1 = df[model.labels_ == 0]
df2 = df[model.labels_ == 1]
df3 = df[model.labels_ == 2]
#df4 = df[model.labels_ == 3]
df5 = df[model.labels_ == -1]
# + [markdown] id="VwsvRh-jc9Ur" colab_type="text"
# ## Scatter plot all clusters
# + id="SQNSlfAlc9Us" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="02f4f645-6d79-4b63-97e1-03518d395924"
plt.scatter(df1.b1,df1.b2, df1.b3, color='green')
plt.scatter(df2.b1,df2.b2, df2.b3, color='red')
plt.scatter(df3.b1,df3.b2, df3.b3, color='blue')
#plt.scatter(df4.b1,df4.b2, df4.b3, color='yellow')
plt.scatter(df5.b1,df5.b2, df5.b3, color='black')
# + [markdown] id="c5K2SX8uc9Uu" colab_type="text"
# ## End
# + [markdown] id="_8MyV-rgc9Uv" colab_type="text"
# ### Generic python code to choose best value of eps and minimum samples
# + id="os8ltgjkc9Uv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="a57eae5c-38ae-48bc-cce6-466c58961444"
# Find optimal value of eps
for ep in np.arange(1,10):
print("ep - " + str(ep))
model = DBSCAN(eps = ep, min_samples=4).fit(df)
labels=model.labels_
n_clusters_ = len(set(labels)) - (1 if -1 in labels else 0)
n_noise_ = list(labels).count(-1)
print (" eps : "+str(ep) +" No. of cluster : "+str(n_clusters_) + " Outliers : "+str(n_noise_))
# + id="ba85ZJpnc9Uy" colab_type="code" colab={}
| DBSCAN_ForestType_EDP_Practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Lists
#
# Earlier when discussing strings we introduced the concept of a *sequence* in Python. Lists can be thought of the most general version of a *sequence* in Python. Unlike strings, the are mutable, meaning the elements inside a list can be changed!
#
# In this section we will learn about:
#
# 1.) Creating lists
# 2.) Indexing and Slicing Lists
# 3.) Basic List Methods
# 4.) Nesting Lists
# 5.) Introduction to List Comprehensions
#
# Lists are constructed with brackets [] and commas seperating every element in the list.
#
# Let's go ahead and see how we can construct lists!
# Assign a list to an variable named my_list
my_list = [1,2,3]
# We just created a list of integers, but lists can actually hold different object types. For example:
my_list = ['A string',23,100.232,'o']
# Just like strings, the len() frunction will tell you how many items are in the sequence of the list.
len(my_list)
# ### Indexing and Slicing
# Indexing and slicing works just like in strings. Let's make a new list to remind ourselves of how this works:
my_list = ['one','two','three',4,5]
# Grab element at index 0
my_list[0]
# Grab index 1 and everything past it
my_list[1:]
# Grab everything UP TO index 3
my_list[:3]
# We can also use + to concatenate lists, just like we did for strings.
my_list + ['new item']
# Note: This doesn't actually change the original list!
my_list
# You would have to reassign the list to make the change permanent.
# Reassign
my_list = my_list + ['add new item permanently']
my_list
# We can also use the * for a duplication method similar to strings:
# Make the list double
my_list * 2
# Again doubling not permanent
my_list
# ## Basic List Methods
#
# If you are familiar with another programming language, you might start to draw parallels between arrays in another language and lists in Python. Lists in Python however, tend to be more flexible than arrays in other languages for a two good reasons: they have no fixed size (meaning we don't have to specify how big a list will be), and they have no fixed type constraint (like we've seen above).
#
# Let's go ahead and explore some more special methods for lists:
# Create a new list
l = [1,2,3]
# Use the **append** method to permanently add an item to the end of a list:
# Append
l.append('append me!')
# Show
l
# Use **pop** to "pop off" an item form the list. By default pop takes off the last index, but you can also specify whic index to pop off. Let's see an example:
# Pop off the 0 indexed item
l.pop(0)
# Show
l
# Assign the popped element, remember default popped index is -1
popped_item = l.pop()
popped_item
# Show remaining list
l
# It should also be noted that lists indexing will return an error if there is no element at that index. For example:
l[100]
# We can use the **sort** method and the **reverse** methods to also effect your lists:
new_list = ['a','e','x','b','c']
#Show
new_list
# Use reverse to reverse order (this is permanent!)
new_list.reverse()
new_list
# Use sort to sort the list (in this case alphabetical order, but for numbers it will go ascending)
new_list.sort()
new_list
# ## Nesting Lists
# A great feature of of Python data structures is that they support *nesting*. This means we can data structures within data structures. For example: A list inside a list.
#
# Let's see how this works!
# +
# Let's make three lists
lst_1=[1,2,3]
lst_2=[4,5,6]
lst_3=[7,8,9]
# Make a list of lists to form a matrix
matrix = [lst_1,lst_2,lst_3]
# -
# Show
matrix
# Now we can again use indexing to grab elements, but now there are two levels for the index. The items in the matrix object, and then the items inside that list!
# Grab first item in matrix object
matrix[0]
# Grab first item of the first item in the matrix object
matrix[0][0]
# # List Comprehensions
# Python has an advanced feature called list comprehensions. They allow for quick construction of lists. To fully understand list comprehensions we need to understand for loops. So don't worry if you don't completely understand this section, and feel free to just skip it since we wil return to this topic later.
#
# But in case you want to know now, here are a few examples!
# Build a list comprehension by deconstructing a for loop within a []
first_col = [row[0] for row in matrix]
first_col
# We used list comprehension here to grab the first element of every row in the matrix object. We will cover this in much more detail later on!
#
# For more advanced methods and features of lists in Python, check out the advanced list section later on in this course!
| Django2.0-Python-Full-Stack-Web-Developer/TravlersDayOneandTwo/2- Python Overview/2 - Statements Methods Functions/Lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%% a collection of of notebook computations associated with v2e\n", "is_executing": false}
# + pycharm={"name": "#%% plot lin-log\n", "is_executing": false}
import numpy as np
from matplotlib import pyplot as plt
from v2e.emulator import lin_log
y=np.array(range(255), float)
l=lin_log(y,threshold=5)
fnt=16
plt.subplot(211)
plt.rcParams.update({'font.size': fnt})
plt.plot(y,l)
plt.xlabel('log Y [DN]')
plt.ylabel('L')
plt.tight_layout()
plt.xscale('log')
plt.subplot(212)
plt.rcParams.update({'font.size': fnt})
plt.plot(y,l)
plt.xlabel('Y [DN]')
plt.ylabel('L')
plt.tight_layout()
plt.xscale('linear')
plt.savefig('../output/linlog.pdf')
plt.savefig('../media/linlog.png')
# + pycharm={"name": "#%% noise sample\n", "is_executing": false}
import numpy as np
from matplotlib import pyplot as plt
t=np.array(range(100),float)
n=np.random.randn(len(t))
fig=plt.figure(3)
plt.plot(t,n)
plt.axis('off')
plt.savefig('../output/noise.pdf')
# + pycharm={"name": "#%% erf function for noise rate - (scheme not currently used in code, instead uses Poisson generator)\n", "is_executing": false}
# compute the expected number of temporal noise events per sample for a
# given sigma of gaussian noise sig and threshold th
import numpy as np
from math import erf, sqrt
# https://docs.python.org/3.2/library/math.html
def phi(x):
'Cumulative distribution function for the standard normal distribution'
return (1.0 + erf(x / sqrt(2.0))) / 2.0
RnHzDesired=0.1 # desired rate of noise events per second
fsHz=100 # sample rate Hz
theta=.15 # event threshold in log_e units, but it doesn't matter what it is
# dumb search for best sigma
for sig in np.linspace(0,theta,1000):
n=0
for k in range(1,100):
th0=k*theta
th1=(k+1)*theta
n+=k*(phi(th1/sig)-phi(th0/sig)) # weight expected # events times probability
Rn=fsHz*n
if Rn>RnHzDesired:
print('sig={:6.2f}\tn={:8.3g}ev/samp\tRn={:8.3g}ev/s'.format(sig,n,Rn))
print('desired rate: {}'.format(RnHzDesired))
break
# + pycharm={"name": "#%% compute time constants and update factors for lowpass filter\n", "is_executing": false}
import numpy as np
cutoffhz=10.
tau=1/(2*np.pi*cutoffhz)
fs=300.
dt=1/fs
eps=dt/tau
# + pycharm={"name": "#%% 2nd order lowpass for Fig. 1\n", "is_executing": false}
import numpy as np
f3dbhz=100.
f=np.logspace(-2,4,100)
s=2*np.pi*f
tau=1/(2*np.pi*f3dbhz)
a=tau*s+1
b=a*a
h=np.reciprocal(b)
from matplotlib import pyplot as plt
plt.rcParams.update({'font.size': 16})
fig=plt.figure(5)
plt.plot(f,h)
plt.yscale('log')
plt.xscale('log')
plt.xlabel('f [Hz]')
plt.ylabel('|H|')
plt.savefig('../output/lowpass.pdf')
# + pycharm={"is_executing": false, "name": "#%% simulate effect of lowpass and dark current on photoreceptor output for figure 3\n"}
import numpy as np
from scipy.signal import square
dark=1 # dark current level
sigdc1=10 # DC photocurrent for bright half
sigdc2=1 # and dark half
cont=1 # contrast in each half
tau=.01 # time constant of lowpass
nper=4 # how many periods to simulate for each half bright/dark
thr=.1 # dvs threshold
dt=np.pi/500 # timesteps per half period
eps=dt/tau
if eps>1:
print('eps={:.3g} for tau={}, increase the number of time steps per cycle'.format(eps,tau))
eps=1
t=np.arange(np.pi,nper*2*2*np.pi,dt)
npts=len(t)
npts2=int(npts/2)
sq=(square(t)+1)/2
sq=np.convolve(sq,np.ones(10)/10,mode='same')
sig=np.zeros_like(sq)
sig[:npts2]=sigdc1*(1+cont*sq[:npts2])
sig[npts2:]=sigdc2*(1+cont*sq[npts2:])
noiseFactor=.2
noise=noiseFactor*np.random.randn(t.shape[0])*np.sqrt(sig)
sig=sig+noise
darknoise=dark+noiseFactor*np.random.randn(t.shape[0])*np.sqrt(dark)
cur=sig+darknoise
logcur=np.log(cur) # log photoreceptor on sum of signal + dark current
ph1=np.zeros_like(logcur)
sf=np.zeros_like(logcur)
ph1[0]=logcur[0]
sf[0]=logcur[0]
for i in range(0,len(t)-1):
tauactual=tau*max(cur)/cur[i] # nonlinear lowpass with time const tau for largest current
e=dt/tauactual
ph1[i+1]=(1-e)*ph1[i]+e*logcur[i] # only photorecptor tau changes
sf[i+1]=(1-eps)*sf[i]+eps*ph1[i] # src follower tau is constant
# include DVS events
lmem=sf[0]
ons=np.zeros_like(t)
offs=np.zeros_like(t)
for i in range(1,len(t)):
if sf[i]-lmem>=thr:
ons[i]=1
lmem=sf[i]
elif sf[i]-lmem<=-thr:
offs[i]=-1
lmem=sf[i]
from matplotlib import pyplot as plt
plt.rcParams.update({'font.size': 16})
fig,ax1=plt.subplots(sharex=True)
ax1.plot(t,cur,'g',t,darknoise,'g')
ax1.set_ylim([0,None])
ax1.set_yscale('linear')
ax1.set_xscale('linear')
ax1.tick_params(axis='y', colors='green')
ax2=ax1.twinx()
ax2.plot(t,sf,'r-')
ax2.set_ylim([0,None])
ax2.set_yscale('linear')
ax2.set_xscale('linear')
ax2.tick_params(axis='y', colors='red')
fig,ax3=plt.subplots(sharex=True)
offset=8
ax3.plot(t,offset+ons,'g', t,offset+offs,'r')
ax3.get_xaxis().set_ticks([])
ax3.get_yaxis().set_ticks([])
ax3.set_aspect(3)
# plt.xlabel('t [s]')
# plt.ylabel('[log]')
# plt.grid('on')
plt.savefig('../output/photo_dark_lowpass_curves.pdf')
# + pycharm={"is_executing": false, "name": "#%% compute scene illuminance from chip illuminance\n"}
f=2.8 # focal length
T=0.9 # lens transmittance
R=.18 # avg scene reflectance
lchip=np.logspace(0,4,5,endpoint=True)
factor=(4*f**2/T/R)
lscene=factor*lchip
print(lchip)
print(lscene)
# + pycharm={"name": "#%% https://stackoverflow.com/questions/33933842/how-to-generate-noise-in-frequency-range-with-numpy\n", "is_executing": false}
import numpy as np
def fftnoise(f):
f = np.array(f, dtype='complex')
Np = (len(f) - 1) // 2
phases = np.random.rand(Np) * 2 * np.pi
phases = np.cos(phases) + 1j * np.sin(phases)
f[1:Np+1] *= phases
f[-1:-1-Np:-1] = np.conj(f[1:Np+1])
return (np.fft.ifft(f).real)
def band_limited_noise(min_freq, max_freq, samples=1024, samplerate=1):
freqs = np.abs(np.fft.fftfreq(samples, 1/samplerate))
f = np.zeros(samples)
idx = np.where(np.logical_and(freqs>=min_freq, freqs<=max_freq))[0]
f[idx] = 1
return fftnoise(f)
from scipy.io import wavfile
x = band_limited_noise(0, 10000, 44100, 44100)
x = np.int16(x * (2**15 - 1))
wavfile.write("test.wav", 44100, x)
mean=np.mean(x)
power=(np.mean(x*x)-mean*mean)
print('total power={}'.format(power))
from matplotlib import pyplot as plt
plt.plot(x)
plt.cla()
freqs=np.fft.fftfreq(44100,1./44100)
ps = np.abs(np.fft.fft(x))**2
idx = np.argsort(freqs)
plt.plot(freqs[idx],ps[idx])
# + pycharm={"name": "#%% load DVS events from txt file, plot distribution of spike rates for ON and OFF\n", "is_executing": false}
from v2e.v2e_utils import _inputFileDialog as ifd
fname=ifd([("AEDAT txt files", ".txt"),('Any type','*')])
from v2e.v2e_utils import read_aedat_txt_events
ev=read_aedat_txt_events(fname)
import numpy as np
sx=346
sy=260
npix=sx*sy
ntot=ev.shape[0]
non=np.count_nonzero(ev[:,3]==1)
noff=np.count_nonzero(ev[:,3]==0)
ttot=max(ev[:,0])-min(ev[:,0])
onrate=non/npix/ttot
offrate=noff/npix/ttot
onidx= np.where(ev[:,3]==1)
offidx=np.where(ev[:,3]==0)
# for each pixel, compute the spike rate
onev= ev[onidx, :][0]
offev=ev[offidx,:][0]
onx=onev[:,1]
ony=onev[:,2]
offx=offev[:,1]
offy=offev[:,2]
histrange = [(0, v) for v in (sx, sy)]
onhist,_,_= np.histogram2d(onx, ony, bins=(sx, sy), range=histrange)
offhist,_,_= np.histogram2d(offx, offy, bins=(sx, sy), range=histrange)
onhist=onhist.flatten()
offhist=offhist.flatten()
maxonhist=max(onhist)
maxoffhist=max(offhist)
onrates=np.flip(np.sort(onhist/ttot))
offrates=np.flip(np.sort(offhist/ttot))
logonrates=np.log10(onrates)
logoffrates=np.log10(offrates)
raterange=(-2,3)
nbins=100
onratehist,onbins=np.histogram(logonrates,bins=nbins, range=raterange)
offratehist,offbins=np.histogram(logoffrates,bins=nbins, range=raterange)
onratehist=onratehist
offratehist=offratehist
from matplotlib import pyplot as plt
plt.plot(onbins[:-1],(onratehist),'g',offbins[:-1],-(offratehist),'r')
plt.xlabel('event rate (log10(Hz))')
plt.ylabel('frequency')
figfname=fname.replace('.txt','.pdf')
plt.savefig(figfname)
# + pycharm={"name": "#%% simulate noise and bandwidth of noise\n"}
import numpy as np
from engineering_notation import EngNumber as eng
dt=1e-4 #
fs=1/dt
totaltime=30
t=np.arange(0,totaltime,dt)
npts=len(t)
def blnoise(f3db,t):
dc=1 # and dark half
tau=1/(2*np.pi*f3db) # time constant of lowpass
eps=dt/tau
if eps>1:
print('eps={:.3g} for tau={}, decrease timestep dt'.format(eps,tau))
eps=1
sig=dc*np.ones_like(t)
noiseFactor=.1
noise=noiseFactor*np.random.randn(t.shape[0])*np.sqrt(dc)
sig=sig+noise
logcur=np.log(sig) # log photoreceptor on sum of signal + dark current
ph1=np.zeros_like(logcur)
# sf=np.zeros_like(logcur)
ph1[0]=np.log(dc)
# sf[0]=ph1[0]
for i in range(0,len(t)-1):
e=eps
ph1[i+1]=(1-e)*ph1[i]+e*logcur[i] # only photorecptor tau changes
# sf[i+1]=(1-eps)*sf[i]+eps*ph1[i] # src follower tau is constant
# print('x={:.2f} y1={:.2f} y2={:.2f}'.format(logcur[i],ph1[i+1],sf[i+1]))
mean=np.mean(ph1)
power=(np.mean(ph1*ph1)-mean*mean)
ph1/=np.sqrt(power)
mean=np.mean(ph1)
power=(np.mean(ph1*ph1)-mean*mean)
print('total power={}'.format(eng(power)))
return ph1
ph1=blnoise(10,t)
ph2=blnoise(100,t)
ph3=blnoise(1000,t)
# + pycharm={"name": "#%% compute spectogram of signal\n"}
import numpy as np
import scipy.signal as signal
import matplotlib.pyplot as plt
def genev(ph,t,thr=1):
refr=5*dt
nons=0
noffs=0
tev=-refr # last event time
# # include DVS events
lmem=ph[0]
ons=np.zeros_like(t)
offs=np.zeros_like(t)
for i in range(1,len(t)):
if ph[i]-lmem>=thr:
ons[i]=1
nons+=1
lmem=ph[i]
elif ph[i]-lmem<=-thr:
offs[i]=-1
noffs+=1
lmem=ph[i]
return ons,offs,np.array(nons),np.array(noffs)
# + pycharm={"name": "#%% estimate power spectum\n"}
ons1,offs1,_,_=genev(ph1,t)
ons2,offs2,_,_=genev(ph2,t)
ons3,offs3,_,_=genev(ph3,t)
# + pycharm={"name": "#%% estimate power spectum\n"}
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.signal.welch.html#scipy.signal.welch
freqs, pow3 = signal.welch(ph3, fs, scaling='spectrum', average='median',window='boxcar')
freqs, pow2 = signal.welch(ph2, fs, scaling='spectrum', average='median',window='boxcar')
freqs, pow1 = signal.welch(ph1, fs, scaling='spectrum', average='median',window='boxcar')
# freqs=np.fft.fftfreq(len(t),1./len(t))
#%% plot results
from matplotlib import pyplot as plt
plt.rcParams.update({'font.size': 16})
plt.rcParams['pdf.fonttype'] = 42
plt.subplots(2,1,sharex=True)
plt.subplot(211)
# ax1.set_ylim([0,None])
# ax1.set_ylim((-3,3))
# ax1.tick_params(axis='y', colors='green')
# ax1.plot(t,ph3+3,'r',t,ph2,'g-',t,ph1-3,'b-')
# ax1.set_xlabel('time (s)')
# ax1.set_ylabel('signal')
plt.plot(t,ph3+3,'r',t,ph2,'g-',t,ph1-3,'b-')
plt.xlim((0,2000*dt))
plt.subplot(212)
plt.xlim((0,2000*dt))
plt.plot(t,ons3+3, t,offs3+3)
plt.plot(t,ons2, t,offs2)
plt.plot(t,ons1-3, t,offs1-3)
plt.savefig('output/noise-time.pdf')
plt.figure(2)
plt.loglog(freqs,pow3,'r-',alpha=.7)
plt.loglog(freqs,pow2,'g-',alpha=.7)
plt.loglog(freqs,pow1,'b-',alpha=.7)
plt.grid(True)
plt.xlabel('freq (Hz)')
plt.ylabel('power (sig^2/Hz)')
plt.ylim(1e-4,0)
# plt.gca().set_aspect('equal', adjustable='box')
plt.savefig('output/lognoise.pdf')
# + pycharm={"name": "#%% study haw fake event rate depends on threshold\n"}
import numpy as np
import matplotlib.pyplot as plt
thrs=np.logspace(-1,1,40)
nons=[]
noffs=[]
for thr in thrs:
_,_,non,noff=genev(ph3,t,thr)
nons.append(non)
noffs.append(noff)
# + pycharm={"name": "#%% study haw fake event rate depends on threshold\n"}
plt.figure(4)
plt.loglog(thrs,nons,thrs,noffs)
plt.xlabel('DVS threshold')
plt.ylabel('event rate (Hz)')
# + pycharm={"name": "#%% plot log normal distribution of generated bias currents for normal variation of threshold voltage\n"}
import numpy as np
import matplotlib.pyplot as plt
svt=0.1
ut=0.25
kappa=0.8
vts=np.random.normal(scale=svt,size=10000)
ids=np.exp(-kappa*vts/ut)
plt.figure(5)
plt.hist(ids,bins=100)
plt.savefig('output/lognormal.pdf')
plt.show()
# + pycharm={"name": "#%%\n"}
| v2e/v2e/computations_and_graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1YwMUyt9LHG1"
# # Generates images from text prompts with CLIP guided diffusion.
#
# By <NAME> (https://github.com/crowsonkb, https://twitter.com/RiversHaveWings). It uses OpenAI's 256x256 unconditional ImageNet diffusion model (https://github.com/openai/guided-diffusion) together with CLIP (https://github.com/openai/CLIP) to connect text prompts with images.
# + cellView="form" id="XIqUfrmvLIhg"
# @title Licensed under the MIT License
# Copyright (c) 2021 <NAME>
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
# THE SOFTWARE.
# + id="qZ3rNuAWAewx"
# Check the GPU status
# !nvidia-smi
# + id="-_UVMZCIAq_r"
# Install dependencies
# !git clone https://github.com/openai/CLIP
# !git clone https://github.com/crowsonkb/guided-diffusion
# !pip install -e ./CLIP
# !pip install -e ./guided-diffusion
# + id="7zAqFEykBHDL"
# Download the diffusion model
# !curl -OL 'https://openaipublic.blob.core.windows.net/diffusion/jul-2021/256x256_diffusion_uncond.pt'
# + id="JmbrcrhpBPC6"
# Imports
import math
import io
import sys
from IPython import display
from PIL import Image
import requests
import torch
from torch import nn
from torch.nn import functional as F
from torchvision import transforms
from torchvision.transforms import functional as TF
from tqdm.notebook import tqdm
sys.path.append('./CLIP')
sys.path.append('./guided-diffusion')
import clip
from guided_diffusion.script_util import create_model_and_diffusion, model_and_diffusion_defaults
# + id="YHOj78Yvx8jP"
# Define necessary functions
def fetch(url_or_path):
if str(url_or_path).startswith('http://') or str(url_or_path).startswith('https://'):
r = requests.get(url_or_path)
r.raise_for_status()
fd = io.BytesIO()
fd.write(r.content)
fd.seek(0)
return fd
return open(url_or_path, 'rb')
class MakeCutouts(nn.Module):
def __init__(self, cut_size, cutn, cut_pow=1.):
super().__init__()
self.cut_size = cut_size
self.cutn = cutn
self.cut_pow = cut_pow
def forward(self, input):
sideY, sideX = input.shape[2:4]
max_size = min(sideX, sideY)
min_size = min(sideX, sideY, self.cut_size)
cutouts = []
for _ in range(self.cutn):
size = int(torch.rand([])**self.cut_pow * (max_size - min_size) + min_size)
offsetx = torch.randint(0, sideX - size + 1, ())
offsety = torch.randint(0, sideY - size + 1, ())
cutout = input[:, :, offsety:offsety + size, offsetx:offsetx + size]
cutouts.append(F.adaptive_avg_pool2d(cutout, self.cut_size))
return torch.cat(cutouts)
def spherical_dist_loss(x, y):
x = F.normalize(x, dim=-1)
y = F.normalize(y, dim=-1)
return (x - y).norm(dim=-1).div(2).arcsin().pow(2).mul(2)
def tv_loss(input):
"""L2 total variation loss, as in Mahendran et al."""
input = F.pad(input, (0, 1, 0, 1), 'replicate')
x_diff = input[..., :-1, 1:] - input[..., :-1, :-1]
y_diff = input[..., 1:, :-1] - input[..., :-1, :-1]
return (x_diff**2 + y_diff**2).mean([1, 2, 3])
# + id="Fpbody2NCR7w"
# Model settings
model_config = model_and_diffusion_defaults()
model_config.update({
'attention_resolutions': '32, 16, 8',
'class_cond': False,
'diffusion_steps': 1000,
'rescale_timesteps': True,
'timestep_respacing': '1000',
'image_size': 256,
'learn_sigma': True,
'noise_schedule': 'linear',
'num_channels': 256,
'num_head_channels': 64,
'num_res_blocks': 2,
'resblock_updown': True,
'use_fp16': True,
'use_scale_shift_norm': True,
})
# + id="VnQjGugaDZPJ"
# Load models
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
print('Using device:', device)
model, diffusion = create_model_and_diffusion(**model_config)
model.load_state_dict(torch.load('256x256_diffusion_uncond.pt', map_location='cpu'))
model.requires_grad_(False).eval().to(device)
for name, param in model.named_parameters():
if 'qkv' in name or 'norm' in name or 'proj' in name:
param.requires_grad_()
if model_config['use_fp16']:
model.convert_to_fp16()
clip_model = clip.load('ViT-B/16', jit=False)[0].eval().requires_grad_(False).to(device)
clip_size = clip_model.visual.input_resolution
normalize = transforms.Normalize(mean=[0.48145466, 0.4578275, 0.40821073],
std=[0.26862954, 0.26130258, 0.27577711])
# + [markdown] id="9zY-8I90LkC6"
# ## Settings for this run:
# + id="U0PwzFZbLfcy"
prompt = 'alien friend by <NAME>'
batch_size = 1
clip_guidance_scale = 1000
tv_scale = 150
cutn = 16
n_batches = 1
init_image = None
skip_timesteps = 0
seed = 0
# + [markdown] id="Nf9hTc8YLoLx"
# ### Actually do the run...
# + id="X5gODNAMEUCR"
if seed is not None:
torch.manual_seed(seed)
text_embed = clip_model.encode_text(clip.tokenize(prompt).to(device)).float()
init = None
if init_image is not None:
init = Image.open(fetch(init_image)).convert('RGB')
init = init.resize((model_config['image_size'], model_config['image_size']), Image.LANCZOS)
init = TF.to_tensor(init).to(device).unsqueeze(0).mul(2).sub(1)
make_cutouts = MakeCutouts(clip_size, cutn)
cur_t = None
def cond_fn(x, t, y=None):
with torch.enable_grad():
x = x.detach().requires_grad_()
n = x.shape[0]
my_t = torch.ones([n], device=device, dtype=torch.long) * cur_t
out = diffusion.p_mean_variance(model, x, my_t, clip_denoised=False, model_kwargs={'y': y})
fac = diffusion.sqrt_one_minus_alphas_cumprod[cur_t]
x_in = out['pred_xstart'] * fac + x * (1 - fac)
clip_in = normalize(make_cutouts(x_in.add(1).div(2)))
image_embeds = clip_model.encode_image(clip_in).float().view([cutn, n, -1])
dists = spherical_dist_loss(image_embeds, text_embed.unsqueeze(0))
losses = dists.mean(0)
tv_losses = tv_loss(x_in)
loss = losses.sum() * clip_guidance_scale + tv_losses.sum() * tv_scale
return -torch.autograd.grad(loss, x)[0]
if model_config['timestep_respacing'].startswith('ddim'):
sample_fn = diffusion.ddim_sample_loop_progressive
else:
sample_fn = diffusion.p_sample_loop_progressive
for i in range(n_batches):
cur_t = diffusion.num_timesteps - skip_timesteps - 1
samples = sample_fn(
model,
(batch_size, 3, model_config['image_size'], model_config['image_size']),
clip_denoised=False,
model_kwargs={},
cond_fn=cond_fn,
progress=True,
skip_timesteps=skip_timesteps,
init_image=init,
randomize_class=True,
)
for j, sample in enumerate(samples):
cur_t -= 1
if j % 100 == 0 or cur_t == -1:
print()
for k, image in enumerate(sample['pred_xstart']):
filename = f'progress_{i * batch_size + k:05}.png'
TF.to_pil_image(image.add(1).div(2).clamp(0, 1)).save(filename)
tqdm.write(f'Batch {i}, step {j}, output {k}:')
display.display(display.Image(filename))
| CLIP_Guided_Diffusion_HQ.ipynb |
# ---
# jupyter:
# jupytext:
# notebook_metadata_filter: all
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# language_info:
# codemirror_mode:
# name: ipython
# version: 3
# file_extension: .py
# mimetype: text/x-python
# name: python
# nbconvert_exporter: python
# pygments_lexer: ipython3
# version: 3.6.5
# plotly:
# description: How to upload data to Plotly from Python with the Plotly Grid API.
# display_as: chart_studio
# has_thumbnail: true
# language: python
# layout: base
# name: Plots from Grids
# order: 5
# page_type: u-guide
# permalink: python/data-api/
# thumbnail: thumbnail/table.jpg
# title: Plotly Data API
# v4upgrade: true
# ---
# #### Creating a Plotly Grid
# You can instantiate a grid with data by either uploading tabular data to Plotly or by creating a Plotly `grid` using the API. To upload the grid we will use `plotly.plotly.grid_ops.upload()`. It takes the following arguments:
# - `grid` (Grid Object): the actual grid object that you are uploading.
# - `filename` (str): name of the grid in your plotly account,
# - `world_readable` (bool): if `True`, the grid is `public` and can be viewed by anyone in your files. If `False`, it is private and can only be viewed by you.
# - `auto_open` (bool): if determines if the grid is opened in the browser or not.
#
# You can run `help(py.grid_ops.upload)` for a more detailed description of these and all the arguments.
# +
import chart_studio
import chart_studio.plotly as py
import chart_studio.tools as tls
import plotly.graph_objects as go
from chart_studio.grid_objs import Column, Grid
from datetime import datetime as dt
import numpy as np
from IPython.display import IFrame
column_1 = Column(['a', 'b', 'c'], 'column 1')
column_2 = Column([1, 2, 3], 'column 2') # Tabular data can be numbers, strings, or dates
grid = Grid([column_1, column_2])
url = py.grid_ops.upload(grid,
filename='grid_ex_'+str(dt.now()),
world_readable=True,
auto_open=False)
print(url)
# -
# #### View and Share your Grid
# You can view your newly created grid at the `url`:
IFrame(src= url.rstrip('/') + ".embed", width="100%",height="200px", frameBorder="0")
# You are also able to view the grid in your list of files inside your [organize folder](https://plotly.com/organize).
# #### Upload Dataframes to Plotly
# Along with uploading a grid, you can upload a Dataframe as well as convert it to raw data as a grid:
# +
import chart_studio.plotly as py
import plotly.figure_factory as ff
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/2014_apple_stock.csv')
df_head = df.head()
table = ff.create_table(df_head)
py.iplot(table, filename='dataframe_ex_preview')
# -
# #### Making Graphs from Grids
# Plotly graphs are usually described with data embedded in them. For example, here we place `x` and `y` data directly into our `Histogram2dContour` object:
# +
x = np.random.randn(1000)
y = np.random.randn(1000) + 1
data = [
go.Histogram2dContour(
x=x,
y=y
)
]
py.iplot(data, filename='Example 2D Histogram Contour')
# -
# We can also create graphs based off of references to columns of grids. Here, we'll upload several `column`s to our Plotly account:
# +
column_1 = Column(np.random.randn(1000), 'column 1')
column_2 = Column(np.random.randn(1000)+1, 'column 2')
column_3 = Column(np.random.randn(1000)+2, 'column 3')
column_4 = Column(np.random.randn(1000)+3, 'column 4')
grid = Grid([column_1, column_2, column_3, column_4])
url = py.grid_ops.upload(grid, filename='randn_int_offset_'+str(dt.now()))
print(url)
# -
IFrame(src= url.rstrip('/') + ".embed", width="100%",height="200px", frameBorder="0")
# #### Make Graph from Raw Data
# Instead of placing data into `x` and `y`, we'll place our Grid columns into `xsrc` and `ysrc`:
# +
data = [
go.Histogram2dContour(
xsrc=grid[0],
ysrc=grid[1]
)
]
py.iplot(data, filename='2D Contour from Grid Data')
# -
# So, when you view the data, you'll see your original grid, not just the columns that compose this graph:
# #### Attaching Meta Data to Grids
# In [Chart Studio Enterprise](https://plotly.com/product/enterprise/), you can upload and assign free-form JSON `metadata` to any grid object. This means that you can keep all of your raw data in one place, under one grid.
#
# If you update the original data source, in the workspace or with our API, all of the graphs that are sourced from it will be updated as well. You can make multiple graphs from a single Grid and you can make a graph from multiple grids. You can also add rows and columns to existing grids programatically.
# +
meta = {
"Month": "November",
"Experiment ID": "d3kbd",
"Operator": "<NAME>",
"Initial Conditions": {
"Voltage": 5.5
}
}
grid_url = py.grid_ops.upload(grid, filename='grid_with_metadata_'+str(dt.now()), meta=meta)
print(url)
# -
# #### Reference
help(py.grid_ops)
| _posts/python/chart-studio/data-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext lab_black
import ccxt
import time
from pprint import pprint
exchange = ccxt.huobipro(
{
"apiKey": "3f9161a1-8ce4eb53-hrf5gdfghe-29179",
"secret": "b15e6f29-7215fce0-5f0ebd13-09c14",
}
)
# while True:
# data = exchange.fetchTicker(symbol = 'BTC/USDT')
# buy_price = data['ask']
# print('newest ask:', buy_price)
#
# if buy_price < 10000:
# order_info = exchange.create_limit_buy_order('BTC/USDT', 1)
# print('buy BTC at:', buy_price)
# break
# else:
# print('continue check\n')
# time.sleep(2)
## 获取 BTC 最新价格
BTC_Last = exchange.fetch_ticker("BTC/USDT")["last"]
print("BTC 最新价格:", BTC_Last)
## 定义下单参数
order_symbol = "BTC/USDT"
order_type = "limit"
order_side = "buy"
order_amount = 0.0001
order_price = BTC_Last - 5000
## 下单
take_order = exchange.create_order(order_symbol, order_type, order_side, order_amount, order_price)
## 获取线上订单 ID and order_side
takeorder_id = take_order['id']
print(takeorder_id)
takeorder_side = take_order['side']
print(takeorder_side)
takeorder_price = take_order['price']
print(takeorder_price)
## 查询订单状态
order_status = exchange.fetch_order_status(takeorder_id, order_symbol)
print(order_status)
## 单个网格的交易程序
## 需求: 下一个订单,循环监控订单状态,如果买单成交,则下一个卖单,如果卖单成交,就下一个买单。
X = True
while X:
print('*'*10)
# 获取 BTC 最新价格
BTC_Last = exchange.fetch_ticker(order_symbol)['last']
print('BTC 最新价格:', BTC_Last)
# 查询订单状态
order_status = exchange.fetch_order_status(takeorder_id, order_symbol)
print(order_status)
# 如果买单成交,则下一个卖单
if order_status == 'canceled' and takeorder_side == 'buy':
sell_side = 'sell'
sell_price = takeorder_price + 7000
take_sell_order = exchange.create_order(order_symbol, order_type, sell_side, order_amount, sell_price)
takeorder_id = take_sell_order['id']
takeorder_side = take_sell_order['side']
#time.sleep(1)
# 如果卖单成交,则下一个买单
elif order_status == 'canceled' and takeorder_side == 'sell':
buy_side = 'buy'
buy_price = BTC_Last - 2000
take_buy_order = exchange.create_order(order_symbol, order_type, buy_side, order_amount, buy_price)
takeorder_id = take_buy_order['id']
takeorder_price = take_buy_order['price']
#time.sleep(1)
# 其他
else:
print('Order in other status.')
#time.sleep(1)
| AlgoTrading/ParameterAnalysis/Grid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division
import pandas as pd
import numpy as np
from sklearn import cluster, datasets, mixture
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from torch.distributions.multivariate_normal import MultivariateNormal
from torch.distributions import LogisticNormal
from math import pi
import os
import sys
from pathlib import Path
import gc
from tqdm import tqdm
pd.set_option('display.max_columns', None)
rootdir = r'C:\\Users\\bened\\Documents\\UNIVERSITY\\SchoenStats\\PyTorch Working Directory\\Undergrad Stats Project\\Crime Models\\Crime Data Devon and Cornwall'
# + tags=[]
# collect dataframes from files
dat_2018_list = []
dat_2019_list = []
dat_2020_list = []
dat_2021_list = []
for root, dirs, files in os.walk(rootdir):
for file in tqdm(files):
if file.endswith('.csv'):
df = pd.read_csv(os.path.join(root, file))
if file.startswith('2018'):
dat_2018_list.append(df)
elif file.startswith('2019'):
dat_2019_list.append(df)
elif file.startswith('2020'):
dat_2020_list.append(df)
else:
dat_2021_list.append(df)
# -
def clean_dat_fn(dat_list):
df = pd.concat(dat_list)
df['Month'] = [int(i.split('-')[1]) for i in df['Month']]
df['Crime type'] = df['Crime type'].astype('category')
df_out = df[['Month', 'Longitude', 'Latitude', 'Crime type']]
return(df_out)
mllc18 = clean_dat_fn(dat_2018_list)
mllc19 = clean_dat_fn(dat_2019_list)
mllc20 = clean_dat_fn(dat_2020_list)
mllc21 = clean_dat_fn(dat_2021_list)
# +
data = mllc18[['Longitude', 'Latitude']].dropna()
# random sample from dataset
X = data.sample(2000).reset_index(drop=True)
X_train = torch.tensor(X.values)
X = data.sample(2000).reset_index(drop=True)
X_test = torch.tensor(X.values)
# +
# plot generated dataset
k = mllc18
k = k[(k.Longitude.notna()) & (k.Longitude < -2.8) & (k.Longitude > -5.8)]
k = k[(k.Latitude.notna()) & (k.Latitude < 51.5) & (k.Latitude > 49.5)]
long, lat = k['Longitude'], k['Latitude']
plt.figure(figsize=(12, 8))
plt.scatter(long, lat, s=10, alpha=0.1)
# -
#n_samples = 2000
#noisy_moons = datasets.make_moons(n_samples=n_samples, noise=.05)
X = np.array(X_train)
y = np.array(X_test)
# normalize
X = StandardScaler().fit_transform(X)
y = StandardScaler().fit_transform(y)
# +
# replica functions for StandardScaler().fit_transform(.)
x_mu = np.mean(np.array(X_train[:,0]))
x_sd = np.std(np.array(X_train[:,0]))
y_mu = np.mean(np.array(X_train[:,1]))
y_sd = np.std(np.array(X_train[:,1]))
# test if scaling works, should produce identical plots superimposed
scale_x = (X_train[:, 0] - x_mu) / x_sd
scale_y = (X_train[:, 1] - y_mu) / y_sd
plt.scatter(scale_x, scale_y, alpha=0.2)
plt.scatter(X[:, 0], X[:, 1], alpha=0.1)
# -
# # R-NVP
base_mu, base_cov = torch.zeros(2), torch.eye(2) * 2
base_dist = MultivariateNormal(base_mu, base_cov)
Z = base_dist.rsample(sample_shape=(2000,))
plt.scatter(Z[:, 0], Z[:, 1], alpha=0.3, s=10)
plt.show()
class R_NVP(nn.Module):
def __init__(self, d, k, hidden):
super().__init__()
self.d, self.k = d, k
self.sig_net = nn.Sequential(
nn.Linear(k, hidden),
nn.LeakyReLU(),
nn.Linear(hidden, d - k))
self.mu_net = nn.Sequential(
nn.Linear(k, hidden),
nn.LeakyReLU(),
nn.Linear(hidden, d - k))
def forward(self, x, flip=False):
x1, x2 = x[:, :self.k], x[:, self.k:]
if flip:
x2, x1 = x1, x2
# forward
sig = self.sig_net(x1)
z1, z2 = x1, x2 * torch.exp(sig) + self.mu_net(x1)
if flip:
z2, z1 = z1, z2
z_hat = torch.cat([z1, z2], dim=-1)
log_pz = base_dist.log_prob(z_hat)
log_jacob = sig.sum(-1)
return z_hat, log_pz, log_jacob
def inverse(self, Z, flip=False):
z1, z2 = Z[:, :self.k], Z[:, self.k:]
if flip:
z2, z1 = z1, z2
x1 = z1
x2 = (z2 - self.mu_net(z1)) * torch.exp(-self.sig_net(z1))
if flip:
x2, x1 = x1, x2
return torch.cat([x1, x2], -1)
class stacked_NVP(nn.Module):
def __init__(self, d, k, hidden, n):
super().__init__()
self.bijectors = nn.ModuleList([
R_NVP(d, k, hidden=hidden) for _ in range(n)
])
self.flips = [True if i%2 else False for i in range(n)]
def forward(self, x):
log_jacobs = []
for bijector, f in zip(self.bijectors, self.flips):
x, log_pz, lj = bijector(x, flip=f)
log_jacobs.append(lj)
return x, log_pz, sum(log_jacobs)
def inverse(self, z):
for bijector, f in zip(reversed(self.bijectors), reversed(self.flips)):
z = bijector.inverse(z, flip=f)
return z
# ## Training Step Function
def train(model, epochs, batch_size, optim, scheduler):
losses = []
for _ in tqdm(range(epochs)):
# get batch
X= X_train
X = torch.from_numpy(StandardScaler().fit_transform(X)).float()
optim.zero_grad()
z, log_pz, log_jacob = model(X)
loss = (-log_pz - log_jacob).mean()
losses.append(loss)
loss.backward()
optim.step()
scheduler.step()
return losses
def view(model, losses, y):
plt.plot(losses)
plt.title("Model Loss vs Epoch")
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.show()
X_hat = model.inverse(Z).detach().numpy()
plt.scatter(X_hat[:, 0], X_hat[:, 1], alpha=0.1, s=10)
#plt.scatter(y[:, 0], y[:, 1], alpha=0.1, s=10)
plt.title("Inverse of Normal Samples Z: X = F^-1(Z)")
plt.ylim((-2.5, 2.5))
plt.xlim((-2.5, 2.5))
plt.show()
n_samples = 2000
#X = X_train
X = X_test
X = torch.from_numpy(StandardScaler().fit_transform(X)).float()
z, _, _ = model(X)
z = z.detach().numpy()
plt.scatter(z[:, 0], z[:, 1], alpha=0.2, s=20)
plt.title("Transformation of Data Samples X: Z = F(X)")
plt.show()
return(z, X_hat)
# ## Model Params
d = 2
k = 1
# # Single Layer R_NVP
model = R_NVP(d, k, hidden=512)
optim = torch.optim.Adam(model.parameters(), lr=1e-3)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 1000, n_samples, optim, scheduler)
z_res, x_hat_res = view(model, losses, y)
# # 3 Layer R_NVP
# +
model = stacked_NVP(d, k, hidden=512, n=3)
optim = torch.optim.Adam(model.parameters(), lr=1e-3)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 1000, n_samples, optim, scheduler)
# -
z_res, x_hat_res = view(model, losses, y)
# # 5 Layer R_NVP
# +
model = stacked_NVP(d, k, hidden=512, n=5)
optim = torch.optim.Adam(model.parameters(), lr=1e-3)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 1000, n_samples, optim, scheduler)
# -
z_res, x_hat_res = view(model, losses, y)
# # 10 Layer R_NVP
# + tags=[]
model = stacked_NVP(d, k, hidden=512, n=10)
optim = torch.optim.Adam(model.parameters(), lr=1e-4)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 2000, n_samples, optim, scheduler)
# -
z_res, x_hat_res = view(model, losses, y)
# # 15 Layer R_NVP
# + tags=[]
model = stacked_NVP(d, k, hidden=512, n=15)
optim = torch.optim.Adam(model.parameters(), lr=1e-4)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 1000, n_samples, optim, scheduler)
# -
z_res, x_hat_res = view(model, losses, y)
# ## 20 layer R_NVP
# + tags=[]
model = stacked_NVP(d, k, hidden=512, n=20)
optim = torch.optim.Adam(model.parameters(), lr=1e-5)
scheduler = torch.optim.lr_scheduler.ExponentialLR(optim, 0.999)
n_samples = 512
# training loop
losses = train(model, 2000, n_samples, optim, scheduler)
# -
z_res, x_hat_res = view(model, losses, y)
# # Model Tests; generate Test Data
# + tags=[]
def gen_sine_wave_sample(n=2000, s=1, amp=1, wl=1,
ns_mean=0, ns_var=1, lim_y=True, lim_x=True, x_type='rand',
x_range=(0,1), y_range=(0,1), set_input=None,
r_rad=0, swap_axis=False, formula=2):
# n - number of points to generate
# s - slope gradient of wave
# amp - amplitude of wave
# wl - wavelength equivalent parameter
# ns_mean, ns_var - noise mean and variance
# set how the domain is generated
if x_type == 'rand':
x = np.random.rand(n)
elif x_type == 'lin' :
x = np.linspace(0, 1, n, endpoint=True)
elif x_type == 'self' :
x = set_input
#set x limits
if lim_x:
x = np.interp(x, (x.min(), x.max()), x_range)
epsilon = ns_mean + ns_var * np.random.randn(n)
# apply to formula [y = s*x + mag*sin(wl*x)]
if formula==0:
y = s*x + amp*x*np.sin(wl*x) + epsilon
elif formula==1:
y = s*x + amp*x*np.sin(wl*x) + epsilon*x
elif formula==2:
y = s*x + amp*x*np.sin(wl*x)
else:
y = s*x + amp*np.sin(wl*x) + epsilon
# create rotation matrix
rM = np.array((np.cos(r_rad), -np.sin(r_rad), np.sin(r_rad), np.cos(r_rad))).reshape((-1, 2))
t = np.array((x, y))
# set y limits
if lim_y:
t = t[:, t[1] < y_range[1]]
t = t[:, t[1] > y_range[0]]
# swap x and y axis
if (swap_axis==True):
t = t[[1, 0], :]
# rotation matrix
output = np.matmul(rM, t)
# note that (no. of output) <= n
return(output)
# +
# scaling x_hat_res to return to original
pred_scaled_x = (x_hat_res[:, 0] * x_sd) + x_mu
pred_scaled_y = (x_hat_res[:, 1] * y_sd) + y_mu
# once scaled, we can input the data into the actual toy data generator
t = gen_sine_wave_sample(s=0.9, amp=0.25, wl=13, formula=1,
lim_x=False, lim_y=False, ns_var=0.09,
swap_axis=False,
x_type='self', set_input=pred_scaled_x)
# limit x
# - this may be turned off to see predictions outside of training range
results_scaled = np.array((pred_scaled_x, pred_scaled_y, t[1,:]))
results_scaled = results_scaled[:, results_scaled[0, :] <= 1]
results_scaled = results_scaled[:, results_scaled[0, :] >= 0]
# -
plt.scatter(results_scaled[0,:], results_scaled[2,:], alpha=0.2, s=10, label='target')
plt.scatter(results_scaled[0,:], results_scaled[1,:], alpha=0.2, s=10, label='prediction')
plt.xlim((0, 1))
plt.ylim((0, 1))
plt.legend()
| NF-Models/NF_CrimeDat_Basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Topic extraction with Non-negative Matrix Factorization and Latent Dirichlet Allocation
#
#
# This is an example of applying Non-negative Matrix Factorization
# and Latent Dirichlet Allocation on a corpus of documents and
# extract additive models of the topic structure of the corpus.
# The output is a list of topics, each represented as a list of terms
# (weights are not shown).
#
# The default parameters (n_samples / n_features / n_topics) should make
# the example runnable in a couple of tens of seconds. You can try to
# increase the dimensions of the problem, but be aware that the time
# complexity is polynomial in NMF. In LDA, the time complexity is
# proportional to (n_samples * iterations).
#
# +
# Author: <NAME> <<EMAIL>>
# <NAME>
# <NAME> <<EMAIL>>
# License: BSD 3 clause
from __future__ import print_function
from time import time
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.decomposition import NMF, LatentDirichletAllocation
from sklearn.datasets import fetch_20newsgroups
n_samples = 2000
n_features = 1000
n_topics = 10
n_top_words = 20
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
print("Topic #%d:" % topic_idx)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]]))
print()
# Load the 20 newsgroups dataset and vectorize it. We use a few heuristics
# to filter out useless terms early on: the posts are stripped of headers,
# footers and quoted replies, and common English words, words occurring in
# only one document or in at least 95% of the documents are removed.
print("Loading dataset...")
t0 = time()
dataset = fetch_20newsgroups(shuffle=True, random_state=1,
remove=('headers', 'footers', 'quotes'))
data_samples = dataset.data[:n_samples]
print("done in %0.3fs." % (time() - t0))
# Use tf-idf features for NMF.
print("Extracting tf-idf features for NMF...")
tfidf_vectorizer = TfidfVectorizer(max_df=0.95, min_df=2,
max_features=n_features,
stop_words='english')
t0 = time()
tfidf = tfidf_vectorizer.fit_transform(data_samples)
print("done in %0.3fs." % (time() - t0))
# Use tf (raw term count) features for LDA.
print("Extracting tf features for LDA...")
tf_vectorizer = CountVectorizer(max_df=0.95, min_df=2,
max_features=n_features,
stop_words='english')
t0 = time()
tf = tf_vectorizer.fit_transform(data_samples)
print("done in %0.3fs." % (time() - t0))
# Fit the NMF model
print("Fitting the NMF model with tf-idf features, "
"n_samples=%d and n_features=%d..."
