code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="i4vFDhvn_Bdn"
# ##### Copyright 2018 The TensorFlow Constrained Optimization Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# > http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# + [markdown] id="RpUmH2nk_Bdo"
# ## PR-AUC Maximization
# In this colab, we'll show how to use the TF Constrained Optimization (TFCO) library to train a model to maximize the *Area Under the Precision-Recall Curve (PR-AUC)*. We'll show how to train the model both with (i) plain TensorFlow (in eager mode), and (ii) with a custom tf.Estimator.
#
# We start by importing the relevant modules.
# + id="FoYVEXPA_Bdp"
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import shutil
from sklearn import metrics
from sklearn import model_selection
import tensorflow.compat.v2 as tf
# + id="Ea1KIDRgC4eq"
# Tensorflow constrained optimization library
# !pip install git+https://github.com/google-research/tensorflow_constrained_optimization
import tensorflow_constrained_optimization as tfco
# + [markdown] id="BLxZD5uH_Bds"
# ## Communities and Crimes
#
# We will use the *Communities and Crimes* dataset from the UCI Machine Learning repository for our illustration. This dataset contains various demographic and racial distribution details (aggregated from census and law enforcement data sources) about different communities in the US, along with the per capita crime rate in each commmunity.
#
#
# We begin by downloading and preprocessing the dataset.
# + id="MgoeyhS0_Bds"
# List of column names in the dataset.
column_names = ["state", "county", "community", "communityname", "fold", "population", "householdsize", "racepctblack", "racePctWhite", "racePctAsian", "racePctHisp", "agePct12t21", "agePct12t29", "agePct16t24", "agePct65up", "numbUrban", "pctUrban", "medIncome", "pctWWage", "pctWFarmSelf", "pctWInvInc", "pctWSocSec", "pctWPubAsst", "pctWRetire", "medFamInc", "perCapInc", "whitePerCap", "blackPerCap", "indianPerCap", "AsianPerCap", "OtherPerCap", "HispPerCap", "NumUnderPov", "PctPopUnderPov", "PctLess9thGrade", "PctNotHSGrad", "PctBSorMore", "PctUnemployed", "PctEmploy", "PctEmplManu", "PctEmplProfServ", "PctOccupManu", "PctOccupMgmtProf", "MalePctDivorce", "MalePctNevMarr", "FemalePctDiv", "TotalPctDiv", "PersPerFam", "PctFam2Par", "PctKids2Par", "PctYoungKids2Par", "PctTeen2Par", "PctWorkMomYoungKids", "PctWorkMom", "NumIlleg", "PctIlleg", "NumImmig", "PctImmigRecent", "PctImmigRec5", "PctImmigRec8", "PctImmigRec10", "PctRecentImmig", "PctRecImmig5", "PctRecImmig8", "PctRecImmig10", "PctSpeakEnglOnly", "PctNotSpeakEnglWell", "PctLargHouseFam", "PctLargHouseOccup", "PersPerOccupHous", "PersPerOwnOccHous", "PersPerRentOccHous", "PctPersOwnOccup", "PctPersDenseHous", "PctHousLess3BR", "MedNumBR", "HousVacant", "PctHousOccup", "PctHousOwnOcc", "PctVacantBoarded", "PctVacMore6Mos", "MedYrHousBuilt", "PctHousNoPhone", "PctWOFullPlumb", "OwnOccLowQuart", "OwnOccMedVal", "OwnOccHiQuart", "RentLowQ", "RentMedian", "RentHighQ", "MedRent", "MedRentPctHousInc", "MedOwnCostPctInc", "MedOwnCostPctIncNoMtg", "NumInShelters", "NumStreet", "PctForeignBorn", "PctBornSameState", "PctSameHouse85", "PctSameCity85", "PctSameState85", "LemasSwornFT", "LemasSwFTPerPop", "LemasSwFTFieldOps", "LemasSwFTFieldPerPop", "LemasTotalReq", "LemasTotReqPerPop", "PolicReqPerOffic", "PolicPerPop", "RacialMatchCommPol", "PctPolicWhite", "PctPolicBlack", "PctPolicHisp", "PctPolicAsian", "PctPolicMinor", "OfficAssgnDrugUnits", "NumKindsDrugsSeiz", "PolicAveOTWorked", "LandArea", "PopDens", "PctUsePubTrans", "PolicCars", "PolicOperBudg", "LemasPctPolicOnPatr", "LemasGangUnitDeploy", "LemasPctOfficDrugUn", "PolicBudgPerPop", "ViolentCrimesPerPop"]
# + colab={"base_uri": "https://localhost:8080/", "height": 246} id="gJ_JcV-V_Bdu" outputId="f0ac9073-186e-47ca-c8ab-85d8012ec0ce"
dataset_url = "http://archive.ics.uci.edu/ml/machine-learning-databases/communities/communities.data"
# Read dataset from the UCI web repository and assign column names.
data_df = pd.read_csv(dataset_url, sep=",", names=column_names,
na_values="?")
data_df.head()
# + [markdown] id="MWHnYDmL_Bdx"
# The 'ViolentCrimesPerPop' column contains the per capita crime rate for each community. We label the communities with a crime rate above the 70-th percentile as 'high crime' and the others as 'low crime'. These would serve as our binary target labels.
# + id="fJtOkt90_Bdy"
# Make sure there are no missing values in the "ViolentCrimesPerPop" column.
assert(not data_df["ViolentCrimesPerPop"].isna().any())
# Binarize the "ViolentCrimesPerPop" column and obtain labels.
crime_rate_70_percentile = data_df["ViolentCrimesPerPop"].quantile(q=0.7)
labels_df = (data_df["ViolentCrimesPerPop"] >= crime_rate_70_percentile)
# Now that we have assigned binary labels,
# we drop the "ViolentCrimesPerPop" column from the data frame.
data_df.drop(columns="ViolentCrimesPerPop", inplace=True)
# + [markdown] id="n8-rQaGH_Bd2"
# We drop all categorical columns, and use only the numerical/boolean features.
# + id="WC12I50e_Bd3"
data_df.drop(columns=["state", "county", "community", "communityname", "fold"],
inplace=True)
# + [markdown] id="2eJcNA1B_Bd5"
# Some of the numerical columns contain missing values (denoted by a NaN). For each feature that has at least one value missing, we append an additional boolean "is_missing" feature indicating that the value was missing, and fill the missing value with 0.
# + id="8_RgukXn_Bd5"
feature_names = data_df.columns
for feature_name in feature_names:
# Which rows have missing values?
missing_rows = data_df[feature_name].isna()
if missing_rows.any(): # Check if at least one row has a missing value.
data_df[feature_name].fillna(0.0, inplace=True) # Fill NaN with 0.
missing_rows.rename(feature_name + "_is_missing", inplace=True)
data_df = data_df.join(missing_rows) # Append "is_missing" feature.
# + [markdown] id="S8U5yENt_Bd-"
# Finally, we divide the dataset randomly into two-thirds for training and one-thirds for testing.
# + id="bixDM2G0Bspg"
# Set random seed so that the results are reproducible.
np.random.seed(123456)
# Train and test indices.
train_indices, test_indices = model_selection.train_test_split(
np.arange(data_df.shape[0]), test_size=1./3.)
# Train and test data.
x_train_df = data_df.loc[train_indices].astype(np.float32)
y_train_df = labels_df.loc[train_indices].astype(np.float32)
x_test_df = data_df.loc[test_indices].astype(np.float32)
y_test_df = labels_df.loc[test_indices].astype(np.float32)
# Convert data frames to NumPy arrays.
x_train = x_train_df.values
y_train = y_train_df.values
x_test = x_test_df.values
y_test = y_test_df.values
# + [markdown] id="gXyyoSrG_BeA"
# ## (i) PR-AUC Training with Plain TF
# + id="hoqCCpf0oTIY"
batch_size = 128 # first fix the batch size for mini-batch training
# + [markdown] id="wqo2mpRi_BeA"
#
# We will work with a linear classification model and define the data and model tensors.
#
# + id="i1fxMQmy_BeB"
# Create linear Keras model.
layers = []
layers.append(tf.keras.Input(shape=(x_train.shape[-1],)))
layers.append(tf.keras.layers.Dense(1))
model = tf.keras.Sequential(layers)
# Create nullary functions that return labels and logits from the current
# batch. In eager mode, TFCO requires these to be provided via nullary function.
# We will maintain a running array of batch indices.
batch_indices = np.arange(batch_size)
labels_fn = lambda: tf.constant(y_train[batch_indices], dtype=tf.float32)
logits_fn = lambda: model(x_train[batch_indices, :])
# + [markdown] id="gmFBVhsQ_BeD"
# We next set up the constraint optimization problem to optimize PR-AUC.
#
# + id="8Swn_y0T_BeD"
# Create context with labels and predictions.
context = tfco.rate_context(logits_fn, labels_fn)
# Create optimization problem with PR-AUC as the objective. The library
# expects a minimization objective, so we negate the PR-AUC.
# We use the pr_auc rate helper which uses a Riemann approximation to the area
# under the precision-recall curve (recall on the horizontal axis, precision on
# the vertical axis). We would need to specify the the number of bins
# ("rectangles") to use for the Riemann approximation. We also can optionally
# specify the surrogate to be used to approximate the PR-AUC.
pr_auc_rate = tfco.pr_auc(
context, bins=10, penalty_loss=tfco.SoftmaxCrossEntropyLoss())
problem = tfco.RateMinimizationProblem(-pr_auc_rate)
# + [markdown] id="OwpTHgqg_BeG"
# We then create a loss function from the `problem` and optimize it to train the model.
# + id="nkq2aVIn_BeG"
# Create Lagrangian loss for `problem`. What we get back is a loss function, a
# a nullary function that returns a list of update_ops that need to be run
# before every gradient update, and the Lagrange multiplier variables internally
# maintained by the loss function. The argument `dual_scale` is a
# hyper-parameter that specifies the relative importance placed on updates on
# the Lagrange multipliers.
loss_fn, update_ops_fn, multipliers = tfco.create_lagrangian_loss(
problem, dual_scale=1.0)
# Set up optimizer and the list of variables to optimize.
optimizer = tf.keras.optimizers.Adagrad(learning_rate=0.1)
var_list = (
model.trainable_weights + list(problem.trainable_variables) + [multipliers])
# + [markdown] id="Ra-Cog9C_BeI"
# Before proceeding to solving the training problem, we write an evaluation function.
# + id="gFUyqbVx_BeI"
def pr_auc(model, features, labels):
# Returns the PR-AUC for given model, features and binary labels.
scores = model.predict(features)
return metrics.average_precision_score(labels, scores)
