code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GPU-Jupyter
#
# This Jupyterlab Instance is connected to the GPU via CUDA drivers. In this notebook, we test the installation and perform some basic operations on the GPU.
# ## Test GPU connection
#
# #### Using the following command, your GPU type and its NVIDIA-SMI driver version should be listed:
# !nvidia-smi
# #### Now, test if PyTorch can access the GPU via CUDA:
import torch
torch.cuda.is_available()
import tensorflow as tf
from tensorflow.python.client import device_lib
print(tf.config.list_physical_devices('XLA_GPU'))
device_lib.list_local_devices()
from __future__ import print_function
import numpy as np
import torch
a = torch.rand(5, 3)
a
# ## Performance test
#
# #### Now we want to know how much faster a typical operation is using GPU. Therefore we do the same operation in numpy, PyTorch and PyTorch with CUDA. The test operation is the calculation of the prediction matrix that is done in a linear regression.
# ### 1) Numpy
x = np.random.rand(10000, 256)
# %%timeit
H = x.dot(np.linalg.inv(x.transpose().dot(x))).dot(x.transpose())
# ### 2) PyTorch
x = torch.rand(10000, 256)
# %%timeit
# Calculate the projection matrix of x on the CPU
H = x.mm( (x.t().mm(x)).inverse() ).mm(x.t())
# ### 3) PyTorch on GPU via CUDA
# let us run this cell only if CUDA is available
# We will use ``torch.device`` objects to move tensors in and out of GPU
if torch.cuda.is_available():
device = torch.device("cuda") # a CUDA device object
x = torch.rand(10000, 256, device=device) # directly create a tensor on GPU
y = x.to(device) # or just use strings ``.to("cuda")``
print(x[0:5, 0:5])
print(y.to("cpu", torch.double)[0:5, 0:5])
# %%timeit
# Calculate the projection matrix of x on the GPU
H = x.mm( (x.t().mm(x)).inverse() ).mm(x.t())
# ## Exhaustive Testing on GPU
# let us run this cell only if CUDA is available
# We will use ``torch.device`` objects to move tensors in and out of GPU
import torch
if torch.cuda.is_available():
device = torch.device("cuda") # a CUDA device object
x = torch.rand(10000, 10, device=device) # directly create a tensor on GPU
if torch.cuda.is_available():
y = x.to(device) # or just use strings ``.to("cuda")``
print(x[0:5, 0:5])
if torch.cuda.is_available():
# Here is the memory of the GPU a border.
# A matrix with 100000 lines requires 37 GB, but only 8 GB are available.
H = x.mm( (x.t().mm(x)).inverse() ).mm(x.t())
if torch.cuda.is_available():
print(H[0:5, 0:5])
if torch.cuda.is_available():
# This operation is difficult, as an symmetric matrix is transferred
# back to the CPU. Is possible up to 30000 rows.
print(H.to("cpu", torch.double)[0:5, 0:5])
| extra/Getting_Started/GPU-processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # COCO Reader
#
# Reader operator that reads a COCO dataset (or subset of COCO), which consists of an annotation file and the images directory.
# +
from __future__ import print_function
from nvidia.dali.pipeline import Pipeline
import nvidia.dali.ops as ops
import nvidia.dali.types as types
import numpy as np
from time import time
subset = "val"
file_root = "/data/coco/coco-2017/coco2017/" + subset + "2017"
annotations_file = "/data/coco/coco-2017/coco2017/annotations/instances_" + subset + "2017.json"
num_gpus = 1
batch_size = 16
# -
class COCOPipeline(Pipeline):
def __init__(self, batch_size, num_threads, device_id):
super(COCOPipeline, self).__init__(batch_size, num_threads, device_id, seed = 15)
self.input = ops.COCOReader(file_root = file_root, annotations_file = annotations_file,
shard_id = device_id, num_shards = num_gpus, ratio=True)
self.decode = ops.nvJPEGDecoder(device = "mixed", output_type = types.RGB)
def define_graph(self):
inputs, bboxes, labels = self.input()
images = self.decode(inputs)
return (images, bboxes, labels)
start = time()
pipes = [COCOPipeline(batch_size=batch_size, num_threads=2, device_id = device_id) for device_id in range(num_gpus)]
for pipe in pipes:
pipe.build()
total_time = time() - start
print("Computation graph built and dataset loaded in %f seconds." % total_time)
# +
pipe_out = [pipe.run() for pipe in pipes]
images_cpu = pipe_out[0][0].asCPU()
bboxes_cpu = pipe_out[0][1]
labels_cpu = pipe_out[0][2]
# -
# Bounding boxes returned by the operator are lists of floats containing composed of **\[x, y, width, height]** (`ltrb` is set to `False` by default).
bboxes = bboxes_cpu.at(4)
bboxes
# Let's see the ground truth bounding boxes drawn on the image.
# +
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import random
img_index = 4
img = images_cpu.at(img_index)
H = img.shape[0]
W = img.shape[1]
fig,ax = plt.subplots(1)
ax.imshow(img)
bboxes = bboxes_cpu.at(img_index)
labels = labels_cpu.at(img_index)
categories_set = set()
for label in labels:
categories_set.add(label[0])
category_id_to_color = dict([ (cat_id , [random.uniform(0, 1) ,random.uniform(0, 1), random.uniform(0, 1)]) for cat_id in categories_set])
for bbox, label in zip(bboxes, labels):
rect = patches.Rectangle((bbox[0]*W,bbox[1]*H),bbox[2]*W,bbox[3]*H,linewidth=1,edgecolor=category_id_to_color[label[0]],facecolor='none')
ax.add_patch(rect)
plt.show()
| docs/examples/coco_reader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Are we in SWAN?
# Working in the right path
# %cd /eos/project/d/da-and-diffusion-studies/DA_Studies/Simulations/Models/da_sixtrack/for_martin
# Install the libraries
import sys
# !{sys.executable} -m pip install --user sixtrackwrap
# !export PYTHONPATH=$CERNBOX_HOME/.local/lib/python3.7/site-packages:$PYTHONPATH
# For this "presentation" only!
import warnings
warnings.filterwarnings('ignore')
# # Taking advantage of the step-by-step tracking
# SixTrackLib allows the creation of track jobs properly allocated on the GPU device. With these jobs, we can efficiently gather the characteristics of every particle turn after turn efficiently without loosing the GPU parallel capabilites, as the data can be gathered from the GPU directly with an optimized memory access.
#
# Thanks to that, we were able to implement some first full-track analysis of particle transversal dynamics, for inspecting the presence of resonances for specific initial conditions.
#
# In this notebook, we show just the most "colorful" plots we have made in our preliminary analysis.
#
# ## Backend setup and libraries
# %matplotlib widget
# +
# Base libraries
import math
import numpy as np
import scipy.integrate as integrate
from scipy.special import erf
import pickle
import itertools
from scipy.optimize import curve_fit
from numba import njit, prange
# Personal libraries
#import sixtrackwrap_light as sx
import sixtrackwrap as sx
from tqdm.notebook import tqdm
import time
import matplotlib.pyplot as plt
import ipywidgets as widgets
from IPython.display import display
import matplotlib
import matplotlib.ticker as ticker
from math import gcd
from scipy.special import lambertw
# -
# ## Some step-by-step trackings
# Here we have performed in a separate instance a radial scan of various angular coordinates. For each angle, given a value $N_\text{max}$ of turns, we look for the last stable particle after $N_\text{max}$ turns.
#
# Once we have found it, we re-track the particle while saving its whole transversal path. We then analyze its motion characteristics in the 4D space considering normalized polar coordinates $(r, \alpha, \theta_1, \theta_2)$.
#
# In this specific plot, we analyze how the particle "explores" the $(\theta_1, \theta_2)$ space by considering a 2D binning of the $N_\text{max}$ steps the particle takes before becoming lost.
#
# In the left plot, we show for each bin the average radial distance the particle has for that specific $(\theta_1, \theta_2)$ interval. The white bins indicates a NaN value, meaning that the particle has not visited that specific interval.
#
# In the right plot, we show for each bin the number of times the particle has visited that specific $(\theta_1, \theta_2)$ interval.
#
# With the slider, you can select the $\alpha_0$ angle as starting considition. In this setup, both $\theta$ angles are set to 0 as starting condition.
with open("data/matrices_2.pkl", 'rb') as f:
count_matrix, avg_matrix = pickle.load(f)
samples = 2049
n_subdivisions = 128
max_turns = 10000
count_total, average_total, result_total, validity_total = sx.recursive_accumulation(count_matrix, avg_matrix)
alpha_preliminary_values = np.linspace(-1.0, 1.0, samples)
alpha_values = np.arccos(alpha_preliminary_values) / 2
# +
fig2 = plt.figure()
def update_scheme(j):
fig2.clear()
axs2 = fig2.subplots(1, 2)
i = 0
j = len(count_total[0]) - 1 - j
coso = axs2[0].imshow(average_total[i][j], origin="lower", extent=(0, np.pi*2, 0, np.pi*2))
axs2[0].set_title("Average radius measured\n$\\alpha = {:.4}\pi$".format(alpha_values[j]/np.pi))
axs2[0].set_xlabel("$\\theta_1$")
axs2[0].set_ylabel("$\\theta_2$")
cb = fig2.colorbar(coso, ax=axs2[0])
#cb.ax.plot([0, 100], [DA_2[(samples, 'refined', 'mc')][0]]*2, 'w')
axs2[0].xaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: str(int(x/np.pi)) + "$\\pi$"))
axs2[0].xaxis.set_major_locator(ticker.MultipleLocator(base=np.pi))
axs2[0].yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: str(int(x/np.pi)) + "$\\pi$"))
axs2[0].yaxis.set_major_locator(ticker.MultipleLocator(base=np.pi))
coso = axs2[1].imshow(count_total[i][j], origin="lower", extent=(0, np.pi*2, 0, np.pi*2), vmin=0)
axs2[1].set_title("Number of samples\n$\\alpha = {:.4}\pi$".format(alpha_values[j]/np.pi))
axs2[1].set_xlabel("$\\theta_1$")
axs2[1].set_ylabel("$\\theta_2$")
fig2.colorbar(coso, ax=axs2[1])
axs2[1].xaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: str(int(x/np.pi)) + "$\\pi$"))
axs2[1].xaxis.set_major_locator(ticker.MultipleLocator(base=np.pi))
axs2[1].yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: str(int(x/np.pi)) + "$\\pi$"))
axs2[1].yaxis.set_major_locator(ticker.MultipleLocator(base=np.pi))
fig2.suptitle("2D binning $(128\\times 128)$ over the $(\\theta_1, \\theta_2)$ space of a particle tracked for 10000 turns.")
k=widgets.IntSlider(value=0, min=0, max=len(count_total[0]) - 1, step=1)
ui2 = widgets.VBox([
widgets.Label("Sample index to visualize"), k,
])
out2 = widgets.interactive_output(
update_scheme,
{"j":k}
)
display(ui2, out2)
# -
# As you can see from the plot above, different $\alpha$ starting conditions implies very different behaviours for the particle transversal dynamics. And it's in our interest to inspect the charatcerstics of these resonances.
#
# A first qualitative measurement of these resonance behaviours is to evaluate the percentage of empty bins in the left plots above: more empty plots implies less uniform diffusion in the $(\theta_1, \theta_2)$ space and, therefore, stronger resonace effects.
#
# We plot this measure down here.
# +
count_total, average_total, result_total, validity_total = sx.recursive_accumulation(count_matrix, avg_matrix)
alpha_preliminary_values = np.linspace(-1.0, 1.0, samples)
alpha_values = np.arccos(alpha_preliminary_values) / 2
nan_data = []
for j in range(0, len(count_total[2])):
nan_data.append(np.count_nonzero(np.isnan(average_total[2][j])) / ((n_subdivisions / 4) ** 2))
nan_data = np.asarray(nan_data)
x = np.cos(alpha_values[::4]) * nan_data[::4]
y = np.sin(alpha_values[::4]) * nan_data[::4]
x = np.append(x, [0.0])
y = np.append(y, [0.0])
#plt.fill(x, y)
fig, ax = plt.subplots(figsize=(16,9))
ax.plot(alpha_values, nan_data, linewidth=0.2, c="C0", alpha=0.5)
ax.scatter(alpha_values, nan_data, s=0.75)
ax.xaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: "{:2}".format(x/np.pi) + "$\\ \\pi$"))
ax.xaxis.set_major_locator(ticker.MultipleLocator(base=np.pi/8))
ax.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: "{}".format(int(x * 100)) + "$\\%$"))
ax.yaxis.set_major_locator(ticker.MultipleLocator(base=0.2))
ax.set_xlabel("$\\alpha$ angle")
ax.set_ylabel("Percentage of empty bins")
ax.set_title("Percentage of empty bins for different intial $\\alpha$ angles. $N$ bins $= ({}\\times{}) = {}$, $N$ turns $= {}$\n(Higher percentage implies less `diffusion')".format(n_subdivisions // 4, n_subdivisions // 4, (n_subdivisions // 4) ** 2, max_turns))
ax.set_xlim(0, np.pi/2)
# -
# These analysis are made possible by the fact that SixTrackLib makes the execution of single parallelized tracking steps extremely easy and well optimized.
| 20200617_hss_section_meeting_sixtracklib/example_01_da_stability_resonance/resonance_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Test run of the PReLIM Net API.
# Author: <NAME>
# -
from PReLIM import PReLIM
import numpy as np
import time
from tqdm import tqdm
import matplotlib.pyplot as plt
# %matplotlib inline
# Automatic reloading so we don't have to restart the kernel each time we make a change
# to CpGNet when we are debuggin
# %load_ext autoreload
# %autoreload 2
# ### Step 1: Collect some data
# +
# Some bins with 2 cpgs
bin1 = np.array([[1,0],[0,-1],[-1,1],[0,0]],dtype=float)
bin2 = np.array([[1,0],[1,0],[-1,1],[0,0],[0,1],[1,1],[0,0]],dtype=float)
bin3 = np.array([[-1,1],[0,-1],[-1,1],[0,0]],dtype=float)
bin4 = np.array([[1,-1],[0,1],[-1,1],[1,0]],dtype=float)
bin5 = np.array([[1,1],[0,0],[0,1],[1,1]],dtype=float)
bin6 = np.array([[1,1],[1,1],[0,1],[1,0]],dtype=float)
bin7 = np.array([[0,0],[1,0],[0,1],[1,1]],dtype=float)
# Collection of bins
bins = [bin1,bin2,bin3,bin4, bin5, bin6, bin7]
# -
# ### Step 2: Create a model and train on the data. CpGNet will take care of feature extraction and all that other boring stuff, it just works!
# +
model = PReLIM(cpgDensity=2)
# 3 options for training/saving model
model.train(bins, model_file="no") # don't want a model file, must use "no"
#model.train(bins, model_file=None) # use a generic title w/ timestamp for the model file
#model.train(bins, model_file="my custom file") # custom file name
# -
# ### Step 3: Impute! (imputation result might look strange here since training data size is small)
model.impute(bin1)
bin3
model.impute(bin3)
binA = np.array([[0,0],[0,0],[0,-1],[0,0],[0,0]],dtype=float)
binB = np.array([[1,1],[1,1],[1,-1],[1,1],[1,1]],dtype=float)
binB
model.impute(binB)
X,y = model.get_X_y(bins)
model.predict(X)
model.model.predict_proba
predicted_matrices = model.impute_many(bins)
# # Is batch imputation faster?
test_bins_100 = bins[:4] * 25
test_bins_1000 = bins[:4] * 250
len(test_bins_100)
# +
individual_data = []
for i in tqdm(range(20)):
start = time.time()
for bin_ in test_bins_1000:
model.impute(bin_)
end = time.time()
duration = end-start
individual_data.append(duration)
# -
batch_data
# +
batch_data = []
for i in tqdm(range(20)):
start = time.time()
model.impute_many(test_bins_1000)
end = time.time()
end = time.time()
duration = end-start
batch_data.append(duration)
# -
plt.boxplot([individual_data,batch_data]);
plt.xticks([1,2],["Individual","Batch"])
#plt.ylim([0,2.5])
plt.ylabel("Seconds")
plt.title("Imputation Speed Comparison on 1000 bins (20 trials)")
#plt.savefig("batch_speed.png",dpi=500)
np.mean(individual_data)
np.mean(batch_data)
np.mean(individual_data)/np.mean(batch_data)
| .ipynb_checkpoints/PReLIM APITest-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
#import seaborn as sns
#sns.set(color_codes=True)
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
import numpy as np
import h5py
import cv2
from sklearn.decomposition import PCA
from scipy.stats.stats import pearsonr
#read movie data sky
def read_movie_from_h5(filename):
h5f = h5py.File(filename,'r')
movie_bgr_h5=h5f['movie_bgr_real'][:]
h5f.close()
return movie_bgr_h5
movie_bgr_h5=np.concatenate((read_movie_from_h5('Data/all_image_patches_lower_1f_28x28_bg_20200512.h5'),\
read_movie_from_h5('Data/all_image_patches_upper_1f_28x28_bg_20200512.h5')),axis=0)
print (movie_bgr_h5.shape)
#
def img_real2view(img):
gamma_correction=lambda x:np.power(x,1.0/2.2)
img_shape=img.shape
# gray image
if np.size(img_shape)==2:
#uint8
if np.max(img)>1:
temp_view=np.zeros_like(img,dtype=np.float32)
temp_view=np.float32(img)/255.0#float32, 1.0
temp_view=gamma_correction(temp_view)
temp_view2=np.zeros_like(img,dtype=np.uint8)
temp_view2=np.uint8(temp_view*255)
return temp_view2
#float
if np.max(img)<2:
return gamma_correction(img)
#color image
if np.size(img_shape)==3:
#uint8
if np.max(img)>1:
temp_view=np.zeros_like(img,dtype=np.float32)
temp_view=np.float32(img)/255.0#1.0
temp_view=gamma_correction(temp_view)
temp_view2=np.zeros_like(img,dtype=np.uint8)
temp_view2=np.uint8(temp_view*255)#255
return temp_view2
#float
if np.max(img)<2:
return gamma_correction(img)
#
fig, axes = plt.subplots(nrows=1, ncols=10,figsize=(12.5,2.5))
for ii in range(10):
temp=np.zeros((28,28,3),np.uint8)
temp[...,1:]=movie_bgr_h5[5000+ii,:,:,::-1]
axes[ii].imshow(img_real2view(temp))
axes[ii].grid(False)#axes[ii,jj].axis('tight')
#input: X, shape (samplesize*nfeatures)
#output: the specific eigenvectors
def pca_image_crops(X,n_components,seed=99):
pca = PCA(n_components=n_components, svd_solver='randomized',random_state=seed)
pca.fit(X)
eigvecs=[]
for ii in range(n_components):
eigvecs.append(pca.components_[ii])
eigvecs=np.asarray(eigvecs)
eigvals=pca.explained_variance_ratio_
return eigvecs,eigvals
def pre_images(images):
temp2 = images - images.mean(axis=0) #global centering
temp2 -= temp2.mean(axis=1).reshape(images.shape[0], -1) #local centering
return temp2
def zca_whitening_matrix(X):
"""
Function to compute ZCA whitening matrix (aka Mahalanobis whitening).
INPUT: X: [M x N] matrix; shape (nfeatures*samplesize)
Rows: Variables
Columns: Observations
OUTPUT: ZCAMatrix: [M x M] matrix
"""
sigma = np.cov(X, rowvar=True) # [M x M]
U,S,V = np.linalg.svd(sigma)
epsilon = 1e-8
ZCAMatrix = np.dot(U, np.dot(np.diag(1.0/np.sqrt(S + epsilon)), U.T)) # [M x M]
return ZCAMatrix
# ## <font color='red'>Upper</font>
#get 50000 small image crops
num_size=50000
np.random.seed(66)
fnum=np.random.randint(10000,19000,size=num_size)#frame start number
np.random.seed(88)
hnum=np.random.randint(0,28-9,size=num_size)#height start number
np.random.seed(99)
wnum=np.random.randint(0,28-9,size=num_size)#weight start number
#
all_images=[]
for ii in range(num_size):
all_images.append(movie_bgr_h5[fnum[ii],hnum[ii]:hnum[ii]+9,wnum[ii]:wnum[ii]+9,:])
all_images=np.asarray(all_images)
print (all_images.shape)
#
fig, axes = plt.subplots(nrows=1, ncols=10,figsize=(12.5,2.5))
for ii in range(10):
temp=np.zeros((9,9,3),np.uint8)
temp[...,1:]=all_images[5000+ii*100,:,:,::-1]
axes[ii].imshow(img_real2view(temp))
axes[ii].grid(False)
#pca
all_images=np.reshape(all_images,(len(all_images),-1))
if np.max(all_images)>1:
all_images=all_images/255.0
print (all_images.shape)
eigvecs,eigvals=pca_image_crops(all_images-np.mean(all_images,axis=0,keepdims=True),162)
print (eigvecs.shape)
print (eigvals)
#
#visualize weights
eigvecs=np.reshape(eigvecs,(162,9,9,2))
print (eigvecs.shape)
print (eigvecs[:6,0,0,0])
#
fig, axes = plt.subplots(nrows=2, ncols=16,figsize=(16,2))
for ii in range(16):
temp=eigvecs[ii]
vmax=np.max(np.abs(temp[:,:,:]))
axes[0,ii].imshow(temp[:,:,0],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[0,ii].set_xticks([])
axes[0,ii].set_yticks([])
#vmax=np.max(np.abs(temp[:,:,1]))
axes[1,ii].imshow(temp[:,:,1],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[1,ii].set_xticks([])
axes[1,ii].set_yticks([])
# + tags=[]
#CC between UV and G weights
print (eigvals)
CCs=np.zeros((len(eigvecs)))
pVs=np.zeros((len(eigvecs)))
for ii in range(len(eigvecs)):
CCs[ii],pVs[ii]=pearsonr(eigvecs[ii,...,0].flatten(),eigvecs[ii,...,1].flatten())
#
print (CCs)
print (pVs)
opp_ind=np.where(CCs<0)[0].tolist()
pV_ind =np.where(pVs<0.05)[0].tolist()
opp_vP_ind=[opp_ind,pV_ind]
#
opp_pV=set(opp_vP_ind[0]).intersection(*opp_vP_ind)
print (len(opp_pV))
print (opp_pV)
print ([x for x in opp_pV if x<16])
print (np.array([eigvals[x] for x in opp_pV]).sum())
# -
#zca
all_images=np.reshape(all_images,(len(all_images),-1))
if np.max(all_images)>1:
all_images=all_images/255.0
print (all_images.shape)
eigvecs=zca_whitening_matrix((all_images-np.mean(all_images,axis=0,keepdims=True)).T)
print (eigvecs.shape)
print ('Symmetrical?: {}'.format(np.all(np.abs(eigvecs-eigvecs.T) < 1e-8)))
#
#visualize weights,
eigvecs=np.reshape(eigvecs,(162,9,9,2))
print (eigvecs.shape)
print (eigvecs[:6,0,0,0])
#
np.random.seed(22)
nums=np.random.choice(len(eigvecs),16,replace=False)
fig, axes = plt.subplots(nrows=2, ncols=16,figsize=(16,2))
for ii in range(16):
temp=eigvecs[nums[ii]]
vmax=np.max(np.abs(temp[:,:,:]))
axes[0,ii].imshow(temp[:,:,0],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[0,ii].set_xticks([])
axes[0,ii].set_yticks([])
axes[1,ii].imshow(temp[:,:,1],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[1,ii].set_xticks([])
axes[1,ii].set_yticks([])
#CC between UV and G weights
CCs=np.zeros((len(eigvecs)))
pVs=np.zeros((len(eigvecs)))
for ii in range(len(eigvecs)):
CCs[ii],pVs[ii]=pearsonr(eigvecs[ii,...,0].flatten(),eigvecs[ii,...,1].flatten())
#
print (CCs)
print (pVs)
opp_ind=np.where(CCs<0)[0].tolist()
pV_ind =np.where(pVs<0.05)[0].tolist()
opp_vP_ind=[opp_ind,pV_ind]
opp_pV=set(opp_vP_ind[0]).intersection(*opp_vP_ind)
print (len(opp_pV))
print (opp_pV)
# ## <font color='red'>Lower</font>
#get 50000 small image crops (9x9 pixels)
num_size=50000
np.random.seed(66)
fnum=np.random.randint(0,9000,size=num_size)#frame start number
np.random.seed(88)
hnum=np.random.randint(0,28-9,size=num_size)#height start number
np.random.seed(99)
wnum=np.random.randint(0,28-9,size=num_size)#weight start number
#
all_images=[]
for ii in range(num_size):
all_images.append(movie_bgr_h5[fnum[ii],hnum[ii]:hnum[ii]+9,wnum[ii]:wnum[ii]+9,:])
all_images=np.asarray(all_images)
print (all_images.shape)
#
fig, axes = plt.subplots(nrows=1, ncols=10,figsize=(12.5,2.5))
for ii in range(10):
temp=np.zeros((9,9,3),np.uint8)
temp[...,1:]=all_images[5000+ii*100,:,:,::-1]
axes[ii].imshow(img_real2view(temp))
axes[ii].grid(False)
#pca
all_images=np.reshape(all_images,(len(all_images),-1))
if np.max(all_images)>1:
all_images=all_images/255.0
print (all_images.shape)
eigvecs,eigvals=pca_image_crops(all_images-np.mean(all_images,axis=0,keepdims=True),162)
print (eigvecs.shape)
print (eigvals)
#
#visualize weights
eigvecs=np.reshape(eigvecs,(162,9,9,2))
print (eigvecs.shape)
print (eigvecs[:6,0,0,0])
#
fig, axes = plt.subplots(nrows=2, ncols=16,figsize=(16,2))
for ii in range(16):
temp=eigvecs[ii]
vmax=np.max(np.abs(temp[:,:,:]))
axes[0,ii].imshow(temp[:,:,0],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[0,ii].set_xticks([])
axes[0,ii].set_yticks([])
axes[1,ii].imshow(temp[:,:,1],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[1,ii].set_xticks([])
axes[1,ii].set_yticks([])
# + tags=[]
#CC between UV and G weights
print (eigvals)
CCs=np.zeros((len(eigvecs)))
pVs=np.zeros((len(eigvecs)))
for ii in range(len(eigvecs)):
CCs[ii],pVs[ii]=pearsonr(eigvecs[ii,...,0].flatten(),eigvecs[ii,...,1].flatten())
#
print (CCs)
print (pVs)
opp_ind=np.where(CCs<0)[0].tolist()
pV_ind =np.where(pVs<0.05)[0].tolist()
opp_vP_ind=[opp_ind,pV_ind]
opp_pV=set(opp_vP_ind[0]).intersection(*opp_vP_ind)
print (len(opp_pV))
print (opp_pV)
print ([x for x in opp_pV if x<16])
print (np.array([eigvals[x] for x in opp_pV]).sum())
# -
#zca
all_images=np.reshape(all_images,(len(all_images),-1))
if np.max(all_images)>1:
all_images=all_images/255.0
print (all_images.shape)
eigvecs=zca_whitening_matrix((all_images-np.mean(all_images,axis=0,keepdims=True)).T)
print (eigvecs.shape)
print ('Symmetrical?: {}'.format(np.all(np.abs(eigvecs-eigvecs.T) < 1e-8)))
#
#visualize weights,
eigvecs=np.reshape(eigvecs,(162,9,9,2))
print (eigvecs.shape)
print (eigvecs[:6,0,0,0])
#
np.random.seed(22)
nums=np.random.choice(len(eigvecs),16,replace=False)
fig, axes = plt.subplots(nrows=2, ncols=16,figsize=(16,2))
for ii in range(16):
temp=eigvecs[nums[ii]]
vmax=np.max(np.abs(temp[:,:,:]))
axes[0,ii].imshow(temp[:,:,0],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[0,ii].set_xticks([])
axes[0,ii].set_yticks([])
axes[1,ii].imshow(temp[:,:,1],vmax=vmax,vmin=-vmax,cmap='bwr')
axes[1,ii].set_xticks([])
axes[1,ii].set_yticks([])
#CC between UV and G weights
CCs=np.zeros((len(eigvecs)))
pVs=np.zeros((len(eigvecs)))
for ii in range(len(eigvecs)):
CCs[ii],pVs[ii]=pearsonr(eigvecs[ii,...,0].flatten(),eigvecs[ii,...,1].flatten())
#
print (CCs)
print (pVs)
opp_ind=np.where(CCs<0)[0].tolist()
pV_ind =np.where(pVs<0.05)[0].tolist()
opp_vP_ind=[opp_ind,pV_ind]
opp_pV=set(opp_vP_ind[0]).intersection(*opp_vP_ind)
print (len(opp_pV))
print (opp_pV)
| code/pca_zca_image_crops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
import requests
import pdfplumber
def download_file(url):
local_filename = url.split('/')[-1]
with requests.get(url) as r:
with open(local_filename, 'wb') as f:
f.write(r.content)
return local_filename
invoice_url = 'https://isaw.nyu.edu/guide/forms/sample-vendor-invoice'
invoice_url = download_file(invoice_url)
with pdfplumber.open(invoice_url) as pdf:
page = pdf.pages[0]
text = page.extract_text()
print(text)
for row in text.split('\n'):
if row.startswith('TOTAL'):
balance = row.split()
total = balance[-2]+balance[-1]
print('$', total)
| Accounting Internship/Extract data from PDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Suggested admonitions
#
# The following admonitions are suggested to be used in Sabueso's documentation
#
#
# ## Note
#
# The "Note" messages are used to include notes to the main text in the documentation. These notes could be, for instance, ...
#
# Notes shouldn't be skipped by the reader. Please **do not use the 'dropdown' option**.
#
# ````
# ```{admonition} Note
# :class: note
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
# ````
#
# ```{admonition} Note
# :class: note
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
#
#
# ## Did you know...?
#
# This kind of messages includes "academic" information related with a topic mentioned in the documentation. The content will not be about Sabueso's syntaxis, methods, or use, but with scientific comments which might be useful to understand the context or enrich the users' culture.
#
# Given that these messages are not 'essential' to follow the documentation, their text will not be shown by default. The box can be unfolded using the "Click to show" button on the right.
#
# ````md
# ```{admonition} Did you know...?
# :class: tip, dropdown
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
# ````
#
# ```{admonition} Did you know...?
# :class: tip, dropdown
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
#
#
# ## It is suggested...
#
# This kind of messages includes "academic" or "tecnical" information the reader should check, or activities or experiments the reader should try.
#
# Given that these messages are not 'essential' to follow the documentation, their text will not be shown by default. The box can be unfolded using the "Click to show" button on the right.
#
# ````md
# ```{admonition} It is suggested...
# :class: tip, dropdown
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
# ````
#
# ```{admonition} It is suggested...
# :class: tip, dropdown
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
#
#
# ## See also
#
# The "See also" messages are used in Sabueso to report those classes, methods, attributes from Sabueso's API, or those sections of Sabueso's documentations, the reader should check at the end of a section.
#
# The "See also" message must be explicitly shown in the documentation (**Do not use the 'dropdown' option**):
#
#
# ````md
# ```{admonition} See also
# :class: attention
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
# ````
#
# ```{admonition} See also
# :class: attention
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
#
#
# ## Warning
#
# Use the "Warning" boxes to communicate messages the users should know to avoid errors, the missuse of sabueso's classes, methods and attributes, or the missinterpretations of Sabueso's output.
#
# These messages must be read by every user. Please **do not use the 'dropdown' option**.
#
# ````
# ```{admonition} Warning
# :class: danger
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
# ````
#
# ```{admonition} Warning
# :class: danger
# Lorem ipsum dolor sit amet, consectetur adipiscing elit.
# ```
| docs/contents/developer/documentation/suggested_admonitions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Examples of Multicore Applications:
# # Shared-Memory Multiprocess Applications
# In this notebook we look at multiprocess applications in IoTPy. The processes share memory. Associated with each process is an agent. The application can also have:
# <ol>
# <li> source threads that acquire data from external sources and </li>
# <li> actuator threads the get data from output queues. </li>
# </ol>
# <b> The central idea is that an application is specified by connecting outputs of processes to inputs of processes.</b>
#
# ## The Agent associated with a Process
# A process in a multicore application executes an agent with the following signature:
# <br>
# <br>
# <b>f(in_streams, out_streams)</b>
# <br>
# <br>
# where:
# <ol>
# <li> <i>f</i> is a function. </li>
# <li> <i>in_streams</i> is a list of input streams. </li>
# <li> <i>out_streams</i> is a list of output streams. </li>
# </ol>
# +
import sys
import time
import threading
sys.path.append("../")
from IoTPy.core.stream import Stream, StreamArray, run
from IoTPy.agent_types.op import map_element, map_list, map_window
from IoTPy.helper_functions.recent_values import recent_values
from IoTPy.helper_functions.print_stream import print_stream
from IoTPy.concurrency.multicore import get_processes, get_processes_and_procs
from IoTPy.concurrency.multicore import terminate_stream
from IoTPy.concurrency.multicore import get_proc_that_inputs_source
from IoTPy.concurrency.multicore import extend_stream
# -
# Next we show a collection of agents, <i>f</i>, <i>g</i>, <i>h</i>, and <i>r</i>, with this signature. We will use these agents in the examples of multicore programs.
# +
def f(in_streams, out_streams):
map_element(lambda v: v+100, in_streams[0], out_streams[0])
def g(in_streams, out_streams):
s = Stream('s')
map_element(lambda v: v*2, in_streams[0], s)
print_stream(s, 's')
def h(in_streams, out_streams):
map_element(lambda v: v*2, in_streams[0], out_streams[0])
def r(in_streams, out_streams):
t = Stream('t')
map_element(lambda v: v*3, in_streams[0], t)
print_stream(t, 't')
# -
# ## Threads
# A process may execute an arbitrary number of threads. You can use any thread target.
# <br>
# <br>
# Most threads in IoTPY applications pass data to the application or get data from the application. A thread that passes data from an external source, such as a sensor or a Twitter stream, to an IoTPy process is called a <b>source thread</b>.
# ### Source Threads
# A source thread calls the following function to put data into a stream:
# <br>
# <br>
# <b>extend_stream(procs, data, stream_name)</b>
# <br>
# <br>
# where
# <ol>
# <li> <i>procs</i> is a list of process metadata created from the specification of a multicore program. <i>procs</i> is passed as a parameter to the thread target. We will discuss <i>procs</i> later. </li>
# <li> <i>data</i> is a list or an array. </li>
# <li> <i>stream_name</i> is a string which is the name of a stream.</li>
# </ol>
# In the example, <i>source_thread_target</i>, the function has a single argument <i>procs</i>. All thread targets that extend streams must have <i>procs</i> as one of its arguments.
# <br>
# <br>
# This function executes a loop in which puts specified data into a stream called <i>x</i> and then sleeps thus yielding the thread.
# <br>
# <br>
# <b>terminate_stream</b>
#
# ### Sources
# A source in a multiprocess application is associated with a process. A source <i>s</i> in a process <i>p</i> is essentially an output stream of <i>p</i>; it differs from output streams created by <i>p</i> in the sense that it is fed by a thread rather than computed by <i>p</i>. However, we don't include <i>s</i> in the list of <b>outputs</b> of <i>p</i>; instead we include <i>s</i> in the list of <b>sources</b> of <i>p</i>.
# <br>
# <br>
# Note that a source thread that generates a source <i>s</i> in a process <i>p</i> can run in a different process <i>p'</i>. You want to choose the process in which to run a thread to balance the computational load across processes. If the output of a source <i>s</i> feeds input streams of exactly one process <i>r</i> then efficiency suggests that <i>s</i> should be a source of <i>r</i>; however, you can make <i>s</i> the source of any process.
#
# ## Steps to create a multiprocess application
# You may find the following steps helpful in creating a multiprocess application. You don't have to follow exactly these steps in this order.
# <br>
# <br>
# <i>Step 0</i>: <b>Define agent functions and source thread targets.</b>
# <ol>
# <li> <i>Step 0.0</i>: Each process has an agent associated with it, as described earlier. Specify the agent functions for each process. Recall that an agent function has the form:
# <br>
# <b>f(in_streams, out_streams)</b>.</li>
# <li><i>Step 0.1 </i> Define the thread targets for each source. These thread targets typically extend a source stream, and finally terminate the stream.</li>
# </ol>
# <i>Step 1: </i> <b>Give the multicore_specification of streams and processes.</b> The multicore specification specifies a list of streams and a list of agents.
# <br>
# <i>Step 2: </i> <b>Create processes</b> by calling:
# <br>
# processes, procs = get_processes_and_procs(multicore_specification)
# <br>
# <i>Step 3: </i> <b>Create threads</b> (if any). An example creation of a thread is:
# <br>
# thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# <br>
# <i>Step 4: </i> <b>Specify which process each thread runs in.</b> An example:
# <br>
# procs['p1'].threads = [thread_0]
# <br>
# <i>Step 5: </i>. <b>Start, join and terminate processes </b> by calling
# <br>
# for process in processes: process.start()
# for process in processes: process.join()
# for process in processes: process.terminate()
#
# Step 0: Define source thread target (if any).
# We will use this thread target for the next few examples.
def source_thread_target(procs):
for i in range(3):
extend_stream(procs, data=list(range(i*2, (i+1)*2)), stream_name='x')
time.sleep(0.001)
terminate_stream(procs, stream_name='x')
# ## Simple example of a multicore program
# 
#
# ### Multicore specification: Processes and their connecting streams
# Look at <b>multicore_specification</b>. The specification states that the program has two processes called p0 and p1. Process p0 has a single input stream <i>x</i> and a single output stream <i>y</i>. Process p1 has a single input stream <i>y</i> and no output streams. Thus, the output <i>y</i> of process p0 is the input of process p1.
# <br>
# ### Multicore specification: Streams
# Streams are specified by a list of pairs where each pair is a stream name and a stream type. The stream type 'i' identifies integers, 'f' floats and 'd' double. We use stream types to allow processes to share memory in Python 2.7+. In this example, the pair ('x', 'i') says that the program has a stream <i>x</i> of type int.
# <br>
# ### Multicore specification: Sources
# Process p0 has a <b>source_functions</b> called <i>h</i>. Function <i>h</i> executes in its own thread within process p0; this thread is started when the process is started. Function <i>h</i> has a single argument called <i>proc</i> which is a dummy argument that represents a process.
# <br>
# <br>
# Function <i>h</i> puts data into stream <i>x</i> when it executes <b>proc.copy_stream()</b>. The thread executing <i>h</i> then sleeps for 0.001 seconds before appending more data to stream <i>x</i>. Finally, the thread signals that the source has terminated appending data to stream <i>x</i> by calling <b>proc.finished_source('x')</b>.
# ### Process Structure
# The source <i>h</i> outputs a stream <i>x</i> which is an input of process p0. The output <i>y</i> of process p0 is an input to process p1.
# ### Process Computations
# The computation of a process is specified by a function with two arguments <i>in_streams</i> and <i>out_streams</i>. The computation carried out by p0 is specified by function <i>f</i> which reads a single input stream, <i>in_streams[0]</i> and write a single output stream, <i>out_streams[0]</i>. This agent makes:
# <br>
# <br>
# <b> y[n] = x[n] + 100 </b>
# <br>
# <br>
# The computation carried out by process p1 is specified by function <i>g</i> which prints <b>2 * y[n]</b> for all n.
# <br>
# <br>
# The source function <i>h</i> sets x[n] to n, and so this multicore process prints:
# <br>
# <br>
# <b> 2 * (n + 100) </b>
# +
# Step 1: multicore_specification of streams and processes.
multicore_specification = [
# Streams
[('x', 'i'), ('y', 'i')],
# Processes
[{'name': 'p0', 'agent': f, 'inputs':['x'], 'outputs': ['y'], 'sources':['x']},
{'name': 'p1', 'agent': g, 'inputs': ['y']}]]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any).
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread (if any) runs in.
# thread_0 runs in the process called 'p1'
procs['p1'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
# +
# Step 1: multicore_specification of streams and processes.
multicore_specification = [
# Streams
[('x', 'i'), ('y', 'i')],
# Processes
[{'name': 'p0', 'agent': f, 'inputs':['x'], 'outputs': ['y']},
{'name': 'p1', 'agent': g, 'inputs': ['y'], 'sources':['x']}]]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any).
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread (if any) runs in.
# thread_0 runs in the process called 'p1'
procs['p1'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
# -
# ## Example of Three Processes in a Row
# 
# This example is the same as the previous one except that it has a third process attached to process p2. The source thread <i>h</i> feeds stream <i>x</i> which is the input to process p0. The output of p0 is stream <i>y</i> which is the input to process p1. The output of p1 is stream <i>z</i> which is the input to process p2.
# <br>
# ### Streams
# [('x', 'i'), ('y', 'i'), ('z', 'i')]
# This specifies that this system has three streams called 'x', 'y' and 'z' which contain ints.
# ### Sources
# <b>Source Function</b> <i>h</i>
# <br>
# This function runs in its own thread. The function puts [0, 1, 2] into the stream called <i>x</i>, then sleeps, and then puts [3, 4, 5] into <i>x</i>. The function then calls <i>finished_source</i> to indicate that it has finished executing and so no further values will be appended to <i>x</i>.
# <br>
# <br>
# This function executes in a thread that runs in process <i>p0</i> because <i>h</i> appears in the specification for <i>p0</i>:
# <br>
# {'name': 'p0', 'agent': f, 'inputs':['x'], 'outputs': ['y'], 'sources': ['x'], <b>'source_functions':[h]</b>}
# <br>
# <b>Stream Sources</b> Stream <i>x</i> is a source in process <i>p0</i> because it appears in the specification of process <i>p0</i>.
# ### Process Structure
# <ol>
# <li>Source function <i>h</i> feeds stream <i>x</i> which is an input of process <i>p0</i>. </li>
# <li> Output stream <i>y</i> of process <i>p0</i> is an input stream of process <i>p1</i>.</li>
# <li> Output stream <i>z</i> of process <i>p1</i> is an input stream of process <i>p2</i>.</li>
# <li> Process <i>p2</i> has no output streams. </li>
# </ol>
#
# ### Process Functions
# Each process function has parameters <i>in_streams</i>, <i>out_streams</i> and possibly additional keyword or positional arguments. The process functions associated with processes <i>p0</i>, <i>p1</i>, and <i>p2</i> are <i>f</i>, <i>g</i> and <i>r</i>, respectively. The process function for a process is in the processes part of <i>multicore_specification</i>.
# <br>
# <ol>
# <li> The source extends stream <i>x</i> with [0, 1, 2, 3, 4, 5] and then calls <i>finished_source</i>. Thus <b>x[n] = n </b> for n less than 6. </li>
# <li> Process function <i>f</i> of <i>p0</i> adds 100 to its <i>in_streams[0]</i> which is stream <i>x</i> and puts the result in its <i>out_streams[0]</i> which is stream <i>y</i>. Thus <b>y[n] = x[n]+100 = n + 100 for </b> </li>.
# <li> Process function <i>g</i> of <i>p1</i> multiplies 2 to its <i>in_streams[0]</i> which is stream <i>y</i> and puts the result in its <i>out_streams[0]</i> which is stream <i>z</i>. Thus <b>z[n] = 2*y[n] = 2n + 200 for </b> </li>.
# <li> Process function <i>r</i> of <i>p2</i> creates a stream <i>s</i> and multiplies 3 to its <i>in_streams[0]</i> which is stream <i>z</i> and and puts the result stream <i>s</i>. This function also prints stream <i>s</i>. Thus it prints <b>3*z[n] = 6n + 600 for </b> </li>.
# </ol>
# +
# Step 1: multicore_specification of streams and processes.
multicore_specification = [
# Streams
[('x', 'i'), ('y', 'i'), ('z', 'i')],
# Processes
[{'name': 'p0', 'agent': f, 'inputs':['x'], 'outputs': ['y']},
{'name': 'p1', 'agent': h, 'inputs': ['y'], 'outputs': ['z'], 'sources': ['x']},
{'name': 'p2', 'agent': r, 'inputs': ['z']}]
]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any)
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread runs in.
# thread_0 runs in the process called 'p1'
procs['p1'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
# -
# ## Example of Multicore with NumPy Arrays
# This example illustrates the use of <b>StreamArray</b> which is a stream treated as a NumPy array with an arbitrarily large number of rows. Using <i>StreamArray</i> can be more efficient than using <i>Stream</i> for large computations.
# <br>
# <br>
# These examples are simple and small; however, in most applications each process function would convert an input stream to a <i>StreamArray</i> and carry out a lot computation as arrays before sending the results as output streams.
# <br>
# <br>
# The streams, sources, and process structure are similar to the previous two examples. The process functions differ in that the functions in this example use <i>StreamArray</i> whereas the earlier examples used <i>Stream</i>.
# <br>
# <br>
# You convert a Stream of numbers to a StreamArray of ints, floats, or doubles by calling the functions <b>dtype_int</b>, <b>dtype_float</b>, and <b>dtype_double</b> respectively.
# <br>
# <br>
# In this example, the agent functions <i>f</i> and <i>g</i> operate on StreamArrays of floats though the source function <i>h</i> generates a stream of int.
# 
# +
import numpy as np
from IoTPy.helper_functions.type import dtype_float
def test_multicore_with_arrays():
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# f_numpy is the agent function for processor called 'p0'.
def f_numpy(in_streams, out_streams):
map_window(
np.mean, dtype_float(in_streams[0]), out_streams[0],
window_size=2, step_size=2)
# g_numpy is the agent function for processor called 'p1'.
def g_numpy(in_streams, out_streams):
t = StreamArray('t')
map_window(max, dtype_float(in_streams[0]), t,
window_size=2, step_size=2)
print_stream(t)
# Step 0.1: Define source thread targets (if any).
def thread_target_numpy(procs):
for i in range(3):
extend_stream(procs, data=list(range(i*10, (i+1)*10)),
stream_name='x')
# Sleep to simulate an external data source.
time.sleep(0.001)
# Terminate stream because this stream will not be extended.
terminate_stream(procs, stream_name='x')
# Step 1: multicore_specification of streams and processes.
# Specify Streams: list of pairs (stream_name, stream_type).
# Specify Processes: name, agent function,
# lists of inputs and outputs, additional arguments.
multicore_specification = [
# Streams
[('x', 'i'), ('y', 'f')],
# Processes
[{'name': 'p0', 'agent': f_numpy, 'inputs':['x'],
'outputs': ['y'], 'sources': ['x']},
{'name': 'p1', 'agent': g_numpy, 'inputs': ['y']}]
]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any)
thread_0 = threading.Thread(target=thread_target_numpy, args=(procs,))
# Step 4: Specify which process each thread runs in.
# thread_0 runs in the process called 'p1'
procs['p1'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
test_multicore_with_arrays()
# -
# ## Example of Merging Streams from Multiple Processes
# This example shows a slightly more complex process structure. The example has four processes
# called <i>coordinator</i>, <i>sine</i>, <i>cosine</i>, and <i>tangent</i>. The <i>coordinator</i> generates a sequence of values that are sent to other processes which compute sines, cosines and tangents of these values and send the results back to the <i>coordinator</i>. The <i>coordinator</i> then computes the square of the error --- the difference between tangent and sine/cosine.
# <br>
# <br>
# This example gives names to agents. This is helpful in debugging because the error statements identify the agent that caused the error. We haven't given names to agents in some examples for brevity.
# 
#
# ### Process Structure
# <ol>
# <li> A source function <i>h</i> extends stream <i>x</i> with a sequence of 10 values between 0.0 and pi. This source function executes in a thread in the process called <i>coordinator</i>. Stream <i>x</i> is an input for all processes.
# </li>
# <li> Agents <i>sine</i>, <i>cosine</i>, and <i>tangent</i> read stream <i>x</i> and output streams <i>sines</i>, <i>cosines</i>, and <i>tangents</i> respectively. These streams are inputs to process <i>coordinate</i>.
# </li>
# <ol>
# +
from IoTPy.agent_types.merge import zip_map
def example_merging_streams_from_multiple_processes():
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# sine is the agent function for the process called 'sine'.
def sine(in_streams, out_streams):
map_element(np.sin, dtype_float(in_streams[0]), out_streams[0],
name='sine')
# cosine is the agent function for the process called 'cosine'.
def cosine(in_streams, out_streams):
map_element(np.cos, dtype_float(in_streams[0]), out_streams[0],
name='cosine')
# tangent is the agent function for the process called 'tangent'.
def tangent(in_streams, out_streams):
map_element(np.tan, dtype_float(in_streams[0]), out_streams[0],
name='tangent')
# coordinate is the agent function for the process called 'coordinate'.
def coordinate(in_streams, out_streams):
x, sines, cosines, tangents = in_streams
def f(lst): return lst[0]/lst[1]
def g(lst):
error_squared= (lst[0] - lst[1])**2
return error_squared
ratios = Stream('ratios')
errors = Stream('errors')
zip_map(f, [sines, cosines], ratios, name='sine / cosine')
zip_map(g, [ratios, tangents], errors, name='compute error')
print_stream(errors, 'error')
# # Step 0.1: Define source thread target (if any).
def source_thread_target(procs):
extend_stream(procs, data = np.linspace(0.0, np.pi, 10), stream_name='x')
terminate_stream(procs, stream_name='x')
# Step 1: multicore_specification of streams and processes.
# Specify Streams: list of pairs (stream_name, stream_type).
# Specify Processes: name, agent function,
# lists of inputs and outputs and sources, additional arguments.
multicore_specification = [
# Streams
[('x', 'f'), ('sines', 'f'), ('cosines', 'f'), ('tangents', 'f')],
# Processes
[{'name': 'sine', 'agent': sine, 'inputs':['x'], 'outputs': ['sines']},
{'name': 'cosine', 'agent': cosine, 'inputs':['x'], 'outputs': ['cosines']},
{'name': 'tanget', 'agent': tangent, 'inputs':['x'], 'outputs': ['tangents']},
{'name': 'coordinator', 'agent': coordinate, 'inputs':['x', 'sines', 'cosines', 'tangents'],
'sources': ['x']}]
]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any)
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread runs in.
# thread_0 runs in the process called 'coordinator'
procs['coordinator'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
example_merging_streams_from_multiple_processes()
# -
# ## Passing Data to and from Multiprocessing Blocks
# Non-IoTPy processes or threads can interact concurrently with IoTPy by extending input streams, getting data from queues fed by output streams, and by putting data into, and getting data from, multiprocessing blocks.
# 
# This example illustrates how to pass data to a multiprocessing block and get data from the block. This example is the same as the previous one except that the variables <b>total</b> and <b>num</b> are passed to the multiprocessing block which returns updated values of these variables.
# <br>
# <br>
# total = multiprocessing.Value('f')
# <br>
# num = multiprocessing.Value('i')
# <br>
# <br>
# creates <i>total</i> a wrapper for a float, and <i>num</i> a wrapper for int.
# <br>
# <br>
# These variables can be passed to any collection of processes. In this example they are passed only to the process <i>coordinator</i>.
# These variables are assigned initial values from a computation that is not shown here. The multiprocessing block shown updates these values. For example, the value of <i>num</i> is 25 before the block is executed and 45 after it terminates.
#
# ### Passing variables as keyword or positional arguments
# In this example, variables are passed to the process <i>coordinator</i> as keyword arguments.
# The keyword arguments are specified as a dict with the name of an argument (e.g. 'total') and its initial value (<i>total</i>).
# <br>
# <br>
# {'name': 'coordinator', 'agent': coordinate, 'inputs':['x', 'sines', 'cosines', 'tangents'],
# <br>
# 'sources': ['x'], 'source_functions':[sequence],
# <br>
# <b>'keyword_args'</b> : {'total' : total, 'num' : num},}
# +
import multiprocessing
def example_passing_data_to_multicore():
total = multiprocessing.Value('f')
num = multiprocessing.Value('i')
# Values computed from an earlier computation which is not shown.
# total and num are passed to the multiprocessing block.
total.value = 4.0e-13
num.value = 25
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# sine is the agent function for the process called 'sine'.
def sine(in_streams, out_streams):
map_element(np.sin, dtype_float(in_streams[0]),
out_streams[0], name='sine')
# cosine is the agent function for the process called 'cosine'.
def cosine(in_streams, out_streams):
map_element(np.cos, dtype_float(in_streams[0]), out_streams[0], name='cosine')
# tangent is the agent function for the process called 'tangent'.
def tangent(in_streams, out_streams):
map_element(np.tan, dtype_float(in_streams[0]),
out_streams[0], name='tangent')
# coordinate is the agent function for the process called 'coordinate'.
def coordinate(in_streams, out_streams, total, num):
x, sines, cosines, tangents = in_streams
def f(lst): return lst[0]/lst[1]
def g(lst):
error_squared= (lst[0] - lst[1])**2
return error_squared
ratios = Stream('ratios')
errors = Stream('errors')
zip_map(f, [sines, cosines], ratios, name='sine / cosine')
zip_map(g, [ratios, tangents], errors, name='compute error')
print_stream(errors, 'error')
# Step 0.1: Define source thread target (if any).
def source_thread_target(procs):
extend_stream(procs, data=np.linspace(0.0, np.pi, 10), stream_name='x')
terminate_stream(procs, stream_name='x')
# Step 1: multicore_specification of streams and processes.
# Specify Streams: list of pairs (stream_name, stream_type).
# Specify Processes: name, agent function,
# lists of inputs and outputs and sources, additional arguments.
multicore_specification = [
# Streams
[('x', 'f'), ('sines', 'f'), ('cosines', 'f'), ('tangents', 'f')],
# Processes
[{'name': 'sine', 'agent': sine, 'inputs':['x'], 'outputs': ['sines']},
{'name': 'cosine', 'agent': cosine, 'inputs':['x'], 'outputs': ['cosines']},
{'name': 'tanget', 'agent': tangent, 'inputs':['x'], 'outputs': ['tangents']},
{'name': 'coordinator', 'agent': coordinate, 'inputs':['x', 'sines', 'cosines', 'tangents'],
'sources': ['x'], 'keyword_args' : {'total' : total, 'num' : num}}]
]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any)
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread runs in.
# thread_0 runs in the process called 'coordinator'
procs['coordinator'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
example_passing_data_to_multicore()
# -
# ## Actuators
# A multiprocessing block may need to interact asynchronously with some external device. To do so, the block puts data into a queue and uses threads responsible for interfacing between the queue and the device. This simple example illustrates the simplest actuator: a printer. Indeed printing can be done synchronously by the multiprocessing block. Printing doesn't need a queue to interface between it and the block. We use the printer in this example to illustrate the idea.
# <br>
# <br>
# Function <i>g</i> of process <i>p1</i> has an agent called 'copy_stream_s_to_queue_q' which copies stream <i>s</i> to queue <i>q</i>. A thread, <b>my_thread</b> in <i>p1</i> prints values from the queue; this thread represents the thread that interfaces with an external actuator device. This thread is in addition to any source threads that may exist.
# <br>
# <br>
# Queue <i>q</i> is specified as an <b>output queue</b>. An output queue gets a special message <b>'_finished'</b> when the multiprocess block terminates.
# <br>
# <br>
# Threads (apart from source threads) and output queues are specified in <i>multicore_specifications</i>. See
# <br>
# <br>
# {'name': 'p1', 'agent': g, 'inputs': ['y'],
# <br>
# 'args': [q], <b>'output_queues'</b>: [q], <b>'threads'</b>: [my_thread]}
# <br>
# <br>
# The thread, <i>my_thread</i>, terminates when it receives a '_finished' message. We want this thread to terminate so that process <i>p1</i> terminates, and then the entire multiprocessing block can terminate as well.
# 
# +
import threading
from IoTPy.agent_types.sink import stream_to_queue
def example_output_thread_with_queue():
q = multiprocessing.Queue()
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# g is the agent function for the process called 'p1'.
def g(in_streams, out_streams, q):
s = Stream('s')
map_element(lambda v: v*2, in_streams[0], s)
stream_to_queue(s, q, name='copy_stream_s_to_queue_q')
# Step 0.1: Define source thread target (if any).
def source_thread_target(procs):
for i in range(3):
extend_stream(procs, data=list(range(i*2, (i+1)*2)),
stream_name='x')
time.sleep(0.001)
terminate_stream(procs, stream_name='x')
# Define the actuator thread target. This thread target is
# used to create a thread (output) which is run in the process
# called 'p0'.
def get_data_from_output_queue(q):
while True:
v = q.get()
if v == '_finished': break
else: print ('q.get() = ', v)
multicore_specification = [
# Streams
[('x', 'i'), ('y', 'i')],
# Processes
[{'name': 'p0', 'agent': f, 'inputs':['x'], 'outputs': ['y'], 'sources': ['x']},
{'name': 'p1', 'agent': g, 'inputs': ['y'],
'args': [q], 'output_queues': [q]}]
]
processes, procs = get_processes_and_procs(multicore_specification)
source_thread = threading.Thread(target=source_thread_target, args=(procs,))
output_thread = threading.Thread(target=get_data_from_output_queue, args=(q,))
procs['p0'].threads = [source_thread, output_thread]
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
example_output_thread_with_queue()
# -
# ## Example of Process Structure with Feedback
# The example shows a process structure with feedback. This example creates an echo from a spoken sound. (You can write more efficient and succinct code to compute echoes. The code in this example is here merely because it illustrates a concept.)
# 
# <br>
# ### Streams
# <ol>
# <li><b>sound_made</b>: This is the sound made by a speaker in a large spherical space.</li>
# <li><b>attenuated</b>: This is the sound made multiplied by an attenuation factor.</li>
# <li><b>echo</b>: This is the echo of the sound made heard at the center of the room. The echo is a delay followed by an attenuation of the sound heard. </li>
# <li><b>sound_heard</b>: This is the sound that is heard by the speaker. The heard sound is the sound made by the speaker plus the echo.</li>
# </ol>
# The equations that define the streams are:
# <ol>
# <li>
# <b>attentuated[n] = sound_heard[n]*attenuation</b>
# </li>
# <li>
# <b>echo[n] = attentuated[n-delay]</b> for n > delay.
# </li>
# <li>
# <b>sound_heard[n] = sound_made[n] + echo[n]</b> for n > delay.
# </li>
# </ol>
#
# ### Process Structure
# Process <i>p0</i> has a source which feeds one of its input streams <i>sound_made</i> with a stream of measurements obtained from a microphone. In this example, the stream is generated with numbers so that we can see how streams are processed.
# <br>
# <br>
# Process <i>p1</i> contains a single input stream which is the sound heard and a single output stream which is an attenuation of the sound heard.
#
# ### Process Functions
# The function <i>f</i> of <i>p0</i> computes <i>echo</i> from <i>sound_made</i>. The first 4 , i.e., <b>delay</b>, units of the echo are empty (i.e. 0).
# <br>
# <b>map_element(lambda v: v, attenuated, echo)</b>
# <br>
# copies the attenuated stream to the echo stream; however, since the first 4 (i.e. delay) values of the echo stream are 0, the echo stream will consist of 4 zeroes followed by the attenuated stream.
# <br>
# <i>out_streams[0]</i> of process <i>p0</i> is <i>sound_heard</i>. Function <i>f</i> makes <i>sound_heard</i> the sum of the echo and the sound made.
# <br>
# The function <i>g</i> of process <i>p1</i> <i>p0</i> puts elements of its input stream (i.e. <i>sound_heard</i> on queue <i>q</i> and returns the elements multiplied by <i>attenuation</i>.
# +
from IoTPy.agent_types.basics import *
def example_echo_two_cores():
# This is the delay from when the made sound hits a
# reflecting surface.
delay = 6
# This is the attenuation of the reflected wave.
attenuation = 0.5
# The results are put in this queue. A thread reads this
# queue and feeds a speaker or headphone.
q = multiprocessing.Queue()
# ----------------------------------------------
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# Agent function for process named 'p0'
# echo is a delay of zeroes followed by attenuated heard sound.
# out_streams[0], which is the same as sound_heard, is
# echo + sound_made
def f_echo(in_streams, out_streams, delay):
sound_made, attenuated = in_streams
echo = StreamArray('echo', dtype='float')
echo.extend(np.zeros(delay, dtype='float'))
map_element(lambda v: v, attenuated, echo)
# The zip_map output is the sound heard which is
# the sound heard plus the echo.
zip_map(sum, [sound_made, echo], out_streams[0])
# Agent function for process named 'p1'
# This process puts the sound heard into the output queue
# and returns an attenuated version of the sound_heard as
# its output stream.
def g_echo(in_streams, out_streams, attenuation, q):
def gg(v):
# v is the sound heard
q.put(v)
# v*attenuation is the echo
print ('in g_echo; v is ', v)
return v*attenuation
map_element(gg, in_streams[0], out_streams[0])
def source_thread_target(procs):
data=list(range(10))
extend_stream(procs, data=np.array(np.arange(10.0)), stream_name='sound_made')
time.sleep(0.0001)
extend_stream(procs, data=np.array([0.0]*10), stream_name='sound_made')
terminate_stream(procs, stream_name='sound_made')
# Thread that gets data from the output queue
# This thread is included in 'threads' in the specification.
# Thread target
def get_data_from_output_queue(q):
finished_getting_output = False
while not finished_getting_output:
v = q.get()
if v == '_finished': break
print ('heard sound = spoken + echo: ', v)
multicore_specification = [
# Streams
[('sound_made', 'f'), ('attenuated', 'f'), ('sound_heard', 'f')],
# Processes
[{'name': 'p0', 'agent': f_echo, 'inputs': ['sound_made', 'attenuated'],
'outputs': ['sound_heard'], 'keyword_args' : {'delay' : delay}, 'sources': ['sound_made']},
{'name': 'p1', 'agent': g_echo, 'inputs': ['sound_heard'], 'outputs': ['attenuated'],
'args': [attenuation, q], 'output_queues': [q] } ]]
processes, procs = get_processes_and_procs(multicore_specification)
source_thread = threading.Thread(target=source_thread_target, args=(procs,))
output_thread = threading.Thread(target=get_data_from_output_queue, args=(q,))
procs['p0'].threads = [source_thread, output_thread]
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
example_echo_two_cores()
# -
# ## Example source and actuator thread with single process
# This example is the same as the previous one except that the computation is carried out in a single process rather in two processes. The example illustrates an actuator thread and a source thread in the same process.
# 
# +
def example_echo_single_core():
# This is the delay from when the made sound hits a
# reflecting surface.
delay = 4
# This is the attenuation of the reflected wave.
attenuation = 0.5
# The results are put in this queue. A thread reads this
# queue and feeds a speaker or headphone.
q = multiprocessing.Queue()
# ----------------------------------------------
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agent functions.
# Agent function for process named 'p0'
# echo is a delay of zeroes followed by attenuated heard sound.
# out_streams[0], which is the same as sound_heard is
# echo + sound_made
def f_echo(in_streams, out_streams, delay, attenuation, q):
echo = StreamArray('echo', dtype='float')
echo.extend(np.zeros(delay, dtype='float'))
#echo = Stream('echo', initial_value=[0]*delay)
#Note: sound_made = in_streams[0]
sound_heard = in_streams[0] + echo
map_element(lambda v: v*attenuation, sound_heard, echo)
stream_to_queue(sound_heard, q)
def source_thread_target(procs):
extend_stream(procs, data=list(range(10)), stream_name='sound_made')
time.sleep(0.0001)
extend_stream(procs=procs, data=[0]*10, stream_name='sound_made')
terminate_stream(procs, stream_name='sound_made')
# Thread that gets data from the output queue
# This thread is included in 'threads' in the specification.
# Thread target
def get_data_from_output_queue(q):
finished_getting_output = False
while not finished_getting_output:
v = q.get()
if v == '_finished': break
print ('heard sound = spoken + echo: ', v)
multicore_specification = [
# Streams
[('sound_made', 'f')],
# Processes
[{'name': 'p0', 'agent': f_echo, 'inputs': ['sound_made'],
'args' : [delay, attenuation, q], 'sources': ['sound_made'],'output_queues': [q]}]]
processes, procs = get_processes_and_procs(multicore_specification)
source_thread = threading.Thread(target=source_thread_target, args=(procs,))
output_thread = threading.Thread(target=get_data_from_output_queue, args=(q,))
procs['p0'].threads = [source_thread, output_thread]
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
example_echo_single_core()
# -
# ## Example of a grid computation
# Grid computations are used in science, for example in computing the temperature of a metal plate. The grid is partitioned into regions with a process assigned to simulate each region. On the n-th step, each process reads the values of relevant parts of the grid and updates its own value.
# <br>
# <br>
# This example uses two copies of the grid; the two copies are <b>even</b> and <b>odd</b>.
# <ol>
# <li>On <b>even</b> steps (i.e., steps 0, 2, 4,..) the <i>j</i>-th
# proces <b>reads</b> the <i>even</i> grid and <b>writes</b> the
# <i>j</i>-th element of the <i>odd</i> grid. </li>
# <li>On <b>odd</b> steps, the <i>j</i>-th proces <b>reads</b> the
# <i>odd</i> grid and <b>writes</b> the <i>j</i>-th element of the
# <i>even</i> grid. </li>
# </ol>
# So, each portion of the grid is modified by only one process. And no process reads a value while it is modified.
#
# ### The example problem
# A linear metal bar of length <i>N</i> is partitioned into a grid of <i>N</i> continuous regions. Grid 0 is kept at a constant temperature of 0 degrees while grid <i>N-1</i> is kept at a constant temperature of <i>N-1</i> degrees. Initially, the temperature at intermediate grid points is arbitrary; in the code below, the temperature at grid point <i>i</i> exceeds <i>i</i> by <i>DELTA</i>.
# <br>
# <br>
# Let <b>TEMP[i][k]</b> be the temperature of the <i>i</i>-th region on step <i>k</i>. Then, for all <i>k</i>:
# <ol>
# <li>TEMP[0][k] = 0 </li>
# <li>TEMP[N-1][k] = N-1 </li>
# <li>TEMP[i][k] = (TEMP[i-1][k] + TEMP[i][k] + TEMP[i+1][k])/3 i in [1, ..,N-2] </li>
# </ol>
#
#
# ### Processes
# The computation uses <i>N-2</i> processes. The <i>i</i>-th process is called 'grid_i' and is responsible for simulating the <i>i</i>-th region.
# <br>
# Each process takes the <i>k + 1</i>-th step after it has finished the <i>k</i>-th step and it has determined that its neighbors have also finished the <i>k</i>-th step.
# <br>
#
#
# ### Streams
# The system has one stream, <b>s_i</b> for the <i>i</i>-th process. This stream contains the elements [0, 1, .. , k] after the <i>i</i>-th process has completed <i>k</i>-th steps.
# <br>
# Process <i>grid_i</i> outputs stream <i>s_i</i> and inputs streams from its neighboring processes which are <i>grid_(i-1)</i> if <i>i</i> exceeds 1 and <i>grid_(i+1)</i> if <i>i</i> is less than <i>N-1</i>.
#
# ### Process Structure
# The process structure is linear with each process getting input streams from each of its neighbors and sending its output stream to all its neighbors.
# 
#
# ### Process Function
# The process begins by sending 0 on its output stream to indicate that it has finished its 0-th step.
# <br>
# <br>
# The <i>k</i>-th value of <i>in_streams[j]</i> is <i>k</i> when the <i>j</i>-th neighboring process has completed its <i>k</i>-th step.
# <br>
# <br>
# <b>synch_stream</b> is an internal stream of the process. The <i>k</i>-th element of this stream is <i>k</i> after all neighboring processes have completed their <i>k</i>-th step.
# <br>
# <br>
# The zip_map function <i>r</i> operates on a list with one element from each neighbor. All the elements of the list will be <i>k</i> on the <i>k</i>-th step. The zip_map function returns <i>k</i> which is any element of the list. In this example it returns the 0-th element.
# <br>
# <br>
# Thus the zip_map function acts as a synchronizer. It waits until all neighbors have completed the <i>k</i>-step and then it outputs <i>k</i>.
# <br>
# <br>
# Function <i>g</i> is called for the <i>k</i>-th time when this process and all its neighbors have completed <i>k - 1</i> steps. Function <i>g</i> does the grid computation. Function <i>r</i> and the zip_map agent are used merely for synchronizing.
# ### run()
# Function <i>f</i> calls <b>run</b> after it has declared all its agents. Without calling run() the function will take no action.
# <br>
# <br>
# Note that when using external source threads, you should not call <i>run</i> because the source threads are responsible for starting and stopping the main computational thread. This example has no source threads so you must call <i>run</i> to start the system.
# +
from IoTPy.core.stream import _no_value
def test_grid():
# N is the size of the grid
N = 5
# M is the number of steps of execution.
M = 5
# DELTA is the deviation from the final solution.
DELTA = 0.01
# even, odd are the grids that will be returned
# by this computation
even = multiprocessing.Array('f', N)
odd = multiprocessing.Array('f', N)
# Set up initial values of the grid.
for i in range(1, N-1):
even[i] = i + DELTA
even[N-1] = N-1
odd[N-1] = N-1
def f(in_streams, out_streams, index, even, odd):
def g(v):
if (0 < index) and (index < N-1):
if v%2 == 0:
odd[index] = (even[index-1] + even[index] + even[index+1])/3.0
else:
even[index] = (odd[index-1] + odd[index] + odd[index+1])/3.0
return v+1
def r(lst, state):
if state < M:
return lst[0], state+1
else:
return _no_value, state
for out_stream in out_streams: out_stream.extend([0])
synch_stream = Stream('synch_stream')
zip_map(r, in_streams, synch_stream, state=0, name='zip_map_'+str(index))
map_element(g, synch_stream, out_streams[0], name='grid'+str(index))
run()
multicore_specification = [
# Streams
[('s_'+str(index), 'i') for index in range(1, N-1)],
# Processes
[{'name': 'grid_'+str(index), 'agent': f,
'inputs':['s_'+str(index+1), 's_'+str(index-1)],
'outputs':['s_'+str(index)],
'args': [index, even, odd]} for index in range(2, N-2)] + \
[{'name': 'grid_'+str(1), 'agent': f,
'inputs':['s_'+str(2)], 'outputs':['s_'+str(1)],
'args': [1, even, odd]}] + \
[{'name': 'grid_'+str(N-2), 'agent': f,
'inputs':['s_'+str(N-3)], 'outputs':['s_'+str(N-2)],
'args': [N-2, even, odd]}]
]
# Execute processes (after including your own non IoTPy processes)
processes = get_processes(multicore_specification)
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
print ('Grid after ', M, ' steps is: ')
if M%2 == 0:
print (even[:])
else:
print (odd[:])
test_grid()
# -
# # Counting Elements in Streams
# The next example uses algorithms to count elements in streams (see IoTPy/examples/Counting). This example uses a Bloom Filter and a count-min-sketch algorithm. The data stream consists of pairs where a pair is either ('add', object) or ('check', object). The pair ('add', z) in the input stream states that object z was added to the data stream. The pair ('check', z) in the input stream is a command to check whether z was added earlier.
# <br>
# 
from examples.Counting.bloom_filter import bloom_filter_stream
from examples.Counting.bloom_filter import BloomFilter
from examples.Counting.count_min_sketch import count_min_sketch_stream
from examples.Counting.count_min_sketch import CountMinSketch
from IoTPy.agent_types.merge import merge_asynch
def test_multiprocessing_counting_algorithms():
# ----------------------------------------------
# Step 0: Define agent functions, source threads
# and actuator threads (if any).
# Step 0.0: Define agents
def bloom_filter_agent(in_streams, out_streams):
bloom_filter = BloomFilter(
est_elements=1000, false_positive_rate=0.05)
bloom_filter_stream(in_streams[0], out_streams[0],
bloom_filter=bloom_filter)
def count_min_sketch_agent(in_streams, out_streams):
count_min_sketch = CountMinSketch(width=1000, depth=20)
count_min_sketch_stream(in_streams[0], out_streams[0],
count_min_sketch=count_min_sketch)
def merge_agent(in_streams, out_streams):
s = Stream('print stream')
def g(pair):
index, value = pair
if index == 0:
print ('bloom_filter. value: ', value)
else:
print ('count_min_sketch. value: ', value)
merge_asynch(g, in_streams, s)
# Step 0.1: Define source thread target (if any).
def source_thread_target(procs):
data=[('add', 'a'), ('add', 'b'), ('add', 'a'),
('check', 'c'), ('add', 'd'), ('check','a')]
extend_stream(procs, data, stream_name='data')
time.sleep(0.001)
data=[('add', 'c'), ('check', 'b'), ('check', 'a'),
('check', 'c'), ('check', 'e'), ('add', 'a')]
extend_stream(procs, data, stream_name='data')
terminate_stream(procs, stream_name='data')
# Step 1: multicore_specification of streams and processes.
# Specify Streams: list of pairs (stream_name, stream_type).
# Specify Processes: name, agent function,
# lists of inputs and outputs and sources, additional arguments.
multicore_specification = [
# Streams
[('data', 'x'), ('bloom_results', 'x'),
('count_min_sketch_results', 'x')],
# Processes
[{'name': 'bloom_filter_process', 'agent': bloom_filter_agent,
'inputs':['data'], 'outputs': ['bloom_results'],
'sources': ['data']},
{'name': 'count_min_sketch_process', 'agent': count_min_sketch_agent,
'inputs':['data'], 'outputs': ['count_min_sketch_results']},
{'name': 'merge_process', 'agent': merge_agent,
'inputs': ['bloom_results', 'count_min_sketch_results']}
]]
# Step 2: Create processes.
processes, procs = get_processes_and_procs(multicore_specification)
# Step 3: Create threads (if any)
thread_0 = threading.Thread(target=source_thread_target, args=(procs,))
# Step 4: Specify which process each thread runs in.
# thread_0 runs in the process called 'coordinator'
procs['bloom_filter_process'].threads = [thread_0]
# Step 5: Start, join and terminate processes.
for process in processes: process.start()
for process in processes: process.join()
for process in processes: process.terminate()
test_multiprocessing_counting_algorithms()
| examples/ExamplesOfMulticore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Check this tensorflow zoo model for checkpoint you wanted, https://github.com/tensorflow/models/tree/master/research/slim
#
# I chose [Inception-Resnet-V2](http://arxiv.org/abs/1602.07261), highest accuracy.
# !wget http://download.tensorflow.org/models/inception_resnet_v2_2016_08_30.tar.gz
# !tar zxf inception_resnet_v2_2016_08_30.tar.gz
| grab-aiforsea/computer-vision/download-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden" slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt
import scipy.stats as ss
import pandas as pd
import numpy as np
from auxiliary import plot_individual_specific_effects
from auxiliary import get_lalonde_data
# -
# # Potential outcome model
# ### Introduction
# + [markdown] slideshow={"slide_type": "slide"}
# Given what we know from the introduction about the potential outcome model, we will initially prepare the Lalonde Dataset to fit the framework and use it as a running example going forward.
#
# What are this example's ...
#
# * potential outcomes
# * counterfactual state
# * treatment
#
# + slideshow={"slide_type": "slide"}
df = get_lalonde_data()
df.head()
# -
# We are dealing with a binary treatment here: $D = 1$ if the individual did participate in the training program and $D = 0$ if it did not. However, in practice assigning **treatment** is never that easy. We lump a lot of heterogeneity together (e.g. different sites, content of curriculum) that might affect the success of program participation. Maybe we should stratify the analysis by site?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Individual-specific effect of treatment
# + [markdown] slideshow={"slide_type": "slide"}
# It would be great if we could get our hands on the individual-specific effect of treatment.
#
# \begin{align*}
# \delta_i = y_i^1 - y_i^0
# \end{align*}
#
# * Why do individuals have potentially different effects of treatment?
# + slideshow={"slide_type": "slide"}
fig, ax = plt.subplots()
x = np.linspace(-5, 5, 5000)
pdf = ss.norm.pdf(x, 0, 1)
ax.plot(x, pdf)
ax.set_xlabel(r"$\delta = Y^1 - Y^0$")
ax.set_ylabel("Density")
ax.set_xticklabels(["", "", "", 0.5, "", "", ""])
ax.set_xlim([-3, 3])
ax.set_ylim([0, 0.5])
# + [markdown] slideshow={"slide_type": "slide"}
# There might be considerable heterogeneity in the benefits of treatment among the population. And summarizing the distribution of benefits with a single number, for example $E[\delta]$, might result in a loss of information.
#
# **Examples**
#
# * medical treatment
# * ..
# + [markdown] slideshow={"slide_type": "slide"}
# Give our definitions of $(Y^1, Y^0, D)$ and their individual realizations $(y^1_i, y^0_i, d_i)$ we can now define the observed outcome $Y$ in terms of them.
#
# \begin{align*}
# Y = \begin{cases}
# Y^1 & \quad\text{if}\quad D = 1 \\
# Y^0 & \quad\text{if}\quad D = 0
# \end{cases}
# \end{align*}
#
# or more compactly in switching-regime notation
#
# \begin{align*}
# Y = D Y^1 + (1 - D) Y^0.
# \end{align*}
# + [markdown] slideshow={"slide_type": "slide"}
# This leads Holland (1986) to describe the fundamental problem of causal inference:
#
# <img src="material/fig-fundamental-problem.png" width="500">
#
# $\rightarrow$ as only the diagonal of the table is observable we cannot simply compute $\delta_i$ by taking the difference in potential outcomes $(y^1_i, y^0_i)$.
# + slideshow={"slide_type": "slide"}
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Population-level parameters
# + [markdown] slideshow={"slide_type": "slide"}
# It looks like we need to give up any hope of obtaining the individual-specific effect of treatment. But what can we still hope for?
#
# $\rightarrow$ population-level parameters
#
# * What are common examples?
#
# * What are the policy questions they address?
#
# * What is their relationship to each other?
# + [markdown] slideshow={"slide_type": "slide"}
# \begin{align*}
# \begin{array}{lll}
# E[Y^1 - Y^0] & \qquad ATE & \qquad\text{average effect of treatment} \\
# E[Y^1 - Y^0 \mid D = 1] & \qquad ATT & \qquad\text{average effect on treated} \\
# E[Y^1 - Y^0 \mid D = 0] & \qquad ATC & \qquad\text{average effect on control}
# \end{array}
# \end{align*}
# + slideshow={"slide_type": "slide"}
plot_individual_specific_effects(with_parameters=[0, 0.7, -0.5])
# + slideshow={"slide_type": "slide"}
plot_individual_specific_effects(with_parameters=[0, -0.7, 0.5])
# + slideshow={"slide_type": "slide"}
plot_individual_specific_effects(with_parameters=[0, 0, 0])
# + [markdown] slideshow={"slide_type": "slide"}
# ### Stable unit treatment value assumption
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true toc-nb-collapsed=true
# The potential outcome model gets its empirical tractability when combined with the **Stable Unit Treatment Value Assumption (SUTVA)** of which there exist many formulations. We will go with the one from Imbens and Rubin (2015):
#
# > The potential outcomes for any unit do not vary with the treatments assigned to other units, and, for each unit there are no different forms or versions of each treatment level, which lead to different potential outcomes.
# + [markdown] slideshow={"slide_type": "slide"}
# The table below shows all possible assignment patterns for a hypothetical treatment where the only constraint is that at least one individual remains in the treatment and control group. As we increase participation from one to two individuals, the potential outcome $Y_1$ declines.
#
# <img src="material/fig-stable-unit-treatment.png" width="500">
# -
# * When do we need to expect this is violated?
#
# * **influence patterns** that result from contact across individuals in social or physical space
# * **dilution / concentration patterns** that one can assume would result from changes in the prevalence of treatment
# + [markdown] slideshow={"slide_type": "slide"}
# ### Treatment assignment and observational studies
# + [markdown] slideshow={"slide_type": "slide"}
# * randomized experiment
#
# \begin{align*}
# (Y^0, Y^1) \perp \!\!\! \perp D
# \end{align*}
#
# * observational study
#
# > A *observational study* is an empirical investigation of treatments, policies, or exposures and the effects they cause, but it differs from an experiment in that the investigator cannot control the assignment of treatments to subjects. (Rosenbaum (2002))
# -
# ### Naive estimation of average causal effects
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# We will now first outline the problem with the naive estimation of average causal effects. Then we take a closer look at the different sources of biases involved and finally discuss the set of assumptions used to ***solve*** these issues.
#
# \begin{align*}
# \hat{\delta}_{NAIVE} \equiv E_N[y_i\mid d_i = 1] - E_N[y_i\mid d_i = 0]
# \end{align*}
#
# We can further decompose the average treatment effect by treatment status as the individual assignment is mutually exclusive.
#
#
# \begin{align*}
# E[Y^1 - Y^0] & = E[\delta] & = \{\pi E[Y^1\mid D = 1] + (1 - \pi)E[Y^1\mid D = 0]\} \\
# &&- \{\pi E[Y^0\mid D = 1] + (1 - \pi)E[Y^0\mid D = 0]\}
# \end{align*}
#
# The average treatment effect is a function of five unknowns. Which components can be easily computed from data?
#
# \begin{align*}
# E_N[y_i\mid d_i = 1] \xrightarrow{p} E[Y^1\mid D = 1] \neq E[Y^1]\\
# E_N[y_i\mid d_i = 0] \xrightarrow{p} E[Y^0\mid D = 0] \neq E[Y^0]
# \end{align*}
# -
# #### Biases
#
# \begin{align*}
# E[Y^1 \mid D = 1] - E[Y^0 \mid D = 0] & = E[\delta] + \underbrace{\{E[Y^0\mid D= 1] - E[Y^0\mid D= 0]\}}_{\text{Baseline bias}} \\
# & + (1 - \pi) \underbrace{\left\{E[\delta \mid D = 1] -E[\delta \mid D = 0]\right\}}_{\text{Differential treatment effect bias}}
# \end{align*}
#
# <img src="material/fig-bias-example.png" width="500">
#
# The additional information provided in the text states that $\pi = 0.3$ meaning that 30\% of the sample participate in the treatment.
#
# \begin{align*}
# ATT & = E[Y_1 - Y_0 \mid D = 1]= 10 - 6 = 4 \\
# ATC & = E[Y_1 - Y_0 \mid D = 0] = 8 - 5 = 3 \\
# \delta^{NAIVE} & = E[Y_1\mid D = 1] - E[Y_0\mid D = 0] = 10 - 5 = 5
# \end{align*}
#
# Now we are ready to calculate the average treatment effect:
#
# \begin{align*}
# ATE = E[Y_1 - Y_0] & = \pi\, E[Y_1 - Y_0 \mid D = 1] + (1 - \pi)\, E[Y_1 - Y_0 \mid D = 0] \\
# & = 0.3 \times 4 + 0.7 \times 3 = 3.3
# \end{align*}
#
# Next, we can determine the different components of the bias.
#
# \begin{align*}
# \Delta^{\text{base}} = E[Y^0\mid D= 1] - E[Y^0\mid D= 0] = 6 - 5 = 1 \\
# \Delta^{\text{diff}} = (1 - \pi)\left( E[\delta \mid D = 1] - E[\delta \mid D = 0]\right) = 0.7 \left( (10 - 6) - (8 - 5 )\right) = 0.7
# \end{align*}
#
# There are several different representation of the bias when using the naive estimator of mean difference in observed outcomes by treatment status as an estimate for the effect of treatment. We continue with the exposition in Frölich & Sperlich (2019) and Heckman, Urzua, & Vytlacil (2006).
#
# \begin{align*}
# E[Y\mid D = 1] - E[Y\mid D = 0] & = E[Y^1\mid D = 1] - E[Y^0\mid D = 0] \\
# & =E[Y^1\mid D = 1] - E[Y^0\mid D = 1] \\
# & + E[Y^0\mid D = 1] - E[Y^0\mid D = 0] \\
# & = \underbrace{E[Y^1 - Y^0\mid D = 1]}_{TT} + \underbrace{E[Y^0\mid D= 1]- E[Y^0 \mid D = 0]}_{\text{Selection bias}}
# \end{align*}
#
# Now we can simply add and subtract $E[Y_1 - Y_0]$ to get the more economic version.
#
# \begin{align*}
# E[Y\mid D = 1] - E[Y\mid D = 0] & = \underbrace{E[Y^1 - Y^0]}_{ATE} \\
# & + \underbrace{E[Y^1 - Y^0 \mid D = 1] - E[Y^1 - Y^0]}_{\text{Sorting on gains}} \\
# & + \underbrace{E[Y^0\mid D = 1] - E[Y^0 \mid D = 0]}_{\text{Sorting on levels}}
# \end{align*}
#
# Sorting on levels is simply a different phrase for selection bias.
#
#
# The exposition in our core textbook is slightly different. Here the term **bias** has two separate components which are **baseline bias** and **differential treatment effect bias**. See the discussion in the book in the subsection on the typical inconsistency and bias of the naive estimator. The term baseline bias refers to the concept of sorting and levels and selection bias.
#
# Differential treatment bias is defined as:
#
# \begin{align*}
# E[Y \mid D = 1] - E[Y \mid D = 0] & = \underbrace{E[\delta]}_{ATE} + \underbrace{\{E[Y^0\mid D= 1] - E[Y^0\mid D= 0]\}}_{\text{Baseline bias}} \\
# & + \underbrace{(1 - \pi) \{E[\delta \mid D = 1] -E[\delta \mid D = 0]\}}_{\text{Differential treatment effect bias}}
# \end{align*}
#
# The last term is derived derived from the term describing selection on gains by the following decomposition.
#
# \begin{align*}
# E[Y^1 - Y^0] = \pi E [Y^1 - Y^0 \mid D = 1] + (1 - \pi) E [Y^1 - Y^0 \mid D = 0]
# \end{align*}
#
# It is interpreted as the difference in effects between treated and control weighted by the share of control individuals. It is probably best thought of as an increment to the first term describing the average effect of treatment.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Assumptions
#
# So, the SUTVA assumption is only necessary but not sufficient to learn about the effect of treatment in light of the biases discussed above. We are still stuck with several unknowns that we need to compute the average effect of treatment.
#
# Consider the following two assumptions:
#
# \begin{align*}
# E[Y^1\mid D = 1] = E[Y^1\mid D = 0] \\
# E[Y^0\mid D = 1] = E[Y^0\mid D = 0] \\
# \end{align*}
#
# and recall our naive estimate
#
#
# \begin{align*}
# \hat{\delta}_{NAIVE} & = E_N[y_i\mid d_i = 1] - E_N[y_i\mid d_i = 0] \\
# & \xrightarrow{p} E[Y^1\mid D = 1] - E[Y^0\mid D = 0]
# \end{align*}
#
# * What assumptions suffice to estimate the ATE with the naive estimator?
#
# * about potential outcomes for subsets of the population
# * about the treatment selection / assignment process
# + [markdown] slideshow={"slide_type": "slide"}
# ### Missing data and imputation
# + [markdown] slideshow={"slide_type": "slide"}
# This is an adopted example from Imbens & Rubin (2015).
# + slideshow={"slide_type": "-"}
df = get_lalonde_data()
df.head()
# +
# There are some observations that miss values
df.loc[661]
# + [markdown] slideshow={"slide_type": "slide"}
# We can impute the missing values simply by their average counterpart.
# + slideshow={"slide_type": "-"}
is_treated = df["D"] == 1
df.loc[~is_treated, "Y_1"] = df["Y_1"].mean()
df.loc[is_treated, "Y_0"] = df["Y_0"].mean()
# -
df.head()
initial_stat = (df["Y_1"] - df["Y_0"]).mean()
print(f"Our estimated treatment effect is {initial_stat:10.2f}")
# + [markdown] slideshow={"slide_type": "slide"}
# However, this does not really account for any uncertainty in our estimate. Can we do better?
# -
df["D"].value_counts().to_dict()
# + slideshow={"slide_type": "-"}
np.random.seed(123) # set seed to ensure reproducibility
df = get_lalonde_data() # get the original data
status_counts = df["D"].value_counts().to_dict()
stats = list()
for _ in range(1000):
y_1_sampled = df["Y_1"].dropna().sample(n=status_counts[0], replace=True).values
y_0_sampled = df["Y_0"].dropna().sample(n=status_counts[1], replace=True).values
df_boot = df.copy()
is_treated = df_boot["D"] == 1
df_boot.loc[is_treated, "Y_0"] = y_0_sampled
df_boot.loc[~is_treated, "Y_1"] = y_1_sampled
stat = (df_boot["Y_1"] - df_boot["Y_0"]).mean()
stats.append(stat)
print(f"Our estimated treatment effect is {np.mean(stats):10.2f}")
# -
df_boot
# + [markdown] slideshow={"slide_type": "slide"}
# How does the full distribution of estimates look like?
# + slideshow={"slide_type": "-"}
fig, ax = plt.subplots()
ax.hist(stats)
ax.set_xlabel("Statistic")
ax.set_ylabel("Frequency")
ax.vlines(initial_stat, 0, 30, linestyles="--", label="Initial", color="lightgrey")
ax.legend()
# + [markdown] slideshow={"slide_type": "slide"}
# Still some limitations remains. For example, we do sample from the empirical distribution of the observed outcomes and not the actual distribution. Phrased differently, we treat the distribution of potential outcomes as known and abstract from any uncertainty in our knowledge about it.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Extensions of the binary potential outcome model
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true toc-nb-collapsed=true
# * over-time potential outcomes and causal effects
# * a single unit over time (time series data)
# * many units over time (panel data)
#
# * many-valued treatments
# + [markdown] slideshow={"slide_type": "slide"}
# #### Over-time potential outcomes
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true toc-nb-collapsed=true
# We explore the case of a single unit over time.
#
# **Ingredients**
#
# * discrete time periods, $t\in\{1, ..., T\}$
# * sequence of observed values, $\{y_1, y_2, ..., y_T\}$
# * treatment initiated in $t^*$
# * duration of treatment $k$
# + [markdown] slideshow={"slide_type": "slide"}
#
# Setting up the potential outcome model to explore the basic features of before-and-after designs for a single unit of analsysis.
#
# * before the treatment is introduced (for $t < t^*$):
#
# \begin{align*}
# D_t & = 0 \\
# Y_t & = Y^0_t\\
# \end{align*}
#
# * while the treatment is in place (from $t^*$ through $t^* + k$):
#
# \begin{align*}
# D_t & = 1 \\
# Y_t & = Y^1_t\\
# Y^0_t & \text{exists but is counterfactual}
# \end{align*}
#
# * after the treatment ends (for time periods $t > (t^* + k)$):
# \begin{align*}
# D_t & = 0 \\
# Y_t & = Y^0_t\\
# Y^1_t & \text{exists but is counterfactual}
# \end{align*}
# + [markdown] slideshow={"slide_type": "slide"}
# The following is an adapted example from our textbook.
#
# #### Year of the fire horse
#
# We study the effect of Japanese folk belief that families who give birth to babies will suffer untold miseries. This example does not only illustrative the versatility of the potential outcome framework but also serves as an example that different approaches (informed by domain-expertise) can result in different reasonable imputations for the counterfactual outcome.
#
# <img src="material/fig-birth-rates.png" width="500">
#
# The treatment indicator is as follows: $D_{1966} = 1$ and $D_{\neq 1966} = 0$ and we are interested in its effect on the birth rate in Japan
#
# \begin{align*}
# \delta_{1966} = y^1_{1966} - y^0_{1966}.
# \end{align*}
#
# A reasonable approach is to estimate it by:
#
# \begin{align*}
# \hat{\delta}_{1966} & = y_{1966} - y^0_{1966}\\
# \end{align*}
# +
df = pd.read_csv("material/world_bank.csv", skiprows=4)
df.set_index("Country Code", inplace=True)
df.drop(["Indicator Name", "Indicator Code"], axis=1, inplace=True)
df = df.loc["JPN", "1960":"2017"]
df = df.to_frame()
df.index.name = "Year"
df.columns = ["Birth rate"]
df.sort_index(inplace=True)
df.index = df.index.astype(int)
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# Let's get to work.
# + slideshow={"slide_type": "-"}
fig, ax = plt.subplots()
ax.plot(df["Birth rate"].index, df["Birth rate"])
ax.set_ylabel("Birth rate")
ax.set_xlabel("Year")
# + slideshow={"slide_type": "slide"}
df.loc[slice(1960, 1970), "Birth rate"]
# -
# We can just take the year before or after treatment?
estimates = list()
for label, year in [("before", 1965), ("after", 1967)]:
y_0 = df.loc[year, "Birth rate"]
y_1 = df.loc[1966, "Birth rate"]
print(f" Using the year {label}, the treatment effect is {y_1 - y_0:10.5f}")
estimates.append(y_1 - y_0)
# Among demographers, there is the consensus that taking the average of 1963 and 1969 the way to go instead.
# + slideshow={"slide_type": "slide"}
y_0 = df.loc[[1963, 1969], "Birth rate"].mean()
y_1 = df.loc[1966, "Birth rate"]
print(" Another treatment effect is {:10.5f}".format(y_1 - y_0))
estimates.append(y_1 - y_0)
# -
# Now we have multiple effects of treatment. Which is it?
labels = ["Before", "After", "Average"]
fig, ax = plt.subplots()
ax.bar(labels, estimates)
ax.set_ylabel("Effect")
# ### Additional resources
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# * **<NAME>. (2020)**. *Potential outcome and directed acyclic graph approaches to causality: Relevance for empirical practice in economics*. https://arxiv.org/abs/1907.07271
# -
# ### Resources
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# * **<NAME>., and <NAME>. (2019)** . *Impact evaluation: Treatment effects and causal analysis*. Cambridge, England: Cambridge University Press.
#
#
# * **<NAME>., <NAME>. and <NAME>. (2006)**. Understanding instrumental variables in models with essential heterogeneity. In *Review of Economics and Statistics*, 88(3), 389–432.
#
#
# * **<NAME>. (1986)**. Statistics and causal inference. In *Journal of the American Statistical Association*, 81(396), 945–960.
#
#
# * **<NAME>., and <NAME>. (2015)**. *Causal inference in statistics, social, and biomedical sciences*. New York, NY: Cambridge University Press.
#
#
# * **<NAME>. (2002)**. Overt bias in observational studies. In *Observational studies* (pp. 71–104). New York, NY: Springer.
#
| lectures/potential-outcome-model/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center>
# <img width=800 src="./figures/ladisk.png">
# <br>
# <font size="8" color="f00e0e" face="garamond"><b>High-speed Image Based Experimental Modal Analysis & Open Source Tools</b></font>
# <font size="6" color="f00e0e" face="garamond"><br><br><b>Free Online Course</b></font>
# <img src="">
# <font size="5">June 29th, July 1st 2020</font>
#
# <b><NAME>$^a$, <NAME>$^a$ and <NAME>$^a$</b>
#
# $^a$ Faculty of Mechanical Engineering, University of Ljubljana
#
# \* Corresponding email: <EMAIL>
# </center>
# <b style="font-size:30px">``https://github.com/ladisk/sdPy``</b>
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><ul class="toc-item"><li><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Requirements" data-toc-modified-id="Requirements-0.0.0.1">Requirements</a></span></li></ul></li></ul></li></ul></li><li><span><a href="#Part-1:-How-to-start-with-Python-for-structural-dynamics" data-toc-modified-id="Part-1:-How-to-start-with-Python-for-structural-dynamics-1">Part 1: How to start with Python for structural dynamics</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#The-Anaconda-distribution" data-toc-modified-id="The-Anaconda-distribution-1.0.1">The Anaconda distribution</a></span></li><li><span><a href="#Jupyter-notebook" data-toc-modified-id="Jupyter-notebook-1.0.2">Jupyter notebook</a></span></li></ul></li><li><span><a href="#Before-the-course" data-toc-modified-id="Before-the-course-1.1">Before the course</a></span><ul class="toc-item"><li><span><a href="#Python-for-beginner-programmers" data-toc-modified-id="Python-for-beginner-programmers-1.1.1">Python for beginner programmers</a></span></li><li><span><a href="#Python-for-Matlab-users" data-toc-modified-id="Python-for-Matlab-users-1.1.2">Python for Matlab users</a></span></li></ul></li><li><span><a href="#Python's-scientific-and-numerical-libraries" data-toc-modified-id="Python's-scientific-and-numerical-libraries-1.2">Python's scientific and numerical libraries</a></span><ul class="toc-item"><li><span><a href="#SciPy" data-toc-modified-id="SciPy-1.2.1">SciPy</a></span><ul class="toc-item"><li><span><a href="#NumPy" data-toc-modified-id="NumPy-1.2.1.1"><a href="http://www.numpy.org/" target="_blank">NumPy</a></a></span></li><li><span><a href="#Matplotlib" data-toc-modified-id="Matplotlib-1.2.1.2"><a href="https://matplotlib.org/" target="_blank">Matplotlib</a></a></span></li><li><span><a href="#The-SciPy-library" data-toc-modified-id="The-SciPy-library-1.2.1.3"><a href="https://docs.scipy.org/doc/scipy/reference/" target="_blank">The SciPy library</a></a></span></li><li><span><a href="#Installing-additional-packages" data-toc-modified-id="Installing-additional-packages-1.2.1.4">Installing additional packages</a></span></li></ul></li></ul></li><li><span><a href="#pyFRF" data-toc-modified-id="pyFRF-1.3">pyFRF</a></span></li><li><span><a href="#pyEMA" data-toc-modified-id="pyEMA-1.4">pyEMA</a></span></li><li><span><a href="#Some-other-useful-packages" data-toc-modified-id="Some-other-useful-packages-1.5">Some other useful packages</a></span><ul class="toc-item"><li><span><a href="#LVM_read" data-toc-modified-id="LVM_read-1.5.1">LVM_read</a></span></li><li><span><a href="#FLife" data-toc-modified-id="FLife-1.5.2">FLife</a></span></li></ul></li></ul></li></ul></div>
# -
# ---
# #### Requirements
# For a list of the required software tools and instructions on how to use this template, please see the
#
# **[README document on the course repository landing page](https://github.com/ladisk/sdPy)**.
# ---
# # Part 1: How to start with Python for structural dynamics
# [Python](https://www.python.org/) is an **open-source**, interpreted, dynamically typed programming language.
#
# Its **availability** and **intuitive syntax** make it one of the most popular programming language choises in the scientific community.
# ### The Anaconda distribution
# [Anaconda](https://www.anaconda.com/products/individual) is one of the many distributions of Python that makes starting out with Python very easy, as it comes with:
#
# * More than 1,400 packages pre-installed
# * Anaconda navigator (desktop GUI)
# * **conda** package manager for additional package installation: https://anaconda.org/anaconda/repo
# ### Jupyter notebook
#
# Jupyter notebook is an interactive programming environment, and is included in the Anaconda distribution. Some of the characteristics of Jupyter notebook:
#
# * cell based programming (similar to Mathematica),
# * file extension **.ipynb**,
# * running each individual cell,
# * includes **Markdown** and **Code** type cells.
#
# **Start Jupyter notebook by running:**
# ```cmd
# $ jupyter notebook
# ```
#
# in a comand prompt / terminal window in your local directory.
# <img src="figures/jupyter_notebook_gif.gif" width="1000px">
# ---
# ## Before the course
# ### Python for beginner programmers
# If you are just starting out with programming and you have already successfully installed a version of Python on your machine, we suggest going over a beginner Python tutorial before following this course.
#
# [Scipy lecture notes](http://scipy-lectures.org/index.html) are a great resource for aspiring Python users, focused more on the scientific community.
# Sections [1.1](http://scipy-lectures.org/intro/intro.html) and [1.2](http://scipy-lectures.org/intro/language/python_language.html) of Scipy lecture notes will provide more than enough info to get you started.
#
# If you want to learn even more about Python, you will find countless tutorials available online, but the [official Python tutorial](https://docs.python.org/3/tutorial/index.html) is a good place to start.
#
# You can also find many more great resources in the [official Python beginners guide](https://wiki.python.org/moin/BeginnersGuide).
# ### Python for Matlab users
# Many researchers and engineers learning Python come from a background of Matlab or a similar commercial software package. A lot of effort has gone into making this transition as simple as possible.
#
# The [Scipy](https://www.scipy.org/) project includes all the tools required to make Python a powerful alternative to most comercially available numerical packages. If you are an experienced Matlab user, the [NumPy for Matlab users](https://numpy.org/doc/stable/user/numpy-for-matlab-users.html) page contains important information to quickly get you started.
# ---
# ## Python's scientific and numerical libraries
# Python comes with some very useful features [out of-the-box](https://docs.python.org/3/library/):
#
# * numerical and mathematical datatypes and functions,
# * filesystem management tools,
# * data compression tools,
# * support for various networking and internet protocols ([http](https://docs.python.org/3/library/http.html), [email](https://docs.python.org/3/library/email.html), [websockets](https://docs.python.org/3/library/socket.html), ...),
# text processing and binary data manipulation,
# * development tools (documentation, unit-testing, ...)
# * The **[pip package maager](https://pip.pypa.io/en/stable/)**,
# * ...
# With the Anaconda distribution of Python you also get Python's essential numerical packages pre-installed:
# ### SciPy
#
# The [SciPy](https://www.scipy.org/index.html) stack is a collection of open-source Python software for science and engineering. To get more information on how to use Scientific Python, see the [Getting started with SciPy page](https://www.scipy.org/getting-started.html).
#
# The SciPy stack consists of six core packages, including:
# ---
# #### [NumPy](http://www.numpy.org/)
#
# * Adds support for the efficient multidimentional ``ndarray`` data object for numerical data representation.
# * Functions for *numerical data manipulation*, *linear algebra* and *the Fourier transform*.
import numpy as np
L = [1, 2, 3, 4, 5] # This is a list
a = np.array(L) # This is an array
a
# Note the difference:
2*L
2*a
# A range of numbers:
a1 = np.arange(0, 10, 2)
a1
# Linearly spaced numbers:
a2 = np.linspace(0, 10, 6, dtype=complex)
a2
# Multidimensional arrays are used to represent matrices:
M = np.random.rand(3, 3) # This is a matrix
M
# **For more information on the NumPy library, see the [official NumPy tutorial](https://numpy.org/devdocs/user/basics.html).**
# ---
# #### [Matplotlib](https://matplotlib.org/)
#
# * A plotting library for producing publication-ready data visualizations.
# * A MATLAB-inspired user interface.
#
import matplotlib.pyplot as plt
# %matplotlib inline
# +
x = np.linspace(0, 1, 40)
y1 = np.sin(2*np.pi*2*x)
y2 = np.cos(2*np.pi*2*x)
plt.plot(x, y1, 'o', label='sin')
plt.plot(x, y2, 'o', label='cos')
plt.legend()
plt.xlabel('x')
plt.ylabel('y')
plt.title('Sin and Cos')
plt.grid(True)
plt.show()
# -
# **For more information on the Matplotlib library, see the [official Matplotlib tutorials page](https://matplotlib.org/tutorials/index.html).**
# ---
# #### [The SciPy library](https://docs.scipy.org/doc/scipy/reference/)
#
# provides user-friendly and efficient numerical routines for:
#
# * Signal processing,
# * numerical integration,
# * optimization,
# * interpolation.
#
# all based on the NumPy `ndarray` class.
from scipy.interpolate import InterpolatedUnivariateSpline
# +
x_int = np.linspace(0, 1, 1000)
spl = InterpolatedUnivariateSpline(x, y1, k=3)
plt.plot(x_int, spl(x_int), '.', label='Interpolated')
plt.plot(x, y1, 'o', label='Original')
plt.legend()
plt.xlabel('x')
plt.ylabel('y')
# -
# **For more information on the SciPy library, see the [official Scipy tutorial](https://docs.scipy.org/doc/scipy/reference/tutorial/).**
# ---
# #### Installing additional packages
# For more specific tasks (such as experimental modal analysis), there is a good chance that you will be able to find an existing package in the [Python Package Index (PyPI)](https://pypi.org/) database of over 200 000 open source packages.
#
# To install a chosen package, use the `pip` package manager by running the following in the command prompt / terminal:
#
# ```cmd
# $ pip install <package name>
# ```
# You can also run commands from a Jupyter Notebook code cell by prefixing them with the `!` character:
# !pip install --upgrade pyFRF
# You can also easily install all the required packages defined in a `requirements.txt` file by running:
# pip_response = !pip install -r requirements.txt
# ## pyFRF
# pyFRF is an open-source package, designed for frequency resposne function measurements in structural dynamics.
#
# The inputs are time signals of excitation and response, the outputs are FRF estimators (H1, H2, Hv, Vector or ODS) and coherence.
#
# You can find more info on the [pyFRF GitHub repository webpage](https://github.com/openmodal/pyFRF).
from pyFRF import FRF
# To automaticlly draw matplotlib figures under code cells
# %matplotlib inline
# **Prepare synthetic FRF data**
# +
C = 0.5+0.1j # modal constant
eta = 5e-3 # damping loss factor
f0 = 320 # natural frequency
df = 1 # freq resolution
D = 1e-8*(1-.1j) # residual
f = 1*np.arange(0, 1400, step=df) # / frequency range
w0 = f0 * 2 * np.pi #to rad/s
w = f * 2 * np.pi
H1_syn = C / (w0 ** 2 - w ** 2 + 1.j * eta * w0 ** 2) + \
+0.5*np.conj(C) / ((w0*2)** 2 - w ** 2 + 1.j * eta * (w0*2)** 2)\
+0.25*C / ((w0*3)** 2 - w ** 2 + 1.j * eta * (w0*3)** 2)\
+ D
# -
# Visualize prepared FRF
# +
fig, ax1 = plt.subplots(figsize=(10, 5))
ax1.semilogy(f,np.abs(H1_syn), 'b')
ax1.set_xlabel('Frequency [Hz]')
ax1.set_ylabel('H1', color='b')
for tl in ax1.get_yticklabels():
tl.set_color('b')
ax2 = ax1.twinx()
ax2.plot(f,np.angle(H1_syn), 'r', alpha=0.2)
ax2.set_ylabel('Angle', color='r')
for tl in ax2.get_yticklabels():
tl.set_color('r')
# -
# **Prepare synthetic impulse response function** by IFFT of the synthetic FRF
h = np.fft.irfft(H1_syn)
l = len(H1_syn)*2-2
t = np.linspace(0, 1, num=l)
exc = np.zeros_like(t)
exc[0] = 1
# +
fig, ax1 = plt.subplots(figsize=(10, 5))
ax1.plot(t, exc, 'b');
ax1.set_xlabel('Time [s]')
ax1.set_ylabel('Excitation', color='b')
ax1.set_xlim(left=0, right=1)
for tl in ax1.get_yticklabels():
tl.set_color('b')
ax2 = ax1.twinx()
ax2.plot(t, h, 'r', alpha=0.7)
ax2.set_ylabel('Response', color='r')
for tl in ax2.get_yticklabels():
tl.set_color('r')
# -
# **Transform the temporal data back into frequency domain via pyFRF**
frf = FRF(sampling_freq=1/t[1], exc=exc, resp=h, resp_type='d', exc_window='None', resp_window='None')
freq = frf.get_f_axis()
H1 = frf.get_FRF(type='H1')
plt.figure(figsize=(10, 5))
plt.semilogy(freq, np.abs(H1), '.', label='via pyFRF')
plt.semilogy(f, np.abs(H1_syn), label='Synthetic')
plt.title('FRF H1')
plt.xlabel('Frequency [Hz]')
plt.legend();
# Different FRF types
plt.figure(figsize=(10, 5))
plt.semilogy(freq, np.abs(frf.get_FRF(form='accelerance')), '.', label='Accelerance')
plt.semilogy(freq, np.abs(frf.get_FRF(form='mobility')), '.', label='Mobility')
plt.semilogy(freq, np.abs(frf.get_FRF(form='receptance')), '.', label='Receptance')
plt.title('FRF H1')
plt.xlabel('Frequency [Hz]')
plt.legend();
# **Multiple measurement with noise and averaging**
averages = 10
frf = FRF(sampling_freq=1/t[1], fft_len=len(h), exc_window='None', \
resp_window='None', resp_type='d', weighting='Linear', n_averages=averages)
k = 0.02 # rate of noise
for i in range(averages):
noise = k * (np.random.rand(len(h))-0.5) * np.std(h)
frf.add_data(exc, h + noise)
fig, ax = plt.subplots(2, 1, figsize=(10, 10))
ax[0].semilogy(frf.get_f_axis(), np.abs(frf.get_H1()), '.', label='via FRF')
ax[0].semilogy(f, np.abs(H1_syn), label='Synthetic')
ax[0].set_xlabel('Frequency [Hz]')
ax[0].set_ylabel('H1 FRF')
ax[0].legend()
ax[1].plot(frf.get_f_axis(), frf.get_coherence(), '.')
ax[1].set_xlabel('Frequency [Hz]')
ax[1].set_ylabel('coherence');
# ---
# ## pyEMA
# pyEMA is an open source experimental modal analysis package for Python.
#
# It implements the Least-Squares Complex Frequency (LSCF) modal identification method, as well as the Least-Squares Frequency Domain (LSFD) modal constant identification algorithm.
#
# Visit the [pyEMA GitHub repository page](https://github.com/ladisk/pyEMA) for more info.
import pyEMA
# The experiment is shown in the figure below: the beam was excited at 6 locations with an impact hammer, while the response was measured at 7 locations using piezo accelerometers (camera data is not used in this showcase):
# <img width=500 src="./figures/experiment_1_pyEMA.jpg">
# Loading experimental data:
freq, H1_main = np.load("./data/acc_data.npy", allow_pickle=True)
H1_main.shape
# From the experimental FRF array `H1_main` of dimensions: `(#inputs, #outputs, frequency)` only the response accelerometer position at index 1 will be used for analysis:
selected_response = 1
FRF = H1_main[:, selected_response, :]
plt.figure(figsize=(10, 5))
plt.semilogy(freq, np.abs(FRF.T));
plt.ylabel('FRF H1')
plt.xlabel('Frequency [Hz]')
plt.xlim(0, 1000);
# Initiate the pyEMA object:
acc = pyEMA.Model(frf=FRF,
freq=freq,
lower=10,
upper=5000,
pol_order_high=60)
# Get the complex system poles:
acc.get_poles()
# Select stable poles from stabilisation chart:
acc.select_poles()
# FRF reconstruction:
frf_rec, modal_const = acc.get_constants(whose_poles='own', FRF_ind='all')
# The selected natural frequencies and corresponding damping factors:
acc.print_modal_data()
# Modal constants are of shape (`n_locations` $\times$ `n_frequencies`):
acc.A.shape
# Show normalized modes (first three):
plt.figure(figsize=(10, 5))
plt.plot(acc.normal_mode()[:, :3]);
plt.xlabel('DOF')
plt.ylabel('mode shape')
# Check out the Auto Modal Assurance Criterion:
autoMAC = acc.autoMAC()
plt.figure(figsize=(6, 6))
plt.imshow(autoMAC);
# Showing the reconstructed FRF:
# +
freq_a = acc.freq
select_loc = 0
plt.figure(figsize = ((10, 10)))
plt.subplot(211)
plt.semilogy(freq, np.abs(FRF[select_loc]), label='Experiment')
plt.semilogy(freq_a, np.abs(frf_rec[select_loc]),'--', label='LSCF')
plt.xlim(0,freq[-1])
plt.ylabel(r"abs($\alpha$)")
plt.legend(loc = 'best')
plt.subplot(212)
plt.plot(freq, np.angle(FRF[select_loc],deg = 1), label='Experiment')
plt.plot(freq_a, np.angle(frf_rec[select_loc],deg = 1),'--',label='LSCF')
plt.xlim(0,freq[-1])
plt.ylabel(r"angle($\alpha$)")
plt.legend(loc = 'best');
# -
# ---
# ## Some other useful packages
# ### LVM_read
#
# LVM_read is an open source tool used to efficiently work with NI LabView `.lvm` files in Python.
#
# Visit the [lvm_read GitHub repository page](https://github.com/openmodal/lvm_read) for more info.
import lvm_read
# Read the sample file with `lvm_read`
filename = 'short.lvm'
lvm = lvm_read.read('./data/' + filename)
# Header of the lvm file:
lvm.keys()
# Number of measurement segments in the lvm file:
lvm['Segments']
# Segments (measurements) are accessed by index:
lvm[0]
# Segment data:
lvm[0]['data']
dt = lvm[0]['Delta_X'][0]
time = np.arange(lvm[0]['data'].shape[0]) * dt
plt.figure(figsize=(10, 5))
plt.plot(time, lvm[0]['data'][:, 0], label=lvm[0]['Channel names'][0]);
plt.plot(time, lvm[0]['data'][:, 1], label=lvm[0]['Channel names'][1]);
plt.xlabel('time [s]')
plt.ylabel('measurement')
plt.legend();
# ---
# ### FLife
# FLife is an open-source Python package for obtaining vibration fatigue life in the spectral domain.
#
# For more information, see the [FLife GitHub repository page](https://github.com/ladisk/FLife).
import FLife
# Generate time history data:
dt = 1e-4
x = np.random.normal(scale=100, size=10000)
time = np.arange(len(x)) * dt
plt.figure(figsize=(10, 5))
plt.plot(time, x)
plt.xlabel('time [s]')
plt.ylabel('input signal');
# Get spectral data:
sd = FLife.SpectralData(input=x, dt=dt)
# Initialize methods:
rf = FLife.Rainflow(sd) # Rainflow reference fatigue life
# Spectral fatigue methods:
dirlik = FLife.Dirlik(sd)
tb = FLife.TovoBenasciutti(sd)
# Set material fatigue parameters:
C = 1.8e+22
k = 7.3
# Calculate predicted fatigue life:
life_rf = rf.get_life(C=C, k=k)
life_dirlik = dirlik.get_life(C=C, k=k)
life_tb = tb.get_life(C=C, k=k)
print(f'{"Rainflow":>20s}: {life_rf:4.0f} s')
print(f'{"Dirlik":>20s}: {life_dirlik:4.0f} s')
print(f'{"Tovo Benasciutti 2":>20s}: {life_tb:4.0f} s')
| Part 1 - Open-source Python tools for structural dynamics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="6mmkhyF-nU5m" outputId="8ea64613-9a04-4a14-f9ee-171314ecbacd"
# !mkdir ~/.kaggle
# !cp kaggle.json ~/.kaggle/
# !chmod 600 ~/.kaggle/kaggle.json
# !kaggle datasets download -d mlg-ulb/creditcardfraud
# !unzip creditcardfraud.zip
# + id="lBBSPc6DrXvi" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="6db42504-676b-4049-ebf7-0714ca124657"
import pandas as pd
df = pd.read_csv('creditcard.csv')
df
# + id="n-0kbp6ssnTa" colab={"base_uri": "https://localhost:8080/"} outputId="a9ec6e5a-93f8-4f48-d138-c8da28d8abf8"
df['Class'].value_counts()
# + id="TSAtn7dUukSM" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="cf8b0c21-601a-44a9-e8ff-47531158bca3"
df.hist(bins=30, figsize=(30, 30))
# + id="j47A5v2MvVsm" colab={"base_uri": "https://localhost:8080/", "height": 317} outputId="19141553-2c1a-45d0-ffe7-3dc4408be5c5"
df.describe()
# + id="LCy-CQ9OvZfL" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="61f1095c-e1cf-4e5b-d53d-8f15e34ccb15"
from sklearn.preprocessing import RobustScaler
new_df = df.copy()
new_df['Amount'] = RobustScaler().fit_transform(new_df['Amount'].to_numpy().reshape(-1, 1))
time = new_df['Time']
new_df['Time'] = (time - time.min()) / (time.max() - time.min())
new_df
# + id="304Tn5bE2vs8" colab={"base_uri": "https://localhost:8080/"} outputId="74a08c7a-d5b1-446b-9dff-11333dd6fbd1"
new_df['Amount'].describe()
# + id="DbuxlisR2ze9" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="3225541b-03ba-4b1f-b01c-53eaca9b19f1"
new_df = new_df.sample(frac=1, random_state=1)
new_df
# + id="kziAdyqs3iW_" colab={"base_uri": "https://localhost:8080/"} outputId="46018ed2-380a-4212-ac5e-5e900cca996a"
train, test, val = new_df[:240000], new_df[240000:262000], new_df[262000:]
train['Class'].value_counts(), test['Class'].value_counts(), val['Class'].value_counts()
# + id="CuIIpKQE4zX8" colab={"base_uri": "https://localhost:8080/"} outputId="e489aba3-5454-4b73-9741-d720cbec5346"
train_np, test_np, val_np = train.to_numpy(), test.to_numpy(), val.to_numpy()
train_np.shape, test_np.shape, val_np.shape
# + id="bLaGQw7S5DWh" colab={"base_uri": "https://localhost:8080/"} outputId="b2a52719-9457-43d5-d118-9ca62148d684"
x_train,y_train= train_np[:,:-1] , train_np[:,-1]
x_test,y_test= train_np[:,:-1] , train_np[:,-1]
x_val,y_val= train_np[:,:-1] , train_np[:,-1]
x_train.shape, y_train.shape, x_test.shape, y_test.shape, x_val.shape, y_val.shape
# + colab={"base_uri": "https://localhost:8080/"} id="5jlMkPisQheQ" outputId="ac7eb8de-0d66-4911-bf13-0e9c3ee59c46"
from sklearn.linear_model import LogisticRegression
logistic_model = LogisticRegression()
logistic_model.fit(x_train, y_train)
logistic_model.score(x_train,y_train)
# + id="RhOihg_fVvM6" colab={"base_uri": "https://localhost:8080/"} outputId="d7697b02-7e15-4de1-9dfc-a3bb5a8c7d8a"
from sklearn.metrics import classification_report
print(classification_report(y_val, logistic_model.predict(x_val), target_names=['Not Fraud','Fraud']))
# + id="L48bOJfMbnJH"
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import InputLayer, Dense, BatchNormalization
from tensorflow.keras.callbacks import ModelCheckpoint
shallow_nn = Sequential()
shallow_nn.add(InputLayer((x_train.shape[1],)))
shallow_nn.add(Dense(2, 'relu'))
shallow_nn.add(BatchNormalization())
shallow_nn.add(Dense(1, 'sigmoid'))
checkpoint = ModelCheckpoint('shallow_nn', save_best_only=True)
shallow_nn.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
# + id="iph4AioIo1PU" colab={"base_uri": "https://localhost:8080/"} outputId="9f5f72b7-67e7-4d41-8913-5bd4bd9130c3"
shallow_nn.fit(x_train,y_train,validation_data=(x_val,y_val),epochs=5,callbacks=checkpoint)
# + id="SXighoHAiDb-" colab={"base_uri": "https://localhost:8080/"} outputId="af365e49-dca7-4b67-f4c9-4c99d66e0a75"
def neural_net_predictions(model, x):
return (model.predict(x).flatten() > 0.5).astype(int)
neural_net_predictions(shallow_nn, x_val)
# + id="wTG8l1qgrKJF" colab={"base_uri": "https://localhost:8080/"} outputId="2c3ef925-084e-4150-db59-b7d9eb60569f"
print(classification_report(y_val, neural_net_predictions(shallow_nn, x_val), target_names=['Not Fraud', 'Fraud']))
# + id="u5F_4sNLjN6w" colab={"base_uri": "https://localhost:8080/"} outputId="0750162d-5cff-4708-fed9-a0f8f507eed4"
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(max_depth=2, n_jobs=-1)
rf.fit(x_train, y_train)
print(classification_report(y_val, rf.predict(x_val), target_names=['Not Fraud', 'Fraud']))
# + id="gr2nkk5xyJVS" colab={"base_uri": "https://localhost:8080/"} outputId="c3673906-ff77-4f6a-b864-b261c5aeff03"
from sklearn.ensemble import GradientBoostingClassifier
gbc = GradientBoostingClassifier(n_estimators=50,learning_rate=1.0,max_depth=1,random_state=0)
gbc.fit(x_train,y_train)
print(classification_report(y_val, gbc.predict(x_val), target_names=['Not Fraud', 'Fraud']))
# + id="g7q-wlFA7MX_" colab={"base_uri": "https://localhost:8080/"} outputId="16352b18-9c26-49de-c215-6ad984d1e4aa"
from sklearn.svm import LinearSVC
svc = LinearSVC(class_weight='balanced')
svc.fit(x_train, y_train)
print(classification_report(y_val, svc.predict(x_val), target_names=['Not Fraud', 'Fraud']))
# + id="_k37FNeH8CKj" colab={"base_uri": "https://localhost:8080/"} outputId="462fbb18-27cb-4b65-f375-f64b703bc352"
#The data set that we have is not balanced at all.The amount of fraud Entries is very less Thus we need to establish some sort of balance
not_frauds = new_df.query('Class==0')
frauds = new_df.query('Class==1')
not_frauds['Class'].value_counts(), frauds['Class'].value_counts()
# + id="rrUyOUFy-LkW" colab={"base_uri": "https://localhost:8080/"} outputId="ac005bf0-7960-4a60-9964-81275289b1bb"
balanced_df = pd.concat([frauds, not_frauds.sample(len(frauds), random_state=1)])
balanced_df['Class'].value_counts()
# + id="geiUxgH4u61N" colab={"base_uri": "https://localhost:8080/", "height": 439} outputId="d31ca85c-74a5-4f45-8f53-60ecbba2cbb3"
balanced_df = balanced_df.sample(frac=1, random_state=1)
balanced_df
# + id="v54scR_lvA8k" colab={"base_uri": "https://localhost:8080/"} outputId="051e6511-4a97-4b93-9d2e-b5181226533b"
balanced_df_np = balanced_df.to_numpy()
x_train_b, y_train_b = balanced_df_np[:700, :-1], balanced_df_np[:700, -1].astype(int)
x_test_b, y_test_b = balanced_df_np[700:842, :-1], balanced_df_np[700:842, -1].astype(int)
x_val_b, y_val_b = balanced_df_np[842:, :-1], balanced_df_np[842:, -1].astype(int)
x_train_b.shape, y_train_b.shape, x_test_b.shape, y_test_b.shape, x_val_b.shape, y_val_b.shape
# + id="9cs221ua-sx0" colab={"base_uri": "https://localhost:8080/"} outputId="7f3f09d6-db61-47a1-b8b8-fa23291a8355"
pd.Series(y_train_b).value_counts(), pd.Series(y_test_b).value_counts(), pd.Series(y_val_b).value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="r3CPWOa7xX3n" outputId="e92962f7-8732-4ae7-ba04-80e84bb4fcfc"
logistic_model_b = LogisticRegression()
logistic_model_b.fit(x_train_b, y_train_b)
print(classification_report(y_val_b, logistic_model_b.predict(x_val_b), target_names=['Not Fraud', 'Fraud']))
# + colab={"base_uri": "https://localhost:8080/"} id="RzCOUXjVxuV8" outputId="d0af1d72-d659-4674-9cbd-b86bb31ca0a4"
shallow_nn_b = Sequential()
shallow_nn_b.add(InputLayer((x_train.shape[1],)))
shallow_nn_b.add(Dense(2, 'relu'))
shallow_nn_b.add(BatchNormalization())
shallow_nn_b.add(Dense(1, 'sigmoid'))
checkpoint = ModelCheckpoint('shallow_nn_b', save_best_only=True)
shallow_nn_b.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
shallow_nn_b.fit(x_train_b, y_train_b, validation_data=(x_val_b, y_val_b), epochs=40, callbacks=checkpoint)
# + colab={"base_uri": "https://localhost:8080/"} id="5arYFLmayluv" outputId="3f57ef64-5ff0-4d2b-8661-264767295c99"
shallow_nn_b.fit(x_train_b, y_train_b, validation_data=(x_val_b, y_val_b), epochs=40, callbacks=checkpoint)
# + colab={"base_uri": "https://localhost:8080/"} id="wPNYLI0Dy7nv" outputId="c9090f83-a9ab-457b-8c14-c885a0f51575"
print(classification_report(y_val_b, neural_net_predictions(shallow_nn_b, x_val_b), target_names=['Not Fraud', 'Fraud']))
# + colab={"base_uri": "https://localhost:8080/"} id="ins6fP42zOxr" outputId="c27a0bd4-707d-4d8d-ba91-b0a8db6b2e23"
shallow_nn_b1 = Sequential()
shallow_nn_b1.add(InputLayer((x_train.shape[1],)))
shallow_nn_b1.add(Dense(1, 'relu'))
shallow_nn_b1.add(BatchNormalization())
shallow_nn_b1.add(Dense(1, 'sigmoid'))
checkpoint = ModelCheckpoint('shallow_nn_b1', save_best_only=True)
shallow_nn_b1.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
shallow_nn_b1.fit(x_train_b, y_train_b, validation_data=(x_val_b, y_val_b), epochs=40, callbacks=checkpoint)
# + colab={"base_uri": "https://localhost:8080/"} id="JgSJmPLCzdGI" outputId="5e4136c0-544a-4134-ae12-a89e59b662e8"
shallow_nn_b1.fit(x_train_b, y_train_b, validation_data=(x_val_b, y_val_b), epochs=40, callbacks=checkpoint)
# + colab={"base_uri": "https://localhost:8080/"} id="vxuDOTFE0DKB" outputId="ce26fd75-d433-4113-c915-17d9612178b9"
print(classification_report(y_val_b, neural_net_predictions(shallow_nn_b1, x_val_b), target_names=['Not Fraud', 'Fraud']))
# + colab={"base_uri": "https://localhost:8080/"} id="L6__IX5K0NGV" outputId="681bcafb-0c9c-4f5c-b77f-f4eb6e8ba308"
rf_b = RandomForestClassifier(max_depth=2, n_jobs=-1)
rf_b.fit(x_train_b, y_train_b)
print(classification_report(y_val_b, rf.predict(x_val_b), target_names=['Not Fraud', 'Fraud']))
# + colab={"base_uri": "https://localhost:8080/"} id="0SSGFY_g0R6Y" outputId="65ddbdf0-63e1-45d3-de3b-f09218f69691"
gbc_b = GradientBoostingClassifier(n_estimators=50, learning_rate=1.0, max_depth=2, random_state=0)
gbc_b.fit(x_train_b, y_train_b)
print(classification_report(y_val_b, gbc.predict(x_val_b), target_names=['Not Fraud', 'Fraud']))
# + id="IvZfLE6N0oRH"
| Detecting_Credit_card_Fraud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Lab 4 - Model Training with AutomatedML
#
# In this lab you will use the automated machine learning (*AutomatedML*) capabilities within the Azure Machine Learning service.
#
# Automated machine learning picks an algorithm and hyperparameters for you and generates a model ready for deployment.
#
#
#
# 
#
#
#
# We will continue with the same scenario as in Lab 1 and Lab 2.
# ## Connect to the workspace
# Verify AML SDK Installed
# view version history at https://pypi.org/project/azureml-sdk/#history
import azureml.core
print("SDK Version:", azureml.core.VERSION)
# +
from azureml.core import Workspace
# Read the workspace config from file
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\n')
# -
# ## Train a model using AutomatedML
#
#
# To train a model using AutoML you need only provide a configuration for AutoML that defines items such as the type of model (classification or regression), the performance metric to optimize, exit criteria in terms of max training time and iterations and desired performance, any algorithms that should not be used, and the path into which to output the results. This configuration is specified using the AutomMLConfig class, which is then used to drive the submission of an experiment via experiment.submit. When AutoML finishes the parent run, you can easily get the best performing run and model from the returned run object by using run.get_output().
#
# ### Create/Get Azure ML Compute cluster
#
# We are reusing the cluster created in Lab 1. In case you removed the cluster the below code snippet is going to re-create it.
# +
# Create an Azure ML Compute cluster
# Create Azure ML cluster
from azureml.core.compute import AmlCompute
from azureml.core.compute import ComputeTarget
import os
# choose a name for your cluster
cluster_name = "cpu-cluster"
cluster_min_nodes = 1
cluster_max_nodes = 3
vm_size = "STANDARD_DS11_V2"
# Check if the cluster exists. If yes connect to it
if cluster_name in ws.compute_targets:
compute_target = ws.compute_targets[cluster_name]
if compute_target and type(compute_target) is AmlCompute:
print('Found existing compute target, using this compute target instead of creating: ' + cluster_name)
else:
print("Error: A compute target with name ",cluster_name," was found, but it is not of type AmlCompute.")
else:
print('Creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,
min_nodes = cluster_min_nodes,
max_nodes = cluster_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current BatchAI cluster status, use the 'status' property
print(compute_target.status.serialize())
# -
# ### Create Get Data script
#
# If you are using a remote compute to run your Automated ML experiments - which is our scenario, the data fetch must be wrapped in a separate python script that implements get_data() function. This script is run on the remote compute where the automated ML experiment is run. get_data() eliminates the need to fetch the data over the wire for each iteration.
import os
project_folder = './project'
script_name = 'get_data.py'
os.makedirs(project_folder, exist_ok=True)
# +
# %%writefile $project_folder/get_data.py
import pandas as pd
import numpy as np
import os
def get_data():
# Load the dataset
data_folder = os.environ["AZUREML_DATAREFERENCE_workspaceblobstore"]
file_name = os.path.join(data_folder, 'banking_train.csv')
df = pd.read_csv(file_name)
# Preprocess the data
feature_columns = [
# Demographic
'age',
'job',
'education',
'marital',
'housing',
'loan',
# Previous campaigns
'month',
'campaign',
'poutcome',
# Economic indicators
'emp_var_rate',
'cons_price_idx',
'cons_conf_idx',
'euribor3m',
'nr_employed']
df = df[feature_columns + ['y']]
features = df.drop(['y'], axis=1)
# Flatten labes
labels = np.ravel(df.y)
return { "X" : features, "y" : labels}
# -
# ### Configure datastore and data reference
#
# The training files have been uploaded to the workspace's default datastore during the previous labs. We will configure AutomatedML to automatically download the files onto the nodes of the cluster.
# +
from azureml.core import Datastore
from azureml.core.runconfig import DataReferenceConfiguration
ds = ws.get_default_datastore()
print("Using the default datastore for training data: ")
print(ds.name, ds.datastore_type, ds.account_name, ds.container_name)
dr = DataReferenceConfiguration(datastore_name=ds.name,
path_on_datastore='datasets',
path_on_compute='datasets',
mode='download', # download files from datastore to compute target
overwrite=True)
# -
# ### Create Docker run configuration
# We will run Automated ML jobs in a custom docker image that will include dependencies required by get_data() script.
# +
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core import Run
from azureml.core import ScriptRunConfig
# create a new RunConfig object
run_config = RunConfiguration(framework="python")
# Azure ML Compute cluster for Automated ML jobs require docker.
run_config.environment.docker.enabled = True
# Set compute target to Azure ML Compute cluster
run_config.target = compute_target
# Set data references
run_config.data_references = {ds.name: dr}
# -
#
# ### Configure Automated ML run.
#
# Automated ML runs can be controlled using a number of configuration parameters.
#
#
# For our run we will use the following configuration:
# - Train a classification task
# - Execute at most 25 iterations
# - Use *normalized macro recall* as a primary performance metrics
# - Use 5-fold cross validation for model evaluation
# - Run the iterations on 3 nodes of a cluster
# - Use 1 core per iteration
# - Automatically pre-process data
# - Exit if the primary metrics is higher than 0.9
# - Limit the model selection to *SVM*, *LogisticRegression*, *LightGBM*, *TensorFlowDNN* and *RandomForest* models
#
# We configured the last setting to demonstrate *white listing* capabilities of *AutomatedML*. Unless you have a strong basis for excluding or choosing certain models you are usually better of leaving the decision to *AutomatedML* - assuming that you have enough time and resources for running through many (more than 100) iterations.
#
#
# We have configured our run to automatically pre-process data.
#
# As a result, the following data preprocessing steps are performed automatically:
# 1. Drop high cardinality or no variance features
# * Drop features with no useful information from training and validation sets. These include features with all values missing, same value across all rows or with extremely high cardinality (e.g., hashes, IDs or GUIDs).
# 1. Missing value imputation
# * For numerical features, impute missing values with average of values in the column.
# * For categorical features, impute missing values with most frequent value.
# 1. Generate additional features
# * For DateTime features: Year, Month, Day, Day of week, Day of year, Quarter, Week of the year, Hour, Minute, Second.
# * For Text features: Term frequency based on word unigram, bi-grams, and tri-gram, Count vectorizer.
# 1. Transformations and encodings
# * Numeric features with very few unique values transformed into categorical features.
# * Depending on cardinality of categorical features, perform label encoding or (hashing) one-hot encoding.
#
# +
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
import logging
automl_config = AutoMLConfig(run_configuration = run_config,
task = 'classification',
debug_log = 'automl_errors.log',
primary_metric = 'norm_macro_recall',
iterations = 25,
n_cross_validations = 5,
max_concurrent_iterations = cluster_max_nodes,
max_cores_per_iteration = 1,
preprocess = True,
experiment_exit_score = 0.99,
#blacklist_models = ['KNN','MultinomialNaiveBayes', 'BernoulliNaiveBayes'],
whitelist_models = ['LogisticRegression', 'RandomForest', 'LightGBM', 'SVM', 'TensorFlowDNN'],
verbosity = logging.INFO,
path = project_folder,
data_script = os.path.join(project_folder, script_name))
# -
# ### Run AutomatedML job.
# +
from azureml.core import Experiment
experiment_name = "propensity_to_buy_classifier_automatedml"
exp = Experiment(ws, experiment_name)
tags = {"Desc": "automated ml"}
run = exp.submit(config=automl_config, tags=tags)
run
# -
# The call to experiment returns `AutoMLRun` object that can be used to track the run.
#
# Since the call is asynchronous, it reports a **Preparing** or **Running** state as soon as the job is started.
#
# Here is what's happening while you wait:
#
# - **Image creation**: A Docker image is created matching the Python environment specified by the RunConfiguration. The image is uploaded to the workspace. This happens only once for each Python environment since the container is cached for subsequent runs. During image creation, logs are streamed to the run history. You can monitor the image creation progress using these logs.