% (n_samples, n_features))
t0 = time()
nmf = NMF(n_components=n_topics, random_state=1,
alpha=.1, l1_ratio=.5).fit(tfidf)
print("done in %0.3fs." % (time() - t0))
print("\nTopics in NMF model:")
tfidf_feature_names = tfidf_vectorizer.get_feature_names()
print_top_words(nmf, tfidf_feature_names, n_top_words)
print("Fitting LDA models with tf features, "
"n_samples=%d and n_features=%d..."
% (n_samples, n_features))
lda = LatentDirichletAllocation(n_topics=n_topics, max_iter=5,
learning_method='online',
learning_offset=50.,
random_state=0)
t0 = time()
lda.fit(tf)
print("done in %0.3fs." % (time() - t0))
print("\nTopics in LDA model:")
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, n_top_words)
| Sources/issuse18OnlineMLFlink/proteus-solma-development/src/main/resources/topics_extraction_with_nmf_lda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !pip install git+git://github.com/jsignell/dask-geopandas.git
# # !pip install git+git://github.com/geopandas/geopandas.git
# +
from dask.distributed import Client, LocalCluster
import time
import dask_geopandas as dask_geopandas
import dask.dataframe as dd
import geopandas
import pygeos
import numpy as np
import pandas as pd
import momepy
from tqdm.notebook import tqdm
from libpysal.weights import Queen
from momepy_utils import _circle_radius, get_corners, squareness, elongation, centroid_corner, solar_orientation_poly
# -
client = Client(LocalCluster(n_workers=14))
client
# +
import warnings
warnings.filterwarnings('ignore', message='.*initial implementation of Parquet.*')
warnings.filterwarnings('ignore', message='.*Assigning CRS to a GeoDataFrame without a geometry*')
# + jupyter={"outputs_hidden": true}
for chunk_id in tqdm(range(103), total=103):
blg = geopandas.read_parquet(f"../../urbangrammar_samba/spatial_signatures/buildings/blg_{chunk_id}.pq")
tess = geopandas.read_parquet(f"../../urbangrammar_samba/spatial_signatures/tessellation/tess_{chunk_id}.pq")
blg = blg.rename_geometry('buildings')
tess = tess.rename_geometry('tessellation')
df = tess.merge(blg, on='uID', how='left')
ddf = dask_geopandas.from_geopandas(df, npartitions=14)
ddf['sdbAre'] = ddf.buildings.area
ddf['sdbPer'] = ddf.buildings.length
exterior_area = ddf.buildings.map_partitions(lambda series: pygeos.area(pygeos.polygons(series.exterior.values.data)), meta='float')
ddf['sdbCoA'] = exterior_area - ddf['sdbAre']
hull = ddf.buildings.convex_hull.exterior
radius = hull.apply(lambda g: _circle_radius(list(g.coords)) if g is not None else None, meta='float')
ddf['ssbCCo'] = ddf['sdbAre'] / (np.pi * radius ** 2)
ddf['ssbCor'] = ddf.buildings.apply(lambda g: get_corners(g), meta='float')
ddf['ssbSqu'] = ddf.buildings.apply(lambda g: squareness(g), meta='float')
bbox = ddf.buildings.apply(lambda g: g.minimum_rotated_rectangle if g is not None else None, meta=geopandas.GeoSeries())
ddf['ssbERI'] = (ddf['sdbAre'] / bbox.area).pow(1./2) * (bbox.length / ddf['sdbPer'])
ddf['ssbElo'] = bbox.map_partitions(lambda s: elongation(s), meta='float')
def _centroid_corner(series):
ccd = series.apply(lambda g: centroid_corner(g))
return pd.DataFrame(ccd.to_list(), index=series.index)
ddf[['ssbCCM', 'ssbCCD']] = ddf.buildings.map_partitions(_centroid_corner, meta=pd.DataFrame({0: [0.1], 1: [1.1]}))
ddf['stbOri'] = bbox.apply(lambda g: solar_orientation_poly(g), meta='float')
hull = ddf.tessellation.convex_hull.exterior
ddf['sdcLAL'] = hull.apply(lambda g: _circle_radius(list(g.coords)), meta='float') * 2
ddf['sdcAre'] = ddf.tessellation.area
radius = hull.apply(lambda g: _circle_radius(list(g.coords)), meta='float')
ddf['sscCCo'] = ddf['sdcAre'] / (np.pi * radius ** 2)
bbox = ddf.tessellation.apply(lambda g: g.minimum_rotated_rectangle, meta=geopandas.GeoSeries())
ddf['sscERI'] = (ddf['sdcAre'] / bbox.area).pow(1./2) * (bbox.length / ddf.tessellation.length)
ddf['stcOri'] = bbox.apply(lambda g: solar_orientation_poly(g), meta='float')
ddf['sicCAR'] = ddf['sdbAre'] / ddf['sdcAre']
ddf['stbCeA'] = (ddf['stbOri'] / ddf['stcOri']).abs()
df = ddf.compute()
df.to_parquet(f"../../urbangrammar_samba/spatial_signatures/morphometrics/cells/cells_{chunk_id}.pq")
client.restart()
time.sleep(5)
# +
# %%time
encl = dask_geopandas.read_parquet("../../urbangrammar_samba/spatial_signatures/enclosures/encl_*.pq")
encl['ldeAre'] = encl.geometry.area
encl['ldePer'] = encl.geometry.length
hull = encl.geometry.convex_hull.exterior
radius = hull.apply(lambda g: _circle_radius(list(g.coords)) if g is not None else None, meta='float')
encl['lseCCo'] = encl['ldeAre'] / (np.pi * radius ** 2)
bbox = encl.geometry.apply(lambda g: g.minimum_rotated_rectangle if g is not None else None, meta=geopandas.GeoSeries())
encl['lseERI'] = (encl['ldeAre'] / bbox.area).pow(1./2) * (bbox.length / encl['ldePer'])
longest_axis = hull.apply(lambda g: _circle_radius(list(g.coords)), meta='float') * 2
encl['lseCWA'] = longest_axis * ((4 / np.pi) - (16 * encl['ldeAre']) / ((encl['ldePer']) ** 2))
encl['lteOri'] = bbox.apply(lambda g: solar_orientation_poly(g), meta='float')
encl_df = encl.compute()
# make 3d geometry 2d
coords = pygeos.get_coordinates(encl_df.geometry.values.data)
counts = pygeos.get_num_coordinates(encl_df.geometry.values.data)
encl_df['geometry'] = geopandas.GeoSeries([pygeos.polygons(c) for c in np.split(coords, np.cumsum(counts)[:-1])], crs=encl_df.crs)
inp, res = encl_df.sindex.query_bulk(encl_df.geometry, predicate='intersects')
indices, counts = np.unique(inp, return_counts=True)
encl_df['neighbors'] = counts - 1
encl_df['lteWNB'] = encl_df['neighbors'] / encl_df['ldePer']
tess = dd.read_parquet("../../urbangrammar_samba/spatial_signatures/tessellation/tess_*.pq")
encl_counts = tess.groupby('enclosureID').count().compute()
merged = encl_df[['enclosureID', 'ldeAre']].merge(encl_counts[['geometry']], how='left', on='enclosureID')
encl_df['lieWCe'] = merged['geometry'] / merged['ldeAre']
encl_df.drop(columns='geometry').to_parquet("../../urbangrammar_samba/spatial_signatures/morphometrics/enclosures.pq")
# -
client.close()
| docs/_sources/measuring/_morphometrics_dask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BRIJNANDA1979/CNN-Sentinel/blob/master/BigEarthNet_Dataset_Populate_unique_labelled_classes_subfolders_with_12_tiff_images.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="7nakWhsSyYsZ"
# -*- coding: utf-8 -*-
"""
Created on Fri Jun 11 14:30:56 2021
@author: BRIJB
"""
import os
import pandas as pd
from os import path
import re
import glob
import shutil
srcdir = 'E:\Test BiGdata'
path='E:\\New BigEarthNet_Labelled 12925 Data.csv'
df=pd.read_csv(path)
dirs = os.listdir(srcdir)
for row in df.itertuples():
des_folder = 'D:' + '//' + 'temp' +'//' + row[4]
s=row[5]
s=s.replace('[','') # cleaning data
s=s.replace(']','')
s=s.replace('\'','')
s=s.strip() #removes blank spaces if any
sub_folders_list = s.split(',')
sub_folders_list = [x.strip() for x in sub_folders_list] #remove blank spaces in list of strings
for sub in sub_folders_list:
for dir in dirs:
if str(sub) == str(dir):
print('Match found')
print(sub)
print(row[4])
ss= srcdir + '\\' + dir + '\\'
sss = ss + '*.tif'
files = glob.glob(sss) # all tif files in sub folder
for f in files: # copy tif files to destination folder
print('copying')
destination_of_file = des_folder + '//' + str(sub)
shutil.copy(os.path.join(ss,f),destination_of_file)
| BigEarthNet_Dataset_Populate_unique_labelled_classes_subfolders_with_12_tiff_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pangeo-notebook
# language: python
# name: pangeo-notebook
# ---
# # Getting started
#
# Let's with NECOFS data, available at [SMAST archive](http://www.smast.umassd.edu:8080/thredds/catalog/models/fvcom/NECOFS/Archive/NECOFS_GOM/catalog.html).
# +
#importing main modules
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
#importing modules used as auxiliary
from netCDF4 import Dataset
from dateutil import parser
from datetime import datetime,timedelta
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# We need to drop `siglay` and `siglev` coordinates. For some reason the data have the same name for coordinates and dimensions, e.g `siglay` is a coordinate that depends on `(siglay,node)`. [Many others](https://github.com/pydata/xarray/issues/2233) had the same problem with this model output. There is also some problem with the time formatting, we will need to decode it after loading the data.
# +
# define year and month to be read
year = 2019
month = 5
# we could use this to run a loop through the years/months we need
# list problematic coordinates
drop_variables = ['siglay','siglev']
# base url for openDAP server
url = "".join(["http://www.smast.umassd.edu:8080/thredds/dodsC/models/fvcom/",
f"NECOFS/Archive/NECOFS_GOM/{year}/gom4_{year}{month:02d}.nc?"])
# lazy load of the data
ds = xr.open_dataset(url,drop_variables=drop_variables,decode_times=False)
# convert lon/c, lat/c to coordinates
ds = ds.assign_coords({var:ds[var] for var in ['lon','lat','lonc','latc']})
# -
# Solving the problem with `siglay` and `siglev`. We need to workaround using `netCDF4` and renaming the coordinates.
# +
# load data with netCDF4
nc = Dataset(url)
# load the problematic coordinates
coords = {name:nc[name] for name in drop_variables}
# function to extract ncattrs from `Dataset()`
get_attrs = lambda name: {attr:coords[name].getncattr(attr) for attr in coords[name].ncattrs()}
# function to convert from `Dataset()` to `xr.DataArray()`
nc2xr = lambda name: xr.DataArray(coords[name],attrs=get_attrs(name),name=f'{name}_coord',dims=(f'{name}','node'))
# apply `nc2xr()` and merge `xr.DataArray()` objects
coords = xr.merge([nc2xr(name) for name in coords.keys()])
# reassign to the main `xr.Dataset()`
ds = ds.assign_coords(coords)
# -
# We now can assign the `z` coordinate for the data.
z = (ds.siglay_coord*ds.h)
z.attrs = dict(long_name='nodal z-coordinate',units='meters')
ds = ds.assign_coords(z=z)
# The time coordinate still needs some fixing. We will parse it and reassign to the dataset.
# the first day
dt0 = parser.parse(ds.time.attrs['units'].replace('days since ',''))
# parse dates summing days to the origin
ds = ds.assign(time=[dt0+timedelta(seconds=day*86400) for day in ds.time.values])
# Now let's try to make some simple plots.
# +
# url for GEBCO data
url = "https://tds.marine.rutgers.edu/thredds/dodsC/other/bathymetry/GEBCO_2019/GEBCO_2019.nc"
# load data from min-buffer to max+buffer
buffer = 2
gebco = xr.open_dataset(url,decode_times=False).sel(
lon=slice(ds.lon.min()-buffer,ds.lon.max()+buffer),
lat=slice(ds.lat.min()-buffer,ds.lat.max()+buffer)
).compute() #compute loads data into memory (faster plots)
# +
# formatting function for the units
fmt = lambda txt: txt.replace('degrees_','$^\circ$').replace('1e-3','10$^{-3}$')
# key argument function that defines vlim from data quantiles
kw = lambda da,q: {"s":25,"vmin":da.quantile(q[0]).values,"vmax":da.quantile(q[1]).values}
# select the surface layer and the first time output
isel = dict(siglay=0,time=0)
fig,ax = plt.subplots(1,2,figsize=(8,3))
for a in np.ravel(ax):
a.axis('scaled')
gebco['elevation'].plot.contour(ax=a,
levels=[-3000,-200],add_colorbar=False,add_labels=False,colors='0.3',linestyles='solid',linewidths=1,zorder=1e3)
gebco['elevation'].plot.contourf(ax=a,
levels=[0,1e5],add_colorbar=False,add_labels=False,zorder=1e6+1,extend='neither',colors='0.2')
da = ds.temp.isel(**isel)
C = ax[0].scatter(da.lon,da.lat,c=da,**kw(da,[0.01,0.99]),cmap='plasma')
fig.colorbar(C,ax=ax[0],label=f"{da.long_name.capitalize()} [{fmt(da.units)}]",fraction=0.05,orientation="horizontal")
da = ds.salinity.isel(**isel)
C = ax[1].scatter(da.lon,da.lat,c=da,**kw(da,[0.2,0.99]))
fig.colorbar(C,ax=ax[1],label=f"{da.long_name.capitalize()} [{fmt(da.units)}]",fraction=0.05,orientation="horizontal")
_ = [a.grid(True, ls='--',alpha=0.5) for a in ax]
lablat = lambda lat: f'{lat:.0f}$^\\circ$' if lat==0 else f'{lat:.0f}$^\\circ$N' if lat>0 else f'{-lat:.0f}$^\\circ$S'
lablon = lambda lat: f'{lat:.0f}$^\\circ$' if lat==0 else f'{lat:.0f}$^\\circ$E' if lat>0 else f'{-lat:.0f}$^\\circ$W'
xlim = np.array([gebco.lon.min().values,gebco.lon.max().values])
ylim = np.array([gebco.lat.min().values,gebco.lat.max().values])
xticks = np.arange(*(np.round(xlim/10)*10).astype('int'),5)
yticks = np.arange(*(np.round(ylim/10)*10).astype('int'),5)
for a in np.ravel(ax):
a.set(
xticks=xticks,
yticks=yticks,
xticklabels=list(map(lablon,xticks)),
yticklabels=list(map(lablat,yticks)),
xlabel='',ylabel='',
xlim=xlim,
ylim=ylim,
)
_ = [a.grid(True,linestyle='--',alpha=0.5,zorder=1e7) for a in np.ravel(ax)]
fig.savefig('../img/scatter_temperature_salinity.png',dpi=150,bbox_inches='tight')
# -
| examples/00-GettingStarted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from matplotlib import pyplot as plt
plt.rcParams['figure.figsize'] = [15, 5]
# +
_min=1
_max=6
m=6*10**4
a=np.random.randint(low=1,high=7,size=m)
hist,edge=np.histogram(a,bins=_max-_min+1,range=(0.5,6.5))
plt.bar(edge[:-1]+0.5,hist);
# +
n=2
_min=1
_max=6
m=6*10**4
a=np.zeros(shape=m)
for i in range(n):
a+=np.random.randint(low=1,high=7,size=m)
hist,edge=np.histogram(a,bins=n*(_max-_min)+1,range=(_min*n-0.5,_max*n+0.5))
plt.bar(edge[:-1]+0.5,hist);
# +
n=10
_min=1
_max=6
m=6*10**4
a=np.zeros(shape=m)
for i in range(n):
a+=np.random.randint(low=1,high=7,size=m)
hist,edge=np.histogram(a,bins=n*(_max-_min)+1,range=(_min*n-0.5,_max*n+0.5))
plt.bar(edge[:-1]+0.5,hist);
#cumulative probability of a between -2std and 2std
len(a [ np.logical_and( mean-2*std < a, a <= mean+2*std) ])/ len(a)
# -
len(a)
mean=np.mean(a) #mu
mean
std=np.std(a) #sigma
std
a [ mean-std < a ]
a [ np.logical_and( mean-std < a, a <= mean+std) ]
#cumulative probability of a between -std and std
len(a [ np.logical_and( mean-std < a, a <= mean+std) ])/ len(a)
#cumulative probability of a between -2std and 2std
len(a [ np.logical_and( mean-2*std < a, a <= mean+2*std) ])/ len(a)
#cumulative probability of a between -3std and 3std
len(a [ np.logical_and( mean-3*std < a, a <= mean+3*std) ])/ len(a)
| dsi200_demo/_extra probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: adapter
# language: python
# name: adapter
# ---
# ## This ipynb help you to transfer the CONLL NER dataset to the backend's DB. So the trainer can train NER model base on the data.
from core.config import API_PORT
import pandas as pd
df = pd.read_csv("./NER_multilabel_data_v2.csv")
df[90:100]
df.loc[:, "Sentence #"] = df["Sentence #"].fillna(method="ffill")
import numpy as np
sentences = list(set(df["Sentence #"]))
import requests
words = df[df["Sentence #"] == sentence]
def process_data(row):
return {
"text": row.Word,
"labels": row.newTag.split("|")
}
def post_data(sentence):
data = {
"user": "CONLL_0818@admin",
"tags": ["setup"]
"texts": list(df[df["Sentence #"] == sentence].apply(process_data, axis = 1))
}
headers = {
'accept': 'application/json',
'Content-Type': 'application/json'
}
return requests.post(f"http://localhost:{API_PORT}/api/v1/data/labeledText", data=json.dumps(data), headers = headers)
# put datas into DB
import json
from tqdm import tqdm
import threading
tasks = []
for sentence in tqdm(sentences):
t = threading.Thread(target=post_data, args={sentence})
t.start()
tasks.append(t)
for task in tqdm(tasks):
task.join()
| Setup_Operation/post_train_data_to_db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from hcipy import *
import numpy as np
from matplotlib import pyplot as plt
N = 64
D = 9.96
aperture = circular_aperture(D)
pupil_grid = make_pupil_grid(N, D)
sps = 40 * N // 128
pupsep = 1
outgrid_size = int(np.ceil(sps * (pupsep + 1)))
D_grid = 3.6e-3
pyramid_grid = make_pupil_grid(N, D_grid)
outgrid = np.zeros((N, N)).tolist()
buffer = (N - 2 * outgrid_size)//2
# +
def get_sub_images(electric_field):
pyramid_grid = make_pupil_grid(N, D_grid)
images = Field(np.asarray(electric_field).ravel(), pyramid_grid)
pysize = int(np.sqrt(images.size))
images.shape = (pysize, pysize)
sub_images = [images[pysize-sps-1:pysize-1, 0:sps], images[pysize-sps-1:pysize-1, pysize-sps-1:pysize-1],
images[0:sps, 0:sps], images[0:sps, pysize-sps-1:pysize-1]]
subimage_grid = make_pupil_grid(sps, D_grid * sps / N)
for count, img in enumerate(sub_images):
img = img.ravel()
img.grid = subimage_grid
sub_images[count] = img
return sub_images
def pyramid_prop(wf):
# Given a wavefront, returns the result of a pyramid propagation and splitting into sub-images,
# as a list of hcipy Field objects.
keck_pyramid = PyramidWavefrontSensorOptics(pupil_grid, pupil_separation=pupsep, num_pupil_pixels=sps)
return get_sub_images(keck_pyramid.forward(wf).electric_field)
def estimate(images_list):
EstimatorObject = PyramidWavefrontSensorEstimator(aperture, make_pupil_grid(sps*2, D_grid*sps*2/N))
I_b = images_list[0]
I_a = images_list[1]
I_c = images_list[2]
I_d = images_list[3]
norm = I_a + I_b + I_c + I_d
I_x = (I_a + I_b - I_c - I_d) / norm
I_y = (I_a - I_b - I_c + I_d) / norm
pygrid = make_pupil_grid(sps)
return Field(I_x.ravel(), pygrid), Field(I_y.ravel(), pygrid)
def make_slopes(wf):
x, y = estimate(pyramid_prop(wf))
return np.concatenate((x, y))
# +
def aberration_basis(N):
wfref = Wavefront(aperture(pupil_grid))
wfref.electric_field.shape = (N, N)
wfrefl = wfref.electric_field.tolist()
aberration_mode_basis = []
for i in range(N):
for j in range(N):
l = np.zeros((N, N), dtype=complex)
if np.complex(wfrefl[i][j]) != 0:
l[i][j] = (1+0.1j)
aberration_mode_basis.append(Wavefront(Field(np.round(np.asarray(l).ravel(), 3) * aperture(pupil_grid), wfref.grid)))
return aberration_mode_basis
basis = aberration_basis(N)
# -
pt = sps
test = Wavefront(basis[0].electric_field + basis[pt].electric_field)
imshow_field(test.electric_field)
plt.show()
test_slopes = make_slopes(test)
spsg = make_pupil_grid(sps)
test_x = Field(test_slopes[0:sps*sps], spsg)
test_y = Field(test_slopes[sps*sps:2*sps*sps], spsg)
slopes_basis = [make_slopes(basis_element) for basis_element in basis]
# +
#Comparing the x slopes
imshow_field(test_x)
plt.show()
imshow_field(slopes_basis[0][0:sps*sps] + slopes_basis[pt][0:sps*sps], spsg)
plt.show()
# -
# Comparing the y slopes
imshow_field(test_y, vmax=0.05, vmin=-0.05)
plt.show()
imshow_field(slopes_basis[0][sps*sps:2*sps*sps] + slopes_basis[pt][sps*sps:2*sps*sps], spsg)
plt.show()
| pwfs-keck/t/LinearityTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pyFIRS]
# language: python
# name: conda-env-pyFIRS-py
# ---
import os
import shutil
import glob
import subprocess
import pandas as pd
import geopandas as gpd
import dask
from dask.distributed import Client, progress, LocalCluster
from pyFIRS.wrappers import lastools
# Launch a parallel computing cluster.
cluster=LocalCluster(scheduler_port=7001, diagnostics_port=7002)
c = Client(cluster)
num_cores = len(c.ncores()) # identify how many workers we have
# At this point, you should also be able to view an interactive dashboard on port 7002. If you're executing this on a remote server, you'll need to set up port forward so you can view the dashboard on your local machine's browser. Once you've done that, or if you're processing on your own machine, you can view the dashboard at [http://localhost:7002/status](http://localhost:7002/status).
las = lastools.useLAStools('/storage/lidar/LAStools/bin')
# +
# where the imported lidar data is currently stored
workdir = os.path.abspath('/storage/lidar/swinomish_2012/')
# define data handling directories
processed = os.path.join(workdir,'processed')
# the coordinate reference system we'll be working with
target_epsg = 26910 # utm 10 N
# -
# push our working directories and wrapper classes to the workers on the cluster as well
c.scatter([processed, las, target_epsg, num_cores], broadcast=True);
def fname(path):
"""returns the filename as basename split from extension.
Parameters
-----------
path : str, path to file
filepath from which filename will be sliced
Returns
--------
filename : str
name of file, split from extension
"""
filename = os.path.basename(path).split('.')[0]
return filename
# +
tiles_to_merge = [fname(tile) for tile in
glob.glob(os.path.join(processed, 'points', '*.laz'))
]
print('Found {:,d} tiles to merge derivative products from.'.format(len(tiles_to_merge)))
# -
# # Merge tiled derivative outputs together
# Merge all the tiled GeoTiffs and Shapefiles into single overview files.
# We'll produce a shapefile showing the layout of the non-buffered tiles as a single shapefile. This is a single process that takes a few seconds to run, so no need to distribute it using `dask`.
@dask.delayed
def tile_boundaries(*args, **kwargs):
odir = os.path.join(processed, 'vectors')
if os.path.exists(os.path.join(processed, 'vectors', 'tiles.shp')):
pass
else:
proc = las.lasboundary(i=os.path.join(processed, 'points', '*.laz'),
use_bb=True, # use bounding box of tiles
overview=True,
labels=True,
cores=num_cores, # use parallel processing
oshp=True,
o=os.path.join(processed, 'vectors', 'tiles.shp'))
return
@dask.delayed
def make_footprint(*args, **kwargs):
if os.path.exists(os.path.join(processed, 'vectors', 'footprint.shp')):
pass
else:
gdf = gpd.read_file(os.path.join(processed, 'vectors', 'tiles.shp'))
gdf['mil_points'] = gdf['num_points'] / 1000000.
buffered = gdf.drop(['file_name', 'point_size', 'point_type', 'num_points'], axis=1)
buffered.geometry = gdf.buffer(0.01) # buffer by 1cm
try:
union = gpd.GeoDataFrame(geometry=list(buffered.unary_union), crs=buffered.crs)
except TypeError: # line above will fail if there is only one polygon for the footprint
union = gpd.GeoDataFrame(geometry=[buffered.unary_union], crs=buffered.crs)
union['footprint_id'] = union.index + 1
buffered = gpd.tools.sjoin(buffered, union, how='left').drop('index_right', axis=1)
aggfuncs={'mil_points':'sum',
'version':'first',
'min_x':'min',
'min_y':'min',
'min_z':'min',
'max_x':'max',
'max_y':'max',
'max_z':'max'}
dissolved = buffered.dissolve(by='footprint_id', aggfunc=aggfuncs)
dissolved.to_file(os.path.join(processed, 'vectors', 'footprint.shp'))
return 'footprint'
def merge_chunks(infiles, outfile):
"""Merges a list of rasters, one chunk at a time.
Parameters
-----------
infiles : list
list containing paths to input files
outfile : string, path to file
the output file that will be created by merging all the input files
"""
if os.path.exists(outfile):
pass
else:
outname = os.path.basename(outfile).split('.')[0]
os.makedirs(os.path.join(processed, 'rasters','chunks'), exist_ok=True)
# break the list of input files into chunks of 500
for i in range(0,len(infiles),500):
chunk_infiles = infiles[i:i+500]
chunk_outfile = os.path.join(processed, 'rasters', 'chunks', 'chunk{}_{}.tif'.format(i, outname))
proc = subprocess.run(['rio', 'merge', *chunk_infiles, chunk_outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', '--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
chunked_outfiles = glob.glob(os.path.join(processed, 'rasters', 'chunks', 'chunk*{}*.tif'.format(outname)))
proc = subprocess.run(['rio', 'merge', *chunked_outfiles, outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', '--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
return
# Merge the bare earth tiles into a single GeoTiff.
@dask.delayed
def merge_dem(*args, **kwargs):
infiles = glob.glob(os.path.join(processed, 'rasters', 'DEM_tiles', '*.tif'))
outfile = os.path.join(processed, 'rasters', 'dem.tif')
if os.path.exists(outfile):
pass
elif len(infiles) < 500:
return subprocess.run(['rio', 'merge', *infiles, outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', '--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
else:
merge_chunks(infiles, outfile)
return
# Now merge the hillshade tiles into a single raster formatted as GeoTiff.
@dask.delayed
def merge_hillshade(*args, **kwargs):
infiles = glob.glob(os.path.join(processed, 'rasters', 'hillshade_tiles', '*.tif'))
outfile = os.path.join(processed, 'rasters', 'hillshade.tif')
if os.path.exists(outfile):
pass
elif len(infiles) < 500:
return subprocess.run(['rio', 'merge', *infiles, outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', '--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
else:
merge_chunks(infiles, outfile)
return
# Merge the trimmed canopy height model tiles into a single raster.
@dask.delayed
def merge_chm(*args, **kwargs):
infiles = glob.glob(os.path.join(processed, 'rasters', 'chm_tiles', '*.tif'))
outfile = os.path.join(processed, 'rasters', 'chm.tif')
if os.path.exists(outfile):
pass
elif len(infiles) < 500:
proc = subprocess.run(['rio', 'merge', *infiles, outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', '--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
else:
merge_chunks(infiles, outfile)
return
# Merge the cleaned tiles of building footprints together into a single shapefile. We'll use `geopandas` to concatenate all the polygons together into a single geodataframe and then write out to a new shapefile.
@dask.delayed
def merge_bldgs(*args, **kwargs):
if os.path.exists(os.path.join(processed,'vectors','buildings.shp')):
pass
else:
building_tiles = glob.glob(os.path.join(processed, 'vectors', 'building_tiles', '*.shp'))
# create a list of geodataframes containing the tiles of building footprints
gdflist = [gpd.read_file(tile) for tile in building_tiles]
# merge them all together
merged = gpd.GeoDataFrame(pd.concat(gdflist, ignore_index=True))
# using pandas' concat caused us to lose projection information, so let's add that back in
merged.crs = gdflist[0].crs
# and write the merged data to a new shapefile
merged.to_file(os.path.join(processed,'vectors','buildings.shp'))
return
# identify the variety of rasters produced by gridmetrics
example_tile = tiles_to_merge[0]
grid_rasters = [os.path.basename(file).split(example_tile)[-1][1:-4] for file in
glob.glob(os.path.join(processed, 'rasters', 'gridmetrics_tiles', example_tile + '*'))
]
print('{:d} different types of rasters from gridmetrics to process for each tile:\r\n'.format(len(grid_rasters)))
for i, raster in enumerate(grid_rasters):
print('{}. {}'.format(i+1, raster))
# +
@dask.delayed
def merge_gridmetric(metric):
infiles = glob.glob(os.path.join(processed, 'rasters', 'gridmetrics_tiles', '*{}*.tif'.format(metric)))
outfile = os.path.join(processed, 'rasters', '{}.tif'.format(metric))
if os.path.exists(outfile):
pass
elif len(infiles) < 500:
proc = subprocess.run(['rio', 'merge', *infiles, outfile, '--co', 'compress=LZW',
'--co', 'tiled=true', #'--co', 'blockxsize=256', '--co', 'blockysize=256',
'--co', 'bigtiff=YES'],
stderr=subprocess.PIPE, stdout=subprocess.PIPE)
else:
merge_chunks(infiles, outfile)
# print(metric)
return metric
# -
# A single state that will depend upon the completion of the merged rasters and vectors.
@dask.delayed
def merge_done(*args, **kwargs):
return
# +
# building the computation receipe
merge_dsk = {}
merge_dsk['tile_boundaries'] = (tile_boundaries,)
merge_dsk['footprint'] = (make_footprint, 'tile_boundaries')
merge_dsk['merge_bldgs'] = (merge_bldgs,)
merge_dsk['merge_hill'] = (merge_hillshade,)
merge_dsk['merge_dem'] = (merge_dem,)
merge_dsk['merge_chm'] = (merge_chm,)
for raster in grid_rasters:
merge_dsk['merge_gridmetric-{}'.format(raster)] = (merge_gridmetric, raster)
merge_dsk['merge_done']=(merge_done, ['tile_boundaries', 'merge_bldgs', 'footprint'] +
['merge_hill', 'merge_dem', 'merge_chm'] +
['merge_gridmetric-{}'.format(raster) for raster in grid_rasters])
# -
merge_graph = c.get(merge_dsk, 'merge_done') # build the computation graph
merge_graph.visualize()
merge_results = c.compute(merge_graph) # this might take a while...
# +
# merge_results.result()
# -
progress(merge_results)
# progress(gridm_res)
# +
# procs = []
# for raster in grid_rasters:
# procs.append(merge_gridmetric(raster))
# +
# c.cancel(merge_results)
# +
# c.close()
# cluster.close()
| notebooks/05_Merge Tiled Derivatives Together.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parsers
#
# ## Parsing functions
#
# The `dataclass` `Parser` contains a single field, `func`.
# `func` is expected to be a function with the following signature:
#
# $$
# f: (Cursor, Aux) \longrightarrow (data, Cursor, Aux)
# $$
#
# Wait a second! Why is the function returning something else than the `data` itself?
#
# The reason is that, apart from the obvious output of `data`, it is very convenient to return an **updated** `Cursor` and an **updated** `Aux`.
# Typically, the updated `Cursor` will contain only the remaining, non-parsed content of the data.
# This is very convenient if we want to concatenate different parsers, passing the output of the first to the next one... and so on.
#
# Let's see an example:
#
# In this example we will create a very simple parser, that just reads the first letter of a text string.
from byteparsing import Cursor
# +
# Create some data to be parsed
data = b"Hello world!"
# Initialize the Cursor
c = Cursor(data, begin=0, end=0)
# Use an empty auxiliary variable
a = []
# Create a parsing function
def read_one(c, a):
c = c.increment() # Increase end index by one
x = c.content_str # Read content
c = c.flush() # Flush (i.e.: move begin to end)
return x, c, a
# -
# In the snippet below we see why it is convenient to use the updated `Cursor`.
while c:
x, c, a = read_one(c, a)
print(x)
# We can even write a new parsing function, based on the previous one, that parses to the end of the string:
def read_all(c, a):
x = [] # Initialize as empty list
while c:
temp, c, a = read_one(c, a)
x.append(temp)
return x, c, a
# Let's try it:
# +
# Restart the Cursor
c = Cursor(data, begin=0, end=0)
x, c, a = read_all(c, a)
print(x)
print(c)
# -
# Notice that the composition of two parsing functions (say, $f$ and $g$) is slightly more complicated than $f \circ g$, because the input and the output spaces of parsing functions are slightly different.
#
# For these and other reasons, it is advisable to manage parsing functions with a more flexible data structure.
#
# We introduce the Parser (data) class.
# ## The Parser class
#
# This section is work in progress.
#
# We'll manage parsing functions using a `Parser` class.
# `Parser` is a `dataclass` that contains a single field, `func`, representing a parser function.
#
# Let's build a `Parser` class from the `read_one` parsing function defined in the previous section:
from byteparsing.trampoline import Parser, parser
# +
# Initialize the Cursor
c = Cursor(data, begin=0, end=0)
# Use an empty auxiliary variable
a = []
# Create a function
def read_one(c, a):
c = c.increment() # Increase end index by one
x = c.content_str # Read content
c = c.flush() # Flush (i.e.: move begin to end)
return x, c, a
read_one_p = Parser(read_one)
# -
# Note: the lines above are entirely equivalent to:
#
# ```python
# @parser
# def read_one_p(c, a):
# c = c.increment() # Increase end index by one
# x = c.content_str # Read content
# c = c.flush() # Flush (i.e.: move begin to end)
# return x, c, a
# ```
# Parsers are callable, but they don't return anything informative until they are invoked:
print(read_one_p(c, a)) # Whithout invoking
x, c, a = read_one_p(c, a).invoke()
print(x)
print(c)
| docs/parsers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: drlnd
# language: python
# name: drlnd
# ---
# # Navigation
#
# ---
#
# Congratulations for completing the first project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893)! In this notebook, you will learn how to control an agent in a more challenging environment, where it can learn directly from raw pixels! **Note that this exercise is optional!**
#
# ### 1. Start the Environment
#
# We begin by importing some necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
from unityagents import UnityEnvironment
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/VisualBanana.app"`
# - **Windows** (x86): `"path/to/VisualBanana_Windows_x86/Banana.exe"`
# - **Windows** (x86_64): `"path/to/VisualBanana_Windows_x86_64/Banana.exe"`
# - **Linux** (x86): `"path/to/VisualBanana_Linux/Banana.x86"`
# - **Linux** (x86_64): `"path/to/VisualBanana_Linux/Banana.x86_64"`
# - **Linux** (x86, headless): `"path/to/VisualBanana_Linux_NoVis/Banana.x86"`
# - **Linux** (x86_64, headless): `"path/to/VisualBanana_Linux_NoVis/Banana.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `VisualBanana.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="VisualBanana.app")
# ```
env = UnityEnvironment(file_name="VisualBanana.app")
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 2. Examine the State and Action Spaces
#
# The simulation contains a single agent that navigates a large environment. At each time step, it has four actions at its disposal:
# - `0` - walk forward
# - `1` - walk backward
# - `2` - turn left
# - `3` - turn right
#
# The environment state is an array of raw pixels with shape `(1, 84, 84, 3)`. *Note that this code differs from the notebook for the project, where we are grabbing **`visual_observations`** (the raw pixels) instead of **`vector_observations`**.* A reward of `+1` is provided for collecting a yellow banana, and a reward of `-1` is provided for collecting a blue banana.
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents in the environment
print('Number of agents:', len(env_info.agents))
# number of actions
action_size = brain.vector_action_space_size
print('Number of actions:', action_size)
# examine the state space
state = env_info.visual_observations[0]
print('States look like:')
plt.imshow(np.squeeze(state))
plt.show()
state_size = state.shape
print('States have shape:', state.shape)
# -
# ### 3. Take Random Actions in the Environment
#
# In the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.
#
# Once this cell is executed, you will watch the agent's performance, if it selects an action (uniformly) at random with each time step. A window should pop up that allows you to observe the agent, as it moves through the environment.
#
# Of course, you'll have to change the code so that the agent is able to use its experience to gradually choose better actions when interacting with the environment!
# +
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.visual_observations[0] # get the current state
score = 0 # initialize the score
while True:
action = np.random.randint(action_size) # select an action
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.visual_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
# -
# When finished, you can close the environment.
env.close()
# ### 4. It's Your Turn!
#
# Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
# ```python
# env_info = env.reset(train_mode=True)[brain_name]
# ```
# +
from unityagents import UnityEnvironment
import numpy as np
# env = UnityEnvironment(file_name="VisualBanana_Windows_x86_64/Banana.exe")
env = UnityEnvironment(file_name="VisualBanana.app")
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# +
# Import the necessary package
import random
import torch
import numpy as np
from collections import deque
from skimage.transform import resize
import matplotlib.pyplot as plt
# %matplotlib inline
is_ipython = 'inline' in plt.get_backend()
if is_ipython:
from IPython import display
plt.ion()
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
torch.cuda.is_available()
# +
# initialize the Agent and reset enviornment
from dqn_agent_Pixels import Agent
agent = Agent(in_channels=3, action_size=4, seed=0)
# +
# Preprocess data
image_size = 28
def process_state(x):
x = np.squeeze(x)
x_resize = resize(x, (image_size, image_size, 3), anti_aliasing=True)
x_modified = torch.from_numpy(x_resize).float().to(device).view(3,image_size,image_size)
return x_modified
def process_stack(frames):
frames_list = list(frames)
torch_stack = torch.stack(frames_list).unsqueeze(dim=0)
torch_stack = torch_stack.view( torch_stack.size(0), torch_stack.size(2), torch_stack.size(1), torch_stack.size(3), torch_stack.size(4) )
return torch_stack
# +
#Train the Agent with DQN
def dqn(n_episodes = 2000, max_t = 1000, eps_start=1.0, eps_end=0.01, eps_decay=0.999):
"""Deep Q-Learning
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = []
scores_window = deque(maxlen=100)
eps = eps_start
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
state = env_info.visual_observations[0] # get the current state
score = 0 # initialize the score
stack_ready = False
frame_stack= deque()
stack_size = 3
is_state_processed = False
for t in range(max_t):
if not is_state_processed:
state = process_state(state)
if len(frame_stack) < stack_size:
for i in range(stack_size):
frame_stack.append(state)
else:
frame_stack.popleft()
frame_stack.append(state)
processed_stack = process_stack(frame_stack)
action = agent.act(processed_stack, eps) #.astype(int)
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.visual_observations[0] # get the next state
# process the next state
next_state = process_state(next_state)
next_fram_stack = frame_stack.copy()
next_fram_stack.popleft()
next_fram_stack.append(next_state)
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state=processed_stack, action=action,
reward=reward, next_state=process_stack(next_fram_stack),
done=done)
state = next_state # roll over the state to next time step
is_state_processed = True
score += reward # update the score
if done: # exit loop if episode finished
break
scores_window.append(score)
scores.append(score)
eps = max(eps_end, eps_decay*eps)
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Socre: {:.2f}'.format(i_episode, np.mean(scores_window)))
if np.mean(scores_window)>13.0:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
# Save the qnetwork_local weights to 'checkpointpixels.pth'
torch.save(agent.qnetwork_local.state_dict(), 'checkpointpixels.pth')
break
return scores
scores = dqn()
#plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# +
# load the weights from file
agent.qnetwork_local.load_state_dict(torch.load('checkpointpixels.pth'))
env_info = env.reset(train_mode=False)[brain_name]
state = env_info.vector_observations[0]
score = 0 # initialize the score
while True:
action = agent.act(state)
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state, action, reward, next_state, done) # the agent interact with environment
score += reward # update the score
state = next_state # roll over the state to next time step
if done: # exit loop if episode finished
break
print("Score: {}".format(score))
env.close()
# -
| (Optional) Challenge Learning from Pixels/Navigation_Pixels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Alternative model 4
#
# Working on the variables.
#
# Similar than model 3, but will use geometric mean to the definition of the amount of layers.
#
# Similarly with model 3:
#
# 1) Recoding the *Income amount* ("INCOME_AMT") variable to numerical. For this I will:
#
# a) use the middle amount in the range as representative in the category
#
# b) for missing values, will apply the median of the known values recoded as in point a.
#
# 2) Dropping redundant columns of data. For that, I will use get_dummies and apply drop_first.
#
#
# Imports
import pandas as pd
import numpy as np
from pathlib import Path
import tensorflow as tf
from tensorflow.keras.layers import Dense
from tensorflow.keras.models import Sequential
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from datetime import date
# ---
#
# ## Prepare the data to be used on a neural network model
# ### Step 1: Read the `applicants_data.csv` file into a Pandas DataFrame. Review the DataFrame, looking for categorical variables that will need to be encoded, as well as columns that could eventually define your features and target variables.
#
# +
# Read the applicants_data.csv file from the Resources folder into a Pandas DataFrame
applicant_data_df = pd.read_csv(
Path('../Resources/applicants_data.csv',)
)
# Review the DataFrame
applicant_data_df.head()
# -
# Review the data types associated with the columns
applicant_data_df.dtypes
# ### Step 2: Drop the “EIN” (Employer Identification Number) and “NAME” columns from the DataFrame, because they are not relevant to the binary classification model.
# +
# Drop the 'EIN' and 'NAME' columns from the DataFrame
applicant_data_df = applicant_data_df.drop(columns=['EIN','NAME'])
# Review the DataFrame
applicant_data_df.head()
# -
applicant_data_df['INCOME_AMT'].hist(xrot=90)
original_income_data_list=list(applicant_data_df['INCOME_AMT'].unique())
original_income_data_list
# Recoding income
recode_list=[0,5000,17500,300000,30000000,62500,50000000,3000000,7500000]
#Recoding Income amount
applicant_data_df_new=applicant_data_df['INCOME_AMT'].replace(original_income_data_list,recode_list)
median_income=applicant_data_df_new[applicant_data_df_new!=0].median()
applicant_data_df_new=applicant_data_df_new.replace(0,median_income)
applicant_data_df['INCOME_AMT']=applicant_data_df_new
applicant_data_df[['INCOME_AMT', 'ASK_AMT','IS_SUCCESSFUL']]
# ### Step 3: Encode the dataset’s categorical variables into a new DataFrame.
# +
# Create a list of categorical variables
categorical_variables = list(applicant_data_df.dtypes[applicant_data_df.dtypes=='object'].index)
# Display the categorical variables list
categorical_variables
# -
# Encode the variables and drop redundant columns
encoded_df = pd.get_dummies(
applicant_data_df[categorical_variables],
drop_first=True
)
encoded_df.head()
# ### Step 4: Add the original DataFrame’s numerical variables to the DataFrame containing the encoded variables.
#
# > **Note** To complete this step, you will employ the Pandas `concat()` function that was introduced earlier in this course.
non_categorical_variables = list(applicant_data_df.dtypes[applicant_data_df.dtypes!='object'].index)
non_categorical_variables
# +
# Add the numerical variables from the original DataFrame to the get_dummies encoding DataFrame
encoded_df =pd.concat([applicant_data_df[non_categorical_variables], encoded_df], axis=1)
# Review the Dataframe
encoded_df.head()
# -
# ### Step 5: Using the preprocessed data, create the features (`X`) and target (`y`) datasets. The target dataset should be defined by the preprocessed DataFrame column “IS_SUCCESSFUL”. The remaining columns should define the features dataset.
#
#
# +
# Define the target set y using the IS_SUCCESSFUL column
y = encoded_df['IS_SUCCESSFUL']
# Display a sample of y
y.head()
# +
# Define features set X by selecting all columns but IS_SUCCESSFUL
X = encoded_df.drop(columns=['IS_SUCCESSFUL'])
# Review the features DataFrame
X.head()
# -
# ### Step 6: Split the features and target sets into training and testing datasets.
#
# Split the preprocessed data into a training and testing dataset
# Assign the function a random_state equal to 1
X_train, X_test, y_train, y_test = train_test_split(X,y)
# ### Step 7: Use scikit-learn's `StandardScaler` to scale the features data.
# +
# Create a StandardScaler instance
scaler = StandardScaler()
# Fit the scaler to the features training dataset
X_scaler = scaler.fit(X_train)
# Fit the scaler to the features training dataset
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# -
#
# * number of nodes of hidden layers by using aritmetic mean between previous layer (or input) and the output
# +
# Define the the number of inputs (features) to the model
number_input_features = len(X_train.iloc[0])
# Review the number of features
number_input_features
# -
# Define the number of neurons in the output layer
number_output_neurons_A4 = 1
# +
# Define the number of hidden nodes for the first hidden layer
hidden_nodes_layer1_A4 = np.ceil(np.sqrt(number_input_features * number_output_neurons_A4))
# Review the number of hidden nodes in the first layer
hidden_nodes_layer1_A4
# +
# Define the number of hidden nodes for the first hidden layer
hidden_nodes_layer2_A4 = np.ceil(np.sqrt(hidden_nodes_layer1_A4 * number_output_neurons_A4))
# Review the number of hidden nodes in the first layer
hidden_nodes_layer2_A4
# -
# Create the Sequential model instance
nn_A4 = Sequential()
# +
# First hidden layer
nn_A4.add(
Dense(
units=hidden_nodes_layer1_A4,
activation='relu',
input_dim=number_input_features,
use_bias=True
)
)
#Second hidden layer
nn_A4.add(
Dense(
units=hidden_nodes_layer2_A4,
activation='relu',
use_bias=True
)
)
# Output layer
nn_A4.add(
Dense(
units=1,
activation='sigmoid'
)
)
# Check the structure of the model
nn_A4.summary()
# -
# Compile the Sequential model
nn_A4.compile(
loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
# Fit the model using 50 epochs and the training data
epochs=50
fit_model_A4 = nn_A4.fit(
X_train_scaled,
y_train,
epochs=epochs
)
# Evaluate the model loss and accuracy metrics using the evaluate method and the test data
model_loss, model_accuracy = nn_A4.evaluate(
X_test_scaled,
y_test,
verbose=2
)
# +
# Display the model loss and accuracy results
results_df=pd.DataFrame(
data={
'Date': [date.today()],
'Model' : [4],
'Observations': ['Recoding of data and Geometric Mean for Amount of Hidden Layers Definition'],
'Loss' : [model_loss],
'Accuracy': [model_accuracy],
'Hidden layers': 2,
'Input Features':[number_input_features],
'Output Neurons': [number_output_neurons_A4],
'Neurons in Hidden Layers1' : [hidden_nodes_layer1_A4],
'Neurons in Hidden Layers2' : [hidden_nodes_layer2_A4],
'Epochs' : [epochs],
'Activation Hidden layer 1': ['relu'],
'Activation Hidden layer 2': ['relu'],
'Activation Output layer 1': ['sigmoid'],
'loss':['binary_crossentropy'],
'optimizer':['adam'],
'metrics':['accuracy']
})
print(results_df.T)
# -
results_df.to_csv('../Resources/results.csv', mode='a')
# ### Step 3: Save each of your alternative models as an HDF5 file.
#
# +
# Set the model's file path
file_path = Path('../Resources/AlphabetSoup_Model_4.h5')
# Export your model to a HDF5 file
nn_A4.save(file_path)
# -
| Alternative models/4venture_funding_with_deep_learning_Alternative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Валидация расчета длительностей раундов
#
# В этом ноутбуке найдем и сравним оценки длительностей раундов при заданной вероятности битовой ошибки (BER) с помощью метода Монте-Карло и аналитической модели, построенной на урновой схеме.