# + [markdown] id="MID6ChJn_BeK"
# We are now ready to train our model.
# + colab={"base_uri": "https://localhost:8080/", "height": 261} id="ZWUWtSp8NzK5" outputId="bd5488b6-030c-4cb7-e276-8788ec81a3d5"
num_steps = 250
num_examples = x_train.shape[0]
train_objectives = []
test_objectives = []
for ii in range(num_steps):
# Indices for current batch; cycle back once we reach the end of stream.
batch_indices = np.arange(ii * batch_size, (ii + 1) * batch_size)
batch_indices = [ind % num_examples for ind in batch_indices]
# First run update ops, and then gradient update.
update_ops_fn()
optimizer.minimize(loss_fn, var_list=var_list)
# Record train and test objectives once every 10 steps.
if ii % 10 == 0:
train_objectives.append(pr_auc(model, x_train, y_train))
test_objectives.append(pr_auc(model, x_test, y_test))
# Plot training and test objective as a function of steps.
fig, ax = plt.subplots(1, 2, figsize=(7, 3.5))
ax[0].plot(np.arange(1, num_steps + 1, 10), train_objectives)
ax[0].set_title('Train PR-AUC')
ax[0].set_xlabel('Steps')
ax[1].plot(np.arange(1, num_steps + 1, 10), test_objectives)
ax[1].set_title('Test PR-AUC')
ax[1].set_xlabel('Steps')
fig.tight_layout()
# + [markdown] id="BTGOcqe0pO5a"
# # (ii) PR-AUC Training with Custom Estimators
# + [markdown] id="uppYKix2-SdA"
# We next show how one can use TFCO to optimize PR-AUC using custom tf.Estimators.
#
# We first create `feature_columns` to convert the dataset into a format that can be processed by an estimator.
# + id="_klY8Dqueag_"
feature_columns = []
for feature_name in x_train_df.columns:
feature_columns.append(
tf.feature_column.numeric_column(feature_name, dtype=tf.float32))
# + [markdown] id="jyaRvuOXAGt-"
# We next construct the input functions that return the data to be used by the estimator for training/evaluation.
# + id="HoEGUpg9pTdD"
def make_input_fn(
data_df, label_df, num_epochs=10, shuffle=True, batch_size=32):
def input_fn():
ds = tf.data.Dataset.from_tensor_slices((dict(data_df), label_df))
if shuffle:
ds = ds.shuffle(1000)
ds = ds.batch(batch_size).repeat(num_epochs)
return ds
return input_fn
train_input_fn = make_input_fn(x_train_df, y_train_df, num_epochs=25)
test_input_fn = make_input_fn(x_test_df, y_test_df, num_epochs=1, shuffle=False)
# + [markdown] id="QSw7IHgKA5NC"
# We then write the model function that is used by the estimator to create the model, loss, optimizers and metrics.
# + id="fUD8PAddptO7"
def make_model_fn(feature_columns):
# Returns model_fn.
def model_fn(features, labels, mode):
# Create model from features.
layers = []
layers.append(tf.keras.layers.DenseFeatures(feature_columns))
layers.append(tf.keras.layers.Dense(1))
model = tf.keras.Sequential(layers)
logits = model(features)
# Baseline cross-entropy loss.
baseline_loss_fn = tf.keras.losses.BinaryCrossentropy(from_logits=True)
baseline_loss = baseline_loss_fn(labels, logits)
# As a slight variant from the above previous training, we will optimize a
# weighted combination of PR-AUC and the baseline loss.
baseline_coef = 0.2
train_op = None
if mode == tf.estimator.ModeKeys.TRAIN:
# Set up PR-AUC optimization problem.
# Create context with labels and predictions.
context = tfco.rate_context(logits, labels)
# Create optimization problem with PR-AUC as the objective. The library
# expects a minimization objective, so we negate the PR-AUC. We optimize
# a convex combination of (negative) PR-AUC and the baseline loss (wrapped
# in a rate object).
pr_auc_rate = tfco.pr_auc(
context, bins=10, penalty_loss=tfco.SoftmaxCrossEntropyLoss())
problem = tfco.RateMinimizationProblem((1 - baseline_coef) *
(-pr_auc_rate) + baseline_coef *
tfco.wrap_rate(baseline_loss))
# Create Lagrangian loss for `problem`. What we get back is a loss
# function, a nullary function that returns a list of update_ops that
# need to be run before every gradient update, and the Lagrange
# multipliers maintained internally by the loss.
# The argument `dual_scale` is a hyper-parameter that specifies the
# relative importance placed on updates on the Lagrange multipliers.
loss_fn, update_ops_fn, multipliers = tfco.create_lagrangian_loss(
problem, dual_scale=1.0)
# Set up optimizer and the list of variables to optimize the loss.
optimizer = tf.keras.optimizers.Adagrad(learning_rate=0.1)
optimizer.iterations = tf.compat.v1.train.get_or_create_global_step()
# Get minimize op and group with update_ops.
var_list = (
model.trainable_weights + list(problem.trainable_variables) +
[multipliers])
minimize_op = optimizer.get_updates(loss_fn(), var_list)
update_ops = update_ops_fn()
train_op = tf.group(*update_ops, minimize_op)
# Evaluate PR-AUC.
pr_auc_metric = tf.keras.metrics.AUC(curve='PR')
pr_auc_metric.update_state(labels, tf.sigmoid(logits))
# We do not use the Lagrangian loss for evaluation/bookkeeping
# purposes as it depends on some internal variables that may not be
# set properly during evaluation time. We instead pass loss=baseline_loss.
return tf.estimator.EstimatorSpec(
mode=mode,
predictions=logits,
loss=baseline_loss,
train_op=train_op,
eval_metric_ops={'PR-AUC': pr_auc_metric})
return model_fn
# + [markdown] id="4kc3_ftIBC6Y"
# We are now ready to train the estimator.
# + colab={"base_uri": "https://localhost:8080/", "height": 676} id="TImVz7WMp-Nb" outputId="d8f64525-dba6-46b2-a7e6-39d05c0a5d85"
# Create a temporary model directory.
model_dir = "tfco_tmp"
if os.path.exists(model_dir):
shutil.rmtree(model_dir)
# Train estimator.
estimator_lin = tf.estimator.Estimator(
make_model_fn(feature_columns), model_dir=model_dir)
estimator_lin.train(train_input_fn, steps=250)
# + [markdown] id="iYkWYCgvBGJA"
# Finally, we evaluate the trained model on the test set.
# + colab={"base_uri": "https://localhost:8080/", "height": 230} id="iiB6oG2fqC7S" outputId="043ce405-99f7-48a7-d2de-d60708017a30"
estimator_lin.evaluate(test_input_fn)
# + [markdown] id="6xU1eK4GVKd_"
# ## Closing Remarks
# + [markdown] id="zjzlHyFnC7d3"
# Before closing, we point out that there are three main hyper-paramters you may want to tune to improve the PR-AUC training:
#
# - `learning_rate`
# - `dual_scale`
# - `baseline_coeff`
#
# You may also be interested in exploring helpers for other similar metrics that TFCO allows you to optimize:
# - `tfco.precision_at_recall`
# - `tfco.recall_at_precision`
# - `tfco.inverse_precision_at_recall`
| examples/colab/PRAUC_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="rK1pP01MMuU1"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="gtl722MvjuSf"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="F9AnjBfz22gq"
# # Save and load Keras models
# + [markdown] colab_type="text" id="TrNGttwSFElt"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/keras/save_and_serialize"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/keras-team/keras-io/blob/master/tf/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/keras-team/keras-io/blob/master/guides/serialization_and_saving.py"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/keras-io/tf/save_and_serialize.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="PlYTaLGGOlmx"
# ## Introduction
#
# A Keras model consists of multiple components:
#
# - An architecture, or configuration, which specifies what layers the model
# contain, and how they're connected.
# - A set of weights values (the "state of the model").
# - An optimizer (defined by compiling the model).
# - A set of losses and metrics (defined by compiling the model or calling
# `add_loss()` or `add_metric()`).
#
# The Keras API makes it possible to save all of these pieces to disk at once,
# or to only selectively save some of them:
#
# - Saving everything into a single archive in the TensorFlow SavedModel format
# (or in the older Keras H5 format). This is the standard practice.
# - Saving the architecture / configuration only, typically as a JSON file.
# - Saving the weights values only. This is generally used when training the model.
#
# Let's take a look at each of these options: when would you use one or the other?
# How do they work?
# + [markdown] colab_type="text" id="EKhPbck9E82N"
# ## The short answer to saving & loading
#
# If you only have 10 seconds to read this guide, here's what you need to know.
#
# **Saving a Keras model:**
#
# ```python
# model = ... # Get model (Sequential, Functional Model, or Model subclass)
# model.save('path/to/location')
# ```
#
# **Loading the model back:**
#
# ```python
# from tensorflow import keras
# model = keras.models.load_model('path/to/location')
# ```
#
# Now, let's look at the details.
# + [markdown] colab_type="text" id="bT80eTSUngCU"
# ## Setup
# + colab_type="code" id="BallmpGiEbXD"
import numpy as np
import tensorflow as tf
from tensorflow import keras
# + [markdown] colab_type="text" id="rZ6eEK8ekthu"
# ## Whole-model saving & loading
#
# You can save an entire model to a single artifact. It will include:
#
# - The model's architecture/config
# - The model's weight values (which were learned during training)
# - The model's compilation information (if `compile()`) was called
# - The optimizer and its state, if any (this enables you to restart training
# where you left)
#
# #### APIs
#
# - `model.save()` or `tf.keras.models.save_model()`
# - `tf.keras.models.load_model()`
#
# There are two formats you can use to save an entire model to disk:
# **the TensorFlow SavedModel format**, and **the older Keras H5 format**.
# The recommended format is SavedModel. It is the default when you use `model.save()`.
#
# You can switch to the H5 format by:
#
# - Passing `save_format='h5'` to `save()`.
# - Passing a filename that ends in `.h5` or `.keras` to `save()`.
# + [markdown] colab_type="text" id="HUg5WkAAObZn"
# ### SavedModel format
#
# **Example:**
# + colab_type="code" id="MsqSBTGkkGma"
def get_model():
# Create a simple model.
inputs = keras.Input(shape=(32,))
outputs = keras.layers.Dense(1)(inputs)
model = keras.Model(inputs, outputs)
model.compile(optimizer="adam", loss="mean_squared_error")
return model
model = get_model()
# Train the model.
test_input = np.random.random((128, 32))
test_target = np.random.random((128, 1))
model.fit(test_input, test_target)
# Calling `save('my_model')` creates a SavedModel folder `my_model`.
model.save("my_model")
# It can be used to reconstruct the model identically.
reconstructed_model = keras.models.load_model("my_model")
# Let's check:
np.testing.assert_allclose(
model.predict(test_input), reconstructed_model.predict(test_input)
)
# The reconstructed model is already compiled and has retained the optimizer
# state, so training can resume:
reconstructed_model.fit(test_input, test_target)