#
# - **Scaling**: If the remote cluster requires more nodes to execute the run than currently available, additional nodes are added automatically.
#
# - **Running**: In this stage, the Automated ML takes over and starts running experiments
#
#
#
# You can check the progress of a running job in multiple ways: Azure Portal, AML Widgets or streaming logs.
#
# ### Monitor the run.
#
# We will use AML Widget to monitor the run. The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# The widget is asynchronous - it does not block the notebook. You can execute other cells while the widget is running.
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.
from azureml.widgets import RunDetails
RunDetails(run).show()
# ### Cancelling Runs
#
# You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions.
# +
# Cancel the ongoing experiment and stop scheduling new iterations.
# run.cancel()
# Cancel iteration 1 and move onto iteration 2.
# run.cancel_iteration(1)
# -
# ### Analyze the run
#
# You can use SDK methods to fetch all the child runs and see individual metrics that we log.
# +
import pandas as pd
children = list(run.get_children())
metricslist = {}
for child in children:
properties = child.get_properties()
metrics = {k: v for k, v in child.get_metrics().items() if isinstance(v, float)}
metricslist[int(properties['iteration'])] = metrics
rundata = pd.DataFrame(metricslist).sort_index(1)
rundata
# -
# ### Waiting until the run finishes
#
# `wait_for_complettion` method will block till the run finishes.
# +
# Wait until the run finishes.
# run.wait_for_completion(show_output = True)
# -
# ## Explore the results
#
# ### Retrieve the best model
#
# Below we select the best pipeline from our iterations. The get_output method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on get_output allow you to retrieve the best run and fitted model for any logged metric or for a particular iteration.
best_run, fitted_model = run.get_output()
print(best_run)
print(fitted_model)
# #### Best model on any other metric
#
# Show the run and the model which has the smallest log_loss value:
lookup_metric = "log_loss"
specific_run, specific_model = run.get_output(metric = lookup_metric)
print(specific_run)
print(specific_model)
# #### Model from a Specific Iteration
iteration = 3
third_run, third_model = run.get_output(iteration=iteration)
print(third_run)
print(third_model)
# ### Test the model
# Load the test data
#
#
# +
import pandas as pd
import os
folder = '../datasets'
filename = 'banking_test.csv'
pathname = os.path.join(folder, filename)
df = pd.read_csv(pathname, delimiter=',')
feature_columns = [
# Demographic
'age',
'job',
'education',
'marital',
'housing',
'loan',
# Previous campaigns
'month',
'campaign',
'poutcome',
# Economic indicators
'emp_var_rate',
'cons_price_idx',
'cons_conf_idx',
'euribor3m',
'nr_employed']
df_test = df[feature_columns + ['y']]
df_test.head()
# -
# Test the best model
# +
from sklearn.metrics import accuracy_score, recall_score
y_pred = fitted_model.predict(df_test.values)
print("Accuracy: ", accuracy_score(df_test.y, y_pred))
print("Recall: ", recall_score(df_test.y, y_pred))
# -
# ## Register the best performing model for later use and deployment
#
# The best model can now be registered into *Model Registry*.
#
# If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered.
#
# You can annotate the model with arbitrary tags.
#
# notice the use of the root run (not best_run) to register the best model
tags = {"Department": "Marketing"}
model = run.register_model(description='AutoML trained propensity to buy classifier',
tags=tags)
| ML-AML-Walkthrough/04-AutomatedML/04-AutomatedML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1D Elastic Wave Equation with a Discontinuous Galerkin-FEM in the Time Domain with Free Surface Boundary Conditions
# ## Notebook setup
import numpy
from matplotlib import pyplot
# %matplotlib inline
# ## Governing Equation
# ### Displacement Formulation
#
# Consider the elastic wave equation in 1D written in its displacement form:
#
# $$
# \rho\partial_{tt}u = \partial_{x}(\mu\partial_{x}u) + f
# $$
#
# with $x \in \Omega = [x_{min},x_{max}]$ and $t \in [0,t_{max}]$, and where $u(x,t)$ is the displacement field, $\rho(x)$ is the material density, $\mu(x)$ is the material shear modulus, and $f(x,t)$ is a source term. The displacement field is subjected to free surface (or stress free) boundary conditions
#
# \begin{align}
# \left.\mu\partial_{x}u(x,t)\right|_{x=x_{min}} &= 0, \\
# \left.\mu\partial_{x}u(x,t)\right|_{x=x_{max}} &= 0,
# \end{align}
#
# and initial conditions
#
# $$
# \left.\partial_{t}u(x,t)\right|_{t=0} = \left.\partial_{tt}u(x,t)\right|_{t=0} = 0.
# $$
# ### Velocity-Stress Formulation
#
# For a Discontinuous Galerkin (DG) approach, it will be convenient to rewrite the displacement form of the elastic wave equation in a velocity-stress formulation in order to reduce the time and space derivatives to first order. For that, we define two other variables namely velocity $v$ and stress $\sigma$:
#
# \begin{align}
# &\partial_{t}u = v, \\
# &\sigma = \mu\partial_{x}u.
# \end{align}
#
# Upon substitution into the displacement form, these variables will give rise to the velocity-stress formulation of the elastic wave equation
#
# \begin{align}
# \partial_{t}v &= 1/\rho\:\partial_{x}\sigma + f/\rho, \\
# \partial_{t}\sigma &= \mu\partial_{x}v.
# \end{align}
#
# or, in vector notation
#
# $$
# \frac{\partial}{\partial t} \left[
# \begin{array}{c}
# v \\
# \sigma
# \end{array}
# \right] = \left[
# \begin{array}{cc}
# 0 & 1/\rho \\
# \mu & 0
# \end{array}
# \right] \frac{\partial}{\partial x} \left[
# \begin{array}{c}
# v \\
# \sigma
# \end{array}
# \right] + \left[
# \begin{array}{c}
# f/\rho \\
# 0
# \end{array}
# \right]
# $$
# As before, this system of PDEs will be endowed with free surface boundary conditions
#
# \begin{align}
# \left.\partial_{x}v(x,t)\right|_{x=x_{min}} =& \ 0, \ \ \left.\sigma(x,t)\right|_{x=x_{min}} = 0, \\
# \left.\partial_{x}v(x,t)\right|_{x=x_{max}} =& \ 0, \ \ \left.\sigma(x,t)\right|_{x=x_{max}} = 0,
# \end{align}
#
# as well as initial conditions
#
# $$
# \left.v(x,t)\right|_{t=0} = \left.\sigma(x,t)\right|_{t=0} = 0.
# $$
# ## Discontinuous Galerkin-FEM Discretization
#
# We approximate $\Omega$ by $K$ nonoverlapping elements, $x \in [x^{k}_{l},x^{k}_{r}] = \Omega^{k}$. On each of these elements we express the local displacement solution as a polynomial of order N
#
# $$
# x \in \Omega^{k}: u^{k}_{h}(x,t) = \sum^{N+1}_{j=1}\hat{u}^{k}_{j}(t)\psi_{j}(x) = \sum^{N+1}_{i=1}u^{k}_{h}(x^{k}_{i},t)\ell^{k}_{i}(x).
# $$
#
# In the last statement, we have introduced two complementary expressions for the local solution known as the modal form, which uses the local polynomial basis, $\psi_{j}(x)$, and the nodal form, which is defined through $N_{p} = N+1$ local grid points, $x^{k}_{i} \in \Omega^{k}$, and uses the interpolating Lagrange polynomials, $\ell^{k}_{i}(x)$, as basis functions. These two representations are connected by the definition of the expansion coefficients, $\hat{u}^{k}_{j}$.
#
# From now on, we will choose the modal representation for the local solution but we should keep in mind that any of the two are mathematically equivalent (although not computationally!) and can be consistently used to discretize the elastic wave equation.
#
# The global solution $u(x,t)$ is assumed to be approximated by the piecewise N-th order polynomial approximation $u_{h}(x,t)$,
#
# $$
# u(x,t) \approx u_{h}(x,t) = \bigoplus^{K}_{k=1}u^{k}_{h}(x,t),
# $$
#
# defined as the direct sum of the $K$ local polynomial solutions $u^{k}_{h}(x,t)$.
# In exactly the same way, for the velocity-stress formulation of the elastic wave equation, we express the local velocity and local stress solutions as
#
# \begin{align}
# v^{k}_{h}(x,t) &= \sum^{N+1}_{j=1}\hat{v}^{k}_{j}(t)\psi_{j}(x), \\
# \sigma^{k}_{h}(x,t) &= \sum^{N+1}_{j=1}\hat{\sigma}^{k}_{j}(t)\psi_{j}(x).
# \end{align}
#
# Going back to the differential form of the velocity-stress formulation, we proceed as in the Galerkin projection forming the residual
#
# $$
# R_{h}(x,t) = \frac{\partial}{\partial t} \left[
# \begin{array}{c}
# v_{h} \\
# \sigma_{h}
# \end{array}
# \right] - \left[
# \begin{array}{cc}
# 0 & 1/\rho \\
# \mu & 0
# \end{array}
# \right] \frac{\partial}{\partial x} \left[
# \begin{array}{c}
# v_{h} \\
# \sigma_{h}
# \end{array}
# \right] - \left[
# \begin{array}{c}
# f/\rho \\
# 0
# \end{array}
# \right]
# $$
#
# and requiring it to be orthogonal to the space of test functions $V_{h} = \oplus^{K}_{k=1}V^{k}_{h}$ where $V^{k}_{h} = span\{\psi_{j}(\Omega^{k})\}^{N+1}_{j=1}$
#
# $$
# \int_{\Omega^{k}} R_{h}(x,t)\psi_{i}(x) dx = 0, \ 1 \leq i \leq N+1,
# \label{orthog-condition}
# $$
#
# on all $K$ elements. This yields exactly $N+1$ equations for the $N+1$ unknowns on each element.
# Integrating by parts once for the velocity field and twice for the stress field we get the local weak form of the elastic wave equation
#
# \begin{align}
# \int_{\Omega^{k}} \left(\frac{\partial v^{k}_{h}}{\partial t}\psi_{i} + \frac{1}{\rho}\sigma^{k}_{h}\frac{\partial \psi_{i}}{\partial x} - \frac{f^{k}}{\rho}\psi_{i}\right) dx &= \int_{\partial \Omega^{k}} \mathbf{\hat{n}} \cdot \left(\frac{1}{\rho}\sigma^{k}_{h}\psi_{i}\right) dx = \left[\frac{1}{\rho}\sigma^{k}_{h}\psi_{i}\right]^{x^{k}_{r}}_{x^{k}_{l}}, \ 1 \leq i \leq N+1, \\
# \int_{\Omega^{k}} \left(\frac{\partial \sigma^{k}_{h}}{\partial t}\psi_{i} + \mu v^{k}_{h}\frac{\partial \psi_{i}}{\partial x} + \frac{\partial \mu}{\partial x} v^{k}_{h}\psi_{i} \right) dx &= \int_{\partial \Omega^{k}} \mathbf{\hat{n}} \cdot (\mu v^{k}_{h}\psi_{i}) dx = \left[\mu v^{k}_{h}\psi_{i}\right]^{x^{k}_{r}}_{x^{k}_{l}}, \ 1 \leq i \leq N+1.
# \end{align}
#
# At this point, using the standard Discontinuous Galerkin (DG) approach, we replace the physical fluxes represented by the line integrals computed along the interfaces between elements by some numerical fluxes in order to recover the global solution from the $K$ local solutions. This leads to the following semidiscrete scheme
#
# \begin{align}
# \int_{\Omega^{k}} \left(\frac{\partial v^{k}_{h}}{\partial t}\psi_{i} + \frac{1}{\rho}\sigma^{k}_{h}\frac{\partial \psi_{i}}{\partial x} - \frac{f^{k}}{\rho}\psi_{i}\right) dx &= \left[\frac{1}{\rho}(\sigma_{h})^{*}\psi_{i}\right]^{x^{k}_{r}}_{x^{k}_{l}}, \ 1 \leq i \leq N+1,\\
# \int_{\Omega^{k}} \left(\frac{\partial \sigma^{k}_{h}}{\partial t}\psi_{i} + \mu v^{k}_{h}\frac{\partial \psi_{i}}{\partial x} + \frac{\partial \mu}{\partial x} v^{k}_{h}\psi_{i} \right) dx &= \left[\mu (v_{h})^{*}\psi_{i}\right]^{x^{k}_{r}}_{x^{k}_{l}}, \ 1 \leq i \leq N+1.
# \end{align}
#
# Now, we have a total of $K\times(N+1)$ equations for the same number of unkowns. It is worth noting that the numerical fluxes are responsible for recovering the global solution from the local solutions as well as imposing (weakly) the boundary conditions.
# The expressions above are known as the DG weak form of the velocity-stress formulation of the 1D elastic wave equations.
# To get the matrix form of the semidiscrete scheme, we continue by substituting the approximations for the velocity and the stress into the DG weak form
# \begin{align}
# \sum^{N+1}_{j=1} \frac{d}{dt}\hat{v}^{k}_{j} \int_{\Omega^{k}} \psi_{j}\psi_{i} dx &+ \frac{1}{\rho} \sum^{N+1}_{j=1} \hat{\sigma}^{k}_{j} \int_{\Omega^{k}} \psi_{j}\frac{\partial \psi_{i}}{\partial x} dx - \frac{1}{\rho}\int_{\Omega^{k}} f^{k}\psi_{i} dx = \frac{1}{\rho}\left[(\sigma_{h})^{*}\psi_{i}(x^{k}_{r})-(\sigma_{h})^{*}\psi_{i}(x^{k}_{l})\right], \\
# \sum^{N+1}_{j=1} \frac{d}{dt}\hat{\sigma}^{k}_{j} \int_{\Omega^{k}} \psi_{j}\psi_{i} dx &+ \sum^{N+1}_{j=1} \hat{v}^{k}_{j} \int_{\Omega^{k}}\mu \psi_{j}\frac{\partial \psi_{i}}{\partial x} dx + \sum^{N+1}_{j=1} \hat{v}^{k}_{j} \int_{\Omega^{k}} \frac{\partial \mu}{\partial x} \psi_{j}\psi_{i} dx = \mu(x^{k}_{r})(v_{h})^{*}\psi_{i}(x^{k}_{r})-\mu(x^{k}_{l})(v_{h})^{*}\psi_{i}(x^{k}_{l}).
# \end{align}
#
# where $\rho$ is assumed to be constant. These equations can be expressed in matrix form in the following way
#
# \begin{align}
# \hat{\mathcal{M}}^{k}\frac{d}{dt}\hat{\boldsymbol{v}}^{k}_{h} &+ \frac{1}{\rho}\hat{\mathcal{S}}^{k}_{v}\hat{\boldsymbol{\sigma}}^{k}_{h} - \frac{1}{\rho}\hat{\mathbf{f}}^{k} = \frac{1}{\rho}\left[(\sigma_{h})^{*}\boldsymbol{\psi}(x^{k}_{r})-(\sigma_{h})^{*}\boldsymbol{\psi}(x^{k}_{l})\right],\\
# \hat{\mathcal{M}}^{k}\frac{d}{dt}\hat{\boldsymbol{\sigma}}^{k}_{h} &+ \hat{\mathcal{S}}^{k}_{s}\hat{\boldsymbol{v}}^{k}_{h} + \hat{\mathcal{S}}^{k}_{m}\hat{\boldsymbol{v}}^{k}_{h} = \mu(x^{k}_{r})(v_{h})^{*}\boldsymbol{\psi}(x^{k}_{r})-\mu(x^{k}_{l})(v_{h})^{*}\boldsymbol{\psi}(x^{k}_{l}).
# \end{align}
#
# where, for $1 \leq i,j \leq N+1$,
#
# \begin{align*}
# \hat{\mathcal{M}}^{k} &= \int_{\Omega^{k}} \psi_{i}\psi_{j} dx, \\
# \hat{\mathcal{S}}^{k}_{v} &= \int_{\Omega^{k}} \psi_{j}\frac{\partial \psi_{i}}{\partial x} dx, \\
# \hat{\mathcal{S}}^{k}_{s} &= \int_{\Omega^{k}}\mu \psi_{j}\frac{\partial \psi_{i}}{\partial x} dx, \\
# \hat{\mathcal{S}}^{k}_{m} &=\int_{\Omega^{k}} \frac{\partial \mu}{\partial x} \psi_{j}\psi_{i} dx, \\
# \hat{\mathbf{f}}^{k} &= \int_{\Omega^{k}} f^{k}\psi_{i} dx.
# \end{align*}
#
# are the local mass, velocity and stress stiffness matrices and source vector, respectively. Furthermore, we have
#
# \begin{align*}
# \hat{\boldsymbol{v}}^{k}_{h} &= [\hat{v}^{k}_{1},\ldots,\hat{v}^{k}_{N+1}]^{T}, \\
# \hat{\boldsymbol{\sigma}}^{k}_{h} &= [\hat{\sigma}^{k}_{1},\ldots,\hat{\sigma}^{k}_{N+1}]^{T}, \\
# \boldsymbol{\psi} &= [\psi_{1}(x),\ldots,\psi_{N+1}(x)]^{T}.
# \end{align*}
#
# as the vectors of local velocity and stress solutions and the local test functions, respectively.
# In order to perform the integrals over each element numerically, we transform coordinates by introducing the affine mapping
#
# $$
# x \in \Omega^{k}: x(\xi) = x^{k}_{l} + \frac{1+\xi}{2}h^{k}, h^{k} = x^{k}_{r} - x^{k}_{l},
# $$
#
# with the reference coordinate $\xi \in \hat{\Omega} = \left[-1,1\right]$. Now, the local polynomial representations can be rewritten as
#
# $$
# x \in \Omega^{k}: u^{k}_{h}(x(\xi),t) = \sum^{N+1}_{j=1}\hat{u}^{k}_{j}(t)\hat{\psi}_{j}(\xi) = \sum^{N+1}_{i=1}u^{k}_{h}(x^{k}_{i},t)l^{k}_{i}(\xi).
# $$
#
# and in exactly the same way for the approximations $v^{k}_{h}$ and $\sigma^{k}_{h}$. We should note that the Jacobian of the previous mapping is given by
# $$
# J = \left|\frac{dx}{d\xi}\right| = \frac{h^{k}}{2}
# $$
#
# and the derivatives of the local test functions are computed using the chain rule
#
# $$
# \frac{d\psi(x(\xi))}{dx} = \frac{d\psi}{d\xi}\frac{d\xi}{dx} = \frac{2}{h^{k}}\frac{d\hat{\psi}}{d\xi}.
# $$
# After the development of a suitable local representation of the solution using the reference element $\hat{\Omega}$, we return to the discrete equations and rewrite them as follows
#
# \begin{align}
# \sum^{N+1}_{j=1} \frac{d}{dt}\hat{v}^{k}_{j} \ \frac{h^{k}}{2}\int_{\hat{\Omega}} \hat{\psi}_{j}\hat{\psi}_{i} d\xi &+ \frac{1}{\rho} \sum^{N+1}_{j=1} \hat{\sigma}^{k}_{j} \int_{\hat{\Omega}} \hat{\psi}_{j}\frac{\partial \hat{\psi}_{i}}{\partial \xi} d\xi - \frac{1}{\rho}\frac{h^{k}}{2}\int_{\hat{\Omega}} \hat{f}\hat{\psi}_{i} d\xi = \frac{1}{\rho}\left[(\sigma_{h})^{*}\hat{\psi}_{i}(1)-(\sigma_{h})^{*}\hat{\psi}_{i}(-1)\right], \\
# \sum^{N+1}_{j=1} \frac{d}{dt}\hat{\sigma}^{k}_{j} \ \frac{h^{k}}{2}\int_{\hat{\Omega}} \hat{\psi}_{j}\hat{\psi}_{i} d\xi &+ \sum^{N+1}_{j=1} \hat{v}^{k}_{j} \int_{\hat{\Omega}}\mu \hat{\psi}_{j}\frac{\partial \hat{\psi}_{i}}{\partial \xi} d\xi + \sum^{N+1}_{j=1} \hat{v}^{k}_{j} \ \frac{h^{k}}{2}\int_{\hat{\Omega}} \frac{\partial \mu}{\partial x} \hat{\psi}_{j}\hat{\psi}_{i} d\xi = \mu(x^{k}_{r})(v_{h})^{*}\hat{\psi}_{i}(1)-\mu(x^{k}_{l})(v_{h})^{*}\hat{\psi}_{i}(-1).
# \end{align}
#
# or in matrix form
#
# \begin{align}
# \frac{h^{k}}{2}\mathcal{M}\frac{d}{dt}\hat{\boldsymbol{v}}^{k}_{h} &+ \frac{1}{\rho}\mathcal{S}_{v}\hat{\boldsymbol{\sigma}}^{k}_{h} - \frac{1}{\rho}\frac{h^{k}}{2}\mathbf{f} = \frac{1}{\rho}\left[(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(1)-(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(-1)\right], \\
# \frac{h^{k}}{2}\mathcal{M}\frac{d}{dt}\hat{\boldsymbol{\sigma}}^{k}_{h} &+ \mathcal{S}_{s}\hat{\boldsymbol{v}}^{k}_{h} + \frac{h^{k}}{2}\mathcal{S}_{m}\hat{\boldsymbol{v}}^{k}_{h} = \mu(x^{k}_{r})(v_{h})^{*}\boldsymbol{\hat{\psi}}(1)-\mu(x^{k}_{l})(v_{h})^{*}\boldsymbol{\hat{\psi}}(-1).
# \end{align}
#
# where, for $1 \leq i,j \leq N+1$,
#
# \begin{align*}
# \mathcal{M} &= \int_{\hat{\Omega}} \hat{\psi}_{i}\hat{\psi}_{j} d\xi, \\
# \mathcal{S}_{v} &= \int_{\hat{\Omega}} \hat{\psi}_{j}\frac{\partial \hat{\psi}_{i}}{\partial \xi} d\xi, \\
# \mathcal{S}_{s} &= \int_{\hat{\Omega}}\mu \hat{\psi}_{j}\frac{\partial \hat{\psi}_{i}}{\partial \xi} d\xi, \\
# \mathcal{S}_{m} &=\int_{\hat{\Omega}} \frac{\partial \mu}{\partial x} \hat{\psi}_{j}\hat{\psi}_{i} d\xi, \\
# \mathbf{f} &= \int_{\hat{\Omega}} \hat{f}\hat{\psi}_{i} d\xi.
# \end{align*}
#
# are the elemental mass, velocity and stress stiffness matrices and source vector defined over the reference element, respectively. The vectors $\hat{\boldsymbol{v}}^{k}_{h}$ and $\hat{\boldsymbol{\sigma}}^{k}_{h}$ are defined exactly as before. The vector of elemental test functions is defined as
#
# $$
# \boldsymbol{\hat{\psi}} = [\hat{\psi}_{1}(\xi),\ldots,\hat{\psi}_{N+1}(\xi)]^{T}.
# $$
#
# As usual, integration on the reference element $\hat{\Omega} = [-1,1]$ is performed using Gaussian quadrature
#
# $$
# \int^{1}_{-1} f(\xi) d\xi = \sum^{N_{i}}_{l=1} w_{l} f(\xi_{l}),
# $$
#
# where $w_{l}$ and $\xi_{l}$ are the quadrature weights and points, respectively, and $N_{i}$ is total number of integration points needed to exactly integrate a polynomial of order at most $2N$ which is the polynomial order associated to the mass matrix coefficients. In the present DG approach, we considered two options for the distribution of quadrature points: standard Gauss-Legendre (GL) and Gauss-Legendre-Lobatto (GLL) points. The first option considers only internal points on the reference element while the second includes its boundaries $-1$ and $1$.
# Now we are in a position to choose the numerical fluxes $(v_{h})^{*}$ and $(\sigma_{h})^{*}$. The role of these fluxes is to guarantee stability of the formulation by mimicking the flow of information in the underlying partial differential equation. A simple choice that preserves stability in the case of the elastic wave equation is the Lax-Friedrichs flux
#
# \begin{align}
# (v_{h})^{*} &= \{\{v_{h}\}\} + \frac{Z}{2}[[\sigma_{h}]], \ \ \text{where} \ \ Z = -(\rho \: \mu)^{-1/2}, \\
# (\sigma_{h})^{*} &= \{\{\sigma_{h}\}\} + \frac{Y}{2}[[v_{h}]], \ \ \text{where} \ \ Y = -(\rho \: \mu)^{1/2}.
# \end{align}
#
# In the expressions above we have used the average and jump operators which are usually defined as follows
#
# $$
# \{\{u\}\} = \frac{u^{-}+u^{+}}{2}
# $$
#
# where $u$ can be both a scalar or a vector quantity and
#
# \begin{align}
# [[u]] &= \hat{\mathbf{n}}^{-}u^{-} + \hat{\mathbf{n}}^{+}u^{+}, \\
# [[\mathbf{u}]] &= \hat{\mathbf{n}}^{-}\cdot\mathbf{u}^{-} + \hat{\mathbf{n}}^{+}\cdot\mathbf{u}^{+}
# \end{align}
#
# where $\hat{\mathbf{n}}$ is defined as the normal vector along the left or right boundaries of the $k$ element. We observe that the jump operator is defined differently depending on whether $u$ is a scalar or a vector, $\mathbf{u}$. The superscripts "$-$" and "$+$" stand for the interior and exterior information of the element, respectively.
#
# It is worth noting that
# $$
# -\mu Z = -\frac{Y}{\rho} = c
# $$
#
# where $c$ is the propagation wave speed.
# Upon substitution of the expressions of the Lax-Friedrichs flux at the right hand side of the matrix equations, we come up with the final matrix form of the proposed DG scheme. Let us consider the numerical flux, $(\sigma_{h})^{*}$, first
#
# \begin{align*}
# \frac{1}{\rho}\left[(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(1)-(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(-1)\right] =& \frac{1}{\rho} \left[\left(\{\{\sigma^{k}_{h}\}\} + \frac{Y}{2}[[v^{k}_{h}]]\right)\boldsymbol{\hat{\psi}}(1) - \left(\{\{\sigma^{k}_{h}\}\} + \frac{Y}{2}[[v^{k}_{h}]]\right)\boldsymbol{\hat{\psi}}(-1)\right] \\
# =& \frac{1}{\rho}\left[\left(\frac{1}{2}(\sigma^{-}_{h}+\sigma^{+}_{h})+\frac{Y}{2}(\hat{\mathbf{n}}^{-}_{r}v^{-}_{h} + \hat{\mathbf{n}}^{+}_{r}v^{+}_{h})\right)\boldsymbol{\hat{\psi}}(1)\right. \\
# &-\left.\left(\frac{1}{2}(\sigma^{-}_{h}+\sigma^{+}_{h})+\frac{Y}{2}(\hat{\mathbf{n}}^{-}_{l}v^{-}_{h} + \hat{\mathbf{n}}^{+}_{l}v^{+}_{h})\right)\boldsymbol{\hat{\psi}}(-1)\right]
# \end{align*}
#
# where the subscripts $r$ or $l$ for the normal vector $\hat{\mathbf{n}}$ indicates that we are taking the normal vector at the right or left side of the $k$ element, respectively.
# Substituting the expressions for the approximations $\sigma_{h}$ and $v_{h}$ into the equation above and remembering that $\hat{\mathbf{n}}^{-} = -\hat{\mathbf{n}}^{+}$ yields
#
# \begin{align}
# \frac{1}{\rho}\left[(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(1)-(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(-1)\right] =& \: \frac{1}{\rho}\left[\frac{1}{2}\left((\hat{\boldsymbol{\sigma}}^{k}_{h})^{T}\boldsymbol{\hat{\psi}}(1)+(\hat{\boldsymbol{\sigma}}^{k+1}_{h})^{T}\boldsymbol{\hat{\psi}}(-1)\right)\boldsymbol{\hat{\psi}}(1)\right. \\
# &+\frac{Y}{2}\hat{\mathbf{n}}^{-}_{r}\left((\hat{\boldsymbol{v}}^{k}_{h})^{T}\boldsymbol{\hat{\psi}}(1)-(\hat{\boldsymbol{v}}^{k+1}_{h})^{T}\boldsymbol{\hat{\psi}}(-1)\right)\boldsymbol{\hat{\psi}}(1) \nonumber \\
# &-\frac{1}{2}\left((\hat{\boldsymbol{\sigma}}^{k}_{h})^{T}\boldsymbol{\hat{\psi}}(-1)+(\hat{\boldsymbol{\sigma}}^{k-1}_{h})^{T}\boldsymbol{\hat{\psi}}(1)\right)\boldsymbol{\hat{\psi}}(-1) \nonumber \\
# &-\left.\frac{Y}{2}\hat{\mathbf{n}}^{-}_{l}\left((\hat{\boldsymbol{v}}^{k}_{h})^{T}\boldsymbol{\hat{\psi}}(-1)-(\hat{\boldsymbol{v}}^{k-1}_{h})^{T}\boldsymbol{\hat{\psi}}(1)\right)\boldsymbol{\hat{\psi}}(-1)\right] \\
# \\
# =& \: \frac{1}{\rho}\left[\frac{1}{2}\left([\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(1)]\:\hat{\boldsymbol{\sigma}}^{k}_{h}+[\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(-1)]\:\hat{\boldsymbol{\sigma}}^{k+1}_{h}\right)\right. \\
# &+\frac{Y}{2}\hat{\mathbf{n}}^{-}_{r}\left([\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(1)]\:\hat{\boldsymbol{v}}^{k}_{h}-[\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(-1)]\:\hat{\boldsymbol{v}}^{k+1}_{h}\right) \\
# &-\frac{1}{2}\left([\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(-1)]\:\hat{\boldsymbol{\sigma}}^{k}_{h}+[\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(1)]\:\hat{\boldsymbol{\sigma}}^{k-1}_{h}\right) \nonumber \\
# &-\left.\frac{Y}{2}\hat{\mathbf{n}}^{-}_{l}\left([\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(-1)]\:\hat{\boldsymbol{v}}^{k}_{h}-[\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(1)]\:\hat{\boldsymbol{v}}^{k-1}_{h}\right)\right]
# \end{align}
#
# Tensor multiplication of vectors of elemental test functions gives rise to matrices which we will call flux matrices. They are defined as follows
#
# \begin{align*}
# \mathcal{F}^{k}_{r} &= [\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(1)], \\
# \mathcal{F}^{k+1}_{l} &= [\boldsymbol{\hat{\psi}}(1)\otimes\boldsymbol{\hat{\psi}}(-1)], \\
# \mathcal{F}^{k}_{l} &= [\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(-1)], \\
# \mathcal{F}^{k-1}_{r} &= [\boldsymbol{\hat{\psi}}(-1)\otimes\boldsymbol{\hat{\psi}}(1)].
# \end{align*}
# We now use the flux matrices definitions and rearrange terms to write the final matricial form for the velocity and stress equations
#
# \begin{align}
# \frac{1}{\rho}\left[(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(1)-(\sigma_{h})^{*}\boldsymbol{\hat{\psi}}(-1)\right] = \:
# \frac{1}{\rho}\left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{\sigma}}^{k+1}_{h}-(\hat{\mathbf{n}}^{-}_{r}\cdot Y)\hat{\boldsymbol{v}}^{k+1}_{h}\right) \right. &+ \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}+(\hat{\mathbf{n}}^{-}_{r}\cdot Y)\hat{\boldsymbol{v}}^{k}_{h}\right) \nonumber \\
# -\frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{\sigma}}^{k-1}_{h}-(\hat{\mathbf{n}}^{-}_{l}\cdot Y)\hat{\boldsymbol{v}}^{k-1}_{h}\right) &- \left.\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}+(\hat{\mathbf{n}}^{-}_{l}\cdot Y)\hat{\boldsymbol{v}}^{k}_{h}\right)\right].
# \end{align}
# Repeating exactly the same steps for the numerical flux, $(v_{h})^{*}$, yields
#
# \begin{align}
# \mu(x^{k}_{r})(v_{h})^{*}\boldsymbol{\hat{\psi}}(1)-\mu(x^{k}_{l})(v_{h})^{*}\boldsymbol{\hat{\psi}}(-1) = \:
# &\mu(x^{k}_{r})\left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{v}}^{k+1}_{h}-(\hat{\mathbf{n}}^{-}_{r}\cdot Z)\hat{\boldsymbol{\sigma}}^{k+1}_{h}\right) + \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{v}}^{k}_{h}+(\hat{\mathbf{n}}^{-}_{r}\cdot Z)\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right] \\
# -&\mu(x^{k}_{l}) \left[\frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{v}}^{k-1}_{h}-(\hat{\mathbf{n}}^{-}_{l}\cdot Z)\hat{\boldsymbol{\sigma}}^{k-1}_{h}\right)+\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{v}}^{k}_{h}+(\hat{\mathbf{n}}^{-}_{l}\cdot Z)\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right].
# \end{align}
# Noting that the exterior normal vector, $\hat{\mathbf{n}}^{-}$, assumes only two values, $\hat{\mathbf{n}}^{-}_{r} = 1$ and $\hat{\mathbf{n}}^{-}_{l} = -1$, at the right and the left boundaries, respectively, of any $k$ element and the variables, $Z$, and, $Y$, can be computed from the expressions for the fluxes, the matrix form of the proposed DG scheme for the velocity-stress formulation of the elastic wave equation can finally be written as
#
# \begin{align}
# \frac{h^{k}}{2}\mathcal{M}\frac{d}{dt}\hat{\boldsymbol{v}}^{k}_{h} + \frac{1}{\rho}\mathcal{S}_{v}\hat{\boldsymbol{\sigma}}^{k}_{h} - \frac{1}{\rho}\frac{h^{k}}{2}\mathbf{f} =& \:
# \frac{1}{\rho}\left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{\sigma}}^{k+1}_{h}+(\rho \: \mu(x^{k}_{r}))^{1/2}\hat{\boldsymbol{v}}^{k+1}_{h}\right) + \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}-(\rho \: \mu(x^{k}_{r}))^{1/2}\hat{\boldsymbol{v}}^{k}_{h}\right)\right. \\
# &- \ \frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{\sigma}}^{k-1}_{h}-(\rho \: \mu(x^{k}_{l}))^{1/2}\hat{\boldsymbol{v}}^{k-1}_{h}\right) - \left.\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}+(\rho \: \mu(x^{k}_{l}))^{1/2}\hat{\boldsymbol{v}}^{k}_{h}\right)\right], \\
# \frac{h^{k}}{2}\mathcal{M}\frac{d}{dt}\hat{\boldsymbol{\sigma}}^{k}_{h} + \mathcal{S}_{s}\hat{\boldsymbol{v}}^{k}_{h} + \frac{h^{k}}{2}\mathcal{S}_{m}\hat{\boldsymbol{v}}^{k}_{h} =& \:
# \mu(x^{k}_{r})\left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{v}}^{k+1}_{h}+(\rho \: \mu(x^{k}_{r}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k+1}_{h}\right) + \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{v}}^{k}_{h}-(\rho \: \mu(x^{k}_{r}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right] \\
# -& \: \mu(x^{k}_{l}) \left[\frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{v}}^{k-1}_{h}-(\rho \: \mu(x^{k}_{l}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k-1}_{h}\right)+\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{v}}^{k}_{h}+(\rho \: \mu(x^{k}_{l}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right].
# \end{align}
# It is important to observe some important properties of the DG formulation:
#
# - The scheme is completely local (there is no global assembly).
# - Boundary conditions and interface continuity are enforced weakly via numerical fluxes.
# - The solutions are piecewise smooth, polynomial in general, but discontinuous between elements.
# - The scheme is well suited for order and element sizes adaption.
# - Parallelism is very natural because of the strictly local formulation.
# The source term $f$ in the original displacement form of the elastic wave equation is defined as a point source
#
# $$
# f(x,t) = \theta(t)\delta(x-x_{\text{source}})
# $$
#
# where $x_{\text{source}}$ is chosen to be any quadrature point inside any $k$ element within the domain $\Omega$. We do not allow $x_{\text{source}}$ to be chosen at the element interfaces, which is a possible situation when using GLL quadrature points, in order to avoid the definition of a double valued Dirac delta function. To consider this possibility we should include some upwind feature in the definition of the Dirac delta function but we postponed this tricky theoretical approach to the future. Having defined the source term $f^{k}$ on the $k$-th element, we can write the final form of the vector source term $\hat{\mathbf{f}}^{k}$ as
#
# \begin{align}
# \hat{\mathbf{f}}^{k} &= \int_{\hat{\Omega}}\theta(t)\delta(x-x_{\text{source}})\psi_{i}(x) \: dx, \ i = 1,\ldots,N+1, \\
# &= \theta(t)\psi_{i}(x_{\text{source}}), \ i = 1,\ldots,N+1.
# \end{align}
#
# We observe that the vector form of the source term as defined above will have non zero entries on one single element, $k$. It is also worth noting that when using the Lagrange basis functions, the resulting source vector on the $k$-th element, will have just a single non zero entry $\mathbf{f}^{k}_{i}$ corresponding to the coefficient associated to the $i^{th}$-Lagrange polynomial $\ell^{k}_{i}(x_{\text{source}})$ defined at the collocation point $x_{\text{source}}$ which must be one of the GLL nodes. In this case the source term simplifies to $\mathbf{f}^{k}_{i} = \theta(t)$ because of the cardinality property of the Lagrange polynomial.
# ## Time Marching
#
# The DG-FEM spatial discretization of the elastic wave equation produces a system of first order ODEs in terms of the velocity and stress fields of the problem.
#
# Before discretizing the semidiscrete problem given, we define the right hand side operator $\mathcal{L}_{h}$ for the variables $\boldsymbol{v}_{h}$ and $\boldsymbol{\sigma}_{h}$ as
#
# \begin{align}
# \mathcal{L}_{h}(\boldsymbol{v}_{h},t) =& \mathcal{M}^{-1} \! \left\{\frac{1}{\rho}\mathbf{f} - \frac{2}{h^{k}}\frac{1}{\rho}\mathcal{S}_{v}\hat{\boldsymbol{\sigma}}^{k}_{h} + \frac{2}{h^{k}}\frac{1}{\rho}\left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{\sigma}}^{k+1}_{h}+(\rho\mu(x^{k}_{r}))^{1/2}\hat{\boldsymbol{v}}^{k+1}_{h}\right) + \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}-(\rho\mu(x^{k}_{r}))^{1/2}\hat{\boldsymbol{v}}^{k}_{h}\right)\right.\right. \\
# -& \left.\ \frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{\sigma}}^{k-1}_{h}-(\rho\mu(x^{k}_{l}))^{1/2}\hat{\boldsymbol{v}}^{k-1}_{h}\right) - \left.\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{\sigma}}^{k}_{h}+(\rho\mu(x^{k}_{l}))^{1/2}\hat{\boldsymbol{v}}^{k}_{h}\right)\right]\right\}, \\
# \mathcal{L}_{h}(\boldsymbol{\sigma}_{h},t) \! =& \mathcal{M}^{-1}\!\left\{ \! -\frac{2}{h^{k}}\mathcal{S}_{s}\hat{\boldsymbol{v}}^{k}_{h} - \mathcal{S}_{m}\hat{\boldsymbol{v}}^{k}_{h} + \frac{2}{h^{k}}\mu(x^{k}_{r}) \! \left[\frac{1}{2}\mathcal{F}^{k+1}_{l}\left(\hat{\boldsymbol{v}}^{k+1}_{h}+(\rho\mu(x^{k}_{r}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k+1}_{h}\right) + \frac{1}{2}\mathcal{F}^{k}_{r}\left(\hat{\boldsymbol{v}}^{k}_{h}-(\rho\mu(x^{k}_{r}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right]\right. \\
# -& \left.\: \frac{2}{h^{k}}\mu(x^{k}_{l}) \left[\frac{1}{2}\mathcal{F}^{k-1}_{r}\left(\hat{\boldsymbol{v}}^{k-1}_{h}-(\rho\mu(x^{k}_{l}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k-1}_{h}\right)+\frac{1}{2}\mathcal{F}^{k}_{l}\left(\hat{\boldsymbol{v}}^{k}_{h}+(\rho\mu(x^{k}_{l}))^{-1/2}\hat{\boldsymbol{\sigma}}^{k}_{h}\right)\right]\right\}.
# \end{align}
# As we are also interested in the displacement field of the problem, we can add this field to the system and write it in a new vector variable, $\boldsymbol{\phi}$,
#
# $$
# \boldsymbol{\phi}_{h} = \left\{
# \begin{array}{c}
# \boldsymbol{u}_{h} \\
# \boldsymbol{v}_{h} \\
# \boldsymbol{\sigma}_{h}
# \end{array}
# \right\}.
# $$
#
# Now, the semidiscrete problem can be rewritten in the form
#
# \begin{align}
# \frac{d\boldsymbol{\phi}_{h}}{dt} = \mathcal{L}_{h}(\boldsymbol{\phi}_{h},t),
# \end{align}
#
# or, in vector notation
#
# $$
# \frac{d\boldsymbol{\phi}_{h}}{dt} = \frac{d}{dt}\left\{
# \begin{array}{c}
# \boldsymbol{u}_{h} \\
# \boldsymbol{v}_{h} \\
# \boldsymbol{\sigma}_{h}
# \end{array}
# \right\} = \left\{
# \begin{array}{c}
# \boldsymbol{v}_{h} \\
# \mathcal{L}_{h}(\boldsymbol{v}_{h},t) \\
# \mathcal{L}_{h}(\boldsymbol{\sigma}_{h},t)
# \end{array}
# \right\}.
# $$
#
# In the above expression, we are assuming that the right hand side operator $\mathcal{L}_{h}$ evaluated at the displacement variable $\mathbf{u}_{h}$ simply returns the variable $\boldsymbol{v}_{h}$, i.e., $\mathcal{L}_{h}(\boldsymbol{u}_{h},t) = \boldsymbol{v}_{h}$.
# To march this system of equations in time, we can use the classical and reliable forth order four stage explicit Runge-Kutta method (ERK)
#
# \begin{align}
# &\boldsymbol{k}^{(1)} = \mathcal{L}_{h}(\boldsymbol{\phi}^{n}_{h},t^{n}), \\
# &\boldsymbol{k}^{(2)} = \mathcal{L}_{h}\left(\boldsymbol{\phi}^{n}_{h}+\frac{1}{2}\mathit{\Delta} t\boldsymbol{k}^{(1)},t^{n}+\frac{1}{2}\mathit{\Delta} t\right), \\
# &\boldsymbol{k}^{(3)} = \mathcal{L}_{h}\left(\boldsymbol{\phi}^{n}_{h}+\frac{1}{2}\mathit{\Delta} t\boldsymbol{k}^{(2)},t^{n}+\frac{1}{2}\mathit{\Delta} t\right), \\
# &\boldsymbol{k}^{(4)} = \mathcal{L}_{h}\left(\boldsymbol{\phi}^{n}_{h}+\mathit{\Delta} t\boldsymbol{k}^{(3)},t^{n}+\mathit{\Delta} t\right), \\
# &\boldsymbol{\phi}^{n+1}_{h} = \boldsymbol{\phi}^{n}_{h}+\frac{1}{6}\mathit{\Delta} t\left(\boldsymbol{k}^{(1)}+2\boldsymbol{k}^{(2)}+2\boldsymbol{k}^{(3)}+\boldsymbol{k}^{(4)}\right),
# \end{align}
#
# to advance from $\boldsymbol{\phi}^{n}_{h}$ to $\boldsymbol{\phi}^{n+1}_{h}$ which are separated by the timestep, $\mathit{\Delta} t$. It is important to note that the ERK method is conditionally stable and requires the timestep size $\mathit{\Delta} t$ be estimated using the $CFL$ (Courant-Friedrichs-Lewy) condition. We define the CFL as
#
# $$
# CFL := \max_{k}\frac{c^{k}\mathit{\Delta} t}{\mathit{\Delta} x^{k}},
# $$
#
# where $c^{k}$ is the local propagation wave speed on element $k$ and $\mathit{\Delta} x^{k} = h^{k}$ is the local spatial grid resolution. The specific value of the $CFL$ (constant) number depends on the method used for the space and time discretizations but, in general, its order of magnitude is about 1. For the present case, we consider $CFL = 0.4$. As a last remark, we observe that, for high order methods such as the DG scheme described in this text, we have to consider the minimum grid space as the minimum distance between the mapped quadrature points (GL or GLL) on the mesh elements.
# ## Python implementation
# ### Simulation parameters
#
# The first step to use the Python DG framework is the definition of the simulation parameters. There is a Python class specifically designed for this, called _SimulationData_, implemented on `simulation_data.py`. Also on the same file are two service classes, _QuadratureNodes_ and _BasisType_ used to represent minemonically the types of quadrature points and basis available in the framework.
#
# #### QuadratureNodes
# There are two variables defined in _QuadratureNodes_, both related to Gaussian integration:
#
# - `QuadratureNodes.GL`, corresponding to the Legendre integration points, and
# - `QuadratureNodes.GLL`, associated with the Legendre-Lobatto points.
#
# #### BasisType
# Here we have three variables:
#
# - `BasisType.NODAL`, used to indicate we are using a nodal basis;
# - `BasisType.MODAL`, the same as above, for modal basis, and
# > Remember to use `%load simulation_data.py` to bring the file to the notebook! Remember also to __run__ the cell!
# +
# # %load simulation_data.py
#
# Common simulation data
# ======================
# It presents a struct to store simulation data common to all problemas, like
# number of elements, polynomial order and so on.
#
# by <NAME> Jr. (Matlab and Python versions)
# <NAME> (Python version)
#
from math import ceil, floor
#
# Used to specify the chosen basis.
#
class BasisType(object):
"""The available basis types."""
NODAL = 0
MODAL = 1
MODAL_ORTHOGONAL = 2
MODAL_ORTHONORMAL = 3
#
# Used to specify the chosen type of points distribution for Gaussian quadrature
#
class QuadratureNodes(object):
"""The available quadrature nodes distribution."""
GL = 0 # Gauss-Legendre
GLL = 1 # Gauss-Legendre-Lobatto
class SimulationData:
"""Creates a simulation data structure."""
def __init__(self,
n_elements = 1,
poly_order = 0,
spatial_domain = (0.0, 1.0),
temporal_domain = (0.0, 1.0),
basis_type = BasisType.NODAL,
node_dist = QuadratureNodes.GL,
non_linear = False):
"""
Parameters
----------
n_elements
Number of elements
poly_order
Polynomial order of approximation
spatial_domain
Spatial domain
temporal_domain
Temporal domain
basis_type
The basis used in the approximation
node_dist
The quadrature points distribution
"""
self.n_elements = n_elements
self.poly_order = poly_order
self.spatial_domain = spatial_domain
self.temporal_domain = temporal_domain
self.basis_type = basis_type
self.node_dist = node_dist
self.non_linear = non_linear
def mass_order(self):
"""Number of integration points needed for mass matrix assembly."""
return self.poly_order*2
def nonlinear_order(self):
"""Number of integration points needed for nonlinear terms."""
return self.poly_order*3
def spatial_domain_length(self):
"""Length of the spatial domain."""
return self.spatial_domain[1] - self.spatial_domain[0]
def n_local_dof(self):
"""Local degrees of freedom.
In 1D we have the polynomial order plus 1, to account for the constant
term."""
return self.poly_order+1
def n_global_dof(self):
"""Global degrees of freedom."""
return self.n_local_dof()*self.n_elements
def nip(self):
order = self.mass_order() if self.non_linear == False else self.nonlinear_order()
if self.node_dist == QuadratureNodes.GL:
return int(ceil(0.5*(order+2)))
elif self.node_dist == QuadratureNodes.GLL:
# floor(nip) will keep the nodal spectral approach Np = N+1
return int(floor((order+3)/2))
else:
raise AssertionError( "Wrong quadrature node distribution!\n" )
def t_min(self):
return self.temporal_domain[0]
def t_max(self):
return self.temporal_domain[1]
# Boilerplate code
def __str__(self):
return ("SimulationData:\n"
" Number of elements: %s\n"
" Polynomial order : %s\n"
" Spatial domain : %s\n"
" Temporal domain : %s") % (self.n_elements,
self.poly_order,
self.spatial_domain,
self.temporal_domain)
#-- simulation_data.py ---------------------------------------------------------
# -
# - Elements: 10
# - Polynomial order: 8
# - Spatial domain: [0, 2]
# - Temporal domain: [0, 10]
# - Gaussian integration with Legendre-Lobatto points
# - Modal basis (orthonormal)
sim_data = SimulationData(n_elements=20, poly_order=8, spatial_domain=[0.0, 9000.0],
temporal_domain=[0, 10], node_dist=QuadratureNodes.GL,
basis_type=BasisType.MODAL)
# ### The finite element mesh
#
# Given that we are on a 1D setting, a very simple mesh generator is easy to implement. Of course, on 2D or 3D we would resort to a third-party application. Note that the mesh is created based on the simulation data previously defined. The mesh is also responsible for evaluating the Jacobian.
#
# The mesh class is _Mesh1D_, defined in `mesh_gen_1d.py`, consists -- at this point -- of two data members:
#
# - `coord`, $x$ coordinate of every node on the mesh, and
# - `conn`, connectivity matrix, associating elements and nodes
# +
# # %load mesh_gen_1d.py
#
# 1D mesh generation
# ==================
# Generate simple equidistant grid with a given number of elements elements
#
# by <NAME>. (Matlab and Python versions)
# <NAME> (Python version)
#
from simulation_data import SimulationData
import numpy as np
class Mesh1D:
"""Simple one-dimensional mesh.
Arguments:
sim_data - SimulationData object
"""
def __init__(self, sim_data):
# Generate node coordinates
# - Note that we have n_elements+1 points on the domain.
# - The name 'coord' is classic in FEM codes.
self.coord = numpy.linspace( sim_data.spatial_domain[0],
sim_data.spatial_domain[1],
sim_data.n_elements+1 )
# Element to node connectivity
# - The name 'conn' is classic in FEM codes.
self.conn = numpy.array([numpy.arange(0, sim_data.n_elements, 1),
numpy.arange(1, sim_data.n_elements+1, 1)]).T
def jacobian(self, sim_data, quad_data):
nip = sim_data.nip()
# The Jacobian
self.J = 0.5*(self.coord[1:]-self.coord[0:-1])
# Elements' mid-points
B = 0.5*(self.coord[1:]+self.coord[0:-1])
# We repeat and reshape the array. Why?
B = numpy.repeat(B, nip).reshape(sim_data.n_elements, nip).T
# The transformed points. The construct 'numpy.newaxis' is needed here
# because we want a de facto matricial product (rank-1 update).
#
# x = xi^T * J
self.x = quad_data.xi[numpy.newaxis].T.dot(self.J[numpy.newaxis]) + B
#-- mesh_gen_1d.py -------------------------------------------------------------
# -
# Generate simple mesh
mesh = Mesh1D(sim_data)
# Lets plot the mesh
pyplot.figure(1)
pyplot.clf()
pyplot.xlim(sim_data.spatial_domain[0]-0.1, sim_data.spatial_domain[1]+0.1)
pyplot.plot(mesh.coord, numpy.zeros(mesh.coord.shape), "ro")
pyplot.title("FEM mesh")
pyplot.show()
# ### Gaussian quadrature points and weights
#
# Having defined the mesh, now we proceed with the creation of the Gaussian quadrature points and integration weights. For this purpose we can resort to the _JacobiGaussQuad_ class, defined in `jacobi_gauss_quad.py`. Objects of this class have two data members:
#
# - `xi`, the integration points, and
# - `w`, the integration weights
# +
# # %load jacobi_gauss_quad.py
#
# Points and weights for Gaussian quadrature
#
# by <NAME>. (Matlab and Python versions)
# <NAME> (Python version)
#
import numpy
from math import gamma
from jacobi_p import jacobi_p
from djacobi_p import djacobi_p
from jacobi_roots import jacobi_roots
from simulation_data import QuadratureNodes
class JacobiGaussQuad:
"""Points and weights for Gaussian quadrature based on Jacobi polynomials"""
def __init__(self, sim_data):
# Sets parameters to obtain quadrature points from Legendre polynomials
# Legendre == Jacobi(0,0)
alpha = 0.0
beta = 0.0
nip = sim_data.nip()
# Case 1: Gauss-Legendre quadrature
if sim_data.node_dist == QuadratureNodes.GL:
self.xi, self.w = self._gl(nip, alpha, beta)
# Case 2: Gauss-Legendre-Lobato quadrature
elif sim_data.node_dist == QuadratureNodes.GLL:
self.xi, self.w = self._gll(nip, alpha, beta)
else:
raise AssertionError("Unknown quadrature type!")
# Case 1: Gauss-Legendre quadrature
def _gl(self, nip, alpha, beta):
xi = jacobi_roots(nip, alpha, beta)
C1 = (2.0**(alpha+beta+1.0))*gamma(alpha+nip+1.0)*gamma(beta+nip+1.0)
C2 = gamma(nip+1.0)*gamma(alpha+beta+nip+1.0)*(1.0-xi**2 )
DPm = djacobi_p(xi, nip, alpha, beta)
w = C1*DPm**(-2)/C2
return xi, w
# Case 2: Gauss-Legendre-Lobato quadrature
def _gll(self, nip, alpha, beta):
r = jacobi_roots(nip-2, alpha+1.0, beta+1.0)
xi = numpy.empty(r.shape[0]+2)
xi[0] = -1.0
xi[1:-1] = r
xi[-1] = 1.0
C1 = (2.0**(alpha+beta+1.0))*gamma(alpha+nip)*gamma(beta+nip)
C2 = (nip-1)*gamma(nip)*gamma(alpha+beta+nip+1.0)
Pm = jacobi_p(xi, nip-1, alpha, beta)
w = C1*Pm**(-2)/C2
w[ 0] = w[ 0]*(beta+1.0)
w[-1] = w[-1]*(alpha+1.0)
return xi, w
def n(self):
"""Number of integration points / weights."""
return self.xi.shape[0]
#-- jacobi_gauss_quad.py -------------------------------------------------------
# -
# Creating the quadrature points and weights
quad_data = JacobiGaussQuad(sim_data)
# Lets check the distribution of points and respective weights
pyplot.figure(2)
pyplot.clf()
pyplot.xlim(-1.1, 1.1)
pyplot.plot(quad_data.xi, quad_data.w, "bo")
pyplot.plot(quad_data.xi, numpy.zeros(quad_data.xi.shape), "ro")
pyplot.show()
# ### Basis functions
#
# The _BasisFunctions_ class implements the three possible choices of basis functions (nodal, modal e modal-orthogonal) in a convenient matricial form accessible through the data member `psi`. Basis functions derivatives are also evaluated (data member `Dpsi`). Beyond the _BasisFunctions_ class itself, the file `basis_functions.py` brings the individual implementations of Lagrange, Legendre and Bubble functions.
# +
# # %load basis_functions.py
#
# Basis functions and respective derivatives
#
# by <NAME>. (Matlab and Python versions)
# <NAME> (Python version)
#
import numpy
from sys import float_info
from math import fabs, sqrt
from jacobi_p import jacobi_p
from djacobi_p import djacobi_p, d_pq, d2_pq
from jacobi_roots import jacobi_roots
from simulation_data import QuadratureNodes, BasisType
def gll_nodes(n_local_dof):
"""Gauss-Legendre-Lobatto auxiliar points.
Parameters
----------
n_local_dof
Per-element number of degrees of freedom
"""
if n_local_dof < 2:
raise AssertionError("Local degress of freedom must be >= 2!")
# Setting GLL nodes
alpha = 0.0
beta = 0.0
xj = numpy.empty(n_local_dof)
# GLL points are the roots of a Jacobi polynomial with alpha = beta = 1,
# plus the interval extrema, -1 and 1.
xj[0] = -1.0
xj[1:-1] = jacobi_roots(n_local_dof-2, alpha+1.0, beta+1.0)
xj[-1] = 1.0
return xj
def lagrange(n_local_dof, xi, xj):
"""Lagrange basis function.
Parameters
----------
n_local_dof
Number of local degrees of freedom
xi
Distribution of integration points on the element defining the
Lagrangian basis
xj
Points where we evaluate the basis
"""
# Number of Lagrange basis functions
Q = n_local_dof
QQ1 = Q*(Q-1)
# Number of evaluation points
npts = xi.shape[0]
# Allocates matrix to store Lagrange basis evaluated at npts points
Hj = numpy.ones([npts, Q])
Hj[:, 0] = ((-1.0)**(Q-1)/QQ1)*(xi-1.0)*(0.5*Q*jacobi_p(xi, Q-2, 1.0, 1.0))
Hj[:,-1] = (1.0/QQ1)*(xi+1.0)*(0.5*Q*jacobi_p(xi, Q-2, 1.0, 1.0))
Lp = jacobi_p(xj, Q-1, 0.0, 0.0)
DLp = djacobi_p(xi, Q-1, 0.0, 0.0)
for j in range(1, Q-1): # from the second to the second-last column
for i in range(0, npts):
if fabs(xi[i]-xj[j]) > float_info.epsilon:
Hj[i, j] = (xi[i]**2-1.0)*DLp[i]/(QQ1*Lp[j]*(xi[i]-xj[j]))
return Hj
def bubble_basis(n_local_dof, xi):
"""Modal bubble functions.
n_local_dof
Number of local degrees of freedom
xi
Distribution of integration points on the element
"""
# Number of bubble basis functions
Q = n_local_dof
# Number of evaluation points
npts = xi.shape[0]
# Allocates matrix to store Lagrange basis evaluated at npts points
Bj = numpy.empty([npts, Q])
# Boundary nodal basis functions (necessary to enforce C^(0) continuity)
Bj[:, 0] = 0.5*(1.0-xi)
Bj[:,-1] = 0.5*(1.0+xi)
# Bubble modes
for j in range(1, Q-1):
Lj = jacobi_p(xi, j+1, 0.0, 0.0)
Ljm2 = jacobi_p(xi, j-1, 0.0, 0.0)
Bj[:, j] = (Lj-Ljm2)/sqrt(2.0*(2.0*j+1.0))
return Bj
def legendre_basis(n_local_dof, xi):
"""Legendre basis functions.
n_local_dof
Number of local degrees of freedom
xi
Distribution of integration points on the element
"""
# Number of basis functions
Q = n_local_dof
# Number of evaluation points
npts = xi.shape[0]
# Allocates matrix to store Lagrange basis evaluated at npts points
Lj = numpy.empty([npts, Q])
# Legendre basis
for j in range(0, Q):
Lj[:, j] = jacobi_p(xi, j, 0.0, 0.0)
return Lj
def orthonormal_legendre_basis(n_local_dof, xi):
"""Legendre basis functions.
n_local_dof
Number of local degrees of freedom
xi
Distribution of integration points on the element
"""
# Number of basis functions
Q = n_local_dof
# Number of evaluation points
npts = xi.shape[0]
# Allocates matrix to store Lagrange basis evaluated at npts points
Lj = numpy.empty([npts, Q])
# Legendre basis with the orthonormalization factor
for j in range(0, Q):
Lj[:, j] = jacobi_p(xi, j, 0.0, 0.0)/numpy.sqrt(2.0/(2.0*j+1.0))
return Lj
class BasisFunctions:
"""Basis functions and derivatives on quadrature points."""
def __init__(self, sim_data, gauss_quad):
"""
Parameters
----------
sim_data
SimulationData object
"""
n_local_dof = sim_data.n_local_dof()
# Constructing the basis functions PHI
if sim_data.basis_type == BasisType.NODAL:
xj = gll_nodes(n_local_dof)
self.psi = lagrange(n_local_dof, gauss_quad.xi, xj)
elif sim_data.basis_type == BasisType.MODAL:
self.psi = bubble_basis(n_local_dof, gauss_quad.xi)
elif sim_data.basis_type == BasisType.MODAL_ORTHOGONAL:
self.psi = legendre_basis(n_local_dof, gauss_quad.xi)
elif sim_data.basis_type == BasisType.MODAL_ORTHONORMAL:
self.psi = orthonormal_legendre_basis(n_local_dof, gauss_quad.xi)
else:
raise AssertionError( "Wrong basis type!\n" )
# Now we construct the derivative of the basis functions, DPHI/DX
nip = sim_data.nip()
self.D = numpy.zeros([nip, nip])
if nip > 1:
dp = d_pq(gauss_quad.xi, nip, 0.0, 0.0, sim_data.node_dist)
dp2 = d2_pq(gauss_quad.xi, nip, 0.0, 0.0, sim_data.node_dist)
for i in range(0, nip):
for j in range(0, nip):
# Never do "ifs" inside loops! Just for pedagogic reasons.
if i == j:
self.D[i, j] = dp2[i]/(2.0*dp[i])
else:
dx = gauss_quad.xi[i] - gauss_quad.xi[j]
self.D[i, j] = dp[i]/(dp[j]*dx)
self.dpsi = self.D.dot(self.psi)
#-- basis_functions.py ---------------------------------------------------------
# -
# Basis functions and derivatives evaluated at quadrature points
basis = BasisFunctions(sim_data, quad_data)
# ### Jacobian
#
# As we have seen in the theoretical part, there is a Jacobian involved in the numerical integration due to the transformation of all integrals from the real elements to the standard one. The Jacobian is evaluated by the same _Mesh1D_ class used before. After Jacobian evaluation two more data members become available:
#
# - `J`, the Jacobian matrix, and
# - `x`, a mapping of the integration points (remember, they are all in $[-1,1]$) to the real elements
mesh.jacobian(sim_data, quad_data)
# ### Material properties
#
# In order to be able to run the simulation, we need to evaluate the properties associated with the medium. They are
#
# - Density, $\rho(x)$ in $kg/m^3$
# - Shear modulus, $\mu(x)$ in $Pa$ and its derivative $d\mu/dx$
# - Wavefield velocity, $c$ in $m/s$
#
# The class _MaterialProperties_ is responsible for the evaluation and book keeping of all these properties.
# +
# # %load material_properties.py
#
# Sets several material properties for the elastic wave equation
#
# by <NAME> Jr. (Matlab and Python versions)
# <NAME> (Python version)
#
import numpy
from simulation_data import QuadratureNodes
from math import ceil
class MaterialProperties:
"""Material properties for the elastic wave equation"""
def __init__(self, sim_data, quad_data, mesh, x_source):
"""MaterialProperties constructor
Arguments:
sim_data - SimulationData object
quad_data - Quadrature points and weights
mesh - 1D Finite Elements mesh
x_source - Ricker wavelet point source location
"""
nip = sim_data.nip()
nldof = sim_data.n_local_dof()
dx = sim_data.spatial_domain_length()
self.rho = numpy.zeros(sim_data.n_elements) # Material density [kg/m^3]
self.mu = numpy.zeros([nip, sim_data.n_elements]) # Material shear modulus [Pa]
self.c = numpy.zeros([nip, sim_data.n_elements]) # Material wavefield velocity [m/s]
self.dmudx = numpy.zeros([nip, sim_data.n_elements]) # Derivative of shear modulus
# mu evaluated at the boundaries of each element including ghosts
self.mub = numpy.zeros([nldof, sim_data.n_elements+2])
# We could have set that above, but here is clearer
self.rho.fill(1000.0)
distance = 0.5*dx-mesh.x
# Signal changes
sg = numpy.sign(distance)
distance = numpy.exp(-(numpy.fabs(distance)-500.0)/100.0)
self.mu = 1.2e10-5.96e9/(1.0+distance)
self.dmudx = 5.96E9*(distance*sg)/(100.0*(1.0+distance)**2)
self.c = numpy.sqrt(self.mu/numpy.reshape(numpy.repeat(self.rho, nip), (nip, sim_data.n_elements)))
distance1 = numpy.fabs(mesh.coord[0:-1]-0.5*dx)
distance2 = numpy.fabs(mesh.coord[1:]-0.5*dx)
self.mub[ 0, 1:-1] = 1.2e10-5.96e9/(1.0+numpy.exp(-(distance1-500.0)/100.0))
self.mub[-1, 1:-1] = 1.2e10-5.96e9/(1.0+numpy.exp(-(distance2-500.0)/100.0))
self.mub[-1, 0] = self.mub[0, 1]
self.mub[ 0, -1] = self.mub[-1, -2]
d = numpy.fabs(mesh.x.T.flatten()-x_source)
dist = numpy.amin(d) # The mininum
globnode = numpy.argmin(d) # The index
self.el = int(ceil(globnode/nldof))
self.locid = globnode-nldof*self.el
if sim_data.node_dist == QuadratureNodes.GLL:
if self.locid == 0:
self.globid = globnode+1
self.locid = self.locid+1
elif locid == nldof-1:
self.globid = globnode-1
self.locid = self.locid-1
else:
self.globid = globnode
elif sim_data.node_dist == QuadratureNodes.GL:
self.globid = globnode
#-- material_properties.py -----------------------------------------------------
# -
# Setting material properties
x_source = 3000.0
mat_prop = MaterialProperties(sim_data, quad_data, mesh, x_source)
# ### The source term
#
# The `generate_source` function creates the source term $f(x)$, as well as the time step and the number of simulation steps.
# +
# # %load generate_source.py
#
# Sets several material properties for the elastic wave equation
#
# by <NAME>. (Matlab and Python versions)
# <NAME> (Python version)
#
import numpy
from math import floor
def generate_source(sim_data, mesh, mat_prop):
v_max = numpy.amax(mat_prop.c)
Delta_x_min = numpy.amin(numpy.fabs(mesh.x[0, :]-mesh.x[1, :]))
CFL = 0.4
dt = CFL*Delta_x_min/v_max
Nsteps = int(floor(sim_data.t_max()/dt))
dt = sim_data.t_max()/float(Nsteps)
t_source = 0.1*sim_data.t_max()
sigma = t_source
# Ricker wavelet over the domain [-5,+5]
t_eval = numpy.linspace(-5.0, 5.0, floor(t_source/dt))
Ricker_wavelet = numpy.zeros(Nsteps)
Ricker_wavelet[0:t_eval.shape[0]] = (2.0/(numpy.sqrt(3.0*sigma)*numpy.pi**0.25))*\
(1.0-(t_eval/sigma)**2)*numpy.exp(-0.5*(t_eval/sigma)**2)
# Note: the 6e7 is an arbitrary reescaling
return 6e7*Ricker_wavelet, dt, Nsteps
#-- generate_source.py ---------------------------------------------------------
# -
# Calculates dt and number of time steps
theta, dt, Nsteps = generate_source(sim_data, mesh, mat_prop)
# ### Making room for the solution
#
# We will record every step of the solution so we can make a movie showing the temporal evolution.
# +
# We will store the solution at every time step, so we need to make room for it
u = numpy.zeros([sim_data.n_global_dof(), Nsteps+1])
# Projection of the initial condition onto the elements
phi = numpy.zeros(3*sim_data.n_global_dof())
# Adjusts source term for modal basis functions
s = numpy.zeros(sim_data.n_global_dof())
if sim_data.basis_type == BasisType.MODAL or sim_data.basis_type == BasisType.MODAL_ORTHOGONAL:
idx = sim_data.n_local_dof()*mat_prop.el
s[idx:idx+sim_data.n_local_dof()] = basis.psi[mat_prop.locid, :]
# -
# ### Auxiliary structures
# +
# # %load local_matrices.py
#
# Local mass and stiffness matrices
# =================================
#
# by <NAME> Jr. (Matlab and Python versions)
# <NAME> (Python version)
#
import numpy
def local_mass(quad_data, basis):
"""Constructs the elemental mass matrix
Arguments:
quad_data - Quadrature points and weights
basis - Basis and respective derivatives
Returns:
Mass matrix M, where m_ij = \int_k psi_i psi_j
"""
return numpy.dot(quad_data.w*basis.psi.T, basis.psi)
def local_mass_diagonal(quad_data, basis):
"""Constructs the elemental mass matrix, diagonal version
Arguments:
quad_data - Quadrature points and weights
basis - Basis and respective derivatives
Returns:
Mass matrix M, where m_ii = \int_k psi_i psi_i
"""
return numpy.sum(quad_data.w*basis.psi.T**2, axis=1)
def local_stiffness(quad_data, basis):
"""Constructs the elemental stiffness matrix
Arguments:
quad_data - Quadrature points and weights
basis - Basis and respective derivatives
Returns:
Stiffness matrix S, where s_ij = \int_k psi_i Dpsi_j/dx
"""
return numpy.dot(quad_data.w*basis.psi.T, basis.dpsi)
def stress_stiffness(quad_data, basis, mat_prop, n):
"""Constructs the elemental stress stiffness matrix
Arguments:
quad_data - Quadrature points and weights
basis - Basis and respective derivatives
n - the element number
"""
return (numpy.dot(quad_data.w*mat_prop.mu[:, n]*basis.psi.T, basis.dpsi).T, \
numpy.dot(quad_data.w*mat_prop.dmudx[:, n]*basis.psi.T, basis.psi).T)
#-- local_matrices.py ----------------------------------------------------------
# +
# # %load eval_k_wave.py
#
# Runge-Kutta 4 elemental evaluation (wave equation version)
# ==========================================================
#
# by <NAME> Jr. (Matlab and Python versions)
# <NAME> (Python version)
#
from simulation_data import BasisType
from local_matrices import local_mass, local_stiffness, stress_stiffness
from flux import Fluxes
import numpy
def eval_k_wave(sim_data, quad_data, basis, mesh, mat_prop, phi, s, t):
ng = sim_data.n_global_dof()
nl = sim_data.n_local_dof()
K = sim_data.n_elements # saving some typing...
# Allocates local vectors v and sigma
v_t = numpy.zeros([(K+2)*nl, 2]) # size: K elements + 1 ghost
sigma_t = numpy.zeros(v_t.shape) # size: K elements + 1 ghost
# Gets v and sigma from previous stage of RK4
v_t[nl:(K+1)*nl, 0] = phi[ng:2*ng]
sigma_t[nl:(K+1)*nl,0] = phi[2*ng:3*ng]
# Applies Neumann BC on v (v_ghost = v_element1)
# d/dx v(x_bound) = 0 -> (v_element1 - v_ghost)/delta_x = 0 -> v_ghost = v_element1
v_t[0:nl, 0] = v_t[nl:2*nl, 0] # left side of the domain
v_t[ng+nl:ng+2*nl, 0] = v_t[ng:ng+nl, 0] # right side of the domain
# Gets Mass, Stiffness and Flux matrices
# Note that we multiply S by 2*a
M = local_mass(quad_data, basis)
S = local_stiffness(quad_data, basis)
flux = Fluxes(sim_data)
# Loop over total number of elements
for i in range(1, sim_data.n_elements+1):
Ss, Ssm = stress_stiffness(quad_data, basis, mat_prop, i-1)
idx = numpy.arange(nl*i, nl*(i+1))
m1 = 0.5*flux.FRp1.dot(sigma_t[idx+nl, 0]+numpy.sqrt(mat_prop.rho[i-1]*mat_prop.mub[nl-1, i])*v_t[idx+nl, 0] )
m2 = 0.5*flux.FR.dot(sigma_t[idx, 0]-numpy.sqrt(mat_prop.rho[i-1]*mat_prop.mub[nl-1, i])*v_t[idx, 0])
m3 = 0.5*flux.FL.dot(sigma_t[idx, 0]+numpy.sqrt(mat_prop.rho[i-1]*mat_prop.mub[0, i])*v_t[idx, 0])
m4 = 0.5*flux.FLm1.dot(sigma_t[idx-nl, 0]-numpy.sqrt(mat_prop.rho[i-1]*mat_prop.mub[0, i])*v_t[idx-nl, 0])
tmp = (s[idx-nl]-S.T.dot(sigma_t[idx, 0])+m1+m2-m3-m4)/(mesh.J[i-1]*mat_prop.rho[i-1])
v_t[idx, 1] = numpy.linalg.solve(M, \
tmp )
m1 = 0.5*flux.FRp1.dot(v_t[idx+nl, 0]+(mat_prop.rho[i-1]*mat_prop.mub[nl-1, i])**(-0.5)*sigma_t[idx+nl, 0] )
m2 = 0.5*flux.FR.dot(v_t[idx, 0]-(mat_prop.rho[i-1]*mat_prop.mub[nl-1, i])**(-0.5)*sigma_t[idx, 0])
m3 = 0.5*flux.FL.dot(v_t[idx, 0]+(mat_prop.rho[i-1]*mat_prop.mub[0, i])**(-0.5)*sigma_t[idx, 0])
m4 = 0.5*flux.FLm1.dot(v_t[idx-nl, 0]-(mat_prop.rho[i-1]*mat_prop.mub[0, i])**(-0.5)*sigma_t[idx-nl, 0])
tmp = (-Ss.dot(v_t[idx, 0])-mesh.J[i-1]*Ssm.dot(v_t[idx, 0])+ \
mat_prop.mub[nl-1, i]*(m1+m2)+mat_prop.mub[0, i]*(-m3-m4))/mesh.J[i-1]
sigma_t[idx, 1] = numpy.linalg.solve(M, tmp)
# Assigns local vectors u, v and sigma to the new global vector phi
ki = numpy.zeros(3*ng)
ki[0:ng] = v_t[nl:(K+1)*nl, 0]
ki[ng:2*ng] = v_t[nl:(K+1)*nl, 1]
ki[2*ng:3*ng] = sigma_t[nl:(K+1)*nl, 1]
return ki
#-- eval_k_wave.py -------------------------------------------------------------
# -
# ### Finally, the time iteration!
#
# Load the `rk4_wave.py` file and be happy!
# +
# # %load rk4_wave.py
#
# Runge-Kutta 4 steps method (wave equation version)
# ==================================================
#
# by <NAME>. (Matlab and Python versions)
# <NAME> (Python version)
#
from simulation_data import BasisType
from eval_k_wave import eval_k_wave
def RK4_wave(sim_data, quad_data, basis, mesh, mat_prop, tstep, t, dt, u, src, theta, phi):
"""Runge-Kutta 4 steps.
Arguments:
sim_data - SimulationData object
quad_data - Quadrature points and weights
basis - Basis and respectivederivatives
mesh - 1D Finite Elements mesh
tstep - present time step (integer)
t - simulation time
dt - delta t
u - solution
s -
phi -
Returns:
A step of the RK4 method
"""
ngdof = sim_data.n_global_dof()
nldof = sim_data.n_local_dof()
# Sets source term
s = src.copy()
if sim_data.basis_type == BasisType.NODAL:
s[mat_prop.globid] = theta[tstep-1]
else:
s[nldof*mat_prop.el:nldof*(mat_prop.el+1)] = s[nldof*mat_prop.el:nldof*(mat_prop.el+1)]*theta[tstep-1]
# RK4
k1 = eval_k_wave(sim_data, quad_data, basis, mesh, mat_prop, phi, s, t)
k2 = eval_k_wave(sim_data, quad_data, basis, mesh, mat_prop, phi+0.5*dt*k1, s, t)
k3 = eval_k_wave(sim_data, quad_data, basis, mesh, mat_prop, phi+0.5*dt*k2, s, t)
k4 = eval_k_wave(sim_data, quad_data, basis, mesh, mat_prop, phi+dt*k3, s, t)
phi = phi+dt*(k1+2.0*k2+2.0*k3+k4)/6.0
# Recovers displacement solution u for modal type basis
if sim_data.basis_type == BasisType.NODAL:
u[:, tstep] = phi[0:ngdof]
else:
nip = sim_data.nip()
nldof = sim_data.n_local_dof()
u_hat = phi[0:ngdof]
for k in range(0, sim_data.n_elements):
u[nip*k:nip*(k+1), tstep] = \
u_hat[nldof*k:nldof*(k+1)].dot(basis.psi.T)
return (u, phi)
#-- rk4_wave.py ----------------------------------------------------------------
# +
# Time marching (RK4 time integrator)
t = sim_data.t_min()
for tstep in range(1, Nsteps):
u, phi = RK4_wave(sim_data, quad_data, basis, mesh, mat_prop, tstep, t, dt, u, s, theta, phi)
t = t+dt
# -
# ### Viewing the solution
#
# If you have installed the JSAnimation library you can generate a small movie showing the evolution of the solution along the time. If you get a message __`ImportError: No module named JSAnimation.IPython_display`__ do not dispair! We can still resort to matplotlib to plot a few steps.
# #### Creating a movie
from matplotlib import animation
from JSAnimation.IPython_display import display_animation
# +
# Lets do some movies!
# From https://jakevdp.github.io/blog/2012/08/18/matplotlib-animation-tutorial/
# First set up the figure, the axis, and the plot element we want to animate
fig = pyplot.figure()
ax = pyplot.axes(xlim=(sim_data.spatial_domain[0], sim_data.spatial_domain[1]),ylim=(numpy.min(u), numpy.max(u)))
ax.grid(True)
line, = ax.plot([], [], lw=2)
# Initialization function: plot the background of each frame
def init():
line.set_data([], [])
return line,
# Animation function. This is called sequentially
def animate(i):
global u, mesh
x = mesh.x.T.flat
y = u[:, i]
line.set_data(x, y)
return line,
# call the animator. blit=True means only re-draw the parts that have changed.
anim = animation.FuncAnimation(fig, animate, init_func=init,frames=Nsteps, interval=20, blit=True)
display_animation(anim, default_mode='once')
#pyplot.show()
# -
# #### Plotting some steps of the solution
#
# The solution _u_ has _Nsteps_ columns, precisely one for each time step. In order to view some is simple, as shown in the code below.
# Plotting a few samples of the solution along the time
pyplot.figure(5)
pyplot.clf()
x = mesh.x.T.flat
pyplot.plot(x, u[:,150]) # tstep = 150
pyplot.plot(x, u[:,300]) # tstep = 300
pyplot.plot(x, u[:,600]) # tstep = 600
pyplot.plot(x, u[:,2100]) # tstep = 900
pyplot.show()
| lessons/08_dg/08_03_Elastic_Wave_1D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:snorkeling]
# language: python
# name: conda-env-snorkeling-py
# ---
# # Generate Compound Binds Gene Candidates
# This notebook is designed to construct a table that contains compound and gene pairs with various statistics (number of sentences, if contained in hetionet, if the edge has sentences and which training category each pair belongs to).
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from collections import Counter
from itertools import product
import os
import pickle
import sys
import pandas as pd
# +
#Set up the environment
username = "danich1"
password = "<PASSWORD>"
dbname = "pubmeddb"
#Path subject to change for different os
database_str = "postgresql+psycopg2://{}:{}@/{}?host=/var/run/postgresql".format(username, password, dbname)
# -
compound_url = "https://raw.githubusercontent.com/dhimmel/drugbank/7b94454b14a2fa4bb9387cb3b4b9924619cfbd3e/data/drugbank.tsv"
gene_url = "https://raw.githubusercontent.com/dhimmel/entrez-gene/a7362748a34211e5df6f2d185bb3246279760546/data/genes-human.tsv"
cbg_url = "https://raw.githubusercontent.com/dhimmel/integrate/93feba1765fbcd76fd79e22f25121f5399629148/compile/CbG-binding.tsv"
# ## Read in Gene and Compound Entities
entrez_gene_df = pd.read_table(gene_url).rename(index=str, columns={"GeneID": "entrez_gene_id", "Symbol":"gene_symbol"})
entrez_gene_df.head(2)
drugbank_df = pd.read_table(compound_url).rename(index=str, columns={'name':'drug_name'})
drugbank_df.head(2)
# ## Read in Compound Binds/Regulates Gene Tables
compound_binds_gene_df = pd.read_table(cbg_url, dtype={'entrez_gene_id': int})
compound_binds_gene_df.head(2)
# ## Read in Sentences with Edge Pair
# +
query = '''
SELECT "Compound_cid" AS drugbank_id, "Gene_cid" AS entrez_gene_id, count(*) AS n_sentences
FROM compound_gene
GROUP BY "Compound_cid", "Gene_cid";
'''
compound_gene_sentence_df = (
pd
.read_sql(query, database_str)
.astype({"entrez_gene_id":int})
.merge(drugbank_df[["drugbank_id", "drug_name"]], on="drugbank_id")
.merge(entrez_gene_df[["entrez_gene_id", "gene_symbol"]], on="entrez_gene_id")
)
compound_gene_sentence_df.head(2)
# -
# ## Merge Edges Into a Unified Table
compound_binds_gene_df = (
compound_binds_gene_df
.merge(compound_gene_sentence_df, on=["drugbank_id", "entrez_gene_id"], how="outer")
)
compound_binds_gene_df=(
compound_binds_gene_df
.assign(hetionet=compound_binds_gene_df.sources.notnull().astype(int))
.assign(has_sentence=(compound_binds_gene_df.n_sentences > 0).astype(int))
)
compound_binds_gene_df.head(2)
# Make sure all existing edges are found
# 11571 is determined from neo4j to be all DaG Edges
assert compound_binds_gene_df.hetionet.value_counts()[1] == 24687
compound_binds_gene_df.query("hetionet==1&has_sentence==1").shape
# Make note that 18741 edges in Hetionet do not have sentences
# ## Sort Edges into categories
def partitioner(df):
"""
This function creates a parition rank for the current dataset.
This algorithm assigns a rank [0-1) for each datapoint inside each group (outlined below):
1,1 -in hetionet and has sentences
1,0 - in hetionet and doesn't have sentences
0,1 - not in hetionet and does have sentences
0,0, - not in hetionet and doesn't have sentences
This ranking will be used in the get split function to assign each datapoint
into its corresponding category (train, dev, test)
"""
partition_rank = pd.np.linspace(0, 1, num=len(df), endpoint=False)
pd.np.random.shuffle(partition_rank)
df['partition_rank'] = partition_rank
return df
def get_split(partition_rank, training=0.7, dev=0.2, test=0.1):
"""
This function partitions the data into training, dev, and test sets
The partitioning algorithm is as follows:
1. anything less than 0.7 goes into training and receives an appropiate label
2. If not less than 0.7 subtract 0.7 and see if the rank is less than 0.2 if not assign to dev
3. Lastly if the rank is greater than 0.9 (0.7+0.2) assign it to test set.
return label that corresponds to appropiate dataset cateogories
"""
if partition_rank < training:
return 6
partition_rank -= training
if partition_rank < dev:
return 7
partition_rank -= dev
assert partition_rank <= test
return 8
pd.np.random.seed(100)
cbg_map_df = compound_binds_gene_df.groupby(['hetionet', 'has_sentence']).apply(partitioner)
cbg_map_df.head(2)
cbg_map_df['split'] = cbg_map_df.partition_rank.map(get_split)
cbg_map_df.split.value_counts()
cbg_map_df.sources.unique()
cbg_map_df = cbg_map_df[[
"drugbank_id", "drug_name",
"entrez_gene_id", "gene_symbol",
"sources", "n_sentences",
"hetionet", "has_sentence",
"split", "partition_rank"
]]
cbg_map_df.head(2)
cbg_map_df.to_csv("output/compound_binds_gene.tsv.xz", sep="\t", compression="xz", index=False)
| compound_gene/compound_binds_gene/datafile/compound_gene_datafile_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2
#
# ## Video 6: Lists and List Operations
# **Python for the Energy Industry**
#
# ## Lists
#
# We often want to keep track of many bits of data at once. One way of doing this in python is with a 'list'. Lists are defined by enclosing multiple comma-separated values inside square brackets.
# +
# list of strings
vessels = ['ABIOLA', 'ACACIA']
# list of floats
prices = [1.23, 1.98, 1.45, 1.67]
# lists of integers
quantities1 = [18, 22, 21, 32]
quantities2 = [12, 11, 16, 18]
# -
# ### Looping Over Lists
# loop over prices
for price in prices:
print(price)
# ### List Indexing
#
# We can access individual items in a list in the following way:
# +
prices = [1.23, 1.98, 1.45, 1.67, 2.40, 1.89]
print('The 1st price: ', prices[0] )
print('The 2nd price: ', prices[1] )
print('The 3rd price: ', prices[2] )
# -
# The number inside the square brackets is the 'index' of the item. Note that Python uses 'zero indexing', so the first item in a list has an index of 0.
#
# You can also access items 'from the end' using a negative index. So an index of -1 gives the last item, -2 gives the second last item, and so on.
print('The last price: ', prices[-1] )
print('The 2nd last price: ', prices[-2] )
# The index can also be used to overwrite an item in the list:
print(prices)
prices[0] = 2.41
prices[-1] = 0.99
print(prices)
# You can also get a range of items from a list:
print(prices[1:4] )
# This is called a 'slice'. Which indices from the prices list are included in this slice?
# ## List Operations
#
# Lists have some built in functionality that makes them easier to work with.
#
# ### Adding to / Removing from Lists
# +
countries = ['USA', 'UK', 'France', 'Germany']
# adding on to the end of a list
countries.append('Spain')
print(countries)
# -
# inserting into a specific index of a list
countries.insert(2, 'Australia')
print(countries)
# removing from a specific index
countries.pop(3)
print(countries)
# removing a particular value from a list
countries.remove('Germany')
print(countries)
# ## Examining Lists
# +
primes = [2,3,5,7,11,13]
# check if a value is in a list
print(3 in primes)
print(4 in primes)
# -
# find the index of a value in a list
print(primes.index(5))
# find the minimum, maximum, and sum
print('minimum:', min(primes))
print('maximum:', max(primes))
print('sum:', sum(primes))
# ### Other List Operations
#
# The addition and multiplication operators also work on lists:
print(quantities1 + quantities2)
print(quantities1 * 2)
# ### Exercise
#
# Python allows you to have lists containing multiple different types of variable, as in the below example. Try to transform this list into: [1,2,3] using the operations you've learned.
| docs/examples/academy/6. Lists and List Operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inverted encoding models, revisited
import numpy as np
import matplotlib.pyplot as plt
import pymc3 as pm
from scipy.stats import pearsonr
from sklearn.base import RegressorMixin, BaseEstimator
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import OneHotEncoder
import seaborn as sns
from scipy.ndimage import gaussian_filter
from scipy.linalg import toeplitz
from sklearn.discriminant_analysis import _cov, LinearDiscriminantAnalysis
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from scipy import stats
import pymc3 as pm
# %matplotlib inline
# +
class Iem(BaseEstimator, RegressorMixin):
""" Base class for Inverted Encoding Model. """
def __init__(self, classify=True, score_func=None):
""" Initializes base class. """
self.W = None # estimated parameters
self.classify = classify
def _preproc(self, S):
""" Preprocesses stimulus features (S). """
if self.classify and S.ndim == 1:
S = OneHotEncoder(sparse=False).fit_transform(S[:, np.newaxis])
elif not self.classify and S.ndim == 1:
S = np.c_[np.ones(S.shape[0]), S]
return S
def _classify(self, S_pred):
""" Makes predictions categorical. """
return np.argmax(S_pred, axis=0)
def fit(self, R, S):
""" Fits model (should be defined in child class). """
raise NotImplementedError
def predict(self, R):
""" Predicts new stimuli based on responses
(should be defined in child class). """
raise NotImplementedError
def score(self, R, S):
""" Scores prediction. """
S_pred = self.predict(R)
if self.classify:
return np.mean(S_pred == S)
class OlsIem(Iem):
def fit(self, R, S):
S = self._preproc(S)
self.W = np.linalg.inv(S.T @ S) @ S.T @ R
return self
def predict(self, R):
S_pred = np.linalg.pinv(self.W @ self.W.T) @ self.W @ R.T
if self.classify:
S_pred = self._classify(S_pred)
return S_pred
class WlsIem(Iem):
def fit(self, R, S):
S = self._preproc(S)
self.W = np.linalg.inv(S.T @ S) @ S.T @ R
resids = R - S @ self.W
var_err = np.var(resids, axis=0)
omega = np.eye(resids.shape[1])
np.fill_diagonal(omega, var_err)
self.omega = np.linalg.inv(omega)
return self
def predict(self, R):
W, omega = self.W, self.omega
S_pred = np.linalg.pinv(W @ omega @ W.T) @ W @ omega @ R.T
if self.classify:
S_pred = self._classify(S_pred)
return S_pred
class GlsIem(Iem):
def __init__(self, shrink_cov='auto', classify=True):
self.shrink_cov = shrink_cov
super().__init__(classify=classify)
def fit(self, R, S):
S = self._preproc(S)
self.W = np.linalg.inv(S.T @ S) @ S.T @ R
resids = R - S @ self.W
cov_err = _cov(resids, shrinkage=self.shrink_cov)
self.omega = np.linalg.inv(cov_err)
return self
def predict(self, R):
W, omega = self.W, self.omega
S_pred = np.linalg.pinv(W @ omega @ W.T) @ W @ omega @ R.T
if self.classify:
S_pred = self._classify(S_pred)
return S_pred
class RidgeGlsIem(GlsIem):
def __init__(self, alpha=1, classify=True, shrink_cov='auto'):
self.alpha = alpha
super().__init__(classify=classify, shrink_cov=shrink_cov)
def predict(self, R):
W, omega = self.W, self.omega
S_pred = (np.linalg.pinv(W @ omega @ W.T) + self.alpha*np.eye(W.shape[0])) @ W @ omega @ R.T
if self.classify:
S_pred = self._classify(S_pred)
return S_pred
# +
class DataGenerator:
def __init__(self, categorical=True, N=100, P=4, K=50, sig_sq=1,
rho=0.9, max_var=10, noise_model='ols', param_model='unif'):
self.categorical = categorical
self.N = N
self.P = P
self.K = K
self.sig_sq = sig_sq
self.rho = rho # ar1 param
self.max_var = max_var
self.noise_model = noise_model
self.param_model = param_model
def generate(self):
N, P, K = self.N, self.P, self.K
S = self._generate_design()
eps = self._generate_noise()
W = self._generate_params()
R = S.dot(W) + eps
if self.categorical:
S = np.argmax(S, axis=1)
return S, R
def _generate_design(self):
N, P = self.N, self.P
if self.categorical:
S_tmp = np.repeat(np.arange(P), N / P)[:, np.newaxis]
S = OneHotEncoder(sparse=False).fit_transform(S_tmp)
else:
S = np.random.normal(0, 1, size=(N, P))
return S
def _generate_noise(self):
N, K = self.N, self.K
noise_mu = np.zeros(K)
if self.noise_model == 'ols':
noise_cov = np.identity(K)
elif self.noise_model in ['wls', 'gls', 'wgls']:
if self.noise_model == 'gls':
# assuming equal variance, but with non-zero covariance
noise_cov = self.rho ** toeplitz(np.arange(K))
else:
varz = np.random.uniform(0, self.max_var, size=K)
if self.noise_model == 'wls':
noise_cov = np.diag(varz)
else:
corr_cov = self.rho ** toeplitz(np.arange(K))
varz = varz[:, np.newaxis]
noise_cov = np.sqrt(varz.dot(varz.T))
noise_cov *= corr_cov
noise = np.random.multivariate_normal(noise_mu, self.sig_sq*noise_cov, size=N)
return noise
def _generate_params(self):
P, K = self.P, self.K
params_mu = np.zeros(P)
if self.param_model == 'unif':
W = np.random.uniform(-.5, .5, size=(P, K))
elif self.param_model == 'ols':
params_cov = np.identity(P) / 10
W = np.random.multivariate_normal(params_mu, params_cov, size=K).T
elif self.param_model == 'gls':
params_cov = 0.5 ** toeplitz(np.arange(P))
W = np.random.multivariate_normal(params_mu, params_cov, size=K).T
elif self.param_model == 'wls':
varz = np.random.uniform(0, 1, size=P)
params_cov = np.diag(varz)
W = np.random.multivariate_normal(params_mu, params_cov, size=K).T
elif self.param_model == 'wgls':
varz = np.random.uniform(0, 1, size=P)[:, np.newaxis]
params_cov = np.sqrt(varz.dot(varz.T))
params_cov *= 0.5 ** toeplitz(np.arange(P))
W = np.random.multivariate_normal(params_mu, params_cov, size=K).T
return W
for categorical in [True, False]:
# print("categorical: %s" % categorical)
for noise_model in ['ols', 'wls', 'gls', 'wgls']:
# print('\t noise_model: %s' % noise_model)
for param_model in ['unif', 'ols', 'wls', 'gls', 'wgls']:
# print('\t\t param_model: %s' % param_model)
dgn = DataGenerator(categorical=categorical, N=100, P=2, K=50,
sig_sq=1, noise_model=noise_model, param_model=param_model)
S, R = dgn.generate()
# +
N = 200
P = 2
K = 100
sig_sq = 10
iters = 500
fig, axes = plt.subplots(ncols=4, figsize=(20, 5), sharex=True, sharey=True)
clfs = [OlsIem(), WlsIem(), GlsIem(), GaussianNB(), LinearDiscriminantAnalysis(shrinkage='auto', solver='lsqr')]
for i, noise_model in enumerate(['ols', 'wls', 'gls', 'wgls']):
scores = np.zeros((iters, len(clfs)))
for ii in range(iters):
S, R = DataGenerator(categorical=True, N=N, P=P, K=K, sig_sq=sig_sq,
noise_model=noise_model).generate()
for iii, clf in enumerate(clfs):
scores[ii, iii] = cross_val_score(estimator=clf, X=R, y=S, cv=10).mean()
for ii in range(scores.shape[1]):
sns.distplot(scores[:, ii], ax=axes[i], hist=False, label=clfs[ii].__class__.__name__,
kde_kws={'lw': 4})
axes[i].set_title('Noise model: %s' % noise_model)
sns.despine()
fig.tight_layout()
fig.show()
| iem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import sklearn
import matplotlib
from matplotlib.pyplot import plot
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.cluster import KMeans
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import MultinomialNB
from sklearn import svm
import nltk
import xgboost
import os
from sklearn.decomposition import PCA
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import metrics
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import mean_squared_error
from sklearn.tree import DecisionTreeClassifier
import random
import time
from datetime import date
from statsmodels.tsa.stattools import adfuller, acf, pacf
from statsmodels.tsa.arima_model import ARIMA
from statsmodels.tsa.seasonal import seasonal_decompose
import seaborn as sns
import math
import sys
from numpy.random import randn
from scipy.cluster.hierarchy import dendogram, linkage
from copy import deepcopy
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV
from sklearn.datasets import load_breast_cancer
from sklearn.datasets import make_blobs
from mlxtend.frequent_patterns import apriori
from mlxtend.frequent_patterns import association_rules
from sklearn.metrics import pairwise_distances
from math import sqrt
from io import StringIO
import json
from matplotlib.pylab import rcParams
from matplotlib.pylab import plt
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import preprocessing
from sklearn.preprocessing import LabelEncoder
import base64
from numpy.linalg import *
from matplotlib.pyplot import *
from sklearn.cluster import AgglomerativeClustering
import pylab as pb
import networkx as nx
import warnings
warnings.filterwarnings('ignore')
from pandas import DataFrame
import statsmodels.api as sm
import statsmodels.tsa.api as smt
import statsmodels.formula.api as smf
| Python libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to use pyHomogeneity package functions : a brief example
# The **Daily Female Births Dataset** is used here for demonstrating the use of this package. This dataset describes the number of daily female births in California in 1959. This dataset is available in [here](https://raw.githubusercontent.com/jbrownlee/Datasets/master/daily-total-female-births.csv).
#
# +
import numpy as np
import pandas as pd
import pyhomogeneity as hg
import matplotlib.pyplot as plt
import statsmodels.api as sm
# %matplotlib inline
# read all datasets
Birth_data = pd.read_csv("daily-total-female-births.csv")
Birth_data.index = pd.to_datetime(Birth_data.Date,format='%d/%m/%Y')
Birth_data.drop('Date', axis=1,inplace=True)
# -
Birth_data.head()
Birth_data.plot(figsize=(16,6));
pettitt_res = hg.pettitt_test(Birth_data, alpha=0.05)
pettitt_res
snht_res = hg.snht_test(Birth_data, sim=10000)
snht_res
buishand_res = hg.buishand_u_test(Birth_data)
buishand_res
# All homogeneity test shows that there is a significant change-point in this dataset. Because the **p-value** is smaller than **alpha=0.05** and **h=True**. The change-point is located at **1959-06-29**. The mean value of before and after change-point is **39.638888888888886** and **44.25945945945946**.
# +
result = pettitt_res
mn = Birth_data.index[0]
mx = Birth_data.index[-1]
loc = pd.to_datetime(result.cp)
mu1 = result.avg.mu1
mu2 = result.avg.mu2
plt.figure(figsize=(16,6))
plt.plot(Birth_data, label="Observation")
plt.hlines(mu1, xmin=mn, xmax=loc, linestyles='--', colors='orange',lw=1.5, label='mu1 : ' + str(round(mu1,2)))
plt.hlines(mu2, xmin=loc, xmax=mx, linestyles='--', colors='g', lw=1.5, label='mu2 : ' + str(round(mu2,2)))
plt.axvline(x=loc, linestyle='-.' , color='red', lw=1.5, label='Change point : '+ loc.strftime('%Y-%m-%d') + '\n p-value : ' + str(result.p))
plt.title('Daily Female Births')
plt.xlabel('Date')
plt.ylabel('Births')
plt.legend(loc='upper right')
# plt.savefig("F:/aaaaaa.jpg", dpi=600)
# -
| Examples/Example_pyHomogeneity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: new_name
# language: python
# name: python3
# ---
# +
import os
import csv
csvpath = os.path.join("Resources", "budget_data.csv")
with open(csvpath, newline="") as csvfile:
csv_header = next(csvfile)
row_count = sum(1 for row in csvfile)
with open(csvpath, newline="") as csvfile2:
csv_header = next(csvfile2)
profit_losses = sum(int(r[1]) for r in csv.reader(csvfile2))
average_changes = int(profit_losses / row_count)
with open(csvpath, "r") as csvfile3:
csv_reader = csv.DictReader(csvfile3)
data = list(csv_reader)
maxProfit = max(data, key=lambda x:x['Profit/Losses'])
minProfit = min(data, key=lambda x:x['Profit/Losses'])
print("Financial Analysis")
print("--------------------------")
print("Total Months:", row_count)
print("Total: $", profit_losses)
print("Average Change: $", average_changes)
print("Greatest Increase in Profits:", maxProfit)
print("Greatest Decrease in Profits:", minProfit)
print("Financial Analysis", file=open("output.txt", "w"))
print("--------------------------", file=open("output.txt", "a"))
print("Total Months:", row_count, file=open("output.txt", "a"))
print("Total: $", profit_losses, file=open("output.txt", "a"))
print("Average Change: $", average_changes, file=open("output.txt", "a"))
print("Greatest Increase in Profits:", maxProfit, file=open("output.txt", "a"))
print("Greatest Decrease in Profits:", minProfit, file=open("output.txt", "a"))
# -
| PyBank/Main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
# -
matplotlib.__version__
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot([1,2,3,4,5], [1,2,3,4,5])
import matplotlib.pyplot as plt
plt.plot([1, 2, 3, 4, 5], [1, 2, 3, 4, 5])
plt.show()
# +
import numpy as np
x = np.linspace(0, 10, 50)
y = x
plt.title("Линейная зависимость y = x")
plt.xlabel("x")
plt.ylabel("y")
plt.grid()
plt.plot(x, y)
# +
plt.title("Линейная зависимость y = x")
plt.xlabel("x")
plt.ylabel("y")
plt.grid()
plt.plot(x, y,"r--" )
# +
x = np.linspace(0, 10, 50)
y1 = x
y2 = [i**2 for i in x]
plt.title("Зависимости: y1 = x, y2 = x^2")
plt.xlabel("x")
plt.ylabel("y1, y2")
plt.grid()
plt.plot(x, y1, x, y2)
# +
x = np.linspace(0, 10, 50)
y1 = x
y2 = [i**2 for i in x]
plt.figure(figsize=(9, 9))
plt.subplot(2, 1, 1)
plt.plot(x, y1)
plt.title("Зависимости: y1 = x, y2 = x^2")
plt.ylabel("y1", fontsize=14)
plt.grid(True)
plt.subplot(2, 1, 2)
plt.plot(x, y2)
plt.xlabel("x", fontsize=14)
plt.ylabel("y2", fontsize=14)
plt.grid(True)
# +
fruits = ['apple','peach','orange','bannana','melon']
counts = [34,25,43,31,17]
plt.bar(fruits,counts)
plt.title('Fruits!')
plt.xlabel('Fruit')
plt.ylabel('Count')
# +
import matplotlib.pyplot as plt
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator)
import numpy as np
x = np.linspace(0, 10, 10)
y1 = 4*x
y2 = [i**2 for i in x]
fig, ax = plt.subplots(figsize=(8, 6))
ax.set_title("Графики зависимостей: y1=4*x, y2=x^2", fontsize=16)
ax.set_xlabel("x", fontsize=14)
ax.set_ylabel("y1, y2", fontsize=14)
ax.grid(which="major", linewidth=1.2)
ax.grid(which="minor", linestyle="--", color="gray", linewidth=0.5)
ax.scatter(x, y1, c="red", label="y1 = 4*x")
ax.plot(x, y2, label="y2 = x^2")
ax.legend()
ax.xaxis.set_minor_locator(AutoMinorLocator())
ax.yaxis.set_minor_locator(AutoMinorLocator())
ax.tick_params(which='major', length=10, width=2)
ax.tick_params(which='minor', length=5, width=1)
plt.show()
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot()
plt.plot([1,7,3,5,11,1])
plt.plot([1, 5, 10, 15, 20], [1, 7, 3, 5, 11])
# +
x = [1, 5, 10, 15, 20]
y = [1, 7, 3, 5, 11]
plt.plot(x, y, label='steel price')
plt.title('Chart price', fontsize=15)
plt.xlabel('Day', fontsize=12, color='blue')
plt.ylabel('Price', fontsize=12, color='blue')
plt.legend()
plt.grid(True)
plt.text(15, 4, 'grow up!')
# -
x = [1, 5, 10, 15, 20]
y = [1, 7, 3, 5, 11]
plt.plot(x, y, '--')
x = [1, 5, 10, 15, 20]
y = [1, 7, 3, 5, 11]
line = plt.plot(x, y)
plt.setp(line, linestyle='--')
# +
x = [1, 5, 10, 15, 20]
y1 = [1, 7, 3, 5, 11]
y2 = [i*1.2 + 1 for i in y1]
y3 = [i*1.2 + 1 for i in y2]
y4 = [i*1.2 + 1 for i in y3]
plt.plot(x, y1, '-', x, y2, '--', x, y3, '-.', x, y4, ':')
# -
plt.plot(x, y, 'ro')
plt.plot(x, y, 'bx')
| united.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# GitHub
# https://github.com/clovaai/CRAFT-pytorch
# 論文
# https://arxiv.org/abs/1904.01941
#
# <a href="https://colab.research.google.com/github/kaz12tech/ai_demos/blob/master/CRAFT_demo.ipynb" target="_blank"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="4TP5UEjJuRjL"
# ## GitHubからCRAFTのソースコードを取得
# + colab={"base_uri": "https://localhost:8080/"} id="JEUMCU9tpZCw" outputId="2071fb47-624e-444d-d3a0-91d2f34ada30"
# %cd /content/
# !git clone https://github.com/clovaai/CRAFT-pytorch.git
# + [markdown] id="hRwgjpBkurB5"
# ## ライブラリのインストール
# + colab={"base_uri": "https://localhost:8080/"} id="KE1PSdNdpqAc" outputId="d203fc5b-e3e3-42eb-f344-5f5ef426410d"
# %cd /content/CRAFT-pytorch/
# !pip uninstall torch torchtext torchaudio -y
# !pip install -r requirements.txt
# + [markdown] id="vopQQhndvxrb"
# ## ライブラリのインポート
# + id="T-cN5xI4vz_q"
import os
import gdown
import glob
from PIL import Image
import matplotlib.pyplot as plt
import random
import shutil
import torch
# + [markdown] id="u1D8w4gcutPM"
# ## 学習済みモデルのダウンロード
# + colab={"base_uri": "https://localhost:8080/"} id="-8KqTN4hsgvK" outputId="f729305b-c70f-4f07-cd1a-e7aaa155351b"
# %cd /content/CRAFT-pytorch/
# !mkdir pretrained
# %cd /content/CRAFT-pytorch/pretrained
for google_drive_file_id in ['1Jk4eGD7crsqCCg9C9VjCLkMN3ze8kutZ']:
gdown.download(
url='https://drive.google.com/uc?id={}'.format(google_drive_file_id),
output="./craft_mlt_25k.pth",
quiet=False
)
# + [markdown] id="2m4eUC5S1_NK"
# ## データセットのダウンロード
# + colab={"base_uri": "https://localhost:8080/"} id="KD1Xz-SfvTo1" outputId="5d230647-a8f4-4ba4-d4ae-161ace396d27"
# %cd /content/CRAFT-pytorch/
# !mkdir dataset
# %cd /content/CRAFT-pytorch/dataset
for google_drive_file_id in ['1sptDnAomQHFVZbjvnWt2uBvyeJ-gEl-A']:
gdown.download(
url='https://drive.google.com/uc?id={}'.format(google_drive_file_id),
output="./icdar2013.zip",
quiet=False
)
# ダウンロードしたzipファイルの解凍
# 解凍したファイルすべてが出力されることを避けるため > /dev/null で出力を捨てる
# !unzip ./icdar2013.zip > /dev/null
# icdar2013 testファイルリストを取得
test_path = "/content/CRAFT-pytorch/dataset/icdar2013/test_images/"
test_path_list = glob.glob(test_path + "*.jpg")
# + [markdown] id="iKViCIIm2BVs"
# ## 文字検出(Character Detection)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="qDYlc-qB2Toz" outputId="14be8ca8-2e58-4106-9ecf-83fecb641c67"
# 文字検出対象のファイル選定
target_path = "/content/CRAFT-pytorch/inputs/"
if os.path.exists(target_path):
shutil.rmtree(target_path)
os.makedirs(target_path, exist_ok=True)
else:
os.makedirs(target_path, exist_ok=True)
# 文字検出用にランダムに5枚選択
target = random.sample(test_path_list, 5)
for file_path in target:
shutil.copy(file_path, target_path)
target_path_list = glob.glob(target_path + "*.jpg")
# 文字検出対象画像の表示
for t_path in target_path_list:
image = Image.open(t_path).convert("RGB")
plt.figure(figsize=(6, 6))
plt.imshow(image)
# + colab={"base_uri": "https://localhost:8080/"} id="dZY-mjjO2rnD" outputId="725fbb91-bd75-4863-e624-aea15301a800"
# %cd /content/CRAFT-pytorch/
# GPUが使用可能であればTrue、不可であればFalse
device = torch.cuda.is_available()
test_img = test_path_list[0]
# !python3 test.py \
# --trained_model=/content/CRAFT-pytorch/pretrained/craft_mlt_25k.pth \
# --test_folder=$target_path \
# --cuda=$device
# + [markdown] id="UFlkF9riEgkF"
# ## 文字検出結果の表示
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="xQF0ESav4xzr" outputId="61ae4c2d-8065-4ee0-ec6d-4bd9b9e8b578"
results = glob.glob("/content/CRAFT-pytorch/result/*.jpg")
results = [f for f in results if not 'mask' in f ]
# 文字検出対象画像の表示
for result in results:
image = Image.open(result).convert("RGB")
plt.figure(figsize=(6, 6))
plt.imshow(image)
| CRAFT_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Machine Learning for Engineers: [SplitData](https://www.apmonitor.com/pds/index.php/Main/SplitData)
# - [Split Data: Train, Validate, Test](https://www.apmonitor.com/pds/index.php/Main/SplitData)
# - Source Blocks: 3
# - Description: Splitting data ensures that there are independent sets for training, testing, and validation.
# - [Course Overview](https://apmonitor.com/pds)
# - [Course Schedule](https://apmonitor.com/pds/index.php/Main/CourseSchedule)
#
# +
import pandas as pd
from sklearn.model_selection import train_test_split
data = pd.read_csv('http://apmonitor.com/pds/uploads/Main/tclab_data6.txt')
data.set_index('Time',inplace=True)
# Split into train and test subsets (20% for test)
train, test = train_test_split(data, test_size=0.2, shuffle=False)
print('Train: ', len(train))
print(train.head())
print('Test: ', len(test))
print(test.head())
# +
from sklearn.datasets import make_classification
X, y = make_classification(n_samples=5000, n_features=20, n_informative=15)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, shuffle=True)
print(len(X),len(X_train),len(X_test))
# +
from sklearn.model_selection import KFold
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.datasets import make_classification
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# define dataset
X, y = make_classification(n_samples=5000, n_features=20, n_informative=15)
# Set up K-fold cross validation
kf = KFold(n_splits=5,shuffle=True)
# Initialize model
dtc = DecisionTreeClassifier()
# Array to store accuracy scores
scores = np.zeros(5)
# Initialize plot
plt.figure(figsize=(12,2))
for i,(train_index, test_index) in enumerate(kf.split(X)):
Xtrain, Xtest = X[train_index], X[test_index]
ytrain, ytest = y[train_index], y[test_index]
dtc.fit(Xtrain,ytrain)
yp = dtc.predict(Xtest)
acc = accuracy_score(ytest,yp)
scores[i] = acc
plt.subplot(1,5,i+1)
cm = confusion_matrix(yp,ytest)
sns.heatmap(cm,annot=True)
plt.show()
print('Accuracy: %.2f%%' %(np.mean(scores*100)))
| All_Source_Code/SplitData/SplitData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3.1 Comments and Docstrings
# Comments are used to explain the code. The interpreter doesn't execute the comments. There are 3 types of comments in Python:
# * Single Line
# * In-Line
# * Multi Line
# ## Single Line
#
# Single line comments are written in a single line. Single line comments start with `#`
#
# ```Python
# # Here's a single line comment
# ```
# ## In-Line
#
# In-Line comments are written beside the code.
#
# ```Python
# print("Hello") # Here's a In-Line comment
# ```
# ## Multi Line
#
# Sometimes we need to write a huge explanation using comments, in those cases we do use multi-line comments. multiline comments are enclosed in `""" """` or `''' '''`
#
#
# ```Python
# """
# Here's a
# multiline comment.
# """
# ```
# ## Docstrings
# Docstrings are specific type of comments that are stored as a attribute to the module, class, method or function.
#
# Docstrings are written similar to the multi-line comments using `""" """` or `''' '''`, the only difference would be they are written exactly at the start(first statement) of the module, class, method or function.
# Docstrings can be programatically acccessed using the `__doc__` method or through the built-in function `help`. Let's give a try 😎.
def double_the_value(value: int):
"""Doubles the integer value passed to the function and returns it."""
return value * 2
# ### Using help
# `help` function provides the docstrings as well as the information about the module, class, method or function.
help(double_the_value)
# ### Using `__doc__`
print(double_the_value.__doc__)
# Can we use the single line comments instead of multi-line docstrings 🤔? Let's try this as well.
def test_single_line_comment_as_docstring():
# This is a single-line comment
pass
print(test_single_line_comment_as_docstring.__doc__)
# We can see that `None` is printed, which explains that we can't use single-line comments as docstrings 🙂
# ## Docstrings for documentation of code.
# [PEP-257](https://www.python.org/dev/peps/pep-0257/) defines two types of docstrings.
# * One-Line docstring
# * Multi-Line docstring
# ### [One-Line docstring](https://www.python.org/dev/peps/pep-0257/#one-line-docstrings)
# One-line docstrings are suited for short and simple Modules, classes, methods or functions.
def one_line_docstring():
"""This is a one-line docstring"""
pass
# ### [Multi-Line docstring](https://www.python.org/dev/peps/pep-0257/#multi-line-docstrings)
# Multi-line docstrings are suited for long, complex Modules, classes, methods or functions
def multi_line_docstring(arg1: int, arg2: str) -> None:
"""
This is a multi-line docstring.
Arguments:
arg1 (int): Argument 1 is an integer.
arg2 (str): Argument 2 is a string.
"""
pass
# ## Styles of docstrings
# There are multiple styles of writing docstrings such as [reStructuredText](https://docutils.sourceforge.io/rst.html), [Google Python Style Guide](https://google.github.io/styleguide/pyguide.html#38-comments-and-docstrings), [Numpy style](https://google.github.io/styleguide/pyguide.html#38-comments-and-docstrings).
#
# We could use any of the above docstrings style as long as we stay consistent.
#
# [Sphinx](https://www.sphinx-doc.org/en/master/) is a tool that generated beautiful HTML based documentation 📜 from the docstrings we provide in our code. reStructuredText is the default style, for other styles like Google Python style, numpy we could use plugins like [Napoleon](https://www.sphinx-doc.org/en/master/usage/extensions/napoleon.html).
#
# Sphinx also provides various templates we can choose from to create the HTML documentation out of it. 😎♥️
# ### A meme on Documentation 😂
# 
# It's always good and professional to have our code documented 🙂.
| Chapters_1-5/Chapter_3/1_Comments_and_docstrings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 13
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### Code from previous chapters
# `make_system`, `plot_results`, and `calc_total_infected` are unchanged.
def make_system(beta, gamma):
"""Make a system object for the SIR model.
beta: contact rate in days
gamma: recovery rate in days
returns: System object
"""
init = State(S=89, I=1, R=0)
init /= np.sum(init)
t0 = 0
t_end = 7 * 14
return System(init=init, t0=t0, t_end=t_end,
beta=beta, gamma=gamma)
def plot_results(S, I, R):
"""Plot the results of a SIR model.
S: TimeSeries
I: TimeSeries
R: TimeSeries
"""
plot(S, '--', label='Susceptible')
plot(I, '-', label='Infected')
plot(R, ':', label='Recovered')
decorate(xlabel='Time (days)',
ylabel='Fraction of population')
def calc_total_infected(results):
"""Fraction of population infected during the simulation.
results: DataFrame with columns S, I, R
returns: fraction of population
"""
return get_first_value(results.S) - get_last_value(results.S)
# Here's an updated version of `run_simulation` that uses `unpack`.
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: TimeFrame
"""
unpack(system)
frame = TimeFrame(columns=init.index)
frame.row[t0] = init
for t in linrange(t0, t_end):
frame.row[t+1] = update_func(frame.row[t], t, system)
return frame
# **Exercise:** Write a version of `update_func` that uses `unpack`.
# +
# Original
def update_func(state, t, system):
"""Update the SIR model.
state: State (s, i, r)
t: time
system: System object
returns: State (sir)
"""
s, i, r = state
infected = system.beta * i * s
recovered = system.gamma * i
s -= infected
i += infected - recovered
r += recovered
return State(S=s, I=i, R=r)
# -
# Solution goes here
def update_func(state, t, system):
"""Update the SIR model.
state: State (s, i, r)
t: time
system: System object
returns: State (sir)
"""
unpack (system)
s, i, r = state
infected = beta * i * s
recovered = gamma * i
s -= infected
i += infected - recovered
r += recovered
return State(S=s, I=i, R=r)
# Test the updated code with this example.
system = make_system(0.333, 0.25)
results = run_simulation(system, update_func)
results.head()
plot_results(results.S, results.I, results.R)
# ### Sweeping beta
# Make a range of values for `beta`, with constant `gamma`.
beta_array = linspace(0.1, 1.1, 11)
gamma = 0.25
# Run the simulation once for each value of `beta` and print total infections.
for beta in beta_array:
system = make_system(beta, gamma)
results = run_simulation(system, update_func)
print(system.beta, calc_total_infected(results))
# Wrap that loop in a function and return a `SweepSeries` object.
def sweep_beta(beta_array, gamma):
"""Sweep a range of values for beta.
beta_array: array of beta values
gamma: recovery rate
returns: SweepSeries that maps from beta to total infected
"""
sweep = SweepSeries()
for beta in beta_array:
system = make_system(beta, gamma)
results = run_simulation(system, update_func)
sweep[system.beta] = calc_total_infected(results)
return sweep
# Sweep `beta` and plot the results.
infected_sweep = sweep_beta(beta_array, gamma)
# +
label = 'gamma = ' + str(gamma)
plot(infected_sweep, label=label)
decorate(xlabel='Contacts per day (beta)',
ylabel='Fraction infected')
savefig('figs/chap06-fig01.pdf')
# -
# ### Sweeping gamma
# Using the same array of values for `beta`
beta_array
# And now an array of values for `gamma`
gamma_array = [0.2, 0.4, 0.6, 0.8]
# For each value of `gamma`, sweep `beta` and plot the results.
# +
for gamma in gamma_array:
infected_sweep = sweep_beta(beta_array, gamma)
label = 'γ = ' + str(gamma)
plot(infected_sweep, label=label)
decorate(xlabel='Contacts per day (beta)',
ylabel='Fraction infected',
loc='upper left')
savefig('figs/chap06-fig02.pdf')
# -
# ** Exercise:** Suppose the infectious period for the Freshman Plague is known to be 2 days on average, and suppose during one particularly bad year, 40% of the class is infected at some point. Estimate the time between contacts.
# +
# Solution goes here
# -
def slope_func(state, t, system):
"""Update the SIR model.
state: State (s, i, r)
t: time
system: System object
returns: State (sir)
"""
unpack (system)
s, i, r = state
dsdt = -1*beta * s * 1
didt = beta * i * s
drdt = gamma * i
return dsdt, didt, drdt
system = make_system(0.333, 0.25)
results, details=run_ode_solver(system, slope_func, max_step=2)
details
plot_results(results.S, results.I, results.R)
| code/chap13-Mine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Multidimensional GP
#
# Until now, our examples have been of 1-dimensional Gaussian processes, where there is just a single predictor variable thought to have a non-linear relationship to the outcome. Let's look at a real-world dataset that involves two predictors. We will use the famous **Walker Lake dataset (Isaaks & Srivistava 1989)** that involves spatial sampling of minerals and other variables over space. The data consist of two spatial coordinates and three measured outcomes. The outcomes are anonymously labeled as U, V (continuous variables, such as concentrarion) and T (discrete variable, such as the presence of a particular element).
# +
import os
os.environ["MKL_NUM_THREADS"] = "1"
os.environ["OPENBLAS_NUM_THREADS"] = "1"
# +
# %matplotlib inline
import warnings
import arviz as az
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
import seaborn as sns
warnings.simplefilter("ignore")
# -
walker_data = pd.read_table(
"../data/walker.txt",
sep=r"\s+",
index_col=0,
skiprows=8,
header=None,
names=["ID", "Xloc", "Yloc", "V", "U", "T"],
)
walker_data.head()
# The samples are taken regularly over a coarse grid across the entire area, and then irregularly over portions of the area, presumably where there were positive samples on the coarser grid.
# +
nx = 40
x1, x2 = np.meshgrid(np.linspace(0, 300, nx), np.linspace(0, 300, nx))
X = np.concatenate([x1.reshape(nx * nx, 1), x2.reshape(nx * nx, 1)], 1)
X_obs = walker_data[["Xloc", "Yloc"]].values
y_obs = walker_data.V.values
with sns.axes_style("white"):
plt.figure(figsize=(10, 8))
plt.scatter(
X_obs[:, 0], X_obs[:, 1], s=50, c=y_obs, marker="s", cmap=plt.cm.viridis
);
# -
# We need a sparse grid of inducing points:
nd = 15
xu1, xu2 = np.meshgrid(np.linspace(0, 300, nd), np.linspace(0, 300, nd))
Xu = np.concatenate([xu1.reshape(nd * nd, 1), xu2.reshape(nd * nd, 1)], 1)
with pm.Model() as spatial_model:
l = pm.HalfCauchy("l", beta=3, shape=(2,))
sf2 = pm.HalfCauchy("sf2", beta=3)
sn2 = pm.HalfCauchy("sn2", beta=3)
K = pm.gp.cov.ExpQuad(2, l) * sf2 ** 2
gp_spatial = pm.gp.MarginalSparse(cov_func=K, approx="FITC")
obs = gp_spatial.marginal_likelihood("obs", X=X_obs, Xu=Xu, y=y_obs, noise=sn2)
mp = pm.find_MAP()
nd = 30
z1, z2 = np.meshgrid(np.linspace(0, 300, nd), np.linspace(0, 300, nd))
Z = np.concatenate([z1.reshape(nd * nd, 1), z2.reshape(nd * nd, 1)], 1)
with spatial_model:
f_pred = gp_spatial.conditional("f_pred", Z)
samples = pm.sample_posterior_predictive([mp], vars=[f_pred], samples=100)
with sns.axes_style("white"):
plt.figure(figsize=(10, 8))
ax = sns.heatmap(samples["f_pred"].mean(0).reshape(nd, nd), cmap="viridis")
ax.invert_yaxis()
ax.set_yticklabels([])
ax.set_xticklabels([])
| notebooks/multidimensional_gp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# TC
from PyQuantum.TC.Cavity import Cavity
from PyQuantum.TC.Hamiltonian import Hamiltonian
from PyQuantum.TC.WaveFunction import WaveFunction
from PyQuantum.TC.Evolution import run_wf
# +
# Common
from PyQuantum.Common.LoadPackage import *
from PyQuantum.Common.STR import *
from PyQuantum.Common.Tools import mkdir
from PyQuantum.Common.Print import *
from PyQuantum.Common.PyPlot import PyPlot3D
# from shutil import copyfile
# from numpy.random import rand
# +
config = load_pkg("config", "PyQuantum/TC/config.py")
mkdir(config.path)
# copyfile("PyQuantum/TC/config.py", config.path + '/config.py')
# +
cavity = Cavity(wc=config.wc, wa=config.wa, g=config.g,
n_atoms=config.n_atoms, n_levels=config.n_levels)
print()
cavity.print()
# +
print("T:", config.T)
print("nt:", config.nt)
print("dt:", config.dt)
print()
# +
H = HamiltonianL(capacity=config.capacity,
cavity=cavity, RWA=False, reduced=True)
H.iprint()
H.df
# +
w_0 = WaveFunction(states=H.states, init_state=config.init_state)
w_0.print()
# -
run_wf(w_0=w_0, H=H, dt=config.dt, nt=config.nt,
config=config, fidelity_mode=True)
# +
from PyQuantum.TC.PlotBuilder3D import PlotBuilder3D
plt = PlotBuilder3D()
plt.set_width(950)
plt.set_height(800)
# ---------------------------------------------- TITLE --------------------------------------------
title = ""
if config.capacity - config.n_atoms > 0:
title += "<b>" + str(config.capacity - config.n) + \
" фотонов в полости" + "</b>" + "<br><br>"
else:
title += "<b>" + "empty cavity" + "</b>" + "<br><br>"
title += "<b>"
title += "n_atoms = " + str(config.n_atoms)
# title += "<br>atoms state: |Ψ<sub>0</sub> i = |11...1>A<sub>0</sub> |00...0>A<sub>1</sub> |vaki<sub>p</sub>" + \
# str(config.init_state)
title += "<br>"
title += "<br>w<sub>c</sub> = " + wc_str(config.wc)
title += "<br>w<sub>a</sub> = " + wa_str(config.wa)
title += "<br><br> g/hw<sub>c</sub> = " + str(config.g/config.wc)
title += "<br>"
title += "<br>"
title += "</b>"
plt.set_title(title)
# ---------------------------------------------- TITLE --------------------------------------------
# ---------------------------------------------- SCALE --------------------------------------------
y_scale = 1
if config.T < 0.25 * config.mks:
y_scale = 0.1
elif config.T <= 0.5 * config.mks:
y_scale = 0.025
elif config.T == 0.5 * config.mks:
y_scale = 0.01
elif config.T == 1 * config.mks:
y_scale = 7.5
# y_scale = 10
elif config.T == 5 * config.mks:
y_scale = 1
plt.set_yscale(y_scale)
plt.set_xaxis("states")
plt.set_yaxis("time, " + T_str_mark(config.T))
plt.set_zaxis("prob.\t\t\t\t\t\t.")
# ---------------------------------------------- SCALE --------------------------------------------
plt.iplot(
x_csv=config.x_csv,
y_csv=config.y_csv,
z_csv=config.z_csv,
)
| .ipynb_checkpoints/TC-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
data = pd.read_csv('new_data.csv', index_col='id')
cs_data = pd.DataFrame(data[data['730'] > 60]['730'])
cs_data.columns = ['time']
cs_data.reset_index(inplace=True)
import requests
def get_user_csgo_data(userid):
url = "http://peace-data-team.ru:4001/user/stats/csgo"
payload={}
headers = {
'steam_id': str(userid)
}
response = requests.request("GET", url, headers=headers, data=payload)
return response.text
cs_data['response'] = cs_data['id'].apply(get_user_csgo_data)
# +
import json
import numpy as np
for j in json.loads(cs_data['response'][1])['playerstats']['stats']:
cs_data[j['name']] = np.nan
# -
for i in range(len(cs_data)):
try:
for j in json.loads(cs_data['response'][i])['playerstats']['stats']:
cs_data[j['name']][i] = j['value']
except Exception:
continue
cs_data.to_csv('cs.csv')
cs_data = cs_data.drop('response',axis=1)
cs_data
for col in cs_data.columns:
if len(cs_data[cs_data[col].isnull()]) > 100:
cs_data = cs_data.drop(col, axis=1)
cs_data = cs_data[~cs_data['total_time_played'].isnull()]
cs_data = pd.read_csv('cs.csv', index_col=0)
cs_data=cs_data.drop('time',axis=1)
cs_data = cs_data.drop('Unnamed: 0', axis=1)
cs_data = cs_data.set_index('id', drop=True)
cs_data.to_csv('cs.csv')
cs_data = pd.read_csv('cs.csv', index_col=0)
cs_data
for col in cs_data.columns:
print(col, len(cs_data[cs_data[col].isnull()]))
cs_data['KD'] = cs_data['total_kills']/cs_data['total_deaths']
for col in cs_data.columns:
cs_data[cs_data[col].isnull()] = 0
cs_data
from sklearn.cluster import KMeans, DBSCAN, FeatureAgglomeration, Birch, MiniBatchKMeans
from sklearn.preprocessing import StandardScaler
cs_data
# +
from sklearn.manifold import TSNE
data_embedded = TSNE(n_components=2).fit_transform(cs_data[['total_kills', 'total_deaths', 'total_time_played','total_planted_bombs', 'KD']])
# -
import seaborn as sns
tmp = pd.DataFrame(data_embedded, columns=['x', 'y'])
sns.scatterplot(data=tmp, x='x', y='y')
cl_model = MiniBatchKMeans(3)
cl_model.fit(cs_data)
cl_model.labels_
cs_data['cluster'] = cl_model.labels_
import seaborn as sns
tmp = pd.DataFrame(data_embedded, columns=['x', 'y'])
sns.scatterplot(data=tmp, x='x', y='y', hue=cs_data.reset_index()['cluster'])
from sklearn.neighbors import NearestNeighbors
knn = NearestNeighbors(n_neighbors=20)
knn.fit(cs_data)
cs_data
# +
cols = ['total_kills', 'total_deaths', 'total_time_played','total_planted_bombs', 'KD']
def get_cs_team(user_id):
user_data = pd.DataFrame({'0': 0}, index=[0])
res = json.loads(get_user_csgo_data(user_id))
for j in res['playerstats']['stats']:
user_data[j['name']] = j['value']
user_data['KD'] = user_data['total_kills']/user_data['total_deaths']
try:
user_data = user_data[cols]
except Exception:
missed = list(set(cols) - set(user_data.columns))
user_data[missed] = 0
user_data = user_data[cols]
for col in user_data.columns:
user_data[user_data[col].isnull()] = 0
user_data['cluster'] = cl_model.predict(user_data)[0]
res = knn.kneighbors(user_data)[1][0][1:]
return json.dumbs(json.loads(pd.DataFrame(cs_data.iloc[res].index).to_json())['id'])
# -
get_cs_team(76561197972876528)
import pickle
pickle.dump(cl_model, open('cl_model.sav', 'wb'))
pickle.dump(knn, open('knn.sav', 'wb'))
cs_data.to_csv('cs.csv')
| ml/CS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="laa9tRjJ59bl"
# ##### Copyright 2020 The TensorFlow Hub Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="form" id="T4ZHtBpK6Dom"
#@title Copyright 2020 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="hk5u_9KN1m-t"
# <table class="tfo-notebook-buttons" align="left">
# <td> <a target="_blank" href="https://www.tensorflow.org/hub/tutorials/yamnet"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org で表示</a> </td>
# <td> <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/hub/tutorials/yamnet.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab で実行</a> </td>
# <td> <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/hub/tutorials/yamnet.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png"> GitHub でソースを表示</a> </td>
# <td> <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/hub/tutorials/yamnet.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード</a> </td>
# <td> <a href="https://tfhub.dev/google/yamnet/1"><img src="https://www.tensorflow.org/images/hub_logo_32px.png">TF Hub モデルを参照</a> </td>
# </table>
# + [markdown] id="x2ep-q7k_5R-"
# # YAMNet によるサウンドの分類
#
# YAMNet は、521 個のオーディオイベント[クラス](https://github.com/tensorflow/models/blob/master/research/audioset/yamnet/yamnet_class_map.csv)を、YAMNet がトレーニングに使用した [AudioSet-YouTube コーパス](http://g.co/audioset)から予測するディープネットです。[Mobilenet_v1](https://arxiv.org/pdf/1704.04861.pdf) という Depthwise-Separable Convolution(深さ方向に分離可能な畳み込み)アーキテクチャを使用しています。
# + id="Bteu7pfkpt_f"
import tensorflow as tf
import tensorflow_hub as hub
import numpy as np
import csv
import matplotlib.pyplot as plt
from IPython.display import Audio
from scipy.io import wavfile
# + [markdown] id="YSVs3zRrrYmY"
# TensorFlow Hub からモデルを読み込みます。
#
# 注意: ドキュメントを読むには、モデルの [url](https://tfhub.dev/google/yamnet/1) に従ってください。
# + id="VX8Vzs6EpwMo"
# Load the model.
model = hub.load('https://tfhub.dev/google/yamnet/1')
# + [markdown] id="lxWx6tOdtdBP"
# models アセットから labels ファイルが読み込まれます。これは `model.class_map_path()` にあります。`class_names` 変数で読み込みます。
# + id="EHSToAW--o4U"
# Find the name of the class with the top score when mean-aggregated across frames.
def class_names_from_csv(class_map_csv_text):
"""Returns list of class names corresponding to score vector."""
class_names = []
with tf.io.gfile.GFile(class_map_csv_text) as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
class_names.append(row['display_name'])
return class_names
class_map_path = model.class_map_path().numpy()
class_names = class_names_from_csv(class_map_path)
# + [markdown] id="mSFjRwkZ59lU"
# 読み込まれたオーディオが適切な sample_rate(16K)であることを確認して変換するメソッドを追加します。これがなければ、モデルの結果に影響があります。
# + id="LizGwWjc5w6A"
def ensure_sample_rate(original_sample_rate, waveform,
desired_sample_rate=16000):
"""Resample waveform if required."""
if original_sample_rate != desired_sample_rate:
desired_length = int(round(float(len(waveform)) /
original_sample_rate * desired_sample_rate))
waveform = scipy.signal.resample(waveform, desired_length)
return desired_sample_rate, waveform
# + [markdown] id="AZEgCobA9bWl"
# ## サウンドファイルのダウンロードと準備
#
# ここでは、wav ファイルをダウンロードして聴くことができるようにします。利用できるファイルがある場合は、Colab にアップロードしてそれを使用してください。
#
# 注意: 期待されるオーディオファイルは、サンプリングレートが 16kHz の mono wav ファイルである必要があります。
# + id="WzZHvyTtsJrc"
# !curl -O https://storage.googleapis.com/audioset/speech_whistling2.wav
# + id="D8LKmqvGzZzr"
# !curl -O https://storage.googleapis.com/audioset/miaow_16k.wav
# + id="Wo9KJb-5zuz1"
# wav_file_name = 'speech_whistling2.wav'
wav_file_name = 'miaow_16k.wav'
sample_rate, wav_data = wavfile.read(wav_file_name, 'rb')
sample_rate, wav_data = ensure_sample_rate(sample_rate, wav_data)
# Show some basic information about the audio.
duration = len(wav_data)/sample_rate
print(f'Sample rate: {sample_rate} Hz')
print(f'Total duration: {duration:.2f}s')
print(f'Size of the input: {len(wav_data)}')
# Listening to the wav file.
Audio(wav_data, rate=sample_rate)
# + [markdown] id="P9I290COsMBm"
# `wav_data` を、`[-1.0, 1.0]` の値に正規化する必要があります(モデルの[ドキュメント](https://tfhub.dev/google/yamnet/1)で指示されています)。
# + id="bKr78aCBsQo3"
waveform = wav_data / tf.int16.max
# + [markdown] id="e_Xwd4GPuMsB"
# ## モデルを実行する
#
# これは簡単なステップです。準備済みのデータを使用して、モデルを呼び出し、スコア、埋め込み、およびスペクトログラムを取得します。
#
# 使用するメインの結果は、スコアです。スペクトログラムについては、後で視覚化を行うために使用します。
# + id="BJGP6r-At_Jc"
# Run the model, check the output.
scores, embeddings, spectrogram = model(waveform)
# + id="Vmo7griQprDk"
scores_np = scores.numpy()
spectrogram_np = spectrogram.numpy()
infered_class = class_names[scores_np.mean(axis=0).argmax()]
print(f'The main sound is: {infered_class}')
# + [markdown] id="Uj2xLf-P_ndS"
# ## 視覚化
#
# YAMNet は、視覚化に使用できる追加情報も返します。波形、スペクトログラム、および推論された上位クラスを確認してみましょう。
# + id="_QSTkmv7wr2M"
plt.figure(figsize=(10, 6))
# Plot the waveform.
plt.subplot(3, 1, 1)
plt.plot(waveform)
plt.xlim([0, len(waveform)])
# Plot the log-mel spectrogram (returned by the model).
plt.subplot(3, 1, 2)
plt.imshow(spectrogram_np.T, aspect='auto', interpolation='nearest', origin='lower')
# Plot and label the model output scores for the top-scoring classes.
mean_scores = np.mean(scores, axis=0)
top_n = 10
top_class_indices = np.argsort(mean_scores)[::-1][:top_n]
plt.subplot(3, 1, 3)
plt.imshow(scores_np[:, top_class_indices].T, aspect='auto', interpolation='nearest', cmap='gray_r')
# patch_padding = (PATCH_WINDOW_SECONDS / 2) / PATCH_HOP_SECONDS
# values from the model documentation
patch_padding = (0.025 / 2) / 0.01
plt.xlim([-patch_padding-0.5, scores.shape[0] + patch_padding-0.5])
# Label the top_N classes.
yticks = range(0, top_n, 1)
plt.yticks(yticks, [class_names[top_class_indices[x]] for x in yticks])
_ = plt.ylim(-0.5 + np.array([top_n, 0]))
| site/ja/hub/tutorials/yamnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from time import time
from main import main
from main_multi import main_multi
from evolution.chromosome import *
from utils.data_manipulators import *
from problems.knapsack_generator import knapsack_generator
# %matplotlib notebook
# +
def get_fitness(results):
fitnesses = np.zeros_like(results)
for i, rep in enumerate(results):
for j, gen in enumerate(rep):
if gen.any() is not None:
fitnesses[i, j, :] = Chromosome.fitness_to_numpy(gen)
return fitnesses
def time_concat(time_hist):
for i in range(1, len(time_hist)):
time_hist[i] += time_hist[i-1]
return time_hist
# -
class args:
tsamples = 10
src_version = 'v3'
stop_condition = True
reps = 0
transfer = True
delta = 2
buildmodel = True
s1_psize = 50
s2_psize = 1
sample_size = 50
sub_sample_size = 50
version = 'ea_time_scale'
mutation_strength = 1
injection_type = 'full'
to_repititon_num = 10
selection_version = 'v1'
c = np.sqrt(1.5)
efficient_version = True
now = time()
results_v1 = main(args)
end = time()
print("duration: ", str((end - now)/60))
now = time()
results_v1 = main_multi(args)
end = time()
print("duration: ", str((end - now)/60))
# +
# now = time()
# results_v2_selv1_tor10_efficient = main(args)
# end = time()
# print("duration: ", str((end - now)/60))
# +
# args.efficient_version = False
# now = time()
# results_v2_selv1_tor10 = main(args)
# end = time()
# print("duration: ", str((end - now)/60))
# -
args.version = 'ea_time_scale'
now = time()
results_ea = main(args)
end = time()
print("duration: ", str((end - now)/60))
args.version = 'to'
now = time()
results_to = main(args)
end = time()
print("duration: ", str((end - now)/60))
Tools.save_to_file('data/to_results', results_to)
Tools.save_to_file('data/ea_results', results_ea)
v1_results = results_v1[]
results_v1
results_ea = Tools.load_from_file('data/ea_results')
results_to = Tools.load_from_file('data/to_results')
v1_results = Tools.load_from_file('data/results_v1')
v1_asynch_results = Tools.load_from_file('data/results_v1_async')
# fitness_v2_selv1_tor10 = get_fitness(results_v2_selv1_tor10[0]).mean(axis=0).mean(axis=1)
# fitness_v2_selv1_tor10_efficient = get_fitness(results_v2_selv1_tor10_efficient[0]).mean(axis=0).mean(axis=1)
fitness_v1_async = np.array([result[1] for result in v1_asynch_results]).mean(axis=0).mean(axis=1)
fitness_v1 = np.array([result[1] for result in v1_results]).mean(axis=0).mean(axis=1)
fitness_ea = results_ea[0].mean(axis=0).mean(axis=1)
fitness_to = results_to[0].mean(axis=0).mean(axis=1)
# time_v2_selv1_tor10 = time_concat(results_v2_selv1_tor10[3].mean(axis=0).mean(axis=1))
# time_v2_selv1_tor10_efficient = time_concat(results_v2_selv1_tor10_efficient[3].mean(axis=0).mean(axis=1))
time_v1_async = time_concat(np.array([result[0] for result in v1_asynch_results]).mean(axis=0))
time_v1 = time_concat(np.array([result[0] for result in v1_results]).mean(axis=0))
time_ea = time_concat(results_ea[1].mean(axis=0))
time_to = time_concat(results_to[2].mean(axis=0))
plt.plot(time_v1, fitness_v1, '#aa0a0a', label='our idea')
plt.plot(time_v1_async, fitness_v1_async, '#a000fa', label='our idea Async')
# plt.plot(time_v2_selv1_tor10, fitness_v2_selv1_tor10, '#bcabcc', label='our idea not efficient 10 pop_s2')
plt.plot(time_to, fitness_to,label='transfer idea (delta=2)')
plt.plot(time_ea, fitness_ea,label='ea idea')
plt.legend()
plt.xlabel('Time (second)')
plt.ylabel('fitness')
plt.title("Average of population's fitness during 30 repetition of Algorithm")
plt.show()
plt.plot(range(100), fitness_v1, '#aa0a0a', label='our idea')
plt.plot(range(100), fitness_v1_async, '#a000fa', label='our idea Async')
# plt.plot(time_v2_selv1_tor10, fitness_v2_selv1_tor10, '#bcabcc', label='our idea not efficient 10 pop_s2')
plt.plot(range(100), fitness_to,label='transfer idea (delta=2)')
plt.plot(range(100), fitness_ea,label='ea idea')
plt.legend()
plt.xlabel('Time (second)')
plt.ylabel('fitness')
plt.title("Average of population's fitness during 30 repetition of Algorithm")
plt.show()
s
| Low Computation Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
# $$
# \newcommand\bs[1]{\boldsymbol{#1}}
# $$
# # Introduction
#
# We will explain how to create and use vectors and matrices through examples.
# # Scalars, Vectors, Matrices and Tensors
#
# Let's start with some basic definitions:
#
# <img src="images/scalar-vector-matrix-tensor.png" width="400" alt="An example of a scalar, a vector, a matrix and a tensor" title="Difference between a scalar, a vector, a matrix and a tensor">
# <em>Difference between a scalar, a vector, a matrix and a tensor</em>
#
# - A scalar is a single number or a matrix with a single entry.
# - A vector is a 1-d array of numbers. Another way to think of vectors is identifying points in space with each element giving the coordinate along a different axis.