# %load_ext autoreload
# %autoreload 2
# +
from rfidam.inventory import RoundModel, SlotModel, estimate_rounds_props, SlotValues
from rfidam.protocol.protocol import Protocol, LinkProps
from rfidam.utils import bcolors, get_err, fmt_err, highlight, pluralize
from rfidam.protocol.symbols import TagEncoding, DR
from tabulate import tabulate
from tqdm.notebook import tqdm
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
from itertools import product
from utility.jupyter_helpers import savefig, get_color, setup_matplotlib
# -
# %matplotlib inline
# Настроим Matplotlib: выберем цветовую схему и сделаем шрифты крупнее
setup_matplotlib()
# Определим вспомогательные методы для однотипной подстветки таблиц и форматирования строк.
# +
def fmt_header(s):
return highlight(s, bcolors.OKBLUE)
def fmt_delim(s):
return highlight(s, bcolors.OKCYAN)
def fmt_float(x):
return f'{x:.4f}'
# -
# ## Параметры модели
#
# Определим два сценария идентификации меток:
#
# - идентификация по EPCID
# - идентификация по TID и EPCID
#
# В обоих случаях будем использовать идентичные настройки протокола:
#
# - `Tari = 12.5us`
# - `RTcal = 37.5us`
# - `TRcal = 50us`
# - `DR = 64/3`
# - `TRext = 0`
# - `Q = 4`
# - `M = 2`
#
# Также будем считать, что длина EPCID равна 12 байтам, а TID - 8 байтам (4 словам).
# +
PROTOCOL_SETTINGS = {
'tari': 12.5e-6,
'rtcal': 37.5e-6,
'trcal': 50e-6,
'm': TagEncoding.M2,
'dr': DR.DR_643,
'trext': False,
'q': 4,
't_off': 0.1,
'n_epcid_bytes': 12,
'n_data_words': 4
}
LINK_PROPS = {
'epcid': LinkProps(**PROTOCOL_SETTINGS, use_tid=False),
'tid': LinkProps(**PROTOCOL_SETTINGS, use_tid=True)
}
PROTOCOLS = {key: Protocol(LINK_PROPS[key]) for key in LINK_PROPS}
# -
# Число меток будем брать из множества $N = 0, 1, 2, 3, 4, 10, 20$.
N_TAGS = np.arange(21)
BER = [.001, .005, .01, .02]
# ## Вычисление параметров раундов методом Монте-Карло
#
# Рассчитаем длины слотов и раундов методом Монте-Карло. Для этого будем моделировать процесс идентификации меток, собирать длительности слотов без ответов меток, слотов с попытками ответа одной метки и слотов с коллизиями.
# +
# We will store values for each protocol in matrices,
# where rows correspond to different BER values and columns - number of tags:
ESTIMATED_ROUND_DURATIONS = {key: [] for key in PROTOCOLS}
ESTIMATED_SLOTS_DURATIONS = {key: [] for key in PROTOCOLS}
ESTIMATED_SLOTS_COUNTS = {key: [] for key in PROTOCOLS}
TQDM_PROTOCOLS = tqdm(PROTOCOLS)
for protocol_name in TQDM_PROTOCOLS:
TQDM_PROTOCOLS.set_description(protocol_name)
TQDM_BER = tqdm(BER, leave=False)
protocol = PROTOCOLS[protocol_name]
for ber in TQDM_BER:
TQDM_BER.set_description(f'BER={ber}')
estimated_round_durations = []
estimated_slots_durations = []
estimated_slots_counts = []
TQDM_TAGS = tqdm(N_TAGS, leave=False)
for n_tags in TQDM_TAGS:
TQDM_TAGS.set_description(f'{n_tags} tags')
# Run Monte-Carlo:
round_duration, slots_durations, slots_counts = \
estimate_rounds_props(protocol, n_tags, ber, n_iters=range(2000))
estimated_round_durations.append(round_duration)
estimated_slots_durations.append(slots_durations)
estimated_slots_counts.append(slots_counts)
# Store results:
ESTIMATED_ROUND_DURATIONS[protocol_name].append(estimated_round_durations)
ESTIMATED_SLOTS_DURATIONS[protocol_name].append(estimated_slots_durations)
ESTIMATED_SLOTS_COUNTS[protocol_name].append(estimated_slots_counts)
# -
# ## Вычисление параметров раундов с помощью аналитической модели
#
# Теперь вычислим длительности раундов и слотов с помощью аналитической модели. Эта модель использует урновую схему (см. `rfidam.baskets`) для расчета среднего количества пустых слотов, слотов с одним ответом и слотов с коллизиями.
# +
# As above, we store models in dictionaries with the same keys sets as protocols.
# At each key, we store a matrix, in which:
# - rows correspond to BER values
# - columns correspond to different numbers of tags
ROUND_MODELS = {key: [] for key in PROTOCOLS}
# Also store round durations separately.
# We do so, since `round_durations` property is cached, so calling it will run estimation.
ANALYTIC_ROUND_DURATIONS = {key: [] for key in PROTOCOLS}
TQDM_PROTOCOLS = tqdm(PROTOCOLS)
for protocol_name in TQDM_PROTOCOLS:
TQDM_PROTOCOLS.set_description(protocol_name)
TQDM_BERS = tqdm(BER, leave=False)
protocol = PROTOCOLS[protocol_name]
for ber in TQDM_BERS:
TQDM_BERS.set_description(f'BER={ber}')
models = []
round_durations = []
for n_tags in N_TAGS:
# Build and store the model:
model = RoundModel(protocol, n_tags, ber)
models.append(model)
round_durations.append(model.round_duration)
# Store models for the given BER:
ROUND_MODELS[protocol_name].append(models)
ANALYTIC_ROUND_DURATIONS[protocol_name].append(round_durations)
# -
# ## Сравнение длин слотов
#
# Сначала сравним длительности слотов, рассчитанные из аналитической модели и методом Монте-Карло.
# +
# Compare slots durations estimated with Monte-Carlo method and analytic approach
BER_INDEX = -2
PROTOCOL_NAME = 'tid'
rows = []
for i, n_tags in enumerate(N_TAGS):
rounds_model = ROUND_MODELS[PROTOCOL_NAME][BER_INDEX][i]
estimated_round_duration = ESTIMATED_ROUND_DURATIONS[PROTOCOL_NAME][BER_INDEX][i]
estiamted_slots_durations = ESTIMATED_SLOTS_DURATIONS[PROTOCOL_NAME][BER_INDEX][i]
estimated_slots_counts = ESTIMATED_SLOTS_COUNTS[PROTOCOL_NAME][BER_INDEX][i]
# Compute slot probabilities:
estimated_slots_probs = SlotValues(
empty=(estimated_slots_counts.empty / protocol.props.n_slots),
collided=(estimated_slots_counts.collided / protocol.props.n_slots),
reply=(estimated_slots_counts.reply / protocol.props.n_slots))
# Build table rows:
value_rows = [
[fmt_header('round duration'), rounds_model.round_duration, estimated_round_duration],
[fmt_header('empty slot duration'), rounds_model.slots.durations.empty, estiamted_slots_durations.empty],
[fmt_header('collided slot duration'), rounds_model.slots.durations.collided, estiamted_slots_durations.collided],
[fmt_header('reply slot duration'), rounds_model.slots.durations.reply, estiamted_slots_durations.reply],
[fmt_header('empty slot probability'), rounds_model.slots.probs.empty, estimated_slots_probs.empty],
[fmt_header('collided slot probability'), rounds_model.slots.probs.collided, estimated_slots_probs.collided],
[fmt_header('reply slot probability'), rounds_model.slots.probs.reply, estimated_slots_probs.reply],
]
# Append errors estimations to each row:
for row in value_rows:
row.append(fmt_err(row[-1], row[-2]))
# Add rows to the main table:
title_row = [fmt_delim(f'{n_tags} tag{pluralize(n_tags)}')] + [''] * 3
rows.append(title_row)
rows.extend(value_rows)
headers = [fmt_header(s) for s in ['', 'Analytic', 'Simulation', 'Error']]
print(tabulate(rows, headers=headers))
# -
# ## Сравнение длительностей раундов
# +
fig, (ax1, ax2) = plt.subplots(figsize=(12, 4), nrows=1, ncols=2)
MARKERS = ['X', 'd', 'o', "s", 'v']
LINES = ['-', '-', '-', '-']
for i, ber in enumerate(BER):
color = get_color(i/len(BER) * 0.8 + 0.1)
marker = MARKERS[i % len(MARKERS)]
linestyle = LINES[i % len(LINES)]
ax1.plot(N_TAGS, ESTIMATED_ROUND_DURATIONS['epcid'][i],
label=f'BER={ber}, Монте-Карло', marker=marker,
color=color, linestyle="None", markersize=8)
ax1.plot(N_TAGS, ANALYTIC_ROUND_DURATIONS['epcid'][i],
label=f'BER={ber}, аналитика', color=color, linestyle=linestyle,
linewidth=2)
ax2.plot(N_TAGS, ESTIMATED_ROUND_DURATIONS['tid'][i],
label=f'BER={ber}, Монте-Карло', marker=marker, color=color,
linestyle="None", markersize=8)
ax2.plot(N_TAGS, ANALYTIC_ROUND_DURATIONS['tid'][i],
label=f'BER={ber}, аналитика', color=color, linestyle=linestyle,
linewidth=2)
lgd = fig.legend(*ax1.get_legend_handles_labels(), loc='lower center',
bbox_to_anchor=(0.5, -0.29), ncol=3)
ax1.set_title('Чтение EPCID')
ax2.set_title('Чтение TID')
# fig.suptitle('Round durations')
for ax in (ax1, ax2):
ax.grid()
ax.set_xlabel('Число меток')
ax.set_ylabel(f'Длительность, сек.')
plt.tight_layout()
savefig("ch3_round_durations_validation")
# plt.savefig('images/round_durations_validation.pdf', bbox_extra_artists=(lgd,), bbox_inches='tight')
# -
# ## Экспериментальный код
| experiments/2. Round durations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import matplotlib
import pandas
import abcutils
# ## Load and Synthesize Data from CSV
df = abcutils.load_and_synthesize_csv('sample_summaries.csv.gz')
df.head()
# Demonstrate how basic data filters are applied and filtered views are presented to this analysis script.
# +
filters = [
df['darshan_app'] == 'ior', # only want IOR jobs
df['darshan_biggest_api_bytes'] > 2.0**40, # only want jobs that did more than 1 TiB of I/O
]
pandas.options.display.max_rows = 11
abcutils.apply_filters(df, filters, verbose=True)
# -
# Demonstrate how columns can be normalized. This is the routine used to calculate the fraction of peak performance metric that is calculated for every job.
# +
target_col = 'darshan_agg_perf_by_slowest_posix'
group_by_cols = ['darshan_app', '_file_system', 'darshan_fpp_or_ssf_job', 'darshan_read_or_write_job']
new_col_base = 'darshan_normalized_perf'
# modifies the dataframe in-place; returns nothing
abcutils.normalize_column(
dataframe=df,
target_col=target_col,
group_by_cols=group_by_cols,
new_col_base=new_col_base)
# -
df[group_by_cols + [target_col]].head()
df[group_by_cols + [target_col] + [new_col_base + "_by_max"]].head()
# ## Correlation Matrix
ax, correlations = abcutils.plot.correlation_matrix(df, fontsize=14)
ax.get_figure().set_size_inches(30, 30)
# ## Correlation Table
# +
pandas.options.display.max_rows = 20
correlation = abcutils.correlation.calc_correlation_vector(df, correlate_with='darshan_normalized_perf_by_max')
filtered_correlations = abcutils.apply_filters(correlation, [correlation['p-value'] < 1.0e-5], verbose=True)
filtered_correlations.sort_values('coefficient')
# -
ax = abcutils.plot.correlation_vector_table(filtered_correlations, row_name_map=abcutils.CONFIG['metric_labels'])
ax.get_figure().set_size_inches(4, 0.4 * len(filtered_correlations))
# ## Boxplots
boxplot_settings = {
'fontsize': 20,
'darshan_normalized_perf_by_max': {
'output_file': "perf-boxplots.pdf",
'ylabel': "Fraction of\nPeak Performance",
'title_pos': [
{'x': 0.04, 'y': 0.02, 'horizontalalignment': 'left', 'fontsize': 14},
{'x': 0.04, 'y': 0.02, 'horizontalalignment': 'left', 'fontsize': 14}]
},
}
# Plot a single boxplot using the `plot.grouped_boxplot` function
ax = abcutils.plot.grouped_boxplot(df[df["_file_system"] == 'scratch1'],
'darshan_normalized_perf_by_max',
fontsize=16)
# Boxplots can also be inserted into existing figures with a little more effort. This can be handy for creating compact publication-ready diagrams.
# +
NUM_ROWS = 2
NUM_COLS = 2
fig, axes = matplotlib.pyplot.subplots(nrows=NUM_ROWS,
ncols=NUM_COLS,
# sharex causes problems if not all axes contain data
#sharex=True,
sharey=True)
fig.set_size_inches(8,6)
SUBPLOT_ARRANGEMENT = {
'scratch1': axes[0, 0],
'scratch2': axes[1, 0],
'scratch3': axes[1, 1],
}
NULL_SUBPLOTS = [
axes[0, 1],
]
### Draw subplots that contain data
for index, fs in enumerate(sorted(SUBPLOT_ARRANGEMENT.keys())):
irow = index / NUM_COLS
ax = SUBPLOT_ARRANGEMENT[fs]
abcutils.plot.grouped_boxplot(df[df["_file_system"] == fs],
'darshan_normalized_perf_by_max',
ax=ax,
fontsize=16)
title = ax.set_title(fs, **(boxplot_settings['darshan_normalized_perf_by_max']['title_pos'][irow]))
title.set_bbox({'color': 'white', 'alpha': 0.5})
### Hide subplots that do not contain data
for ax in NULL_SUBPLOTS:
ax.set_visible(False)
### Set global figure labels
fig.suptitle("")
fig.text(0.0, 0.5,
boxplot_settings['darshan_normalized_perf_by_max']['ylabel'],
verticalalignment='center',
horizontalalignment='center',
rotation='vertical',
fontsize=boxplot_settings['fontsize'])
fig.subplots_adjust(hspace=0.05, wspace=0.05)
# -
# ## Umami Diagrams
import time
import datetime
import tokio.tools.umami
# +
umami_diagrams = [
# The "I/O contention" case study figure
{
'filters': [
df['_file_system'] == 'scratch2',
df['darshan_app'] == 'hacc_io_write',
df['darshan_read_or_write_job'] == 'write',
df['_datetime_start'] > datetime.datetime(2017, 2, 14),
df['_datetime_start'] < datetime.datetime(2017, 3, 3, 0, 0, 0),
],
'rows': [
'darshan_agg_perf_by_slowest_posix',
'coverage_factor_bw',
'coverage_factor_nodehrs',
'fs_ave_mds_cpu',
'fs_tot_open_ops',
'topology_job_max_radius',
],
},
# The "storage capacity" case study figure
{
'filters': [
df['_file_system'] == 'scratch3',
df['darshan_app'] == 'hacc_io_write',
df['darshan_read_or_write_job'] == 'write',
df['_datetime_start'] > datetime.datetime(2017, 2, 21, 0, 0, 0),
df['_datetime_start'] < datetime.datetime(2017, 3, 15, 0, 0, 0),
],
'rows': [
'darshan_agg_perf_by_slowest_posix',
'coverage_factor_bw',
'fs_max_oss_cpu',
'fshealth_ost_most_full_pct',
],
},
]
pandas.options.display.max_rows = 11
filtered_df = abcutils.apply_filters(df, umami_diagrams[0]['filters'], verbose=True)
filtered_df.head().T
# -
for umami_diagram in umami_diagrams:
filtered_df = abcutils.apply_filters(df, umami_diagram['filters'], verbose=True)
fig = abcutils.plot.generate_umami(filtered_df, umami_diagram['rows'])
| abcutils_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from binance.client import Client
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime as dt
from matplotlib.pyplot import figure
# -
client = Client("", "")
symb='BTCUSDT_211231'
symb_spot = 'BTCUSDT'
# + jupyter={"outputs_hidden": true} tags=[]
data_futures = client.futures_historical_klines(symbol=symb, interval='1d',start_str = '1633039200000')
data_spot = client.get_historical_klines(symbol=symb_spot,interval='1d',start_str = '1633039200000',end_str='1640908800000')
print(data_futures)
print(data_spot)
print(float(data_futures[0][1])/float(data_spot[0][1]))
print(float(data_futures[-1][4])/float(data_spot[-1][4]))
# -
len(data_futures)
len(data_spot)
spread_open = [float(i[1])/float(z[1]) for i,z in zip(data_futures,data_spot)]
spread_time_diff = spread_close = [float(i[0])-float(z[0]) for i,z in zip(data_futures,data_spot)] #check that timestamps are equal
plt.plot(spread_time_diff)
spread_open_time = [dt.fromtimestamp(float(i[0])/1000) for i in data_futures]
spread_close = [float(i[4])/float(z[4]) for i,z in zip(data_futures,data_spot)]
spread_close_time = [dt.fromtimestamp(float(i[0])/1000) for i in data_futures]
figure(figsize=(8, 8), dpi=80)
plt.plot(spread_open_time,spread_open)
plt.ylabel('spread')
plt.xlabel('time')
plt.xticks(rotation=90)
plt.legend()
plt.show()
plt.plot(spread_close)
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Language Processing and Python
# ## Computing with Language: Texts and Words
#
from nltk.book import *
text1
# ### Searching Text
# shows every occurrence of a given word, together with somecontext
text1.concordance("monstrous")
text2.concordance("affection")
# **other words appear in a similar range of contexts**
text1.similar("monstrous")
# **common_contexts allows us to examine just the contexts that are shared by
# two or more words, such as monstrous and very.**
text2.common_contexts(["monstrous", "very"])
# Using a dispersion plot to show the location of a word in the text: how many words from the beginning it appears. Each stripe represents
# an instance of a word, and each row represents the entire text.
text4.dispersion_plot(["citizens", "democracy", "freedom", "duties", "America"])
# **generating some random text in the various styles we have just seen**
text3.generate()
# ### Counting Vocabulary
len(text3)
sorted(set(text3))
len(set(text3))
# **lexical richness of the text**
from __future__ import division
len(text3) / len(set(text3))
text3.count("smote")
# **compute what percentage of the text is taken up by a specific word**
100 * text4.count('a') / len(text4)
# **How many times does the word lol appear in text5? How
# much is this as a percentage of the total number of words in this text?**
text5.count("lol")
100 * text5.count('lol') / len(text5)
# +
def lexical_diversity(text):
return len(text) / len(set(text))
def percentage(count, total):
return 100 * count / total
# -
lexical_diversity(text3)
lexical_diversity(text5)
percentage(4, 5)
percentage(text4.count('a'), len(text4))
# ## A Closer Look at Python: Texts as Lists of Words##
# ### List ###
sent1 = ['Call', 'me', 'Ishmael', '.']
lexical_diversity(sent1)
sent1.append("Some")
sent1
# ### Indexing Lists ###
text4[173]
text4.index('awaken')
text5[16715:16735]
sent = ['word1', 'word2', 'word3', 'word4', 'word5', 'word6', 'word7', 'word8', 'word9', 'word10']
sent[5:8]
sent[7]
sent[:3] #sent 0 1 2
# ### String ###
name = 'Monty'
name[0]
name[:4]
name * 2
name + '!'
' '.join(['Monty', 'Python'])
'Monty Python'.split()
# ## Computing with Language: Simple Statistics ##
saying = ['After', 'all', 'is', 'said', 'and', 'done','more', 'is', 'said', 'than', 'done']
tokens = set(saying)
tokens
tokens = sorted(tokens)
tokens
tokens[-2:]
# ## Frequency Distributions ##
fdist1 = FreqDist(text1)
fdist1
vocabulary1 = fdist1.keys()
fdist1['whale']
fdist1.plot(50, cumulative=True)
#words that occur once only
fdist1.hapaxes()
#find the words from the vocabulary of the text that are more than 15 characters long
V = set(text1)
long_words = [w for w in V if len(w) > 15]
sorted(long_words)
# all words from the chat corpus that are longer than seven characters, that occur more than seven times
fdist5 = FreqDist(text5)
sorted([w for w in set(text5) if len(w) > 7 and fdist5[w] > 7])
# ## Collocations and Bigrams ##
# **a list of word pairs, also known as bigrams**
bigram = bigrams(['more', 'is', 'said', 'than', 'done'])
# fix bug with code "text4.collocations()"
print('; '.join(text4.collocation_list()))
# ## Counting Other Things ##
[len(w) for w in text1]
fdist = FreqDist([len(w) for w in text1])
fdist
fdist.keys()
fdist.items()
fdist.max()
fdist[3]
fdist.freq(3)
# |function|explaination|
# |----|-----|
# |fdist = FreqDist(samples)| Create a frequency distribution containing the given samples|
# |fdist.inc(sample) |Increment the count for this sample|
# |fdist['monstrous'] |Count of the number of times a given sample occurred|
# |fdist.freq('monstrous') |Frequency of a given sample|
# |fdist.N() |Total number of samples|
# |fdist.keys() |The samples sorted in order of decreasing frequency|
# |for sample in fdist: |Iterate over the samples, in order of decreasing frequency|
# |fdist.max() |Sample with the greatest count|
# |fdist.tabulate() |Tabulate the frequency distribution|
# |fdist.plot()| Graphical plot of the frequency distribution|
# |fdist.plot(cumulative=True) |Cumulative plot of the frequency distribution|
# |fdist1 |< fdist2 Test if samples in fdist1 occur less frequently than in fdist2|
# ### Some word comparison operators ###
# |function|explaination|
# |----|-----|
# |s.startswith(t) |Test if s starts with t|
# |s.endswith(t) |Test if s ends with t|
# |t in s |Test if t is contained inside s|
# |s.islower() |Test if all cased characters in s are lowercase|
# |s.isupper() |Test if all cased characters in s are uppercase|
# |s.isalpha() |Test if all characters in s are alphabetic|
# |s.isalnum() |Test if all characters in s are alphanumeric|
# |s.isdigit() |Test if all characters in s are digits|
# |s.istitle() |Test if s is titlecased (all words in s have initial capitals)|
sorted([w for w in set(text1) if w.endswith('ableness')])
sorted([term for term in set(text4) if 'gnt' in term])
sorted([item for item in set(text6) if item.istitle()])
sorted([item for item in set(sent7) if item.isdigit()])
[len(w) for w in text1]
[w.upper() for w in text1]
len(set([word.lower() for word in text1]))
#eliminate numbers and punctuation from the vocabulary count by filtering out any non-alphabetic items
len(set([word.lower() for word in text1 if word.isalpha()]))
| NLP with Python/Chapter01/Chapter1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="t09eeeR5prIJ"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="GCCk8_dHpuNf"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="ovpZyIhNIgoq"
# # RNN によるテキスト生成
# + [markdown] colab_type="text" id="hcD2nPQvPOFM"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/text/text_generation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/tutorials/text/text_generation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/tutorials/text/text_generation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/tutorials/text/text_generation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="RoRi17kuCYFs"
# Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [<EMAIL> メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。
# + [markdown] colab_type="text" id="BwpJ5IffzRG6"
# このチュートリアルでは、文字ベースの RNN を使ってテキストを生成する方法を示します。ここでは、<NAME> の [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) からのシェイクスピア作品のデータセットを使います。このデータからの文字列("Shakespear")を入力にして、文字列中の次の文字("e")を予測するモデルを訓練します。このモデルを繰り返し呼び出すことで、より長い文字列を生成することができます。
#
# Note: このノートブックの実行を速くするために GPU による高速化を有効にしてください。Colab では、*ランタイム > ランタイムのタイプを変更 > ハードウェアアクセラレータ > GPU* を選択します。ローカルで実行する場合には、TensorFlow のバージョンが 1.11 以降であることを確認してください。
#
# このチュートリアルには、[tf.keras](https://www.tensorflow.org/programmers_guide/keras) と [eager execution](https://www.tensorflow.org/programmers_guide/eager) を使ったコードが含まれています。下記は、このチュートリアルのモデルを 30 エポック訓練したものに対して、文字列 "Q" を初期値とした場合の出力例です。
#
# <pre>
# QUEENE:
# I had thought thou hadst a Roman; for the oracle,
# Thus by All bids the man against the word,
# Which are so weak of care, by old care done;
# Your children were in your holy love,
# And the precipitation through the bleeding throne.
#
# BISHOP OF ELY:
# Marry, and will, my lord, to weep in such a one were prettiest;
# Yet now I was adopted heir
# Of the world's lamentable day,
# To watch the next way with his father with his face?
#
# ESCALUS:
# The cause why then we are all resolved more sons.
#
# VOLUMNIA:
# O, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, no, it is no sin it should be dead,
# And love and pale as any will to that word.
#
# QUEEN ELIZABETH:
# But how long have I heard the soul for this world,
# And show his hands of life be proved to stand.
#
# PETRUCHIO:
# I say he look'd on, if I must be content
# To stay him from the fatal of our country's bliss.
# His lordship pluck'd from this sentence then for prey,
# And then let us twain, being the moon,
# were she such a case as fills m
# </pre>
#
# いくつかは文法にあったものがある一方で、ほとんどは意味をなしていません。このモデルは、単語の意味を学習していませんが、次のことを考えてみてください。
#
# * このモデルは文字ベースです。訓練が始まった時に、モデルは英語の単語のスペルも知りませんし、単語がテキストの単位であることも知らないのです。
#
# * 出力の構造は戯曲に似ています。だいたいのばあい、データセットとおなじ大文字で書かれた話し手の名前で始まっています。
#
# * 以下に示すように、モデルはテキストの小さなバッチ(各100文字)で訓練されていますが、一貫した構造のより長いテキストのシーケンスを生成できます。
# + [markdown] colab_type="text" id="srXC6pLGLwS6"
# ## 設定
# + [markdown] colab_type="text" id="WGyKZj3bzf9p"
# ### TensorFlow 等のライブラリインポート
# + colab={} colab_type="code" id="yG_n40gFzf9s"
import tensorflow as tf
import numpy as np
import os
import time
# + [markdown] colab_type="text" id="EHDoRoc5PKWz"
# ### シェイクスピアデータセットのダウンロード
#
# 独自のデータで実行するためには下記の行を変更してください。
# + colab={} colab_type="code" id="pD_55cOxLkAb"
path_to_file = tf.keras.utils.get_file('shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
# + [markdown] colab_type="text" id="UHjdCjDuSvX_"
# ### データの読み込み
#
# まずはテキストをのぞいてみましょう。
# + colab={} colab_type="code" id="aavnuByVymwK"
# 読み込んだのち、Python 2 との互換性のためにデコード
text = open(path_to_file, 'rb').read().decode(encoding='utf-8')
# テキストの長さは含まれる文字数
print ('Length of text: {} characters'.format(len(text)))
# + colab={} colab_type="code" id="Duhg9NrUymwO"
# テキストの最初の 250文字を参照
print(text[:250])
# + colab={} colab_type="code" id="IlCgQBRVymwR"
# ファイル中のユニークな文字の数
vocab = sorted(set(text))
print ('{} unique characters'.format(len(vocab)))
# + [markdown] colab_type="text" id="rNnrKn_lL-IJ"
# ## テキストの処理
# + [markdown] colab_type="text" id="LFjSVAlWzf-N"
# ### テキストのベクトル化
#
# 訓練をする前に、文字列を数値表現に変換する必要があります。2つの参照テーブルを作成します。一つは文字を数字に変換するもの、もう一つは数字を文字に変換するものです。
# + colab={} colab_type="code" id="IalZLbvOzf-F"
# それぞれの文字からインデックスへの対応表を作成
char2idx = {u:i for i, u in enumerate(vocab)}
idx2char = np.array(vocab)
text_as_int = np.array([char2idx[c] for c in text])
# + [markdown] colab_type="text" id="tZfqhkYCymwX"
# これで、それぞれの文字を整数で表現できました。文字を、0 から`len(unique)` までのインデックスに変換していることに注意してください。
# + colab={} colab_type="code" id="FYyNlCNXymwY"
print('{')
for char,_ in zip(char2idx, range(20)):
print(' {:4s}: {:3d},'.format(repr(char), char2idx[char]))
print(' ...\n}')
# + colab={} colab_type="code" id="l1VKcQHcymwb"
# テキストの最初の 13 文字がどのように整数に変換されるかを見てみる
print ('{} ---- characters mapped to int ---- > {}'.format(repr(text[:13]), text_as_int[:13]))
# + [markdown] colab_type="text" id="bbmsf23Bymwe"
# ### 予測タスク
# + [markdown] colab_type="text" id="wssHQ1oGymwe"
# ある文字、あるいは文字列が与えられたとき、もっともありそうな次の文字はなにか?これが、モデルを訓練してやらせたいタスクです。モデルへの入力は文字列であり、モデルが出力、つまりそれぞれの時点での次の文字を予測をするようにモデルを訓練します。
#
# RNN はすでに見た要素に基づく内部状態を保持しているため、この時点までに計算されたすべての文字を考えると、次の文字は何でしょうか?
# + [markdown] colab_type="text" id="hgsVvVxnymwf"
# ### 訓練用サンプルとターゲットを作成
#
# つぎに、テキストをサンプルシーケンスに分割します。それぞれの入力シーケンスは、元のテキストからの `seq_length` 個の文字を含みます。
#
# 入力シーケンスそれぞれに対して、対応するターゲットは同じ長さのテキストを含みますが、1文字ずつ右にシフトしたものです。
#
# そのため、テキストを `seq_length+1` のかたまりに分割します。たとえば、 `seq_length` が 4 で、テキストが "Hello" だとします。入力シーケンスは "Hell" で、ターゲットシーケンスは "ello" となります。
#
# これを行うために、最初に `tf.data.Dataset.from_tensor_slices` 関数を使ってテキストベクトルを文字インデックスの連続に変換します。
# + colab={} colab_type="code" id="0UHJDA39zf-O"
# ひとつの入力としたいシーケンスの文字数としての最大の長さ
seq_length = 100
examples_per_epoch = len(text)//(seq_length+1)
# 訓練用サンプルとターゲットを作る
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
for i in char_dataset.take(5):
print(idx2char[i.numpy()])
# + [markdown] colab_type="text" id="-ZSYAcQV8OGP"
# `batch` メソッドを使うと、個々の文字を求める長さのシーケンスに簡単に変換できます。
# + colab={} colab_type="code" id="l4hkDU3i7ozi"
sequences = char_dataset.batch(seq_length+1, drop_remainder=True)
for item in sequences.take(5):
print(repr(''.join(idx2char[item.numpy()])))
# + [markdown] colab_type="text" id="UbLcIPBj_mWZ"
# シーケンスそれぞれに対して、`map` メソッドを使って各バッチに単純な関数を適用することで、複製とシフトを行い、入力テキストとターゲットテキストを生成します。
# + colab={} colab_type="code" id="9NGu-FkO_kYU"
def split_input_target(chunk):
input_text = chunk[:-1]
target_text = chunk[1:]
return input_text, target_text
dataset = sequences.map(split_input_target)
# + [markdown] colab_type="text" id="hiCopyGZymwi"
# 最初のサンプルの入力とターゲットを出力します。
# + colab={} colab_type="code" id="GNbw-iR0ymwj"
for input_example, target_example in dataset.take(1):
print ('Input data: ', repr(''.join(idx2char[input_example.numpy()])))
print ('Target data:', repr(''.join(idx2char[target_example.numpy()])))
# + [markdown] colab_type="text" id="_33OHL3b84i0"
# これらのベクトルのインデックスそれぞれが一つのタイムステップとして処理されます。タイムステップ 0 の入力として、モデルは "F" のインデックスを受け取り、次の文字として "i" のインデックスを予測しようとします。次のタイムステップでもおなじことをしますが、`RNN` は現在の入力文字に加えて、過去のステップのコンテキストも考慮します。
# + colab={} colab_type="code" id="0eBu9WZG84i0"
for i, (input_idx, target_idx) in enumerate(zip(input_example[:5], target_example[:5])):
print("Step {:4d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
# + [markdown] colab_type="text" id="MJdfPmdqzf-R"
# ### 訓練用バッチの作成
#
# `tf.data` を使ってテキストを分割し、扱いやすいシーケンスにします。しかし、このデータをモデルに供給する前に、データをシャッフルしてバッチにまとめる必要があります。
# + colab={} colab_type="code" id="p2pGotuNzf-S"
# バッチサイズ
BATCH_SIZE = 64
# データセットをシャッフルするためのバッファサイズ
# (TF data は可能性として無限長のシーケンスでも使えるように設計されています。
# このため、シーケンス全体をメモリ内でシャッフルしようとはしません。
# その代わりに、要素をシャッフルするためのバッファを保持しています)
BUFFER_SIZE = 10000
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
dataset
# + [markdown] colab_type="text" id="r6oUuElIMgVx"
# ## モデルの構築
# + [markdown] colab_type="text" id="m8gPwEjRzf-Z"
# `tf.keras.Sequential` を使ってモデルを定義します。この簡単な例では、モデルの定義に3つのレイヤーを使用しています。
#
# * `tf.keras.layers.Embedding`: 入力レイヤー。それぞれの文字を表す数を `embedding_dim` 次元のベクトルに変換する、訓練可能な参照テーブル。
# * `tf.keras.layers.GRU`: サイズが `units=rnn_units` のRNNの一種(ここに LSTM レイヤーを使うこともできる)。
# * `tf.keras.layers.Dense`: `vocab_size` の出力を持つ、出力レイヤー。
# + colab={} colab_type="code" id="zHT8cLh7EAsg"
# 文字数で表されるボキャブラリーの長さ
vocab_size = len(vocab)
# 埋め込みベクトルの次元
embedding_dim = 256
# RNN ユニットの数
rnn_units = 1024
# + colab={} colab_type="code" id="MtCrdfzEI2N0"
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape=[batch_size, None]),
tf.keras.layers.GRU(rnn_units,
return_sequences=True,
stateful=True,
recurrent_initializer='glorot_uniform'),
tf.keras.layers.Dense(vocab_size)
])
return model
# + colab={} colab_type="code" id="wwsrpOik5zhv"
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
# + [markdown] colab_type="text" id="RkA5upJIJ7W7"
# 1文字ごとにモデルは埋め込みベクトルを検索し、その埋め込みベクトルを入力として GRU を 1 タイムステップ実行します。そして Dense レイヤーを適用して、次の文字の対数尤度を予測するロジットを生成します。
#
# 
# + [markdown] colab_type="text" id="-ubPo0_9Prjb"
# ## モデルを試す
#
# 期待通りに動作するかどうかを確認するためモデルを動かしてみましょう。
#
# 最初に、出力の shape を確認します。
# + colab={} colab_type="code" id="C-_70kKAPrPU"
for input_example_batch, target_example_batch in dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(example_batch_predictions.shape, "# (batch_size, sequence_length, vocab_size)")
# + [markdown] colab_type="text" id="Q6NzLBi4VM4o"
# 上記の例では、入力のシーケンスの長さは `100` ですが、モデルはどのような長さの入力でも実行できます。
# + colab={} colab_type="code" id="vPGmAAXmVLGC"
model.summary()
# + [markdown] colab_type="text" id="uwv0gEkURfx1"
# モデルから実際の予測を得るには出力の分布からサンプリングを行う必要があります。この分布は、文字ボキャブラリー全体のロジットで定義されます。
#
# Note: この分布から _サンプリング_ するということが重要です。なぜなら、分布の _argmax_ をとったのでは、モデルは簡単にループしてしまうからです。
#
# バッチ中の最初のサンプルで試してみましょう。
# + colab={} colab_type="code" id="4V4MfFg0RQJg"
sampled_indices = tf.random.categorical(example_batch_predictions[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices,axis=-1).numpy()
# + [markdown] colab_type="text" id="QM1Vbxs_URw5"
# これにより、タイムステップそれぞれにおいて、次の文字のインデックスの予測が得られます。
# + colab={} colab_type="code" id="YqFMUQc_UFgM"
sampled_indices
# + [markdown] colab_type="text" id="LfLtsP3mUhCG"
# これらをデコードすることで、この訓練前のモデルによる予測テキストをみることができます。
# + colab={} colab_type="code" id="xWcFwPwLSo05"
print("Input: \n", repr("".join(idx2char[input_example_batch[0]])))
print()
print("Next Char Predictions: \n", repr("".join(idx2char[sampled_indices ])))
# + [markdown] colab_type="text" id="LJL0Q0YPY6Ee"
# ## モデルの訓練
# + [markdown] colab_type="text" id="YCbHQHiaa4Ic"
# ここまでくれば問題は標準的な分類問題として扱うことができます。これまでの RNN の状態と、いまのタイムステップの入力が与えられ、次の文字のクラスを予測します。
# + [markdown] colab_type="text" id="trpqTWyvk0nr"
# ### オプティマイザと損失関数の付加
# + [markdown] colab_type="text" id="UAjbjY03eiQ4"
# この場合、標準の `tf.keras.losses.sparse_categorical_crossentropy` 損失関数が使えます。予測の最後の次元に適用されるからです。
#
# このモデルはロジットを返すので、`from_logits` フラグをセットする必要があります。
# + colab={} colab_type="code" id="4HrXTACTdzY-"
def loss(labels, logits):
return tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True)
example_batch_loss = loss(target_example_batch, example_batch_predictions)
print("Prediction shape: ", example_batch_predictions.shape, " # (batch_size, sequence_length, vocab_size)")
print("scalar_loss: ", example_batch_loss.numpy().mean())
# + [markdown] colab_type="text" id="jeOXriLcymww"
# `tf.keras.Model.compile` を使って、訓練手順を定義します。既定の引数を持った `tf.keras.optimizers.Adam` と、先ほどの loss 関数を使用しましょう。
# + colab={} colab_type="code" id="DDl1_Een6rL0"
model.compile(optimizer='adam', loss=loss)
# + [markdown] colab_type="text" id="ieSJdchZggUj"
# ### チェックポイントの構成
# + [markdown] colab_type="text" id="C6XBUUavgF56"
# `tf.keras.callbacks.ModelCheckpoint` を使って、訓練中にチェックポイントを保存するようにします。
# + colab={} colab_type="code" id="W6fWTriUZP-n"
# チェックポイントが保存されるディレクトリ
checkpoint_dir = './training_checkpoints'
# チェックポイントファイルの名称
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
checkpoint_callback=tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_prefix,
save_weights_only=True)
# + [markdown] colab_type="text" id="3Ky3F_BhgkTW"
# ### 訓練の実行
# + [markdown] colab_type="text" id="IxdOA-rgyGvs"
# 訓練時間を適切に保つために、10エポックを使用してモデルを訓練します。Google Colab を使用する場合には、訓練を高速化するためにランタイムを GPU に設定します。
# + colab={} colab_type="code" id="7yGBE2zxMMHs"
EPOCHS=10
# + colab={} colab_type="code" id="UK-hmKjYVoll"
history = model.fit(dataset, epochs=EPOCHS, callbacks=[checkpoint_callback])
# + [markdown] colab_type="text" id="kKkD5M6eoSiN"
# ## テキスト生成
# + [markdown] colab_type="text" id="JIPcXllKjkdr"
# ### 最終チェックポイントの復元
# + [markdown] colab_type="text" id="LyeYRiuVjodY"
# 予測ステップを単純にするため、バッチサイズ 1 を使用します。
#
# RNN が状態をタイムステップからタイムステップへと渡す仕組みのため、モデルは一度構築されると固定されたバッチサイズしか受け付けられません。
#
# モデルを異なる `batch_size` で実行するためには、モデルを再構築し、チェックポイントから重みを復元する必要があります。
# + colab={} colab_type="code" id="zk2WJ2-XjkGz"
tf.train.latest_checkpoint(checkpoint_dir)
# + colab={} colab_type="code" id="LycQ-ot_jjyu"
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model.build(tf.TensorShape([1, None]))
# + colab={} colab_type="code" id="71xa6jnYVrAN"
model.summary()
# + [markdown] colab_type="text" id="DjGz1tDkzf-u"
# ### 予測ループ
#
# 下記のコードブロックでテキストを生成します。
#
# * 最初に、開始文字列を選択し、RNN の状態を初期化して、生成する文字数を設定します。
#
# * 開始文字列と RNN の状態を使って、次の文字の予測分布を得ます。
#
# * つぎに、カテゴリー分布を使用して、予測された文字のインデックスを計算します。この予測された文字をモデルの次の入力にします。
#
# * モデルによって返された RNN の状態はモデルにフィードバックされるため、1つの文字だけでなく、より多くのコンテキストを持つことになります。つぎの文字を予測した後、更新された RNN の状態が再びモデルにフィードバックされます。こうしてモデルは以前に予測した文字からさらにコンテキストを得ることで学習するのです。
#
# 
#
# 生成されたテキストを見ると、モデルがどこを大文字にするかや、段落の区切り方、シェークスピアらしい書き言葉を真似ることを知っていることがわかります。しかし、訓練のエポック数が少ないので、まだ一貫した文章を生成するところまでは学習していません。
# + colab={} colab_type="code" id="WvuwZBX5Ogfd"
def generate_text(model, start_string):
# 評価ステップ(学習済みモデルを使ったテキスト生成)
# 生成する文字数
num_generate = 1000
# 開始文字列を数値に変換(ベクトル化)
input_eval = [char2idx[s] for s in start_string]
input_eval = tf.expand_dims(input_eval, 0)
# 結果を保存する空文字列
text_generated = []
# 低い temperature は、より予測しやすいテキストをもたらし
# 高い temperature は、より意外なテキストをもたらす
# 実験により最適な設定を見つけること
temperature = 1.0
# ここではバッチサイズ == 1
model.reset_states()
for i in range(num_generate):
predictions = model(input_eval)
# バッチの次元を削除
predictions = tf.squeeze(predictions, 0)
# カテゴリー分布をつかってモデルから返された文字を予測
predictions = predictions / temperature
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# 過去の隠れ状態とともに予測された文字をモデルへのつぎの入力として渡す
input_eval = tf.expand_dims([predicted_id], 0)
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
# + colab={} colab_type="code" id="ktovv0RFhrkn"
print(generate_text(model, start_string=u"ROMEO: "))
# + [markdown] colab_type="text" id="AM2Uma_-yVIq"
# この結果を改善するもっとも簡単な方法は、もっと長く訓練することです(`EPOCHS=30` を試してみましょう)。
#
# また、異なる初期文字列を使ったり、モデルの精度を向上させるためにもうひとつ RNN レイヤーを加えたり、temperature パラメータを調整して、よりランダム性の強い、あるいは、弱い予測を試してみたりすることができます。
# + [markdown] colab_type="text" id="Y4QwTjAM6A2O"
# ## 上級編: 訓練のカスタマイズ
#
# 上記の訓練手順は単純ですが、制御できるところがそれほどありません。
#
# モデルを手動で実行する方法を見てきたので、訓練ループを展開し、自分で実装してみましょう。このことが、たとえばモデルのオープンループによる出力を安定化するための _カリキュラム学習_ を実装するための出発点になります。
#
# 勾配を追跡するために `tf.GradientTape` を使用します。このアプローチについての詳細を学ぶには、 [eager execution guide](https://www.tensorflow.org/guide/eager) をお読みください。
#
# この手順は下記のように動作します。
#
# * 最初に、RNN の状態を初期化する。`tf.keras.Model.reset_states` メソッドを呼び出すことでこれを実行する。
#
# * つぎに、(1バッチずつ)データセットを順番に処理し、それぞれのバッチに対する*予測値*を計算する。
#
# * `tf.GradientTape` をオープンし、そのコンテキストで、予測値と損失を計算する。
#
# * `tf.GradientTape.grads` メソッドを使って、モデルの変数に対する損失の勾配を計算する。
#
# * 最後に、オプティマイザの `tf.train.Optimizer.apply_gradients` メソッドを使って、逆方向の処理を行う。
# + colab={} colab_type="code" id="_XAm7eCoKULT"
model = build_model(
vocab_size = len(vocab),
embedding_dim=embedding_dim,
rnn_units=rnn_units,
batch_size=BATCH_SIZE)
# + colab={} colab_type="code" id="qUKhnZtMVpoJ"
optimizer = tf.keras.optimizers.Adam()
# + colab={} colab_type="code" id="b4kH1o0leVIp"
@tf.function
def train_step(inp, target):
with tf.GradientTape() as tape:
predictions = model(inp)
loss = tf.reduce_mean(
tf.keras.losses.sparse_categorical_crossentropy(
target, predictions, from_logits=True))
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
return loss
# + colab={} colab_type="code" id="d4tSNwymzf-q"
# 訓練ステップ
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
# 各エポックの最初に、隠れ状態を初期化する
# 最初は隠れ状態は None
hidden = model.reset_states()
for (batch_n, (inp, target)) in enumerate(dataset):
loss = train_step(inp, target)
if batch_n % 100 == 0:
template = 'Epoch {} Batch {} Loss {}'
print(template.format(epoch+1, batch_n, loss))
# 5エポックごとにモデル(のチェックポイント)を保存する
if (epoch + 1) % 5 == 0:
model.save_weights(checkpoint_prefix.format(epoch=epoch))
print ('Epoch {} Loss {:.4f}'.format(epoch+1, loss))
print ('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
model.save_weights(checkpoint_prefix.format(epoch=epoch))
| site/ja/tutorials/text/text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
url = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv'
url1 = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv'
url2 = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv'
confirmed_data = pd.read_csv(url)
death_data = pd.read_csv(url1)
# -
recovered_data=pd.read_csv(url2)
confirmed_data.head()
death_data.head()
recovered_data.head()
confirmed_data.info()
confirmed_data.describe()
confirmed_data.isna().sum()
df=confirmed_data.iloc[:,4:]
df.columns
dic={}
for i in df.columns:
key = i
values = [confirmed_data['Province/State'].values.tolist(),confirmed_data['Country/Region'].values.tolist(),confirmed_data[i].values.tolist(),death_data[i].values.tolist()]
dic[key] = values
d=pd.DataFrame(dic)
d=d.T
d.columns = ["Province/State",'Country',"confirmed_case",'Death_case']
d.index.name='Date'
d
d.reset_index(level=0, inplace=True)
pd.set_option('display.max_columns',None)
[1,2,3]==[1,2,3]
d=d.set_index(['Date']).apply(pd.Series.explode).reset_index()
d
main_data = d[['Province/State', 'Country','Date', "confirmed_case",'Death_case']]
main_data
main_data['Date']=pd.to_datetime(main_data['Date'])
main_data
main_data.to_csv("main.csv")
#Data for Nepal
main_data.query('Country=="Nepal"').groupby("Date")[['confirmed_case', 'Death_case']].sum().reset_index()
# +
#Overall worldwide Data
overall_data=main_data.groupby("Date")['confirmed_case','Death_case'].sum().reset_index()
overall_data
# +
import matplotlib.pyplot as plt
plt.plot(overall_data['Date'],overall_data['confirmed_case'],label="Confirmed")
plt.plot(overall_data['Date'],overall_data['Death_case'],label="Death")
plt.legend()
# -
# ### Confirmed Case
overall_confirmed = overall_data.iloc[:,0:-1]
overall_confirmed
| notebooks/Data creating.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Envío de valores
# Para comunicarse con el exterior, las funciones no sólo pueden devolver valores, también pueden recibir información:
def suma(a,b): # valores que se reciben
return a+b
# #### Ahora podemos enviar dos valores a la función:
r = suma(2,5) # valores que se envían
r
# ## Parámetros y argumentos
# En la definición de una función, los valores que se reciben se denominan **parámetros**, y en la llamada se denominan **argumentos**.
| MaterialCursoPython/Fase 2 - Manejo de datos y optimizacion/Tema 06 - Programacion de funciones/Apuntes/Leccion 3 (Apuntes) - Envio de valores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
from datasets import Dataset0
from presets import SegmentationPresetTrain, SegmentationPresetEval
from matplotlib.pyplot import imshow, subplot
import utils
import torch
import numpy as np
ds1 = Dataset0("../BIO_data/RetinaDataset", transforms=SegmentationPresetEval(100, 100))
print(len(ds1))
img, target = ds1[1]
imshow(target)
logger = utils.MetricLogger()
train_sampler = torch.utils.data.RandomSampler(ds1)
data_loader = torch.utils.data.DataLoader(
ds1, batch_size=32,
sampler=train_sampler, num_workers=4,
collate_fn=utils.collate_fn, drop_last=True)
tmp = None
for i in data_loader:
tmp = i
break
len(tmp)
# +
ax1 = subplot(212)
ax1.margins(0.05) # Default margin is 0.05, value 0 means fit
ax1.imshow(tmp[0][0][0])
ax2 = subplot(221)
ax2.margins(2, 2) # Values >0.0 zoom out
ax2.imshow(tmp[1][0][0])
ax3 = subplot(222)
ax3.margins(2, 2) # Values >0.0 zoom out
ax3.imshow(tmp[1][0][1])
# +
import os
root = "../BIO_data/DB_UoA"
objs = list(filter(os.path.isdir, [os.path.join(root, d) for d in os.listdir(root)]))
objs.sort()
images = []
targets = []
target_type = ("idk", "idk2", "idk3")
for obj in objs:
files = [os.path.join(obj, f) for f in os.listdir(obj)]
files.sort()
images.append(files[-1])
targets.append(files[:-1])
# +
import os
root = "../BIO_data/Database"
split = "test"
dataset_path = {
"train" : "a. Training Set",
"test" : "b. Testing Set"
}
if split not in dataset_path:
assert "err"
images = os.listdir(os.path.join(root, "A. Segmentation", "1. Original Images", dataset_path[split] ) )
images = [ os.path.join(root, "A. Segmentation", "1. Original Images", dataset_path[split], f) for f in images]
images.sort()
targets = {
"1. Microaneurysms" : None,
"2. Haemorrhages" : None,
"3. Hard Exudates" : None,
"4. Soft Exudates" : None,
"5. Optic Disc" : None,
}
for t in targets:
targets[t] = os.listdir( os.path.join(root , "A. Segmentation" , "2. All Segmentation Groundtruths" , dataset_path[split] , t) )
targets[t] = [ os.path.join(root , "A. Segmentation" , "2. All Segmentation Groundtruths" , dataset_path[split] , t, f) for f in targets[t]]
targets[t].sort()
# -
for i in targets:
print (targets[i][:1])
| testing_ntb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrdbourke/tensorflow-deep-learning/blob/main/04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ObwiuiGkZb87"
# # 04. Transfer Learning with TensorFlow Part 1: Feature Extraction
#
# We've built a bunch of convolutional neural networks from scratch and they all seem to be learning, however, there is still plenty of room for improvement.