# + [markdown] colab_type="text" id="onibKMsFZ4Bk"
# #### What the SavedModel contains
#
# Calling `model.save('my_model')` creates a folder named `my_model`,
# containing the following:
# + colab_type="code" id="o0kniAGdvEmH"
# !ls my_model
# + [markdown] colab_type="text" id="9gZk3nwEHKCt"
# The model architecture, and training configuration
# (including the optimizer, losses, and metrics) are stored in `saved_model.pb`.
# The weights are saved in the `variables/` directory.
#
# For detailed information on the SavedModel format, see the
# [SavedModel guide (*The SavedModel format on disk*)](
# https://www.tensorflow.org/guide/saved_model#the_savedmodel_format_on_disk).
#
#
# #### How SavedModel handles custom objects
#
# When saving the model and its layers, the SavedModel format stores the
# class name, **call function**, losses, and weights (and the config, if implemented).
# The call function defines the computation graph of the model/layer.
#
# In the absence of the model/layer config, the call function is used to create
# a model that exists like the original model which can be trained, evaluated,
# and used for inference.
#
# Nevertheless, it is always a good practice to define the `get_config`
# and `from_config` methods when writing a custom model or layer class.
# This allows you to easily update the computation later if needed.
# See the section about [Custom objects](save_and_serialize.ipynb#custom-objects)
# for more information.
#
# Below is an example of what happens when loading custom layers from
# he SavedModel format **without** overwriting the config methods.
# + colab_type="code" id="PPIAXT8BFSf9"
class CustomModel(keras.Model):
def __init__(self, hidden_units):
super(CustomModel, self).__init__()
self.dense_layers = [keras.layers.Dense(u) for u in hidden_units]
def call(self, inputs):
x = inputs
for layer in self.dense_layers:
x = layer(x)
return x
model = CustomModel([16, 16, 10])
# Build the model by calling it
input_arr = tf.random.uniform((1, 5))
outputs = model(input_arr)
model.save("my_model")
# Delete the custom-defined model class to ensure that the loader does not have
# access to it.
del CustomModel
loaded = keras.models.load_model("my_model")
np.testing.assert_allclose(loaded(input_arr), outputs)
print("Original model:", model)
print("Loaded model:", loaded)
# + [markdown] colab_type="text" id="WnESi1jRVLHz"
# As seen in the example above, the loader dynamically creates a new model class
# that acts like the original model.
# + [markdown] colab_type="text" id="STywDB8VW8gu"
# ### Keras H5 format
#
# Keras also supports saving a single HDF5 file containing the model's architecture,
# weights values, and `compile()` information.
# It is a light-weight alternative to SavedModel.
#
# **Example:**
# + colab_type="code" id="gRIvOIfqWhQJ"
model = get_model()
# Train the model.
test_input = np.random.random((128, 32))
test_target = np.random.random((128, 1))
model.fit(test_input, test_target)
# Calling `save('my_model.h5')` creates a h5 file `my_model.h5`.
model.save("my_h5_model.h5")
# It can be used to reconstruct the model identically.
reconstructed_model = keras.models.load_model("my_h5_model.h5")
# Let's check:
np.testing.assert_allclose(
model.predict(test_input), reconstructed_model.predict(test_input)
)
# The reconstructed model is already compiled and has retained the optimizer
# state, so training can resume:
reconstructed_model.fit(test_input, test_target)
# + [markdown] colab_type="text" id="bjxsX8XdS4Oj"
# #### Limitations
#
# Compared to the SavedModel format, there are two things that don't
# get included in the H5 file:
#
# - **External losses & metrics** added via `model.add_loss()`
# & `model.add_metric()` are not saved (unlike SavedModel).
# If you have such losses & metrics on your model and you want to resume training,
# you need to add these losses back yourself after loading the model.
# Note that this does not apply to losses/metrics created *inside* layers via
# `self.add_loss()` & `self.add_metric()`. As long as the layer gets loaded,
# these losses & metrics are kept, since they are part of the `call` method of the layer.
# - The **computation graph of custom objects** such as custom layers
# is not included in the saved file. At loading time, Keras will need access
# to the Python classes/functions of these objects in order to reconstruct the model.
# See [Custom objects](save_and_serialize.ipynb#custom-objects).
#
# + [markdown] colab_type="text" id="cY3tXyZyk4Ws"
# ## Saving the architecture
#
# The model's configuration (or architecture) specifies what layers the model
# contains, and how these layers are connected*. If you have the configuration of a model,
# then the model can be created with a freshly initialized state for the weights
# and no compilation information.
#
# *Note this only applies to models defined using the functional or Sequential apis
# not subclassed models.
# + [markdown] colab_type="text" id="rIjTX1Z0ljoo"
# ### Configuration of a Sequential model or Functional API model
#
# These types of models are explicit graphs of layers: their configuration
# is always available in a structured form.
#
# #### APIs
#
# - `get_config()` and `from_config()`
# - `tf.keras.models.model_to_json()` and `tf.keras.models.model_from_json()`
# + [markdown] colab_type="text" id="F7V8jN9nt9hB"
# #### `get_config()` and `from_config()`
#
# Calling `config = model.get_config()` will return a Python dict containing
# the configuration of the model. The same model can then be reconstructed via
# `Sequential.from_config(config)` (for a `Sequential` model) or
# `Model.from_config(config)` (for a Functional API model).
#
# The same workflow also works for any serializable layer.
#
# **Layer example:**
# + colab_type="code" id="E4H3XIDY91oy"
layer = keras.layers.Dense(3, activation="relu")
layer_config = layer.get_config()
new_layer = keras.layers.Dense.from_config(layer_config)
# + [markdown] colab_type="text" id="2orPhGTaHZRX"
# **Sequential model example:**
# + colab_type="code" id="F09I6yvGV2uf"
model = keras.Sequential([keras.Input((32,)), keras.layers.Dense(1)])
config = model.get_config()
new_model = keras.Sequential.from_config(config)
# + [markdown] colab_type="text" id="Q9SuxM15lEUr"
# **Functional model example:**
# + colab_type="code" id="HHIVpEKSsT8o"
inputs = keras.Input((32,))
outputs = keras.layers.Dense(1)(inputs)
model = keras.Model(inputs, outputs)
config = model.get_config()
new_model = keras.Model.from_config(config)
# + [markdown] colab_type="text" id="NDjRR6fO4GS6"
# #### `to_json()` and `tf.keras.models.model_from_json()`
#
# This is similar to `get_config` / `from_config`, except it turns the model
# into a JSON string, which can then be loaded without the original model class.
# It is also specific to models, it isn't meant for layers.
#
# **Example:**
# + colab_type="code" id="J7jcVOpdPRie"
model = keras.Sequential([keras.Input((32,)), keras.layers.Dense(1)])
json_config = model.to_json()
new_model = keras.models.model_from_json(json_config)
# + [markdown] colab_type="text" id="WE6kPB1B8Xy5"
# ### Custom objects
#
# **Models and layers**
#
# The architecture of subclassed models and layers are defined in the methods
# `__init__` and `call`. They are considered Python bytecode,
# which cannot be serialized into a JSON-compatible config
# -- you could try serializing the bytecode (e.g. via `pickle`),
# but it's completely unsafe and means your model cannot be loaded on a different system.
#
# In order to save/load a model with custom-defined layers, or a subclassed model,
# you should overwrite the `get_config` and optionally `from_config` methods.
# Additionally, you should use register the custom object so that Keras is aware of it.
#
# **Custom functions**
#
# Custom-defined functions (e.g. activation loss or initialization) do not need
# a `get_config` method. The function name is sufficient for loading as long
# as it is registered as a custom object.
#
# **Loading the TensorFlow graph only**
#
# It's possible to load the TensorFlow graph generated by the Keras. If you
# do so, you won't need to provide any `custom_objects`. You can do so like
# this:
# + colab_type="code" id="znOcN8keiaaD"
model.save("my_model")
tensorflow_graph = tf.saved_model.load("my_model")
x = np.random.uniform(size=(4, 32)).astype(np.float32)
predicted = tensorflow_graph(x).numpy()
# + [markdown] colab_type="text" id="Ovu5chswcHzn"
# Note that this method has several drawbacks:
# * For traceability reasons, you should always have access to the custom
# objects that were used. You wouldn't want to put in production a model
# that you cannot re-create.
# * The object returned by `tf.saved_model.load` isn't a Keras model. So it's
# not as easy to use. For example, you won't have access to `.predict()` or `.fit()`
#
# Even if its use is discouraged, it can help you if you're in a tight spot,
# for example, if you lost the code of your custom objects or have issues
# loading the model with `tf.keras.models.load_model()`.
#
# You can find out more in
# the [page about `tf.saved_model.load`](https://www.tensorflow.org/api_docs/python/tf/saved_model/load)
# + [markdown] colab_type="text" id="B5p8XgNCi0Sm"
# #### Defining the config methods
#
# Specifications:
#
# * `get_config` should return a JSON-serializable dictionary in order to be
# compatible with the Keras architecture- and model-saving APIs.
# * `from_config(config)` (`classmethod`) should return a new layer or model
# object that is created from the config.
# The default implementation returns `cls(**config)`.
#
# **Example:**
# + colab_type="code" id="YeVMs9Rs5ojC"
class CustomLayer(keras.layers.Layer):
def __init__(self, a):
self.var = tf.Variable(a, name="var_a")
def call(self, inputs, training=False):
if training:
return inputs * self.var
else:
return inputs
def get_config(self):
return {"a": self.var.numpy()}
# There's actually no need to define `from_config` here, since returning
# `cls(**config)` is the default behavior.
@classmethod
def from_config(cls, config):
return cls(**config)
layer = CustomLayer(5)
layer.var.assign(2)
serialized_layer = keras.layers.serialize(layer)
new_layer = keras.layers.deserialize(
serialized_layer, custom_objects={"CustomLayer": CustomLayer}
)
# + [markdown] colab_type="text" id="OlbIz9cmWDsr"
# #### Registering the custom object
#
# Keras keeps a note of which class generated the config.
# From the example above, `tf.keras.layers.serialize`
# generates a serialized form of the custom layer:
#
# ```
# {'class_name': 'CustomLayer', 'config': {'a': 2}}
# ```
#
# Keras keeps a master list of all built-in layer, model, optimizer,
# and metric classes, which is used to find the correct class to call `from_config`.
# If the class can't be found, then an error is raised (`Value Error: Unknown layer`).
# There are a few ways to register custom classes to this list:
#
# 1. Setting `custom_objects` argument in the loading function. (see the example
# in section above "Defining the config methods")
# 2. `tf.keras.utils.custom_object_scope` or `tf.keras.utils.CustomObjectScope`
# 3. `tf.keras.utils.register_keras_serializable`
# + [markdown] colab_type="text" id="5X5chZaxYpC2"
# #### Custom layer and function example
# + colab_type="code" id="MdYdOM5u4NJ9"
class CustomLayer(keras.layers.Layer):
def __init__(self, units=32, **kwargs):
super(CustomLayer, self).__init__(**kwargs)
self.units = units
def build(self, input_shape):
self.w = self.add_weight(
shape=(input_shape[-1], self.units),
initializer="random_normal",
trainable=True,
)
self.b = self.add_weight(
shape=(self.units,), initializer="random_normal", trainable=True
)
def call(self, inputs):
return tf.matmul(inputs, self.w) + self.b
def get_config(self):
config = super(CustomLayer, self).get_config()
config.update({"units": self.units})
return config
def custom_activation(x):
return tf.nn.tanh(x) ** 2
# Make a model with the CustomLayer and custom_activation
inputs = keras.Input((32,))
x = CustomLayer(32)(inputs)
outputs = keras.layers.Activation(custom_activation)(x)
model = keras.Model(inputs, outputs)
# Retrieve the config
config = model.get_config()
# At loading time, register the custom objects with a `custom_object_scope`:
custom_objects = {"CustomLayer": CustomLayer, "custom_activation": custom_activation}
with keras.utils.custom_object_scope(custom_objects):
new_model = keras.Model.from_config(config)