#
# $$
# \bs{x} =\begin{bmatrix}
# x_1 \\\\
# x_2 \\\\
# \cdots \\\\
# x_n
# \end{bmatrix}
# $$
#
# - A matrix is a 2-D array where each element is identified by two indices (ROW then COLUMN).
#
# $$
# \bs{A}=
# \begin{bmatrix}
# A_{1,1} & A_{1,2} & \cdots & A_{1,n} \\\\
# A_{2,1} & A_{2,2} & \cdots & A_{2,n} \\\\
# \cdots & \cdots & \cdots & \cdots \\\\
# A_{m,1} & A_{m,2} & \cdots & A_{m,n}
# \end{bmatrix}
# $$
#
# - A tensor is a $n$-dimensional array with $n>2$
#
# ## We will follow the following conventions :
# - `scalars` are written in `lowercase and italics`. For instance: $n$
# - `vectors` are written in `lowercase, italics and bold type`. For instance: $\bs{x}$
# - `matrices` are written in `uppercase, italics and bold`. For instance: $\bs{X}$
# ### Example 1.
#
# #### Create a vector with Python and Numpy
#
# *Coding tip*: Unlike the `matrix()` function which necessarily creates $2$-dimensional matrices, you can create $n$-dimensionnal arrays with the `array()` function. The main advantage to use `matrix()` is the useful methods (conjugate transpose, inverse, matrix operations...). We will use the `array()` function in this series.
#
# We will start by creating a vector. This is just a $1$-dimensional array:
x = np.array([1, 2, 3, 4])
x
# ### Example 2.
#
# #### Create a (3x2) matrix with nested brackets
#
# The `array()` function can also create $2$-dimensional arrays with nested brackets:
A = np.array([[1, 2], [3, 4], [5, 6]])
A
# ### Shape
#
# The shape of an array (that is to say its dimensions) tells you the number of values for each dimension. For a $2$-dimensional array it will give you the number of rows and the number of columns. Let's find the shape of our preceding $2$-dimensional array `A`. Since `A` is a Numpy array (it was created with the `array()` function) you can access its shape with:
A.shape
# We can see that $\bs{A}$ has 3 rows and 2 columns.
#
# Let's check the shape of our first vector:
x.shape
# As expected, you can see that $\bs{x}$ has only one dimension. The number corresponds to the length of the array:
len(x)
# # Transposition
#
# With transposition you can convert a row vector to a column vector and vice versa:
#
# <img src="images/vector-transposition.png" alt="Transposition of a vector" title="Vector transposition" width="200">
# <em>Vector transposition</em>
#
# The transpose $\bs{A}^{\text{T}}$ of the matrix $\bs{A}$ corresponds to the mirrored axes. If the matrix is a square matrix (same number of columns and rows):
#
# <img src="images/square-matrix-transposition.png" alt="Transposition of a square matrix" title="Square matrix transposition" width="300">
# <em>Square matrix transposition</em>
#
# If the matrix is not square the idea is the same:
#
# <img src="images/non-squared-matrix-transposition.png" alt="Transposition of a square matrix" title="Non square matrix transposition" width="300">
# <em>Non-square matrix transposition</em>
#
#
# The superscript $^\text{T}$ is used for transposed matrices.
#
# $$
# \bs{A}=
# \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}
# $$
#
# $$
# \bs{A}^{\text{T}}=
# \begin{bmatrix}
# A_{1,1} & A_{2,1} & A_{3,1} \\\\
# A_{1,2} & A_{2,2} & A_{3,2}
# \end{bmatrix}
# $$
#
# The shape ($m \times n$) is inverted and becomes ($n \times m$).
#
# <img src="images/dimensions-transposition-matrix.png" alt="Dimensions of matrix transposition" title="Dimensions of matrix transposition" width="300">
# <em>Dimensions of matrix transposition</em>
# ### Example 3.
#
# #### Create a matrix A and transpose it
A = np.array([[1, 2], [3, 4], [5, 6]])
A
A_t = A.T
A_t
# We can check the dimensions of the matrices:
A.shape
A_t.shape
# We can see that the number of columns becomes the number of rows with transposition and vice versa.
# # Addition
#
# <img src="images/matrix-addition.png" alt="Addition of two matrices" title="Addition of two matrices" width="300">
# <em>Addition of two matrices</em>
#
# Matrices can be added if they have the same shape:
#
# $$\bs{A} + \bs{B} = \bs{C}$$
#
# Each cell of $\bs{A}$ is added to the corresponding cell of $\bs{B}$:
#
# $$\bs{A}_{i,j} + \bs{B}_{i,j} = \bs{C}_{i,j}$$
#
# $i$ is the row index and $j$ the column index.
#
# $$
# \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}+
# \begin{bmatrix}
# B_{1,1} & B_{1,2} \\\\
# B_{2,1} & B_{2,2} \\\\
# B_{3,1} & B_{3,2}
# \end{bmatrix}=
# \begin{bmatrix}
# A_{1,1} + B_{1,1} & A_{1,2} + B_{1,2} \\\\
# A_{2,1} + B_{2,1} & A_{2,2} + B_{2,2} \\\\
# A_{3,1} + B_{3,1} & A_{3,2} + B_{3,2}
# \end{bmatrix}
# $$
#
# The shape of $\bs{A}$, $\bs{B}$ and $\bs{C}$ are identical. Let's check that in an example:
# ### Example 4.
#
# #### Create two matrices A and B and add them
#
# With Numpy you can add matrices just as you would add vectors or scalars.
A = np.array([[1, 2], [3, 4], [5, 6]])
A
B = np.array([[2, 5], [7, 4], [4, 3]])
B
# Add matrices A and B
C = A + B
C
# It is also possible to add a scalar to a matrix. This means adding this scalar to each cell of the matrix.
#
# $$
# \alpha+ \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}=
# \begin{bmatrix}
# \alpha + A_{1,1} & \alpha + A_{1,2} \\\\
# \alpha + A_{2,1} & \alpha + A_{2,2} \\\\
# \alpha + A_{3,1} & \alpha + A_{3,2}
# \end{bmatrix}
# $$
# ### Example 5.
#
# #### Add a scalar to a matrix
A
# Exemple: Add 4 to the matrix A
C = A+4
C
# # Broadcasting
#
# Numpy can handle operations on arrays of different shapes. The smaller array will be extended to match the shape of the bigger one. The advantage is that this is done in `C` under the hood (like any vectorized operations in Numpy). Actually, we used broadcasting in the example 5. The scalar was converted in an array of same shape as $\bs{A}$.
#
# Here is another generic example:
#
# $$
# \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}+
# \begin{bmatrix}
# B_{1,1} \\\\
# B_{2,1} \\\\
# B_{3,1}
# \end{bmatrix}
# $$
#
# is equivalent to
#
# $$
# \begin{bmatrix}
# A_{1,1} & A_{1,2} \\\\
# A_{2,1} & A_{2,2} \\\\
# A_{3,1} & A_{3,2}
# \end{bmatrix}+
# \begin{bmatrix}
# B_{1,1} & B_{1,1} \\\\
# B_{2,1} & B_{2,1} \\\\
# B_{3,1} & B_{3,1}
# \end{bmatrix}=
# \begin{bmatrix}
# A_{1,1} + B_{1,1} & A_{1,2} + B_{1,1} \\\\
# A_{2,1} + B_{2,1} & A_{2,2} + B_{2,1} \\\\
# A_{3,1} + B_{3,1} & A_{3,2} + B_{3,1}
# \end{bmatrix}
# $$
#
# where the ($3 \times 1$) matrix is converted to the right shape ($3 \times 2$) by copying the first column. Numpy will do that automatically if the shapes can match.
# ### Example 6.
#
# #### Add two matrices of different shapes
A = np.array([[1, 2], [3, 4], [5, 6]])
A
B = np.array([[2], [4], [6]])
B
# Broadcasting
C=A+B
C
# *Coding tip*: Sometimes row or column vectors are not in proper shape for broadcasting. We need to imploy a trick ( a`numpy.newaxis` object) to help fix this issue.
x = np.arange(4)
x.shape
# Adds a new dimension
x[:, np.newaxis]
A = np.random.randn(4,3)
A
# This will throw an error
try:
A - x
except ValueError:
print("Operation cannot be completed. Dimension mismatch")
# But this works -- subtract each column of A by the column vector x
A - x[:, np.newaxis]
| Python/3. Computational Sciences and Mathematics/Linear Algebra/Excellent Topics-Specific Examples and Visualizations - Jupyters/Scalars, Vectors, Matrices, and Tensors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + colab={} colab_type="code" id="gE76T8J7IsGC"
#@title Copyright 2019 The Lifetime Value Authors.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ============================================================================
# + [markdown] colab_type="text" id="sswTFWDv7HZd"
# # KDD Cup 98 LTV Prediction
# + [markdown] colab_type="text" id="PSr1mSJP7O1J"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/google/lifetime_value/blob/master/notebooks/kdd_cup_98/regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/google/lifetime_value/blob/master/notebooks/kdd_cup_98/regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + colab={} colab_type="code" id="pBXE3Dz3NI4A"
import os
import numpy as np
import pandas as pd
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
import tensorflow_probability as tfp
from typing import Sequence
# install and import ltv
# !pip install -q git+https://github.com/google/lifetime_value
import lifetime_value as ltv
# + colab={} colab_type="code" id="Bq0Ah16lBmgV"
tfd = tfp.distributions
tf.enable_eager_execution()
# %config InlineBackend.figure_format='retina'
sns.set_style('whitegrid')
# + [markdown] colab_type="text" id="2qN319qZK3IG"
# ## Configs
# + colab={} colab_type="code" id="hNy_ybw_K19n"
LOSS = 'ziln' # @param { isTemplate: true, type: 'string'} ['mse', 'ziln']
LEARNING_RATE = 0.001 # @param { isTemplate: true}
VERSION = 0 # @param { isTemplate: true, type: 'integer'}
OUTPUT_CSV_FOLDER = '/tmp/lifetime-value/kdd_cup_98/result' # @param { isTemplate: true, type: 'string'}
# + [markdown] colab_type="text" id="mDSR921CCEcL"
# ## Load data
# + [markdown] colab_type="text" id="lHxp4rOGI02Q"
# Download kdd_cup_98 data to /tmp/lifetime-value/kdd_cup_98
# + colab={} colab_type="code" id="Dg3qtgJyJpdi"
# %%shell
# mkdir -p /tmp/lifetime-value/kdd_cup_98
wget https://kdd.ics.uci.edu/databases/kddcup98/epsilon_mirror/cup98lrn.zip -P /tmp/lifetime-value/kdd_cup_98/
wget https://kdd.ics.uci.edu/databases/kddcup98/epsilon_mirror/cup98val.zip -P /tmp/lifetime-value/kdd_cup_98/
wget https://kdd.ics.uci.edu/databases/kddcup98/epsilon_mirror/valtargt.txt -P /tmp/lifetime-value/kdd_cup_98/
# cd /tmp/lifetime-value/kdd_cup_98/
unzip cup98lrn.zip
unzip cup98val.zip
# + colab={} colab_type="code" id="a_LnLmQQRlYF"
df_train = pd.read_csv('/tmp/lifetime-value/kdd_cup_98/cup98LRN.txt')
num_train = df_train.shape[0]
df_eval = pd.read_csv('/tmp/lifetime-value/kdd_cup_98/cup98VAL.txt')
df_eval_target = pd.read_csv('/tmp/lifetime-value/kdd_cup_98/valtargt.txt')
df_eval = df_eval.merge(df_eval_target, on='CONTROLN')
# + colab={} colab_type="code" id="ggQmy9wiP5M6"
df = pd.concat([df_train, df_eval], axis=0, sort=True)
# + [markdown] colab_type="text" id="0rgxHpIyjaMH"
# ## Label distribution
# + colab={} colab_type="code" id="Xmpu_d3YjcFC"
y = df['TARGET_D'][:num_train]
# + colab={} colab_type="code" id="yMr2EDRyK5Sb"
def plot_hist_log_scale(y):
max_val = y.max() + 1.
ax = pd.Series(y).hist(
figsize=(8, 5), bins = 10 ** np.linspace(0., np.log10(max_val), 20))
plt.xlabel('Donation ($)')
plt.ylabel('Count')
# plt.title('Histogram of LTV')
plt.xticks(rotation='horizontal')
plt.legend(loc='upper left')
ax.set_xscale('log')
ax.grid(False)
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
plt.show()
fig = ax.get_figure()
output_file = tf.io.gfile.GFile(
'/tmp/lifetime-value/kdd_cup_98/histogram_kdd98_log_scale.pdf',
'wb')
fig.savefig(output_file, bbox_inches='tight', format='pdf')
# + colab={"height": 355} colab_type="code" executionInfo={"elapsed": 371, "status": "ok", "timestamp": 1575669899758, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="KbwCzGkBOWhH" outputId="c9de8cf1-0d2e-476a-9361-b755257baa20"
plot_hist_log_scale(y[y>0])
# + [markdown] colab_type="text" id="1XXMLbnlCdlN"
# ## Preprocess features
# + [markdown] colab_type="text" id="L1sBf_RSU3pR"
# ### Vocab
# + colab={} colab_type="code" id="xB_ddsd_U_4e"
VOCAB_FEATURES = [
'ODATEDW', # date of donor's first gift (YYMM)
'OSOURCE', # donor acquisition mailing list
'TCODE', # donor title code
'STATE',
'ZIP',
'DOMAIN', # urbanicity level and socio-economic status of the neighborhood
'CLUSTER', # socio-economic status
'GENDER',
'MAXADATE', # date of the most recent promotion received
'MINRDATE',
'LASTDATE',
'FISTDATE',
'RFA_2A',
]
# + colab={} colab_type="code" id="f2oPZGVLRSPe"
df['ODATEDW'] = df['ODATEDW'].astype('str')
df['TCODE'] = df['TCODE'].apply(
lambda x: '{:03d}'.format(x // 1000 if x > 1000 else x))
df['ZIP'] = df['ZIP'].str.slice(0, 5)
df['MAXADATE'] = df['MAXADATE'].astype('str')
df['MINRDATE'] = df['MINRDATE'].astype('str')
df['LASTDATE'] = df['LASTDATE'].astype('str')
df['FISTDATE'] = df['FISTDATE'].astype('str')
# + colab={} colab_type="code" id="isL9Ofv9JLAP"
def label_encoding(y, frequency_threshold=100):
value_counts = pd.value_counts(y)
categories = value_counts[
value_counts >= frequency_threshold].index.to_numpy()
# 0 indicates the unknown category.
return pd.Categorical(y, categories=categories).codes + 1
# + colab={} colab_type="code" id="BgXGO5D0OdJP"
for key in VOCAB_FEATURES:
df[key] = label_encoding(df[key])
# + [markdown] colab_type="text" id="kZkmnJ93Zrjw"
# ### Indicator
# + colab={} colab_type="code" id="tGBpMfaGhCD0"
MAIL_ORDER_RESPONSES = [
'MBCRAFT',
'MBGARDEN',
'MBBOOKS',
'MBCOLECT',
'MAGFAML',
'MAGFEM',
'MAGMALE',
'PUBGARDN',
'PUBCULIN',
'PUBHLTH',
'PUBDOITY',
'PUBNEWFN',
'PUBPHOTO',
'PUBOPP',
'RFA_2F',
]
# + colab={} colab_type="code" id="4V-DeOZFZhjB"
INDICATOR_FEATURES = [
'AGE', # age decile, 0 indicates unknown
'NUMCHLD',
'INCOME',
'WEALTH1',
'HIT',
] + MAIL_ORDER_RESPONSES
# + colab={} colab_type="code" id="U9y5qA1vZ0kz"
df['AGE'] = pd.qcut(df['AGE'].values, 10).codes + 1
df['NUMCHLD'] = df['NUMCHLD'].apply(lambda x: 0 if np.isnan(x) else int(x))
df['INCOME'] = df['INCOME'].apply(lambda x: 0 if np.isnan(x) else int(x))
df['WEALTH1'] = df['WEALTH1'].apply(lambda x: 0 if np.isnan(x) else int(x) + 1)
df['HIT'] = pd.qcut(df['HIT'].values, q=50, duplicates='drop').codes
for col in MAIL_ORDER_RESPONSES:
df[col] = pd.qcut(df[col].values, q=20, duplicates='drop').codes + 1
# + [markdown] colab_type="text" id="8DOO_2a-U6gr"
# ### Numeric
# + colab={} colab_type="code" id="rqVteSLDiLVr"
NUMERIC_FEATURES = [
# binary
'MAILCODE', # bad address
'NOEXCH', # do not exchange
'RECINHSE', # donor has given to PVA's in house program
'RECP3', # donor has given to PVA's P3 program
'RECPGVG', # planned giving record
'RECSWEEP', # sweepstakes record
'HOMEOWNR', # home owner
'CHILD03',
'CHILD07',
'CHILD12',
'CHILD18',
# continuous
'CARDPROM',
'NUMPROM',
'CARDPM12',
'NUMPRM12',
'RAMNTALL',
'NGIFTALL',
'MINRAMNT',
'MAXRAMNT',
'LASTGIFT',
'AVGGIFT',
]
# + colab={} colab_type="code" id="xMRP05Ztic0A"
df['MAILCODE'] = (df['MAILCODE'] == 'B').astype('float32')
df['PVASTATE'] = df['PVASTATE'].isin(['P', 'E']).astype('float32')
df['NOEXCH'] = df['NOEXCH'].isin(['X', '1']).astype('float32')
df['RECINHSE'] = (df['RECINHSE'] == 'X').astype('float32')
df['RECP3'] = (df['RECP3'] == 'X').astype('float32')
df['RECPGVG'] = (df['RECPGVG'] == 'X').astype('float32')
df['RECSWEEP'] = (df['RECSWEEP'] == 'X').astype('float32')
df['HOMEOWNR'] = (df['HOMEOWNR'] == 'H').astype('float32')
df['CHILD03'] = df['CHILD03'].isin(['M', 'F', 'B']).astype('float32')
df['CHILD07'] = df['CHILD07'].isin(['M', 'F', 'B']).astype('float32')
df['CHILD12'] = df['CHILD12'].isin(['M', 'F', 'B']).astype('float32')
df['CHILD18'] = df['CHILD18'].isin(['M', 'F', 'B']).astype('float32')
df['CARDPROM'] = df['CARDPROM'] / 100
df['NUMPROM'] = df['NUMPROM'] / 100
df['CARDPM12'] = df['CARDPM12'] / 100
df['NUMPRM12'] = df['NUMPRM12'] / 100
df['RAMNTALL'] = np.log1p(df['RAMNTALL'])
df['NGIFTALL'] = np.log1p(df['NGIFTALL'])
df['MINRAMNT'] = np.log1p(df['MINRAMNT'])
df['MAXRAMNT'] = np.log1p(df['MAXRAMNT'])
df['LASTGIFT'] = np.log1p(df['LASTGIFT'])
df['AVGGIFT'] = np.log1p(df['AVGGIFT'])
# + [markdown] colab_type="text" id="GoLg1PvWuCT_"
# ### All
# + colab={} colab_type="code" id="lSnNgjBCuJdb"
CATEGORICAL_FEATURES = VOCAB_FEATURES + INDICATOR_FEATURES
ALL_FEATURES = CATEGORICAL_FEATURES + NUMERIC_FEATURES
# + [markdown] colab_type="text" id="8HJBvvCxRPg3"
# ## Train/eval split
# + colab={} colab_type="code" id="N7BXLB1eHovl"
def dnn_split(df):
df_train = df.iloc[:num_train]
df_eval = df.iloc[num_train:]
def feature_dict(df):
features = {k: v.values for k, v in dict(df[CATEGORICAL_FEATURES]).items()}
features['numeric'] = df[NUMERIC_FEATURES].astype('float32').values
return features
x_train, y_train = feature_dict(df_train), df_train['TARGET_D'].astype(
'float32').values
x_eval, y_eval = feature_dict(df_eval), df_eval['TARGET_D'].astype(
'float32').values
return x_train, x_eval, y_train, y_eval
# + [markdown] colab_type="text" id="4yw6fekBtX7X"
# ## Model
# + colab={} colab_type="code" id="_rIuO0XYtZH2"
def embedding_dim(x):
return int(x**.25) + 1
def embedding_layer(vocab_size):
return tf.keras.Sequential([
tf.keras.layers.Embedding(
input_dim=vocab_size,
output_dim=embedding_dim(vocab_size),
input_length=1),
tf.keras.layers.Flatten(),
])
def dnn_model(output_units):
numeric_input = tf.keras.layers.Input(
shape=(len(NUMERIC_FEATURES),), name='numeric')
embedding_inputs = [
tf.keras.layers.Input(shape=(1,), name=key, dtype=np.int64)
for key in CATEGORICAL_FEATURES
]
embedding_outputs = [
embedding_layer(vocab_size=df[key].max() + 1)(input)
for key, input in zip(CATEGORICAL_FEATURES, embedding_inputs)
]
deep_input = tf.keras.layers.concatenate([numeric_input] + embedding_outputs)
deep_model = tf.keras.Sequential([
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(units=output_units),
])
return tf.keras.Model(
inputs=[numeric_input] + embedding_inputs, outputs=deep_model(deep_input))
# + [markdown] colab_type="text" id="G5h7X6botcHl"
# ## Loss
# + colab={} colab_type="code" id="iJ9gpkC6tgP0"
if LOSS == 'mse':
loss = tf.keras.losses.MeanSquaredError()
output_units = 1
if LOSS == 'ziln':
loss = ltv.zero_inflated_lognormal_loss
output_units = 3
# + colab={} colab_type="code" id="_afFfIritjCM"
x_train, x_eval, y_train, y_eval = dnn_split(df)
model = dnn_model(output_units)
# + colab={} colab_type="code" id="Qj3kI7pyVwzO"
model.compile(optimizer=tf.keras.optimizers.Nadam(lr=LEARNING_RATE), loss=loss)
# + [markdown] colab_type="text" id="KZSYxgWdwiXC"
# ## Train
# + colab={} colab_type="code" id="Nwj9h5ysQDLp"
callbacks = [
tf.keras.callbacks.ReduceLROnPlateau(monitor='val_loss', min_lr=1e-6),
tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=10),
]
# + colab={"height": 672} colab_type="code" executionInfo={"elapsed": 25814, "status": "ok", "timestamp": 1574346431087, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="Vb5Tnld6hsfx" outputId="b102f5e8-e550-4fd7-9e75-5f6e43199cef"
history = model.fit(
x=x_train,
y=y_train,
batch_size=2048,
epochs=200,
verbose=2,
callbacks=callbacks,
validation_data=(x_eval, y_eval)).history
# + colab={"height": 279} colab_type="code" executionInfo={"elapsed": 145, "status": "ok", "timestamp": 1574346431309, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="J1sLSUdgvfa6" outputId="57feb0a2-a7b7-470d-f06f-ce74c8945ccd"
pd.DataFrame(history)[['loss', 'val_loss']].plot();
# + [markdown] colab_type="text" id="jRKuZBqhvhT9"
# ## Eval
# + colab={} colab_type="code" id="q9_zNMd3vjNk"
if LOSS == 'mse':
y_pred = model.predict(x=x_eval, batch_size=1024).flatten()
if LOSS == 'ziln':
logits = model.predict(x=x_eval, batch_size=1024)
y_pred = ltv.zero_inflated_lognormal_pred(logits).numpy().flatten()
# + [markdown] colab_type="text" id="SkfkUMUvUu_E"
# ### Total Profit
# + colab={} colab_type="code" id="AwfWAp8WQuns"
unit_costs = [0.4, 0.5, 0.6, 0.68, 0.7, 0.8, 0.9, 1.0]
# + colab={"height": 34} colab_type="code" executionInfo={"elapsed": 49, "status": "ok", "timestamp": 1574346432560, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="zqi91dfCUxpx" outputId="8085071c-7ce8-4dcc-8b56-33eddc4ecfac"
num_mailed = [np.sum(y_pred > v) for v in unit_costs]
num_mailed
# + colab={"height": 34} colab_type="code" executionInfo={"elapsed": 55, "status": "ok", "timestamp": 1574346432670, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="ZgFjZUcuhScv" outputId="b101ec33-e695-4591-dc04-b977baa51a44"
baseline_total_profit = np.sum(y_eval - 0.68)
baseline_total_profit
# + colab={"height": 151} colab_type="code" executionInfo={"elapsed": 80, "status": "ok", "timestamp": 1574346432799, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="VwsFnin5U-R9" outputId="b1f2727d-79f5-4f9a-a6ce-a09b650b3b18"
total_profits = [np.sum(y_eval[y_pred > v] - v) for v in unit_costs]
total_profits
# + [markdown] colab_type="text" id="zROhsEWxnA5u"
# ### Gini Coefficient
# + colab={} colab_type="code" id="gRsJ7y-632h_"
gain = pd.DataFrame({
'lorenz': ltv.cumulative_true(y_eval, y_eval),
'baseline': ltv.cumulative_true(y_eval, x_eval['numeric'][:, 19]),
'model': ltv.cumulative_true(y_eval, y_pred),
})
# + colab={} colab_type="code" id="yg-ndbve4AL_"
num_customers = np.float32(gain.shape[0])
gain['cumulative_customer'] = (np.arange(num_customers) + 1.) / num_customers
# + colab={"height": 361} colab_type="code" executionInfo={"elapsed": 246, "status": "ok", "timestamp": 1574346433299, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="WEoAvuCj4OVy" outputId="031558ef-8a9d-48d8-980e-62fd2dda0017"
ax = gain[[
'cumulative_customer',
'lorenz',
'baseline',
'model',
]].plot(
x='cumulative_customer', figsize=(8, 5), legend=True)
ax.legend(['Groundtruth', 'Baseline', 'Model'], loc='lower right')
ax.set_xlabel('Cumulative Fraction of Customers')
ax.set_xticks(np.arange(0, 1.1, 0.1))
ax.set_xlim((0, 1.))
ax.set_ylabel('Cumulative Fraction of Total Lifetime Value')
ax.set_yticks(np.arange(0, 1.1, 0.1))
ax.set_ylim((0, 1.05))
ax.set_title('Gain Chart');
# + colab={"height": 136} colab_type="code" executionInfo={"elapsed": 630, "status": "ok", "timestamp": 1574346434025, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="kzPqaiNO4iWC" outputId="2fa8db69-ebb8-4681-9e54-78c98a81708c"
gini = ltv.gini_from_gain(gain[['lorenz', 'baseline', 'model']])
gini
# + [markdown] colab_type="text" id="S84RitIa9PBu"
# ### Calibration
# + colab={"height": 373} colab_type="code" executionInfo={"elapsed": 79, "status": "ok", "timestamp": 1574346434119, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="X7sKbsEf6RvF" outputId="e03f8f82-1240-4562-c3e4-b1ac576231da"
df_decile = ltv.decile_stats(y_eval, y_pred)
df_decile
# + colab={"height": 306} colab_type="code" executionInfo={"elapsed": 165, "status": "ok", "timestamp": 1574346434331, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="DHdLqUqdL4hf" outputId="9b917c76-e741-433e-bfe2-50efea763b8c"
ax = df_decile[['label_mean', 'pred_mean']].plot.bar(rot=0)
ax.set_title('Decile Chart')
ax.set_xlabel('Prediction bucket')
ax.set_ylabel('Average bucket value')
ax.legend(['Label', 'Prediction'], loc='upper left');
# + [markdown] colab_type="text" id="nK6DQ89xU-d4"
# ### Rank Correlation
# + colab={"height": 34} colab_type="code" executionInfo={"elapsed": 98, "status": "ok", "timestamp": 1574346575024, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="I9qWGyY3WePz" outputId="32d8a79b-8903-420b-d816-3d89b9d42b15"
def spearmanr(
x1: Sequence[float],
x2: Sequence[float]) -> float:
"""Calculates spearmanr rank correlation coefficient.
See https://docs.scipy.org/doc/scipy/reference/stats.html.
Args:
x1: 1D array_like.
x2: 1D array_like.
Returns:
correlation: float.
"""
return stats.spearmanr(x1, x2, nan_policy='raise')[0]
spearman_corr = spearmanr(y_eval, y_pred)
spearman_corr
# + [markdown] colab_type="text" id="-i_AbqhXcurk"
# ### All metrics together
# + colab={} colab_type="code" id="Umqg1-0Bc1HS"
df_metrics = pd.DataFrame({
'model': MODEL,
'loss_function': LOSS,
'train_loss': history['loss'][-1],
'eval_loss': history['val_loss'][-1],
'label_positive': np.mean(y_eval > 0),
'label_mean': y_eval.mean(),
'pred_mean': y_pred.mean(),
'decile_mape': df_decile['decile_mape'].mean(),
'baseline_gini': gini['normalized'][1],
'gini': gini['normalized'][2],
'spearman_corr': spearman_corr,
}, index=[VERSION])
# + colab={} colab_type="code" id="C_cM2Mc2SB3W"
for unit_cost, total_profit in zip(unit_costs, total_profits):
df_metrics['total_profit_{:02d}'.format(int(unit_cost * 100))] = total_profit
# + colab={"height": 610} colab_type="code" executionInfo={"elapsed": 99, "status": "ok", "timestamp": 1574346578846, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 300} id="iyMvsOtbRrXZ" outputId="e39f2617-7279-4776-ad01-2723ee8c8539"
df_metrics.T
# + [markdown] colab_type="text" id="8uHtLKk1x0IE"
# ## Save
# + colab={} colab_type="code" id="L-fMkqWIm6X6"
output_path = OUTPUT_CSV_FOLDER
# + colab={} colab_type="code" id="jpJJAbWEm94h"
if not os.path.isdir(output_path):
os.makedirs(output_path)
# + colab={} colab_type="code" id="y4LcrTLOm_4B"
output_file = os.path.join(output_path, '{}_regression_{}_{}.csv'.format(MODEL, LOSS, VERSION))
# + colab={} colab_type="code" id="4WOF7a-dnENp"
df_metrics.to_csv(output_file, index=False)
| notebooks/kdd_cup_98/regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Regular Expression Examples
# These examples are adapted from the online book Python For Everyone Chapter 11, authored by <NAME>.
# https://www.py4e.com/html3/11-regex
#
# The sample data are some emails.
# https://www.py4e.com/code3/mbox-short.txt
# Import the regular expression package in python
import re
# Search for lines that contain 'From'
hand = open('mbox-short.txt') # remember to change the filepath to the path on your computer
for line in hand:
line = line.rstrip() #remove the white spaces etc at the end of the line
if re.search('From:', line):
print(line)
# Search for lines that start with 'From'
line = "edu From: <EMAIL>"
if re.search('^From:', line):
print("line starts with 'From:'")
else:
print("line does not start with 'From:'")
# Search for lines that start with From and have an at sign
line = "From: <EMAIL>"
if re.search('^From:.+@', line): # "." can match any character, "+" means match at least one character, "*" means match zero to more chracters
print("line starts with From and have an at sign")
else:
print("line does not contain the pattern")
# +
# Extract data using regular expression
# Extracting email addresses with a non-whitespace string before @ and another after @
# In regular expression, the pattern is \S+@\S+
# "@2PM" does not match with this regular expression because there is no string before @
line = 'A message from <EMAIL> to <EMAIL> about meeting @2PM'
items = re.findall('\S+@\S+', line) #"\S" means a non-whitespace character
print(items)
# -
# Search and extract data using regular expression
# If you are only interested in the string after @, but you need the entire pattern to find it,
# you can add parentheses around the string using regular expression \S+@(\S+).
# It will return the string after @
line = 'A message from <EMAIL> to <EMAIL> about meeting @2PM'
items = re.findall('\S+@(\S+)', line) #"\S" means a non-whitespace character
print(items)
# +
# common regular expressions
# '^' - beginning of line
# '$' - end of line
# '.' - any character
# '\d' - one digit number
# '*' - zero or more occurrences
# '+' - one or more occurrences
# '\S' - non-whitespace character
# '[a-z]' - all lowercase letters
# '[A-Z]' - all uppercase letters
line = "123abc456DEF"
# find the entire line
items = re.findall('^.*$', line)
print(items)
# find all numbers
items = re.findall('(\d+)', line)
print(items)
# find all strings that begin with one or more digits and end with one or more letters
items = re.findall('(\d+[a-zA-Z]+)', line)
print(items)
# -
# ## Exercise
# +
# find all strings that begin with one or more digits and end with one or more lowercase letters
# the answer is '123abc'
line = "123abc456DEF"
# your code starts here
items = re.findall('(\d+[a-z]+)', line)
print(items)
# your code ends here
# +
# find the digits at the beginning of the line
# the answer is '123'
line = "123abc456DEF"
# your code starts here
items = re.findall('(^\d+)', line)
print(items)
# your code ends here
# +
# find the digits between letters
# the answer is '456'
line = "123abc456DEF"
# your code starts here
items = re.findall('[a-z](\d+)', line)
print(items)
# your code ends here
# -
| assets/all_html/2019_10_15_python-regular-expression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
# %matplotlib inline
class Interpolation:
def __call__(self, img, py=32, px=32):
sy = py / img.shape[0]
sx = px / img.shape[1]
return self.bilinear(img, sy, sx)
def bilinear(self, img, sy=1.5, sx=1.5):
H, W = img.shape[:2]
ch = 1 if len(img.shape) < 3 else img.shape[2]
H_big, W_big = int(H * sy), int(W * sx)
output_img = np.zeros((H_big, W_big, ch))
for i in range(H_big):
for j in range(W_big):
y, x = min(H-2, int(i/sy)), min(W-2, int(j/sx))
dy, dx = i/sy - y, j/sx - x
D = [(1-dy)*(1-dx), dy*(1-dx), (1-dy)*dx, dy*dx]
if len(img.shape) == 3:
I = [img[y, x, :], img[y+1, x, :], img[y, x+1, :], img[y+1, x+1, :]]
output_img[i, j, :] = sum(d*z for (d, z) in zip(D, I))
else:
I = [img[y, x], img[y+1, x], img[y, x+1], img[y+1, x+1]]
output_img[i, j] = sum(d*z for (d, z) in zip(D, I))
# 仮に幅や高さが 1 になるような画像であると成立しないので注意
output_img = np.squeeze(np.clip(output_img, 0, 255).astype("uint8"))
return output_img
class HOGFeatureValue:
def quantize(self, value):
res = np.floor(value / 20)
if res >= 9: res = 8
return res
# reference solution は差分計算でオーバーフローをおこしていそう
def calc_gx_gy(self, img):
input_img = img.copy().astype(np.int32)
H, W = input_img.shape
gx = np.zeros_like(input_img).astype(np.int32)
gy = np.zeros_like(input_img).astype(np.int32)
for y in range(H):
for x in range(W):
gx[y, x] = input_img[y, min(x+1, W-1)] - input_img[y, max(x-1, 0)]
gy[y, x] = input_img[min(y+1, H-1), x] - input_img[max(y-1, 0), x]
return gx, gy
def calc_mag_ang(self, gx, gy):
H, W = gx.shape
mag = np.zeros_like(gx, dtype=np.float32)
ang = np.zeros_like(gy, dtype=np.float32)
for y in range(H):
for x in range(W):
mag[y, x] = np.sqrt(gx[y, x]**2 + gy[y, x]**2)
ang[y, x] = np.arctan2(gy[y, x], gx[y, x])
if ang[y, x] < 0: ang[y, x] += np.pi
ang[y, x] = self.quantize(ang[y, x] / np.pi * 180.0)
ang = ang.astype(np.uint8)
return mag, ang
def get_colorized_ang(self, ang):
color = [
[255, 0, 0],
[0, 255, 0],
[0, 0, 255],
[255, 255, 0],
[255, 0, 255],
[0, 255, 255],
[127, 127, 0],
[127, 0, 127],
[0, 127, 127],
]
H, W = ang.shape
colorized_ang = np.zeros((H, W, 3), dtype=np.uint8)
for i in range(9):
colorized_ang[ang == i] = color[i]
return colorized_ang
# reference solution で x, y に掛ける数が 4 なのはおそらく間違い
def gradient_histogram(self, mag, ang, N):
H, W = mag.shape
cell_H, cell_W = H // N, W // N
histogram = np.zeros((cell_H, cell_W, 9), dtype=np.float32)
for y in range(cell_H):
for x in range(cell_W):
for j in range(N):
for i in range(N):
histogram[y, x, ang[y*N+j, x*N+i]] += mag[y*N+j, x*N+i]
return histogram
def normalization(self, histogram, epsilon=1):
cell_H, cell_W, _ = histogram.shape
result_histogram = histogram.copy().astype(np.float32)
for y in range(cell_H):
for x in range(cell_W):
result_histogram[y, x] /= np.sqrt(
np.sum(
histogram[max(y-1, 0):min(y+2, cell_H),
max(x-1, 0):min(x+2, cell_W)]**2
) + epsilon
)
return result_histogram
def draw_feature_value(self, img_gray, histogram, N=8):
H, W = img_gray.shape
cell_H, cell_W, _ = histogram.shape
output_img = img_gray.copy().astype(np.uint8)
for y in range(cell_H):
for x in range(cell_W):
cx = x*N + N//2
cy = y*N + N//2
x1, y1, x2, y2 = cx + N//2 - 1, cy, cx - N//2 + 1, cy
h = histogram[y, x] / np.sum(histogram[y, x])
h /= h.max()
for c in range(9):
theta = (20 * c + 10) / 180.0 * np.pi
rx = int(np.sin(theta)*(x1-cx) + np.cos(theta)*(y1-cy) + cx)
ry = int(np.cos(theta)*(x1-cx) - np.cos(theta)*(y1-cy) + cy)
lx = int(np.sin(theta)*(x2-cx) + np.cos(theta)*(y2-cy) + cx)
ly = int(np.cos(theta)*(x2-cx) - np.cos(theta)*(y2-cy) + cy)
color = int(255.0 * h[c])
cv2.line(output_img, (lx, ly), (rx, ry), (color, color, color))
return output_img
def get_hog_matrix(self, img, cell_size=8):
if len(img.shape) == 3:
img_gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
elif len(img.shape) == 2:
img_gray = img.copy()
else:
assert False, "invlaid image dimension."
gx, gy = self.calc_gx_gy(img_gray)
mag, ang = self.calc_mag_ang(gx, gy)
histogram = self.normalization(self.gradient_histogram(mag, ang, cell_size))
return histogram
class Metrics:
# rectangle: [y1, x1, y2, x2] (y1 <= y2, x1 <= x2)
def iou(self, r1, r2):
oy = max(0, min(r1[2], r2[2]) - max(r1[0], r2[0]))
ox = max(0, min(r1[3], r2[3]) - max(r1[1], r2[1]))
R1 = (r1[2] - r1[0]) * (r1[3] - r1[1])
R2 = (r2[2] - r2[0]) * (r2[3] - r2[1])
Rol = oy * ox
return abs(Rol) / abs(R1 + R2 - Rol)
def accuracy(self, num_of_samples, correct):
return correct / num_of_samples
class Solver:
def __init__(self, seed=0):
self.hog = HOGFeatureValue()
self.resize = Interpolation()
def get_bounding_boxes(self, img, stride=4):
recs = np.array(((42, 42), (56, 56), (70, 70)), dtype=np.float32)
H, W = img.shape[:2]
num_boxes = ((H+stride-1)//stride) * ((W+stride-1)//stride) * recs.shape[0]
boxes = np.zeros((num_boxes, 4))
i = 0
for y in range(0, H, stride):
for x in range(0, W, stride):
for dy, dx in recs:
y1, x1 = y, x
y2, x2 = min(H-1, y+dy), min(W-1, x+dx)
boxes[i] = np.array([y1, x1, y2, x2])
i += 1
return boxes
def get_hog_matrices(self, img, boxes, img_size=32, cell_size=8):
vec_size = (img_size//cell_size)**2 * 9
data = np.zeros((boxes.shape[0], vec_size))
for i, box in enumerate(tqdm(boxes)):
y1, x1, y2, x2 = box.astype(np.int)
resized_img = self.resize(img[y1:y2, x1:x2], img_size, img_size)
data[i] = self.hog.get_hog_matrix(resized_img, cell_size).ravel().astype(np.float32)
return data
def get_test_data(self, img, stride=4, img_size=32, cell_size=8):
boxes = self.get_bounding_boxes(img, stride)
test_data = self.get_hog_matrices(img, boxes, img_size, cell_size)
return test_data
def problem_97(self, img_path):
input_img = cv2.imread(img_path)
# stride = 4 だと 5 分掛かるので妥協。並列化したらマシになるかもしれないけど
test_data = self.get_test_data(input_img, stride=8)
solver = Solver()
solver.problem_97("../imori_many.jpg")
| Question_91_100/solutions_py/solution_097.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: DeepLearning Python
# language: python
# name: dl
# ---
# <font size="+5">#01 | Why Neural Networks Deeply Learn a Mathematical Formula</font>
# - Subscribe to my [Blog ↗](https://blog.pythonassembly.com/)
# - Let's keep in touch on [LinkedIn ↗](www.linkedin.com/in/jsulopz) 😄
# # Discipline to Search Solutions in Google
# > Apply the following steps when **looking for solutions in Google**:
# >
# > 1. **Necesity**: How to load an Excel in Python?
# > 2. **Search in Google**: by keywords
# > - `load excel python`
# > - ~~how to load excel in python~~
# > 3. **Solution**: What's the `function()` that loads an Excel in Python?
# > - A Function to Programming is what the Atom to Phisics.
# > - Every time you want to do something in programming
# > - **You will need a `function()`** to make it
# > - Theferore, you must **detect parenthesis `()`**
# > - Out of all the words that you see in a website
# > - Because they indicate the presence of a `function()`.
# # Load the Data
import tensorflow as tf
fashion_mnist = tf.keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
train_images = train_images / 255
test_images = test_images / 255
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# ## Get to Know the Data
# ### Visualize some Samples
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# -
# ### Visualize One Sample/Row/Image/Explanatory Variables
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
# #### Target Variable Value
idx_label = train_labels[0]
class_names[idx_label]
# # Neural Network Concepts in Python
# ## Initializing the `Weights`
# > - https://keras.io/api/layers/initializers/
# ### How to `kernel_initializer` the weights?
from tensorflow.keras import Sequential, Input
from tensorflow.keras.layers import Dense, Flatten
train_images.shape
model = Sequential()
model.add(Flatten(input_shape=(28, 28)))
model.add(layer=Dense(units=128, kernel_initializer='zeros'))
model.add(layer=Dense(units=10))
# #### Make a Prediction with the Neural Network
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
train_images[0].shape
# #### Observe the numbers for the `weights`
model.get_weights()
# #### Predictions vs Reality
# > 1. Calculate the Predicted Accidents and
# > 2. Compare it with the Real Total Accidents
# #### `fit()` the `model` and compare again
model.compile(loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# + tags=[]
model.fit(train_images, train_labels, epochs=20, verbose=0)
# + [markdown] tags=[]
# ##### Observe the numbers for the `weights`
# -
# + [markdown] tags=[]
# ##### Predictions vs Reality
# -
y_pred = model.predict(train_images)
(y_pred.argmax(axis=1) == train_labels).mean()
# ### How to `kernel_initializer` the weights to 1?
# ### How to `kernel_initializer` the weights to `glorot_uniform` (default)?
# ## Play with the Activation Function
# > - https://keras.io/api/layers/activations/
# %%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/IHZwWFHWa-w?start=558" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# ### Use `sigmoid` activation in last layer
model = Sequential()
model.add(Flatten(input_shape=(28, 28)))
model.add(layer=Dense(units=128, kernel_initializer='zeros'))
model.add(layer=Dense(units=10))
model.compile(loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# #### `fit()` the Model
model.fit(train_images, train_labels, epochs=20, verbose=0)
# #### Predictions vs Reality
y_pred = model.predict(train_images)
(y_pred.argmax(axis=1) == train_labels).mean()
# #### Observe the numbers for the `weights`
#
# > - Have they changed?
model.get_weights()
# ### Use `linear` activation in last layer
# ### Use `tanh` activation in last layer
# ### Use `relu` activation in last layer
# ### How are the predictions changing? Why?
# ## Optimizer
# > - https://keras.io/api/optimizers/#available-optimizers
# Optimizers comparison in GIF → https://mlfromscratch.com/optimizers-explained/#adam
# Tesla's Neural Network Models is composed of 48 models trainned in 70.000 hours of GPU → https://tesla.com/ai
# 1 Year with a 8 GPU Computer → https://twitter.com/thirdrowtesla/status/1252723358342377472
# ### Use Gradient Descent `SGD`
model = Sequential()
model.add(Flatten(input_shape=(28, 28)))
model.add(layer=Dense(units=128, kernel_initializer='zeros'))
model.add(layer=Dense(units=10))
# #### `compile()` the model
model.compile(optimizer='sgd', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# #### `fit()` the Model
history = model.fit(X, y, epochs=20, verbose=0)
# #### Predictions vs Reality
y_pred = model.predict(train_images)
(y_pred.argmax(axis=1) == train_labels).mean()
# #### Observe the numbers for the `weights`
#
# > - Have they changed?
model.get_weights()
#
# #### View History
import matplotlib.pyplot as plt
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# ### Use `ADAM`
# ### Use `RMSPROP`
# ### Does it take different times to get the best accuracy? Why?
# ## Loss Functions
# > - https://keras.io/api/losses/
# ### `binary_crossentropy`
# ### `sparse_categorical_crossentropy`
# ### `mean_absolute_error`
# ### `mean_squared_error`
# ## In the end, what should be a feasible configuration of the Neural Network for this data?
# # Common Errors
# ## The `kernel_initializer` Matters
# ## The `activation` Function Matters
# ## The `optimizer` Matters
# ## The Number of `epochs` Matters
# ## The `loss` Function Matters
# ## The Number of `epochs` Matters
# # Neural Network's importance to find **Non-Linear Patterns** in the Data
#
# > - The number of Neurons & Hidden Layers
# https://towardsdatascience.com/beginners-ask-how-many-hidden-layers-neurons-to-use-in-artificial-neural-networks-51466afa0d3e
# https://playground.tensorflow.org/#activation=tanh&batchSize=10&dataset=circle®Dataset=reg-plane&learningRate=0.03®ularizationRate=0&noise=0&networkShape=4,2&seed=0.87287&showTestData=false&discretize=false&percTrainData=50&x=true&y=true&xTimesY=false&xSquared=false&ySquared=false&cosX=false&sinX=false&cosY=false&sinY=false&collectStats=false&problem=classification&initZero=false&hideText=false
# ## Summary
#
# - Mathematical Formula
# - Weights / Kernel Initializer
# - Loss Function
# - Activation Function
# - Optimizers
# ## What cannot you change arbitrarily of a Neural Network?
#
# - Input Neurons
# - Output Neurons
# - Loss Functions
# - Activation Functions
| II Machine Learning & Deep Learning/02_Why Neural Networks Deeply Learn a Mathematical Formula/02practice_neural-networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Amazon Kinesis Data Stream
#
# Amazon Kinesis Data Streams ingests a large amount of data in real time, durably stores the data, and makes the data available for consumption. The unit of data stored by Kinesis Data Streams is a data record. A data stream represents a group of data records. The data records in a data stream are distributed into shards.
#
# A shard has a sequence of data records in a stream. When you create a stream, you specify the number of shards for the stream. The total capacity of a stream is the sum of the capacities of its shards. You can increase or decrease the number of shards in a stream as needed. However, you are charged on a per-shard basis.
#
# The producers continually push data to Kinesis Data Streams, and the consumers process the data in real time. Consumers (such as a custom application running on Amazon EC2 or an Amazon Kinesis Data Firehose delivery stream) can store their results using an AWS service such as Amazon DynamoDB, Amazon Redshift, or Amazon S3.
#
# <img src="img/kinesis_data_stream_docs.png" width="80%" align="left">
# +
import boto3
import sagemaker
import pandas as pd
import json
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sm = boto3.Session().client(service_name="sagemaker", region_name=region)
kinesis = boto3.Session().client(service_name="kinesis", region_name=region)
sts = boto3.Session().client(service_name="sts", region_name=region)
# -
# # Create a Kinesis Data Stream
# <img src="img/kinesis-data-stream.png" width="90%" align="left">
# %store -r stream_name
try:
stream_name
except NameError:
print("+++++++++++++++++++++++++++++++")
print("[ERROR] Please run all previous notebooks in this section before you continue.")
print("+++++++++++++++++++++++++++++++")
print(stream_name)
shard_count = 2
# +
from botocore.exceptions import ClientError
try:
response = kinesis.create_stream(StreamName=stream_name, ShardCount=shard_count)
print("Data Stream {} successfully created.".format(stream_name))
print(json.dumps(response, indent=4, sort_keys=True, default=str))
except ClientError as e:
if e.response["Error"]["Code"] == "ResourceInUseException":
print("Data Stream {} already exists.".format(stream_name))
else:
print("Unexpected error: %s" % e)
# +
import time
status = ""
while status != "ACTIVE":
r = kinesis.describe_stream(StreamName=stream_name)
description = r.get("StreamDescription")
status = description.get("StreamStatus")
time.sleep(5)
print("Stream {} is active".format(stream_name))
# -
# ## _This may take a minute. Please be patient._
# +
stream_response = kinesis.describe_stream(StreamName=stream_name)
print(json.dumps(stream_response, indent=4, sort_keys=True, default=str))
# -
stream_arn = stream_response["StreamDescription"]["StreamARN"]
print(stream_arn)
# %store stream_arn
# # Review Kinesis Data Stream
# +
from IPython.core.display import display, HTML
display(
HTML(
'<b>Review <a target="blank" href="https://console.aws.amazon.com/kinesis/home?region={}#/streams/details/{}/details"> Kinesis Data Stream</a></b>'.format(
region, stream_name
)
)
)
# -
# # Store Variables for the Next Notebooks
# %store
# # Release Resources
# + language="html"
#
# <p><b>Shutting down your kernel for this notebook to release resources.</b></p>
# <button class="sm-command-button" data-commandlinker-command="kernelmenu:shutdown" style="display:none;">Shutdown Kernel</button>
#
# <script>
# try {
# els = document.getElementsByClassName("sm-command-button");
# els[0].click();
# }
# catch(err) {
# // NoOp
# }
# </script>
# + language="javascript"
#
# try {
# Jupyter.notebook.save_checkpoint();
# Jupyter.notebook.session.delete();
# }
# catch(err) {
# // NoOp
# }
| 11_stream/04_Create_Kinesis_Data_Stream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import os
import feedparser
from features import get_words, entry_features
from services.fisher_classifier_service import FisherClassifierService
feeds = "".join(open(os.path.join(os.getcwd(), "search.xml")).readlines())
entries = feedparser.parse(feeds)['entries']
entry_features(entries[0])
sentence = "This is a simple test to check if only words between 2 and 20 characters are returned"
get_words(sentence)
# # Playfield
classifier = FisherClassifierService(get_features=get_words, persistent_storage=True)
# If we are using "get_words"
classifier.train('Nobody owns the water.', 'good')
classifier.train('the quick rabbit jumps fences', 'good')
classifier.train('buy pharmaceuticals now', 'bad')
classifier.train('make quick money at the online casino', 'bad')
classifier.train('the quick brown fox jumps', 'good')
classifier = FisherClassifierService(get_features=entry_features, persistent_storage=False)
# If we are using "entry_features"
for entry in entries[:5]:
# Print the contents of the entry
print 'Title: ' + entry['title'].encode('utf-8')
print 'Publisher: ' + entry['publisher'].encode('utf-8')
print
print entry['summary'].encode('utf-8')
# Combine all the text to create one item for the classifier
fulltext = "\n".join([entry['title'], entry['publisher'], entry['summary']])
# Print the best guess at the current category
print 'Guess: ' + str(classifier.classify(entry))
# Ask the user to specify the correct category and train on that
cl = raw_input('Enter category: ')
classifier.train(entry, cl)
if not classifier.persistent_storage:
classifier.fc, classifier.cc
classifier.fprob("money", "good"), classifier.fprob("money", "bad")
classifier.wprob("money", "good", assumed_probability=1), classifier.wprob("money", "bad", assumed_probability=1),
classifier.classify(entries[4])
#classifier.classify(entries[])
# +
| chapter6/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# get_ipython().magic('matplotlib notebook')
get_ipython().magic('matplotlib inline')
get_ipython().magic('load_ext autoreload')
get_ipython().magic('autoreload 2')
#___________________________________________________________________________________________________________________
import os
import tripyview as tpv
import shapefile as shp
import numpy as np
# + tags=["parameters"]
# Parameters
# mesh_path ='/work/ollie/projects/clidyn/FESOM2/meshes/core2/'
mesh_path = '/work/ollie/pscholz/mesh_fesom2.0/core2_srt_dep@node/'
save_path = None #'~/figures/test_papermill/'
save_fname= None
#_____________________________________________________________________________________
which_cycl= 3
which_mode= 'xmoc'
#_____________________________________________________________________________________
input_paths= list()
input_paths.append('/home/ollie/pscholz/results/trr181_tke_ctrl_ck0.1/')
# input_paths.append('/home/ollie/pscholz/results/trr181_tke_ctrl_ck0.3/')
# input_paths.append('/home/ollie/pscholz/results/trr181_tke+idemix_orig_ck0.1/')
input_paths.append('/home/ollie/pscholz/results/trr181_tke+idemix_jayne09_ck0.1/')
input_paths.append('/home/ollie/pscholz/results/trr181_tke+idemix_nycander05_ck0.1/')
input_paths.append('/home/ollie/pscholz/results/trr181_tke+idemix_stormtide2_ck0.1/')
# input_paths.append('/home/ollie/pscholz/results/trr181_tke+idemix_jayne09_ck0.3/')
# input_paths.append('/home/ollie/pscholz/results/old_trr181_tke+idemix_nycander05_ck0.3/')
# input_paths.append('/home/ollie/pscholz/results/old_trr181_tke+idemix_stormtide2_ck0.3/')
input_names= list()
input_names.append('TKE, ck=0.1')
# input_names.append('TKE, ck=0.3')
# input_names.append('TKE+IDEMIX, ck=0.1, jayne (old param)')
input_names.append('TKE+IDEMIX, ck=0.1, jayne (new param)')
input_names.append('TKE+IDEMIX, ck=0.1, nycander (new param)')
input_names.append('TKE+IDEMIX, ck=0.1, stormtide (new param)')
# input_names.append('TKE+IDEMIX, ck=0.3, jayne (new param)')
# input_names.append('TKE+IDEMIX, ck=0.3, nycander (new param)')
# input_names.append('TKE+IDEMIX, ck=0.3, stormtide (new param)')
vname = 'amoc'
year = [1979,2019]
mon, day, record, box, depth = None, None, None, None, None
#_____________________________________________________________________________________
# do anomaly plots in case ref_path is not None
ref_path = None #'/home/ollie/pscholz/results/trr181_tke_ctrl_ck0.1/' # None
ref_name = None # 'TKE, ck=0.1' # None
ref_year = None # [2009,2019]
ref_mon, ref_day, ref_record = None, None, None
#_____________________________________________________________________________________
cstr = 'blue2red'
cnum = 20
cref = 0
crange, cmin, cmax, cfac, climit = None, None, None, None, None
#_____________________________________________________________________________________
ncolumn = 3
do_rescale= None
which_dpi = 300
do_contour= True
# +
#___LOAD FESOM2 MESH___________________________________________________________________________________
mesh=tpv.load_mesh_fesom2(mesh_path, do_rot='None', focus=0, do_info=True, do_pickle=True,
do_earea=True, do_narea=True, do_eresol=[True,'mean'], do_nresol=[True,'eresol'])
#______________________________________________________________________________________________________
if which_cycl is not None:
for ii,ipath in enumerate(input_paths):
input_paths[ii] = os.path.join(ipath,'{:d}/'.format(which_cycl))
print(ii, input_paths[ii])
if ref_path is not None:
ref_path = os.path.join(ref_path,'{:d}/'.format(which_cycl))
print('R', ref_path)
#______________________________________________________________________________________________________
cinfo=dict({'cstr':cstr, 'cnum':cnum})
if crange is not None: cinfo['crange']=crange
if cmin is not None: cinfo['cmin' ]=cmin
if cmax is not None: cinfo['cmax' ]=cmax
if cref is not None: cinfo['cref' ]=cref
if cfac is not None: cinfo['cfac' ]=cfac
if climit is not None: cinfo['climit']=climit
if ref_path is not None: cinfo['cref' ]=0.0
#______________________________________________________________________________________________________
# in case of diff plots
if ref_path is not None:
if ref_year is None: ref_year = year
if ref_mon is None: ref_mon = mon
if ref_record is None: ref_record = record
# +
aux_vname, aux_vnamebv = 'w', 'bolus_w'
#___LOAD FESOM2 REFERENCE DATA________________________________________________________________________
if ref_path is not None:
print(ref_path)
data_ref = tpv.load_data_fesom2(mesh, ref_path, vname=aux_vname, year=ref_year, descript=ref_name , do_info=False)
data_ref[aux_vname].data = data_ref[aux_vname].data+tpv.load_data_fesom2(mesh, ref_path, vname=aux_vnamebv, year=ref_year, do_info=False)[aux_vnamebv].data
#___LOAD FESOM2 DATA___________________________________________________________________________________
data_list = list()
for datapath, descript in zip(input_paths, input_names):
print(datapath, descript)
data = tpv.load_data_fesom2(mesh, datapath, vname=aux_vname, year=year, descript=descript , do_info=False)
data[aux_vname].data = data[aux_vname].data+tpv.load_data_fesom2(mesh, datapath, vname=aux_vnamebv, year=year, do_info=False)[aux_vnamebv].data
data_list.append(data)
del(data)
#___COMPUTE XMOC_______________________________________________________________________________________
if ref_path is not None:
moc_ref = tpv.calc_xmoc(mesh, data_ref, dlat=1.0, which_moc=vname, do_info=False)
del(data_ref)
moc_list = list()
for data in data_list:
moc = tpv.calc_xmoc(mesh, data, dlat=1.0, which_moc=vname, do_info=False)
if ref_path is not None: moc_list.append( tpv.do_anomaly(moc, moc_ref) )
else: moc_list.append( moc )
del(data_list, data, moc)
# -
#___PLOT FESOM2 DATA___________________________________________________________________________________
spath = save_path
sname = vname
slabel = moc_list[0]['moc'].attrs['str_lsave']
if spath is not None: spath = os.path.join(spath,'{}_{}_{}.png'.format(which_mode, sname, slabel))
nrow = np.ceil(len(moc_list)/ncolumn).astype('int')
if save_fname is not None: spath = save_fname
#____________________________________________________________________________________________________
fig, ax, cbar=tpv.plot_xmoc(moc_list, which_moc=vname, cinfo=cinfo, figsize=[ncolumn*7, nrow*3.5], n_rc=[nrow, ncolumn],
do_save = spath, save_dpi=which_dpi, do_contour = do_contour)
| templates_notebooks/template_xmoc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
x = 10
nome = 'aluno'
nota = 8.75
fez_inscricao = True
print(type(x))
print(type(nome))
print(type(nota))
print(type(fez_inscricao))
# +
# Qual o resultado armazenado na variável operacao_1: 25 ou 17? 17
operacao_1 = 2 + 3 * 5
# Qual o resultado armazenado na variável operacao_2 25 ou 17? 25
operacao_2 = (2 + 3) * 5
# Qual o resultado armazenado na variável operacao_3: 4 ou 1? 1
operacao_3 = 4 / 2 ** 2
# Qual o resultado armazenado na variável operacao_4: 1 ou 5? 5
operacao_4 = 13 % 3 + 4
print(f'Resultado em operacao_1 = {operacao_1}')
print(f'Resultado em operacao_2 = {operacao_2}')
print(f'Resultado em operacao_3 = {operacao_3}')
print(f'Resultado em operacao_4 = {operacao_4}')
# +
a = 2
b = 0.5
c = 1
x = int(input('Digite o valor de x: '))
y = a * x ** 2 + b * x + c
print(f'O resultado de y para x = {x} é {y}')
# +
c = 200 # Valor da constante
mes = int(input('Digite o mês que deseja saber o resultado: '))
r = c * mes
print(f'A quantidade de peças para o mês {mes} será {r}')
# -
from time import sleep
cont = 0
texto = """
A inserção de comentários no código do programa é uma prática normal.
Em função disso, toda linguagem de programação tem alguma maneira de permitir que comentários sejam inseridos nos programas.
O objetivo é adicionar descrições em partes do código, seja para documentá-lo ou para adicionar uma descrição do algoritmo implementado (BANIN, p. 45, 2018)."
"""
for i, c in enumerate(texto):
if c == 'a':
cont += 1
else:
continue
print(f'A vogal "a" foi encontrada no texto {cont} vezes')
# # FUNÇÕES EM PYTHON
#
# +
def converter_mes_para_extenso(data):
mes = '''Janeiro Fevereiro Março Abril Maio Junho
Julho Agosto Setembro Outubro Novembro Dezembro'''.split()
d, m, a = data.split('/')
mes_extenso = mes[int(m) - 1] # Isso porque o mês 1 está na posição 0
return f'{d} de {mes_extenso} de {a}'
print(converter_mes_para_extenso('18/09/1978'))
# -
data = '10.04.2020'.split('.')
print(data)
# +
def calcular_valor(valor_prod, qtde, moeda = "real", desconto = None, acrescimo = None):
v_bruto = valor_prod * qtde
if desconto:
v_liquido = v_bruto - (v_bruto * (desconto / 100))
elif acrescimo:
v_liquido = v_bruto + (v_bruto * (acrescimo / 100))
else:
v_liquido = v_bruto
if moeda == 'real':
return v_liquido
elif moeda == 'dolar':
return v_liquido * 5
elif moeda == 'euro':
return v_liquido * 5.7
else:
print('Moeda não caddastrada')
return 0
valor_a_pagar = calcular_valor(valor_prod = 32, qtde = 5, desconto = 5)
print(f'O valor final da conta é {valor_a_pagar}')
# -
# # Listas
# +
linguagens = '''Python Java JavaScript C C# c++ Swift Go Kotlin'''.split()
print('Antes da listcomp = ', linguagens)
linguagens = [item.lower() for item in linguagens]
print('\nDepois da listcomp = ', linguagens)
# +
linguagens = '''Python Java JavaScript C C# c++ Swift Go Kotlin'''.split()
linguagens_java = [item for item in linguagens if 'Java' in item]
print(linguagens_java)
# +
print('Exemplo 1')
linguagens = '''Python Java JavaScript C C# c++ Swift Go Kotlin'''.split()
nova_lista = map(lambda x: x.lower(), linguagens)
print(f'A nova lista é = {nova_lista}')
nova_lista = list(nova_lista)
print(f'Agora sim, a nova lista é = {nova_lista}')
print('Exemplo 2')
numeros = [0, 1, 2, 3, 4, 5]
quadrados = list(map(lambda x: x*x, numeros))
print(f'Lista com o número elevado a ele mesmo = {quadrados}')
# +
numeros = list(range(0, 21))
numeros_pares = list(filter(lambda x: x % 2 == 0, numeros))
print(numeros_pares)
# +
vogais = ('a', 'e', 'i', 'o', 'u')
for item in enumerate(vogais):
print(item)
print(tuple(enumerate(vogais)))
print(list(enumerate(vogais)))
# -
# # Set
# +
def create_report():
componentes_verificados = set(['caixas de som', 'cooler', 'dissipador de calor', 'cpu', 'hd', 'estabilizador', 'gabinete', 'hub', 'impressora', 'joystick', 'memória ram', 'microfone', 'modem', 'monitor', 'mouse', 'no-break', 'placa de captura', 'placa de som', 'placa de vídeo', 'placa mãe', 'scanner', 'teclado', 'webcam' ])
componentes_com_defeito = set(['hd', 'monitor', 'placa de som','scanner'])
qtde_componentes_verificados = len(componentes_verificados)
qtde_componentes_com_defeito = len(componentes_com_defeito)
componentes_ok = componentes_verificados.difference(componentes_com_defeito)
print(f'Foram verificados {qtde_componentes_verificados} componentes\n')
print(f'{qtde_componentes_com_defeito} componentes apresentaram defeito')
print('Os componentes que podem ser vendidos são: ')
for item in componentes_ok:
print(item)
create_report()
# +
def extrair_lista_email(dict_1, dict_2):
lista_1 = list(zip(dict_1['nome'], dict_1['email'], dict_1['enviado']))
print(f'Amostra da lista 1 = {lista_1[0]}')
lista_2 = list(zip(dict_2['nome'], dict_2['email'], dict_2['enviado']))
print(f'Amostra da lista 2 = {lista_2[0]}')
dados = lista_1 + lista_2
print(f'Amostra dos dados = {dados[:3]}')
# Queremos uma lista com o e-mail de quem ainda não recebeu o aviso
emails = [item[1] for item in dados if not item[2]]
return emails
dados_1 = {
'nome': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'email': ['<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>'],
'enviado': [False, False, False, False, False, False, False, True, False, False]
}
dados_2 = {
'nome': ['<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>'],
'email': ['<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>', '<EMAIL>'],
'enviado': [False, False, False, True, True, True, False, True, True, False]
}
emails = extrair_lista_email(dict_1 = dados_1, dict_2 = dados_2)
print(f'\nE-mails a serem enviados')
# -
# # Algoritmos de Busca
# +
lista = [10, 4, 1, 15, -3]
lista_ordenada1 = sorted(lista)
lista_ordenada2 = lista.sort()
print('lista = ', lista, '\n')
print('lista_ordenada1 = ', lista_ordenada1)
print('lista_ordenada2 = ', lista_ordenada2)
print('lista = ', lista)
# -
# # SELECTION SORT
#
# +
def executar_selection_sort(lista):
n = len(lista)
for i in range(0, n):
index_menor = i
for j in range(i+1, n):
if lista[j] < lista[index_menor]:
index_menor = j
lista[i], lista[index_menor] = lista[index_menor], lista[i]
return lista
lista = [10, 9, 5, 8, 11, -1, 3]
executar_selection_sort(lista)
# +
def executar_selection_sort_2(lista):
lista_ordenada = []
while lista:
minimo = min(lista)
lista_ordenada.append(minimo)
lista.remove(minimo)
return lista_ordenada
lista = [10, 9, 5, 8, 11, -1, 3]
executar_selection_sort_2(lista)
# -
# # BUBBLE SORT
# +
def executar_bubble_sort(lista):
n = len(lista)
for i in range(n-1):
for j in range(n-1):
if lista[j] > lista[j+1]:
lista[j], lista[j+1] = lista[j+1], lista[j]
return lista
lista = [10, 9, 5, 8, 11, -1, 3]
executar_bubble_sort(lista)
# -
# # INSERTION SORT
# +
def executar_insertion_sort(lista):
n = len(lista)
for i in range(1, n):
valor_inserir = lista[i]
j = i - 1
while j >= 0 and lista[j] > valor_inserir:
lista[j + 1] = lista[j]
j -= 1
lista[j + 1] = valor_inserir
return lista
lista = [10, 9, 5, 8, 11, -1, 3]
executar_insertion_sort(lista)
# -
# # MERGE SORT
# +
def executar_merge_sort(lista):
if len(lista) <= 1:
return lista
else:
meio = len(lista) // 2
esquerda = executar_merge_sort(lista[:meio])
direita = executar_merge_sort(lista[meio:])
return executar_merge(esquerda, direita)
def executar_merge(esquerda, direita):
sub_lista_ordenada = []
topo_esquerda, topo_direita = 0, 0
while topo_esquerda < len(esquerda) and topo_direita < len(direita):
if esquerda[topo_esquerda] <= direita[topo_direita]:
sub_lista_ordenada.append(esquerda[topo_esquerda])
topo_esquerda += 1
else:
sub_lista_ordenada.append(direita[topo_direita])
topo_direita += 1
sub_lista_ordenada += esquerda[topo_esquerda:]
sub_lista_ordenada += direita[topo_direita:]
return sub_lista_ordenada
lista = [10, 9, 5, 8, 11, -1, 3]
executar_merge_sort(lista)
# -
# # QUICK SORT
# +
def executar_quicksort(lista, inicio, fim):
if inicio < fim:
pivo = executar_particao(lista, inicio, fim)
executar_quicksort(lista, inicio, pivo - 1)
executar_quicksort(lista, pivo + 1, fim)
return lista
def executar_particao(lista, inicio, fim):
pivo = lista[fim]
esquerda = inicio
for direita in range(inicio, fim):
if lista[direita] <= pivo:
lista[direita], lista[esquerda] = lista[esquerda], lista[direita]
esquerda += 1
lista[esquerda], lista[fim] = lista[fim], lista[esquerda]
return esquerda
lista = [10, 9, 5, 8, 11, -1, 3]
executar_quicksort(lista, inicio = 0, fim = len(lista) - 1)
# +
def executar_quicksort_2(lista):
if len(lista) <= 1:
return lista
pivo = lista[0]
iguais = [valor for valor in lista if valor == pivo]
menores = [valor for valor in lista if valor < pivo]
maiores = [valor for valor in lista if valor > pivo]
return executar_quicksort_2(menores) + iguais + executar_quicksort_2(maiores)
lista = [10, 9, 5, 8, 11, -1, 3]
executar_quicksort_2(lista)
# -
# # Exercício
# +
# Parte 1 - Implementar o algoritmo de ordenação merge sort
def executar_merge_sort(lista, inicio = 0, fim = None):
if not fim:
fim = len(lista)
if fim - inicio > 1:
meio = (inicio + fim) // 2
executar_merge_sort(lista, inicio, meio)
executar_merge_sort(lista, meio, fim)
executar_merge(lista, inicio, meio, fim)
return lista
def executar_merge(lista, inicio, meio, fim):
esquerda = lista[inicio:meio]
direita = lista[meio:fim]
topo_esquerda = topo_direita = 0
for p in range(inicio, fim):
if topo_esquerda >= len(esquerda):
lista[p] = direita[topo_direita]
topo_direita += 1
elif topo_direita >= len(direita):
lista[p] = esquerda[topo_esquerda]
topo_esquerda += 1
elif esquerda[topo_esquerda] < direita[topo_direita]:
lista[p] = esquerda[topo_esquerda]
topo_esquerda += 1
else:
lista[p] = direita[topo_direita]
topo_direita += 1
# Parte 2 - Implementar o algoritmo de busca binária
def executar_busca_binaria(lista, valor):
minimo = 0
maximo = len(lista) - 1
while minimo <= maximo:
meio = (minimo + maximo) // 2
if valor < lista[meio]:
maximo = meio - 1
elif valor > lista[meio]:
minimo = meio + 1
else:
return True
return False
# Parte 3 - Implementar a função que faz a verificação do CPF, o dedup e devolve o resultado esperado
def criar_lista_dedup_ordenada(lista):
lista = [str(cpf).replace('.', '').replace('-', '') for cpf in lista]
lista = [cpf for cpf in lista if len(cpf) == 11]
lista = executar_merge_sort(lista)
lista_dedup = []
for cpf in lista:
if not executar_busca_binaria(lista_dedup, cpf):
lista_dedup.append(cpf)
return lista_dedup
# Parte 4 - Criar uma função de teste
def testar_funcao(lista_cpfs):
lista_dedup = criar_lista_dedup_ordenada(lista_cpfs)
print(lista_dedup)
lista_cpfs = ['44444444444', '111.111.111-11', '11111111111', '222.222.222-22', '333.333.333-33', '22222222222', '444.44444']
testar_funcao(lista_cpfs)
# -
| Anhanguera.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 2: "Introduction" to Python (Intermediate)
# ## 2/20/19
#
# ### Table Of Contents
# * [Q1: Worst Walk on Campus](#section1)
# * [Q2: Solving Sudoku](#section2)
# * [Q3: Alien Language](#section3)
#
#
#
# ### Hosted by and maintained by the [Statistics Undergraduate Students Association (SUSA)](https://susa.berkeley.edu). Originally authored by [<NAME>](mailto:<EMAIL>).
#
# Note: CS 61A is a pre-req for this lecture.
# <a id='section1'></a>
# ## Q1: Worst Walk on Campus
#
# References:
# - https://access.berkeley.edu/navigating-berkeley/campus-buildings
# Some documentation:
# - https://docs.python.org/3/tutorial/datastructures.html#list-comprehensions
# - https://docs.python.org/2/library/itertools.html#itertools.combinations
# - https://docs.python.org/2/library/functions.html
from geopy.geocoders import Nominatim
geolocator = Nominatim()
building_names = []
with open('berkeley/buildings.txt', 'r') as f:
for line in f.readlines():
building_names.append(line.strip())
names_and_coordinates = {}
for i, building_name in enumerate(building_names):
location = geolocator.geocode(building_name)
if location and 'Berkeley' in location.address:
names_and_coordinates[building_name] = (location.latitude, location.longitude)
print('Halls scanned: {0}/{1}'.format(i + 1, len(building_names)), end='\r')
# If the code in above cell fails in any way, run the cell below instead.
# +
import json
names_and_coordinates = json.load(open('berkeley/cal_halls.json'))
# -
from math import sin, cos, sqrt, atan2, radians
def haversine(lat1, lon1, lat2, lon2):
"""Calculates the straight line distance between two latitude-longitude coordinates"""
# approximate radius of earth in miles
R = 3959.0
lat1_r = radians(lat1)
lon1_r = radians(lon1)
lat2_r = radians(lat2)
lon2_r = radians(lon2)
dlon = lon2_r - lon1_r
dlat = lat2_r - lat1_r
a = sin(dlat / 2)**2 + cos(lat1_r) * cos(lat2_r) * sin(dlon / 2)**2
c = 2 * atan2(sqrt(a), sqrt(1 - a))
return R * c
# You are trying to create the most evil schedule on campus, which contains back-to-back classes in buildings that are the furthest apart on campus. You are given the buildings and coordinates of those locations in a dictionary that can be queried as follows:
names_and_coordinates['<NAME>'], names_and_coordinates['<NAME>']
# **Write a function that returns a tuple of the furthest two buildings on campus, and the distance in miles between the two.**
# +
from itertools import combinations
def furthest_walk_on_campus(names_and_coordinates):
def key_fn(entry):
(bname1, (lat1, lon1)), (bname2, (lat2, lon2)) = entry
return haversine(lat1, lon1, lat2, lon2)
(bname1, (lat1, lon1)), (bname2, (lat2, lon2)) = max(
combinations(
names_and_coordinates.items(),
2
),
key=key_fn
)
return bname1, bname2, haversine(lat1, lon1, lat2, lon2)
# -
print('The furthest walk on campus is {0} to {1}, which is {2} miles'.format(*furthest_walk_on_campus(names_and_coordinates)))
# **Extension**: Distance isn't always the best measure of the worst walk. Use Google Maps Matrix API (https://developers.google.com/maps/documentation/distance-matrix/start)'s time to destination function to rank the walks instead.
# <a id='section2'></a>
# ## Q2: Solving Sudoku
# References:
# - http://lipas.uwasa.fi/~timan/sudoku/
# +
def read_board(filename):
with open(filename, 'r') as f:
lines = f.readlines()
board = []
for line in lines:
board.append(line.split(' ')[:-1])
return board[:-1]
read_board('sudoku-puzzles/puzzle1.txt')
# -
def check_solution(original_board, board):
"""Checks that the board has a valid solution based on Sudoku rules.
This does NOT test all of the rules! Don't use it to help you with the other parts.
"""
for i in range(len(original_board)):
for j in range(len(original_board[0])):
if original_board[i][j] != '0' and original_board[i][j] != board[i][j]:
return False
for row in board:
if sum(int(digit) for digit in row) != 45:
return False
for col in range(len(original_board[0])):
if sum([int(original_board[i][col]) for i in range(len(original_board))]) != 45:
return False
return True
# In this problem, you will write a solver for the classic puzzle game, Sudoku. Here are the rules if you are unfamiliar: http://www.counton.org/sudoku/rules-of-sudoku.php
#
# You will be implementing a solver using depth-first backtracking search. Don't worry about how complicated it sounds: it's a fancy way of saying, try random numbers in each box, and if you break the rules, erase and start over.
#
# We will be representing our board as a list of lists, where each list is a row on the Sudoku board. '0' represents a blank space on the board.
#
# **Complete the following functions to create your Sudoku solver.**
def get_first_empty_cell(board):
"""Gets the first empty cell on the board, left to right, up to down"""
for i, row in enumerate(board):
for j, col in enumerate(row):
if board[i][j] == '0':
return i, j
# if none available, return -1, -1
return -1, -1
def is_placement_valid(board, i, j, val):
"""Checks if placing <val> at position (i, j) on the board is a valid move"""
"""Checks if placing <val> at position (i, j) on the board is valid"""
row_count = sum(1 for k in range(9) if board[k][j] == val)
col_count = sum(1 for l in range(9) if board[i][l] == val)
if row_count + col_count > 0:
return False
box_start_i, box_start_j = i - (i % 3), j - (j % 3)
box_count = 0
for i in range(3):
for j in range(3):
if board[box_start_i + i][box_start_j + j] == val:
box_count += 1
return box_count == 0
def solve(board):
"""Solves a Sudoku board: here are the steps:
1. Find an empty cell, if none exist (-1, -1), return board (because all spaces are filled)
2. Find a value to place (try all 9 possible values) in that empty cell
3. Place the value in the empty cell
4. Recursively call solve(board):
a. If a solution is found, return it
b. If no solution is found with this placement, replace the value with '0', and try a new value
5. Return None if there is no value to place
"""
i, j = get_first_empty_cell(board)
if i == -1 and j == -1:
return board
for p in range(1, 10):
if is_placement_valid(board, i, j, str(p)):
board[i][j] = str(p)
if solve(board):
return board
board[i][j] = '0'
return None
# +
# Run this cell to test your code on the sample puzzles
puzzles = ['sudoku-puzzles/puzzle1.txt', 'sudoku-puzzles/puzzle2.txt']
for puzzle in puzzles:
original_board = read_board(puzzle)
print('Testing', puzzle)
if not check_solution(original_board, solve(original_board)):
print('Failed on', puzzle)
# -
# <a id='section3'></a>
# ## Q3: Alien Language
# +
from collections import defaultdict
import random
from numpy import allclose
alphabet = ''.join([chr(97 + i) for i in range(26)])
english_frequencies = {
'a': 0.0892,
'b': 0.0251,
'c': 0.0389,
'd': 0.0274,
'e': 0.1144,
'f': 0.0251,
'g': 0.0228,
'h': 0.0434,
'i': 0.0732,
'k': 0.0091,
'l': 0.0343,
'm': 0.0411,
'n': 0.0549,
'o': 0.0846,
'p': 0.0160,
'q': 0.0022,
'r': 0.0549,
's': 0.0709,
't': 0.0961,
'u': 0.0366,
'v': 0.0091,
'w': 0.0045,
'y': 0.0228,
'z': 0.0022,
}
# -
alphabet
# You've received an alien transmission from above! It might be the answer: to your life, to the planet, to the universe, to everything!
secret = open('alien/secret.txt').readline().strip()
secret
# That's alien language, all right. Fortunately, they've left you the following note:
partial_mapping = {
'f': 'r',
'g': 'f',
'i': 'n',
'n': 'y',
'u': 'z',
'm': 'v'
}
# You realize that this alien language isn't so foreign at all, it is a simple **substitution cipher** of the English language, which means that every letter in the alien language corresponds to exactly one letter in the English alphabet.
#
# As a short-sighted mammal, you believe that alien life is similar to life on Earth. As a minor linguist yourself, you have the frequencies that each English letter appears in literature. To discover the rest of the mapping, you decide to find the frequencies that each letter appears in the alien text, map it to the closest frequency in the English alphabet.
#
# **Complete the following functions to translate the text:**
def get_frequencies(text):
"""Creates a dictionary where each key is a letter of the alphabet, and each corresponding value is
the frequency (number of times the letter appears / total number of letters in the passage).
Don't forget to ignore spaces and punctuation!"""
counts = defaultdict(int)
for c in text:
if c.lower() in alphabet:
counts[c.lower()] += 1
total = sum(counts.values())
return {k: v / total for k, v in counts.items()}
def permute(text, mapping):
"""Converts every letter in <text> to its substitution in <mapping>, and return the mapped text"""
def switch(c):
if c.lower() not in alphabet:
return c
return mapping[c.lower()]
return ''.join([switch(c) for c in text])
def get_mapping(partial_mapping, english_frequencies, alien_frequencies):
"""Complete <partial_mapping> by matching each unmapped letter in <alien_frequencies> to the corresponding
English letter with closest frequencies"""
mapping = partial_mapping.copy()
for c, freq in alien_frequencies.items():
if c not in partial_mapping:
mapping[c] = min(english_frequencies, key=lambda english_c: abs(english_frequencies[english_c] - freq))
return mapping
def translate(alien_text, partial_mapping):
"""Translate the alien text, by completing the mapping and substituting."""
alien_frequencies = get_frequencies(alien_text)
mapping = get_mapping(partial_mapping, english_frequencies, alien_frequencies)
return permute(alien_text, mapping)
print('The answer to life itself is: ', translate(secret, partial_mapping))
| Lectures/Lecture2_Python/intermediate/Python Intermediate (Solution).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EPA Comments
# +
# import packages
import pandas as pd
import numpy as np
import requests
import urllib
import json
import time
import os
# datetime package too: https://docs.python.org/3/library/datetime.html
from datetime import datetime
from datetime import timedelta
# -
# ## API Retrieval
# +
# Specify the path of the folder where the data are saved
filePath = "DESIGNATE_FILE_PATH"
# general variables for setting parameters
APIkey = "INSERT_API_KEY"
rpp = 1000
pageIndex = 0
po = pageIndex * rpp
agency = 'EPA'
# -
# ### Public Submissions: Jan 2020 - June 2020
# +
# variables for setting parameters
# document type = Public Submission
# Rulemaking dockets only
# range: receivedDate
baseURL_PS = "https://api.data.gov/regulations/v3/documents.json?encoded=1&countsOnly=0&dkt=R&so=ASC&sb=postedDate"
dctType = 'PS'
dateRangeStart = '01/01/20'
dateRangeEnd = '06/30/20'
# set parameters to retrieve PS documents
params = {'po': po,
'rpp': rpp,
'api_key': APIkey,
'rd': dateRangeStart+'-'+dateRangeEnd,
'dct': dctType,
'a': agency}
# +
# ----- RETRIEVE COMMENTS ----- #
# retrieve comments using Requests library and check GET request response
dcts_response = requests.get(baseURL_PS, params=params)
RL_remaining = int(dcts_response.headers['X-RateLimit-Remaining'])
print("Status Code: "+str(dcts_response.status_code),
'Request URL: '+str(dcts_response.request.url)+'\n',sep='\n')
# nested list: separate 'documents' from 'totalNumRecords'
# confirm total requested and number of documents retrieved
numPS = dcts_response.json()['totalNumRecords']
dctsPS = dcts_response.json()['documents']
print('Total number of records requested: '+str(numPS), 'Number retrieved: '+str(len(dctsPS)), sep='\n')
# if requested == retrieved, then export as JSON
if len(dctsPS)==numPS:
with open(filePath+'endpoint_documents_PS_2020Jan01_2020May31.json', 'w', encoding='utf-8') as outfile:
json.dump(dctsPS, outfile, ensure_ascii=False, indent=4)
print('Exported as JSON!')
else:
print('\n''Determine how many pages of records need to be combined via the extend method...',
'Start with: '+str(numPS // rpp + 1),
'That would be enough to retrieve '+str(rpp * (numPS // rpp + 1))+' records'
' -- a margin of '+str(rpp * (numPS // rpp + 1) - numPS)+' records.',sep='\n')
# +
# %%time
# define empty object to put extended data
dctsPS_all = []
totalNumPages = numPS//rpp + (1 if (numPS%rpp>0) else 0)
print('Initial length of data: '+str(len(dctsPS_all)))
# define time objects for avoiding rate limit
initialNextTime = datetime.now()
nextAllowableTime = []
pagesPerHour = 1000 ## regulations.gov rate limit of 1000
# fill array of allowable times
for index in range(0,pagesPerHour):
nextAllowableTime.append(initialNextTime)
print('Time array length: '+str(len(nextAllowableTime)))
# retrieve additional pages of documents and extend object
for pageIndex in range (0,totalNumPages): ## remember range is non-inclusive
if RL_remaining < 10:
print('Rate Limit remaining: '+str(RL_remaining),
"sleeping 5 minutes...", sep='\n')
time.sleep(300)
elif (RL_remaining <= 100) & (RL_remaining%25==0):
print('Rate Limit remaining: '+str(RL_remaining))
nextAllowableTimeIndex = pageIndex % pagesPerHour
currentTime = datetime.now()
if pageIndex%100 == 0:
print("nextAllowableTimeIndex = "+str(nextAllowableTimeIndex),
"nextAllowableTime = "+str(nextAllowableTime[nextAllowableTimeIndex]),
"currentTime = "+str(currentTime), sep=" ")
if currentTime < nextAllowableTime[nextAllowableTimeIndex]:
waitTime = nextAllowableTime[nextAllowableTimeIndex] - currentTime
print("sleeping " + str(waitTime.total_seconds()) + " seconds...")
time.sleep(waitTime.total_seconds() + 0.01)
if nextAllowableTime[nextAllowableTimeIndex] <= datetime.now():
nextAllowableTime[nextAllowableTimeIndex] = datetime.now() + timedelta(seconds = 3600) ## add one hour to nextAllowableTime
try:
po = pageIndex * rpp
params.update({'po': po})
temp_response = requests.get(baseURL_PS, params=params)
RL_remaining = int(temp_response.headers['X-RateLimit-Remaining'])
if temp_response.status_code != 200: ## status code = 429 means over rate limit
print('code '+str(temp_response.status_code)+' for page #'+str(pageIndex),
temp_response.text, sep='\n')
data_this_page = temp_response.json()['documents']
dctsPS_all.extend(data_this_page)
if pageIndex%100 == 0:
print("request made (pageIndex = " + str(pageIndex) + ")")
print('Retrieved: '+str(len(dctsPS_all)),'\n')
except:
print('missing page: '+str(pageIndex))
continue
else:
print("request failed")
print("too soon -- breaking (pageIndex = "+str(pageIndex)+")")
break
print('If this works, we should have retrieved all the requested documents: '+str(len(dctsPS_all)))
# -
# if requested == retrieved, then export as JSON
if len(dctsPS_all)==numPS:
dataFile = 'EPA_endpoint_documents_PS_2020Jan_2020Jun.json'
with open(filePath+dataFile, 'w', encoding='utf-8') as outfile:
json.dump(dctsPS_all, outfile, ensure_ascii=False, indent=4)
print('Exported as JSON!')
else:
print('Export unsuccessful. Check your code.')
print('0',dctsPS_all[0],'\n',
'last',dctsPS_all[-1], sep='\n')
# convert to pandas DataFrame
df2020PS = pd.DataFrame(dctsPS_all)
df2020PS.info()
# create column with document URL
df2020PS['documentURL'] = "https://www.regulations.gov/document?D="
df2020PS.loc[:,'documentURL'] = df2020PS['documentURL']+df2020PS['documentId']
print(df2020PS.loc[0,'documentURL'],
df2020PS.loc[1,'documentURL'], sep='\n')
# print columns list -- determine which ones to write to CSV
dfColumns = df2020PS.columns.tolist()
print(dfColumns)
# +
write_columns = ['agencyAcronym','docketId','docketType','rin',
'documentId','documentType','numberOfCommentsReceived','postedDate',
'title','commentText','attachmentCount','documentURL']
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'EPA_endpoint_documents_PS_2020.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
df2020PS.to_csv(outfile, index_label='index', line_terminator='\n', columns=write_columns)
print('Saved as CSV!')
# -
# ## Data Cleaning
# ### Public Submissions data
# +
# load CSV of Public Submissions data
savePath = 'DESIGNATE_FILE_PATH'
fileName = 'EPA_endpoint_documents_PS_2020.csv'
with open(savePath+fileName,'r',encoding='utf-8') as loadfile:
dfPS = pd.read_csv(loadfile, index_col='index')
dfPS.info()
# +
# create column for commentsPosted (==1 for each obs)
dfPS['commentsPosted'] = 1
# rename column
dfPS = dfPS.rename(columns={'numberOfCommentsReceived': 'commentsReceived'})
# create strings that will convert to naive datetime
dfPS['postedDateNaive'] = dfPS['postedDate'].str.slice(start=0,stop=10)
# convert to datetime format
dfPS['dtPosted'] = pd.to_datetime(dfPS['postedDateNaive'], infer_datetime_format=True)
# generate year and month columns
dfPS['postedMonth'] = dfPS['dtPosted'].dt.month
dfPS['postedYear'] = dfPS['dtPosted'].dt.year
dfPS.loc[:,['dtPosted','postedMonth','postedYear','commentsPosted','commentsReceived']]
# -
# #### Get receivedDate for Top 30 Dockets
# from: Export Docket Folder (export all as csv)
Top30Received_by_Docket = pd.pivot_table(dfPS, values=['commentsPosted','commentsReceived','postedMonth'], index=['docketId'],
aggfunc={'commentsPosted': np.sum,
'commentsReceived': np.sum,
'postedMonth': np.max}
).sort_values('commentsReceived', ascending=False).head(30)
Top30Received_by_Docket['docketURL'] = 'https://www.regulations.gov/docket?D='+Top30Received_by_Docket.index
Top30Received_by_Docket['exportURL'] = 'https://www.regulations.gov/exportdocket?docketId='+Top30Received_by_Docket.index
top30DktList = Top30Received_by_Docket.index.tolist()
rdSample = Top30Received_by_Docket.sum(0)['commentsPosted']
print(rdSample)
Top30Received_by_Docket
for n in range(len(top30DktList)):
print('Docket '+str(n)+': '+Top30Received_by_Docket['exportURL'][n])
# +
# set variables outside the for loop
savePath = 'DESIGNATE_FILE_PATH'
docId = []
docType = []
docSub = []
rDate = []
pmDate = []
for dktId in range(len(top30DktList)):
# update dktFile pointer
dktFile = 'DOCKET_'+str(top30DktList[dktId])+'.csv'
# load csv
with open(savePath+dktFile,'r', encoding='utf-8') as loadfile:
dfTopDkt = pd.read_csv(loadfile, skiprows=list(range(0,5)),
usecols=['Document ID','Document Type','Document SubType','Received Date','Post Mark Date'],
dtype={'Document ID': 'str', 'Document Type': 'str', 'Document SubType': 'str'})
# print length of documents for dktId
print(str(top30DktList[dktId])+': '+str(len(dfTopDkt)))
# narrow DataFrame and fix column names
dfTopDkt = dfTopDkt.rename(columns={'Document ID': 'documentId',
'Document Type': 'documentType',
'Document SubType': 'documentSubType',
'Received Date': 'receivedDate',
'Post Mark Date': 'postmarkDate'})
docId.extend(dfTopDkt['documentId'].tolist())
docType.extend(dfTopDkt['documentType'].tolist())
docSub.extend(dfTopDkt['documentSubType'].tolist())
rDate.extend(dfTopDkt['receivedDate'].tolist())
pmDate.extend(dfTopDkt['postmarkDate'].tolist())
# print length of longest list
print(max([len(docId), len(docType), len(docSub), len(rDate), len(pmDate)]),'\n')
dfTopDktcombo = pd.DataFrame(zip(docId, docType, docSub, rDate, pmDate),
columns=['documentId','documentType','documentSubType','receivedDate','postmarkDate'])
# remove obs missing documentId (e.g., "withdrawn" documents)
print(len(dfTopDktcombo))
dfTopDktcombo = dfTopDktcombo[dfTopDktcombo['documentId'].notna()]
print(len(dfTopDktcombo))
# merge dataframes on documentId
print(rdSample,'\n') ## compare length to sample of top 20 dockets
dfPSrd = dfPS.merge(dfTopDktcombo, how='left', on=['documentId'], indicator=True, validate="1:1")
print(dfPSrd['_merge'].value_counts(),'\n')
dfPSrd = dfPSrd.rename(columns={'_merge': '_mergeTop30'}) ## rename _merge column
dfPSrd.info()
# +
print(dfPSrd['documentType_x'].value_counts(),
dfPSrd['documentType_y'].value_counts() ,sep='\n')
dfPSrd = dfPSrd.drop(columns=['documentType_y'], errors='ignore')
# -
# #### Get receivedDate for remaining comments
# from: API document endpoint
# distribution of comments missing receivedDate by month
dfPSrd.loc[dfPSrd['receivedDate'].isna(),
['documentId','postedDate','postedMonth']].groupby('postedMonth').documentId.nunique()
# +
# list of docIds for API request
bool_missingRD = dfPSrd['receivedDate'].isna()
missingRD = dfPSrd.loc[bool_missingRD,'documentId'].tolist()
print(bool_missingRD.value_counts(),
len(missingRD), sep='\n')
# +
# ----- Create new DataFrame for Remaining Comments ----- #
# ----- Retrieve receivedDates for comments ----- #
import requests
# general variables for setting parameters
APIkey = "INSERT_API_KEY"
baseURL = "https://api.data.gov:443/regulations/v3/document.json?"
dctId = ""
# set parameters
params = {'api_key': APIkey,
'documentId': dctId}
rangeRD = len(missingRD)
listRD = [] # list for adding receivedDate of each entry
# retrieve comments using Requests library and check GET request response
for d in range(rangeRD):
dctId = missingRD[d]
params.update({'documentId': dctId})
dct_response = requests.get(baseURL, params=params)
RL_remaining = int(dct_response.headers['X-RateLimit-Remaining'])
if dct_response.status_code != 200:
print('code '+str(dct_response.status_code)+' for page #'+str(pageIndex),
dct_response.text, sep='\n')
if RL_remaining < 10:
print('Rate Limit remaining: '+str(RL_remaining),
"sleeping 1 minute...", sep='\n')
time.sleep(60)
this_receivedDate = dct_response.json()['receivedDate']
listRD.append(this_receivedDate)
if d%100==0:
print("Number of comments retrieved: "+str(d))
print('Length of receivedDate list is '+str(len(listRD)))
# ----- Generate df from the lists ----- #
remainingList = list(zip(missingRD, listRD))
dfRemaining = pd.DataFrame(remainingList, columns = ['documentId', 'receivedDate'])
dfRemaining.info()
# -
# #### Concatenate and Merge DataFrames
# +
# concatenate dfs so we have one df with receivedDate
dfRD = pd.concat([dfTopDktcombo, dfRemaining],
axis=0, join='outer', ignore_index=True, verify_integrity=True)
# merge dataframes on documentId
dfPS2020 = dfPS.merge(dfRD, how='left', on=['documentId'], indicator=True, validate="1:1")
print(dfPS2020['_merge'].value_counts(),'\n')
dfPS2020 = dfPS2020.rename(columns={'_merge': '_mergeRD'})
dfPS2020.info()
# +
# convert to datetime format
dfPS2020['receivedDateNaive'] = dfPS2020['receivedDate'].str.slice(start=0,stop=10)
dfPS2020['dtReceived'] = pd.to_datetime(dfPS2020['receivedDateNaive'])
dfPS2020['receivedMonth'] = dfPS2020['dtReceived'].dt.month
dfPS2020.loc[:,['receivedDate','dtReceived','receivedMonth']]
# -
pd.pivot_table(dfPS2020, values=['commentsPosted'], index=['receivedMonth'], columns=['postedMonth'],
aggfunc={'commentsPosted': np.sum}, fill_value=0, margins=True)
# #### Drop Select Documents
# +
# create list for dropping PS documents
dropPS = []
### remove documents posted before 2020
dropPS = dfPS2020[dfPS2020['dtPosted']<datetime(2020,1,1)].index.tolist()
print(len(dropPS),'\n')
### remove documents posted in July 2020
dropPS.extend(dfPS2020[dfPS2020['dtReceived']>=datetime(2020,7,1)].index.tolist())
print(len(dropPS),'\n')
# drop entries
print(len(dfPS2020))
dfPS2020 = dfPS2020.drop(index=dropPS, errors='ignore') ## ignore → only existing labels are dropped
print(len(dfPS2020))
# -
# #### Results: Clean Public Submissions Dataset
pd.pivot_table(dfPS2020, values=['commentsPosted','commentsReceived'], index=['receivedMonth'],
aggfunc=np.sum, fill_value=0, margins=True)
# https://stackoverflow.com/questions/15411158/pandas-countdistinct-equivalent
print(dfPS2020.groupby('postedMonth').docketId.nunique(),
dfPS2020.groupby('receivedMonth').docketId.nunique(), sep='\n')
# +
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'EPA_cleaned_PS_2020.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
dfPS2020.to_csv(outfile, index_label='index', line_terminator='\n')
print('Saved as CSV!')
# -
# ## Data for Analysis
# +
# load CSV of Public Submissions data
savePath = 'DESIGNATE_FILE_PATH'
fileName = 'EPA_cleaned_PS_2020.csv'
with open(savePath+fileName,'r',encoding='utf-8') as loadfile:
dfPS2020 = pd.read_csv(loadfile, index_col='index')
dfPS2020.info()
# -
# create filter for Science Rule
bool_science = dfPS2020['docketId']=='EPA-HQ-OA-2018-0259'
dfPS2020.loc[bool_science,'scienceRule'] = 1
dfPS2020.loc[~bool_science,'scienceRule'] = 0
dfPS2020['scienceRule'].value_counts()
# ### Public Submissions per Month
print(len(dfPS2020.groupby(['receivedMonth','docketId'])))
print(sum(dfPS2020.groupby('receivedMonth').docketId.nunique().tolist()))
# #### Include all dockets
# +
# create new DataFrame with sum of comments per month
dfCommentsMonthly = dfPS2020.groupby('receivedMonth')[['commentsPosted','commentsReceived']].sum().reset_index()
# add new column with unique dockets receiving comments per month
dfCommentsMonthly['docketsUnique'] = dfPS2020.groupby('receivedMonth')['docketId'].nunique().tolist()
# add column for month labels
dfCommentsMonthly.insert(1,'labelMonth',['Jan','Feb','Mar','Apr','May','Jun'])
# rename columns
dfCommentsMonthly = dfCommentsMonthly.rename(columns={'commentsPosted': 'commentsUnique', 'commentsReceived': 'commentsAll'})
# calculate two new columns: comments per unique dockets
dfCommentsMonthly['unq_per_dkts'] = dfCommentsMonthly['commentsUnique']/dfCommentsMonthly['docketsUnique']
dfCommentsMonthly['all_per_dkts'] = dfCommentsMonthly['commentsAll']/dfCommentsMonthly['docketsUnique']
# view returned df
dfCommentsMonthly
# +
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'data_for_analysis_monthly.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
dfCommentsMonthly.to_csv(outfile, line_terminator='\n', index=False)
print('Saved as CSV!')
# -
# #### Science rule filter
# +
# create new DataFrame with sum of comments per month
dfCommentsMonthly_filterSR = dfPS2020.groupby(['scienceRule','receivedMonth'])[['commentsPosted','commentsReceived']].sum().reset_index()
# add new column with unique dockets receiving comments per month
dfCommentsMonthly_filterSR['docketsUnique'] = dfPS2020.groupby(['scienceRule','receivedMonth'])['docketId'].nunique().tolist()
# add column for month labels
dfCommentsMonthly_filterSR.insert(2,'labelMonth',['Jan','Feb','Mar','Apr','May','Jun','Jan','Mar','Apr','May','Jun'])
# rename columns
dfCommentsMonthly_filterSR = dfCommentsMonthly_filterSR.rename(columns={'commentsPosted': 'commentsUnique', 'commentsReceived': 'commentsAll'})
# calculate two new columns: comments per unique dockets
exclude_science_rule = dfCommentsMonthly_filterSR['scienceRule']==0
dfCommentsMonthly_filterSR.loc[exclude_science_rule,'unq_per_dkts'] = dfCommentsMonthly_filterSR['commentsUnique']/dfCommentsMonthly_filterSR['docketsUnique']
dfCommentsMonthly_filterSR.loc[exclude_science_rule,'all_per_dkts'] = dfCommentsMonthly_filterSR['commentsAll']/dfCommentsMonthly_filterSR['docketsUnique']
# view returned df
dfCommentsMonthly_filterSR
# +
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'data_for_analysis_monthly_filterSR.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
dfCommentsMonthly_filterSR.to_csv(outfile, line_terminator='\n', index=False)
print('Saved as CSV!')
# -
# ### Public Submissions per Day
# +
print(len(dfPS2020.groupby('dtReceived')[['commentsPosted','commentsReceived']]))
print(len(dfPS2020.groupby('dtReceived')['docketId'].nunique().tolist()))
print(len(dfPS2020.groupby(['scienceRule','dtReceived'])[['commentsPosted','commentsReceived']]))
print(len(dfPS2020.groupby(['scienceRule','dtReceived'])['docketId'].nunique().tolist()))
# -
# #### Include all dockets
# +
# create new DataFrame with sum of comments per month
dfCommentsDaily = dfPS2020.groupby('dtReceived')[['commentsPosted','commentsReceived']].agg(np.sum).reset_index()
# add new column with unique dockets receiving comments per month
dfCommentsDaily['docketsUnique'] = dfPS2020.groupby('dtReceived')['docketId'].nunique().tolist()
# rename columns
dfCommentsDaily = dfCommentsDaily.rename(columns={'commentsPosted': 'commentsUnique', 'commentsReceived': 'commentsAll'})
# calculate two new columns: comments per unique dockets
dfCommentsDaily['unq_per_dkts'] = dfCommentsDaily['commentsUnique']/dfCommentsDaily['docketsUnique']
dfCommentsDaily['all_per_dkts'] = dfCommentsDaily['commentsAll']/dfCommentsDaily['docketsUnique']
# view returned df
dfCommentsDaily
# +
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'data_for_analysis_daily.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
dfCommentsDaily.to_csv(outfile, line_terminator='\n', index=False)
print('Saved as CSV!')
# -
# #### Science rule filter
# +
# create new DataFrame with sum of comments per month
dfCommentsDaily_filterSR = dfPS2020.groupby(['scienceRule','dtReceived'])[['commentsPosted','commentsReceived']].agg(np.sum).reset_index()
# add new column with unique dockets receiving comments per month
dfCommentsDaily_filterSR['docketsUnique'] = dfPS2020.groupby(['scienceRule','dtReceived'])['docketId'].nunique().tolist()
# rename columns
dfCommentsDaily_filterSR = dfCommentsDaily_filterSR.rename(columns={'commentsPosted': 'commentsUnique', 'commentsReceived': 'commentsAll'})
# calculate two new columns: comments per unique dockets
exclude_science_rule = dfCommentsDaily_filterSR['scienceRule']==0
dfCommentsDaily_filterSR.loc[exclude_science_rule,'unq_per_dkts'] = dfCommentsDaily_filterSR['commentsUnique']/dfCommentsDaily_filterSR['docketsUnique']
dfCommentsDaily_filterSR.loc[exclude_science_rule,'all_per_dkts'] = dfCommentsDaily_filterSR['commentsAll']/dfCommentsDaily_filterSR['docketsUnique']
# view returned df
dfCommentsDaily_filterSR
# +
savePath = 'DESIGNATE_FILE_PATH'
saveFile = 'data_for_analysis_daily_filterSR.csv'
# write to csv, reference: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
with open(savePath+saveFile, 'w', encoding='utf-8') as outfile:
dfCommentsDaily_filterSR.to_csv(outfile, line_terminator='\n', index=False)
print('Saved as CSV!')
# -
# ## Docket Analysis
# +
# load CSV of Public Submissions data
savePath = 'DESIGNATE_FILE_PATH'
fileName = 'EPA_cleaned_PS_2020.csv'
with open(savePath+fileName,'r',encoding='utf-8') as loadfile:
dfPS2020 = pd.read_csv(loadfile, index_col='index')
dfPS2020.info()
# -
# ### Top 5 Dockets
# view top 5 dockets in terms of total comments
dfTop5Dkts = pd.pivot_table(dfPS2020, values=['commentsPosted','commentsReceived'], index=['docketId'],
aggfunc=np.sum, fill_value=0).sort_values('commentsReceived', ascending=False).head(5)
dfTop5Dkts
# +
# view commentsReceived per month on top 5 dockets
select_dockets = dfTop5Dkts.index.tolist()
bool_select = [True if doc in select_dockets else False for doc in dfPS2020['docketId'].tolist()]
print(bool_select.count(True))
pd.pivot_table(dfPS2020[bool_select], values=['commentsPosted','commentsReceived'], index=['docketId','receivedMonth'],
aggfunc=np.sum, fill_value=0)
| API Retrieval/Public Commenting and COVID-19 - EPA Comments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Emulator: Gaussian Process (`george`)
#
# #### Index<a name="index"></a>
# 1. [Import packages](#imports)
# 2. [Load data](#loadData)
# 1. [Load train data](#loadTrainData)
# 2. [Load test data](#loadTestData)
# 3. [Emulator method](#emulator)
# 1. [Scale data](#scaleData)
# 2. [Train emulator](#trainEmu)
# 3. [Plot results](#plotEmu)
#
# ## 1. Import packages<a name="imports"></a>
# +
import george
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pickle
import scipy.optimize as op
import seaborn as sns
from sklearn.preprocessing import StandardScaler
# -
# #### Aesthetic settings
# +
# %matplotlib inline
sns.set(font_scale=1.3, style="ticks")
# -
# ## 2. Load data<a name="loadData"></a>
#
# Read the training data from a `.npy` file:
#
# ### 2.1. Load train data<a name="loadTrainData"></a>
# For the full demo, we'll use 1d data (a single input parameter/feature), but you can also try it the full 3d data; this just takes a long time to train, so you might want to load in our already saved results below to view it. Remember to load in the corresponding test data below.
path_train = '../data/cosmology_train_1d.pickle'
#path_train = '../data/cosmology_train.pickle'
#path_train = '../data/cosmology_train_big.pickle'
with open(path_train, 'rb') as input_file:
data_train = pickle.load(input_file)
input_train = data_train['input_data']
output_train = data_train['output_data']
number_train = input_train.shape[0]
number_param = input_train.shape[1] - 1
n_values = output_train.shape[1]-1
print("Number of datapoints:", number_train)
print("Number of input parameters:", number_param) # remove the `object_id` column
extra_train = data_train['extra_input']
r_vals = extra_train['r_vals']
xs_train = input_train.drop(columns=['object_id'])
ys_train = output_train.drop(columns=['object_id'])
# ### 2.2. Load test data<a name="loadTestData"></a>
path_test = '../data/cosmology_test_1d.pickle'
#path_test = '../data/cosmology_test.pickle'
with open(path_test, 'rb') as input_file:
data_test = pickle.load(input_file)
input_test = data_test['input_data']
output_test = data_test['output_data']
number_test = input_test.shape[0]
print("Number of datapoints:", number_test)
xs_test = input_test.drop(columns=['object_id'])
ys_test = output_test.drop(columns=['object_id'])
# ## 3. Emulator method<a name="emulator"></a>
#
# ### 3.1. Scale data<a name="scaleData"></a>
#
# Let's first scale our input parameters, to make training easier:
scaler = StandardScaler()
scaler.fit(xs_train)
xs_train.iloc[:] = scaler.transform(xs_train)
xs_test.iloc[:] = scaler.transform(xs_test)
y_mean = np.mean(ys_train, axis=0)
ys_train = ys_train/y_mean
ys_test = ys_test/y_mean
# ### 3.2. Train emulator<a name="trainEmu"></a>
def fit_gp(kernel, xs, ys, xs_new):
def neg_log_like(p): # Objective function: negative log-likelihood
gp.set_parameter_vector(p)
loglike = gp.log_likelihood(ys, quiet=True)
return -loglike if np.isfinite(loglike) else 1e25
def grad_neg_log_like(p): # Gradient of the objective function.
gp.set_parameter_vector(p)
return -gp.grad_log_likelihood(ys, quiet=True)
gp = george.GP(kernel)
gp.compute(xs)
results = op.minimize(neg_log_like, gp.get_parameter_vector(),
jac=grad_neg_log_like, method="L-BFGS-B", tol=1e-6)
gp.set_parameter_vector(results.x)
gp_mean, gp_cov = gp.predict(ys, xs_new)
return gp_mean
# Here we are going to train and predict the value straight away. (If you're loading in saved results, comment out the next 2 cells.)
number_outputs = np.shape(ys_test)[1]
print(number_outputs)
ys_test_preds = ys_test.copy()
ys_train_0 = ys_train.iloc[:, 0]
for i in np.arange(number_outputs):
print(i)
ys_train_i = ys_train.iloc[:, i]
kernel = np.var(ys_train_0) * george.kernels.ExpSquaredKernel(0.5, ndim=number_param)
ys_pred = fit_gp(kernel=kernel, xs=xs_train,
ys=ys_train_i, xs_new=xs_test)
ys_test_preds.iloc[:, i] = ys_pred
# Undo all the normalizations.
ys_test = ys_test*y_mean
ys_test_preds = ys_test_preds*y_mean
# Save results. (Commented out as results have already been saved.)
path_save_results = f'emulator_results/output_pred_big_train_{number_param}d.pickle'
#ys_test_preds.to_pickle(path_save_results)
# Verify the results were well saved. (If you're looking at the 3d data, you'll want to load this in here.)
# +
#ys_test_preds_saved = pd.read_pickle(path_save_results)
#np.allclose(ys_test_preds_saved, ys_test_preds)
#ys_test_preds = ys_test_preds_saved
# -
# ### 3.3. Plot results<a name="plotEmu"></a>
#
# We compare our predictions to the truth (choosing a subset for visual clarity).
n_plot = int(0.2*number_test)
idxs = np.random.choice(np.arange(number_test), n_plot)
color_idx = np.linspace(0, 1, n_plot)
colors = np.array([plt.cm.rainbow(c) for c in color_idx])
plt.figure(figsize=(8,6))
for i in range(n_plot):
ys_test_i = ys_test.iloc[idxs[i], :]
ys_pred_i = ys_test_preds.iloc[idxs[i], :]
if i==0:
label_test = 'truth'
label_pred = 'emu_prediction'
else:
label_test = None
label_pred = None
plt.plot(r_vals, ys_test_i, alpha=0.8, label=label_test,
marker='o', markerfacecolor='None', ls='None', color=colors[i])
plt.plot(r_vals, ys_pred_i, alpha=0.8, label=label_pred, color=colors[i])
plt.xlabel('$r$')
plt.ylabel(r'$\xi(r)$')
plt.legend()
# We plot the fractional error of all test set statistics:
color_idx = np.linspace(0, 1, number_test)
colors = np.array([plt.cm.rainbow(c) for c in color_idx])
plt.figure(figsize=(8,6))
frac_errs = np.empty((number_test, n_values))
for i in range(number_test):
ys_test_i = ys_test.iloc[i, :]
ys_pred_i = ys_test_preds.iloc[i, :]
frac_err = (ys_pred_i-ys_test_i)/ys_test_i
frac_errs[i] = frac_err
plt.plot(r_vals, frac_err, alpha=0.8, color=colors[i])
plt.axhline(0.0, color='k')
plt.xlabel('$r$')
plt.ylabel(r'fractional error')
# We show the spread of these fractional errors:
# +
color_idx = np.linspace(0, 1, number_test)
colors = np.array([plt.cm.rainbow(c) for c in color_idx])
plt.figure(figsize=(8,6))
frac_errs_stdev = np.std(frac_errs, axis=0)
plt.plot(r_vals, frac_errs_stdev, alpha=0.8, color='blue', label='standard deviation')
frac_errs_p16 = np.percentile(frac_errs, 16, axis=0)
frac_errs_p84 = np.percentile(frac_errs, 84, axis=0)
frac_errs_percentile = np.mean([np.abs(frac_errs_p16), np.abs(frac_errs_p84)], axis=0)
plt.plot(r_vals, frac_errs_percentile, alpha=0.8, color='green', label="mean of 16/84 percentile")
plt.xlabel('$r$')
plt.ylabel(r'spread of fractional errors')
plt.legend()
# -
# The GP is doing incredibly well at accurately emulating the correlation function!
# [Go back to top.](#index)
| emulator_examples/emulator_gp_george.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Converting a Keras model to ONNX
#
# In the steps that follow, you will convert Keras model you just trained to the ONNX format. This will enable you to use this model for classification in a very broad range of environments, outside of Azure Databricks including:
#
# - Web services
# - iOS and Android mobile apps
# - Windows apps
# - IoT devices
#
# Furthermore, ONNX runtimes and libraries are also designed to maximize performance on some of the best hardware in the industry. In this lab, we will compare the Inference performance of the ONNX vs Keras models.
#
# First we will load the trained Keras model from file, and then convert the model to ONNX.
# ## Load the Keras Model
#
# Load the saved Keras model. We will convert the Keras model to ONNX format.
# +
import os
import numpy as np
import pandas as pd
np.random.seed(125)
from keras.models import load_model
import joblib
output_folder = './output'
model_filename = 'final_model.hdf5'
keras_model = load_model(os.path.join(output_folder, model_filename))
print(keras_model.summary())
# -
# ## Convert to ONNX
#
# Convert the loaded Keras model to ONNX format, and save the ONNX model to the deployment folder.
# +
import onnxmltools
deployment_folder = 'deploy'
onnx_export_folder = 'onnx'
# Convert the Keras model to ONNX
onnx_model_name = 'claim_classifier.onnx'
converted_model = onnxmltools.convert_keras(keras_model, onnx_model_name, target_opset=7)
# Save the model locally...
onnx_model_path = os.path.join(deployment_folder, onnx_export_folder)
os.makedirs(onnx_model_path, exist_ok=True)
onnxmltools.utils.save_model(converted_model, os.path.join(onnx_model_path,onnx_model_name))
# -
# ## Make Inference using the ONNX Model
#
# - Create an ONNX runtime InferenceSession
# - Review the expected input shape to make inferences
# - Prepare test data
# - Make inferences using both the ONNX and the Keras Model on the test data
# ### ONNX Runtime InferenceSession
import onnxruntime
# Load the ONNX model and observe the expected input shape
onnx_session = onnxruntime.InferenceSession(
os.path.join(os.path.join(deployment_folder, onnx_export_folder), onnx_model_name))
input_name = onnx_session.get_inputs()[0].name
output_name = onnx_session.get_outputs()[0].name
print('Expected input shape: ', onnx_session.get_inputs()[0].shape)
# ### Prepare test data
# **Load the GloVe word vectors**
# +
word_vectors_dir = './word_vectors'
dictonary = np.load(os.path.join(word_vectors_dir, 'wordsList.npy'))
dictonary = dictonary.tolist()
dictonary = [word.decode('UTF-8') for word in dictonary]
print('Loaded the dictonary! Dictonary size: ', len(dictonary))
word_vectors = np.load(os.path.join(word_vectors_dir, 'wordVectors.npy'))
print ('Loaded the word vectors! Shape of the word vectors: ', word_vectors.shape)
# -
# **Create the word contractions map**
contractions_url = ('https://quickstartsws9073123377.blob.core.windows.net/'
'azureml-blobstore-0d1c4218-a5f9-418b-bf55-902b65277b85/glove50d/contractions.xlsx')
contractions_df = pd.read_excel(contractions_url)
contractions = dict(zip(contractions_df.original, contractions_df.expanded))
# **Setup the helper functions to process the test data**
# +
import re
import string
def remove_special_characters(token):
pattern = re.compile('[{}]'.format(re.escape(string.punctuation)))
filtered_token = pattern.sub('', token)
return filtered_token
def convert_to_indices(corpus, dictonary, c_map, unk_word_index = 399999):
sequences = []
for i in range(len(corpus)):
tokens = corpus[i].split()
sequence = []
for word in tokens:
word = word.lower()
if word in c_map:
resolved_words = c_map[word].split()
for resolved_word in resolved_words:
try:
word_index = dictonary.index(resolved_word)
sequence.append(word_index)
except ValueError:
sequence.append(unk_word_index) #Vector for unkown words
else:
try:
clean_word = remove_special_characters(word)
if len(clean_word) > 0:
word_index = dictonary.index(clean_word)
sequence.append(word_index)
except ValueError:
sequence.append(unk_word_index) #Vector for unkown words
sequences.append(sequence)
return sequences
# -
# **Preprocess the test data**
# +
from keras.preprocessing.sequence import pad_sequences
maxSeqLength = 125
test_claim = ['I crashed my car into a pole.']
test_claim_indices = convert_to_indices(test_claim, dictonary, contractions)
test_data = pad_sequences(test_claim_indices, maxlen=maxSeqLength, padding='pre', truncating='post')
# convert the data type to float
test_data_float = np.reshape(test_data.astype(np.float32), (1,maxSeqLength))
# -
# ### Make Inferences
#
# Make inferences using both the ONNX and the Keras Model on the test data
# +
# Run an ONNX session to classify the sample.
print('ONNX prediction: ', onnx_session.run([output_name], {input_name : test_data_float}))
# Use Keras to make predictions on the same sample
print('Keras prediction: ', keras_model.predict(test_data_float))
# -
# ## Compare Inference Performance: ONNX vs Keras
#
# Evaluate the performance of ONNX and Keras by running the same sample 1,000 times. Run the next three cells and compare the performance in your environment.
# Next we will compare the performance of ONNX vs Keras
import timeit
n = 1000
start_time = timeit.default_timer()
for i in range(n):
keras_model.predict(test_data_float)
keras_elapsed = timeit.default_timer() - start_time
print('Keras performance: ', keras_elapsed)
start_time = timeit.default_timer()
for i in range(n):
onnx_session.run([output_name], {input_name : test_data_float})
onnx_elapsed = timeit.default_timer() - start_time
print('ONNX performance: ', onnx_elapsed)
print('ONNX is about {} times faster than Keras'.format(round(keras_elapsed/onnx_elapsed)))
# # Deploy ONNX model to Azure Container Instance (ACI)
# ## Create and connect to an Azure Machine Learning Workspace
#
# Review the workspace config file saved in the previous notebook.
# !cat .azureml/config.json
# **Create the `Workspace` from the saved config file**
# +
import azureml.core
print(azureml.core.VERSION)
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
print(ws)
# -
# ## Register the model with Azure Machine Learning
#
# In the following, you register the model with Azure Machine Learning (which saves a copy in the cloud).
# +
#Register the model and vectorizer
from azureml.core.model import Model
registered_model_name = 'claim_classifier_onnx'
onnx_model_path = os.path.join(os.path.join(deployment_folder, onnx_export_folder), onnx_model_name)
registered_model = Model.register(model_path = onnx_model_path, # this points to a local file
model_name = registered_model_name, # this is the name the model is registered with
description = "Claims classification model.",
workspace = ws)
print(registered_model.name, registered_model.description, registered_model.version)
# -
# ## Create the scoring web service
#
# When deploying models for scoring with Azure Machine Learning services, you need to define the code for a simple web service that will load your model and use it for scoring. By convention this service has two methods init which loads the model and run which scores data using the loaded model.
#
# This scoring service code will later be deployed inside of a specially prepared Docker container.
# **Save the scoring web service Python file**
#
# Note that the scoring web service needs the registered model: the ONNX model to make inferences.
# +
# %%writefile scoring_service.py
import string
import re
import os
import numpy as np
import pandas as pd
import urllib.request
import json
import keras
from keras.preprocessing.sequence import pad_sequences
import tensorflow as tf
from azureml.core.model import Model
import onnxruntime
def init():
global onnx_session
global dictonary
global contractions
try:
words_list_url = ('https://quickstartsws9073123377.blob.core.windows.net/'
'azureml-blobstore-0d1c4218-a5f9-418b-bf55-902b65277b85/glove50d/wordsList.npy')
word_vectors_dir = './word_vectors'
os.makedirs(word_vectors_dir, exist_ok=True)
urllib.request.urlretrieve(words_list_url, os.path.join(word_vectors_dir, 'wordsList.npy'))
dictonary = np.load(os.path.join(word_vectors_dir, 'wordsList.npy'))
dictonary = dictonary.tolist()
dictonary = [word.decode('UTF-8') for word in dictonary]
print('Loaded the dictonary! Dictonary size: ', len(dictonary))
contractions_url = ('https://quickstartsws9073123377.blob.core.windows.net/'
'azureml-blobstore-0d1c4218-a5f9-418b-bf55-902b65277b85/glove50d/contractions.xlsx')
contractions_df = pd.read_excel(contractions_url)
contractions = dict(zip(contractions_df.original, contractions_df.expanded))
print('Loaded contractions!')
# Retrieve the path to the model file using the model name
onnx_model_name = 'claim_classifier_onnx'
onnx_model_path = Model.get_model_path(onnx_model_name)
print('onnx_model_path: ', onnx_model_path)
onnx_session = onnxruntime.InferenceSession(onnx_model_path)
print('Onnx Inference Session Created!')
except Exception as e:
print(e)
def remove_special_characters(token):
pattern = re.compile('[{}]'.format(re.escape(string.punctuation)))
filtered_token = pattern.sub('', token)
return filtered_token
def convert_to_indices(corpus, dictonary, c_map, unk_word_index = 399999):
sequences = []
for i in range(len(corpus)):
tokens = corpus[i].split()
sequence = []
for word in tokens:
word = word.lower()
if word in c_map:
resolved_words = c_map[word].split()
for resolved_word in resolved_words:
try:
word_index = dictonary.index(resolved_word)
sequence.append(word_index)
except ValueError:
sequence.append(unk_word_index) #Vector for unkown words
else:
try:
clean_word = remove_special_characters(word)
if len(clean_word) > 0:
word_index = dictonary.index(clean_word)
sequence.append(word_index)
except ValueError:
sequence.append(unk_word_index) #Vector for unkown words
sequences.append(sequence)
return sequences
def run(raw_data):
try:
print("Received input: ", raw_data)
maxSeqLength = 125
print('Processing input...')
input_data_raw = np.array(json.loads(raw_data))
input_data_indices = convert_to_indices(input_data_raw, dictonary, contractions)
input_data_padded = pad_sequences(input_data_indices, maxlen=maxSeqLength, padding='pre', truncating='post')
# convert the data type to float
input_data = np.reshape(input_data_padded.astype(np.float32), (1,maxSeqLength))
print('Done processing input.')
# Run an ONNX session to classify the input.
result = onnx_session.run(None, {onnx_session.get_inputs()[0].name: input_data})[0].argmax(axis=1).item()
# return just the classification index (0 or 1)
return result
except Exception as e:
print(e)
error = str(e)
return error
# -
# ## Package Model and deploy to ACI
#
# Your scoring service can have dependencies install by using a Conda environment file. Items listed in this file will be conda or pip installed within the Docker container that is created and thus be available to your scoring web service logic.
#
# The recommended deployment pattern is to create a deployment configuration object with the `deploy_configuration` method and then use it with the deploy method of the [Model](https://docs.microsoft.com/python/api/azureml-core/azureml.core.model.model?view=azure-ml-py) class as performed below. In this case, we use the `AciWebservice`'s `deploy_configuration` and specify the CPU cores and memory size.
#
# You will see output similar to the following when your web service is ready: `Succeeded - ACI service creation operation finished, operation "Succeeded"`
#
# Run the following cell. This may take between 5-10 minutes to complete.
# +
# create a Conda dependencies environment file
print("Creating conda dependencies file locally...")
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core import Environment
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice, Webservice
conda_packages = ['numpy==1.16.4', 'xlrd==1.2.0', 'pandas==0.25.1', 'scikit-learn==0.21.3']
pip_packages = ['azureml-defaults', 'azureml-sdk', 'tensorflow==1.13.1', 'keras==2.3.1', 'onnxruntime==1.0.0']
environment = Environment('my-environment')
environment.python.conda_dependencies = CondaDependencies.create(conda_packages=conda_packages, pip_packages=pip_packages)
execution_script = 'scoring_service.py'
service_name = "claimclassservice"
inference_config = InferenceConfig(entry_script=execution_script, environment=environment)
aci_config = AciWebservice.deploy_configuration(
cpu_cores=1,
memory_gb=1,
tags = {'name': 'Claim Classification'},
description = "Classifies a claim as home or auto.")
service = Model.deploy(workspace=ws,
name=service_name,
models=[registered_model],
inference_config=inference_config,
deployment_config=aci_config)
# wait for the deployment to finish
service.wait_for_deployment(show_output=True)
# -
# ## Test Deployment
# ### Make direct calls on the service object
# +
import json
test_claims = ['I crashed my car into a pole.',
'The flood ruined my house.',
'I lost control of my car and fell in the river.']
for i in range(len(test_claims)):
result = service.run(json.dumps([test_claims[i]]))
print('Predicted label for test claim #{} is {}'.format(i+1, result))
# -
# ### Make HTTP calls to test the deployed Web Service
#
# In order to call the service from a REST client, you need to acquire the scoring URI. Take a note of printed scoring URI, you will need it in the last notebook.
#
# The default settings used in deploying this service result in a service that does not require authentication, so the scoring URI is the only value you need to call this service.