#
# To improve our model(s), we could spend a while trying different configurations, adding more layers, changing the learning rate, adjusting the number of neurons per layer and more.
#
# However, doing this is very time consuming.
#
# Luckily, there's a technique we can use to save time.
#
# It's called **transfer learning**, in other words, taking the patterns (also called weights) another model has learned from another problem and using them for our own problem.
#
# There are two main benefits to using transfer learning:
# 1. Can leverage an existing neural network architecture proven to work on problems similar to our own.
# 2. Can leverage a working neural network architecture which has **already learned** patterns on similar data to our own. This often results in achieving great results with less custom data.
#
# What this means is, instead of hand-crafting our own neural network architectures or building them from scratch, we can utilise models which have worked for others.
#
# And instead of training our own models from scratch on our own datasets, we can take the patterns a model has learned from datasets such as [ImageNet](http://www.image-net.org/) (millions of images of different objects) and use them as the foundation of our own. Doing this often leads to getting great results with less data.
#
# Over the next few notebooks, we'll see the power of transfer learning in action.
#
# ## What we're going to cover
#
# We're going to go through the following with TensorFlow:
#
# - Introduce transfer learning (a way to beat all of our old self-built models)
# - Using a smaller dataset to experiment faster (10% of training samples of 10 classes of food)
# - Build a transfer learning feature extraction model using TensorFlow Hub
# - Introduce the TensorBoard callback to track model training results
# - Compare model results using TensorBoard
#
# ## How you can use this notebook
#
# You can read through the descriptions and the code (it should all run, except for the cells which error on purpose), but there's a better option.
#
# Write all of the code yourself.
#
# Yes. I'm serious. Create a new notebook, and rewrite each line by yourself. Investigate it, see if you can break it, why does it break?
#
# You don't have to write the text descriptions but writing the code yourself is a great way to get hands-on experience.
#
# Don't worry if you make mistakes, we all do. The way to get better and make less mistakes is to **write more code**.
# + [markdown] id="UTWetPM7AWfY"
# ## Using a GPU
#
# To begin, let's check to see if we're using a GPU. Using a GPU will make sure our model trains faster than using just a CPU.
#
# + id="Nq4kxIpQMpZT" colab={"base_uri": "https://localhost:8080/"} outputId="5a53217a-7674-437c-df90-8e9ae37a54ed"
# Are we using a GPU?
# !nvidia-smi
# + [markdown] id="Ol3NDTVlRLSv"
# If the cell above doesn't output something which looks like:
#
# ```
# Fri Sep 4 03:35:21 2020
# # +-----------------------------------------------------------------------------+
# | NVIDIA-SMI 450.66 Driver Version: 418.67 CUDA Version: 10.1 |
# |-------------------------------+----------------------+----------------------+
# | GPU Name Persistence-M| Bus-Id Disp.A | Volatile Uncorr. ECC |
# | Fan Temp Perf Pwr:Usage/Cap| Memory-Usage | GPU-Util Compute M. |
# | | | MIG M. |
# |===============================+======================+======================|
# | 0 Tesla P100-PCIE... Off | 00000000:00:04.0 Off | 0 |
# | N/A 35C P0 26W / 250W | 0MiB / 16280MiB | 0% Default |
# | | | ERR! |
# # +-------------------------------+----------------------+----------------------+
#
# # +-----------------------------------------------------------------------------+
# | Processes: |
# | GPU GI CI PID Type Process name GPU Memory |
# | ID ID Usage |
# |=============================================================================|
# | No running processes found |
# # +-----------------------------------------------------------------------------+
# ```
#
# Go to Runtime -> Change Runtime Type -> Hardware Accelerator and select "GPU", then rerun the cell above.
# + [markdown] id="7NY09457NKv4"
# ## Transfer leanring with TensorFlow Hub: Getting great results with 10% of the data
#
# If you've been thinking, "surely someone else has spent the time crafting the right model for the job..." then you're in luck.
#
# For many of the problems you'll want to use deep learning for, chances are, a working model already exists.
#
# And the good news is, you can access many of them on TensorFlow Hub.
#
# [TensorFlow Hub](https://tfhub.dev/) is a repository for existing model components. It makes it so you can import and use a fully trained model with as little as a URL.
#
# Now, I really want to demonstrate the power of transfer learning to you.
#
# To do so, what if I told you we could get much of the same results (or better) than our best model has gotten so far with only 10% of the original data, in other words, 10x less data.
#
# This seems counterintuitive right?
#
# Wouldn't you think more examples of what a picture of food looked like led to better results?
#
# And you'd be right if you thought so, generally, more data leads to better results.
#
# However, what if you didn't have more data? What if instead of 750 images per class, you had 75 images per class?
#
# Collecting 675 more images of a certain class could take a long time.
#
# So this is where another major benefit of transfer learning comes in.
#
# **Transfer learning often allows you to get great results with less data.**
#
# But don't just take my word for it. Let's download a subset of the data we've been using, namely 10% of the training data from the `10_food_classes` dataset and use it to train a food image classifier on.
#
# 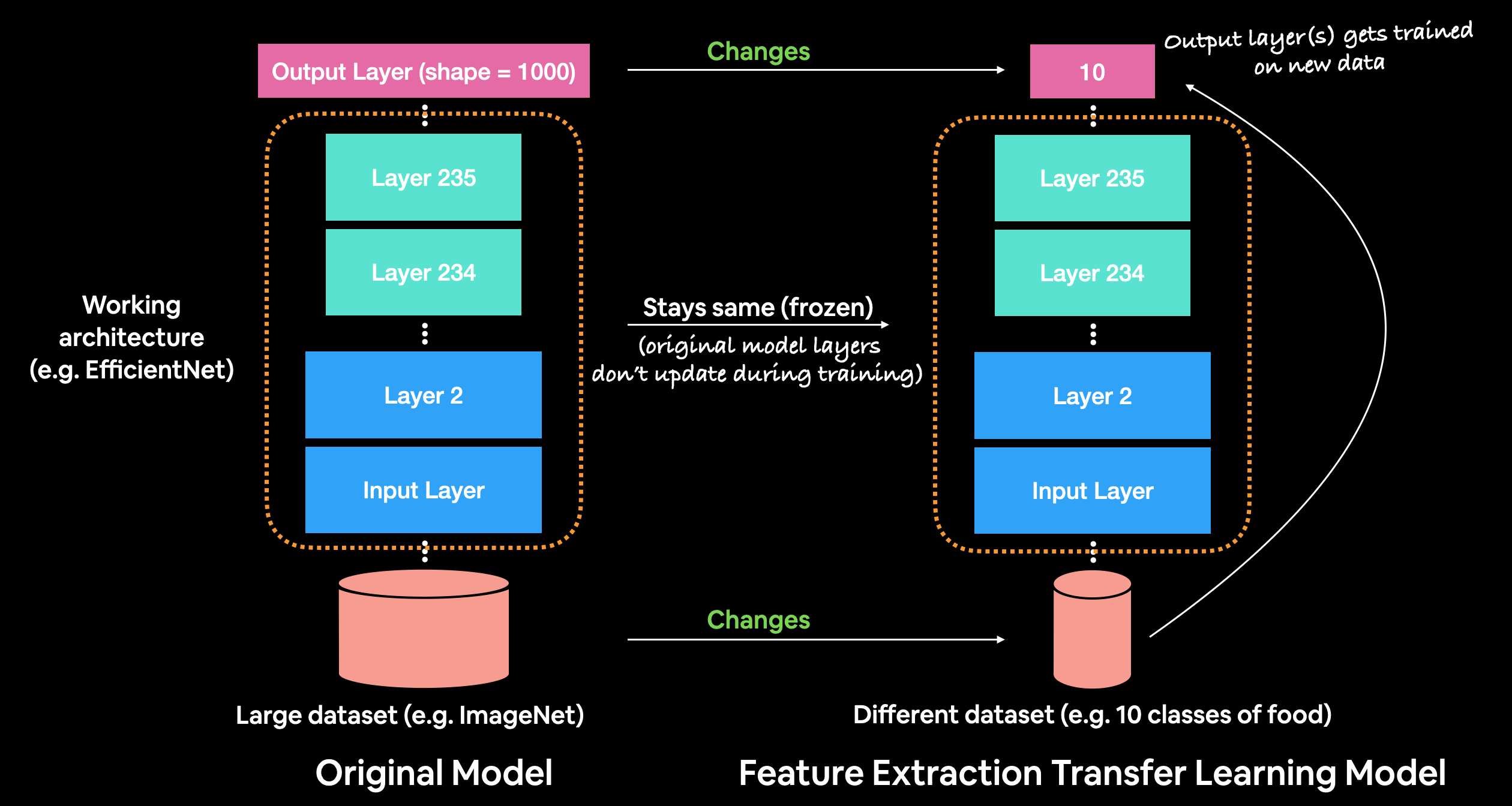
# *What we're working towards building. Taking a pre-trained model and adding our own custom layers on top, extracting all of the underlying patterns learned on another dataset our own images.*
#
#
# + [markdown] id="UIwVrX6vXb4z"
# ## Downloading and becoming one with the data
# + id="qwWwP657Szfv" colab={"base_uri": "https://localhost:8080/"} outputId="46b1d719-f47b-4918-92e8-92a18c1935b7"
# Get data (10% of labels)
import zipfile
# Download data
# !wget https://storage.googleapis.com/ztm_tf_course/food_vision/10_food_classes_10_percent.zip
# Unzip the downloaded file
zip_ref = zipfile.ZipFile("10_food_classes_10_percent.zip", "r")
zip_ref.extractall()
zip_ref.close()
# + id="agzJYtfFBl6I" colab={"base_uri": "https://localhost:8080/"} outputId="f347e60c-72ed-49c6-e706-c8be3439bea4"
# How many images in each folder?
import os
# Walk through 10 percent data directory and list number of files
for dirpath, dirnames, filenames in os.walk("10_food_classes_10_percent"):
print(f"There are {len(dirnames)} directories and {len(filenames)} images in '{dirpath}'.")
# + [markdown] id="F0r-zyagV7Qa"
# Notice how each of the training directories now has 75 images rather than 750 images. This is key to demonstrating how well transfer learning can perform with less labelled images.
#
# The test directories still have the same amount of images. This means we'll be training on less data but evaluating our models on the same amount of test data.
# + [markdown] id="EES-NoeaXfYT"
# ## Creating data loaders (preparing the data)
#
# Now we've downloaded the data, let's use the [`ImageDataGenerator`](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator) class along with the `flow_from_directory` method to load in our images.
# + id="YAp0GN60S-rK" colab={"base_uri": "https://localhost:8080/"} outputId="bb4a2dc3-fad9-416b-b81a-b2a0d3abcb45"
# Setup data inputs
from tensorflow.keras.preprocessing.image import ImageDataGenerator
IMAGE_SHAPE = (224, 224)
BATCH_SIZE = 32
train_dir = "10_food_classes_10_percent/train/"
test_dir = "10_food_classes_10_percent/test/"
train_datagen = ImageDataGenerator(rescale=1/255.)
test_datagen = ImageDataGenerator(rescale=1/255.)
print("Training images:")
train_data_10_percent = train_datagen.flow_from_directory(train_dir,
target_size=IMAGE_SHAPE,
batch_size=BATCH_SIZE,
class_mode="categorical")
print("Testing images:")
test_data = train_datagen.flow_from_directory(test_dir,
target_size=IMAGE_SHAPE,
batch_size=BATCH_SIZE,
class_mode="categorical")
# + [markdown] id="6QWuVeSvQPoK"
# Excellent! Loading in the data we can see we've got 750 images in the training dataset belonging to 10 classes (75 per class) and 2500 images in the test set belonging to 10 classes (250 per class).
# + [markdown] id="6Qcwii2uYjOx"
# ## Setting up callbacks (things to run whilst our model trains)
#
# Before we build a model, there's an important concept we're going to get familiar with because it's going to play a key role in our future model building experiments.
#
# And that concept is **callbacks**.
#
# [Callbacks](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks) are extra functionality you can add to your models to be performed during or after training. Some of the most popular callbacks include:
# * [**Experiment tracking with TensorBoard**](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) - log the performance of multiple models and then view and compare these models in a visual way on [TensorBoard](https://www.tensorflow.org/tensorboard) (a dashboard for inspecting neural network parameters). Helpful to compare the results of different models on your data.
# * [**Model checkpointing**](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/ModelCheckpoint) - save your model as it trains so you can stop training if needed and come back to continue off where you left. Helpful if training takes a long time and can't be done in one sitting.
# * [**Early stopping**](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/EarlyStopping) - leave your model training for an arbitrary amount of time and have it stop training automatically when it ceases to improve. Helpful when you've got a large dataset and don't know how long training will take.
#
# We'll explore each of these overtime but for this notebook, we'll see how the TensorBoard callback can be used.
#
# The TensorBoard callback can be accessed using [`tf.keras.callbacks.TensorBoard()`](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard).
#
# Its main functionality is saving a model's training performance metrics to a specified `log_dir`.
#
# By default, logs are recorded every epoch using the `update_freq='epoch'` parameter. This is a good default since tracking model performance too often can slow down model training.
#
# To track our modelling experiments using TensorBoard, let's create a function which creates a TensorBoard callback for us.
#
# > 🔑 **Note:** We create a function for creating a TensorBoard callback because as we'll see later on, each model needs its own TensorBoard callback instance (so the function will create a new one each time it's run).
#
#
# + id="2yamhJ8xJA5x"
# Create tensorboard callback (functionized because need to create a new one for each model)
import datetime
def create_tensorboard_callback(dir_name, experiment_name):
log_dir = dir_name + "/" + experiment_name + "/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(
log_dir=log_dir
)
print(f"Saving TensorBoard log files to: {log_dir}")
return tensorboard_callback
# + [markdown] id="11TjBJQXdCyZ"
# Because you're likely to run multiple experiments, it's a good idea to be able to track them in some way.
#
# In our case, our function saves a model's performance logs to a directory named `[dir_name]/[experiment_name]/[current_timestamp]`, where:
# * `dir_name` is the overall logs directory
# * `experiment_name` is the particular experiment
# * `current_timestamp` is the time the experiment started based on Python's [`datetime.datetime().now()`](https://docs.python.org/3/library/datetime.html#datetime.datetime.now)
#
# > 🔑 **Note:** Depending on your use case, the above experimenting tracking naming method may work or you might require something more specific. The good news is, the TensorBoard callback makes it easy to track modelling logs as long as you specify where to track them. So you can get as creative as you like with how you name your experiments, just make sure you or your team can understand them.
#
#
# + [markdown] id="8UP8vA_nYoI7"
# ## Creating models using TensorFlow Hub
#
# In the past we've used TensorFlow to create our own models layer by layer from scratch.
#
# Now we're going to do a similar process, except the majority of our model's layers are going to come from [TensorFlow Hub](https://tfhub.dev/).
#
# In fact, we're going to use two models from TensorFlow Hub:
# 1. [ResNetV2](https://arxiv.org/abs/1603.05027) - a state of the art computer vision model architecture from 2016.
# 2. [EfficientNet](https://arxiv.org/abs/1905.11946) - a state of the art computer vision architecture from 2019.
#
# State of the art means that at some point, both of these models have achieved the lowest error rate on [ImageNet (ILSVRC-2012-CLS)](http://www.image-net.org/), the gold standard of computer vision benchmarks.
#
# You might be wondering, how do you find these models on TensorFlow Hub?
#
# Here are the steps I took:
#
# 1. Go to [tfhub.dev](https://tfhub.dev/).
# 2. Choose your problem domain, e.g. "Image" (we're using food images).
# 3. Select your TF version, which in our case is TF2.
# 4. Remove all "Problem domanin" filters except for the problem you're working on.
# * **Note:** "Image feature vector" can be used alongside almost any problem, we'll get to this soon.
# 5. The models listed are all models which could potentially be used for your problem.
#
# > 🤔 **Question:** *I see many options for image classification models, how do I know which is best?*
#
# You can see a list of state of the art models on [paperswithcode.com](https://www.paperswithcode.com), a resource for collecting the latest in deep learning paper results which have code implementations for the findings they report.
#
# Since we're working with images, our target are the [models which perform best on ImageNet](https://paperswithcode.com/sota/image-classification-on-imagenet).
#
# You'll probably find not all of the model architectures listed on paperswithcode appear on TensorFlow Hub. And this is okay, we can still use what's available.
#
# To find our models, let's narrow down our search using the Architecture tab.
#
# 6. Select the Architecture tab on TensorFlow Hub and you'll see a dropdown menu of architecture names appear.
# * The rule of thumb here is generally, names with larger numbers means better performing models. For example, EfficientNetB4 performs better than EfficientNetB0.
# * However, the tradeoff with larger numbers can mean they take longer to compute.
# 7. Select EfficientNetB0 and you should see [something like the following](https://tfhub.dev/s?module-type=image-classification,image-feature-vector&network-architecture=efficientnet-b0&tf-version=tf2):
# 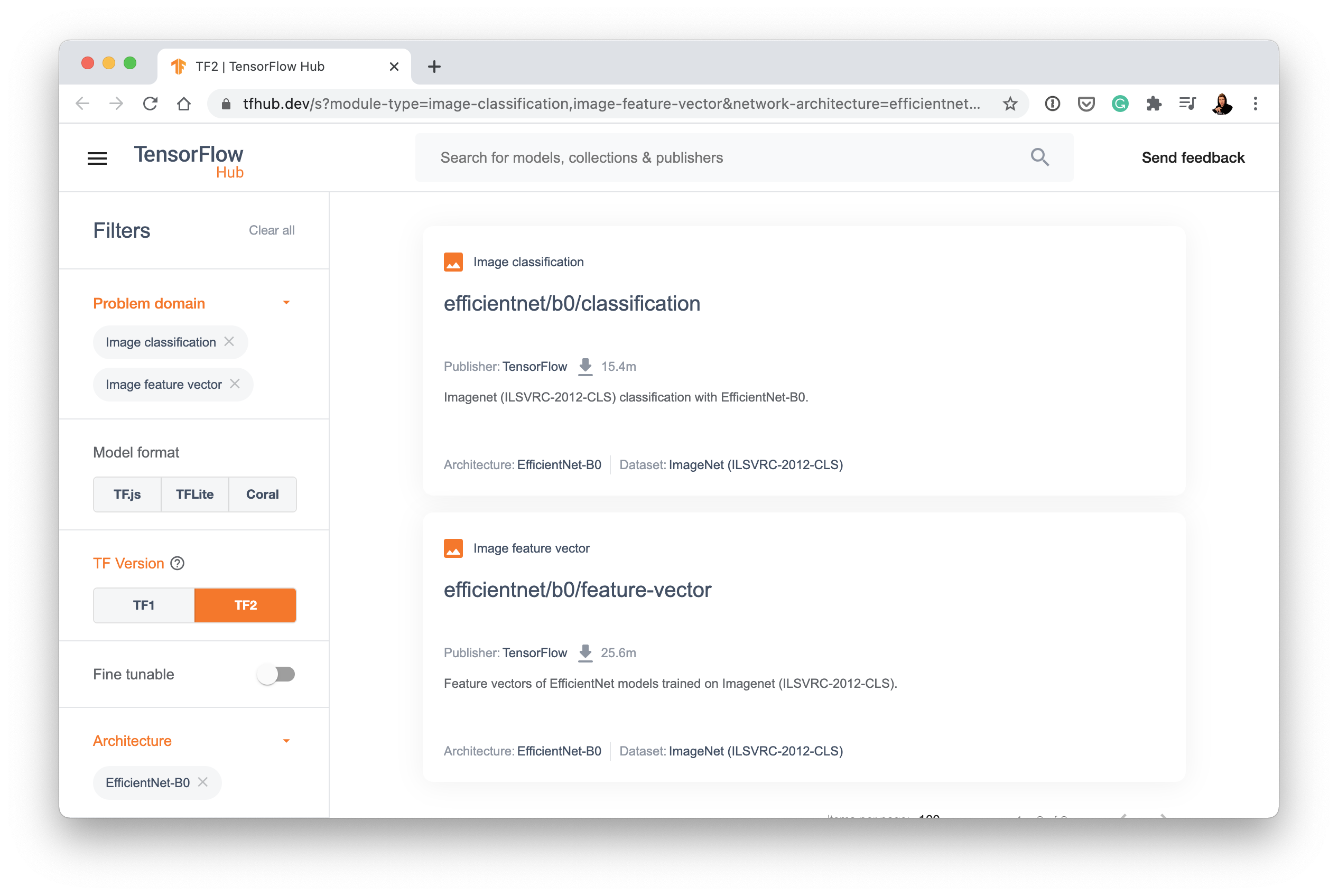
# 8. Clicking the one titled "[efficientnet/b0/feature-vector](https://tfhub.dev/tensorflow/efficientnet/b0/feature-vector/1)" brings us to a page with a button that says "Copy URL". That URL is what we can use to harness the power of EfficientNetB0.
# * Copying the URL should give you something like this: https://tfhub.dev/tensorflow/efficientnet/b0/feature-vector/1
#
# > 🤔 **Question:** *I thought we were doing image classification, why do we choose feature vector and not classification?*
#
# Great observation. This is where the differnet types of transfer learning come into play, as is, feature extraction and fine-tuning.
#
# 1. **"As is" transfer learning** is when you take a pretrained model as it is and apply it to your task without any changes.
#
# * For example, many computer vision models are pretrained on the ImageNet dataset which contains 1000 different classes of images. This means passing a single image to this model will produce 1000 different prediction probability values (1 for each class).
#
# * This is helpful if you have 1000 classes of image you'd like to classify and they're all the same as the ImageNet classes, however, it's not helpful if you want to classify only a small subset of classes (such as 10 different kinds of food). Model's with `"/classification"` in their name on TensorFlow Hub provide this kind of functionality.
#
# 2. **Feature extraction transfer learning** is when you take the underlying patterns (also called weights) a pretrained model has learned and adjust its outputs to be more suited to your problem.
#
# * For example, say the pretrained model you were using had 236 different layers (EfficientNetB0 has 236 layers), but the top layer outputs 1000 classes because it was pretrained on ImageNet. To adjust this to your own problem, you might remove the original activation layer and replace it with your own but with the right number of output classes. The important part here is that **only the top few layers become trainable, the rest remain frozen**.
#
# * This way all the underlying patterns remain in the rest of the layers and you can utilise them for your own problem. This kind of transfer learning is very helpful when your data is similar to the data a model has been pretrained on.
#
# 3. **Fine-tuning transfer learning** is when you take the underlying patterns (also called weights) of a pretrained model and adjust (fine-tune) them to your own problem.
#
# * This usually means training **some, many or all** of the layers in the pretrained model. This is useful when you've got a large dataset (e.g. 100+ images per class) where your data is slightly different to the data the original model was trained on.
#
# A common workflow is to "freeze" all of the learned patterns in the bottom layers of a pretrained model so they're untrainable. And then train the top 2-3 layers of so the pretrained model can adjust its outputs to your custom data (**feature extraction**).
#
# After you've trained the top 2-3 layers, you can then gradually "unfreeze" more and more layers and run the training process on your own data to further **fine-tune** the pretrained model.
#
# > 🤔 **Question:** *Why train only the top 2-3 layers in feature extraction?*
#
# The lower a layer is in a computer vision model as in, the closer it is to the input layer, the larger the features it learn. For example, a bottom layer in a computer vision model to identify images of cats or dogs might learn the outline of legs, where as, layers closer to the output might learn the shape of teeth. Often, you'll want the larger features (learned patterns are also called features) to remain, since these are similar for both animals, where as, the differences remain in the more fine-grained features.
#
# 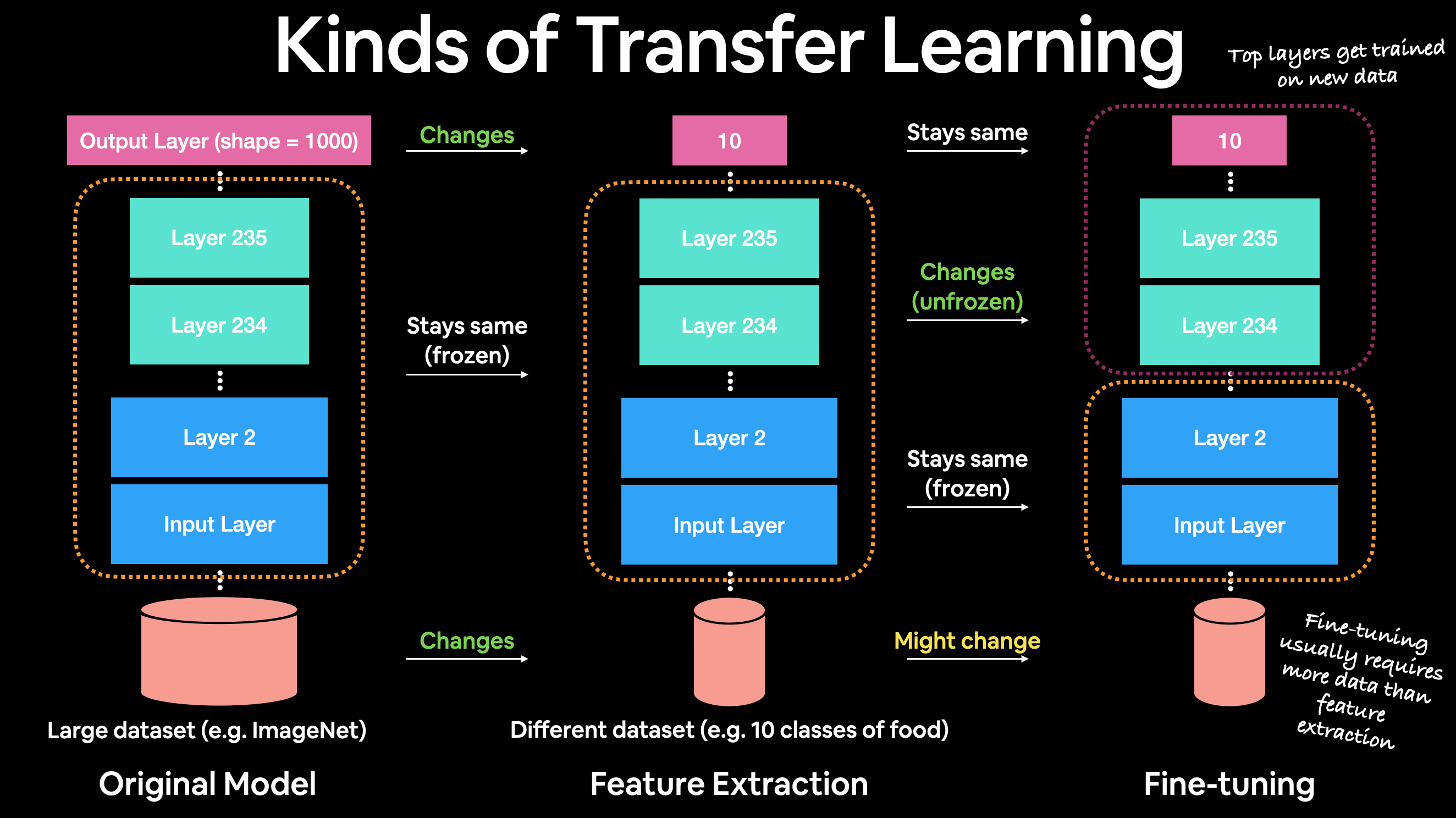
# *The different kinds of transfer learning. An original model, a feature extraction model (only top 2-3 layers change) and a fine-tuning model (many or all of original model get changed).*
#
# Okay, enough talk, let's see this in action. Once we do, we'll explain what's happening.
#
# First we'll import TensorFlow and TensorFlow Hub.
# + id="xsoE9nUJNN6s"
import tensorflow as tf
import tensorflow_hub as hub
from tensorflow.keras import layers
# + [markdown] id="nvGge7Xevt_F"
# Now we'll get the feature vector URLs of two common computer vision architectures, [EfficientNetB0 (2019)](https://tfhub.dev/tensorflow/efficientnet/b0/feature-vector/1) and [ResNetV250 (2016)](https://tfhub.dev/google/imagenet/resnet_v2_50/feature_vector/4) from TensorFlow Hub using the steps above.
#
# We're getting both of these because we're going to compare them to see which performs better on our data.
#
# > 🔑 **Note:** Comparing different model architecture performance on the same data is a very common practice. The simple reason is because you want to know which model performs best for your problem.
#
# > **Update:** As of 14 August 2021, [EfficientNet V2 pretrained models are available on TensorFlow Hub](https://tfhub.dev/google/collections/efficientnet_v2/1). The original code in this notebook uses EfficientNet V1, it has been left unchanged. In [my experiments with this dataset](https://github.com/mrdbourke/tensorflow-deep-learning/discussions/166), V1 outperforms V2. Best to experiment with your own data and see what suits you.
# + id="LZfUivHxOCbP"
# Resnet 50 V2 feature vector
resnet_url = "https://tfhub.dev/google/imagenet/resnet_v2_50/feature_vector/4"
# Original: EfficientNetB0 feature vector (version 1)
efficientnet_url = "https://tfhub.dev/tensorflow/efficientnet/b0/feature-vector/1"
# # New: EfficientNetB0 feature vector (version 2)
# efficientnet_url = "https://tfhub.dev/google/imagenet/efficientnet_v2_imagenet1k_b0/feature_vector/2"
# + [markdown] id="bdwjFaCRwdCX"
# These URLs link to a saved pretrained model on TensorFlow Hub.
#
# When we use them in our model, the model will automatically be downloaded for us to use.
#
# To do this, we can use the [`KerasLayer()`](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer) model inside the TensorFlow hub library.
#
# Since we're going to be comparing two models, to save ourselves code, we'll create a function `create_model()`. This function will take a model's TensorFlow Hub URL, instatiate a Keras Sequential model with the appropriate number of output layers and return the model.
# + id="p7vXoqSjId0f"
def create_model(model_url, num_classes=10):
"""Takes a TensorFlow Hub URL and creates a Keras Sequential model with it.
Args:
model_url (str): A TensorFlow Hub feature extraction URL.
num_classes (int): Number of output neurons in output layer,
should be equal to number of target classes, default 10.
Returns:
An uncompiled Keras Sequential model with model_url as feature
extractor layer and Dense output layer with num_classes outputs.
"""
# Download the pretrained model and save it as a Keras layer
feature_extractor_layer = hub.KerasLayer(model_url,
trainable=False, # freeze the underlying patterns
name='feature_extraction_layer',
input_shape=IMAGE_SHAPE+(3,)) # define the input image shape
# Create our own model
model = tf.keras.Sequential([
feature_extractor_layer, # use the feature extraction layer as the base
layers.Dense(num_classes, activation='softmax', name='output_layer') # create our own output layer
])
return model
# + [markdown] id="IirF2Ohlz-6i"
# Great! Now we've got a function for creating a model, we'll use it to first create a model using the ResNetV250 architecture as our feature extraction layer.
#
# Once the model is instantiated, we'll compile it using `categorical_crossentropy` as our loss function, the Adam optimizer and accuracy as our metric.
# + id="-KVRwwbDT-HL"
# Create model
resnet_model = create_model(resnet_url, num_classes=train_data_10_percent.num_classes)
# Compile
resnet_model.compile(loss='categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
# + [markdown] id="ZinVcxBi0jsv"
# 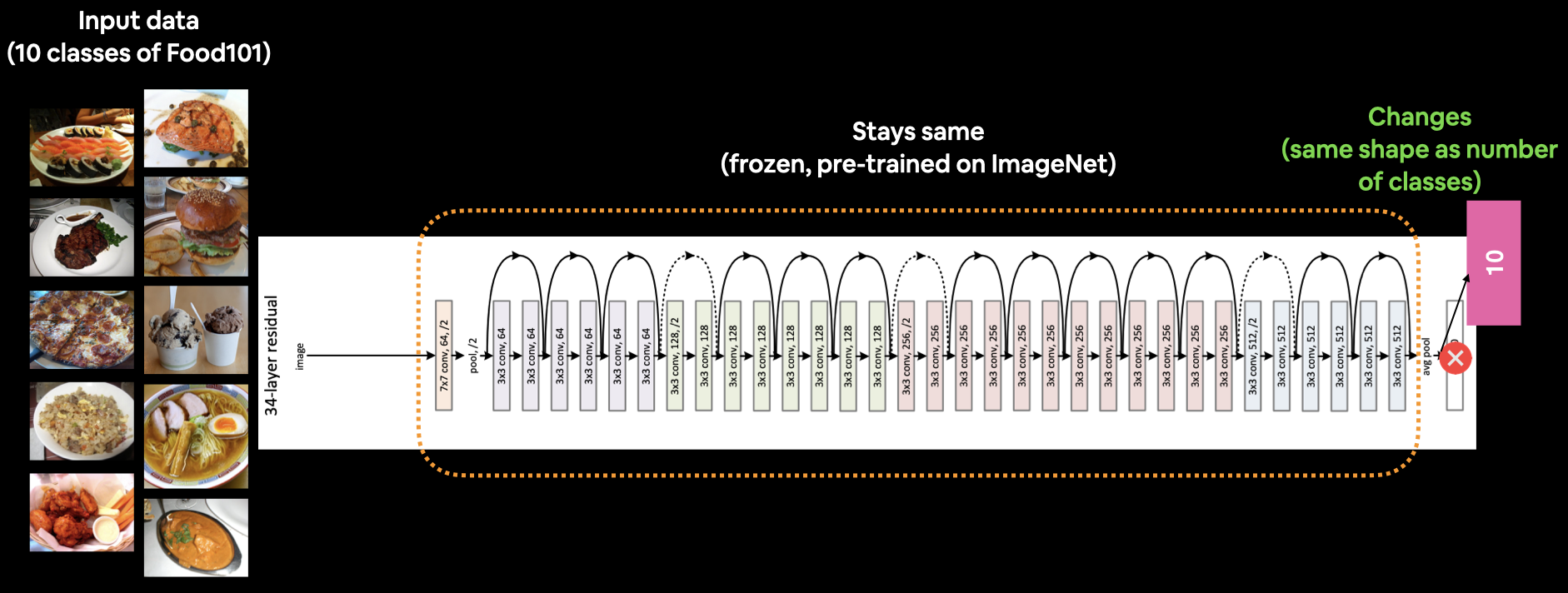
# *What our current model looks like. A ResNet50V2 backbone with a custom dense layer on top (10 classes instead of 1000 ImageNet classes). **Note:** The Image shows ResNet34 instead of ResNet50. **Image source:** https://arxiv.org/abs/1512.03385.*
#
# Beautiful. Time to fit the model.
#
# We've got the training data ready in `train_data_10_percent` as well as the test data saved as `test_data`.
#
# But before we call the fit function, there's one more thing we're going to add, a callback. More specifically, a TensorBoard callback so we can track the performance of our model on TensorBoard.
#
# We can add a callback to our model by using the `callbacks` parameter in the fit function.
#
# In our case, we'll pass the `callbacks` parameter the `create_tensorboard_callback()` we created earlier with some specific inputs so we know what experiments we're running.
#
# Let's keep this experiment short and train for 5 epochs.
# + id="2GTl0fwE0Hx6" colab={"base_uri": "https://localhost:8080/"} outputId="4cf373f4-7143-45b0-ddfc-d1cb009e555a"
# Fit the model
resnet_history = resnet_model.fit(train_data_10_percent,
epochs=5,
steps_per_epoch=len(train_data_10_percent),
validation_data=test_data,
validation_steps=len(test_data),
# Add TensorBoard callback to model (callbacks parameter takes a list)
callbacks=[create_tensorboard_callback(dir_name="tensorflow_hub", # save experiment logs here
experiment_name="resnet50V2")]) # name of log files
# + [markdown] id="i5SuOe672UJi"
# Wow!
#
# It seems that after only 5 epochs, the ResNetV250 feature extraction model was able to blow any of the architectures we made out of the water, achieving around 90% accuracy on the training set and nearly 80% accuracy on the test set...**with only 10 percent of the training images!**
#
# That goes to show the power of transfer learning. And it's one of the main reasons whenever you're trying to model your own datasets, you should look into what pretrained models already exist.
#
# Let's check out our model's training curves using our `plot_loss_curves` function.
# + id="Ot2QPj41ODCQ"
# If you wanted to, you could really turn this into a helper function to load in with a helper.py script...
import matplotlib.pyplot as plt
# Plot the validation and training data separately
def plot_loss_curves(history):
"""
Returns separate loss curves for training and validation metrics.
"""
loss = history.history['loss']
val_loss = history.history['val_loss']
accuracy = history.history['accuracy']
val_accuracy = history.history['val_accuracy']
epochs = range(len(history.history['loss']))
# Plot loss
plt.plot(epochs, loss, label='training_loss')
plt.plot(epochs, val_loss, label='val_loss')
plt.title('Loss')
plt.xlabel('Epochs')
plt.legend()
# Plot accuracy
plt.figure()
plt.plot(epochs, accuracy, label='training_accuracy')
plt.plot(epochs, val_accuracy, label='val_accuracy')
plt.title('Accuracy')
plt.xlabel('Epochs')
plt.legend();
# + id="ywQ9Wr9UODJ_" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="a225ebca-4308-4175-95e9-849c4bd21ab4"
plot_loss_curves(resnet_history)
# + [markdown] id="5BGVFEIi3_CT"
# And what about a summary of our model?
# + id="aps1FV4qWrZb" colab={"base_uri": "https://localhost:8080/"} outputId="306849a0-a3a2-4404-d046-eceb7186871f"
# Resnet summary
resnet_model.summary()
# + [markdown] id="okdbmBA0SgCb"
# You can see the power of TensorFlow Hub here. The feature extraction layer has 23,564,800 parameters which are prelearned patterns the model has already learned on the ImageNet dataset. Since we set `trainable=False`, these patterns remain frozen (non-trainable) during training.
#
# This means during training the model updates the 20,490 parameters in the output layer to suit our dataset.
#
# Okay, we've trained a ResNetV250 model, time to do the same with EfficientNetB0 model.
#
# The setup will be the exact same as before, except for the `model_url` parameter in the `create_model()` function and the `experiment_name` parameter in the `create_tensorboard_callback()` function.
# + id="MrGi-CpMXHav" colab={"base_uri": "https://localhost:8080/"} outputId="251ab535-7852-4e05-c356-dba75c37ba34"
# Create model
efficientnet_model = create_model(model_url=efficientnet_url, # use EfficientNetB0 TensorFlow Hub URL
num_classes=train_data_10_percent.num_classes)
# Compile EfficientNet model
efficientnet_model.compile(loss='categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
# Fit EfficientNet model
efficientnet_history = efficientnet_model.fit(train_data_10_percent, # only use 10% of training data
epochs=5, # train for 5 epochs
steps_per_epoch=len(train_data_10_percent),
validation_data=test_data,
validation_steps=len(test_data),
callbacks=[create_tensorboard_callback(dir_name="tensorflow_hub",
# Track logs under different experiment name
experiment_name="efficientnetB0")])
# + [markdown] id="iMbtls4C57Dr"
# Holy smokes! The EfficientNetB0 model does even better than the ResNetV250 model! Achieving over 85% accuracy on the test set...again **with only 10% of the training data**.
#
# How cool is that?
#
# With a couple of lines of code we're able to leverage state of the art models and adjust them to our own use case.
#
# Let's check out the loss curves.
# + id="8UzgNYFdODRB" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="24ffc097-9536-4ff7-f739-51bf2cf883a8"
plot_loss_curves(efficientnet_history)
# + [markdown] id="pDi4i0H16tSa"
# From the look of the EfficientNetB0 model's loss curves, it looks like if we kept training our model for longer, it might improve even further. Perhaps that's something you might want to try?
#
# Let's check out the model summary.
# + id="clJzUbKpODXA" colab={"base_uri": "https://localhost:8080/"} outputId="3538504b-1cc9-4569-db9e-b191ea37d20f"
efficientnet_model.summary()
# + [markdown] id="KHTMjJG07ElO"
# It seems despite having over four times less parameters (4,049,564 vs. 23,564,800) than the ResNet50V2 extraction layer, the EfficientNetB0 feature extraction layer yields better performance. Now it's clear where the "efficient" name came from.
# + [markdown] id="YV_ZWKC8SkE_"
# ## Comparing models using TensorBoard
#
# Alright, even though we've already compared the performance of our two models by looking at the accuracy scores. But what if you had more than two models?
#
# That's where an experiment tracking tool like [TensorBoard](https://www.tensorflow.org/tensorboard) (preinstalled in Google Colab) comes in.
#
# The good thing is, since we set up a TensorBoard callback, all of our model's training logs have been saved automatically. To visualize them, we can upload the results to [TensorBoard.dev](https://tensorboard.dev/).
#
# Uploading your results to TensorBoard.dev enables you to track and share multiple different modelling experiments. So if you needed to show someone your results, you could send them a link to your TensorBoard.dev as well as the accompanying Colab notebook.
#
# > 🔑 **Note:** These experiments are public, do not upload sensitive data. You can delete experiments if needed.
#
# ### Uploading experiments to TensorBoard
#
# To upload a series of TensorFlow logs to TensorBoard, we can use the following command:
#
# ```
# Upload TensorBoard dev records
#
# # # !tensorboard dev upload --logdir ./tensorflow_hub/ \
# # # --name "EfficientNetB0 vs. ResNet50V2" \
# # # --description "Comparing two different TF Hub feature extraction models architectures using 10% of training images" \
# # # --one_shot
# ```
#
# Where:
# * `--logdir` is the target upload directory
# * `--name` is the name of the experiment
# * `--description` is a brief description of the experiment
# * `--one_shot` exits the TensorBoard uploader once uploading is finished
#
# Running the `tensorboard dev upload` command will first ask you to authorize the upload to TensorBoard.dev. After you've authorized the upload, your log files will be uploaded.
# + id="tbKgWdIVNncW" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="e344cf7c-7064-484d-f31b-4079735dc921"
# Upload TensorBoard dev records
# !tensorboard dev upload --logdir ./tensorflow_hub/ \
# --name "EfficientNetB0 vs. ResNet50V2" \
# --description "Comparing two different TF Hub feature extraction models architectures using 10% of training images" \
# --one_shot
# + [markdown] id="FlVfmBdBOPvf"
# Every time you upload something to TensorBoad.dev you'll get a new experiment ID. The experiment ID will look something like this: https://tensorboard.dev/experiment/73taSKxXQeGPQsNBcVvY3g/ (this is the actual experiment from this notebook).
#
# If you upload the same directory again, you'll get a new experiment ID to go along with it.
#
# This means to track your experiments, you may want to look into how you name your uploads. That way when you find them on TensorBoard.dev you can tell what happened during each experiment (e.g. "efficientnet0_10_percent_data").
#
# ### Listing experiments you've saved to TensorBoard
#
# To see all of the experiments you've uploaded you can use the command:
#
# ```tensorboard dev list```
# + id="sDamroaMOFJx" colab={"base_uri": "https://localhost:8080/", "height": 496} outputId="d086e0da-e091-4504-87d6-a50aa4636577"
# Check out experiments
# !tensorboard dev list
# + [markdown] id="mdLUjm-xADQ4"
# ### Deleting experiments from TensorBoard
#
# Remember, all uploads to TensorBoard.dev are public, so to delete an experiment you can use the command:
#
# `tensorboard dev delete --experiment_id [INSERT_EXPERIMENT_ID]`
#
#
# + id="qj69wuAlT-xS" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="fcc4a104-85eb-4f66-aa43-8bc25b75a51a"
# Delete an experiment
# !tensorboard dev delete --experiment_id n6kd8XZ3Rdy1jSgSLH5WjA
# + id="Oov6qKvbU1lL" colab={"base_uri": "https://localhost:8080/", "height": 309} outputId="9d2997d3-7278-4290-e8b8-f4fb2c9aab76"
# Check to see if experiments still exist
# !tensorboard dev list
# + [markdown] id="KLvUjrL2Y1Ed"
# ## 🛠 Exercises
#
# 1. Build and fit a model using the same data we have here but with the MobileNetV2 architecture feature extraction ([`mobilenet_v2_100_224/feature_vector`](https://tfhub.dev/google/imagenet/mobilenet_v2_100_224/feature_vector/4)) from TensorFlow Hub, how does it perform compared to our other models?
# 2. Name 3 different image classification models on TensorFlow Hub that we haven't used.
# 3. Build a model to classify images of two different things you've taken photos of.
# * You can use any feature extraction layer from TensorFlow Hub you like for this.
# * You should aim to have at least 10 images of each class, for example to build a fridge versus oven classifier, you'll want 10 images of fridges and 10 images of ovens.
# 4. What is the current best performing model on ImageNet?
# * Hint: you might want to check [sotabench.com](https://www.sotabench.com) for this.
# + [markdown] id="w_YxwuhfRzD5"
# ## 📖 Extra-curriculum
#
# * Read through the [TensorFlow Transfer Learning Guide](https://www.tensorflow.org/tutorials/images/transfer_learning) and define the main two types of transfer learning in your own words.
# * Go through the [Transfer Learning with TensorFlow Hub tutorial](https://www.tensorflow.org/tutorials/images/transfer_learning_with_hub) on the TensorFlow website and rewrite all of the code yourself into a new Google Colab notebook making comments about what each step does along the way.