# + [markdown] colab_type="text" id="Ia1JUuCjy70o"
# ### In-memory model cloning
#
# You can also do in-memory cloning of a model via `tf.keras.models.clone_model()`.
# This is equivalent to getting the config then recreating the model from its config
# (so it does not preserve compilation information or layer weights values).
#
# **Example:**
# + colab_type="code" id="16KQFlItCZf2"
with keras.utils.custom_object_scope(custom_objects):
new_model = keras.models.clone_model(model)
# + [markdown] colab_type="text" id="wq1Dgi9eZUrR"
# ## Saving & loading only the model's weights values
#
# You can choose to only save & load a model's weights. This can be useful if:
#
# - You only need the model for inference: in this case you won't need to
# restart training, so you don't need the compilation information or optimizer state.
# - You are doing transfer learning: in this case you will be training a new model
# reusing the state of a prior model, so you don't need the compilation
# information of the prior model.
# + [markdown] colab_type="text" id="dRJgbG8Zq7WB"
# ### APIs for in-memory weight transfer
#
# Weights can be copied between different objects by using `get_weights`
# and `set_weights`:
#
# * `tf.keras.layers.Layer.get_weights()`: Returns a list of numpy arrays.
# * `tf.keras.layers.Layer.set_weights()`: Sets the model weights to the values
# in the `weights` argument.
#
# Examples below.
#
#
# ***Transfering weights from one layer to another, in memory***
# + colab_type="code" id="xXT0h7yxAA4e"
def create_layer():
layer = keras.layers.Dense(64, activation="relu", name="dense_2")
layer.build((None, 784))
return layer
layer_1 = create_layer()
layer_2 = create_layer()
# Copy weights from layer 2 to layer 1
layer_2.set_weights(layer_1.get_weights())
# + [markdown] colab_type="text" id="IvCxdjmy6eKA"
# ***Transfering weights from one model to another model with a
# compatible architecture, in memory***
# + colab_type="code" id="CleccO1um5WU"
# Create a simple functional model
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = keras.layers.Dense(10, name="predictions")(x)
functional_model = keras.Model(inputs=inputs, outputs=outputs, name="3_layer_mlp")
# Define a subclassed model with the same architecture
class SubclassedModel(keras.Model):
def __init__(self, output_dim, name=None):
super(SubclassedModel, self).__init__(name=name)
self.output_dim = output_dim
self.dense_1 = keras.layers.Dense(64, activation="relu", name="dense_1")
self.dense_2 = keras.layers.Dense(64, activation="relu", name="dense_2")
self.dense_3 = keras.layers.Dense(output_dim, name="predictions")
def call(self, inputs):
x = self.dense_1(inputs)
x = self.dense_2(x)
x = self.dense_3(x)
return x
def get_config(self):
return {"output_dim": self.output_dim, "name": self.name}
subclassed_model = SubclassedModel(10)
# Call the subclassed model once to create the weights.
subclassed_model(tf.ones((1, 784)))
# Copy weights from functional_model to subclassed_model.
subclassed_model.set_weights(functional_model.get_weights())
assert len(functional_model.weights) == len(subclassed_model.weights)
for a, b in zip(functional_model.weights, subclassed_model.weights):
np.testing.assert_allclose(a.numpy(), b.numpy())
# + [markdown] colab_type="text" id="V42tpJDicL4v"
# ***The case of stateless layers***
#
# Because stateless layers do not change the order or number of weights,
# models can have compatible architectures even if there are extra/missing
# stateless layers.
# + colab_type="code" id="TWVjoCuVP6to"
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = keras.layers.Dense(10, name="predictions")(x)
functional_model = keras.Model(inputs=inputs, outputs=outputs, name="3_layer_mlp")
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
# Add a dropout layer, which does not contain any weights.
x = keras.layers.Dropout(0.5)(x)
outputs = keras.layers.Dense(10, name="predictions")(x)
functional_model_with_dropout = keras.Model(
inputs=inputs, outputs=outputs, name="3_layer_mlp"
)
functional_model_with_dropout.set_weights(functional_model.get_weights())
# + [markdown] colab_type="text" id="tUrgZcDAYaML"
# ### APIs for saving weights to disk & loading them back
#
# Weights can be saved to disk by calling `model.save_weights`
# in the following formats:
#
# * TensorFlow Checkpoint
# * HDF5
#
# The default format for `model.save_weights` is TensorFlow checkpoint.
# There are two ways to specify the save format:
#
# 1. `save_format` argument: Set the value to `save_format="tf"` or `save_format="h5"`.
# 2. `path` argument: If the path ends with `.h5` or `.hdf5`,
# then the HDF5 format is used. Other suffixes will result in a TensorFlow
# checkpoint unless `save_format` is set.
#
# There is also an option of retrieving weights as in-memory numpy arrays.
# Each API has its pros and cons which are detailed below.
# + [markdown] colab_type="text" id="de8G1QVux2za"
# ### TF Checkpoint format
#
# **Example:**
# + colab_type="code" id="1W82BZuskILz"
# Runnable example
sequential_model = keras.Sequential(
[
keras.Input(shape=(784,), name="digits"),
keras.layers.Dense(64, activation="relu", name="dense_1"),
keras.layers.Dense(64, activation="relu", name="dense_2"),
keras.layers.Dense(10, name="predictions"),
]
)
sequential_model.save_weights("ckpt")
load_status = sequential_model.load_weights("ckpt")
# `assert_consumed` can be used as validation that all variable values have been
# restored from the checkpoint. See `tf.train.Checkpoint.restore` for other
# methods in the Status object.
load_status.assert_consumed()
# + [markdown] colab_type="text" id="CUDB1dkiecxZ"
# #### Format details
#
# The TensorFlow Checkpoint format saves and restores the weights using
# object attribute names. For instance, consider the `tf.keras.layers.Dense` layer.
# The layer contains two weights: `dense.kernel` and `dense.bias`.
# When the layer is saved to the `tf` format, the resulting checkpoint contains the keys
# `"kernel"` and `"bias"` and their corresponding weight values.
# For more information see
# ["Loading mechanics" in the TF Checkpoint guide](https://www.tensorflow.org/guide/checkpoint#loading_mechanics).
#
# Note that attribute/graph edge is named after **the name used in parent object,
# not the name of the variable**. Consider the `CustomLayer` in the example below.
# The variable `CustomLayer.var` is saved with `"var"` as part of key, not `"var_a"`.
# + colab_type="code" id="wwjjEg7zQ29O"
class CustomLayer(keras.layers.Layer):
def __init__(self, a):
self.var = tf.Variable(a, name="var_a")
layer = CustomLayer(5)
layer_ckpt = tf.train.Checkpoint(layer=layer).save("custom_layer")
ckpt_reader = tf.train.load_checkpoint(layer_ckpt)
ckpt_reader.get_variable_to_dtype_map()
# + [markdown] colab_type="text" id="tfdbha2TvYWH"
# #### Transfer learning example
#
# Essentially, as long as two models have the same architecture,
# they are able to share the same checkpoint.
#
# **Example:**
# + colab_type="code" id="6Xqhxo35q0qj"
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = keras.layers.Dense(10, name="predictions")(x)
functional_model = keras.Model(inputs=inputs, outputs=outputs, name="3_layer_mlp")
# Extract a portion of the functional model defined in the Setup section.
# The following lines produce a new model that excludes the final output
# layer of the functional model.
pretrained = keras.Model(
functional_model.inputs, functional_model.layers[-1].input, name="pretrained_model"
)
# Randomly assign "trained" weights.
for w in pretrained.weights:
w.assign(tf.random.normal(w.shape))
pretrained.save_weights("pretrained_ckpt")
pretrained.summary()
# Assume this is a separate program where only 'pretrained_ckpt' exists.
# Create a new functional model with a different output dimension.
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = keras.layers.Dense(5, name="predictions")(x)
model = keras.Model(inputs=inputs, outputs=outputs, name="new_model")
# Load the weights from pretrained_ckpt into model.
model.load_weights("pretrained_ckpt")
# Check that all of the pretrained weights have been loaded.
for a, b in zip(pretrained.weights, model.weights):
np.testing.assert_allclose(a.numpy(), b.numpy())
print("\n", "-" * 50)
model.summary()
# Example 2: Sequential model
# Recreate the pretrained model, and load the saved weights.
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
pretrained_model = keras.Model(inputs=inputs, outputs=x, name="pretrained")
# Sequential example:
model = keras.Sequential([pretrained_model, keras.layers.Dense(5, name="predictions")])
model.summary()
pretrained_model.load_weights("pretrained_ckpt")
# Warning! Calling `model.load_weights('pretrained_ckpt')` won't throw an error,
# but will *not* work as expected. If you inspect the weights, you'll see that
# none of the weights will have loaded. `pretrained_model.load_weights()` is the
# correct method to call.
# + [markdown] colab_type="text" id="eCsRvSzqMJ0s"
# It is generally recommended to stick to the same API for building models. If you
# switch between Sequential and Functional, or Functional and subclassed,
# etc., then always rebuild the pre-trained model and load the pre-trained
# weights to that model.
# + [markdown] colab_type="text" id="a9EmwUaZBTeW"
# The next question is, how can weights be saved and loaded to different models
# if the model architectures are quite different?
# The solution is to use `tf.train.Checkpoint` to save and restore the exact layers/variables.
#
# **Example:**
# + colab_type="code" id="j6jE9sz7yQ9b"
# Create a subclassed model that essentially uses functional_model's first
# and last layers.
# First, save the weights of functional_model's first and last dense layers.
first_dense = functional_model.layers[1]
last_dense = functional_model.layers[-1]
ckpt_path = tf.train.Checkpoint(
dense=first_dense, kernel=last_dense.kernel, bias=last_dense.bias
).save("ckpt")
# Define the subclassed model.
class ContrivedModel(keras.Model):
def __init__(self):
super(ContrivedModel, self).__init__()
self.first_dense = keras.layers.Dense(64)
self.kernel = self.add_variable("kernel", shape=(64, 10))
self.bias = self.add_variable("bias", shape=(10,))
def call(self, inputs):
x = self.first_dense(inputs)
return tf.matmul(x, self.kernel) + self.bias
model = ContrivedModel()
# Call model on inputs to create the variables of the dense layer.
_ = model(tf.ones((1, 784)))
# Create a Checkpoint with the same structure as before, and load the weights.
tf.train.Checkpoint(
dense=model.first_dense, kernel=model.kernel, bias=model.bias
).restore(ckpt_path).assert_consumed()
# + [markdown] colab_type="text" id="1R9zCAelVexH"
# ### HDF5 format
#
# The HDF5 format contains weights grouped by layer names.
# The weights are lists ordered by concatenating the list of trainable weights
# to the list of non-trainable weights (same as `layer.weights`).
# Thus, a model can use a hdf5 checkpoint if it has the same layers and trainable
# statuses as saved in the checkpoint.
#
# **Example:**
# + colab_type="code" id="J2LictZSclDh"
# Runnable example
sequential_model = keras.Sequential(
[
keras.Input(shape=(784,), name="digits"),
keras.layers.Dense(64, activation="relu", name="dense_1"),
keras.layers.Dense(64, activation="relu", name="dense_2"),
keras.layers.Dense(10, name="predictions"),
]
)
sequential_model.save_weights("weights.h5")
sequential_model.load_weights("weights.h5")
# + [markdown] colab_type="text" id="rCy09yfqXQT8"
# Note that changing `layer.trainable` may result in a different
# `layer.weights` ordering when the model contains nested layers.
# + colab_type="code" id="VX8hFyI9HgYT"
class NestedDenseLayer(keras.layers.Layer):
def __init__(self, units, name=None):
super(NestedDenseLayer, self).__init__(name=name)
self.dense_1 = keras.layers.Dense(units, name="dense_1")
self.dense_2 = keras.layers.Dense(units, name="dense_2")
def call(self, inputs):
return self.dense_2(self.dense_1(inputs))
nested_model = keras.Sequential([keras.Input((784,)), NestedDenseLayer(10, "nested")])
variable_names = [v.name for v in nested_model.weights]
print("variables: {}".format(variable_names))
print("\nChanging trainable status of one of the nested layers...")