# +
import requests
url = service.scoring_uri
print('ACI Service: Claim Classification scoring URI is: {}'.format(url))
headers = {'Content-Type':'application/json'}
for i in range(len(test_claims)):
response = requests.post(url, json.dumps([test_claims[i]]), headers=headers)
print('Predicted label for test claim #{} is {}'.format(i+1, response.text))
| Hands-on lab/notebooks/04 Deploy Classifier Web Service.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:DSO570]
# language: python
# name: conda-env-DSO570-py
# ---
from gensim.test.utils import common_texts, get_tmpfile # get_tmpfile: save a model for us
from gensim.models import Word2Vec
from nltk import word_tokenize
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
data = pd.read_csv("occ_cleaned.csv")
data.head()
# +
import nltk
from nltk.tokenize import RegexpTokenizer
from nltk.stem import WordNetLemmatizer,PorterStemmer
from nltk.corpus import stopwords
import re
lemmatizer = WordNetLemmatizer()
stemmer = PorterStemmer()
def preprocess(sentence):
sentence=str(sentence)
sentence = sentence.lower()
cleanr = re.compile('<.*?>')
cleantext = re.sub(cleanr, '', sentence)
rem_num = re.sub('[0-9]+', '', cleantext)
tokenizer = RegexpTokenizer(r'\w+')
tokens = tokenizer.tokenize(rem_num)
filtered_words = [w for w in tokens if len(w) > 2 if not w in stopwords.words('english')]
stem_words=[stemmer.stem(w) for w in filtered_words]
return " ".join(stem_words)
data['cleanText']=data['description'].map(lambda s:preprocess(s))
# -
pd.set_option('max_colwidth', 400)
data[['description','cleanText']].head()
data.shape
data.describe()
data.attribute_value.unique()
all_value = data.groupby('product_id')['attribute_value'].unique().reset_index()
# ## label=1
wkd = all_value[all_value['attribute_value'].apply(lambda x: 'weekend' in x)]
wkd = pd.merge(wkd, data, how='left', on='product_id')
wkd = wkd.iloc[:,3:]
wkd.columns = ['label','brand','description','details','name']
wkd = wkd[wkd.label=='weekend']
wkd.drop_duplicates(inplace=True)
dtn = all_value[all_value['attribute_value'].apply(lambda x: 'daytonight' in x)]
dtn = pd.merge(dtn, data, how='left', on='product_id')
dtn = dtn.iloc[:,3:]
dtn.columns = ['label','brand','description','details','name']
dtn = dtn[dtn.label=='daytonight']
dtn.drop_duplicates(inplace=True)
work = all_value[all_value['attribute_value'].apply(lambda x: 'work' in x)]
work = pd.merge(work, data, how='left', on='product_id')
work = work.iloc[:,3:]
work.columns = ['label','brand','description','details','name']
work = work[work.label=='work']
work.drop_duplicates(inplace=True)
ntot = all_value[all_value['attribute_value'].apply(lambda x: 'nightout' in x)]
ntot = pd.merge(ntot, data, how='left', on='product_id')
ntot = ntot.iloc[:,3:]
ntot.columns = ['label','brand','description','details','name']
ntot = ntot[ntot.label=='nightout']
ntot.drop_duplicates(inplace=True)
vc = all_value[all_value['attribute_value'].apply(lambda x: 'vacation' in x)]
vc = pd.merge(vc, data, how='left', on='product_id')
vc = vc.iloc[:,3:]
vc.columns = ['label','brand','description','details','name']
vc = vc[vc.label=='vacation']
vc.drop_duplicates(inplace=True)
wkot = all_value[all_value['attribute_value'].apply(lambda x: 'workout' in x)]
wkot = pd.merge(wkot, data, how='left', on='product_id')
wkot = wkot.iloc[:,3:]
wkot.columns = ['label','brand','description','details','name']
wkot = wkot[wkot.label=='workout']
wkot.drop_duplicates(inplace=True)
cw = all_value[all_value['attribute_value'].apply(lambda x: 'coldweather' in x)]
cw = pd.merge(cw, data, how='left', on='product_id')
cw = cw.iloc[:,3:]
cw.columns = ['label','brand','description','details','name']
cw = cw[cw.label=='coldweather']
cw.drop_duplicates(inplace=True)
# ## label = 0
nwkd = all_value[all_value['attribute_value'].apply(lambda x: 'weekend' not in x)]
nwkd = pd.merge(nwkd, data, how='left', on='product_id')
nwkd = nwkd.iloc[:,3:]
nwkd.columns = ['label','brand','description','details','name']
nwkd['label'] = 0
nwkd.drop_duplicates(inplace=True)
ndtn = all_value[all_value['attribute_value'].apply(lambda x: 'daytonight' not in x)]
ndtn = pd.merge(ndtn, data, how='left', on='product_id')
ndtn = ndtn.iloc[:,3:]
ndtn.columns = ['label','brand','description','details','name']
ndtn['label'] = 0
ndtn.drop_duplicates(inplace=True)
nvc = all_value[all_value['attribute_value'].apply(lambda x: 'vacation' not in x)]
nvc = pd.merge(nvc, data, how='left', on='product_id')
nvc = nvc.iloc[:,3:]
nvc.columns = ['label','brand','description','details','name']
nvc['label'] = 0
nvc.drop_duplicates(inplace=True)
nwkot = all_value[all_value['attribute_value'].apply(lambda x: 'workout' not in x)]
nwkot = pd.merge(nwkot, data, how='left', on='product_id')
nwkot = nwkot.iloc[:,3:]
nwkot.columns = ['label','brand','description','details','name']
nwkot['label'] = 0
nwkot.drop_duplicates(inplace=True)
ncw = all_value[all_value['attribute_value'].apply(lambda x: 'coldweather' not in x)]
ncw = pd.merge(ncw, data, how='left', on='product_id')
ncw = ncw.iloc[:,3:]
ncw.columns = ['label','brand','description','details','name']
ncw['label'] = 0
ncw.drop_duplicates(inplace=True)
# ## wkd
# ## description
from numpy import array
from keras.preprocessing.text import one_hot
from keras.preprocessing.sequence import pad_sequences
from numpy import asarray
from numpy import zeros
from keras.preprocessing.text import Tokenizer
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import Embedding
from keras.utils import to_categorical
from sklearn.preprocessing import LabelEncoder
import spacy
import en_core_web_md
from sklearn.model_selection import train_test_split
num_1 = len(wkd['description'])
num_0 = len(nwkd['description'])
docs = wkd['description'].tolist() + nwkd['description'].tolist()
labels = np.concatenate([np.ones(num_1), np.zeros(num_0)])
nlp = en_core_web_md.load()
stopwords_removed_docs = list(
map(lambda doc: " ".join([token.text for token in nlp(doc) if not token.is_stop]), docs))
tokenizer = Tokenizer(num_words=5000, oov_token="<PASSWORD>")
tokenizer.fit_on_texts(stopwords_removed_docs)
# +
def integer_encode_documents(docs, tokenizer):
return tokenizer.texts_to_sequences(docs)
# integer encode the documents
encoded_docs = integer_encode_documents(stopwords_removed_docs, tokenizer)
# this is a list of lists, the numbers represent the index position of that word.
MAX_SEQUENCE_LENGTH = len(max(stopwords_removed_docs,key=len))
padded_docs = pad_sequences(encoded_docs, maxlen=MAX_SEQUENCE_LENGTH, padding='post')
# -
encoder = LabelEncoder()
labels = to_categorical(encoder.fit_transform(labels))
X_train, X_test, y_train, y_test = train_test_split(padded_docs, labels, test_size=0.2)
# ### Glove
from random import randint
from numpy import array, argmax, asarray, zeros
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Embedding
from keras import optimizers
# +
def load_glove_vectors():
embeddings_index = {}
with open('glove.6B.100d.txt') as f:
for line in f:
values = line.split()
word = values[0]
coefs = asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
print('Loaded %s word vectors.' % len(embeddings_index))
return embeddings_index
embeddings_index = load_glove_vectors()
# -
VOCAB_SIZE = int(len(tokenizer.word_index) * 1.1)
len(tokenizer.word_index)
embedding_matrix = zeros((VOCAB_SIZE, 100))
for word, i in tokenizer.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None: # check that it is an actual word that we have embeddings for
embedding_matrix[i] = embedding_vector
embedding_matrix.shape
# +
from keras.layers.recurrent import SimpleRNN, LSTM
from keras.layers import Flatten, Masking
# define model
def make_binary_classification_rnn_model(plot=False):
model = Sequential()
# embedding(input_dim, output_dim)
model.add(Embedding(VOCAB_SIZE, 100, weights=[embedding_matrix], input_length=MAX_SEQUENCE_LENGTH, trainable=False))
model.add(Masking(mask_value=0.0)) # masking layer, masks any words that don't have an embedding as 0s.
model.add(SimpleRNN(units=32, input_shape=(1, MAX_SEQUENCE_LENGTH))) # set hidden state dimensionality to 64
model.add(Dense(16))
model.add(Dense(2, activation='softmax')) # 2 nodes, prediction for positive/negative, two classes
# Compile the model
model.compile(
optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# summarize the model
model.summary()
if plot:
plot_model(model, to_file='model.png', show_shapes=True)
return model
def make_lstm_classification_model(plot=False):
model = Sequential()
model.add(Embedding(VOCAB_SIZE, 100, weights=[embedding_matrix], input_length=MAX_SEQUENCE_LENGTH, trainable=False))
model.add(Masking(mask_value=0.0)) # masking layer, masks any words that don't have an embedding as 0s.
model.add(LSTM(units=32, input_shape=(1, MAX_SEQUENCE_LENGTH), return_sequences=True))
#model.add(Dense(16))
model.add(Dense(2, activation='softmax'))
opt = optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999, amsgrad=False)
# Compile the model
model.compile(
optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# summarize the model
model.summary()
if plot:
plot_model(model, to_file='model.png', show_shapes=True)
return model
# -
MAX_SEQUENCE_LENGTH = len(max(stopwords_removed_docs,key=len))
model = make_lstm_classification_model()
#model = make_binary_classification_rnn_model()
history = model.fit(X_train, y_train,validation_split = 0.1, epochs=5, verbose=1)
# +
import keras
from matplotlib import pyplot as plt
def plot_fit_history(history):
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
plot_fit_history(history)
# -
loss, accuracy = model.evaluate(X_test, y_test, verbose=1)
print('Accuracy: %f' % (accuracy*100))
output = model.predict(X_train)
print(output, '\n shape:', output.shape)
# ### Train own embedding
num_1 = len(wkd['description'])
num_0 = len(nwkd['description'])
docs = wkd['description'].tolist() + nwkd['description'].tolist()
labels = np.concatenate([np.ones(num_1), np.zeros(num_0)])
nlp = en_core_web_md.load()
stopwords_removed_docs = list(
map(lambda doc: " ".join([token.text for token in nlp(doc) if not token.is_stop]), docs))
# +
def integer_encode_documents(docs, tokenizer):
return tokenizer.texts_to_sequences(docs)
vocab_size = 5000
MAX_SEQUENCE_LENGTH = len(max(stopwords_removed_docs,key=len))
EMBEDDING_SIZE = 100
tokenizer = Tokenizer()
tokenizer.fit_on_texts(stopwords_removed_docs)
encoded_docs = integer_encode_documents(stopwords_removed_docs, tokenizer)
padded_docs = pad_sequences(encoded_docs, maxlen=MAX_SEQUENCE_LENGTH, padding='post')
encoder = LabelEncoder()
labels = to_categorical(encoder.fit_transform(labels))
X_train, X_test, y_train, y_test = train_test_split(padded_docs, labels, test_size=0.2)
# -
model = make_lstm_classification_model()
history = model.fit(X_train, y_train,validation_split = 0.1, epochs=25, verbose=1)
plot_fit_history(history)
loss, accuracy = model.evaluate(X_test, y_test, verbose=1)
print('Accuracy: %f' % (accuracy*100))
# ## Other features
# +
# from https://radimrehurek.com/gensim/models/word2vec.html
from gensim.test.utils import common_texts, get_tmpfile # get_tmpfile: save a model for us
from gensim.models import Word2Vec
path = get_tmpfile("word2vec.model")
model = Word2Vec(size=100, window=3, min_count=5, workers=4)
model.save("word2vec.model")
# -
wkd['other features'] = wkd[['brand','details','name']].apply(lambda x: " ".join(x.astype(str)),axis=1)
nwkd['other features'] = nwkd[['brand','details','name']].apply(lambda x: " ".join(x.astype(str)),axis=1)
num_1 = len(wkd['other features'])
num_0 = len(nwkd['other features'])
features = wkd['other features'].tolist() + nwkd['other features'].tolist()
labels = np.concatenate([np.ones(num_1), np.zeros(num_0)])
nlp = en_core_web_md.load()
stopwords_removed_docs = list(
map(lambda doc: " ".join([token.text for token in nlp(doc) if not token.is_stop]), features))
tokenizer = Tokenizer(num_words=5000, oov_token="UNKNOWN_TOKEN")
tokenizer.fit_on_texts(stopwords_removed_docs)
tok_word_num = len(tokenizer.word_index)
# +
def integer_encode_documents(docs, tokenizer):
return tokenizer.texts_to_sequences(docs)
encoded_docs = integer_encode_documents(stopwords_removed_docs, tokenizer)
# this is a list of lists, the numbers represent the index position of that word.
MAX_SEQUENCE_LENGTH = len(max(stopwords_removed_docs,key=len))
padded_docs = pad_sequences(encoded_docs, maxlen=MAX_SEQUENCE_LENGTH, padding='post')
# -
encoder = LabelEncoder()
labels = to_categorical(encoder.fit_transform(labels))
X_train, X_test, y_train, y_test = train_test_split(padded_docs, labels, test_size=0.2)
X_train.shape
y_train.shape
w2vdocs = [word_tokenize(f) for f in features]
w2vmodel = Word2Vec(w2vdocs, size=100, window=3, min_count=5, workers=4)
len(w2vmodel.wv.vocab)
VOCAB_SIZE = int(len(tokenizer.word_index) * 1.3)
#VOCAB_SIZE = len(w2vmodel.wv.vocab)
#VOCAB_SIZE = min(5000, tok_word_num)+1
embedding_matrix = zeros((VOCAB_SIZE, 100))
for word, i in tokenizer.word_index.items():
if word in w2vmodel:
embedding_matrix[i] = w2vmodel[word]
print('Null word embeddings: %d' % np.sum(np.sum(embedding_matrix, axis=1) == 0))
embedding_matrix.shape
VOCAB_SIZE
# +
from keras.layers.recurrent import SimpleRNN, LSTM
from keras.layers import Flatten, Masking
from keras import optimizers
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.models import load_model, Sequential
# define model
def make_binary_classification_rnn_model(plot=False):
model = Sequential()
# embedding(input_dim, output_dim)
model.add(Embedding(VOCAB_SIZE, 100, weights=[embedding_matrix], input_length=MAX_SEQUENCE_LENGTH, trainable=False))
model.add(Masking(mask_value=0.0)) # masking layer, masks any words that don't have an embedding as 0s.
model.add(SimpleRNN(units=32, input_shape=(1, MAX_SEQUENCE_LENGTH))) # set hidden state dimensionality to 64
model.add(Dense(16))
model.add(Dense(2, activation='softmax')) # 2 nodes, prediction for positive/negative, two classes
# Compile the model
model.compile(
optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# summarize the model
model.summary()
if plot:
plot_model(model, to_file='model.png', show_shapes=True)
return model
def make_lstm_classification_model(plot=False):
model = Sequential()
model.add(Embedding(VOCAB_SIZE, 100, weights=[embedding_matrix], input_length=MAX_SEQUENCE_LENGTH, trainable=True))
model.add(Masking(mask_value=0.0)) # masking layer, masks any words that don't have an embedding as 0s.
model.add(LSTM(units=32, input_shape=(1, MAX_SEQUENCE_LENGTH)))
model.add(Dense(16))
model.add(Dense(2, activation='softmax'))
opt = optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999, amsgrad=False)
# early stopping
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=30)
mc = ModelCheckpoint('best_model_feature.h5', monitor='val_accuracy', mode='max', verbose=1, save_best_only=True)
# Compile the model
model.compile(
optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# summarize the model
model.summary()
if plot:
plot_model(model, to_file='model.png', show_shapes=True)
return model, es, mc
# -
MAX_SEQUENCE_LENGTH = len(max(stopwords_removed_docs,key=len))
model, es, mc = make_lstm_classification_model()
history = model.fit(X_train, y_train, validation_split = 0.1, epochs=100, verbose=1, callbacks=[es,mc])
# +
import keras
from matplotlib import pyplot as plt
def plot_fit_history(history):
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
plot_fit_history(history)
# -
saved_model = load_model('best_model_feature.h5')
loss, accuracy = saved_model.evaluate(X_test, y_test, verbose=1)
print('Accuracy: %f' % (accuracy*100))
def make_nn_classification_model(plot=False):
model = Sequential()
model.add(Embedding(VOCAB_SIZE, 100, weights=[embedding_matrix], input_length=MAX_SEQUENCE_LENGTH, trainable=True))
model.add(Masking(mask_value=0.0)) # masking layer, masks any words that don't have an embedding as 0s.
#model.add(LSTM(units=32, input_shape=(1, MAX_SEQUENCE_LENGTH)))
model.add(Dense(32, activation='relu', kernel_initializer='random_normal', input_dim=MAX_SEQUENCE_LENGTH))
model.add(Dense(2, activation='softmax'))
opt = optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999, amsgrad=False)
# early stopping
#es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=30)
#mc = ModelCheckpoint('best_model_feature.h5', monitor='val_accuracy', mode='max', verbose=1, save_best_only=True)
# Compile the model
model.compile(
optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# summarize the model
model.summary()
if plot:
plot_model(model, to_file='model.png', show_shapes=True)
return model#, es, mc
| project/Model - all inputs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Importing things
import pandas as pd
import numpy
import os
survey_2018 = pd.read_csv('./resources/04_Kaggle_Survey_2018.csv')
survey_2017 = pd.read_csv('./resources/04_Kaggle_Survey_2018.csv')
survey_2018.shape
# +
kaggle_frame= kaggle_frame[['Queried_Salary','Job_Type','Company_Industry','python','sql','machine learning','r','hadoop','tableau','sas','spark','java','Others','CA','NY','VA','TX','MA','IL','WA','MD','DC','NC','Other_states','Consulting and Business Services','Internet and Software','Banks and Financial Services','Health Care','Insurance','Other_industries']]
kaggle_frame.head()
# -
| .ipynb_checkpoints/survey_comparison-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import pandas as pd, json, numpy as np
import matplotlib.pyplot as plt
from bs4 import BeautifulSoup
# %matplotlib inline
# Load airports of each country
L=json.loads(file('../json/L.json','r').read())
M=json.loads(file('../json/M.json','r').read())
N=json.loads(file('../json/N.json','r').read())
import requests
AP={}
for c in M:
if c not in AP:AP[c]={}
for i in range(len(L[c])):
AP[c][N[c][i]]=L[c][i]
# record schedules for 2 weeks, then augment count with weekly flight numbers.
# seasonal and seasonal charter will count as once per week for 3 months, so 12/52 per week. TGM separate, since its history is in the past.
# parse Departures
baseurl='https://www.airportia.com/'
import requests, urllib2
def urlgetter(url):
s = requests.Session()
cookiesopen = s.get(url)
cookies=str(s.cookies)
fcookies=[[k[:k.find('=')],k[k.find('=')+1:k.find(' for ')]] for k in cookies[cookies.find('Cookie '):].split('Cookie ')[1:]]
#push token
opener = urllib2.build_opener()
for k in fcookies:
opener.addheaders.append(('Cookie', k[0]+'='+k[1]))
#read html
return s.get(url).content
# good dates
SD={}
SC=json.loads(file('../json/SC2.json','r').read())
#pop out last - if applicable
try: SD.pop(c)
except: pass
for h in range(len(AP.keys())):
c=AP.keys()[h]
#country not parsed yet
if c in SC:
if c not in SD:
SD[c]=[]
print h,c
airportialinks=AP[c]
sch={}
#all airports of country, where there is traffic
for i in airportialinks:
if i in SC[c]:
print i,
if i not in sch:sch[i]={}
url=baseurl+airportialinks[i]
m=urlgetter(url)
for d in range (3,17):
#date not parsed yet
if d not in sch[i]:
url=baseurl+airportialinks[i]+'departures/201704'+str(d)
m=urlgetter(url)
soup = BeautifulSoup(m, "lxml")
#if there are flights at all
if len(soup.findAll('table'))>0:
sch[i][d]=pd.read_html(m)[0]
else: print '--W-',d,
SD[c]=sch
print
# Save
dbpath='E:/Dropbox/Public/datarepo/aviation/' #large file db path
file(dbpath+"json/SD_dest.json",'w').write(repr(SD))
cnc_path='../../universal/countries/'
cnc=pd.read_excel(cnc_path+'cnc.xlsx').set_index('Name')
MDF=pd.DataFrame()
for c in SD:
sch=SD[c]
mdf=pd.DataFrame()
for i in sch:
for d in sch[i]:
df=sch[i][d].drop(sch[i][d].columns[3:],axis=1).drop(sch[i][d].columns[0],axis=1)
df['From']=i
df['Date']=d
mdf=pd.concat([mdf,df])
mdf=mdf.replace('Hahn','Frankfurt')
mdf=mdf.replace('Hahn HHN','Frankfurt HHN')
if len(sch)>0:
mdf['City']=[i[:i.rfind(' ')] for i in mdf['To']]
mdf['Airport']=[i[i.rfind(' ')+1:] for i in mdf['To']]
cpath=str(cnc.T.loc[c]['ISO2']).lower()
if cpath=='nan':cpath='na'
file('../countries/'+cpath+"/json/mdf_dest.json",'w').write(json.dumps(mdf.reset_index().to_json()))
MDF=pd.concat([MDF,mdf])
print c,
dbpath='E:/Dropbox/Public/datarepo/aviation/' #large file db path
MDF.reset_index().to_json(dbpath+'json/MDF_dest.json')
| code/.ipynb_checkpoints/airport_dest_parser-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fbocqoWfpVPg"
# The Meterstick package provides a concise and flexible syntax to describe and execute
# routine data analysis tasks. The easiest way to learn to use Meterstick is by example.
# + [markdown] id="1KBTLgUrzmS7"
# # For External users
#
# You can open this notebook in [Google Colab](https://colab.research.google.com/github/google/meterstick/blob/master/meterstick_demo.ipynb).
# + [markdown] id="H9ojnghz0b2N"
# ## Installation
#
# You can install from pip for the stable version
# + id="ohmnh0qRz6bS"
#@test {"skip": true}
# !pip install meterstick
# + [markdown] id="MZXKtCHy0CEo"
# or from GitHub for the latest version.
# + id="uQRaNJ2h0NvF"
#@test {"skip": true}
# !git clone https://github.com/google/meterstick.git
import sys, os
sys.path.append(os.getcwd())
# + [markdown] id="te-lKCw20P41"
# # Demo Starts
# + executionInfo={"elapsed": 165, "status": "ok", "timestamp": 1634584765965, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="0UI9rAtZnBUG"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from meterstick import *
# + colab={"height": 204} executionInfo={"elapsed": 297, "status": "ok", "timestamp": 1634584770460, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="hyDE-bxMBxQY" outputId="a97737a8-32df-4fed-cc2f-90a88861dc55"
np.random.seed(42)
platform = ('Desktop', 'Mobile', 'Tablet')
exprs = ('ctrl', 'expr')
country = ('US', 'non-US')
size = 1000
impressions = np.random.randint(10, 20, size)
clicks = impressions * 0.1 * np.random.random(size)
df = pd.DataFrame({'impressions': impressions, 'clicks': clicks})
df['platform'] = np.random.choice(platform, size=size)
df['expr_id'] = np.random.choice(exprs, size=size)
df['country'] = np.random.choice(country, size=size)
df['cookie'] = np.random.choice(range(5), size=size)
df.loc[df.country == 'US', 'clicks'] *= 2
df.loc[(df.country == 'US') & (df.platform == 'Desktop'), 'impressions'] *= 4
df.head()
# + [markdown] id="M4nEYlPhudKL"
# # Simple Metrics
# There are many built-in simple Metrics in Meterstick. They directly operate on a DataFrame.
# + [markdown] id="xv090eQ6vJsA"
# ## Sum
# + colab={"height": 80} executionInfo={"elapsed": 274, "status": "ok", "timestamp": 1595026545506, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="TVyts3vuno80" outputId="f0065028-d348-473a-b0bf-218d482a1584"
Sum('clicks').compute_on(df)
# + [markdown] id="imbzELxnuwLN"
# ## Count
# + colab={"height": 80} executionInfo={"elapsed": 439, "status": "ok", "timestamp": 1602800472205, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="cTww4tuHuzMC" outputId="b2fb4b2b-9dca-4c73-a3f5-0c5ea029c066"
Count('country').compute_on(df)
# + colab={"height": 80} executionInfo={"elapsed": 483, "status": "ok", "timestamp": 1602800482735, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="QWylIectWgu_" outputId="4db7f426-0190-4907-a5db-dd2439383080"
Count('country', distinct=True).compute_on(df)
# + [markdown] id="tbpIWm7_5z2H"
# ## Dot (inner product)
# + colab={"height": 80} executionInfo={"elapsed": 327, "status": "ok", "timestamp": 1616410947877, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="LvRyrQx85ttF" outputId="4d3108e3-11c5-4f2c-dc59-ea97a5b88303"
Dot('clicks', 'impressions').compute_on(df)
# + [markdown] id="EyyDXIbgmjzI"
# It can also be normalized.
# + colab={"height": 80} executionInfo={"elapsed": 340, "status": "ok", "timestamp": 1616410961294, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="glQsTDVgla6x" outputId="9aa20256-b02c-4085-ec62-dccfd994af18"
Dot('clicks', 'clicks', True).compute_on(df)
# + [markdown] id="T1hPtrBliRSa"
# ## Max
# + colab={"height": 80} executionInfo={"elapsed": 515, "status": "ok", "timestamp": 1600991606672, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="XzMvl52ZiSiz" outputId="e3b2aa26-1936-4f60-eebc-3223332173ce"
Max('clicks').compute_on(df)
# + [markdown] id="hUlXQ1IjiV5b"
# ## Min
# + colab={"height": 80} executionInfo={"elapsed": 427, "status": "ok", "timestamp": 1600991623794, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="7kyjKainiWzp" outputId="7abe9e3c-529b-4c88-b673-b7c9c629ca7f"
Min('clicks').compute_on(df)
# + [markdown] id="om2mS_pKu_fg"
# ## Mean
# + colab={"height": 80} executionInfo={"elapsed": 368, "status": "ok", "timestamp": 1594365621883, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="segsW0glvB3K" outputId="b3e5e0b8-27a6-4acb-f059-745bf00f25d2"
Mean('clicks').compute_on(df)
# + [markdown] id="VjMoDhvNvmyn"
# ###Weighted Mean
# + colab={"height": 80} executionInfo={"elapsed": 415, "status": "ok", "timestamp": 1594365622359, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="2JagqbG7vonz" outputId="c9ad7f21-e471-40cc-96ea-0303cbf97104"
Mean('clicks', 'impressions').compute_on(df)
# + [markdown] id="baVEO52pvM6M"
# ##Quantile
# + colab={"height": 80} executionInfo={"elapsed": 393, "status": "ok", "timestamp": 1594365622826, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="FxBSzQFkvXpi" outputId="6cf32f81-40a3-4021-cfb8-b2f8934cabd9"
Quantile('clicks').compute_on(df) # Default is median.
# + colab={"height": 80} executionInfo={"elapsed": 388, "status": "ok", "timestamp": 1594365623293, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="J0ojPu4wv1j9" outputId="746363a7-f671-42da-8a20-33ee73a39f36"
Quantile('clicks', 0.2).compute_on(df)
# + colab={"height": 80} executionInfo={"elapsed": 388, "status": "ok", "timestamp": 1594365623764, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="rO7n3brmv8Q8" outputId="045b3b41-6e98-4d88-88f5-733e1e808487"
Quantile('clicks', (0.2, 0.5)).compute_on(df) # Quantile can take multiple quantiles.
# + [markdown] id="8jcluN5LwgCt"
# ### Interpolation
# You can specify how you want to interpolate the quantile. It could be any of (‘linear’, ‘lower’, ‘higher’, ‘midpoint’, ‘nearest’).
# + colab={"height": 80} executionInfo={"elapsed": 384, "status": "ok", "timestamp": 1594365624220, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="4LFIT6MIwhaT" outputId="e54cbea3-2537-40ee-fd99-f5fd5a2a2d06"
Quantile('clicks', 0.5, interpolation='higher').compute_on(df)
# + [markdown] id="-JXvzK1TwGqC"
# ### Weighted Quantile
# + colab={"height": 80} executionInfo={"elapsed": 383, "status": "ok", "timestamp": 1594365624656, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="rquZ6eL5wL8_" outputId="e13dbc33-fd10-4682-9b66-950e4427c1d3"
Quantile('clicks', weight='impressions').compute_on(df)
# + [markdown] id="4ds1nTgZvPCn"
# ##Variance
# + colab={"height": 80} executionInfo={"elapsed": 385, "status": "ok", "timestamp": 1594365625093, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="LrSqbbtVw-So" outputId="cabb28b4-23e5-4bf2-a96d-c02fe1e76ec2"
Variance('clicks').compute_on(df)
# + [markdown] id="GePCapHoxVb-"
# ###Biased Variance
# The default Variance is unbiased, namely, the divisor used in calculations is N - 1. You could set unbiased=False to use N as the divisor.
# + colab={"height": 80} executionInfo={"elapsed": 368, "status": "ok", "timestamp": 1594365625540, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="bxwlTyxrxusg" outputId="36f588c5-8089-4b52-8cdb-4c19f64e89a4"
Variance('clicks', unbiased=False).compute_on(df)
# + [markdown] id="jr9jIf0ox1eM"
# ### Weighted Variance
# + colab={"height": 80} executionInfo={"elapsed": 377, "status": "ok", "timestamp": 1594365625988, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="0ZxNnhcZx5fz" outputId="740f3f38-5d42-4d73-b572-8bf7d6e5f28c"
Variance('clicks', weight='impressions').compute_on(df)
# + [markdown] id="IHozqoZ2vQIf"
# ##Standard Deviation
# + colab={"height": 80} executionInfo={"elapsed": 406, "status": "ok", "timestamp": 1594365626510, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="v15AY43ByekK" outputId="dfb0aaa9-22f0-4467-ccb5-3d7235bbe806"
StandardDeviation('clicks').compute_on(df)
# + [markdown] id="N1rDr4SGyu72"
# ### Biased Standard Deviation
# Similar to biased Variance, it's possible to compute biased standard deviation.
# + colab={"height": 80} executionInfo={"elapsed": 375, "status": "ok", "timestamp": 1594365626961, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="fHRfub38zAGZ" outputId="964be900-5acd-4a3a-fa63-26c7c10115aa"
StandardDeviation('clicks', False).compute_on(df)
# + [markdown] id="245lDve-zD_s"
# ### Weighted Standard Deviation
# + colab={"height": 80} executionInfo={"elapsed": 378, "status": "ok", "timestamp": 1594365627393, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="e0ELsbRDzGcy" outputId="a030b1fe-576a-4ee0-ac4a-73ff8d89b7cf"
StandardDeviation('clicks', weight='impressions').compute_on(df)
# + [markdown] id="FVCJgHe7vRnD"
# ##Coefficient of Variation
# + colab={"height": 80} executionInfo={"elapsed": 378, "status": "ok", "timestamp": 1594365627823, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="xSWoWiQLzcz0" outputId="bae8e13d-07fa-4da1-9b5d-f85ef7809fe5"
CV('clicks').compute_on(df)
# + [markdown] id="3QOKrzf5vTdX"
# ##Correlation
# + colab={"height": 80} executionInfo={"elapsed": 378, "status": "ok", "timestamp": 1594365628254, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="wfLRzJ87zlyR" outputId="329f1a51-cde3-4147-f641-3bc1e7b68730"
Correlation('clicks', 'impressions').compute_on(df)
# + [markdown] id="ofd39aQs0Xwn"
# ###Weighted Correlation
# + colab={"height": 80} executionInfo={"elapsed": 382, "status": "ok", "timestamp": 1594365628689, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="jivKl_cZ0aM_" outputId="1514c4ed-1caf-419e-cc34-f9bba59bdb3c"
Correlation('clicks', 'impressions', weight='impressions').compute_on(df)
# + [markdown] id="Pwoz9GUwvVUr"
# ##Covariance
# + colab={"height": 80} executionInfo={"elapsed": 373, "status": "ok", "timestamp": 1594365629117, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="CYR8RCP50l4y" outputId="5521eb32-1cd2-4823-9661-a43f24e73a7f"
Cov('clicks', 'impressions').compute_on(df)
# + [markdown] id="Sez3EB8J0l44"
# ###Weighted Covariance
# + colab={"height": 80} executionInfo={"elapsed": 373, "status": "ok", "timestamp": 1594365629545, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="GfZmRTQx0l45" outputId="a2a8a8f5-5826-411a-e3c5-9ac9ae740e81"
Cov('clicks', 'impressions', weight='impressions').compute_on(df)
# + [markdown] id="qbGHSpls0u8e"
# # Slicing
# You can group your DataFrame and compute the Metrics on slices.
# + colab={"height": 142} executionInfo={"elapsed": 386, "status": "ok", "timestamp": 1594365629982, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="xd3vQWs617ow" outputId="6330ca80-bc55-4b09-8aab-2f120678057e"
Sum('clicks').compute_on(df, 'country')
# + colab={"height": 266} executionInfo={"elapsed": 373, "status": "ok", "timestamp": 1601683238341, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6kOyjbDs2Dhj" outputId="ad6e13cf-20c0-496a-b33e-d5707de3dba5"
Mean('clicks').compute_on(df, ['platform', 'country'])
# + [markdown] id="6nHJIFFO3kN9"
# # Multiple Metrics
# You can put multiple Metrics into a MetricList and compute them together. It's not only makes your codes terser, it might make the computation much faster. See Caching section for more infomation.
# + colab={"height": 80} executionInfo={"elapsed": 354, "status": "ok", "timestamp": 1594365630811, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="9GX9SxKG3mKO" outputId="6f8252bc-57b5-4f9c-ac05-dd9de7126da2"
MetricList((Sum('clicks'), Count('clicks'))).compute_on(df)
# + [markdown] id="U9oidu9eELbG"
# #Arithmetic of Metrics
# You can do many arithmetic operations on Metrics. It can also be between a Metric and a scalar. You can call set_name() to give your composite Metric a new name. Internally, we operate on the results returned by Metrics with return_dataframe=False to avoid incompatible DataFrame columns names. However, if both Metrics return DataFrames even when return_dataframe is set to False, you might get lots of NAs. The solution is use rename_columns() to unify the column names. See section "Compare the standard errors between Jackknife and Bootstrap" for an example.
# + [markdown] id="rsF4N5WJE3SA"
# ### Add
# + colab={"height": 80} executionInfo={"elapsed": 373, "status": "ok", "timestamp": 1594365631252, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="JQmHZgKtE8P8" outputId="48609b1c-0d15-4634-a78f-b0f4fef291bd"
(Sum('clicks') + 1).compute_on(df)
# + colab={"height": 80} executionInfo={"elapsed": 385, "status": "ok", "timestamp": 1594365631686, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="s6nVzOXXGFln" outputId="25fcc58a-b190-4cb4-9f2f-9dda13e78b9c"
sum((Sum('clicks'), Sum('impressions'), 1)).compute_on(df)
# + colab={"height": 80} executionInfo={"elapsed": 393, "status": "ok", "timestamp": 1594365632129, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="qYYsIXTYHpgk" outputId="fd8cfdc6-a373-446f-8a04-d2bbe5c99f86"
sum((Sum('clicks'), Sum('impressions'), 1)).set_name('meaningless sum').compute_on(df)
# + [markdown] id="ZnRbSjz6HOZ6"
# ### Divide
# + colab={"height": 80} executionInfo={"elapsed": 377, "status": "ok", "timestamp": 1594365633901, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="MGSIaravHSZL" outputId="a6a8696b-68a9-4c5b-8180-b97e02d92020"
(Sum('clicks') / Sum('impressions')).compute_on(df)
# + [markdown] id="ymR7ZEbzHySV"
# ####Ratio
# Since division between two Sums is common, we make a Ratio() Metric as a syntax sugar. Its third arg is the name for the Metric and is optional.
# + colab={"height": 80} executionInfo={"elapsed": 380, "status": "ok", "timestamp": 1594365634330, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="GbFTWys7H7aN" outputId="53f40838-bc15-41f3-8ef6-4eee1c15403e"
Ratio('clicks', 'impressions', 'ctr').compute_on(df)
# + [markdown] id="TJ9P3C1yGCEZ"
# We also support many other common arithmetic operations.
# + colab={"height": 266} executionInfo={"elapsed": 513, "status": "ok", "timestamp": 1595023588993, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="bspYv-_kGDLj" outputId="af2d8e6e-018d-4ba8-ff2c-f57dabe17230"
MetricList(
(Sum('clicks') - 1,
-Sum('clicks'),
2 * Sum('clicks'),
Sum('clicks')**2,
2**Mean('clicks'),
(Mean('impressions')**Mean('clicks')).set_name('meaningless power'))
).compute_on(df, melted=True)
# + [markdown] id="ReIMkRtT0rko"
# # Output Format
# There are two options for you to control the format of the return.
# 1. return_dataframe: Default True, if False, we try to return a scalar or pd.Series. For complex Metrics it might have no effect and a DataFrame is always returned. For example, all Metrics in the Operations section below always return a DataFrame.
#
# return_dataframe has different effect on MetricList. If False, MetricList will return a list of DataFrames instead of trying to concat them. This is a convenient way to compute incompatible Metrics together to maximize caching (see section Caching also). There is an attribute "children_return_dataframe" in MetricList which will be passed to children Metrics as their return_dataframe so you can get a list of numbers or pd.Series.
# 2. melted: Dafault False. It decides if the returned DataFrame is in wide/unmelted or long/melted form. It doesn't have effect if the return is not a DataFrame.
# - Long/melted means the leftmost index is 'Metric' so
#
# `MetricList((m1, m2)).compute_on(df, melted=True).loc[m1.name] ≡ m1.compute_on(df, melted=True)`
#
# - Wide/unmelted means the outermost column index is 'Metric' so
#
# `MetricList((m1, m2)).compute_on(df)[m1.name] ≡ m1.compute_on(df)`
# + colab={"height": 34} executionInfo={"elapsed": 351, "status": "ok", "timestamp": 1594365636630, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="BPqNffOz3QXO" outputId="bfb357db-2644-4245-9215-14a7a8504b5e"
Sum('clicks').compute_on(df, return_dataframe=False)
# + colab={"height": 153} executionInfo={"elapsed": 333, "status": "ok", "timestamp": 1594365637064, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="ECg_e0NE3Usb" outputId="3806fb1d-580e-4d99-d71a-3c723ab57aad"
Count('clicks').compute_on(df, ['platform', 'country'], return_dataframe=False)
# + colab={"height": 111} executionInfo={"elapsed": 379, "status": "ok", "timestamp": 1594365637526, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="k1Wktj2S3ah1" outputId="05af8497-045a-4feb-e14f-e7bd35e0e215"
Mean('clicks').compute_on(df, melted=True)
# + colab={"height": 142} executionInfo={"elapsed": 385, "status": "ok", "timestamp": 1594365637961, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="eYv2KDsQ4gRD" outputId="1408a743-ec4f-48ea-b5b3-22b727ee303d"
MetricList((Sum('clicks'), Count('clicks'))).compute_on(df, 'country')
# + colab={"height": 204} executionInfo={"elapsed": 378, "status": "ok", "timestamp": 1594365638394, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="a87pRDem3hH8" outputId="c80441ac-d636-4814-d358-f7f4e4861a03"
Quantile('clicks', [0.2, 0.7]).compute_on(df, 'country', melted=True)
# + colab={"height": 173} executionInfo={"elapsed": 452, "status": "ok", "timestamp": 1594365638903, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="WH48W49s9Sni" outputId="3c12a85e-2b24-404f-87ce-7676d838d25e"
# Don't worry. We will talk more about the pipeline operator "|" later.
(MetricList((Sum('clicks'), Count('clicks')))
| Jackknife('cookie')
| compute_on(df, 'country'))
# + colab={"height": 204} executionInfo={"elapsed": 1907, "status": "ok", "timestamp": 1594365640862, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="EiNhXrmM9dIX" outputId="ae3a3654-12f2-4efd-af3b-5794ccdb5427"
(MetricList((Sum('clicks'), Count('clicks')))
| Bootstrap(n_replicates=100)
| compute_on(df, 'country', melted=True))
# + [markdown] id="aeceO97t2-rc"
# # Operations
#
# An Operation is a special type of Metric that is built on top of another Metric (called a "child"). A Metric is anything that has the compute_on() method, so the child doesn't need to be a simple Metric like Sum. It could be a MetricList, a composite Metric, or even another Operation.
# + [markdown] id="zVRs_VKe7Bqb"
# ##Distribution
# Compute the child Metric on a DataFrame grouped by a column, then normalize the numbers to 1 within group.
# + colab={"height": 142} executionInfo={"elapsed": 685, "status": "ok", "timestamp": 1603228234351, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="qU27HLT60n03" outputId="93461f9c-e82d-456c-be06-2487c9d9dad8"
Distribution('country', Sum('clicks')).compute_on(df)
# + [markdown] id="ecTg-LNF---8"
# It's equal to
# + colab={"height": 142} executionInfo={"elapsed": 382, "status": "ok", "timestamp": 1594365641750, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="1-srLwAU9Zkk" outputId="189a6edf-a252-4a87-f7d2-8c64da834c41"
(Sum('clicks').compute_on(df, 'country') /
Sum('clicks').compute_on(df, return_dataframe=False))
# + [markdown] id="Ijmv6fpnyN_U"
# Distribution has an alias Normalize.
# + colab={"height": 142} executionInfo={"elapsed": 711, "status": "ok", "timestamp": 1603228243421, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="p7ZZXDJizEr1" outputId="e1adad30-1cf0-4748-db30-c572ef1a73f0"
Normalize('country', Sum('clicks')).compute_on(df)
# + [markdown] id="QV9REkDR_G2h"
# ##Cumulative Distribution
# Similar to Distribution except that it returns the cumulative sum after normalization, but unlike Distribution the order of the cumulating column matters. As the result, we always sort the column and there is an 'order' arg for you to customize the ordering.
# + colab={"height": 142} executionInfo={"elapsed": 404, "status": "ok", "timestamp": 1596620965224, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="mNEvEQu--8Qn" outputId="40467f8c-64b3-4691-a95b-0d29be621c87"
CumulativeDistribution('country', MetricList(
(Sum('clicks'), Sum('impressions')))).compute_on(df)
# + colab={"height": 266} executionInfo={"elapsed": 419, "status": "ok", "timestamp": 1596620997849, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="sbg5eHCv_eqc" outputId="e468abbe-e204-47ec-d1c5-9a371851c8f6"
CumulativeDistribution(
'country', Sum('clicks'), order=('non-US', 'US')).compute_on(df, 'platform')
# + colab={"height": 204} executionInfo={"elapsed": 538, "status": "ok", "timestamp": 1596621061031, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="VvRhl_vQAABY" outputId="cc2edffc-4248-467a-daa8-47cf494b0c47"
CumulativeDistribution(
'country', MetricList((Sum('clicks'), Sum('impressions')))
).compute_on(df, melted=True)
# + [markdown] id="5PDDFqXZAa_S"
# ##PercentChange
# Computes the percent change to a certain group on the DataFrame returned by the child Metric. The returned value is the # of percent points.
# + colab={"height": 111} executionInfo={"elapsed": 404, "status": "ok", "timestamp": 1594365643465, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="cbmM9OW9AJ0c" outputId="decb86ec-9dd5-4711-8672-710436bdbafa"
PercentChange('country', 'US', Mean('clicks')).compute_on(df)
# + colab={"height": 51} executionInfo={"elapsed": 367, "status": "ok", "timestamp": 1594365643891, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="0o7pYZaPBTiB" outputId="c36e8073-eb33-49e4-d27f-90109c1d5d97"
mean = Mean('clicks').compute_on(df, 'country')
(mean.loc['non-US'] / mean.loc['US'] - 1) * 100
# + [markdown] id="855lw27MB2Nf"
# You can include the base group in your result.
# + colab={"height": 266} executionInfo={"elapsed": 391, "status": "ok", "timestamp": 1594365644334, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6YTYgQ7GBfUu" outputId="f9e77b75-6c7e-4c5e-e0f5-b81852c2ba8a"
PercentChange(
'country',
'US',
MetricList((Count('clicks'), Count('impressions'))),
include_base=True).compute_on(df, 'platform')
# + [markdown] id="jPMAQUt6HsXt"
# You can also specify multiple columns as the condition columns, then your base value should be a tuple.
# + colab={"height": 266} executionInfo={"elapsed": 500, "status": "ok", "timestamp": 1601353784290, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="Fe8o-qoDHy-A" outputId="8deaf5ea-d64c-47ab-e19e-08ea388a79fc"
PercentChange(
['country', 'platform'],
('US', 'Desktop'),
MetricList((Count('clicks'), Count('impressions'))),
include_base=True).compute_on(df)
# + [markdown] id="y6PJdEMOCMVV"
# ##Absolute Change
# Very similar to PercentChange, but the absolute difference is returned.
# + colab={"height": 111} executionInfo={"elapsed": 372, "status": "ok", "timestamp": 1594365644758, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="jl3tnA7iCDdC" outputId="a31985be-b3c1-49ba-b9a0-e8d52296b42e"
AbsoluteChange('country', 'US', Mean('clicks')).compute_on(df)
# + [markdown] id="w_gMScULCglZ"
# You can also include the base group in your result.
# + colab={"height": 266} executionInfo={"elapsed": 373, "status": "ok", "timestamp": 1594365645175, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="mHuDueBQCglb" outputId="92769455-bf2a-44a5-e55e-99a1f7333997"
AbsoluteChange(
'country', 'US', Count('clicks'), include_base=True).compute_on(
df, 'platform', melted=True)
# + [markdown] id="ProNeuPcCpvt"
# ##Cochran-Mantel-Haenszel statistics
# Please refer to the Wikepedia [page](https://en.wikipedia.org/wiki/Cochran%E2%80%93Mantel%E2%80%93Haenszel_statistics) for its definition. Besides the condition column and baseline key that PercentChange and AbsoluteChange take, CMH also needs a column to stratify. The child Metric must be a ratio of two single-column Metrics or CMH doesn't make sense. So instead of passing
#
# AbsoluteChange(MetricList([a, b])) / AbsoluteChange(MetricList([c, d])),
# please use
#
# MetricList([AbsoluteChange(a) / AbsoluteChange(c),
# AbsoluteChange(b) / AbsoluteChange(d)]).
# + colab={"height": 111} executionInfo={"elapsed": 364, "status": "ok", "timestamp": 1594365645596, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="rz_U1FrfCV66" outputId="f4a5379b-1cad-40c0-a7b4-1b88c448db38"
ctr = Ratio('clicks', 'impressions')
MH('country', 'US', 'platform', ctr).compute_on(df) # stratified by platform
# + [markdown] id="uvQrrW7NY0Vj"
# ## Standard Errors
# + [markdown] id="vH3Id5U9JOqZ"
# ###Jackknife
# Unlike all Metrics we have seen so far, Jackknife returns a multiple-column DataFrame because by default we return point estimate and standard error.
# + colab={"height": 111} executionInfo={"elapsed": 442, "status": "ok", "timestamp": 1594365646085, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="2ND79IibD6Pg" outputId="c56f0562-9572-44e5-d525-86e84da32044"
Jackknife('cookie', MetricList((Sum('clicks'), Sum('impressions')))).compute_on(df)
# + colab={"height": 204} executionInfo={"elapsed": 520, "status": "ok", "timestamp": 1594366655230, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="837hcADhOYFY" outputId="ae3926ec-70d2-47eb-b2f1-bcd7889535dd"
metrics = MetricList((Sum('clicks'), Sum('impressions')))
Jackknife('cookie', metrics).compute_on(df, 'country', True)
# + [markdown] id="eE3e5FAgRPf2"
# You can also specify a confidence level, the we'll return the confidence interval. The returned DataFrame also comes with a display() method for visualization which will highlight significant changes. To customize the display(), please take a look at confidence_interval_display_demo.ipynb.
# + colab={"height": 111} executionInfo={"elapsed": 510, "status": "ok", "timestamp": 1594366671346, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="yA8-xkyXOfH8" outputId="045b37a1-9894-4272-e94a-f6331af139dc"
Jackknife('cookie', metrics, 0.9).compute_on(df)
# + colab={"height": 349} executionInfo={"elapsed": 699, "status": "ok", "timestamp": 1603245233872, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="53NI01DoqyDe" outputId="2141ad51-672e-4bef-800f-7e1b703ea394"
#@test {"skip": true}
res = (
MetricList((Ratio('clicks', 'impressions', 'ctr'), Sum('clicks')))
| PercentChange('country', 'US')
| Jackknife('cookie', confidence=0.9)
| compute_on(df, 'platform'))
res.display()
# + [markdown] id="QzG196CeUxQP"
# ###Bootstrap
# The output is similar to Jackknife. The different args are
# - unit: If None, we bootstrap on rows. Otherwise we do a [block bootstrap](https://en.wikipedia.org/wiki/Bootstrapping_(statistics)#Block_bootstrap). The unique values in unit column will be used as the resampling buckets.
# - n_replicates: The number of resamples. Default to 10000, which is recommended in Tim Hesterberg's [What Teachers Should Know About the Bootstrap](https://amstat.tandfonline.com/doi/full/10.1080/00031305.2015.1089789). Here we use a smaller number for faster demonstration.
# + colab={"height": 111} executionInfo={"elapsed": 1141, "status": "ok", "timestamp": 1595024950827, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="9zdiyrk-SoGl" outputId="34bf82c2-983d-42fa-f4df-bd921195b5ce"
np.random.seed(42)
Bootstrap(None, Sum('clicks'), 100).compute_on(df)
# + colab={"height": 173} executionInfo={"elapsed": 2537, "status": "ok", "timestamp": 1595024957016, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="jUoTT2dNV-m9" outputId="464018ac-5a9e-4e75-d6c0-7f10ba579b97"
np.random.seed(42)
Bootstrap('cookie', Sum('clicks'), 100).compute_on(df, 'country')
# + colab={"height": 173} executionInfo={"elapsed": 2034, "status": "ok", "timestamp": 1594365718636, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="Bm5c-nM1WZ1M" outputId="21c29fe3-6428-4dcd-e332-4fea902392b3"
np.random.seed(42)
Bootstrap('cookie', Sum('clicks'), 100, 0.95).compute_on(df, 'country')
# + colab={"height": 349} executionInfo={"elapsed": 5167, "status": "ok", "timestamp": 1603245244409, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="uKBRJlBBqskw" outputId="6b850c3e-be8b-4a27-89fc-1040813e5cf0"
#@test {"skip": true}
np.random.seed(42)
res = (
MetricList((Ratio('clicks', 'impressions', 'ctr'), Sum('impressions')))
| AbsoluteChange('country', 'US')
| Bootstrap(None, n_replicates=100, confidence=0.9)
| compute_on(df, 'platform'))
res.display()
# + [markdown] id="QI5njXEUg_G1"
# # Models
#
# Meterstick also has built-in support for model fitting. The module is not imported by default, so you need to manually import it.
# + executionInfo={"elapsed": 429, "status": "ok", "timestamp": 1634584662059, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="TajdScHljI56"
with adhoc_import.Google3():
from meterstick.v2.models import *
# + [markdown] id="7zzOX_bNiwoq"
# ## Linear Regression
# + colab={"height": 80} executionInfo={"elapsed": 113, "status": "ok", "timestamp": 1634530013559, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="McLNFF3wi0JE" outputId="f0fa2bf5-82e2-4b03-ade8-1b6c0565e639"
m = LinearRegression(Mean('clicks'), Mean('impressions'), 'platform')
m.compute_on(df)
# + [markdown] id="kRqbmZ2Ryugp"
# What `Model(y, x, groupby).compute_on(data)` does is
# 1. Computes `MetricList((y, x)).compute_on(data, groupby)`.
# 2. Fits the underlying sklearn model on the result from #1.
# + executionInfo={"elapsed": 93, "status": "ok", "timestamp": 1634528151126, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="DAoznoAYkkoS" outputId="a1f81219-81dd-4458-ea54-bdc77a9309af"
from sklearn import linear_model
x = Mean('impressions').compute_on(df, 'platform')
y = Mean('clicks').compute_on(df, 'platform')
m = linear_model.LinearRegression().fit(x, y)
print(m.coef_, m.intercept_)
# + [markdown] id="VYe66X0gyOt6"
# ## Ridge Regression
# + colab={"height": 173} executionInfo={"elapsed": 125, "status": "ok", "timestamp": 1634528195346, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="_TzTVLOpxgAL" outputId="829e8455-0d50-4824-eb06-c1fc19b6e43c"
# x can also be a list of Metrics or a MetricList.
m = Ridge(
Mean('clicks'),
[Mean('impressions'), Variance('clicks')],
'platform',
alpha=2)
m.compute_on(df, melted=True)
# + [markdown] id="Gul9O8xyyQ5t"
# ## Lasso Regression
# + colab={"height": 142} executionInfo={"elapsed": 13, "status": "ok", "timestamp": 1634528265646, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="yaGsAMfKyQ5u" outputId="46feb304-0537-468a-eb3b-7d4ee708fd5e"
m = Lasso(
Mean('clicks'),
Mean('impressions'),
'platform',
fit_intercept=False,
alpha=5)
m.compute_on(df, 'country')
# + [markdown] id="aTC8os9tyRfo"
# ## Logistic Regression
# + colab={"height": 142} executionInfo={"elapsed": 82, "status": "ok", "timestamp": 1634584819513, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="FzErH-Ib16FJ" outputId="f361c589-d223-4d03-e8c9-23f8318042ec"
m = LogisticRegression(Count('clicks'), Mean('impressions'), 'country')
m.compute_on(df, melted=True)
# + [markdown] id="5LwB8nBTfNkR"
# If y is not binary, by default a multinomal model is fitted. The behavior can be controlled via the '[multinomial](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html)' arg.
# + colab={"height": 266} executionInfo={"elapsed": 22, "status": "ok", "timestamp": 1634528278429, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="34kbNeq5yRfo" outputId="4f770e8c-d844-42ed-ac9a-cc84a3176a77"
m = LogisticRegression(Count('clicks'), Mean('impressions'), 'platform', name='LR')
m.compute_on(df, melted=True)
# + [markdown] id="zjDtn_3-e-xa"
# Classes are the unique values of y.
# + colab={"height": 173} executionInfo={"elapsed": 200, "status": "ok", "timestamp": 1634528368772, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="X5ZbvavOe0w1" outputId="1e38f267-d6f6-4185-f98f-f673a8b9a39d"
Count('clicks').compute_on(df, 'platform')
# + [markdown] id="UXRS4p9jy8PR"
# Wrapping sklearn models into Meterstick provides the ability to combine Models with other built-in Metrics and Operations. For example, you can Jackknife the Model to get the uncertainty of coefficients.
# + colab={"height": 314} executionInfo={"elapsed": 319, "status": "ok", "timestamp": 1634577563356, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="ezICFP4rzDEb" outputId="1fc420e6-6889-4403-f235-74a0664860eb"
#@test {"skip": true}
(LinearRegression(
Mean('clicks'),
[Mean('impressions'), Variance('impressions')],
'country',
name='lm')
| AbsoluteChange('platform', 'Desktop')
| Jackknife('cookie', confidence=0.9)
| compute_on(df)).display()
# + [markdown] id="H-bQTkuaYHNb"
# #Pipeline
# You have already seen this. Instead of
#
# Jackknife(PercentChange(MetricList(...))).compute_on(df)
# you can write
#
# MetricList(...) | PercentChange() | Jackknife() | compute_on(df)
#
# which is more intuitive. We overwrite the "|" operator on Metric and the \_\_call\_\_() of Operation so a Metric can be pipelined to an Operation. As Operation is a special kind of Metric, so it can bu further pipelined to another Operation. At last, compute_on() takes a Metric from the pipeline and is equavalent to calling metric.compute_on().
# + [markdown] id="d-OgV6EFCv-7"
# #Filter
#
# There is a "where" arg for Metric. It'll be passed to df.query() at the beginning of compute_on(df). By default the filter is not reflected in the name of Metric so same Metrics with different filters would have same column names in the returned DataFrames. It makes combining them easy.
# + colab={"height": 80} executionInfo={"elapsed": 421, "status": "ok", "timestamp": 1594365726294, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="mY1Fi24TDVqG" outputId="2abe5bc0-be81-4868-ce09-ccded55a5e67"
clicks_us = Sum('clicks', where='country == "US"')
clicks_not_us = Sum('clicks', where='country != "US"')
(clicks_not_us - clicks_us).compute_on(df)
# + [markdown] id="gqeuyU8HE8Ei"
# It's equivalent to
# + colab={"height": 111} executionInfo={"elapsed": 1167, "status": "ok", "timestamp": 1594365727507, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="hJfOsMVyEr8r" outputId="1456f744-7b95-4b37-e655-86a1c8cf6fdf"
Sum('clicks') | AbsoluteChange('country', 'US') | compute_on(df)
# + [markdown] id="T_HRZXdYrcf-"
# #SQL
#
# You can easily get SQL query for all built-in Metrics and Operations, except for weighted Quantile/CV/Correlation/Cov, by calling
#
# > to_sql(sql_table, split_by).
#
# You can also directly execute the query by calling
# > compute_on_sql(sql_table, split_by, execute, melted),
#
# where `execute` is a function that can execute SQL queries. The return is very similar to compute_on().
#
# The dialect it uses is the [standard SQL](https://cloud.google.com/bigquery/docs/reference/standard-sql) in Google Cloud's BigQuery.
#
# + executionInfo={"elapsed": 380, "status": "ok", "timestamp": 1608033025528, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="i9wHC71rred8" outputId="cea68e98-192e-44f2-bb5f-fd20886f5f35"
MetricList((Sum('X', where='Y > 0'), Sum('X'))).to_sql('T', 'grp')
# + colab={"height": 173} executionInfo={"elapsed": 477, "status": "ok", "timestamp": 1607761548536, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="LjAANaq0Zx4d" outputId="9d674807-ebf3-4f76-e48c-b0fc7cce3b54"
m = MetricList((Sum('clicks'), Mean('impressions')))
m = AbsoluteChange('country', 'US', m)
m.compute_on(df, 'platform')
# + colab={"height": 173} executionInfo={"elapsed": 410, "status": "ok", "timestamp": 1607761538409, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="hkayduNpC4UC" outputId="2511d051-b84a-4017-ccc9-dc0b52a51c18"
from sqlalchemy import create_engine
engine = create_engine('sqlite://', echo=False)
df.to_sql('T', con=engine)
# Meterstick uses a different SQL dialect from SQLAlchemy, so this doesn't
# always work.
m.compute_on_sql('T', 'platform', execute=lambda sql: pd.read_sql(sql, engine))
# + [markdown] id="AQjJAr3YcQB2"
# #Custom Metric
# We provide many Metrics out of box but we understand there are cases you need more, so we make it easy for you to write you own Metrics.
# First you need to understand the dataflow of a DataFrame when it's passed to compute_on(). The dataflow looks like this.
#
# <-------------------------------------------compute_on(handles caching)---------------------------------------------->
# <-------------------------------------compute_through-----------------------------------> |
# | <------compute_slices------> | |
# | |-> slice1 -> compute | | | |
# df -> df.query(where) -> precompute -> split_data -|-> slice2 -> compute | -> pd.concat -> postcompute -> manipulate -> final_compute
# |-> ... |
#
# In summary, compute() operates on a slice of data and hence only takes one arg, df. While precompute(), postcompute(), compute_slices(), compute_through() and final_compute() operate on the whole DataFrame so they take the df that has been processed by the dataflow till them and the split_by passed to compute_on(). final_compute() also has access to the original df passed to compute_on() for you to make additional manipulation. manipulate() does common data manipulation like melting and cleaning. Besides wrapping all the computations above, compute_on() also caches the result from compute_through(). Please refer to the section of Caching for more details.
#
# Depending on your case, you can overwrite all the methods, but we suggest you NOT to overwrite compute_on() because it might mess up the caching mechanism, nor manipulate(), because it might not work well with other Metrics' data manipulation. Here are some rules to help you decide.
# 1. If your Metric has no vectorization over slices, overwrite compute() which only takes one arg, df. To overwrite, you can either create a new class inheriting from Metric or just pass a lambda function into Metric.
# 2. If you have vectorization logic over slices, overwrite compute_slices().
# 3. As compute() operates on a slice of data, it doesn't have access to the columns to split_by and the index value of the slice. If you need them, overwrite compute_with_split_by(self, df, split_by, slice_name), which is just a wrapper of compute(), but has access to split_by and the value of current slice, slice_name.
# 4. The data passed into manipulate() should be a number, a pd.Series, or a wide/unmelted pd.DataFrame.
# 5. split_data() returns (sub_dataframe, corresponding slice value). You might want to overwrite it for non-vectorized Operations. See section Linear Regression for examples.
#
# Also there are some requirements.
# 1. Your Metric shouldn't change the input DataFrame inplace or it might not work with other Metrics.
# 2. Your Metric shouldn't rely on the index of the input DataFrame if you want it to work with Jackknife. The reason is Jackknife might reset the index.
#
# + [markdown] id="a_imRCi1gYa6"
# ## No Vectorization
# + colab={"height": 142} executionInfo={"elapsed": 521, "status": "ok", "timestamp": 1595025555146, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="uAgrTxLDfh3z" outputId="d7c34236-7960-45bc-f853-0911e45aed12"
class CustomSum(Metric):
def __init__(self, var):
name = 'custom sum(%s)' % var
super(CustomSum, self).__init__(name)
self.var = var
def compute(self, df):
return df[self.var].sum()
CustomSum('clicks').compute_on(df, 'country')
# + colab={"height": 142} executionInfo={"elapsed": 361, "status": "ok", "timestamp": 1594365728719, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="XgYXgTOTgPwo" outputId="82b69916-096d-4ba1-89f2-52bc3767804f"
Sum('clicks').compute_on(df, 'country')
# + [markdown] id="-hvgLLKbglrP"
# CustomSum doesn't have vectorization. It loops through the DataFrame and sum on every slice. As the result, it's slower than vectorized summation.
# + colab={"height": 34} executionInfo={"elapsed": 2958, "status": "ok", "timestamp": 1595025560571, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6MVo9EtTgg0e" outputId="017b05a1-882a-49a3-9d43-3fe2768b1f75"
#@test {"skip": true}
# %%timeit
CustomSum('clicks').compute_on(df, 'country')
# + colab={"height": 34} executionInfo={"elapsed": 1341, "status": "ok", "timestamp": 1595025567402, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="cLrEAl2Tgi_W" outputId="44f05eec-f3c8-42f5-ac74-24c23506ca02"
#@test {"skip": true}
# %%timeit
Sum('clicks').compute_on(df, 'country')
# + colab={"height": 34} executionInfo={"elapsed": 1065, "status": "ok", "timestamp": 1595025569063, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="qdhczptcg6VO" outputId="f091e1e0-71c7-4a03-ad96-ddb9a2e01f65"
#@test {"skip": true}
# %%timeit
df.groupby('country')['clicks'].sum()
# + [markdown] id="5tUfc996hNBm"
# ## With Vectorization
#
# We can do better. Let's implement a Sum with vectorization.
# + colab={"height": 142} executionInfo={"elapsed": 538, "status": "ok", "timestamp": 1594366815541, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="1ny3uHTuhXAJ" outputId="14cb7c24-140f-4ed0-d7cb-7bce888b0493"
class VectorizedSum(Metric):
def __init__(self, var):
name = 'vectorized sum(%s)' % var
super(VectorizedSum, self).__init__(name = name)
self.var = var
def compute_slices(self, df, split_by):
if split_by:
return df.groupby(split_by)[self.var].sum()
return df[self.var].sum()
VectorizedSum('clicks').compute_on(df, 'country')
# + colab={"height": 34} executionInfo={"elapsed": 7884, "status": "ok", "timestamp": 1594365752825, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="VekZlay-hoZ6" outputId="235f9aca-2e47-4dfa-ecde-bd57f7fc0b38"
#@test {"skip": true}
# %%timeit
VectorizedSum('clicks').compute_on(df, 'country')
# + [markdown] id="FtRTh1lLiA2W"
# ## Precompute, postcompute and final_compute
#
# They are useful when you need to preprocess and postprocess the data.
# + colab={"height": 264} executionInfo={"elapsed": 370, "status": "ok", "timestamp": 1594366947511, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="OrcVt-gviQv5" outputId="f8af947a-a643-45ec-b153-edb4b6bbec35"
class USOnlySum(Sum):
def precompute(self, df, split_by):
return df[df.country == 'US']
def postcompute(self, data, split_by):
print('Inside postcompute():')
print('Input data: ', data)
print('Input split_by: ', split_by)
print('\n')
return data
def final_compute(self, res, melted, return_dataframe, split_by, df):
# res is the result processed by the dataflow till now. df is the original
# DataFrme passed to compute_on().
print('Inside final_compute():')
for country in df.country.unique():
if country not in res.index:
print('Country "%s" is missing!' % country)
return res
USOnlySum('clicks').compute_on(df, 'country')
# + [markdown] id="xjT67nhPlMng"
# ## Overwrite using Lambda Functions
# For one-off Metrics, you can also overwrite precompute, compute, postcompute, compute_slices and final_compute by passing them to Metric() as lambda functions.
# + colab={"height": 142} executionInfo={"elapsed": 215, "status": "ok", "timestamp": 1634528716676, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="IqhEw210f7RG" outputId="9d0fc2fd-6ad1-4de6-f3d3-ba76a1975eb1"
normalize = metrics.Sum('clicks', postcompute=lambda res, split_by: res / res.sum())
normalize.compute_on(df, 'country')
# + colab={"height": 142} executionInfo={"elapsed": 111, "status": "ok", "timestamp": 1634528743698, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="rkUUmBajgQjb" outputId="9e110344-e3c1-4183-f677-0135091c10dd"
# The above is equivalent to Normalize by 'country'.
Normalize('country', Sum('clicks')).compute_on(df)
# + [markdown] id="8TJkDyF6o2aW"
# ##Custom Operation
# Writing a custom Operation is a bit more complex. Take a look at the Caching section below as well. Typically an Operation first computes its children Metrics with expanded split_by. Here are some rules to keep in mind.
# 1. Always use compute_on and compute_child to compute the children Metrics. They handle caching so your Operation can interact with other Metrics correctly.
# 2. If the Operation extends the split_by when computing children Metrics, you need to register the extra columns added in the \_\_init__().
# 3. The extra columns should come after the original split_by.
# 4. If you really cannot obey #2 or #3, you need to overwrite Operation.flush_children(), or it won't work with Jackknife and Bootstrap.
# 5. Try to vectorize the Operation as much as possible. At least you can compute the children Metrics in a vectorized way by calling compute_child(). It makes the caching of the children Metrics more available.
# 6. Jackknife takes shortcuts when computing leave-one-out (LOO) estimates for Sum, Mean and Count, so if you want your Operation to work with Jackknife fast, delegate computations to Sum, Mean and Count as much as possible. See section Linear Regression for a comparison.
# 7. For the same reason, you computation logic should avoid using input df other than in compute_on() and compute_child(). When cutting corners, Jackknife emits None as the input df for LOO estimation. The compute_on() and compute_child() functions know to read from cache but other functions may not know what to do. If your Operation uses df outside the compute_on() and compute_child() functions, you have either to
# * ensure that your computation doesn't break when df is None.
# * set attribute 'precomputable_in_jk' to False (which will force the jackknife to be computed the manual way, which is slower).
#
# Let's see Distribution for an example.
# + id="QWsivJZMpvgO"
class Distribution(Operation):
"""Computes the normalized values of a Metric over column(s).
Attributes:
extra_index: A list of column(s) to normalize over.
children: A tuple of a Metric whose result we normalize on. And all other
attributes inherited from Operation.
"""
def __init__(self,
over: Union[Text, List[Text]],
child: Optional[Metric] = None,
**kwargs):
self.over = over
# The 3rd argument is the extra column that will be added to split_by. It'll
# be converted to a list then assigned to self.extra_index.
super(Distribution, self).__init__(child, 'Distribution of {}', over,
**kwargs)
def compute_slices(self, df, split_by=None):
# extra_index is after the split_by.
lvls = split_by + self.extra_index if split_by else self.extra_index
res = self.compute_child(df, lvls)
total = res.groupby(level=split_by).sum() if split_by else res.sum()
return res / total
# + [markdown] id="5adfiJUjxIfa"
# ## SQL Generation
#
# If you want the custom Metric to generate SQL query, you need to implement to_sql() or get_sql_and_with_clause(). The latter is more common and recommended. Please refer to built-in Metrics to see how it should be implemented. Here we show two examples, one for Metric and the other for Operation.
# + id="Neiqbr-wxlej"
class SumWithSQL(SimpleMetric):
def __init__(self,
var: Text,
name: Optional[Text] = None,
where: Optional[Text] = None,
**kwargs):
super(SumWithSQL, self).__init__(var, name, 'sum({})', where, **kwargs)
self._sum = Sum(var, name, where, **kwargs)
def compute_slices(self, df, split_by):
return self._sum.compute_slices(df, split_by)
# All the SQL-related classes, like Datasource, Filters, Columns, and so on,
# are defined in sql.py.
def get_sql_and_with_clause(self, table: Datasource, split_by: Columns,
global_filter: Filters, indexes: Columns,
local_filter: Filters, with_data: Datasources):
del indexes # unused
# Always starts with this line unless you know what you are doing.
local_filter = Filters([self.where, local_filter]).remove(global_filter)
columns = Column(self.var, 'SUM({})', self.name, local_filter)
# Returns a Sql instance and the WITH clause it needs.
return Sql(columns, table, global_filter, split_by), with_data
# + colab={"height": 173} executionInfo={"elapsed": 1364, "status": "ok", "timestamp": 1628296647413, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "12474546967758012552"}, "user_tz": 420} id="rEFXXMFtCgh5" outputId="e9184c8e-7356-44ef-cc79-cbee0aa469d0"
m = Sum('clicks') - SumWithSQL('clicks', 'custom_sum')
m.compute_on_sql('T', 'platform', execute=lambda sql: pd.read_sql(sql, engine))
# + [markdown] id="rzUpMSEr8n4z"
# For an Operation, you ususally call the child metrics' get_sql_and_with_clause() to get the subquery you need.
# + executionInfo={"elapsed": 545, "status": "ok", "timestamp": 1615428210116, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="k8p1y7UG8mg1" outputId="c955b932-fd47-4e5e-fc9e-a992ef5ace4c"
class DistributionWithSQL(Operation):
def __init__(self,
over: Union[Text, List[Text]],
child: Optional[Metric] = None,
**kwargs):
super(DistributionWithSQL, self).__init__(child, 'Distribution of {}', over,
**kwargs)
def compute_slices(self, df, split_by=None):
lvls = split_by + self.extra_index if split_by else self.extra_index
res = self.compute_child(df, lvls)
total = res.groupby(level=split_by).sum() if split_by else res.sum()
return res / total
def get_sql_and_with_clause(self,
table: Datasource,
split_by: Columns,
global_filter: Filters,
indexes: Columns,
local_filter: Filters,
with_data: Datasources):
"""Gets the SQL query and WITH clause.
The query is constructed by
1. Get the query for the child metric.
2. Keep all indexing/groupby columns unchanged.
3. For all value columns, get
value / SUM(value) OVER (PARTITION BY split_by).
Args:
table: The table we want to query from.
split_by: The columns that we use to split the data.
global_filter: The Filters that can be applied to the whole Metric tree.
indexes: The columns that we shouldn't apply any arithmetic operation.
local_filter: The Filters that have been accumulated so far.
with_data: A global variable that contains all the WITH clauses we need.
Returns:
The SQL instance for metric, without the WITH clause component.
The global with_data which holds all datasources we need in the WITH
clause.
"""
# Always starts with this line unless you know what you are doing.
local_filter = Filters([self.where, local_filter]).remove(global_filter)
# The intermediate tables needed by child metrics will be added to with_data
# in-place.
child_sql, with_data = self.children[0].get_sql_and_with_clause(
table, indexes, global_filter, indexes, local_filter, with_data)
child_table = sql.Datasource(child_sql, 'DistributionRaw')
# Always use the alias returned by with_data.add(), because if the with_data
# already holds a different table that also has 'DistributionRaw' as its
# alias, we'll use a different alias for the child_table, which is returned
# by with_data.add().
child_table_alias = with_data.add(child_table)
groupby = sql.Columns(indexes.aliases, distinct=True)
columns = sql.Columns()
for c in child_sql.columns:
if c.alias in groupby:
continue
col = sql.Column(c.alias) / sql.Column(
c.alias, 'SUM({})', partition=split_by.aliases)
col.set_alias('Distribution of %s' % c.alias_raw)
columns.add(col)
return sql.Sql(groupby.add(columns), child_table_alias), with_data
m = DistributionWithSQL('country', Sum('clicks'))
m.to_sql('T')
# + [markdown] id="aaaU_93icO6e"
# # Caching
#
# tl;dr: Reuse Metrics as much as possible and compute them together.
#
# Computation can be slow so it'd nice if we pass in the same DataFrame multiple
# times the computation is actually only done once. The difficulty is that
# DataFrame is mutable so it's hard to decide whether we really saw this DataFrame
# before. However, in one round of compute_on(), the DataFrame shouldn't change
# (our Metrics never change the original DataFrame and your custom Metrics
# shouldn't either), so we can cache the result, namely, a Metric appearing in
# multiple places will only be computed once. This all happens automatically so
# you don't need to worry about it. If you really cannot compute all your Metrics
# in one round, there is a "cache_key" arg in compute_on(). What it does is
#
# 1. if the key is in cache, just read the cache;
# 2. if not, compute and save the result to cache under the key.
#
# Note:
# 1. All we check is cache_key, nothing more, so it's your responsibility to
# make sure same key really corresponds to the same input DataFrame AND split_by.
# 2. The caching and retrieving happen in all levels of Metrics, so
#
# `PercentChange(..., Sum('x')).compute_on(df, cache_key='foo')`
#
# not only cache the percent change to PercentChange's cache, but also cache
#
# `Sum('x').compute_through(df)`
#
# to Sum('x')'s cache. Note it's the output of compute_through() is cached so we
# do't need to re-compute just because you change "melted" from True to False.
# 3. Anything that can be a key of a dict can be used as cache_key, except '_RESERVED' and tuples like ('_RESERVED', ...).
#
# First, let's illustrate that when we don't reuse Metrics, everything gets
# computed once as expected.
# + colab={"height": 131} executionInfo={"elapsed": 393, "status": "ok", "timestamp": 1600044635745, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="Y5EXq9oacJUm" outputId="13b9216b-e029-4356-b465-b0dbde2624b9"
class SumWithTrace(Sum):
def compute_through(self, data, split_by):
print('Computing %s...' % self.name)
return super(SumWithTrace, self).compute_through(data, split_by)
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
ctr = SumWithTrace('clicks') / SumWithTrace('impressions')
MetricList((sum_clicks, ctr)).compute_on(df)
# + [markdown] id="fMYFs2ZwvwCx"
# Now let's see what heppens if we reuse sum_clicks.
# + colab={"height": 114} executionInfo={"elapsed": 380, "status": "ok", "timestamp": 1600044639096, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="EiBXMLQbu-Eg" outputId="de516266-32f7-4767-a26e-358e3e435801"
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
ctr = sum_clicks / SumWithTrace('impressions')
MetricList((sum_clicks, ctr)).compute_on(df)
# + [markdown] id="7kXxnS41wOwj"
# Then sum_clicks only gets computed once. For Metics that are not quite compatible, you can still put them in a MeticList and set return_dataframe to False to maximize the caching.
# + colab={"height": 145} executionInfo={"elapsed": 781, "status": "ok", "timestamp": 1600044820809, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="VesfUvBoF00L" outputId="d930aac3-06bf-4d52-e0f2-67cd996cef4c"
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
jk, s = MetricList(
[Jackknife('cookie', sum_clicks), sum_clicks],
children_return_dataframe=False).compute_on(
df, return_dataframe=False)
print(s)
jk
# + [markdown] id="pvcgzS3EGYj8"
# If you really cannot compute Metrics together, you can use a cache_key.
# + colab={"height": 176} executionInfo={"elapsed": 589, "status": "ok", "timestamp": 1596160622607, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="jywqs9-PvXXP" outputId="e99a88c2-3b00-4148-d41c-bbd5b0f40e87"
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
ctr = sum_clicks / SumWithTrace('impressions')
sum_clicks.compute_on(df, 'country', cache_key='foo')
ctr.compute_on(df, 'country', cache_key='foo')
# + [markdown] id="dEwtDv2ayFzW"
# The resulte are cached in ctr, a composite Metric, as well as its children, the Sum Metrics.
# + colab={"height": 148} executionInfo={"elapsed": 499, "status": "ok", "timestamp": 1594367514617, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="SjXfLwQ2wgU1" outputId="d68f15c2-39dc-4b2b-df64-3718bca6ffba"
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
ctr = sum_clicks / SumWithTrace('impressions')
MetricList((sum_clicks, ctr)).compute_on(df, cache_key='foo')
print('sum_clicks cached: ', sum_clicks.get_cached('foo'))
print('ctr cached: ', ctr.get_cached('foo'))
ctr.compute_on(None, cache_key='foo')
# + [markdown] id="7xpo6Taoybcj"
# You can flush the cache by calling flush_cache(key, split_by=None, recursive=True, prune=True), where "recursive" means if you want to flush the cache of the children Metrics as well, and "prune" means if the key is not found in current Metric, do you still want to flush the children Metrics or stop early. It's useful when a high level Metric appears in several places then during the flushing we will hit it multiple times. We can save time by stop early.
# + colab={"height": 51} executionInfo={"elapsed": 414, "status": "ok", "timestamp": 1594367740852, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="sXC0q_DEyVqR" outputId="23f6dfb6-1a5a-49b6-a1fa-166fec85f9c8"
sum_clicks = SumWithTrace('clicks', 'sum of clicks')
ctr = sum_clicks / SumWithTrace('impressions')
MetricList((sum_clicks, ctr)).compute_on(df, cache_key='foo')
ctr.flush_cache('foo', recursive=False)
# + colab={"height": 80} executionInfo={"elapsed": 469, "status": "ok", "timestamp": 1594367707179, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="ArsM6q90uH8d" outputId="e96bfcb2-2226-4c1b-8154-70b271f16f93"
sum_clicks.compute_on(None, cache_key='foo') # sum is not flushed.
# + colab={"height": 34} executionInfo={"elapsed": 334, "status": "ok", "timestamp": 1594365757078, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="NTHaPN4uz2Tk" outputId="5524f682-423d-4c91-f02a-c6781f5fd016"
ctr.in_cache('foo')
# + [markdown] id="fgsrP8_21cPX"
# Though ctr's cache has been flushed, we can still compute ctr from cache because all its children are cached.
# + colab={"height": 80} executionInfo={"elapsed": 380, "status": "ok", "timestamp": 1594365757514, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="c-ZFVlsVzyQx" outputId="1bec4d29-3def-4ebb-a12b-98f3a71cb5c2"
ctr.compute_on(None, cache_key='foo')
# + colab={"height": 34} executionInfo={"elapsed": 367, "status": "ok", "timestamp": 1594365757924, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="9Wy-0jye1jpQ" outputId="baf8b6a5-28c7-4cc9-a354-76cbda9daeeb"
ctr.in_cache('foo')
# + [markdown] id="JgEIhwL01vPr"
# We won't be able to re-compute ctr if we recursively flush its cache.
# + colab={"height": 341} executionInfo={"elapsed": 1724, "status": "error", "timestamp": 1594365759687, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="10p7-ZYpzlSQ" outputId="08b94fab-fd7f-40ab-9777-b3b498ec91e9"
#@test {"skip": true}
ctr.flush_cache('foo')
sum_clicks.compute_on(None, cache_key='foo') # sum is flushed too.
# + [markdown] id="ejgnC-PGep2j"
# However, the behavior becomes subtle when Operation is involved.
# + colab={"height": 159} executionInfo={"elapsed": 488, "status": "ok", "timestamp": 1596161299762, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="8b9JEg6Je-Xl" outputId="824906ef-dd41-46db-f1cc-e5a59f9a1c03"
sum_clicks = SumWithTrace('clicks')
PercentChange('country', 'US', sum_clicks).compute_on(df, cache_key=42)
sum_clicks.compute_on(None, 'country', cache_key=42)
# + [markdown] id="0VbUCE3KfJcV"
# Note that it's sum_clicks.compute_on(df, 'country') instead of sum_clicks.compute_on(df) got saved in the cache. The reason is we need the former not the latter to compute the PercentChange. Using sum_clicks.compute_on(df, cache_key=42) will always give you the right result so it's not a big issue, just might confuse you sometime.
# + colab={"height": 97} executionInfo={"elapsed": 381, "status": "ok", "timestamp": 1596161302605, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="qlxl_hqIkU-Z" outputId="8df8ba88-84b5-4d2a-845c-de43265df207"
sum_clicks.compute_on(df, cache_key=42)
# + colab={"height": 142} executionInfo={"elapsed": 432, "status": "ok", "timestamp": 1596161868474, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "12474546967758012552"}, "user_tz": 420} id="tW__O9JZkfEC" outputId="70208a20-2d74-417d-c66c-7aa4e884d9f4"
sum_clicks.compute_on(df, 'country', cache_key=42)
# + [markdown] id="H6MJwaLlZ79Y"
# #Advanced Examples
# + [markdown] id="gcmROLlqcIYa"
# ## Click Split
# + colab={"height": 173} executionInfo={"elapsed": 689, "status": "ok", "timestamp": 1594367859739, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6lDnUPdzcJuX" outputId="a1dcc94f-7ccf-4ad1-e8f7-d7dd99c7df90"
np.random.seed(42)
df['duration'] = np.random.random(len(df)) * 200
long_clicks = Sum('clicks', where='duration > 60')
short_clicks = Sum('clicks', where='duration < 30')
click_split = (long_clicks / short_clicks).set_name('click split')
click_split | Jackknife('cookie') | compute_on(df, 'country')
# + [markdown] id="8Vplq4TDzwBY"
# ##Difference in differences
# + colab={"height": 173} executionInfo={"elapsed": 581, "status": "ok", "timestamp": 1601684321443, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="NPiALqlMG0aM" outputId="4f305de8-6891-4031-f1c9-958440a40e72"
np.random.seed(42)
df['period'] = np.random.choice(('preperiod', 'postperiod'), size=size)
sum_clicks = Sum('clicks')
ctr = sum_clicks / Sum('impressions')
metrics = (sum_clicks, ctr)
preperiod_clicks = MetricList(metrics, where='period == "preperiod"')
postperiod_clicks = MetricList(metrics, where='period == "postperiod"')
pct = PercentChange('platform', 'Desktop')
did = (pct(postperiod_clicks) - pct(preperiod_clicks)).rename_columns(
['clicks% DID', 'ctr% DID'])
Jackknife('cookie', did).compute_on(df)
# + [markdown] id="3ItlGoihbYRz"
# ## Compare the standard errors between Jackknife and Bootstrap
# + colab={"height": 173} executionInfo={"elapsed": 3716, "status": "ok", "timestamp": 1595026875847, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="wU8M4nTA0dgD" outputId="fbc207e4-9724-4060-93a9-6516f42c7169"
np.random.seed(42)
sum_clicks = Sum('clicks')
ctr = sum_clicks / Sum('impressions')
metrics = MetricList((sum_clicks, ctr))
(Jackknife('cookie', metrics) /
Bootstrap('cookie', metrics, 100)).rename_columns(
pd.MultiIndex.from_product(
(('sum(clicks)', 'ctr'), ('Value', 'SE')))).compute_on(df, 'country')
# + [markdown] id="GqiuYVJGGsPT"
# ## Linear Regression
#
# Here we fit a linear regression on mean values of groups. We show two versions, the former delgates computations to Mean so its Jackknife is faster than the latter which doesn't delegate.
# + id="yPn6aNBzAbC_"
np.random.seed(42)
size = 1000000
df_lin = pd.DataFrame({'grp': np.random.choice(range(10), size=size)})
df_lin['x'] = df_lin.grp + np.random.random(size=size)
df_lin['y'] = 2 * df_lin.x + np.random.random(size=size)
df_lin['cookie'] = np.random.choice(range(20), size=size)
df_lin_mean = df_lin.groupby('grp').mean()
# + colab={"height": 274} executionInfo={"elapsed": 809, "status": "ok", "timestamp": 1600055117412, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="qQZV_quPGvct" outputId="1ab1ebae-3ec8-46ae-da2a-b5a68e187207"
#@test {"skip": true}
plt.scatter(df_lin_mean.x, df_lin_mean.y)
plt.show()
# + colab={"height": 142} executionInfo={"elapsed": 1732, "status": "ok", "timestamp": 1600048693850, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="q7InRdD7HsoD" outputId="7cb8de2c-0d3a-4945-edff-f4373ebea9da"
from sklearn import linear_model
class LinearReg(Operation):
def __init__(self, x, y, grp):
self.lm = linear_model.LinearRegression()
# Delegate most of the computations to Mean Metrics.
child = MetricList((Mean(x), Mean(y)))
self.grp = grp
# Register grp as the extra_index.
super(LinearReg, self).__init__(child, '%s ~ %s' % (y, x), grp)
def split_data(self, df, split_by=None):
"""The 1st element in yield will be passed to compute()."""
if not split_by:
yield self.compute_child(df, self.grp), None
else:
# grp needs to come after split_by.
child = self.compute_child(df, split_by + [self.grp])
keys, indices = list(zip(*child.groupby(split_by).groups.items()))
for i, idx in enumerate(indices):
yield child.loc[idx.unique()].droplevel(split_by), keys[i]
def compute(self, df):
self.lm.fit(df.iloc[:, [0]], df.iloc[:, 1])
return pd.Series((self.lm.coef_[0], self.lm.intercept_))
lr = LinearReg('x', 'y', 'grp')
Jackknife('cookie', lr, 0.95).compute_on(df_lin)
# + colab={"height": 142} executionInfo={"elapsed": 2532, "status": "ok", "timestamp": 1600048697347, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6PdjGmjqTmrl" outputId="a6282960-fbe7-4293-b88f-d0e0e1a9d370"
class LinearRegSlow(Metric):
def __init__(self, x, y, grp):
self.lm = linear_model.LinearRegression()
# Doesn't delegate.
self.x = x
self.y = y
self.grp = grp
super(LinearRegSlow, self).__init__('%s ~ %s' % (y, x))
def split_data(self, df, split_by=None):
"""The 1st element in yield will be passed to compute()."""
idx = split_by + [self.grp] if split_by else self.grp
mean = df.groupby(idx).mean()
if not split_by:
yield mean, None
else:
keys, indices = list(zip(*mean.groupby(split_by).groups.items()))
for i, idx in enumerate(indices):
yield mean.loc[idx.unique()].droplevel(split_by), keys[i]
def compute(self, df):
self.lm.fit(df.iloc[:, [0]], df.iloc[:, 1])
return pd.Series((self.lm.coef_[0], self.lm.intercept_))
lr_slow = LinearRegSlow('x', 'y', 'grp')
Jackknife('cookie', lr_slow, 0.95).compute_on(df_lin)
# + colab={"height": 34} executionInfo={"elapsed": 7068, "status": "ok", "timestamp": 1600048714606, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="aNLbzk-CUgqm" outputId="bb18f495-e310-43cb-cc51-85da74273eb5"
#@test {"skip": true}
# %%timeit
Jackknife('cookie', lr, 0.95).compute_on(df_lin)
# + colab={"height": 34} executionInfo={"elapsed": 9558, "status": "ok", "timestamp": 1600048724249, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="a3BBhzbdUUwo" outputId="9498d586-421a-4812-9a4d-1648688c33ec"
#@test {"skip": true}
# %%timeit
Jackknife('cookie', lr_slow, 0.95).compute_on(df_lin)
# + [markdown] id="QFjhj96EdK-r"
# ## LOWESS
# + colab={"height": 359} executionInfo={"elapsed": 433, "status": "ok", "timestamp": 1594367896139, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="2e_ttZzta7JH" outputId="c3735e57-0b88-4f25-8b87-8615b38ba5dc"
# Mimics that measurements, y, are taken repeatedly at a fixed grid, x.
np.random.seed(42)
size = 10
x = list(range(5))
df_sin = pd.DataFrame({'x': x * size, 'cookie': np.repeat(range(size), len(x))})
df_sin['y'] = np.sin(df_sin.x) + np.random.normal(scale=0.5, size=len(df_sin.x))
df_sin.head(10)
# + id="Eei8Kd0wd-Gt"
import statsmodels.api as sm
lowess = sm.nonparametric.lowess
class Lowess(Metric):
def __init__(self, x, y, name=None, where=None):
self.x = x
self.y = y
name = name or 'LOWESS(%s ~ %s)' % (y, x)
super(Lowess, self).__init__(name, where=where)
def compute(self, data):
lowess_fit = pd.DataFrame(
lowess(data[self.y], data[self.x]), columns=[self.x, self.y])
return lowess_fit.drop_duplicates().reset_index(drop=True)
# + colab={"height": 204} executionInfo={"elapsed": 383, "status": "ok", "timestamp": 1594367897604, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="qvp8ihsnlY-d" outputId="112dda48-2b3c-4169-bc9e-9bb4b1513ea0"
Lowess('x', 'y') | compute_on(df_sin)
# + colab={"height": 274} executionInfo={"elapsed": 735, "status": "ok", "timestamp": 1594367899224, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="3eR0fV8feyqu" outputId="a868fae4-9d36-42f0-c4c2-130cff77a74c"
#@test {"skip": true}
jk = Lowess('x', 'y') | Jackknife('cookie', confidence=0.9) | compute_on(df_sin)
point_est = jk[('y', 'Value')]
ci_lower = jk[('y', 'Jackknife CI-lower')]
ci_upper = jk[('y', 'Jackknife CI-upper')]
plt.scatter(df_sin.x, df_sin.y)
plt.plot(x, point_est, c='g')
plt.fill_between(
x, ci_lower,
ci_upper,
color='g',
alpha=0.5)
plt.show()
# + [markdown] id="bEweF3aFZUlq"
# ## Coefficient Shrikage
# + colab={"height": 464} executionInfo={"elapsed": 1374, "status": "ok", "timestamp": 1634531316077, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="zqBCQAu4OBR2" outputId="2d73d8dc-d1fe-4d96-8420-77a190c91edb"
#@test {"skip": true}
from plotnine import ggplot, geom_point, geom_ribbon, aes, ylab
y = Mean('clicks')
x = [Mean('impressions'), Variance('impressions')]
grpby = 'platform'
baseline = LinearRegression(y, x, grpby, fit_intercept=False)
shrinkage = [(Ridge(y, x, grpby, a, False) / baseline).rename_columns(
('%s::mean(impressions)' % a, '%s::var(impressions)' % a))
for a in range(10)]
jk = (MetricList(shrinkage)
| Jackknife('cookie', confidence=0.95)
| compute_on(df, melted=True)).reset_index()
jk[['penalty', 'X']] = jk.Metric.str.split('::', expand=True)
jk.penalty = jk.penalty.astype(int)
(ggplot(jk, aes('penalty', 'Value', color='X'))
+ ylab('Shrinkage')
+ geom_point()
+ geom_ribbon(
aes(ymin='Jackknife CI-lower', ymax='Jackknife CI-upper', fill='X'),
alpha=0.1))
| meterstick_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
# ### Trim a Binary Search Tree
#
# Given a binary search tree and the lowest and highest boundaries as L and R, trim the tree so that all its elements lies in [L, R] (R >= L). You might need to change the root of the tree, so the result should return the new root of the trimmed binary search tree.
#
# See [The LeetCode example problem](https://leetcode.com/problems/trim-a-binary-search-tree/description/)
# +
debugging = False
debugging = True
def dprint(f, *args):
if debugging:
print((' DBG:' + f).format(*args))
# +
# Definition for a binary tree node.
# class TreeNode(object):
class TreeNode():
""" A tree node """
def __init__(self, value):
self.value = value
self.left = None
self.right = None
def link(self, left, right):
self.left = left
self.right = right
def minv(self):
if self.left: return self.left.minv()
if self.value: return self.value
return self.right.minv()
def maxv(self):
if self.right: return self.right.maxv()
if self.value: return self.value
return self.left.maxv()
def depth(self, current=0):
ldepth = current if self.left is None else self.left.depth(current+1)
rdepth = current if self.right is None else self.right.depth(current+1)
d = max(ldepth, rdepth)
dprint("--- [{0}] c={1} d={2}", self.value, current, d)
return d
def ldepth(self):
return 0 if self.left is None else self.left.depth(1)
def rdepth(self):
return 0 if self.right is None else self.right.depth(1)
def insert(self, v):
rootv = self.value
if rootv is None:
self.value = v
elif v < rootv:
if self.left is None:
self.left = TreeNode(v)
else:
self.left.insert(v)
else:
if self.right is None:
self.right = TreeNode(v)
else:
self.right.insert(v)
def __str__(self):
if not self:
return "()"
else:
leftstr = '.' if self.left is None else "{0}, ".format(str(self.left))
rightstr = '.' if self.right is None else ", {0}".format(str(self.right))
return "({0}[{1}]{2}])".format(leftstr, str(self.value), rightstr)
def liststr(self):
if not self:
return "(*)"
else:
leftstr = '' if self.left is None else "{0}".format(self.left.liststr())
rightstr = '' if self.right is None else "{0}".format(self.right.liststr())
return "{0}{1}, {2}".format(leftstr, str(self.value), rightstr)
def tolist(self):
llist = [] if self.left == None else self.left.tolist();
rlist = [] if self.right == None else self.right.tolist();
return llist + [self.value] + rlist
def inorder(self):
""" A recursive generator that generates Tree leaves in in-order """
if self:
if self.left:
for x in self.left.inorder():
yield x
yield self.value
if self.right:
for x in self.right.inorder():
yield x
__repr__ = __str__
# -
y = TreeNode(None)
y.insert(3)
y.insert(12)
y.insert(5)
y.insert(1)
y.insert(9)
print(y)
y.depth()
print([x for x in y.inorder()])
[3, 4, 5] + [55,] + [6,]
import turtle
# #### A Tree Viewer
#
# [See <NAME>'s code](https://discuss.leetcode.com/topic/16600/tree-deserializer-and-visualizer-for-python)
# +
# External turtle window
import turtle
# -- Limited function turtle that draws directly in Jupyter
# from mobilechelonian import Turtle
# +
class TreeNode:
def __init__(self, val, left=None, right=None):
self.val = val
self.left = left
self.right = right
def __repr__(self):
return 'TreeNode({})'.format(self.val)
def deserialize(string):
if string == '{}':
return None
nodes = [None if val == 'null' else TreeNode(int(val))
for val in string.strip('[]{}').split(',')]
kids = nodes[::-1]
root = kids.pop()
for node in nodes:
if node:
if kids: node.left = kids.pop()
if kids: node.right = kids.pop()
return root
def drawtree(root):
def height(root):
return 1 + max(height(root.left), height(root.right)) if root else -1
def jumpto(x, y):
t.penup()
t.goto(x, y)
t.pendown()
def draw(node, x, y, dx):
if node:
t.goto(x, y)
jumpto(x, y-20)
t.write(node.val, align='center', font=('Arial', 12, 'normal'))
draw(node.left, x-dx, y-60, dx/2)
jumpto(x, y-20)
draw(node.right, x+dx, y-60, dx/2)
t = turtle.Turtle()
#t = Turtle()
turtle_window = turtle.Screen() # JF
print("Click on Turtle Screen to Continue.")
t.speed(0)
turtle.delay(0)
h = height(root)
jumpto(0, 30*h)
draw(root, 0, 30*h, 40*h)
t.hideturtle()
turtle_window.exitonclick() # JF
try:
turtle.mainloop()
except turtle.Terminator as ex:
print("Done.".format(ex))
# -
drawtree(deserialize('[1,2,3,null,null,4,null,null,5]'))
drawtree(deserialize('[2,1,3,0,7,9,1,2,null,1,0,null,null,8,8,null,null,null,null,7]'))
# ETE code is fully functional under IPython Notebook environments, providing inline image visualization.
#
# Inline plots can be generated by calling the tree.render() function with the "%%inline" word as its first argument. This will produce a SVG image of your tree that is directly shown by the browser.
#
# tree.render("%%inline")
# In addition, ETE's Graphical User Interface (GUI) can be launched at any moment for a complete interactive interaction with your tree objects. Just call the tree.show() method available in every node. Browse, explore or even modify your tree. When the GUI is closed, changes will be reflected in your Ipython notebook code.
#
# Layout functions, node faces, styles and all drawing features from ETE treeview module will also be accessible. Phylogenetic trees can even been displayed together with their alignments:
from ete3 import Tree
t = Tree("((A, B)Internal_1:0.7, (C, D)Internal_2:0.5)root:1.3;", format=1)
t.add_features(size=11)
print(t.get_ascii(attributes=["name", "dist", "size"]))
t.render("%%inline")
t.render('')
| BSTTrim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning Project
# ###### The current project relies on using CNNs in order to process sperm data.
#
# The goal is to develop a classifier able to identify different morphologies of head spearm cells.
#
#
#
# #### CONTEXT :
#
# Due to our background in the areas of biology and health, it was in our interest to use this project to develop our knowledge of neural network, in microscopic images.
# The chosen theme is a classification problem taking into account the morphology of the sperm cell heads.
#
# #### There are two datasets available, which we have chosen to aggregate into one.
#
# - SCIAN-MorphoSpermGS - https://cimt.uchile.cl/gold10/ - with 1132 image, each image has 35 x 35 pixels and has been classified by 3 experts. We will use majority vote result as target.
#
#
# - Human Sperm head Morphology dataset – HuSHeM- https://easy.dans.knaw.nl/ui/datasets/id/easy-dataset:77214/tab/2 - with 725 images were taken, each image has 576×720 pixels and have the respective classification
# <h3><center> The heads of sperm cells will have the following classification:</center></h3>
#
#
# 
#
#
# * We don't include the "small" class !
#
# Ref: https://doi.org/10.1016/j.compbiomed.2019.103342
# -> The process of adapting the data for use on CNN for reasons of operating system compatibility has been removed. But they can be found in [this document.][mylink]
#
# [mylink]: DL_Project_LoadImg.html
# ###### In this pipeline, 4 models were developed:
#
# - The first model was produced by us, based on heuristics.
# - The second model was developed based on the Alex Net network architecture
# - The third model was based on the Resnet50 model.
# - The last model was based on an Efficient Net which, according to the current literature, has very promising results.
# +
#base libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import random
import matplotlib.image as mpimg
# %matplotlib inline
from timeit import default_timer as timer
#image manipulation packages
import cv2
import os
from tqdm import tqdm
from glob import glob
from sklearn.metrics import confusion_matrix,classification_report
#Classification
from sklearn.metrics import classification_report, confusion_matrix, multilabel_confusion_matrix
sns.set()
#import keras
from tensorflow import keras
import glob
import cv2
import shutil
import random
import Augmentor
#Data Augmentation
from keras.preprocessing.image import ImageDataGenerator
# Callbacks
from tensorflow.keras import callbacks
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.callbacks import ReduceLROnPlateau
import keras
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout, Flatten, Conv2D, MaxPooling2D
from keras.layers.normalization import BatchNormalization
## RESNET50
from keras.applications.resnet50 import preprocess_input
from tensorflow.keras.applications.resnet50 import ResNet50
### Efficient
from tensorflow.keras.applications import EfficientNetB7
from tensorflow.keras import models
from tensorflow.keras import layers
from tensorflow.keras import callbacks
# -
# # 1- Import Images and Dataframe
#
dataframe=pd.read_csv("Majority_Vote.csv")
os.chdir('../CNN_SpermCells')
path = os.getcwd()
files = os.listdir(path)
path
# +
#Count the number of images by class
def fileCount(folder):
"count the number of files in a directory"
count = 0
for filename in os.listdir(folder):
path = os.path.join(folder, filename)
if os.path.isfile(path):
count += 1
elif os.path.isdir(path):
count += fileCount(path)
return count
count_0 = fileCount(path+'/path/class0')
count_1 = fileCount(path+'/path/class1')
count_2 = fileCount(path+'/path/class2')
count_3 = fileCount(path+'/path/class3')
print(f'Images Normal: {count_0}\n' +
f'Images Tapered: {count_1}\n' +
f'Images Pyriform: {count_2}\n' +
f'Images Amorphous: {count_3}\n')
# -
# # 2- Preprocessing
# ### 2.1 DATA AUGMENTATION
# #### 2.1 Offline Data Augmentation
#
# Options are limited since we're working with microscopy data
#
#
# +
# Define augmentation pipelines
class_0 = Augmentor.Pipeline(path+'/path/train/class0')
class_1 = Augmentor.Pipeline(path+'/path/train/class1')
class_2 = Augmentor.Pipeline(path+'/path/train/class2')
class_3 = Augmentor.Pipeline(path+'/path/train/class3')
#rotate by a maximum of 90 degrees
class_0.rotate(probability=0.7, max_left_rotation=25, max_right_rotation=25)
class_1.rotate(probability=0.7, max_left_rotation=25, max_right_rotation=25)
class_2.rotate(probability=0.7, max_left_rotation=25, max_right_rotation=25)
class_3.rotate(probability=0.7, max_left_rotation=25, max_right_rotation=25)
#mirroring, vertical or horizontal, randomly
class_0.flip_random(probability=0.7)
class_1.flip_random(probability=0.7)
class_2.flip_random(probability=0.7)
class_3.flip_random(probability=0.7)
# Augment images to the same proportion as existing ones in class 4 (majority class - get to 1000 in each class)
class_0.sample(6400 - count_0)
class_1.sample(6400 - count_1)
class_2.sample(6400 - count_2)
class_3.sample(6400 - count_3)
# -
# #### 2.2 Define Callbacks
# - Checkpoint
# - Early Stopping
# - Reduce Learning Rate
# - Timer
# +
checkpoint_filepath = path+'/Models/'
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=5, min_lr=0, verbose = 1)
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=20)
mc = ModelCheckpoint(
filepath=checkpoint_filepath,
verbose = 1,
monitor='val_accuracy',
mode='max',
save_best_only=True)
class TimingCallback(keras.callbacks.Callback):
def __init__(self, logs={}):
self.logs=[]
def on_epoch_begin(self, epoch, logs={}):
self.starttime = timer()
def on_epoch_end(self, epoch, logs={}):
self.logs.append(timer()-self.starttime)
cb = TimingCallback()
# -
# ## Models:
# - <a href='#1'>First Model</a>
# - <a href='#2'>Alex Net</a>
# - <a href='#3'>ResNet50</a>
# - <a href='#4'>EfficientNet B7</a>
# # 3. First Model
# <a id='1'></a>
checkpoint_filepath = path+'/Models/model_HM.h5'
# #### 3.1 Preprocess input
# Online Data Augmentation in GrayScale
# +
cnn_train_datagen = ImageDataGenerator(rescale = 1./255,
rotation_range = 5,
width_shift_range = 0.1,
height_shift_range = 0.1,
vertical_flip = True,
horizontal_flip = True,
brightness_range=[0.2,1.5],
fill_mode='nearest',
zoom_range = 0.2,
)
cnn_val_datagen = ImageDataGenerator(rescale = 1./255)
cnn_test_datagen = ImageDataGenerator(rescale = 1./255)
#test different color maps - class modes and cross validation types
cnn_training = cnn_train_datagen.flow_from_directory(path+'/path/train',
target_size = (32, 32),
batch_size = 64,
shuffle = True,
class_mode="categorical",
color_mode = 'grayscale')
cnn_val = cnn_val_datagen.flow_from_directory(path+'/path/val',
target_size = (32, 32),
batch_size = 64,
shuffle = True,
class_mode="categorical",
color_mode = 'grayscale')
cnn_test = cnn_test_datagen.flow_from_directory(path+'/path/test',
target_size = (32, 32),
batch_size = 1,
shuffle = True,
class_mode="categorical",
color_mode = 'grayscale')
# -
# #### 3.2 Build model
# +
cnn_model = Sequential()
#convolutional layer with 32 3x3 filters - again, arbitrary
cnn_model.add(Conv2D(filters=32, kernel_size=(3,3), padding = 'same', activation='relu'))
cnn_model.add(BatchNormalization())
#MaxPooling - takes the max value of each 2x2 pool in the feature map
cnn_model.add(MaxPooling2D(pool_size=(2,2)))
cnn_model.add(BatchNormalization())
#3rd convolution with 64 filters
cnn_model.add(Conv2D(filters=64, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
#4rd convolution with 128 filters
cnn_model.add(Conv2D(filters=128, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
#Second MaxPool2D - to check with other options
cnn_model.add(MaxPooling2D(pool_size=(2,2)))
cnn_model.add(BatchNormalization())
#5th convolutional layer with 24 3x3 filters - again, arbitrary
cnn_model.add(Conv2D(filters=32, kernel_size=(4,4), padding = 'same', activation='relu'))
cnn_model.add(BatchNormalization())
#second convolution with 36 filters
cnn_model.add(Conv2D(filters=64, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
cnn_model.add(Conv2D(filters=128, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
#Second MaxPool2D - to check with other options
cnn_model.add(MaxPooling2D(pool_size=(2,2)))
cnn_model.add(BatchNormalization())
#convolutional layer with 24 3x3 filters - again, arbitrary,
cnn_model.add(Conv2D(filters=128, kernel_size=(4,4), padding = 'same', activation='relu'))
cnn_model.add(BatchNormalization())
#second convolution with 36 filters
cnn_model.add(Conv2D(filters=128, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
cnn_model.add(Conv2D(filters=128, kernel_size=(3,3), padding = 'same', activation= 'relu'))
cnn_model.add(BatchNormalization())
#Second MaxPool2D - to check with other options
cnn_model.add(MaxPooling2D(pool_size=(2,2)))
cnn_model.add(BatchNormalization())
#the result of kthe CNN is then flattened and placed into the
cnn_model.add(Flatten())
cnn_model.add(Dense(128, activation='relu'))
cnn_model.add(BatchNormalization())
#Add Dropout
cnn_model.add(Dropout(0.4))
cnn_model.add(Dense(256, activation='relu'))
cnn_model.add(BatchNormalization())
#final layer, is output, 1 out of 5 possible results
#0 Normal, 1 Tapered, 2 Pyriform, 3 Amorphous
cnn_model.add(Dense(4))
cnn_model.add(Activation('softmax'))
# -
# #### 3.3 Compile model
# +
cnn_model.compile(optimizer=keras.optimizers.Adam(learning_rate=1e-3),
loss='categorical_crossentropy',
metrics=['accuracy'])
# -
# #### 3.4 Fit model
# +
history = cnn_model.fit(cnn_training,
epochs=200,
validation_data=cnn_val,
verbose = 1,
callbacks = [mc, reduce_lr, es,cb])
# -
# #### 3.5 Evaluate model
# +
_, train_acc = cnn_model.evaluate(cnn_training, verbose=1)
_, val_acc = cnn_model.evaluate(cnn_val, verbose=1)
print('Train: %.3f, Val: %.3f' % (train_acc, val_acc))
# plot training history
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='val')
plt.legend()
plt.show()
# -
# #### 3.6 Make Predictions
# +
#get predicted probality
prediction = cnn_model.predict(cnn_test,verbose=1)
#Get class of prediction
predicted_class = np.argmax(prediction,axis=1)
#get trueclass
true_classes = cnn_test.classes
# get names of pictures
filenames = cnn_test.filenames
#store info in dataframe
df_predictions = pd.DataFrame({'Filename': filenames, 'Label': true_classes, 'Test': predicted_class})
#store info in dataframe
cnn_model_predictions = pd.DataFrame({'Filename': filenames,'cnn_model': predicted_class})
# -
# #### 3.7 Confusion matrix and classification report
# +
class_labels = list(cnn_test.class_indices.keys())
print(confusion_matrix(true_classes, predicted_class))
report = classification_report(true_classes, predicted_class, target_names=class_labels)
print(report)
# -
# <a id='2'></a>
#
# # 4. ALEX NET
#
checkpoint_filepath = path+'/Models/AlexNet.h5'
# #### 4.1 Preprocess input
# Online Data Augmentation in RGB
# +
Alex_train_datagen = ImageDataGenerator(rescale = 1./255,
rotation_range = 5,
width_shift_range = 0.1,
height_shift_range = 0.1,
vertical_flip = True,
horizontal_flip = True,
brightness_range=[0.2,1.5],
fill_mode='nearest',
zoom_range = 0.2,
)
Alex_val_datagen = ImageDataGenerator(rescale = 1./255)
Alex_test_datagen = ImageDataGenerator(rescale = 1./255)
#test different color maps - class modes and cross validation types
Alex_training = Alex_train_datagen.flow_from_directory(path+'/path/train',
target_size = (32, 32),
batch_size = 64,
shuffle = True,
class_mode="categorical",
color_mode = 'rgb')
Alex_val = Alex_val_datagen.flow_from_directory(path+'/path/val',
target_size = (32, 32),
batch_size = 64,
shuffle = True,
class_mode="categorical",
color_mode = 'rgb')
Alex_test = Alex_test_datagen.flow_from_directory(path+'/path/test',
target_size = (32, 32),
batch_size = 1,
shuffle = True,
class_mode="categorical",
color_mode = 'rgb')
# -
# #### 4.2 Build model
# +
#Instantiation
AlexNet = Sequential()
#1st Convolutional Layer
AlexNet.add(Conv2D(filters=96, input_shape=(32,32,3), kernel_size=(11,11), strides=(4,4), padding='same'))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
AlexNet.add(MaxPooling2D(pool_size=(2,2), strides=(2,2), padding='same'))
#2nd Convolutional Layer
AlexNet.add(Conv2D(filters=256, kernel_size=(5, 5), strides=(1,1), padding='same'))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
AlexNet.add(MaxPooling2D(pool_size=(2,2), strides=(2,2), padding='same'))
#3rd Convolutional Layer
AlexNet.add(Conv2D(filters=384, kernel_size=(3,3), strides=(1,1), padding='same'))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
#4th Convolutional Layer
AlexNet.add(Conv2D(filters=384, kernel_size=(3,3), strides=(1,1), padding='same'))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
#5th Convolutional Layer
AlexNet.add(Conv2D(filters=256, kernel_size=(3,3), strides=(1,1), padding='same'))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
AlexNet.add(MaxPooling2D(pool_size=(2,2), strides=(2,2), padding='same'))
#Passing it to a Fully Connected layer
AlexNet.add(Flatten())
# 1st Fully Connected Layer
AlexNet.add(Dense(4096, input_shape=(32,32,3,)))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
# Add Dropout to prevent overfitting
AlexNet.add(Dropout(0.4))
#2nd Fully Connected Layer
AlexNet.add(Dense(4096))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
#Add Dropout
AlexNet.add(Dropout(0.4))
#3rd Fully Connected Layer
AlexNet.add(Dense(1000))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('relu'))
#Add Dropout
AlexNet.add(Dropout(0.4))
#Output Layer
AlexNet.add(Dense(4))
AlexNet.add(BatchNormalization())
AlexNet.add(Activation('softmax'))
#Model Summary
AlexNet.summary()
# -
# #### 4.3 Compile model
AlexNet.compile(loss = keras.losses.categorical_crossentropy, optimizer= 'adam', metrics=['accuracy'])
# #### 4.4 Fit model
history = AlexNet.fit(Alex_training,
epochs=200,
validation_data=Alex_val,
verbose = 1,
callbacks = [mc, reduce_lr, es,cb])
# #### 4.5 Evaluate model
# +
_, train_acc = AlexNet.evaluate(Alex_training, verbose=1)
_, val_acc = AlexNet.evaluate(Alex_val, verbose=1)
print('Train: %.3f, Val: %.3f' % (train_acc, val_acc))
# plot training history
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='val')
plt.legend()
plt.show()
# +
test_loss, test_acc = AlexNet.evaluate(Alex_test)
print('Test loss:', test_loss)
print('Test accuracy:', test_acc)
print("\nTime:",sum(cb.logs)/60,"min")
# -
# #### 4.6 Make Predictions
# +
#get predicted probality
prediction = AlexNet.predict(Alex_test,verbose=1)
#Get class of prediction
predicted_class = np.argmax(prediction,axis=1)
#get trueclass
true_classes = Alex_test.classes
# get names of pictures
filenames = Alex_test.filenames
#store info in dataframe
alex_predictions = pd.DataFrame({'Filename': filenames,'AlexNet': predicted_class})
# +
#join prediction to dataframe
df_predictions = df_predictions.merge(alex_predictions, on = 'Filename', how = 'inner')
del alex_predictions
# -
df_predictions
# #### 4.7 Confusion matrix and classification report
# +
class_labels = list(Alex_test.class_indices.keys())
print(confusion_matrix(true_classes, predicted_class))
report = classification_report(true_classes, predicted_class, target_names=class_labels)
print(report)
# -
# <a id='3'></a>
# # 5. RESNET50
#
#
checkpoint_filepath = path+'/Models/ResNet50.h5'
# #### 5.1 Preprocess input
# resnet50 requires its own preprocessing
# +
#to play around with these
res_train_datagen = ImageDataGenerator(rescale = 1./255,
preprocessing_function=preprocess_input,
rotation_range = 5,
width_shift_range = 0.06,
height_shift_range = 0.06,
vertical_flip = True,
horizontal_flip = True,
brightness_range=[0.2,1.2],
fill_mode='nearest',
zoom_range = 0.2,
)
res_val_datagen = ImageDataGenerator(rescale = 1./255, preprocessing_function=preprocess_input)
res_test_datagen = ImageDataGenerator(rescale = 1./255, preprocessing_function=preprocess_input)
#test different color maps - class modes and cross validation types
res_train = res_train_datagen.flow_from_directory(path+'/path/train',
target_size = (32, 32),
batch_size = 64,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
res_val = res_val_datagen.flow_from_directory(path+'/path/val',
target_size = (32, 32),
batch_size = 64,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
res_test = res_test_datagen.flow_from_directory(path+'/path/test',
target_size = (32, 32),
batch_size = 1,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
# -
# #### 5.2 Build model
# +
res50model = Sequential()
#Layer 1: RES50 without top layer
res50model.add(ResNet50(weights='imagenet', input_shape= (32,32,3),
include_top = False, classes=5,))
#Passing it to a Fully Connected layer
res50model.add(Flatten())
# 1st Fully Connected Layer
res50model.add(Dense(4096, input_shape=(32,32,3,)))
res50model.add(BatchNormalization())
res50model.add(Activation('relu'))
# Add Dropout to prevent overfitting
res50model.add(Dropout(0.4))
#2nd Fully Connected Layer
res50model.add(Dense(4096))
res50model.add(BatchNormalization())
res50model.add(Activation('relu'))
#Add Dropout
res50model.add(Dropout(0.4))
res50model.add(Dense(4, activation = 'softmax'))
#freeze layers in resnet - weights obtained with IMAGENET challenge, we only train final layer
res50model.layers[0].trainable = False
res50model.summary()
# -
# #### 5.3 Compile the model
res50model.compile(loss = keras.losses.categorical_crossentropy, optimizer= 'adam', metrics=['accuracy'])
# #### 5.4 Fit model
history = res50model.fit(res_train,
epochs=200,
validation_data=res_val,
verbose = 1,
callbacks = [mc, reduce_lr, es,cb])
# #### 5.5 Evaluate the model
# +
_, train_acc = res50model.evaluate(res_train, verbose=1)
_, val_acc = res50model.evaluate(res_val, verbose=1)
print('Train: %.3f, Val: %.3f' % (train_acc, val_acc))
# plot training history
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='val')
plt.legend()
plt.show()
test_loss, test_acc = res50model.evaluate(res_test)
print('Test loss:', test_loss)
print('Test accuracy:', test_acc)
print("\nTime:",sum(cb.logs)/60,"min")
# -
# #### 5.6 Make Predictions
# +
#get predicted probality
prediction = res50model.predict(res_test,verbose=1)
#Get class of prediction
predicted_class = np.argmax(prediction,axis=1)
#get trueclass
true_classes = res_test.classes
# get names of pictures
filenames = res_test.filenames
#store info in dataframe
df_predictions = pd.DataFrame({'Filename': filenames, 'Label': true_classes, 'Test': predicted_class})
#store info in dataframe
res50_prebuilt_predictions = pd.DataFrame({'Filename': filenames,'ResNet50': predicted_class})
# -
# #### 5.7 Confusion matrix and classification report
# +
class_labels = list(res_test.class_indices.keys())
print(confusion_matrix(true_classes, predicted_class))
report = classification_report(true_classes, predicted_class, target_names=class_labels)
print(report)
# -
# <a id='4'></a>
#
# # 6. EFFICIENT NET B7
#
#
#
checkpoint_filepath = path+'/Models/EffnetB7.h5'
# #### 6.1 Preprocess input
#
# +
#to play around with these
EffNetB7train_datagen = ImageDataGenerator(rescale = 1./255,
rotation_range = 5,
width_shift_range = 0.06,
height_shift_range = 0.06,
vertical_flip = True,
horizontal_flip = True,
brightness_range=[0.2,1.2],
fill_mode='nearest',
zoom_range = 0.2,
)
EffNetB7val_datagen = ImageDataGenerator(rescale = 1./255, preprocessing_function=preprocess_input)
EffNetB7test_datagen = ImageDataGenerator(rescale = 1./255, preprocessing_function=preprocess_input)
#test different color maps - class modes and cross validation types
EffNetB7_train = EffNetB7train_datagen.flow_from_directory(path+'/path/train',
target_size = (32, 32),
batch_size = 64,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
EffNetB7_val = EffNetB7val_datagen.flow_from_directory(path+'/path/val',
target_size = (32, 32),
batch_size = 64,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
EffNetB7_test = EffNetB7test_datagen.flow_from_directory(path+'/path/test',
target_size = (32, 32),
batch_size = 1,
class_mode="categorical",
color_mode = 'rgb',
shuffle = True)
# -
# #### 6.1Build model
# +
dropout_rate = 0.2
EffNetB7model = Sequential()
#Layer 1
EffNetB7model.add(EfficientNetB7(weights='imagenet', input_shape= (32,32,3),
include_top = False, classes=4))
EffNetB7model.add(layers.GlobalMaxPooling2D(name="gap")) ### mais eficiente qie o flatten
#EffNetB7.add(Flatten())
if dropout_rate > 0:
EffNetB7model.add(Dropout(dropout_rate, name="dropout_out"))
EffNetB7model.add(Dense(4, activation="softmax"))
EffNetB7model.layers[0].trainable = False
# -
EffNetB7model.summary()
# #### 6.2 Compile the model
EffNetB7model.compile(loss = keras.losses.categorical_crossentropy, optimizer= 'adam', metrics=['accuracy'])
# #### 6.3 Fit model
EffNet_history = EffNetB7model.fit(EffNetB7_train,
epochs=200,
validation_data=EffNetB7_val,
verbose = 1,
validation_steps = len(EffNetB7_val),
callbacks = [reduce_lr,cb,es])
# #### 6.4 Evaluate the model
# +
# evaluate the model
_, train_acc = EffNetB7model.evaluate(EffNetB7_train, verbose=1)
_, val_acc = EffNetB7model.evaluate(EffNetB7_val, verbose=1)
print('Train: %.3f, Val: %.3f' % (train_acc, val_acc))
# plot training history
plt.plot(EffNet_history.history['loss'], label='train')
plt.plot(EffNet_history.history['val_loss'], label='val')
plt.legend()
plt.show()
# +
test_loss, test_acc = EffNetB7model.evaluate(EffNetB7_test)
print('Test loss:', test_loss)
print('Test accuracy:', test_acc)
print("\nTime:",sum(cb.logs)/60,"min")
# -
# #### 6.5 Make Predictions
# +
#get predicted probality
prediction = EffNetB7model.predict(EffNetB7_test,verbose=1)
#Get class of prediction
predicted_class = np.argmax(prediction,axis=1)
#get trueclass
true_classes = EffNetB7_test.classes
# get names of pictures
filenames = EffNetB7_test.filenames
#store info in dataframe
EffNetB7_predictions = pd.DataFrame({'Filename': filenames,'EffNetB7': predicted_class})
# +
#join prediction to dataframe
df_predictions = df_predictions.merge(EffNetB7_predictions, on = 'Filename', how = 'inner')
del EffNetB7_predictions
# -
df_predictions
# #### 6.6 Confusion matrix and classification report
# +
class_labels = list(EffNetB7_test.class_indices.keys())
print(confusion_matrix(true_classes, predicted_class))
report = classification_report(true_classes, predicted_class, target_names=class_labels)
print(report)
# -
| .ipynb_checkpoints/CNN_SpermCells-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# <div class="alert alert-warning">
#
# <b>Disclaimer:</b>
#
# The main objective of the <i>Jupyter</i> notebooks is to show how to use the models of the <i>QENS library</i> by
#
# - building a fitting model: composition of models, convolution with a resolution function
# - setting and running the fit
# - extracting and displaying information about the results
#
# These steps have a minimizer-dependent syntax. That's one of the reasons why different minimizers have been used in the notebooks provided as examples.
# But, the initial guessed parameters might not be optimal, resulting in a poor fit of the reference data.
#
# </div>
#
# # Two Lorentzian ∗ resolution with lmfit
#
# ## Table of Contents
#
# - [Introduction](#Introduction)
# - [Importing the required libraries](#Importing-the-required-libraries)
# - [Importing the reference data](#Importing-the-reference-data)
# - [Setting and fitting](#Setting-and-fitting)
# - [Plotting the results](#Plotting-the-results)
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Introduction
#
# <div class="alert alert-info">
#
# The objective of this notebook is to show how to use one of the models of
# the <a href="https://github.com/QENSlibrary/QENSmodels">QENSlibrary</a>, <b>Lorentzian</b>, to perform some fits.
# <a href="https://lmfit.github.io/lmfit-py/">lmfit</a> is used for fitting.
# </div>
#
# The following example uses the data from IRIS:
# - workspace_index=0, file: `irs26176_graphite002_red.nxs`
# - related instrument resolution data `irs26173_graphite002_res.nxs`
#
# The ISIS sample datasets can be downloaded from [Mantid's website](http://download.mantidproject.org/).
# The data used for this example are in the sample datafile: `data_2lorentzians.dat` and the instrument resolution datafile `irf_iris.dat`, respectively.
#
# This example is based on a [Mantid "Fitting QENS Peaks" tutorial](https://www.mantidproject.org/Fitting_QENS_Peaks).
#
# The implementation with `lmfit` is based on https://lmfit.github.io/lmfit-py/model.html
#
# This example requires an additional Python module `scipy.interpolate` to interpolate the tabulated data of the instrument resolution.
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Importing the required librairies
# + tags=["import_cell"]
# import python modules for plotting, fitting
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
# + tags=["remove_cell"]
import ipywidgets
# + tags=["remove_cell"]
# install QENSmodels (if not already installed)
import pkgutil
import sys
if not pkgutil.find_loader("QENSmodels"):
buttonY = ipywidgets.Button(description='Yes', button_style='success')
buttonN = ipywidgets.Button(description='No', button_style='danger')
choice_installation = ipywidgets.VBox(
[ipywidgets.Label("Do you want to install the QENSmodels' library?"), ipywidgets.HBox([buttonY, buttonN])],
layout=ipywidgets.Layout(width='50%', height='80px'))
display(choice_installation)
def on_buttonY_clicked(b):
# !{sys.executable} -m pip install git+https://github.com/QENSlibrary/QENSmodels#egg=QENSmodels
def on_buttonN_clicked(b):
print("You will not be able to run some of the remaining parts of this notebook")
buttonY.on_click(on_buttonY_clicked)
buttonN.on_click(on_buttonN_clicked)
# + tags=["remove_cell"]
# install lmfit (if not already installed)
if not pkgutil.find_loader("lmfit"):
lmfitY = ipywidgets.Button(description='Yes', button_style='success')
lmfitN = ipywidgets.Button(description='No', button_style='danger')
choice_installation = ipywidgets.VBox(
[ipywidgets.Label("Do you want to install lmfit?"), ipywidgets.HBox([lmfitY, lmfitN])],
layout=ipywidgets.Layout(width='30%', height='80px'))
display(choice_installation)
def on_lmfitY_clicked(b):
# !{sys.executable} -m pip install lmfit
def on_lmfitN_clicked(b):
print("You will not be able to run some of the remaining parts of this notebook")
lmfitY.on_click(on_lmfitY_clicked)
lmfitN.on_click(on_lmfitN_clicked)
# +
from lmfit import Model, CompositeModel
from scipy.interpolate import interp1d
# import model from QENS library
import QENSmodels
# + [markdown] tags=["remove_cell"]
# ### Physical units
# For information about unit conversion, please refer to the jupyter notebook called `Convert_units.ipynb` in the `tools` folder.
#
# The dictionary of units defined in the cell below specify the units of the refined parameters adapted to the convention used in the experimental datafile.
# -
# Units of parameters for selected QENS model and experimental data
dict_physical_units = {'omega': "meV",
'q': "1/Angstrom",
'hwhm': "meV",
'scale': "unit_of_signal.meV",
'center': "meV"}
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Importing the reference data
# -
path_to_data = './data/'
# Create model - 2 lorentzians convoluted with instrument resolution - 6 parameters
# Step 1 load instrument resolution data
irf_iris = np.loadtxt(path_to_data + 'irf_iris.dat')
x_irf = irf_iris[:, 0]
y_irf = irf_iris[:, 1]
# +
# Step 2: create function for instrument resolution data
# (cubic interpolation between tabulated data points)
f = interp1d(x_irf, y_irf, kind='cubic', bounds_error=False, fill_value='extrapolate')
def irf_gate(x):
""" Function defined from the interpolation of instrument resolution data
Used to define fitting model and plot """
return f(x)
# plot tabulated data and interpolated data
xx = np.linspace(-.25, .25, 500)
fig0, ax0 = plt.subplots()
ax0.plot(x_irf, y_irf, 'b.', label='tabulated data')
ax0.plot(xx, irf_gate(xx), 'g--', label='extrapolated data')
ax0.legend()
ax0.set_xlabel('Energy transfer (meV)')
ax0.set_title('Instrument resolution: plot tabulated data and interpolated data')
ax0.grid();
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Setting and fitting
# -
# Step 3: create "double lorentzian" profile
def model_2lorentzians(x, scale1, center1, hwhm1, scale2, center2, hwhm2):
model = QENSmodels.lorentzian(x, scale1, center1, hwhm1)
model += QENSmodels.lorentzian(x, scale2, center2, hwhm2)
return model
# +
# Step 4: create convolution function
# code from https://lmfit.github.io/lmfit-py/model.html
def convolve(arr, kernel):
# simple convolution of two arrays
npts = min(len(arr), len(kernel))
pad = np.ones(npts)
tmp = np.concatenate((pad * arr[0], arr, pad * arr[-1]))
out = np.convolve(tmp, kernel, mode='valid')
noff = int((len(out) - npts) / 2)
return out[noff:noff + npts]
# +
# Create model for the fit
gmodel = CompositeModel(Model(irf_gate), Model(model_2lorentzians), convolve)
print('Names of parameters:', gmodel.param_names)
print('Independent variable(s):', gmodel.independent_vars)
# -
# Load reference data - extract x and y values
two_lorentzians_iris = np.loadtxt(path_to_data + 'data_2lorentzians.dat')
xx = two_lorentzians_iris[:, 0]
yy = two_lorentzians_iris[:, 1]
# +
# Fit
result = gmodel.fit(yy, x=xx, scale1=1., center1=0., hwhm1=0.25, scale2=1., center2=1., hwhm2=0.25)
fig1, ax1 = plt.subplots()
ax1.plot(xx, yy, '+', label='experimental data')
ax1.plot(xx, result.init_fit, 'k--', label='model with initial guesses')
ax1.legend()
ax1.set(xlabel='Energy transfer (meV)',
title='Plot before fitting: experimental data and mode with initial guesses')
ax1.grid();
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Plotting results
# +
# display result
print('Result of fit:\n', result.fit_report())
# plot selected results: experimental data and best fit
fig2, ax2 = plt.subplots()
ax2.plot(xx, yy, '+', label='experimental data')
ax2.plot(xx, result.best_fit, 'r-', label='best fit')
ax2.grid()
ax2.set(xlabel='Energy transfer (meV)',
title='Plot selected fitting results: experimental data and best fit')
ax2.legend();
# -
# other option to plot result using lmfit's features
result.plot()
for item in result.params.keys():
print(item[:-1],
result.params[item].value,
'+/-',
result.params[item].stderr,
dict_physical_units[item[:-1]])
| docs/examples/lmfit_two_lorentzian_fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import os
import sys
import logging
import librosa
import sys
# import tensorflow as tf
import plotly.graph_objects as go
import plotly.express as px
import re
sns.set_theme()
# +
module_path =os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path+"/scripts")
if module_path not in sys.path:
sys.path.append(module_path+"/model")
logging.basicConfig(filename='../logs/notebook.log', filemode='w', format='%(asctime)s - %(name)s - %(levelname)s - %(message)s',level=logging.INFO)
# -
#we import our module script that we shall be using for preprocessing
logging.info(" ===== Loading created scripts ===== ")
try:
import audio
logging.info(" ==== Succefully Loaded the Modules ===== ")
print(" ===== Succefully loaded modules, modules are now ready to use ===== ")
except ImportError as error:
logging.info(" !!!! Import Error !!!! ")
print(" !!!! The system ran into an Import Error \n ")
print(error.__class__ )
sys.exit(1)
data=pd.read_csv('../data/duration.csv')
data
#we read the data
try:
print("===== Reading Dataset ===== \n ")
logging.info("====== Redaing Dataset ====== ")
data=pd.read_csv('../data/merged_data.csv')
print(" ==== Successfuly read dataset ===== \n")
except FileNotFoundError as e:
logging.info(" !!!! File Not Found Error !!!! ")
print(" !!!! The system ran into a File Not Found Erro !!! \n ")
print(e.__class__.__name__ )
sys.exit(1)
# + tags=[]
#we create a function to add the root folder to the location of each wav file to make a complete path
location_directory='../data/alldatas'
def add_root(x):
x=os.path.join(location_directory,x)
return x
data['Location']=data['wav_file'].apply(lambda x:add_root(x) )
data
# -
#call the plot wav function from the audio script to plot a sample wav file
audio.plot_wav(data['Location'][1],44100)
#we begin by making all text lower case
data['text']=data['text'].apply(lambda x: x.lower())
# +
#we remove any punctuation marks that might hinder our model
import string
data['text']=data['text'].apply(lambda x: re.sub('[%s]' % re.escape(string.punctuation), '', x))
# -
data
#remove any extra padding and whitespaces
data['text']=data['text'].apply(lambda x: re.sub(' +',' ',x))
# +
# def char_map(text):
# charlist=[]
# for chars in text:
# charlist.append(ord(chars))
# return charlist
# data['Textmap']=data['text'].apply(lambda x: char_map(x))
# +
"""
Defines two dictionaries for converting
between text and integer sequences.
"""
char_map_str = """
' 0
<SPACE> 1
a 2
b 3
c 4
d 5
e 6
f 7
g 8
h 9
i 10
j 11
k 12
l 13
m 14
n 15
o 16
p 17
q 18
r 19
s 20
t 21
u 22
v 23
w 24
x 25
y 26
z 27
N 28
U 29
K 30
1 31
2 32
3 33
4 34
5 35
6 36
7 37
8 38
9 39
0 40
< 41
> 42
, 43
. 45
? 46
- 47
_ 48
"""
# the "blank" character is mapped to 28
char_map = {}
index_map = {}
for line in char_map_str.strip().split('\n'):
ch, index = line.split()
char_map[ch] = int(index)
index_map[int(index)+1] = ch
index_map[2] = ' '
# -
char_map
newdata=data[['Location','text','duration']]
newdata.columns=['key','text','duration']
# +
train_data=newdata.iloc[:8000,:]
valid_data=newdata.iloc[8001:,:]
print("Shape of new dataframes - {} , {}".format(train_data.shape, valid_data.shape))
# -
train_data.to_json('../data/train_corpus.json', orient='records', lines=True)
valid_data.to_json('../data/valid_corpus.json', orient='records', lines=True)
# +
import soundfile
from numpy.lib.stride_tricks import as_strided
def feat_dimen(window,max_freq):
return int(0.001 * window * max_freq)+1
# +
"""
Defines various functions for processing the data.
"""
import numpy as np
import soundfile
from numpy.lib.stride_tricks import as_strided
def calc_feat_dim(window, max_freq):
return int(0.001 * window * max_freq) + 1
def conv_output_length(input_length, filter_size, border_mode, stride,
dilation=1):
""" Compute the length of the output sequence after 1D convolution along
time. Note that this function is in line with the function used in
Convolution1D class from Keras.
Params:
input_length (int): Length of the input sequence.
filter_size (int): Width of the convolution kernel.
border_mode (str): Only support `same` or `valid`.
stride (int): Stride size used in 1D convolution.
dilation (int)
"""
if input_length is None:
return None
assert border_mode in {'same', 'valid'}
dilated_filter_size = filter_size + (filter_size - 1) * (dilation - 1)
if border_mode == 'same':
output_length = input_length
elif border_mode == 'valid':
output_length = input_length - dilated_filter_size + 1
return (output_length + stride - 1) // stride
def spectrogram(samples, fft_length=256, sample_rate=2, hop_length=128):
"""
Compute the spectrogram for a real signal.