# * We haven't covered fine-tuning with TensorFlow Hub in this notebook, but if you'd like to know more, go through the [fine-tuning a TensorFlow Hub model tutorial](https://www.tensorflow.org/hub/tf2_saved_model#fine-tuning) on the TensorFlow homepage.How to fine-tune a tensorflow hub model:
# * Look into [experiment tracking with Weights & Biases](https://www.wandb.com/experiment-tracking), how could you integrate it with our existing TensorBoard logs?
| 04_transfer_learning_in_tensorflow_part_1_feature_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **[Time Series With Siraj Course Home Page](https://kaggle.com/learn/time-series-with-siraj)**
#
# ---
#
# + _uuid="3144c4e3acf6570cccdee57ec7205ad6140814bd"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from datetime import datetime
from pandas import Series
import warnings
warnings.filterwarnings("ignore")
plt.style.use('fivethirtyeight')
# + _uuid="c4b3c73366a1fdfde6860bb3cd93ffe65ee6dc25"
train = pd.read_csv("../input/siraj2/Train_SU63ISt.csv")
test = pd.read_csv("../input/siraj2/Test_0qrQsBZ.csv")
train_original = train.copy()
test_original = test.copy()
# + [markdown] _uuid="5f90fbfaea3ef1777354018241e084e2a2a957c3"
# # Extracting Features
# + _uuid="4b04f41f3fd4669ea32f2457a104b09c982cf39a"
train['Datetime'] = pd.to_datetime(train.Datetime, format = '%d-%m-%Y %H:%M')
test['Datetime'] = pd.to_datetime(test.Datetime, format = '%d-%m-%Y %H:%M')
train_original['Datetime'] = pd.to_datetime(train_original.Datetime, format = '%d-%m-%Y %H:%M')
test_original['Datetime'] = pd.to_datetime(test_original.Datetime, format = '%d-%m-%Y %H:%M')
for i in (train, test, train_original, test_original):
i['year'] = i.Datetime.dt.year
i['month'] = i.Datetime.dt.month
i['day']= i.Datetime.dt.day
i['Hour']=i.Datetime.dt.hour
train['Day of week'] = train['Datetime'].dt.dayofweek
temp = train['Datetime']
def applyer(row):
if row.dayofweek == 5 or row.dayofweek == 6:
return 1
else:
return 0
temp2 = train['Datetime'].apply(applyer)
train['weekend'] = temp2
train.index = train['Datetime']
df = train.drop('ID',1)
ts = df['Count']
plt.figure(figsize = (16,8))
plt.plot(ts)
plt.title("Time Series")
plt.xlabel("Time (year-month)")
plt.ylabel("Passenger Count")
plt.legend(loc = 'best')
test.Timestamp = pd.to_datetime(test.Datetime, format='%d-%m-%Y %H:%M')
test.index = test.Timestamp
#Converting to Daily mean
test = test.resample('D').mean()
train.Timestamp = pd.to_datetime(train.Datetime, format='%d-%m-%Y %H:%M')
train.index = train.Timestamp
#Converting to Daily mean
train = train.resample('D').mean()
Train = train.ix['2012-08-25':'2014-06-24']
valid = train.ix['2014-06-25':'2014-09-25']
# + [markdown] _uuid="b83aaa27665d43bf56947c74e02fa618622f4ee5"
# # Divide data into training and validation
# + _uuid="8cf94d181df8bc29de7b03d1ee0e12d02f91b384"
Train.Count.plot(figsize = (15,8), title = 'Daily Ridership', fontsize = 14, label = 'Train')
valid.Count.plot(figsize = (15,8), title = 'Daily Ridership', fontsize =14, label = 'Valid')
plt.xlabel('Datetime')
plt.ylabel('Passenger Count')
plt.legend(loc = 'best')
# + [markdown] _uuid="96fd3baf36e9c893d4eca4657f12958f4511cbdf"
# # Predicting Timeseries: Naive Approach
#
# In this approach you're simply using the last data point to make a prediction. Try modifying `y_hat['naive']` and see what happens. You could also try making it more complex or maybe even more Naive.
# + _uuid="284e39826fe7da226aadbc4ff974d04bb9533318"
dd = np.asarray(Train.Count)
y_hat =valid.copy()
y_hat['naive']= dd[len(dd)- 1]
plt.figure(figsize = (12,8))
plt.plot(Train.index, Train['Count'],label = 'Train')
plt.plot(valid.index, valid['Count'], label = 'Validation')
plt.plot(y_hat.index, y_hat['naive'], label = 'Naive')
plt.legend(loc = 'best')
plt.title('Naive Forecast')
# + [markdown] _uuid="9dd07c502432ca73e0d18d09a497d266a3eb434c"
# ### Calculate the RMSE for a Naive Approach
# + _uuid="0a4a0fb6e10e58e4402c466a3d253d44c7da93e4"
from sklearn.metrics import mean_squared_error
from math import sqrt
rmse = sqrt(mean_squared_error(valid.Count, y_hat.naive))
rmse
# + [markdown] _uuid="3a13ba71186bf20d8d619fc028ee0f25213b1034"
# In this section, try increasgin or decreasing the size of the rolling average widow to see how it affects your Naive predictions.
# + _uuid="5a4084d95ec0e5229b43fa0e926ca42dd8b070a5"
y_hat_avg = valid.copy()
y_hat_avg['moving_average_forecast'] = Train['Count'].rolling(10).mean().iloc[-1]
plt.figure(figsize = (15,5))
plt.plot(Train['Count'], label = 'Train')
plt.plot(valid['Count'], label = 'Validation')
plt.plot(y_hat_avg['moving_average_forecast'], label = 'Moving Average Forecast with 10 Observations')
plt.legend(loc = 'best')
plt.show()
y_hat_avg = valid.copy()
y_hat_avg['moving_average_forecast'] = Train['Count'].rolling(20).mean().iloc[-1]
plt.figure(figsize = (15,5))
plt.plot(Train['Count'], label = 'Train')
plt.plot(valid['Count'], label = 'Validation')
plt.plot(y_hat_avg['moving_average_forecast'],label = 'Moving Average Forecast with 20 Observations')
plt.legend(loc = 'best')
plt.show()
y_hat_avg = valid.copy()
y_hat_avg['moving_average_forecast']= Train['Count'].rolling(50).mean().iloc[-1]
plt.figure(figsize = (15,5))
plt.plot(Train['Count'], label = 'Train')
plt.plot(valid['Count'], label = 'Validation')
plt.plot(y_hat_avg['moving_average_forecast'], label = "Moving Average Forecast with 50 Observations")
plt.legend(loc = 'best')
plt.show()
# + [markdown] _uuid="5a3d08c495e53b2c760a6818c2a8f68c38374063"
# ### Calculate RMSE for a moving average
# + _uuid="826687f45fe3613552454b8c6fcca0f4f1c800ac"
rmse = sqrt(mean_squared_error(valid['Count'], y_hat_avg['moving_average_forecast']))
rmse
# + [markdown] _uuid="55c436a2ff8f8bdc03fcd7a5c9b8c526bb1a2e49"
# # Predicting Timeseries: Simple Exponential Smoothing
#
# Simple Exponential Smoothing is another fun method. Try out different for `smoothing_level` and `optimized`
# + _uuid="5802001e255f13be3c84bfac5b289cc4de7f9782"
from statsmodels.tsa.api import ExponentialSmoothing,SimpleExpSmoothing, Holt
y_hat = valid.copy()
fit2 = SimpleExpSmoothing(np.asarray(Train['Count'])).fit(smoothing_level = 0.6,optimized = False)
y_hat['SES'] = fit2.forecast(len(valid))
plt.figure(figsize =(15,8))
plt.plot(Train['Count'], label = 'Train')
plt.plot(valid['Count'], label = 'Validation')
plt.plot(y_hat['SES'], label = 'Simple Exponential Smoothing')
plt.legend(loc = 'best')
# + [markdown] _uuid="45214d58cead3a838a629086a2cf4f72d7c330df"
# ### Calculate RMSE for Simple Exponential Smoothing
# + _uuid="ff18cf53c3ba294219f7ef855b35ac0eac20aa5e"
rmse = sqrt(mean_squared_error(valid.Count, y_hat['SES']))
rmse
# + [markdown] _uuid="3630634f01b68dcb7bf71f951815f4708057ceae"
# # Predicting Timeseris: Holt's Linear Trend Model
#
# For this method, modify the `smoothing_level` and `smoothing_slope` parameters to see how it affects your predictions.
# + _uuid="25b15d3f0471132dfd5d7f1e87f8f47c8f52706c"
plt.style.use('default')
plt.figure(figsize = (16,8))
import statsmodels.api as sm
sm.tsa.seasonal_decompose(Train.Count).plot()
result = sm.tsa.stattools.adfuller(train.Count)
plt.show()
# + _uuid="511fec7e253db745d083be532e0418435fec0e9d"
y_hat_holt = valid.copy()
fit1 = Holt(np.asarray(Train['Count'])).fit(smoothing_level = 0.3, smoothing_slope = 0.1)
y_hat_holt['Holt_linear'] = fit1.forecast(len(valid))
plt.style.use('fivethirtyeight')
plt.figure(figsize = (15,8))
plt.plot(Train.Count, label = 'Train')
plt.plot(valid.Count, label = 'Validation')
plt.plot(y_hat_holt['Holt_linear'], label = 'Holt Linear')
plt.legend(loc = 'best')
# + _uuid="e2cfd3bd7a5e2968c2ef1b69bb33e4e0e4d0b2c1"
rmse = sqrt(mean_squared_error(valid.Count, y_hat_holt.Holt_linear))
rmse
# + _uuid="8aa473bc92b6e8d1d4d4d446b4077bfd5a91fccb"
predict = fit1.forecast(len(test))
test['prediction'] = predict
# + _uuid="61596ea4e2e6cae2ce5ddaac3970abc38404a868"
#Calculating hourly ration of count
train_original['ratio'] = train_original['Count']/train_original['Count'].sum()
#Grouping hourly ratio
temp = train_original.groupby(['Hour']) ['ratio'].sum()
#Group by to csv format
pd.DataFrame(temp, columns= ['Hour', 'ratio']).to_csv('Groupby.csv')
temp2 = pd.read_csv("Groupby.csv")
temp2 =temp2.drop('Hour.1',1)
#Merge test and test_original on day, month and year
merge = pd.merge(test, test_original, on = ('day', 'month','year'), how = 'left')
merge['Hour'] = merge['Hour_y']
merge = merge.drop(['year','month','day','Hour_x','Datetime','Hour_y'], axis =1)
#Predicting by merging temp2 and merge
prediction = pd.merge(merge, temp2, on = 'Hour',how = 'left')
#Converting the ration to original scale
prediction['Count'] = prediction['prediction'] * prediction['ratio'] * 24
prediction['ID'] = prediction['ID_y']
prediction.head()
# + _uuid="89b0a6ba6252d6ed932bbb74e91d3706b572f0c3"
submission = prediction.drop(['ID_x','ID_y','prediction','Hour','ratio'], axis =1)
pd.DataFrame(submission, columns = ['ID','Count']).to_csv('Holt_Linear.csv')
# + [markdown] _uuid="c194c5e1990d9b25e15ae94ab24008885985f6a9"
# # Predicting Timeseries: Holt Winter's Model
#
# In this method, you can change the `ExponentialSmoothing` parameters. How many season_periods gives the best predictions?
# + _uuid="fb06270c7863e22af1cc011875f65312a0054ca5"
y_hat_avg = valid.copy()
fit1 = ExponentialSmoothing(np.asarray(Train['Count']), seasonal_periods= 7, trend = 'add', seasonal= 'add').fit()
y_hat_avg['Holt_Winter'] = fit1.forecast(len(valid))
plt.figure(figsize = (16,8))
plt.plot(Train['Count'], label = 'Train')
plt.plot(valid['Count'], label = 'Validation')
plt.plot(y_hat_avg.Holt_Winter, label = 'Holt Winters')
plt.legend(loc = 'best')
# + _uuid="2da66bdd9426862bd3be720be50c55cbac5c127e"
rmse = sqrt(mean_squared_error(valid['Count'], y_hat_avg['Holt_Winter']))
rmse
# + _uuid="85a664f8ee45b2a5ee8338885adf81665df8a25e"
predict = fit1.forecast(len(test))
test['prediction'] = predict
# + _uuid="3dfe9d683e6cfa5551b35f537c24e02dcb5dbf0a"
#Merge test and test_original on day,month and year
merge = pd.merge(test, test_original, on = ('day', 'month', 'year'), how = 'left')
merge['Hour']= merge['Hour_y']
merge.head()
merge = merge.drop(['year', 'month', 'Datetime','Hour_x', 'Hour_y'], axis =1)
#Predicting by merge and temp2
prediction = pd.merge(merge, temp2 , on = 'Hour', how = 'left')
#Converting the ration to original scale
prediction['Count'] = prediction['prediction'] * prediction['ratio'] *24
prediction.head()
# + _uuid="ff3484ebae9aae6698e94c8655c3bb468837f12e"
prediction['ID']= prediction['ID_y']
submission = prediction.drop(['ID_x','ID_y','day','Hour','prediction','ratio'], axis =1)
pd.DataFrame(submission, columns = ['ID','Count']).to_csv('Holt winters.csv')
# + [markdown] _uuid="7aa6285a78164bc3640d4ce35d61cee89faa09fb"
# # Predicting Time Series: ARIMA Model
#
# Experiment with this ARIMA model by adjusting the rolling average and standard deviation windows. How do they affect the results of the Dickey-Fuller test?
# + _uuid="3f16dbe5920b5c6acf59e8658357d4504241cb8b"
from statsmodels.tsa.stattools import adfuller
def test_stationary(timeseries):
#Determine rolling statistics
rolmean = timeseries.rolling(24).mean()
rolstd = timeseries.rolling(24).std()
#Plot rolling Statistics
orig = plt.plot(timeseries, color = "blue", label = "Original")
mean = plt.plot(rolmean, color = "red", label = "Rolling Mean")
std = plt.plot(rolstd, color = "black", label = "Rolling Std")
plt.legend(loc = "best")
plt.title("Rolling Mean and Standard Deviation")
plt.show(block = False)
#Perform Dickey Fuller test
print("Results of Dickey Fuller test: ")
dftest = adfuller(timeseries, autolag = 'AIC')
dfoutput = pd.Series(dftest[0:4], index = ['Test Statistics', 'p-value', '# Lag Used', 'Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)' %key] = value
print(dfoutput)
# + _uuid="5fca38363901ee82aa2d2705ab23eb4ef4dc73e6"
from matplotlib.pylab import rcParams
rcParams['figure.figsize']=(20,10)
test_stationary(train_original['Count'])
# + [markdown] _uuid="a6adc15ab0baffe09faac7f2eab965a33862a1d0"
# ### Remove trends
# + _uuid="c8b1e0e36f11a1d6e41f7c04bc735b10e7606f46"
Train_log = np.log(Train['Count'])
valid_log = np.log(valid['Count'])
# + _uuid="b385377a96eee9622ca2f2e9d62d135f8f454061"
moving_avg = Train_log.rolling(24).mean()
plt.plot(Train_log)
plt.plot(moving_avg, color = 'red')
# + _uuid="2c20f04bcb3820d7a7c900f870bef98eea05fa6f"
train_log_moving_diff = Train_log - moving_avg
train_log_moving_diff.dropna(inplace = True)
test_stationary(train_log_moving_diff)
# + [markdown] _uuid="4546b34fa6fd8c30bb7bfb285b5f217ce15a95ef"
# ### Differencing can help to make the series stable and eliminate trends
# + _uuid="7691022b05bafb82ee7a602feff7c1a7602842ad"
train_log_diff = Train_log - Train_log.shift(1)
test_stationary(train_log_diff.dropna())
# + [markdown] _uuid="af98f5d4eb829b09e07e53573a411cdd7ec42997"
# ### Removing Seasonailty
# + _uuid="072d280550d248010be412c5a536188e43482201"
from statsmodels.tsa.seasonal import seasonal_decompose
plt.figure(figsize = (16,10))
decomposition = seasonal_decompose(pd.DataFrame(Train_log).Count.values, freq = 24)
plt.style.use('default')
trend = decomposition.trend
seasonal = decomposition.seasonal
residual = decomposition.resid
plt.subplot(411)
plt.plot(Train_log, label = 'Original')
plt.legend(loc = 'best')
plt.subplot(412)
plt.plot(trend, label = 'Trend')
plt.legend(loc = 'best')
plt.subplot(413)
plt.plot(seasonal, label = 'Seasonal')
plt.legend(loc = 'best')
plt.subplot(414)
plt.plot(residual, label = 'Residuals')
plt.legend(loc = 'best')
plt.tight_layout()
# + [markdown] _uuid="0c8c8b03d48e528f2e10b28a755dc3846275f148"
# ### Check stationarity of residuals
# + _uuid="367f440768793a2f2c082bd0654758a983cf8f7e"
plt.figure(figsize = (16,8))
train_log_decompose = pd.DataFrame(residual)
train_log_decompose['date'] = Train_log.index
train_log_decompose.set_index('date', inplace = True)
train_log_decompose.dropna(inplace = True)
test_stationary(train_log_decompose[0])
# + _uuid="87e2d816a34f8e19ccaad6d2170b031ffed1308f"
from statsmodels.tsa.stattools import acf, pacf
lag_acf = acf(train_log_diff.dropna(), nlags = 25)
lag_pacf = pacf(train_log_diff.dropna(), nlags = 25, method= "ols")
# + _uuid="4d74c1ddd58bd69656225ba4be989a85f36dbf3d"
plt.figure(figsize = (15,8))
plt.style.use("fivethirtyeight")
plt.plot(lag_acf)
plt.axhline( y = 0, linestyle = "--", color = "gray")
plt.axhline( y= -1.96/np.sqrt(len(train_log_diff.dropna())), linestyle = "--", color = "gray")
plt.axhline(y = 1.96 /np.sqrt(len(train_log_diff.dropna())), linestyle = "--", color = "gray")
plt.title("Autocorrelation Function")
plt.show()
# PACF
plt.figure(figsize = (15,8))
plt.plot(lag_pacf)
plt.axhline(y = 0, linestyle = "--", color = "gray")
plt.axhline(y = -1.96/np.sqrt(len(train_log_diff.dropna())), linestyle = "--", color = "gray")
plt.axhline( y = 1.96/np.sqrt(len(train_log_diff.dropna())), linestyle = "--", color = "gray")
plt.title("Partial Autocorrelation Function")
plt.show()
# + [markdown] _uuid="2b6f8aa10b137871365390c6aaef23fd23066adc"
# # Predicting Timeseries: AR Model
#
# If you're unfamiliar with the ARIMA model, it might be helpful to see what parameters and arguments you can use to improve your predictions. Read up on `p,d,q`.<br>
# [https://www.statsmodels.org/dev/generated/statsmodels.tsa.arima_model.ARIMA.html](https://www.statsmodels.org/dev/generated/statsmodels.tsa.arima_model.ARIMA.html)
# + _uuid="136116404e359ae60f0190b11d7a4032f5af6070"
from statsmodels.tsa.arima_model import ARIMA
plt.figure(figsize = (15,8))
model = ARIMA(Train_log, order = (2,1,0)) #here q value is zero since it is just AR Model
results_AR = model.fit(disp=-1)
plt.plot(train_log_diff.dropna(), label = "Original")
plt.plot(results_AR.fittedvalues, color = 'red', label = 'Predictions')
plt.legend(loc = 'best')
# + _uuid="b674ac36018574661a2801e976340c350f0511c2"
AR_predict = results_AR.predict(start="2014-06-25", end="2014-09-25")
AR_predict = AR_predict.cumsum().shift().fillna(0)
AR_predict1 = pd.Series(np.ones(valid.shape[0])* np.log(valid['Count'])[0], index = valid.index)
AR_predict1=AR_predict1.add(AR_predict,fill_value=0)
AR_predict = np.exp(AR_predict1)
# + _uuid="4cf9fe9ba34ef3d76d515c1bce6794b84ef3dc74"
plt.figure(figsize = (15,8))
plt.plot(valid['Count'], label = "Validation")
plt.plot(AR_predict, color = "red", label = "Predict")
plt.legend(loc = "best")
plt.title('RMSE: %.4f'% (np.sqrt(np.dot(AR_predict, valid['Count']))/valid.shape[0]))
plt.show()
# + [markdown] _uuid="20326483786cc61f08b62ad98fc90e9dd208118e"
# # Predicting Timeseries: Moving Average Model
#
# For this model, you can modify the same parameters (p,d,q) because the base model relies on ARIMA.
# + _uuid="96e5ed7e83199c7f8ddc528b1a45b459b4f75785"
plt.figure(figsize = (15,8))
model = ARIMA(Train_log, order = (0,1,2)) # here the p value is 0 since it is moving average model
results_MA = model.fit(disp = -1)
plt.plot(train_log_diff.dropna(), label = "Original")
plt.plot(results_MA.fittedvalues, color = "red", label = "Prediction")
plt.legend(loc = "best")
# + _uuid="42c1ea42d899794463764392cb763e613b6529a9"
MA_predict = results_MA.predict(start="2014-06-25", end="2014-09-25")
MA_predict=MA_predict.cumsum().shift().fillna(0)
MA_predict1=pd.Series(np.ones(valid.shape[0]) * np.log(valid['Count'])[0], index = valid.index)
MA_predict1=MA_predict1.add(MA_predict,fill_value=0)
MA_predict = np.exp(MA_predict1)
# + _uuid="b878152fffa01bff5914636a8aa66c47ab737313"
plt.figure(figsize = (15,8))
plt.plot(valid['Count'], label = "Valid")
plt.plot(MA_predict, color = 'red', label = "Predict")
plt.legend(loc= 'best')
plt.title('RMSE: %.4f'% (np.sqrt(np.dot(MA_predict, valid['Count']))/valid.shape[0]))
plt.show()
# + [markdown] _uuid="790469da0d7bb3fd1c3828eda1e63b75ff4e03aa"
# ### Combining Models
# + _uuid="a9ba2862b1027e02852e9687f819bb93e9ec0cf0"
plt.figure(figsize = (16,8))
model = ARIMA(Train_log, order=(2, 1, 2))
results_ARIMA = model.fit(disp=-1)
plt.plot(train_log_diff.dropna(), label='Original')
plt.plot(results_ARIMA.fittedvalues, color='red', label='Predicted')
plt.legend(loc='best')
plt.show()
# + [markdown] _uuid="9252435f4d4860e79bdef6f838c0050b02462240"
# ### Scale model to original scale
# + _uuid="f428106f716c917bfcdb19e395cabf6c55d20aec"
def check_prediction_diff(predict_diff, given_set):
predict_diff= predict_diff.cumsum().shift().fillna(0)
predict_base = pd.Series(np.ones(given_set.shape[0]) * np.log(given_set['Count'])[0], index = given_set.index)
predict_log = predict_base.add(predict_diff,fill_value=0)
predict = np.exp(predict_log)
plt.plot(given_set['Count'], label = "Given set")
plt.plot(predict, color = 'red', label = "Predict")
plt.legend(loc= 'best')
plt.title('RMSE: %.4f'% (np.sqrt(np.dot(predict, given_set['Count']))/given_set.shape[0]))
plt.show()
# + _uuid="368e8110a2b29a6d0cd62a0b91382f563b751acf"
def check_prediction_log(predict_log, given_set):
predict = np.exp(predict_log)
plt.plot(given_set['Count'], label = "Given set")
plt.plot(predict, color = 'red', label = "Predict")
plt.legend(loc= 'best')
plt.title('RMSE: %.4f'% (np.sqrt(np.dot(predict, given_set['Count']))/given_set.shape[0]))
plt.show()
# + _uuid="b7968788888244c49b7b910fe4a875e91ef381b9"
ARIMA_predict_diff=results_ARIMA.predict(start="2014-06-25", end="2014-09-25")
# + _uuid="91a7a3fa847c7a3b239e9f73e0dfbb8f599d0700"
plt.figure(figsize = (16,8))
check_prediction_diff(ARIMA_predict_diff, valid)
# + [markdown] _uuid="bd74c9861565749513e454de2d977bd883eedace"
# # Predicting Timeseries: SARIMAX Model
#
# By this point, you've probably got a pretty good idea about which parameters you can modify to make improvements to your predictions. Try out what you've learned using the SARIMAX model.
# + _uuid="f56a45c3f85eebfcf4483ae0af9a4f5a2d4eeddb"
import statsmodels.api as sm
# + _uuid="7bd1508f578f7bf63daaefd3db4dbd823f2c6422"
y_hat_avg = valid.copy()
fit1 = sm.tsa.statespace.SARIMAX(Train.Count, order = (2,1,4), seasonal_order =(0,1,1,7)).fit()
y_hat_avg['SARIMA'] = fit1.predict(start="2014-6-25", end="2014-9-25", dynamic=True)
plt.figure(figsize=(16,8))
plt.plot(Train['Count'], label = "Train")
plt.plot(valid.Count, label = "Validation")
plt.plot(y_hat_avg['SARIMA'], label ="SARIMA")
plt.legend(loc = "best")
plt.title("SARIMAX Model")
# + _uuid="9ec2369438ecf5eec22a64b0b8e2bbcefb5d0f09"
rms = sqrt(mean_squared_error(valid.Count, y_hat_avg.SARIMA))
print(rms)
# + [markdown] _uuid="50e3d0ad2675331d44df151cec2eb6ebd9d06285"
# ### Covert to Hourly Predictions
# + _uuid="854f50e8aa413157080f322357c65c585b1a4bb0"
predict = fit1.predict(start="2014-9-26", end="2015-4-26", dynamic=True)
# + _uuid="646970748036902d0be7479e094505b8ab1ee7f0"
test['prediction']=predict
# + _uuid="c34a8802998888583942b5303edb63b453aa972e"
#Merge test and test_original on day,month and year
merge = pd.merge(test,test_original, on = ('day', 'month', 'year'), how = 'left')
merge['Hour'] = merge['Hour_y']
#Predicting by merging merge and temp2
prediction = pd.merge(merge, temp2, on = 'Hour', how = 'left')
#Converting the ratio to original scale
prediction['Count'] = prediction['prediction'] * prediction['ratio'] * 24
# + _uuid="16c3cfd3f649383ed43fbbffb667ba299da9f8dc"
prediction['ID']=prediction['ID_y']
submission=prediction.drop(['day','Hour','ratio','prediction', 'ID_x', 'ID_y'],axis=1)
# Converting the final submission to csv format
pd.DataFrame(submission, columns=['ID','Count']).to_csv('SARIMAX.csv')
# -
# ---
#
# **[Time Series With Siraj Course Home Page](https://kaggle.com/learn/time-series-with-siraj)**
#
| learntools/time_series_with_siraj/nbs/ex2_time_series_modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import collections
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import clear_output
import utils
# +
params = {'growth_rate_real_estate': [i/100 for i in list(range(3, 10))],
'growth_rate_stocks': [i/100 for i in list(range(3, 10))],
'initial_price_stocks': [100],
'mortgage_overpayment_amount': list(range(0, 20001, 5000)),
'investment_amount': list(range(0, 20001, 5000))}
scenarios = []
for gre in params['growth_rate_real_estate']:
for gs in params['growth_rate_stocks']:
for ps in params['initial_price_stocks']:
for ma in params['mortgage_overpayment_amount']:
for ia in params['investment_amount']:
scenarios.append(utils.Scenario(utils.RealEstate(4.2e6, utils.Mortgage(0.0305, 3.6e6, 30, 12)), utils.Portfolio(), gre, gs, ps, ma, ia))
sim = utils.Simulation(scenarios)
# -
# %%time
sim.simulate()
history = sim.history
history.head()
history.info()
totals = sim.profit
totals.head()
totals.info()
real_estate_price_15_y = 77339
real_estate_price_0_y = 27308
snp_price_15_y = 3235
snp_price_0_y = 1186
growth_rate_real_estate = math.pow(real_estate_price_15_y/real_estate_price_0_y, 1/15) - 1
growth_rate_stocks = math.pow(snp_price_15_y/snp_price_0_y, 1/15) - 1
print(f'Growth rate real estate for the last 15 years: {growth_rate_real_estate:.2%}, growth rate S&P500 for the last 15 years: {growth_rate_stocks:.2%}.')
| prototype.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from scipy import stats
import os
import glob
import numpy as np
path = r'./dataFolders/Output/Step6_v5/'
outpath = r'./dataFolders/Output/Step6_v5/'
def in_ranges(x,bins):
return [((x>=y[0])&(x<=y[1])) for y in bins]
notoutliers = pd.read_csv(path + 'AllLight_EveryMoth_notOutliers.csv')
notoutliers['DiscoveryTime'] = notoutliers.DiscoveryTime.div(100,axis = 'index')
notoutliers = notoutliers.drop(columns = ['Unnamed: 0', 'Unnamed: 0.1', 'Unnamed: 0.1.1'])
notoutliers.head()
Bins = [(0, 10), (5, 15), (10, 20), (15, 25), (20, 30)]
label = ['(0,10)', '(5,15)', '(10,20)', '(15,25)', '(20,30)']
binned = notoutliers['trialNum'].apply(lambda x: pd.Series(in_ranges(x,Bins), label))
binned
notoutliers = notoutliers.join(binned)
test = notoutliers.name.str.split('_', expand = True)
notoutliers['lightLevel'] = test[0]
notoutliers.head()
notoutliers.columns
for bin_label in label:
sub_df = notoutliers[notoutliers[bin_label] == True]
for l, df in sub_df.groupby('lightLevel'):
n = len(df)
print('%s and %s bin has %i visits' %(l, bin_label, n))
# ### mark early and late visits
notoutliers.loc[notoutliers.lightLevel == 'L50', 'trialNum'].max()
Bins = [(0, 4), (14,24)]
label = ['early', 'late']
earlyLate_binned = notoutliers['trialNum'].apply(lambda x: pd.Series(in_ranges(x,Bins), label))
test = notoutliers.join(earlyLate_binned)
test.head()
for l, df in test.groupby('lightLevel'):
n = len(df[df.early == True])
print('%s has %s early visits' %(l, n))
n = len(df[df.late == True])
print('%s has %s early visits' %(l, n))
# ## get bins for first and last n visits
# +
high_mend = test.loc[test.lightLevel == 'L50', 'trialNum'].max()
high_mstrt = test.loc[test.lightLevel == 'L50', 'trialNum'].max()-10
low_mend = test.loc[test.lightLevel == 'L0.1', 'trialNum'].max()
low_mstrt = test.loc[test.lightLevel == 'L0.1', 'trialNum'].max()-10
# -
high_mend, high_mstrt
low_mend, low_mstrt
# +
Bins = [(0, 2)]
label = ['early3']
early3 = test['trialNum'].apply(lambda x: pd.Series(in_ranges(x,Bins), label))
test = test.join(early3)
# +
Bins = [(high_mstrt, high_mend)]
label = ['late10_high']
Late10_binned_high = test.loc[test.lightLevel == 'L50', 'trialNum'].apply(lambda x: pd.Series(in_ranges(x,Bins), label))
# +
Bins = [(low_mstrt, low_mend)]
label = ['late10_low']
Late10_binned_low = test.loc[test.lightLevel == 'L0.1', 'trialNum'].apply(lambda x: pd.Series(in_ranges(x,Bins), label))
# -
new_test = test.join(Late10_binned_high, how='outer')
final_test = pd.concat([new_test, Late10_binned_low], axis=1, sort=False)
final_test.head()
sub = final_test.loc[final_test.lightLevel == 'L0.1', :]
len(sub[sub['late10_low'] == True])
sub = final_test.loc[final_test.lightLevel == 'L50', :]
len(sub[sub['late10_high'] == True])
sub = final_test.loc[final_test.lightLevel == 'L0.1', :]
len(sub[sub['early3'] == True])
sub = final_test.loc[final_test.lightLevel == 'L50', :]
len(sub[sub['early3'] == True])
final_test.to_csv(outpath + 'AllLight_EveryMoth_notOutliers_withBins.csv')
# ## generate and store the pde and data for plotting later
from scipy.stats import gaussian_kde
bin_center = np.linspace(0,40,100)
delta = np.diff(bin_center)[0]
notoutliers = pd.read_csv(outpath + 'AllLight_EveryMoth_notOutliers_withBins.csv')
# ### store the sliding window pde
# +
label = ['(0,10)', '(5,15)', '(10,20)', '(15,25)', '(20,30)']
SampleSize = pd.DataFrame(columns = ['L0.1', 'L50'], index = label)
pde_df = pd.DataFrame(columns = label, index = bin_center)
for l, subdf in notoutliers.groupby('lightLevel'):
for bin_label in label:
df = subdf[subdf[bin_label] == True]
data = df.DiscoveryTime
SampleSize.loc[bin_label,l] = len(data)
kde = gaussian_kde(data)
temp = kde.pdf(bin_center)
temp /= delta * np.sum(temp)
pde_df.loc[:,bin_label]=temp
pde_df.to_csv(outpath+ l + '_pde.csv')
SampleSize.to_csv(outpath+ 'samplesize.csv')
# -
# ### Store the pde for early and late visits
# +
label = ['early', 'late']
SampleSize = pd.DataFrame(columns = ['L0.1', 'L50'], index = label)
pde_df = pd.DataFrame(columns = label, index = bin_center)
for l, subdf in notoutliers.groupby('lightLevel'):
for bin_label in label:
df = subdf[subdf[bin_label] == True]
data = df.DiscoveryTime
SampleSize.loc[bin_label,l] = len(data)
kde = gaussian_kde(data)
temp = kde.pdf(bin_center)
temp /= delta * np.sum(temp)
pde_df.loc[:,bin_label]=temp
pde_df.to_csv(outpath+ l + '_earlyLate_pde.csv')
SampleSize.to_csv(outpath+ 'earlyLateSamplesize.csv')
# -
# ## store the pde for first 3 and last 10 visits
# +
superlabel = ['early3', 'last10']
SampleSize = pd.DataFrame(columns = ['L0.1', 'L50'], index = superlabel)
pde_df = pd.DataFrame(columns = superlabel, index = bin_center)
for l, subdf in notoutliers.groupby('lightLevel'):
if l == 'L0.1':
label = ['early3', 'late10_low']
else:
label = ['early3', 'late10_high']
for bin_label, bl in zip(label, superlabel):
df = subdf[subdf[bin_label] == True]
data = df.DiscoveryTime
SampleSize.loc[bl,l] = len(data)
kde = gaussian_kde(data)
temp = kde.pdf(bin_center)
temp /= delta * np.sum(temp)
pde_df.loc[:,bl]=temp
pde_df.to_csv(outpath+ l + '_firstlast_pde.csv')
SampleSize.to_csv(outpath+ 'firstlastSamplesize.csv')
# -
| Step6b_Learning_Curves_And_Probability_Distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/mnslarcher/cs224w-slides-to-code/blob/main/notebooks/06-graph-neural-networks-1-gnn-model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PJSONe-045tN"
try:
# Check if PyTorch Geometric is installed:
import torch_geometric
except ImportError:
# If PyTorch Geometric is not installed, install it.
# %pip install -q torch-scatter -f https://pytorch-geometric.com/whl/torch-1.7.0+cu101.html
# %pip install -q torch-sparse -f https://pytorch-geometric.com/whl/torch-1.7.0+cu101.html
# %pip install -q torch-geometric
# + [markdown] id="oV56fjpNYjCR"
# # Graph Neural Networks 1: GNN Model
# + colab={"base_uri": "https://localhost:8080/"} id="2OtSkdGl48E0" outputId="7efa478a-75df-4efb-991a-a3ac877f6469"
from typing import Callable, List, Optional, Tuple
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn.functional as F
import torch_geometric.transforms as T
from torch import Tensor
from torch.optim import Optimizer
from torch_geometric.data import Data
from torch_geometric.datasets import Planetoid
from torch_geometric.nn import GCNConv
from torch_geometric.utils import accuracy
from typing_extensions import Literal, TypedDict
# + [markdown] id="KYGSasdA5ziu"
# ## Cora Dataset
# + [markdown] id="_iseKiaT5GZ7"
# > From The [Papers With Code page of the Cora Dataset](https://paperswithcode.com/dataset/cora): "The Cora dataset consists of 2708 scientific publications classified into one of seven classes. The citation network consists of 5429 links. Each publication in the dataset is described by a 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary. The dictionary consists of 1433 unique words."
# + [markdown] id="z518Aul25JZv"
# > From [<NAME> (ICLR 2017)](https://arxiv.org/abs/1609.02907): "[...] evaluate prediction accuracy on a test set of 1,000 labeled examples. [...] validation set of 500 labeled examples for hyperparameter optimization (dropout rate for all layers, L2 regularization factor for the first GCN layer and number of hidden units). We do not use the validation set labels for training."
# + colab={"base_uri": "https://localhost:8080/"} id="6Gg7oGlS5NRX" outputId="b8d10e40-275d-4b51-9907-b66d23465c6b"
dataset = Planetoid("/tmp/Cora", name="Cora")
num_nodes = dataset.data.num_nodes
# For num. edges see:
# - https://github.com/pyg-team/pytorch_geometric/issues/343
# - https://github.com/pyg-team/pytorch_geometric/issues/852
num_edges = dataset.data.num_edges // 2
train_len = dataset[0].train_mask.sum()
val_len = dataset[0].val_mask.sum()
test_len = dataset[0].test_mask.sum()
other_len = num_nodes - train_len - val_len - test_len
print(f"Dataset: {dataset.name}")
print(f"Num. nodes: {num_nodes} (train={train_len}, val={val_len}, test={test_len}, other={other_len})")
print(f"Num. edges: {num_edges}")
print(f"Num. node features: {dataset.num_node_features}")
print(f"Num. classes: {dataset.num_classes}")
print(f"Dataset len.: {dataset.len()}")
# + [markdown] id="hBzvCryI5Uba"
# > From [Kipf & Welling (ICLR 2017)](https://arxiv.org/abs/1609.02907): "We initialize weights using the initialization described in Glorot & Bengio (2010) and accordingly (row-)normalize input feature vectors."
# + colab={"base_uri": "https://localhost:8080/"} id="nljKuAA85XDE" outputId="48bc3cd5-4d3b-4c1f-90ff-e435c28962da"
dataset = Planetoid("/tmp/Cora", name="Cora")
print(f"Sum of row values without normalization: {dataset[0].x.sum(dim=-1)}")
dataset = Planetoid("/tmp/Cora", name="Cora", transform=T.NormalizeFeatures())
print(f"Sum of row values with normalization: {dataset[0].x.sum(dim=-1)}")
# + [markdown] id="c0Jw_TT35AiO"
# ## Graph Convolutional Networks
# + [markdown] id="b19trEwa5Y-W"
# > From [Kipf & Welling (ICLR 2017)](https://arxiv.org/abs/1609.02907): "We used the following sets of hyperparameters for Citeseer, Cora and Pubmed: 0.5 (dropout rate), $5\cdot10^{-4}$ (L2 regularization) and 16 (number of hidden units);"
# + id="-m_i0aCa5eM4"
class GCN(torch.nn.Module):
def __init__(
self,
num_node_features: int,
num_classes: int,
hidden_dim: int = 16,
dropout_rate: float = 0.5,
) -> None:
super().__init__()
self.dropout1 = torch.nn.Dropout(dropout_rate)
self.conv1 = GCNConv(num_node_features, hidden_dim)
self.relu = torch.nn.ReLU(inplace=True)
self.dropout2 = torch.nn.Dropout(dropout_rate)
self.conv2 = GCNConv(hidden_dim, num_classes)
def forward(self, x: Tensor, edge_index: Tensor) -> torch.Tensor:
x = self.dropout1(x)
x = self.conv1(x, edge_index)
x = self.relu(x)
x = self.dropout2(x)
x = self.conv2(x, edge_index)
return x
# + colab={"base_uri": "https://localhost:8080/"} id="jKQH-HKS5hVt" outputId="e98433b7-97c9-4469-e409-6aa8b07dea4e"
print("Graph Convolutional Network (GCN):")
GCN(dataset.num_node_features, dataset.num_classes)
# + [markdown] id="mN5NPwag6GPT"
# ## Training and Evaluation
# +
LossFn = Callable[[Tensor, Tensor], Tensor]
Stage = Literal["train", "val", "test"]
def train_step(
model: torch.nn.Module, data: Data, optimizer: torch.optim.Optimizer, loss_fn: LossFn
) -> Tuple[float, float]:
model.train()
optimizer.zero_grad()
mask = data.train_mask
logits = model(data.x, data.edge_index)[mask]
preds = logits.argmax(dim=1)
y = data.y[mask]
loss = loss_fn(logits, y)
# + L2 regularization to the first layer only
# for name, params in model.state_dict().items():
# if name.startswith("conv1"):
# loss += 5e-4 * params.square().sum() / 2.0
acc = accuracy(preds, y)
loss.backward()
optimizer.step()
return loss.item(), acc
@torch.no_grad()
def eval_step(model: torch.nn.Module, data: Data, loss_fn: LossFn, stage: Stage) -> Tuple[float, float]:
model.eval()
mask = getattr(data, f"{stage}_mask")
logits = model(data.x, data.edge_index)[mask]
preds = logits.argmax(dim=1)
y = data.y[mask]
loss = loss_fn(logits, y)
# + L2 regularization to the first layer only
# for name, params in model.state_dict().items():
# if name.startswith("conv1"):
# loss += 5e-4 * params.square().sum() / 2.0
acc = accuracy(preds, y)
return loss.item(), acc
# + [markdown] id="oMqTMjZD5mTy"
# > From [<NAME> (ICLR 2017)](https://arxiv.org/abs/1609.02907): "We train all models for a maximum of 200 epochs (training iterations) using Adam (Kingma & Ba, 2015) with a learning rate of 0.01 and early stopping with a window size of 10, i.e. we stop training if the validation loss does not decrease for 10 consecutive epochs."