nested_model.get_layer("nested").dense_1.trainable = False
variable_names_2 = [v.name for v in nested_model.weights]
print("\nvariables: {}".format(variable_names_2))
print("variable ordering changed:", variable_names != variable_names_2)
# + [markdown] colab_type="text" id="V4GHHReOFEGq"
# #### Transfer learning example
#
# When loading pretrained weights from HDF5, it is recommended to load
# the weights into the original checkpointed model, and then extract
# the desired weights/layers into a new model.
#
# **Example:**
# + colab_type="code" id="YcgjA7yYG49d"
def create_functional_model():
inputs = keras.Input(shape=(784,), name="digits")
x = keras.layers.Dense(64, activation="relu", name="dense_1")(inputs)
x = keras.layers.Dense(64, activation="relu", name="dense_2")(x)
outputs = keras.layers.Dense(10, name="predictions")(x)
return keras.Model(inputs=inputs, outputs=outputs, name="3_layer_mlp")
functional_model = create_functional_model()
functional_model.save_weights("pretrained_weights.h5")
# In a separate program:
pretrained_model = create_functional_model()
pretrained_model.load_weights("pretrained_weights.h5")
# Create a new model by extracting layers from the original model:
extracted_layers = pretrained_model.layers[:-1]
extracted_layers.append(keras.layers.Dense(5, name="dense_3"))
model = keras.Sequential(extracted_layers)
model.summary()
| site/en-snapshot/guide/keras/save_and_serialize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Dataset
# For this demo, I will be using Tripadvisor Hotel Review datatset.
# ### Preliminaries
#
# #### WordCloud Package - pip install wordcloud
# +
from wordcloud import WordCloud, STOPWORDS
# For Data Manipulation
import pandas as pd
# For cleaning the texts
import re
# For the Mask
from PIL import Image
import numpy as np
import urllib
import requests
# For Visualization
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Import Data
df = pd.read_csv('tripadvisor_hotel_reviews.csv')
print(f'The row and column sizes, respectively of the table are {df.shape}')
df.head()
# ### Data Preprocessing
# <p>To process best wordcloud we can produce, here are few things to do in the preprocessing stage:</p>
# <ul>
# <li>combine all texts into one string</li>
# <li>turn all texts to lower case(the count of the different case letters are counted differently</li>
# <li>remove STOPWORDS</li>
# </ul>
# #### Preprocessing - Combining texts and Converting to Lower Case
text = df['Review'].str.cat(sep=', ').lower()
# #### Preprocessing - Stop Words
# <p>WordCloud comes with a decent list of stopwords such as "the","a","or" and much more. There are ways to eyeball potential words that may appear more frequently than desired and we can always add them to the list of stop words that our WordCloud would exclude. While one can split the text and a do value count, I choose to simply generate the WordCloud first and remove those that we do not want to be part of the image.</p>
# Adding the list of stopwords
stopwords = list(set(STOPWORDS)) + ["n't"]
# ### Visualization
# +
wordcloud = WordCloud(width=800, height=800,
background_color='white',
stopwords=stopwords,
min_font_size=10).generate(text)
# Plot the WordCloud Image
plt.figure(figsize=(8, 8), facecolor=None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# -
# ### Masking
# <p>We can improve the above wordcloud by adding some mask, which we can do so by choosing a PNG image. For the demo, lets choose the airplane icon as it is associated with hotels and the tourism industry.</p>
# <p>In choosing a mask, remember to choose a png or jpeg image with white background. A colored background is treated as a separate object by the computer so it may not capture the shape that we want.</p>
# <p>Now, in masking functions, the white portion of the image should be of value 255 and not 0.</p>
mask = np.array(Image.open(requests.get('https://www.freeiconspng.com/uploads/airplane-icon-image-gallery-1.png', stream=True).raw))
# #### Eyeballing the mask objects:
mask
# #### WordCloud With Mask
# +
wordcloud = WordCloud(background_color ='white',
mask=mask,
stopwords = stopwords,
min_font_size = 10,
width=mask.shape[1],
height=mask.shape[0],
contour_width=1,
contour_color='#000080').generate(text)
plt.figure(figsize = (8, 8), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
# -
# ### Color Functions for The WordCloud
# <p>The color function parameter accepts a function and then ouputs a particular or variant of a color(scheme) for each word depending on its characteristics.</p>
# <p>Many prefer the use of monotone color instead of the default color schemes for plotting software.</p>
def one_color_func(word=None, font_size=None, position=None,
orientation=None, font_path=None,
random_state=None):
h = 204 #0-360
s = 100 #0-100
l = random_state.randint(30, 70)#0-100 As we want to randomize it per color, let's randomize this
return f"hsl({h},{s}%, {l}%)"
wordcloud = WordCloud(background_color ='white',
mask=mask,
stopwords = stopwords,
min_font_size = 10,
width=mask.shape[1],
height=mask.shape[0],
color_func=one_color_func,
contour_width=1,
contour_color='#000080').generate(text)
plt.figure(figsize = (8, 8), facecolor = None)
plt.imshow(wordcloud)
plt.axis("off")
plt.tight_layout(pad = 0)
plt.show()
| Tripadvisor_Hotel_Review_WordCloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to 'saturation'
#
# This is a small example to introduce the technique of saturation. This is an advanced modelling technique in ASP, that allows you to model [coNP](https://en.wikipedia.org/wiki/Co-NP)-type problems.
#
# Let's start by setting up the basics.
# +
import clingo
def print_answer_sets(program):
control = clingo.Control()
control.add("base", [], program)
control.ground([("base", [])])
def on_model(model):
sorted_model = [str(atom) for atom in model.symbols(shown=True)]
sorted_model.sort()
print("Answer set: {{{}}}".format(", ".join(sorted_model)))
control.configuration.solve.models = 0
answer = control.solve(on_model=on_model)
if answer.satisfiable == False:
print("No answer sets")
# -
# ## Example
#
# To explain the technique of saturation, we'll use the example problem of deciding if a given propositional logic formula in CNF is **un**satisfiable.
#
# The typical approach is to express satisfiability, in the sense that we construct an program *P* whose answer sets correspond to satisfying assignments, and thus there is at least one answer set if and only if the formula is satisfiable.
#
# We will now use saturation to construct a program *P* that has an answer set if and only if the propositional logic formula is **unsatisfiable**.
#
# "What's the point?" you may ask. Indeed, for the simple example of satisfiability we wouldn't need this complicated solution. But there are some more complicated problems that we can only express using this more complicated method (in combination with the typical modelling of NP-type problems).
#
# So let's take a simple example propositional formula $\varphi$ in CNF:
#
# $$ \varphi = (x_1 \vee x_2) \wedge (x_1 \vee \neg x_2) \wedge (\neg x_1 \vee x_2) \wedge (\neg x_1 \vee \neg x_2), $$
#
# and let's encode this formula using some facts.
asp_program = """
var(1;2).
clause(1,(1;2)).
clause(2,(1;-2)).
clause(3,(-1;2)).
clause(4,(-1;-2)).
clause(C) :- clause(C,_).
"""
# ## Saturation in a nutshell
#
# The main ideas behind the technique of saturation are the following.
# - We use rules with disjunction in the head to generate a search space that includes all candidate solutions.
# - We introduce some rules that check, for everything in this search space, whether it is *invalid* or *incorrect*.
# - Invalid means that it does not correspond to a candidate solution.
# - Incorrect means that it does correspond to a candidate solution, but not an actual solution.
# - We add some rules that enforce that for every invalid or incorrect option, all atoms involved in this entire process are true.
# - This is what the word 'saturation' refers to: making all involved atoms/facts true.
# - Finally, we add a constraint that states that any answer set must be saturated.
#
# This has the effect that there can only be an answer set if there is **no solution**.
#
# If there is no solution (in other words, the search space contains only invalid and incorrect options), then the rules in the program enforce saturation. We must select at least one option in the search space, and whichever one we pick will lead to saturation.
#
# If there is some solution (in other words, the search space contains some valid and correct option), then there cannot be an answer set. Why is this? This is because the definition of answer sets says that an answer set must be a *minimal* model of its own reduct. Because of the last constraint, the only candidate for an answer set is the saturated set (i.e., the set with all atoms true). But this is not a minimal model, because there is a smaller model (namely, the one that corresponds to some valid and correct option in the search space).
#
# ## Back to our example
#
# Let's run through the steps involved in saturation for our example of unsatisfiability.
#
# The first step is to use some rules with disjunction in the head to generate a search space. Our candidate solutions are the truth assignments to the variables in $\varphi$, so we'll generate a search space that includes this.
asp_program += """
assign(V) ; assign(-V) :- var(V).
"""
# Now let's deduce what options in this search space are invalid. In our case that means any set that assigns some variable both to true and to false.
asp_program += """
invalid :- assign(V), assign(-V), var(V).
"""
# Then, the options that are incorrect are those that do not satisfy some clause. In other words, options that for some clause $c$ set the negation $\neg l$ of each literal $l$ in this clause to true. Let's add a rule for this.
asp_program += """
incorrect :- clause(C), assign(-L) : clause(C,L).
"""
# Now it's time to start the actual saturation. For each invalid or incorrect option, we saturate the whole set.
asp_program += """
saturate :- invalid.
saturate :- incorrect.
assign(V) :- var(V), saturate.
assign(-V) :- var(V), saturate.
invalid :- saturate.
incorrect :- saturate.
"""
# And to make sure that only saturated sets remain as answer sets, we add a constraint that enforces this.
asp_program += """
:- not saturate.
"""
# Finally, let's add some `#show` statements to hide the entire saturated set from answer sets.
asp_program += """
unsat :- saturate.
#show unsat/0.
"""
print_answer_sets(asp_program)
# ## A satisfiable input
#
# And just to show that this indeed gives answer sets only for unsatisfiable formulas, let's try the whole thing again with a satisfiable formula.
# +
asp_program2 = """
var(1;2).
clause(1,(1;2)).
clause(2,(1;-2)).
clause(3,(-1;2)).
clause(C) :- clause(C,_).
assign(V) ; assign(-V) :- var(V).
invalid :- assign(V), assign(-V), var(V).
incorrect :- clause(C), assign(-L) : clause(C,L).
saturate :- invalid.
saturate :- incorrect.
assign(V) :- var(V), saturate.
assign(-V) :- var(V), saturate.
invalid :- saturate.
incorrect :- saturate.
:- not saturate.
unsat :- saturate.
#show unsat/0.
"""
print_answer_sets(asp_program2)
# -
# Let's consider why there is no answer set for this last program ($P = $ `asp_program2`).
# - The only candidate answer set, due to the line `:- not saturate.` is the set that contains `saturate` and therefore (by lines 16–19) all other atoms involved. Let's call this set $A$.