The parameters follow the naming convention of
matplotlib.mlab.specgram
Args:
samples (1D array): input audio signal
fft_length (int): number of elements in fft window
sample_rate (scalar): sample rate
hop_length (int): hop length (relative offset between neighboring
fft windows).
Returns:
x (2D array): spectrogram [frequency x time]
freq (1D array): frequency of each row in x
Note:
This is a truncating computation e.g. if fft_length=10,
hop_length=5 and the signal has 23 elements, then the
last 3 elements will be truncated.
"""
assert not np.iscomplexobj(samples), "Must not pass in complex numbers"
window = np.hanning(fft_length)[:, None]
window_norm = np.sum(window**2)
# The scaling below follows the convention of
# matplotlib.mlab.specgram which is the same as
# matlabs specgram.
scale = window_norm * sample_rate
trunc = (len(samples) - fft_length) % hop_length
x = samples[:len(samples) - trunc]
# "stride trick" reshape to include overlap
nshape = (fft_length, (len(x) - fft_length) // hop_length + 1)
nstrides = (x.strides[0], x.strides[0] * hop_length)
x = as_strided(x, shape=nshape, strides=nstrides)
# window stride sanity check
assert np.all(x[:, 1] == samples[hop_length:(hop_length + fft_length)])
# broadcast window, compute fft over columns and square mod
x = np.fft.rfft(x * window, axis=0)
x = np.absolute(x)**2
# scale, 2.0 for everything except dc and fft_length/2
x[1:-1, :] *= (2.0 / scale)
x[(0, -1), :] /= scale
freqs = float(sample_rate) / fft_length * np.arange(x.shape[0])
return x, freqs
def spectrogram_from_file(filename, step=10, window=20, max_freq=None,
eps=1e-14):
""" Calculate the log of linear spectrogram from FFT energy
Params:
filename (str): Path to the audio file
step (int): Step size in milliseconds between windows
window (int): FFT window size in milliseconds
max_freq (int): Only FFT bins corresponding to frequencies between
[0, max_freq] are returned
eps (float): Small value to ensure numerical stability (for ln(x))
"""
with soundfile.SoundFile(filename) as sound_file:
audio = sound_file.read(dtype='float32')
sample_rate = sound_file.samplerate
if audio.ndim >= 2:
audio = np.mean(audio, 1)
if max_freq is None:
max_freq = sample_rate / 2
if max_freq > sample_rate / 2:
raise ValueError("max_freq must not be greater than half of "
" sample rate")
if step > window:
raise ValueError("step size must not be greater than window size")
hop_length = int(0.001 * step * sample_rate)
fft_length = int(0.001 * window * sample_rate)
pxx, freqs = spectrogram(
audio, fft_length=fft_length, sample_rate=sample_rate,
hop_length=hop_length)
ind = np.where(freqs <= max_freq)[0][-1] + 1
return np.transpose(np.log(pxx[:ind, :] + eps))
def text_to_int_sequence(text):
""" Convert text to an integer sequence """
int_sequence = []
for c in text:
if c == ' ':
ch = char_map['<SPACE>']
else:
ch = char_map[c]
int_sequence.append(ch)
return int_sequence
def int_sequence_to_text(int_sequence):
""" Convert an integer sequence to text """
text = []
for c in int_sequence:
ch = index_map[c]
text.append(ch)
return text
# +
import json
import numpy as np
import random
from python_speech_features import mfcc
import librosa
import scipy.io.wavfile as wav
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
RNG_SEED = 123
class AudioGenerator():
def __init__(self, step=10, window=20, max_freq=8000, mfcc_dim=13,
minibatch_size=4000, desc_file=None, spectrogram=True, max_duration=10.0,
sort_by_duration=False):
"""
Params:
step (int): Step size in milliseconds between windows (for spectrogram ONLY)
window (int): FFT window size in milliseconds (for spectrogram ONLY)
max_freq (int): Only FFT bins corresponding to frequencies between
[0, max_freq] are returned (for spectrogram ONLY)
desc_file (str, optional): Path to a JSON-line file that contains
labels and paths to the audio files. If this is None, then
load metadata right away
"""
self.feat_dim = calc_feat_dim(window, max_freq)
self.mfcc_dim = mfcc_dim
self.feats_mean = np.zeros((self.feat_dim,))
self.feats_std = np.ones((self.feat_dim,))
self.rng = random.Random(RNG_SEED)
if desc_file is not None:
self.load_metadata_from_desc_file(desc_file)
self.step = step
self.window = window
self.max_freq = max_freq
self.cur_train_index = 0
self.cur_valid_index = 0
self.cur_test_index = 0
self.max_duration=max_duration
self.minibatch_size = minibatch_size
self.spectrogram = spectrogram
self.sort_by_duration = sort_by_duration
def get_batch(self, partition):
""" Obtain a batch of train, validation, or test data
"""
if partition == 'train':
audio_paths = self.train_audio_paths
cur_index = self.cur_train_index
texts = self.train_texts
elif partition == 'valid':
audio_paths = self.valid_audio_paths
cur_index = self.cur_valid_index
texts = self.valid_texts
elif partition == 'test':
audio_paths = self.test_audio_paths
cur_index = self.test_valid_index
texts = self.test_texts
else:
raise Exception("Invalid partition. "
"Must be train/validation")
features = [self.normalize(self.featurize(a)) for a in
audio_paths[cur_index:cur_index+self.minibatch_size]]
# calculate necessary sizes
max_length = max([features[i].shape[0]
for i in range(0, self.minibatch_size)])
max_string_length = max([len(texts[cur_index+i])
for i in range(0, self.minibatch_size)])
# initialize the arrays
X_data = np.zeros([self.minibatch_size, max_length,
self.feat_dim*self.spectrogram + self.mfcc_dim*(not self.spectrogram)])
labels = np.ones([self.minibatch_size, max_string_length]) * 28
input_length = np.zeros([self.minibatch_size, 1])
label_length = np.zeros([self.minibatch_size, 1])
for i in range(0, self.minibatch_size):
# calculate X_data & input_length
feat = features[i]
input_length[i] = feat.shape[0]
X_data[i, :feat.shape[0], :] = feat
# calculate labels & label_length
label = np.array(text_to_int_sequence(texts[cur_index+i]))
labels[i, :len(label)] = label
label_length[i] = len(label)
# return the arrays
outputs = {'ctc': np.zeros([self.minibatch_size])}
inputs = {'the_input': X_data,
'the_labels': labels,
'input_length': input_length,
'label_length': label_length
}
return (inputs, outputs)
def shuffle_data_by_partition(self, partition):
""" Shuffle the training or validation data
"""
if partition == 'train':
self.train_audio_paths, self.train_durations, self.train_texts = shuffle_data(
self.train_audio_paths, self.train_durations, self.train_texts)
elif partition == 'valid':
self.valid_audio_paths, self.valid_durations, self.valid_texts = shuffle_data(
self.valid_audio_paths, self.valid_durations, self.valid_texts)
else:
raise Exception("Invalid partition. "
"Must be train/validation")
def sort_data_by_duration(self, partition):
""" Sort the training or validation sets by (increasing) duration
"""
if partition == 'train':
self.train_audio_paths, self.train_durations, self.train_texts = sort_data(
self.train_audio_paths, self.train_durations, self.train_texts)
elif partition == 'valid':
self.valid_audio_paths, self.valid_durations, self.valid_texts = sort_data(
self.valid_audio_paths, self.valid_durations, self.valid_texts)
else:
raise Exception("Invalid partition. "
"Must be train/validation")
def next_train(self):
""" Obtain a batch of training data
"""
while True:
ret = self.get_batch('train')
self.cur_train_index += self.minibatch_size
if self.cur_train_index >= len(self.train_texts) - self.minibatch_size:
self.cur_train_index = 0
self.shuffle_data_by_partition('train')
yield ret
def next_valid(self):
""" Obtain a batch of validation data
"""
while True:
ret = self.get_batch('valid')
self.cur_valid_index += self.minibatch_size
if self.cur_valid_index >= len(self.valid_texts) - self.minibatch_size:
self.cur_valid_index = 0
self.shuffle_data_by_partition('valid')
yield ret
def next_test(self):
""" Obtain a batch of test data
"""
while True:
ret = self.get_batch('test')
self.cur_test_index += self.minibatch_size
if self.cur_test_index >= len(self.test_texts) - self.minibatch_size:
self.cur_test_index = 0
yield ret
def load_train_data(self, desc_file='../data/trainprep_corpus.json'):
self.load_metadata_from_desc_file(desc_file, 'train')
self.fit_train()
if self.sort_by_duration:
self.sort_data_by_duration('train')
def load_validation_data(self, desc_file='../data/validprep_corpus.json'):
self.load_metadata_from_desc_file(desc_file, 'validation')
if self.sort_by_duration:
self.sort_data_by_duration('valid')
def load_test_data(self, desc_file='../data/testprep_corpus.json'):
self.load_metadata_from_desc_file(desc_file, 'test')
def load_metadata_from_desc_file(self, desc_file, partition):
""" Read metadata from a JSON-line file
(possibly takes long, depending on the filesize)
Params:
desc_file (str): Path to a JSON-line file that contains labels and
paths to the audio files
partition (str): One of 'train', 'validation' or 'test'
"""
audio_paths, durations, texts = [], [], []
with open(desc_file) as json_line_file:
for line_num, json_line in enumerate(json_line_file):
try:
spec = json.loads(json_line)
if float(spec['duration']) > self.max_duration:
continue
audio_paths.append(spec['key'])
durations.append(float(spec['duration']))
texts.append(spec['text'])
except Exception as e:
# Change to (KeyError, ValueError) or
# (KeyError,json.decoder.JSONDecodeError), depending on
# json module version
print('Error reading line #{}: {}'
.format(line_num, json_line))
if partition == 'train':
self.train_audio_paths = audio_paths
self.train_durations = durations
self.train_texts = texts
elif partition == 'validation':
self.valid_audio_paths = audio_paths
self.valid_durations = durations
self.valid_texts = texts
elif partition == 'test':
self.test_audio_paths = audio_paths
self.test_durations = durations
self.test_texts = texts
else:
raise Exception("Invalid partition to load metadata. "
"Must be train/validation/test")
def fit_train(self, k_samples=100):
""" Estimate the mean and std of the features from the training set
Params:
k_samples (int): Use this number of samples for estimation
"""
k_samples = min(k_samples, len(self.train_audio_paths))
samples = self.rng.sample(self.train_audio_paths, k_samples)
feats = [self.featurize(s) for s in samples]
feats = np.vstack(feats)
self.feats_mean = np.mean(feats, axis=0)
self.feats_std = np.std(feats, axis=0)
def featurize(self, audio_clip):
""" For a given audio clip, calculate the corresponding feature
Params:
audio_clip (str): Path to the audio clip
"""
if self.spectrogram:
return spectrogram_from_file(
audio_clip, step=self.step, window=self.window,
max_freq=self.max_freq)
else:
(rate, sig) = wav.read(audio_clip)
return mfcc(sig, rate, numcep=self.mfcc_dim)
def normalize(self, feature, eps=1e-14):
""" Center a feature using the mean and std
Params:
feature (numpy.ndarray): Feature to normalize
"""
return (feature - self.feats_mean) / (self.feats_std + eps)
def shuffle_data(audio_paths, durations, texts):
""" Shuffle the data (called after making a complete pass through
training or validation data during the training process)
Params:
audio_paths (list): Paths to audio clips
durations (list): Durations of utterances for each audio clip
texts (list): Sentences uttered in each audio clip
"""
p = np.random.permutation(len(audio_paths))
audio_paths = [audio_paths[i] for i in p]
durations = [durations[i] for i in p]
texts = [texts[i] for i in p]
return audio_paths, durations, texts
def sort_data(audio_paths, durations, texts):
""" Sort the data by duration
Params:
audio_paths (list): Paths to audio clips
durations (list): Durations of utterances for each audio clip
texts (list): Sentences uttered in each audio clip
"""
p = np.argsort(durations).tolist()
audio_paths = [audio_paths[i] for i in p]
durations = [durations[i] for i in p]
texts = [texts[i] for i in p]
return audio_paths, durations, texts
def vis_train_features(index=0):
""" Visualizing the data point in the training set at the supplied index
"""
# obtain spectrogram
audio_gen = AudioGenerator(spectrogram=True)
audio_gen.load_train_data()
vis_audio_path = audio_gen.train_audio_paths[index]
vis_spectrogram_feature = audio_gen.normalize(audio_gen.featurize(vis_audio_path))
# obtain mfcc
audio_gen = AudioGenerator(spectrogram=False)
audio_gen.load_train_data()
vis_mfcc_feature = audio_gen.normalize(audio_gen.featurize(vis_audio_path))
# obtain text label
vis_text = audio_gen.train_texts[index]
# obtain raw audio
vis_raw_audio, _ = librosa.load(vis_audio_path)
# print total number of training examples
print('There are %d total training examples.' % len(audio_gen.train_audio_paths))
# return labels for plotting
return vis_text, vis_raw_audio, vis_mfcc_feature, vis_spectrogram_feature, vis_audio_path
def plot_raw_audio(vis_raw_audio):
# plot the raw audio signal
fig = plt.figure(figsize=(12,3))
ax = fig.add_subplot(111)
steps = len(vis_raw_audio)
ax.plot(np.linspace(1, steps, steps), vis_raw_audio)
plt.title('Audio Signal')
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.show()
def plot_mfcc_feature(vis_mfcc_feature):
# plot the MFCC feature
fig = plt.figure(figsize=(12,5))
ax = fig.add_subplot(111)
im = ax.imshow(vis_mfcc_feature, cmap=plt.cm.jet, aspect='auto')
plt.title('Normalized MFCC')
plt.ylabel('Time')
plt.xlabel('MFCC Coefficient')
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
ax.set_xticks(np.arange(0, 13, 2), minor=False);
plt.show()
def plot_spectrogram_feature(vis_spectrogram_feature):
# plot the normalized spectrogram
fig = plt.figure(figsize=(12,5))
ax = fig.add_subplot(111)
im = ax.imshow(vis_spectrogram_feature, cmap=plt.cm.jet, aspect='auto')
plt.title('Normalized Spectrogram')
plt.ylabel('Time')
plt.xlabel('Frequency')
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
plt.colorbar(im, cax=cax)
plt.show()
# -
model=AudioGenerator()
vis_text, vis_raw_audio, vis_mfcc_feature, vis_spectrogram_feature, vis_audio_path = vis_train_features()
model.load_train_data()
model.load_validation_data()
model.fit_train()
a,b=model.get_batch('train')
gx = pd.DataFrame({'the_input': list(a['the_input']), 'the_labels': list(a['the_labels']), 'input_length': list(a['input_length']), 'label_length': list(a['label_length'])})
gx.to_csv('../data/prep_model.csv',index=False)
dx.to_csv('../data/procGen2.csv',index=False)
data.to_json('../data/train.json', orient='records', lines=True)
data.to_json('../data/train.json', orient='records', lines=True)
a,b=model.get_batch('valid')
dz = pd.DataFrame({'the_input': list(a['the_input']), 'the_labels': list(a['the_labels']), 'input_length': list(a['input_length']), 'label_length': list(a['label_length'])})
# !pip3 install --user --upgrade tensorflow
#from keras.backend.tensorflow_backend import set_session
#from tensorflow.keras.backend.tensorflow_backend import set_session
from tensorflow.compat.v1 import keras
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
tf.compat.v1.disable_eager_execution()
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.5
tf.keras.backend.set_session(tf.Session(config=config))
# +
from keras import backend as K
from keras.models import Model
from keras.layers import (BatchNormalization, Conv1D, Dense, Input,
TimeDistributed, Activation, Bidirectional, SimpleRNN, GRU, LSTM)
def simple_rnn_model(input_dim, output_dim=50):
""" Build a recurrent network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# Add recurrent layer
simp_rnn = GRU(output_dim, return_sequences=True,
implementation=2, name='rnn')(input_data)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(simp_rnn)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: x
print(model.summary())
return model
def rnn_model(input_dim, units, activation, output_dim=50):
""" Build a recurrent network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# Add recurrent layer
simp_rnn = GRU(units, activation=activation,
return_sequences=True, implementation=2, name='rnn')(input_data)
# TODO: Add batch normalization
bn_rnn = BatchNormalization()(simp_rnn)
# TODO: Add a TimeDistributed(Dense(output_dim)) layer
time_dense = TimeDistributed(Dense(output_dim))(bn_rnn)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(time_dense)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: x
print(model.summary())
return model
def cnn_rnn_model(input_dim, filters, kernel_size, conv_stride,
conv_border_mode, units, output_dim=50):
""" Build a recurrent + convolutional network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# Add convolutional layer
conv_1d = Conv1D(filters, kernel_size,
strides=conv_stride,
padding=conv_border_mode,
activation='relu',
name='conv1d')(input_data)
# Add batch normalization
bn_cnn = BatchNormalization(name='bn_conv_1d')(conv_1d)
# Add a recurrent layer
simp_rnn = SimpleRNN(units, activation='relu',
return_sequences=True, implementation=2, name='rnn')(bn_cnn)
# TODO: Add batch normalization
bn_rnn = BatchNormalization(name='bn_norm')(simp_rnn)
# TODO: Add a TimeDistributed(Dense(output_dim)) layer
time_dense = TimeDistributed(Dense(output_dim))(bn_rnn)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(time_dense)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: cnn_output_length(
x, kernel_size, conv_border_mode, conv_stride)
print(model.summary())
return model
def cnn_output_length(input_length, filter_size, border_mode, stride,
dilation=1):
""" Compute the length of the output sequence after 1D convolution along
time. Note that this function is in line with the function used in
Convolution1D class from Keras.
Params:
input_length (int): Length of the input sequence.
filter_size (int): Width of the convolution kernel.
border_mode (str): Only support `same` or `valid`.
stride (int): Stride size used in 1D convolution.
dilation (int)
"""
if input_length is None:
return None
assert border_mode in {'same', 'valid'}
dilated_filter_size = filter_size + (filter_size - 1) * (dilation - 1)
if border_mode == 'same':
output_length = input_length
elif border_mode == 'valid':
output_length = input_length - dilated_filter_size + 1
return (output_length + stride - 1) // stride
def deep_rnn_model(input_dim, units, recur_layers, output_dim=50):
""" Build a deep recurrent network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# TODO: Add recurrent layers, each with batch normalization
simp_rnn = SimpleRNN(units, activation='relu',
return_sequences=True, implementation=2, name='rnn')(input_data)
# TODO: Add batch normalization
bn_rnn = BatchNormalization(name='bn_norm')(simp_rnn)
## the loop should be here
for i in range(1, recur_layers):
print("+Layer", i)
#simp_rnn = LSTM(units, name='rnn' + str(i), return_sequences=True)(bn_rnn)
#simp_rnn = SimpleRNN(units, name='rnn' + str(i), return_sequences=True, activation='relu')(bn_rnn)
simp_rnn = GRU(units, name='rnn' + str(i), return_sequences=True, activation='relu')(bn_rnn)
# TODO: Add batch normalization
bn_rnn = BatchNormalization(name='bnorm' + str(i))(simp_rnn)
time_dense = TimeDistributed(Dense(output_dim))(bn_rnn)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(time_dense)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: x
print(model.summary())
return model
def bidirectional_rnn_model(input_dim, units, output_dim=50):
""" Build a bidirectional recurrent network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# TODO: Add bidirectional recurrent layer
bidir_rnn = Bidirectional(LSTM(units, return_sequences=True))(input_data)
# bidir_rnn = GRU(units, activation='relu',
# return_sequences=True, implementation=2, name='rnn')(input_data)
bn_rnn = BatchNormalization(name='bnorm')(bidir_rnn)
# TODO: Add a TimeDistributed(Dense(output_dim)) layer
time_dense = TimeDistributed(Dense(output_dim))(bn_rnn)
# Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(time_dense)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
model.output_length = lambda x: x
print(model.summary())
return model
def final_model(input_dim, units, output_dim=50):
""" Build a deep network for speech
"""
# Main acoustic input
input_data = Input(name='the_input', shape=(None, input_dim))
# TODO: Specify the layers in your network
bidir_rnn1 = Bidirectional(LSTM(units, return_sequences=True, dropout_W=0.5, dropout_U=0.5,
name='bidir_rnn1'))(input_data)
bn_rnn1 = BatchNormalization(name='bnorm1')(bidir_rnn1)
bidir_rnn2 = Bidirectional(LSTM(units, return_sequences=True,
name='bidir_rnn2'))(bn_rnn1)
bn_rnn2 = BatchNormalization(name='bnorm2')(bidir_rnn2)
time_dense = TimeDistributed(Dense(output_dim))(bn_rnn2)
# TODO: Add softmax activation layer
y_pred = Activation('softmax', name='softmax')(time_dense)
# Specify the model
model = Model(inputs=input_data, outputs=y_pred)
# TODO: Specify model.output_length
model.output_length = model.output_length = lambda x: x
print(model.summary())
return model
# -
model_1 = rnn_model(input_dim=161, # change to 13 if you would like to use MFCC features
units=200,
activation='relu')
# +
"""
Defines a functions for training a NN.
"""
import _pickle as pickle
from keras import backend as K
from keras.models import Model
from keras.layers import (Input, Lambda)
from tensorflow.keras.optimizers import SGD
from keras.callbacks import ModelCheckpoint
import os
def ctc_lambda_func(args):
y_pred, labels, input_length, label_length = args
return K.ctc_batch_cost(labels, y_pred, input_length, label_length)
def add_ctc_loss(input_to_softmax):
the_labels = Input(name='the_labels', shape=(None,), dtype='float32')
input_lengths = Input(name='input_length', shape=(1,), dtype='int64')
label_lengths = Input(name='label_length', shape=(1,), dtype='int64')
output_lengths = Lambda(input_to_softmax.output_length)(input_lengths)
# CTC loss is implemented in a lambda layer
loss_out = Lambda(ctc_lambda_func, output_shape=(1,), name='ctc')(
[input_to_softmax.output, the_labels, output_lengths, label_lengths])
model = Model(
inputs=[input_to_softmax.input, the_labels, input_lengths, label_lengths],
outputs=loss_out)
return model
def train_model(input_to_softmax,
pickle_path,
save_model_path,
train_json='../data/trainprep_corpus.json',
valid_json='../data/validprep_corpus.json',
minibatch_size=20,
spectrogram=True,
mfcc_dim=13,
optimizer='adam',
epochs=20,
verbose=1,
sort_by_duration=False,
max_duration=10.0):
# create a class instance for obtaining batches of data
audio_gen = AudioGenerator(minibatch_size=minibatch_size,
spectrogram=spectrogram, mfcc_dim=mfcc_dim, max_duration=max_duration,
sort_by_duration=sort_by_duration)
# add the training data to the generator
audio_gen.load_train_data(train_json)
audio_gen.load_validation_data(valid_json)
# calculate steps_per_epoch
num_train_examples=len(audio_gen.train_audio_paths)
steps_per_epoch = num_train_examples//minibatch_size
# calculate validation_steps
num_valid_samples = len(audio_gen.valid_audio_paths)
validation_steps = num_valid_samples//minibatch_size
# add CTC loss to the NN specified in input_to_softmax
model = add_ctc_loss(input_to_softmax)
# CTC loss is implemented elsewhere, so use a dummy lambda function for the loss
model.compile(loss={'ctc': lambda y_true, y_pred: y_pred}, optimizer=optimizer)
# make results/ directory, if necessary
if not os.path.exists('results'):
os.makedirs('results')
# add checkpointer
checkpointer = ModelCheckpoint(filepath='results/'+save_model_path, verbose=0)
# train the model
hist = model.fit_generator(generator=audio_gen.next_train(), steps_per_epoch=steps_per_epoch,
epochs=epochs, validation_data=audio_gen.next_valid(), validation_steps=validation_steps,
callbacks=[checkpointer], verbose=verbose)
# save model loss
with open('results/'+pickle_path, 'wb') as f:
pickle.dump(hist.history, f)
# -
train_model(input_to_softmax=model_1,
pickle_path='model_12.pickle',
save_model_path='model_12.h5',
spectrogram=True) # change to False if you would like to use MFCC features
# +
# import tensorflow as tf
# rnnmodel = tf.keras.models.load_model('../notebooks/results/model_1.h5')
loaded_model = pickle.load(open("../notebooks/results/model_1.pickle",'rb'))
# + jupyter={"outputs_hidden": true} tags=[]
# !pip install mlflow
# +
import mlflow
import mlflow.sklearn
import matplotlib.pyplot as plt
mlflow.end_run()
mlflow.set_experiment("Simple RNN")
for itr in range(20):
with mlflow.start_run():
mlflow.log_param("Model", "Simple RNN" )
ls=loaded_model.get('loss')[itr]
val=loaded_model.get('val_loss')[itr]
mlflow.log_metric("CTC_LOSS",ls)
mlflow.log_metric("Validatio_Loss",val)
vals=pd.DataFrame.from_dict(loaded_model)
print (vals)
plt.figure(figsize=(9,5))
plt.plot(vals['loss'],label="ctc_loss")
plt.plot(vals['val_loss'],label="validation_loss")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/RNN.png")
mlflow.log_artifact("../plots/RNN.png")
with open("metx.txt",'w') as outfile:
outfile.write("values",vals)
# -
vals['loss']
# +
with open("../metrics/loss.txt",'w') as outfile:
for ind in range(20):
outfile.write("loss: %s\n"%str(vals['loss'][ind]))
with open("../metrics/val_loss.txt",'w') as outfile:
for ind in range (20):
outfile.write("val_loss: %s \n"%str(vals['val_loss'][ind]))
# +
plt.figure(figsize=(10,9))
plt.plot(vals['loss'],label="ctc_loss")
plt.plot(vals['val_loss'],label="validation_loss")
plt.title("Losses per Epoch")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/RNN.png")
# -
plt.figure(figsize=(10,8))
sns.scatterplot(data=vals)
plt.title("Losses per Epoch")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/SCRNN.png")
model_2 = cnn_rnn_model(input_dim=13,
filters=200,
kernel_size=11,
conv_stride=2,
conv_border_mode='valid',
units=200)
train_model(input_to_softmax=model_2,
pickle_path='model_2.pickle',
save_model_path='model_2.h5',
spectrogram=False)
import pickle
loaded_model = pickle.load(open("../notebooks/results/model_2.pickle",'rb'))
# +
import mlflow
import mlflow.sklearn
import matplotlib.pyplot as plt
mlflow.set_experiment("CNN _ RNN")
for itr in range(20):
with mlflow.start_run():
mlflow.log_param("Model", "CNN RNN" )
ls=loaded_model.get('loss')[itr]
val=loaded_model.get('val_loss')[itr]
mlflow.log_metric("CTC_LOSS",ls)
mlflow.log_metric("ValidatioN_Loss",val)
vals=pd.DataFrame.from_dict(loaded_model)
print (vals)
plt.figure(figsize=(9,5))
plt.plot(vals['loss'],label="ctc_loss")
plt.plot(vals['val_loss'],label="validation_loss")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/CNN.png")
mlflow.log_artifact("../plots/CNN.png")
# +
with open("../metrics/CNNloss.txt",'w') as outfile:
for ind in range(20):
outfile.write("loss: %s\n"%str(vals['loss'][ind]))
with open("../metrics/CNNval_loss.txt",'w') as outfile:
for ind in range (20):
outfile.write("val_loss: %s \n"%str(vals['val_loss'][ind]))
# +
plt.figure(figsize=(10,9))
plt.plot(vals['loss'],label="ctc_loss")
plt.plot(vals['val_loss'],label="validation_loss")
plt.title("Losses per Epoch")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/CNN.png")
# +
model4=pickle.load(open("../notebooks/results/model_4.pickle",'rb'))
vals=pd.DataFrame.from_dict(model4)
# +
import mlflow
import mlflow.sklearn
import matplotlib.pyplot as plt
mlflow.end_run()
mlflow.set_experiment("Tuned RNN")
for itr in range(20):
with mlflow.start_run():
mlflow.log_param("Model", " RNN" )
ls=model4.get('loss')[itr]
val=model4.get('val_loss')[itr]
mlflow.log_metric("CTC_LOSS",ls)
mlflow.log_metric("Validatio_Loss",val)
vals=pd.DataFrame.from_dict(model4)
print (vals)
plt.figure(figsize=(9,5))
plt.plot(vals['loss'],label="ctc_loss")
plt.plot(vals['val_loss'],label="validation_loss")
plt.xlabel("EPOCH")
plt.ylabel("LOSS")
plt.legend()
plt.show()
plt.savefig("../plots/TunedRNN.png")
mlflow.log_artifact("../plots/TunedRNN.png")
# with open("metx.txt",'w') as outfile:
# outfile.write("values",vals)
# -
vals
| notebooks/ModelTraining.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib
import numpy
import pandas
import random
import sklearn
import xgboost
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import (
feature_extraction, feature_selection, decomposition, linear_model,
model_selection, metrics, svm
)
#pandas.options.display.max_rows=1000
#pandas.options.display.max_columns=300
# -
df=pandas.read_csv("numerai_training_data.csv")
df.head()
# There are 501808 rows grouped into eras, and a single target (target)
df.shape
# There's 310 features
features = [c for c in df if c.startswith("feature")]
df["erano"] = df.era.str.slice(3).astype(int)
eras = df.erano
target = "target"
len(features)
# The features are grouped together into 6 types
feature_groups = {
g: [c for c in df if c.startswith(f"feature_{g}")]
for g in ["intelligence", "wisdom", "charisma", "dexterity", "strength", "constitution"]
}
feature_groups
# +
# The models should be scored based on the rank-correlation (spearman) with the target
def numerai_score(y_true, y_pred):
rank_pred = y_pred.groupby(eras).apply(lambda x: x.rank(pct=True, method="first"))
return numpy.corrcoef(y_true, rank_pred)[0,1]
# It can also be convenient while working to evaluate based on the regular (pearson) correlation
def correlation_score(y_true, y_pred):
return numpy.corrcoef(y_true, y_pred)[0,1]
# -
# There are 120 eras numbered from 1 to 120
eras.describe()
# The earlier eras are smaller, but generally each era is 4000-5000 rows
df.groupby(eras).size().plot()
# The target is discrete and takes on 5 different values
df.groupby(target).size()
# # Some of the features are very correlated
# Especially within feature groups
feature_corrs = df[features].corr()
feature_corrs.stack().head()
tdf = feature_corrs.stack()
tdf = tdf[tdf.index.get_level_values(0) < tdf.index.get_level_values(1)]
tdf.sort_values()
# ### The correlation can change over time
# You can see this by comparing feature correlations on the first half and second half on the training set
df1 = df[eras<=eras.median()]
df2 = df[eras>eras.median()]
corr1 = df1[features].corr().unstack()
corr1 = corr1[corr1.index.get_level_values(0) < corr1.index.get_level_values(1)]
corr2 = df2[features].corr().unstack()
corr2 = corr2[corr2.index.get_level_values(0) < corr2.index.get_level_values(1)]
tdf = pandas.DataFrame({
"corr1": corr1,
"corr2": corr2,
})
tdf["corr_diff"] = tdf.corr2 - tdf.corr1
tdf.sort_values(by="corr_diff")
# ## Some features are predictive on their own
feature_scores = {
feature: numerai_score(df[target], df[feature])
for feature in features
}
pandas.Series(feature_scores).sort_values()
# Single features do not work consistently though
by_era_correlation = pandas.Series({
era: numpy.corrcoef(tdf[target], tdf["feature_strength34"])[0,1]
for era, tdf in df.groupby(eras)
})
by_era_correlation.plot()
# With a rolling 10 era average you can see some trends
by_era_correlation.rolling(10).mean().plot()
# # Gotcha: MSE looks worse than correlation out of sample
# Models will generally be overconfident, so even if they are good at ranking rows, the Mean-Squared-Error of the residuals could be larger than event the Mean-Squared-Error of the target (r-squared<0)
df1 = df[eras<=eras.median()]
df2 = df[eras>eras.median()]
linear1 = linear_model.LinearRegression()
linear1.fit(df1[features], df1[target])
linear2 = linear_model.LinearRegression()
linear2.fit(df2[features], df2[target])
# Note in particular that the R-squared of (train_on_1, eval_on_2) is slightly negative!
r2 = [
[
model.score(dfX[features], dfX[target])
for dfX in [df1, df2]
]
for model in [linear1, linear2]
]
pandas.DataFrame(r2, columns=["eval_on_1", "eval_on_2"], index=["train_on_1", "train_on_2"])
# Note in particular that the correlation of (train_on_1, eval_on_2) is quite decent
corrs = [
[
numerai_score(dfX[target], pandas.Series(model.predict(dfX[features]), index=dfX.index))
for dfX in [df1, df2]
]
for model in [linear1, linear2]
]
pandas.DataFrame(corrs, columns=["eval_on_1", "eval_on_2"], index=["train_on_1", "train_on_2"])
# +
# This can be be run with XGB as well
# -
xgb1 = xgboost.XGBRegressor()
xgb1.fit(df1[features], df1[target])
xgb2 = xgboost.XGBRegressor()
xgb2.fit(df2[features], df2[target])
r2 = [
[
model.score(dfX[features], dfX[target])
for dfX in [df1, df2]
]
for model in [xgb1, xgb2]
]
pandas.DataFrame(r2, columns=["eval_on_1", "eval_on_2"], index=["train_on_1", "train_on_2"])
corrs = [
[
numerai_score(dfX[target], pandas.Series(model.predict(dfX[features]), index=dfX.index))
for dfX in [df1, df2]
]
for model in [xgb1, xgb2]
]
pandas.DataFrame(corrs, columns=["eval_on_1", "eval_on_2"], index=["train_on_1", "train_on_2"])
# # Gotcha: {0, 1} are noticeably different from {0.25, 0.75}
# This makes training a classifier one-versus-rest behave counterintuitively.
#
# Specifically, the 0-vs-rest and 1-vs-rest classifiers seem to learn how to pick out extreme targets, and their predictions are the most correlated
# Train a standard logistic regression as a classifier
logistic = linear_model.LogisticRegression()
logistic.fit(df[features], (df[target]*4).astype(int))
logistic.score(df[features], (df[target]*4).astype(int))
# The first and last class are highly correlated
corrs=numpy.corrcoef(logistic.predict_proba(df[features]).T)
plt.imshow(corrs, vmin=-1, vmax=1, cmap="RdYlGn")
corrs
# In-sample correlation is 5.4%
preds = pandas.Series(logistic.predict_proba(df[features]).dot(logistic.classes_), index=df.index)
numerai_score(df[target], preds)
# A standard linear model has a slightly higher correlation
linear = linear_model.LinearRegression()
linear.fit(df[features], df[target])
linear.score(df[features], df[target])
preds = pandas.Series(linear.predict(df[features]), index=df.index)
numerai_score(df[target], preds)
# # Gotcha: eras are homogenous, but different from each other
# ## Random cross-validation will look much better than cross-validating by era
#
# Even for a simple linear model, taking a random shuffle reports a correlation of 4.3%, but a time series split reports a lower score of 3.4%
crossvalidators = [
model_selection.KFold(5),
model_selection.KFold(5, shuffle=True),
model_selection.GroupKFold(5),
model_selection.TimeSeriesSplit(5)
]
def correlation_score(y_true, y_pred):
return numpy.corrcoef(y_true, y_pred)[0,1]
for cv in crossvalidators:
print(cv)
print(numpy.mean(
model_selection.cross_val_score(
linear_model.LinearRegression(),
df[features],
df[target],
cv=cv,
n_jobs=1,
groups=eras,
scoring=metrics.make_scorer(correlation_score, greater_is_better=True)
)))
print()
# ## Eras can be more or less applicable to other eras
# You can test this be splitting the eras into blocks of 10, training on each block, and evaluating on each other block.
eras10 = (eras // 10) * 10
eras10.value_counts()
results10 = []
for train_era, tdf in df[eras10<120].groupby(eras10):
print(train_era)
model = linear_model.LinearRegression()
model.fit(tdf[features], tdf[target])
for test_era, tdf in df[eras10<120].groupby(eras10):
results10.append([
train_era,
test_era,
correlation_score(tdf[target], model.predict(tdf[features]))
])
results_df = pandas.DataFrame(
results10,
columns=["train_era", "test_era", "score"]
).pivot(index="train_era", columns="test_era", values="score")
results_df
# Each row here is the training block of eras, each column is a testing block of eras.
# Note that there is a period in the middle that does not seem to be relevant to other eras, and the
# overall performance seems to decrease a bit over time.
plt.figure(figsize=(15,15))
plt.imshow(results_df, vmin=-0.04, vmax=0.04, cmap="RdYlGn")
# + [markdown] pycharm={"name": "#%% md\n"}
# Here is an advanced paper that talks about generalization.
# Eras can be thought about in the same way that "distributions" or "environments" are talked about here
# https://arxiv.org/pdf/1907.02893.pdf
# -
# ## Gotcha: Since the signal-to-noise ratio is so low, models can take many more iterations than expected, and have scarily high in-sample performance
# +
def our_score(preds, dtrain):
return "score", -numpy.corrcoef(preds, dtrain.get_label())[0,1]
dtrain = xgboost.DMatrix(df1[features], df1[target])
dtest = xgboost.DMatrix(df2[features], df2[target])
dall = xgboost.DMatrix(df[features], df[target])
# +
param = {
'max_depth':3,
'eta':0.1,
'silent':1,
'objective':'reg:linear',
'eval_metric':'rmse',
'nthread': -1,
}
evals_result = {}
bst = xgboost.train(
params=param,
dtrain=dtrain,
feval=our_score,
num_boost_round=1000,
evals=[(dtrain, 'train'), (dtest, 'test')],
evals_result=evals_result,
verbose_eval=10,
)
(0.5 - 0.57*pandas.DataFrame({k: v['score'] for k,v in evals_result.items()})).plot()
# -
(-pandas.DataFrame({k: v['score'] for k,v in evals_result.items()})).plot(ylim=[0,0.045])
# # The results are sensitive to the choice of parameters, which should be picked through cross-validation
df1 = df[eras<=eras.median()]
df2 = df[eras>eras.median()]
models = [
linear_model.LinearRegression(),
] + [
linear_model.ElasticNet(alpha=alpha)
for alpha in [0.01, 0.005, 0.002, 0.001, 0.0005, 0.0002, 0.0001, 0.00005, 0.00002, 0.00001]
] + [
xgboost.XGBRegressor(n_jobs=-1),
xgboost.XGBRegressor(n_jobs=-1, learning_rate=0.01, n_estimators=1000),
xgboost.XGBRegressor(n_jobs=-1, colsample_bytree=0.1, learning_rate=0.01, n_estimators=1000),
xgboost.XGBRegressor(n_jobs=-1, colsample_bytree=0.1, learning_rate=0.01, n_estimators=1000, max_depth=5),
xgboost.XGBRegressor(n_jobs=-1, colsample_bytree=0.1, learning_rate=0.001, n_estimators=10000, max_depth=5),
]
for model in models:
print(" -- ", model)
model.fit(df1[features], df1[target])
outsample = numerai_score(df2[target], pandas.Series(model.predict(df2[features]), index=df2.index))
insample = numerai_score(df1[target], pandas.Series(model.predict(df1[features]), index=df1.index))
print(
f"outsample: {outsample}, insample: {insample}"
)
print()
# ## Gotcha: Models with large exposures to individual features tend to perform poorly or inconsistently out of sample ##
#
import numpy as np
import scipy
# Train a standard xgboost on half the train eras
xgb = xgboost.XGBRegressor(n_estimators=1000, max_depth=5, learning_rate=0.01, n_jobs=-1)
xgb.fit(df1[features], df1[target])
xgb_preds = xgb.predict(df2[features])
# ### Our predictions have correlation > 0.2 in either direction for some single features!
# Sure hope those features continue to act as they have in the past!
corr_list = []
for feature in features:
corr_list.append(numpy.corrcoef(df2[feature], xgb_preds)[0,1])
corr_series = pandas.Series(corr_list, index=features)
corr_series.describe()
from sklearn.preprocessing import MinMaxScaler
def _neutralize(df, columns, by, proportion=1.0):
scores = df[columns]
exposures = df[by].values
scores = scores - proportion * exposures.dot(numpy.linalg.pinv(exposures).dot(scores))
return scores / scores.std(ddof=0)
def _normalize(df):
X = (df.rank(method="first") - 0.5) / len(df)
return scipy.stats.norm.ppf(X)
def normalize_and_neutralize(df, columns, by, proportion=1.0):
# Convert the scores to a normal distribution
df[columns] = _normalize(df[columns])
df[columns] = _neutralize(df, columns, by, proportion)
return df[columns]
df2["preds"] = xgb_preds
df2["preds_neutralized"] = df2.groupby("era").apply(
lambda x: normalize_and_neutralize(x, ["preds"], features, 0.5) # neutralize by 50% within each era
)
scaler = MinMaxScaler()
df2["preds_neutralized"] = scaler.fit_transform(df2[["preds_neutralized"]]) # transform back to 0-1
# ### Now our single feature exposures are much smaller
corr_list2 = []
for feature in features:
corr_list2.append(numpy.corrcoef(df2[feature], df2["preds_neutralized"])[0,1])
corr_series2 = pandas.Series(corr_list2, index=features)
corr_series2.describe()
# ### Our overall score goes down, but the scores are more consistent than before. This leads to a higher sharpe
# +
unbalanced_scores_per_era = df2.groupby("era").apply(lambda d: np.corrcoef(d["preds"], d[target])[0,1])
balanced_scores_per_era = df2.groupby("era").apply(lambda d: np.corrcoef(d["preds_neutralized"], d[target])[0,1])
print(f"score for high feature exposure: {unbalanced_scores_per_era.mean()}")
print(f"score for balanced feature expo: {balanced_scores_per_era.mean()}")
print(f"std for high feature exposure: {unbalanced_scores_per_era.std(ddof=0)}")
print(f"std for balanced feature expo: {balanced_scores_per_era.std(ddof=0)}")
print(f"sharpe for high feature exposure: {unbalanced_scores_per_era.mean()/unbalanced_scores_per_era.std(ddof=0)}")
print(f"sharpe for balanced feature expo: {balanced_scores_per_era.mean()/balanced_scores_per_era.std(ddof=0)}")
# -
balanced_scores_per_era.describe()
unbalanced_scores_per_era.describe()
| example_scripts/analysis_and_tips.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Explicit Models Tutorial
# This tutorial will show you how to create and use `ExplicitOpModel` objects. `Model` objects are fundamental to pyGSTi, as each represents a set of quantum gates along with state preparation and measurement (i.e. POVM) operations. In pyGSTi, a "state space" refers to a Hilbert space of *pure* quantum states (often thought of as length-$d$ vectors, where $d=2^N$ for $N$ qubits). A "density matrix space" refers to a Hilbert space of density matrices, which while often thought of as $d \times d$ matrices can also be represented by length $d^2$ vectors. Mathematically, these vectors live in Hilbert-Schmidt space, the space of linear operators on the original $d\times d$ density matrix space. pyGSTi uses the "Liouville" vector-representation for density matrices and POVM effects, since this allows quantum gates to be represented by $d^2 \times d^2$ matrices which act on Hilbert-Schmidt vectors.
#
# `ExplicitOpModel` objects are the simplest type of `Model` objects in pyGSTi. They have the look and feel of Python dictionaries which hold $d^2\times d^2$ operation matrices, length-$d^2$ state preparation vectors, and sets of length-$d^2$ effect vectors which encode positive operator value measures (POVMs). State preparation and POVM effect vectors are both generically referred to as "SPAM" (state preparation and measurement) vectors.
import pygsti
import pygsti.construction as pc
# ## Creating models
# Before getting to `ExplicitOpModels` in particular, lets explain two quantites that *all* `Model` objects posess: a *basis* and *state space labels*:
# - A model's `.state_space_labels` member (a `StateSpaceLabels` object) describes the model's state space as the direct sum and tensor product of labelled *factors*. Typically, this is just a tensor product of one or more 2-dimensional qubit spaces labelled by the integers 0 to $N_{qubits}-1$ or `"Q0"`, `"Q1"`, etc. We specify a 1-qubit state space using `["Q0"]` below (the "Q" tells pyGSTi it's a 2-dimensional *qubit* space). If you had two qubits you could use `["Q0","Q1"]` or `[0,1]` to describe the tensor product of two qubit spaces, as pyGSTi assumes integer labels stand for qubit spaces too. To learn more about the `StateSpaceLabels` object, see the [state space labelling tutorial](advanced/StateSpaceLabels.ipynb).
# - A model's `.basis` member (a `Basis` object) describes how any dense representations (matrices or vectors) of the the operations in a `Model` should be interpreted. We'll be using the "Pauli product" basis, which is named `"pp"` in pyGSTi and consists of the tensor products of Pauli matrices (since our example has just a 1-qubit state space the `"pp"` basis is just the 4 Pauli matrices $\{\sigma_0,\sigma_X,\sigma_Y,\sigma_Z\}$). To learn more about `Basis` objects see the [Basis object tutorial](advanced/MatrixBases.ipynb)).
#
#
# ## Creating explicit models
# There are more or less four ways to create `ExpicitOpModel` objects in pyGSTi:
#
# * By creating an empty `ExpicitOpModel` and setting its elements directly, possibly with the help of `pygsti.construction`'s `build_operation` and `build_vector` functions.
# * By a single call to `create_explicit_model`, which automates the above approach.
# * By loading from a text-format model file using `pygsti.io.load_model`.
# * By copying one from one of the `pygsti.construction.std*` modules.
# ### Creating a `ExplicitOpModel` from scratch
#
# Layer operations (often called "gates" in a 1- or even 2-qubit context) and SPAM vectors can be assigned to a `ExplicitOpModel` object as to an ordinary python dictionary. Internally a `ExpicitOpModel` holds these quantities as `LinearOperator`- and `SPAMVec`- and `POVM`-derived objects, but you may assign lists, Numpy arrays, or other types of Python iterables to a `ExplicitOpModel` key and a conversion will be performed automatically. To keep gates, state preparations, and POVMs separate, the `ExplicitOpModel` object looks at the beginning of the dictionary key being assigned: keys beginning with `rho`, `M`, and `G` are categorized as state preparations, POVMs, and gates, respectively. To avoid ambiguity, each key *must* begin with one of these three prefixes.
#
# To separately access (set or get) the state preparations, POVMs, and operations contained in a `ExplicitOpModel` use the `preps`, `povms`, and `operations` members respectively. Each one provides dictionary-like access to the underlying objects. For example, `myModel.operations['Gx']` accesses the same underlying `LinearOperator` object as `myModel['Gx']`, and similarly for `myModel.preps['rho0']` and `myModel['rho0']`. The values of operations and state preparation vectors can be read and written in this way.
#
# A POVM object acts similarly to dictionary of `SPAMVec`-derived effect vectors, but typically requires all such vectors to be initialized at once, that is, you cannot assign individual effect vectors to a `POVM`. The string-valued keys of a `POVM` label the outcome associated with each effect vector, and are therefore termed *effect labels* or *outcome labels*. The outcome labels also designate data within a `DataSet` object (see the [DataSet tutorial](DataSet.ipynb)), and thereby associate modeled POVMs with experimental measurements.
#
#
#
# The below cell illustrates how to create a `ExplicitOpModel` from scratch.
# +
from math import sqrt
import numpy as np
#Initialize an empty Model object
#Designate the basis being used for the matrices and vectors below
# as the "Pauli product" basis of dimension 2 - i.e. the four 2x2 Pauli matrices I,X,Y,Z
model1 = pygsti.objects.ExplicitOpModel(['Q0'],'pp')
#Populate the Model object with states, effects, gates,
# all in the *normalized* Pauli basis: { I/sqrt(2), X/sqrt(2), Y/sqrt(2), Z/sqrt(2) }
# where I, X, Y, and Z are the standard Pauli matrices.
model1['rho0'] = [ 1/sqrt(2), 0, 0, 1/sqrt(2) ] # density matrix [[1, 0], [0, 0]] in Pauli basis
model1['Mdefault'] = pygsti.objects.UnconstrainedPOVM(
{'0': [ 1/sqrt(2), 0, 0, 1/sqrt(2) ], # projector onto [[1, 0], [0, 0]] in Pauli basis
'1': [ 1/sqrt(2), 0, 0, -1/sqrt(2) ] }) # projector onto [[0, 0], [0, 1]] in Pauli basis
model1['Gi'] = np.identity(4,'d') # 4x4 identity matrix
model1['Gx'] = [[1, 0, 0, 0],
[0, 1, 0, 0],
[0, 0, 0,-1],
[0, 0, 1, 0]] # pi/2 X-rotation in Pauli basis
model1['Gy'] = [[1, 0, 0, 0],
[0, 0, 0, 1],
[0, 0, 1, 0],
[0,-1, 0, 0]] # pi/2 Y-rotation in Pauli basis
pygsti.io.write_model(model1, "../tutorial_files/Example_gatesetFromScratch.txt", title="My Model")
# -
# Check out the model file that was written [here](../tutorial_files/Example_gatesetFromScratch.txt).
# ### Creating a `ExplicitOpModel` from scratch using `modelconstruction._basis_create_operation` and `modelconstruction._basis_create_spam_vector`
# The `modelconstruction._basis_create_operation` and `modelconstruction._basis_create_spam_vector` functions take a human-readable string representation of a gate or SPAM vector, and return a `LinearOperator` or `SPAMVector` object that gets stored in a dictionary-like `ExplicitOpModel` or `POVM` object. To use these functions, you must specify what state space you're working with, and the basis for that space - so the `.state_space_labels` and `.basis` member of your `Model` object, as described above.
#
# `build_vector` currently only understands strings which are integers (e.g. "1"), for which it creates a vector performing state preparation of (or, equivalently, a state projection onto) the $i^{th}$ state of the Hilbert space, that is, the state corresponding to the $i^{th}$ row and column of the $d\times d$ density matrix.
#
# `build_operation` accepts a wider range of descriptor strings, which take the form of *functionName*(*args*) and include:
# - `I(label0, label1, ...)` : the identity on the spaces labeled by `label0`, `label1`, etc.
# - `X(theta,Qlabel)`, `Y(theta,Qlabel)`, `Z(theta,Qlabel)` : single qubit X-, Y-, and Z-axis rotations by angle `theta` (in radians) on the qubit labeled by `Qlabel`. Note that `pi` can be used within an expression for `theta`, e.g. `X(pi/2,Q0)`.
# - `CX(theta, Qlabel1, Qlabel2)`, `CY(theta, Qlabel1, Qlabel2)`, `CZ(theta, Qlabel1, Qlabel2)` : two-qubit controlled rotations by angle `theta` (in radians) on qubits `Qlabel1` (the control) and `Qlabel2` (the target).
# +
#Initialize an empty Model object
model2 = pygsti.objects.ExplicitOpModel(['Q0'],'pp') # single qubit labelled 'Q0'; Pauli basis
spaceLabels = model2.state_space_labels
basis = model2.basis
#Populate the Model object with states, effects, and gates using
# build_vector, build_operation, and create_identity_vec.
model2['rho0'] = pc.modelconstruction._basis_create_spam_vector("0",basis)
model2['Mdefault'] = pygsti.objects.UnconstrainedPOVM(
{ '0': pc.modelconstruction._basis_create_spam_vector("0",basis),
'1': pc.modelconstruction._basis_create_spam_vector("1",basis) } )
model2['Gi'] = pc.modelconstruction._basis_create_operation(spaceLabels,"I(Q0)",basis)
model2['Gx'] = pc.modelconstruction._basis_create_operation(spaceLabels,"X(pi/2,Q0)",basis)
model2['Gy'] = pc.modelconstruction._basis_create_operation(spaceLabels,"Y(pi/2,Q0)",basis)
# -
# ### Create a `ExplicitOpModel` in a single call to create_explicit_model
# The approach illustrated above using calls to `build_vector` and `build_operation` can be performed in a single call to `create_explicit_model`. You will notice that all of the arguments to `create_explicit_model` corrspond to those used to construct a model using `build_vector` and `build_operation`; the `create_explicit_model` function is merely a convenience function which allows you to specify everything at once. These arguments are:
# - Arg 1 : the state-space-labels, as described above.
# - Args 2 & 3 : list-of-gate-labels, list-of-gate-expressions (labels *must* begin with 'G'; "expressions" being the descriptor strings passed to `build_operation`)
# - Args 4 & 5 : list-of-prep-labels, list-of-prep-expressions (labels *must* begin with 'rho'; "expressions" being the descriptor strings passed to `build_vector`)
# - Args 6 & 7 : list-of-effect-labels, list-of-effect-expressions (labels can be anything; "expressions" being the descriptor strings passed to `build_vector`). These effect vectors will comprise a single POVM named `"Mdefault"` by default, but which can be changed via the `povmLabels` argument (see doc string for details).
#
# The optional argument `basis` can be set to any of the known built-in basis *names* (e.g. `"gm"`, `"pp"`, `"qt"`, or `"std"`) to select the basis for the Model as described above. By default, `"pp"` is used when possible (if the state space corresponds to an integer number of qubits), `"qt"` if the state space has dimension 3, and `"gm"` otherwise. The optional argument `parameterization` is used to specify the parameterization used for the created gates (see below).
model3 = pc.create_explicit_model(['Q0'],
['Gi','Gx','Gy'], [ "I(Q0)","X(pi/2,Q0)", "Y(pi/2,Q0)"],
prep_labels=['rho0'], prep_expressions=["0"],
effect_labels=['0','1'], effect_expressions=["0","1"] )
# In this case, the parameters to `create_explicit_model`, specify:
#
# - The state space has dimension 2 and is interpreted as that of a single qubit labeled "Q0" (label must begin with 'Q' or be an integer if we don't want to create a full `StateSpaceLabels` object that contains the dimensions too.)
#
# - there are three gates: Idle, $\pi/2$ x-rotation, $\pi/2$ y-rotation, labeled `Gi`, `Gx`, and `Gy`.
#
# - there is one state prep operation, labeled `rho0`, which prepares the 0-state (the first basis element of the 2D state space)
#
# - there is one POVM (~ measurement), named `Mdefault` with two effect vectors: `'0'` projects onto the 0-state (the first basis element of the 2D state space) and `'1'` projects onto the 1-state.
#
# Note that **by default**, there is a single state prep, `"rho0"`, that prepares the 0-state and a single POVM, `"Mdefault"`, which consists of projectors onto each standard basis state that are labelled by their integer indices (so just `'0'` and `'1'` in the case of 1-qubit). Thus, all but the first four arguments used above just specify the default behavior and can be omitted:
model4 = pc.create_explicit_model( ['Q0'], ['Gi','Gx','Gy'], [ "I(Q0)","X(pi/2,Q0)", "Y(pi/2,Q0)"] )
# ### Load a `ExplicitOpModel` from a file
# You can also construct a `ExplicitOpModel` object from a file using `pygsti.io.load_model`. The format of the text file should be fairly self-evident given the above discussion. Note that vector and matrix elements need not be simple numbers, but can be any mathematical expression parseable by the Python interpreter, and in addition to numbers can include "sqrt" and "pi".
# +
#3) Write a text-format model file and read it in.
model5_txt = \
"""
# Example text file describing a model
PREP: rho0
LiouvilleVec
1/sqrt(2) 0 0 1/sqrt(2)
POVM: Mdefault
EFFECT: 0
LiouvilleVec
1/sqrt(2) 0 0 1/sqrt(2)
EFFECT: 1
LiouvilleVec
1/sqrt(2) 0 0 -1/sqrt(2)
END POVM
GATE: Gi
LiouvilleMx
1 0 0 0
0 1 0 0
0 0 1 0
0 0 0 1
GATE: Gx
LiouvilleMx
1 0 0 0
0 1 0 0
0 0 0 -1
0 0 1 0
GATE: Gy
LiouvilleMx
1 0 0 0
0 0 0 1
0 0 1 0
0 -1 0 0
STATESPACE: Q0(4)
BASIS: pp
"""
with open("../tutorial_files/Example_ExplicitModel.txt","w") as gsetfile:
gsetfile.write(model5_txt)
model5 = pygsti.io.load_model("../tutorial_files/Example_ExplicitModel.txt")
# -
#All four of the above models are identical. See this by taking the frobenius differences between them:
assert(model1.frobeniusdist(model2) < 1e-8)
assert(model1.frobeniusdist(model3) < 1e-8)
assert(model1.frobeniusdist(model4) < 1e-8)
assert(model1.frobeniusdist(model5) < 1e-8)
# ## Viewing models
# In the cells below, we demonstrate how to print and access information within a `ExplicitOpModel`.
#Printing the contents of a Model is easy
print("Model 1:\n", model1)
# +
#You can also access individual gates like they're numpy arrays:
Gx = model1['Gx'] # a LinearOperator object, but behaves like a numpy array
#By printing a gate, you can see that it's not just a numpy array
print("Gx = ", Gx)
#But can be accessed as one:
print("Array-like printout\n", Gx[:,:],"\n")
print("First row\n", Gx[0,:],"\n")
print("Element [2,3] = ",Gx[2,3], "\n")
Id = np.identity(4,'d')
Id_dot_Gx = np.dot(Id,Gx)
print("Id_dot_Gx\n", Id_dot_Gx, "\n")
# -
# ## Basic Operations with Explicit Models
#
# `ExplicitOpModel` objects have a number of methods that support a variety of operations, including:
#
# * Depolarizing or rotating every gate
# * Writing the model to a file
# * Computing products of operation matrices
# * Printing more information about the model
# +
#Add 10% depolarization noise to the gates
depol_model3 = model3.depolarize(op_noise=0.1)
#Add a Y-axis rotation uniformly to all the gates
rot_model3 = model3.rotate(rotate=(0,0.1,0))
# -
#Writing a model as a text file
pygsti.io.write_model(depol_model3, "../tutorial_files/Example_depolarizedModel.txt", title="My Model")
# +
print("Probabilities of outcomes of the gate\n sequence GxGx (rho0 and Mdefault assumed)= ",
depol_model3.probabilities( ("Gx", "Gx")))
print("Probabilities of outcomes of the \"complete\" gate\n sequence rho0+GxGx+Mdefault = ",
depol_model3.probabilities( ("rho0", "Gx", "Gx", "Mdefault")))
# -
#Printing more detailed information about a model
depol_model3.print_info()
# It is also possible to manipulate the underlying operations by accessing the forward simulator in the `Model` class. For example, using the `matrix` simulator type, one can compute the product of two gate operations. For more details, see the [ForwardSimulationTypes tutorial](../algorithms/advanced/ForwardSimulationTypes.ipynb).
# Computing the product of operation matrices (only allowed with the matrix simulator type)
print("Product of Gx * Gx = \n",depol_model3.sim.product(("Gx", "Gx")), end='\n\n')
# ## Explicit Model Parameterizations
# In addition to specifying a set of $d^2 \times d^2$ operation matrices and length-$d^2$ SPAM vectors, every `Model` encapsulates a *parametrization*, that is, a function mapping a set of real-valued *parameters* to its set of operation matrices and SPAM vectors. A `Model`'s contents must always correspond to a valid set of parameters, which can be obtained by its `to_vector` method, and can always be initialized from a vector of parameters via its `from_vector` method. The number of parameters (obtained via `num_params`) is independent (and need not equal!) the total number of gate-matrix and SPAM-vector elements comprising the `Model`. For example, in a "TP-parameterized" model, the first row of each operation matrix is fixed at `[1,0,...0]`, regardless to what the `Model`'s underlying parameters are. When pyGSTi generates `Model` estimates the *parameters* of an initial `Model` (often times the "target" model) supplied by the caller are optimized. Thus, by its parameterization a single `Model` can determine the space of possible `Model`s that are searched for a best-fit estimate.
#
# Each gate and SPAM vector within a `ExplicitOpModel` have independent paramterizations, so that each `pygsti.objects.LinearOperator`-derived gate object and `pygsti.objects.SPAMVec`-derived SPAM vector has its own `to_vector`, `from_vector`, and `num_params` method. A `Model`'s parameter vector is just the concatenation of the parameter vectors of its contents, in the order: 1) state preparation vectors, 2) measurement vectors, 3) gates.
#
# Users are able to create their own gate parameterizations by deriving from `pygsti.objects.LinearOperator` or `pygsti.objects.DenseOperator` (which itself derives from `LinearOperator`). Included in pyGSTi are several convenient gate parameterizations which are worth knowing about:
#
# - The `FullArbitraryOp` class defines a gate which has a parameter for every element, and thus optimizations using this gate class allow the operation matrix to be completely arbitrary.
# - The `FullTPOp` class defines a gate whose first row must be `[1,0,...0]`. This corresponds to a trace-preserving (TP) gate in the Gell-Mann and Pauli-product bases. Each element in the remaining rows is a separate parameter, similar to a fully parameterized gate. Optimizations using this gate type are used to constrain the estimated gate to being trace preserving.
# - The `LindbladOp` (or `LindbladDenseOp`) class defines a gate whose logarithm take a particular Lindblad form. This class is fairly flexible, but is predominantly used to constrain optimizations to the set of infinitesimally-generated CPTP maps.
#
# Similarly, there are `FullSPAMVec` and `TPSPAMVec` classes, the latter which fixes its first element to $\sqrt{d}$, where $d^2$ is the vector length, as this is the appropriate value for a unit-trace state preparation.
#
# We now illustrate how one map specify the type of paramterization in `create_explicit_model`, and change the parameterizations of all of a `ExplicitOpModel`'s contents using its `set_all_parameterizaions` method.
# +
# Speciy basis as 'gm' for Gell-Mann (could also be 'pp' for Pauli-Product)
# and parameterization to 'TP', so that gates are TPParameterizedGates
model6 = pc.create_explicit_model(['Q0'], ['Gi',], [ "I(Q0)"],
basis='pp', parameterization="TP")
#See that gates and prep vectors are TP, whereas previous Model's have
# fully parameterized elements
print("model6 gate type = ", type(model6['Gi']))
print("model6 prep type = ", type(model6['rho0']))
print("model5 gate type = ", type(model5['Gi']))
print("model5 prep type = ", type(model5['rho0']))
#Switch parameterization to CPTP gates
model6.set_all_parameterizations('CPTP')
print("\nAfter setting all parameterizations to CPTP:")
print("model6 gate type = ", type(model6['Gi']))
print("model6 prep type = ", type(model6['rho0']))
# -
# To alter an individual gate or SPAM vector's parameterization, one can simply construct a `LinearOperator` or `SPAMVec` object with the desired parameterization and assign it to the `Model`.
newOp = pygsti.objects.FullTPOp(model6['Gi'])
model6['Gi'] = newOp
print("model6['Gi'] =",model6['Gi'])
# **NOTE:** When a `LinearOperator` or `SPAMVec`-derived object is assigned as an element of an `ExplicitOpModel` (as above), the object *replaces* any existing object with the given key. However, if any other type of object is assigned to an `ExplicitOpModel` element, an attempt is made to initialize or update the existing existing gate using the assigned data (using its `set_matrix` function internally). For example:
# +
import numpy as np
numpy_array = np.array( [[1, 0, 0, 0],
[0, 0.5, 0, 0],
[0, 0, 0.5, 0],
[0, 0, 0, 0.5]], 'd')
model6['Gi'] = numpy_array # after assignment with a numpy array...
print("model6['Gi'] =",model6['Gi']) # this is STILL a FullTPOp object
#If you try to assign a gate to something that is either invalid or it doesn't know how
# to deal with, it will raise an exception
invalid_TP_array = np.array( [[2, 1, 3, 0],
[0, 0.5, 0, 0],
[0, 0, 0.5, 0],
[0, 0, 0, 0.5]], 'd')
try:
model6['Gi'] = invalid_TP_array
except ValueError as e:
print("ERROR!! " + str(e))
# -
| jupyter_notebooks/Tutorials/objects/ExplicitModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-hsic_align]
# language: python
# name: conda-env-.conda-hsic_align-py
# ---
# # Demo I - Different Cases
#
# In this document, I will be looking at the motivation behind this study and why we would like to pursue this further.
# +
import sys, os
import warnings
import tqdm
import random
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# Insert path to model directory,.
cwd = os.getcwd()
path = f"{cwd}/../../src"
sys.path.insert(0, path)
# Insert path to package,.
pysim_path = f"/home/emmanuel/code/pysim/"
sys.path.insert(0, pysim_path)
# toy datasets
from data.toy import generate_dependence_data
from data.distribution import DataParams
from dataclasses import dataclass
# Kernel Dependency measure
from sklearn.preprocessing import StandardScaler
from sklearn.gaussian_process.kernels import RBF
from models.dependence import HSICModel
from pysim.kernel.utils import get_init_gammas, get_gamma_grid, estimate_sigma
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use(['seaborn-talk'])
# %matplotlib inline
warnings.filterwarnings('ignore') # get rid of annoying warnings
# %load_ext autoreload
# %autoreload 2
# -
FIG_PATH = "/home/emmanuel/projects/2019_hsic_align/results/figures/1d_dataset/demo/"
# ##### Estimating Sigma & HSIC
# +
def standardize_data(X, Y, standardize: bool=False):
X = StandardScaler().fit_transform(X)
Y = StandardScaler().fit_transform(Y)
return X, Y
def get_sigma(X, Y, method: str='silverman', per_dimension: bool=False, separate_scales: bool=False):
# sigma parameters
subsample = None
percent = 0.20
random_state = 123
sigma_X = estimate_sigma(
X,
subsample=subsample,
method=method,
percent=percent,
random_state=random_state,
per_dimension=per_dimension
)
sigma_Y = estimate_sigma(
Y,
subsample=subsample,
method=method,
percent=percent,
random_state=random_state,
per_dimension=per_dimension
)
if separate_scales:
sigma_X = np.mean([sigma_X, sigma_Y])
sigma_Y = np.mean([sigma_X, sigma_Y])
return sigma_X, sigma_Y
def get_hsic(X, Y, scorer: str, sigma_X=None, sigma_Y=None):
# init hsic model class
hsic_model = HSICModel()
# hsic model params
if sigma_X is not None:
hsic_model.kernel_X = RBF(sigma_X)
hsic_model.kernel_Y = RBF(sigma_Y)
# get hsic score
hsic_val = hsic_model.get_score(X, Y, scorer)
return hsic_val
# -
# ## Data I - 1D Dataset
#
# +
# data params
dataset = 'sine'
num_points = 1000
seed = 123
noise = 0.1
# get dataset
X, Y = generate_dependence_data(
dataset=dataset,
num_points=num_points,
seed=seed,
noise_x=noise,
noise_y=noise
)
# plot
fig, ax = plt.subplots()
ax.scatter(X[:100,:], Y[:100,:])
plt.tight_layout()
fig.savefig(FIG_PATH + f"demo_{dataset}.png")
plt.show()
# -
# Let's take a simple 1D distribution: a sine curve. It is clear that there is a nonlinear relationship between them that cannot be captured (well) by linear methods. We are interested in looking at the dependence between $X$ and $Y$. We have the HSIC family of methods: HSIC, kernel alignment and centered kernel alignment. They are all very similar but there are some subtle differences. We will highlight them as we go through the overview. Let's take a generic approach and use the default HSIC, KA and CKA methods to try and estimate the dependence between $X,Y$. If we run the algorithm, we get the following results.
# ### Question I - Which Algorithm?
results_df = pd.DataFrame()
# +
method = 'scott'
per_dimension = False
separate_scales = False
# sigma_X, sigma_y = get_sigma(
# X, Y,
# method=method,
# per_dimension=per_dimension,
# separate_scales=separate_scales
# )
method = 'default'
sigma_X, sigma_Y = None, None
scorer = 'hsic'
results_df = results_df.append(pd.DataFrame({
"hsic": [get_hsic(X, Y, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"ka": [get_hsic(X, Y, 'ka', sigma_X, sigma_Y)], # Estimate KA
"cka": [get_hsic(X, Y, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q1']),)
print(results_df.to_markdown())
# -
# Notice how all of the values are slightly difference. This is because of the composition of the methods. We can highlight the differences with a simple table.
#
# <center>
#
# | **Method** | **Centered Kernel** | **Normalized** |
# | ------------------------- | ------------------- | -------------- |
# | HSIC | Yes | No |
# | Kernel Alignment | No | Yes |
# | Centered Kernel Alignment | Yes | No |
#
# </center>
#
# So each method has a slightly different formulation but they are mostly the same. So now the next question is: how do we estimate the parameters of the kernel used? Well the default is simply $\sigma=1.0$ but we know that this won't do as the kernel depends on the parameters of the kernel. In this case we are using the most commonly used kernel: the Radial Basis Function (RBF). Since this is a 1D example, I will use some generic estimators called the "Silverman Rule" and "Scott Rule". These are very commonly found in packages like `scipy.stats.gaussian_kde` or `statsmodels.nonparametric.bandwidth`. They are mostly used for the Kernel Density Estimation (KDE) where we need a decent parameter to approximate the kernel to get a decent density estimate.
#
# So what happens with the methods and the results?
# ### Question II - Which Parameter Estimator?
# +
methods = ['scott', 'silverman', 'median']
per_dimension = False
separate_scales = True
results_df = pd.DataFrame()
for imethod in methods:
sigma_X, sigma_Y = get_sigma(
X, Y,
method=imethod,
per_dimension=per_dimension,
separate_scales=separate_scales
)
results_df = results_df.append(pd.DataFrame({
# "sigma_x": [sigma_X],
# "sigma_y": [sigma_Y],
'Estimator': [imethod],
"hsic": [get_hsic(X, Y, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"ka": [get_hsic(X, Y, 'ka', sigma_X, sigma_Y)], # Estimate KA
"cka": [get_hsic(X, Y, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q2']),)
print(results_df.to_markdown())
# -
# ### Question III - How do we estimate the length scale?
#
# * Use the same length scale?
# * Use different length scales?
# * Use a length scale per dimension (D>1)
# +
methods = ['scott', 'silverman', 'median']
per_dimension = False
separate_scales = [True, False]
results_df = pd.DataFrame()
for iscaler in separate_scales:
for imethod in methods:
sigma_X, sigma_Y = get_sigma(
X, Y,
method=imethod,
per_dimension=per_dimension,
separate_scales=iscaler
)
results_df = results_df.append(pd.DataFrame({
# "sigma_x": [sigma_X],
"separate": [iscaler],
'Estimator': [imethod],
"hsic": [get_hsic(X, Y, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"ka": [get_hsic(X, Y, 'ka', sigma_X, sigma_Y)], # Estimate KA
"cka": [get_hsic(X, Y, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q3']),)
print(results_df.to_markdown())
# -
# ### Question IV - Standardize Data?
#
# We could also standardize our data... This could actually change the size of each of the features which could eliminate the need to apply separate length scales.
# +
standardize = [True, False]
methods = ['scott', 'silverman', 'median']
per_dimension = False
separate_scales = [True, False]
results_df = pd.DataFrame()
for istandard in standardize:
X_, Y_ = standardize_data(X, Y, istandard)
for iscaler in separate_scales:
for imethod in methods:
sigma_X, sigma_Y = get_sigma(
X_, Y_,
method=imethod,
per_dimension=per_dimension,
separate_scales=iscaler
)
results_df = results_df.append(pd.DataFrame({
"standardize": [istandard],
"separate": [iscaler],
'Estimator': [imethod],
"hsic": [get_hsic(X_, Y_, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"ka": [get_hsic(X_, Y_, 'ka', sigma_X, sigma_Y)], # Estimate KA
"cka": [get_hsic(X_, Y_, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q4']),)
print(results_df.to_markdown())
# -
# Now we see that the values you get are quite different for all methods. What happens if we use different sigmas?
# #### Verdict
#
# Well, hard to say as it depends on the parameters. Every researcher I've met who dealt with kernel methods seems to have a suggestion that they swear by but I never know who to follow. My thoughts is that we should use dedicated sigma values per dataset however, that still leaves us with other methods that we may want to try. So we're going to repeat the same experiment but with a 2D dataset and we will see that the difficult will increase again.
# ## 2D Example
# For this experiment, we're going to take two 2D datasets each generated from a T-Student distribution. We will apply the same sequence as we did above and we will end the section by adding another option for picking the parameters.
# + jupyter={"source_hidden": true}
# initialize Data Params class
dataset = 'tstudent'
samples = 1_000
dimensions = 2
std = 5
nu = 8
trial = 1
standardize = False
# initialize params
example_params = DataParams(
dataset=dataset,
samples=samples,
dimensions=dimensions,
std=std,
nu=nu,
trial=trial,
standardize=standardize
)
# generate some parameters
inputs = example_params.generate_data()
# + jupyter={"source_hidden": true}
sns.jointplot(x=inputs.X, y=inputs.Y)
plt.tight_layout()
plt.savefig(FIG_PATH + f"demo_{dataset}.png")
# -
# **Fig I**: An example 2D T-Student distribution.
# ### Question III (Revisited) - Different Length Scales?
#
# Now we can revisit this question because we actually could estimate a different length scale depending upon the dimensionality. One problem with scott or Silverman's method is that it takes into account the entire dataset instead of having one estimate per feature.
# +
methods = ['scott', 'silverman', 'median']
per_dimension = False
separate_scales = [True, False]
separate_dimensions = [True, False]
results_df = pd.DataFrame()
for iscaler in separate_scales:
for idim in separate_dimensions:
for imethod in methods:
sigma_X, sigma_Y = get_sigma(
X, Y,
method=imethod,
per_dimension=idim,
separate_scales=iscaler
)
results_df = results_df.append(pd.DataFrame({
"standardize": [istandard],
"Separate Dimensions": [idim],
"Separate Length Scales": [iscaler],
'Param Estimator': [imethod],
"HSIC": [get_hsic(X, Y, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"KA": [get_hsic(X, Y, 'ka', sigma_X, sigma_Y)], # Estimate KA
"CKA": [get_hsic(X, Y, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q3']),)
print(results_df.to_markdown())
# -
# ### Q1-Q4
#
# So now, let's look at all questions for the 2D data distribution
# +
standardize = [True, False]
methods = ['scott', 'silverman', 'median']
per_dimension = False
separate_scales = [True, False]
separate_dimensions = [True, False]
results_df = pd.DataFrame()
for istandard in standardize:
X_, Y_ = standardize_data(X, Y, istandard)
for iscaler in separate_scales:
for idim in separate_dimensions:
for imethod in methods:
sigma_X, sigma_Y = get_sigma(
X_, Y_,
method=imethod,
per_dimension=idim,
separate_scales=iscaler
)
results_df = results_df.append(pd.DataFrame({
"standardize": [istandard],
"Separate Dimensions": [idim],
"Separate Length Scales": [iscaler],
'Param Estimator': [imethod],
"HSIC": [get_hsic(X_, Y_, 'hsic', sigma_X, sigma_Y)], # Estimate HSIC
"KA": [get_hsic(X_, Y_, 'ka', sigma_X, sigma_Y)], # Estimate KA
"CKA": [get_hsic(X_, Y_, 'cka', sigma_X, sigma_Y)], # Estimate CKA
},index=['Q4']),)
print(results_df.to_markdown())
# -
# ### Verdict
#
# For the distributions, it seemed to be a little more consistent but with higher dimensions and more samples, these estimators start to fail. But then, we still don't have good alternative estimators.
# ## What Now?
#
# I will be looking at the following:
#
# <center>
#
# | | Options |
# | ------------------- | ---------------------------- |
# | Standardize | Yes / No |
# | Parameter Estimator | Mean, Median, Silverman, etc |
# | Center Kernel | Yes / No |
# | Normalized Score | Yes / No |
#
# </center>
| notebooks/1_1D_parameter/1.0_motivation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Image preprocessing**
#
#
# The algorithm should be able to process an image like this:
#
# 
#
# into the MNIST-like image.
# ## **Step-by-step experiments**
# +
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import torch
import torchvision.transforms as T
# -
im = T.Grayscale()(Image.open("image_example.png"))
npim = np.array(im)
ptim = T.ToTensor()(im)
ptim[:4, :5]
mean, std = ptim.mean(), ptim.std()
mean, std
ptim = T.Normalize(ptim.mean(), ptim.std())(ptim)
ptim = ptim.view(*ptim.size()[1:])
fill_val = min(ptim.flatten()).item()
npim.shape
ptim[271:281, 465:475]
im
np.amin(np.where(npim > 0), axis=1)
torch.amin(torch.stack(torch.where(ptim > 0)), dim=1)
top, left = torch.amin(torch.stack(torch.where(ptim > 0)), dim=1)
bottom, right = torch.amax(torch.stack(torch.where(ptim > 0)), dim=1)
plt.imshow(ptim[top:bottom, left:right], cmap='gray')
plt.xticks([])
plt.yticks([])
center = torch.tensor([(top.item() + bottom.item()) // 2, (left.item() + right.item()) // 2])
center
imrad = max(bottom - center[0], right - center[1])
imrad
center + imrad < torch.tensor(ptim.size())
torch.tensor(ptim.size()) - (center + imrad)
# +
if sum(center + imrad < torch.tensor(ptim.size())) < 2 or sum(center - imrad > 0) < 2:
br_offsite = abs(min(torch.tensor(ptim.size()) - (center + imrad)))
tl_offsite = abs(min(center - imrad))
pad = max(br_offsite, tl_offsite)
ptim_2 = T.Pad(padding=pad.item(), fill=fill_val)(ptim)
center += pad
else:
ptim_2 = ptim
plt.imshow(ptim_2, cmap='gray')
plt.xticks([])
plt.yticks([])
# -
ptim.size(), ptim_2.size()
topc, leftc = center - imrad
bottomc, rightc = center + imrad
topc, leftc
ptim_3 = ptim_2[topc:bottomc, leftc:rightc]
plt.imshow(ptim_3, cmap='gray')
plt.xticks([])
plt.yticks([])
ptim_3p = T.GaussianBlur(kernel_size=(7, 7), sigma=(0.1, 7))(ptim_3.unsqueeze(dim=0).unsqueeze(dim=0)).squeeze()
plt.imshow(ptim_3p, cmap='gray')
plt.xticks([])
plt.yticks([])
ptim_4 = T.Resize(size=[26, 26])(ptim_3p.view(1, 1, *ptim_3p.size())).view(26, 26)
ptim_4 = T.Pad(padding=1, fill=fill_val)(ptim_4)
plt.imshow(ptim_4, cmap='gray')
plt.xticks([])
plt.yticks([])
ptim_4.size()
# ## **Complete function**
def prepare_image(im, output_size=28):
"""Prepares an image for training
Parameters
----------
im : PIL image
An input image.
output_size : integer, optional
The size of the output image. Default 28.
"""
# Convert to grayscale PyTorch Tensor
ptim = T.ToTensor()(T.Grayscale()(im))
ptim = ptim.view(*ptim.size()[1:])
back_val = min(ptim.flatten()).item()
# Find the digit in the image
top, left = torch.amin(torch.stack(
torch.where(ptim > back_val)), dim=1)
bottom, right = torch.amax(torch.stack(
torch.where(ptim > back_val)), dim=1)
center = torch.tensor([
(top.item() + bottom.item()) // 2,
(left.item() + right.item()) // 2,
])
imrad = max(bottom - center[0], right - center[1])
# Make a padding if the (center + radius) is biger
# then an input image or (center - radius) < 0.
if (sum(center + imrad < torch.tensor(ptim.size())) < 2
or sum(center - imrad > 0) < 2):
br_offsite = abs(min(torch.tensor(ptim.size()) - (center + imrad)))
tl_offsite = abs(min(center - imrad))
pad = max(br_offsite, tl_offsite)
ptim = T.Pad(padding=pad.item(), fill=back_val)(ptim)
center += pad
# Crop, resize and normalize the image
topc, leftc = center - imrad
bottomc, rightc = center + imrad
ptim = ptim[topc:bottomc, leftc:rightc]
ptim = ptim.view(1, 1, *ptim.size())
ptim = T.Resize(size=[output_size-4, output_size-4])(ptim)
ptim = T.Pad(padding=2, fill=back_val)(ptim)
mean, std = ptim.mean(), ptim.std()
ptim = T.Normalize(mean, std)(ptim)
return ptim
# ## **Exhibition**
# + tags=[]
im1 = Image.open("image_example.png")
im1
# + tags=[]
plt.imshow(prepare_image(im1)[0][0], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.show()
# -
im2 = Image.open("image_example_v2.png")
im2
plt.imshow(prepare_image(im2)[0][0], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.show()
im3 = Image.open("image_example_v3.png")
im3
plt.imshow(prepare_image(im3)[0][0], cmap='gray')
plt.xticks([])
plt.yticks([])
plt.show()
| src/handwritten-digits/Notebooks/image_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # Google Search - Perform search
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Google%20Search/Google_Search_Perform_search.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #googlesearch #snippet
# + [markdown] papermill={} tags=["naas"]
# **Author:** [<NAME>](https://www.linkedin.com/in/ACoAAAJHE7sB5OxuKHuzguZ9L6lfDHqw--cdnJg/)
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Import library
# + papermill={} tags=[]
try:
from googlesearch import search
except:
# !pip install google
from googlesearch import search
# + [markdown] papermill={} tags=[]
# ### Variables
# + papermill={} tags=[]
query = "telsa"
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# Parameters :
#
# - query: This is the text that you want to search for.
#
# - tld: This refers to the top level domain value like co.in or com which will specify which Google website we want to use.
#
# - lang: This parameter stands for language.
#
# - num: This is used to specify the number of results we want.
#
# - start: This is to specify from where to start the results. We should keep it 0 to begin from the very start.
#
# - stop: The last result to retrieve. Use None to keep searching forever.
#
# - pause: This parameter is used to specify the number of seconds to pause between consecutive HTTP requests because if we hit too many requests, Google can block our IP address.
#
# The above function will return a python generator (iterator) which has the search result URLs.
# + [markdown] papermill={} tags=[]
# Source : https://www.studytonight.com/post/how-to-perform-google-search-using-python#
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Display the result of the search
# + papermill={} tags=[]
for i in search(query, tld="co.in", num=30, stop=10, pause=2):
print(i)
| Google Search/Google_Search_Perform_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get Familiar with Python
# ## Functions
# +
# function
def get_weather_info():
return 'Weather is good today!'
def get_weather_info():
a = 2 + 2
# print(a)
return
# -
get_weather_info()
def update_model(model, *args, **kwargs):
print(model, args, kwargs)
update_model('my model')
update_model('my model', 9, 'a', False, sep=',', header=[])
def update_model2(model, *a, **b):
print(model, a, b)
update_model2('my model', 9, 'a', False, sep=',', header=[])
my_lambda_function = lambda x, y, z: round((x / y) ** z, 2)
my_lambda_function
update_model2
my_lambda_function(1,2,3)
(lambda: print('yey'))()
# ## Classes
class Person:
kind = 'Human' # class attributes
def __init__(self, firstname, lastname, birth_year): # the word `self` means nothing. represents an instance of the class.
self.birth_year = birth_year # instance attribute
self.firstname = firstname
self.lastname = lastname
def printname(self):
print(self.firstname, self.lastname)
def get_age(self):
return 2021 - self.birth_year
Person.kind
# will fail without instantiation
Person.birth_year
# instantiation will make instance attributes available
# to instance(copy) of this class
p = Person('Metin', 'Senturk', 1989)
p2 = Person('Kusum', 'Singh', 1995)
p.birth_year, p.firstname, p.lastname
p2.birth_year, p2.firstname, p2.lastname
p.printname(), p2.printname()
p.ssn = '123121234'
p.ssn
p.get_age(), p2.get_age()
def get_age2(year):
return 2021 - year
p.get_age2 = get_age2
p.get_age2(p.birth_year)
# ## Generators
# %time sum(range(100000000))
my_long_list = list(range(100000000))
# 1. Create the definition of generator function
def get_index():
a = 1
yield a # once called, it stops its execution at this point
print('after yield', a)
# return 1
# 2. Create an instance from generator defintion
index_gen = get_index()
# +
# calling with next will fail if generator has no more values
# next(index_gen)
# -
# for loop will understand and not fail if gen has no values
for idx in index_gen:
print(idx)
def get_index2():
a = 1
yield a
a += 1
print('after yield', a)
index_gen = get_index2()
for idx in index_gen:
print(idx)
def get_index3(stop):
a = 1
while True:
if a == stop:
return a
yield a
a += 1
print('after yield', a)
index_gen = get_index3(10)
for idx in index_gen:
print(idx)
# ## Modules and Packages
#
# - Can import a standard module (comes with python)
# - A third party package (you install with pip)
# - A local file on your computer
import pathlib
pathlib
from module1 import greeting
greeting('Ramoji')
from module1 import PERSON1, FolderOperations
my_folder = FolderOperations('My Local Folder')
my_folder
if not my_folder.exists():
my_folder.create()
my_folder.add_file()
| contents/python-warmup/Python Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from IPython import display
# %matplotlib inline
def plot_map():
plt.rcParams["figure.figsize"] = (6,6)
plt.clf();
gray = {}
gray[0]=100
gray[100]=0
gray[-1]=80
m = np.array([gray[pix] for pix in og])
m = np.flipud(m.reshape((4000, 4000)))
plt.imshow(m, cmap='gray');
#plt.imshow(m, origin='lower', cmap='gray');
# -
import rospy
from nav_msgs.srv import GetMap
rospy.init_node('listener', anonymous=True)
rospy.wait_for_service('dynamic_map')
get_map = rospy.ServiceProxy('dynamic_map', GetMap)
data = get_map()
og = data.map.data
plot_map()
plt.axis('equal');
plt.axis([1500,1700,2100,2300]);
import matplotlib.image as mpimg
img=mpimg.imread('my_map.pgm')
plt.imshow(img, cmap='gray');
plt.axis('equal');
plt.axis([1500,1700,2100,2300]);
| Notebooks/slam_tutorials/Plot the Map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + papermill={"duration": 1.531146, "end_time": "2020-11-22T16:07:49.329729", "exception": false, "start_time": "2020-11-22T16:07:47.798583", "status": "completed"} tags=[]
"""
[V6]
Blend 4 Models:
* kibuna-nn-hs-1024-last-train (aka. 2stage-NN, LB: 0.01822)
* simpleNN-oldcv (LB: 0.01836)
* deepinsight-efficientnet-v7-b3-infer (LB: 0.01850)
* deepinsight_resnest_lightning_v2_infer (LB: 0.01854)
Removed for now due to low weights:
* 503-203-tabnet-with-nonscored-features (LB: 0.01836)
* fork-of-2heads-looper-super-puper-markpeng-infer (LB: 0.1836)
"""
kernel_mode = False
import os
import numpy as np
import pandas as pd
import time
import random
import math
import datetime
import pickle
from pickle import dump, load
import glob
from numba import njit
from scipy.optimize import minimize, fsolve
import optuna
import warnings
warnings.filterwarnings('ignore')
import gc
gc.enable()
rand_seed = 1120
search_mode = False
run_submit_script = True
# method = "CV"
method = "scipy_per_target"
# method = "scipy"
# method = "optuna"
study_name = "blend_search_optuna_v6_per_target"
# n_trials = 500
n_trials = 3000
# n_trials = 5000
# + papermill={"duration": 7.831926, "end_time": "2020-11-22T16:07:57.173865", "exception": false, "start_time": "2020-11-22T16:07:49.341939", "status": "completed"} tags=[]
# # !mkdir -p /root/.cache/torch/hub/checkpoints/
# # !cp ../input/gen-efficientnet-pretrained/tf_efficientnet_*.pth /root/.cache/torch/hub/checkpoints/
# # !cp ../input/deepinsight-resnest-v1-resnest50/*.pth /root/.cache/torch/hub/checkpoints/
# # !cp ../input/deepinsight-resnest-v2-resnest50-output/*.pth /root/.cache/torch/hub/checkpoints/
# # !ls -la /root/.cache/torch/hub/checkpoints/
# + papermill={"duration": 2.211524, "end_time": "2020-11-22T16:07:59.432178", "exception": false, "start_time": "2020-11-22T16:07:57.220654", "status": "completed"} tags=[]
# # !cp ../input/kaggle-moa-team/scripts/* .
# # !cp ../input/kaggle-moa-team/blends/*.pkl .
# # !ls -la
# + papermill={"duration": 0.034902, "end_time": "2020-11-22T16:07:59.488095", "exception": false, "start_time": "2020-11-22T16:07:59.453193", "status": "completed"} tags=[]
dataset_folder = "../input/lish-moa" if kernel_mode else "/workspace/Kaggle/MoA/"
# Add your model inference script here
# Tuple Format: (script, oof_filename, output_filename, weight)
model_list = [
("2stageNN_with_ns_oldcv.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/2stageNN_with_ns_oldcv.py",
"../../Github/kaggle_moa_team/oof/oof_2stageNN_ns_oldcv.npy",
"submission_2stageNN_with_ns_oldcv_0.01822.csv",
0.323528084383917),
("script_simpleNN_oldcv.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/script_simpleNN_oldcv.py",
"../../Github/kaggle_moa_team/oof/oof_script_simpleNN_oldcv.npy",
"submission_script_simpleNN_oldcv_0.01836.csv",
0.08786476491118465),
# ("fork-of-2heads-looper-super-puper-markpeng-infer.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/fork-of-2heads-looper-super-puper-markpeng-infer.py",
# "../../Github/kaggle_moa_team/oof/oof_fork-of-2heads-looper-super-puper-markpeng.npy",
# "submission_2heads-looper-super-puper_0.01836.csv",
# 0.018966959973949222),
("deepinsight_efficientnet_lightning_v7_b3_infer.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/deepinsight_efficientnet_lightning_v7_b3_infer.py",
"../../Github/kaggle_moa_team/oof/oof_deepinsight_efficientnet_lightning_v7_b3_0.01850.npy",
"submission_effnet_v7_b3_0.01850.csv",
0.21849845883367852),
# ("script_tabnet_ns_oldcv.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/script_tabnet_ns_oldcv.py",
# "../../Github/kaggle_moa_team/oof/oof_tabnet_ns_oldcv.npy",
# "submission_tabnet_ns_oldcv_0.01836.csv",
# 0.0013224625996093413),
("deepinsight_resnest_lightning_v2_infer.py" if kernel_mode else "../../Github/kaggle_moa_team/scripts/deepinsight_resnest_lightning_v2_infer.py",
"../../Github/kaggle_moa_team/oof/oof_deepinsight_ResNeSt_v2_resnest50_0.01854.npy",
"submission_resnest_v2_0.01854.csv",
0.3704230222796271),
]
model_path = "." if kernel_mode else dataset_folder
# + papermill={"duration": 0.797221, "end_time": "2020-11-22T16:08:00.303937", "exception": false, "start_time": "2020-11-22T16:07:59.506716", "status": "completed"} tags=[]
train_features = pd.read_csv(f"{dataset_folder}/train_features.csv",
engine='c')
train_labels = pd.read_csv(f'{dataset_folder}/train_targets_scored.csv',
engine='c')
train_classes = [c for c in train_labels.columns if c != "sig_id"]
non_control_group_rows = train_features["cp_type"] == "trt_cp"
non_control_group_train_labels = train_labels.loc[
non_control_group_rows, :].copy().reset_index(drop=True)
submission = pd.read_csv(f'{dataset_folder}/sample_submission.csv')
submission.iloc[:, 1:] = 0
# + papermill={"duration": 0.02445, "end_time": "2020-11-22T16:08:00.342614", "exception": false, "start_time": "2020-11-22T16:08:00.318164", "status": "completed"} tags=[]
def mean_logloss(y_pred, y_true):
logloss = (1 - y_true) * np.log(1 - y_pred +
1e-15) + y_true * np.log(y_pred + 1e-15)
return np.nanmean(-logloss)
def save_pickle(obj, folder, name):
dump(obj, open(f"{folder}/{name}.pkl", 'wb'), pickle.HIGHEST_PROTOCOL)
def load_pickle(path):
return load(open(path, 'rb'))
# +
# Reference: https://www.kaggle.com/gogo827jz/optimise-blending-weights-with-bonus-0/notebook
# CPMP's logloss from https://www.kaggle.com/c/lish-moa/discussion/183010
def log_loss_numpy(y_pred, y_true):
y_true_ravel = np.asarray(y_true).ravel()
y_pred = np.asarray(y_pred).ravel()
y_pred = np.clip(y_pred, 1e-15, 1 - 1e-15)
loss = np.where(y_true_ravel == 1, -np.log(y_pred),
-np.log(1 - y_pred))
return loss.mean()
def func_numpy_metric(weights):
oof_blend = np.tensordot(weights, all_oof, axes=((0), (0)))
return log_loss_numpy(oof_blend, y_true)
@njit
def grad_func_jit(weights):
oof_clip = np.minimum(1 - 1e-15, np.maximum(all_oof, 1e-15))
gradients = np.zeros(all_oof.shape[0])
for i in range(all_oof.shape[0]):
a, b, c = y_true, oof_clip[i], np.zeros(
(all_oof.shape[1], all_oof.shape[2]))
for j in range(oof.shape[0]):
if j != i:
c += weights[j] * oof_clip[j]
gradients[i] = -np.mean(
(-a * b + (b**2) * weights[i] + b * c) /
((b**2) *
(weights[i]**2) + 2 * b * c * weights[i] - b * weights[i] +
(c**2) - c))
return gradients
# -
# ## Bayesian Optimization and Sequential Least Squares Programming (SLSQP)
# https://optuna.readthedocs.io/en/stable/reference/generated/optuna.samplers.TPESampler.html#optuna.samplers.TPESampler
#
# https://docs.scipy.org/doc/scipy/reference/optimize.minimize-slsqp.html
def run_inference_scripts(submission, weights=None, target_weights=None):
for i, (script, oof_filename, output_filename, weight) in enumerate(model_list):
print(f"Generating submission file from {script} ......")
infer_start = time.time()
# !python {model_path}/{script}
infer_elapsed = time.time() - infer_start
print(f"Time spent on inference: {infer_elapsed/60:.2f} minutes.")
model_submit = pd.read_csv(output_filename, engine='c')
print(model_submit.head(5))
print(model_submit.shape)
if target_weights is not None:
for j, target in enumerate(train_classes):
print(f"Blending {script} for {target} with weight: {optimized_target_weights[j][i]} ......")
submission.iloc[:, j+1] += model_submit.iloc[:, j+1] * optimized_target_weights[j][i]
elif weights is None:
print(f"Blending {script} with weight: {weight} ......")
submission.iloc[:, 1:] += weight * model_submit.iloc[:, 1:]
else:
print(f"Blending {script} with weight: {weights[i]} ......")
submission.iloc[:, 1:] += weights[i] * model_submit.iloc[:, 1:]
return submission
# + papermill={"duration": 1377.882267, "end_time": "2020-11-22T16:30:58.240001", "exception": false, "start_time": "2020-11-22T16:08:00.357734", "status": "completed"} tags=[]
total_start = time.time()
if not search_mode and run_submit_script:
if method == "scipy_per_target":
weights_path = glob.glob(f'{model_path}/{study_name}_*.pkl')[0]
print(f"Loading target-wise optimized weights from {weights_path} ......")
optimized_target_weights = load_pickle(weights_path)
# For 206 target weights
submission = run_inference_scripts(
submission, target_weights=optimized_target_weights)
else:
submission = run_inference_scripts(submission)
elif search_mode and method == "CV":
y_true = non_control_group_train_labels[train_classes].values
all_oof = np.zeros(
(len(model_list), non_control_group_train_labels.shape[0], 206))
blend_oof = np.zeros((non_control_group_train_labels.shape[0], 206))
print(all_oof.shape)
for i, (script, oof_filename, output_filename,
weight) in enumerate(model_list):
print(f"Loading OOF from {oof_filename} ......")
oof = np.load(f"{dataset_folder}/{oof_filename}")
if oof.shape[0] == 23814:
oof = oof[non_control_group_rows, :]
all_oof[i, :, :] = oof
blend_oof += oof * weight
oof_loss = mean_logloss(oof, y_true)
print(f"OOF Validation Loss of {script}: {oof_loss:.6f}\n")
blend_oof_loss = mean_logloss(blend_oof, y_true)
print(f"Blend OOF Validation Loss: {blend_oof_loss:.6f}\n")
elif search_mode and method == "optuna":
print("[Optuna]")
## Search Best Blend Weights by Optuna ##
model_oofs = []
for i, (script, oof_filename, output_filename,
weight) in enumerate(model_list):
print(f"Loading OOF from {oof_filename} ......")
oof = np.load(f"{dataset_folder}/{oof_filename}")
if oof.shape[0] == 23814:
oof = oof[non_control_group_rows, :]
oof_loss = mean_logloss(
oof, non_control_group_train_labels[train_classes].values)
print(f"OOF Validation Loss of {script}: {oof_loss:.6f}\n")
model_oofs.append(oof)
def objective(trial):
weights = []
for i in range(len(model_list)):
weights.append(trial.suggest_float(f"w{i}", 0, 1.0))
blend = np.zeros(model_oofs[0].shape)
for i in range(len(model_list)):
blend += weights[i] * model_oofs[i]
blend = np.clip(blend, 0, 1.0)
loss = mean_logloss(
blend, non_control_group_train_labels[train_classes].values)
return loss
pruner = optuna.pruners.MedianPruner(
n_startup_trials=5,
n_warmup_steps=0,
interval_steps=1,
)
sampler = optuna.samplers.TPESampler(seed=rand_seed)
study = optuna.create_study(direction="minimize",
pruner=pruner,
sampler=sampler,
study_name=study_name,
storage=f'sqlite:///{study_name}.db',
load_if_exists=True)
study.optimize(objective,
n_trials=n_trials,
timeout=None,
gc_after_trial=True,
n_jobs=-1)
trial = study.best_trial
if run_submit_script:
optimal_weights = []
for i, (script, oof_filename, output_filename,
_) in enumerate(model_list):
optimal_weights.append(trial.params[f"w{i}"])
submission = run_inference_scripts(submission, weights=optimal_weights)
print("\n[Optuna]")
print("Number of finished trials: {}".format(len(study.trials)))
print("Best trial:")
print(" Value: {}".format(trial.value))
print(" Params: ")
for key, value in trial.params.items():
print(" {}: {}".format(key, value))
elif search_mode and method == "scipy":
print("[Scipy SLSQP]")
# Optimise Blending Weights with Bonus
# https://www.kaggle.com/gogo827jz/optimise-blending-weights-with-bonus-0/notebook
model_oofs = []
y_true = non_control_group_train_labels[train_classes].values
all_oof = np.zeros(
(len(model_list), non_control_group_train_labels.shape[0], 206))
print(all_oof.shape)
for i, (script, oof_filename, output_filename,
weight) in enumerate(model_list):
print(f"Loading OOF from {oof_filename} ......")
oof = np.load(f"{dataset_folder}/{oof_filename}")
if oof.shape[0] == 23814:
oof = oof[non_control_group_rows, :]
all_oof[i, :, :] = oof
oof_loss = mean_logloss(oof, y_true)
print(f"OOF Validation Loss of {script}: {oof_loss:.6f}\n")
model_oofs.append(oof)
tol = 1e-10
init_guess = [1 / all_oof.shape[0]] * all_oof.shape[0]
bnds = [(0, 1) for _ in range(all_oof.shape[0])]
cons = {
'type': 'eq',
'fun': lambda x: np.sum(x) - 1,
'jac': lambda x: [1] * len(x)
}
print('Inital Blend OOF:', func_numpy_metric(init_guess))
start_time = time.time()
res_scipy = minimize(
fun=func_numpy_metric,
x0=init_guess,
method='SLSQP',
# jac=grad_func_jit, # grad_func
bounds=bnds,
constraints=cons,
tol=tol)
print("\n[Scipy SLSQP]")
print(
f'[{str(datetime.timedelta(seconds = time.time() - start_time))[2:7]}] Optimised Blend OOF:',
res_scipy.fun)
print(f'Optimised Weights: {res_scipy.x}\n')
if run_submit_script:
submission = run_inference_scripts(submission, weights=res_scipy.x)
# Target-wise Weight Optimization #
elif search_mode and method == "scipy_per_target":
print("[Scipy SLSQP]")
# Optimise Blending Weights with Bonus
# https://www.kaggle.com/gogo827jz/optimise-blending-weights-with-bonus-0/notebook
model_oofs = []
y_true = non_control_group_train_labels[train_classes].values
all_oof = np.zeros(
(len(model_list), non_control_group_train_labels.shape[0], 206))
print(all_oof.shape)
for i, (script, oof_filename, output_filename,
weight) in enumerate(model_list):
print(f"Loading OOF from {oof_filename} ......")
oof = np.load(f"{dataset_folder}/{oof_filename}")
if oof.shape[0] == 23814:
oof = oof[non_control_group_rows, :]
all_oof[i, :, :] = oof
oof_loss = mean_logloss(oof, y_true)
print(f"OOF Validation Loss of {script}: {oof_loss:.6f}\n")
model_oofs.append(oof)
print("\n[Scipy SLSQP Per Target]")
optimized_target_weights = []
for i, target in enumerate(train_classes):
tol = 1e-10
init_guess = [1 / all_oof.shape[0]] * all_oof.shape[0]
bnds = [(0, 1) for _ in range(all_oof.shape[0])]
cons = {
'type': 'eq',
'fun': lambda x: np.sum(x) - 1,
'jac': lambda x: [1] * len(x)
}
def func_numpy_metric_targes(weights):
oof_blend = np.tensordot(weights,
all_oof[:, :, i],
axes=((0), (0)))
return log_loss_numpy(oof_blend, y_true[:, i])
start_time = time.time()
res_scipy = minimize(
fun=func_numpy_metric_targes,
x0=init_guess,
method='SLSQP',
# jac=grad_func_jit, # grad_func
bounds=bnds,
constraints=cons,
tol=tol)
print(
f'[{str(datetime.timedelta(seconds = time.time() - start_time))[2:7]}] ' + \
f'Optimised Blend OOF for {target}:', res_scipy.fun)
print(f'Optimised Weights for {target}: {res_scipy.x}\n')
optimized_target_weights.append(res_scipy.x)
blend_targets_oof = np.zeros(
(non_control_group_train_labels.shape[0], 206))
for i, (script, oof_filename, output_filename,
weight) in enumerate(model_list):
print(f"Loading OOF from {oof_filename} ......")
oof = np.load(f"{dataset_folder}/{oof_filename}")
if oof.shape[0] == 23814:
oof = oof[non_control_group_rows, :]
for j in range(206):
blend_targets_oof[:,
j] += oof[:, j] * optimized_target_weights[j][i]
oof_loss = mean_logloss(oof, y_true)
print(f"OOF Validation Loss of {script}: {oof_loss:.6f}\n")
blend_targets_oof_loss = mean_logloss(blend_targets_oof, y_true)
print(
f"Blend Target-Wise OOF Validation Loss: {blend_targets_oof_loss:.6f}\n"
)
# Save optimized weights per target
save_pickle(optimized_target_weights, model_path,
f"{study_name}_{blend_targets_oof_loss}")
if run_submit_script:
# For 206 target weights
submission = run_inference_scripts(
submission, target_weights=optimized_target_weights)
# + papermill={"duration": 0.268265, "end_time": "2020-11-22T16:30:58.782670", "exception": false, "start_time": "2020-11-22T16:30:58.514405", "status": "completed"} tags=[]
total_elapsed = time.time() - total_start
print(f"Total time spent: {total_elapsed/60:.2f} minutes.")
# +
# [V6 - without TabNet, 2heads]
# Total time spent: 0.68 minutes.
# Blend Target-Wise OOF Validation Loss: 0.015044
# -
# +
# [V6 - without TabNet, 2heads]
# [Optuna]
# Number of finished trials: 3000
# Best trial:
# Value: 0.015171999561900233
# Params:
# w0: 0.323528084383917
# w1: 0.08786476491118465
# w2: 0.21849845883367852
# w3: 0.3704230222796271
# [Scipy SLSQP]
# [00:22] Optimised Blend OOF: 0.015172004593585666
# Optimised Weights: [0.32020133 0.09043987 0.22122948 0.36812932]
# +
# [V6 - without TabNet]
# [Optuna]
# Number of finished trials: 3000
# Best trial:
# Value: 0.015172424601530761
# Params:
# w0: 0.3138176484100186
# w1: 0.07850519440561339
# w2: 0.0007183363099561991
# w3: 0.23849563017967007
# w4: 0.3694870328388392
# [Scipy SLSQP]
# [00:21] Optimised Blend OOF: 0.015172004898867827
# Optimised Weights: [0.32045559 0.09026525 0. 0.22069638 0.36858278]
# +
# [V6]
# [Optuna]
# Number of finished trials: 5000
# Best trial:
# Value: 0.015173437622007157
# Params:
# w0: 0.30923325055652684
# w1: 0.09831493504786226
# w2: 0.018966959973949222
# w3: 0.19863369862866234
# w4: 0.0013224625996093413
# w5: 0.3728865483320761
# [Scipy SLSQP]
# [00:36] Optimised Blend OOF: 0.015172005464591968
# Optimised Weights: [3.20472642e-01 9.01191588e-02 1.78893358e-18 2.20448482e-01
# 3.27971157e-18 3.68959717e-01]
# -
# +
# [V5]
# Number of finished trials: 3000
# Best trial:
# Value: 0.015344701181290615
# Params:
# w0: 0.5141433844379889
# w1: 0.11747776562133813
# w2: 0.3668324643717302
# [00:14] Optimised Blend OOF: 0.015344695215068541
# Optimised Weights: [0.51922623 0.11292509 0.36784869]
# -
# +
# [V4]
# [Optuna]
# Number of finished trials: 3000
# Best trial:
# Value: 0.015331901615194453
# Params:
# w0: 0.4505928450756189
# w1: 0.13010257032841785
# w2: 0.06308933354044946
# w3: 0.35639153615958885
#
# [Scipy]
# [00:23] Optimised Blend OOF: 0.015331777381591449
# Optimised Weights: [0.44090106 0.14508641 0.05945655 0.35455598]
# -
# +
# [V3]
# improving-mark-s-2-heads-model-infer
# Number of finished trials: 3000
# Best trial:
# Value: 0.01515466145873492
# Params:
# w0: 0.0002980690037490555
# w1: 0.29771381784976886
# w2: 0.1569191862042946
# w3: 0.18156875605872544
# w4: 0.36371774630338105
# +
# [V3]
# fork-of-2heads-looper-super-puper-markpeng-infer
# Number of finished trials: 3000
# Best trial:
# Value: 0.015170138066049686
# Params:
# w0: 0.00019903389488299251
# w1: 0.3853752127955825
# w2: 0.015968332256452233
# w3: 0.22945916769823432
# w4: 0.3711290150522236
# -
if search_mode and method == "scipy_per_target":
# OOF scores per target
target_oof_losses = []
for i, target in enumerate(train_classes):
print(target)
# print(y_true[:, i])
oof_loss = mean_logloss(blend_targets_oof[:, i], y_true[:, i])
target_oof_losses.append(oof_loss)
print(f"Blend OOF Validation Loss of {target}: {oof_loss:.6f}\n")
target_loss_df = pd.DataFrame(
data={
"target": train_classes,
"oof_logloss": target_oof_losses
},
columns=["target", "oof_logloss"
]).sort_values(by="oof_logloss",
ascending=False).reset_index(drop=True)
print(target_loss_df)
# + papermill={"duration": 0.294364, "end_time": "2020-11-22T16:31:00.767175", "exception": false, "start_time": "2020-11-22T16:31:00.472811", "status": "completed"} tags=[]
if run_submit_script:
print(submission.shape)
print(submission)
submission.to_csv('submission.csv', index=False)
# + [markdown] papermill={"duration": 0.239792, "end_time": "2020-11-22T16:31:03.876454", "exception": false, "start_time": "2020-11-22T16:31:03.636662", "status": "completed"} tags=[]
# ## EOF
# + papermill={"duration": 1.595866, "end_time": "2020-11-22T16:31:05.717490", "exception": false, "start_time": "2020-11-22T16:31:04.121624", "status": "completed"} tags=[]
if kernel_mode:
# !rm ./*.py
# !ls -la
# + papermill={"duration": 0.243405, "end_time": "2020-11-22T16:31:06.199770", "exception": false, "start_time": "2020-11-22T16:31:05.956365", "status": "completed"} tags=[]
| blends/blend_search_optuna_v6_per_target.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Dependencies
import pandas as pd
import os
import sqlalchemy
from sqlalchemy import create_engine
from sqlalchemy.ext.declarative import declarative_base
Base = declarative_base()
from sqlalchemy import Column, Integer, String, Float, BigInteger
from sqlalchemy.orm import Session
#Cleaned dataframe paths from dataframes cleaned in Part 1
cleaned_measurements_path = os.path.join("Resources", "cleaned_hawaii_measurements.csv")
cleaned_stations_path = os.path.join("Resources", "cleaned_hawaii_stations.csv")
#Check Dataframe 1
cleaned_measurements_df = pd.read_csv(cleaned_measurements_path)
cleaned_measurements_df.head()
#Check Dataframe 2
cleaned_stations_df = pd.read_csv(cleaned_stations_path)
cleaned_stations_df.head()
#Create engine , connect to engine, set base to declarative_base
engine = create_engine('sqlite:///Resources/hawaii.sqlite')
conn = engine.connect()
Base = declarative_base()
# +
#Create the two tables in the database
class Measurements(Base):
__tablename__ = 'measurements_table'
id = Column(Integer, primary_key=True)
station = Column(String(500))
date = Column(String(500))
prcp = Column(Float(10))
tobs = Column(Float(10))
class Stations(Base):
__tablename__ = 'stations_table'
id = Column(Integer, primary_key=True)
station = Column(String(500))
name = Column(String(500))
latitude = Column(Float(15))
longitude = Column(Float(15))
elevation = Column(Float(15))
# +
#Create metadata and session instances
Base.metadata.create_all(engine)
session = Session(bind=engine)
#Create variables to store all data from a single column in the dataframes we cleaned
stations = cleaned_measurements_df['station']
dates = cleaned_measurements_df['date']
prcps = cleaned_measurements_df['prcp']
tobs = cleaned_measurements_df['tobs']
#Check values for consistancy (column lengths should be the same)
print(type(tobs[0]))
print(type(stations))
print(len(tobs))
print(len(stations))
#Ids represents the row number in the dataframe, since all columns are the same length we can arbitrarily assign it to station
ids = len(cleaned_measurements_df['station'])
#Add each row using a for loop that loops through all rows and uses session.add() to add each row individually
for x in range(0, ids):
session.add(Measurements(station=stations[x], date=dates[x], prcp=prcps[x], tobs=tobs[x]))
session.commit()
# +
#Same method to add stations table to database
stations = cleaned_stations_df['station']
names = cleaned_stations_df['name']
latitudes = cleaned_stations_df['latitude']
longitudes = cleaned_stations_df['longitude']
elevations = cleaned_stations_df['elevation']
ids = len(cleaned_stations_df['station'])
for x in range(0, ids):
session.add(Stations(station=stations[x], name=names[x], latitude=latitudes[x], longitude=longitudes[x], elevation=elevations[x]))
session.commit()
# +
#Test query database table
measurements_list = session.query(Measurements)
for measurement in measurements_list:
print(measurement.tobs)
# -
| database_engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem?
# solve this?:
# $$ \frac{acc_{direction}}{\sin(\theta+\Delta\theta)} = \frac{acc_{reverse}}{\sin(\theta-\Delta\theta)}$$
#
# - vi denne lighed hvor vi prøver at gøre delta theta så lille som mulight ved at difference nedenfor er mindre end (0.000001).
# - begge accelerationer, og theta er normalfordelinger med en sigma, mens deltatheta er ukendt.
# # Code as is (more or less)
# +
import numpy as np
r = np.random
r.seed(42)
#N_exp =1000
optimize_max_steps=1000
print_steps = optimize_max_steps/10
for iexp in range(N_exp):
# Generate random experiment?
a_norm = r.normal(1.5, 0.1)
a_rev = r.normal(1.4, 0.1)
# Why is theta different.
# isn't theta (the inclination of the table the same)
# and we try to find a Delta theta that will align it better
# (ie. correct for the accellation difference)?
theta_norm = r.normal(13.4, 0.2)
theta_rev = r.normal(13.9,0.3)
diff = 1.0
# These are in radian ie. sin(3.14) = 0, sin(3.14/2 = 1)
dtheta= 1
ddtheta = 0.03
N=0
diff_new = 1
while (diff_new > 1e-6):
# calc both sides of equation
norm = a_norm/np.sin(theta_norm+dtheta)
rev = a_rev/np.sin(theta_rev-dtheta)
# find diference
diff_new = abs(norm-rev)
# find ratio of difference compared to original distance
# Quite unorthodox but seems to work :)
ratio = diff_new/diff
# subtract a smaller and smaller amount
# Note since diff_abs is always positive you only subtract values from it
# Since sinus is cyclical it works in this case but would not generalize
dtheta -= ddtheta*ratio
#diff = diff_new
if iexp == 0 and N%print_steps == 0:
print(f'Difference = {diff_new}, dtheta= {dtheta}, iteration = {N}')
N += 1
if N==optimize_max_steps:
break
# -
np.sin(np.pi/2)
# # Emil notes
# The "proper" way to solve optimization is to frase it as a minimization problem
#
# $$ \frac{acc_{direction}}{\sin(\theta+\Delta\theta)} = \frac{acc_{reverse}}{\sin(\theta-\Delta\theta)}$$
# - becomes this
# $$ f(\Delta\theta) = \frac{acc_{direction}}{\sin(\theta+\Delta\theta)} - \frac{acc_{reverse}}{\sin(\theta-\Delta\theta)} = 0$$
# - will be fullfiled if we minimize
# $$ min(f(\Delta\theta)^2)_{\Delta\theta}$$
# - then you can use standard methods such as
# - Newton rapton: https://en.wikipedia.org/wiki/Newton's_method
# - gradient decent: https://en.wikipedia.org/wiki/Gradient_descent
#
#
# # Emils meta notes
# - You probably should only one $\theta$ (and not both $\theta_{norm}$ and $\theta_{reverse}$) (you only have 1 table)
# - I am not sure you are solving the right problem. Ie. I suspect it would be more around hypothesis testing so given different accelations generate some distributions and do calculus / simulations with the distributions
#
# (you seem to be solving the same problem a 1000 times with different random generated values. but I could be wrong :)
# - suspected real approach
# - 1. generate 1000 data points for each distribution
# - 2. Calculate the means and std_of_means
# - 3. Given these calculate the possible ranges for $\Delta\theta$ once
| kjertan_stats/.ipynb_checkpoints/Kjertans_problems-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework: Sentiment Analysis of Reddit Stories By Title
#
# ## The Problem
#
# In this assignment you're tasked with performing a sentiment analysis on Reddit story titles within a sub-reddit. The goal is to provide a overall "gist" as to whether that subreddit is a positive or negative one.
#
# If you are unfamiliar with Reddit, here are some API urls to a few sub-reddits:
#
# - `worldnews` URL: https://www.reddit.com/r/worldnews/top.json
# - `writingprompts` URL: https://www.reddit.com/r/writingprompts/top.json
# - `todayilearned` URL: https://www.reddit.com/r/todayilearned/top.json
# - `explainlikeimfive` URL: https://www.reddit.com/r/explainlikeimfive/top.json
# - `politics` URL: https://www.reddit.com/r/politics/top.json
#
# From these URLs you should be able to figure out the pattern URL for any given subreddit name such as `news`, or `ama`. Somewhere within the bowles of the API output, you can see the title of each story.
#
# Use IPython interact to provide a list of subreddits to choose from (a list will be provided, but you are free to add your own). When an item from the list is selected:
#
# - Use the Reddit API to get the titles of the top 25 stories in that subreddit
# - Combine the titles from all 25 stories into a single paragraph of text
# - Pass the text to the sentiment analysis API we used in the lab. It is important to send all titles to the sentiment analysis with a single request. If you don't the serive will lock us out.
# - Print the overall sentiment, each probability (pos, neg, neutral) and the title of each story in the subreddit.
#
#
#
#
# HINTS and ADVICE:
#
# - Take a bottom up approach; work on the pieces, then assemble them.
# - Start with sentiment.
# - Get that working as a function similar to the lab. You can copy the code from the lab.
# - Make sure you can output the overall sentiment, and probababilities for positive, negative and neutral
# - Next work on fetching the Reddit story titles from the API
# - The Reddit api requires you to use a unique `User-Agent` key in the headers of the `get()` request. The value of this key should be `ist256.lesson10.homework:v1.0 (by /u/netid)` where `netid` is your netid, of course.
# - Make sure to put a `limit` key on the query string with a value of `25`
# - Pick a single subreddit like `https://www.reddit.com/r/worldnews/top.json` and work with that until you can figure out how to fetch the stories.
# - Once you've got the stories, figure out how to just get the titles form the stories.
# - With everything working, refactor into a function which inputs a subreddit name and outputs the story titles as a list.
# - Once you have the story titles and sentiment, its time build the main program
# - DO NOT get sentiment for each title. If you do this the sentiment API will block us. Instead combine all the titles into a single paragraph with `S.join()`
# - Once it all works, feel free to add your own subreddits to the list!
#
#
#
#
#
# + [markdown] label="problem_analysis_cell"
# ## Part 1: Problem Analysis
#
# Inputs:
#
# ```
# TODO: Inputs
# ```
#
# Outputs:
#
# ```
# TODO: Outputs
# ```
#
# Algorithm (Steps in Program):
#
# ```
# TODO:Steps Here
#
# ```
# -
# ## Part 2: Code Solution
#
# You may write your code in several cells, but place the complete, final working copy of your code solution within this single cell below. Only the within this cell will be considered your solution. Any imports or user-defined functions should be copied into this cell.
# + label="code_solution_cell"
subreddits=['ama','aww','news','worldnews', 'politics','todayilearned','explainlikeimfive','writingprompts','upliftingnews','wholesomememes','freecompliments','happy','financialadvice','breadit']
subreddits.sort()
# Step 2: Write code here
# + [markdown] label="homework_questions_cell"
# ## Part 3: Questions
#
# 1. What happens to this program when you do not have connectivity to the Internet? How can this code be modified to correct the issue?
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 2. Most of the subreddits come back with a neutral sentiment score. Does this surprise you? Explain your answer.
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
# 3. In what ways could we improve this program to provide more accurate sentiment?
#
# `--== Double-Click and Write Your Answer Below This Line ==--`
#
#
#
# + [markdown] label="reflection_cell"
# ## Part 4: Reflection
#
# Reflect upon your experience completing this assignment. This should be a personal narrative, in your own voice, and cite specifics relevant to the activity as to help the grader understand how you arrived at the code you submitted. Things to consider touching upon: Elaborate on the process itself. Did your original problem analysis work as designed? How many iterations did you go through before you arrived at the solution? Where did you struggle along the way and how did you overcome it? What did you learn from completing the assignment? What do you need to work on to get better? What was most valuable and least valuable about this exercise? Do you have any suggestions for improvements?
#
# To make a good reflection, you should journal your thoughts, questions and comments while you complete the exercise.
#
# Keep your response to between 100 and 250 words.
#
# `--== Double-Click and Write Your Reflection Below Here ==--`
#
# -
# run this code to turn in your work!
from coursetools.submission import Submission
Submission().submit()
| content/lessons/10-HTTP/HW-HTTP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot
# ensure the plots are inside this notebook, not an external window
# %matplotlib inline
#
# +
img = cv2.imread('images/hand.png',0)
# threshold
blur = cv2.GaussianBlur(img,(5,5),0)
ret3,th3 = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
# invert color
th4 = cv2.bitwise_not(th3)
# plot different versions of the hand
f, axarr = matplotlib.pyplot.subplots(1,3, figsize = (18,5))
axarr[0].imshow(img, cmap='Greys_r', aspect='auto', interpolation='nearest')
axarr[1].imshow(th3, cmap='Greys_r', aspect='auto', interpolation='nearest')
axarr[2].imshow(th4, cmap='Greys_r', aspect='auto', interpolation='nearest')
# +
_,contours,_ = cv2.findContours(th4, 2, 1)
cnt = contours[0]
M = cv2.moments(cnt)
hull = cv2.convexHull(cnt, returnPoints = False)
matplotlib.pyplot.imshow(th4)
# -
matplotlib.pyplot.imshow(th5, cmap='Greys_r')
https://www.pyimagesearch.com/2016/04/11/finding-extreme-points-in-contours-with-opencv/
https://arxiv.org/pdf/1312.7560.pdf
| Samples/.ipynb_checkpoints/ObjectContours-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Measure of 'Happiness'
# Note: Instead of using subjective estimate, I changed the data proprecessing - dealing with missing value part using the average value.