# +
class HistoryDict(TypedDict):
loss: List[float]
acc: List[float]
val_loss: List[float]
val_acc: List[float]
def train(
model: torch.nn.Module,
data: Data,
optimizer: torch.optim.Optimizer,
loss_fn: LossFn = torch.nn.CrossEntropyLoss(),
max_epochs: int = 200,
early_stopping: int = 10,
print_interval: int = 20,
verbose: bool = True,
) -> HistoryDict:
history = {"loss": [], "val_loss": [], "acc": [], "val_acc": []}
for epoch in range(max_epochs):
loss, acc = train_step(model, data, optimizer, loss_fn)
val_loss, val_acc = eval_step(model, data, loss_fn, "val")
history["loss"].append(loss)
history["acc"].append(acc)
history["val_loss"].append(val_loss)
history["val_acc"].append(val_acc)
# The official implementation in TensorFlow is a little different from what is described in the paper...
if epoch > early_stopping and val_loss > np.mean(history["val_loss"][-(early_stopping + 1) : -1]):
if verbose:
print("\nEarly stopping...")
break
if verbose and epoch % print_interval == 0:
print(f"\nEpoch: {epoch}\n----------")
print(f"Train loss: {loss:.4f} | Train acc: {acc:.4f}")
print(f" Val loss: {val_loss:.4f} | Val acc: {val_acc:.4f}")
test_loss, test_acc = eval_step(model, data, loss_fn, "test")
if verbose:
print(f"\nEpoch: {epoch}\n----------")
print(f"Train loss: {loss:.4f} | Train acc: {acc:.4f}")
print(f" Val loss: {val_loss:.4f} | Val acc: {val_acc:.4f}")
print(f" Test loss: {test_loss:.4f} | Test acc: {test_acc:.4f}")
return history
# -
def plot_history(history: HistoryDict, title: str, font_size: Optional[int] = 14) -> None:
plt.suptitle(title, fontsize=font_size)
ax1 = plt.subplot(121)
ax1.set_title("Loss")
ax1.plot(history["loss"], label="train")
ax1.plot(history["val_loss"], label="val")
plt.xlabel("Epoch")
ax1.legend()
ax2 = plt.subplot(122)
ax2.set_title("Accuracy")
ax2.plot(history["acc"], label="train")
ax2.plot(history["val_acc"], label="val")
plt.xlabel("Epoch")
ax2.legend()
# +
SEED = 42
MAX_EPOCHS = 200
LEARNING_RATE = 0.01
WEIGHT_DECAY = 5e-4
EARLY_STOPPING = 10
torch.manual_seed(SEED)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = GCN(dataset.num_node_features, dataset.num_classes).to(device)
data = dataset[0].to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE, weight_decay=WEIGHT_DECAY)
history = train(model, data, optimizer, max_epochs=MAX_EPOCHS, early_stopping=EARLY_STOPPING)
plt.figure(figsize=(12, 4))
plot_history(history, "GCN")
| notebooks/06-graph-neural-networks-1-gnn-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Summary
# # Imports
# +
import importlib
import os
import sys
from collections import Counter
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pyarrow as pa
import pyarrow.parquet as pq
import torch
import torch.nn as nn
import torch.nn.functional as F
# +
# %matplotlib inline
pd.set_option("max_columns", 100)
# +
SRC_PATH = Path.cwd().joinpath('..', 'src').resolve(strict=True)
if SRC_PATH.as_posix() not in sys.path:
sys.path.insert(0, SRC_PATH.as_posix())
import helper
importlib.reload(helper)
# -
# # Parameters
NOTEBOOK_PATH = Path(os.getenv("CI_JOB_NAME", "add_adjacency_distances_test"))
NOTEBOOK_PATH
OUTPUT_PATH = Path(os.getenv('OUTPUT_DIR', NOTEBOOK_PATH.name)).resolve()
OUTPUT_PATH.mkdir(parents=True, exist_ok=True)
OUTPUT_PATH
# +
DEBUG = "CI" not in os.environ
TASK_ID = os.getenv("SLURM_ARRAY_TASK_ID")
TASK_COUNT = os.getenv("ORIGINAL_ARRAY_TASK_COUNT") or os.getenv("SLURM_ARRAY_TASK_COUNT")
TASK_ID = int(TASK_ID) if TASK_ID is not None else None
TASK_COUNT = int(TASK_COUNT) if TASK_COUNT is not None else None
TASK_ID, TASK_COUNT
# -
if DEBUG:
# %load_ext autoreload
# %autoreload 2
# # `DATAPKG`
DATAPKG = {
'training_dataset':
Path(os.environ['DATAPKG_OUTPUT_DIR']).joinpath(
"adjacency-net-v2", "master", "training_dataset"),
'training_dataset_wdistances':
Path(os.environ['DATAPKG_OUTPUT_DIR']).joinpath(
"adjacency-net-v2", "master", "training_dataset_wdistances"),
'pdb_mmcif_ffindex':
Path(os.environ['DATAPKG_OUTPUT_DIR']).joinpath(
"pdb-ffindex", "master", "pdb_mmcif_ffindex", "pdb-mmcif"),
}
# # Load data
parquet_files = list(
DATAPKG['training_dataset_wdistances'].joinpath("adjacency_matrix.parquet").glob("*/*.parquet")
)
# +
dfs = []
for i, parquet_file in enumerate(parquet_files[:10]):
print(i)
file_obj = pq.ParquetFile(parquet_file)
df = file_obj.read_row_group(0).to_pandas()
dfs.append(df)
# -
master_df = pd.concat(dfs)
# ## Save
# +
output_file = OUTPUT_PATH.joinpath("example_rows.parquet")
if not output_file.is_file():
table = pa.Table.from_pandas(master_df, preserve_index=False)
pq.write_table(table, output_file)
# -
# # Process data
# ## Get distances
# +
output_file = OUTPUT_PATH.joinpath("example_rows.parquet")
master_df = pq.read_table(output_file).to_pandas()
# -
master_df = master_df[
(master_df['residue_idx_1_corrected'].notnull())
].copy()
# ### `get_aa_distances`
def get_aa_distances(seq, residue_idx_1_corrected, residue_idx_2_corrected):
arr1 = np.array(residue_idx_1_corrected)
arr2 = np.array(residue_idx_2_corrected)
aa_distances = np.hstack([np.zeros(len(seq), dtype=np.int), np.abs(arr1 - arr2)])
return aa_distances
master_df["aa_distances"] = [
get_aa_distances(seq, idx1, idx2)
for seq, idx1, idx2
in master_df[["sequence", "residue_idx_1_corrected", "residue_idx_2_corrected"]].values
]
master_df["aa_distances"].head()
aa_distances = np.hstack(master_df["aa_distances"].values)
# ### `get_cart_distances`
def get_cart_distances(seq, distances):
cart_distances = np.hstack([np.zeros(len(seq), dtype=np.float), distances])
return cart_distances
master_df["cart_distances"] = [
get_cart_distances(seq, distances)
for seq, distances
in master_df[["sequence", "distances"]].values
]
master_df["cart_distances"].head()
cart_distances = np.hstack(master_df["cart_distances"].values)
# ## Validate
assert len(aa_distances) == len(cart_distances)
# ## Shuffle
indices = np.arange(len(aa_distances))
np.random.RandomState(42).shuffle(indices)
np.random.RandomState(42).shuffle(aa_distances)
np.random.RandomState(42).shuffle(cart_distances)
assert (cart_distances[:10_000][aa_distances[:10_000] == 0] == 0).all()
# ## Save
np.save(OUTPUT_PATH.joinpath("aa_distances.npy"), aa_distances)
np.save(OUTPUT_PATH.joinpath("cart_distances.npy"), cart_distances)
# # Machine learning
# ## Load data
aa_distances = np.load(OUTPUT_PATH.joinpath("aa_distances.npy"))
cart_distances = np.load(OUTPUT_PATH.joinpath("cart_distances.npy"))
# ## Functions
# ### `gen_barcode`
# +
from numba import njit, prange
@njit(parallel=True)
def gen_barcode(distances, bins):
barcode = np.zeros((len(distances), len(bins)), dtype=np.int32)
for i in prange(len(distances)):
a = distances[i]
for j in range(len(bins)):
if a < bins[j]:
barcode[i, j] = 1
break
return barcode
# -
# ## Normalize seq distances
plt.hist(np.clip(aa_distances, 0, 100), bins=50)
plt.xlabel("Amino acid distance")
plt.label("Number of amino acid pairs")
None
def normalize_seq_distances(aa_distances):
aa_distances_log_mean = 3.5567875815104903
aa_distances_log_std = 1.7065822763411669
with np.errstate(divide='ignore'):
aa_distances_log = np.where(aa_distances > 0, np.log(aa_distances) + 1, 0)
aa_distances_corrected = (aa_distances_log - aa_distances_log_mean) / aa_distances_log_std
return aa_distances_corrected
aa_distances_corrected = normalize_seq_distances(aa_distances)
assert np.isclose(aa_distances_corrected.mean(), 0)
assert np.isclose(aa_distances_corrected.std(), 1)
plt.hist(aa_distances_corrected, bins=50)
plt.xlabel("Amino acid distance (normalized)")
plt.ylabel("Number of amino acid pairs")
None
# ## Bin seq distances
aa_quantile_custom = normalize_seq_distances(np.array([1, 4, 8, 14, 32, 100_000])).tolist()
aa_quantile_custom
# +
aa_quantile_2 = np.quantile(aa_distances_corrected, np.linspace(0, 1, 3)[1:]).tolist()
aa_quantile_4 = np.quantile(aa_distances_corrected, np.linspace(0, 1, 5)[1:]).tolist()
aa_quantile_6 = np.quantile(aa_distances_corrected, np.linspace(0, 1, 7)[1:]).tolist()
aa_quantile_2[-1] = aa_quantile_custom[-1]
aa_quantile_4[-1] = aa_quantile_custom[-1]
aa_quantile_6[-1] = aa_quantile_custom[-1]
aa_quantile_2, aa_quantile_4, aa_quantile_6
# -
# %timeit -n 1 -r 3 gen_barcode(aa_distances_corrected, aa_quantile_custom)
# %timeit -n 1 -r 3 gen_barcode(aa_distances_corrected, aa_quantile_6)
# ## Normalize cart distances
plt.hist(np.clip(cart_distances, 0, 14), bins=50)
plt.xlabel("Euclidean distance")
plt.ylabel("Number of amino acid pairs")
None
def normalize_cart_distances(cart_distances):
cart_distances_mean = 6.9936892028873965
cart_distances_std = 3.528368101492991
cart_distances_corrected = (cart_distances - cart_distances_mean) / cart_distances_std
return cart_distances_corrected
cart_distances_corrected = normalize_cart_distances(cart_distances)
assert np.isclose(cart_distances_corrected.mean(), 0)
assert np.isclose(cart_distances_corrected.std(), 1)
plt.hist(cart_distances_corrected, bins=50)
plt.xlabel("Euclidean distance (normalized)")
plt.ylabel("Number of amino acid pairs")
None
# ## Bin cart distances
cart_quantile_custom = normalize_cart_distances(np.array([1.0, 2.0, 4.0, 6.2, 8.5, 100_000.0])).tolist()
cart_quantile_custom
# +
cart_quantile_2 = np.quantile(cart_distances_corrected, np.linspace(0, 1, 3)[1:]).tolist()
cart_quantile_4 = np.quantile(cart_distances_corrected, np.linspace(0, 1, 5)[1:]).tolist()
cart_quantile_6 = np.quantile(cart_distances_corrected, np.linspace(0, 1, 7)[1:]).tolist()
cart_quantile_2[-1] = cart_quantile_custom[-1]
cart_quantile_4[-1] = cart_quantile_custom[-1]
cart_quantile_6[-1] = cart_quantile_custom[-1]
cart_quantile_2, cart_quantile_4, cart_quantile_6
# -
# %timeit -n 1 -r 3 gen_barcode(cart_distances, cart_quantile_6)
# ## Networks
# +
class DistanceNet(nn.Module):
def __init__(self, barcode_size, hidden_layer_size=32):
super().__init__()
self.barcode_size = barcode_size
self.hidden_layer_size = hidden_layer_size
self.linear1 = nn.Linear(1, self.hidden_layer_size)
self.linear2 = nn.Linear(self.hidden_layer_size, self.barcode_size)
# self.reset_parameters()
def reset_parameters(self):
stdv = np.sqrt(self.hidden_layer_size)
self.linear1.weight.data.normal_(0, stdv)
self.linear1.bias.data.normal_(0, stdv)
def forward(self, x):
x = self.linear1(x)
x = F.leaky_relu(x)
x = self.linear2(x)
x = F.leaky_relu(x)
return x
# +
class TwoDistanceNet(nn.Module):
def __init__(self, barcode_size, hidden_layer_size=64):
super().__init__()
self.hidden_layer_size = hidden_layer_size
self.barcode_size = barcode_size
self.linear1 = nn.Linear(2, self.hidden_layer_size)
self.linear2 = nn.Linear(self.hidden_layer_size, self.barcode_size)
# self.reset_parameters()
def reset_parameters(self):
stdv = np.sqrt(self.hidden_layer_size)
self.linear1.weight.data.normal_(0, stdv)
self.linear1.bias.data.normal_(0, stdv)
def forward(self, x):
x = self.linear1(x)
x = F.leaky_relu(x)
x = self.linear2(x)
x = F.leaky_relu(x)
return x
# -
np.sqrt(2 / 64)
# +
aa_distances_onehot = gen_barcode(aa_distances_corrected, aa_quantile_custom).astype(np.float32)
assert (aa_distances_onehot.sum(axis=1) == 1).all()
cart_distances_onehot = gen_barcode(cart_distances_corrected, cart_quantile_custom).astype(np.float32)
assert (cart_distances_onehot.sum(axis=1) == 1).all()
# +
model_a = DistanceNet(6)
model_c = DistanceNet(6)
model_ac = TwoDistanceNet(12)
learning_rate = 1e-4 #0.00005
betas = (0.5, 0.9)
optimizer_a = torch.optim.Adam(model_a.parameters(), lr=learning_rate, betas=betas)
optimizer_c = torch.optim.Adam(model_c.parameters(), lr=learning_rate, betas=betas)
optimizer_ac = torch.optim.Adam(model_ac.parameters(), lr=learning_rate, betas=betas)
# loss_fn = nn.BCELoss()
loss_fn = nn.MSELoss()
batch_size = 64
#
losses_a = []
losses_c = []
losses_ac = []
# -
for t in range(30_000):
if t % 2_000 == 0:
print(t)
t_slice = slice(t * batch_size, (t + 1) * batch_size)
X_a = torch.from_numpy(aa_distances_corrected[t_slice]).to(torch.float32).unsqueeze(1)
X_c = torch.from_numpy(cart_distances_corrected[t_slice]).to(torch.float32).unsqueeze(1)
X_ac = torch.cat([X_a, X_c], 1)
Y_a = torch.from_numpy(aa_distances_onehot[t_slice, :])
Y_c = torch.from_numpy(cart_distances_onehot[t_slice, :])
Y_ac = torch.cat([Y_a, Y_c], 1)
Y_a_pred = model_a(X_a)
Y_c_pred = model_c(X_c)
Y_ac_pred = model_ac(X_ac)
loss_a = loss_fn(Y_a_pred, Y_a)
losses_a.append(loss_a.detach().data.numpy())
loss_c = loss_fn(Y_c_pred, Y_c)
losses_c.append(loss_c.detach().data.numpy())
loss_ac = loss_fn(Y_ac_pred, Y_ac)
losses_ac.append(loss_ac.detach().data.numpy())
optimizer_a.zero_grad()
optimizer_c.zero_grad()
optimizer_ac.zero_grad()
loss_a.backward()
loss_c.backward()
loss_ac.backward()
optimizer_a.step()
optimizer_c.step()
optimizer_ac.step()
# + code_folding=[0]
# Sequence distance
test_seq_distances = normalize_seq_distances(np.linspace(0, 50, 50)).astype(np.float32).reshape(-1, 1)
img = model_a(torch.from_numpy(test_seq_distances)).data.numpy()
with plt.rc_context(rc={"font.size": 12}):
fig, ax = plt.subplots(figsize=(14, 2.5))
im = ax.imshow(img.T, aspect=2, extent=[-0.5, 50 - 0.5, 5 - 0.5, - 0.5])
fig.colorbar(im)
plt.xlabel("Sequence distance")
plt.ylabel("One-hot encoding")
# + code_folding=[0]
# Euclidean distance
test_cart_distances = normalize_cart_distances(np.linspace(0, 12, 50)).astype(np.float32).reshape(-1, 1)
img = model_c(torch.from_numpy(test_cart_distances)).data.numpy()
with plt.rc_context(rc={"font.size": 12}):
fig, ax = plt.subplots(figsize=(14, 2.5))
im = ax.imshow(img.T, aspect=12/50*2, extent=[-0.5, 12 - 0.5, 5 - 0.5, - 0.5])
fig.colorbar(im)
plt.xlabel("Euclidean distance")
plt.ylabel("One-hot encoding")
# -
for i in range(13):
# Sequence - Euclidean distance
test_seq_cart_distances = np.hstack([
test_seq_distances,
np.ones((len(test_seq_distances), 1), dtype=np.float32) * normalize_cart_distances(i),
])
img = model_ac(torch.from_numpy(test_seq_cart_distances)).data.numpy()
with plt.rc_context(rc={"font.size": 12}):
fig, ax = plt.subplots(figsize=(14, 5))
im = ax.imshow(img.T, aspect=12/50*8, extent=[-0.5, 50 - 0.5, 11 - 0.5, - 0.5])
fig.colorbar(im)
plt.xlabel("Euclidean distance")
plt.ylabel("One-hot encoding")
plt.show()
# +
torch.save(
model_a.state_dict(),
"/home/kimlab1/database_data/datapkg/adjacency-net-v2/src/model_data/seq_barcode_model.state",
)
torch.save(
model_c.state_dict(),
"/home/kimlab1/database_data/datapkg/adjacency-net-v2/src/model_data/cart_barcode_model.state",
)
torch.save(
model_ac.state_dict(),
"/home/kimlab1/database_data/datapkg/adjacency-net-v2/src/model_data/seq_cart_barcode_model.state",
)
# -
# ### `linear1`
# +
fg, axs = plt.subplots(1, 2, figsize=(14, 4))
axs[0].hist(model_a.linear1.weight.data.numpy().reshape(-1))
axs[1].hist(model_a.linear1.bias.data.numpy().reshape(-1))
print(model_a.linear1.weight.data.numpy().std())
print(model_a.linear1.bias.data.numpy().std())
# +
fg, axs = plt.subplots(1, 2, figsize=(14, 4))
axs[0].hist(model_ac.linear1.weight.data.numpy().reshape(-1))
axs[1].hist(model_ac.linear1.bias.data.numpy().reshape(-1))
print(model_ac.linear1.weight.data.numpy().std())
print(model_ac.linear1.bias.data.numpy().std())
# -
# ### `linear2`
# +
fg, axs = plt.subplots(1, 2, figsize=(14, 4))
axs[0].hist(model_a.linear2.weight.data.numpy().reshape(-1))
axs[1].hist(model_a.linear2.bias.data.numpy().reshape(-1))
print(model_a.linear2.weight.data.numpy().std())
print(model_a.linear2.bias.data.numpy().std())
# +
fg, axs = plt.subplots(1, 2, figsize=(14, 4))
axs[0].hist(model_ac.linear2.weight.data.numpy().reshape(-1))
axs[1].hist(model_ac.linear2.bias.data.numpy().reshape(-1))
print(model_ac.linear2.weight.data.numpy().std())
print(model_ac.linear2.bias.data.numpy().std())
# -
losses_array = np.hstack(losses_ac)
plt.plot(losses_array)
t = torch.eye(10, dtype=torch.float32).unsqueeze(0)
t
mp = nn.MaxPool1d(3, ceil_mode=True)
mp(t)
t.unsqueeze(0).shape
| notebooks2/choosing_value_mapping_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").appName("ProjectOnTransaction_datasets").getOrCreate()
df2 = spark.read.format("csv")\
.option('header', True)\
.option('multiline', True)\
.load("C:\\Users\\Sudip\Desktop\\transaction_data.csv")
df2.show()
df2.take(5)
# count records in datasets
print(f'Record count is: {df2.count()}')
| PySpark- Data Load from CSV file formats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Imports" data-toc-modified-id="Imports-1"><span class="toc-item-num">1 </span>Imports</a></span></li><li><span><a href="#Experiment-environment-/-system-metrics" data-toc-modified-id="Experiment-environment-/-system-metrics-2"><span class="toc-item-num">2 </span>Experiment environment / system metrics</a></span></li><li><span><a href="#Notebook-options" data-toc-modified-id="Notebook-options-3"><span class="toc-item-num">3 </span>Notebook options</a></span></li><li><span><a href="#Source" data-toc-modified-id="Source-4"><span class="toc-item-num">4 </span>Source</a></span><ul class="toc-item"><li><span><a href="#Train-/-Valid-/-Test-lists" data-toc-modified-id="Train-/-Valid-/-Test-lists-4.1"><span class="toc-item-num">4.1 </span>Train / Valid / Test lists</a></span></li></ul></li><li><span><a href="#DataSets-config" data-toc-modified-id="DataSets-config-5"><span class="toc-item-num">5 </span>DataSets config</a></span></li><li><span><a href="#Datasets" data-toc-modified-id="Datasets-6"><span class="toc-item-num">6 </span>Datasets</a></span></li><li><span><a href="#Transforms" data-toc-modified-id="Transforms-7"><span class="toc-item-num">7 </span>Transforms</a></span><ul class="toc-item"><li><span><a href="#Define-Label-remaping" data-toc-modified-id="Define-Label-remaping-7.1"><span class="toc-item-num">7.1 </span>Define Label remaping</a></span></li><li><span><a href="#Define-transforms" data-toc-modified-id="Define-transforms-7.2"><span class="toc-item-num">7.2 </span>Define transforms</a></span></li><li><span><a href="#Apply-transforms" data-toc-modified-id="Apply-transforms-7.3"><span class="toc-item-num">7.3 </span>Apply transforms</a></span></li><li><span><a href="#Points-per-voxel-distribution" data-toc-modified-id="Points-per-voxel-distribution-7.4"><span class="toc-item-num">7.4 </span>Points per voxel distribution</a></span></li></ul></li><li><span><a href="#DataBunch" data-toc-modified-id="DataBunch-8"><span class="toc-item-num">8 </span>DataBunch</a></span><ul class="toc-item"><li><span><a href="#Dataloader-idle-run-speed-measurement" data-toc-modified-id="Dataloader-idle-run-speed-measurement-8.1"><span class="toc-item-num">8.1 </span>Dataloader idle run speed measurement</a></span></li></ul></li><li><span><a href="#Model" data-toc-modified-id="Model-9"><span class="toc-item-num">9 </span>Model</a></span><ul class="toc-item"><li><span><a href="#learner-creation" data-toc-modified-id="learner-creation-9.1"><span class="toc-item-num">9.1 </span>learner creation</a></span></li><li><span><a href="#Learning-Rate-finder" data-toc-modified-id="Learning-Rate-finder-9.2"><span class="toc-item-num">9.2 </span>Learning Rate finder</a></span></li></ul></li><li><span><a href="#Train" data-toc-modified-id="Train-10"><span class="toc-item-num">10 </span>Train</a></span></li><li><span><a href="#Results" data-toc-modified-id="Results-11"><span class="toc-item-num">11 </span>Results</a></span></li><li><span><a href="#Save" data-toc-modified-id="Save-12"><span class="toc-item-num">12 </span>Save</a></span></li><li><span><a href="#Validate" data-toc-modified-id="Validate-13"><span class="toc-item-num">13 </span>Validate</a></span><ul class="toc-item"><li><span><a href="#Load-weights" data-toc-modified-id="Load-weights-13.1"><span class="toc-item-num">13.1 </span>Load weights</a></span></li><li><span><a href="#Validate" data-toc-modified-id="Validate-13.2"><span class="toc-item-num">13.2 </span>Validate</a></span></li></ul></li></ul></div>
# -
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
# # Imports
# +
import os, sys
import time
import numpy as np
import pandas as pd
import glob
from os.path import join, exists, basename, splitext
from pathlib import Path
from matplotlib import pyplot as plt
from matplotlib import cm
import shutil
from functools import partial
from tqdm import tqdm
from joblib import Parallel, delayed, cpu_count
from IPython.display import display, HTML, FileLink
import torch
import torch.nn as nn
import torch.optim as optim
import sparseconvnet as scn
from fastai_sparse import utils, visualize
from fastai_sparse.utils import log
from fastai_sparse.data import DataSourceConfig, MeshesDataset, SparseDataBunch
from fastai_sparse.learner import SparseModelConfig, Learner
from fastai_sparse.callbacks import TimeLogger, SaveModelCallback, CSVLogger
# autoreload python modules on the fly when its source is changed
# %load_ext autoreload
# %autoreload 2
# -
# Importing modules specific to this task (standard 3D semantic markers, ScanNet)
# +
from fastai_sparse.metrics import IouMean
from metrics import IouMeanFiltred
from callbacks import CSVLoggerIouByClass
from data import merge_fn
import transforms as T
# -
import neptune
from neptune_callbacks import NeptuneMonitor
# # Experiment environment / system metrics
# +
params={'n_epoch': 256+128,
'max_lr': 2,
'wd':0.0001
}
params
with open('NEPTUNE_API_TOKEN.txt','r') as f:
NEPTUNE_API_TOKEN = f.readline().splitlines()[0]
neptune.init(api_token=NEPTUNE_API_TOKEN,
project_qualified_name='roma-goodok/fastai-sparse-scannet')
# create experiment in the project defined above
exp = neptune.create_experiment(params=params)
print(exp.id)
exp.append_tag('study')
exp.append_tag('unet24')
exp.append_tag('1cycle')
# -
experiment_name = exp.id
utils.watermark(pandas=True)
# !git log1 -n3
# TODO: write to file versions, and so on
fn_commit = Path('results', experiment_name, 'commit.txt')
fn_commit.parent.mkdir(parents=True, exist_ok=True)
# !git log -n1 --stat > {str(fn_commit)}
# !nvidia-smi
# # Notebook options
utils.wide_notebook()
# uncomment this lines if you want switch off interactive and save visaulisation as screenshoots:
# For rendering run command in terminal: `chromium-browser --remote-debugging-port=9222`
if False:
visualize.options.interactive = False
visualize.options.save_images = True
visualize.options.verbose = True
visualize.options.filename_pattern_image = Path('images', experiment_name, 'fig_{fig_number}')
# # Source
# +
SOURCE_DIR = Path('data', 'scannet_merged_ply')
assert SOURCE_DIR.exists(), "Run prepare_data.ipynb"
definition_of_spliting_dir = Path('data', 'ScanNet_Tasks_Benchmark')
assert definition_of_spliting_dir.exists()
os.listdir(SOURCE_DIR / 'scene0000_01')
# -
# ## Train / Valid / Test lists
#
# MeshesDataset uses the pandas DataFrame as a datasource to determine the splitting into testing/valid as well as files locations.
# +
def find_files(path, ext='merged.ply'):
pattern = str(path / '*' / ('*' + ext))
fnames = glob.glob(pattern)
return fnames
def get_df_list(verbose=0):
# train /valid / test splits
fn_lists = {}
fn_lists['train'] = definition_of_spliting_dir / 'scannetv1_train.txt'
fn_lists['valid'] = definition_of_spliting_dir / 'scannetv1_val.txt'
fn_lists['test'] = definition_of_spliting_dir / 'scannetv1_test.txt'
for datatype in ['train', 'valid', 'test']:
assert fn_lists[datatype].exists(), datatype
dfs = {}
total = 0
for datatype in ['train', 'valid', 'test']:
df = pd.read_csv(fn_lists[datatype], header=None, names=['example_id'])
df = df.assign(datatype=datatype)
df = df.assign(subdir=df.example_id)
df = df.sort_values('example_id')
dfs[datatype] = df
if verbose:
print(f"{datatype:5} counts: {len(df):>4}")
total += len(df)
if verbose:
print(f"total: {total}")
return dfs
# -
df_list = get_df_list(verbose=1)
df_list['valid'].head(10)
# # DataSets config
# You can create MeshDataset using the configuration.
train_source_config = DataSourceConfig(root_dir=SOURCE_DIR,
df_item_list=df_list['train'],
batch_size=32,
num_workers=8,
ply_label_name='label',
file_ext='.merged.ply',
)
train_source_config
valid_source_config = DataSourceConfig(root_dir=SOURCE_DIR,
df_item_list=df_list['valid'],
batch_size=2,
num_workers=2,
ply_label_name='label',
file_ext='.merged.ply',
#init_numpy_random_seed=False,
)
valid_source_config
# # Datasets
# +
train_items = MeshesDataset.from_source_config(train_source_config)
valid_items = MeshesDataset.from_source_config(valid_source_config)
train_items.check()
valid_items.check()
# -
# Let's see what we've done with one example.
#
# In interactive mode, you can switch the color parameters (labels, rgb) to display labels or scan colors.
o = train_items[0]
o.describe()
o.show()
# short representation
train_items
# # Transforms
# ## Define Label remaping
df_classes = pd.read_csv(definition_of_spliting_dir / 'classes_SemVoxLabel-nyu40id.txt', header=None, names=['class_id', 'name'], delim_whitespace=True)
df_classes.T
# Manual definition
assert (np.array([1,2,3,4,5,6,7,8,9,10,11,12,14,16,24,28,33,34,36,39]) == df_classes.class_id).all()
# Map relevant classes to {0,1,...,19}, and ignored classes to -100
remapper = np.ones(3000, dtype=np.int32) * (-100)
for i, x in enumerate([1,2,3,4,5,6,7,8,9,10,11,12,14,16,24,28,33,34,36,39]):
remapper[x] = i
# ## Define transforms
#
# In order to reproduce the [example of SparseConvNet](https://github.com/facebookresearch/SparseConvNet/tree/master/examples/ScanNet), the same transformations have been redone, but in the manner of fast.ai transformations.
#
# The following cells define the transformations: preprocessing (PRE_TFMS); augmentation (AUGS_); and transformation to convert the points cloud to a sparse representation (SPARSE_TFMS). Sparse representation is the input format for the SparseConvNet model and contains a list of voxels and their features
# +
PRE_TFMS = [T.to_points_cloud(method='vertices', normals = False),
T.remap_labels(remapper=remapper, inplace=False),
T.colors_normalize(),
T.normalize_spatial(),
]
_scale = 20
AUGS_TRAIN = [
T.noise_affine(amplitude=0.1),
T.flip_x(p=0.5),
T.scale(scale=_scale),
T.rotate_XY(),
T.elastic(gran=6 * _scale // 50, mag=40 * _scale / 50),
T.elastic(gran=20 * _scale // 50, mag=160 * _scale / 50),
T.specific_translate(full_scale=4096),
T.crop_points(low=0, high=4096),
T.colors_noise(amplitude=0.1),
]
AUGS_VALID = [
T.noise_affine(amplitude=0.1),
T.flip_x(p=0.5),
T.scale(scale=_scale),
T.rotate_XY(),
T.translate(offset=4096 / 2),
T.rand_translate(offset=(-2, 2, 3)), # low, high, dimention
T.specific_translate(full_scale=4096),
T.crop_points(low=0, high=4096),
T.colors_noise(amplitude=0.1),
]
SPARSE_TFMS = [
T.merge_features(ones=False, colors=True, normals=False),
T.to_sparse_voxels(),
]
# train/valid transforms
tfms = (
PRE_TFMS + AUGS_TRAIN + SPARSE_TFMS,
PRE_TFMS + AUGS_VALID + SPARSE_TFMS,
)
# -
# ## Apply transforms
# Now we will apply the transformation to the data sets.
# +
train_items.transform(tfms[0])
pass
valid_items.transform(tfms[1])
pass
# -
# Let's see what we got in results of train and valid tranformations for the first example:
# ##### Train
o = train_items[0]
o.describe()
o.show(colors=o.data['features'])
# ##### Valid
o = valid_items[0]
o.describe()
o.show(colors=o.data['features'])
# ## Points per voxel distribution
# + deletable=false editable=false run_control={"frozen": true}
# # iterations are in one thread
# ppv_valid = []
# for o in tqdm(valid_items, total=len(valid_items)):
# n_voxels = o.num_voxels()
# n_points = len(o.data['coords'])
# ppv_valid.append(n_points / n_voxels)
#
# ppv_train = []
# for o in tqdm(train_items, total=len(train_items)):
# n_voxels = o.num_voxels()
# n_points = len(o.data['coords'])
# ppv_train.append(n_points / n_voxels)
#
# + deletable=false editable=false run_control={"frozen": true}
# fig, ax = plt.subplots()
# ax.hist(ppv_train, bins=100, label='train')
# ax.hist(ppv_valid, bins=100, label='valid')
# ax.legend()
# ax.set_title('Points per voxel distribution')
# ax.set_xlabel('Points per voxel')
# ax.set_ylabel('Number of examples')
# pass
# -
# # DataBunch
# In fast.ai the data is represented DataNunch which contains train, valid and optionally test data loaders.
data = SparseDataBunch.create(train_ds=train_items,
valid_ds=valid_items,
collate_fn=merge_fn)
data.describe()
# ## Dataloader idle run speed measurement
# + deletable=false editable=false run_control={"frozen": true}
# print("cpu_count:", cpu_count())
# !lscpu | grep "Model"
# + deletable=false editable=false run_control={"frozen": true}
# # train
# t = tqdm(enumerate(data.train_dl), total=len(data.train_dl))
# for i, batch in t:
# pass
# + deletable=false editable=false run_control={"frozen": true}
# # valid
# t = tqdm(enumerate(data.valid_dl), total=len(data.valid_dl))
# for i, batch in t:
# pass
# -
# # Model
# U-Net SparseConvNet implemenation ([link](https://github.com/facebookresearch/SparseConvNet/blob/master/examples/ScanNet/unet.py)):
# spatial_size is full_scale
model_config = SparseModelConfig(spatial_size=4096, num_classes=20, num_input_features=3, mode=4,
m=16, num_planes_coeffs=[1, 2, 3, 4, 5, 6, 7])
model_config
# +
class Model(nn.Module):
def __init__(self, cfg):
nn.Module.__init__(self)
self.sparseModel = scn.Sequential(
scn.InputLayer(cfg.dimension, cfg.spatial_size, mode=cfg.mode),
scn.SubmanifoldConvolution(cfg.dimension, nIn=cfg.num_input_features, nOut=cfg.m, filter_size=3, bias=cfg.bias),
scn.UNet(cfg.dimension, cfg.block_reps, cfg.num_planes, residual_blocks=cfg.residual_blocks, downsample=cfg.downsample),
scn.BatchNormReLU(cfg.m),
scn.OutputLayer(cfg.dimension),
)
self.linear = nn.Linear(cfg.m, cfg.num_classes)
def forward(self, xb):
x = [xb['coords'], xb['features']]
x = self.sparseModel(x)
x = self.linear(x)
return x
model = Model(model_config)
# -
utils.print_trainable_parameters(model)
# ## learner creation
# Learner is core fast.ai class which contains model architecture, databunch and optimizer options and implement train loop and prediction
learn = Learner(data, model,
opt_func=partial(optim.SGD),
wd=params['wd'],
true_wd=False,
path=str(Path('results', experiment_name)))
# ## Learning Rate finder
# We use Learning Rate Finder provided by fast.ai library to find the optimal learning rate
# + deletable=false editable=false run_control={"frozen": true}
# learn.lr_find()
# + deletable=false editable=false run_control={"frozen": true}
# learn.recorder.plot()
# -
# # Train
# To visualize the learning process, we specify some additional callbacks
# +
learn.callbacks = []
cb_iouf = IouMeanFiltred(learn, model_config.num_classes)
learn.callbacks.append(cb_iouf)
learn.callbacks.append(TimeLogger(learn))
learn.callbacks.append(CSVLogger(learn))
learn.callbacks.append(CSVLoggerIouByClass(learn, cb_iouf, class_names=list(df_classes.name), filename='iouf_by_class'))
learn.callbacks.append(SaveModelCallback(learn, every='epoch', name='weights', overwrite=True))
learn.callbacks.append(NeptuneMonitor(learn, exp))
# -
learn.fit_one_cycle(cyc_len=params['n_epoch'], max_lr=params['max_lr'],pct_start=0.1, div_factor=1000)
# It takes 512 epochs to get IoU ~ 0.44 by this simple model.
# + deletable=false editable=false run_control={"frozen": true}
# learn.fit(512)
# -
# # Results
learn.recorder.plot()
learn.recorder.plot_losses()
learn.recorder.plot_lr()
learn.recorder.plot_metrics()
cb = learn.find_callback(CSVLoggerIouByClass)
cb.read_logged_file().iloc[-1:].T
# # Save
# Save manually
fn_checkpoint = learn.path / learn.model_dir / 'learn_model.pth'
#torch.save(epoch, 'epoch.pth')
torch.save(model.state_dict(), fn_checkpoint)
# Check saved weights
model.load_state_dict(torch.load(fn_checkpoint))
# # Validate
# ## Load weights
fn_checkpoint = learn.path / learn.model_dir / 'weights.pth'
print(fn_checkpoint)
assert fn_checkpoint.exists()
assert os.path.isfile(fn_checkpoint)
fn_epoch = learn.path / learn.model_dir / 'weights_epoch.pth'
print("Epoch:", torch.load(fn_epoch))
learn.model.load_state_dict(torch.load(fn_checkpoint)['model'])
# ## Validate
# +
# remove callback, because CSVLogger not worked with `learn.validate`
learn.callbacks = []
cb_iouf = IouMeanFiltred(learn, model_config.num_classes)
validation_callbacks = [cb_iouf, TimeLogger(learn)]
# -
np.random.seed(42)
last_metrics = learn.validate(callbacks=validation_callbacks)
last_metrics
# Internal validation details
print(repr(cb_iouf._d['valid']['cm']))
cb_iouf._d['valid']['iou']
cb_iouf._d['valid']['cm']
cb_iouf._d['valid']['iou_per_class']
cb_iouf._d['valid']['iou']
# send some metrics to your experiment
exp.send_metric('valid_iou', cb_iouf._d['valid']['iou'])
exp.stop()
| scannet/neptune_unet_24_exp_06_one_cycle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (text_env)
# language: python
# name: text_env
# ---
# <h3>ローカルでモデルをトレーニングする</h3>
# <h4>エクスポートしたデータを読込みます</h4>
import numpy as np
npz = np.load('docdata1.npz')
print(npz.files)
x = npz['arr_0']
y = npz['arr_1']
# <h4>読込んだ内容を確認します</h4>
print(x.shape)
print(y.shape)
print(x[0])
print(y[0])
# <h4>モデル学習のためのデータ準備をします</h4>
#
# - torch 関連のパッケージをインポートします
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchmetrics.functional import accuracy
import torch.utils.data
from torch.utils.data import DataLoader
# -
# <h4>torch tensor に変換します</h4>
x = torch.tensor(x, dtype=torch.int64)
y = torch.tensor(y, dtype=torch.int64)
print(len(x))
print(x)
print(len(y))
print(y)
print(type(x), x.dtype)
print(type(y), y.dtype)
# <h3>単語ID表現の文章の分散表現を試してみます</h3>
# <h4>参考資料</h4>
#
# - [EMBEDDING](https://pytorch.org/docs/stable/generated/torch.nn.Embedding.html)
# <br></br>
# <br>この例では一つの単語 ID が 10次元のベクトルに変換されてます
sample_embeddings = nn.Embedding(num_embeddings=7295, embedding_dim=10, padding_idx=0)
sample = sample_embeddings(x[0])
print(sample)
print('\n文章のサイズ', len(sample))
# <h3>データセットを作成します</h3>
dataset = torch.utils.data.TensorDataset(x, y)
dataset
len(dataset)
# <h3>トレーニング、検証、テスト、それぞれのデータセットに分割します</h3>
num_train = int(len(dataset) * 0.6)
num_validation = int(len(dataset) * 0.2)
num_test = len(dataset) - num_train - num_validation
torch.manual_seed(0)
train, validation, test = torch.utils.data.random_split(dataset, [num_train, num_validation, num_test])
len(train), len(validation), len(test)
# <h3>Dataloader を作成します</h3>
# <h4>参考資料</h4>
#
# - [TORCH.UTILS.DATA](https://pytorch.org/docs/stable/data.html)
# - [LIGHTNINGDATAMODULE](https://pytorch-lightning.readthedocs.io/en/stable/extensions/datamodules.html)
# - [TORCHTEXT](https://pytorch.org/text/stable/index.html)
# +
batch_size = 128
num_workers = 4
train_dataloader = DataLoader(train, batch_size=batch_size, shuffle=True, num_workers=num_workers)
val_dataloader = DataLoader(validation, batch_size=batch_size, shuffle=False, num_workers=num_workers)
test_dataloader = DataLoader(test, batch_size=batch_size, shuffle=False, num_workers=num_workers)
# -
# <h3>モデル学習に使うクラスを定義します</h3>
# <h4>参考資料</h4>
#
# - [LIGHTNINGMODULE](https://pytorch-lightning.readthedocs.io/en/stable/common/lightning_module.html)
# - [LOGGING](https://pytorch-lightning.readthedocs.io/en/stable/extensions/logging.html)
# - [TORCHMETRICS](https://torchmetrics.readthedocs.io/en/latest/?_ga=2.242351115.847291179.1621688579-221285708.1621323678)
import pytorch_lightning as pl
from pytorch_lightning import Trainer
from pytorch_lightning.callbacks.early_stopping import EarlyStopping
from pytorch_lightning.loggers import MLFlowLogger
# verify version
pl.__version__
class LitTrainClassifier(pl.LightningModule):
def training_step(self, batch, batch_idx):
x, y = batch
y_hat = self.forward(x)
loss = F.cross_entropy(y_hat, y)
self.log('train_loss', loss, on_step=False, on_epoch=True, prog_bar=False, logger=True)
return loss
class LitValidationClassifier(pl.LightningModule):
def validation_step(self, batch, batch_idx):
x, y = batch
y_hat = self.forward(x)
y_pred = torch.argmax(y_hat, dim=1)
loss = F.cross_entropy(y_hat, y)
acc = accuracy(y_pred, y)
self.log('val_loss', loss, on_step=False, on_epoch=True, prog_bar=False, logger=True)
self.log('val_acc', acc, on_step=False, on_epoch=True, prog_bar=False, logger=True)
return loss
class LitTestClassifier(pl.LightningModule):
def test_step(self, batch, batch_idx):
x, y = batch
y_hat = self.forward(x)
y_pred = torch.argmax(y_hat, dim=1)
loss = F.cross_entropy(y_hat, y)
acc = accuracy(y_pred, y)
self.log('test_loss', loss, on_step=False, on_epoch=True, prog_bar=False, logger=True)
self.log('test_acc', acc, on_step=False, on_epoch=True, prog_bar=False, logger=True)
return loss
class LSTMModel(LitTrainClassifier, LitValidationClassifier, LitTestClassifier):
def __init__(self, vocab_size=7295 , embedding_dim=200, hidden_dim=100, layer_dim=2, output_dim=9, drop_out=0.3):
super(LSTMModel, self).__init__()
self.hidden_dim = hidden_dim
self.layer_dim = layer_dim
self.lstm = torch.nn.LSTM(input_size = embedding_dim,
hidden_size = hidden_dim,
num_layers = layer_dim,
dropout = drop_out,
batch_first=True)
self.fc = nn.Linear(hidden_dim, output_dim)
self.embeddings = nn.Embedding(vocab_size, embedding_dim, padding_idx=0)
def forward(self, x):
x = self.embeddings(x)
lstm_out, _ = self.lstm(x)
out = self.fc(lstm_out[:, -1, :])
return out
def configure_optimizers(self):
return torch.optim.Adam(self.parameters(), lr=0.01)
# <h4>モデル学習を開始します</h4>
# +
mlf_logger = MLFlowLogger(
experiment_name="default",
tracking_uri="file:./ml-runs"
)
#torch.backends.cudnn.deterministic = True
#torch.backends.cudnn.benchmark = False
#torch.cuda.empty_cache()
torch.manual_seed(0)
net = LSTMModel()
#trainer = Trainer(gpus=1, max_epochs=20, logger=mlf_logger, callbacks=[EarlyStopping(monitor='val_loss')])
trainer = Trainer(max_epochs=20, logger=mlf_logger, callbacks=[EarlyStopping(monitor='val_loss')])
trainer.fit(net, train_dataloader=train_dataloader, val_dataloaders=val_dataloader)
# -
# <h4>検証データによるメトリックを確認します</h4>
val_metric = trainer.callback_metrics
print(val_metric)
print('val_loss: ', val_metric['val_loss'].item())
print('val_acc: ', val_metric['val_acc'].item())
# <h4>テストデータによる精度を確認します</h4>
test_metric = trainer.test(test_dataloaders=test_dataloader)
metrics = trainer.callback_metrics
print('val_loss: ', metrics['val_loss'].item())
print('val_acc: ', metrics['val_acc'].item())
print('test_loss: ', metrics['test_loss'].item())
print('test_acc: ', metrics['test_acc'].item())
# <h4>モデルを保存します</h4>
os.makedirs('./models', exist_ok=True)
torch.save(net.state_dict(), './models/text_classifier_lstm_local.pt')
| 11_text_classification_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ___Imports___
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from scipy import stats
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.metrics import mean_squared_error, confusion_matrix, r2_score, accuracy_score, recall_score, precision_score, classification_report
from sklearn.pipeline import make_pipeline
# # ___Load Data___
# +
# load data
data = pd.read_csv('../data/ALL_DATA.csv')
print(data.columns)
print(data.head())
print(np.shape(data))
print('\n--------\n')
# Turn into Dataframe
print("Converted to Dataframe: \n")
all_data = pd.DataFrame(data=data)
print(all_data)
# -
# # ___Scaling (10%)___
# +
# *****************************
# HIGHLY IMPORTANT
# *****************************
# Sample data
print("Original Data Stats: \n")
print(all_data.describe())
print('\n--------\n')
print("New Sample Data Stats: \n")
all_data = all_data.sample(frac=0.1) # 10% sample set
print(all_data.describe())
# -
# # ___Target and Feature Variables___
# +
# target variable
target = data.year
# features
features = data.drop(['year'], axis=1)
# print(data.head())
features.columns
# -
# # ___Encoding (One-Hot)___
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
# ### Categorical data to be converted to numeric data
# class_data = list(all_data['class'])
# fall_data = list(all_data['fall'])
class_data = list(features['class'])
fall_data = list(features['fall'])
### integer mapping using LabelEncoder
le = LabelEncoder()
class_encoded = le.fit_transform(class_data)
fall_encoded = le.fit_transform(fall_data)
class_encoded = class_encoded.reshape(len(class_encoded), 1)
fall_encoded = fall_encoded.reshape(len(fall_encoded), 1)
### One hot encoding
onehot_encoder = OneHotEncoder(sparse=False)
onehot_encoded_class = onehot_encoder.fit_transform(class_encoded)
onehot_encoded_fall = onehot_encoder.fit_transform(fall_encoded)
# print(onehot_encoded_class)
# all_data['class'] = onehot_encoded_class
# print(all_data['class'])
# print('\n\n\n')
# print(onehot_encoded_fall)
# all_data['fall'] = onehot_encoded_fall
# print(all_data['fall'])
print(onehot_encoded_class)
features['class'] = onehot_encoded_class
print(features['class'])
print('\n\n\n')
print(onehot_encoded_fall)
features['fall'] = onehot_encoded_fall
print(features['fall'])
# -
# # ___Build Model___
def data_model(data, target): #x,y
X_train, X_test, y_train, y_test = train_test_split(data, target, test_size=0.25, random_state=0)
pipeline = make_pipeline(LogisticRegression())
model = pipeline.fit(X_train, y_train)
return (X_test, y_test, model)
# +
# print("Value distribution of features: ")
# print(list(all_data.iloc[0]))
# min_max = MinMaxScaler()
# data_min_max = min_max.fit_transform(all_data)
# print('\n')
# print("Value distribution after min max: ")
# print(list(data_min_max[0]))
# std = StandardScaler()
# data_std = std.fit_transform(all_data)
# print('\n')
# print("Value distribution after std: ")
# print(list(data_std[0]))
print("Value distribution of features: ")
print(list(features.iloc[0]))
min_max = MinMaxScaler()
data_min_max = min_max.fit_transform(features)
print('\n')
print("Value distribution after min max: ")
print(list(data_min_max[0]))
std = StandardScaler()
data_std = std.fit_transform(features)
print('\n')
print("Value distribution after std: ")
print(list(data_std[0]))
# -
# # ___Model Evaluation___
#Data Variable
# x = all_data[all_data.columns[0]]
x = features
print(x)
#Target Variable
# y = all_data[all_data.columns[1]]
y = target
print(y)
# +
# print("Base:")
# X_test, y_test, model = data_model(x, y)
# prediction = model.predict(X_test)
# print("MSE: {}".format(mean_squared_error(y_test, prediction)))
# print("R Squared: {}".format(r2_score(y_test, prediction)))
# print('\n')
print("MinMax:")
X_test, y_test, model = data_model(data_min_max, y)
prediction = model.predict(X_test)
print("MSE: {}".format(mean_squared_error(y_test, prediction)))
print("R Squared: {}".format(r2_score(y_test, prediction)))
print("Confusion Matrix: {}".format(confusion_matrix(y_test, prediction.round())))
print("Accuracy: {}".format(accuracy_score(y_test, prediction.round(), normalize=False)))
print("Recall Score: {}".format(recall_score(y_test, prediction.round(), average=None)))
print("Precision Score: {}".format(precision_score(y_test, prediction.round(), average=None)))
print("Classification Report: {}".format(classification_report(y_test, prediction.round())))
print('\n')
print("Std:")
X_test, y_test, model = data_model(data_std, y)
prediction = model.predict(X_test)
print("MSE: {}".format(mean_squared_error(y_test, prediction)))
print("R Squared: {}".format(r2_score(y_test, prediction)))
print("Confusion Matrix: {}".format(confusion_matrix(y_test, prediction.round())))
print("Accuracy: {}".format(accuracy_score(y_test, prediction.round(), normalize=False)))
print("Recall Score: {}".format(recall_score(y_test, prediction.round(), average=None)))
print("Precision Score: {}".format(precision_score(y_test, prediction.round(), average=None)))
print("Classification Report: {}".format(classification_report(y_test, prediction.round())))
# -
# # ___Stop here as this model is meant for classification not regression___
| machine-learning/logistic-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cross-Entropy Methods (CEM) on MountainCarContinuous-v0
#
# > In this post, We will take a hands-on-lab of Cross-Entropy Methods (CEM for short) on openAI gym MountainCarContinuous-v0 environment. This is the coding exercise from udacity Deep Reinforcement Learning Nanodegree.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Reinforcement_Learning, PyTorch, Udacity]
# - image: images/MountainCarContinuous-v0.gif
# ## Cross-Entropy Methods (CEM)
# ---
# In this notebook, you will implement CEM on OpenAI Gym's MountainCarContinuous-v0 environment. For summary, The **cross-entropy method** is sort of Black box optimization and it iteratively suggests a small number of neighboring policies, and uses a small percentage of the best performing policies to calculate a new estimate.
#
# ### Import the Necessary Packages
# +
import gym
import math
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import base64, io
# For visualization
from gym.wrappers.monitoring import video_recorder
from IPython.display import HTML
from IPython import display
import glob
# -
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
# ### Instantiate the Environment and Agent
#
# MountainCar environment has two types: Discrete and Continuous. In this notebook, we used Continuous version of MountainCar. That is, we can move the car to the left (or right) precisely.
# +
env = gym.make('MountainCarContinuous-v0')
env.seed(101)
np.random.seed(101)
print('observation space:', env.observation_space)
print('action space:', env.action_space)
print(' - low:', env.action_space.low)
print(' - high:', env.action_space.high)
# -
# ### Define Agent
class Agent(nn.Module):
def __init__(self, env, h_size=16):
super(Agent, self).__init__()
self.env = env
# state, hidden layer, action sizes
self.s_size = env.observation_space.shape[0]
self.h_size = h_size
self.a_size = env.action_space.shape[0]
# define layers (we used 2 layers)
self.fc1 = nn.Linear(self.s_size, self.h_size)
self.fc2 = nn.Linear(self.h_size, self.a_size)
def set_weights(self, weights):
s_size = self.s_size
h_size = self.h_size
a_size = self.a_size
# separate the weights for each layer
fc1_end = (s_size * h_size) + h_size
fc1_W = torch.from_numpy(weights[:s_size*h_size].reshape(s_size, h_size))
fc1_b = torch.from_numpy(weights[s_size*h_size:fc1_end])
fc2_W = torch.from_numpy(weights[fc1_end:fc1_end+(h_size*a_size)].reshape(h_size, a_size))
fc2_b = torch.from_numpy(weights[fc1_end+(h_size*a_size):])
# set the weights for each layer
self.fc1.weight.data.copy_(fc1_W.view_as(self.fc1.weight.data))
self.fc1.bias.data.copy_(fc1_b.view_as(self.fc1.bias.data))
self.fc2.weight.data.copy_(fc2_W.view_as(self.fc2.weight.data))
self.fc2.bias.data.copy_(fc2_b.view_as(self.fc2.bias.data))
def get_weights_dim(self):
return (self.s_size + 1) * self.h_size + (self.h_size + 1) * self.a_size
def forward(self, x):
x = F.relu(self.fc1(x))
x = torch.tanh(self.fc2(x))
return x.cpu().data
def act(self, state):
state = torch.from_numpy(state).float().to(device)
with torch.no_grad():
action = self.forward(state)
return action
def evaluate(self, weights, gamma=1.0, max_t=5000):
self.set_weights(weights)
episode_return = 0.0
state = self.env.reset()
for t in range(max_t):
state = torch.from_numpy(state).float().to(device)
action = self.forward(state)
state, reward, done, _ = self.env.step(action)
episode_return += reward * math.pow(gamma, t)
if done:
break
return episode_return
# ### Cross Entropy Method
def cem(agent, n_iterations=500, max_t=1000, gamma=1.0, print_every=10, pop_size=50, elite_frac=0.2, sigma=0.5):
"""PyTorch implementation of the cross-entropy method.