# - This set $A$ can only be an answer set (by definition) if it is a (subset-)minimal model of the reduct $P^A$.
# - The reduct $P^A$ is almost exactly the same as the program $P$—only line 21 is removed.
# - To see why $A$ is not a subset-minimal model of $P^A$, consider the following set $A'$:
# * $A' = \{$ `assign(1)`, `assign(2)` $\} \cup B'$, where $B'$ contains all statement using `var/1`, `clause/1` and `clause/2` needed to satisfy lines 2–6.
# - In other words, $A'$ corresponds to the counterexample $\{ x_1 \mapsto 1, x_2 \mapsto 1 \}$.
# - Since $A' \subsetneq A$, the only thing remaining to show is that $A'$ satisfies all rules of $P^A$. And this one can do straightforwardly by going over all rules one-by-one. (Try this yourself!)
#
# This also helps to clarify why the set $A$ is an answer set if and only if the formula $\varphi$ were unsatisfiable. If the formula is satisfiable, we can always find such an $A' \subsetneq A$ that witnesses that $A$ is not a subset-minimal model of $P^A$. And if the formula is unsatisfiable, then this is not possible.
#
# ## A combined variant
#
# If we remove the rule `:- not saturate.`, we get a variant of the program that either has an answer set containing `unsat` if the formula is unsatisfiable, or that has answer sets corresponding to the satisfying truth assignments.
#
# Let's do this, and start with our original unsatisfiable formula.
# +
asp_program3 = """
var(1;2).
clause(1,(1;2)).
clause(2,(1;-2)).
clause(3,(-1;2)).
clause(4,(-1;-2)).
clause(C) :- clause(C,_).
assign(V) ; assign(-V) :- var(V).
invalid :- assign(V), assign(-V), var(V).
incorrect :- clause(C), assign(-L) : clause(C,L).
saturate :- invalid.
saturate :- incorrect.
assign(V) :- var(V), saturate.
assign(-V) :- var(V), saturate.
invalid :- saturate.
incorrect :- saturate.
unsat :- saturate.
#show unsat/0.
#show assign/1.
"""
print_answer_sets(asp_program3)
# -
# And if we try the same program, but then based on the satisfiable formula where the fourth clause is thrown out:
# +
asp_program3 = """
var(1;2).
clause(1,(1;2)).
clause(2,(1;-2)).
clause(3,(-1;2)).
clause(C) :- clause(C,_).
assign(V) ; assign(-V) :- var(V).
invalid :- assign(V), assign(-V), var(V).
incorrect :- clause(C), assign(-L) : clause(C,L).
saturate :- invalid.
saturate :- incorrect.
assign(V) :- var(V), saturate.
assign(-V) :- var(V), saturate.
invalid :- saturate.
incorrect :- saturate.
unsat :- saturate.
#show unsat/0.
#show assign/1.
"""
print_answer_sets(asp_program3)
# -
# ## Final notes
# Saturation is a tricky and often counterintuitive method, so play around with this and practice with using saturation to get a good feel for it.
#
# This method traces back to a theoretical paper from 1995 (["On the computational cost of disjunctive logic programming: Propositional case"](https://link.springer.com/article/10.1007/BF01536399) by <NAME> and <NAME>). If you find such theoretical work insightful, have a look at (the proof of) Theorem 3.1, where they use the method of saturation.
| advanced/saturation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Shapiro Diagram with RSJ in Si Unit
# ### Cythonized
# ### power in dBm with a guess attenuation
import numpy as np
import matplotlib.pyplot as plt
from datetime import *
from scipy.io import savemat
from scipy.integrate import odeint
# %load_ext Cython
# ### Resistively Shunted Model:
#
# $\frac{d\phi}{dt}=\frac{2eR_N}{\hbar}[I_{DC}+I_{RF}\sin(2\pi f_{RF}t)-I_C\sin\phi]$
#
# Solving $\phi(t)$, then you can get the voltage difference between the superconducting leads:
#
# $V=\frac{\hbar}{2e}\langle\frac{d\phi}{dt}\rangle$
#
# After Normalizing:
# $I_{DC}\leftrightarrow \tilde{I_{DC}}=I_{DC}/I_C$,
#
# $I_{RF} \leftrightarrow \tilde{I_{RF}}=I_{RF}/I_C$,
#
# $ V \leftrightarrow \tilde{V}=\frac{V}{I_CR_N}$,
#
# $ R=\frac{dV}{dI} \leftrightarrow \tilde{R}=\frac{R}{R_N}$,
#
#
# $\because f_0=2eI_CR_N/h$,
#
# $f_{RF} \leftrightarrow \tilde{f_{RF}}=f_{RF}/f_0$,
#
# $t \leftrightarrow \tilde{t}=f_0t$,
#
# The Josephson voltage quantized at $\frac{V}{hf_{RF}f_0/2e}=n \leftrightarrow \frac{V}{f_{RF}f_0}=n$
#
# Here, we can set $f_0=1$ or $\frac{I_CR_N}{hf_0/2e}=1$, without loss of generality
#
# The RSJ model simply becomes (omitting $\tilde{}$):
#
# $\frac{d\phi}{dt}=[I_{DC}+I_{RF}\sin(2\pi f_{RF}t)-\sin\phi]$
#
# At equilibrium, $V=\frac{\hbar}{2e}\langle\frac{d\phi}{dt}\rangle \leftrightarrow \tilde{V}=\frac{1}{2\pi}\langle\frac{d\phi}{d\tilde{t}}\rangle$ would also quantized at integers in the Shapiro step regime.
#
#
#
# ### Cython codes here is to speed up the simulation because python is slower than C:
# + magic_args="--pgo" language="cython"
#
# #Use GNU compiler gcc-10 specified in .bash_profile
# cimport numpy as np
# from libc.math cimport sin, pi
# import numpy as np
#
#
# cdef double Qe=1.608e-19
# cdef double hbar=6.626e-34/2/pi
#
#
# ### cdef is faster but can only be used for cython in this cell
# #cpdef can be used for python outside this cell
#
# cdef double CPR(double A, double G, double C):
# '''
# Current-phase relation for the junction
# '''
# return sin(G) + A*sin(2*G +C*pi)
#
# cdef double i(double t,double i_dc,double i_ac,double f_rf):
# '''
# Applied current
# '''
# return i_dc + i_ac * sin(2*pi*f_rf*t)
#
# cpdef dGdt(y,double t,double i_dc,double i_ac,double f_rf,double A, double C, double Ic, double Rn):
# '''
# Define RSJ model
# '''
# der = 2*Qe*Rn/hbar*(-Ic*CPR(A,y,C) + i(t,i_dc,i_ac,f_rf))
# return der
# +
Qe=1.608e-19
h=6.626e-34
hbar=6.626e-34/2/np.pi
Ic=2e-6
Rn=13.0
w0=2*Qe*Ic*Rn/hbar
f0=w0/2/np.pi
attenuation =-40 # A guess value
#C_array = np.array([0.16])*np.pi
f_array=np.array([1.5])#,1.1,0.9,0.6,0.5])
A=0.909
C_array=[0.16] # as a unit of pi
IDC_step=0.05
IDC_array=np.arange(-5,5+IDC_step/2,IDC_step)*Ic
PRF_step=1
PRF_array=np.arange(-25+attenuation,-24+attenuation+PRF_step/2,PRF_step)
IRF_array = np.sqrt(10**(PRF_array/10)/Rn/1000)/Ic
print("DC array size: "+str(len(IDC_array)))
print("RF array size: "+str(len(PRF_array)))
print("Charecteristic frequency fc = "+str(w0/1e9/2/np.pi)+" GHz")
print("Driving RF frequency f_rf = "+str(f_array*w0/1e9/2/np.pi)+" GHz")
print("C = "+str(C_array)+"*pi")
# -
# ### Test at one RF current
# +
t=np.arange(0,300.01,0.01)/f0/f_array[0]
V=np.empty([len(IDC_array)])
for i in range(0,len(IDC_array)):
G_array= odeint(dGdt,0,t,args=(IDC_array[i],2e-6,f_array[0]*f0,A,C_array[0],Ic,Rn))
V[i]=np.mean(np.gradient(G_array[:-1501,0]))/(0.01/f0/f_array[0])*hbar/2/Qe
JV=h*f_array[0]*f0/2/Qe
# -
plt.figure()
plt.plot(IDC_array/Ic,V/JV)
plt.grid()
# ### Plot Shapiro diagram with loops of f and C
for f in f_array:
for C in C_array:
_name_file = "f_" +str(f)+"f0_A"+str(np.round(A,3))+"_C"+str(np.round(C,2))+"pi"
_name_title = "f= " +str(f)+"*f0, A= "+str(np.round(A,3))+", C= "+str(np.round(C,2))+"pi"
print(_name_title)
T1=datetime.now()
print (T1)
WB_Freq=np.empty([len(IRF_array),len(IDC_array)])
for i in range(0,len(IRF_array)):
print("RF power now: "+str(i)+" of "+str(len(IRF_array))+" ,"+str(datetime.now()),end="\r")
for j in range(0,len(IDC_array)):
t=np.arange(0,300.01,0.01)/f/f0
G_array= odeint(dGdt,0,t,args=(IDC_array[j],IRF_array[i],f*f0,A,C,Ic,Rn))
WB_Freq[i,j]=np.mean(np.gradient(G_array[:-1501,0]))/(0.01/f0/f)*hbar/2/Qe # in the unit of V
DVDI=np.gradient(WB_Freq,IDC_step*Ic,axis=1)
print ("It takes " + str(datetime.now()-T1))
plt.figure()
plt.pcolormesh(IDC_array, PRF_array, DVDI, cmap = 'inferno', vmin = 0,linewidth=0,rasterized=True,shading="auto")
plt.xlabel("DC Current(I/Ic)")
plt.ylabel("RF power (a.u.)")