# +
import os
import time
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn import metrics
from datetime import datetime
import matplotlib.pyplot as plt
from sklearn.metrics import roc_auc_score, roc_curve, mean_squared_error,mean_absolute_error, f1_score
import lightgbm as lgb
import xgboost as xgb
from sklearn.ensemble import RandomForestRegressor as rfr
from sklearn.ensemble import ExtraTreesRegressor as etr
from sklearn.linear_model import BayesianRidge as br
from sklearn.ensemble import GradientBoostingRegressor as gbr
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import LinearRegression as lr
from sklearn.linear_model import ElasticNet as en
from sklearn.kernel_ridge import KernelRidge as kr
from sklearn.model_selection import KFold, StratifiedKFold,GroupKFold, RepeatedKFold
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn import preprocessing
import logging
import warnings
warnings.filterwarnings('ignore')
# -
# ### Dataset
# +
train = pd.read_csv("/Users/Melodie/Downloads/2021Spring/Study/DataWhale/April_Ensembled_Learning/Notes_Ensemble_Learning/Data/case_1/train.csv", parse_dates=['survey_time'],encoding='latin-1')
test = pd.read_csv("/Users/Melodie/Downloads/2021Spring/Study/DataWhale/April_Ensembled_Learning/Notes_Ensemble_Learning/Data/case_1/test.csv", parse_dates=['survey_time'],encoding='latin-1') #latin-1向下兼容ASCII
#Remove the 'happiness' variable
train = train[train["happiness"]!=-8].reset_index(drop=True)
train_data_copy = train.copy()
#'Happiness' variable
target_col = "happiness"
target = train_data_copy[target_col]
del train_data_copy[target_col]
data = pd.concat([train_data_copy,test],axis=0,ignore_index=True)
# -
# EDA 'Happiness' variable
train.happiness.describe()
data.columns
# ## Data Preprocessing
# There are some negative values. We wants to figure out how many such negative values in each row.
# +
def getres1(row):
return len([x for x in row.values if type(x)==int and x<0])
def getres2(row):
return len([x for x in row.values if type(x)==int and x==-8])
def getres3(row):
return len([x for x in row.values if type(x)==int and x==-1])
def getres4(row):
return len([x for x in row.values if type(x)==int and x==-2])
def getres5(row):
return len([x for x in row.values if type(x)==int and x==-3])
data['neg1'] = data[data.columns].apply(lambda row:getres1(row),axis=1)
data['neg2'] = data[data.columns].apply(lambda row:getres2(row),axis=1)
data['neg3'] = data[data.columns].apply(lambda row:getres3(row),axis=1)
data['neg4'] = data[data.columns].apply(lambda row:getres4(row),axis=1)
data['neg5'] = data[data.columns].apply(lambda row:getres5(row),axis=1)
#When there are more than 20 negative values in one observation, we set it to be 20 (Why?)
data.loc[data['neg1']>20,'neg1'] = 20
# -
# Filling the Missing values
# +
data['work_status'] = data['work_status'].fillna(0)
data['work_yr'] = data['work_yr'].fillna(0)
data['work_manage'] = data['work_manage'].fillna(0)
data['work_type'] = data['work_type'].fillna(0)
data['edu_yr'] = data['edu_yr'].fillna(0)
data['edu_status'] = data['edu_status'].fillna(0)
data['s_work_type'] = data['s_work_type'].fillna(0)
data['s_work_status'] = data['s_work_status'].fillna(0)
data['s_political'] = data['s_political'].fillna(0)
data['s_hukou'] = data['s_hukou'].fillna(0)
data['s_income'] = data['s_income'].fillna(0)
data['s_birth'] = data['s_birth'].fillna(0)
data['s_edu'] = data['s_edu'].fillna(0)
data['s_work_exper'] = data['s_work_exper'].fillna(0)
data['minor_child'] = data['minor_child'].fillna(0)
data['marital_now'] = data['marital_now'].fillna(0)
data['marital_1st'] = data['marital_1st'].fillna(0)
data['social_neighbor']=data['social_neighbor'].fillna(0)
data['social_friend']=data['social_friend'].fillna(0)
data['hukou_loc']=data['hukou_loc'].fillna(1)
#Use mean to fill
data['family_income']=data['family_income'].fillna(66365)
# +
data['survey_time'] = pd.to_datetime(data['survey_time'], format='%Y-%m-%d',errors='coerce')
data['survey_time'] = data['survey_time'].dt.year
#The age while doing the survey
data['age'] = data['survey_time']-data['birth']
#Categorize the age to be 6 categories: 0-17/18-26/27-34/....
bins = [0,17,26,34,50,63,100]
data['age_bin'] = pd.cut(data['age'], bins, labels=[0,1,2,3,4,5])
# -
# Note: when dealing with the missing value, here used some subjective standards
data['height_cm'].min()
# +
#Religion, if less than 0 - no religion
data.loc[data['religion']<0,'religion'] = 1
data.loc[data['religion_freq']<0,'religion_freq'] = 1
#Education level - if less than 0, set to be middle school. (But there are people especially cohort born before 1950, it is likely that their highest education level is primary school)
data.loc[data['edu']<0,'edu'] = 4 #Middle school
data.loc[data['edu_status']<0,'edu_status'] = 0
data.loc[data['edu_yr']<0,'edu_yr'] = 0
#Why there are negative value for income? 'Prefer not to say' or 'no income'?
data.loc[data['income']<0,'income'] = 0
data.loc[data['political']<0,'political'] = 1 #Default not in the party
data.loc[(data['weight_jin']<=80)&(data['height_cm']>=160),'weight_jin']= data['weight_jin']*2
# data.loc[data['weight_jin']<=60,'weight_jin']= data['weight_jin']*2 #Data integrity
# Minimum value of height_cm is 100, which makes sense
# data.loc[data['height_cm']<150,'height_cm'] = 150
#Health
data.loc[data['health']<0,'health'] = 3
data.loc[data['health_problem']<0,'health_problem'] = 3
data.loc[data['depression']<0,'depression'] = 3
data.loc[data['media_1']<0,'media_1'] = 1
data.loc[data['media_2']<0,'media_2'] = 1
data.loc[data['media_3']<0,'media_3'] = 1
data.loc[data['media_4']<0,'media_4'] = 1
data.loc[data['media_5']<0,'media_5'] = 1
data.loc[data['media_6']<0,'media_6'] = 1
data.loc[data['leisure_1']<0,'leisure_1'] = 1
data.loc[data['leisure_2']<0,'leisure_2'] = 5
data.loc[data['leisure_3']<0,'leisure_3'] = 3
# -
# Dealing with missing value in categorical data - mode
# +
data.loc[data['leisure_4']<0,'leisure_4'] = data['leisure_4'].mode()
data.loc[data['leisure_5']<0,'leisure_5'] = data['leisure_5'].mode()
data.loc[data['leisure_6']<0,'leisure_6'] = data['leisure_6'].mode()
data.loc[data['leisure_7']<0,'leisure_7'] = data['leisure_7'].mode()
data.loc[data['leisure_8']<0,'leisure_8'] = data['leisure_8'].mode()
data.loc[data['leisure_9']<0,'leisure_9'] = data['leisure_9'].mode()
data.loc[data['leisure_10']<0,'leisure_10'] = data['leisure_10'].mode()
data.loc[data['leisure_11']<0,'leisure_11'] = data['leisure_11'].mode()
data.loc[data['leisure_12']<0,'leisure_12'] = data['leisure_12'].mode()
data.loc[data['socialize']<0,'socialize'] = data['socialize'].mode()
data.loc[data['relax']<0,'relax'] = data['relax'].mode()
data.loc[data['learn']<0,'learn'] = data['learn'].mode()
data.loc[data['social_neighbor']<0,'social_neighbor'] = 0
data.loc[data['social_friend']<0,'social_friend'] = 0
data.loc[data['socia_outing']<0,'socia_outing'] = 1
data.loc[data['neighbor_familiarity']<0,'neighbor_familiarity']= 4
data.loc[data['equity']<0,'equity'] = 4
data.loc[data['class_10_before']<0,'class_10_before'] = 3
data.loc[data['class']<0,'class'] = 5
data.loc[data['class_10_after']<0,'class_10_after'] = 5
data.loc[data['class_14']<0,'class_14'] = 2
data.loc[data['work_status']<0,'work_status'] = 0
data.loc[data['work_yr']<0,'work_yr'] = 0
data.loc[data['work_manage']<0,'work_manage'] = 0
data.loc[data['work_type']<0,'work_type'] = 0
data.loc[data['insur_1']<0,'insur_1'] = 1
data.loc[data['insur_2']<0,'insur_2'] = 1
data.loc[data['insur_3']<0,'insur_3'] = 1
data.loc[data['insur_4']<0,'insur_4'] = 1
data.loc[data['insur_1']==0,'insur_1'] = 0
data.loc[data['insur_2']==0,'insur_2'] = 0
data.loc[data['insur_3']==0,'insur_3'] = 0
data.loc[data['insur_4']==0,'insur_4'] = 0
# -
# Dealing with missing continuous data - mean
data[data['family_income'] >=0]['family_income'].mean()
# +
data.loc[data['family_income']<0,'family_income'] = data[data['family_income'] >=0]['family_income'].mean()
data.loc[data['family_m']<0,'family_m'] = 1
data.loc[data['family_status']<0,'family_status'] = data['family_status'].mode()
data.loc[data['house']<0,'house'] = 1
data.loc[data['car']<0,'car'] = 0
data.loc[data['car']==2,'car'] = 0
data.loc[data['son']<0,'son'] = 0
data.loc[data['daughter']<0,'daughter'] = 0
data.loc[data['minor_child']<0,'minor_child'] = 0
data.loc[data['marital_1st']<0,'marital_1st'] = 0
data.loc[data['marital_now']<0,'marital_now'] = 0
data.loc[data['s_birth']<0,'s_birth'] = 0
data.loc[data['s_edu']<0,'s_edu'] = 0
data.loc[data['s_political']<0,'s_political'] = 0
data.loc[data['s_hukou']<0,'s_hukou'] = 0
data.loc[data['s_income']<0,'s_income'] = 0
data.loc[data['s_work_type']<0,'s_work_type'] = 0
data.loc[data['s_work_status']<0,'s_work_status'] = 0
data.loc[data['s_work_exper']<0,'s_work_exper'] = 0
data.loc[data['f_birth']<0,'f_birth'] = data[data['f_birth'] >=0]['f_birth'].mean()
data.loc[data['f_edu']<0,'f_edu'] = 1
data.loc[data['f_political']<0,'f_political'] = 1
data.loc[data['f_work_14']<0,'f_work_14'] = 2
data.loc[data['m_birth']<0,'m_birth'] = data[data['m_birth'] >=0]['m_birth'].mean()
data.loc[data['m_edu']<0,'m_edu'] = 1
data.loc[data['m_political']<0,'m_political'] = 1
data.loc[data['m_work_14']<0,'m_work_14'] = 2
data.loc[data['status_peer']<0,'status_peer'] = 2
data.loc[data['status_3_before']<0,'status_3_before'] = 2
data.loc[data['view']<0,'view'] = data[data['view'] >=0]['view'].mean()
data.loc[data['inc_ability']<=0,'inc_ability']= data[data['inc_ability'] >=0]['inc_ability'].mean()
data.loc[data['inc_exp']<=0,'inc_exp']= data[data['inc_exp'] >=0]['inc_exp'].mean()
for i in range(1,9+1):
data.loc[data['public_service_'+str(i)]<0,'public_service_'+str(i)] = data['public_service_'+str(i)].dropna().mode().values
for i in range(1,13+1):
data.loc[data['trust_'+str(i)]<0,'trust_'+str(i)] = data['trust_'+str(i)].dropna().mode().values
# -
# ## Data Augmentation
# +
data['marital_1stbir'] = data['marital_1st'] - data['birth']
data['marital_nowtbir'] = data['marital_now'] - data['birth']
data['mar'] = data['marital_nowtbir'] - data['marital_1stbir']
data['marital_sbir'] = data['marital_now']-data['s_birth']
data['age_'] = data['marital_nowtbir'] - data['marital_sbir']
data['income/s_income'] = data['income']/(data['s_income']+1)
data['income+s_income'] = data['income']+(data['s_income']+1)
data['income/family_income'] = data['income']/(data['family_income']+1)
data['all_income/family_income'] = (data['income']+data['s_income'])/(data['family_income']+1)
data['income/inc_exp'] = data['income']/(data['inc_exp']+1)
data['family_income/m'] = data['family_income']/(data['family_m']+0.01)
data['income/m'] = data['income']/(data['family_m']+0.01)
data['income/floor_area'] = data['income']/(data['floor_area']+0.01)
data['all_income/floor_area'] = (data['income']+data['s_income'])/(data['floor_area']+0.01)
data['family_income/floor_area'] = data['family_income']/(data['floor_area']+0.01)
data['floor_area/m'] = data['floor_area']/(data['family_m']+0.01)
data['class_10_diff'] = (data['class_10_after'] - data['class'])
data['class_diff'] = data['class'] - data['class_10_before']
data['class_14_diff'] = data['class'] - data['class_14']
leisure_fea_lis = ['leisure_'+str(i) for i in range(1,13)]
data['leisure_sum'] = data[leisure_fea_lis].sum(axis=1) #skew
public_service_fea_lis = ['public_service_'+str(i) for i in range(1,10)]
data['public_service_sum'] = data[public_service_fea_lis].sum(axis=1) #skew
trust_fea_lis = ['trust_'+str(i) for i in range(1,14)]
data['trust_sum'] = data[trust_fea_lis].sum(axis=1) #skew
data['province_income_mean'] = data.groupby(['province'])['income'].transform('mean').values
data['province_family_income_mean'] = data.groupby(['province'])['family_income'].transform('mean').values
data['province_equity_mean'] = data.groupby(['province'])['equity'].transform('mean').values
data['province_depression_mean'] = data.groupby(['province'])['depression'].transform('mean').values
data['province_floor_area_mean'] = data.groupby(['province'])['floor_area'].transform('mean').values
data['province_health_mean'] = data.groupby(['province'])['health'].transform('mean').values
data['province_class_10_diff_mean'] = data.groupby(['province'])['class_10_diff'].transform('mean').values
data['province_class_mean'] = data.groupby(['province'])['class'].transform('mean').values
data['province_health_problem_mean'] = data.groupby(['province'])['health_problem'].transform('mean').values
data['province_family_status_mean'] = data.groupby(['province'])['family_status'].transform('mean').values
data['province_leisure_sum_mean'] = data.groupby(['province'])['leisure_sum'].transform('mean').values
data['province_public_service_sum_mean'] = data.groupby(['province'])['public_service_sum'].transform('mean').values
data['province_trust_sum_mean'] = data.groupby(['province'])['trust_sum'].transform('mean').values
#city mean 181+13=194
data['city_income_mean'] = data.groupby(['city'])['income'].transform('mean').values #按照city分组
data['city_family_income_mean'] = data.groupby(['city'])['family_income'].transform('mean').values
data['city_equity_mean'] = data.groupby(['city'])['equity'].transform('mean').values
data['city_depression_mean'] = data.groupby(['city'])['depression'].transform('mean').values
data['city_floor_area_mean'] = data.groupby(['city'])['floor_area'].transform('mean').values
data['city_health_mean'] = data.groupby(['city'])['health'].transform('mean').values
data['city_class_10_diff_mean'] = data.groupby(['city'])['class_10_diff'].transform('mean').values
data['city_class_mean'] = data.groupby(['city'])['class'].transform('mean').values
data['city_health_problem_mean'] = data.groupby(['city'])['health_problem'].transform('mean').values
data['city_family_status_mean'] = data.groupby(['city'])['family_status'].transform('mean').values
data['city_leisure_sum_mean'] = data.groupby(['city'])['leisure_sum'].transform('mean').values
data['city_public_service_sum_mean'] = data.groupby(['city'])['public_service_sum'].transform('mean').values
data['city_trust_sum_mean'] = data.groupby(['city'])['trust_sum'].transform('mean').values
data['county_income_mean'] = data.groupby(['county'])['income'].transform('mean').values
data['county_family_income_mean'] = data.groupby(['county'])['family_income'].transform('mean').values
data['county_equity_mean'] = data.groupby(['county'])['equity'].transform('mean').values
data['county_depression_mean'] = data.groupby(['county'])['depression'].transform('mean').values
data['county_floor_area_mean'] = data.groupby(['county'])['floor_area'].transform('mean').values
data['county_health_mean'] = data.groupby(['county'])['health'].transform('mean').values
data['county_class_10_diff_mean'] = data.groupby(['county'])['class_10_diff'].transform('mean').values
data['county_class_mean'] = data.groupby(['county'])['class'].transform('mean').values
data['county_health_problem_mean'] = data.groupby(['county'])['health_problem'].transform('mean').values
data['county_family_status_mean'] = data.groupby(['county'])['family_status'].transform('mean').values
data['county_leisure_sum_mean'] = data.groupby(['county'])['leisure_sum'].transform('mean').values
data['county_public_service_sum_mean'] = data.groupby(['county'])['public_service_sum'].transform('mean').values
data['county_trust_sum_mean'] = data.groupby(['county'])['trust_sum'].transform('mean').values
data['income/province'] = data['income']/(data['province_income_mean'])
data['family_income/province'] = data['family_income']/(data['province_family_income_mean'])
data['equity/province'] = data['equity']/(data['province_equity_mean'])
data['depression/province'] = data['depression']/(data['province_depression_mean'])
data['floor_area/province'] = data['floor_area']/(data['province_floor_area_mean'])
data['health/province'] = data['health']/(data['province_health_mean'])
data['class_10_diff/province'] = data['class_10_diff']/(data['province_class_10_diff_mean'])
data['class/province'] = data['class']/(data['province_class_mean'])
data['health_problem/province'] = data['health_problem']/(data['province_health_problem_mean'])
data['family_status/province'] = data['family_status']/(data['province_family_status_mean'])
data['leisure_sum/province'] = data['leisure_sum']/(data['province_leisure_sum_mean'])
data['public_service_sum/province'] = data['public_service_sum']/(data['province_public_service_sum_mean'])
data['trust_sum/province'] = data['trust_sum']/(data['province_trust_sum_mean']+1)
data['income/city'] = data['income']/(data['city_income_mean'])
data['family_income/city'] = data['family_income']/(data['city_family_income_mean'])
data['equity/city'] = data['equity']/(data['city_equity_mean'])
data['depression/city'] = data['depression']/(data['city_depression_mean'])
data['floor_area/city'] = data['floor_area']/(data['city_floor_area_mean'])
data['health/city'] = data['health']/(data['city_health_mean'])
data['class_10_diff/city'] = data['class_10_diff']/(data['city_class_10_diff_mean'])
data['class/city'] = data['class']/(data['city_class_mean'])
data['health_problem/city'] = data['health_problem']/(data['city_health_problem_mean'])
data['family_status/city'] = data['family_status']/(data['city_family_status_mean'])
data['leisure_sum/city'] = data['leisure_sum']/(data['city_leisure_sum_mean'])
data['public_service_sum/city'] = data['public_service_sum']/(data['city_public_service_sum_mean'])
data['trust_sum/city'] = data['trust_sum']/(data['city_trust_sum_mean'])
data['income/county'] = data['income']/(data['county_income_mean'])
data['family_income/county'] = data['family_income']/(data['county_family_income_mean'])
data['equity/county'] = data['equity']/(data['county_equity_mean'])
data['depression/county'] = data['depression']/(data['county_depression_mean'])
data['floor_area/county'] = data['floor_area']/(data['county_floor_area_mean'])
data['health/county'] = data['health']/(data['county_health_mean'])
data['class_10_diff/county'] = data['class_10_diff']/(data['county_class_10_diff_mean'])
data['class/county'] = data['class']/(data['county_class_mean'])
data['health_problem/county'] = data['health_problem']/(data['county_health_problem_mean'])
data['family_status/county'] = data['family_status']/(data['county_family_status_mean'])
data['leisure_sum/county'] = data['leisure_sum']/(data['county_leisure_sum_mean'])
data['public_service_sum/county'] = data['public_service_sum']/(data['county_public_service_sum_mean'])
data['trust_sum/county'] = data['trust_sum']/(data['county_trust_sum_mean'])
data['age_income_mean'] = data.groupby(['age'])['income'].transform('mean').values
data['age_family_income_mean'] = data.groupby(['age'])['family_income'].transform('mean').values
data['age_equity_mean'] = data.groupby(['age'])['equity'].transform('mean').values
data['age_depression_mean'] = data.groupby(['age'])['depression'].transform('mean').values
data['age_floor_area_mean'] = data.groupby(['age'])['floor_area'].transform('mean').values
data['age_health_mean'] = data.groupby(['age'])['health'].transform('mean').values
data['age_class_10_diff_mean'] = data.groupby(['age'])['class_10_diff'].transform('mean').values
data['age_class_mean'] = data.groupby(['age'])['class'].transform('mean').values
data['age_health_problem_mean'] = data.groupby(['age'])['health_problem'].transform('mean').values
data['age_family_status_mean'] = data.groupby(['age'])['family_status'].transform('mean').values
data['age_leisure_sum_mean'] = data.groupby(['age'])['leisure_sum'].transform('mean').values
data['age_public_service_sum_mean'] = data.groupby(['age'])['public_service_sum'].transform('mean').values
data['age_trust_sum_mean'] = data.groupby(['age'])['trust_sum'].transform('mean').values
data['income/age'] = data['income']/(data['age_income_mean'])
data['family_income/age'] = data['family_income']/(data['age_family_income_mean'])
data['equity/age'] = data['equity']/(data['age_equity_mean'])
data['depression/age'] = data['depression']/(data['age_depression_mean'])
data['floor_area/age'] = data['floor_area']/(data['age_floor_area_mean'])
data['health/age'] = data['health']/(data['age_health_mean'])
data['class_10_diff/age'] = data['class_10_diff']/(data['age_class_10_diff_mean'])
data['class/age'] = data['class']/(data['age_class_mean'])
data['health_problem/age'] = data['health_problem']/(data['age_health_problem_mean'])
data['family_status/age'] = data['family_status']/(data['age_family_status_mean'])
data['leisure_sum/age'] = data['leisure_sum']/(data['age_leisure_sum_mean'])
data['public_service_sum/age'] = data['public_service_sum']/(data['age_public_service_sum_mean'])
data['trust_sum/age'] = data['trust_sum']/(data['age_trust_sum_mean'])
# -
print('shape',data.shape)
data.head()
del_list=['id','survey_time','edu_other','invest_other','property_other','join_party','province','city','county']
use_feature = [clo for clo in data.columns if clo not in del_list]
data.fillna(0,inplace=True)
train_shape = train.shape[0]
features = data[use_feature].columns
X_train_263 = data[:train_shape][use_feature].values
y_train = target
X_test_263 = data[train_shape:][use_feature].values
X_train_263.shape
# 49 important features
imp_fea_49 = ['equity','depression','health','class','family_status','health_problem','class_10_after',
'equity/province','equity/city','equity/county',
'depression/province','depression/city','depression/county',
'health/province','health/city','health/county',
'class/province','class/city','class/county',
'family_status/province','family_status/city','family_status/county',
'family_income/province','family_income/city','family_income/county',
'floor_area/province','floor_area/city','floor_area/county',
'leisure_sum/province','leisure_sum/city','leisure_sum/county',
'public_service_sum/province','public_service_sum/city','public_service_sum/county',
'trust_sum/province','trust_sum/city','trust_sum/county',
'income/m','public_service_sum','class_diff','status_3_before','age_income_mean','age_floor_area_mean',
'weight_jin','height_cm',
'health/age','depression/age','equity/age','leisure_sum/age'
]
train_shape = train.shape[0]
X_train_49 = data[:train_shape][imp_fea_49].values
X_test_49 = data[train_shape:][imp_fea_49].values
X_train_49.shape
# +
cat_fea = ['survey_type','gender','nationality','edu_status','political','hukou','hukou_loc','work_exper','work_status','work_type',
'work_manage','marital','s_political','s_hukou','s_work_exper','s_work_status','s_work_type','f_political','f_work_14',
'm_political','m_work_14']
noc_fea = [clo for clo in use_feature if clo not in cat_fea]
onehot_data = data[cat_fea].values
enc = preprocessing.OneHotEncoder(categories = 'auto')
oh_data=enc.fit_transform(onehot_data).toarray()
oh_data.shape
X_train_oh = oh_data[:train_shape,:]
X_test_oh = oh_data[train_shape:,:]
X_train_oh.shape
X_train_383 = np.column_stack([data[:train_shape][noc_fea].values,X_train_oh])
X_test_383 = np.column_stack([data[train_shape:][noc_fea].values,X_test_oh])
X_train_383.shape
# -
# ## Feature Engineering - 263 features
# ### LightGBM - 5 fold CV
# +
lgb_263_param = {
'num_leaves': 7,
'min_data_in_leaf': 20,
'objective':'regression',
'max_depth': -1,
'learning_rate': 0.003,
"boosting": "gbdt",
"feature_fraction": 0.18,
"bagging_freq": 1,
"bagging_fraction": 0.55,
"bagging_seed": 14,
"metric": 'mse',
"lambda_l1": 0.1,
"lambda_l2": 0.2,
"verbosity": -1}
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=4)
oof_lgb_263 = np.zeros(len(X_train_263))
predictions_lgb_263 = np.zeros(len(X_test_263))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_263, y_train)):
print("fold n°{}".format(fold_+1))
trn_data = lgb.Dataset(X_train_263[trn_idx], y_train[trn_idx])
val_data = lgb.Dataset(X_train_263[val_idx], y_train[val_idx])#train:val=4:1
num_round = 10000
lgb_263 = lgb.train(lgb_263_param, trn_data, num_round, valid_sets = [trn_data, val_data], verbose_eval=500, early_stopping_rounds = 800)
oof_lgb_263[val_idx] = lgb_263.predict(X_train_263[val_idx], num_iteration=lgb_263.best_iteration)
predictions_lgb_263 += lgb_263.predict(X_test_263, num_iteration=lgb_263.best_iteration) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_lgb_263, target)))
# -
# Feature Importance
# +
pd.set_option('display.max_columns', 1000)
pd.set_option('display.max_rows', 1000)
pd.set_option('max_colwidth',100)
df = pd.DataFrame(data[use_feature].columns.tolist(), columns=['feature'])
df['importance']=list(lgb_263.feature_importance())
df = df.sort_values(by='importance',ascending=False)
plt.figure(figsize=(14,28))
sns.barplot(x="importance", y="feature", data=df.head(50))
plt.title('Features importance (averaged/folds)')
plt.tight_layout()
# -
# ### XGBoot
# +
xgb_263_params = {'eta': 0.02,
'max_depth': 6,
'min_child_weight':3,
'gamma':0,
'subsample': 0.7,
'colsample_bytree': 0.3,
'lambda':2,
'objective': 'reg:linear',
'eval_metric': 'rmse',
'silent': True,
'nthread': -1}
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=2019)
oof_xgb_263 = np.zeros(len(X_train_263))
predictions_xgb_263 = np.zeros(len(X_test_263))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_263, y_train)):
print("fold n°{}".format(fold_+1))
trn_data = xgb.DMatrix(X_train_263[trn_idx], y_train[trn_idx])
val_data = xgb.DMatrix(X_train_263[val_idx], y_train[val_idx])
watchlist = [(trn_data, 'train'), (val_data, 'valid_data')]
xgb_263 = xgb.train(dtrain=trn_data, num_boost_round=3000, evals=watchlist, early_stopping_rounds=600, verbose_eval=500, params=xgb_263_params)
oof_xgb_263[val_idx] = xgb_263.predict(xgb.DMatrix(X_train_263[val_idx]), ntree_limit=xgb_263.best_ntree_limit)
predictions_xgb_263 += xgb_263.predict(xgb.DMatrix(X_test_263), ntree_limit=xgb_263.best_ntree_limit) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_xgb_263, target)))
# -
# ### Random Forest
# +
folds = KFold(n_splits=5, shuffle=True, random_state=2019)
oof_rfr_263 = np.zeros(len(X_train_263))
predictions_rfr_263 = np.zeros(len(X_test_263))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_263, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_263[trn_idx]
tr_y = y_train[trn_idx]
rfr_263 = rfr(n_estimators=1600,max_depth=9, min_samples_leaf=9, min_weight_fraction_leaf=0.0,
max_features=0.25,verbose=1,n_jobs=-1)
#verbose = 0
#verbose = 1
#verbose = 2
rfr_263.fit(tr_x,tr_y)
oof_rfr_263[val_idx] = rfr_263.predict(X_train_263[val_idx])
predictions_rfr_263 += rfr_263.predict(X_test_263) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_rfr_263, target)))
# -
# ### GradientBoostingRegressor
# +
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=2018)
oof_gbr_263 = np.zeros(train_shape)
predictions_gbr_263 = np.zeros(len(X_test_263))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_263, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_263[trn_idx]
tr_y = y_train[trn_idx]
gbr_263 = gbr(n_estimators=400, learning_rate=0.01,subsample=0.65,max_depth=7, min_samples_leaf=20,
max_features=0.22,verbose=1)
gbr_263.fit(tr_x,tr_y)
oof_gbr_263[val_idx] = gbr_263.predict(X_train_263[val_idx])
predictions_gbr_263 += gbr_263.predict(X_test_263) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_gbr_263, target)))
# -
# ### ExtraTreesRegressor
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_etr_263 = np.zeros(train_shape)
predictions_etr_263 = np.zeros(len(X_test_263))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_263, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_263[trn_idx]
tr_y = y_train[trn_idx]
etr_263 = etr(n_estimators=1000,max_depth=8, min_samples_leaf=12, min_weight_fraction_leaf=0.0,
max_features=0.4,verbose=1,n_jobs=-1)
etr_263.fit(tr_x,tr_y)
oof_etr_263[val_idx] = etr_263.predict(X_train_263[val_idx])
predictions_etr_263 += etr_263.predict(X_test_263) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_etr_263, target)))
# -
# ## Modeling & Feature Engineering - 49 features
# ### Light GBM
# +
lgb_49_param = {
'num_leaves': 9,
'min_data_in_leaf': 23,
'objective':'regression',
'max_depth': -1,
'learning_rate': 0.002,
"boosting": "gbdt",
"feature_fraction": 0.45,
"bagging_freq": 1,
"bagging_fraction": 0.65,
"bagging_seed": 15,
"metric": 'mse',
"lambda_l2": 0.2,
"verbosity": -1}
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=9)
oof_lgb_49 = np.zeros(len(X_train_49))
predictions_lgb_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
trn_data = lgb.Dataset(X_train_49[trn_idx], y_train[trn_idx])
val_data = lgb.Dataset(X_train_49[val_idx], y_train[val_idx])
num_round = 12000
lgb_49 = lgb.train(lgb_49_param, trn_data, num_round, valid_sets = [trn_data, val_data], verbose_eval=1000, early_stopping_rounds = 1000)
oof_lgb_49[val_idx] = lgb_49.predict(X_train_49[val_idx], num_iteration=lgb_49.best_iteration)
predictions_lgb_49 += lgb_49.predict(X_test_49, num_iteration=lgb_49.best_iteration) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_lgb_49, target)))
# -
# ### XGBoost
# +
xgb_49_params = {'eta': 0.02,
'max_depth': 5,
'min_child_weight':3,
'gamma':0,
'subsample': 0.7,
'colsample_bytree': 0.35,
'lambda':2,
'objective': 'reg:linear',
'eval_metric': 'rmse',
'silent': True,
'nthread': -1}
folds = KFold(n_splits=5, shuffle=True, random_state=2019)
oof_xgb_49 = np.zeros(len(X_train_49))
predictions_xgb_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
trn_data = xgb.DMatrix(X_train_49[trn_idx], y_train[trn_idx])
val_data = xgb.DMatrix(X_train_49[val_idx], y_train[val_idx])
watchlist = [(trn_data, 'train'), (val_data, 'valid_data')]
xgb_49 = xgb.train(dtrain=trn_data, num_boost_round=3000, evals=watchlist, early_stopping_rounds=600, verbose_eval=500, params=xgb_49_params)
oof_xgb_49[val_idx] = xgb_49.predict(xgb.DMatrix(X_train_49[val_idx]), ntree_limit=xgb_49.best_ntree_limit)
predictions_xgb_49 += xgb_49.predict(xgb.DMatrix(X_test_49), ntree_limit=xgb_49.best_ntree_limit) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_xgb_49, target)))
# -
# ### GradientBoostingRegressor
# +
folds = StratifiedKFold(n_splits=5, shuffle=True, random_state=2018)
oof_gbr_49 = np.zeros(train_shape)
predictions_gbr_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_49[trn_idx]
tr_y = y_train[trn_idx]
gbr_49 = gbr(n_estimators=600, learning_rate=0.01,subsample=0.65,max_depth=6, min_samples_leaf=20,
max_features=0.35,verbose=1)
gbr_49.fit(tr_x,tr_y)
oof_gbr_49[val_idx] = gbr_49.predict(X_train_49[val_idx])
predictions_gbr_49 += gbr_49.predict(X_test_49) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_gbr_49, target)))
# +
train_stack3 = np.vstack([oof_lgb_49,oof_xgb_49,oof_gbr_49]).transpose()
test_stack3 = np.vstack([predictions_lgb_49, predictions_xgb_49,predictions_gbr_49]).transpose()
#
folds_stack = RepeatedKFold(n_splits=5, n_repeats=2, random_state=7)
oof_stack3 = np.zeros(train_stack3.shape[0])
predictions_lr3 = np.zeros(test_stack3.shape[0])
for fold_, (trn_idx, val_idx) in enumerate(folds_stack.split(train_stack3,target)):
print("fold {}".format(fold_))
trn_data, trn_y = train_stack3[trn_idx], target.iloc[trn_idx].values
val_data, val_y = train_stack3[val_idx], target.iloc[val_idx].values
#Kernel Ridge Regression
lr3 = kr()
lr3.fit(trn_data, trn_y)
oof_stack3[val_idx] = lr3.predict(val_data)
predictions_lr3 += lr3.predict(test_stack3) / 10
mean_squared_error(target.values, oof_stack3)
# -
# KernelRidge
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_kr_49 = np.zeros(train_shape)
predictions_kr_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_49[trn_idx]
tr_y = y_train[trn_idx]
kr_49 = kr()
kr_49.fit(tr_x,tr_y)
oof_kr_49[val_idx] = kr_49.predict(X_train_49[val_idx])
predictions_kr_49 += kr_49.predict(X_test_49) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_kr_49, target)))
# -
# Ridge
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_ridge_49 = np.zeros(train_shape)
predictions_ridge_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_49[trn_idx]
tr_y = y_train[trn_idx]
ridge_49 = Ridge(alpha=6)
ridge_49.fit(tr_x,tr_y)
oof_ridge_49[val_idx] = ridge_49.predict(X_train_49[val_idx])
predictions_ridge_49 += ridge_49.predict(X_test_49) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_ridge_49, target)))
# -
# BayesianRidge
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_br_49 = np.zeros(train_shape)
predictions_br_49 = np.zeros(len(X_test_49))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_49[trn_idx]
tr_y = y_train[trn_idx]
br_49 = br()
br_49.fit(tr_x,tr_y)
oof_br_49[val_idx] = br_49.predict(X_train_49[val_idx])
predictions_br_49 += br_49.predict(X_test_49) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_br_49, target)))
# -
# ElasticNet
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_en_49 = np.zeros(train_shape)
predictions_en_49 = np.zeros(len(X_test_49))
#
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_49, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_49[trn_idx]
tr_y = y_train[trn_idx]
en_49 = en(alpha=1.0,l1_ratio=0.05)
en_49.fit(tr_x,tr_y)
oof_en_49[val_idx] = en_49.predict(X_train_49[val_idx])
predictions_en_49 += en_49.predict(X_test_49) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_en_49, target)))
# +
train_stack2 = np.vstack([oof_lgb_263,oof_xgb_263,oof_gbr_263,oof_rfr_263,oof_etr_263]).transpose()
test_stack2 = np.vstack([predictions_lgb_263, predictions_xgb_263,predictions_gbr_263,predictions_rfr_263,predictions_etr_263]).transpose()
folds_stack = RepeatedKFold(n_splits=5, n_repeats=2, random_state=7)
oof_stack2 = np.zeros(train_stack2.shape[0])
predictions_lr2 = np.zeros(test_stack2.shape[0])
for fold_, (trn_idx, val_idx) in enumerate(folds_stack.split(train_stack2,target)):
print("fold {}".format(fold_))
trn_data, trn_y = train_stack2[trn_idx], target.iloc[trn_idx].values
val_data, val_y = train_stack2[val_idx], target.iloc[val_idx].values
#Kernel Ridge Regression
lr2 = kr()
lr2.fit(trn_data, trn_y)
oof_stack2[val_idx] = lr2.predict(val_data)
predictions_lr2 += lr2.predict(test_stack2) / 10
mean_squared_error(target.values, oof_stack2)
# +
train_stack4 = np.vstack([oof_br_49,oof_kr_49,oof_en_49,oof_ridge_49]).transpose()
test_stack4 = np.vstack([predictions_br_49, predictions_kr_49,predictions_en_49,predictions_ridge_49]).transpose()
folds_stack = RepeatedKFold(n_splits=5, n_repeats=2, random_state=7)
oof_stack4 = np.zeros(train_stack4.shape[0])
predictions_lr4 = np.zeros(test_stack4.shape[0])
for fold_, (trn_idx, val_idx) in enumerate(folds_stack.split(train_stack4,target)):
print("fold {}".format(fold_))
trn_data, trn_y = train_stack4[trn_idx], target.iloc[trn_idx].values
val_data, val_y = train_stack4[val_idx], target.iloc[val_idx].values
#LinearRegression
lr4 = lr()
lr4.fit(trn_data, trn_y)
oof_stack4[val_idx] = lr4.predict(val_data)
predictions_lr4 += lr4.predict(test_stack1) / 10
mean_squared_error(target.values, oof_stack4)
# -
# ### Modeling and Feature Engineering - 383 Features
# Kernel Ridge Regression
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_kr_383 = np.zeros(train_shape)
predictions_kr_383 = np.zeros(len(X_test_383))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_383, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_383[trn_idx]
tr_y = y_train[trn_idx]
kr_383 = kr()
kr_383.fit(tr_x,tr_y)
oof_kr_383[val_idx] = kr_383.predict(X_train_383[val_idx])
predictions_kr_383 += kr_383.predict(X_test_383) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_kr_383, target)))
# -
# Ridge Regression
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_ridge_383 = np.zeros(train_shape)
predictions_ridge_383 = np.zeros(len(X_test_383))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_383, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_383[trn_idx]
tr_y = y_train[trn_idx]
ridge_383 = Ridge(alpha=1200)
ridge_383.fit(tr_x,tr_y)
oof_ridge_383[val_idx] = ridge_383.predict(X_train_383[val_idx])
predictions_ridge_383 += ridge_383.predict(X_test_383) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_ridge_383, target)))
# -
# ElasticNet
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_en_383 = np.zeros(train_shape)
predictions_en_383 = np.zeros(len(X_test_383))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_383, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_383[trn_idx]
tr_y = y_train[trn_idx]
en_383 = en(alpha=1.0,l1_ratio=0.06)
en_383.fit(tr_x,tr_y)
oof_en_383[val_idx] = en_383.predict(X_train_383[val_idx])
predictions_en_383 += en_383.predict(X_test_383) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_en_383, target)))
# -
# BayesianRidge
# +
folds = KFold(n_splits=5, shuffle=True, random_state=13)
oof_br_383 = np.zeros(train_shape)
predictions_br_383 = np.zeros(len(X_test_383))
for fold_, (trn_idx, val_idx) in enumerate(folds.split(X_train_383, y_train)):
print("fold n°{}".format(fold_+1))
tr_x = X_train_383[trn_idx]
tr_y = y_train[trn_idx]
br_383 = br()
br_383.fit(tr_x,tr_y)
oof_br_383[val_idx] = br_383.predict(X_train_383[val_idx])
predictions_br_383 += br_383.predict(X_test_383) / folds.n_splits
print("CV score: {:<8.8f}".format(mean_squared_error(oof_br_383, target)))
# +
train_stack1 = np.vstack([oof_br_383,oof_kr_383,oof_en_383,oof_ridge_383]).transpose()
test_stack1 = np.vstack([predictions_br_383, predictions_kr_383,predictions_en_383,predictions_ridge_383]).transpose()
folds_stack = RepeatedKFold(n_splits=5, n_repeats=2, random_state=7)
oof_stack1 = np.zeros(train_stack1.shape[0])
predictions_lr1 = np.zeros(test_stack1.shape[0])
for fold_, (trn_idx, val_idx) in enumerate(folds_stack.split(train_stack1,target)):
print("fold {}".format(fold_))
trn_data, trn_y = train_stack1[trn_idx], target.iloc[trn_idx].values
val_data, val_y = train_stack1[val_idx], target.iloc[val_idx].values
# LinearRegression简单的线性回归
lr1 = lr()
lr1.fit(trn_data, trn_y)
oof_stack1[val_idx] = lr1.predict(val_data)
predictions_lr1 += lr1.predict(test_stack1) / 10
mean_squared_error(target.values, oof_stack1)
# -
# ## Ensemble Model
# +
train_stack5 = np.vstack([oof_stack1,oof_stack2,oof_stack3,oof_stack4]).transpose()
test_stack5 = np.vstack([predictions_lr1, predictions_lr2,predictions_lr3,predictions_lr4]).transpose()
folds_stack = RepeatedKFold(n_splits=5, n_repeats=2, random_state=7)
oof_stack5 = np.zeros(train_stack5.shape[0])
predictions_lr5= np.zeros(test_stack5.shape[0])
for fold_, (trn_idx, val_idx) in enumerate(folds_stack.split(train_stack5,target)):
print("fold {}".format(fold_))
trn_data, trn_y = train_stack5[trn_idx], target.iloc[trn_idx].values
val_data, val_y = train_stack5[val_idx], target.iloc[val_idx].values
#LinearRegression
lr5 = lr()
lr5.fit(trn_data, trn_y)
oof_stack5[val_idx] = lr5.predict(val_data)
predictions_lr5 += lr5.predict(test_stack5) / 10
mean_squared_error(target.values, oof_stack5)
# -
# ### Submission
# +
submit_example = pd.read_csv('/Users/Melodie/Downloads/2021Spring/Study/DataWhale/April_Ensembled_Learning/Notes_Ensemble_Learning/Data/case_1/submit_example.csv',sep=',',encoding='latin-1')
submit_example['happiness'] = predictions_lr5
submit_example.happiness.describe()
# +
submit_example.loc[submit_example['happiness']>4.96,'happiness']= 5
submit_example.loc[submit_example['happiness']<=1.04,'happiness']= 1
submit_example.loc[(submit_example['happiness']>1.96)&(submit_example['happiness']<2.04),'happiness']= 2
submit_example.to_csv("/Users/Melodie/Downloads/2021Spring/Study/DataWhale/April_Ensembled_Learning/Notes_Ensemble_Learning/Data/case_1/submision.csv",index=False)
submit_example.happiness.describe()
| Task14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.00977, "end_time": "2021-04-20T21:01:31.533883", "exception": false, "start_time": "2021-04-20T21:01:31.524113", "status": "completed"} tags=[]
# # Experiments: LSOT in high dimensions
# Entropy and L2 regularization approaches.
# DenseICNNs for fitting transport maps between gaussians.
# **GPU-only implementation.**
# + papermill={"duration": 1.218422, "end_time": "2021-04-20T21:01:32.761829", "exception": false, "start_time": "2021-04-20T21:01:31.543407", "status": "completed"} tags=[]
import os, sys
sys.path.append("..")
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import torch
import gc
from sklearn.decomposition import PCA
from src import distributions
from src import map_benchmark
from src.icnn import DenseICNN
from src.tools import train_identity_map, unfreeze
from src.plotters import plot_benchmark_emb, plot_W2, plot_benchmark_metrics
import src.map_benchmark as mbm
from src.metrics import score_fitted_maps, score_baseline_maps, metrics_to_dict
import pandas as pd
from tqdm import tqdm
from IPython.display import clear_output
# + [markdown] papermill={"duration": 0.021505, "end_time": "2021-04-20T21:01:32.798067", "exception": false, "start_time": "2021-04-20T21:01:32.776562", "status": "completed"} tags=[]
# ## Changable Config
# + papermill={"duration": 0.015288, "end_time": "2021-04-20T21:01:32.821957", "exception": false, "start_time": "2021-04-20T21:01:32.806669", "status": "completed"} tags=["parameters"]
DIM = 16
BATCH_SIZE = 1024
GPU_DEVICE = 3
# + [markdown] papermill={"duration": 0.015242, "end_time": "2021-04-20T21:01:32.875603", "exception": false, "start_time": "2021-04-20T21:01:32.860361", "status": "completed"} tags=[]
# ## Hardcoded Config
# + papermill={"duration": 0.052556, "end_time": "2021-04-20T21:01:32.940671", "exception": false, "start_time": "2021-04-20T21:01:32.888115", "status": "completed"} tags=[]
BENCHMARK = 'Mix3toMix10'
MAX_ITER = 100001
LR = 1e-3
EPSILON = 3 * 1e-2
REG = 'L2' #Ent
D_HYPERPARAMS = {
'dim' : DIM,
'rank' : 1,
'hidden_layer_sizes' : [max(2*DIM, 64), max(2*DIM, 64), max(DIM, 32)],
'strong_convexity' : 1e-4
}
OUTPUT_PATH = '../logs/' + BENCHMARK
if not os.path.exists(OUTPUT_PATH):
os.makedirs(OUTPUT_PATH)
METHOD = 'LS'
assert torch.cuda.is_available()
torch.cuda.set_device(GPU_DEVICE)
torch.manual_seed(0x000000); np.random.seed(0x000000)
# + [markdown] papermill={"duration": 0.008239, "end_time": "2021-04-20T21:01:32.959506", "exception": false, "start_time": "2021-04-20T21:01:32.951267", "status": "completed"} tags=[]
# ## Benchmark Setup
# + papermill={"duration": 2.671162, "end_time": "2021-04-20T21:01:35.638453", "exception": false, "start_time": "2021-04-20T21:01:32.967291", "status": "completed"} tags=[]
benchmark = mbm.Mix3ToMix10Benchmark(DIM)
emb_X = PCA(n_components=2).fit(benchmark.input_sampler.sample(2**14).cpu().detach().numpy())
emb_Y = PCA(n_components=2).fit(benchmark.output_sampler.sample(2**14).cpu().detach().numpy())
# + [markdown] papermill={"duration": 0.013472, "end_time": "2021-04-20T21:01:35.666206", "exception": false, "start_time": "2021-04-20T21:01:35.652734", "status": "completed"} tags=[]
# ## Potentials Setup
# + papermill={"duration": 2.295732, "end_time": "2021-04-20T21:01:37.975498", "exception": false, "start_time": "2021-04-20T21:01:35.679766", "status": "completed"} tags=[]
D = DenseICNN(**D_HYPERPARAMS).cuda()
D_conj = DenseICNN(**D_HYPERPARAMS).cuda()
pretrain_sampler = distributions.StandardNormalSampler(dim=DIM)
print('Pretraining identity potential. Final MSE:', train_identity_map(D, pretrain_sampler, convex=False, blow=3))
D_conj.load_state_dict(D.state_dict())
del pretrain_sampler
# + [markdown] papermill={"duration": 0.012747, "end_time": "2021-04-20T21:01:38.000121", "exception": false, "start_time": "2021-04-20T21:01:37.987374", "status": "completed"} tags=[]
# ## Main training
# + papermill={"duration": 0.018241, "end_time": "2021-04-20T21:01:38.028604", "exception": false, "start_time": "2021-04-20T21:01:38.010363", "status": "completed"} tags=[]
D_opt = torch.optim.Adam(D.parameters(), lr=LR)
D_conj_opt = torch.optim.Adam(D_conj.parameters(), lr=LR)
# + papermill={"duration": 0.300194, "end_time": "2021-04-20T21:01:38.337515", "exception": false, "start_time": "2021-04-20T21:01:38.037321", "status": "completed"} tags=[]
W2_history = []
metrics = dict(L2_UVP_fwd=[], cos_fwd=[], L2_UVP_inv=[], cos_inv=[])
baselines = {
baseline : metrics_to_dict(*score_baseline_maps(benchmark, baseline))
for baseline in ['identity', 'constant', 'linear']
}
# + papermill={"duration": 2146.75902, "end_time": "2021-04-20T21:37:25.127337", "exception": false, "start_time": "2021-04-20T21:01:38.368317", "status": "completed"} tags=[]
for iteration in tqdm(range(MAX_ITER)):
unfreeze(D); unfreeze(D_conj)
X = benchmark.input_sampler.sample(BATCH_SIZE)
X.requires_grad_(True)
Y = benchmark.output_sampler.sample(BATCH_SIZE)
Y.requires_grad_(True)
# Negative Wasserstein distance
D_opt.zero_grad(); D_conj_opt.zero_grad()
D_X = D(X); D_conj_Y = D_conj(Y)
W_loss = (D_X + D_conj_Y).mean()
# Non-backpropagated part
with torch.no_grad():
W_loss_nograd = -.5 * ((X ** 2).sum(dim=1).mean() + (Y ** 2).sum(dim=1).mean())
if REG == 'L2':
reg_loss = ((torch.relu((X * Y).sum(dim=1) - D_X.flatten() - D_conj_Y.flatten())) ** 2).mean() / (4 * EPSILON)
W_loss += reg_loss
elif REG == 'Ent':
reg_loss = EPSILON * torch.exp(((X * Y).sum(dim=1) - D_X.flatten() - D_conj_Y.flatten()) / EPSILON).mean()
W_loss += reg_loss
else:
raise Exception('Unknown Reg')
W2_history.append(-W_loss.item() - W_loss_nograd.item())
W_loss.backward()
D_opt.step(); D_conj_opt.step();
if iteration % 100 == 0:
L2_UVP_fwd, cos_fwd, L2_UVP_inv, cos_inv = score_fitted_maps(benchmark, D, D_conj)
metrics['L2_UVP_fwd'].append(L2_UVP_fwd); metrics['cos_fwd'].append(cos_fwd)
metrics['L2_UVP_inv'].append(L2_UVP_inv); metrics['cos_inv'].append(cos_inv)
if iteration % 1000 == 0:
clear_output(wait=True)
print("Iteration", iteration)
fig, axes = plot_benchmark_emb(benchmark, emb_X, emb_Y, D, D_conj)
plt.show(fig); plt.close(fig)
fig, axes = plot_W2(benchmark, W2_history)
plt.show(fig); plt.close(fig)
fig, axes = plot_benchmark_metrics(benchmark, metrics, baselines)
plt.show(fig); plt.close(fig)
# + papermill={"duration": 0.049133, "end_time": "2021-04-20T21:37:25.203331", "exception": false, "start_time": "2021-04-20T21:37:25.154198", "status": "completed"} tags=[]
print(f'Logging Metrics Evolution to {OUTPUT_PATH}')
results = pd.DataFrame(metrics)
results.to_csv(os.path.join(OUTPUT_PATH, f'{DIM}-{METHOD}.csv'))
torch.save(D.state_dict(), os.path.join(OUTPUT_PATH, f'{DIM}-{METHOD}.pt'))
torch.save(D_conj.state_dict(), os.path.join(OUTPUT_PATH, f'{DIM}-{METHOD}_conj.pt'))
print('Done!')
# + papermill={"duration": 0.020276, "end_time": "2021-04-20T21:37:25.252139", "exception": false, "start_time": "2021-04-20T21:37:25.231863", "status": "completed"} tags=[]
| notebooks/LS_test_hd_benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Sample jup2py notebook file
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import sklearn
from sklearn.datasets import load_iris
print("Sample jup2py notebook file")
data = load_iris()
data.keys()
df = pd.DataFrame(data["data"], columns=data["feature_names"])
df["target_names"] = data["target"]
df.head()
# #### Some Graphs
df.plot(figsize=(15, 8), title="Flower attributes")
df.target_names.astype("O").value_counts().plot.pie()
| tests/jup2py_sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.9 64-bit
# name: python3
# ---
import pandas as pd
# +
# get the initial number of columns
initialColumns = 399
my_cols = [str(i) for i in range(initialColumns)]
# +
# read the dataframe
combined_df = pd.read_csv('Inequality_Datafiles/CombinedInequality.txt', sep='\t', names=my_cols, dtype=str)
# +
# a sample of the initial dataframe:
pd.read_csv('Inequality_Datafiles/CombinedInequality.txt', sep='\t', names=my_cols, dtype=str, nrows=5)
# +
# make a function to filter the ngram datasets
def filter_ngram(df, badWords=[]):
# get number of columns starting at 0
initColumns = len(df.columns)-1
# make a copy of dataframe at least for now to not change originak\l
df=df.copy()
# remove rows where headers match rows (probably one read in from both Lower and Upper)
df = df[df[str(initColumns)]!=str(initColumns)]
# remove all rows with "_"
df = df[~df['0'].str.contains("_")]
return df
# +
badWords = []
# filter out the dataframe which combined the upper and lowercase "silly"
df = filter_ngram(combined_df)
df
# -
# make a new column called 'Sentence ID, which is a unique identifier which corresponds to each sentence in the list
df['Sentence ID']=df['0'].copy()
# change column names
df = df.rename(columns = {'0':'5grams'})
# +
# get sum of NA values in each row
naSum = df.isna().sum(axis=1)
# get the row index of the minimum occurance of NA values (axis=1 means row, axis=0 means columns)
minNaIndex = (naSum == min(df.isna().sum(axis=1)))
# get the number of columns that has at least one non NA value:
numColumnsKeep = initialColumns - min(df.isna().sum(axis=1))
# Show the row that has the minummumn number of NA values
df[minNaIndex]
# +
# remove columns with all NaN values
# columns to keep (in list form)
colKeep = [str(i) for i in range(1,numColumnsKeep)]
# since the first column is called '5grams', rename the first element of the list from 0 to '5grams'
colKeep.insert(0, '5grams')
# Keep the Sentence ID column
colKeep.append('Sentence ID')
df = df[colKeep]
df = df.reset_index(drop=True)
df
# +
del df['Sentence ID']
# unpivot the dataframe, all columns except for '5grams'
df = pd.melt(df, id_vars=['5grams'], var_name='Sentence Order', value_name='Sentence Info')
# drop NA values
df = df.dropna().reset_index(drop=True)
# drop Sentence Order column
del df['Sentence Order']
df
# +
# get the numBooks, Num Occurances, and Year as its own data frame
allSentenceInfo = pd.DataFrame(df['Sentence Info'].str.split(',').tolist(),
columns = ['Year','Num Occurances', 'Num books'])
# Combine the 5grams column from the original dataframe with the dataframe about the 5gram information (year, num books, etc.)
df = pd.concat([df['5grams'], allSentenceInfo], axis=1)
df
# +
# make a copy of df, since the df dataframe could be used some other way than this yearGroupsDF
yearGroupsDF = df.copy()
splitWordsDF = pd.DataFrame(df["5grams"].str.split(' ').tolist(),
columns = ['first','second', 'third', 'fourth', 'fith'])
yearGroupsDF = pd.concat([splitWordsDF, yearGroupsDF], axis=1)
yearGroupsDF = yearGroupsDF.drop(['5grams', 'Num Occurances'], 1)
yearGroupsDF['Year'] = pd.to_numeric(yearGroupsDF['Year'], errors='coerce')
yearGroupsDF['Num books'] = pd.to_numeric(yearGroupsDF['Num books'], errors='coerce')
yearGroupsDF
# +
def WordsUsedInYearRange(df, minYear=0, maxYear=2019):
# get the values of df that are in range
df = df[(df['Year'] >= minYear) & (df['Year'] <= maxYear)]
# since this is in the pre 1600 year range, just need to get occurances of words in book
df = df.drop('Year', 1)
# unpivot by all columns except 'Num books'
df = pd.melt(df, id_vars=['Num books'], var_name='Sentence Order', value_name='Words')
# drop the sentence order column
df = df.drop('Sentence Order', 1)
# get the sum of each occurance of a word
df = df.groupby(['Words']).sum()
df = df.sort_values('Num books', ascending=False)
# return the modified dataframe
return df
# -
# an example of this function
yearRangeDF = WordsUsedInYearRange(yearGroupsDF, minYear=1400, maxYear=1999)
yearRangeDF.head(3)
# use loc, not iloc for when there is a categorical index (this is an example)
yearRangeDF.loc['and']
# +
# export these dataframes to csv files
yearRanges = [1475, 1700, 1815, 1914, 1957, 1989, 2019]
fullDF = pd.DataFrame()
for i in range(len(yearRanges)-1):
yearRangeDF = WordsUsedInYearRange(yearGroupsDF, minYear=yearRanges[i], maxYear=yearRanges[i+1])
yearRangeDF = yearRangeDF.reset_index(level=0)
yearRangeDF['Year Range'] = str(yearRanges[i]) + "-" + str(yearRanges[i+1])
fullDF = pd.concat([yearRangeDF, fullDF])
fullDF.to_csv('Inequality_Datafiles/FullYearRange.csv', index=False)
fullDF.head(3)
# +
# get number of books that refrence 'Silly' over time
numBooksOverTime = yearGroupsDF[['Year', 'Num books']]
numBooksOverTime = numBooksOverTime.groupby(['Year']).sum().reset_index()
numBooksOverTime.to_csv('Inequality_Datafiles/BooksPerYear.csv', index=False)
numBooksOverTime.head(3)
# -
| SecondFiltering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''hw2'': conda)'
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
from torchsummary import summary
from tensorboardX import SummaryWriter
from torch import optim
from torch.optim.lr_scheduler import StepLR
import os
from torchvision import datasets
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
import numpy as np
import time
import copy
from tqdm.notebook import tqdm
import torch.backends.cudnn as cudnn
# -
os.environ["CUDA_VISIBLE_DEVICES"] = '0' # GPU Number
start_time = time.time()
batch_size = 128
learning_rate = 0.003
default_directory = './save_models'
writer = SummaryWriter('./log/resnet_50_origin') #!#
# +
transformer_train = transforms.Compose([
transforms.RandomCrop(32, padding=4), # Random Position Crop
transforms.RandomHorizontalFlip(), # right and left flip
#transforms.ColorJitter(brightness=(0.2, 2), contrast=(0.3, 2), saturation=(0.2, 2), hue=(-0.3, 0.3)),
transforms.ToTensor(), # change [0,255] Int value to [0,1] Float value
transforms.Normalize(mean=(0.4914, 0.4824, 0.4467), # RGB Normalize MEAN
std=(0.2471, 0.2436, 0.2616)) # RGB Normalize Standard Deviation
])
transform_test = transforms.Compose([
transforms.ToTensor(), # change [0,255] Int value to [0,1] Float value
transforms.Normalize(mean=(0.4914, 0.4824, 0.4467), # RGB Normalize MEAN
std=(0.2471, 0.2436, 0.2616)) # RGB Normalize Standard Deviation
])
training_dataset = datasets.CIFAR10('./data', train=True, download=True, transform=transformer_train)
#training_dataset_2 = datasets.CIFAR10('./data', train=True, download=True, transform=transformer_train)
#training_dataset_3 = datasets.CIFAR10('./data', train=True, download=True, transform=transformer_train)
validation_dataset = datasets.CIFAR10('./data', train=False, download=True, transform=transform_test)
training_loader = torch.utils.data.DataLoader(dataset=training_dataset, batch_size=batch_size, shuffle=True)
#training_loader_2 = torch.utils.data.DataLoader(dataset=training_dataset_2, batch_size=batch_size, shuffle=True)
#training_loader_3 = torch.utils.data.DataLoader(dataset=training_dataset_3, batch_size=batch_size, shuffle=True)
validation_loader = torch.utils.data.DataLoader(dataset=validation_dataset, batch_size=batch_size, shuffle=False)
# +
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, in_planes, planes, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = out + self.shortcut(x)
out = F.relu(out)
return out
class BottleNeck(nn.Module):
expansion = 4
def __init__(self, in_channels, out_channels, stride=1):
super().__init__()
self.residual_function = nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel_size=1, stride=1, bias=False),
nn.BatchNorm2d(out_channels),
nn.ReLU(),
nn.Conv2d(out_channels, out_channels, kernel_size=3, stride=stride, padding=1, bias=False),
nn.BatchNorm2d(out_channels),
nn.ReLU(),
nn.Conv2d(out_channels, out_channels * BottleNeck.expansion, kernel_size=1, stride=1, bias=False),
nn.BatchNorm2d(out_channels * BottleNeck.expansion),
)
self.shortcut = nn.Sequential()
self.relu = nn.ReLU()
if stride != 1 or in_channels != out_channels * BottleNeck.expansion:
self.shortcut = nn.Sequential(
nn.Conv2d(in_channels, out_channels*BottleNeck.expansion, kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(out_channels*BottleNeck.expansion)
)
def forward(self, x):
x = self.residual_function(x) + self.shortcut(x)
x = self.relu(x)
return x
# +
class ResNet(nn.Module):
def __init__(self, block, num_blocks, num_classes=10):
super(ResNet, self).__init__()
self.in_planes = 64
self.conv1 = nn.Conv2d(3, 64, kernel_size=3, stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.layer1 = self._make_layer(block, 64, num_blocks[0], stride=1)
self.layer2 = self._make_layer(block, 128, num_blocks[1], stride=2)
self.layer3 = self._make_layer(block, 256, num_blocks[2], stride=2)
self.layer4 = self._make_layer(block, 512, num_blocks[3], stride=2)
self.linear = nn.Linear(512*block.expansion, num_classes)
def _make_layer(self, block, planes, num_blocks, stride):
strides = [stride] + [1]*(num_blocks-1)
layers = []
for stride in strides:
layers.append(block(self.in_planes, planes, stride))
self.in_planes = planes * block.expansion
return nn.Sequential(*layers)
def forward(self, x):
#self.dropblock.step()
out = F.relu(self.bn1(self.conv1(x)))
out = self.layer1(out)
out = self.layer2(out)
#out = self.dropblock(out)
out = self.layer3(out)
out = self.layer4(out)
#out = self.dropblock(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
model = ResNet(BottleNeck, [3, 4, 6, 3])
# +
if torch.cuda.device_count() > 0:
print("USE", torch.cuda.device_count(), "GPUs!")
model = nn.DataParallel(model).cuda()
cudnn.benchmark = True
else:
print("USE ONLY CPU!")
summary(model, (3, 32,32))
# -
optimizer = optim.SGD(model.parameters(), learning_rate,
momentum=0.9,
weight_decay=1e-4,
nesterov=True)
#scheduler = torch.optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer, T_0=50, T_mult=3, eta_min=0.001)
criterion = nn.CrossEntropyLoss()
def train(epoch):
model.train()
train_loss = 0
total = 0
correct = 0
iters = len(training_loader)
for batch_idx, (data, target) in enumerate(training_loader):
if torch.cuda.is_available():
data, target = Variable(data.cuda()), Variable(target.cuda())
else:
data, target = Variable(data), Variable(target)
optimizer.zero_grad()
output = model(data)
loss = criterion(output, target)
loss.backward()
optimizer.step()
#scheduler.step(epoch + batch_idx / iters)
train_loss += loss.item()
_, predicted = torch.max(output.data, 1)
total += target.size(0)
correct += predicted.eq(target.data).cpu().sum()
if batch_idx % 10 == 0:
print('Epoch: {} | Batch_idx: {} | Loss_1: ({:.4f}) | Acc_1: ({:.2f}%) ({}/{})'
.format(epoch, batch_idx, train_loss / (batch_idx + 1), 100. * correct / total, correct, total))
writer.add_scalar('training loss', (train_loss / (batch_idx + 1)) , epoch * len(training_loader) + batch_idx) #!#
writer.add_scalar('training accuracy', (100. * correct / total), epoch * len(training_loader) + batch_idx) #!#
writer.add_scalar('lr', optimizer.param_groups[0]['lr'], epoch * len(training_loader) + batch_idx) #!#
# +
def test(epoch):
model.eval()
test_loss = 0
correct = 0
total = 0
for batch_idx, (data, target) in enumerate(validation_loader):
if torch.cuda.is_available():
data, target = Variable(data.cuda()), Variable(target.cuda())
else:
data, target = Variable(data), Variable(target)
outputs = model(data)
loss = criterion(outputs, target)
test_loss += loss.item()
_, predicted = torch.max(outputs.data, 1)
total += target.size(0)
correct += predicted.eq(target.data).cpu().sum()
writer.add_scalar('test loss', test_loss / (batch_idx + 1), epoch * len(validation_loader)+ batch_idx) #!#
writer.add_scalar('test accuracy', 100. * correct / total, epoch * len(validation_loader)+ batch_idx) #!#
print('# TEST : Loss: ({:.4f}) | Acc: ({:.2f}%) ({}/{})'
.format(test_loss / (batch_idx + 1), 100. * correct / total, correct, total))
# +
def save_checkpoint(directory, state, filename='latest_1.tar.gz'):
if not os.path.exists(directory):
os.makedirs(directory)
model_filename = os.path.join(directory, filename)
torch.save(state, model_filename)
print("=> saving checkpoint")
def load_checkpoint(directory, filename='latest_1.tar.gz'):
model_filename = os.path.join(directory, filename)
if os.path.exists(model_filename):
print("=> loading checkpoint")
state = torch.load(model_filename)
return state
else:
return None
# +
start_epoch = 0
checkpoint = load_checkpoint(default_directory, filename='resnet_50_origin.tar.gz')
if not checkpoint:
pass
else:
start_epoch = checkpoint['epoch'] + 1
model.load_state_dict(checkpoint['state_dict'])
for epoch in range(start_epoch, 200):
train(epoch)
save_checkpoint(default_directory, {
'epoch': epoch,
'model': model,
'state_dict': model.state_dict(),
'optimizer': optimizer.state_dict(),
}, filename='resnet_50_origin.tar.gz')
test(epoch)
now = time.gmtime(time.time() - start_time)
print('{} hours {} mins {} secs for training'.format(now.tm_hour, now.tm_min, now.tm_sec))
| resnet50_origin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **LassoCV tuned model**
# For this competition, the objective is to use the test dataset to predict whether a rookie NBA player will be playing in the NBA after 5 years. In a never ending quest to find the next Michael Jordan or Kobe Bryant, NBA teams are looking to invest in young players and want to know, with evidence-based analysis, whether their investments will pay off. This vignette will detail the steps undertaken to create a predictive model and the accuracy of the model against the stated aims.
#Initial imports
import pandas as pd
import numpy as np
import seaborn as sb
import matplotlib.pyplot as plt
import os
import sys
sys.path.append(os.path.abspath('..'))
from src.common_lib import DataReader, NBARawData
# ## **Data input and cleansing**
# The nominated dataset for training and testing was supplied by the kaggle competition. The training dataset is called train.csv and the test dataset is called test.csv.
#Load dataset using common function DataReader.read_data()
data_reader = DataReader()
# Load Raw Train Data
df_train = data_reader.read_data(NBARawData.TRAIN)
# Load Test Raw Data
df_test = data_reader.read_data(NBARawData.TEST)
#For train dataframe, remove redundant column 'Id_old'
df_train.drop('Id_old', axis=1, inplace=True)
df_train.columns = df_train.columns.str.strip()
df_train.describe
#For test dataframe, remove redundant column 'Id_old'
df_test.drop('Id_old', axis=1, inplace=True)
df_test.columns = df_test.columns.str.strip()
df_test.describe
# ## **Feature Correlation and Selection**
# When there are two or more correlated features, multicollinearity can occur and this can lead to skewed or misleading results [1]. In order to reduce this from happening, the features have to be examined for colinearity. Pearson correlation was used for this analysis.
# +
#Use Pearson Correlation to determine feature correlation
pearsoncorr = df_train.corr('pearson')
#Create heatmap of pearson correlation factors
fig, ax = plt.subplots(figsize=(10,10))
sb.heatmap(pearsoncorr,
xticklabels=pearsoncorr.columns,
yticklabels=pearsoncorr.columns,
cmap='RdBu_r',
annot=True,
linewidth=0.2)
# -
#Drop correlated features w/ score over 0.9 - retain "MINS", "3P MADE", "REB"
cols_drop = ["PTS", "FGM", "FGA", "TOV", "3PA", "FTM", "FTA", "REB"]
df_train.drop(cols_drop, axis=1, inplace=True)
df_test.drop(cols_drop, axis=1, inplace=True)
# Split the training dataset using common function data_reader.splitdata
X_train, X_val, y_train, y_val = data_reader.split_data(df_train)
# ## **Model Selection and Training**
# Instead of trial-and-error selection of models, a quick regression model selector was run on common models to see which would yield
# the lowest mean squared error for the training/validation dataset:
#Classifier imports
from sklearn.metrics import f1_score
from sklearn.metrics import mean_squared_error
from math import sqrt
from sklearn.svm import SVC, LinearSVC, NuSVC
from sklearn import linear_model
from sklearn.linear_model import ElasticNetCV
from sklearn.svm import SVR
from sklearn.cluster import KMeans
from sklearn.linear_model import LassoCV
#Select classifiers and display RMSE score:
regressors = [
linear_model.LassoCV(alphas=None, cv=5, n_jobs=-1),
linear_model.LinearRegression(),
ElasticNetCV(alphas=None, copy_X=True, cv=5, eps=0.001, fit_intercept=True,
l1_ratio=0.5, max_iter=5000, n_alphas=100, n_jobs=None,
normalize=False, positive=False, precompute='auto', random_state=0,
selection='cyclic', tol=0.0001, verbose=0),
SVR(C=1.0, cache_size=200, coef0=0.0, degree=3, epsilon=0.1,
gamma='auto', kernel='rbf', max_iter=-1, shrinking=True,
tol=0.001, verbose=False),
linear_model.Ridge(alpha=.05)
]
for regressor in regressors:
model = regressor
model.fit(X_train, y_train)
y_pred_b = model.predict(X_val)
print(regressor)
print("mean squared error: %.3f" % sqrt(mean_squared_error(y_val, y_pred_b)))
# From the above evaluation, it can be seen that all the models except the SVR model returned the same mean squared error (0.355). For this version, the LassoCV model was selected for evaluation. In order to boost accuracy, paramater tuning is employed on the LassoCV to find the optimal alpha value.
#Tune LassoCV
model = LassoCV(alphas=np.arange(0, 1, 0.001), cv=7, n_jobs=-1)
model.fit(X_train, y_train);
#Store model in /models
from joblib import dump
dump(model, '../models/LassoCV.joblib')
#Print optimised model parameters
print('alpha: %f' % model.alpha_)
#Create predictions for train and validation
y_train_preds = model.predict(X_train)
y_val_preds = model.predict(X_val)
#Clip any values > 1, to 1
y_train_preds = np.clip(y_train_preds, a_min=0, a_max=1)
y_val_preds = np.clip(y_val_preds, a_min=0, a_max=1)
# +
#Evaluate train predictions
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
print(mse(y_train, y_train_preds, squared=False))
print(mae(y_train, y_train_preds))
accuracy_score(y_train, y_train_preds.round())
# -
#Train confusion matrix
confusion_matrix(y_train, y_train_preds.round())
#Validation set scores
y_val_prediction = model.predict(X_val)
print(mse(y_val, y_val_preds, squared=False))
print(mae(y_val, y_val_preds))
accuracy_score(y_val, y_val_preds.round())
#Validation confusion matrix
confusion_matrix(y_val, y_val_prediction.round())
# After training and validation, the LassoCV model achieved an accuracy of 83.2% and 83.9% respectively. It is expected to do well in testing against the final target output.
# ## **Test output**
# The output needs to be formatted to comply with the kaggle competition input requirements. The final steps detail this formatting,
# as well as the outcome of the model's testing.
#Output predictions
X_test = df_test
y_test_preds = model.predict(X_test)
y_test_preds.size
y_test_preds = np.clip(y_test_preds, a_min=0, a_max=1)
output = pd.DataFrame({'Id': range(0,3799), 'TARGET_5Yrs': [p for p in y_test_preds]})
output.to_csv("../reports/dt_submission_lasso_03.csv", index=False)
# ## **Outcome**
# After outputting the predictions into kaggle, the final accuracy was 70.6%. This was not the highest achieved by Jump Shot (Hnin Pwint's logistic regression model scored 71.13%) so some further improvements can be made. The removal of the correlated features improved the accuracy, so that will be retained go forwards. Also, from the model evaluation, the Ridge regression model scored the same mean square error as the other models tested (LassoCV and Elasticnet), so a parameter-tuned Ridge regression model should be evaluated in the future.
| notebooks/tay_donovan_12964300_week1_lasso02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import math
import scipy.stats as stats
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, LabelEncoder
# +
# df = pd.read_csv('C:/Users/govindd1/Desktop/Code/VR-Project/ACC_data_from_different_subjects/E4_ACC_data_3.csv')
df = pd.read_csv('./E4_ACC_data_3.csv')
df['Magnitude'] = np.sqrt(df['acc_x']**2 + df['acc_y']**2 + df['acc_z']**2)
df.head()
# -
plt.figure(figsize=(16,8))
plt.title('Data provided by each user', fontsize=20)
sns.countplot(x='subject_id',hue='label', data = df)
plt.show()
df.shape
df2 = df.copy()
df2
# +
group_by_label = df.groupby(by=['subject_id'])
gpdf3 = group_by_label.get_group(3)
X = gpdf3[['acc_x', 'acc_y', 'acc_z']]
y = gpdf3['label']
from imblearn.over_sampling import SMOTE
from matplotlib import pyplot
from collections import Counter
y = LabelEncoder().fit_transform(y)
# transform the dataset
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
# summarize distribution
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
# +
group_by_label = df.groupby(by=['subject_id'])
gpdf6 = group_by_label.get_group(6)
# gpdf6
# plt.figure(figsize=(16,8))
plt.title('Data provided by each user', fontsize=20)
sns.countplot(x='subject_id',hue='label', data = gpdf6)
X = gpdf6[['acc_x', 'acc_y', 'acc_z']]
y = gpdf6['label']
from imblearn.over_sampling import SMOTE
from matplotlib import pyplot
from collections import Counter
y = LabelEncoder().fit_transform(y)
# transform the dataset
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
# summarize distribution
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
plt.show()
# +
group_by_label = df.groupby(by=['subject_id'])
gpdf7 = group_by_label.get_group(7)
# gpdf7
# plt.figure(figsize=(16,8))
plt.title('Data provided by each user', fontsize=20)
sns.countplot(x='subject_id',hue='label', data = gpdf7)
X = gpdf7[['acc_x', 'acc_y', 'acc_z']]
y = gpdf7['label']
from imblearn.over_sampling import SMOTE
from matplotlib import pyplot
from collections import Counter
y = LabelEncoder().fit_transform(y)
# transform the dataset
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
# summarize distribution
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
plt.show()
# +
group_by_label = df.groupby(by=['subject_id'])
gpdf8 = group_by_label.get_group(8)
# plt.figure(figsize=(16,8))
plt.title('Data provided by each user', fontsize=20)
sns.countplot(x='subject_id',hue='label',data = gpdf8)
X = gpdf8[['acc_x', 'acc_y', 'acc_z']]
y = gpdf8['label']
from imblearn.over_sampling import SMOTE
from matplotlib import pyplot
from collections import Counter
y = LabelEncoder().fit_transform(y)
# transform the dataset
oversample = SMOTE()
X, y = oversample.fit_resample(X, y)
# summarize distribution
counter = Counter(y)
for k,v in counter.items():
per = v / len(y) * 100
print('Class=%d, n=%d (%.3f%%)' % (k, v, per))
# plot the distribution
pyplot.bar(counter.keys(), counter.values())
pyplot.show()
plt.show()
# -
frames = [gpdf6, gpdf3, gpdf7, gpdf8]
df3 = pd.concat(frames)
df3
df3['Magnitude'] = np.sqrt(df3['acc_x']**2 + df3['acc_y']**2 + df3['acc_z']**2)
df3['Magnitude']
df3
columns = ['acc_x', 'acc_y', 'acc_z', 'Magnitude', 'type', 'subject_id']
acc_df = pd.DataFrame(data = df3, columns = columns)
acc_df
# +
from window_slider import Slider
def make_windows(df, bucket_size, overlap_count):
window_list = []
final = pd.DataFrame()
activity_list = list(df['type'].unique()) #list of the four activities
sub_id_list = list(df['subject_id'].unique())#list of the subject ids
# co_variance_x_y = list(df['cov_x_y'].unique())
# co_variance_y_z = list(df['cov_y_z'].unique())
# co_variance_x_z = list(df['cov_x_z'].unique())
# round_list = list(df['round'].unique())
df_list = []
for i in sub_id_list:
df_subject = df[df['subject_id'] == i] #isolate a single subject id
for j in activity_list:
df_subject_activity = df_subject[df_subject['type'] == j] #isolate by activity
# for k in co_variance_x_y:
# df_cov_x_y = df_subject_activity[df_subject_activity['cov_x_y'] == k]
# for l in co_variance_y_z:
# df_cov_y_z = df_cov_x_y[df_cov_x_y['cov_y_z'] == l]
# for m in co_variance_x_z:
# df_cov_x_z = df_cov_y_z[df_cov_y_z['cov_x_z'] == m]
final_df = pd.DataFrame()
if df_subject.empty:
pass
else:
df_flat = df_subject[['acc_x', 'acc_y', 'acc_z', 'Magnitude']].T.values #array of arrays, each row is every single reading in an array for a sensor in that isolation
slider = Slider(bucket_size,overlap_count)
slider.fit(df_flat)
while True:
window_data = slider.slide()
if slider.reached_end_of_list(): break
window_list.append(list(window_data))
final_df = final.append(window_list)
final_df.columns = [['acc_x', 'acc_y', 'acc_z', 'Magnitude']]
final_df.insert(4, "subject_id", [i]*len(final_df), True)
final_df.insert(5, "type", [j]*len(final_df), True)
# final_df.insert(6, 'cov_x_y', [k]*len(final_df), True)
# final_df.insert(7, 'cov_y_z', [l]*len(final_df), True)
# final_df.insert(8, 'cov_x_z', [m]*len(final_df), True)
df_list.append(final_df)
window_list = []
final = pd.DataFrame(columns = df_list[0].columns)
for l in df_list:
final = final.append(l)
final
final.columns = final.columns.map(''.join)
return final
# -
df4 = make_windows(acc_df, 64, 32)
df4
df5 = df4.copy()
for i in df5[['acc_x', 'acc_y', 'acc_z', 'Magnitude']].columns:
df5[i] = df5[i].map(lambda x: np.mean(x))
df5.columns = df5.columns + "_mean"
df5
df6 = df4.copy()
for i in df6[['acc_x', 'acc_y', 'acc_z', 'Magnitude']].columns:
df6[i] = df6[i].map(lambda x: np.std(x))
df6.columns = df6.columns + "_std"
df6
frames = [df5, df6]
new_df = pd.concat(frames, axis =1)
new_df = pd.DataFrame(data= new_df)
# +
new_df.columns
# +
# new_df.iloc[:, :12]
# new_df = ['acc_x_mean', 'acc_y_mean', 'acc_z_mean', 'acc_x_std', 'acc_y_std', 'acc_z_std', "Magnitude_mean", 'cov_x_y_mean', 'cov_y_z_mean', 'cov_x_z_mean', 'label_mean']
# new_df
# -
columns = ['acc_x_mean', 'acc_y_mean', 'acc_z_mean', 'acc_x_std', 'acc_y_std', 'acc_z_std', "Magnitude_mean", 'subject_id_mean', 'type_mean']
new_df = pd.DataFrame(data = new_df, columns = columns)
new_df
# +
# df[['Magnitude_mean']].plot(subplots=True, figsize=(33, 10), color = 'red')
new_df['Magnitude_mean'].plot(subplots=True, figsize=(33, 10), color = 'red')
# -
group_by_label = new_df.groupby(by=['subject_id_mean'])
group_by_label
for label_mean, label_mean_df in group_by_label:
print(label_mean)
print(label_mean_df)
# +
# standing_df = group_by_label.get_group(1)
# standing_df
# # standing_df['Magnitude_mean'].plot(subplots=True, figsize=(33, 10), color = 'red')
# standing_df['Magnitude_mean'].plot(figsize = (25,10))
# -
new_df
new_df = new_df.rename(columns={'acc_x_mean': 'accXmean', 'acc_y_mean': 'accYmean', 'acc_z_mean': 'accZmean', 'acc_x_std': 'accXstd', 'acc_y_std': 'accYstd', 'acc_z_std':'accZstd', 'Magnitude_mean': 'MagnitudeMean', 'subject_id_mean': 'subjectID', 'type_mean': 'label' })
new_df
# +
# X = new_df[['accXmean', 'accYmean', 'accZmean', 'accXstd', 'accYstd', 'accZstd', 'MagnitudeMean', 'covXYmean', 'covYZmean', 'covXZmean']]
X = new_df[['accXmean', 'accYmean', 'accZmean', 'accXstd', 'accYstd', 'accZstd', 'MagnitudeMean']]
y = new_df['label']
# -
# scaler = StandardScaler()
# X = scaler.fit_transform(X)
# scaled_X = pd.DataFrame(data = X, columns = ['accXmean', 'accYmean', 'accZmean', 'accXstd', 'accYstd', 'accZstd', 'MagnitudeMean', 'covXYmean', 'covYZmean', 'covXZmean'])
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
X_train
# +
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC, LinearSVC, NuSVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, GradientBoostingClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score
classifiers = [
KNeighborsClassifier(3)
# SVC(kernel="rbf", C=0.025, probability=True),
# NuSVC(probability=True),
# RandomForestClassifier()
# DecisionTreeClassifier()
# AdaBoostClassifier()
# GradientBoostingClassifier()
]
for classifier in classifiers:
# pipe = Pipeline(steps=[('preprocessor', preprocessor),
# ('classifier', classifier)])
# pipe.fit(X_train, y_train)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
print("Accuracy:", accuracy_score(y_test, y_pred))
print("\n ------------------------------------------", classifier , "Classification Report-------------------------------------------------\n")
print(classification_report(y_test, y_pred))
# K-Fold
kfold = 4
results = cross_val_score(classifier, X_train, y_train, cv=kfold)
print(" ------------------------------------------------------ Confusion Matrix ----------------------------------------------------------")
# labels = ['walking', 'sitting', 'standing', 'upstairs', 'downstairs']
labels = ['InMotion', 'MotionLess']
_y_test = y_test
_y_pred = y_pred
confusion_matrix = confusion_matrix(_y_test, _y_pred)
confusion_matrix = confusion_matrix.astype('float') / confusion_matrix.sum(axis=1)[:, np.newaxis]
sns.heatmap(confusion_matrix, xticklabels=labels, yticklabels=labels, annot=True,linewidths = 0.1, cmap = "YlGnBu")
plt.title("Confusion matrix", fontsize = 15)
plt.ylabel("True label")
plt.xlabel("Predicted label")
plt.show()
# print(" ------------------------------------------------------ AUC SCORE -------------------------------------------------------------------")
# skplt.metrics.plot_roc_curve(y_test, y_pred)
# plt.show()
print("---------------------------------------------- ", classifier , "K-Fold Report ------------------------------------------------------", "\n")
# print("model score: %.3f" % pipe.score(X_test, y_test))
# print('------------------------------------ REPORT ----------------------------------', "\n")
print(' * Results: {}'.format(results))
print('\n * Average acuracy: {}'.format(np.mean(results)), "\n")
print('<-------------------------------------------------------- END ------------------------------------------------------------------>')
# -
| ACC_data_from_different_subjects/Notebooks/final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Remember Shift+ENTER is your friend XD :)
# ### First you gotta login to do something amazing!
# +
from selenium import webdriver
import sys
import os
from numpy.random import choice
import numpy as np
from time import sleep
from random import choice, randint
import traceback
import regex
import datetime
import re
from selenium.webdriver.common.keys import Keys
from time import sleep, strftime
from random import randint
import random
import IPython
location=os.getcwd()
print(location)
# +
#direc="C:/Users/farha/Downloads/chromedriver_win32_new/"
#os.chdir(direc)
#sys.path.append(direc)
from selenium.webdriver.chrome.options import Options
mobile_emulation = {
"deviceMetrics": { "width": 360, "height": 640, "pixelRatio": 3.0 },
"userAgent": "Mozilla/5.0 (Linux; Android 4.2.1; en-us; Nexus 5 Build/JOP40D) AppleWebKit/535.19 (KHTML, like Gecko) Chrome/18.0.1025.166 Mobile Safari/535.19" }
chrome_options = Options()
chrome_options.add_experimental_option("mobileEmulation", mobile_emulation)
driver = webdriver.Chrome(chrome_options = chrome_options)
#driver = webdriver.Chrome(options = chrome_options)
webdriver=driver
cd=driver
#chrome driver
# -
def login(usr,passwrd):
cd.get('https://www.instagram.com/accounts/login/?source=auth_switcher')
sleep(3)
username = webdriver.find_element_by_name('username')
username.send_keys(usr)
password = webdriver.find_element_by_name('password')
password.send_keys(<PASSWORD>)
sleep(randint(2,4))
button_login = webdriver.find_element_by_xpath('//div[text()="Log In"]')
button_login.click()
#try increasing sleep to increase low crash issues
sleep(5)
try:
notnow = webdriver.find_element_by_xpath('//button[text()="Not Now"]')
notnow.click() #comment these last 2 lines out, if you don't get a pop up asking about notifications
except:
pass
#try increasing sleep to increase low crash issues
sleep(5)
try:
cancel = webdriver.find_element_by_xpath('//button[text()="Cancel"]')
cancel.click() #comment these last 2 lines out, if you don't get a pop up asking about notifications
except:
pass
username='i.am.the.swagger'
password='<PASSWORD>'
login(username,password)
# +
#run this cell if you want to change the location to a pre-specified one.
#location="C:/Users/farha/Desktop/DESKTOP_FILES/ColorSplash/"
# -
def fetch_url(url):
if (cd.current_url==url):
return 0
else :
cd.get(url)
return 1
# ### Basically the above cells are for logging in only. We will now consider deeper functions!
# ## Running the below cells gets the number of followers and followings that you have.
# +
def get_int(followers):
thou=False
if(re.search('k',followers)):
#followers=followers.replace('k','000')
thou=True
followers=followers.replace(',','')
temp = re.findall(r'\d+', followers)
res = list(map(int, temp))
if thou:
res[0]=res[0]*1000
try:
return res[0]+res[1]*100
except:
return res[0]
def print_list(lst):
for i in lst:
print(i)
# -
def get_num(what="followers",username="xyz"):
profile_link="https://www.instagram.com/"+username
fetch_url(profile_link)
if(what=="followers"):
mini_followers_link="/"+username+"/followers/"
followers_searc_str="//a[@href='{}']".format(mini_followers_link)
followers_element=webdriver.find_element_by_xpath(followers_searc_str)
followers=get_int(followers_element.text)
return followers
else:
mini_following_link="/"+username+"/following/"
following_searc_str="//a[@href='{}']".format(mini_following_link)
following_element=webdriver.find_element_by_xpath(following_searc_str)
following=get_int(following_element.text)
return following
#for following follower list from id
#profile_link="https://www.instagram.com/"+"suvradipghosh01/"
username="i.am.the.swagger"
#username="suvradipghosh01"
profile_link="https://www.instagram.com/"+username
fetch_url(profile_link)
followers=get_num(what="followers",username=username)
print("Followers :",followers)
# +
following=get_num(what="following",username=username)
print("Following :",following)
# -
# ### A few utility functions
#https://stackoverflow.com/questions/20986631/how-can-i-scroll-a-web-page-using-selenium-webdriver-in-python
def ids_scroll(ids):
num=50*ids
#1 id=150 pixels
script="window.scrollTo(0, {});".format(num)
driver.execute_script(script)
def scroll_down(num):
for i in range(int(num)):
ids_scroll(100000)
SCROLL_PAUSE_TIME = 0.5
sleep(SCROLL_PAUSE_TIME)
# +
#small val -- ids_scroll
# -
#https://stackoverflow.com/questions/20986631/how-can-i-scroll-a-web-page-using-selenium-webdriver-in-python
def scroll_till_end():
SCROLL_PAUSE_TIME = 0.5
# Get scroll height
last_height = driver.execute_script("return document.body.scrollHeight")
while True:
# Scroll down to bottom
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
# Wait to load page
sleep(SCROLL_PAUSE_TIME)
# Calculate new scroll height and compare with last scroll height
new_height = driver.execute_script("return document.body.scrollHeight")
if new_height == last_height:
break
last_height = new_height
# +
#progress bar
#https://stackoverflow.com/questions/3160699/python-progress-bar
import time, sys
# update_progress() : Displays or updates a console progress bar
## Accepts a float between 0 and 1. Any int will be converted to a float.
## A value under 0 represents a 'halt'.
## A value at 1 or bigger represents 100%
def update_progress(progress):
barLength = 80 # Modify this to change the length of the progress bar
status = ""
if isinstance(progress, int):
progress = float(progress)
if not isinstance(progress, float):
progress = 0
status = "error: progress var must be float\r\n"
if progress < 0:
progress = 0
status = "Halt...\r\n"
if progress >= 1:
progress = 1
status = "Done...\r\n"
block = int(round(barLength*progress))
text = "\rCompleted: [{0}] {1}% {2}".format( "#"*block + "-"*(barLength-block), round(progress*100,2), status)
sys.stdout.write(text)
sys.stdout.flush()
# -
def get_list(lim,allow=20,e=0,l=0):
list_users=[]
#IPython.display.clear_output()
scroll_down(lim/8)
elems = cd.find_elements_by_xpath("//a[@href]")
if(len(elems)<int(lim-allow)):
e+=1
text="Failed to scroll down completely. Retrying : "+str(e)
IPython.display.clear_output()
sys.stdout.write(text)
sys.stdout.flush()
list_users=get_list(lim,allow,e,l)
count=0
length=len(elems)
for elem in elems:
try:
id_link=elem.get_attribute("href")
id_link=id_link[26:]
id_link=id_link.strip('/')
list_users.append(id_link)
count+=1
update_progress(count/length)
except:
continue
list_users=list(set(list_users))
if(len(list_users)<int(lim-allow)):
l+=1
IPython.display.clear_output()
text="Failed to get all users. Retrying : "+str(l)
sys.stdout.write(text)
text="Current List Length : "+str(len(list_users))
sys.stdout.write(text)
sys.stdout.flush()
#print("Failed to get all users. Retrying : ",l)
#print("Current List Length :",len(list_users))
list_users=get_list(lim,allow,e,l)
print("Returned List Length :",len(list_users))
return list_users
# # Below cell gets all your following list.
# **This may take some long time depending on your number of following**<br>
# Todo optimizer.
followers_link="https://www.instagram.com/"+username+"/followers/"
following_link="https://www.instagram.com/"+username+"/following/"
fetch_url(following_link)
print(following_link)
try:
ids_scroll(5)
following_href="/"+username+"/following/"
url=following_href
following_xpath_complete='//a[@href="'+url+'"]'
action_following=webdriver.find_element_by_xpath(following_xpath_complete)
action_following.click()
sleep(5)
except:
pass
list_following=get_list(following)
# +
def write_txt(username,list_action,filepost="following"):
filename=username+"_"+filepost+".txt"
#filename=os.path.join(location,filename)
file_action=open(filename,"a+")
for each in list_action:
file_action.write(each+'\n')
file_action.close()
def rewrite_txt(username,list_action,filepost="following"):
filename=username+"_"+filepost+".txt"
#filename=os.path.join(location,filename)
file_action=open(filename,"w")
for each in list_action:
file_action.write(each+'\n')
file_action.close()
# -
rewrite_txt(username,list_following,"following")
if(len(list_following)>=following):
print("Processed following successfully!!")
"""
lenOfPage = cd.execute_script("window.scrollTo(0, document.body.scrollHeight);var lenOfPage=document.body.scrollHeight;return lenOfPage;")
match=False
while(match==False):
lastCount = lenOfPage
time.sleep(3)
lenOfPage = driver.execute_script("window.scrollTo(0, document.body.scrollHeight);var lenOfPage=document.body.scrollHeight;return lenOfPage;")
if lastCount==lenOfPage:
match=True
"""
"""from selenium.webdriver.common.keys import Keys
html = driver.find_element_by_tag_name('html')
html.send_keys(Keys.END)
"""
# # Below cell gets all your followers list.
# **This may take some long time depending on your number of followers**<br>
# Todo optimizer.
followers_link="https://www.instagram.com/"+username+"/followers/"
following_link="https://www.instagram.com/"+username+"/following/"
webdriver.get(followers_link)
print(followers_link)
try:
ids_scroll(5)
followers_href="/"+username+"/followers/"
url=followers_href
followers_xpath_complete='//a[@href="'+url+'"]'
action_followers=webdriver.find_element_by_xpath(followers_xpath_complete)
action_followers.click()
sleep(5)
except:
pass
list_followers=get_list(followers)
rewrite_txt(username,list_followers,"followers")
if(len(list_followers)>=followers):
print("Processed followers successfully!!")
# # Run these cells to load your followers from the saved txt file.
def print_list(lines):
for e in lines:
print (e)
def filter_list(lst):
new_lst=[]
for e in lst:
if(e=="\n"):
continue
else:
#print(e.strip())
new_lst.append(e.strip())
return new_lst
def load_txt(username,filepost="following"):
filename=username+"_"+filepost+".txt"
#filename=os.path.join(location,filename)
text_file = open(filename, "r")
lines = text_file.readlines()
filtered_lst=filter_list(lines)
list_action=list(set(filtered_lst))
text_file.close()
return list_action
followers_all=load_txt(username,filepost="followers")
print_list(followers_all)
# # Run these cells to load your following from the saved txt file.
# +
following_all=load_txt(username,filepost="following")
print_list(following_all)
# +
# Python code to get difference of two lists
# Using set()
def Diff(li1, li2):
return (list(set(li1) - set(li2)))
not_following_back=Diff(following_all,followers_all)
print_list(not_following_back)
# -
len(not_following_back)
def write_txt(username,list_action,filepost="following"):
filename=username+"_"+filepost+".txt"
#filename=os.path.join(location,filename)
file_action=open(filename,"a+")
for each in list_action:
file_action.write(each+'\n')
file_action.close()
write_txt(username,not_following_back,"not_following_back")
# +
#don't run the following cell
# +
#load the unfollowed list
location="C:/Users/farha/Desktop/DESKTOP_FILES/ColorSplash/"
username="i.am.the.swagger"
filename=username+"_unfollowed.txt"
filename=os.path.join(location,filename)
text_file = open(filename, "r")
lines = text_file.readlines()
lines=filter_list(lines)
unfollowed=list(set(lines))
print_list(unfollowed)
# -
#todo take another pareameter isopen=False
def get_both(username="xyz"):
profile_link="https://www.instagram.com/"+username
webdriver.get(profile_link)
mini_followers_link="/"+username+"/followers/"
followers_searc_str="//a[@href='{}']".format(mini_followers_link)
followers_element=webdriver.find_element_by_xpath(followers_searc_str)
followers=get_int(followers_element.text)
mini_following_link="/"+username+"/following/"
following_searc_str="//a[@href='{}']".format(mini_following_link)
following_element=webdriver.find_element_by_xpath(following_searc_str)
following=get_int(following_element.text)
return (followers,following)
#unfollowed=[]
unfollowed=load_txt(username,filepost="unfollowed")
people=Diff(not_following_back,unfollowed)
print(len(people))
# +
#unfollowed=[]
unfollowed=load_txt(username,filepost="unfollowed")
people=Diff(not_following_back,unfollowed)
counter=0
for each in people:
try:
current_followers,current_following=get_both(username=each)
#margin=20
#criteria=True
#this is the criteria
#if(current_following>=(current_followers+margin)):
#criteria=True
#cd.get("www.instagram.com/"+each+'/')
pre_unfollow = webdriver.find_element_by_xpath("//span[@aria-label='Following']")
pre_unfollow.click()
sleep(randint(1,2))
#try with longer sleep duration to avoid blocking Use sleep(randint(10,200))
unfollow = webdriver.find_element_by_xpath('//button[text()="Unfollow"]')
unfollow.click()
counter+=1
print(counter,each)
if (counter>=200):
print("Limit reached")
break
unfollowed.append(each)
sleep(randint(6,20))
write_txt(username,unfollowed,"unfollowed")
except (KeyboardInterrupt, SystemExit):
write_txt(username,unfollowed,"unfollowed")
break
except:
pass
#save filter list
write_txt(username,unfollowed,"unfollowed")
# -
# # Now filter people on the basis of a specefic criteria
#
# The *people* variable contains what type of people we want to filter the conditions from.<br>
# typically followers following and not_following_back
#
people=not_following_back
filtered_list=[]
for each in people:
try:
current_followers,current_following=get_both(username=each)
margin=200
criteria=False
#this is the criteria
if(current_following>=(current_followers+margin)):
criteria=True
filtered_list.append(each)
sleep(randint(1,2))
except:
continue
# +
#save filter list
filename=username+"_filter_criteria.txt"
location="C:/Users/farha/Desktop/DESKTOP_FILES/ColorSplash/"
filename=os.path.join(location,filename)
file_followers=open(filename,"a+")
for each in filtered_list:
file_followers.write(each+'\n')
file_followers.close()
# -
print("Go check the filtered file at : ",filename)
# # Warning : you might want to visit and edit your filter_criteria.txt file.
# ## The below code will unfollow people from the filter_criteria.txt file.
# # Now it's time for some action.
# *Let's unfollow those guys straight!*
location="C:/Users/farha/Desktop/DESKTOP_FILES/ColorSplash/"
filename=username+"_filter_criteria.txt"
filename=os.path.join(location,filename)
text_file = open(filename, "r")
lines = text_file.readlines()
#load the filtered data+manual filter if needed
lines=filter_list(lines)
filtered_list=list(set(lines))
print_list(filtered_list)
unfollow_list=[]
counter=0
for each in not_following_back[200:300]:
try:
criteria=True
if(criteria):
#unfollow_list.append(people)
#comment out the following lines by putting a # in front of them
#to only get a list and not unfollow the filtered person by criteria
#id_link='www.instagram.com/'+each
#webdriver.get(id_link)
profile_link="https://www.instagram.com/"+each
webdriver.get(profile_link)
pre_unfollow = webdriver.find_element_by_xpath("//span[@aria-label='Following']")
pre_unfollow.click()
sleep(randint(1,20))
#try with longer sleep duration to avoid blocking Use sleep(randint(10,200))
unfollow = webdriver.find_element_by_xpath('//button[text()="Unfollow"]')
unfollow.click()
counter+=1
print(counter,":",each)
unfollow_list.append(each)
except:
continue
# +
#save the finally unfollowed list
filename=username+"_unfollowed.txt"
location="C:/Users/farha/Desktop/DESKTOP_FILES/ColorSplash/"
filename=os.path.join(location,filename)
file_followers=open(filename,"a+")
for each in unfollow_list:
file_followers.write(each+'\n')
file_followers.close()
# -
webdriver.quit()
# +
#unfollow = webdriver.find_element_by_xpath('//button[text()="Unfollow"]')\
#username = driver.find_element_by_xpath("//input[@name='username']")
#/html/body/div[1]/section/main/div/header/section/div[2]/div[2]/span/span[1]/button/div/span
# -
# +
list_followers=[]
for i in range(2000):
elems = webdriver.find_elements_by_xpath("//a[@href]")
for elem in elems:
try:
id_link=elem.get_attribute("href")
except:
continue
#print(id_link)
id_link=id_link[26:]
id_link=id_link.strip('/')
if id_link not in list_followers:
list_followers.append(id_link)
#print(id_link)
webdriver.execute_script("window.scrollTo(0,document.body.scrollHeight)")
if len(list_followers)>=followers:
break
list_followers=set(list_followers)
print(len(list_followers))
#print_list(list_following)
# -
file_follower=open("suvradip_followers.txt","a+")
for each in list_followers:
file_follower.write(each+'\n')
file_follower.close()
# +
def Diff(li1, li2):
return (list(set(li1) - set(li2)))
def common_member(a, b):
a_set = set(a)
b_set = set(b)
if (a_set & b_set):
#print(a_set & b_set)
return a_set & b_set
else:
print("No common elements")
not_following_back=Diff(list_following,list_followers)
file_not_following_back=open("suvradip_not_following_back.txt","a+")
for each in not_following_back:
file_not_following_back.write(each+'\n')
file_not_following_back.close()
mutual=common_member(list_following,list_followers)
file_mutual=open("suvradip_mutual.txt","a+")
for each in mutual:
file_mutual.write(each+'\n')
file_mutual.close()
# -
#get the complete list in the given page
#stopiter -- max number of iterations the loop will run for - experimental , 200/2000 works well
#limit -- the number of followers/following (expected termination value)
#adjust-- the number of allowed missing components /ids
#optimizer -- the optimal value for how many values does one loop catch
def get_list2(stopiter=2000,limit=2000,optimizer=10,adjust=20)
list_users=[]
counter=0
for i in range(round(limit/optimizer)):
elems = cd.find_elements_by_xpath("//a[@href]")
counter+=1
if counter==stopiter:
return list(set(list_users))
for elem in elems:
try:
id_link=elem.get_attribute("href")
except:
continue
#print(id_link)
id_link=id_link[26:]
id_link=id_link.strip('/')
if id_link not in list_following:
list_following.append(id_link)
#might try sleep here for weak internet connections
#sleep(3)
#print(id_link)
cd.execute_script("window.scrollTo(0,document.body.scrollHeight)")
if len(list_users)>=limit-adjust:
break
list_users=set(list_users)
return list_users
#print(len(list_users))
# +
filtered=[]
for each_user in following:
webdriver.get("https://www.instagram.com/"+str(user)+"/")
#for following follower list from id
#get followers and following
try:
followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/header/section/ul/li[2]/a')
followers=get_int(followers_element.text)
except:
followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/ul/li[2]/a/span')
followers=get_int(followers_element.text)
print("Followers :",followers)
try:
following_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/ul/li[3]/a')
except:
#followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/header/section/ul/li[3]/a/span')
print("Err")
following=get_int(following_element.text)
print("Following :",following)
#here comes the criteria
if (((following-200)>followers) or ((following<2000))):
#then filterout the user
filtered.append(each_user)
# -
filtered=set(filtered)
file_filtered=open("suvradip_filtered.txt","a+")
for each in filtered:
file_filtered.write(each+'\n')
file_filtered.close()
webdriver.close()
# +
#no cell will run after this
# +
def get_int(followers):
thou=False
if(re.search('k',followers)):
#followers=followers.replace('k','000')
thou=True
followers=followers.replace(',','')
temp = re.findall(r'\d+', followers)
res = list(map(int, temp))
if thou:
res[0]=res[0]*1000
try:
return res[0]+res[1]*100
except:
return res[0]
def print_list(lst):
for i in lst:
print(i)
#for following follower list from id
profile_link="https://www.instagram.com/"+"suvradipghosh01/"
webdriver.get(profile_link)
#get followers and following
try:
followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/header/section/ul/li[2]/a')
followers=get_int(followers_element.text)
except:
followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/ul/li[2]/a/span')
followers=get_int(followers_element.text)
print("Followers :",followers)
try:
following_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/header/section/ul/li[3]/a/span')
except:
#followers_element=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/div/header/section/ul/li[3]/a/span')
print("Err")
following=get_int(following_element.text)
print("Following :",following)
# -
string1 = 'AJYF'
>>> string2 = 'AJ'
>>> if string2 in string1:
... string1.replace(string2,'')
hashtag_list = ['photography', 'deeplearning']
webdriver.get('https://www.instagram.com/explore/tags/'+ hashtag_list[1] + '/')
sleep(randint(4,9))
#thumb_str='/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[1]/a/div[1]/div[2]'
#thumb_str='/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[2]/a/div/div[2]'
#thumb_str='/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[2]/a/div/div[2]'
#thumb_str='/html/body/div[1]/section/main/article/div[1]/div/div/div[long_text_1]/div[long_text_2]/a/div/div[2]'
#thumb_str = '//*[@id="react-root"]/section/main/article/div[1]/div/div/div[long_text_1]/div[long_text_2]/a/div/div[2]'
row,col=3,2
element=thumb_str.replace("long_text_1",str(row))
element=element.replace("long_text_2",str(col))
print(element)
element='/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[1]/a/div/div[2]'
thumb_element=webdriver.find_element_by_xpath(element)
thumb_element.click()
#/html/body/div[1]/section/main/article/div[1]/div/div/div[3]/div[2]/a/div/div[2]
first_thumbnail = webdriver.find_element_by_xpath('//*[@id="react-root"]/section/main/article/div[1]/div/div/div[1]/div[1]/a/div')
first_thumbnail.click()
first_th=webdriver.find_element_by_xpath('/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[1]/a/div/div[2]')
second_th= webdriver.find_element_by_xpath('/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[2]/a/div/div[2]')
third_th= webdriver.find_element_by_xpath('/html/body/div[1]/section/main/article/div[1]/div/div/div[1]/div[3]/a/div/div[2]')
sleep(randint(3,6))
webdriver.quit()
# +
list_following=[]
for i in range(2):
elems = webdriver.find_elements_by_xpath("//a[@href]")
for elem in elems:
try:
id_link=elem.get_attribute("href")
except:
continue
#print(id_link)
id_link=id_link.strip()
#string1-string2
string1=str(id_link)
string2="https://www.instagram.com/"
if string2 in string1:
print(True)
string1.replace(string2,'')
print(string1)
id_link=string1
id_link=id_link.strip('/')
if id_link not in list_following:
list_following.append(id_link)
#print(id_link)
webdriver.execute_script("window.scrollTo(0,document.body.scrollHeight)")
if len(list_following)>=1000:
break
print(len(list_following))
| Main_Insta_Fast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
file_name=input('enter file name ')
g1=pd.read_csv(file_name)
g2=g1.iloc[:,3:]
g2
g2=g2.drop(g2.columns[[4, 5, 0]], axis=1)
g2=g2.fillna(g2.mean())
g2
file_name1=file_name+'clean'
g2.to_csv(file_name, sep=',')
| backend/plot/extra/cleaning data/nasa delhi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../imgs/logo.png" width="20%" align="right" style="margin:0px 20px">
#
#
# # Evolutionary Computation
#
# ## 3.2 NSGA-II
#
# <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/"><img alt="Creative Commons License" align="left" src="https://i.creativecommons.org/l/by-sa/4.0/80x15.png" /></a> | <NAME> | <a href="https://d9w.github.io/evolution/">https://d9w.github.io/evolution/</a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Outline
#
# 1. [The Knapsack Problem](#knapsack)
# 2. [Evolution](#evolution)
# 3. [Visualization](#visualization)
# + slideshow={"slide_type": "slide"}
using NSGAII
using JuMP
import GLPK
using Plots
default(show=:ijulia)
# + [markdown] slideshow={"slide_type": "fragment"}
# To experiment with NSGAII we'll be using a rather new implementation of it from Github: https://github.com/gsoleilhac/NSGAII.jl/.
#
# This implementation isn't so complicated and if we look at the code, we can see that much of it resembles our [notebook code](https://github.com/d9w/evolution/blob/master/2_ga/1_GAs.ipynb) on Genetic Algorithms. Let's look at the [functions](https://github.com/gsoleilhac/NSGAII.jl/blob/master/src/functions.jl) code which contains the main Genetic Algorithm loop, as well as the [NSGA-II](http://repository.ias.ac.in/83498/1/2-a.pdf) Fast Non-Dominated Sort and Crowding Distance Algorithm. Notice the structure of individuals in the code and how rank is stored; we will use this information later.
# + [markdown] slideshow={"slide_type": "slide"}
# ## <a id="knapsack"></a>The Knapsack Problem
# + [markdown] slideshow={"slide_type": "fragment"}
# <img src="https://imgs.xkcd.com/comics/np_complete.png">
# <div style="text-align:center"><a href="https://xkcd.com/287/">XKCD on the Knapsack Problem</a></div>
#
# + [markdown] slideshow={"slide_type": "slide"}
# Imagine we're travelling and our bag has a maximum weight limit. We want to fill this bag with items of different value and maximize the total value in our bag, as long as it stays in the weight limit. This is known as the "knapsack problem".
# <img src="../imgs/knapsack.png" width="30%" height="auto">
# Instead of a bag with items, you could imagine a scenario like the comic. You want to order different appetizers, but you only want to spend a fixed maximum amount. How can you get the best appetizers?
# + [markdown] slideshow={"slide_type": "slide"}
# Let's look at an example, specifically using the version of the Knapsack problem known as the "0-1 knapsack problem" where we can have at most 1 copy of each item. We'll use 5 items and randomly geneate values for them.
# + slideshow={"slide_type": "fragment"}
n = 5
values = rand(0:100, n)
println(values, " total value: ", sum(values))
# + slideshow={"slide_type": "fragment"}
weights = rand(10:20, n)
max_weight = 55
println(weights, " total weight: ", sum(weights))
# + [markdown] slideshow={"slide_type": "fragment"}
# We won't be able to take all items because of the maximum weight of 55. We'll express our choice of items using a binary string, 1 if we take the item and 0 otherwise.
# + slideshow={"slide_type": "fragment"}
using Random: bitrand
choices = bitrand(n)
println(choices)
# + [markdown] slideshow={"slide_type": "slide"}
# The total value of our knapsack is therefore the sum of the element-wise multiplication of our choices and the individual item values.
# + slideshow={"slide_type": "fragment"}
sum(choices .* values)
# + [markdown] slideshow={"slide_type": "fragment"}
# In other words, the dot product of the two vectors.
# + slideshow={"slide_type": "fragment"}
using LinearAlgebra: dot
dot(choices, values)
# + [markdown] slideshow={"slide_type": "fragment"}
# We can also use the dot product to check if our choices fall under the weight limit
# + slideshow={"slide_type": "fragment"}
dot(choices, weights), max_weight
# + [markdown] slideshow={"slide_type": "slide"}
# Let's say you and a friend are travelling together and to make it cheaper, you're sharing a bag. You and your friend assign different values to each item based on what you want to bring on the trip. Your preferences are in `values` and your friends preferences are:
# + slideshow={"slide_type": "fragment"}
values2 = rand(0:100, n)
println("You:\t", values, " total value: ", sum(values))
println("Friend:\t", values2, " total value: ", sum(values2))
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's see who is happier with the choices we made
# + slideshow={"slide_type": "fragment"}
choices = bitrand(n)
println(choices)
println("weight: ", dot(choices, weights))
println("You: ", dot(choices, values), " Friend: ", dot(choices, values2))
# + [markdown] slideshow={"slide_type": "fragment"}
# This is the multi-objective version of the Knapsack problem where we use $K$ different value vectors for $K$ objectives.
# + [markdown] slideshow={"slide_type": "slide"}
# [The Knapsack problem](https://en.wikipedia.org/wiki/Knapsack_problem) has been thoroughly studied in theoretical computer science and evolutionary computation is not the only choice for solving it; dynamic programming and mixed integer programming are what [JuMP](https://github.com/JuliaOpt/JuMP.jl), an optimization library, uses.
# + slideshow={"slide_type": "fragment"}
m = Model(GLPK.Optimizer)
@variable(m, x[1:n], Bin)
# Objective: maximize profit
@objective(m, Max, dot(values, x))
# Constraint: can carry all
@constraint(m, dot(weights, x) <= max_weight)
# Solve problem using MIP solver
optimize!(m);
# -
objective_function(m)
# + slideshow={"slide_type": "fragment"}
println("Objective is: ", objective_value(m))
println("Solution is:")
for i in 1:n
print("x[$i] = ", value(x[i]))
println(", p[$i]/w[$i] = ", values[i]/weights[i])
end
# + [markdown] slideshow={"slide_type": "slide"}
# ## <a id="evolution"></a>Evolution
# + [markdown] slideshow={"slide_type": "fragment"}
# We will apply NSGA-II to the Knapsack problem. First, let's scale up the number of items to 100.
# + slideshow={"slide_type": "fragment"}
n = 100
n_objectives = 2
values = rand(n, n_objectives)
# + [markdown] slideshow={"slide_type": "slide"}
# Let's normalize our fitness values by their sum, so that the total maximum value is 1.0. This will help with plotting later.
# + slideshow={"slide_type": "fragment"}
values = values ./ sum(values, dims=1);
sum(values, dims=1)
# + [markdown] slideshow={"slide_type": "slide"}
# Similarly, let's normalize the weights to sum to 1 and set a maximum weight of 0.5, meaning we can keep only half the items on average. This will let us change the number of items more easily later on but doesn't impact the problem in any way.
# + slideshow={"slide_type": "fragment"}
weights = rand(n)
weights = weights ./ sum(weights)
max_weight = 0.5
# + [markdown] slideshow={"slide_type": "slide"}
# Let's write our objective function which will compute the value of our knapsack using each different column of profit values.
# + slideshow={"slide_type": "fragment"}
function objective(genes::AbstractArray)
fits = zeros(n_objectives)
for i in 1:n_objectives
fits[i] = dot(genes, values[:, i])
end
tuple(fits...)
end
# + [markdown] slideshow={"slide_type": "slide"}
# We'll add in our weight constraint separately. This is used in the evaluation of individuals and is considered a part of our objective function, but is handled separately. Our lecture on evolution with constraints will go into more detail on that. For this NSGA-II implementation, we return 0 if the condition passes and the distance from meeting the condition otherwise.
# + slideshow={"slide_type": "fragment"}
function constraint(genes::AbstractArray)
sum_weights = dot(genes, weights)
return sum_weights <= max_weight ? 0 : sum_weights - max_weight
end
# + [markdown] slideshow={"slide_type": "slide"}
# That's all, so now we're ready to run NSGA-II. Let's run it for 100 generations on a population of 100. The function returns the set of non-dominated individuals at the end
# + slideshow={"slide_type": "fragment"}
popsize = 100
nbgen = 100
init() = bitrand(n)
non_dominated = nsga_max(popsize, nbgen, objective, init, fCV=constraint)
non_dominated[1]
# + [markdown] slideshow={"slide_type": "slide"}
# Let's see what type of total profit values we have from our two objectives.
# + slideshow={"slide_type": "fragment"}
unique([collect(i.y) for i in non_dominated])
# + [markdown] slideshow={"slide_type": "slide"}
# Finally, we can compare this value to a value obtained by JuMP. Note that JuMP can only currently perform single objective optimization, so we need to solve the different objectives separately.
# + slideshow={"slide_type": "fragment"}
m = Model(GLPK.Optimizer)
@variable(m, x[1:n], Bin)
# Objective: maximize profit
@objective(m, Max, dot(values[:, 1], x))
# Constraint: can carry all
@constraint(m, dot(weights, x) <= max_weight)
# Solve problem using MIP solver
optimize!(m)
println("Objective 1: ", objective_value(m))
print("Objective 2: ")
o2 = sum([value(x[i])*values[i, 2] for i in 1:n])
print(o2)
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success">
# <b>Exercise 1</b>
# <br/>
# Increase the problem difficulty by increasing the number of items and reducing the weight limit. How does NSGA-II do?
# <br/>
# </div>
# -
# <div class="alert alert-success">
# <b>Exercise 2</b>
# <br/>
# Increase the number of objectives to 5. What do you observe about the solutions found by NSGA-II? How do they compare with the solutions found by JuMP?
# <br/>
# </div>
# + [markdown] slideshow={"slide_type": "slide"}
# ## <a id="visualization"></a>Visualization
# + [markdown] slideshow={"slide_type": "fragment"}
# To better understand NSGA-II's evolution, let's observe the population over time.
# + slideshow={"slide_type": "slide"}
function plot_pop(P)
scatter([i.y[1] for i in P], [i.y[2] for i in P], xlims=(0.0, 1.0), ylims=(0.0, 1.0), legend=:none)
ND = filter(indiv -> indiv.rank == 1, P)
scatter!([i.y[1] for i in ND], [i.y[2] for i in ND], legend=:none)
end
nsga_max(popsize, nbgen, objective, init, fCV=constraint, fplot=plot_pop, plotevery=1, showprogress=true);
# + [markdown] slideshow={"slide_type": "slide"}
# This type of visualization, plotting one objective function as X and another as Y, allows us to see the Pareto front for 2 dimensions easily. However, once we increase the number of objectives, this plot is much less useful as we'll always be constrained to looking at 2 dimensions, or at best 3.
# + [markdown] slideshow={"slide_type": "fragment"}
# <div class="alert alert-success">
# <b>Exercise 3</b>
# <br/>
# Change the number of objectives to 3. Then use a <tt>scatter3d</tt> plot instead of a <tt>scatter</tt> plot to plot the evolution over 3 objectives.
# <br/>
# <ul>
# <li>Can you still observe the Pareto front?</li>
# <li>Can you identify non-dominated individuals easily</li>
# <li>Do you see the performance for each objective</li>
# </ul>
# <br/>
# </div>
# + [markdown] slideshow={"slide_type": "slide"}
# Another way to look at the different objectives is to plot the different objectives on the `x` axis and the fitness values on the `y` axis. Each individual in a population is then represented as a line. This is known as a Parallel Coordinate Plot. Let's try this method, increasing the number of objectives to 10.
# + slideshow={"slide_type": "fragment"}
n = 100
n_objectives = 10
values = rand(n, n_objectives)
values = values ./ sum(values, dims=1)
weights = rand(n)
weights = weights ./ sum(weights)
max_weight = 0.5
popsize = 100
nbgen = 50
init() = bitrand(n)
# + slideshow={"slide_type": "slide"}
ymin = Inf
ymax = -Inf
function plot_pop(P)
global ymin = min(ymin, minimum([minimum(i.y) for i in P]))
global ymax = max(ymax, maximum([maximum(i.y) for i in P]))
plot(xlims=(1, n_objectives), ylims=(ymin, ymax))
ND = filter(indiv -> indiv.rank == 1, P)
dominated = filter(indiv->indiv.rank != 1, P)
for i in dominated
plot!(1:n_objectives, collect(i.y), legend=:none, color=:blue)
end
for i in ND
plot!(1:n_objectives, collect(i.y), legend=:none, color=:orange)
end
sleep(2.0)
end
non_dominated = nsga_max(popsize, 50, objective, init, fCV=constraint,
fplot=plot_pop, plotevery=10, showprogress=true);
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success">
# <b>Exercise 4</b>
# <br/>
# Gradually increase $K$, the number of objectives, and watch the number of non-dominated individuals in the population over evolution. What do you notice as $K$ increases?
# <br/>
# </div>
# -
| 3_moo/2_NSGAII.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"} toc-hr-collapsed=false
# # Testing Configurations
#
# The behavior of a program is not only governed by its data. The _configuration_ of a program – that is, the settings that govern the execution of a program on its (regular) input data, as set by options or configuration files – just as well influences behavior, and thus can and should be tested. In this chapter, we explore how to systematically _test_ and _cover_ software configurations. By _automatically inferring configuration options_, we can apply these techniques out of the box, with no need for writing a grammar. Finally, we show how to systematically cover _combinations_ of configuration options, quickly detecting unwanted interferences.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# **Prerequisites**
#
# * You should have read the [chapter on grammars](Grammars.ipynb).
# * You should have read the [chapter on grammar coverage](GrammarCoverageFuzzer.ipynb).
# -
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from fuzzingbook.ConfigurationFuzzer import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter provides two classes:
#
# * `OptionRunner` automatically extract command-line options from a Python program;
# * `OptionFuzzer` uses these to automatically test a Python program with a large variety of options.
#
# `OptionRunner` runs a program up to the point where it parses its arguments, and then extracts a grammar that describes its invocations:
#
# ```python
# >>> autopep8_runner = OptionRunner("autopep8", "foo.py")
# ```
# The grammar can be extracted via the method `ebnf_grammar()`:
#
# ```python
# >>> option_ebnf_grammar = autopep8_runner.ebnf_grammar()
# >>> print(option_ebnf_grammar)
# {'<start>': ['(<option>)*<arguments>'], '<option>': [' -h', ' --help', ' --version', ' -v', ' --verbose', ' -d', ' --diff', ' -i', ' --in-place', ' --global-config <filename>', ' --ignore-local-config', ' -r', ' --recursive', ' -j <n>', ' --jobs <n>', ' -p <n>', ' --pep8-passes <n>', ' -a', ' --aggressive', ' --experimental', ' --exclude <globs>', ' --list-fixes', ' --ignore <errors>', ' --select <errors>', ' --max-line-length <n>', ' --line-range <line> <line>', ' --range <line> <line>', ' --indent-size <int>', ' --hang-closing'], '<arguments>': [' foo.py'], '<str>': ['<char>+'], '<char>': ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9', 'a', 'b', 'c', 'd', 'e', 'f', 'g', 'h', 'i', 'j', 'k', 'l', 'm', 'n', 'o', 'p', 'q', 'r', 's', 't', 'u', 'v', 'w', 'x', 'y', 'z', 'A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K', 'L', 'M', 'N', 'O', 'P', 'Q', 'R', 'S', 'T', 'U', 'V', 'W', 'X', 'Y', 'Z', '!', '"', '#', '$', '%', '&', "'", '(', ')', '*', '+', ',', '-', '.', '/', ':', ';', '<', '=', '>', '?', '@', '[', '\\', ']', '^', '_', '`', '{', '|', '}', '~'], '<filename>': ['<str>'], '<int>': ['(-)?<digit>+'], '<digit>': ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9'], '<n>': ['<int>'], '<globs>': ['<str>'], '<errors>': ['<str>'], '<line>': ['<int>']}
#
# ```
# The grammar can be immediately used for fuzzing. A `GrammarCoverageFuzzer` will ensure all options are covered:
#
# ```python
# >>> from Grammars import convert_ebnf_grammar
# >>> fuzzer = GrammarCoverageFuzzer(convert_ebnf_grammar(option_ebnf_grammar))
# >>> [fuzzer.fuzz() for i in range(3)]
# [' foo.py',
# ' --indent-size 54 --diff --global-config k --select &, --list-fixes -a --hang-closing --range 0 72 --ignore-local-config -p 8 --version -d --experimental foo.py',
# ' --ignore i --jobs -16 --verbose -v --line-range -3 9 -r --help --max-line-length 8 -h --aggressive --recursive --exclude qE" --in-place -j -979 -i --pep8-passes 4 --version --in-place --aggressive --version foo.py']
# ```
# The `OptionFuzzer` class summarizes these steps. Its constructor takes an `OptionRunner` to automatically extract the grammar; it does the necessary steps to extract the grammar and fuzz with it.
#
# ```python
# >>> autopep8_runner = OptionRunner("autopep8", "foo.py")
# >>> autopep8_fuzzer = OptionFuzzer(autopep8_runner)
# >>> [autopep8_fuzzer.fuzz() for i in range(3)]
# [' foo.py',
# ' --range 46 -1 --recursive -d --select <6 --exclude :" --global-config UVE --help --aggressive --experimental -r --line-range -7 -9 --version -i -h --indent-size -05 --max-line-length 8 --in-place --verbose --jobs -32 --ignore-local-config -v -p -1 --hang-closing -j 38 -a --list-fixes --pep8-passes 67 --diff --ignore v --select I --ignore (1NJ --ignore Km --ignore ? --select ^kZ --global-config y --select ia]9 --exclude o --ignore R!4GP.x8/ --ignore D --exclude 7 --exclude Bd -a --recursive --verbose foo.py',
# " --ignore \\ --global-config l --global-config @ --ignore ,CM~& --ignore nb --select c --global-config zgW --ignore $`s{H --global-config - --exclude 2| --select O --exclude 0 --exclude * --ignore qA'F}X --global-config p>_r+ --global-config eQ --exclude [ --ignore t --select h) --select %f --exclude u#3;=TL --global-config w --ignore j5 --exclude Y --ignore S --ignore ]J --global-config 1 --ignore-local-config --max-line-length 36693 -i foo.py"]
# ```
# The final step in testing would now to invoke the program with these arguments.
#
# Note that `OptionRunner` is experimental: It assumes that the Python program in question uses the `argparse` module; and not all `argparse` features are supported. Still, it does a pretty good job even on nontrivial programs.
#
#
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Configuration Options
#
# When we talk about the input to a program, we usually think of the _data_ it processes. This is also what we have been fuzzing in the past chapters – be it with [random input](Fuzzer.ipynb), [mutation-based fuzzing](MutationFuzzer.ipynb), or [grammar-based fuzzing](GrammarFuzzer.ipynb). However, programs typically have several input sources, all of which can and should be tested – and included in test generation.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# One important source of input is the program's _configuration_ – that is, a set of inputs that typically is set once when beginning to process data and then stays constant while processing data, while the program is running, or even while the program is deployed. Such a configuration is frequently set in _configuration files_ (for instance, as key/value pairs); the most ubiquitous method for command-line tools, though, are _configuration options_ on the command line.
# + [markdown] slideshow={"slide_type": "subslide"}
# As an example, consider the `grep` utility to find textual patterns in files. The exact mode by which `grep` works is governed by a multitude of options, which can be listed by providing a `--help` option:
# + slideshow={"slide_type": "fragment"}
# !grep --help
# + [markdown] slideshow={"slide_type": "fragment"}
# All these options need to be tested for whether they operate correctly. In security testing, any such option may also trigger a yet unknown vulnerability. Hence, such options can become _fuzz targets_ on their own. In this chapter, we analyze how to systematically test such options – and better yet, how to extract possible configurations right out of given program files, such that we do not have to specify anything.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Options in Python
#
# Let us stick to our common programming language here and examine how options are processed in Python. The `argparse` module provides a parser for command-line arguments (and options) with great functionality – and great complexity. You start by defining a parser (`argparse.ArgumentParser()`) to which individual arguments with various features are added, one after another. Additional parameters for each argument can specify the type (`type`) of the argument (say, integers or strings), or the number of arguments (`nargs`).
# + [markdown] slideshow={"slide_type": "subslide"}
# By default, arguments are stored under their name in the `args` object coming from `parse_args()` – thus, `args.integers` holds the `integers` arguments added earlier. Special actions (`actions`) allow to store specific values in given variables; the `store_const` action stores the given `const` in the attribute named by `dest`. The following example takes a number of integer arguments (`integers`) as well as an operator (`--sum`, `--min`, or `--max`) to be applied on these integers. The operators all store a function reference in the `accumulate` attribute, which is finally invoked on the integers parsed:
# + slideshow={"slide_type": "skip"}
import argparse
# + slideshow={"slide_type": "subslide"}
def process_numbers(args=[]):
parser = argparse.ArgumentParser(description='Process some integers.')
parser.add_argument('integers', metavar='N', type=int, nargs='+',
help='an integer for the accumulator')
group = parser.add_mutually_exclusive_group(required=True)
group.add_argument('--sum', dest='accumulate', action='store_const',
const=sum,
help='sum the integers')
group.add_argument('--min', dest='accumulate', action='store_const',
const=min,
help='compute the minimum')
group.add_argument('--max', dest='accumulate', action='store_const',
const=max,
help='compute the maximum')
args = parser.parse_args(args)
print(args.accumulate(args.integers))
# + [markdown] slideshow={"slide_type": "subslide"}
# Here's how `process_numbers()` works. We can, for instance, invoke the `--min` option on the given arguments to compute the minimum:
# + slideshow={"slide_type": "fragment"}
process_numbers(["--min", "100", "200", "300"])
# + [markdown] slideshow={"slide_type": "fragment"}
# Or compute the sum of three numbers:
# + slideshow={"slide_type": "fragment"}
process_numbers(["--sum", "1", "2", "3"])
# + [markdown] slideshow={"slide_type": "subslide"}
# When defined via `add_mutually_exclusive_group()` (as above), options are mutually exclusive. Consequently, we can have only one operator:
# + slideshow={"slide_type": "skip"}
import fuzzingbook_utils
# + slideshow={"slide_type": "skip"}
from ExpectError import ExpectError
# + slideshow={"slide_type": "fragment"}
with ExpectError(print_traceback=False):
process_numbers(["--sum", "--max", "1", "2", "3"])
# + [markdown] slideshow={"slide_type": "slide"}
# ## A Grammar for Configurations
#
# How can we test a system with several options? The easiest answer is to write a grammar for it. The grammar `PROCESS_NUMBERS_EBNF_GRAMMAR` reflects the possible combinations of options and arguments:
# + slideshow={"slide_type": "skip"}
from Grammars import crange, srange, convert_ebnf_grammar, extend_grammar, is_valid_grammar
from Grammars import START_SYMBOL, new_symbol
# + slideshow={"slide_type": "fragment"}
PROCESS_NUMBERS_EBNF_GRAMMAR = {
"<start>": ["<operator> <integers>"],
"<operator>": ["--sum", "--min", "--max"],
"<integers>": ["<integer>", "<integers> <integer>"],
"<integer>": ["<digit>+"],
"<digit>": crange('0', '9')
}
assert is_valid_grammar(PROCESS_NUMBERS_EBNF_GRAMMAR)
# + slideshow={"slide_type": "fragment"}
PROCESS_NUMBERS_GRAMMAR = convert_ebnf_grammar(PROCESS_NUMBERS_EBNF_GRAMMAR)
# + [markdown] slideshow={"slide_type": "subslide"}
# We can feed this grammar into our [grammar coverage fuzzer](GrammarCoverageFuzzer.ipynb) and have it cover one option after another:
# + slideshow={"slide_type": "skip"}
from GrammarCoverageFuzzer import GrammarCoverageFuzzer
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(PROCESS_NUMBERS_GRAMMAR, min_nonterminals=10)
for i in range(3):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# Of course, we can also invoke `process_numbers()` with these very arguments. To this end, we need to convert the string produced by the grammar back into a list of individual arguments, using `split()`:
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(PROCESS_NUMBERS_GRAMMAR, min_nonterminals=10)
for i in range(3):
args = f.fuzz().split()
print(args)
process_numbers(args)
# + [markdown] slideshow={"slide_type": "subslide"}
# In a similar way, we can define grammars for any program to be tested; as well as define grammars for, say, configuration files. Yet, the grammar has to be updated with every change to the program, which creates a maintenance burden. Given that the information required for the grammar is already all encoded in the program, the question arises: _Can't we go and extract configuration options right out of the program in the first place?_
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Mining Configuration Options
#
# In this section, we try to extract option and argument information right out of a program, such that we do not have to specify a configuration grammar. The aim is to have a configuration fuzzer that works on the options and arguments of an arbitrary program, as long as it follows specific conventions for processing its arguments. In the case of Python programs, this means using the `argparse` module.
#
# Our idea is as follows: We execute the given program up to the point where the arguments are actually parsed – that is, `argparse.parse_args()` is invoked. Up to this point, we track all calls into the argument parser, notably those calls that define arguments and options (`add_argument()`). From these, we construct the grammar.
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Tracking Arguments
#
# Let us illustrate this approach with a simple experiment: We define a trace function (see [our chapter on coverage](Coverage.ipynb) for details) that is active while `process_numbers` is invoked. If we have a call to a method `add_argument`, we access and print out the local variables (which at this point are the arguments to the method).
# + slideshow={"slide_type": "skip"}
import sys
# + slideshow={"slide_type": "skip"}
import string
# + slideshow={"slide_type": "fragment"}
def traceit(frame, event, arg):
if event != "call":
return
method_name = frame.f_code.co_name
if method_name != "add_argument":
return
locals = frame.f_locals
print(method_name, locals)
# + [markdown] slideshow={"slide_type": "subslide"}
# What we get is a list of all calls to `add_argument()`, together with the method arguments passed:
# + slideshow={"slide_type": "fragment"}
sys.settrace(traceit)
process_numbers(["--sum", "1", "2", "3"])
sys.settrace(None)
# + [markdown] slideshow={"slide_type": "subslide"}
# From the `args` argument, we can access the individual options and arguments to be defined:
# + slideshow={"slide_type": "fragment"}
def traceit(frame, event, arg):
if event != "call":
return
method_name = frame.f_code.co_name
if method_name != "add_argument":
return
locals = frame.f_locals
print(locals['args'])
# + slideshow={"slide_type": "fragment"}
sys.settrace(traceit)
process_numbers(["--sum", "1", "2", "3"])
sys.settrace(None)
# + [markdown] slideshow={"slide_type": "subslide"}
# We see that each argument comes as a tuple with one (say, `integers` or `--sum`) or two members (`-h` and `--help`), which denote alternate forms for the same option. Our job will be to go through the arguments of `add_arguments()` and detect not only the names of options and arguments, but also whether they accept additional parameters, as well as the type of the parameters.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### A Grammar Miner for Options and Arguments
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us now build a class that gathers all this information to create a grammar.
# + [markdown] slideshow={"slide_type": "fragment"}
# We use the `ParseInterrupt` exception to interrupt program execution after gathering all arguments and options:
# + slideshow={"slide_type": "fragment"}
class ParseInterrupt(Exception):
pass
# + [markdown] slideshow={"slide_type": "fragment"}
# The class `OptionGrammarMiner` takes an executable function for which the grammar of options and arguments is to be mined:
# + slideshow={"slide_type": "fragment"}
class OptionGrammarMiner(object):
def __init__(self, function, log=False):
self.function = function
self.log = log
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `mine_ebnf_grammar()` is where everything happens. It creates a grammar of the form
#
# ```
# <start> ::= <option>* <arguments>
# <option> ::= <empty>
# <arguments> ::= <empty>
# ```
#
# in which the options and arguments will be collected. It then sets a trace function (see [our chapter on coverage](Coverage.ipynb) for details) that is active while the previously defined `function` is invoked. Raising `ParseInterrupt` (when `parse_args()` is invoked) ends execution.
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
OPTION_SYMBOL = "<option>"
ARGUMENTS_SYMBOL = "<arguments>"
def mine_ebnf_grammar(self):
self.grammar = {
START_SYMBOL: ["(" + self.OPTION_SYMBOL + ")*" + self.ARGUMENTS_SYMBOL],
self.OPTION_SYMBOL: [],
self.ARGUMENTS_SYMBOL: []
}
self.current_group = self.OPTION_SYMBOL
old_trace = sys.gettrace()
sys.settrace(self.traceit)
try:
self.function()
except ParseInterrupt:
pass
sys.settrace(old_trace)
return self.grammar
def mine_grammar(self):
return convert_ebnf_grammar(self.mine_ebnf_grammar())
# + [markdown] slideshow={"slide_type": "subslide"}
# The trace function checks for four methods: `add_argument()` is the most important function, resulting in processing arguments; `frame.f_locals` again is the set of local variables, which at this point is mostly the arguments to `add_argument()`. Since mutually exclusive groups also have a method `add_argument()`, we set the flag `in_group` to differentiate.
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that we make no specific efforts to differentiate between multiple parsers or groups; we simply assume that there is one parser, and at any point at most one mutually exclusive group.
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
def traceit(self, frame, event, arg):
if event != "call":
return
if "self" not in frame.f_locals:
return
self_var = frame.f_locals["self"]
method_name = frame.f_code.co_name
if method_name == "add_argument":
in_group = repr(type(self_var)).find("Group") >= 0
self.process_argument(frame.f_locals, in_group)
elif method_name == "add_mutually_exclusive_group":
self.add_group(frame.f_locals, exclusive=True)
elif method_name == "add_argument_group":
# self.add_group(frame.f_locals, exclusive=False)
pass
elif method_name == "parse_args":
raise ParseInterrupt
return None
# + [markdown] slideshow={"slide_type": "subslide"}
# The `process_arguments()` now analyzes the arguments passed and adds them to the grammar:
#
# * If the argument starts with `-`, it gets added as an optional element to the `<option>` list
# * Otherwise, it gets added to the `<argument>` list.
#
# The optional `nargs` argument specifies how many arguments can follow. If it is a number, we add the appropriate number of elements to the grammar; if it is an abstract specifier (say, `+` or `*`), we use it directly as EBNF operator.
#
# Given the large number of parameters and optional behavior, this is a somewhat messy function, but it does the job.
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
def process_argument(self, locals, in_group):
args = locals["args"]
kwargs = locals["kwargs"]
if self.log:
print(args)
print(kwargs)
print()
for arg in args:
self.process_arg(arg, in_group, kwargs)
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
def process_arg(self, arg, in_group, kwargs):
if arg.startswith('-'):
if not in_group:
target = self.OPTION_SYMBOL
else:
target = self.current_group
metavar = None
arg = " " + arg
else:
target = self.ARGUMENTS_SYMBOL
metavar = arg
arg = ""
if "nargs" in kwargs:
nargs = kwargs["nargs"]
else:
nargs = 1
param = self.add_parameter(kwargs, metavar)
if param == "":
nargs = 0
if isinstance(nargs, int):
for i in range(nargs):
arg += param
else:
assert nargs in "?+*"
arg += '(' + param + ')' + nargs
if target == self.OPTION_SYMBOL:
self.grammar[target].append(arg)
else:
self.grammar[target].append(arg)
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `add_parameter()` handles possible parameters of options. If the argument has an `action` defined, it takes no parameter. Otherwise, we identify the type of the parameter (as `int` or `str`) and augment the grammar with an appropriate rule.
# + slideshow={"slide_type": "skip"}
import inspect
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
def add_parameter(self, kwargs, metavar):
if "action" in kwargs:
# No parameter
return ""
type_ = "str"
if "type" in kwargs:
given_type = kwargs["type"]
# int types come as '<class int>'
if inspect.isclass(given_type) and issubclass(given_type, int):
type_ = "int"
if metavar is None:
if "metavar" in kwargs:
metavar = kwargs["metavar"]
else:
metavar = type_
self.add_type_rule(type_)
if metavar != type_:
self.add_metavar_rule(metavar, type_)
param = " <" + metavar + ">"
return param
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `add_type_rule()` adds a rule for parameter types to the grammar. If the parameter is identified by a meta-variable (say, `N`), we add a rule for this as well to improve legibility.
# + slideshow={"slide_type": "fragment"}
class OptionGrammarMiner(OptionGrammarMiner):
def add_type_rule(self, type_):
if type_ == "int":
self.add_int_rule()
else:
self.add_str_rule()
def add_int_rule(self):
self.grammar["<int>"] = ["(-)?<digit>+"]
self.grammar["<digit>"] = crange('0', '9')
def add_str_rule(self):
self.grammar["<str>"] = ["<char>+"]
self.grammar["<char>"] = srange(
string.digits
+ string.ascii_letters
+ string.punctuation)
def add_metavar_rule(self, metavar, type_):
self.grammar["<" + metavar + ">"] = ["<" + type_ + ">"]
# + [markdown] slideshow={"slide_type": "subslide"}
# The method `add_group()` adds a new mutually exclusive group to the grammar. We define a new symbol (say, `<group>`) for the options added to the group, and use the `required` and `exclusive` flags to define an appropriate expansion operator. The group is then prefixed to the grammar, as in
#
# ```
# <start> ::= <group><option>* <arguments>
# <group> ::= <empty>
# ```
#
# and filled with the next calls to `add_argument()` within the group.
# + slideshow={"slide_type": "subslide"}
class OptionGrammarMiner(OptionGrammarMiner):
def add_group(self, locals, exclusive):
kwargs = locals["kwargs"]
if self.log:
print(kwargs)
required = kwargs.get("required", False)
group = new_symbol(self.grammar, "<group>")
if required and exclusive:
group_expansion = group
if required and not exclusive:
group_expansion = group + "+"
if not required and exclusive:
group_expansion = group + "?"
if not required and not exclusive:
group_expansion = group + "*"
self.grammar[START_SYMBOL][0] = group_expansion + \
self.grammar[START_SYMBOL][0]
self.grammar[group] = []
self.current_group = group
# + [markdown] slideshow={"slide_type": "subslide"}
# That's it! With this, we can now extract the grammar from our `process_numbers()` program. Turning on logging again reveals the variables we draw upon.
# + slideshow={"slide_type": "subslide"}
miner = OptionGrammarMiner(process_numbers, log=True)
process_numbers_grammar = miner.mine_ebnf_grammar()
# + [markdown] slideshow={"slide_type": "subslide"}
# Here is the extracted grammar:
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar
# + [markdown] slideshow={"slide_type": "subslide"}
# The grammar properly identifies the group found:
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<start>"]
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<group>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# It also identifies a `--help` option provided not by us, but by the `argparse` module:
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<option>"]
# + [markdown] slideshow={"slide_type": "subslide"}
# The grammar also correctly identifies the types of the arguments:
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<arguments>"]
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<integers>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# The rules for `int` are set as defined by `add_int_rule()`
# + slideshow={"slide_type": "fragment"}
process_numbers_grammar["<int>"]
# + [markdown] slideshow={"slide_type": "subslide"}
# We can take this grammar and convert it to BNF, such that we can fuzz with it right away:
# + slideshow={"slide_type": "fragment"}
assert is_valid_grammar(process_numbers_grammar)
# + slideshow={"slide_type": "fragment"}
grammar = convert_ebnf_grammar(process_numbers_grammar)
assert is_valid_grammar(grammar)
# + slideshow={"slide_type": "fragment"}
f = GrammarCoverageFuzzer(grammar)
for i in range(10):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "fragment"}
# Each and every invocation adheres to the rules as set forth in the `argparse` calls. By mining options and arguments from existing programs, we can now fuzz these options out of the box – without having to specify a grammar.
# + [markdown] slideshow={"slide_type": "slide"} toc-hr-collapsed=true
# ## Testing Autopep8
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us try out the option grammar miner on real-world Python programs. `autopep8` is a tool that automatically converts Python code to the [PEP 8 Style Guide for Python Code](https://www.python.org/dev/peps/pep-0008/). (Actually, all Python code in this book runs through `autopep8` during production.) `autopep8` offers a wide range of options, as can be seen by invoking it with `--help`:
# + slideshow={"slide_type": "subslide"}
# !autopep8 --help
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Autopep8 Setup
#
# We want to systematically test these options. In order to deploy our configuration grammar miner, we need to find the source code of the executable:
# + slideshow={"slide_type": "skip"}
import os
# + slideshow={"slide_type": "fragment"}
def find_executable(name):
for path in os.get_exec_path():
qualified_name = os.path.join(path, name)
if os.path.exists(qualified_name):
return qualified_name
return None
# + slideshow={"slide_type": "fragment"}
autopep8_executable = find_executable("autopep8")
assert autopep8_executable is not None
autopep8_executable
# + [markdown] slideshow={"slide_type": "subslide"}
# Next, we build a function that reads the contents of the file and executes it.
# + slideshow={"slide_type": "fragment"}
def autopep8():
executable = find_executable("autopep8")
# First line has to contain "/usr/bin/env python" or like
first_line = open(executable).readline()
assert first_line.find("python") >= 0
contents = open(executable).read()
exec(contents)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Mining an Autopep8 Grammar
#
# We can use the `autopep8()` function in our grammar miner:
# + slideshow={"slide_type": "fragment"}
autopep8_miner = OptionGrammarMiner(autopep8)
# + [markdown] slideshow={"slide_type": "fragment"}
# and extract a grammar for it:
# + slideshow={"slide_type": "fragment"}
autopep8_ebnf_grammar = autopep8_miner.mine_ebnf_grammar()
# + [markdown] slideshow={"slide_type": "fragment"}
# This works because here, `autopep8` is not a separate process (and a separate Python interpreter), but we run the `autopep8()` function (and the `autopep8` code) in our current Python interpreter – up to the call to `parse_args()`, where we interrupt execution again. At this point, the `autopep8` code has done nothing but setting up the argument parser – which is what we are interested in.
# + [markdown] slideshow={"slide_type": "subslide"}
# The grammar options mined reflect precisely the options seen when providing `--help`:
# + slideshow={"slide_type": "fragment"}
print(autopep8_ebnf_grammar["<option>"])
# + [markdown] slideshow={"slide_type": "subslide"}
# Metavariables like `<n>` or `<line>` are placeholders for integers. We assume all metavariables of the same name have the same type:
# + slideshow={"slide_type": "fragment"}
autopep8_ebnf_grammar["<line>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# The grammar miner has inferred that the argument to `autopep8` is a list of files:
# + slideshow={"slide_type": "fragment"}
autopep8_ebnf_grammar["<arguments>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# which in turn all are strings:
# + slideshow={"slide_type": "fragment"}
autopep8_ebnf_grammar["<files>"]
# + [markdown] slideshow={"slide_type": "subslide"}
# As we are only interested in testing options, not arguments, we fix the arguments to a single mandatory input. (Otherwise, we'd have plenty of random file names generated.)
# + slideshow={"slide_type": "fragment"}
autopep8_ebnf_grammar["<arguments>"] = [" <files>"]
autopep8_ebnf_grammar["<files>"] = ["foo.py"]
assert is_valid_grammar(autopep8_ebnf_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Creating Autopep8 Options
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us now use the inferred grammar for fuzzing. Again, we convert the EBNF grammar into a regular BNF grammar:
# + slideshow={"slide_type": "fragment"}
autopep8_grammar = convert_ebnf_grammar(autopep8_ebnf_grammar)
assert is_valid_grammar(autopep8_grammar)
# + [markdown] slideshow={"slide_type": "fragment"}
# And we can use the grammar for fuzzing all options:
# + slideshow={"slide_type": "subslide"}
f = GrammarCoverageFuzzer(autopep8_grammar, max_nonterminals=4)
for i in range(20):
print(f.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us apply these options on the actual program. We need a file `foo.py` that will serve as input:
# + slideshow={"slide_type": "fragment"}
def create_foo_py():
open("foo.py", "w").write("""
def twice(x = 2):
return x + x
""")
# + slideshow={"slide_type": "fragment"}
create_foo_py()
# + slideshow={"slide_type": "fragment"}
print(open("foo.py").read(), end="")
# + [markdown] slideshow={"slide_type": "fragment"}
# We see how `autopep8` fixes the spacing:
# + slideshow={"slide_type": "fragment"}
# !autopep8 foo.py
# + [markdown] slideshow={"slide_type": "subslide"}
# Let us now put things together. We define a `ProgramRunner` that will run the `autopep8` executable with arguments coming from the mined `autopep8` grammar.