Params
======
Agent (object): agent instance
n_iterations (int): maximum number of training iterations
max_t (int): maximum number of timesteps per episode
gamma (float): discount rate
print_every (int): how often to print average score (over last 100 episodes)
pop_size (int): size of population at each iteration
elite_frac (float): percentage of top performers to use in update
sigma (float): standard deviation of additive noise
"""
n_elite=int(pop_size*elite_frac)
scores_deque = deque(maxlen=100)
scores = []
# Initialize the weight with random noise
best_weight = sigma * np.random.randn(agent.get_weights_dim())
for i_iteration in range(1, n_iterations+1):
# Define the cadidates and get the reward of each candidate
weights_pop = [best_weight + (sigma * np.random.randn(agent.get_weights_dim())) for i in range(pop_size)]
rewards = np.array([agent.evaluate(weights, gamma, max_t) for weights in weights_pop])
# Select best candidates from collected rewards
elite_idxs = rewards.argsort()[-n_elite:]
elite_weights = [weights_pop[i] for i in elite_idxs]
best_weight = np.array(elite_weights).mean(axis=0)
reward = agent.evaluate(best_weight, gamma=1.0)
scores_deque.append(reward)
scores.append(reward)
torch.save(agent.state_dict(), 'checkpoint.pth')
if i_iteration % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(i_iteration, np.mean(scores_deque)))
if np.mean(scores_deque)>=90.0:
print('\nEnvironment solved in {:d} iterations!\tAverage Score: {:.2f}'.format(i_iteration-100, np.mean(scores_deque)))
break
return scores
# ### Run
agent = Agent(env).to(device)
scores = cem(agent)
# ### Plot the learning progress
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# ### Animate it with Video
# +
def show_video(env_name):
mp4list = glob.glob('video/*.mp4')
if len(mp4list) > 0:
mp4 = 'video/{}.mp4'.format(env_name)
video = io.open(mp4, 'r+b').read()
encoded = base64.b64encode(video)
display.display(HTML(data='''<video alt="test" autoplay
loop controls style="height: 400px;">
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii'))))
else:
print("Could not find video")
def show_video_of_model(agent, env_name):
env = gym.make(env_name)
vid = video_recorder.VideoRecorder(env, path="video/{}.mp4".format(env_name))
agent.load_state_dict(torch.load('checkpoint.pth'))
state = env.reset()
done = False
while not done:
vid.capture_frame()
action = agent.act(state)
next_state, reward, done, _ = env.step(action)
state = next_state
if done:
break
env.close()
# -
agent = Agent(env).to(device)
show_video_of_model(agent, 'MountainCarContinuous-v0')
show_video('MountainCarContinuous-v0')
# > Note: While I tried to execute the model with VideoRecorder in Linux, it doesn't show correctly. But in windows, it works! Perhaps, It may be bug in VideoRecoder.
| _notebooks/2021-05-11-CEM-MountainCar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# # Build Classification Models
import pandas as pd
cuisines_df = pd.read_csv("../../data/cleaned_cuisines.csv")
cuisines_df.head()
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import accuracy_score,precision_score,confusion_matrix,classification_report, precision_recall_curve
from sklearn.svm import SVC
import numpy as np
cuisines_label_df = cuisines_df['cuisine']
cuisines_label_df.head()
cuisines_feature_df = cuisines_df.drop(['Unnamed: 0', 'cuisine'], axis=1)
cuisines_feature_df.head()
X_train, X_test, y_train, y_test = train_test_split(cuisines_feature_df, cuisines_label_df, test_size=0.3)
# +
lr = LogisticRegression(multi_class='ovr',solver='liblinear')
model = lr.fit(X_train, np.ravel(y_train))
accuracy = model.score(X_test, y_test)
print ("Accuracy is {}".format(accuracy))
# -
# test an item
print(f'ingredients: {X_test.iloc[50][X_test.iloc[50]!=0].keys()}')
print(f'cuisine: {y_test.iloc[50]}')
# +
#rehsape to 2d array and transpose
test= X_test.iloc[50].values.reshape(-1, 1).T
# predict with score
proba = model.predict_proba(test)
classes = model.classes_
# create df with classes and scores
resultdf = pd.DataFrame(data=proba, columns=classes)
# create df to show results
topPrediction = resultdf.T.sort_values(by=[0], ascending = [False])
topPrediction.head()
# -
y_pred = model.predict(X_test)
print(classification_report(y_test,y_pred))
| 4-Classification/2-Classifiers-1/solution/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spam text Message Classification using Bag of Words:
#
# #### <NAME>
# imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid")
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
data = pd.read_csv('spam.tsv',sep='\t')
data.head() #length indicates character length
data.isnull().sum()
print("No of Data Samples",len(data))
data['label'].value_counts()
# #### The data is imbalanced! This is a scenario where the number of observations belonging to one class is significantly lower than those belonging to the other classes.In order to get better accuracy,let's balance the data.
ham=data[data['label']=='ham']
ham.head()
spam=data[data['label']=='spam']
spam.head()
# Randomly select 747 samples in ham to balance the data
ham=ham.sample(747)
ham.shape,spam.shape
#Append spam and ham
data = ham.append(spam,ignore_index=True)
data.shape
# ## Exploratory Data Analysis:
sns.pairplot(data, hue='label')
# #### Generally,character length of spam messages is greater than that of ham messages.
data.head()
# Train-test split:
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data['message'], data['label'], test_size = 0.3,
random_state=0, shuffle = True, stratify=data['label'])
X_train
y_train
X_test
y_test
# ## Bag of words model:
from sklearn.feature_extraction.text import TfidfVectorizer
# +
###vectorizer = TfidfVectorizer()
##X_train_vector = vectorizer.fit_transform(X_train)
# +
### X_train_vector.shape
# +
### X_train_vector
# -
# ### Pipeline with Randomforest classifier:
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
# Machine learning (ML) pipelines consist of several steps to train a model.Machine learning pipelines are cyclical and iterative as every step is repeated to continuously improve the accuracy of the model and achieve a successful algorithm
clf = Pipeline([('tfidf', TfidfVectorizer()), ('clf', RandomForestClassifier(n_estimators=100, n_jobs=-1))])
#njobs=-1 will use all the cores of CPU
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix
confusion_matrix(y_test, y_pred)
print(classification_report(y_test, y_pred))
accuracy_score(y_test, y_pred)
clf.predict(["IMPORTANT - You could be entitled up to £3,160 in compensation from mis-sold PPI on a credit card or loan. Please reply PPI for info or STOP to opt out."])
clf.predict(["you have won tickets to KGF 2 this summer."])
# ###### This is a spam :(
clf.predict(["My name is thiru"])
# ### Support Vector Machines:
from sklearn.svm import SVC
clf = Pipeline([('tfidf', TfidfVectorizer()), ('clf', SVC(C = 1000, gamma = 'auto'))])
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
confusion_matrix(y_test, y_pred)
print(classification_report(y_test, y_pred))
accuracy_score(y_test, y_pred)
clf.predict(["My name is thiru"])
clf.predict(["You have won tickets to KGF 2 this summer"])
# ### Naive bayes:
from sklearn.naive_bayes import MultinomialNB
clf = Pipeline([('tfidf', TfidfVectorizer()), ('clf', MultinomialNB(alpha=1))])
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
confusion_matrix(y_test, y_pred)
print(classification_report(y_test, y_pred))
accuracy_score(y_test, y_pred)
clf.predict(["You have won tickets to KGF 2 this summer"])
| Spam text Message Classification.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Further information
#
# ## How to pronounce the double underscore?
#
# The double underscore used in magic methods like `__init__` or `__repr__` is
# pronounced "dunder".
#
# ## What is the `self` variable for?
#
# In methods the first variable is used to refer to the instance of a given class.
# It is conventional to use `self`.
#
# As an example let us consider this class:
class PetDog:
"""
A class for a Pet.
Has two methods:
- `bark` which returns "Woof" as a string.
- `give_toy` which gives a toy to the dog in question. This updates the
`toys` attribute.
"""
def __init__(self):
self.toys = []
def bark(self):
"""
Returns the string Woof.
"""
return "Woof"
def give_toy(self, toy):
"""
Updates the instances toys list.
"""
self.toys.append(toy)
# If we now create two dogs:
auraya = PetDog()
riggins = PetDog()
# Both have no toys:
auraya.toys
riggins.toys
# Now when we want to give `riggins` a toy we need to specify which of those two
# empty lists to update:
riggins.give_toy("ball")
riggins.toys
# However `auraya` still has no toys:
auraya.toys
# When running `riggins.give_toy("ball")`, internally the `give_toy` method is
# taking `self` to be `riggins` and so the
# line `self.toys.append(toy)` in fact is running as `riggins.toys.append(toy)`.
#
# The variable name `self` is a convention and not a functional requirement.
# If we modify it (using {ref}`inheritance <how_to_use_inheritance>`):
class OtherPetDog(PetDog):
"""
A class for a Pet.
Has two methods:
- `bark` which returns "Woof" as a string.
- `give_toy` which gives a toy to the dog in question. This updates the
`toys` attribute.
"""
def give_toy(the_dog_in_question, toy):
"""
Updates the instances toys list.
"""
the_dog_in_question.toys.append(toy)
# Then we get the same outcome:
riggins = OtherPetDog()
riggins.toys
riggins.give_toy("ball")
riggins.toys
# Indeed the line `the_dog_in_question.toys.append(toy)` is run as
# `riggins.toys.append(toy)`.
#
# You should however use `self` as it is convention and helps with readability of
# your code.
#
# ## Why do we use `CamelCase` for classes but `snake_case` for functions?
#
# This is specified by the Python convention:
# <https://www.python.org/dev/peps/pep-0008/>
#
# These conventions are important as it helps with readability of code.
#
# ## What is the difference between a method and a function?
#
# A method is a function defined on a class and always takes a first argument
# which is the specific instance from which the method is called.
#
# ## Other resources on objects
#
# - A non programmers tutorial for Python: Object Oriented Programming:
# <https://en.wikibooks.org/wiki/Non-Programmer%27s_Tutorial_for_Python_3/Intro_to_Object_Oriented_Programming_in_Python_3>
# - A tutorial based on a role playing game:
# <https://inventwithpython.com/blog/2014/12/02/why-is-object-oriented-programming-useful-with-a-role-playing-game-example/>
# - A video: <https://www.youtube.com/watch?v=trOZBgZ8F_c>
| book/building-tools/03-objects/why/.main.md.bcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hhx4vnU-qiGj" colab_type="text"
# # **Natural Language Processing**
# by **<NAME>**
#
# # **Part 3: Creating Urdu NER**
#
# + [markdown] id="v9Hltk54qpOG" colab_type="text"
#
#
# **Reading Data from the Google Drive**
#
#
# + id="6gq5T7CCsB0I" colab_type="code" outputId="390c3db4-339c-43e2-c914-11e9cca339f6" executionInfo={"status": "ok", "timestamp": 1581579156250, "user_tz": -300, "elapsed": 68122, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 125}
from google.colab import drive
# This will prompt for authorization.
drive.mount('/content/drive')
# + [markdown] id="4OqN2wfjAlyr" colab_type="text"
#
# + [markdown] id="1Slm21kLsE7N" colab_type="text"
# **Note about the data:**
# ```
# The dataset is a chunk of the dataset retrieved from
# http://ltrc.iiit.ac.in/ner-ssea-08/index.cgi?topic=5
# (Workshop on NER for South and South East Asian Languages in IJCNLP 2008 at Hyderabad, India)
#
# The data is reformatted and enriched. Please refer/acknowledge in your paper/report to the original source, if you use this data.
# ```
#
#
# + id="I4NaatT9sN6m" colab_type="code" outputId="d71e5427-7f57-42a1-9be1-3f6e88ee3520" executionInfo={"status": "ok", "timestamp": 1581579494714, "user_tz": -300, "elapsed": 1256, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
import csv
with open('/content/drive/My Drive/CLT20/NER-Dataset/conll-ner.csv', encoding = 'utf-8') as csvfile:
data = list(csv.reader(csvfile, delimiter=','))
print(data[0])
# + [markdown] id="SSLCvwEVBcp8" colab_type="text"
#
#
# ```
# The format of the above output is:
# word-no, word, lemma/root-word, universal-part-of-speech, other-part-of-speech,empty, IBO-tag, Entity-type
#
# The first word of a named entity is tagged as B(eginning),
# the other words of the named entity are tagged as I(ntermediate),
# and the words not belonging to named entity are tagged as O(ther)
#
# For example
# Bill ... B Person
# Gates ... I Person
# founded ... O
# Microsoft ... B Organization
#
# ```
#
#
#
# + [markdown] id="J8u6EGRG-C-w" colab_type="text"
# Extracting features
# + id="Rf3LyYD_-C-x" colab_type="code" outputId="8184ef61-d987-466b-86f7-3a3bb94cde7d" executionInfo={"status": "ok", "timestamp": 1581583234692, "user_tz": -300, "elapsed": 819, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
def extract_features(words, i):
wid = words[i][0]
token = words[i][1]
upos = words[i][3]
xpos = words[i][4]
prev_token = ""
prev_upos = ""
prev_xpos = ""
next_token = ""
next_upos = ""
next_xpos = ""
if int(wid) != 1:
prev_token = words[i-1][1]
prev_upos = words[i-1][3]
prev_xpos = words[i-1][4]
if i < len(words)-1:
if int(wid) < int(words[i+1][0]):
next_token = words[i+1][1]
next_upos = words[i+1][3]
next_xpos = words[i+1][4]
is_number = False
try:
if float(token):
is_number = True
except:
pass
features_dict = {"token": token
# , "upos": upos
# , "xpos": xpos
, "prev_token": prev_token
# , "prev_upos": next_upos
# , "prev_xpos": next_xpos
, "next_token": next_token
# , "next_upos": next_upos
# , "next_xpos": next_xpos
, "is_number": is_number}
return features_dict
print(data[3:6])
print(extract_features(data, 4))
# + [markdown] id="NjvxI4sRBFXR" colab_type="text"
# Converting feature vector for each word
# + id="5yVk3_Ur-C-1" colab_type="code" outputId="6fd9a326-c064-47fc-fc36-74f3e98ec6de" executionInfo={"status": "ok", "timestamp": 1581583235300, "user_tz": -300, "elapsed": 1414, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
X_features = []
Y = []
for i in range(len(data)):
try:
X_features.append(extract_features(data, i))
Y.append(data[i][6])
except:
pass
print(len(X_features),":",len(Y))
# + id="6sE1Uhtkn1t-" colab_type="code" outputId="0feec8af-6b0f-47e7-8b6d-23f3d6d37892" executionInfo={"status": "ok", "timestamp": 1581583235302, "user_tz": -300, "elapsed": 1406, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 52}
print(X_features[4:6])
print(Y[4:6])
# + [markdown] id="3VZCMYOTEoVc" colab_type="text"
#
#
# ```
# # This is formatted as code
# ```
#
#
# + [markdown] id="TNyTE89OEbc-" colab_type="text"
# Currently many features have string data, we convert it into numeric vectors with a column for each string
# + id="65eAJJUG-C-9" colab_type="code" outputId="34cfd66e-60a8-4dd8-9cf5-a99def70d1a2" executionInfo={"status": "ok", "timestamp": 1581583236043, "user_tz": -300, "elapsed": 2138, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 105}
from sklearn.feature_extraction import DictVectorizer
vectoriser = DictVectorizer(sparse=True)
X = vectoriser.fit_transform(X_features)
print("Shape of the matrix: ", X.get_shape())
print(X[3])
# + [markdown] id="SaIhQp6zgIbz" colab_type="text"
# #Loding the Word2Vec Model
# import gensim
# model = gensim.models.Word2Vec.load("/content/drive/My Drive/CLT20/urdu-w2vec")
# + [markdown] id="dkijmB7PJ2zV" colab_type="text"
# #adding word embedding features
# import numpy
#
# lm = []
# Yf = []
#
# e = 0
# for i in range(len(X)):
# try:
# m = model[X_features[i]['token']]
# #Xv[i] = numpy.append(X[i],m)
# lm.append(numpy.append(X[i],m))
# Yf.append(Y[i])
# except:
# #print(X_features[i]['token'], Y[i])
# e = e + 1
#
# print(e)
#
# X = numpy.array(lm)
# Y = Yf
#
#
# print("number of features: ", len(X[3]))
# print(X[3])
# + [markdown] id="jYXiEeNc-C_B" colab_type="text"
# **Making training set and training the classifier**
# + id="qPbSzUHy-C_C" colab_type="code" outputId="4f5c4a19-fa04-47f9-8b20-23174586dcc0" executionInfo={"status": "ok", "timestamp": 1581583236048, "user_tz": -300, "elapsed": 2132, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
from sklearn import model_selection
from sklearn import svm # import support vector machine
cl = svm.LinearSVC()
validation_size = 0.20
seed = 7
X_train, X_test, Y_train, Y_test = model_selection.train_test_split(X, Y, stratify = Y, test_size=validation_size, random_state=seed)
cl.fit(X_train, Y_train)
# + id="bUt01to4-C_G" colab_type="code" outputId="14ab5700-a3b5-4679-a5df-bb911205b768" executionInfo={"status": "ok", "timestamp": 1581583236050, "user_tz": -300, "elapsed": 2124, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print(cl.predict(X_test[4:7]))
# + id="uWOC-r_Mk1Pq" colab_type="code" colab={}
# + id="1Vs6P8_n-C_L" colab_type="code" outputId="1cd7d97a-00da-4c18-f12d-664cc30f39d4" executionInfo={"status": "ok", "timestamp": 1581583236055, "user_tz": -300, "elapsed": 2111, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
print(Y_test[4:7])
# + id="O5h-rMQT-C_P" colab_type="code" outputId="81af30a0-a472-4d07-a39e-2022b0fdd5ac" executionInfo={"status": "ok", "timestamp": 1581583236058, "user_tz": -300, "elapsed": 2104, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 565}
from sklearn import metrics
X_predict = cl.predict(X_test)
print(metrics.classification_report(X_predict , Y_test))
print(metrics.confusion_matrix(X_predict , Y_test))
# + [markdown] id="LxoxhoWQrqGV" colab_type="text"
# **Exercise 1:** Enable the PoSTag attributes in extract_features, then train train and evaluate the model.
#
# **Exercise 2:** Enable the the Word Embedding Code, then train and evaluate the model.
#
# **Exercise 3:** Include I and B subtags in the target tag, then run the experiment(s) again. (Hint: The Y would have data[i][6] + "-" + data[i][7])
#
# **(further) Exercises:** create other features, then traina and evaluate the model.
# e.g. length of token, 3-letter suffix, gazetter lists etc,
#
# + [markdown] id="flolGWwXqFco" colab_type="text"
# K-fold Cross Validation
# + [markdown] id="HcfGqNPkros-" colab_type="text"
#
# + id="fupXl0PxqEiu" colab_type="code" outputId="c624273e-5e52-42e8-c771-d576ec45a613" executionInfo={"status": "ok", "timestamp": 1581583237105, "user_tz": -300, "elapsed": 3142, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mDN7x3VHp1GGTsALx2iUyEcW7dR3n19shLRfU0Pkg=s64", "userId": "14391780310186458281"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.model_selection import cross_val_score
clf = svm.LinearSVC()
scores = cross_val_score(clf, X, Y, cv=5)
print(scores)
| CLT20 ws3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.cm import coolwarm
from mpl_toolkits.mplot3d import Axes3D
from scipy.ndimage import correlate as im_corr
from scipy.stats import poisson
plt.rcParams['font.size'] = 15
plt.rcParams['axes.spines.right'] = False
plt.rcParams['ytick.right'] = False
plt.rcParams['axes.spines.top'] = False
plt.rcParams['xtick.top'] = False
plt.rcParams['lines.linewidth'] = 2
plt.rcParams['lines.markersize'] = 10
# ### Useful function definitions
# +
from numpy.lib import stride_tricks
# Useful functions
H = lambda x: -np.sum(x[x>0]*np.log2(x[x>0]))
def add_fake_dimension(org_ndarray, win_size):
""" Rolls a window over a vector and extracts the content within the window
Stride_tricks only affect the shape and strides in the array interface.
The memory footprint is therefore equal for both org_ndarray and
fake_ndarray.
Important!!!
The windowed dimension in X must be along the first dimension (axis=0)
Args:
org_ndarray: vector to roll the window over
win_size: window size in vector elements
Returns:
fake_ndarray:
Raises:
"""
n_element = org_ndarray.size
element_size = org_ndarray.itemsize
input_dims = org_ndarray.shape
stride_length = 1
for dims in input_dims[1:]:
stride_length *= dims
org_1darray = org_ndarray.ravel()
shape = (n_element/stride_length - win_size + 1, win_size*stride_length)
strides = (stride_length*element_size, element_size)
fake_2darray = stride_tricks.as_strided(org_1darray, shape=shape, strides=strides)
new_shape = [shape[0], win_size]
for dims in input_dims[1:]:
new_shape.append(dims)
fake_ndarray = fake_2darray.reshape(new_shape)
return fake_ndarray
def getBiasCorrection(words):
""" Returns a bias corrected entropy
See Part 2, entropy and bias correction
Args:
words: 2D array of code words with each row representing one word
Returns:
biasCorrectedEntropy:
Raises:
"""
nReps = 5
nSamples, wordLength = words.shape
fracs = np.logspace(np.log10(0.25), np.log10(1), 5)
entropies = []
for frac in fracs:
nSubSamples = np.int64(np.round(frac*nSamples))
entropyTmp = 0
for rep in range(nReps):
permutation = np.random.permutation(nSamples)
uniqueWords, wordCounts = np.unique(words[permutation[0:nSubSamples], :], axis=0, return_counts=True)
wordProb = wordCounts / np.float64(wordCounts.sum())
entropyTmp += H(wordProb)
entropies.append(entropyTmp/nReps)
# Quadratic fit
A = np.vstack([np.ones(fracs.size), 1./fracs, 1./fracs**2])
w = np.linalg.lstsq(A.T, entropies)[0]
correction = -w[1:].sum()
return correction
def getValidEntropies(entropies, biasCorrections, threshold=0.02):
""" Returns valid entropy estimates
Check which entropies that are valid based on the size of the correction
Args:
entropies: raw entropy estiamtes
biasCorrections: bias corrections
threshold: threshold fraction for determining valid entropies
Returns:
validEntropies: boolean array with valid entries
Raises:
"""
return biasCorrections/entropies < threshold
def getInfiteLengthExtrapolation(entropies, biasCorrections, wordLengths, nIgnore=0):
""" Returns the extrapoalted entropy for an infinite word length
Args:
entropies: raw entropy estiamtes
biasCorrections: bias corrections
wordLengths:
nIgnore: ignore the n smallest word lengths
Returns:
A: linear regression matrix
w: parameters for the linear extrapolation
Raises:
"""
validEntropies = getValidEntropies(entropies, biasCorrections)
A = np.vstack([np.ones(validEntropies.sum()), 1./wordLengths[validEntropies]])
b = (entropies+biasCorrections)[validEntropies]
w = np.linalg.lstsq(A[:, nIgnore:].T, b[nIgnore:])[0]
A = np.hstack([A, np.array([[1], [0]])])
return A, w
# -
# ### Quantifying the amount of information in a spike train
# Neurons communicate through spikes and their spike trains thus convey information, but how much? The concepts of entropy and mutual information can easily be extended to answer this question, by focusing on entropy rates instead of just entropy. For example, if you obtain one hundred independent samples per second from a distribution with an entropy of 1 bit, then the entropy rate is 100 bits/s.
#
# Quantifying the information rates of spike trains thus relies on binning these into vectors of spike counts. The entropy rate is then defined as the entropy of the distribution from which the spike count vector is drawn divided by the duration of time over which the vector was obtained. In the simplest of cases, where no temporal correlations are present, it is enough to just look at the entropy of the one-dimensional distribution underlying the counts in each bin. However, if temporal correlations are present, then one needs to determine the entropy of the multidimensional distribution from which longer spike count vectors are drawn, as we will see further down.
# ### Entropy of Poisson spike trains (theoretical)
# Initially, it is a good idea to first try to get an impression of how the firing rate and the binning width affect the entropy rate. To keep things simple, we assume that spikes are drawn from a Poisson distribution and that the spike count in each bin is independent. With these simplifications, the entropy rate is simply the entropy for a Poisson distribution divided by the bin width. As shown below, the entropy rate increases as a function of both the firing rate and the bin width. Thus indicating that a neurons ability to transmit information increases with higher firing rates and with a higher temporal precision.
# +
nVals = 21
firingRates = np.linspace(0, 100, nVals) # spikes per second (Hz)
binWidths = np.linspace(0.002, 0.02, nVals) # bin width (s)
firingRatesGrid, binWidthsGrid = np.meshgrid(firingRates, binWidths)
# Determine entropy rates
entropyRates = np.zeros(firingRatesGrid.shape)
for i, rate, width in zip(np.arange(nVals*nVals), firingRatesGrid.ravel(), binWidthsGrid.ravel()):
muRate = rate*width # Mean count within each bin corresponding to the mean firing rate
maxVal = muRate + 10*np.sqrt(muRate) # Upper count limit after which the probability is essentially zero
p = poisson.pmf(np.arange(maxVal), muRate)
entropyRates.ravel()[i] = H(p)/width
# Plotting
fig = plt.figure(figsize=(10, 6))
ax = fig.gca(projection='3d')
surf = ax.plot_surface(firingRatesGrid, binWidthsGrid, entropyRates, cmap=coolwarm)
ax.set_xticks(np.linspace(0, 100, 6))
ax.set_xlabel('Firing rate (Hz)', labelpad=10)
ax.set_yticks(np.linspace(0.002, 0.02, 2))
ax.set_ylabel('Bin width (ms)', labelpad=10)
ax.set_zlabel('Entropy rate (bits/s)', labelpad=5)
ax.view_init(20, 140)
# -
# ### Entropy of Poisson spike trains (empirical)
# Real spike trains obviously contain some temporal correlations. We therefore need a method for extracting entropy rates that corrects for any such temporal correlations. Strong et al. (1998) presented such a method for neural spike trains and this method appears to have been used in essentially all studies since. The basic idea is to determine entropy rates for spike count vectors (also called words) of increasing length, and then correct for temporal correlations by extrapolating to an infinite word length. To begin with, we will skip the extrapolation step and instead just familiarize us with the method by looking at its output for the same independent Poisson case as above, but with fixed firing rate and bin width. In short, we will generate a long vector of spike counts and subsequently try to empirically estimate the entropy of the probability distributions for words of varying length.
# +
# Parameters
rate = 30
width = 0.01
nSamples = 10000;
wordLengths = np.arange(1, 16);
# Get the theoretical entropy rate
muRate = rate*width # Mean count within each bin corresponding to the mean firing rate
maxVal = muRate + 10*np.sqrt(muRate) # Upper count limit after which the probability is essentially zero
p = poisson.pmf(np.arange(maxVal), muRate)
theoreticalEntropy = H(p)/width
# Generate a sequence of Poisson data
r = np.random.poisson(np.ones(nSamples)*rate*width)
# Calculate raw and bias corrected entropies for all word lengths
entropies = np.zeros(wordLengths.size)
biasCorrections = np.zeros(wordLengths.size)
for i, wordLength in enumerate(wordLengths):
words = add_fake_dimension(r, wordLength)
uniqueWords, wordCounts = np.unique(words, axis=0, return_counts=True)
wordProb = wordCounts / np.float64(wordCounts.sum())
entropies[i]= H(wordProb)/width/wordLength
biasCorrections[i] = getBiasCorrection(words)/width/wordLength
validEntropies = getValidEntropies(entropies, biasCorrections, threshold=0.02)
print('Theoretical entropy: %2.2f') % theoreticalEntropy
# Plot empirical entropy rates as a function of word length
fig = plt.figure(figsize=(10, 4))
ax = plt.axes()
ax.plot(1./wordLengths, entropies, 'bo-', label='$H_\mathrm{raw}$')
ax.plot(1./wordLengths, entropies+biasCorrections, 'r.-', label='$H_\mathrm{corrected}$')
ax.plot(1./wordLengths[validEntropies], (entropies+biasCorrections)[validEntropies], 'ro')
ax.axhline(theoreticalEntropy, ls=':', color='k')
ax.set_ylim([0, 1.1*theoreticalEntropy])
ax.set_xlim([0, 1])
ax.set_xlabel('(Word length)$^{-1}$')
ax.set_ylabel('Entropy rate (bits/s)');
ax.legend(frameon=False, bbox_to_anchor=(1, 0, 0.4, 1), loc=5);
# -
# The main observation from the figure above is that the estimated entropy rate is essentially identical to the theoretical one for word lengths up to 6-7 bins. For longer words, finite sample effects start affecting our estimates, and the bias gets more severe as the word length increases (more and more data is needed when estimating entropies of higher dimensional distributions). The bias correction mechanism explained in Part 2 can alleviate the problem to some extent, but the improvement is quite modest (compare the red to the blue line above). However, the bias correction mechanism also provides us with a way of checking when finite sample effects start affecting our entropy estimates significantly. The larger red dots above, for example, denotes cases when the correction is predicted to be less than 2%, and thus sorts out the cases where we can reasonable sure that we have a good estimate.
#
# In summary, we thus see that the entropy estimates for words of varying length are identical whenever no temporal correlations are present (as expected).
# ### Mutual information, noiseless transmission
# Next, we will look at an example with temporal correlations. More precisely, we assume that a light sensitive cell is being stimulated by a binary stimulus sequence $\mathbf{x}$, where each element $x$ is drawn from a uniform binary distribution $X\sim U\{0, 1\}$. The cell performs a temporal summation over time and outputs $y_i=\mathbf{w}^T\mathbf{x}_i$ spikes in bin $i$, with $\mathbf{w}$ being a temporal filter and $\mathbf{x}_i$ the recent stimulus history up to time point $i$. We further assume that the output is deterministic, meaning that the noise entropy is zero and that the information rate (mutual information) simply equals the entropy rate.
# +
width = 0.01
nSamples = 10000
w = np.array([1, 2, 4])
wordLengths = np.arange(1, 15);
# Generate a random binary sequence and calculate the spike count in each bin
x = (np.sign(np.random.randn(nSamples))+1) / 2
y = im_corr(x, w, mode='constant', origin=(w.size-1)/2)
# Noiseless transmission -> theoretical I same as the entropy of the binary stimulus x
theoreticalEntropy = 1./width
# Calculate raw and bias corrected entropies for various word lengths
entropies = np.zeros(wordLengths.size)
biasCorrections = np.zeros(wordLengths.size)
for i, wordLength in enumerate(wordLengths):
words = add_fake_dimension(y, wordLength)
uniqueWords, wordCounts = np.unique(words, axis=0, return_counts=True)
wordProb = wordCounts / np.float64(wordCounts.sum())
entropies[i]= H(wordProb)/width/wordLength
biasCorrections[i] = getBiasCorrection(words)/width/wordLength
validEntropies = getValidEntropies(entropies, biasCorrections)
A, w = getInfiteLengthExtrapolation(entropies, biasCorrections, wordLengths, nIgnore=2)
print('Theoretical entropy: %2.2f') % theoreticalEntropy
print('Estiamted entropy: %2.2f') % w[0]
# Plot entropies as a function of the word length
fig = plt.figure(figsize=(10, 4))
ax = plt.axes()
ax.plot(1./wordLengths, entropies, 'bo-', label='$H_\mathrm{raw}$')
ax.plot(1./wordLengths, entropies+biasCorrections, 'r.-', label='$H_\mathrm{corrected}$')
ax.plot(1./wordLengths[validEntropies], (entropies+biasCorrections)[validEntropies], 'ro')
ax.plot(A[1, :], np.dot(A.T, w), 'r:', label='$H_\mathrm{final}$')
ax.axhline(theoreticalEntropy, ls=':', color='k')
ax.set_ylim([0, 1.1*entropies.max()])
ax.set_xlim([0, 1])
ax.set_xlabel('(Word length)$^{-1}$')
ax.set_ylabel('Entropy rate (bits/s)');
ax.legend(frameon=False, bbox_to_anchor=(1, 0, 0.4, 1), loc=5);
# -
# In the example above, where $\mathbf{w}=[1, 2, 4]$, the cell outputs a unique spike count that directly identifies what the stimulus was during the preceding three time steps. However, this means that the value of $x_i$ can be revealed from either of $y_i$, $y_{i+1}$, or $y_{i+2}$, and that the spike output thus conveys the same information during multiple time steps (the output is correlated in time). We will consequently overestimate the entropy rate if we only look at words of length one (300 bits in the figure above), as this approach fails to take the correlations into account. However, if we look at the entropy rates for words of increasing length, we notice that they decrease and that they probably decrease to roughly 100 bits/s if you would extrapolate to an infinite word length. Thus, we see that our estimate of the entropy rate is only correct when extrapolated to an infinite word length, whenever temporal correlations are present. You might wonder whether this was just a coincidence, but Strong et al. (1998) highlight that the leading subextensive correction term is a constant whenever the correlations are of finite length. However, is you play around with different values of $\mathbf{w}$ above, you will notice that the extrapolation is not always this good, and that you might need a lot of data before the extrapolation works well.
# ### Mutual information, noisy transmission
# In real life, the example cell above is obviously not deterministic. In such cases, we need to estimate both the entropy rate as well as the noise entropy rate. The latter can be obtained in the same manner as the former, with the only difference being that the probability distributions are conditioned on the stimulus. We thus need data where a stimulus sequence has been repeated hundreds of times so that the word distributions can be reliably estimated also for longer word lengths. These ideas are illustrated in the example below, where cell responses are modeled as above, but where noise (possibly correlated in time) is added to the spike counts.
# +
width = 0.01
nSamples = 50000
nNoiseSamples = 100
nNoiseReps = 1000
w = np.array([1, 2])
wn = np.array([1])
wordLengths = np.arange(1, 10);
# 1. Entropy first
# Generate a random binary sequence and calculate the spike count in each bin
x = np.random.randint(0, 2, nSamples);
y = im_corr(x, w, mode='constant', origin=(w.size-1)/2)
n = np.random.randint(0, 2, nSamples);
nc = im_corr(n, wn, mode='constant', origin=(wn.size-1)/2)
y += nc;
# Raw and bias corrected entropies for various word lengths
entropies = np.zeros(wordLengths.size)
biasCorrections = np.zeros(wordLengths.size)
for i, wordLength in enumerate(wordLengths):
words = add_fake_dimension(y, wordLength)
uniqueWords, wordCounts = np.unique(words, axis=0, return_counts=True)
wordProb = wordCounts / np.float64(wordCounts.sum())
entropies[i] = H(wordProb)/width/wordLength
biasCorrections[i] = getBiasCorrection(words)/width/wordLength
validEntropies = getValidEntropies(entropies, biasCorrections)
A, w = getInfiteLengthExtrapolation(entropies, biasCorrections, wordLengths, nIgnore=2)
print('Estiamted entropy: %2.2f') % w[0]
# 2. Noise entropy second
# Generate a shorter random binary sequence that is repeated many times, and calculate the spike count in each bin
x = np.random.randint(0, 2, nNoiseSamples);
yTmp = im_corr(x, w, mode='constant', origin=(w.size-1)/2)
n = np.random.randint(0, 2,[nNoiseSamples, nNoiseReps]);
nc = im_corr(n, np.reshape(wn, [wn.size, 1]), mode='constant', origin=(wn.size-1)/2)
yNoise = yTmp[:, np.newaxis] + nc
# Raw and bias corrected noise entropies for various word lengths
noiseEntropies = np.zeros(wordLengths.size)
noiseBiasCorrections = np.zeros(wordLengths.size)
for i, wordLength in enumerate(wordLengths):
entropiesTmp = 0
biasCorrectionsTmp = 0
wordsAll = add_fake_dimension(yNoise, wordLength)
for j in range(wordsAll.shape[0]):
wordsTmp = wordsAll[j, :, :].T
uniqueWords, wordCounts = np.unique(wordsTmp, axis=0, return_counts=True)
wordProb = wordCounts / np.float64(wordCounts.sum())
entropiesTmp += H(wordProb)/width/wordLength
biasCorrectionsTmp += getBiasCorrection(wordsTmp)/width/wordLength
noiseEntropies[i] = entropiesTmp / wordsAll.shape[0]
noiseBiasCorrections[i] = biasCorrectionsTmp / wordsAll.shape[0]
validNoiseEntropies = getValidEntropies(noiseEntropies, noiseBiasCorrections)
ANoise, wNoise = getInfiteLengthExtrapolation(noiseEntropies, noiseBiasCorrections, wordLengths, nIgnore=0)
print('Estiamted noise entropy: %2.2f') % wNoise[0]
print('Estiamted mutual information: %2.2f') % (w[0] - wNoise[0])
# Plot entropies as a function of word length
fig = plt.figure(figsize=(10, 4))
ax = plt.axes()
ax.plot(1./wordLengths, entropies, 'bo-', label='$H_\mathrm{raw}$')
ax.plot(1./wordLengths, entropies+biasCorrections, 'r.-', label='$H_\mathrm{corrected}$')
ax.plot(1./wordLengths[validEntropies], (entropies+biasCorrections)[validEntropies], 'ro')
ax.plot(A[1, :], np.dot(A.T, w), 'r:', label='$H_\mathrm{final}$')
ax.plot(1./wordLengths, noiseEntropies, 'bo-', label='$H_\mathrm{raw}^\mathrm{noise}$')
ax.plot(1./wordLengths, noiseEntropies+noiseBiasCorrections, 'r.-', label='$H_\mathrm{corrected}^\mathrm{noise}$')
ax.plot(1./wordLengths[validNoiseEntropies], (noiseEntropies+noiseBiasCorrections)[validNoiseEntropies], 'ro')
ax.plot(ANoise[1, :], np.dot(ANoise.T, wNoise), 'r:', label='$H_\mathrm{final}^\mathrm{noise}$')
ax.set_ylim([0, 1.1*entropies.max()])
ax.set_xlim([0, 1])
ax.set_xlabel('(Word length)$^{-1}$')
ax.set_ylabel('Entropy rate (bits/s)');
ax.legend(frameon=False, bbox_to_anchor=(1, 0, 0.4, 1), loc=5);
| InformationTheory/Part 4, entropy of spike trains.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Laboratory 13 Probability Modeling
# ## Full name:
# ## R#:
# ## HEX:
# ## Title of the notebook
# ## Date:
# ### Important Terminology:
# __Population:__ In statistics, a population is the entire pool from which a statistical sample is drawn. A population may refer to an entire group of people, objects, events, hospital visits, or measurements. <br>
# __Sample:__ In statistics and quantitative research methodology, a sample is a set of individuals or objects collected or selected from a statistical population by a defined procedure. The elements of a sample are known as sample points, sampling units or observations.<br>
# __Distribution (Data Model):__ A data distribution is a function or a listing which shows all the possible values (or intervals) of the data. It also (and this is important) tells you how often each value occurs. <br>
#
# __*From https://www.investopedia.com/terms*__<br>
# __*https://www.statisticshowto.com/data-distribution/*__
# + [markdown] jupyter={"outputs_hidden": false}
# ### Important Steps:
# 1. __Get descriptive statistics- mean, variance, std. dev.__
# 2. __Use plotting position formulas (e.g., weibull, gringorten, cunnane) and plot the SAMPLES (data you already have)__
# 3. __Use different data models (e.g., normal, log-normal, Gumbell) and find the one that better FITs your samples- Visual or Numerical__
# 4. __Use the data model that provides the best fit to infer about the POPULATION__
# -
# # Estimate the magnitude of the annual peak flow at Spring Ck near Spring, TX.
#
# The file `08068500.pkf` is an actual WATSTORE formatted file for a USGS gage at Spring Creek, Texas. The first few lines of the file look like:
#
# Z08068500 USGS
# H08068500 3006370952610004848339SW12040102409 409 72.6
# N08068500 Spring Ck nr Spring, TX
# Y08068500
# 308068500 19290530 483007 34.30 1879
# 308068500 19390603 838 13.75
# 308068500 19400612 3420 21.42
# 308068500 19401125 42700 33.60
# 308068500 19420409 14200 27.78
# 308068500 19430730 8000 25.09
# 308068500 19440319 5260 23.15
# 308068500 19450830 31100 32.79
# 308068500 19460521 12200 27.97
#
# The first column are some agency codes that identify the station , the second column after the fourth row is a date in YYYYMMDD format, the third column is a discharge in CFS, the fourth and fifth column are not relevant for this laboratory exercise. The file was downloadef from
#
# https://nwis.waterdata.usgs.gov/tx/nwis/peak?site_no=08068500&agency_cd=USGS&format=hn2
#
# In the original file there are a couple of codes that are manually removed:
#
# - 19290530 483007; the trailing 7 is a code identifying a break in the series (non-sequential)
# - 20170828 784009; the trailing 9 identifies the historical peak
#
# The laboratory task is to fit the data models to this data, decide the best model from visual perspective, and report from that data model the magnitudes of peak flow associated with the probebilitiess below (i.e. populate the table)
#
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# The first step is to read the file, skipping the first part, then build a dataframe:
#
# + jupyter={"outputs_hidden": false}
# Read the data file
amatrix = [] # null list to store matrix reads
rowNumA = 0
matrix1=[]
col0=[]
col1=[]
col2=[]
with open('08068500.pkf','r') as afile:
lines_after_4 = afile.readlines()[4:]
afile.close() # Disconnect the file
howmanyrows = len(lines_after_4)
for i in range(howmanyrows):
matrix1.append(lines_after_4[i].strip().split())
for i in range(howmanyrows):
col0.append(matrix1[i][0])
col1.append(matrix1[i][1])
col2.append(matrix1[i][2])
# col2 is date, col3 is peak flow
#now build a datafranem
# + jupyter={"outputs_hidden": false}
import pandas
df = pandas.DataFrame(col0)
df['date']= col1
df['flow']= col2
# + jupyter={"outputs_hidden": false}
df.head()
# -
# Now explore if you can plot the dataframe as a plot of peaks versus date.
# + jupyter={"outputs_hidden": false}
# Plot here
# -
# From here on you can proceede using the lecture notebook as a go-by, although you should use functions as much as practical to keep your work concise
# + jupyter={"outputs_hidden": false}
# Descriptive Statistics
# + jupyter={"outputs_hidden": false}
# Weibull Plotting Position Function
# + jupyter={"outputs_hidden": false}
# Normal Quantile Function
# + jupyter={"outputs_hidden": false}
# Fitting Data to Normal Data Model
# -
# ## Normal Distribution Data Model
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# <hr>
# + jupyter={"outputs_hidden": false}
# Log-Normal Quantile Function
# + jupyter={"outputs_hidden": false}
# Fitting Data to Normal Data Model
# -
# ## Log-Normal Distribution Data Model
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# <hr>
# + jupyter={"outputs_hidden": false}
# Gumbell EV1 Quantile Function
# + jupyter={"outputs_hidden": false}
# Fitting Data to Gumbell EV1 Data Model
# -
# ## Gumbell Double Exponential (EV1) Distribution Data Model
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# <hr>
# + jupyter={"outputs_hidden": false}
# Gamma (Pearson Type III) Quantile Function
# + jupyter={"outputs_hidden": false}
# Fitting Data to Pearson (Gamma) III Data Model
# This is new, in lecture the fit was to log-Pearson, same procedure, but not log transformed
# -
# ## Pearson III Distribution Data Model
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# <hr>
# + jupyter={"outputs_hidden": false}
# Fitting Data to Log-Pearson (Log-Gamma) III Data Model
# -
# ## Log-Pearson III Distribution Data Model
# |Exceedence Probability|Flow Value|Remarks|
# |:---|:---|:---|
# |25% |????| 75% chance of greater value|
# |50% |????| 50% chance of greater value|
# |75% |????| 25% chance of greater value|
# |90% |????| 10% chance of greater value|
# |99% |????| 1% chance of greater value (in flood statistics, this is the 1 in 100-yr chance event)|
# |99.8%|????| 0.002% chance of greater value (in flood statistics, this is the 1 in 500-yr chance event)|
# |99.9%|????| 0.001% chance of greater value (in flood statistics, this is the 1 in 1000-yr chance event)|
#
# <hr>
# # Summary of "Best" Data Model based on Graphical Fit
| 1-Lessons/Lesson13/Lab13/Lab13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: left; margin: 30px 15px 15px 15px;" src="https://pngimage.net/wp-content/uploads/2018/06/logo-iteso-png-5.png" width="300" height="500" />
#
#
# ### <font color='navy'> Simulación de procesos financieros.
#
# **Nombres:** <NAME>, <NAME>
#
# **Fecha:** 07 de mayo de 2021.
#
# **Expediente** : if72622, if721861.
# **Profesor:** <NAME>.
#
# # Tarea 10: Clase 23
# [Liga al repositorio](https://github.com/PintorOD1997/ProyectoConjunto_LEspinosa_DPintor.git)
# ## Enunciado de tarea
# # Tarea
#
# Implementar el método de esquemas del trapecio, para valuar la opción call y put asiática con precio inicial, $S_0 = 100$, precio de ejercicio $K = 100$, tasa libre de riesgo $r = 0.10$, volatilidad $\sigma = 0.20$ y $T = 1$ año. Cuyo precio es $\approx 7.04$. Realizar la simulación en base a la siguiente tabla:
# 
#
# Observe que en esta tabla se encuentran los intervalos de confianza de la aproximación obtenida y además el tiempo de simulación que tarda en encontrar la respuesta cada método.
# - Se debe entonces realizar una simulación para la misma cantidad de trayectorias y número de pasos y construir una Dataframe de pandas para reportar todos los resultados obtenidos.**(70 puntos)**
# - Compare los resultados obtenidos con los resultados arrojados por la función `Riemann_approach`. Concluya. **(30 puntos)**
# ### Solución Lyha Espinosa
#importar los paquetes que se van a usar
import pandas as pd
import pandas_datareader.data as web
import numpy as np
import datetime
from time import time
import matplotlib.pyplot as plt
import scipy.stats as st
import seaborn as sns; sns.set()
# %matplotlib inline
#algunas opciones para Pandas
pd.set_option('display.notebook_repr_html', True)
pd.set_option('display.max_columns', 6)
pd.set_option('display.max_rows', 10)
pd.set_option('display.width', 78)
pd.set_option('precision', 3)
# +
def BSprices(mu,sigma,S0,NbTraj,NbStep):
"""
Expresión de la solución de la ecuación de Black-Scholes
St = S0*exp((r-sigma^2/2)*t+ sigma*DeltaW)
Parámetros
---------
mu : Tasa libre de riesgo
sigma : Desviación estándar de los rendimientos
S0 : Precio inicial del activo subyacente
NbTraj: Cantidad de trayectorias a simular
NbStep: Número de días a simular
"""
# Datos para la fórmula de St
nu = mu-(sigma**2)/2
DeltaT = 1/NbStep
SqDeltaT = np.sqrt(DeltaT)
DeltaW = SqDeltaT*np.random.randn(NbTraj,NbStep-1)
# Se obtiene --> Ln St = Ln S0+ nu*DeltaT + sigma*DeltaW
increments = nu*DeltaT + sigma*DeltaW
concat = np.concatenate((np.log(S0)*np.ones([NbTraj,1]),increments),axis=1)
# Se utiliza cumsum por que se quiere simular los precios iniciando desde S0
LogSt = np.cumsum(concat,axis=1)
# Se obtienen los precios simulados para los NbStep fijados
St = np.exp(LogSt)
# Vector con la cantidad de días simulados
t = np.arange(0,NbStep)
return St.T,t
def calc_daily_ret(closes):
return np.log(closes/closes.shift(1)).iloc[1:]
# -
# ## Call
def call_asiatica_trap(K:'Strike price',r:'Tasa libre de riesgo',S0:'Precio inicial',
NbTraj:'Número trayectorias',NbStep:'Cantidad de pasos a simular',
sigma:'Volatilidad',T:'Tiempo de cierre del contrato'):
# Iniciar Tiempo
tiempo_in = time()
# precios mediante black and scholes
St,t = BSprices(r,sigma,S0,NbTraj,NbStep)
prices = pd.DataFrame(St,index=t) #DataFrame de precios
# precios promedio
h = T/NbStep
Av_t = np.cumsum(prices * (2+r*h+np.random.randn(NbStep,NbTraj)*sigma))*h/(2*T)
strike = K
# Calculamos el call
valuacion = pd.DataFrame({'Prima': np.exp(-r*T)*np.fmax(Av_t - strike, 0).mean(axis=1)}, index=t)
# intervalos de confianza
conf = 0.95
i = st.norm.interval(conf, loc = valuacion.iloc[-1].Prima, scale = valuacion.sem().Prima)
tiempo_fin = time()
tiempo_total = tiempo_fin - tiempo_in
# regresar prima, intervalos de confianza, rango de intervalos y tiempo
return np.array([valuacion.iloc[-1].Prima,i[0],i[1],i[1]-i[0],tiempo_total])
# +
# Valores de la opcion
S0 = 100
K = 100
r = 0.1
sigma = 0.2
T = 1
# Trayectorias monte carlo
NbTraj = [1000,5000,10000,50000,100000,500000,1000000]
# Pasos en el tiempo
NbStep = [10,50,100]
# -
call = list(map(lambda trayectorias:list(map(lambda pasos:
call_asiatica_trap(K,r,S0,trayectorias,pasos,sigma,T),NbStep)), NbTraj))
# +
n = 3 # niveles por index
# indice posicion de cada rango
i1 =list(map(lambda i: int(i/n),range(7*n)))
# posicion de cada sub rango
i2 =list(map(lambda i: int(i%n),range(7*n)))
# index del data frame
indx = pd.MultiIndex(levels=[NbTraj,NbStep], codes=[i1, i2])
call_array = np.array([call[i1[i]][i2[i]] for i in range(len(i1))])
call_aprox = np.array([i[0] for i in call_array]) # aproximacion
# limite inferior
lim_inf = np.array([i[1] for i in call_array])
# limite superior
lim_sup = np.array([i[2] for i in call_array])
# rango de limites
rango = np.array([i[3] for i in call_array])
# obtener tiempos
tiempos_call = np.array([i[4] for i in call_array])
# General DF
tabla_call = pd.DataFrame(index=indx,columns=['Aproximacion','Linferior','Lsuperior','Longitud 95%','Tiempo'])
tabla_call.index.names = (['Tray. Monte Carlo','Num. pasos en el tiempo'])
tabla_call['Aproximacion'] = call_aprox
tabla_call['Linferior'] = lim_inf
tabla_call['Lsuperior'] = lim_sup
tabla_call['Longitud 95%'] = rango
tabla_call['Tiempo'] = np.round(tiempos_call,2)
# -
pd.set_option("display.max_rows", None, "display.max_columns", None)
tabla_call
# ## Put
def put_asiatica_trap(K:'Strike price',r:'Tasa libre de riesgo',S0:'Precio inicial',
NbTraj:'Número trayectorias',NbStep:'Cantidad de pasos a simular',
sigma:'Volatilidad',T:'Tiempo de cierre del contrato'):
# Iniciar Tiempo
tiempo_in = time()
# precios mediante black and scholes
St,t = BSprices(r,sigma,S0,NbTraj,NbStep)
prices = pd.DataFrame(St,index=t) #DataFrame de precios
# precios promedio
h = T/NbStep
Av_t = np.cumsum(prices * (2+r*h+np.random.randn(NbStep,NbTraj)*sigma))*h/(2*T)
strike = K
# Calculamos put
valuacion = pd.DataFrame({'Prima': np.exp(-r*T)*np.fmax(strike - Av_t , 0).mean(axis=1)}, index=t)
# intervalos de confianza
conf = 0.95
i = st.norm.interval(conf, loc = valuacion.iloc[-1].Prima, scale = valuacion.sem().Prima)
tiempo_fin = time()
tiempo_total = tiempo_fin - tiempo_in
# regresar prima, intervalos de confianza, rango de intervalos y tiempo
return np.array([valuacion.iloc[-1].Prima,i[0],i[1],i[1]-i[0],tiempo_total])
put = list(map(lambda trayectorias:list(map(lambda pasos:
put_asiatica_trap(K,r,S0,trayectorias,pasos,sigma,T),NbStep)), NbTraj))
# +
n = 3 # niveles por index
# indice posicion de cada rango
i1 =list(map(lambda i: int(i/n),range(7*n)))
# posicion de cada sub rango
i2 =list(map(lambda i: int(i%n),range(7*n)))
# index del data frame
indx = pd.MultiIndex(levels=[NbTraj,NbStep], codes=[i1, i2])
put_array = np.array([put[i1[i]][i2[i]] for i in range(len(i1))])
put_aprox = np.array([i[0] for i in put_array]) # aproximacion
# limite inferior
lim_inf_p = np.array([i[1] for i in put_array])
# limite superior
lim_sup_p = np.array([i[2] for i in put_array])
# rango de limites
rango = np.array([i[3] for i in put_array])
# obtener tiempos
tiempos_put = np.array([i[4] for i in put_array])
# General DF
tabla_put = pd.DataFrame(index=indx,columns=['Aproximacion','Linferior','Lsuperior','Longitud 95%','Tiempo'])
tabla_put.index.names = (['Tray. Monte Carlo','Num. pasos en el tiempo'])
tabla_put['Aproximacion'] = put_aprox
tabla_put['Linferior'] = lim_inf_p
tabla_put['Lsuperior'] = lim_sup_p
tabla_put['Longitud 95%'] = rango
tabla_put['Tiempo'] = np.round(tiempos_put,2)
# -
tabla_put
| ProyectoConjunto_2_Tarea10_LEspinosa_DPintor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Schelling Segregation
#
# ## Conceptual description
# The basic idea of Schelling's seggregation model is quite straight forward. Imagine we have agents organized on a grid. Agents are either red or blue. Agents prefer to be in an environment with agents of the same color, but they are quite tolerant. If the fraction of neighbors of a different color is higher than this tolerance threshold, the agent moves to another free grid cell. At what level of tolerance does segregation occur?