plt.colorbar(label = "DV/DI")
plt.title(_name_title)
#plt.savefig("DVDI_"+_name_file+".pdf")
plt.show()
JV=h*f*f0/2/Qe
plt.figure()
plt.pcolormesh(IDC_array, PRF_array, WB_Freq/JV, cmap = 'coolwarm',linewidth=0,rasterized=True,shading="auto")#/(np.pi*hbar*f/Qe)
plt.xlabel("DC Current(I/Ic)")
plt.ylabel("RF power(dBm)")
plt.colorbar(label = "$V/(hf/2e)$")
plt.title(_name_title)
#plt.savefig("V_"+_name_file+".pdf")
plt.show()
plt.figure()
plt.plot(IDC_array,WB_Freq[1,:]/JV)#/(np.pi*hbar*f/Qe))
plt.show()
plt.figure()
plt.plot(IDC_array,DVDI[1,:])
plt.show()
#savemat("data"+_name_file+'.mat',mdict={'IDC':IDC_array,'IRF':IRF_array,'PRF':PRF_array,'A':A, 'freq':f_rf,'C':C,'Vmatrix':WB_Freq/w_rf,'DVDI':DVDI})
print('file saved')
| Shapiro/Shapiro_Diagram_RSJ_SI_unit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### \*\*\*needs cleaning***
# +
import pandas as pd
import numpy as np
import sys
import os
import itertools
import time
import random
#import utils
sys.path.insert(0, '../utils/')
from utils_preprocess_v3 import *
from utils_modeling_v9 import *
from utils_plots_v2 import *
#sklearn
from sklearn.metrics import mean_squared_error
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
start_time = time.time()
# -
data = pd.read_csv('../data/datasets_processed/VanAllen_data.csv', index_col = 0)
response = pd.read_csv('../data/datasets_processed/VanAllen_response.csv')
interactome = pd.read_csv('../data/interactomes/inbiomap_processed.txt', sep = '\t')
# +
# get nodes from data and graph
data_nodes = data['node'].tolist()
interactome_nodes = list(set(np.concatenate((interactome['node1'], interactome['node2']))))
# organize data
organize = Preprocessing()
save_location = '../data/reduced_interactomes/reduced_interactome_VanAllen.txt'
organize.transform(data_nodes, interactome_nodes, interactome, data, save_location, load_graph = True)
# +
# extract info from preprocessing
X = organize.sorted_X.T.values
y = response['PFS'].values.reshape(-1,1)
L_norm = organize.L_norm
L = organize.L
g = organize.g
num_to_node = organize.num_to_node
# split for training
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42)
# scaling X
scaler_X = StandardScaler()
X_train = scaler_X.fit_transform(X_train)
X_test = scaler_X.transform(X_test)
# scalying y
scaler_y = StandardScaler()
y_train = scaler_y.fit_transform(y_train).reshape(-1)
y_test = scaler_y.transform(y_test).reshape(-1)
# -
val_1, vec_1 = scipy.linalg.eigh(L_norm.toarray())
val_zeroed = val_1 - min(val_1) + 1e-8
L_rebuild = vec_1.dot(np.diag(val_zeroed)).dot(np.linalg.inv(vec_1))
X_train_lower = np.linalg.cholesky(L_rebuild)
L_rebuild.sum()
# check
X_train_lower.dot(X_train_lower.T).sum()
L_norm.sum()
# # Lasso + LapRidge
# hyperparameters
alpha1_list = np.logspace(-1,0,15)
alpha2_list = np.logspace(-1,2,15)
threshold_list = np.logspace(-3,-1,10)
max_features = 10
alpha_pairs = list(itertools.product(alpha1_list, alpha2_list))
# +
def loss_fn(X,Y, L, alpha1, alpha2, beta):
return 0.5/(len(X)) * cp.norm2(cp.matmul(X, beta) - Y)**2 + \
alpha1 * cp.norm1(beta) + \
alpha2 * cp.sum(cp.quad_form(beta,L))
def run(pair, X_train, y_train, L_norm):
beta = cp.Variable(X_train.shape[1])
alpha1 = cp.Parameter(nonneg=True)
alpha2 = cp.Parameter(nonneg=True)
alpha1.value = pair[0]
alpha2.value = pair[1]
problem = cp.Problem(cp.Minimize(loss_fn(X_train, y_train, L_norm, alpha1, alpha2, beta )))
problem.solve(solver = cp.SCS, verbose=True, max_iters=50000)
return beta.value
# -
betas = Parallel(n_jobs=15, verbose=10)(delayed(run)(alpha_pairs[i],
X_train,
y_train,
L_norm) for i in range(len(alpha_pairs)))
feats = [getFeatures(None, i, threshold=0.001, max_features=10) for i in betas]
regr = LinearRegression()
scores = [getScoring(regr, X_train, y_train, X_test, y_test, i, None) for i in feats]
train_scores = [i[0] for i in scores]
test_scores = [i[1] for i in scores]
gridsearch_results = pd.DataFrame(np.array(test_scores), columns = ['Test MSE'])
getGridsearchPlot(gridsearch_results, alpha1_list, alpha2_list, save_location = None)
np.where(test_scores == min(test_scores))
min(test_scores)
getTranslatedNodes(feats[7], betas[7][feats[7]], num_to_node, g, )
# # MCP + LapRidge
# define training params
alpha1_list = np.logspace(-3,-2,15)
alpha2_list = np.logspace(-1,2,15)
threshold_list = np.logspace(-3,-1,10)
max_features = 10
alpha_list_pairs = list(itertools.product(alpha1_list, alpha2_list))
results = {}
feats_list = []
betas = []
for i in alpha2_list:
X_train_new = np.vstack((X_train, np.sqrt(i)*X_train_lower))
y_train_new = np.concatenate((y_train, np.zeros(len(X_train_lower))))
s = pycasso.Solver(X_train_new, y_train_new, lambdas=alpha1_list, penalty = 'mcp')
s.train()
beta = s.coef()['beta']
betas += [i for i in beta]
feats = [getFeatures(None, i, threshold=0.001, max_features=10) for i in beta]
feats_list += feats
print([len(i) for i in feats])
regr = LinearRegression()
scores = [getScoring(regr, X_train, y_train, X_test, y_test, i, None) for i in feats]
results[i] = scores
train_scores = []
test_scores = []
for k,v in results.items():
train_scores += [i[0] for i in v]
test_scores += [i[1] for i in v]
gridsearch_results = pd.DataFrame(np.array(test_scores), columns = ['Test MSE'])
getGridsearchPlot(gridsearch_results, alpha1_list, alpha2_list, save_location = None)
min(test_scores)
np.where(test_scores == min(test_scores))
getTranslatedNodes(feats_list[215], betas[215][feats_list[215]], num_to_node, g)
# # SCAD + LapRidge
# define training params
alpha1_list = np.logspace(-3,-2,15)
alpha2_list = np.logspace(-1,2,15)
threshold_list = np.logspace(-3,-1,10)
max_features = 10
alpha_list_pairs = list(itertools.product(alpha1_list, alpha2_list))
results = {}
feats_list = []
betas = []
for i in alpha2_list:
X_train_new = np.vstack((X_train, np.sqrt(i)*X_train_lower))
y_train_new = np.concatenate((y_train, np.zeros(len(X_train_lower))))
s = pycasso.Solver(X_train_new, y_train_new, lambdas=alpha1_list, penalty = 'scad')
s.train()
beta = s.coef()['beta']
betas += [i for i in beta]
feats = [getFeatures(None, i, threshold=0.001, max_features=10) for i in beta]
feats_list += feats
print([len(i) for i in feats])
regr = LinearRegression()
scores = [getScoring(regr, X_train, y_train, X_test, y_test, i, None) for i in feats]
results[i] = scores
train_scores = []
test_scores = []
for k,v in results.items():
train_scores += [i[0] for i in v]
test_scores += [i[1] for i in v]
gridsearch_results = pd.DataFrame(np.array(test_scores), columns = ['Test MSE'])
getGridsearchPlot(gridsearch_results, alpha1_list, alpha2_list, save_location = None)
min(test_scores)
np.where(test_scores == min(test_scores))
getTranslatedNodes(feats_list[5], betas[5][feats_list[5]], num_to_node, g)
| notebooks/VanAllen_LapRidge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="n3h9HHyH8Yh4"
# Berdasarkan isu [#141](https://github.com/hidrokit/hidrokit/issues/141): **Uji Chi-Square**
#
# Referensi Isu:
# - <NAME>., <NAME>., Press, U. B., & Media, U. (2017). Rekayasa Statistika untuk Teknik Pengairan. Universitas Brawijaya Press. https://books.google.co.id/books?id=TzVTDwAAQBAJ
# - Soewarno. (1995). hidrologi: Aplikasi Metode Statistik untuk Analisa Data. NOVA.
# - <NAME>. (2018). Rekayasa Hidrologi.
#
# Deskripsi Isu:
# - Melakukan Uji Kecocokan Distribusi menggunakan Uji Chi-Square.
#
# Diskusi Isu:
# - [#182](https://github.com/hidrokit/hidrokit/discussions/182) - Formula pada perhitungan Chi Square (Uji Kecocokan Distribusi).
#
# Strategi:
# - Tidak dibandingkan dengan fungsi `scipy.stats.chisquare`.
# + [markdown] id="nCwAQOWb9U96"
# # PERSIAPAN DAN DATASET
# + id="FG1q9l2Y76tr"
try:
import hidrokit
except ModuleNotFoundError:
# saat dibuat menggunakan cabang @dev/dev0.3.7
# !pip install git+https://github.com/taruma/hidrokit.git@dev/dev0.3.7 -q
# + id="LB6uUJIV9Xbh"
import numpy as np
import pandas as pd
from scipy import stats
from hidrokit.contrib.taruma import hk172, hk124, hk127, hk126
frek_normal, frek_lognormal, frek_gumbel, frek_logpearson3 = hk172, hk124, hk127, hk126
# + colab={"base_uri": "https://localhost:8080/", "height": 394} id="gmxQAUAA9ZRE" outputId="5bb1c7b7-7feb-4a75-ddf8-60f5d8ae6fca"
# contoh data diambil dari buku
# limantara hal. 118
_HUJAN = np.array([85, 92, 115, 116, 122, 52, 69, 95, 96, 105])
_TAHUN = np.arange(1998, 2008) # 1998-2007
data = pd.DataFrame(
data=np.stack([_TAHUN, _HUJAN], axis=1),
columns=['tahun', 'hujan']
)
data.tahun = pd.to_datetime(data.tahun, format='%Y')
data.set_index('tahun', inplace=True)
data
# + [markdown] id="YYwJuJGg95Zy"
# # TABEL
#
# Terdapat 1 tabel untuk modul `hk141` yaitu:
# - `t_chi_lm`: Tabel nilai kritis untuk Distribusi Chi Square ($X^2$) dari buku Rekayasa Hidrologi oleh Limantara.
#
# Dalam modul `hk141` nilai kritis $X^2$ akan dibangkitkan menggunakan fungsi `scipy.stats.chi2.isf` secara `default`. Mohon diperhatikan jika ingin menggunakan nilai $X^2$ yang berasal dari sumber lain.