# + slideshow={"slide_type": "skip"}
from Fuzzer import ProgramRunner
# + [markdown] slideshow={"slide_type": "fragment"}
# Running `autopep8` with the mined options reveals a surprisingly high number of passing runs. (We see that some options depend on each other or are mutually exclusive, but this is handled by the program logic, not the argument parser, and hence out of our scope.) The `GrammarCoverageFuzzer` ensures that each option is tested at least once. (Digits and letters, too, by the way.)
# + slideshow={"slide_type": "subslide"}
f = GrammarCoverageFuzzer(autopep8_grammar, max_nonterminals=5)
for i in range(20):
invocation = "autopep8" + f.fuzz()
print("$ " + invocation)
args = invocation.split()
autopep8 = ProgramRunner(args)
result, outcome = autopep8.run()
if result.stderr != "":
print(result.stderr, end="")
# + [markdown] slideshow={"slide_type": "subslide"}
# Our `foo.py` file now has been formatted in place a number of times:
# + slideshow={"slide_type": "fragment"}
print(open("foo.py").read(), end="")
# + [markdown] slideshow={"slide_type": "skip"}
# We don't need it anymore, so we clean up things:
# + slideshow={"slide_type": "skip"}
import os
# + slideshow={"slide_type": "skip"}
os.remove("foo.py")
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Classes for Fuzzing Configuration Options
#
# Let us now create reusable classes that we can use for testing arbitrary programs. (Okay, make that "arbitrary programs that are written in Python and use the `argparse` module to process command-line arguments.")
# + [markdown] slideshow={"slide_type": "subslide"}
# The class `OptionRunner` is a subclass of `ProgramRunner` that takes care of automatically determining the grammar, using the same steps we used for `autopep8`, above.
# + slideshow={"slide_type": "fragment"}
class OptionRunner(ProgramRunner):
def __init__(self, program, arguments=None):
if isinstance(program, str):
self.base_executable = program
else:
self.base_executable = program[0]
self.find_contents()
self.find_grammar()
if arguments is not None:
self.set_arguments(arguments)
super().__init__(program)
# + [markdown] slideshow={"slide_type": "subslide"}
# First, we find the contents of the Python executable:
# + slideshow={"slide_type": "fragment"}
class OptionRunner(OptionRunner):
def find_contents(self):
self._executable = find_executable(self.base_executable)
first_line = open(self._executable).readline()
assert first_line.find("python") >= 0
self.contents = open(self._executable).read()
def invoker(self):
exec(self.contents)
def executable(self):
return self._executable
# + [markdown] slideshow={"slide_type": "subslide"}
# Next, we determine the grammar using the `OptionGrammarMiner` class:
# + slideshow={"slide_type": "fragment"}
class OptionRunner(OptionRunner):
def find_grammar(self):
miner = OptionGrammarMiner(self.invoker)
self._ebnf_grammar = miner.mine_ebnf_grammar()
def ebnf_grammar(self):
return self._ebnf_grammar
def grammar(self):
return convert_ebnf_grammar(self._ebnf_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# The two service methods `set_arguments()` and `set_invocation()` help us to change the arguments and program, respectively.
# -
from Grammars import unreachable_nonterminals
# + slideshow={"slide_type": "fragment"}
class OptionRunner(OptionRunner):
def set_arguments(self, args):
self._ebnf_grammar["<arguments>"] = [" " + args]
# Delete rules for previous arguments
for nonterminal in unreachable_nonterminals(self._ebnf_grammar):
del self._ebnf_grammar[nonterminal]
def set_invocation(self, program):
self.program = program
# + [markdown] slideshow={"slide_type": "subslide"}
# We can instantiate the class on `autopep8` and immediately get the grammar:
# + slideshow={"slide_type": "fragment"}
autopep8_runner = OptionRunner("autopep8", "foo.py")
# + slideshow={"slide_type": "fragment"}
print(autopep8_runner.ebnf_grammar()["<option>"])
# + [markdown] slideshow={"slide_type": "subslide"}
# An `OptionFuzzer` interacts with the given `OptionRunner` to obtain its grammar, which is then passed to its `GrammarCoverageFuzzer` superclass.
# + slideshow={"slide_type": "fragment"}
class OptionFuzzer(GrammarCoverageFuzzer):
def __init__(self, runner, *args, **kwargs):
assert issubclass(type(runner), OptionRunner)
self.runner = runner
grammar = runner.grammar()
super().__init__(grammar, *args, **kwargs)
# + [markdown] slideshow={"slide_type": "subslide"}
# When invoking `run()`, the `OptionFuzzer` creates a new invocation (using `fuzz()` from its grammar) and runs the now given (or previously set) runner with the arguments from the grammar. Note that the runner specified in `run()` can differ from the one set during initialization; this allows for mining options from one program and applying it in another context.
# + slideshow={"slide_type": "fragment"}
class OptionFuzzer(OptionFuzzer):
def run(self, runner=None, inp=""):
if runner is None:
runner = self.runner
assert issubclass(type(runner), OptionRunner)
invocation = runner.executable() + " " + self.fuzz()
runner.set_invocation(invocation.split())
return runner.run(inp)
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Example: Autopep8
#
# Let us apply our newly defined classes on the `autopep8` runner:
# + slideshow={"slide_type": "fragment"}
autopep8_fuzzer = OptionFuzzer(autopep8_runner, max_nonterminals=5)
# + slideshow={"slide_type": "fragment"}
for i in range(3):
print(autopep8_fuzzer.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# We can now systematically test `autopep8` with these classes:
# + slideshow={"slide_type": "fragment"}
autopep8_fuzzer.run(autopep8_runner)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: MyPy
#
# We can extract options for the `mypy` static type checker for Python:
# + slideshow={"slide_type": "skip"}
assert find_executable("mypy") is not None
# + slideshow={"slide_type": "fragment"}
mypy_runner = OptionRunner("mypy", "foo.py")
print(mypy_runner.ebnf_grammar()["<option>"])
# + slideshow={"slide_type": "subslide"}
mypy_fuzzer = OptionFuzzer(mypy_runner, max_nonterminals=5)
for i in range(10):
print(mypy_fuzzer.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Notedown
#
# Here's the configuration options for the `notedown` Notebook to Markdown converter:
# + slideshow={"slide_type": "fragment"}
assert find_executable("notedown") is not None
# + slideshow={"slide_type": "fragment"}
notedown_runner = OptionRunner("notedown")
# + slideshow={"slide_type": "fragment"}
print(notedown_runner.ebnf_grammar()["<option>"])
# + slideshow={"slide_type": "fragment"}
notedown_fuzzer = OptionFuzzer(notedown_runner, max_nonterminals=5)
for i in range(10):
print(notedown_fuzzer.fuzz())
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Combinatorial Testing
#
# Our `CoverageGrammarFuzzer` does a good job in covering each and every option at least once, which is great for systematic testing. However, as we also can see in our examples above, some options require each other, while others interfere with each other. What we should do as good testers is not only to cover every option individually, but also _combinations_ of options.
# + [markdown] slideshow={"slide_type": "subslide"}
# The Python `itertools` module gives us means to create combinations from lists. We can, for instance, take the `notedown` options and create a list of all pairs.
# + slideshow={"slide_type": "skip"}
from itertools import combinations
# + slideshow={"slide_type": "fragment"}
option_list = notedown_runner.ebnf_grammar()["<option>"]
pairs = list(combinations(option_list, 2))
# + [markdown] slideshow={"slide_type": "fragment"}
# There's quite a number of pairs:
# + slideshow={"slide_type": "fragment"}
len(pairs)
# + slideshow={"slide_type": "fragment"}
print(pairs[:20])
# + [markdown] slideshow={"slide_type": "subslide"}
# Testing every such pair of options frequently suffices to cover all interferences between options. (Programs rarely have conditions involving three or more configuration settings.) To this end, we _change_ the grammar from having a list of options to having a list of _option pairs_, such that covering these will automatically cover all pairs.
# + [markdown] slideshow={"slide_type": "subslide"}
# We create a function `pairwise()` that takes a list of options as occurring in our grammar and returns a list of _pairwise options_ – that is, our original options, but concatenated.
# + slideshow={"slide_type": "fragment"}
def pairwise(option_list):
return [option_1 + option_2
for (option_1, option_2) in combinations(option_list, 2)]
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the first 20 pairs:
# + slideshow={"slide_type": "fragment"}
print(pairwise(option_list)[:20])
# + [markdown] slideshow={"slide_type": "subslide"}
# The new grammar `pairwise_notedown_grammar` is a copy of the `notedown` grammar, but with the list of options replaced with the above pairwise option list.
# + slideshow={"slide_type": "fragment"}
notedown_grammar = notedown_runner.grammar()
pairwise_notedown_grammar = extend_grammar(notedown_grammar)
pairwise_notedown_grammar["<option>"] = pairwise(notedown_grammar["<option>"])
assert is_valid_grammar(pairwise_notedown_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# Using the "pairwise" grammar to fuzz now covers one pair after another:
# + slideshow={"slide_type": "fragment"}
notedown_fuzzer = GrammarCoverageFuzzer(
pairwise_notedown_grammar, max_nonterminals=4)
# + slideshow={"slide_type": "fragment"}
for i in range(10):
print(notedown_fuzzer.fuzz())
# + [markdown] slideshow={"slide_type": "subslide"}
# Can we actually test all combinations of options? Not in practice, as the number of combinations quickly grows as the length increases. It decreases again as the number of options reaches the maximum (with 20 options, there is only 1 combination involving _all_ options), but the absolute numbers are still staggering:
# + slideshow={"slide_type": "fragment"}
for combination_length in range(1, 20):
tuples = list(combinations(option_list, combination_length))
print(combination_length, len(tuples))
# + [markdown] slideshow={"slide_type": "subslide"}
# Formally, the number of combinations of length $k$ in a set of options of length $n$ is the binomial coefficient
# $$
# {n \choose k} = \frac{n!}{k!(n - k)!}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# which for $k = 2$ (all pairs) gives us
#
# $$
# {n \choose 2} = \frac{n!}{2(n - 2)!} = n \times (n - 1)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# For `autopep8` with its 29 options...
# + slideshow={"slide_type": "fragment"}
len(autopep8_runner.ebnf_grammar()["<option>"])
# + [markdown] slideshow={"slide_type": "fragment"}
# ... we thus need 812 tests to cover all pairs:
# + slideshow={"slide_type": "fragment"}
len(autopep8_runner.ebnf_grammar()["<option>"]) * \
(len(autopep8_runner.ebnf_grammar()["<option>"]) - 1)
# + [markdown] slideshow={"slide_type": "subslide"}
# For `mypy` with its 110 options, though, we already end up with 11,990 tests to be conducted:
# + slideshow={"slide_type": "fragment"}
len(mypy_runner.ebnf_grammar()["<option>"])
# + slideshow={"slide_type": "fragment"}
len(mypy_runner.ebnf_grammar()["<option>"]) * \
(len(mypy_runner.ebnf_grammar()["<option>"]) - 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# Even if each pair takes a second to run, we'd still be done in three hours of testing, though.
# + [markdown] slideshow={"slide_type": "subslide"}
# If your program has more options that you all want to get covered in combinations, it is advisable that you limit the number of configurations further – for instance by limiting combinatorial testing to those combinations that possibly can interact with each other; and covering all other (presumably orthogonal) options individually.
# + [markdown] slideshow={"slide_type": "fragment"}
# This mechanism of creating configurations by extending grammars can be easily extended to other configuration targets. One may want to explore a greater number of configurations, or expansions in specific contexts. The [exercises](#Exercises), below, have a number of options ready for you.
# -
# ## Synopsis
#
# This chapter provides two classes:
#
# * `OptionRunner` automatically extract command-line options from a Python program;
# * `OptionFuzzer` uses these to automatically test a Python program with a large variety of options.
# `OptionRunner` runs a program up to the point where it parses its arguments, and then extracts a grammar that describes its invocations:
# + slideshow={"slide_type": "fragment"}
autopep8_runner = OptionRunner("autopep8", "foo.py")
# -
# The grammar can be extracted via the method `ebnf_grammar()`:
# + slideshow={"slide_type": "fragment"}
option_ebnf_grammar = autopep8_runner.ebnf_grammar()
print(option_ebnf_grammar)
# -
# The grammar can be immediately used for fuzzing. A `GrammarCoverageFuzzer` will ensure all options are covered:
from Grammars import convert_ebnf_grammar
fuzzer = GrammarCoverageFuzzer(convert_ebnf_grammar(option_ebnf_grammar))
[fuzzer.fuzz() for i in range(3)]
# The `OptionFuzzer` class summarizes these steps. Its constructor takes an `OptionRunner` to automatically extract the grammar; it does the necessary steps to extract the grammar and fuzz with it.
# + slideshow={"slide_type": "fragment"}
autopep8_runner = OptionRunner("autopep8", "foo.py")
autopep8_fuzzer = OptionFuzzer(autopep8_runner)
# + slideshow={"slide_type": "fragment"}
[autopep8_fuzzer.fuzz() for i in range(3)]
# -
# The final step in testing would now to invoke the program with these arguments.
# Note that `OptionRunner` is experimental: It assumes that the Python program in question uses the `argparse` module; and not all `argparse` features are supported. Still, it does a pretty good job even on nontrivial programs.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * Besides regular input data, program _configurations_ make an important testing target.
# * For a given program using a standard library to parse command-line options and arguments, one can automatically extract these and convert them into a grammar.
# * To cover not only single options, but combinations of options, one can expand the grammar to cover all pairs, or come up with even more ambitious targets.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# ## Next Steps
#
# If you liked the idea of mining a grammar from a program, do not miss:
#
# * [how to mine grammars for input data](GrammarMiner.ipynb)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
# Our next steps in the book focus on:
#
# * [how to parse and recombine inputs](Parser.ipynb)
# * [how to assign weights and probabilities to specific productions](ProbabilisticGrammarFuzzer.ipynb)
# * [how to simplify inputs that cause a failure](Reducer.ipynb)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# Although configuration data is just as likely to cause failures as other input data, it has received relatively little attention in test generation – possibly because, unlike "regular" input data, configuration data is not so much under control of external parties, and because, again unlike regular data, there is little variance in configurations. Creating models for software configurations and using these models for testing is commonplace, as is the idea of pairwise testing. For an overview, see \cite{Pezze2008}; for a discussion and comparison of state-of-the-art techniques, see \cite{Petke2015}.
#
# More specifically, \cite{Sutton2007} also discuss techniques to systematically cover command-line options. Dai et al. \cite{Dai2010} apply configuration fuzzing by changing variables associated with configuration files.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} toc-hr-collapsed=false
# ## Exercises
# + [markdown] slideshow={"slide_type": "subslide"} toc-hr-collapsed=true
# ### Exercise 1: #ifdef Configuration Fuzzing
#
# In C programs, the *C preprocessor* can be used to choose which code parts should be compiled and which ones should not. As an example, in the C code
#
# ```C
# #ifdef LONG_FOO
# long foo() { ... }
# #else
# int foo() { ... }
# #endif
# ```
#
# the compiler will compile the function `foo()` with return type`long` if the _preprocessor variable_ `LONG_FOO` is defined, and with return type `int` if not. Such preprocessor variables are either set in the source files (using `#define`, as in `#define LONG_FOO`) or on the C compiler command line (using `-D<variable>` or `-D<variable>=<value>`, as in `-DLONG_FOO`.
# + [markdown] slideshow={"slide_type": "subslide"}
# Such *conditional compilation* is used to configure C programs towards their environment. System-specific code can contain lots of conditional compilation. As an example, consider this excerpt of `xmlparse.c`, the XML parser that is part of the Python runtime library:
#
# ```c
# #if defined(_WIN32) && !defined(LOAD_LIBRARY_SEARCH_SYSTEM32)
# # define LOAD_LIBRARY_SEARCH_SYSTEM32 0x00000800
# #endif
#
# #if !defined(HAVE_GETRANDOM) && !defined(HAVE_SYSCALL_GETRANDOM) \
# && !defined(HAVE_ARC4RANDOM_BUF) && !defined(HAVE_ARC4RANDOM) \
# && !defined(XML_DEV_URANDOM) \
# && !defined(_WIN32) \
# && !defined(XML_POOR_ENTROPY)
# # error
# #endif
#
# #if !defined(TIOCSWINSZ) || defined(__SCO__) || defined(__UNIXWARE__)
# #define USE_SYSV_ENVVARS /* COLUMNS/LINES vs. TERMCAP */
# #endif
#
# #ifdef XML_UNICODE_WCHAR_T
# #define XML_T(x) (const wchar_t)x
# #define XML_L(x) L ## x
# #else
# #define XML_T(x) (const unsigned short)x
# #define XML_L(x) x
# #endif
#
# int fun(int x) { return XML_T(x); }
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# A typical configuration for the C preprocessor on the above code could be `cc -c -D_WIN32 -DXML_POOR_ENTROPY -DXML_UNICODE_WCHAR_T xmlparse.c`, defining the given preprocessor variables and selecting the appropriate code fragments.
# + [markdown] slideshow={"slide_type": "subslide"}
# Since the compiler can only compile one configuration at a time (implying that we can also only _test_ one resulting executable at a time), your task is to find out which of these configurations actually compile. To this end, proceed in three steps.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 1: Extract Preprocessor Variables
#
# Write a _function_ `cpp_identifiers()` that, given a set of lines (say, from `open(filename).readlines()`), extracts all preprocessor variables referenced in `#if` or `#ifdef` preprocessor instructions. Apply `ifdef_identifiers()` on the sample C input above, such that
#
# ```python
# cpp_identifiers(open("xmlparse.c").readlines())
# ```
#
# returns the set
#
# ```python
# {'_WIN32', 'LOAD_LIBRARY_SEARCH_SYSTEM32', 'HAVE_GETRANDOM', 'HAVE_SYSCALL_GETRANDOM', 'HAVE_ARC4RANDOM_BUF', ...}
#
# ```
# + [markdown] slideshow={"slide_type": "skip"}
# **Solution.** Let us start with creating a sample input file, `xmlparse.c`:
# + slideshow={"slide_type": "skip"}
filename = "xmlparse.c"
open(filename, "w").write(
"""
#if defined(_WIN32) && !defined(LOAD_LIBRARY_SEARCH_SYSTEM32)
# define LOAD_LIBRARY_SEARCH_SYSTEM32 0x00000800
#endif
#if !defined(HAVE_GETRANDOM) && !defined(HAVE_SYSCALL_GETRANDOM) \
&& !defined(HAVE_ARC4RANDOM_BUF) && !defined(HAVE_ARC4RANDOM) \
&& !defined(XML_DEV_URANDOM) \
&& !defined(_WIN32) \
&& !defined(XML_POOR_ENTROPY)
# error
#endif
#if !defined(TIOCSWINSZ) || defined(__SCO__) || defined(__UNIXWARE__)
#define USE_SYSV_ENVVARS /* COLUMNS/LINES vs. TERMCAP */
#endif
#ifdef XML_UNICODE_WCHAR_T
#define XML_T(x) (const wchar_t)x
#define XML_L(x) L ## x
#else
#define XML_T(x) (const unsigned short)x
#define XML_L(x) x
#endif
int fun(int x) { return XML_T(x); }
""");
# + [markdown] slideshow={"slide_type": "skip"}
# To find C preprocessor `#if` directives and preprocessor variables, we use regular expressions matching them.
# + slideshow={"slide_type": "skip"}
import re
# + slideshow={"slide_type": "skip"}
re_cpp_if_directive = re.compile(r"\s*#\s*(el)?if")
re_cpp_identifier = re.compile(r"[a-zA-Z_$]+")
# + slideshow={"slide_type": "skip"}
def cpp_identifiers(lines):
identifiers = set()
for line in lines:
if re_cpp_if_directive.match(line):
identifiers |= set(re_cpp_identifier.findall(line))
# These are preprocessor keywords
identifiers -= {"if", "ifdef", "ifndef", "defined"}
return identifiers
# + slideshow={"slide_type": "skip"}
cpp_ids = cpp_identifiers(open("xmlparse.c").readlines())
cpp_ids
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 2: Derive an Option Grammar
#
# With the help of `cpp_identifiers()`, create a grammar which has C compiler invocations with a list of options, where each option takes the form `-D<variable>` for a preprocessor variable `<variable>`. Using this grammar `cpp_grammar`, a fuzzer
#
# ```python
# g = GrammarCoverageFuzzer(cpp_grammar)
# ```
#
# would create C compiler invocations such as
#
# ```python
# [g.fuzz() for i in range(10)]
# ['cc -DHAVE_SYSCALL_GETRANDOM xmlparse.c',
# 'cc -D__SCO__ -DRANDOM_BUF -DXML_UNICODE_WCHAR_T -D__UNIXWARE__ xmlparse.c',
# 'cc -DXML_POOR_ENTROPY xmlparse.c',
# 'cc -DRANDOM xmlparse.c',
# 'cc -D_WIN xmlparse.c',
# 'cc -DHAVE_ARC xmlparse.c', ...]
# ```
# + [markdown] slideshow={"slide_type": "skip"}
# **Solution.** This is not very difficult:
# + slideshow={"slide_type": "skip"}
from Grammars import new_symbol
# + slideshow={"slide_type": "skip"}
cpp_grammar = {
"<start>": ["cc -c<options> " + filename],
"<options>": ["<option>", "<options><option>"],
"<option>": []
}
for id in cpp_ids:
s = new_symbol(cpp_grammar, "<" + id + ">")
cpp_grammar["<option>"].append(s)
cpp_grammar[s] = [" -D" + id]
cpp_grammar
# + slideshow={"slide_type": "skip"}
assert is_valid_grammar(cpp_grammar)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 3: C Preprocessor Configuration Fuzzing
#
# Using the grammar just produced, use a `GrammarCoverageFuzzer` to
#
# 1. Test each processor variable individually
# 2. Test each pair of processor variables, using `pairwise()`.
#
# What happens if you actually run the invocations?
# + [markdown] slideshow={"slide_type": "skip"}
# **Solution.** We can simply run the coverage fuzzer, as described above.
# + slideshow={"slide_type": "skip"}
g = GrammarCoverageFuzzer(cpp_grammar)
g.fuzz()
# -
from Fuzzer import ProgramRunner
for i in range(10):
invocation = g.fuzz()
print("$", invocation)
# subprocess.call(invocation, shell=True)
cc_runner = ProgramRunner(invocation.split(' '))
(result, outcome) = cc_runner.run()
print(result.stderr, end="")
# To test all pairs, we can use `pairwise()`:
pairwise_cpp_grammar = extend_grammar(cpp_grammar)
pairwise_cpp_grammar["<option>"] = pairwise(cpp_grammar["<option>"])
pairwise_cpp_grammar["<option>"][:10]
for i in range(10):
invocation = g.fuzz()
print("$", invocation)
# subprocess.call(invocation, shell=True)
cc_runner = ProgramRunner(invocation.split(' '))
(result, outcome) = cc_runner.run()
print(result.stderr, end="")
# + [markdown] slideshow={"slide_type": "skip"}
# Some of the compilation errors we get could be expected – for instance, defining `XML_UNICODE_WCHAR_T` when actually, the type is not supported in our environment. Other errors may not be expected – and it is these errors we would find through systematic configuration fuzzing, as described above.
# + [markdown] slideshow={"slide_type": "skip"}
# At the end, don't forget to clean up:
# + slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
os.remove("xmlparse.c")
# + slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
if os.path.exists("xmlparse.o"):
os.remove("xmlparse.o")
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} toc-hr-collapsed=true
# ### Exercise 2: .ini Configuration Fuzzing
#
# Besides command-line options, another important source of configurations are _configuration files_. In this exercise, we will consider the very simple configuration language provided by the Python `ConfigParser` module, which is very similar to what is found in Microsoft Windows _.ini_ files.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# The following example for a `ConfigParser` input file stems right from [the ConfigParser documentation](https://docs.python.org/3/library/configparser.html):
# ```
# [DEFAULT]
# ServerAliveInterval = 45
# Compression = yes
# CompressionLevel = 9
# ForwardX11 = yes
#
# [bitbucket.org]
# User = hg
#
# [topsecret.server.com]
# Port = 50022
# ForwardX11 = no
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# The above `ConfigParser` file can be created programmatically:
# + slideshow={"slide_type": "skip"}
import configparser
# + slideshow={"slide_type": "subslide"}
config = configparser.ConfigParser()
config['DEFAULT'] = {'ServerAliveInterval': '45',
'Compression': 'yes',
'CompressionLevel': '9'}
config['bitbucket.org'] = {}
config['bitbucket.org']['User'] = 'hg'
config['topsecret.server.com'] = {}
topsecret = config['topsecret.server.com']
topsecret['Port'] = '50022' # mutates the parser
topsecret['ForwardX11'] = 'no' # same here
config['DEFAULT']['ForwardX11'] = 'yes'
with open('example.ini', 'w') as configfile:
config.write(configfile)
with open('example.ini') as configfile:
print(configfile.read(), end="")
# + [markdown] slideshow={"slide_type": "subslide"}
# and be read in again:
# + cell_style="center" slideshow={"slide_type": "fragment"}
config = configparser.ConfigParser()
config.read('example.ini')
topsecret = config['topsecret.server.com']
topsecret['Port']
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 1: Read Configuration
#
# Using `configparser`, create a program reading in the above configuration file and accessing the individual elements.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 2: Create a Configuration Grammar
#
# Design a grammar that will automatically create configuration files suitable for your above program. Fuzz your program with it.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Part 3: Mine a Configuration Grammar
#
# By dynamically tracking the individual accesses to configuration elements, you can again extract a basic grammar from the execution. To this end, create a subclass of `ConfigParser` with a special method `__getitem__`:
# + slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
class TrackingConfigParser(configparser.ConfigParser):
def __getitem__(self, key):
print("Accessing", repr(key))
return super().__getitem__(key)
# + [markdown] slideshow={"slide_type": "fragment"}
# For a `TrackingConfigParser` object `p`, `p.__getitem__(key)` will be invoked whenever `p[key]` is accessed:
# + slideshow={"slide_type": "subslide"}
tracking_config_parser = TrackingConfigParser()
tracking_config_parser.read('example.ini')
section = tracking_config_parser['topsecret.server.com']
# + [markdown] slideshow={"slide_type": "fragment"}
# Using `__getitem__()`, as above, implement a tracking mechanism that, while your program accesses the read configuration, automatically saves options accessed and values read. Create a prototype grammar from these values; use it for fuzzing.
# + [markdown] slideshow={"slide_type": "skip"}
# At the end, don't forget to clean up:
# + slideshow={"slide_type": "skip"}
import os
# + slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
os.remove("example.ini")
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Left to the reader. Enjoy!
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} toc-hr-collapsed=true
# ### Exercise 3: Extracting and Fuzzing C Command-Line Options
#
# In C programs, the `getopt()` function are frequently used to process configuration options. A call
#
# ```
# getopt(argc, argv, "bf:")
# ```
#
# indicates that the program accepts two options `-b` and `-f`, with `-f` taking an argument (as indicated by the following colon).
# + [markdown] cell_style="center" slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# #### Part 1: Getopt Fuzzing
#
# Write a framework which, for a given C program, automatically extracts the argument to `getopt()` and derives a fuzzing grammar for it. There are multiple ways to achieve this:
#
# 1. Scan the program source code for occurrences of `getopt()` and return the string passed. (Crude, but should frequently work.)
# 2. Insert your own implementation of `getopt()` into the source code (effectively replacing `getopt()` from the runtime library), which outputs the `getopt()` argument and exits the program. Recompile and run.
# 3. (Advanced.) As above, but instead of changing the source code, hook into the _dynamic linker_ which at runtime links the program with the C runtime library. Set the library loading path (on Linux and Unix, this is the `LD_LIBRARY_PATH` environment variable) such that your own version of `getopt()` is linked first, and the regular libraries later. Executing the program (without recompiling) should yield the desired result.
#
# Apply this on `grep` and `ls`; report the resulting grammars and results.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Left to the reader. Enjoy hacking!
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# #### Part 2: Fuzzing Long Options in C
#
# Same as Part 1, but also hook into the GNU variant `getopt_long()`, which accepts "long" arguments with double dashes such as `--help`. Note that method 1, above, will not work here, since the "long" options are defined in a separately defined structure.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Left to the reader. Enjoy hacking!
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ### Exercise 4: Expansions in Context
#
# In our above option configurations, we have multiple symbols which all expand to the same integer. For instance, the `--line-range` option of `autopep8` takes two `<line>` parameters which both expand into the same `<int>` symbol:
#
# ```
# <option> ::= ... | --line-range <line> <line> | ...
# <line> ::= <int>
# <int> ::= (-)?<digit>+
# <digit> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
# ```
# + cell_style="center" slideshow={"slide_type": "subslide"}
autopep8_runner.ebnf_grammar()["<line>"]
# + cell_style="center" slideshow={"slide_type": "fragment"}
autopep8_runner.ebnf_grammar()["<int>"]
# + slideshow={"slide_type": "fragment"}
autopep8_runner.ebnf_grammar()["<digit>"]
# + [markdown] slideshow={"slide_type": "fragment"}
# Once the `GrammarCoverageFuzzer` has covered all variations of `<int>` (especially by covering all digits) for _one_ option, though, it will no longer strive to achieve such coverage for the next option. Yet, it could be desirable to achieve such coverage for each option separately.
# + [markdown] slideshow={"slide_type": "subslide"}
# One way to achieve this with our existing `GrammarCoverageFuzzer` is again to change the grammar accordingly. The idea is to _duplicate_ expansions – that is, to replace an expansion of a symbol $s$ with a new symbol $s'$ whose definition is duplicated from $s$. This way, $s'$ and $s$ are separate symbols from a coverage point of view and would be independently covered.
# + [markdown] slideshow={"slide_type": "subslide"}
# As an example, consider again the above `--line-range` option. If we want our tests to independently cover all elements of the two `<line>` parameters, we can duplicate the second `<line>` expansion into a new symbol `<line'>` with subsequent duplicated expansions:
# ```
# <option> ::= ... | --line-range <line> <line'> | ...
# <line> ::= <int>
# <line'> ::= <int'>
# <int> ::= (-)?<digit>+
# <int'> ::= (-)?<digit'>+
# <digit> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
# <digit'> ::= 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# Design a function `inline(grammar, symbol)` that returns a duplicate of `grammar` in which every occurrence of `<symbol>` and its expansions become separate copies. The above grammar could be a result of `inline(autopep8_runner.ebnf_grammar(), "<line>")`.
# + [markdown] slideshow={"slide_type": "subslide"}
# When copying, expansions in the copy should also refer to symbols in the copy. Hence, when expanding `<int>` in
#
# ```<int> ::= <int><digit>```
#
# make that
#
# ```<int> ::= <int><digit>
# <int'> ::= <int'><digit'>
# ```
#
# (and not `<int'> ::= <int><digit'>` or `<int'> ::= <int><digit>`).
# + [markdown] slideshow={"slide_type": "subslide"} solution2="hidden" solution2_first=true
# Be sure to add precisely one new set of symbols for each occurrence in the original grammar, and not to expand further in the presence of recursion.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Again, left to the reader. Enjoy!
| notebooks/ConfigurationFuzzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Open Data Cube Sandbox
#
# Welcome to the Open Data Cube (ODC) Sandbox. Here you can access an instance of the ODC that has data indexed and ready to go. This sandbox utilises data products available on AWS' Landsat public dataset bucket on S3 (see: https://docs.opendata.aws/landsat-pds/readme.html).
#
# #### Scenes indexed in yellow.
# <img src="images/indexed_areas.png">
#
#
# Those who code in Python can use this space to explore using the ODC API to interact with indexed products and the potential of accessing earth observation data on demand.
#
# And for the novice users we've developed prototype apps that allow you to explore any location in thw world in a simple graphical user interface that is using stored macro commands.
#
# #### ODC
#
# The [ODC](https://github.com/opendatacube) provides an integrated gridded data analysis environment capable of delivering decades of analysis ready earth observation satellite and related data from multiple satellite and other acquisition systems.
#
# The ODC initiative seeks to increase the value and impact of global Earth obervation satellite data by providing an open and freely accessible exploitation archectecture and to foster a community to develop, sustain, and grow the techonology and the breadth and depth of its application. To read more see the [ODC Initiative Whitepaper](https://github.com/opendatacube/documentation/blob/master/Roadmap/ODC%20White%20Paper.md)
#
# The ODC manual can be found at http://datacube-core.readthedocs.io/en/latest/.
#
# #### To use the notebook -> Click your mouse onto the cell below, and push ctrl+enter, or click the run button from the toolbox above.
# %store -r
# You have just installed our apps. In our app workflow, the first step is to select a case-study area. This app will compile a table of what data products are available over an area and for what timesteps.
#
# If you're familiar with Google Maps then this app will be easy to master, zoom in and out using the (+|-) buttons, once you've navigated to an area of interest.
#
# Please use the pan and zoom tools to find an area of interest and then select the rectangle draw tool from the toolbox panel and draw a rectangle over your desired case-study. You will then see a table of what products are available and for what dates.
#
# _Note that you should select a reasonably small area, otherwise it will take a long time to load._
#
# ## Define an area of interest
#
# To define your area of interest in space and time, do the following:
#
# * Click your mouse the cell below, and push ctrl+enter, or click the run button from the toolbox above.
# * Pan and zoom until you have found an area of interest. Use the drawing tools on the left of the map to draw an outline of your area of interest
# * Click back on the first point of your area of interest to finish drawing
# * Select a date range for satellite data using the blue slider. This will determine how much data will be loaded for your case study.
select_case_study_area
#
# <style>
# table {margin-left: 0 !important;}
# </style>
# You can use any of our apps to explore the area by just typing any of the paragraph heading in a juptyter notebook cell.
#
# |App|Description|Data Product|
# |----|:-----|----|
# |`band_indices_aws_landsat8`|Enables querying the aws landsat 8 product with points/polygons, and visualising responses as band indices derived on line graphs.|
# |`water_stackplot_aws_landsat8`|Enables querying the aws landsat 8 product with points/polygons and classifier landsat cells that water.
#
# ### Test out the apps with the following cells
band_indices_aws_landsat8
water_stackplot_aws_landsat8
water_max_min_aws_landsat8
| Landing_Page.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anamic]
# language: python
# name: conda-env-anamic-py
# ---
# ### Microtubule Simulator
#
# Show how to build and save informations of **ONE** microtubule.
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from pathlib import Path
import sys
sys.path.append("../")
import anamic
import numpy as np
import matplotlib.pyplot as plt
# +
n_pf = 11
mt_length_nm = 2000 # nm
taper_length_nm = 200 # nm
dimers = anamic.simulator.dimers_builder(n_pf, mt_length_nm, taper_length_nm)
# Set parameters for the image generation.
parameters = {}
parameters['labeling_ratio'] = 0.1 # from 0 to 1
parameters['pixel_size'] = 110 # nm/pixel
parameters['x_offset'] = 1500 # nm
parameters['y_offset'] = 1500 # nm
parameters['psf_size'] = 135 # nm
parameters['signal_mean'] = 700
parameters['signal_std'] = 100
parameters['bg_mean'] = 500
parameters['bg_std'] = 24
parameters['noise_factor'] = 1
parameters['snr_line_width'] = 3 # pixel
ms = anamic.simulator.MicrotubuleSimulator(dimers)
ms.parameters.update(parameters)
# Build the geometry.
ms.build_positions(apply_random_z_rotation=True, show_progress=True)
ms.label()
ms.project()
ms.random_rotation_projected()
# Generate the image
ms.discretize_position()
ms.convolve()
snr = ms.calculate_snr()
print(f"SNR is {snr:.02f}")
# Save image, parameters and dimer positions.
#ms.save_positions("/home/hadim/test.csv")
#ms.save_metadata("/home/hadim/md.json")
#ms.save_image("/home/hadim/md.tif")
# -
# Show 3D position
_ = ms.show_positions(size=6, color_feature_name='pf', backend='ipv')
# Show final generated image
ms.show_image(tip_marker_size=80)
# Show projected 2D positions
_ = ms.visualize_2d_positions('x_proj', 'y_proj', show_all=True, show_labeled=True,
color_feature='pf', marker_size=30, x_offset=400)
# Show projected 2D positions after rotation
_ = ms.visualize_2d_positions('y_proj_rotated', 'x_proj_rotated', show_all=True, show_labeled=False,
color_feature='pf', marker_size=20, x_offset=0)
| notebooks/Howto/1_Simulate_One_Microtubule.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import os
qprc_path = os.path.abspath(os.path.join('..'))
if qprc_path not in sys.path:
sys.path.append(qprc_path)
# # A Support Fuzzy-sets Machine, i.e., a SVM with kernel on fuzzy sets
#
# This notebook contains:
# * A data fuzzification procedure
# * kernel on fuzzy sets estimation via the cross-product kernel in fuzzy sets
# * experimental validation with a SVM
# ## Iris dataset
# +
from sklearn.datasets import load_iris
from sklearn.preprocessing import MinMaxScaler
import pandas as pd
import numpy as np
import seaborn as sns
from kernelfuzzy.fuzzyset import FuzzySet
from kernelfuzzy.fuzzification import FuzzyData
#from https://stackoverflow.com/questions/38105539/how-to-convert-a-scikit-learn-dataset-to-a-pandas-dataset/46379878#46379878
def sklearn_to_df(sklearn_dataset):
df = pd.DataFrame(sklearn_dataset.data, columns=sklearn_dataset.feature_names)
df['target'] = pd.Series(sklearn_dataset.target)
return df
#data to pandas dataframe
iris_data = sklearn_to_df(load_iris())
iris_data.head()
#data fuzzification
iris_fuzzy_data = FuzzyData(data=iris_data, target='target')
iris_fuzzy_data.quantile_fuzzification_classification()
#printing epistemic values given my the membsership degrees of the values of the dataset
ax = sns.heatmap(iris_fuzzy_data.get_epistemic_values(), cmap="coolwarm")
# -
# ## Kernel Matrices of the fuzzified version of the Iris dataset
# +
#kernel gram matrix
from sklearn.metrics.pairwise import rbf_kernel, linear_kernel
from kernelfuzzy.kernels import cross_product_kernel, gram_matrix_cross_product_kernel
import matplotlib.pyplot as plt
kernel_bandwidth = [0.05, 0.5, 5, 50]
fig, axn = plt.subplots(2, 2,figsize=(10,10))
for i, ax in enumerate(axn.flat):
K = gram_matrix_cross_product_kernel(iris_fuzzy_data.get_fuzzydata(),
iris_fuzzy_data.get_fuzzydata(),
rbf_kernel,kernel_bandwidth[i],
linear_kernel,'')
sns.heatmap(K, ax=ax)
fig.tight_layout()
# -
# ## A Support fuzzy-set Machine
# +
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
X = iris_fuzzy_data.get_fuzzydata()
y = iris_fuzzy_data.get_target()
from sklearn.model_selection import ShuffleSplit
rs = ShuffleSplit(n_splits=5, test_size=.25, random_state=0)
rs.get_n_splits(X)
acc = []
for train_index, test_index in rs.split(X):
#print("TRAIN:", train_index, "TEST:", test_index)
X_train = X[train_index]
y_train = y[train_index]
X_test = X[test_index]
y_test = y[test_index]
svc_p = SVC(kernel='precomputed')
#training
K_train = gram_matrix_cross_product_kernel(X_train,
X_train,
rbf_kernel,50,
rbf_kernel,50)
sns.heatmap(K_train)
plt.show()
svc_p.fit(K_train,y_train)
#testing
K_test = gram_matrix_cross_product_kernel(X_test,
X_train,
rbf_kernel,50,
rbf_kernel,50)
sns.heatmap(K_test)
plt.show()
y_pred=svc_p.predict(K_test)
acc_test = accuracy_score(y_test, y_pred)
acc.append(acc_test)
print("Test accuracy: {}".format(acc_test))
print("Results: mean: ", np.mean(acc)," std: ", np.std(acc))
# -
| notebooks/5-Support Fuzzy-Set Machines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import shutil
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
data_train_dir = '/kaggle/input/fingers/fingers/train/'
data_test_dir = '/kaggle/input/fingers/fingers/test/'
train_dir = '/kaggle/working/train_dir'
val_dir = '/kaggle/working/val_dir'
test_dir = '/kaggle/working/test_dir'
def create_directory(dir_name):
dirs = [os.path.join(dir_name, "L"), os.path.join(dir_name, "R")]
if os.path.exists(dir_name):
shutil.rmtree(dir_name)
os.makedirs(dir_name)
for dir in dirs:
for i in range(6):
os.makedirs(dir + str(i))
create_directory(train_dir)
create_directory(val_dir)
create_directory(test_dir)
paths_train = []
paths_test = []
count_data = {'0L': 0,
'1L': 0,
'2L': 0,
'3L': 0,
'4L': 0,
'5L': 0,
'0R': 0,
'1R': 0,
'2R': 0,
'3R': 0,
'4R': 0,
'5R': 0}
# +
for dirname, _, filenames in os.walk(data_train_dir):
for filename in filenames:
paths_train.append(os.path.join(dirname, filename))
for dirname, _, filenames in os.walk(data_test_dir):
for filename in filenames:
paths_test.append(os.path.join(dirname, filename))
# +
for filename in paths_train:
for key in count_data.keys():
if filename[-6:-4] == key:
count_data[key] += 1
for filename in paths_test:
for key in count_data.keys():
if filename[-6:-4] == key:
count_data[key] += 1
# -
counts = count_data.values()
groups = count_data.keys()
def copy_images(start_index, end_index, paths, dest_dir):
for i in range(start_index, end_index):
dest_path = os.path.join(dest_dir, paths[i][-5] + paths[i][-6])
shutil.copy2(paths[i], dest_path)
# Part of the validation data set
validation_data_proportion = 0.15
# copying images from input directory to output
copy_images(0, len(paths_train) - int(validation_data_proportion * len(paths_train)),
paths_train, train_dir)
copy_images(len(paths_train) - int(validation_data_proportion * len(paths_train)),
len(paths_train), paths_train, val_dir)
copy_images(0, len(paths_test), paths_test, test_dir)
| .ipynb_checkpoints/finger-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://githubtocolab.com/giswqs/leafmap/blob/master/examples/notebooks/08_whitebox.ipynb)
# [](https://gishub.org/leafmap-binder)
#
# **Using WhiteboxTools with leafmap**
#
# Uncomment the following line to install [leafmap](https://leafmap.org) if needed.
# +
# # !pip install leafmap
# -
import os
import leafmap
# Download a sample DEM dataset.
# +
out_dir = os.path.expanduser('~/Downloads')
dem = os.path.join(out_dir, 'dem.tif')
if not os.path.exists(dem):
dem_url = 'https://drive.google.com/file/d/1vRkAWQYsLWCi6vcTMk8vLxoXMFbdMFn8/view?usp=sharing'
leafmap.download_from_gdrive(dem_url, 'dem.tif', out_dir, unzip=False)
# -
# Create an interactive map.
Map = leafmap.Map()
Map
# Use the built-in toolbox to perform geospatial analysis. For example, you can perform depression filling using the sample DEM dataset downloaded in the above step.
# 
# Display the toolbox using the default mode.
leafmap.whiteboxgui()
# Display the toolbox using the collapsible tree mode. Note that the tree mode does not support Google Colab.
leafmap.whiteboxgui(tree=True)
# Perform geospatial analysis using the [whitebox](https://github.com/giswqs/whitebox-python) package.
import os
import pkg_resources
wbt = leafmap.WhiteboxTools()
wbt.verbose = False
# +
# identify the sample data directory of the package
data_dir = os.path.dirname(pkg_resources.resource_filename("whitebox", 'testdata/'))
wbt.set_working_dir(data_dir)
wbt.feature_preserving_smoothing("DEM.tif", "smoothed.tif", filter=9)
wbt.breach_depressions("smoothed.tif", "breached.tif")
| examples/notebooks/08_whitebox.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3
# language: python
# name: python3
# ---
# # Loop API Example on Hartmann6
#
# The loop API is the most lightweight way to do optimization in Ax. The user makes one call to `optimize`, which performs all of the optimization under the hood and returns the optimized parameters.
#
# For more customizability of the optimization procedure, consider the Service or Developer API.
# +
import numpy as np
from ax.plot.contour import plot_contour
from ax.plot.trace import optimization_trace_single_method
from ax.service.managed_loop import optimize
from ax.metrics.branin import branin
from ax.utils.measurement.synthetic_functions import hartmann6
from ax.utils.notebook.plotting import render, init_notebook_plotting
init_notebook_plotting()
# -
# ## 1. Define evaluation function
#
# First, we define an evaluation function that is able to compute all the metrics needed for this experiment. This function needs to accept a set of parameter values and can also accept a weight. It should produce a dictionary of metric names to tuples of mean and standard error for those metrics.
def hartmann_evaluation_function(parameterization):
x = np.array([parameterization.get(f"x{i+1}") for i in range(6)])
# In our case, standard error is 0, since we are computing a synthetic function.
return {"hartmann6": (hartmann6(x), 0.0), "l2norm": (np.sqrt((x ** 2).sum()), 0.0)}
# If there is only one metric in the experiment – the objective – then evaluation function can return a single tuple of mean and SEM, in which case Ax will assume that evaluation corresponds to the objective. It can also return only the mean as a float, in which case Ax will treat SEM as unknown and use a model that can infer it. For more details on evaluation function, refer to the "Trial Evaluation" section in the docs.
# ## 2. Run optimization
# The setup for the loop is fully compatible with JSON. The optimization algorithm is selected based on the properties of the problem search space.
best_parameters, values, experiment, model = optimize(
parameters=[
{
"name": "x1",
"type": "range",
"bounds": [0.0, 1.0],
"value_type": "float", # Optional, defaults to inference from type of "bounds".
"log_scale": False, # Optional, defaults to False.
},
{
"name": "x2",
"type": "range",
"bounds": [0.0, 1.0],
},
{
"name": "x3",
"type": "range",
"bounds": [0.0, 1.0],
},
{
"name": "x4",
"type": "range",
"bounds": [0.0, 1.0],
},
{
"name": "x5",
"type": "range",
"bounds": [0.0, 1.0],
},
{
"name": "x6",
"type": "range",
"bounds": [0.0, 1.0],
},
],
experiment_name="test",
objective_name="hartmann6",
evaluation_function=hartmann_evaluation_function,
minimize=True, # Optional, defaults to False.
parameter_constraints=["x1 + x2 <= 20"], # Optional.
outcome_constraints=["l2norm <= 1.25"], # Optional.
total_trials=30, # Optional.
)
# And we can introspect optimization results:
best_parameters
means, covariances = values
means
# For comparison, minimum of Hartmann6 is:
hartmann6.fmin
# ## 3. Plot results
# Here we arbitrarily select "x1" and "x2" as the two parameters to plot for both metrics, "hartmann6" and "l2norm".
render(plot_contour(model=model, param_x='x1', param_y='x2', metric_name='hartmann6'))
render(plot_contour(model=model, param_x='x1', param_y='x2', metric_name='l2norm'))
# We also plot optimization trace, which shows best hartmann6 objective value seen by each iteration of the optimization:
# `plot_single_method` expects a 2-d array of means, because it expects to average means from multiple
# optimization runs, so we wrap out best objectives array in another array.
best_objectives = np.array([[trial.objective_mean for trial in experiment.trials.values()]])
best_objective_plot = optimization_trace_single_method(
y=np.minimum.accumulate(best_objectives, axis=1),
optimum=hartmann6.fmin,
title="Model performance vs. # of iterations",
ylabel="Hartmann6",
)
render(best_objective_plot)
| tutorials/gpei_hartmann_loop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hide_input=true
# # Computer vision data
# + hide_input=true
# %matplotlib inline
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
# -
# This module contains the classes that define datasets handling [`Image`](/vision.image.html#Image) objects and their tranformations. As usual, we'll start with a quick overview, before we get in to the detailed API docs.
# ## Quickly get your data ready for training
# To get you started as easily as possible, the fastai provides two helper functions to create a [`DataBunch`](/basic_data.html#DataBunch) object that you can directly use for training a classifier. To demonstrate them you'll first need to download and untar the file by executing the following cell. This will create a data folder containing an MNIST subset in `data/mnist_sample`.
path = untar_data(URLs.MNIST_SAMPLE); path
# There are a number of ways to create an [`ImageDataBunch`](/vision.data.html#ImageDataBunch). One common approach is to use *Imagenet-style folders* (see a ways down the page below for details) with [`ImageDataBunch.from_folder`](/vision.data.html#ImageDataBunch.from_folder):
tfms = get_transforms(do_flip=False)
data = ImageDataBunch.from_folder(path, ds_tfms=tfms, size=24)
# Here the datasets will be automatically created in the structure of *Imagenet-style folders*. The parameters specified:
# - the transforms to apply to the images in `ds_tfms` (here with `do_flip`=False because we don't want to flip numbers),
# - the target `size` of our pictures (here 24).
#
# As with all [`DataBunch`](/basic_data.html#DataBunch) usage, a `train_dl` and a `valid_dl` are created that are of the type PyTorch [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader).
#
# If you want to have a look at a few images inside a batch, you can use [`ImageDataBunch.show_batch`](/vision.data.html#ImageDataBunch.show_batch). The `rows` argument is the number of rows and columns to display.
data.show_batch(rows=3, figsize=(5,5))
# The second way to define the data for a classifier requires a structure like this:
# ```
# path\
# train\
# test\
# labels.csv
# ```
# where the labels.csv file defines the label(s) of each image in the training set. This is the format you will need to use when each image can have multiple labels. It also works with single labels:
pd.read_csv(path/'labels.csv').head()
# You can then use [`ImageDataBunch.from_csv`](/vision.data.html#ImageDataBunch.from_csv):
data = ImageDataBunch.from_csv(path, ds_tfms=tfms, size=28)
data.show_batch(rows=3, figsize=(5,5))
# An example of multiclassification can be downloaded with the following cell. It's a sample of the [planet dataset](https://www.google.com/search?q=kaggle+planet&rlz=1C1CHBF_enFR786FR786&oq=kaggle+planet&aqs=chrome..69i57j0.1563j0j7&sourceid=chrome&ie=UTF-8).
planet = untar_data(URLs.PLANET_SAMPLE)
# If we open the labels files, we seach that each image has one or more tags, separated by a space.
df =pd.read_csv(planet/'labels.csv')
df.head()
data = ImageDataBunch.from_csv(planet, folder='train', size=128, suffix='.jpg', sep=' ',
ds_tfms=get_transforms(flip_vert=True, max_lighting=0.1, max_zoom=1.05, max_warp=0.))
# The `show_batch`method will then print all the labels that correspond to each image.
data.show_batch(rows=3, figsize=(10,8), ds_type=DatasetType.Valid)
# You can find more ways to build an [`ImageDataBunch`](/vision.data.html#ImageDataBunch) without the factory methods in [`data_block`](/data_block.html#data_block).
# + hide_input=true
show_doc(ImageDataBunch)
# -
# This is the same initilialization as a regular [`DataBunch`](/basic_data.html#DataBunch) so you probably don't want to use this directly, but one of the factory methods instead.
# ### Factory methods
# If you quickly want to get a [`ImageDataBunch`](/vision.data.html#ImageDataBunch) and train a model, you should process your data to have it in one of the formats the following functions handle.
# + hide_input=true
show_doc(ImageDataBunch.from_folder)
# -
# "*Imagenet-style*" datasets look something like this (note that the test folder is optional):
#
# ```
# path\
# train\
# clas1\
# clas2\
# ...
# valid\
# clas1\
# clas2\
# ...
# test\
# ```
#
# For example:
data = ImageDataBunch.from_folder(path, ds_tfms=tfms, size=24)
# Note that this (and all factory methods in this section) pass any `kwargs` to [`ImageDataBunch.create`](/vision.data.html#ImageDataBunch.create).
# + hide_input=true
show_doc(ImageDataBunch.from_csv)
# -
# Create [`ImageDataBunch`](/vision.data.html#ImageDataBunch) from `path` by splitting the data in `folder` and labelled in a file `csv_labels` between a training and validation set. Use `valid_pct` to indicate the percentage of the total images for the validation set. An optional `test` folder contains unlabelled data and `suffix` contains an optional suffix to add to the filenames in `csv_labels` (such as '.jpg').
# For example:
data = ImageDataBunch.from_csv(path, ds_tfms=tfms, size=24);
# + hide_input=true
show_doc(ImageDataBunch.from_df)
# -
# Same as [`ImageDataBunch.from_csv`](/vision.data.html#ImageDataBunch.from_csv), but passing in a `DataFrame` instead of a csv file. E.gL
df = pd.read_csv(path/'labels.csv', header='infer')
df.head()
data = ImageDataBunch.from_df(path, df, ds_tfms=tfms, size=24)
# Different datasets are labeled in many different ways. The following methods can help extract the labels from the dataset in a wide variety of situations. The way they are built in fastai is constructive: there are methods which do a lot for you but apply in specific circumstances and there are methods which do less for you but give you more flexibility.
#
# In this case the hierachy is:
#
# 1. [`ImageDataBunch.from_name_re`](/vision.data.html#ImageDataBunch.from_name_re): Gets the labels from the filenames using a regular expression
# 2. [`ImageDataBunch.from_name_func`](/vision.data.html#ImageDataBunch.from_name_func): Gets the labels from the filenames using any function
# 3. [`ImageDataBunch.from_lists`](/vision.data.html#ImageDataBunch.from_lists): Labels need to be provided as an input in a list
# + hide_input=true
show_doc(ImageDataBunch.from_name_re)
# -
# Creates an [`ImageDataBunch`](/vision.data.html#ImageDataBunch) from `fnames`, calling a regular expression (containing one *re group*) on the file names to get the labels, putting aside `valid_pct` for the validation. In the same way as [`ImageDataBunch.from_csv`](/vision.data.html#ImageDataBunch.from_csv), an optional `test` folder contains unlabelled data.
#
# Our previously created dataframe contains the labels in the filenames so we can leverage it to test this new method. [`ImageDataBunch.from_name_re`](/vision.data.html#ImageDataBunch.from_name_re) needs the exact path of each file so we will append the data path to each filename before creating our [`ImageDataBunch`](/vision.data.html#ImageDataBunch) object.
fn_paths = [path/name for name in df['name']]; fn_paths[:2]
pat = r"/(\d)/\d+\.png$"
data = ImageDataBunch.from_name_re(path, fn_paths, pat=pat, ds_tfms=tfms, size=24)
data.classes
# + hide_input=true
show_doc(ImageDataBunch.from_name_func)
# -
# Works in the same way as [`ImageDataBunch.from_name_re`](/vision.data.html#ImageDataBunch.from_name_re), but instead of a regular expression it expects a function that will determine how to extract the labels from the filenames. (Note that `from_name_re` uses this function in its implementation).
#
# To test it we could build a function with our previous regex. Let's try another, similar approach to show that the labels can be obtained in a different way.
def get_labels(file_path): return '3' if '/3/' in str(file_path) else '7'
data = ImageDataBunch.from_name_func(path, fn_paths, label_func=get_labels, ds_tfms=tfms, size=24)
data.classes
# + hide_input=true
show_doc(ImageDataBunch.from_lists)
# -
# The most flexible factory function; pass in a list of `labels` that correspond to each of the filenames in `fnames`.
#
# To show an example we have to build the labels list outside our [`ImageDataBunch`](/vision.data.html#ImageDataBunch) object and give it as an argument when we call `from_lists`. Let's use our previously created function to create our labels list.
labels_ls = list(map(get_labels, fn_paths))
data = ImageDataBunch.from_lists(path, fn_paths, labels=labels_ls, ds_tfms=tfms, size=24)
data.classes
# + hide_input=true
show_doc(ImageDataBunch.create_from_ll)
# -
# Use `bs`, `num_workers`, `collate_fn` and a potential `test` folder. `ds_tfms` is a tuple of two lists of transforms to be applied to the training and the validation (plus test optionally) set. `tfms` are the transforms to apply to the [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader). The `size` and the `kwargs` are passed to the transforms for data augmentation.
# + hide_input=true
show_doc(ImageDataBunch.single_from_classes)
# + hide_input=true
jekyll_note('This method is deprecated, you should use DataBunch.load_empty now.')
# -
# ### Methods
# In the next two methods we will use a new dataset, CIFAR. This is because the second method will get the statistics for our dataset and we want to be able to show different statistics per channel. If we were to use MNIST, these statistics would be the same for every channel. White pixels are [255,255,255] and black pixels are [0,0,0] (or in normalized form [1,1,1] and [0,0,0]) so there is no variance between channels.
path = untar_data(URLs.CIFAR); path
# + hide_input=true
show_doc(channel_view)
# -
data = ImageDataBunch.from_folder(path, ds_tfms=tfms, valid='test', size=24)
def channel_view(x:Tensor)->Tensor:
"Make channel the first axis of `x` and flatten remaining axes"
return x.transpose(0,1).contiguous().view(x.shape[1],-1)
# This function takes a tensor and flattens all dimensions except the channels, which it keeps as the first axis. This function is used to feed [`ImageDataBunch.batch_stats`](/vision.data.html#ImageDataBunch.batch_stats) so that it can get the pixel statistics of a whole batch.
#
# Let's take as an example the dimensions our MNIST batches: 128, 3, 24, 24.
t = torch.Tensor(128, 3, 24, 24)
t.size()
tensor = channel_view(t)
tensor.size()
# + hide_input=true
show_doc(ImageDataBunch.batch_stats)
# -
data.batch_stats()
# + hide_input=true
show_doc(ImageDataBunch.normalize)
# -
# In the fast.ai library we have `imagenet_stats`, `cifar_stats` and `mnist_stats` so we can add normalization easily with any of these datasets. Let's see an example with our dataset of choice: MNIST.
data.normalize(cifar_stats)
data.batch_stats()
# ## Data normalization
# You may also want to normalize your data, which can be done by using the following functions.
# + hide_input=true
show_doc(normalize)
# + hide_input=true
show_doc(denormalize)
# + hide_input=true
show_doc(normalize_funcs)
# -
# On MNIST the mean and std are 0.1307 and 0.3081 respectively (looked on Google). If you're using a pretrained model, you'll need to use the normalization that was used to train the model. The imagenet norm and denorm functions are stored as constants inside the library named <code>imagenet_norm</code> and <code>imagenet_denorm</code>. If you're training a model on CIFAR-10, you can also use <code>cifar_norm</code> and <code>cifar_denorm</code>.
#
# You may sometimes see warnings about *clipping input data* when plotting normalized data. That's because even although it's denormalized when plotting automatically, sometimes floating point errors may make some values slightly out or the correct range. You can safely ignore these warnings in this case.
data = ImageDataBunch.from_folder(untar_data(URLs.MNIST_SAMPLE),
ds_tfms=tfms, size=24)
data.normalize()
data.show_batch(rows=3, figsize=(6,6))
# + hide_input=true
show_doc(get_annotations)
# -
# To use this dataset and collate samples into batches, you'll need to following function:
# + hide_input=true
show_doc(bb_pad_collate)
# -
# Finally, to apply transformations to [`Image`](/vision.image.html#Image) in a [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset), we use this last class.
# ## ItemList specific to vision
# The vision application adds a few subclasses of [`ItemList`](/data_block.html#ItemList) specific to images.
# + hide_input=true
show_doc(ImageItemList, title_level=3)
# -
# Create a [`ItemList`](/data_block.html#ItemList) in `path` from filenames in `items`. `create_func` will default to [`open_image`](/vision.image.html#open_image). `label_cls` can be specified for the labels, `xtra` contains any extra information (usually in the form of a dataframe) and `processor` is applied to the [`ItemList`](/data_block.html#ItemList) after splitting and labelling.
# + hide_input=true
show_doc(ImageItemList.from_folder)
# + hide_input=true
show_doc(ImageItemList.from_df)
# + hide_input=true
show_doc(get_image_files)
# + hide_input=true
show_doc(ImageItemList.open)
# + hide_input=true
show_doc(ImageItemList.show_xys)
# + hide_input=true
show_doc(ImageItemList.show_xyzs)
# + hide_input=true
show_doc(ObjectCategoryList, title_level=3)
# + hide_input=true
show_doc(ObjectItemList, title_level=3)
# + hide_input=true
show_doc(SegmentationItemList, title_level=3)
# + hide_input=true
show_doc(SegmentationLabelList, title_level=3)
# + hide_input=true
show_doc(PointsLabelList, title_level=3)
# + hide_input=true
show_doc(PointsItemList, title_level=3)
# + hide_input=true
show_doc(ImageImageList, title_level=3)
# -
# ## Building your own dataset
# This module also contains a few helper functions to allow you to build you own dataset for image classification.
# + hide_input=true
show_doc(download_images)
# + hide_input=true
show_doc(verify_images)
# -
# It will try if every image in this folder can be opened and has `n_channels`. If `n_channels` is 3 – it'll try to convert image to RGB. If `delete=True`, it'll be removed it this fails. If `resume` – it will skip already existent images in `dest`. If `max_size` is specifided, image is resized to the same ratio so that both sizes are less than `max_size`, using `interp`. Result is stored in `dest`, `ext` forces an extension type, `img_format` and `kwargs` are passed to PIL.Image.save. Use `max_workers` CPUs.
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
show_doc(PointsItemList.get)
show_doc(SegmentationLabelList.new)
show_doc(ImageItemList.from_csv)
show_doc(ObjectCategoryList.get)
show_doc(ImageItemList.get)
show_doc(SegmentationLabelList.reconstruct)
show_doc(ImageImageList.show_xys)
show_doc(ImageImageList.show_xyzs)
show_doc(ImageItemList.open)
show_doc(PointsItemList.analyze_pred)
show_doc(SegmentationLabelList.analyze_pred)
show_doc(PointsItemList.reconstruct)
show_doc(SegmentationLabelList.open)
show_doc(ImageItemList.reconstruct)
show_doc(resize_to)
show_doc(ObjectCategoryList.reconstruct)
show_doc(PointsLabelList.reconstruct)
show_doc(PointsLabelList.analyze_pred)
show_doc(PointsLabelList.get)
# ## New Methods - Please document or move to the undocumented section
# + hide_input=true
show_doc(ObjectCategoryList.analyze_pred)
# -
#
| docs_src/vision.data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LOAD PACKAGES!
import numpy as np
print ("Loading package(s)")
# # PRINT function usages
# +
print ("Hello, world")
# THERE ARE THREE POPULAR TYPES
# 1. INTEGER
x = 3;
print ("Integer: %01d, %02d, %03d, %04d, %05d"
% (x, x, x, x, x))
# 2. FLOAT
x = 123.456;
print ("Float: %.0f, %.1f, %.2f, %1.2f, %2.2f"
% (x, x, x, x, x))
# 3. STRING
x = "Hello, world"
print ("String: [%s], [%3s], [%20s]"
% (x, x, x))
# -
# # FOR + IF/ELSE
# +
dlmethods = ["ANN", "MLP", "CNN", "RNN", "DAE"]
for alg in dlmethods:
if alg in ["ANN", "MLP"]:
print ("We have seen %s" % (alg))
# +
dlmethods = ["ANN", "MLP", "CNN", "RNN", "DAE"];
for alg in dlmethods:
if alg in ["ANN", "MLP", "CNN"]:
print ("%s is a feed-forward network." % (alg))
elif alg in ["RNN"]:
print ("%s is a recurrent network." % (alg))
else:
print ("%s is an unsupervised method." % (alg))
# Little more advanced?
print("\nFOR loop with index.")
for alg, i in zip(dlmethods, range(len(dlmethods))):
if alg in ["ANN", "MLP", "CNN"]:
print ("[%d/%d] %s is a feed-forward network."
% (i, len(dlmethods), alg))
elif alg in ["RNN"]:
print ("[%d/%d] %s is a recurrent network."
% (i, len(dlmethods), alg))
else:
print ("[%d/%d] %s is an unsupervised method."
% (i, len(dlmethods), alg))
# -
# ## Note that, index starts with 0 !
# # Let's make a function in Python
# Function definition looks like this
def sum(a, b):
return a+b
X = 10.
Y = 20.
# Usage
print ("%.1f + %.1f = %.1f" % (X, Y, sum(X, Y)))
# # String operations
# +
head = "Deep learning"
body = "very "
tail = "HARD."
print (head + " is " + body + tail)
# Repeat words
print (head + " is " + body*3 + tail)
print (head + " is " + body*10 + tail)
# It is used in this way
print ("\n" + "="*50)
print (" "*15 + "It is used in this way")
print ("="*50 + "\n")
# Indexing characters in the string
x = "Hello, world"
for i in range(len(x)):
print ("Index: [%02d/%02d] Char: %s"
% (i, len(x), x[i]))
# -
# More indexing
print ""
idx = -2
print ("(%d)th char is %s" % (idx, x[idx]))
idxfr = 0
idxto = 8
print ("String from %d to %d is [%s]"
% (idxfr, idxto, x[idxfr:idxto]))
idxfr = 4
print ("String from %d to END is [%s]"
% (idxfr, x[idxfr:]))
x = "20160607Cloudy"
year = x[:4]
day = x[4:8]
weather = x[8:]
print ("[%s] -> [%s] + [%s] + [%s] "
% (x, year, day, weather))
# # LIST
a = []
b = [1, 2, 3]
c = ["Hello", ",", "world"]
d = [1, 2, 3, "x", "y", "z"]
x = []
print (x)
x.append('a')
print (x)
x.append(123)
print (x)
x.append(["a", "b"])
print (x)
print ("Length of x is %d "
% (len(x)))
for i in range(len(x)):
print ("[%02d/%02d] %s"
% (i, len(x), x[i]))
z = []
z.append(1)
z.append(2)
z.append(3)
z.append('Hello')
for i in range(len(z)):
print (z[i])
# # DICTIONARY
# +
dic = dict()
dic["name"] = "Sungjoon"
dic["age"] = 31
dic["job"] = "Ph.D. Candidate"
print (dic)
# -
# # Class
# +
class Greeter:
# Constructor
def __init__(self, name):
self.name = name # Create an instance variable
# Instance method
def greet(self, loud=False):
if loud:
print ('HELLO, %s!'
% self.name.upper())
else:
print ('Hello, %s'
% self.name)
g = Greeter('Fred') # Construct an instance of the Greeter class
g.greet() # Call an instance method; prints "Hello, Fred"
g.greet(loud=True) # Call an instance method; prints "HELLO, FRED!"
# -
def print_np(x):
print ("Type is %s" % (type(x)))
print ("Shape is %s" % (x.shape,))
print ("Values are: \n%s" % (x))
print
# # RANK 1 ARRAY
# +
x = np.array([1, 2, 3]) # rank 1 array
print_np(x)
x[0] = 5
print_np(x)
# -
# # RANK 2 ARRAY
y = np.array([[1,2,3], [4,5,6]])
print_np(y)
# # ZEROS
a = np.zeros((3, 2))
print_np(a)
# # ONES
b = np.ones((1, 2))
print_np(b)
# # IDENTITY
c = np.eye(2, 2)
print_np(c)
# # RANDOM (UNIFORM)
d = np.random.random((2, 2))
print_np(d)
# # RANDOM (GAUSSIAN)
e = np.random.randn(1, 10)
print_np(e)
# # ARRAY INDEXING
# +
# Create the following rank 2 array with shape (3, 4)
# [[ 1 2 3 4]
# [ 5 6 7 8]
# [ 9 10 11 12]]
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
print_np(a)
print
# Use slicing to pull out the subarray consisting
# of the first 2 rows
# and columns 1 and 2; b is the following array
# of shape (2, 2):
# [[2 3]
# [6 7]]
b = a[:2, 1:3]
print_np(b)
# -
# # GET ROW
# +
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
print_np(a)
row_r1 = a[1, :] # Rank 1 view of the second row of a
row_r2 = a[1:2, :] # Rank 2 view of the second row of a
row_r3 = a[[1], :] # Rank 2 view of the second row of a
print_np(row_r1)
print_np(row_r2)
print_np(row_r3)
# +
a = np.array([[1,2], [3, 4], [5, 6]])
print_np(a)
# An example of integer array indexing.
# The returned array will have shape (3,) and
b = a[[0, 1, 2], [0, 1, 0]]
print_np(b)
# The above example of integer array indexing
# is equivalent to this:
c = np.array([a[0, 0], a[1, 1], a[2, 0]])
print_np(c)
# -
# # DATATYPES
# +
x = np.array([1, 2]) # Let numpy choose the datatype
y = np.array([1.0, 2.0]) # Let numpy choose the datatype
z = np.array([1, 2], dtype=np.int64) # particular datatype
print_np(x)
print_np(y)
print_np(z)
# -
# ## Array math
# +
x = np.array([[1,2],[3,4]], dtype=np.float64)
y = np.array([[5,6],[7,8]], dtype=np.float64)
# Elementwise sum; both produce the array
print (x + y)
print (np.add(x, y))
# -
# Elementwise difference; both produce the array
print (x - y)
print (np.subtract(x, y))
# Elementwise product; both produce the array
print (x * y)
print (np.multiply(x, y))
# Elementwise division; both produce the array
# [[ 0.2 0.33333333]
# [ 0.42857143 0.5 ]]
print (x / y)
print (np.divide(x, y))
# Elementwise square root; produces the array
# [[ 1. 1.41421356]
# [ 1.73205081 2. ]]
print (np.sqrt(x))
# +
x = np.array([[1,2],[3,4]])
y = np.array([[5,6],[7,8]])
v = np.array([9,10])
w = np.array([11, 12])
print_np(x)
print_np(y)
print_np(v)
print_np(w)
# Inner product of vectors; both produce 219
print (v.dot(w))
print (np.dot(v, w)) # <= v * w'
# Matrix / vector product; both produce the rank 1 array [29 67]
print (x.dot(v))
print (np.dot(x, v)) # <= x * v'
# Matrix / matrix product; both produce the rank 2 array
# [[19 22]
# [43 50]]
print (x.dot(y))
print (np.dot(x, y))
# -
x = np.array([[1,2],[3,4]])
print_np(x)
print
print (x)
print (x.T)
print (np.sum(x)) # Compute sum of all elements
print (np.sum(x, axis=0)) # Compute sum of each column
print (np.sum(x, axis=1)) # Compute sum of each row
print (x)
print (x.T)
v = np.array([1,2,3])
print (v)
print (v.T)
v = np.array([[1,2,3]])
print (v)
print (v.T)
# ## Other useful operations
# +
# We will add the vector v to each row of the matrix x,
# storing the result in the matrix y
x = np.array([[1,2,3], [4,5,6], [7,8,9], [10, 11, 12]])
v = np.array([1, 0, 1])
y = np.empty_like(x) # Create an empty matrix
# with the same shape as x
print_np(x)
print_np(v)
print_np(y)
# -
# Add the vector v to each row of the matrix x
# with an explicit loop
for i in range(4):
y[i, :] = x[i, :] + v
print_np(y)
vv = np.tile(v, (4, 1)) # Stack 4 copies of v on top of each other
print_np(vv) # Prints "[[1 0 1]
# [1 0 1]
# [1 0 1]
# [1 0 1]]"
# We will add the vector v to each row of the matrix x,
# storing the result in the matrix y
x = np.array([[1,2,3], [4,5,6], [7,8,9], [10, 11, 12]])
v = np.array([1, 0, 1])
y = x + v # Add v to each row of x using BROADCASTING
print_np(x)
print_np(v)
print_np(y)
# +
# Add a vector to each row of a matrix
x = np.array([[1,2,3], [4,5,6]])
print_np(x)
print_np(v)
print (x + v)
# +
# Add a vector to each column of a matrix
print_np(x)
print_np(w)
print (x.T + w).T
# Another solution is to reshape w
# to be a row vector of shape (2, 1);
print
print (x + np.reshape(w, (2, 1)))
# -
# ## Matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Compute the x and y coordinates for points on a sine curve
x = np.arange(0, 3 * np.pi, 0.1)
y = np.sin(x)
# Plot the points using matplotlib
plt.plot(x, y)
# +
y_sin = np.sin(x)
y_cos = np.cos(x)
# Plot the points using matplotlib
plt.plot(x, y_sin)
plt.plot(x, y_cos)
plt.xlabel('x axis label')
plt.ylabel('y axis label')
plt.title('Sine and Cosine')
plt.legend(['Sine', 'Cosine'])
# Show the figure.
plt.show()
# +
# Compute the x and y coordinates for points
# on sine and cosine curves
x = np.arange(0, 3 * np.pi, 0.1)
y_sin = np.sin(x)
y_cos = np.cos(x)
# Set up a subplot grid that has height 2 and width 1,
# and set the first such subplot as active.
plt.subplot(2, 1, 1)
# Make the first plot
plt.plot(x, y_sin)
plt.title('Sine')
# Set the second subplot as active, and make the second plot.
plt.subplot(2, 1, 2)
plt.plot(x, y_cos)
plt.title('Cosine')
# Show the figure.
plt.show()
# -
tested; Gopal
| tests/tf/tf-basics/basic_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
""" Original Author: <NAME> """
import numpy as np
import ctypes as ct
import cv2
import sys
import os
BASE_DIR = "/mnt/nfs/work1/miyyer/kalpesh/projects/PointCloudLearningACD/visualizer"
showsz = 800
mousex, mousey = 0.5, 0.5
zoom = 1.0
changed = True
def onmouse(*args):
global mousex, mousey, changed
y = args[1]
x = args[2]
mousex = x / float(showsz)
mousey = y / float(showsz)
changed = True
cv2.namedWindow('show3d')
cv2.moveWindow('show3d', 0, 0)
cv2.setMouseCallback('show3d', onmouse)
dll = np.ctypeslib.load_library(os.path.join(BASE_DIR, 'render_balls_so'), '.')
def showpoints(xyz, c_gt=None, c_pred=None, waittime=0, showrot=False, magnifyBlue=0, freezerot=False,
background=(0, 0, 0), normalizecolor=True, ballradius=10):
global showsz, mousex, mousey, zoom, changed
xyz = xyz - xyz.mean(axis=0)
radius = ((xyz ** 2).sum(axis=-1) ** 0.5).max()
xyz /= (radius * 2.2) / showsz
if c_gt is None:
c0 = np.zeros((len(xyz),), dtype='float32') + 255
c1 = np.zeros((len(xyz),), dtype='float32') + 255
c2 = np.zeros((len(xyz),), dtype='float32') + 255
else:
c0 = c_gt[:, 0]
c1 = c_gt[:, 1]
c2 = c_gt[:, 2]
if normalizecolor:
c0 /= (c0.max() + 1e-14) / 255.0
c1 /= (c1.max() + 1e-14) / 255.0
c2 /= (c2.max() + 1e-14) / 255.0
c0 = np.require(c0, 'float32', 'C')
c1 = np.require(c1, 'float32', 'C')
c2 = np.require(c2, 'float32', 'C')
show = np.zeros((showsz, showsz, 3), dtype='uint8')
def render():
rotmat = np.eye(3)
if not freezerot:
xangle = (mousey - 0.5) * np.pi * 1.2
else:
xangle = 0
rotmat = rotmat.dot(np.array([
[1.0, 0.0, 0.0],
[0.0, np.cos(xangle), -np.sin(xangle)],
[0.0, np.sin(xangle), np.cos(xangle)],
]))
if not freezerot:
yangle = (mousex - 0.5) * np.pi * 1.2
else:
yangle = 0
rotmat = rotmat.dot(np.array([
[np.cos(yangle), 0.0, -np.sin(yangle)],
[0.0, 1.0, 0.0],
[np.sin(yangle), 0.0, np.cos(yangle)],
]))
rotmat *= zoom
nxyz = xyz.dot(rotmat) + [showsz / 2, showsz / 2, 0]
ixyz = nxyz.astype('int32')
show[:] = background
dll.render_ball(
ct.c_int(show.shape[0]),
ct.c_int(show.shape[1]),
show.ctypes.data_as(ct.c_void_p),
ct.c_int(ixyz.shape[0]),
ixyz.ctypes.data_as(ct.c_void_p),
c0.ctypes.data_as(ct.c_void_p),
c1.ctypes.data_as(ct.c_void_p),
c2.ctypes.data_as(ct.c_void_p),
ct.c_int(ballradius)
)
if magnifyBlue > 0:
show[:, :, 0] = np.maximum(show[:, :, 0], np.roll(show[:, :, 0], 1, axis=0))
if magnifyBlue >= 2:
show[:, :, 0] = np.maximum(show[:, :, 0], np.roll(show[:, :, 0], -1, axis=0))
show[:, :, 0] = np.maximum(show[:, :, 0], np.roll(show[:, :, 0], 1, axis=1))
if magnifyBlue >= 2:
show[:, :, 0] = np.maximum(show[:, :, 0], np.roll(show[:, :, 0], -1, axis=1))
if showrot:
cv2.putText(show, 'xangle %d' % (int(xangle / np.pi * 180)), (30, showsz - 30), 0, 0.5,
cv2.cv.CV_RGB(255, 0, 0))
cv2.putText(show, 'yangle %d' % (int(yangle / np.pi * 180)), (30, showsz - 50), 0, 0.5,
cv2.cv.CV_RGB(255, 0, 0))
cv2.putText(show, 'zoom %d%%' % (int(zoom * 100)), (30, showsz - 70), 0, 0.5, cv2.cv.CV_RGB(255, 0, 0))
changed = True
while True:
if changed:
render()
changed = False
cv2.imshow('show3d', show)
if waittime == 0:
cmd = cv2.waitKey(10) % 256
else:
cmd = cv2.waitKey(waittime) % 256
if cmd == ord('q'):
break
elif cmd == ord('Q'):
sys.exit(0)
if cmd == ord('t') or cmd == ord('p'):
if cmd == ord('t'):
if c_gt is None:
c0 = np.zeros((len(xyz),), dtype='float32') + 255
c1 = np.zeros((len(xyz),), dtype='float32') + 255
c2 = np.zeros((len(xyz),), dtype='float32') + 255
else:
c0 = c_gt[:, 0]
c1 = c_gt[:, 1]
c2 = c_gt[:, 2]
else:
if c_pred is None:
c0 = np.zeros((len(xyz),), dtype='float32') + 255
c1 = np.zeros((len(xyz),), dtype='float32') + 255
c2 = np.zeros((len(xyz),), dtype='float32') + 255
else:
c0 = c_pred[:, 0]
c1 = c_pred[:, 1]
c2 = c_pred[:, 2]
if normalizecolor:
c0 /= (c0.max() + 1e-14) / 255.0
c1 /= (c1.max() + 1e-14) / 255.0
c2 /= (c2.max() + 1e-14) / 255.0
c0 = np.require(c0, 'float32', 'C')
c1 = np.require(c1, 'float32', 'C')
c2 = np.require(c2, 'float32', 'C')
changed = True
if cmd == ord('n'):
zoom *= 1.1
changed = True
elif cmd == ord('m'):
zoom /= 1.1
changed = True
elif cmd == ord('r'):
zoom = 1.0
changed = True
elif cmd == ord('s'):
cv2.imwrite('show3d.png', show)
if waittime != 0:
break
return cmd
# -
| visualizer/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="cn5_7mqggh2H"
# ## sigMF RF classification; 12 classes
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 2251, "status": "ok", "timestamp": 1548951950015, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11966704463856227449"}, "user_tz": 300} id="r80FflgHhCiH" outputId="143411b2-cc11-47a1-c334-a76291219798"
import os
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.utils.data
import torch.utils.data as data
from torch.autograd import Variable
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
import glob
import json
from os import walk
import pickle
import json
import pathlib
import random
from timeit import default_timer as timer
import time
from tqdm import tqdm
from torch.utils.data import Dataset, DataLoader
global GPU, fft, Fs, center_freq, fft_val, Fs_test, loss, batches, eps, var_noise, mean_noise, top
global c1_coeff, c2_coeff, a, r1_c1, r2_c1, r1_c2, r2_c2, compare
a = 1
r1_c2 = 1
r2_c2 = 10e1
top = .995
var_noise = 8.78e-09
mean_noise = 1 # not used
eps = 1e-15
Fs = 1000000
fft = 1024
center_freq_file = 433.65e6 # when SDR doing 25MSPS with center at 428MHz
center_freq_live = 428.00e6 # when SDR doing 25MSPS with center at 428MHz
batches = 64
plt.style.use('default')
GPU = 0
device = torch.device('cuda:0')
print('Torch version =', torch.__version__, 'CUDA version =', torch.version.cuda)
print('CUDA Device:', device)
print('Is cuda available? =',torch.cuda.is_available())
# -
r2_c2
# !. /home/david/prefix-3.8/setup_env.sh
# + [markdown] colab_type="text" id="2t_9_D3l0Px9"
# #### Machine paths
# -
path_ram = "/home/david/sigMF_ML/RF/ramdisk/"
path_usrp = "/home/david/prefix-3.8/"
path = "/home/david/sigMF_ML/RF/RF_class/data/" # ACE
path_val1 = "/home/david/sigMF_ML/RF/RF_class/data_val1/" # fft_center - class 9
path_val2 = "/home/david/sigMF_ML/RF/RF_class/data_val2/" # light switch - class 10
path_val3 = "/home/david/sigMF_ML/RF/RF_class/data_val3/" # clickndig - class 5
path_val4 = "/home/david/sigMF_ML/RF/RF_class/data_val4/" # GD55 - class 1
path_val5 = "/home/david/sigMF_ML/RF/RF_class/data_val5/" # lora125 - class 0
path_val6 = "/home/david/sigMF_ML/RF/RF_class/data_val6/" # lora250 - class 7
path_val7 = "/home/david/sigMF_ML/RF/RF_class/data_val7/" # NFM - class 2
path_val8 = "/home/david/sigMF_ML/RF/RF_class/data_val8/" # sado - class 6
path_val9 = "/home/david/sigMF_ML/RF/RF_class/data_val9/" # TYT - class 3
path_val10 = "/home/david/sigMF_ML/RF/RF_class/data_val10/" # vodeson - class 4
path_val11 = "/home/david/sigMF_ML/RF/RF_class/data_val11/" # white noise - class 8
path_val12 = "/home/david/sigMF_ML/RF/RF_class/data_val12/" # ysf - class 11
path_fig = "/home/david/sigMF_ML/RF/RF_class/" # ACE
path_val = "/home/david/sigMF_ML/RF/RF_class/testing_data/" # ACE
path_save = "/home/david/sigMF_ML/RF/RF_class/saved/" # ACE
path_test = "/home/david/sigMF_ML/RF/RF_class/testing_data/" # ACE
path_test_1msps = "/home/david/sigMF_ML/RF/RF_class/testing_data_1msps/" # ACE
path_test_5msps = "/home/david/sigMF_ML/RF/RF_class/testing_data_5msps/" # ACE
path_test_10msps = "/home/david/sigMF_ML/RF/RF_class/testing_data_10msps/" # ACE
path_test_25msps = "/home/david/sigMF_ML/RF/RF_class/testing_data_25msps/" # ACE
print(path)
# ## Functions
# +
# START OF FUNCTIONS ****************************************************
def meta_encoder(meta_list, num_classes):
a = np.asarray(meta_list, dtype=int)
# print('a = ', a)
return a
def save_model(epoch,loss):
rf_model = 'ResNet18_v58_20210226_2D_10dB_1e4lr_autosave'
PATH = path_save+rf_model
torch.save({
'epoch': epoch,
'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict(),
'loss': loss,}, PATH)
def load_model():
rf_model = 'ResNet50_v58_20210215_2D_10dB_noise_autosave'
PATH = path_save+rf_model
device = torch.device("cuda:1")
model = resnet50(2, 12)
model.load_state_dict(torch.load(PATH))
model.to(device)
model.eval()
def gpu_test_file(db):
if (msps == 1):
w1 = fft
elif (msps == 5):
w1 = fft*1
elif (msps == 10):
w1 = fft*1
elif (msps == 25):
w1 = fft*1
print('gpu_test file function')
I = db[0::2]
Q = db[1::2]
w = fft*msps
print('Sample Rate = ',w,'MSPS')
den = 2
print('window length = ', w1)
win = torch.hann_window(w1, periodic=True, dtype=None, layout=torch.strided, requires_grad=False).cuda(GPU)
I_stft = torch.stft(torch.tensor(I).cuda(GPU), n_fft=fft, hop_length=fft//den, win_length=w1, window=win, center=True, normalized=True, onesided=True)
Q_stft = torch.stft(torch.tensor(Q).cuda(GPU), n_fft=fft, hop_length=fft//den, win_length=w1, window=win, center=True, normalized=True, onesided=True)
I_mag = torch.abs(torch.sqrt(I_stft[:,:,0]**2+I_stft[:,:,1]**2))
Q_mag = torch.abs(torch.sqrt(Q_stft[:,:,0]**2+Q_stft[:,:,1]**2))
I_mag = torch.unsqueeze(I_mag, dim=2)
Q_mag = torch.unsqueeze(Q_mag, dim=2)
# print('I shape =', I_stft.shape, 'Q shape = ', Q_stft.shape )
# print('I_mag shape =', I_mag.shape, 'Q_mag shape = ', Q_mag.shape )
Z_stft = torch.cat((I_mag,Q_mag),2)
# Z_stft = torch.cat((Z_stft[fft//2:,:,:],Z_stft[:fft//2,:,:])) # NOT SURE I NEED TO DO THIS...
# print('gpu_test file Z shape =', Z_stft.shape)
Z_stft = Z_stft[fft//2:,:,:] # throw bottom 1/2 away
print('FINAL gpu_test FILE IQ shape =', Z_stft.shape)
torch.cuda.empty_cache()
return Z_stft # Returning 2D
def gpu_test_live(db,msps):
if (msps == 1):
w1 = fft
elif (msps == 5):
w1 = fft*1
elif (msps == 10):
w1 = fft*1
elif (msps == 25):
w1 = fft*1
print('gpu_test live function')
# I = db[0:10000000:2]
# Q = db[1:10000000:2]
I = db[0::2]
Q = db[1::2]
print('I length = ', len(I))
print('Q length = ', len(Q))
w = fft*msps
print(w,'MSPS')
den = 2
win = torch.hann_window(w1, periodic=True, dtype=None, layout=torch.strided, requires_grad=False).cuda(GPU)
I_stft = torch.stft(torch.tensor(I).cuda(GPU), n_fft=w, hop_length=w//den, win_length=w1, window=win, center=True, normalized=True, onesided=True)
Q_stft = torch.stft(torch.tensor(Q).cuda(GPU), n_fft=w, hop_length=w//den, win_length=w1, window=win, center=True, normalized=True, onesided=True)
I_mag = torch.abs(torch.sqrt(I_stft[:,:,0]**2+I_stft[:,:,1]**2))
Q_mag = torch.abs(torch.sqrt(Q_stft[:,:,0]**2+Q_stft[:,:,1]**2))
I_mag = torch.unsqueeze(I_mag, dim=2)
Q_mag = torch.unsqueeze(Q_mag, dim=2)
Z_stft = torch.cat((I_mag,Q_mag),2)
print('gpu_test live IQ shape =', Z_stft.shape)
# Z_stft = torch.cat((Z_stft[w//2:,:,:],Z_stft[:w//2,:,:])) # NOT SURE I NEED TO DO THIS...
Z_stft = Z_stft[:w//2,:,:]# throw bottom 1/2 away
print('FINAL gpu_test LIVE IQ shape =', Z_stft.shape)
return Z_stft # Returning 2D and plot
def iq_read(data_files): # USING GPU to perform STFT
print('iq_read function**********')
data_IQ_list = []
data_IQ_temp = []
for file in data_files:
db = np.fromfile(file, dtype="float32")
# stft = gpu(db).detach().cpu().numpy()
print('iq_read function')
stft, stft_plot = gpu_test_file(db)
stft = stft.detach().cpu().numpy()
stft_plot = stft_plot.detach().cpu().numpy()
stft_plot = 10*np.log10(np.abs(stft_plot+eps))
plt.imshow(stft_plot)
plt.pcolormesh(stft_plot)
# plt.imshow(stft, aspect='auto', origin='lower')
plt.show()
data_IQ_temp.append(stft)
data_IQ_list = np.array(data_IQ_temp)
return data_IQ_list
def iq_read_test_file(data_files): # USING GPU to perform STFT
data_IQ_list = []
data_IQ_temp = []
print('iq_read_test file')
for file in data_files:
db = np.fromfile(file, dtype="float32")
stft = gpu_test_file(db)
stft_plot = 20*np.log10(np.abs(stft[:,:,0].detach().cpu().numpy()+eps))
print('imshow method')
plt.imshow(stft_plot, vmin=-70, vmax=5, aspect='auto', origin='lower')
plt.show()
data_IQ_temp.append(stft.detach().cpu().numpy())
data_IQ_list = np.array(data_IQ_temp)
return data_IQ_list
def iq_read_test_live(data_files,msps): # USING GPU to perform STFT
# iq_cpu_plot(data_files) #checking with cpu complex plotting
data_IQ_list = []
data_IQ_temp = []
print('iq_read_test live')
for file in data_files:
db = np.fromfile(file, dtype="float32")
stft = gpu_test_live(db,msps)
# *************************************************************************
stft_plot = 20*np.log10(np.abs(stft[:,:,0].detach().cpu().numpy()+eps))
print('iq_read_test live imshow method')
plt.imshow(stft_plot, vmin=-70, vmax=5, aspect='auto', origin='lower')
plt.show()
# *************************************************************************
data_IQ_temp.append(stft.detach().cpu().numpy())
data_IQ_list = np.array(data_IQ_temp)
return data_IQ_list
def read_meta(meta_files):
meta_list = []
for meta in meta_files:
all_meta_data = json.load(open(meta))
meta_list.append(all_meta_data['global']["core:class"])
meta_list = list(map(int, meta_list))
return meta_list
def read_num_val(x):
x = len(meta_list_val)
return x
#**************************** Print historgram subplots ******************************
def histo_plots(inputs):
fig=plt.figure(figsize=(8,8))
ncols = 2
nrows = 2
print('make torch inputs')
print('inputs shape for histogram1 = ', inputs.shape)
inputs = 10*np.log10(np.abs(inputs.cpu()+eps))
for x in range(4):
# print('x = ', x, 'inputs shape for histogram2 = ', inputs[:,:,x].shape)
flat_inputs = torch.flatten(inputs[:,:,x], start_dim=0, end_dim=-1).numpy()
# print('type = ', type(flat_inputs))
# print('x = ', x, 'flat_input max = ', np.amax(flat_inputs))
# print('inputs are: ', flat_inputs.shape)
fig.add_subplot(nrows, ncols, x+1)
plt.hist(flat_inputs, bins=5000)
plt.gca().set(title='Frequency Histogram', ylabel='Frequency');
plt.xlim(-100, 10)
# plt.ylim(0, 40000)
return flat_inputs
#*************************************************************************************
#**************************** Print historgram subplots ******************************
def histo_stats(inputs):
# print('make torch inputs')
# print('inputs shape for histogram1 = ', inputs.shape)
mean = np.zeros(4)
std = np.zeros(4)
for x in range(4):
# print('x = ', x, 'inputs shape for histogram2 = ', inputs[:,:,x].shape)
flat_inputs = torch.flatten(inputs[:,:,x], start_dim=0, end_dim=-1).numpy()
# print('inputs are: ', flat_inputs.shape)
mean[x] = flat_inputs.mean()
std[x] = flat_inputs.std()
# print('mean = ', mean, 'std = ', std)
return mean, std
#**************************** Print historgram freq stats ******************************
def histo_stats_freq_file(inputs,msps):
mean = inputs.mean()
std = inputs.std()
print("mean Freq = {0:9,.2f}".format(mean))
print("std Freq = {0:9,.2f}".format(std))
print('length of inputs = ', len(inputs))
# plt.hist(inputs, 30, facecolor='blue', align='mid')
if (msps==25):
plt.hist(inputs, 30, range=[428.0, 440.0], facecolor='blue', align='mid')
elif (msps==1):
plt.hist(inputs, 30, range=[433.65, 434.15], facecolor='blue', align='mid')
elif (msps==5):
plt.hist(inputs, 30, range=[433.00, 435.50], facecolor='blue', align='mid')
elif (msps==10):
plt.hist(inputs, 30, range=[433.00, 438.00], facecolor='blue', align='mid')
else:
print('WRONG SAMPLE RATE CHOSEN')
plt.gca().set(title='Frequency Histogram', ylabel='Frequency');
plt.show()
def histo_stats_freq_live(inputs,msps):
mean = inputs.mean()
std = inputs.std()
print("mean Freq = {0:9,.2f}".format(mean))
print("std Freq = {0:9,.2f}".format(std))
print('length of inputs = ', len(inputs))
# plt.hist(inputs, 30, facecolor='blue', align='mid')
if (msps==25):
plt.hist(inputs, 30, range=[428.0, 440.0], facecolor='blue', align='mid')
elif (msps==1):
plt.hist(inputs, 30, range=[433.65, 434.15], facecolor='blue', align='mid')
elif (msps==5):
plt.hist(inputs, 30, range=[433.00, 435.50], facecolor='blue', align='mid')
elif (msps==10):
plt.hist(inputs, 30, range=[433.00, 438.00], facecolor='blue', align='mid')
else:
print('WRONG SAMPLE RATE CHOSEN')
plt.gca().set(title='Frequency Histogram', ylabel='Frequency');
plt.show()
# END OF FUNCTIONS ******************************************************
# -
from functools import partial
from dataclasses import dataclass
from collections import OrderedDict
# +
class Conv2dAuto(nn.Conv2d):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.padding = (self.kernel_size[0] // 2, self.kernel_size[1] // 2) # dynamic add padding based on the kernel_size
conv3x3 = partial(Conv2dAuto, kernel_size=3, bias=False)
# -
class ResidualBlock(nn.Module):
def __init__(self, in_channels, out_channels):
super().__init__()
self.in_channels, self.out_channels = in_channels, out_channels
self.blocks = nn.Identity()
self.shortcut = nn.Identity()
def forward(self, x):
residual = x
if self.should_apply_shortcut: residual = self.shortcut(x)
x = self.blocks(x)
x += residual
return x
@property
def should_apply_shortcut(self):
return self.in_channels != self.out_channels
class ResNetResidualBlock(ResidualBlock):
def __init__(self, in_channels, out_channels, expansion=1, downsampling=1, conv=conv3x3, *args, **kwargs):
super().__init__(in_channels, out_channels)
self.expansion, self.downsampling, self.conv = expansion, downsampling, conv
self.shortcut = nn.Sequential(OrderedDict(
{
'conv' : nn.Conv2d(self.in_channels, self.expanded_channels, kernel_size=1,
stride=self.downsampling, bias=False),
'bn' : nn.BatchNorm2d(self.expanded_channels)
})) if self.should_apply_shortcut else None
@property
def expanded_channels(self):
return self.out_channels * self.expansion
@property
def should_apply_shortcut(self):
return self.in_channels != self.expanded_channels
def conv_bn(in_channels, out_channels, conv, *args, **kwargs):
return nn.Sequential(OrderedDict({'conv': conv(in_channels, out_channels, *args, **kwargs),
'bn': nn.BatchNorm2d(out_channels) }))
class ResNetBasicBlock(ResNetResidualBlock):
expansion = 1
def __init__(self, in_channels, out_channels, activation=nn.ReLU, *args, **kwargs):
super().__init__(in_channels, out_channels, *args, **kwargs)
self.blocks = nn.Sequential(
conv_bn(self.in_channels, self.out_channels, conv=self.conv, bias=False, stride=self.downsampling),
activation(),
conv_bn(self.out_channels, self.expanded_channels, conv=self.conv, bias=False),
)
class ResNetBottleNeckBlock(ResNetResidualBlock):
expansion = 4
def __init__(self, in_channels, out_channels, activation=nn.ReLU, *args, **kwargs):
super().__init__(in_channels, out_channels, expansion=4, *args, **kwargs)
self.blocks = nn.Sequential(
conv_bn(self.in_channels, self.out_channels, self.conv, kernel_size=1),
activation(),
conv_bn(self.out_channels, self.out_channels, self.conv, kernel_size=3, stride=self.downsampling),
activation(),
conv_bn(self.out_channels, self.expanded_channels, self.conv, kernel_size=1),
)
class ResNetLayer(nn.Module):
def __init__(self, in_channels, out_channels, block=ResNetBasicBlock, n=1, *args, **kwargs):
super().__init__()
# 'We perform downsampling directly by convolutional layers that have a stride of 2.'
downsampling = 2 if in_channels != out_channels else 1
self.blocks = nn.Sequential(
block(in_channels , out_channels, *args, **kwargs, downsampling=downsampling),
*[block(out_channels * block.expansion,
out_channels, downsampling=1, *args, **kwargs) for _ in range(n - 1)]
)
def forward(self, x):
x = self.blocks(x)
return x
class ResNetEncoder(nn.Module):
"""
ResNet encoder composed by increasing different layers with increasing features.
"""
def __init__(self, in_channels=3, blocks_sizes=[64, 128, 256, 512], deepths=[2,2,2,2],
activation=nn.ReLU, block=ResNetBasicBlock, *args,**kwargs):
super().__init__()
self.blocks_sizes = blocks_sizes
self.gate = nn.Sequential(
nn.Conv2d(in_channels, self.blocks_sizes[0], kernel_size=7, stride=2, padding=3, bias=False),
nn.BatchNorm2d(self.blocks_sizes[0]),
activation(),
nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
)
self.in_out_block_sizes = list(zip(blocks_sizes, blocks_sizes[1:]))
self.blocks = nn.ModuleList([
ResNetLayer(blocks_sizes[0], blocks_sizes[0], n=deepths[0], activation=activation,
block=block, *args, **kwargs),
*[ResNetLayer(in_channels * block.expansion,
out_channels, n=n, activation=activation,
block=block, *args, **kwargs)
for (in_channels, out_channels), n in zip(self.in_out_block_sizes, deepths[1:])]
])
def forward(self, x):
x = self.gate(x)
for block in self.blocks:
x = block(x)
return x
class ResnetDecoder(nn.Module):
"""
This class represents the tail of ResNet. It performs a global pooling and maps the output to the
correct class by using a fully connected layer.
"""
def __init__(self, in_features, n_classes):
super().__init__()
self.avg = nn.AdaptiveAvgPool2d((1, 1))
self.decoder = nn.Linear(in_features, n_classes)
def forward(self, x):
x = self.avg(x)
x = x.view(x.size(0), -1)
x = self.decoder(x)
return x
class ResNet(nn.Module):
def __init__(self, in_channels, n_classes, *args, **kwargs):
super().__init__()
self.encoder = ResNetEncoder(in_channels, *args, **kwargs)
self.decoder = ResnetDecoder(self.encoder.blocks[-1].blocks[-1].expanded_channels, n_classes)
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
return x
# +
def resnet18(in_channels, n_classes):
return ResNet(in_channels, n_classes, block=ResNetBasicBlock, deepths=[2, 2, 2, 2])
def resnet34(in_channels, n_classes):
return ResNet(in_channels, n_classes, block=ResNetBasicBlock, deepths=[3, 4, 6, 3])
def resnet50(in_channels, n_classes):
return ResNet(in_channels, n_classes, block=ResNetBottleNeckBlock, deepths=[3, 4, 6, 3])
def resnet101(in_channels, n_classes):
return ResNet(in_channels, n_classes, block=ResNetBottleNeckBlock, deepths=[3, 4, 23, 3])
def resnet152(in_channels, n_classes):
return ResNet(in_channels, n_classes, block=ResNetBottleNeckBlock, deepths=[3, 8, 36, 3])
# +
from torchsummary import summary
# model = resnet50(4, 12)
# summary(model.cuda(GPU), (4, 224, 224))
# -
from torch.utils.data import Dataset
class RFDataset(Dataset):
def __init__(self, root_path):
print(root_path)
self.root_path = root_path
self.list_of_all_pickles = sorted(pathlib.Path(root_path).rglob('*/chopped-data-224-224/*.pickle'))
self.get_class = dict()
class_folders = list(pathlib.Path(root_path).glob('*/'))
for class_folder in class_folders:
class_index = -1
metadata_path = list(class_folder.rglob('*.sigmf-meta'))[0]
with open(metadata_path) as fp:
metadata = json.load(fp)
class_index = int(metadata["global"]["core:class"])
self.get_class[str(class_folder.stem)] = class_index
def __len__(self):
return len(self.list_of_all_pickles)
def __getitem__(self, idx):
filepath = self.list_of_all_pickles[idx]
with open(filepath, 'rb') as fp:
tensor = pickle.load(fp)['bounded']
foldername = filepath.parts[7]
label = self.get_class[foldername]
#return (tensor, label) # this is a tuple
return {'data': tensor, 'label': label}
# #### Evaluation dataset loader
rf_dataset1 = RFDataset(path_val1)
val1_data = data.DataLoader(rf_dataset1, batch_size=batches, shuffle=True)
rf_dataset2 = RFDataset(path_val2)
val2_data = data.DataLoader(rf_dataset2, batch_size=batches, shuffle=True)
rf_dataset3 = RFDataset(path_val3)
val3_data = data.DataLoader(rf_dataset3, batch_size=batches, shuffle=True)
rf_dataset4 = RFDataset(path_val4)
val4_data = data.DataLoader(rf_dataset4, batch_size=batches, shuffle=True)
rf_dataset5 = RFDataset(path_val5)
val5_data = data.DataLoader(rf_dataset5, batch_size=batches, shuffle=True)
rf_dataset6 = RFDataset(path_val6)
val6_data = data.DataLoader(rf_dataset6, batch_size=batches, shuffle=True)
rf_dataset7 = RFDataset(path_val7)
val7_data = data.DataLoader(rf_dataset7, batch_size=batches, shuffle=True)
rf_dataset8 = RFDataset(path_val8)
val8_data = data.DataLoader(rf_dataset8, batch_size=batches, shuffle=True)
rf_dataset9 = RFDataset(path_val9)
val9_data = data.DataLoader(rf_dataset9, batch_size=batches, shuffle=True)
rf_dataset10 = RFDataset(path_val10)
val10_data = data.DataLoader(rf_dataset10, batch_size=batches, shuffle=True)
rf_dataset11 = RFDataset(path_val11)
val11_data = data.DataLoader(rf_dataset11, batch_size=batches, shuffle=True)
rf_dataset12 = RFDataset(path_val12)
val12_data = data.DataLoader(rf_dataset12, batch_size=batches, shuffle=True)
# #### list of loaders
val_data_list = [val1_data]
val_data_list.append(val2_data)
val_data_list.append(val3_data)
val_data_list.append(val4_data)
val_data_list.append(val5_data)
val_data_list.append(val6_data)
val_data_list.append(val7_data)
val_data_list.append(val8_data)
val_data_list.append(val9_data)
val_data_list.append(val10_data)
val_data_list.append(val11_data)
val_data_list.append(val12_data)
print('done')
# ## Training
def train_net(total):
test_patch_total = 24000 # 120000
compare = .7
loss_plot = np.zeros(total)
total_plot = np.zeros(total//5+1)
batch_plot = np.zeros(len(training_data)*total//100)
batch_indexer = 0
for epoch in tqdm(range(total), desc="Epoch"):
model.train()
start = timer()
for i, rf_data in enumerate(training_data, 0):
inputs = rf_data['data']
inputs = torch.squeeze(inputs, dim=1)
# print('input1 = ', inputs.shape)
inputs = inputs.permute(0,3,1,2).contiguous()
# print('input before noise add = ', inputs.shape)
batch_dim, b, c, d = inputs.shape
# add som noise
c2 = torch.FloatTensor(a).uniform_(r1_c2, r2_c2)
for batch_num in range(batch_dim):
inputs[batch_num,:,:,:] = inputs[batch_num,:,:,:] + (((var_noise*c2)**0.5)*torch.randn(1, 4, 224, 224))
#**************** take I and Q magnitudue only ***********
I_temp = torch.abs(torch.sqrt(inputs[:,0,:,:]**2+inputs[:,1,:,:]**2))
I_temp = torch.unsqueeze(I_temp, dim=1)
Q_temp = torch.abs(torch.sqrt(inputs[:,2,:,:]**2+inputs[:,3,:,:]**2))
Q_temp = torch.unsqueeze(Q_temp, dim=1)
inputs = torch.cat((I_temp,Q_temp),1)
# print('inputs after noise add = ', inputs.shape)
#*********************************************************
inputs = Variable(inputs.cuda(GPU))
labels = rf_data['label']
labels = labels.cuda(GPU)
outputs = model(inputs)
loss = criterion(outputs, labels)
optimizer.zero_grad()
loss.backward()
optimizer.step()
end = timer()
batch_time = end - start
#***************************************************************************************
print('batch time = ', batch_time)
print('************************* start *************************')
total_correct_patches = grand_total = 0
start_test = timer()
model.eval()
for testing in val_data_list:
t = train_val(testing)
total_correct_patches = total_correct_patches + t
grand_total = total_correct_patches/test_patch_total
batch_plot[batch_indexer] = grand_total*100
batch_indexer = batch_indexer + 1
# print('Batch number = ', i, 'of', len(training_data))
print('Total % correct {:.2f}%'.format(grand_total*100))
model.train
end_test = timer()
test_time = end_test - start_test
print('test time = ', test_time)
print('*************************** end ***************************')
#****************************************************************************************
save_model(epoch,loss)
tqdm.write('___________________________________________')
tqdm.write("Epoch {} Loss {:.10f} ".format(epoch+1, loss.data*1))
tqdm.write('___________________________________________')
return loss_plot, batch_plot
# #### Training Evaluation
def train_val(val_data):
with torch.no_grad():
total = noise = center_fft = target_to_int = accumulated_corrects = percent_correct = 0
c0 = c1 = c2 = c3 = c4 = c5 = c6 = c7 = c8 = c9 = 0
for i, rf_data in enumerate(val_data, 0):
inputs = rf_data['data']
inputs = torch.squeeze(inputs, dim=1)
inputs = inputs.permute(0,3,1,2).contiguous()
batch_dim, b, c, d = inputs.shape
# add som noise
c2 = torch.FloatTensor(a).uniform_(r1_c2, r2_c2)
for batch_num in range(batch_dim):
inputs[batch_num,:,:,:] = inputs[batch_num,:,:,:] + (((var_noise*c2)**0.5)*torch.randn(1, 4, 224, 224)*1)
#**************** take I and Q magnitudue only ***********
I_temp = torch.abs(torch.sqrt(inputs[:,0,:,:]**2+inputs[:,1,:,:]**2))
I_temp = torch.unsqueeze(I_temp, dim=1)
Q_temp = torch.abs(torch.sqrt(inputs[:,2,:,:]**2+inputs[:,3,:,:]**2))
Q_temp = torch.unsqueeze(Q_temp, dim=1)
inputs = torch.cat((I_temp,Q_temp),1)
#*********************************************************
inputs = Variable(inputs.cuda(GPU))
labels = rf_data['label']
labels = labels.cuda(GPU)
optimizer.zero_grad()
outputs = model(inputs)
_, predicted = torch.max(outputs.data, 1)
for b in range(len(predicted)):
labels_temp = labels[b].detach().cpu().numpy()
temp = predicted[b].detach().cpu().numpy()
if (labels_temp==temp):
accumulated_corrects = accumulated_corrects+1
torch.cuda.empty_cache()
return accumulated_corrects
# ## Inference Segmented Dataloader
# Inference Dataloader with labels
class inference_dataloader_segmented_live(data.Dataset):
def __init__(self):
self.dataPath = path
self.num_classes = 12
self.num_examples = 1 # use only 1 for semi-live inferencing
def __getitem__(self, index):
sigmf_data = np.array(data_IQ_list_val[index])
print('sigmf_data = ', sigmf_data.shape)
frequency, time, dims1 = sigmf_data.shape
print('frequency = ', frequency, ' time = ', time)
data_IQ = []
data_IQ_temp2 = []
seg_t = 224
seg_f = 224
seg_time = time//seg_t
seg_freq = frequency//seg_f
print('seg_time = ', seg_time, 'seg_freq = ', seg_freq)
# Segment the time axis
for j in range(seg_time):
# Segment the frequency axis
for k in range(seg_freq):
IQ = sigmf_data[seg_f*k:(seg_f)+seg_f*k,seg_t*j:(seg_t)+seg_t*j]
data_IQ_temp2.append(IQ)
data_IQ = np.array(data_IQ_temp2)
print('data_IQ shape = ', data_IQ.shape)
loop_counter, dim1, dim2, dim3 = data_IQ.shape
TRUTH = meta_encoder(meta_list_val, self.num_classes)
TRUTH = TRUTH.astype(np.float32)
return torch.from_numpy(data_IQ),torch.from_numpy(TRUTH), loop_counter, seg_freq
def __len__(self):
return self.num_examples
# Inference Dataloader with labels
class inference_dataloader_segmented(data.Dataset):
def __init__(self):
self.dataPath = path
self.num_classes = 12
self.num_examples = 1
def __getitem__(self, index):
sigmf_data = np.array(data_IQ_list_val[index])
print('sigmf_data = ', sigmf_data.shape)
frequency, time, dims1 = sigmf_data.shape
print('frequency = ', frequency, ' time = ', time)
data_IQ = []
data_IQ_temp2 = []
seg_t = 224
seg_f = 224
seg_time = time//seg_t
seg_freq = frequency//seg_f
print('seg_time = ', seg_time, 'seg_freq = ', seg_freq)
# Segment the time axis
for j in range(seg_time):
# Segment the frequency axis
for k in range(seg_freq):
IQ = sigmf_data[seg_f*k:(seg_f)+seg_f*k,seg_t*j:(seg_t)+seg_t*j]
data_IQ_temp2.append(IQ)
data_IQ = np.array(data_IQ_temp2)
print('data_IQ shape = ', data_IQ.shape)
loop_counter, dim1, dim2, dim3 = data_IQ.shape
TRUTH = meta_encoder(meta_list_val, self.num_classes)
TRUTH = TRUTH.astype(np.float32)
return torch.from_numpy(data_IQ),torch.from_numpy(TRUTH), loop_counter, seg_freq
def __len__(self):
return self.num_examples
# ### validation functions
def validation_read():
# Inference DATA READING ************************************************
# read in validation IQ and meta data
os.chdir(path_val)
data_files_validation = sorted(glob.glob('*.sigmf-data'))
meta_files_validation = sorted(glob.glob('*.sigmf-meta'))
for meta in meta_files_validation:
all_meta_data = json.load(open(meta))
print("validation file name = ", meta)
# Load validation sigmf-data files
meta_list_val = read_meta(meta_files_validation)
data_IQ_list_val = iq_read(data_files_validation)
return data_IQ_list_val, meta_list_val
def testing_read():
print('testing_read function')
# Inference DATA READING ************************************************
# read in validation IQ and meta data
os.chdir(path_val)
data_files_validation = sorted(glob.glob('*.sigmf-data'))
meta_files_validation = sorted(glob.glob('*.sigmf-meta'))
for meta in meta_files_validation:
all_meta_data = json.load(open(meta))
print("testing file name = ", meta)
meta_list_val = read_meta(meta_files_validation)
data_IQ_list_val = iq_read_test_file(data_files_validation)
return data_IQ_list_val, meta_list_val
# #### Changed to get test data from different directory
def inference_read(msps):
# Inference DATA READING ************************************************
# read in validation IQ and meta data
os.chdir(path_ram)
data_files_validation = sorted(glob.glob('*.sigmf-data'))
meta_files_validation = sorted(glob.glob('*.sigmf-meta'))
for meta in meta_files_validation:
all_meta_data = json.load(open(meta))
print("inference file name = ", meta)
# Load validation sigmf-data files
meta_list_val = read_meta(meta_files_validation)
data_IQ_list_val = iq_read_test_live(data_files_validation,msps)
return data_IQ_list_val, meta_list_val
def inference_read_file(msps,path):
# Inference DATA READING ************************************************
# read in validation IQ and meta data
os.chdir(path)
data_files_validation = sorted(glob.glob('*.sigmf-data'))
meta_files_validation = sorted(glob.glob('*.sigmf-meta'))
for meta in meta_files_validation:
all_meta_data = json.load(open(meta))
print("inference file name = ", meta)
# Load validation sigmf-data files
meta_list_val = read_meta(meta_files_validation)
data_IQ_list_val = iq_read_test_live(data_files_validation,msps)
return data_IQ_list_val, meta_list_val
# inference ************************************************************
def testing_file(msps):
large_width = 400
np.set_printoptions(precision=2,floatmode='fixed', linewidth=large_width)
model.eval()
V = data.DataLoader(inference_dataloader_segmented(), batch_size=1)
start_frequency = (center_freq_file)
match_freq = start_frequency
print('start_frequency = ', start_frequency/1000000)
freq_offset = 0
total = noise = center_fft = target_to_int = accumulated_corrects = percent_correct = 0
c0 = c1 = c2 = c3 = c4 = c5 = c6 = c7 = c8 = c9 = 0
# total = 68
with torch.no_grad():
for i, rf_data in enumerate(V, 0):
accumulated_corrects = 0
percent_correct = 0
target_to_int = 0
inputs, target, counter, seg_freq = rf_data
print('testing counter = ', counter, 'seg_freq =', seg_freq)
#**************************** Print segmented pics ***********************************
# stft_plot = np.squeeze(inputs, axis=0)
# fig=plt.figure(figsize=(8,8))
# ncols = 5
# nrows = 5
# range_plot = 1
# for x in range(1,22): # need to figure out how to not hard code this ********************* <-----
# stft_mean, stft_std = histo_stats(stft_plot[x,:,:,:])
# if (x>=range_plot and x<(range_plot+25)):
# stft_plot1 = 10*np.log10(np.abs(stft_plot[x, :, :, 0]+eps))
# stft_plot1 = np.squeeze(stft_plot1, axis=0)
# fig.add_subplot(nrows, ncols, x-range_plot+1)
# plt.imshow(stft_plot1, vmin=-70, vmax=5)
# plt.show()
#******************************************************************************************
freq_increment = (Fs*msps/2)/seg_freq.detach().cpu().numpy().item()
print('freq_increment = ', freq_increment)
print('TESTING inputs SHAPE = ', inputs.shape)
target = Variable(target.cuda(GPU))
print('input in = ', inputs.shape)
inputs = torch.squeeze(inputs, dim=0)
print('input out = ', inputs.shape)
inputs = inputs.permute(0,3,1,2).contiguous()
print('counter convert stuff = ', counter, type(counter.numpy()))
inputs = Variable(inputs.cuda(GPU))
print('permuted shape = ', inputs.shape)
freq_count = 0 # keep track of array position
freq_histo = np.zeros(counter.numpy())
for j in range(counter):
inputs2 = inputs[j,:,:,:]
inputs2 = torch.unsqueeze(inputs2,0)
outputs = model(inputs2)
_, predicted = torch.max(outputs.data, 1)
#******************************* Print prediction range to match pics above ***********
# if (j>=range_plot and j<(range_plot+25)):
# print("j= ",j,' ',outputs.data.detach().cpu().numpy())
# print('prediction = ', predicted.detach().cpu().numpy())
#*************************************************************************************
total = total +1 # Increment the total count
match_freq = match_freq + freq_offset*freq_increment
if (predicted.detach().cpu().numpy() == 0):
c0 = c0 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 1):
c1 = c1 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 2):
c2 = c2 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 3):
c3 = c3 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 4):
c4 = c4 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 5):
c5 = c5 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 6):
c6 = c6 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 7):
c7 = c7 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 8):
noise = noise + 1
if (predicted.detach().cpu().numpy() == 9):
center_fft = center_fft + 1
if (predicted.detach().cpu().numpy() == 10):
c8 = c8 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 11):
c9 = c9 + 1
freq_histo[j] = match_freq/1000000
freq_offset = freq_offset + 1
if (freq_offset == seg_freq):
freq_offset = 0
match_freq = start_frequency
torch.cuda.empty_cache()
# complete ******************************************************
freq_histo = np.ma.masked_equal(freq_histo, 0)
histo_stats_freq_file(freq_histo,msps)
denom = total-center_fft-noise
print('************************* Probabilities ********************************')
print('----------------------------WAVEFORMS-----------------------------------')
if (denom == 0):
print('Nothing but noise')
else:
print('LoRa 125 = {:.2f}%'.format(c0/denom*100))
print('GD55 DMR = {:.2f}%'.format(c1/denom*100))
print('NFM = {:.2f}%'.format(c2/denom*100))
print('TYT = {:.2f}'.format(c3/denom*100))
print('Vodeson Doorbell = {:.2f}%'.format(c4/denom*100))
print('clickndig = {:.2f}%'.format(c5/denom*100))
print('Sado doorbell = {:.2f}%'.format(c6/denom*100))
print('LoRa 250 = {:.2f}%'.format(c7/denom*100))
print('light switch = {:.2f}%'.format(c8/denom*100))
print('YSF = {:.2f}%'.format(c9/denom*100))
print('------------------------------------------------------------------------')
print('***************************** noise and fft ****************************')
print('noise matches = ', noise)
print('center fft matches = ', center_fft)
print('TOTAL patch count = ', total)
print('***************************** Finished *********************************')
# inference ************************************************************
def testing_live(msps):
large_width = 400
np.set_printoptions(precision=2,floatmode='fixed', linewidth=large_width)
model.eval()
V = data.DataLoader(inference_dataloader_segmented(), batch_size=1)
start_frequency = (center_freq_live)
match_freq = start_frequency
print('start_frequency = ', start_frequency/1000000)
freq_offset = 0
total = noise = center_fft = target_to_int = accumulated_corrects = percent_correct = 0
c0 = c1 = c2 = c3 = c4 = c5 = c6 = c7 = c8 = c9 = 0
with torch.no_grad():
for i, rf_data in enumerate(V, 0):
accumulated_corrects = 0
percent_correct = 0
target_to_int = 0
inputs, target, counter, seg_freq = rf_data
print('testing counter = ', counter, 'seg_freq =', seg_freq)
print('seg_freq = ', seg_freq)
#**************************** Print segmented pics ***********************************
stft_plot = np.squeeze(inputs, axis=0)
fig=plt.figure(figsize=(8,8))
ncols = 5
nrows = 5
range_plot = 1
range_end = range_plot+5
for x in range(1,51): # need to figure out how to not hard code this ********************* <-----
if (x>=range_plot and x<(range_end)):
stft_plot1 = stft_plot[x, :, :, 1]
stft_plot1 = 10*np.log10(np.abs(stft_plot[x, :, :, 0]+eps))
fig.add_subplot(nrows, ncols, x-range_plot+1)
plt.imshow(stft_plot1, vmin=-70, vmax=5)
plt.show()
#******************************************************************************************
freq_increment = (Fs*msps/2)/seg_freq.detach().cpu().numpy().item()
print('freq_increment = ', freq_increment)
print('TESTING inputs SHAPE = ', inputs.shape)
target = Variable(target.cuda(GPU))
print('input in = ', inputs.shape)
inputs = torch.squeeze(inputs, dim=0)
print('input out = ', inputs.shape)
inputs = inputs.permute(0,3,1,2).contiguous()
print('counter convert stuff = ', counter, type(counter.numpy()))
inputs = Variable(inputs.cuda(GPU))
print('permuted shape = ', inputs.shape)
freq_count = 0 # keep track of array position
freq_histo = np.zeros(counter.numpy())
for j in range(counter):
inputs2 = inputs[j,:,:,:]
inputs2 = torch.unsqueeze(inputs2,0)
outputs = model(inputs2)
_, predicted = torch.max(outputs.data, 1)
#******************************* Print prediction range to match pics above ***********
# if (j>=range_plot and j<(range_end)):
# # print("j= ",j,' ',outputs.data.detach().cpu().numpy())
# print('prediction = ', predicted.detach().cpu().numpy())
# print('******************')
#*************************************************************************************
total = total +1 # Increment the total count
match_freq = match_freq + freq_offset*freq_increment
if (predicted.detach().cpu().numpy() == 0):
c0 = c0 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 1):
c1 = c1 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 2):
c2 = c2 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 3):
c3 = c3 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 4):
c4 = c4 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 5):
c5 = c5 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 6):
c6 = c6 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 7):
c7 = c7 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 8):
noise = noise + 1
if (predicted.detach().cpu().numpy() == 9):
center_fft = center_fft + 1
if (predicted.detach().cpu().numpy() == 10):
c8 = c8 + 1
freq_histo[j] = match_freq/1000000
if (predicted.detach().cpu().numpy() == 11):
c9 = c9 + 1
freq_histo[j] = match_freq/1000000
freq_offset = freq_offset + 1
if (freq_offset == seg_freq):
freq_offset = 0
match_freq = start_frequency
torch.cuda.empty_cache()
# complete ******************************************************
freq_histo = np.ma.masked_equal(freq_histo, 0)
histo_stats_freq_live(freq_histo,msps)
denom = total-center_fft-noise
print('************************* Probabilities ********************************')
print('----------------------------WAVEFORMS-----------------------------------')
if (denom == 0):
print('Nothing but noise')
else:
print('LoRa 125 = {:.2f}%'.format(c0/denom*100))
print('GD55 DMR = {:.2f}%'.format(c1/denom*100))
print('NFM = {:.2f}%'.format(c2/denom*100))
print('TYT = {:.2f}'.format(c3/denom*100))
print('Vodeson Doorbell = {:.2f}%'.format(c4/denom*100))
print('clickndig = {:.2f}%'.format(c5/denom*100))
print('Sado doorbell = {:.2f}%'.format(c6/denom*100))
print('LoRa 250 = {:.2f}%'.format(c7/denom*100))
print('light switch = {:.2f}%'.format(c8/denom*100))
print('YSF = {:.2f}%'.format(c9/denom*100))
print('------------------------------------------------------------------------')
print('***************************** noise and fft ****************************')
print('noise matches = ', noise)
print('center fft matches = ', center_fft)
print('TOTAL patch count = ', total)
print('***************************** Finished *********************************')
train_dataloader = RFDataset(path)
training_data = data.DataLoader(train_dataloader, batch_size=batches, shuffle=True)
# ## CUDA initialization
model = resnet18(2, 12) # number of input channels, number of classes
CUDA = torch.cuda.is_available()
if CUDA:
model.cuda(GPU)
CUDA
torch.cuda.empty_cache()
# ## Final training initialization
# momentum = .3
criterion = nn.CrossEntropyLoss()
lr= 1e-4
optimizer = optim.Adam(model.parameters(), lr=lr)
# optimizer = optim.SGD(model.parameters(), lr=lr,momentum=momentum)
# optimizer = optim.RMSprop(model.parameters(), lr=lr,momentum=momentum)
# optimizer = optim.SGD(model.parameters(), lr=lr)
# training_data = data.DataLoader(rf_dataset, batch_size=batches, shuffle=True)
model.train()
# VALIDATION ************************************************************
np.set_printoptions(threshold=np.inf)
calc = np.zeros([6])
averaging = 0
correct = 0
total = 0
V = data.DataLoader(inference_dataloader_segmented(), batch_size=1, shuffle=True)
model.eval()
# ### TRAIN Model
total = 100
loss_plot,total_plot = train_net(total)
path_plot_fig = "/home/david/sigMF_ML/RF/RF_class/plot_data/" # ACE
os.chdir(path_plot_fig)
# num = 20
np.save('resnet18_2D_20210227_10dB_1e4lr', np.asarray(total_plot))
np.save('resnet18_2D_20210227_10dB_1e4lr_2', total_plot)
os.chdir(path_fig)
plt.figure(figsize=(9, 6))
fig = plt.figure()
plt.plot(total_plot[:5],c='r', label='Total patches correct')
plt.legend(loc='lower right')
plt.title('Total % correct vs batches')
plt.xlabel('Batch number')
plt.ylabel('% correct')
plt.grid()
fig.savefig('ResNet50_v58_20210211_2D_10dB_noise_autosave_correct.pdf', format="pdf")
plt.show()
os.chdir(path_fig)
plt.figure(figsize=(9, 6))
fig = plt.figure()
plt.plot(loss_plot,c='r', label='Loss curve')
plt.legend(loc='upper right')
plt.title('Loss vs Epochs')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.grid()
fig.savefig('ResNet50_v58_20210211_2D_10dB_noise_autosave_loss.pdf', format="pdf")
plt.show()
# ### LIVE inferencing
# +
# # !python3 /home/david/sigMF_ML/gnuradio/record_live.py
# -
# #### 2D tensor with 300 noise
# vodeson - 10
# lora250 - 0
# lora125 -
# click - 10
# sado - 4ish
# light -
# tyt -
# GD55
# nfm -
#
# ### TESTING
msps = 25
center_freq_live = 428.0e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
# testing new decimator
# !python3 /home/david/sigMF_ML/gnuradio/record_live_25msps.py
# # !python3 /home/david/sigMF_ML/gnuradio/record_live1msps.py
# usrp_data_collect_1MSPS()
data_IQ_list_val, meta_list_val = inference_read(msps)
testing_live(msps)
torch.cuda.empty_cache()
msps = 5
center_freq_live = 433.0e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
# !python3 /home/david/sigMF_ML/gnuradio/record_live_5msps.py
# # !python3 /home/david/sigMF_ML/gnuradio/record_live1msps.py
# usrp_data_collect_1MSPS()
data_IQ_list_val, meta_list_val = inference_read(msps)
testing_live(msps)
torch.cuda.empty_cache()
msps = 1
center_freq_live = 433.65e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
# testing new decimator
# !python3 /home/david/sigMF_ML/gnuradio/record_live_25to1.py
# # !python3 /home/david/sigMF_ML/gnuradio/record_live_1msps.py
# usrp_data_collect_1MSPS()
data_IQ_list_val, meta_list_val = inference_read(msps)
testing_live(msps)
torch.cuda.empty_cache()
torch.cuda.empty_cache()
# ## testing pre-recorded files in /home/david/sigMF_ML/RF/RF_class/testing_data
msps = 25
center_freq_file = 428.00e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
data_IQ_list_val, meta_list_val = inference_read_file(msps, path_test_25msps)
testing_file(msps)
torch.cuda.empty_cache()
msps = 5
center_freq_file = 433.00e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
data_IQ_list_val, meta_list_val = inference_read_file(msps, path_test_5msps)
testing_file(msps)
torch.cuda.empty_cache()
msps = 1
center_freq_file = 433.65e6 # when SDR doing 25MSPS with center at 428MHz, or 433.65e6, 428.00e6
data_IQ_list_val, meta_list_val = inference_read_file(msps, path_test_1msps)
testing_file(msps)
torch.cuda.empty_cache()
torch.cuda.empty_cache()
# #### Save and Load model data
# #### save
# +
# torch.save(model.state_dict(), PATH)
# -
# #### load
# +
# device = torch.device("cuda:0")
# model = resnet50(2, 12)
# model.load_state_dict(torch.load(PATH))
# model.to(device)
# model.eval()
# +
# # Print model's state_dict
# print("Model's state_dict:")
# for param_tensor in model.state_dict():
# print(param_tensor, "\t", model.state_dict()[param_tensor].size())
# +
# # Print optimizer's state_dict
# print("Optimizer's state_dict:")
# for var_name in optimizer.state_dict():
# print(var_name, "\t", optimizer.state_dict()[var_name])
# +
# # SAVE MODEL
# os.chdir(path_save)
# torch.save({
# 'epoch': epoch,
# 'model_state_dict': model.state_dict(),
# 'optimizer_state_dict': optimizer.state_dict(),
# 'loss': loss,
# }, path_save+rf_model)
# +
# # LOAD MODEL
# checkpoint = torch.load(path_save+rf_model, map_location=device)
# # STATUS
# checkpoint.keys()
# epoch = checkpoint['epoch']
# model_state_dict = checkpoint['model_state_dict']
# optimizer_state_dict = checkpoint['optimizer_state_dict']
# loss = checkpoint['loss']
# optimizer_state_dict.keys()
# optimizer_state_dict['param_groups']
# loss
# model.load_state_dict(model_state_dict)
# +
# loss
# +
# optimizer_state_dict.keys()
# -
| RF_class_v58 rev1_2D RESNET MULE1-long run-1e-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Convolution
#
# As the name implies, convolution operations are an important component of convolutional neural networks. The ability for a CNN to accurately match diverse patterns can be attributed to using convolution operations. These operations require complex input which was shown in the previous section. In this section we'll experiment with convolution operations and the parameters which are available to tune them.
#
# 
#
# <p style="text-align: center;"><i>Convolution operation convolving two input tensors (input and kernel) into a single output tensor which represents information from each input.</i></p>
# <br />
#
# ### Input and Kernel
#
# Convolution operations in TensorFlow are done using `tf.nn.conv2d` in a typical situation. There are other convolution operations available using TensorFlow designed with special use cases. `tf.nn.conv2d` is the preferred convolution operation to begin experimenting with. For example, we can experiment with convolving two tensors together and inspect the result.
# setup-only-ignore
import tensorflow as tf
import numpy as np
# setup-only-ignore
sess = tf.InteractiveSession()
# +
input_batch = tf.constant([
[ # First Input
[[0.0], [1.0]],
[[2.0], [3.0]]
],
[ # Second Input
[[2.0], [4.0]],
[[6.0], [8.0]]
]
])
kernel = tf.constant([
[
[[1.0, 2.0]]
]
])
# -
# The example code creates two tensors. The `input_batch` tensor has a similar shape to the `image_batch` tensor seen in the previous section. This will be the first tensor being convolved and the second tensor will be `kernel`. *Kernel* is an important term that is interchangeable with *weights*, *filter*, *convolution matrix* or *mask*. Since this task is computer vision related, it's useful to use the term kernel because it is being treated as an [image kernel](https://en.wikipedia.org/wiki/Kernel_(image_processing). There is no practical difference in the term when used to describe this functionality in TensorFlow. The parameter in TensorFlow is named `filter` and it expects a set of weights which will be learned from training. The amount of different weights included in the kernel (`filter` parameter) will configure the amount of kernels which will be learned.
#
# In the example code, there is a single kernel which is the first dimension of the `kernel` variable. The kernel is built to return a tensor which will include one channel with the original input and a second channel with the original input doubled. In this case, channel is used to describe the elements in a rank 1 tensor (vector). Channel is a term from computer vision which describes the output vector, for example an RGB image has three channels represented as a rank 1 tensor `[red, green, blue]`. At this time, ignore the `strides` and `padding` parameter which will be covered later and focus on the convolution (`tf.nn.conv2d`) output.
# +
conv2d = tf.nn.conv2d(input_batch, kernel, strides=[1, 1, 1, 1], padding='SAME')
sess.run(conv2d)
# -
# The output is another tensor which is the same rank as the `input_batch` but includes the number of dimensions found in the kernel. Consider if `input_batch` represented an image, the image would have a single channel, in this case it could be considered a grayscale image (see [Working with Colors](#working-with-colors)). Each element in the tensor would represent one pixel of the image. The pixel in the bottom right corner of the image would have the value of `3.0`.
#
# Consider the `tf.nn.conv2d` convolution operation as a combination of the image (represented as `input_batch`) and the `kernel` tenser. The convolution of these two tensors create a feature map. Feature map is a broad term except in computer vision where it relates to the output of operations which work with an image kernel. The feature map now represents the convolution of these tensors by adding new layers to the output.
#
# The relationship between the input images and the output feature map can be explored with code. Accessing elements from the input batch and the feature map are done using the same index. By accessing the same pixel in both the input and the feature map shows how the input was changed when it convolved with the `kernel`. In the following case, the lower right pixel in the image was changed to output the value found by multiplying <span class="math-tex" data-type="tex">\\(3.0 \* 1.0\\)</span> and <span class="math-tex" data-type="tex">\\(3.0 \* 2.0\\)</span>. The values correspond to the pixel value and the corresponding value found in the `kernel`.
# +
lower_right_image_pixel = sess.run(input_batch)[0][1][1]
lower_right_kernel_pixel = sess.run(conv2d)[0][1][1]
lower_right_image_pixel, lower_right_kernel_pixel
# -
# In this simplified example, each pixel of every image is multiplied by the corresponding value found in the kernel and then added to a corresponding layer in the feature map. Layer, in this context, is referencing a new dimension in the output. With this example, it's hard to see a value in convolution operations.
#
# ### Strides
#
# The value of convolutions in computer vision is their ability to reduce the dimensionality of the input, which is an image in this case. An image's dimensionality (2D image) is its width, height and number of channels. A large image dimensionality requires an exponentially larger amount of time for a neural network to scan over every pixel and judge which ones are important. Reducing dimensionality of an image with convolutions is done by altering the `strides` of the kernel.
#
# The parameter `strides`, causes a kernel to skip over pixels of an image and not include them in the output. It's not fair to say the pixels are skipped because they still may affect the output. The `strides` parameter highlights how a convolution operation is working with a kernel when a larger image and more complex kernel are used. As a convolution is sliding the kernel over the input, it's using the strides parameter to change how it walks over the input. Instead of going over every element of an input the `strides` parameter could configure the convolution to skip certain elements.