#
# Given this basic description, the model is composed of a grid, with one agent per grid cell. This grid is not completely filled because we need to have free space to where an agent can move. Each agent has a tolerance level, and checks when asked if the fraction of neighbors that is of the different type is higher than this tolerance level.
#
# ## Implementing a first version of the model
#
# ### Assignment
#
# 1. What classes do you need for this model? And what are the key attributes for each of these classes?
#
#
# <font color = red>IvS: Is dit het antwoord van de assignment hiervoor? Weet niet of het handig is om direct de antwoorden te geven, dan gaan de studenten niet zelf nadenken maar nemen ze dit gewoon over. Als je wilt dat de studenten zelf nadenken maar toch de antwoorden geven omdat deze belangrijk zijn voor de vervolgstappen, zou je dit kunnen verplaatsen naar Assignment 1. Staat het er niet direct onder maar is het wel benoemd. Hier gebruik je ook de classes dus dat geeft ook direct duidelijkheid. </font>
#
# You need <font color = red> the following classes with its key attributes:</font>
#
# **SchellingModel**
# * grid <font color = red> IvS: SingleGrid of gewoon grid? Gebaseerd op de beschrijving zou ik SingleGrid zeggen, triggert de studenten gelijk om weer na te denken over de verschillende mogelijke Grids.</font>
# * schedule
# * initial ratio between agent colors
#
# **SchellingAgent**
# * color
# * tolerance threshold
# ## Schedulers
# A key issue with the Schelling model, and a key way in which this model is different from the various versions of the Axelrod model is that a step by a given agent can change the local environment. Imagine that a given agent checks its neighborhood, decided that the number of neighbors of a different color is higher than its threshold, and thus decides to move to an empty spot. This move changes the neighborhood the agent is leaving. It also changes the neighborhood the agent is moving to. Depending on the order in which agents are activated (i.e. the `step` method is called) different dynamic patterns can occur.
#
# This agent activation is typically handled through a so-called scheduler. In the Axelrod method, we did this ourself by having a list or set of agents as an attribute on the model. However, we might want to use a more sophisticated class to handle agent activation. MESA comes with a number of schedulers.
#
# * **BaseScheduler**; the base scheduler is nothing but a list and we activate the agents based on the unique ID they have. Internally, base scheduler uses an `OrderedDict`, orded by `agent.unique_id`.
# * **RandomActivation**; shuffle all agents before activating them one by one in every time step. This is the most often used scheduler in all agent based models. It does introduce yet another form of stochasticity in the model, thus making it necessary to run multiple replications and carefully control the random seed.
# * **SimultaneousActivation**; a more complicated activation for mimicking all agents moving simultaneously. Rather than having a single step method on an agent, each agent has a staging method and an execution method. You first stage all agents, and then execute all agents. Staging in the context of Schelling Segregation means deciding to move or not. Executing means making the move. So SimultaneousActivation partly solves the path dependency problem mentioned above. Partly, because when moving only a single agent can occupy a given free spot.
# * **StagedActivation**; A more complicated activation system were agents have not a single step method, but multiple phases of action were each phase needs to be completed for all agents, before moving to the next method. Basically, this is an even more sophisticated extension of SimultaneousActivation.
#
# A model typically has only a single scheduler for all agents, but it is also possible to have multiple schedulers for different agent types.
# ## Assignment 1
#
# Below, I have given the outline of the model, we are going to implement this now. First, implement the ``__init__`` of the model. It is indicated below with the `...`. A hint regarding attributes can be found in the docstring. The code is very similar to the initialization used for the spatial version of the Axelrod model. After the code block, there is some test code to help you check if your code is working.
#
# Note that I suggest to use Color.RED for the minority and Color.BLUE for the majority. The reasons for this choice will become clear later in the assignment.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from mesa import Model, Agent
from mesa.time import RandomActivation
from mesa.space import SingleGrid
from mesa.datacollection import DataCollector
from enum import Enum
class Color(Enum):
RED = 1 # minority color
BLUE = 2 # majority color
class Schelling(Model):
"""
Model class for the Schelling segregation model.
Parameters
----------
height : int
height of grid
width : int
height of width
density : float
fraction of grid cells kept empty
minority_fraction : float
fraction of agent of minority color
tolerance_threshold : int
seed : int, optional
Attributes
----------
height : int
width : int
density : float
minority_fraction : float
schedule : RandomActivation instance
grid : SingleGrid instance
"""
def __init__(self, height=20, width=20, density=0.8, minority_fraction=0.2,
tolerance_threshold=4, seed=None):
self.grid = SingleGrid(width, height, torus=True)
self.time.RandomActivation
for cell in self.grid.coord_iter():
if self.random.random() > density:
continue
else:
#create agent
#add agent to grid and scheduler
def step(self):
"""
Run one step of the model.
"""
self.schedule.step()
class SchellingAgent(Agent):
"""
Schelling segregation agent
Parameters
----------
pos : tuple of 2 ints
the x,y coordinates in the grid
model : Model instance
color : {Color.RED, Color.BLUE}
tolerance_threshold : int
"""
def __init__(self, pos, model, color, tolerance_threshold):
super().__init__(pos, model)
self.pos = pos
self.color = color
self.tolerance_threshold = tolerance_threshold
def step(self):
raise NotImplementedError
# -
#
# To test the implementation, we can visualize the grid. To do this, we are creating a numpy array with zeros of the same width and height as the model grid. Next, we iterate over the grid and fill in this empty array. We use a value of 3 for an empty cell, and the color number from the `Color` enum class.
#
# We can easily visualize the filled in numpy array using matplotlib (e.g. matshow) or seaborn's heatmap. To make the figure a bit nicer, I use a few tricks. First, I am getting a color map from seaborn. This color map is generated by the [colorbrewer](https://colorbrewer2.org/) website. As you can see on this website, the `Set1` colormap has red as 1, blue as 2, and 3 as green. This matches perfectly with the numbers used in filling in the numpy array (and why we used 3 for the empty cells). Second, I like to have a nice equally spaced figure, which you can do by setting the aspect of the axes to equal.
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
model = Schelling()
def plot_model(model, ax, colors):
grid = np.zeros((model.height, model.width))
for (cell, i, j) in model.grid.coord_iter():
value = 3
if cell is not None:
value = cell.color.value
grid[i, j] = value
sns.heatmap(grid, ax=ax, cmap=colors)
fig, ax = plt.subplots(figsize=(10, 10))
colors = sns.color_palette('Set1', 3)
ax.set_aspect('equal')
plot_model(model, ax, colors)
plt.show()
# -
# ## Assignment 2
# The next step is to implement the logic in the step method of the agent. This is straight forward, we iterate over our neighbors. If our neighbor is of a different color, we increment a counter. After having checked our entire neighborhood, the agent compares this counter with its tolerance threshold. If the number of neighbors of a different color is higher than the threshold, the agent will move. For now, we will use the `move_to_empty` method on the grid for this.
#
# Regarding neighborhood, assume that we include also our diagonal neighbors.
#
# Go ahead and implement the model.
#
# Test code is provided below.
class SchellingAgent(Agent):
"""
Schelling segregation agent.
Parameters
----------
pos : tuple of 2 ints
the x,y coordinates in the grid
model : Model instance
color : {Color.RED, Color.BLUE}
tolerance_threshold : float
"""
def __init__(self, pos, model, color, tolerance_threshold):
super().__init__(pos, model)
self.pos = pos
self.color = color
self.tolerance_threshold = tolerance_threshold
def step(self):
...
# +
model = Schelling()
for i in range(10):
fig, ax = plt.subplots()
fig.suptitle(i)
ax.set_aspect('equal')
plot_model(model, ax, colors)
plt.show()
model.step()
# -
# We now have a basis version of Schelling seggregation working. We can play around with this model and already learn quite a bit. However, we can also expand the model a bit more.
# ## Assignment 3
# We now have a working model. But we have so far relied only on visual inspection of the model. If we want to explore different tolerance levels, we need some useful statistics from the model as well. Making sense only of the grid is rather hard if we start running this model a few hundred times for different random seeds, different tolerance levels, and different neighborhood rules. A statistic that is easy to check is keeping track of how many agents are happy with their position. That is, they did not want to move to a new location.
#
# Implementing this requires a few changes:
#
# * Agents need a happy attribute. This is set to True if the agent did not move. It is set to False otherwise.
# * The model needs a data collector. This collector is called at each step of the model after all agents have moved and keeps track of the total number of happy agents.
#
# Go ahead and update both the agent and the model.
# +
def count_happy(model):
...
class Schelling(Model):
"""
Model class for the Schelling segregation model.
Parameters
----------
height : int
height of grid
width : int
height of width
density : float
fraction of grid cells kept empty
minority_fraction : float
fraction of agent of minority color
tolerance_threshold : int
Attributes
----------
height : int
width : int
density : float
minority_fraction : float
schedule : RandomActivation instance
grid : SingleGrid instance
"""
def __init__(self, height=20, width=20, density=0.8, minority_fraction=0.2,
tolerance_threshold=4, seed=None):
...
def step(self):
"""
Run one step of the model.
"""
self.schedule.step()
self.datacollector.collect(self)
# -
class SchellingAgent(Agent):
"""
Schelling segregation agent
Parameters
----------
pos : tuple of 2 ints
the x,y coordinates in the grid
model : Model instance
color : {Color.RED, Color.BLUE}
tolerance_threshold : float
"""
def __init__(self, pos, model, color, tolerance_threshold):
super().__init__(pos, model)
self.pos = pos
self.color = color
self.tolerance_threshold = tolerance_threshold
self.happy = True
def step(self):
'''Execute one step of the agent'''
...
model = Schelling()
for _ in range(30):
model.step()
model.datacollector.get_model_vars_dataframe().plot()
plt.grid(True)
plt.show()
# ## Assignment 4
# We used the ``move_to_empty`` method to move our agent. However, this method simply returns a random empty cell. We can make this a bit smarter by checking each free cell to see if moving there makes sense. That is, would the new neighborhood meet our preferences with respect to the different types of neighbors?
#
# Below I have given the basic structure of this refined agent class. You have to fill in the `...` parts. Some more useful methods and attributes you might need are
#
# * *grid.empties*; contains the collection of empty grid cells
# * *grid.move_agent(agent, pos)*; method for moving an agent to a diffent grid cell. It also updates the position of the agent on the agent itself.
# * the agent has a model attribute, the model has a grid attribute
#
# Note that I have moved the checking of the threshold to its own seperate method which takes as input a position. In this way we can use the exact same code for checking whether an agent wants to move (we pass `self.pos`) or whether an empty cell meets the threshold (we pass an entry in `grid.empties`).
#
# Note also that you should keep updating the state of the agent (i.e. `agent.happy`)
class SchellingAgent(Agent):
"""
Schelling segregation agent.
Parameters
----------
pos : tuple of 2 ints
the x,y coordinates in the grid
model : Model instance
color : {Color.RED, Color.BLUE}
tolerance_threshold : float
"""
def __init__(self, pos, model, color, tolerance_threshold):
super().__init__(pos, model)
self.pos = pos
self.color = color
self.tolerance_threshold = tolerance_threshold
self.happy = True
def meets_threshold(self, pos):
'''Check if a given position meets the tolerance threshold
Returns
-------
bool
'''
different = 0
for neighbor in self.model.grid.neighbor_iter(pos, moore=True):
if neighbor.color != self.color:
different += 1
return different <= self.tolerance_threshold
def move_to_empty(self):
'''Move to an empty spot in the grid if the empty spot
meets the tolerance threshold'''
...
def step(self):
'''Execute one step of the agent'''
...
# +
model = Schelling()
for i in range(10):
model.step()
model.datacollector.get_model_vars_dataframe().plot()
plt.grid(True)
plt.show()
# -
# ## Assignment 5
# So far, we have run the model for the default values of the parameters `density`, `minority_fraction`, and `tolerance_threshold`. It is quite common in Agent Based Modelling to perform a parameter sweep where one runs the model over a range of parameterizations and analyse the results.
#
# MESA support this with a so-called BatchRunner. See the [MESA tutorial](https://mesa.readthedocs.io/en/master/tutorials/intro_tutorial.html#batch-run) for details. In short, the BatchRunner allows you to specify a set of values for the parameters you want to vary, fixed values for parameters you don't want to vary, the number of iterations/replications for each run, the number of steps to run the model for, and what statistics to collect at the end of the run. The BatchRunner assumes that a `self.running` attribute is available on the model. This is used to indicate that the model has reached a stable equilibrium and will not change in any subsequent steps. If running is True, the model will still change. If running is False, the model has reached an equilibrium.
#
# 1. Let's update our model with this running attribute. What conditions must hold for the model to have reached equilibirum?
#
# 2. Implement this condition and the running attribute.
# *Hint:* the scheduler has a `get_agent_count` method
#
# 3. Perform a batch run with the following settings. Visualize some results:
# * height and width fixed at 20
# * density varies from 0.8 to 0.9 with 0.025 increments
# * minority_fraction varies from 0.2 to 0.4 with 0.1 increments
# * tolerance_threshold varies from 2 to 4 with increment of 1
# * 5 replications
# * 50 steps
#
# *Note:* there is a bug in the batchrunner were the column names in the dataframe after the batch run don't match the correct labels.
#
class Schelling(Model):
"""
Model class for the Schelling segregation model.
Parameters
----------
height : int
height of grid
width : int
height of width
density : float
fraction of grid cells kept empty
minority_fraction : float
fraction of agent of minority color
tolerance_threshold : int
Attributes
----------
height : int
width : int
density : float
minority_fraction : float
schedule : RandomActivation instance
grid : SingleGrid instance
"""
def __init__(self, height=20, width=20, density=0.8, minority_fraction=0.2,
tolerance_threshold=4, seed=None):
super().__init__(seed=seed)
self.height = height
self.width = width
self.density = density
self.minority_fraction = minority_fraction
self.running = False
self.schedule = RandomActivation(self)
self.grid = SingleGrid(width, height, torus=True)
self.datacollector = DataCollector(model_reporters={'happy':count_happy})
# Set up agents
# We use a grid iterator that returns
# the coordinates of a cell as well as
# its contents. (coord_iter)
for cell in self.grid.coord_iter():
x = cell[1]
y = cell[2]
if self.random.random() < self.density:
if self.random.random() < self.minority_fraction:
agent_color = Color.RED
else:
agent_color = Color.BLUE
agent = SchellingAgent((x, y), self, agent_color, tolerance_threshold)
self.grid.position_agent(agent, (x, y))
self.schedule.add(agent)
def step(self):
"""
Run one step of the model.
"""
self.schedule.step()
self.datacollector.collect(self)
...
# +
from mesa.batchrunner import BatchRunner
...
# -
results = batch_run.get_model_vars_dataframe()
results.head(10)
| src/week_4/Assignment 3 segregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Terminology
#
# Before we start, we need to get a bit of terminology out of the way. Data Scientists come from a variety of backgrounds and techniques have been invented and reinvented under a variety of names in a number of fields. In order to get everyone on the same page, we're going to talk a little bit about terminology.
#
# Artificial Intelligence (AI) is concerned with building algorithms that enable computers to learn, understand, and solve problems. One subset of AI is Machine Learning. It might help to contrast ML with one of the older branches of AI, state space search.
#
# In state space search, a problem is specified terms of permitted states and actions that can be taken in those states. If you specify an end state as a goal, a state space search algorithm will find a series of actions that will get you from the current state to the goal.
#
# In contrast, a *supervised* machine learning algorithm takes data in the form of *examples* of inputs and outputs and returns a model of those examples. You can then supply different inputs to the model and get an appropriate output. Because of the relationship between inputs and an output, supervised machine learning is similar or identical to function approximation or pattern recognition. We can write this in symbols as:
#
# $$f = G(y, X)$$
#
# where $G$ is the machine learning algorithm (we'll assume we're talking about "supervised" for now), $(y, X)$ are a tuple of outputs and inputs, and $f$ is the model.
#
# $$\hat{y} = f(X)$$
#
# we can then use $f$ on new inputs $X$ to estimate values of $y$, which are called (by convention), $\hat{y}$.
#
# $X$ is a matrix of variables. Each column is called a *feature* (attribute, factor, variable, inputs) and $y$ is a vector of output values (target, response). Each column of $X$, $X_i$, can be a different data type (unlike a mathematical matrix) although the user has to be careful. Sometimes an algorithm $G$ will accept types that don't actually make sense for $f$.
#
# If $y$ is a numerical variable, this is called *regression* regardless of the actual $G$ used. If $y$ is a categorical variable, this is called *classification*. It is important to distinguish these uses because we can use can use *linear regression* to solve *regression* problems and we can use *logistic regression* to solve *classification* problems.
#
# In concrete terms, if you want to estimate things like price, age, IQ, or leaf volume, this is a regression problem. If you want to estimate a purchase outcome, religion, animal, or creditworthiness outcome, then this is a classification problem. This is important because not all algorithms, metrics, evaluation techniques are appropriate for both regression and classification.
#
# ## Why Linear and Logistic Regression
#
# So why not dive directly into boosted random forests? Linear and logistic regression regularly top the "must know" lists of algorithms for data scientists...and for good reason.
#
# First, we can go very deeply into these models and understand how they work, their advantages, and disadvantages.
#
# Second, one of the key findings in data science is that with "big" data, simple models can perform extremely well and more time should generally be placed on refining the features (inputs, or in regression-speak, the *regressors*) to the models. Facebook does this for many of its machine learning applications.
#
# Third, simple models are often sufficient to get a project off the ground, to go from 0% to 60% instead of 92% to 93%. A fielded, simple data product with 3% lift is better than a complicated data product with 20% lift that never gets fielded, needs to be constantly re-tuned or nobody understands.
#
# Finally, and specifically with regard to all types of regression, they are useful when we are trying to explore relationships in our process rather than simply prediction.
#
# For example, if we use a neural network to model our system, we may get a very good model of the inputs to the outputs. However, we will not be able to understand *how* the inputs relate to the outputs. With a linear regression model, we may get a poorer model (or have to futz with it more) but we will be able to understand the relationship between inputs and outputs. Ultimately it depends on what we're trying to do with the model (and hence the George Box quote from before). This is a general tension between statistics-based methods and machine learning-based methods.
#
# Nevertheless, we will address more complicated approaches in later chapters (machine learning) because *big* is really relative to the problem space and you may not be able to find or create better features than what you have. In any case, you should always start with the simplest thing first.
#
# Regression as a technique comes out of the field of statistics although people have been working with regression for over 130 years. <NAME> presented regression for the first time in a lecture in [1877](http://www.amstat.org/publications/jse/v9n3/stanton.html).
| fundamentals_2018.9/linear/terms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
import scipy.io
from matplotlib import pyplot
import pandas as pd
import numpy as np
from numpy import mean
from numpy import std
from pandas import read_csv
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_theme(style="ticks", color_codes=True)
import tensorflow as tf
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LeakyReLU
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.utils import plot_model
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
from sklearn.svm import SVC
from sklearn.datasets import make_classification
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from sklearn import svm
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.exceptions import UndefinedMetricWarning
from sklearn.metrics import accuracy_score
from sklearn.metrics import make_scorer
from sklearn.metrics import balanced_accuracy_score
from imbalanced_ensemble.ensemble import SelfPacedEnsembleClassifier as SPE
from sklearn.metrics import balanced_accuracy_score
# %%
print(__doc__)
# Import imbalanced_ensemble
import imbalanced_ensemble as imbens
# Import utilities
from collections import Counter
import sklearn
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from imbalanced_ensemble.ensemble.base import sort_dict_by_key
# Import plot utilities
import matplotlib.pyplot as plt
from imbalanced_ensemble.utils._plot import plot_2Dprojection_and_cardinality
# -
data = scipy.io.loadmat('x.mat')
columns = np.asarray([['Electrode %d - %d/2 Hz'%(i+1, j+1)] for i in range(data['x'].shape[1]) for j in range(data['x'].shape[2])])
data['x'].shape
labels = pd.read_csv("table_withlabels.csv")
foof = pd.read_csv("foof2features.csv")
beh = pd.read_csv("behaviorals.csv")
df = pd.DataFrame(data['x'].reshape((data['x'].shape[0], -1)))
df.columns = columns
df['IDs'] = foof['C1']
df2 = pd.merge(df, labels[['label', 'IDs']], on='IDs', how='inner')
print(df2['label'].value_counts())
# +
# traversing through disease
# column of dataFrame and
# writing values where
# condition matches.
df2.label[df2.label == 'Other Neurodevelopmental Disorders'] = 1
df2.label[df2.label == 'ADHD-Inattentive Type'] = 2
df2.label[df2.label == 'ADHD-Combined Type'] = 3
df2.label[df2.label == 'Anxiety Disorders'] = 4
df2.label[df2.label == 'No Diagnosis Given'] = 5
df2.label[df2.label == 'Depressive Disorders'] = 6
print(df2['label'].unique())
print(df2['label'].value_counts())
df = df2
df.label = df['label'].astype(int)
# -
X = df[df.columns.difference(['IDs', 'label'])]
y = df['label']
print(X.shape)
y.shape
X_train, X_valid, y_train, y_valid = train_test_split(X, y,
test_size=0.5, stratify=y, random_state=42)
# +
# Visualize the training dataset
fig = plot_2Dprojection_and_cardinality(X_train, y_train, figsize=(8, 4))
plt.show()
# Print class distribution
print('Training dataset distribution %s' % sort_dict_by_key(Counter(y_train)))
print('Validation dataset distribution %s' % sort_dict_by_key(Counter(y_valid)))
# +
# Use SVC as the base estimator
clf = SPE(
n_estimators=5,
base_estimator=SVC(probability=True), # Use SVM as the base estimator
random_state=42,
).fit(X_train, y_train)
# Evaluate
balanced_acc_score = balanced_accuracy_score(y_valid, clf.predict(X_valid))
print (f'SPE: ensemble of {clf.n_estimators} {clf.base_estimator_}')
print ('Validation Balanced Accuracy: {:.3f}'.format(balanced_acc_score))
# +
# %% [markdown]
# Enable training log
# -------------------
# (``fit()`` parameter ``train_verbose``: bool, int or dict)
clf = SPE(random_state=42).fit(
X_train, y_train,
train_verbose=True, # Enable training log
)
| archive/ensembles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: deepreinforcement
# language: python
# name: deepreinforcement
# ---
# +
from loss import cemse
import tensorflow as tf
from keras import losses
import keras.backend as K
# -
sess = tf.InteractiveSession()
# +
y_true = tf.constant([[0.9,0,0,1,0,0,0,0,0,0], [0.9,0,0,1,0,0,0,0,0,0]])
y_pred = tf.constant([[-0.5,0.5,-0.1,-0.8,-0.3,-0.1,0.7,0.2,-0.2,-0.2], [0.5,-0.5,0.1,0.8,0.3,0.1,-0.7,-0.2,0.2,0.2]])
v = tf.slice(y_pred, [0,0], [-1,1])
z = tf.slice(y_true, [0,0], [-1,1])
p = tf.slice(y_pred, [0,1], [-1,-1])
pi = tf.slice(y_true, [0,1], [-1,-1])
# -
loss_value = losses.mean_squared_error(z, tf.tanh(v))
loss_value.eval()
loss_actions = tf.nn.softmax_cross_entropy_with_logits(labels = pi, logits = p)
loss_actions.eval()
out = cemse(y_true, y_pred)
out2 = K.square(y_pred - y_true)
out.eval()
out2.eval()
| src/DeepReinforcementLearning/.ipynb_checkpoints/loss checking-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cython
# * Cython is one of python's *dialects* to bridge between C and python.
# * Following code block is an [example](https://stackoverflow.com/questions/35656604/running-cython-in-jupyter-ipython) of the cython code.
# +
# %load_ext Cython
# make cython feature avaiable in ipython
# + magic_args="--annotate" language="cython"
# # with command above, ipython would regard this cell as cython
# # cell magic command "%%" must be on the first line of the cell
# # --annotate option would present c code from the cython code
# # later click on the + signs of the yellow highlight lines
#
#
# # declare a cython function
# def geo_prog_cython(double alpha, int n):
# """
# Sum of a geographic sequence
# Cython version
#
# ==========
# Parameters
# ==========
#
# double alpha : ratio
# int n : number of terms
# """
#
# # local variables for this function
# # with types and initial values
# cdef double current = 1.0
# cdef double sum = current
# cdef int i
#
# # accumulation loop
# for i in range(n):
# # current term of a geometric sequence
# current *= alpha
# # accumulation
# sum += current
#
# # sum of the geometric sequence
# return sum
#
#
# -
# * As you can see, cython version has type information.
# * Let's try calling the cython function
# +
# measure time of computation
# repeating if necessary
# %timeit -n5 -r5 geo_prog_cython(0.5, 5)
# -
# ?timeit
# * To compare the computation time, let's prepare a python version.
# +
def geo_prog_python(alpha, n):
"""
Sum of a geographic sequence
Python version
==========
Parameters
==========
alpha : ratio
n : number of terms
"""
# initialize
current = 1.0
sum = current
# accumulation loop
for i in range(n):
current *= alpha
# accumulation
sum += current
return sum
# +
# measure time of computation
# repeating if necessary
# %timeit -n5 -r5 geo_prog_python(0.5, 5)
# -
# * Can you see the benefit?
# * Here `%%cython` command is doing all the [plumbing](http://blog.yclin.me/gsoc/2016/07/23/Cython-IPython/) for us.
# * Under the hood, [more](https://cython.readthedocs.io/en/latest/src/userguide/source_files_and_compilation.html) is going on as we may see later.
# ## Visualizing
# * `matplotlib` can visualize results from cython functions.
# * Following code presents Euler's method simulation.
# +
# %load_ext Cython
# Repeated to help locating
# +
# simulation parameters
t_start_sec = 0
t_end_sec = 10
delta_t_sec = 1e-3
# + magic_args="--annotate" language="cython"
# # with command above, ipython would regard this cell as cython
# # cell magic command "%%" must be on the first line of the cell
# # --annotate option would present c code from the cython code
#
# # https://stackoverflow.com/questions/29036068/how-to-expose-a-numpy-array-from-c-array-in-cython
# cimport numpy as np
# # for cimport, please see:
# # https://stackoverflow.com/questions/29311207/cython-compilation-import-vs-cimport
# # it would make C API available for np.ndarray[] later
# import numpy as np
# # for np.empty in this case
#
#
# def diff_eq(double xi):
# """
# Differential equation to solve
# x' + x = 0
# x' = - x
# """
#
# # dx/dt + x = 0
# # return value == result of this function
# return -xi
#
#
# def euler_cython_numpy_array(double ti_sec=0.0, double te_sec=10, double delta_t_sec=1e-3):
# """
# Cython + Numpy Simulation
# Euler's method
#
# ===============
# Input arguments
# ===============
# ti_sec : start time
# te_sec : end time
# delta_t_sec : time step
# """
#
# # initial values
# # inital value
# cdef double x0 = 1.0
#
# # time variable
# cdef double t = ti_sec
#
# # number of time steps
# cdef int n = int((te_sec - ti_sec) / delta_t_sec) + 1
# # time step
# cdef int i = 0
#
# # https://stackoverflow.com/questions/25974975/cython-c-array-initialization
# # simulation time
# cdef np.ndarray [double, ndim=1, mode="c"] result_t = np.empty(n)
# # state variable
# cdef np.ndarray [double, ndim=1, mode="c"] result_x = np.empty(n)
#
# # local variable for x'
# cdef double dx_dt = 0
#
# # initial values of time and state
# result_t[0] = ti_sec
# result_x[0] = x0
#
# # time step loop
# for i in range(1, n):
# # slope
# dx_dt = diff_eq(result_x[i-1])
# # state variable of next time step
# result_x[i] = result_x[i-1] + dx_dt * delta_t_sec
# # time step
# result_t[i] = result_t[i-1] + delta_t_sec
#
# # result of simulation
# return result_t, result_x
#
#
# -
# * To compare, following cell prepares an equivalent simulation in python.
# +
def diff_eq_python(xi):
# dx/dt + x = 0
# dx/dt = - x
return -xi
def euler_python(ti_sec = 0.0, te_sec = 10.0, delta_t_sec = 1e-3):
# initial value
x0 = 1.0
# number of time steps
n = int((te_sec - ti_sec) / delta_t_sec) + 1
# prepare for buffers before the loop
result_t = [0.0] * n
result_x = [0.0] * n
# set initial value
result_t[0] = ti_sec
result_x[0] = x0
# time step loop
for i in range(1, n):
dx_dt = diff_eq_python(result_x[i-1])
result_x[i] = result_x[i-1] + dx_dt * delta_t_sec
result_t[i] = result_t[i-1] + delta_t_sec
return result_t, result_x
# -
# * Let's run both cython and python versions
# +
# %%time
# measure time to calculate
t_cy_np, x_cy_np = euler_cython_numpy_array(
t_start_sec,
t_end_sec,
delta_t_sec,
)
# +
# %%time
# measure time to calculate
t_py, x_py = euler_python(
t_start_sec,
t_end_sec,
delta_t_sec,
)
# -
# * Let's plot both results
# +
# to plot properly, may need to separate this line
import pylab as py
# +
py.plot(t_cy_np, x_cy_np, 'o', label='cython numpy')
py.plot(t_py, x_py, '-', label='python')
py.grid(True)
py.xlabel('t')
py.ylabel('y')
py.legend(loc=0)
py.show()
# +
# measure time to calculate
# -o makes the result available as output
# time_cython_dynamic_numpy = %timeit -n5 -r5 -o t_np, x_np = euler_cython_numpy_array(t_start_sec, t_end_sec, delta_t_sec,)
# +
# # %timeit is a magic command to measure time
# -o makes the result available as output
# time_python = %timeit -n5 -r5 -o t_py, x_py = euler_python(t_start_sec, t_end_sec, delta_t_sec,)
# -
# ## Calling C/C++ functions
# * Cython can call C/C++ functions as follows.
# [[ref0](https://cython.readthedocs.io/en/latest/src/userguide/external_C_code.html)]
# , [[ref1](https://stackoverflow.com/questions/37426534/how-can-i-import-an-external-c-function-into-an-ipython-notebook-using-cython)]
# , [[ref2](https://stackoverflow.com/questions/19260253/cython-compiling-error-multiple-definition-of-functions)]
# , [[ref3](https://media.readthedocs.org/pdf/cython/stable/cython.pdf)]
# , [[ref4](http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html)]
#
#
# ### With `numpy` support
# * If we need to use matrices and vectors frequently, combining `numpy` and cython may be helpful.
# * Let's take a look at an example of calculating cosine values.
# * Following is the header file for the C source code.
# +
# %%writefile cos_cython_numpy.h
/*
2.8.5.2. Numpy Support, 2.8.5. Cython, http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html#id13
*/
void cos_cython_numpy_c_func(double * in_array, double * out_array, int size);
# -
# * Following is the source code file to wrap with cython.
# +
# %%writefile cos_cython_numpy.c
/*
2.8.5.2. Numpy Support, 2.8.5. Cython, http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html#id13
*/
#include <math.h>
/* Compute the cosine of each element in in_array, storing the result in
* out_array. */
void cos_cython_numpy_c_func(double * in_array, double * out_array, int size){
int i;
for(i=0;i<size;i++){
out_array[i] = cos(in_array[i]);
}
}
# -
# * Following is the cython `.pyx` file calling the C function.
# +
# %%writefile _cos_cython_numpy.pyx
""" Example of wrapping a C function that takes C double arrays as input using
the Numpy declarations from Cython
<NAME>, 2.8.5.2. Numpy Support, 2.8.5. Cython, Scipy Lectures, Oct 18 2016, [Online]
Available: http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html#id13
"""
""" Example of wrapping a C function that takes C double arrays as input using
the Numpy declarations from Cython """
# cimport the Cython declarations for numpy
cimport numpy as np
# if you want to use the Numpy-C-API from Cython
# (not strictly necessary for this example, but good practice)
np.import_array()
# cdefine the signature of our c function
cdef extern from "cos_cython_numpy.h":
void cos_cython_numpy_c_func (double * in_array, double * out_array, int size)
# create the wrapper code, with numpy type annotations
def cos_cython_numpy_py_func(np.ndarray[double, ndim=1, mode="c"] in_array not None,
np.ndarray[double, ndim=1, mode="c"] out_array not None):
cos_cython_numpy_c_func(<double*> np.PyArray_DATA(in_array),
<double*> np.PyArray_DATA(out_array),
in_array.shape[0])
# -
# * Following is file `setup.py`. Running this `.py` file would build the C function wrapper.
# +
# %%writefile setup.py
# <NAME>, 2.8. Interfacing with C, Scipy Lectures, Oct 18 2016, [Online]
# Available: http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html
from distutils.core import setup, Extension
# distutils : building and installing modules
# https://docs.python.org/3/library/distutils.html
import numpy
from Cython.Distutils import build_ext
print('for NumPy Support of Cython '.ljust(60, '#'))
setup(cmdclass={'build_ext': build_ext},
ext_modules=[Extension("cos_cython_numpy",
sources=['_cos_cython_numpy.pyx', "cos_cython_numpy.c"],
include_dirs=[numpy.get_include()])],
)
# + language="bash"
#
# # this is the command to build C function wrapper
# python setup.py build_ext --inplace
# # with --inplace, the module would be in the same folder.
#
#
# -
# * Now let's import the module with c wrapper function
# +
# the cython module including C function
import cos_cython_numpy
# to visualize the result
import pylab as py
# allocate arrays externally
x = py.arange(0, 2 * py.pi, 0.1)
y = py.empty_like(x)
# c wrapper function
cos_cython_numpy.cos_cython_numpy_py_func(x, y)
# plot the result
py.plot(x, py.cos(x), 'o', label='cos numpy')
py.plot(x, y, '.', label='cos cython')
py.show()
# -
# * Cleanup
# + language="bash"
# rm cos_cython_numpy.h
# rm cos_cython_numpy.c
# rm _cos_cython_numpy.c
# rm _cos_cython_numpy.pyx
# rm setup.py
# rm cos_cython_numpy.*.so
#
#
# -
# * Check results
# +
cos = py.cos(x)
for i in range(len(cos)):
assert y[i] == cos[i], "Cython Result != Pylab Expected"
# -
# ### Revisiting Euler's method
# * Let's try wrapping a C/C++ version of the Euler's method above.
# * Following is the header file for the C source code.
# +
# %%writefile euler_cython_numpy.h
/*
2.8.5.2. Numpy Support, 2.8.5. Cython, http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html#id13
*/
void euler_cython_c_function(
const double t_start_sec, const double t_end_sec,
const double delta_t_sec,
const int size,
const double * result_t, const double * result_x
);
# -
# * Following is a C source code example.
# +
# %%writefile euler_cython_numpy.c
#include <assert.h>
double diff_eq(double xi){
return -xi;
}
void euler_cython_c_function(
const double t_start_sec,
const double t_end_sec,
const double delta_t_sec,
const int size,
double * result_t, double * result_x
){
// Simulation start and end time
// Initial state
double x0 = 1.0;
double dx_dt = 0.0;
int i = 0;
// Length of simulation
const int n = (int) ((t_end_sec - t_start_sec) / delta_t_sec) + 1;
// Check array size
assert(size > n);
// Set initial value
result_t[0] = t_start_sec;
result_x[0] = x0;
// Time step loop
// Watch the last value of i here
for (i=0; (n-1)>i; ++i){
// Calculate derivative
dx_dt = diff_eq(result_x[i]);
// Calculate state value of the next step
result_x[i+1] = result_x[i] + dx_dt * delta_t_sec;
// Calculate time of next step
result_t[i+1] = result_t[i] + delta_t_sec;
}
return;
}
# -
# * Following is file `_euler_cython_numpy.pyx`.
# +
# %%writefile _euler_cython_numpy.pyx
# cimport the Cython declarations for numpy
cimport numpy as np
# to dynamically allocate numpy.ndarray using np.empty()
import numpy as np
# Recommended practice for Numpy-C-API from Cython
np.import_array()
# cdefine the signature of our c function
cdef extern from "euler_cython_numpy.h":
void euler_cython_c_function (
const double t_start_sec,
const double t_end_sec,
const double delta_t_sec,
const int size,
double * result_t, double * result_x,
)
# now we can call euler_cython_c_function()
# create the wrapper code, with numpy type annotations
# a python program would call this function
# and this function would call the C function
def euler_cython_numpy_py_func(
double t_start_sec=0,
double t_end_sec=10,
double delta_t_sec=1e-3,
):
# number of time steps
cdef int n = int((t_end_sec - t_start_sec) / delta_t_sec) + 1
# preallocate output arrays
cdef np.ndarray[double, ndim=1, mode="c"] t_array = np.empty(n)
cdef np.ndarray[double, ndim=1, mode="c"] x_array = np.empty(n)
# calling C function
euler_cython_c_function(
t_start_sec,
t_end_sec,
delta_t_sec,
t_array.shape[0],
<double*> np.PyArray_DATA(t_array),
<double*> np.PyArray_DATA(x_array),
)
# <double*> np.PyArray_DATA(t_array)
# this line would obtain a C double pointer for t_array
# Cython MemoryView can be an alternative
# https://cython.readthedocs.io/en/latest/src/userguide/memoryviews.html
return t_array, x_array
# -
# * Following is file `setup.py`.
# +
# %%writefile setup.py
# <NAME>, 2.8. Interfacing with C, Scipy Lectures, Oct 18 2016, [Online]
# Available: http://www.scipy-lectures.org/advanced/interfacing_with_c/interfacing_with_c.html
from distutils.core import setup, Extension
import numpy
from Cython.Distutils import build_ext
print('for NumPy Support of Cython '.ljust(60, '#'))
setup(cmdclass={'build_ext': build_ext},
ext_modules=[
Extension(
"euler_cython_numpy",
sources=['_euler_cython_numpy.pyx', "euler_cython_numpy.c"],
include_dirs=[numpy.get_include()]
),
],
)
# -
# * Now build the module.
# + language="bash"
# python setup.py build_ext --inplace
#
#
# -
# * Let's import the cython c wrap module, run the simulation, and plot the results.
# +
import pylab as py
import euler_cython_numpy as cwrap
# call the simulation function
t_cy_wrap, x_cy_wrap = cwrap.euler_cython_numpy_py_func(
t_start_sec,
t_end_sec,
delta_t_sec,
)
# plot result from python simulation
py.plot(t_py, x_py, 'o', label='python')
# plot result from cython + numpy
py.plot(t_cy_np, x_cy_np, '.', label='cython numpy')
# plot result from cython c wrapper function
py.plot(t_cy_wrap, x_cy_wrap, '-', label='cython wrap')
py.legend(loc=0)
py.grid(True)
py.show()
# -
# * Let's compare the calculation time again.
# +
# cython wrap with memory allocation
# -o makes the result available as output
# time_c_wrap_malloc = %timeit -n5 -r5 -o t_cy_wrap, x_cy_wrap = cwrap.euler_cython_numpy_py_func(t_start_sec, t_end_sec, delta_t_sec,)
# -
# * Clean up the files.
# + language="bash"
# rm euler_cython_numpy.h
# rm euler_cython_numpy.c
# rm _euler_cython_numpy.c
# rm _euler_cython_numpy.pyx
# rm setup.py
# rm euler_cython_numpy.*.so
#
#
# -
# * Check array type and dimension size
# +
assert isinstance(t_cy_wrap, py.ndarray), "t is not a numpy.ndarray"
assert isinstance(x_cy_wrap, py.ndarray), "x is not a numpy.ndarray"
# for now expect 1-dimensional arrays
assert 1 == t_cy_wrap.ndim, "t dimension > 1"
assert 1 == x_cy_wrap.ndim, "x dimension > 1"
# -
# * Check each element of the results
# +
for i in range(len(t_py)):
assert t_cy_np[i] == t_py[i], f"t[{i}] Cython NumPy Result f{t_cy_np[i]} != Python Result f{t_py[i]}\n"
assert x_cy_np[i] == x_py[i], f"x[{i}] Cython NumPy Result f{x_cy_np[i]} != Python Result f{x_py[i]}\n"
assert t_cy_wrap[i] == t_py[i], f"t[{i}] Cython C Wrap Result f{t_cy_wrap[i]} != Python Result f{t_py[i]}\n"
assert x_cy_wrap[i] == x_py[i], f"x[{i}] Cython C Wrap Result f{x_cy_wrap[i]} != Python Result f{x_py[i]}\n"
assert abs(t_py[-1] - t_end_sec) < (0.5 * delta_t_sec), f"t_end_sec = {t_end_sec}, t_py[-1] = {t_py[-1]}"
assert abs(t_cy_np[-1] - t_end_sec) < (0.5 * delta_t_sec), f"t_end_sec = {t_end_sec}, t_cy_np[-1] = {t_cy_np[-1]}"
assert abs(t_cy_wrap[-1] - t_end_sec) < (0.5 * delta_t_sec), f"t_end_sec = {t_end_sec}, t_cy_wrap[-1] = {t_cy_wrap[-1]}"
# -
# * Let's try different simulation time
# +
import pylab as py
import euler_cython_numpy as cwrap
t_start_2_sec = 0.0
t_end_2_sec = 1.0
delta_t_2_sec = 2.0 ** (-10)
# cython numpy
t_2_np, x_2_np = euler_cython_numpy_array(
t_start_2_sec, t_end_2_sec, delta_t_2_sec,
)
# python list
t_2_py, x_2_py = euler_python(
t_start_2_sec, t_end_2_sec, delta_t_2_sec,
)
# cython C wrap
t_2_cy_wrap, x_2_cy_wrap = cwrap.euler_cython_numpy_py_func(
t_start_2_sec, t_end_2_sec, delta_t_2_sec,
)
# plots
py.plot(t_2_py, x_2_py, 'o', label='python list')
py.plot(t_2_np, x_2_np, '.', label='cython numpy')
py.plot(t_2_cy_wrap, x_2_cy_wrap, '-', label='cython wrap')
py.legend(loc=0)
py.grid(True)
py.show()
# -
# * Let's check if cython numpy result correct
# +
for i in range(len(t_2_py)):
assert t_2_np[i] == t_2_py[i], f"t[{i}] Cython NumPy Result f{t_2_np[i]} != Python Result f{t_2_py[i]}\n"
assert x_2_np[i] == x_2_py[i], f"x[{i}] Cython NumPy Result f{x_2_np[i]} != Python Result f{x_2_py[i]}\n"
assert t_2_cy_wrap[i] == t_2_py[i], f"t[{i}] Cython C Wrap Result f{t_2_cy_wrap[i]} != Python Result f{t_2_py[i]}\n"
assert x_2_cy_wrap[i] == x_2_py[i], f"x[{i}] Cython C Wrap Result f{x_2_cy_wrap[i]} != Python Result f{x_2_py[i]}\n"
assert abs(t_2_py[-1] - t_end_2_sec) < (0.5 * delta_t_2_sec), f"t_end_2_sec = {t_end_2_sec}, t_2_py[-1] = {t_2_py[-1]}"
assert abs(t_2_np[-1] - t_end_2_sec) < (0.5 * delta_t_2_sec), f"t_end_2_sec = {t_end_2_sec}, t_2_np[-1] = {t_2_np[-1]}"
assert abs(t_2_cy_wrap[-1] - t_end_2_sec) < (0.5 * delta_t_2_sec), f"t_end_2_sec = {t_end_2_sec}, t_2_cy_wrap[-1] = {t_2_cy_wrap[-1]}"
# -
# * Let's compare the computation times of different versions:
# +
import IPython.display as disp
disp.display(
disp.Markdown(
'\n'.join([
"| type | result |",
"|:-----:|:-----:|",
f"| cython c wrap (with memory allocation) | {time_c_wrap_malloc} |",
f"| cython dynamic NumPy | {time_cython_dynamic_numpy} |",
f"| python list | {time_python} |",
]
)
)
)
# -
# ## Python Control Open Source Project
# * Nowadays in late 2010s, there is an open source project of [**python-control**](https://github.com/python-control/).
# * The objective is to develop a useful tool for control engineers in python.
# * One of the challenges is a numerical library named [**Slycot**](https://github.com/python-control/Slycot) mostly in Fortran.
# * Slycot uses other numerical libraries such as **BLAS** and **LAPACK** also in Fortran.
# * It would be noteworthy that `numpy` of SciPy Stack also includes **BLAS** and **LAPACK**.
# * Please refer to [**python-control**](https://github.com/python-control/python-control) and [**Slycot**](https://github.com/python-control/Slycot) for more information.
# ## Exercise
# ### 00 Allocation in `.pyx`
# * In the cosine example, see if you can pre-allocate output array in `.pyx` file.
# ### 01 Other numerical methods
# * From the Euler's method example above, implement other numerical methods and compare the performance.
| 10.ipynb |