#
# For example, take the convolution of a larger image and a larger kernel. In this case, it's a convolution between a 6 pixel tall, 6 pixel wide and 1 channel deep image (6x6x1) and a (3x3x1) kernel.
# +
input_batch = tf.constant([
[ # First Input (6x6x1)
[[0.0], [1.0], [2.0], [3.0], [4.0], [5.0]],
[[0.1], [1.1], [2.1], [3.1], [4.1], [5.1]],
[[0.2], [1.2], [2.2], [3.2], [4.2], [5.2]],
[[0.3], [1.3], [2.3], [3.3], [4.3], [5.3]],
[[0.4], [1.4], [2.4], [3.4], [4.4], [5.4]],
[[0.5], [1.5], [2.5], [3.5], [4.5], [5.5]],
],
])
kernel = tf.constant([ # Kernel (3x3x1)
[[[0.0]], [[0.5]], [[0.0]]],
[[[0.0]], [[1.0]], [[0.0]]],
[[[0.0]], [[0.5]], [[0.0]]]
])
# NOTE: the change in the size of the strides parameter.
conv2d = tf.nn.conv2d(input_batch, kernel, strides=[1, 3, 3, 1], padding='SAME')
sess.run(conv2d)
# -
# The `input_batch` was combined with the `kernel` by moving the `kernel` over the `input_batch` striding (or skipping) over certain elements. Each time the `kernel` was moved, it get centered over an element of `input_batch`. Then the overlapping values are multiplied together and the result is added together. This is how a convolution combines two inputs using what's referred to as pointwise multiplication. It may be easier to visualize using the following figure.
#
# 
#
# In this figure, the same logic is done as what is found in the code. Two tensors convolved together while striding over the input. The strides reduced the dimensionality of the output a large amount while the kernel size allowed the convolution to use all the input values. None of the input data was completely removed from striding but now the input is a smaller tensor.
#
# Strides are a way to adjust the dimensionality of input tensors. Reducing dimensionality requires less processing power, and will keep from creating receptive fields which completely overlap. The `strides` parameter follows the same format as the input tensor `[image_batch_size_stride, image_height_stride, image_width_stride, image_channels_stride]`. Changing the first or last element of the stride parameter are rare, they'd skip data in a `tf.nn.conv2d` operation and not take the input into account. The `image_height_stride` and `image_width_stride` are useful to alter in reducing input dimensionality.
#
# A challenge which comes up often with striding over the input is how to deal with a stride which doesn't evenly end at the edge of the input. The uneven striding will come up often due to image size and kernel size not matching the striding. If the image size, kernel size and strides can't be changed then padding can be added to the image to deal with the uneven area.
# ### Padding
#
# When a kernel is overlapped on an image it should be set to fit within the bounds of the image. At times, the sizing may not fit and a good alternative is to fill the missing area in the image. Filling the missing area of the image is known as padding the image. TensorFlow will pad the image with zeros or raise an error when the sizes don't allow a kernel to stride over an image without going past its bounds. The amount of zeros or the error state of `tf.nn.conv2d` is controlled by the parameter `padding` which has two possible values ('VALID', 'SAME').
#
# **SAME:** The convolution output is the **SAME** size as the input. This doesn't take the filter's size into account when calculating how to stride over the image. This may stride over more of the image than what exists in the bounds while padding all the missing values with zero.
#
# **VALID:** Take the filter's size into account when calculating how to stride over the image. This will try to keep as much of the kernel inside the image's bounds as possible. There may be padding in some cases but will avoid.
#
# It's best to consider the size of the input but if padding is necessary then TensorFlow has the option built in. In most simple scenarios, `SAME` is a good choice to begin with. `VALID` is preferential when the input and kernel work well with the strides. For further information, TensorFlow covers this subject well in the [convolution documentation](https://www.tensorflow.org/versions/r0.8/api_docs/python/nn.html#convolution).
# ### Data Format
#
# There's another parameter to `tf.nn.conv2d` which isn't shown from these examples named `data_format`. The [`tf.nn.conv2d` docs](https://www.tensorflow.org/versions/r0.8/api_docs/python/nn.html#conv2d) explain how to change the data format so the `input`, `kernel` and `strides` follow a format other than the format being used thus far. Changing this format is useful if there is an input tensor which doesn't follow the `[batch_size, height, width, channel]` standard. Instead of changing the input to match, it's possible to change the `data_format` parameter to use a different layout.
#
# > data_format: An optional string from: "NHWC", "NCHW". Defaults to "NHWC". Specify the data format of the input and output data. With the default format "NHWC", the data is stored in the order of: [batch, in_height, in_width, in_channels]. Alternatively, the format could be "NCHW", the data storage order of: [batch, in_channels, in_height, in_width].
#
# | Data Format | Definition |
# |:---: | :---: |
# | N | Number of tensors in a batch, the `batch_size`. |
# | H | Height of the tensors in each batch. |
# | W | Width of the tensors in each batch. |
# | C | Channels of the tensors in each batch. |
# ### Kernels in Depth
#
# In TensorFlow the filter parameter is used to specify the kernel convolved with the input. Filters are commonly used in photography to adjust attributes of a picture, such as the amount of sunlight allowed to reach a camera's lens. In photography, filters allow a photographer to drastically alter the picture they're taking. The reason the photographer is able to alter their picture using a filter is because the filter can recognize certain attributes of the light coming in to the lens. For example, a red lens filter will absorb (block) every frequency of light which isn't red allowing only red to pass through the filter.
#
# 
#
# In computer vision, kernels (filters) are used to recognize important attributes of a digital image. They do this by using certain patterns to highlight when features exist in an image. A kernel which will replicate the red filter example image is implemented by using a reduced value for all colors except red. In this case, the reds will stay the same but all other colors matched are reduced.
#
# The example seen at the [start of this chapter](#convolutional-neural-networks) uses a kernel designed to do edge detection. Edge detection kernels are common in computer vision applications and could be implemented using basic TensorFlow operations and a single `tf.nn.conv2d` operation.
# +
# setup-only-ignore
import matplotlib as mil
#mil.use('svg')
mil.use("nbagg")
from matplotlib import pyplot
fig = pyplot.gcf()
fig.set_size_inches(4, 4)
image_filename = "./images/chapter-05-object-recognition-and-classification/convolution/n02113023_219.jpg"
# image_filename = "/Users/erikerwitt/Downloads/images/n02085936-Maltese_dog/n02085936_804.jpg"
filename_queue = tf.train.string_input_producer(tf.train.match_filenames_once(image_filename))
image_reader = tf.WholeFileReader()
_, image_file = image_reader.read(filename_queue)
image = tf.image.decode_jpeg(image_file)
sess.run(tf.global_variables_initializer())
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(coord=coord)
image_batch = tf.image.convert_image_dtype(tf.expand_dims(image, 0), tf.float32, saturate=False)
# +
kernel = tf.constant([
[
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]]
],
[
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ 8., 0., 0.], [ 0., 8., 0.], [ 0., 0., 8.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]]
],
[
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]]
]
])
conv2d = tf.nn.conv2d(image_batch, kernel, [1, 1, 1, 1], padding="SAME")
activation_map = sess.run(tf.minimum(tf.nn.relu(conv2d), 255))
# -
# setup-only-ignore
fig = pyplot.gcf()
pyplot.imshow(activation_map[0], interpolation='nearest')
#pyplot.show()
fig.set_size_inches(4, 4)
fig.savefig("./images/chapter-05-object-recognition-and-classification/convolution/example-edge-detection.png")
# 
#
# The output created from convolving an image with an edge detection kernel are all the areas where and edge was detected. The code assumes a batch of images is already available (`image_batch`) with a real image loaded from disk. In this case, the image is an example image found in the Stanford Dogs Dataset. The kernel has three input and three output channels. The channels sync up to RGB values between <span class="math-tex" data-type="tex">\\([0, 255]\\)</span> with 255 being the maximum intensity. The `tf.minimum` and `tf.nn.relu` calls are there to keep the convolution values within the range of valid RGB colors of <span class="math-tex" data-type="tex">\\([0, 255]\\)</span>.
#
# There are [many other](https://en.wikipedia.org/wiki/Kernel_(image_processing)) common kernels which can be used in this simplified example. Each will highlight different patterns in an image with different results. The following kernel will sharpen an image by increasing the intensity of color changes.
# +
kernel = tf.constant([
[
[[ 0., 0., 0.], [ 0., 0., 0.], [ 0., 0., 0.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ 0., 0., 0.], [ 0., 0., 0.], [ 0., 0., 0.]]
],
[
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ 5., 0., 0.], [ 0., 5., 0.], [ 0., 0., 5.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]]
],
[
[[ 0., 0., 0.], [ 0., 0., 0.], [ 0., 0., 0.]],
[[ -1., 0., 0.], [ 0., -1., 0.], [ 0., 0., -1.]],
[[ 0, 0., 0.], [ 0., 0., 0.], [ 0., 0., 0.]]
]
])
conv2d = tf.nn.conv2d(image_batch, kernel, [1, 1, 1, 1], padding="SAME")
activation_map = sess.run(tf.minimum(tf.nn.relu(conv2d), 255))
# -
# setup-only-ignore
fig = pyplot.gcf()
pyplot.imshow(activation_map[0], interpolation='nearest')
#pyplot.show()
fig.set_size_inches(4, 4)
fig.savefig("./images/chapter-05-object-recognition-and-classification/convolution/example-sharpen.png")
# 
#
# The values in the kernel were adjusted with the center of the kernel increased in intensity and the areas around the kernel reduced in intensity. The change, matches patterns with intense pixels and increases their intensity outputting an image which is visually sharpened. Note that the corners of the kernel are all `0` and don't affect the output which operates in a plus shaped pattern.
#
# These kernels match patterns in images at a rudimentary level. A convolutional neural network matches edges and more by using a complex kernel it learned during training. The starting values for the kernel are usually random and over time they're trained by the CNN's learning layer. When a CNN is complete, it starts running and each image sent in is convolved with a kernel which is then changed based on if the predicted value matches the labeled value of the image. For example, if a Sheepdog picture is considered a Pit Bull by the CNN being trained it will then change the filters a small amount to try and match Sheepdog pictures better.
#
# Learning complex patterns with a CNN involves more than a single layer of convolution. Even the example code included a `tf.nn.relu` layer used to prepare the output for visualization. Convolution layers may occur more than once in a CNN but they'll likely include other layer types as well. These layers combined form the support network required for a successful CNN architecture.
# setup-only-ignore
filename_queue.close(cancel_pending_enqueues=True)
coord.request_stop()
coord.join(threads)
| chapters/05_object_recognition_and_classification/Chapter 5 - 02 Convolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is basically an update on some parts of the overview tutorial from TF Probability.
# I've changed some distributions, and experimented with other bijectors.
import tensorflow as tf
import tensorflow_probability as tfp
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
tfd = tfp.distributions
# +
sns.set()
tfd = tfp.distributions
tfb = tfp.bijectors
# -
# # Distributions
normal = tfd.Normal(loc=0., scale=1.)
print(normal)
samples = normal.sample(1000)
sns.displot(samples)
plt.title("Samples from a standard Normal")
plt.show()
normal.log_prob(0.)
ng_bin=tfd.NegativeBinomial(total_count=12,probs=0.5)
# Given a Bernoulli trial with probability p of success,
# the NegativeBinomial distribution represents the distribution over the number of
# successes s that occur until we observe f failures.
samples2 = ng_bin.sample(10000)
sns.displot(samples2)
plt.title("Samples from a Negative Binomial")
plt.show()
# TensorFlow Probability Distributions have shape semantics -- we partition shapes into semantically distinct pieces, even though the same chunk of memory (Tensor/ndarray) is used for the whole everything.
#
# + Batch shape denotes a collection of Distributions with distinct parameters
# + Event shape denotes the shape of samples from the Distribution.
# Create a batch of 3 normals, and plot 1000 samples from each
normals = tfd.Normal([-2.5, 0., 2.5], 1.) # The scale parameter broadacasts!
print("Batch shape:", normals.batch_shape)
print("Event shape:", normals.event_shape)
samples = normals.sample(1000)
print("Shape of samples:", samples.shape)
sns.displot(samples[:,0])
plt.title("Samples from a Normal(-2.5, 1)")
plt.show()
# A batch of normals gives a batch of log_probs.
print(normals.log_prob([-2.5, 0., 2.5]))
print(normals.log_prob(0.))
xs = np.linspace(-6, 6, 200)[:,np.newaxis]
for i in range(3):
sns.histplot(samples[:, i], kde=False, stat="density")
plt.plot(np.tile(xs, 3), normals.prob(xs), c='k', alpha=.5)
plt.title("Samples from 3 Normals, and their PDF's")
plt.show()
# ## Vector Distribution
covariance_matrix = [[1., 1],[1,2.]]
mvn = tfd.MultivariateNormalTriL(loc=[0., 0.],scale_tril=tf.linalg.cholesky(covariance_matrix))
print("Batch shape:", mvn.batch_shape)
print("Event shape:", mvn.event_shape)
samples = mvn.sample(1000)
print("Samples shape:", samples.shape)
samples[0,1]
sns.jointplot(x=samples[:, 0], y=samples[:, 1], kind='scatter')
plt.show()
# +
mu = [1., 2]
cov = [[ 1, 0.5],
[ 0.5, 1]]
mvn = tfd.MultivariateNormalFullCovariance(
loc=mu,
covariance_matrix=cov)
samples = mvn.sample(1000)
print("Samples shape:", samples.shape)
# -
sns.jointplot(x=samples[:, 0], y=samples[:, 1], kind='scatter')
plt.show()
# ## Matrix Distributions
lkj = tfd.LKJ(dimension=10, concentration=[1.5, 3.0])
print("Batch shape: ", lkj.batch_shape)
print("Event shape: ", lkj.event_shape)
# Event Shape = Dimension of each Distribution
#
# Batch Shape = Number of Distributions
samples = lkj.sample()
print("Samples shape: ", samples.shape)
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(6, 3))
sns.heatmap(samples[0,:], ax=axes[0], cbar=False)
sns.heatmap(samples[1,:], ax=axes[1], cbar=False)
fig.tight_layout()
plt.show()
# # Gaussian Distributions
# ## Prior
kernel = tfp.math.psd_kernels.ExponentiatedQuadratic()
xs = np.linspace(-5., 5., 200).reshape([-1, 1])
gp = tfd.GaussianProcess(kernel, index_points=xs)
print("Batch shape:", gp.batch_shape)
print("Event shape:", gp.event_shape)
upper, lower = gp.mean() + [2 * gp.stddev(), -2 * gp.stddev()]
plt.plot(xs, gp.mean())
plt.fill_between(xs[:, 0], upper, lower, color='k', alpha=.1)
for _ in range(10):
plt.plot(xs, gp.sample(), c='r', alpha=.3)
plt.title(r"GP prior mean, $2\sigma$ intervals, and samples")
plt.show()
# ## GP Regression
# +
# Suppose we have some observed data
obs_x = [[-3.], [0.], [2.]] # Shape 3x1 (3 1-D vectors)
obs_y = [3., -2., 2.] # Shape 3 (3 scalars)
# Here's the regression model
gprm = tfd.GaussianProcessRegressionModel(kernel, xs, obs_x, obs_y)
# -
upper, lower = gprm.mean() + [2 * gprm.stddev(), -2 * gprm.stddev()]
plt.plot(xs, gprm.mean())
plt.fill_between(xs[:, 0], upper, lower, color='k', alpha=.1)
for _ in range(5):
plt.plot(xs, gprm.sample(), c='r', alpha=.3)
plt.scatter(obs_x, obs_y, c='k', zorder=3)
plt.title(r"GP posterior mean, $2\sigma$ intervals, and samples")
plt.show()
# # Bijectors (invertible, smooth functions)
normal_cdf = tfp.bijectors.NormalCDF()
xs = np.linspace(-4., 4., 200)
plt.plot(xs, normal_cdf.forward(xs))
plt.show()
# +
exp_bijector = tfp.bijectors.Exp()
log_normal = exp_bijector(tfd.Normal(0., 1))
samples = log_normal.sample(1000)
xs = np.linspace(1e-10, np.max(samples), 200)
sns.histplot(samples, stat="density", kde=False)
#plt.plot(xs, log_normal.prob(xs), c='k', alpha=.75)
log_normal_direct=tfp.distributions.LogNormal(
loc=0, scale=1)# Exp(N(0,1)) = Log-Normal(0,1)
plt.plot(xs, log_normal_direct.prob(xs), c='r', alpha=.75)
# Works well
plt.show()
# +
square_bijector=tfp.bijectors.Square()
chi_squared=square_bijector(tfd.Normal(0., 1))
samples = chi_squared.sample(1000)
xs = np.linspace(0.1, np.max(samples), 200)
# I changed the mininmum, otherwise, the graph would be a bit ugly...
sns.histplot(samples, stat="density", kde=False)
chi_squared_direct=tfp.distributions.Chi2(
df=1)# (N(0,1))^2 = Chi-Squared(1)
plt.plot(xs, chi_squared_direct.prob(xs), c='r', alpha=.75)
# Works well
plt.show()
# -
# The first bin seems to go over the density of the chi-squared, but that's only because I cut the x axis for the density at 0.1. Change the x_min to 0.01 above, and it will go very easily over it.
# ## Batching Bijectors
softplus = tfp.bijectors.Softplus(
hinge_softness=[1., .5, .1])
print("Hinge softness shape:", softplus.hinge_softness.shape)
# This bijector $g$ with hinge_softness equal to $c$ is such that $$Y = g(X) = c\log\left(1 + \exp\left(\frac{X}{c}\right)\right)$$
xs = np.linspace(-4., 4., 200)[:, np.newaxis]
ys = softplus.forward(xs)
print("Forward shape:", ys.shape)
# +
lines = plt.plot(np.tile(xs, reps=3), ys)
# numpy.tile - Construct an array by repeating A the number of times given by reps.
for line, hs in zip(lines, softplus.hinge_softness):
line.set_label("Softness: %1.1f" % hs)
plt.legend()
plt.show()
# +
bij = tfb.CholeskyOuterProduct()
size = 2500
# Make a big, lower-triangular matrix
big_lower_triangular = tf.eye(size)
# Squaring it gives us a positive-definite matrix
big_positive_definite = bij.forward(big_lower_triangular)
# Caching for the win!
# %timeit bij.inverse(big_positive_definite)
# %timeit tf.linalg.cholesky(big_positive_definite)
# -
# Yes, my pc is that slow (and old).
| Tensorflow-Probability/Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.4 自定义层
# ## 4.4.1 不含模型参数的自定义层
# +
import torch
from torch import nn
print(torch.__version__)
# -
class CenteredLayer(nn.Module):
def __init__(self, **kwargs):
super(CenteredLayer, self).__init__(**kwargs)
def forward(self, x):
return x - x.mean()
layer = CenteredLayer()
layer(torch.tensor([1, 2, 3, 4, 5], dtype=torch.float))
net = nn.Sequential(nn.Linear(8, 128), CenteredLayer())
y = net(torch.rand(4, 8))
y.mean().item()
# ## 4.4.2 含模型参数的自定义层
class MyListDense(nn.Module):
def __init__(self):
super(MyListDense, self).__init__()
self.params = nn.ParameterList([nn.Parameter(torch.randn(4, 4)) for i in range(3)])
self.params.append(nn.Parameter(torch.randn(4, 1)))
def forward(self, x):
for i in range(len(self.params)):
x = torch.mm(x, self.params[i])
return x
net = MyListDense()
print(net)
# +
class MyDictDense(nn.Module):
def __init__(self):
super(MyDictDense, self).__init__()
self.params = nn.ParameterDict({
'linear1': nn.Parameter(torch.randn(4, 4)),
'linear2': nn.Parameter(torch.randn(4, 1))
})
self.params.update({'linear3': nn.Parameter(torch.randn(4, 2))}) # 新增
def forward(self, x, choice='linear1'):
return torch.mm(x, self.params[choice])
net = MyDictDense()
print(net)
# -
x = torch.ones(1, 4)
print(net(x, 'linear1'))
print(net(x, 'linear2'))
print(net(x, 'linear3'))
net = nn.Sequential(
MyDictDense(),
MyListDense(),
)
print(net)
print(net(x))
# + jupyter={"outputs_hidden": true}
| code/chapter04_DL_computation/4.4_custom-layer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Rolling Mean
# **TL;DR** This notebook demonstrates the performance improvement of using a numba JIT compiled algorithm for calculating rolling mean over the Pandas equivalent for some sample data.
# +
from numba import jit
import pandas as pd
import numpy as np
import time
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
from matplotlib import rcParams
from matplotlib import pyplot as plt
rcParams['figure.figsize'] = 16, 8
import os
import sys
nb_dir = os.path.split(os.getcwd())[0]
if nb_dir not in sys.path:
sys.path.append(nb_dir)
# -
from utilities.rolling_stats import rolling_mean # this is the function we're going to test versus pandas
x = np.arange(30).astype(float)
s = pd.Series(x)
s[0] = np.nan
s[6] = np.nan
s[12:18] = np.nan
s[-1] = np.nan
s.values # arbitrary but small input data
s.rolling(window=3).mean().values # pandas output
rolling_mean(s.values, 3) # rolling_sum output
a = s.rolling(window=3).mean().values
b = rolling_mean(s.values, 3)
np.allclose(a, b, equal_nan=True)
def benchmarks():
res = []
for exponent in range(3, 7):
n = 10**exponent
data = np.arange(n).astype(float)
data[3] = np.nan
data[4] = np.nan
data[-1] = np.nan
s = pd.Series(data)
window = int(max(1000, n * 0.1)) # cap window size at 1,000
t1 = time.time()
pandas_output = s.rolling(window=window).mean().values
t2 = time.time()
res.append(('pandas', n, (t2 - t1)))
t1 = time.time()
rmean_output = rolling_mean(s.values, window)
t2 = time.time()
res.append(('rolling_mean', n, (t2 - t1)))
assert np.allclose(pandas_output, rmean_output, equal_nan=True)
return res
# +
data = benchmarks()
df = pd.DataFrame(data, columns = ['fn', 'population', 'time (ms)'])
df['time (ms)'] = df['time (ms)'].apply(lambda x: x * 1000.)
df = pd.pivot_table(df, values='time (ms)', index=['population'], columns=['fn'], aggfunc=np.sum)
df
# -
df.plot(logx=True)
plt.ylabel('time (ms)')
| notebooks/rolling_mean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overwatch Hero Survey (September 2019)
# Firstly, thank you to the 1200 players from [/r/cow](https://reddit.com/r/competitiveoverwatch) who took the time to fill out the survey.
#
# **Links**
# * Responses are based on this [Google Forms survey](https://forms.gle/Rgmn2CdViBApsbP6A)
# * The raw results can be found in this [Google Sheets spreadsheet](https://docs.google.com/spreadsheets/d/1Ax1unE4I2DWW3iz2Bv4c24FTUx_qxNPwrKGZ6LGNux0)
# * Original [reddit post](https://www.reddit.com/r/Competitiveoverwatch/comments/d9cmo9/5_minute_overwatch_hero_survey_fun_vs_perception/)
# * The code used to generate the results can be found on [github](https://github.com/marksimpson82/overwatch_survey).
#
# I encourage folk to check out the [github repo](https://github.com/marksimpson82/overwatch_survey) and examine/fork & modify the code.
#
# ## Caveats
# 1. I'm only human; the survey and analysis may contain bias or errors.
# 2. The survey might tell us _what_ people think, but it doesn't really tell us _why_.
# 3. It also lacks a little context: e.g. do people find hero X unfun to play against because the hero's kit is inherently flawed, or is it because of the meta?
#
# ## Goals
# I'm curious about how the community perceives hero fun (playing as and against a specific hero) vs. balance and whether we can spot any patterns in the responses.
#
# ## Future survey improvements
# If I run the survey again in future, I plan to:
# * Collect platform (PC, PS4, Xbox).
# * Collect whether people enjoy playing _with_ a hero on their team.
# * Collect general comments at the end.
# * Re-word a few questions, specifically:
# * Make main/most-played questions more explicit (it makes more sense to ask about the last season or two rather than historical data collected over 3+ years -- Rein today is not Rein of 2016!)
# +
import pandas as pd
import seaborn as sb
import ipywidgets as widgets
import numpy as np
from ipywidgets import interact, interactive, fixed, interact_manual
from matplotlib import pyplot as plt
from matplotlib.ticker import PercentFormatter
# load flat csv files -- the survey results are a little clunky
hero_names_df = pd.read_csv("../data/overwatch_heroes.csv").loc[:,["Hero_name"]]
basic_info_df = pd.read_csv("../data/table_basic_info.csv")
hero_ratings_df = pd.read_csv("../data/table_hero_ratings.csv")
# join on Response_id such that we can more easily query the result set
joined_df = basic_info_df.set_index('Response_id').join(
hero_ratings_df.set_index('Response_id'),
lsuffix='caller',
rsuffix='_other')
# +
# handy constants
HERO_COUNT=len(hero_names_df)
COLUMN_PLAYING_AS="Playing_as"
COLUMN_PLAYING_VS="Playing_vs"
RATING_MIN=1
RATING_MAX=5
CHART_BAR_FIGSIZE = (6, 12)
CHART_HIST_FIGSIZE = (6, 6)
# -
# helper functions for graphs
def set_graph_format_x_axis(ax):
ax.set(xlim=(RATING_MIN, RATING_MAX))
plt.xticks(np.arange(RATING_MIN, RATING_MAX + 1, 1))
def draw_fun_factor_graph(df, title):
f, ax = plt.subplots(figsize=CHART_BAR_FIGSIZE)
ax = sb.barplot(x="Value", y="Hero", data=df)
set_graph_format_x_axis(ax)
ax.set_title(title)
ax.set(xlabel="Fun factor (higher is more enjoyable)", ylabel="")
# ## General responses
# ### Q: Which game modes do people play the most?
# +
def draw_game_mode_chart(df):
game_modes = (df['Game_mode']
.value_counts(normalize=True, sort=False)
.sort_index())
figsize=(CHART_BAR_FIGSIZE[0], 4)
f, ax = plt.subplots(figsize=figsize)
ax = sb.barplot(x=game_modes.values, y=game_modes.index)
plt.gca().xaxis.set_major_formatter(PercentFormatter(1))
ax.set_title("Most-played game modes")
draw_game_mode_chart(basic_info_df)
# -
# #### Notes
# Pretty self-explanatory. Roughly 40% of respondents play non-competitive game modes.
# ### Q: What rank are they?
# +
def draw_rank_graph(df):
hero_time = (df['Rank']
.value_counts(normalize=True, sort=False))
# re-order based on ascending rank so we can see the distribution
hero_time = hero_time[[
"N/A (don't play comp)",
'Bronze',
'Silver',
'Gold',
'Platinum',
'Diamond',
'Masters',
'Grandmasters',
't500'
]]
palette = sb.color_palette(
palette=[
"red",
"peru",
"darkgrey",
"gold","lightgrey",
"lightskyblue",
"orange",
"yellow",
"paleturquoise"
],
n_colors=9)
f, ax = plt.subplots(figsize=(CHART_BAR_FIGSIZE[0], 4))
ax = sb.barplot(x=hero_time.values, y=hero_time.index, palette=palette, linewidth=0.5, edgecolor=".2")
plt.gca().xaxis.set_major_formatter(PercentFormatter(1))
ax.set_title("Rank")
draw_rank_graph(basic_info_df)
# -
# #### Notes
# This confirms what I'd expect to see -- the members of [/r/cow](https://reddit.com/r/competitiveoverwatch) tend to be higher ranked players compared to the general population. Hero choices and win rates change as players climb the ranks, so this will have a bearing on the answers.
# ### Q: What roles do people play?
# +
def draw_role_chart(df):
roles_played_raw = (df['Most_played_roles']
.value_counts(normalize=True, sort=False)
.sort_index())
TARGET_COLUMNS = ['Damage', 'Support', 'Tank']
# avert your eyes -- this is messy. I'm really just looking for a simple pandas aggregation
# method where => "if an index contains the string 'Damage', add its count to the 'Damage' sum"
roles_played = roles_played_raw[TARGET_COLUMNS].copy()
# zero these out, as we're going to re-count the originals down below
roles_played['Damage'] = 0
roles_played['Support'] = 0
roles_played['Tank'] = 0
for index, value in roles_played_raw.iteritems():
for target_col in TARGET_COLUMNS:
if target_col in index:
roles_played[target_col] += value
figsize=(CHART_BAR_FIGSIZE[0], 2.4)
f, ax = plt.subplots(figsize=figsize)
ax = sb.barplot(x=roles_played.values, y=roles_played.index)
plt.gca().xaxis.set_major_formatter(PercentFormatter(1))
ax.set_title("Most-played roles")
draw_role_chart(basic_info_df)
# -
# #### Notes
# Respondents were able to select multiple roles. The results are weighted much more towards `Tanks` & `Supports` than `Damage` (aka `DPS`), which I wasn't expecting. This colours the survey results somewhat, so bear that in mind.
# ### Q: Which heroes do people main?
# Rationale: Mains and favourites may well differ. While certain heroes are attractive choices due to their strength or skill curve, people may play a hero in competitive to keep the peace ("don't choose Torb or I'm staying in spawn!")
def draw_hero_percent_graph(df, hero_names_df, column_name, title):
hero_time = (df[column_name]
.value_counts(normalize=True))
# fill in missing data where heroes got 0 votes
hero_time = hero_time.reindex(hero_names_df["Hero_name"].values, fill_value=0.0)
hero_time.sort_values(inplace=True, ascending=False)
assert(len(hero_time) == HERO_COUNT)
f, ax = plt.subplots(figsize=CHART_BAR_FIGSIZE)
ax = sb.barplot(x=hero_time.values, y=hero_time.index)
plt.gca().xaxis.set_major_formatter(PercentFormatter(1))
ax.set_title(title)
plt.axvline(1.0/HERO_COUNT, 0.0, 1.0).set_linestyle('dashed')
draw_hero_percent_graph(basic_info_df, hero_names_df, "Hero_time_1st", "Most-played heroes")
# #### Notes
# Caveats:
#
# Firstly, the wording of this question was a ambiguous -- I didn't explicitly ask for the highest overall play time in their prefered game mode _or_ the most-played hero over the last season or two. I.e. respondents would've anwswered based their interpretation of the question. My hunch is that most people took the question literally and chose their most-played hero, but I can't be certain.
#
# Secondly, `Damage` heroes (aka `DPS`) have to compete with many more heroes in their own role, so their playtime is somewhat diluted. E.g. For much of the game's life, `Ana` had to compete with `Lucio`, `Mercy` and `Zenyatta` whereas `Junkrat` had to compete with a large cast of damage dealers.
#
# There's a number of heroes that were released at launch or shortly thereafter that sit low in the responses. That being said, heroes like `Torbjorn`, `Symmetra` and `Bastion` have always been situational.
# ### Q: Which heroes are our favourites?
# Rationale: Again, favourites are not equivalent to main heroes. E.g. I used to main `Tracer`, but she's been weak for over a year. `Tracer` remains my favourite hero, though.
draw_hero_percent_graph(basic_info_df, hero_names_df, "Hero_favourite", "Favourite heroes")
# #### Notes
# ##### We appreciate many of the older heroes
# Older heroes tend to dominate in the favourite category. `Ana` is the runaway favourite; she's 50% more popular than the next challenger!
#
# `<NAME>` is the highest placed favourite 'new' hero by quite some margin (I'm not including `Ana` as a 'new' hero as she's been around forever).
#
# Finally, you can see certain heroes are higher in the pecking order vs. the playtime results. This is to be expected, as while we might love playing a certain hero, it's not always a viable pick.
#
# ##### Brigitte
# I can't really avoid mentioning `Brigitte`. I'm not sure we can easily interpret this result; players may dislike her for any number of reasons:
#
# * Being forced to pick `Brigitte` when she was clearly over-tuned (and this happened for *months*, even pre-GOATS)
# * Residual hate; `Brigitte` destroyed beloved heroes such as `Tracer`, `Genji` and `Winston` while barely breaking a sweat
# * Post-rework, players view `Brigitte` as weak
# * Elitism (limited mechanical requirement)
# * People just haven't warmed to `Brigitte`
#
# I have a few friends who stomped their way up the ranks as `Brigitte` -- you'd think that'd count for *something*?
#
# ##### When choosing favourites, meta's not better
# It sticks out to see just how low `Orisa`, `Reaper` & `Mei` place. All of these heroes have been viable for months and continued to gain in strength with the introduction of `Sigma`. Given that `Orisa` is currently meta but remains unpopular, perhaps the developers might consider a re-work of sorts?
#
# Despite a number of heroes being borderline throw picks (`Tracer` ever since `Brigitte` arrived on the scene) or a form of masochism (`Ana` vs. double shield, `Reinhardt` into `Mei`/`Reaper`/`Doomfist`), the romantics among us still love these heroes.
#
# ##### The problem children
# I'm a little surprised to see `Torbjorn`, `Symmetra`, `Bastion` & `Mei` so clearly at the bottom. [/r/cow](https://reddit.com/r/competitiveoverwatch) used to host endless discussions regarding one-tricks ruining the game. Either the reworks to `Torbjorn` & co largely fixed the problem, or our survey respondents are a different crowd and don't play as those heroes.
#
# It's also odd to see `Soldier: 76` ranking so badly, as he does much better under [Q: Which heroes are fun to play as?](#Q:-Which-heroes-are-fun-play-as?) My best guess is that he's an afterthought -- while it's fun to play the ukelele in quiet room, why even bother when there's a 130dB death metal band drowning you out?
# ## Hero responses
# ### Q: Which heroes are fun play as?
def hero_fun_factor(df, response_type):
return (df.query(f'Response_type == "{response_type}"')
.groupby('Hero', as_index=False)
.mean()
.sort_values(by='Value', ascending=False))
draw_fun_factor_graph(
hero_fun_factor(joined_df, COLUMN_PLAYING_AS),
"Which heroes are fun to play as?")
# #### Notes
# There's a few discrepancies vs responses to the above question ([Q: Which heroes are our favourites?](#Q:-Which-heroes-are-our-favourites?)), but we can make an educated guess as to why.
#
# ##### Oh, egads! My Mercy is ruined!
# While `Mercy`'s in the top half of respondents' favourites, she is considered distinctly un-fun to play as for the majority. My guess is that a subset of players *really* love playing `Mercy`, whereas other players are much less thrilled by the prospect. One person's flying angel is another's heal-bot. There's nothing wrong with that -- you don't have to please all of the people all of the time.
#
# ##### Orisa trots home in last place
# Unfortunately, `Orisa` continues to skulk around the bottom. In addition to respondents stating `Orisa` is un-fun to play, `Orisa` also lacks the niche appeal of `Mercy`. I guess what I'm trying to say is that `Orisa` is neither mainstream like [<NAME>](https://www.youtube.com/watch?v=dQw4w9WgXcQ) nor a golden nugget like [<NAME>](https://www.youtube.com/watch?v=iyRzzmjK5_Q&list=PLKl69KbB34_rPmngjlvqpXJa_Sar-aXZL). In this analogy, `Orisa` is <NAME>.
#
# ##### Other interesting placings
# `Baptiste` does considerably better here vs. his favourites result. This suggests `Baptiste` is fairly fun to play, but hasn't quite ousted our comfort picks just yet.
#
# As previously mentioned, `Soldier: 76` is still hanging in there, somehow.
#
# ### Q: Which heroes are fun play against?
draw_fun_factor_graph(
hero_fun_factor(joined_df, COLUMN_PLAYING_VS),
"Which heroes are fun to play against?")
# #### Notes
# ##### Most fun to play against
#
# To start, let's do a run-down of the Fun to Play Against Gang. They're nearly all 'old' heroes (released with the game plus `Ana`). There are a few exceptions which I'll come to, though.
#
# * On the `Tank` front, `Reinhardt`, `Winston`, `Zarya` and `D.va` score very well.
# * For `Supports`, `Ana`, `Zenyatta`, `Lucio` and `Mercy` all feature.
# * `Soldier: 76`, `McCree`, `Genji` and `Tracer` make up the most fun to play against `DPS` crew.
#
# For 'new' heroes, `Ashe` sneaks into the top 10, and `Wrecking Ball` sits dead-centre in the results.
#
# ##### Least fun to play against
# Next, we'll consider the un-fun crowd. Much of the 'new' cast of heroes are perceived to be un-fun (with exceptions).
#
# * `Doomfist`, `Orisa`, `Sombra` and `Brigitte` are all banished from the Kingdom of Fun.
# * As the least popular fellow by far, `Doomfist` is made to face the wall (probably in case he punches your face into it, then animation cancels your funeral).
# * `Sigma` makes an appearance -- the newest hero is the 7th least fun to play against.
# * `Wrecking Ball` & `Baptiste` are decently received, but can't quite crack the top half.
#
# Overwatch's older 'problem children' remain. I fully expected to see `Symmetra`, `Mei`, `Bastion` and `Widow` lurking around the bottom. I'll come back to some of these fellows in a further question, though.
#
# I had expected to see `Reaper` even lower on the fun chart due to his life-steal and other buffs making him an oppressive figure, but he comes in at 10th least fun to play against.
#
# ##### A Brigitte miracle
# Finally, while still placing in the bottom third, the post-rework `Brigitte` is approaching mid-table in the fun to play *against* stakes. For me, this is a borderline miraculous result. A sincere well-done to the devs is in order, as `Brigitte` is no longer the enfant terrible of Overwatch.
# ### Q: Which heroes are more fun to play as than to play against?
# Rationale: Optimising for the fun of one player at the expense of 6 others might tell us something about why certain heroes are strongly disliked by the playerbase.
#
# E.g. `Doomfist` might be satisfying to play, as it's rewarding to pull off combos that have high mechanical skill requirements. However, the players on the receiving end may feel helpless and demoralised.
#
# On the other hand, players may enjoy fragging out on `Zen`, but this is tempered by `Zen`'s sphere-like hitbox & squishy nature.
#
# **How to interpret this chart**
#
# For each hero, larger values on the x axis (increasing to the right) tell us a particular hero is more fun to play *as* than to play *against*. The reverse is also true; smaller values tell us the hero is more fun to play *against* than to play *as*.
def hero_fun_as_vs(df):
"""
Q: For each hero, how fun are they to play _as_ instead of _against_?
(e.g. a hero might be very fun to play as, but very un-fun to play against)
"""
playing_as_df = (df.query('Response_type == "Playing_as"')
.groupby('Hero')
.mean())
playing_vs_df = (df.query('Response_type == "Playing_vs"')
.groupby('Hero')
.mean())
fun_difference_df = (playing_as_df.join(
playing_vs_df,
lsuffix='_fun_as',
rsuffix='_fun_vs'))
fun_difference_df["Value_fun_difference"] = (
fun_difference_df["Value_fun_as"] -
fun_difference_df["Value_fun_vs"])
fun_difference_df.sort_values(by="Value_fun_difference", inplace=True)
# Initialize the matplotlib figure
f, ax = plt.subplots(figsize=CHART_BAR_FIGSIZE)
ax = sb.barplot(x="Value_fun_difference", y=fun_difference_df.index, data=fun_difference_df)
ax.set(xlabel="Playing as/vs fun difference", ylabel="")
plt.axvline(0.0, 0.0, 1.0).set_linestyle('dashed')
hero_fun_as_vs(joined_df)
# #### Notes
# ##### Oh Mercy
# `Mercy` is the hero that is considered far and away more fun to play *against* than to play *as*. This makes a lot of sense when we consider the results of a previous question (see: [Q:-Which-heroes-are-fun-play-as?](#Q:-Which-heroes-are-fun-play-as?)) -- `Mercy` mains aside, most of the playerbase doesn't enjoy playing `Mercy`. Coupled with the fact that `Mercy` poses no real threat to most of the cast, this makes a lot of sense (unless you're Neptuno popping `Valkyrie`).
#
# ##### CC & one-shots
# Two thirds of the cast are considered more fun to play *as* than to play *against*. It is absolutely no surprise to see many of the following heroes considered to be significantly more fun to play *as* than to play *against*:
# * `Doomfist` (CC & one-shots)
# * `Mei` (CC & cheerily waving)
# * `Sigma` (concerning to see him here so quickly, but he has CC & few weaknesses in the double barrier meta)
# * `Sombra` (CC)
# * `Widowmaker` (one-shots)
# * `Hanzo` (one-shots)
#
# ... and so on. There seems to be a distinct pattern here -- CC & one-shots are fun for the person dishing them out, but less so when the roles are reversed.
# ### Q: Where do heroes fall on the balance scale?
# Rationale: Gives us a feel for heroes that are considered weak/balanced/over-tuned (aka OP). This allows interesting questions to be asked later, too.
def hero_balance(df):
return (joined_df.query('Response_type == "Balance"')
.groupby('Hero', as_index=False)
.mean()
.sort_values(by='Value', ascending=False))
def draw_hero_balance_graph(df):
f, ax = plt.subplots(figsize=CHART_BAR_FIGSIZE)
ax = sb.barplot(x="Value", y="Hero", data=df)
set_graph_format_x_axis(ax)
# plot ideal balance line
balance_line_x = CHART_BAR_FIGSIZE[0] * 0.5
plt.axvline(balance_line_x, 0.0, 1.0).set_linestyle('dashed')
ax.set_title("Which heroes are perceived to be balanced?")
ax.set(xlabel="1 = Weak, 3 = Balanced, 5 = Strong", ylabel="")
draw_hero_balance_graph(hero_balance(joined_df))
# #### Notes
# ##### The over-powered
# All of the meta heroes are accounted for in the OP list (`Orisa`, `Sigma`, `Doomfist` etc.)
#
# ##### The weak
# Similarly, many of the underpowered choices are predictable: `Soldier: 76`, `Winston`, `D.va`, `Genji`, `Tracer`. We've seen many threads on forums about these heroes, so I shan't dwell on them.
#
# Many of the weak heroes lack burst damage or shields. Even the seemingly meta-proof `D.va` is finding things tough.
#
# ##### Brigitte (again)
# `Brigitte` is once again having a rough time. `Brigitte` is not only considered un-fun to play *as* & somewhat un-fun to play *against*, but she's also perceived as being incredibly weak. However, the best evidence I have directly contradicts this result: her [win rate on overbuff](https://www.overbuff.com/heroes/brigitte) is currently a thumping 57 or 58% -- the highest in the game at the moment! Time to hire an image consultant and buy some instagram follows, hen.
#
# ##### Sombra: balanced, yet can't win?
# `Sombra` is considered to be reasonably strong-ish, but her [winrate is in the toilet](https://www.overbuff.com/heroes/sombra) at 43% overall (the worst in the entire game, though it rises to 48% in `Grandmasters`). What's behind the disconnect? Perhaps players find `Sombra`'s mere presence oppressive, so they lose sight of the bigger picture?
#
# The new meta combined with global ultimate charge nerfs pushed `Sombra` out of contention due to her heavy reliance on `EMP`. Either way, it's worrying that `Sombra` can't buy a win at lower ranks and is still perceived as strong-ish. Perhaps `Sombra` is still strong in isolation and will ghost back into contention with a meta change.
#
# ##### Reinhardt mains should form a support group
# `Reinhardt` is amongst the weakest, but this flies in the face of a marginally positive [win rate](https://www.overbuff.com/heroes/reinhardt). I feel this may once again play into how people *feel* while playing `Reinhardt`. He may win more than he loses, but it's a common complaint that he gets pinballed, punched, hacked, frozen, booped etc. to the point where playing him is a masochistic exercise.
# ### Q: Which heroes are fun to play against relative to their perceived balance?
# Rationale: I'd expect to see something approaching an inversely proportional relationship between hero strength and fun playing *against* them (i.e. the more OP a hero is, the less fun it is to play *against* them).
#
# Are there any weak heroes that significantly buck this trend? If so, it suggests the hero design may have inherent problems.
#
# **How is the result calculated?**
#
# I'd suggest [reading the code](https://github.com/marksimpson82/overwatch_survey/blob/35ba66e3f9ef05bcd8804bebfec0c540366c9d3d/overwatch_survey/Analysis.ipynb#L779-L802) in github, but the basic idea is this:
#
# * Take the playing *against* un-fun/fun rating and transform it from the range `[1, 5]` => `[-1, 1]` (larger values are more fun)
# * Take the balance rating and transform it from the range `[1, 5]` => `[-1, 1]` (-1 is weak, 0 is balanced, 1 is strong)
# * Add these two values together.
#
# **Possible interpretations**
#
# I'd say the negative values are way more interesting.
#
# * If a hero is un-fun *and* OP, the result will be close to 0
# * The hero is probably just un-fun due to being OP rather than any inherent design problem. Tuning will resolve it.
# * If a hero is un-fun *and* weak, the result will be strongly negative
# * Not only is the hero weak, but nobody likes them. May suggest a design flaw.
# * If a hero is fun *and* balanced, the result will be positive
# * The hero design is in a great place.
# * If a hero is fun *and* OP, the result will be strongly positive
# * The hero design is in a great place, to the point where they kill us and we say "yes sir, thank you sir".
# +
def hero_fun_vs_balance(df):
"""
Q: which heroes are unpopular relative to their perceived balance level?
(i.e. people dislike playing against them even if they're not considered OP)
"""
playing_vs_df = (df.query('Response_type == "Playing_vs"')
.groupby('Hero')
.mean())
balance_df = (df.query('Response_type == "Balance"')
.groupby('Hero')
.mean())
# our fun and balance ranges go from [1, 5], so let's move them into the range
# 0 to 4, then transform them into the range [-1, +1]. E.g. a 1 => -1, 5 => 1
# this will make it easy to see if they 'cancel out' or not
playing_vs_df = (playing_vs_df - 3.0) / 2.0
balance_df = (balance_df - 3.0) / 2.0
fun_balance_df = (playing_vs_df.join(
balance_df,
lsuffix='_fun',
rsuffix='_balance')
.query('Value_balance.notnull()')) # filter out the invalid joined data; should do this first though
# this is a little arbitrary, but gives us a decent idea of 'fun/unfun' vs balance.
# we have a fun rating range [-1, 1] and a strength rating range [-1, 1].
#
# we'd expect to see something of an inverse relationship between fun and balance,
# so we're going to add them together and plot the whole lot. Adding the two values
# should roughly cancel out, and anything that significantly departs from 0 should
# give us a feel for outliers in a fun/power relationship.
#
# Examples:
#
# 1. hero that is viewed as well-balanced and also OK to play against:
# fun: 0.1, strength: -0.05 (i.e. these values are both close to 0)
# result: 0.1 + (-0.05) = 0.05
# interpretation: fun and balance are in check
#
# 2. hero that is un-fun and also also weak:
# fun: -0.5, strength: -0.3 (i.e. these values are both negative)
# result: -0.5 + (-0.3) = -0.8
# interpretation: the hero design has inherently un-fun elements even though weak.
#
# 3. hero that is fun and also slightly OP
# fun: 0.7, strength: 0.3
# result: 0.7 + 0.3 = 1.0
# interpretation: the hero design is inherently so much fun that we forgive the OPness
fun_balance_df["Value_relative_balance"] = (
fun_balance_df["Value_balance"] +
fun_balance_df["Value_fun"])
fun_balance_df.sort_values(by="Value_relative_balance", inplace=True)
return fun_balance_df
def draw_hero_fun_vs_balance_graph(df):
# Initialize the matplotlib figure
f, ax = plt.subplots(figsize=CHART_BAR_FIGSIZE)
ax = sb.barplot(x="Value_relative_balance", y=df.index, data=df)
ax.set(xlabel="Playing against hero: fun relative to balance", ylabel="")
# plot balance vs. fun baseline. Any hero to the <--- left of this line is considered
# un-fun relative to their balance level.
balance_line_x = 0.0
plt.axvline(balance_line_x, 0.0, 1.0).set_linestyle('dashed')
# -
draw_hero_fun_vs_balance_graph(
hero_fun_vs_balance(joined_df))
# #### Notes
# ##### Negatives
# Let's focus on the negative values. Firstly, poor `Brigitte` can't catch a break. I'm going to move on, as I've discussed her enough.
#
# The rest of the negatives are pretty much what I expected to see -- heroes that are considered to be well-balanced yet un-fun to play against:
#
# * `Bastion` is a niche pick, kills quickly, has a self-heal, often requires his team to build around him and takes a lot more coordination to stop than to use. I can't ever recall a teammate saying, "Yay, they're playing pirate ship!", either. Even if `Bastion` is perceived to be fairly well-balanced, respondents strongly dislike playing against him.
#
#
# * `Sombra`. Despite the perception of her being well-balanced, her balance/fun ratio is out of sorts. `Sombra` appears behind her enemies and silences their abilities. This is especially the case for `Tanks` like `Reinhardt`, `D.Va` and `Wrecking Ball` who need the support of their team to survive a successful hack. I suspect the loss of agency & feeling of helplessness makes `Sombra` unpopular.
#
#
# * `Mei` has the most devilish CC in the game and can also capitalise on a lack of coordination (her wall also happens to perfectly fit most choke points and is difficult to break due to its high health and multiple sections).
#
#
# * `Torbjorn` has an auto-aiming turret, high burst damage and a spammy area denial ultimate. I'm somewhat surprised to see he's neck and neck with `Mei` and `Symmetra`, though. Perhaps people's dislike for `Torbjorn` one-tricks has influenced the voting, or I'm missing something. I'm looking forward to reading your comments.
#
#
# * `Symmetra` has CC turrets that can surprise, slow & quickly kill squishies, a fight-winning (and visually noisy) ult that is almost indestructible *and* has received many buffs to her damage & beam. In the current double shield meta, her damage is strong.
#
#
# * `Junkrat` has a sneaky trap, long-range spam capabilities, two mines and an ult that is strong at lower levels of play.
#
# ##### The rest
# `Doomfist` is so universally disliked at this point that it's hard to read much into his negative rating. Blizzard has a formidable task ahead to balance him, as he tends to be useless and niche, or infuriatingly oppressive (especially when played into heroes that rely on their teammates for peel).
#
# The heroes that have the largest positive ratings are our usual motley crew of old heroes: `Ana`, `Lucio`, `Zenyatta` etc. These heroes are well-balanced and score very well in the fun to play against category.
#
# One interpretation of `Sigma`'s positive score here (despite being considered one of the most OP heroes in the game) is that his kit is not inherently un-fun to play against -- `Sigma` may just need his power pared back via small tweaks.
# ## Over to you
# Thanks again for reading. Have you got any questions or suggestions? I'll be reading the comments, and I intend to run this survey again sometime in future.
#
| overwatch_survey/Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature: "Abhishek's Features"
# Based on <NAME>'s features published on [GitHub](https://github.com/abhishekkrthakur/is_that_a_duplicate_quora_question) and [Kaggle forum](https://www.kaggle.com/c/quora-question-pairs/discussion/31284).
# ## Imports
# This utility package imports `numpy`, `pandas`, `matplotlib` and a helper `kg` module into the root namespace.
from pygoose import *
import os
import warnings
import gensim
from fuzzywuzzy import fuzz
from nltk import word_tokenize
from nltk.corpus import stopwords
from scipy.stats import skew, kurtosis
from scipy.spatial.distance import cosine, cityblock, jaccard, canberra, euclidean, minkowski, braycurtis
# ## Config
# Automatically discover the paths to various data folders and compose the project structure.
project = kg.Project.discover()
# Identifier for storing these features on disk and referring to them later.
feature_list_id = '3rdparty_abhishek'
# The path to the saved GoogleNews Word2Vec model.
google_news_model_path = os.path.join(project.aux_dir, 'word2vec', 'GoogleNews-vectors-negative300.bin.gz')
# ## Read data
# Original question datasets.
df_train = pd.read_csv(project.data_dir + 'train.csv').fillna('').drop(['id', 'qid1', 'qid2'], axis=1)
df_test = pd.read_csv(project.data_dir + 'test.csv').fillna('').drop(['test_id'], axis=1)
stop_words = stopwords.words('english')
# ## Build features
# Raw implementations from Abhishek below (excluding the features we already have in other notebooks):
def wmd(model, s1, s2):
s1 = str(s1).lower().split()
s2 = str(s2).lower().split()
stop_words = stopwords.words('english')
s1 = [w for w in s1 if w not in stop_words]
s2 = [w for w in s2 if w not in stop_words]
return model.wmdistance(s1, s2)
def norm_wmd(model, s1, s2):
s1 = str(s1).lower().split()
s2 = str(s2).lower().split()
stop_words = stopwords.words('english')
s1 = [w for w in s1 if w not in stop_words]
s2 = [w for w in s2 if w not in stop_words]
return model.wmdistance(s1, s2)
def sent2vec(model, s):
words = s.lower()
words = word_tokenize(words)
words = [w for w in words if not w in stop_words]
words = [w for w in words if w.isalpha()]
M = []
for w in words:
try:
M.append(model[w])
except:
continue
M = np.array(M)
v = M.sum(axis=0)
return v / np.sqrt((v ** 2).sum())
def extend_with_features(data):
data['common_words'] = data.apply(lambda x: len(set(str(x['question1']).lower().split()).intersection(set(str(x['question2']).lower().split()))), axis=1)
data['fuzz_qratio'] = data.apply(lambda x: fuzz.QRatio(str(x['question1']), str(x['question2'])), axis=1)
data['fuzz_WRatio'] = data.apply(lambda x: fuzz.WRatio(str(x['question1']), str(x['question2'])), axis=1)
model = gensim.models.KeyedVectors.load_word2vec_format(google_news_model_path, binary=True)
data['wmd'] = data.apply(lambda x: wmd(model, x['question1'], x['question2']), axis=1)
norm_model = gensim.models.KeyedVectors.load_word2vec_format(google_news_model_path, binary=True)
norm_model.init_sims(replace=True)
data['norm_wmd'] = data.apply(lambda x: norm_wmd(norm_model, x['question1'], x['question2']), axis=1)
question1_vectors = np.zeros((data.shape[0], 300))
for i, q in progressbar(enumerate(data.question1.values), total=len(data)):
question1_vectors[i, :] = sent2vec(model, q)
question2_vectors = np.zeros((data.shape[0], 300))
for i, q in progressbar(enumerate(data.question2.values), total=len(data)):
question2_vectors[i, :] = sent2vec(model, q)
question1_vectors = np.nan_to_num(question1_vectors)
question2_vectors = np.nan_to_num(question2_vectors)
data['cosine_distance'] = [cosine(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['cityblock_distance'] = [cityblock(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['jaccard_distance'] = [jaccard(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['canberra_distance'] = [canberra(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['euclidean_distance'] = [euclidean(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['minkowski_distance'] = [minkowski(x, y, 3) for (x, y) in zip(question1_vectors, question2_vectors)]
data['braycurtis_distance'] = [braycurtis(x, y) for (x, y) in zip(question1_vectors, question2_vectors)]
data['skew_q1vec'] = [skew(x) for x in question1_vectors]
data['skew_q2vec'] = [skew(x) for x in question2_vectors]
data['kur_q1vec'] = [kurtosis(x) for x in question1_vectors]
data['kur_q2vec'] = [kurtosis(x) for x in question2_vectors]
warnings.filterwarnings('ignore')
extend_with_features(df_train)
extend_with_features(df_test)
df_train.drop(['is_duplicate', 'question1', 'question2'], axis=1, inplace=True)
df_test.drop(['question1', 'question2'], axis=1, inplace=True)
# ### Build final features
X_train = np.array(df_train.values, dtype='float64')
X_test = np.array(df_test.values, dtype='float64')
print('X_train:', X_train.shape)
print('X_test: ', X_test.shape)
df_train.describe().T
# ## Save features
feature_names = [
'abh_common_words',
'abh_fuzz_qratio',
'abh_fuzz_WRatio',
'abh_wmd',
'abh_norm_wmd',
'abh_cosine_distance',
'abh_cityblock_distance',
'abh_jaccard_distance',
'abh_canberra_distance',
'abh_euclidean_distance',
'abh_minkowski_distance',
'abh_braycurtis_distance',
'abh_skew_q1vec',
'abh_skew_q2vec',
'abh_kur_q1vec',
'abh_kur_q2vec',
]
project.save_features(X_train, X_test, feature_names, feature_list_id)
| notebooks/feature-3rdparty-abhishek.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (12, 9)
plt.rcParams["font.size"] = 18
# + [markdown] deletable=true editable=true
# # Radioactivity
#
# Learning Objectives:
#
# - Explain how radioactivity was discovered
# - Explain the nuclear physical reason for radioactive decay
# - List the sources of natural and man made radiation
# - Read and understand a decay diagram
# - Calculate Q values for various types of decay
# - Describe the physics of various types of decay
# - State the radioactive decay law
# - Derive the radioactive decay law
# - Understand how incorporating sources impacts decay calcuations
# - Calculate decay with production for simple cases
# + [markdown] deletable=true editable=true
# ## Discovery of Radioactivity
# - Radioactivity was first discovered in 1896 by <NAME>, while working on phosphorescent materials.
# - These materials glow in the dark after exposure to light, and he thought that the glow produced in cathode ray tubes by X-rays might be connected with phosphorescence.
# - He wrapped a photographic plate in black paper and placed various phosphorescent minerals on it.
# - All results were negative until he used uranium salts.
#
# <a title="By UnknownUnknown author (http://ihm.nlm.nih.gov/images/B02617) [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Becquerel_in_the_lab.jpg"><img width="256" alt="Becquerel in the lab" src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b6/Becquerel_in_the_lab.jpg/256px-Becquerel_in_the_lab.jpg"></a>
# <center>Becquerel in the lab</center>
#
# <a title="By <NAME> [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Becquerel_plate.jpg"><img width="256" alt="Becquerel plate" src="https://upload.wikimedia.org/wikipedia/commons/1/1e/Becquerel_plate.jpg"></a>
# <center>Photographic plate made by <NAME> showing effects of exposure to radioactivity.</center>
#
# + [markdown] deletable=true editable=true
# **What is nuclear decay?**
# A spontaneous process where the protons and neutrons in a given nucleus are rearranged into a lower energy state.
# The transition may involve levels of the same nucleus (gamma emission, internal conversion) or levels of a different nucleus (alpha, beta decay).
#
# **Why do nuclei decay?** A nucleus decays when it is **unstable**. It undergoes decay in order to become more stable (lower energy state).
#
# **Where do the unstable nuclei come from?**
#
# + [markdown] deletable=true editable=true
# ## Sources of Radiation
#
# > A chart of the public's exposure to ionizing radiation (displayed below)
# > shows that people generally receive a total annual dose of about 620 millirem.
# > Of this total, natural sources of radiation account for about 50 percent,
# > while man-made sources account for the remaining 50 percent. -- US NRC
#
# 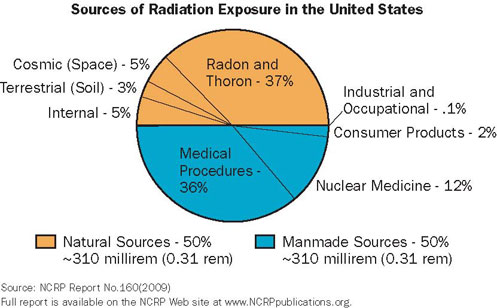
#
# ## Natural Sources
#
#
#
# ### Terrestrial
#
# Uranium, thorium, radium, etc. are all present naturally inside rocks and soil and are part of the four major decay chains. Others are 'primordial.' Primoidal radioactive isotopes include about 20 isotopes that are long lived and not part of any decay chain (K-40, Rubidium-87) Potassium has a quite radioactive isotope, $^{40}K$. (bananas)
#
# <a title="By Offnfopt [Public domain or CC0], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Evolution_of_Earth%27s_radiogenic_heat.svg"><img width="512" alt="Evolution of Earth's radiogenic heat" src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b9/Evolution_of_Earth%27s_radiogenic_heat.svg/512px-Evolution_of_Earth%27s_radiogenic_heat.svg.png"></a>
# <center>Evolution of the earth's radiogenic heat (in the mantle).</center>
#
# ### Internal
#
# Mostly $^{40}K$ and $^{14}C$ inside your body.
#
# ### Cosmic
# Commonly, cosmic radiation includes $^{14}C$, tritium ($^3H$), and others.
#
# <p><a href="https://commons.wikimedia.org/wiki/File:Shower_detection.png#/media/File:Shower_detection.png"><img src="https://upload.wikimedia.org/wikipedia/commons/2/27/Shower_detection.png" alt="Shower detection.png"></a><br><a href="http://creativecommons.org/licenses/by-sa/3.0/" title="Creative Commons Attribution-Share Alike 3.0">CC BY-SA 3.0</a>, <a href="https://commons.wikimedia.org/w/index.php?curid=44271">Link</a></p>
#
#
# 
#
# Air showers ensuing from very-high-energy cosmic rays can enter Earth’s atmosphere from multiple directions. Credit: <NAME>/NASA.
#
#
# ## Man-Made Radiation
#
# Of approximately 3200 known nuclides:
# - **266** are stable,
# - **65** long-lived radioactive isotopes are found in nature,
# - and the remaining **~2900** have been made by humans.
#
# One of the heaviest named elements is Livermorium (Z=116), its most stable isotope $^{293}Lv$ has half-life $t_{1/2} = 60 ms$.
#
# 
# + [markdown] deletable=true editable=true
# ### Types of Decay
# + deletable=true editable=true
# The below IFrame displays Page 121 of your textbook:
# <NAME>. (2016). Fundamentals of Nuclear Science and Engineering Third Edition,
# 3rd Edition. [Vitalsource]. Retrieved from https://bookshelf.vitalsource.com/#/books/9781498769303/
from IPython.display import IFrame
IFrame("https://bookshelf.vitalsource.com/books/9781498769303/pageid/121", width=1000, height=1000)
# + [markdown] deletable=true editable=true
# All elements smaller than Z=83 (Bismuth) have at least one stable isotope.
# Exceptions: Technetium (Z=43) and Promethium (Z=61).
#
# Once nucleus gets past a certain size, it is unstable
# The largest stable nucleus is Pb-208, all isotopes larger than this are unstable
#
# Nuclear conservation laws apply. The following are conserved:
# - charge
# - number of nucleons (proton + neutron)
# - total energy (mass + energy)
# - linear momentum (in inertial frame of reference)
# - angular momentum (spin)
# - alternatively: leptons (electrons + neutrinos)
# + deletable=true editable=true
# The below IFrame displays Page 100 of your textbook:
# <NAME>. (2016). Fundamentals of Nuclear Science and Engineering Third Edition,
# 3rd Edition. [Vitalsource]. Retrieved from https://bookshelf.vitalsource.com/#/books/9781498769303/
from IPython.display import IFrame
IFrame("https://bookshelf.vitalsource.com/books/9781498769303/pageid/122", width=1000, height=1000)
# + [markdown] deletable=true editable=true
# ## Energetics of Decay
#
# <a title="By User:Stannered (Traced from this PNG image.) [GFDL (http://www.gnu.org/copyleft/fdl.html), CC-BY-SA-3.0 (http://creativecommons.org/licenses/by-sa/3.0/) or CC BY 2.5
# (https://creativecommons.org/licenses/by/2.5
# )], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Alfa_beta_gamma_radiation.svg"><img width="256" alt="Alfa beta gamma radiation" src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/d6/Alfa_beta_gamma_radiation.svg/256px-Alfa_beta_gamma_radiation.svg.png"></a>
#
#
# ### Alpha ($\alpha$) Decay
#
# An $\alpha$ particle is emitted.
# The daughter is left with 2 fewer neutrons and 2 fewer protons than the parent.
#
# \begin{align}
# ^{A}_{Z}P \longrightarrow ^{A-4}_{Z-2}D^{2-} + ^4_2\alpha
# \end{align}
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)-\left[ M\left(^{A-4}_{Z-2}D\right) + M\left(^{4}_{2}He\right)\right]
# \end{align}
#
# <a title="By Inductiveload [Public domain], from Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Alpha_Decay.svg"><img width="128" alt="Alpha Decay" src="https://upload.wikimedia.org/wikipedia/commons/thumb/7/79/Alpha_Decay.svg/128px-Alpha_Decay.svg.png"></a>
#
# ### Gamma ($\gamma$) Decay
#
# An excited nucleus decays to its ground state by the emission of a gamma photon.
#
# \begin{align}
# ^{A}_{Z}P^* \longrightarrow ^{A}_{Z}P + \gamma
# \end{align}
#
#
# \begin{align}
# Q = E^*
# \end{align}
#
# <a title="By Inductiveload [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Gamma_Decay.svg"><img width="128" alt="Gamma Decay" src="https://upload.wikimedia.org/wikipedia/commons/thumb/c/c2/Gamma_Decay.svg/128px-Gamma_Decay.svg.png"></a>
#
# ### Negatron ($\beta -$) Decay
# A neutron changes into a proton in the nucleus (nuclear weak force). An electron ($\beta -$) and an antineutrino ($\bar{\nu}$) are emitted.
#
# \begin{align}
# ^{A}_{Z}P \longrightarrow ^{A}_{Z+1}D^{+} + ^0_{-1}e + \bar{\nu}
# \end{align}
#
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)- M\left(^{A}_{Z+1}D\right)
# \end{align}
#
# <a title="By Inductiveload [Public domain], from Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Beta-minus_Decay.svg"><img width="128" alt="Beta-minus Decay" src="https://upload.wikimedia.org/wikipedia/commons/thumb/a/aa/Beta-minus_Decay.svg/128px-Beta-minus_Decay.svg.png"></a>
#
# ### Positron ($\beta +$) Decay
#
# A proton changes into a neutron in the nucleus (nuclear weak force). A positron ($\beta +$) and a neutrino ($\nu$) are emitted.
#
# \begin{align}
# ^{A}_{Z}P \longrightarrow ^{A}_{Z-1}D^{-} + ^0_{+1}e + \nu
# \end{align}
#
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)- \left[ M\left(^{A}_{Z-1}D\right) + 2m_e\right]
# \end{align}
#
# <a title="By Master-m1000 [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Beta-plus_Decay.svg"><img width="128" alt="Beta-plus Decay" src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/3f/Beta-plus_Decay.svg/128px-Beta-plus_Decay.svg.png"></a>
#
# ### Electron Capture
# 1. An orbital electron is absorbed by the nucleus,
# 2. converts a nuclear proton into a neutron and a neutrino ($\nu$),
# 3. and typically leaves the nucleus in an excited state.
#
# \begin{align}
# ^{A}_{Z}P + \left(^0_{-1}e\right) \longrightarrow ^{A}_{Z-1}D + \nu
# \end{align}
#
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)- M\left(^{A}_{Z-1}D\right)
# \end{align}
#
# <a title="By Pamputt [CC BY-SA 4.0
# (https://creativecommons.org/licenses/by-sa/4.0
# )], from Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Atomic_rearrangement_following_an_electron_capture.svg"><img width="256" alt="Atomic rearrangement following an electron capture" src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b1/Atomic_rearrangement_following_an_electron_capture.svg/256px-Atomic_rearrangement_following_an_electron_capture.svg.png"></a>
#
# ### Proton Emission
# A proton is ejected from the nucleus.
#
# \begin{align}
# ^{A}_{Z}P \longrightarrow ^{A-1}_{Z-1}D^{-} + ^1_1p
# \end{align}
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)-\left[ M\left(^{A-1}_{Z-1}D\right) + M\left(^{1}_{1}H\right)\right]
# \end{align}
#
# <a title="By No machine-readable author provided. Abschalom~commonswiki assumed (based on copyright claims). [Public domain], via Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Zerfallsschema.png"><img width="512" alt="Zerfallsschema" src="https://upload.wikimedia.org/wikipedia/commons/1/1f/Zerfallsschema.png"></a>
#
# The decay of a proton rich nucleus A populates excited states of a daughter nucleus B by β+ emission or electron capture (EC). Those excited states that lie below the separation energy for protons (Sp) decay by γ emission towards the groundstate of daughter B. For the higher excited states a competitive decay channel of proton emission to the granddaughter C exists, called β-delayed proton emission.
#
# ### Neutron Emission
# A neutron is ejected from the nucleus.
#
# \begin{align}
# ^{A}_{Z}P \longrightarrow ^{A-1}_{Z}P + ^1_0n
# \end{align}
#
# \begin{align}
# \frac{Q}{c^2} = M\left(^{A}_{Z}P\right)-\left[ M\left(^{A-1}_{Z}P\right) + m_n\right]
# \end{align}
#
# ### Internal Conversion
#
# The excitation energy of a nucleus is used to eject an orbital electron (typically a K-shell) electron.
#
# \begin{align}
# ^{A}_{Z}P^* \longrightarrow ^{A-1}_{Z}P^+ + ^0_{-1}e
# \end{align}
#
# \begin{align}
# Q = E* - BE^K_e
# \end{align}
#
# <a title="By HPaul [CC BY-SA 4.0
# (https://creativecommons.org/licenses/by-sa/4.0
# )], from Wikimedia Commons" href="https://commons.wikimedia.org/wiki/File:Decay_scheme_of_203Hg.jpg"><img width="512" alt="Decay scheme of 203Hg" src="https://upload.wikimedia.org/wikipedia/commons/thumb/d/df/Decay_scheme_of_203Hg.jpg/512px-Decay_scheme_of_203Hg.jpg"></a>
# + [markdown] deletable=true editable=true
# ### An Aside on The Nuclear Weak Force
#
# Beta ($\beta \pm$) decay is a consequence of the **weak force**, which is characterized by long decay times (thousands of years). Nucleons are composed of up quarks and down quarks, and **the weak force allows a quark to change type** by the exchange of a W boson and the creation of an electron/antineutrino or positron/neutrino pair. For example, **a neutron, composed of two down quarks and an up quark, decays to a proton composed of a down quark and two up quarks.**
# + [markdown] deletable=true editable=true
#
#
# ## Radioactive Decay Law
#
#
# **The Law:** The probability that an unstable parent nucleus will decay spontaneously into one or more particles of lower mass/energy is independent of the past history of the nucleus and is the same for all radionuclides of the same type.
#
# ### Decay Constant
# Radioactive decay takes place stochastically in a single atom, and quite predictably in a large group of radioactive atoms of the same type.
#
# The probability that any one of the radionuclides in the sample decays in $\Delta t$ is $\frac{\Delta N}{N}$. This **decay probability** $\frac{\Delta N}{N}$ per unit time for a time interval $\Delta t$ should vary smoothly for large $\Delta t$.
#
# The statistically averaged **decay probability per unit time**, in the limit of infinitely small $\Delta t$, approaches a constant $\lambda$.
# Thus:
#
# \begin{align}
# \lambda &= \mbox{decay constant}\\
# &\equiv \lim_{\Delta t \to 0} \frac{\left(\Delta N/N\right)}{\Delta t}\\
# N(t) &= \mbox{expected number of nuclides in a sample at time t}\\
# \implies \mbox{ } -dN &= \mbox{decrease in number of radionuclides}\\
# &= \lambda N(t) dt\\
# \implies \mbox{ } \frac{dN(t)}{dt} &=-\lambda N(t)\\
# \end{align}
#
# The above is a differential equation. The solution to this differential equation give a definition of N as a function of t. We can now describe the change in radioactive nuclides over time as:
#
# \begin{align}
# \frac{dN}{dt} &= -\lambda N \\
# \Rightarrow N_i(t) &= N_i(0)e^{-\lambda t}\\
# \end{align}
#
# where
#
# \begin{align}
# N_i(t) &= \mbox{number of isotopes i adjusted for decay}\\
# N_i(0)&= \mbox{initial condition}\\
# \end{align}
# + deletable=true editable=true
import math
def n_decay(t, n_initial=100, lam=1):
"""This function describes the decay of an isotope"""
return n_initial*math.exp(-lam*t)
# This code plots the decay of an isotope
import numpy as np
y = np.arange(6.0)
x = np.arange(6.0)
for t in range(0,6):
x[t] = t
y[t] = n_decay(t)
# creates a figure and axes with matplotlib
fig, ax = plt.subplots()
scatter = plt.scatter(x, y, color='blue', s=y*20, alpha=0.4)
ax.plot(x, y, color='red')
# adds labels to the plot
ax.set_ylabel('N_i(t)')
ax.set_xlabel('Time')
ax.set_title('N_i')
# adds tooltips
import mpld3
labels = ['{0}% remaining'.format(i) for i in y]
tooltip = mpld3.plugins.PointLabelTooltip(scatter, labels=labels)
mpld3.plugins.connect(fig, tooltip)
mpld3.display()
# + [markdown] deletable=true editable=true
# ## Half Life
#
# All dynamic processes which decay (or grow) exponentially can be characterized by their half life (or doubling time). In the case of radioactive decay
#
# \begin{align}
# \tau_{1/2}&= \mbox{half-life}\\
# &=\frac{ln(2)}{\lambda} \\
# t &= \mbox{time elapsed [s]}\\
# \tau_{1/2} &= \mbox{half-life [s]} \\
# \end{align}
#
# + deletable=true editable=true
# This code converts decay constant to half life
def half_life(lam):
return math.log(2)/lam
lam = 1.0
# This code plots the decay of an isotope for various half lives
import numpy as np
y = np.arange(8.0)
x = np.arange(8.0)
lives = np.arange(half_life(lam), 9*half_life(lam), half_life(lam))
for i, t in enumerate(lives):
x[i] = float(t)
y[i] = n_decay(t)
# creates a figure and axes with matplotlib
fig, ax = plt.subplots()
scatter = plt.scatter(x, y, color='blue', s=y*20, alpha=0.4)
ax.plot(x, y, color='red')
# adds labels to the plot
ax.set_ylabel('N_i(t)')
ax.set_xlabel('Time')
ax.set_title('N_i')
# adds tooltips
import mpld3
labels = ['{0}% remaining'.format(i) for i in y]
tooltip = mpld3.plugins.PointLabelTooltip(scatter, labels=labels)
mpld3.plugins.connect(fig, tooltip)
mpld3.display()
# + [markdown] deletable=true editable=true
# After n half lives, only $\frac{1}{2^n}$ of the original sample remain.
# \begin{align}
# N(n\tau_{1/2})= \frac{1}{2^n}N_0
# \end{align}
#
# In a sample, $N_0$ has has been reduced to a fraction of $N_0$, $\epsilon$. How many half lives have passed?
#
# \begin{align}
# n = \frac{-ln\epsilon}{ln2}
# \end{align}
#
# Finally, the radioactive decay law can be expressed using the half-life:
#
# \begin{align}
# N(t) = N_0\left(\frac{1}{2}\right)^{t/\tau_{1/2}}
# \end{align}
# + [markdown] deletable=true editable=true
# ## Decay by competing processes
#
# \begin{align}
# \frac{dN(t)}{dt} &= -\lambda_1N(t) - \lambda_2N(t) - \cdots \lambda_nN(t)\\
# &= -\sum_{i=1}^n \lambda_iN(t)\\
# &\equiv-\lambda N(t)
# \end{align}
#
# A nuclide will decay by the $i^{th}$ mode with probability $f_i$.
#
# \begin{align}
# f_i &= \frac{\mbox{decay rate by ith mode}}{\mbox{decay rate by all modes}}\\
# &=\frac{\lambda_i}{\lambda}
# \end{align}
# + [markdown] deletable=true editable=true
# ## Decay with Production
#
# In reactors, isotopes decay into one another and still others are born from fission. Thus, if there is production, we can rewrite the standard decay differential equation as:
#
#
# \begin{align}
# \frac{dN(t)}{dt} &= -\mbox{rate of decay} + \mbox{rate of production}\\
# \implies N(t) &= N_0 e^{-\lambda t} + \int_0^t dt'Q(t')e^{-\lambda (t-t')}\\
# \end{align}
#
# If the production rate is constant, this simplifies:
#
#
# \begin{align}
# N(t) &= N_0 e^{-\lambda t} + \frac{Q_0}{\lambda}\left[1-e^{-\lambda t}\right]\\
# \end{align}
#
# ### Fuel Depletion
#
# Decays, fissions, and absorptions compete throughout the life of the reactor.
#
# 
#
# #### Aside: Reaction Rates
#
# In a reactor, this Q can be characterized via reaction rates.
#
# - The microscopic cross section $\sigma_{i,j}$ is just the likelihood of the event per unit area.
# - The macroscopic cross section $\Sigma_{i,j}$is just the likelihood of the event per unit area of a certain density of target isotopes.
# - The reaction rate is the macroscopic cross section times the flux of incident neutrons.
#
# \begin{align}
# R_{i,j}(\vec{r}) &= N_j(\vec{r})\int dE \phi(\vec{r},E)\sigma_{i,j}(E)\\
# R_{i,j}(\vec{r}) &= \mbox{reactions of type i involving isotope j } [reactions/cm^2s]\\
# N_j(\vec{r}) &= \mbox{number of nuclei participating in the reactions }\\
# E &= \mbox{energy}\\
# \phi(\vec{r},E)&= \mbox{flux of neutrons with energy E at position i}\\
# \sigma_{i,j}(E)&= \mbox{cross section}\\
# \end{align}
#
#
# We said this can be written more simply as $R_x = \sigma_x I N$, where I is intensity of the neutron flux. In the notation of the above equation, we can describe the production of an isotope by neutron absorption by another isotope as :
#
# \begin{align}
# \mbox{isotope i production via neutron absorption in m} = f_{im}\sigma_{am}N_m \phi
# \end{align}
#
#
# ### Total composition evolution
#
# \begin{align}
# \frac{dN_i}{dt} &= \sum_{m=1}^{M}l_{im}\lambda_mN_m + \phi\sum_{m=1}^{M}f_{im}\sigma_mN_m - (\lambda_i + \phi\sigma_i + r_i - c_i)N_i + F_i\Big|_{i\in [1,M]}\\
# \end{align}
# \begin{align}
# N_i &= \mbox{atom density of nuclide i}\\
# M &= \mbox{number of nuclides}\\
# l_{im} &= \mbox{fraction of decays of nuclide m that result in formation of nuclide i}\\
# \lambda_i &= \mbox{radioactive decay constant of nuclide i}\\
# \phi &= \mbox{neutron flux, averaged over position and energy}\\
# f_{im} &= \mbox{fraction of neutron absorption by nuclide m leading to the formation of nuclide i}\\
# \sigma_m &= \mbox{average neutron absorption cross section of nuclide m}\\
# r_i &= \mbox{continuous removal rate of nuclide i from the system}\\
# c_i &= \mbox{continuous feed rate of nuclide i}\\
# F_i &= \mbox{production rate of nuclide i directly from fission}\\
# \end{align}
#
#
# 
#
# 
#
# + [markdown] deletable=true editable=true
# ### Example: $^{135}Xe$
#
# **Discussion: What is interesting about Xenon?**
#
#
# $^{135}Xe$ is produced directly by fission and from the decay of iodine.
#
# \begin{align}
# \frac{dN_{xe}}{dt} &= \sum_{m=1}^{M}l_{Xem}\lambda_mN_m + \phi\sum_{m=1}^{M}f_{Xem}\sigma_mN_m - (\lambda_{Xe} + \phi\sigma_{Xe} + r_{Xe} - c_{Xe})N_{Xe} + F_{Xe}\\
# &= -\lambda_{Xe}N_{Xe} - \sigma_{aXe}\phi N_{Xe} + \lambda_IN_I + F_{Xe}\\
# &= -\lambda_{Xe}N_{Xe} - \sigma_{aXe}\phi N_{Xe} + \lambda_IN_I + \gamma_{Xe}\Sigma_f\phi\\
# \gamma_{Xe} &= 0.003\\
# \gamma_{I} &= 0.061\\
# \end{align}
# + [markdown] deletable=true editable=true
# ### Example: $^{239}Pu$
#
#
# \begin{align}
# \frac{dN_{Pu}}{dt} &= \sum_{m=1}^{M}l_{Pum}\lambda_mN_m + \phi\sum_{m=1}^{M}f_{Pum}\sigma_mN_m - (\lambda_{Pu} + \phi\sigma_{Pu} + r_{Pu} - c_{Pu})N_{Pu} + F_{Pu}\\
# \end{align}
#
#
# Let's formulate this equation together.
#
#
# $$\mathrm{^{238}_{\ 92}U \ + \ ^{1}_{0}n \ \longrightarrow \ ^{239}_{\ 92}U \ \xrightarrow [23.5\ min]{\beta^-} \ ^{239}_{\ 93}Np \ \xrightarrow [2.3565\ d]{\beta^-} \ ^{239}_{\ 94}Pu}$$
#
#
# - Decay of what nuclides result in the formation of $^{239}Pu$?
# - Does $^{239}Pu$ decay?
# - Is there a nuclide that becomes $^{239}Pu$ after it absorbs a neutron?
# - Does $^{239}Pu$ ever absorb neutrons?
# - Is $^{239}Pu$ ever produced directly from fission?
#
# + [markdown] deletable=true editable=true
# ## Burnable Poisons
#
# - Gadolinia ($Gd_2O_3$) or erbia ($Er_2O_3$) common
# - Natural Erbium consists of Er166, Er167, Er168 and Er170 primarily. Er167 has large thermal cross section.
# - Gd is an early life burnable poison, typically gone by 10‐20 GWd
# - Boron also used widely.
# - Can be mixed with the fuel or a coating on the pellet.
#
#
# + [markdown] deletable=true editable=true
# \begin{align}
# \frac{dN^P(t)}{dt} &= -g(t)\sigma_{aP}N^P(t)\phi\\
# g(t) &= \frac{\mbox{average flux inside BP}}{\mbox{average flux in core}}\\
# \sigma_{aP} &=\mbox{neutron absorption cross section of the BP}\\
# N^P(t) &= \mbox{number of atoms of the BP at time t}
# \end{align}
#
# 
#
# 
# + deletable=true editable=true
| radioactivity/00-radioactivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# **<center>Team Leabra: Project Milestone (March 15th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Hardware | 90% | March 14 | 100% |
# | Code | 0% | April 20 | 25% |
# | Demo | 0% | April 25 | 5% |
# | Presentation | 0% | April 28 | 5% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# This is the first milestone in the project. Our main deliverable goal as a team was to get the hardware for the R/C car setup so that we would be able to determine if that was a viable option or if we had to go with a software based model (simulation).
# We set up the car using a pre-built plastic frame to which four independent motors were attached. The motors were connected to an Arduino Uno for motor control and then powered by four AA batteries. We also added a Raspberry PI for to control the entire system. A camera module was attached to the Raspberry PI 4 for image detection. We are now done with the hardware phase of the vehicle and will tackle the software setup in the next phase.
#
# The only hardware involved part that is left to create is the track that the car will run and train on. We are planning to use white paper with colored tape on the edges.
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# We had initially anticipated to get the software set up on the R/C car but we were not able to fully accomplish this. One problem we faced was lack of equipment to solder wires on to the Arduino. This pushed back the timeline for setting up the hardware on the vehicle which was essential for us to move on to the software set up.
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# The main deliverable goals to be met for the next milestone are:
#
# Software Setup on R/C car and lane/track creation -> <NAME> & <NAME>. <br>
# OpenCV code for image detection and finding additional data sets for sign recogniton -> <NAME> & <NAME>. <br>
# Initial code to control vehicle motors -> <NAME> <br>
# Neural Network planning and setup -> Entire Team
#
# This is what we are planning so far; group member duties may change depending on individual capabilites
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (March 21th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Hardware | 100% | March 21 | 100% |
# | Code | 5% | April 20 | 25% |
# | Demo | 0% | April 25 | 0% |
# | Presentation | 0% | April 28 | 0% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# Now that the hardware setup of the car is complete and the arduino issues are resolved we have shifted our focus to installing the necessary software on the car. A operating system is currently installed and the necessary libraries will be installed soon.
#
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
#
# The code part of the project has stagnated a bit as the OS of the Raspberry Pi has only just been installed. However, we have collected some quality datasets. From these datasets we are currently creating Haarcascades which, when finished, can be used for object detection of traffic signs.
#
# also, we have decided the push the demo and presentation aspects of the project forward as we want to prioritize development of the car.
#
#
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# The next step for the people responsible for the hardware would be to set up a serial connection between the raspberry, the arduino, and the motor controllers. -> <NAME> & <NAME>.
#
# For the people that currently do most of the coding, finishing and testing .the haarcascades that are used in object detection would be a good achievement for the next milestone -> <NAME> & <NAME>
#
# Now that we have an operating system running on the RSP start writing the initial code to control vehicle motors. -> <NAME>
#
# Neural Network planning and setup -> Entire Team
#
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (March 28th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Hardware | 100% | March 21 | 100% |
# | Code | 20% | April 20 | 45% |
# | Demo | 0% | April 25 | 0% |
# | Presentation | 0% | April 28 | 0% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# We have made a lot of process on the coding part of the project. The development of the line-detection software has been going well, only thing left for that is to get the cars current position (distance to right&left line) on the track.
#
# Install OS on raspberry pi and physically linked arduino to raspberry.
#
# A test haarcascade has been created for the stopsign using machine learning (adaboost). Perfomance of the cascade isn't optimal yet and will need improvement if we want to deploy it onto the car.
#
# There have been talks on how to implement the neural network and we decided will start working writing the actual code when the car is up and running.
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# Install the necessary libraries on the raspberry. We had some logistic issues because the person who was supposed to work on this didn't have access to the RC car
#
# For the same reason the motor control code hasn't been written just yet. Another thing that should be done before the next milestone is have the arduino and the raspberry talking to eachother
#
# We need to improve performance of the object detection, current haarcascades do not perform well enough.
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# Write motor controller code -> Kwabena
#
# Continue developing linedetection code -> Stijn
#
# Set connection between raspberry and arduino -> Patrick
#
# Set up libraries for raspberry -> Patrick
#
# create haarscascades for stopsign and speel limits -> Stijn
#
#
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (April 4th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Code | 25% | April 20 | 50% |
# | Demo | 0% | April 25 | 0% |
# | Presentation | 0% | April 28 | 0% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# We had a massive setback as we have lost the car because one of our group members has dropped the class. We are now trying to get back on schedule by instead of using the car, we use a simulation that is hosted in unity. The simulation is developed by someone else and is will therefore be easily implementable. This is great as it will allow us to gain back some of the lost time which we used on building and setting up the RC car.
#
# Because we don't have to drive the car in real life, we were already able to capture driving data from the simulation which can be used for training.
#
# Line detection has also been improved, it's still unknown whether this will be useful in the simulation but we are planning on trying.
#
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# All goals that included working on the RC have been dropped and therefore have not been completed. This meant all goals apart from improving the linedetection have not been completed.
#
#
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# Start working on the CNN -> Patrick
#
# create data augmention code -> Stijn
#
# clean training data -> Stijn
#
# write code to connect to simulation -> Patrick
#
#
#
#
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (April 11th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Code | 60% | April 20 | 90% |
# | Demo | 0% | April 25 | 50% |
# | Presentation | 0% | April 28 | 50% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# All the training data has been cleaned and processed for better perfomance while model fitting. Data has also been augmented so that we have more training data to work with. Also started working on the Neural Network and did a couple of test runs with the training data, perfomance of the actual model can't be tested yet as we don't have the code that connects the model with the simulation working yet.
#
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# the code to connect to the simulation isn't finished yet. Should be finished in the next couple of days so that we can start testing the models in the simualtion
#
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# Continue working on the CNN -> Patrick & Stijn
#
# write code to connect to simulation -> Patrick
#
#
#
#
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (April 18th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Code | 60% | April 20 | 100% |
# | Demo | 0% | April 25 | 50% |
# | Presentation | 0% | April 28 | 50% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# We have continued to refine the network to reach a higher training accuracy.
#
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# There are still errors in the code needed to connect to the simulation so the model still can't be tested.
#
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# Continue working on the CNN -> Patrick, Stijn, James
#
# Finish code to connect to simulation -> Patrick
#
# Collect more training data -> James
#
#
#
# <center>_______________________________________________________________________________________________</center>
# **<center>Team Leabra: Project Milestone (April 25th, 2022)</center>**
# | Deliverable | Percent Complete | Estimated Completion Date | Percent Complete By Next Milestone |
# | :- | :- | :- | :- |
# | Code | 90% | May 2 | 100% |
# | Demo | 0% | May 2 | 100% |
# | Presentation | 0% | May 2 | 100% |
# **1. What deliverables goals established in the last milestone report were accomplished to the anticipated percentage?**
#
# We have continued to work on the CNN and connecting to the simulation. All of the data needed for training has been collected
# **2. What deliverables goals established in the last milestone report were not accomplished to the anticipated percentage?**
#
# The code for connecting to the simulation still has bugs that make it impossible to test the model on, the CNN needs further optimization.
# **3. What are the main deliverable goals to meet before the next milestone report, and who is working on them?**
#
# Optimize CNN -> Stijn, Patrick, James, Le'Shawn
#
# Finish code to connect to simulation -> Patrick
#
# Write Demo -> Stijn, Patrick, James, Le'Shawn
#
# Make Presentation -> Stijn, Patrick, James, Le'Shawn
#
#
#
# <center>_______________________________________________________________________________________________</center>
| Project_Milestones.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 009
#
# 二次元平面上に N 個の点が存在し、点 i は座標 (X_i, Y_i) にあります。
#
# あなたは相異なる 3 つの整数 A, B, C (1 ≦ A, B, C ≦ N) を選び、角 ABC (点 A → 点 B → 点 C の折れ線で構成される角) の大きさを最大にしたいです。
#
# そのときの角の大きさを 0 ~ 180 度の範囲で表される度数法で出力してください。
#
# 【制約】
#
# ・3 ≦ N ≦ 2000
#
# ・0 ≦ X[i], Y[i] ≦ 10^9
#
# ・(X[i], Y[i]) ≠ (X[j], Y[j])
#
# ・入力はすべて整数
#
# ・絶対誤差または相対誤差が 10^{-8} 以内であれば正解とみなされる
#
#
# ### 入力形式
# N
#
# X[1] Y[1]
#
# X[2] Y[2]
#
# X[3] Y[3]
#
# :
#
# X[N] Y[N]
#
# +
# 入力例 1
3
0 0
0 10
10 10
# 出力例 1
90.000000000000
# +
# 入力例 2
5
8 6
9 1
2 0
1 0
0 1
# 出力例 2
171.869897645844
# +
# 入力例 3
10
0 0
1 7
2 6
2 8
3 5
5 5
6 7
7 1
7 9
8 8
# 出力例 3
180.000000000000
# +
# 入力例 4
40
298750376 229032640
602876667 944779015
909539868 533609371
231368330 445484152
408704870 850216874
349286798 30417810
807260002 554049450
40706045 380488344
749325840 801881841
459457853 66691229
5235900 8100458
46697277 997429858
827651689 790051948
981897272 271364774
536232393 997361572
449659237 602191750
294800444 346669663
792837293 277667068
997282249 468293808
444906878 702693341
894286137 845317003
27053625 926547765
739689211 447395911
902031510 326127348
582956343 842918193
235655766 844300842
438389323 406413067
862896425 464876303
68833418 76340212
911399808 745744264
551223563 854507876
196296968 52144186
431165823 275217640
424495332 847375861
337078801 83054466
648322745 694789156
301518763 319851750
432518459 772897937
630628124 581390864
313132255 350770227
# 出力例 4
179.983434068423
| 009_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, transpile, Aer, IBMQ
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
from qiskit.providers.aer import QasmSimulator
# Loading your IBM Quantum account(s)
provider = IBMQ.load_account()
# -
# ### Solving Linear Equations
# +
# reference - # https://quantumcomputing.stackexchange.com/questions/20862/hhl-the-result-is-correct-for-one-matrix-but-wrong-for-another-one
# -
# https://www.youtube.com/watch?v=KtIPAPyaPOg
# +
from qiskit import ClassicalRegister, QuantumRegister, QuantumCircuit, Aer, execute, IBMQ
from qiskit.compiler import transpile, assemble
from qiskit.quantum_info import Statevector
from qiskit.algorithms.linear_solvers.hhl import HHL
import numpy as np
import math as m
# decimal to binary
def dec2bin(decimal, bits):
b = bin(decimal)
b = b[2:len(b)]
while len(b) < bits:
b = '0' + b
return b
processor = Aer.backends(name='qasm_simulator')[0]
processor
# -
# 
# +
# 1st linear system; A*x = B
matrix_A = np.array([[1,0,0,0],[0,2,0,0],[0,0,3,0],[0,0,0,4]])
vector_b = np.array([1,4,0.5,3])
# solution = [1, 2, 0.1667, 0.75]
matrix_A
# -
# number of variables (assuming n=2^k)
n = len(matrix_A[2,:])
n
# number of necessary qubits
n_qubits = m.ceil(m.log(n,2))
n_qubits
# norm of right-side
bNorm = np.linalg.norm(vector_b)
bNorm
hhlSolver = HHL(quantum_instance = processor)
hhlSolver
# preparing circuit in general form
hhlCircuit = hhlSolver.construct_circuit(matrix_A, vector_b)
hhlCircuit.draw()
# +
# transpile -- adapt circuit to real quantum processor
circuitToRun = transpile(hhlCircuit, basis_gates=['id', 'rz', 'sx', 'x', 'cx'])
# add measurement
circuitToRun.measure_all()
circuitToRun.draw()
# +
from qiskit.visualization import plot_histogram
# execute the circuit
shots = 8192
counts = execute(circuitToRun, processor, shots=shots).result().get_counts(circuitToRun)
plot_histogram(counts)
# -
# width - num of qubits and classical bit summed
usedQubits = circuitToRun.width()/2
usedQubits
# +
# working qubits
zeros = dec2bin(0, usedQubits - n_qubits - 1)
# ancilla qubit has to be |1>, working qubits |0>, results are in last "n_qubits" qubits
# e.g. if n_qubits and total qubits is 6, then 3 are working qubits
# results are stored in |1 000 00>, |1 000 01>, |1 000 10>, |1 000 11>
for i in range(0,n):
indx = '1' + zeros + dec2bin(i, n_qubits)
# bNorm*m.sqrt(M[indx]/shots) - we need probability amplitudes => sqrt
# and adjust them by norm of vector b to get "unnormalized" solution x
print('x_' + str(i) + ': ' + str(round(bNorm*m.sqrt(counts[indx]/shots),4)))
# -
| Notebooks/Algorithms/.ipynb_checkpoints/QuantumLinearSolver_HHL_1.2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color=maroon> Male Family Planning Analysis </font> #
# #### <font color=indigo><NAME>, Applied Data Science, Spring 2019</font> ####
import pandas as pd
import numpy as np
import seaborn as sns
import statsmodels.formula.api as smf
from sklearn.linear_model import Lasso
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
import matplotlib.pyplot as plt
df = pd.read_stata('Raw_Data_Clipped.dta')
df.head()
| sophiasga/Atik_Proj2_Resub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="eC-Vn7LDUeCk"
# #Clone the repo
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1922, "status": "ok", "timestamp": 1605646197925, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="q8syRlXZUf_o" outputId="933f45dc-76c2-4869-fdbe-ab492db3ca7d"
# !git clone https://github.com/Moda007/MethodicalSplit.git
# + executionInfo={"elapsed": 7468, "status": "ok", "timestamp": 1605646203477, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="m6RA55ybUkCH"
from MethodicalSplit import Functions as Fn
from MethodicalSplit.ExpModel import ExpModel
# + [markdown] id="8Snsq7kuUw8m"
# #Define experiment details
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 20867, "status": "ok", "timestamp": 1605646221353, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="tIetyqWxUz27" outputId="f9775fe7-05f7-4565-d1b5-63d52ea363e5"
DataSet, ModelName, group_idx, exp_idx, Avg, Sub, Rare, stratify, train = Fn.expDetails(Fn.all_exp)
# + [markdown] id="8R5MmBJ4OWEZ"
# #Colab
# + [markdown] id="q5a_NWs0tl69"
# ##Ignore Warnings
# + executionInfo={"elapsed": 1728, "status": "ok", "timestamp": 1605646227246, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="D8HRA1sEseKY"
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="ZCPkvD-LtpQ1"
# ##Mount Drive
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 23605, "status": "ok", "timestamp": 1605646249519, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="nc9LE3cAStjB" outputId="fb8c694d-1605-464e-bb45-5feb4f4c3fa0"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="CsZngWFkkSPv"
# #Importing Packages and Libraries
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 57796, "status": "ok", "timestamp": 1605646306435, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="8jRrVAmpcCqJ" outputId="428e9b92-1af9-421b-b480-9de1a0e98333"
# !pip install hdbscan
# + executionInfo={"elapsed": 55205, "status": "ok", "timestamp": 1605646306437, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="nZHqCupPl7E-"
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
import umap
import hdbscan
import sklearn.cluster as cluster
from sklearn.metrics import adjusted_rand_score, adjusted_mutual_info_score
# + [markdown] id="G4ejsZkTlsaC"
# #Dataset
# + [markdown] id="tY7yX9KHmYpx"
# ##Importing Dataset
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1976, "status": "ok", "timestamp": 1605646308434, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="PkdXdi9HmL-H" outputId="08ba8cb6-e452-46c9-b4c3-2984583b09ee"
from keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1971, "status": "ok", "timestamp": 1605646308435, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="5b32ImdjvZ_9" outputId="03bfc177-5624-48b6-cdc9-f8b00937a1f6"
print("x_train shape:", x_train.shape)
print("x_test shape:", x_test.shape)
print("y_train shape:", y_train.shape)
print("y_test shape:", y_test.shape)
# + [markdown] id="lsOf-c-emcvm"
# ##Concatenate train and test dataset
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1967, "status": "ok", "timestamp": 1605646308436, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="xrUmC6YgjyBw" outputId="ca429cae-cd17-4eaa-8efe-6394d2fdc687"
x_data = np.concatenate([x_train, x_test], axis=0)
y_data = np.concatenate([y_train, y_test], axis=0)
print("x_data shape:", x_data.shape)
print("y_data shape:", y_data.shape)
# + [markdown] id="4FXQx0pzA3J-"
# ##Split test data (holdout)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2947, "status": "ok", "timestamp": 1605646309421, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="bdmANJFNA_c8" outputId="c2d8982f-0687-4d1c-cd24-9a28c7087604"
from sklearn.model_selection import train_test_split
x_data, x_test, y_data, y_test = train_test_split(x_data, y_data, train_size= 0.9, test_size= 0.1)
print("x_data shape:", x_data.shape)
print("x_test shape:", x_test.shape)
print("y_data shape:", y_data.shape)
print("y_test shape:", y_test.shape)
# + [markdown] id="wuftUZ_zmmbq"
# ##Flatten dataset
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2943, "status": "ok", "timestamp": 1605646309422, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="tik1wkfKmPt-" outputId="ea1fe914-098e-427c-af1c-12b0a795f5dd"
flatten_x = x_data.reshape((x_data.shape[0], -1))
print("flatten_x shape:", flatten_x.shape)
x_test = x_test.reshape((x_test.shape[0], -1))
print("x_test shape:", x_test.shape)
# + [markdown] id="yaSdl7Fkm6wR"
# #Model (Loop)
# + [markdown] id="Yzhq8e73HcxL"
# ##>>> Experiment
# + executionInfo={"elapsed": 2941, "status": "ok", "timestamp": 1605646309423, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="xAOfmVErptoo"
looped = True
if looped:
looped = 'looped_'
else:
looped = ''
ds_path = '/content/drive/My Drive/Thesis Notebooks/' + DataSet + '/'
exp_path = ds_path + ModelName + '/Group_' + group_idx + '/Exp' + exp_idx + '/'
# + executionInfo={"elapsed": 2940, "status": "ok", "timestamp": 1605646309424, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="XwuyC_I0sWdN"
def plotResults(DataSet, ModelName, group_idx, exp_idx, history, loop_no=''):
# plot the loss and accuracy
global exp_path
loop_no = str(loop_no)
plot_path = exp_path + 'plot'
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.title('Training and validation accuracy')
plt.plot(epochs, acc, 'red', label='Training acc')
plt.plot(epochs, val_acc, 'blue', label='Validation acc')
plt.legend()
plt.savefig(f'{plot_path}/{loop_no}TrainValidAcc.jpg')
plt.figure()
plt.title('Training and validation loss')
plt.plot(epochs, loss, 'red', label='Training loss')
plt.plot(epochs, val_loss, 'blue', label='Validation loss')
plt.legend()
plt.savefig(f'{plot_path}/{loop_no}TrainValidAcc&Loss.jpg')
plt.show()
# + [markdown] id="-kjmGObDRuBf"
# #Looped Experiment
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 4086818, "status": "ok", "timestamp": 1605651455498, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="EGLKgCtwRy2j" outputId="f6343052-ff58-4aea-e51b-a1f82258e46a"
#Define All Results array
All_results = []
for idx in range(5):
global exp_path
plot_path = exp_path + 'plot'
print('*****************************')
print(f'Experiment {str(idx)} starts')
print('*****************************')
#2D Embedding - UMAP
standard_embedding = umap.UMAP(random_state=42, n_neighbors=30, min_dist=0.0, n_components=2).fit_transform(flatten_x)
#Plotting with Original Labels
##2D - Plotting
plt.scatter(standard_embedding[:, 0], standard_embedding[:, 1], c=y_data, s=0.1, cmap='Spectral')
plt.savefig(f'{plot_path}/{idx}2DOriginalScatter.jpg')
plt. clf()
sns.distplot(standard_embedding, kde=True, rug=True, bins=15)
plt.savefig(f'{plot_path}/{idx}2DOriginalhistogram.jpg')
plt. clf()
#Clustering - HDBSCAN
hdbscan_labels = hdbscan.HDBSCAN(min_samples=10, min_cluster_size=500).fit_predict(standard_embedding)
(adjusted_rand_score(y_data, hdbscan_labels), adjusted_mutual_info_score(y_data, hdbscan_labels))
no_of_clusters = Fn.clustersDet(hdbscan_labels)
##Plotting with Cluster Labels
### 2D - Plotting
plt.scatter(standard_embedding[:, 0], standard_embedding[:, 1], c=hdbscan_labels, s=0.1, cmap='Spectral')
plt.savefig(f'{plot_path}/{idx}2DClustersScatter.jpg')
plt. clf()
#1D Embedding per Cluster on the original Data
#Using UMAP =>>> random_state=42, n_neighbors=30, min_dist=0, n_components=1
clusters_list, labels_list, original_images_list = Fn.embedding1D(no_of_clusters, flatten_x, hdbscan_labels, y_data, False)
#Plotting Clusters Scatter Diagram
##All Clusters
#Plotting scatter diagram for all clusters together
#define a list of colors for clusters
color_maps=['tab10', 'hsv', 'gist_stern', 'Accent', 'Dark2', 'Spectral', 'rainbow', 'brg', 'Pastel1', 'coolwarm']
for u in range(no_of_clusters):
plt.scatter(standard_embedding[hdbscan_labels==u][:, 0], standard_embedding[hdbscan_labels==u][:, 1],\
c=hdbscan_labels[hdbscan_labels==u], s=0.1, cmap=color_maps[u])
plt.savefig(f'{plot_path}/{idx}1DClustersScatter.jpg')
plt. clf()
for u in range(no_of_clusters):
plt.scatter(standard_embedding[hdbscan_labels==u][:, 0], standard_embedding[hdbscan_labels==u][:, 1],\
c=hdbscan_labels[hdbscan_labels==u], s=0.1, cmap=color_maps[u])
plt.savefig(f'{plot_path}/{idx}1DCluster{str(u)}Scatter.jpg')
plt. clf()
#Data Splitting (per cluster) using Quantile
Region_1, Region_2, Region_3,\
Region_1_labels, Region_2_labels, Region_3_labels,\
Region_1_original, Region_2_original, Region_3_original = Fn.splitData(no_of_clusters, Avg, Sub, Rare, clusters_list,\
labels_list, original_images_list)
for u in range(no_of_clusters):
Fn.showSplit(Region_1, Region_2, Region_3, u)
plt.savefig(f'{plot_path}/{idx}Cluster{str(u)}SplitHistogram.jpg')
plt. clf()
Fn.showSamples(Region_1_original[u], Region_2_original[u], Region_3_original[u], Region_1_labels[u], Region_2_labels[u], Region_3_labels[u])
plt.savefig(f'{plot_path}/{idx}Samples{str(u)}.jpg')
plt. clf()
##Data Preperation
###Splitting Data into 70% Train, 30% Validate
x_train_X, y_train_X, x_valid_X, y_valid_X, x_test_X, y_test_X = Fn.prepareData(no_of_clusters, Region_1_original,\
Region_1_labels, Region_2_original,\
Region_2_labels, Region_3_original,\
Region_3_labels, x_test, y_test, train)
print('x_train_X shape:', x_train_X.shape)
print('x_valid_X shape:', x_valid_X.shape)
print('x_test_X shape:', x_test_X.shape)
##Train
thisModel = ExpModel(ModelName, DataSet, x_train_X, y_train_X, x_valid_X, y_valid_X, x_test_X, y_test_X)
model, history, results = thisModel.trainModel()
##Store experiment results
All_results.append(results)
##Model Saving
filename = str(idx) + ModelName + '_' + DataSet + '_' + group_idx + '_'+ exp_idx
model.save(exp_path + filename + '.h5')
# plot the loss and accuracy
plotResults(DataSet, ModelName, group_idx, exp_idx, history, idx)
print('*****************************')
print(f'Experiment {str(idx)} ends')
print('*****************************')
tf.keras.backend.clear_session()
# + [markdown] id="ZdT3trA6YpBV"
# ##Check experiments results
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 832, "status": "ok", "timestamp": 1605651456363, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="xRA4uAr8YtXz" outputId="e87efbb9-9733-4288-a992-0e7f09d64ef9"
if len(All_results)==5:
print('Results Are Complete')
else:
print('Results record has issue!!!')
# + [markdown] id="Kuy8kh87RejV"
# ##Exporting Results to Excel
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 4391, "status": "ok", "timestamp": 1605651459931, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="E55xG3NLR5Ir" outputId="e344d5ad-fa46-42f2-8dd6-b78b15215d63"
# !pip install xlsxwriter
# + [markdown] id="2uFo4BzAWzfc"
# ###Create Excel with columns header
# + executionInfo={"elapsed": 5946, "status": "ok", "timestamp": 1605651461491, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="1HVzxZXQSwF6"
import pandas as pd
from openpyxl import load_workbook
excel_name = ModelName + '.xlsx'
sheet_name = group_idx + '_' + exp_idx
filepath = ds_path + excel_name
column_titles = [['Idx', 'Train Acc', 'Valid Acc', 'Train Loss', 'Valid Loss',\
'Test Acc', 'Test Precision', 'Test Recall', 'Test F-score', 'Hamming Loss']]
df = pd.DataFrame(column_titles)
# + executionInfo={"elapsed": 6500, "status": "ok", "timestamp": 1605651462050, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="2Em1e9YkAK9F"
from pathlib import Path
if not Path(filepath).exists():
# Create writer object with an engine xlsxwriter
writer = pd.ExcelWriter(filepath, engine='xlsxwriter')
# Write data to an excel
df.to_excel(writer, sheet_name=sheet_name, index=False, header=None)
writer.save()
else:
book = load_workbook(filepath)
with pd.ExcelWriter(filepath, engine='openpyxl') as writer:
writer.book = book
writer.sheets = dict((ws.title, ws) for ws in book.worksheets)
##dataframe to append.
df.to_excel(writer, sheet_name=sheet_name, index=False, header=None)
writer.save()
# + [markdown] id="M0OBSspQxCJV"
# ###Parse this experiment results
# + executionInfo={"elapsed": 8024, "status": "ok", "timestamp": 1605651463580, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04085889877448088796"}, "user_tz": -60} id="CuWcIk1qxI0h"
for idx, result in enumerate(All_results):
this_result = [idx] + result['train'] + result['test']
this_result = np.array(this_result).reshape(1,-1)
this_result = pd.DataFrame(this_result)
writer = pd.ExcelWriter(filepath, engine='openpyxl')
# try to open an existing workbook
writer.book = load_workbook(filepath)
# copy existing sheets
writer.sheets = dict((ws.title, ws) for ws in writer.book.worksheets)
# read existing file
reader = pd.read_excel(filepath)
# write out the new sheet
this_result.to_excel(writer, sheet_name=sheet_name, index=False, header=False, startrow=idx+1)
writer.close()
# + id="W6OjfqG2S-U8"
| notebooks/hdbscan/MNIST/VGG/Group_1/Exp4/looped_VGG_MNIST_Exp_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] run_control={"frozen": false, "read_only": false}
# Please go through the "building_strategies" notebook first before you go through this notebook
#
# -
# ## Some Prebuilt Reporting ##
#
# Lets first build the strategy described in that notebook, add it to a portfolio and run the portfolio
# + run_control={"frozen": false, "read_only": false}
from types import SimpleNamespace
import pandas as pd
import numpy as np
import pyqstrat as pq
from pyqstrat.notebooks.support.build_example_strategy import build_example_strategy
strategy = build_example_strategy(SimpleNamespace(lookback_period = 10, num_std = 2))
strategy.run()
# -
# Many objects have functions that return pandas dataframes for ease of use. Any function that returns a dataframe starts with df_ so its easy to tell which dataframes an object returns.
#
# Probably the most common function you will use is df_data on the strategy object. This returns the market data, indicators, signal values and P&L at each market data bar. The last column, i is the integer index of that bar, and can be used to query data in other dataframes or objects for that strategy.
# + run_control={"frozen": false, "read_only": false}
strategy.df_data().iloc[550:560]
# -
# You can also look at just the PNL or just the marketdata by themselves.
# + run_control={"frozen": false, "read_only": false}
strategy.df_pnl().head()
# -
# We can look at orders and trades that were created during this run
# + run_control={"frozen": false, "read_only": false}
strategy.df_orders().head()
# + run_control={"frozen": false, "read_only": false}
strategy.df_trades().head()
# -
# You can also look at the returns at the portfolio level (i.e. summing up several strategies)
# + run_control={"frozen": false, "read_only": false}
strategy.df_returns().head()
# -
# We can also get data as native Python objects as opposed to pandas dataframes.
# + run_control={"frozen": false, "read_only": false}
strategy.trades(start_date = '2019-01-15', end_date = '2019-01-20')
# -
# ## Adding your Own Metrics ##
#
# Each strategy may have metrics that you want to measure that are specific to that strategy. To add these, you can use the Evaluator object which can make things easier.
#
# To evaluate a strategy we use the evaluate returns function.
# + run_control={"frozen": false, "read_only": false}
strategy.evaluate_returns(plot = False);
# -
# What if we want to add some more metrics to this. For example, lets say we want to add a metric that looks at how many long trades we had versus short trades. We can do this using an Evaluator object.
# + run_control={"frozen": false, "read_only": false}
def compute_num_long_trades(trades):
return len([trade for trade in trades if trade.order.reason_code == pq.ReasonCode.ENTER_LONG])
def compute_num_short_trades(trades):
return len([trade for trade in trades if trade.order.reason_code == pq.ReasonCode.ENTER_SHORT])
evaluator = pq.Evaluator(initial_metrics = {'trades' : strategy.trades()})
evaluator.add_metric('num_long_trades', compute_num_long_trades, dependencies = ['trades'])
evaluator.add_metric('num_short_trades', compute_num_short_trades, dependencies = ['trades'])
evaluator.compute()
print('Long Trades: {} Short Trades: {}'.format(evaluator.metric('num_long_trades'), evaluator.metric('num_short_trades')))
# -
# The Evaluator takes care of dependency management so that if you want to compute a metric that relies on other metrics, it will compute the metrics in the right order.
#
# Lets compute Maximum Adverse Execution for each trade. MAE tells you the maximum loss each trade in its lifetime. It's useful for figuring out where to put trailing stops. For example, if most of your profitable trades had a maximum loss during their life up to 5% but many losing trades had losses of 50% and 60%, it might make sense to place a trailing stop around 6% or 7% so you don't get stopped out of your profitable trades but get out of the losing ones quickly. See Jaekle and Tomasini, page 66 for details
#
# To do this we first have to first pair up entry and exit trades into a single "round-trip" trade.
# + run_control={"frozen": false, "read_only": false}
import collections
trades = strategy.trades()
entry_trades = [trade for trade in trades if trade.order.reason_code == pq.ReasonCode.ENTER_LONG]
exit_trades = [trade for trade in trades if trade.order.reason_code == pq.ReasonCode.EXIT_LONG]
def compute_mae(entry_trades, exit_trades, c, timestamps):
mae = np.empty(len(exit_trades)) * np.nan
round_trip_pnl = np.empty(len(exit_trades)) * np.nan
for i, exit in enumerate(exit_trades):
#if i == len(exit_trades): break
entry = entry_trades[i]
_round_trip_pnl = entry.qty * (exit.price - entry.price)
running_price = c[(timestamps >= entry.timestamp) & (timestamps <= exit.timestamp)]
running_pnl = entry.qty * (running_price - entry.price)
_mae = -1 * min(_round_trip_pnl, np.min(running_pnl))
_mae = _mae / entry.price # Get mae in % terms
_mae = max(0, _mae) # If we have no drawdown for this trade, set it to 0
mae[i] = _mae
round_trip_pnl[i] = _round_trip_pnl / entry.price # Also store round trip pnl for this trade since we will have to plot it
return mae, round_trip_pnl
def get_trades(trades, entry):
rc = [pq.ReasonCode.ENTER_LONG, pq.ReasonCode.ENTER_SHORT] if entry else [pq.ReasonCode.EXIT_LONG, pq.ReasonCode.EXIT_SHORT]
return [trade for trade in trades if trade.order.reason_code in rc]
contract_group = strategy.contract_groups[0]
evaluator = pq.Evaluator(initial_metrics = {'trades' : strategy.trades(),
'c' : strategy.indicator_values[contract_group].c,
'timestamps' : strategy.timestamps})
evaluator.add_metric('entry_trades', lambda trades : get_trades(trades, True), dependencies=['trades'])
evaluator.add_metric('exit_trades', lambda trades : get_trades(trades, False), dependencies=['trades'])
evaluator.add_metric('mae', compute_mae, dependencies=['entry_trades', 'exit_trades', 'c', 'timestamps'])
evaluator.compute()
# -
# We could have easily done the same computation without using the Evaluator. The main advantage of using the Evaluator is that you can reuse other metrics you are dependent on without having to recompute them each time, i.e it provides a local cache of metrics.
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## Plotting ##
#
# Now that we have maximum adverse execution, lets plot it.
#
# pyqstrat plots are built on top of matplotlib. Each pyqstrat Plot object contains a list of Subplot objects which in turn each contain a list of the data you want to plot in that subplot.
#
# In this case, we want to plot round trip PNL for each trade versus its max drawdown (MAE). The goal is to find out where to set a stop loss so we keep most of the winning trades while limiting our losses from the biggest losing trades. Since we want to look at the effect of setting a stop loss on both winning and losing trades, we flip the sign of the round trip pnl on the losing trades and mark them in a different color to show the negative P&L.
# + run_control={"frozen": false, "read_only": false}
mae = evaluator.metric('mae')[0]
round_trip_pnl = evaluator.metric('mae')[1]
# Separate out positive trades from negative trades
round_trip_profit = round_trip_pnl[round_trip_pnl >= 0]
mae_profit = mae[round_trip_pnl >= 0]
round_trip_loss = round_trip_pnl[round_trip_pnl <= 0]
mae_loss = mae[round_trip_pnl <= 0]
subplot = pq.Subplot([
pq.XYData('Profitable Trade', mae_profit, round_trip_profit, display_attributes = pq.ScatterPlotAttributes(marker = '^', marker_color = 'green')),
pq.XYData('Losing Trade', mae_loss, -1 * round_trip_loss, display_attributes = pq.ScatterPlotAttributes(marker = 'v', marker_color = 'red'))],
horizontal_lines = [pq.HorizontalLine(y = 0, color = 'black')],
vertical_lines = [pq.VerticalLine(x = 0, color = 'black')],
xlabel = 'Drawdown in %', ylabel = 'Profit / Loss in %')
plot = pq.Plot([subplot])
plot.draw();
# -
# It looks like a good place to put a stop loss so we keep most of the winning trades but don't take big losses might be around 4%. The next step would be to re-test the system with this stop loss inserted. Lets plot the same data with a line showing the stop loss.
# + run_control={"frozen": false, "read_only": false}
subplot = pq.Subplot([
pq.XYData('Profitable Trade', mae_profit, round_trip_profit, display_attributes = pq.ScatterPlotAttributes(marker = '^', marker_color = 'green')),
pq.XYData('Losing Trade', mae_loss, -1 * round_trip_loss, display_attributes = pq.ScatterPlotAttributes(marker = 'v', marker_color = 'red'))],
horizontal_lines = [pq.HorizontalLine(y = 0, color = 'black')],
vertical_lines = [pq.VerticalLine(x = 0, color = 'black'), pq.VerticalLine(name = 'Stop Loss', x = 4, color = 'blue')],
xlabel = 'Drawdown in %', ylabel = 'Profit / Loss in %')
plot = pq.Plot([subplot])
plot.draw();
# -
# There are several different kinds of subplots, including 3d subplots that you can include in the same plot. See the code of the plot function in the Strategy class or the Evaluator class for eamples.
| pyqstrat/notebooks/reporting_and_plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zsMX4_zEhr3Z" colab_type="text"
# # Инициализация
# + id="QYr79QJohReK" colab_type="code" outputId="59a681ee-6fbb-473b-8b74-fcae1ba80934" executionInfo={"status": "ok", "timestamp": 1557569714393, "user_tz": -480, "elapsed": 4246, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
#@markdown - **Монтирование GoogleDrive**
from google.colab import drive
drive.mount('GoogleDrive')
# + id="lASDIhZrjE1Z" colab_type="code" colab={}
# #@markdown - **Размонтирование**
# # !fusermount -u GoogleDrive
# + [markdown] id="3n1Xgh64jG_H" colab_type="text"
# # Область кодов
# + id="3fEu2-mJjI9v" colab_type="code" outputId="f5c60531-ed90-4574-eb84-8e661c403dea" executionInfo={"status": "ok", "timestamp": 1557582181709, "user_tz": -480, "elapsed": 1677, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
#@title Распознавание лиц { display-mode: "both" }
# facial recognition
# В этой программе реализовано распознавание лиц в наборе лица FaceWarehouse
# при использовании TensorFlow для построения четырехслойных сверточных нейронных сетей
#@markdown - **Информации о наборе лица FaceWarehouse** [FaceWarehouse](http://kunzhou.net/zjugaps/facewarehouse/)
# Набор обучающих, тестовых изображений и этикетки хранятся в face_150.mat
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import scipy.io as scio
import os,sys
from keras.utils import to_categorical
tf.logging.set_verbosity(tf.logging.ERROR)
# + id="yQkQEcdHlgmw" colab_type="code" colab={}
#@markdown - **Привязка данных**
class Bunch(dict):
def __init__(self,*args,**kwds):
super(Bunch,self).__init__(*args,**kwds)
self.__dict__ = self
#@markdown - **Определение функции print_progress**
def print_progress(progress, epoch_num, loss, acc):
"""
This function draw an active progress bar.
:param progress: Where we are:
type: float
value: [0,1]
:param epoch_num: number of epochs for training
:param loss: The loss for the specific batch in training phase.
:return: Progressing bar
"""
barLength = 30
assert type(progress) is float, "id is not a float: %r" % id
assert 0 <= progress <= 1, "variable should be between zero and one!"
# Символ статуса
status = ""
# Возврат "\ r \ n" после окончания печати
if progress >= 1:
progress = 1
status = "\r\n"
# Запись статуса
indicator = int(round(barLength*progress))
list = [str(epoch_num), "#"*indicator , "-"*(barLength-indicator), progress*100, loss, acc, status]
text = "\rEpoch {0[0]} {0[1]} {0[2]} {0[3]:.2f}% completed, loss={0[4]:.4f}, acc={0[5]:.2f}%{0[6]}".format(list)
sys.stdout.write(text)
sys.stdout.flush()
# + id="JLjNX3fvmZoM" colab_type="code" colab={}
#@markdown - **Определения нескольких функций**
def weight_variable(shape, name):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial, name=name)
def bias_variable(shape, name):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial, name=name)
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2x2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
# + id="rs6G3hOAm539" colab_type="code" colab={}
#@markdown - **Настройка параметров**
num_epochs = 400 #@param {type: "integer"}
batch_size = 128 #@param {type: "integer"}
decay_steps = 200 #@param {type: "number"}
keep_p = 0.8 #@param {type: "number"}
online_test = True #@param {type: "boolean"}
data_path = 'GoogleDrive/My Drive/MATLAB/face recognition/faces_database/faces_150.mat'
# data_path = 'GoogleDrive/My Drive/MATLAB/face recognition/faces_database/faces_150_equalhis.mat'
events_path = 'GoogleDrive/My Drive/Colab Notebooks/Tensorboard'
checkpoints_path = 'GoogleDrive/My Drive/Colab Notebooks/Checkpoints'
# + id="lPfUyrLTou6z" colab_type="code" outputId="cf3b3c27-a7db-40c1-fcdf-41fac40a8d6f" executionInfo={"status": "ok", "timestamp": 1557582181714, "user_tz": -480, "elapsed": 1648, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 101}
#@markdown - **Считывание данных**
data = scio.loadmat(data_path)
# -------------------Выделение обучающих изображений---------------------------
train_image = data['train_faces']
train_labels = to_categorical(data['train_labels'].flatten(), num_classes=150)
train_data = Bunch(train_image=train_image, train_labels=train_labels)
print('\n', 'Train image set extraction completed... ...\n')
# ----------------Выделение тестовых изображений-------------------------------
test_image = data['test_faces']
test_labels = to_categorical(data['test_labels'].flatten(), num_classes=150)
test_data = Bunch(test_image=test_image, test_labels=test_labels)
print(' Test image set extraction completed... ...\n')
# + id="6Jo32skVsZVQ" colab_type="code" colab={}
#@markdown - **Создание graph**
model_name = 'model.ckpt'
graph = tf.Graph()
with graph.as_default():
global_step = tf.Variable(0, name='global_step', trainable=False)
# decay_steps = 100
decay_rate = 0.8
start_rate = 1e-3 #@param {type: "number"}
learning_rate = tf.train.exponential_decay(start_rate,
global_step=global_step,
decay_steps=decay_steps,
decay_rate=decay_rate,
staircase=True,
name='exponential_decay')
with tf.name_scope('Input'):
x = tf.placeholder("float", shape=[None, 90, 75])
y = tf.placeholder("float", shape=[None, 150])
keep_prob = tf.placeholder("float") # keep_prob указывает на вероятность того, что каждый нейрон активирован
with tf.name_scope('Input_images'):
x_image = tf.reshape(x, [-1, 90, 75, 1])
# --------------conv1-----------------------------------45*38*32
with tf.name_scope('Conv1'):
with tf.name_scope('weights_conv1'):
W_conv1 = weight_variable([3, 3, 1, 32], name='w_conv1')
with tf.name_scope('bias_covn1'):
b_conv1 = bias_variable([32], name='b_conv1')
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
with tf.name_scope('features_conv1'):
h_pool1 = max_pool_2x2(h_conv1)
# --------------conv2-----------------------------------23*19*64
with tf.name_scope('Conv2'):
with tf.name_scope('weights_conv2'):
W_conv2 = weight_variable([3, 3, 32, 64], name='w_conv2')
with tf.name_scope('bias_covn2'):
b_conv2 = bias_variable([64], name='b_conv2')
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
with tf.name_scope('features_conv2'):
h_pool2 = max_pool_2x2(h_conv2)
# --------------conv3-----------------------------------12*10*128
with tf.name_scope('Conv3'):
W_conv3 = weight_variable([3, 3, 64, 128], name='w_conv3')
b_conv3 = bias_variable([128], name='b_conv3')
h_conv3 = tf.nn.relu(conv2d(h_pool2, W_conv3) + b_conv3)
h_pool3 = max_pool_2x2(h_conv3)
# --------------conv4-----------------------------------6*5*256
with tf.name_scope('Conv4'):
W_conv4 = weight_variable([3, 3, 128, 256], name='w_conv4')
b_conv4 = bias_variable([256], name='b_conv4')
h_conv4 = tf.nn.relu(conv2d(h_pool3, W_conv4) + b_conv4)
h_pool4 = max_pool_2x2(h_conv4)
# --------------conv5-----------------------------------3*3*512
with tf.name_scope('Conv5'):
W_conv5 = weight_variable([3, 3, 256, 512], name='w_conv5')
b_conv5 = bias_variable([512], name='b_conv5')
h_conv5 = tf.nn.relu(conv2d(h_pool4, W_conv5) + b_conv5)
h_pool5 = max_pool_2x2(h_conv5)
# --------------fc--------------------------------------
with tf.name_scope('FC1'):
h_pool5_flat = tf.layers.flatten(h_pool5, name='pool5_flatten')
num_f = h_pool5_flat.get_shape().as_list()[-1]
W_fc1 = weight_variable([num_f, 1024], name='w_fc1')
b_fc1 = bias_variable([1024], name='b_fc1')
h_fc1 = tf.nn.relu(tf.matmul(h_pool5_flat, W_fc1) + b_fc1) # y=wx+b или y.T=(x.T)(w.T)+b.T, где у -- вектор столбца
with tf.name_scope('Dropout1'):
h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob)
with tf.name_scope('FC2'):
W_fc2 = weight_variable([1024, 1024], name='w_fc2')
b_fc2 = bias_variable([1024], name='b_fc2')
h_fc2 = tf.nn.relu(tf.matmul(h_fc1_drop, W_fc2) + b_fc2)
with tf.name_scope('Dropout2'):
h_fc2_drop = tf.nn.dropout(h_fc2, keep_prob)
with tf.name_scope('OutPut_layer'):
W_fc3 = weight_variable([1024, 150], name='w_fc2')
b_fc3 = bias_variable([150], name='b_fc2')
y_conv = tf.matmul(h_fc2_drop, W_fc3) + b_fc3
# ---------------------loss-----------------------------
with tf.name_scope('Loss'):
# y_conv = tf.nn.softmax(tf.matmul(h_fc1_drop, W_fc2) + b_fc2)
# cross_entropy = -tf.reduce_sum(y * tf.log(y_conv + 1e-10)) # предотвратить log0
# or like
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y,
logits=y_conv))
with tf.name_scope('Train'):
train_step = tf.train.AdamOptimizer(learning_rate).minimize(cross_entropy, global_step=global_step)
# or like
# with tf.name_scope('Train'):
# optimizer = tf.train.AdamOptimizer(learning_rate=2e-4)
# gradients_vars = optimizer.compute_gradients(cross_entropy)
# train_step = optimizer.apply_gradients(gradients_vars, global_step=global_step)
with tf.name_scope('Accuracy'):
correct_prediction = tf.equal(tf.argmax(y_conv,1), tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
# + id="OdFx8xm4swy-" colab_type="code" outputId="5ffb1cb9-eb47-4a80-fae6-623ad1387501" executionInfo={"status": "ok", "timestamp": 1557582471040, "user_tz": -480, "elapsed": 290953, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 3427}
#@markdown - **Обучение СНС**
max_acc = 101.0 # модели с выше этой точностью будут сохранены
min_cross = 0.1
with tf.Session(graph=graph) as sess:
saver = tf.train.Saver(var_list=tf.global_variables(), max_to_keep=3)
sess.run(tf.global_variables_initializer())
print('Training ========== (。・`ω´・) ========')
for epoch_num in range(num_epochs):
train_s = np.c_[train_data.train_image.reshape((1500,-1)), train_data.train_labels]
np.random.shuffle(train_s)
max_size = train_s.shape[0] // batch_size
for num in range(max_size):
batch = [train_s[num*batch_size:(num+1)*batch_size, :90*75].reshape((-1,90,75)),
train_s[num*batch_size:(num+1)*batch_size, -150:]]
_, acc, loss = sess.run([train_step, accuracy, cross_entropy],
feed_dict={x: batch[0], y: batch[1], keep_prob: keep_p})
acc *= 100
num_iter = max_size * 10
progress = ((epoch_num * max_size + num) % num_iter + 1) / num_iter
num_ep = epoch_num + 1
print_progress(progress, num_ep, loss, acc)
if online_test and (epoch_num + 1) % 10 ==0 :
print(' '*12, 'Online-Testing ========== (。・`ω´・) ========')
imgs_t, labels_t = test_data.test_image.reshape((-1, 90, 75)), test_data.test_labels
test_acc, test_loss = sess.run([accuracy, cross_entropy], feed_dict={x: imgs_t, y: labels_t,
keep_prob: 1.0})
test_acc *= 100
print(' '*10, 'Loss on testing data is %.4f, accuracy is %.2f%%.' %(test_loss, test_acc))
print('\nKeep on training ========== (。・`ω´・) ========')
# 3 модели с высокой точностью сохраняются
if (loss <= min_cross) & (acc >= max_acc) & (epoch_num > 100):
min_cross = loss
max_acc = acc
saver.save(sess, os.path.join(checkpoints_path, model_name), global_step=epoch_num)
test_im, test_lab = train_data.train_image[0].reshape((-1, 90, 75)), train_data.train_labels[0].reshape((-1, 150))
feature_map1 = sess.run(h_pool1, feed_dict={x: test_im, y: test_lab, keep_prob: 1.0})
# feature_map2 = sess.run(h_pool2, feed_dict={x: test_im, y: test_lab, keep_prob: 1.0})
sess.close()
print('\n', 'Training completed.')
# + id="5BxOTSrtrHJI" colab_type="code" colab={}
#@markdown - **Восстановление сохраненной модели**
# with tf.Session() as sess:
# model_path = 'Tensorboard/f_map.ckpt-241'
# saver.restore(sess, model_path)
# acc, loss = sess.run([accuracy, cross_entropy], feed_dict={x: test_data.test_image,
# y: test_data.test_labels, keep_prob: 1.0})
# acc_p = acc*100
# print('Accuracy is %.2f' %(acc_p), '%.')
# sess.close()
# + id="E8axAOGBkR83" colab_type="code" outputId="32e90a37-dc52-4d2f-80ca-22e6624a8534" executionInfo={"status": "ok", "timestamp": 1557582473149, "user_tz": -480, "elapsed": 293049, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 819}
#@markdown - **Представление feature map первого сверточного слоя**
f_map = feature_map1.reshape((45, 38, 32))
num_map = range(f_map.shape[-1])
fig = plt.figure(1,figsize=(24, 14))
G = gridspec.GridSpec(4, 8)
G.hspace,G.wspace = .05,.05
try:
for i in range(4):
for j in range(8):
plt.subplot(G[i, j])
num = i * 5 + j
plt.imshow(f_map[:, :, num], 'gray')
plt.xticks([]), plt.yticks([])
except IndexError:
plt.xticks([]), plt.yticks([])
plt.show()
| notebooks(colab)/Neural_network_models/Supervised_learning_models/Facial_recognition_RU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from merging import merge_datasets
# +
# Define an array to hold dataset sizes
datasets_sizes = np.array([4,5])
# Full matrix of pairwise comparisons including cross-dataset and within dataset comparisons
pwc = np.array([[0, 0, 0, 3, 0, 3, 0, 0, 0],
[6, 0, 0, 0, 0, 6, 0, 0, 5],
[0, 0, 0, 0, 0, 0, 0, 0, 0],
[3, 6, 0, 0, 6, 0, 0, 9, 0],
[0, 0, 0, 4, 0, 2, 3, 2, 3],
[7, 4, 0, 0, 4, 0, 3, 3, 3],
[0, 0, 0, 0, 3, 3, 0, 3, 4],
[0, 0, 0, 1, 4, 3, 3, 0, 2],
[0, 5, 0, 0, 3, 3, 2, 4, 0]])
# Matrix with rating scores, each row is a condition, it's column is the rating measurement trial
mos = np.array([[3, 4, 5, 5, np.nan, np.nan, np.nan, np.nan, np.nan],
[5, 7, 7, 7, np.nan, np.nan, np.nan, np.nan, np.nan],
[9, 7, 8, 9, np.nan, np.nan, np.nan, np.nan, np.nan],
[9, 9, 9, 8, np.nan, np.nan, np.nan, np.nan, np.nan],
[np.nan, np.nan, np.nan, np.nan, 3, 2, 3, 3, 2],
[np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan],
[np.nan, np.nan, np.nan, np.nan, 4, 3, 3, 3, 2],
[np.nan, np.nan, np.nan, np.nan, 3, 3, 3, 4, 2],
[np.nan, np.nan, np.nan, np.nan, 3, 2, 4, 4, 3]])
# Produce the estimates
Q, a, b, nu = merge_datasets(pwc,mos,datasets_sizes)
print("Q = ", Q)
print("a = ", a)
print("b = ", b)
print("nu = ", nu)
# -
| python/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def detectCapitalUse(self, word: str) -> bool:
uppercase = set([chr(x) for x in range(ord('A'), ord('Z')+1)])
lowercase = set([chr(x) for x in range(ord('a'), ord('z')+1)])
s_w = set(word)
if uppercase & s_w == s_w:
return True
if lowercase & s_w == s_w:
return True
s_w = set(word[1:])
if word[0] in uppercase and s_w & lowercase == s_w:
return True
return False
s = Solution()
s.detectCapitalUse('Usb')
| algorithms/520-detect-capital.ipynb |