# + colab={"base_uri": "https://localhost:8080/", "height": 990} id="W3OOs5acDvrL" outputId="3c46d0f1-d6d5-4dfc-b26d-a5b3c269ac84"
# tabel dari limantara hal. 117
# Tabel Nilai Kritis untuk Distribusi Chi Square (X^2)
# KODE: LM
_DATA_LM = [
[0.039, 0.016, 0.698, 0.393, 3.841, 5.024, 6.635, 7.879],
[0.100, 0.201, 0.506, 0.103, 5.991, 0.738, 9.210, 10.597],
[0.717, 0.115, 0.216, 0.352, 7.815, 9.348, 11.345, 12.838],
[0.207, 0.297, 0.484, 0.711, 9.488, 11.143, 13.277, 14.860],
[0.412, 0.554, 0.831, 1.145, 11.070, 12.832, 15.086, 16.750],
[0.676, 0.872, 1.237, 1.635, 12.592, 14.449, 16.812, 18.548],
[0.989, 1.239, 1.690, 2.167, 14.067, 16.013, 18.475, 20.278],
[1.344, 1.646, 2.180, 2.733, 15.507, 17.535, 20.090, 21.955],
[1.735, 2.088, 2.700, 3.325, 16.919, 19.023, 21.666, 23.589],
[2.156, 2.558, 3.247, 3.940, 18.307, 20.483, 23.209, 25.188],
[2.603, 3.053, 3.816, 4.575, 19.675, 21.920, 24.725, 26.757],
[3.074, 3.571, 4.404, 5.226, 21.026, 23.337, 26.217, 28.300],
[3.565, 4.107, 5.009, 5.892, 22.362, 24.736, 27.688, 29.819],
[4.075, 4.660, 5.629, 6.571, 23.685, 26.119, 29.141, 31.319],
[4.601, 5.229, 6.262, 7.261, 24.996, 27.488, 30.578, 32.801],
[5.142, 5.812, 6.908, 7.962, 26.296, 28.845, 32.000, 34.267],
[5.697, 6.408, 7.564, 8.672, 27.587, 30.191, 33.409, 35.718],
[6.265, 7.015, 8.231, 9.390, 28.869, 31.526, 34.805, 37.156],
[6.884, 7.633, 8.907, 10.117, 30.144, 32.852, 36.191, 38.582],
[7.434, 8.260, 9.591, 10.851, 31.410, 34.170, 37.566, 39.997],
[8.034, 8.897, 10.283, 11.591, 32.671, 35.479, 38.932, 41.401],
[8.643, 9.542, 10.982, 12.338, 33.924, 36.781, 40.289, 42.796],
[9.260, 10.196, 11.689, 13.091, 36.172, 38.076, 41.638, 44.181],
[9.886, 10.856, 12.401, 13.848, 36.415, 39.364, 42.980, 45.558],
[10.520, 11.524, 13.120, 14.611, 37.652, 40.646, 44.314, 46.928],
[11.160, 12.198, 13.844, 15.379, 38.885, 41.923, 45.642, 48.290],
[11.808, 12.879, 14.573, 16.151, 40.113, 43.194, 46.963, 49.645],
[12.461, 13.565, 15.308, 16.928, 41.337, 44.461, 48.278, 50.993],
[13.121, 14.256, 16.047, 17.708, 42.557, 45.722, 49.588, 52.336],
[13.787, 14.953, 16.791, 18.493, 43.773, 46.979, 50.892, 53.672],
]
_INDEX_LM = range(1, 31)
_COL_LM = [0.995, .99, .975, .95, .05, .025, 0.01, 0.005]
t_chi_lm = pd.DataFrame(
data=_DATA_LM, index=_INDEX_LM, columns=_COL_LM
)
t_chi_lm
# + [markdown] id="ryR8EYWJGBpM"
# # KODE
# + id="xSbkavoqECGd"
from scipy import interpolate
def _func_interp_bivariate(df):
"Membuat fungsi dari tabel untuk interpolasi bilinear"
table = df[df.columns.sort_values()].sort_index().copy()
x = table.index
y = table.columns
z = table.to_numpy()
# penggunaan kx=1, ky=1 untuk interpolasi linear antara 2 titik
# tidak menggunakan (cubic) spline interpolation
return interpolate.RectBivariateSpline(x, y, z, kx=1, ky=1)
def _as_value(x, dec=4):
x = np.around(x, dec)
return x.flatten() if x.size > 1 else x.item()
# + id="nr0k9aKyGG2Q"
table_source = {
'limantara': t_chi_lm
}
anfrek = {
'normal': frek_normal.calc_x_normal,
'lognormal': frek_lognormal.calc_x_lognormal,
'gumbel': frek_gumbel.calc_x_gumbel,
'logpearson3': frek_logpearson3.calc_x_lp3,
}
def _calc_k(n):
return np.floor(1 + 3.22 * np.log10(n)).astype(int)
def _calc_dk(k, m):
return k - 1 - m
def calc_xcr(alpha, dk, source='scipy'):
alpha = np.array(alpha)
if source.lower() in table_source.keys():
func_table = _func_interp_bivariate(table_source[source.lower()])
return _as_value(func_table(dk, alpha, grid=False), 3)
if source.lower() == 'scipy':
#ref: https://stackoverflow.com/questions/32301698
return stats.chi2.isf(alpha, dk)
def chisquare(
df, col=None, dist='normal', source_dist='scipy',
alpha=0.05, source_xcr='scipy', show_stat=True,
):
source_dist = 'gumbel' if dist.lower() == 'gumbel' else source_dist
col = df.columns[0] if col is None else col
data = df[[col]].copy()
n = len(data)
data = data.rename({col: 'x'}, axis=1)
if dist.lower() in ['lognormal', 'logpearson3']:
data['log_x'] = np.log10(data.x)
k = _calc_k(n)
prob_class = 1 / k
prob_list = np.linspace(0, 1, k+1)[::-1]
prob_seq = prob_list[1:-1]
func = anfrek[dist.lower()]
T = 1 / prob_seq
val_x = func(data.x, return_period=T, source=source_dist)
# Chi Square Table
calc_df = pd.DataFrame()
min = data.x.min()
max = data.x.max()
seq_x = np.concatenate([[min], val_x, [max]])
calc_df['no'] = range(1, k+1)
class_text = []
for i in range(seq_x.size-1):
if i == 0:
class_text += [f'X <= {seq_x[i+1]:.4f}']
elif i == seq_x.size-2:
class_text += [f'X > {seq_x[i]:.4f}']
else:
class_text += [f'{seq_x[i]:.4f} < X <= {seq_x[i+1]:.4f}']
calc_df['batas_kelas'] = class_text
# calculate fe
fe = []
for i in range(seq_x.size-1):
if i == 0:
fe += [(data.x <= seq_x[i+1]).sum()]
elif i == seq_x.size-2:
fe += [(data.x > seq_x[i]).sum()]
else:
fe += [data.x.between(seq_x[i], seq_x[i+1], inclusive='right').sum()]
calc_df['fe'] = fe
ft = prob_class * n
calc_df['ft'] = [ft]*k
if dist.lower() in ['normal', 'gumbel', 'lognormal']:
dk = _calc_dk(k, 2)
elif dist.lower() in ['logpearson3']:
# di buku soetopo nilai m nya diberi angka 3
dk = _calc_dk(k, 2)
X_calc = np.sum(np.power(2, (calc_df.fe-calc_df.ft))/calc_df.ft)
X_critical = calc_xcr(alpha=alpha, dk=dk, source=source_xcr)
result = int(X_calc < X_critical)
result_text = ['Distribusi Tidak Diterima', 'Distribusi Diterima']
calc_df.set_index('no', inplace=True)
if show_stat:
print(f'Periksa Kecocokan Distribusi {dist.title()}')
print(f'Jumlah Kelas = {k}')
print(f'Dk = {dk}')
print(f'X^2_hitungan = {X_calc:.3f}')
print(f'X^2_kritis = {X_critical:.3f}')
print(f'Result (X2_calc < X2_cr) = {result_text[result]}')
return calc_df
# + [markdown] id="Zy9jLuEQIEpp"
# # FUNGSI
# + [markdown] id="xQaXygUAIGj5"
# ## Fungsi `calc_xcr(alpha, dk, ...)`
#
# Function: `calc_xcr(alpha, dk, source='scipy')`
#
# Fungsi `calc_xcr(...)` digunakan untuk mencari nilai $X^2_{kritis}$ dari berbagai sumber berdasarkan nilai derajat kepercayaan $\alpha$ dan nilai $DK$.
#
# - Argumen Posisi:
# - `alpha`: Nilai _level of significance_ $\alpha$. Dalam satuan desimal.
# - `dk`: Nilai $DK$ hasil perhitungan antara $K$ (jumlah kelas) dan parameter distribusi $m$.
# - Argumen Opsional:
# - `source`: sumber nilai $X^2_{kritis}$. `'scipy'` (default). Sumber yang dapat digunakan antara lain: Limantara (`'limantara'`).
# + colab={"base_uri": "https://localhost:8080/"} id="nrukdX0uJ2Vs" outputId="bbb95ace-2ff3-4db9-bb46-6d59f3249046"
calc_xcr(0.05, 3)
# + colab={"base_uri": "https://localhost:8080/"} id="-aUH_AugJ5j_" outputId="e2609ce9-6a4b-4db8-84e2-0159f5d90f16"
calc_xcr([0.05, 0.1, 0.2], 5, source='limantara')
# + colab={"base_uri": "https://localhost:8080/"} id="pM_jik3MKFVC" outputId="9e896f9d-3b38-41e5-979a-821d59888d82"
# perbandingan antara nilai tabel dan fungsi scipy
source_test = ['limantara', 'scipy']
_dk = 5
_alpha = [0.2, 0.15, 0.1, 0.07, 0.05, 0.01]
for _source in source_test:
print(f'Xcr {_source:<12}=', calc_xcr(_alpha, _dk, source=_source))
# + [markdown] id="w4_Zw33TKUWf"
# ## Fungsi `chisquare(df, ...)`
#
# Function: `chisquare(df, col=None, dist='normal', source_dist='scipy', alpha=0.05, source_xcr='scipy', show_stat=True)`
#
# Fungsi `chisquare(...)` merupakan fungsi untuk melakukan uji chi square terhadap distribusi yang dibandingkan. Fungsi ini mengeluarkan objek `pandas.DataFrame`.
#
# - Argumen Posisi:
# - `df`: `pandas.DataFrame`.
# - Argumen Opsional:
# - `col`: nama kolom, `None` (default). Jika tidak diisi menggunakan kolom pertama dalam `df` sebagai data masukan.
# - `dist`: distribusi yang dibandingkan, `'normal'` (distribusi normal) (default). Distribusi yang dapat digunakan antara lain: Log Normal (`'lognormal'`), Gumbel (`'gumbel'`), Log Pearson 3 (`'logpearson3'`).
# - `source_dist`: sumber perhitungan distribusi, `'scipy'` (default). Lihat masing-masing modul analisis frekuensi untuk lebih jelasnya.
# - `alpha`: nilai $\alpha$, `0.05` (default).
# - `source_xcr`: sumber nilai $X^2_{kritis}$, `'scipy'` (default). Sumber yang dapat digunakan antara lain: Limantara (`'limantara'`).
# - `show_stat`: menampilkan hasil luaran uji, `True` (default).
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="2Tv06tUxLC0X" outputId="d090eda4-4bb7-4a3d-e3af-ef61907812ef"
chisquare(data)
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="U2vXMkHRLFhb" outputId="bca954cc-3b35-4c85-d144-e7fa6c697ad1"
chisquare(data, dist='gumbel', source_dist='soetopo')
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="JVogXG-jLJda" outputId="230938a2-b93f-4a95-8c9d-338f780d920c"
chisquare(data, 'hujan', dist='logpearson3', alpha=0.2, source_xcr='limantara')
# + [markdown] id="PjG-gD6kLUnt"
# # Changelog
#
# ```
# - 20220317 - 1.0.0 - Initial
# ```
#
# #### Copyright © 2022 [<NAME>](https://taruma.github.io)
#
# Source code in this notebook is licensed under a [MIT License](https://choosealicense.com/licenses/mit/). Data in this notebook is licensed under a [Creative Common Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0/).
#
| booklet_hidrokit/0.4.0/ipynb/manual/taruma_0_4_0_hk141_chi_square.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pynq.overlays.base import BaseOverlay
base = BaseOverlay("base.bit")
from pynq.lib.arduino import Arduino_Displaycam
test = Arduino_Displaycam(base.ARDUINO)
import time
test.init()
# test.display()
test.gpio()
test.pwm()
print(test.microblaze.read_mailbox(0))
# -
test.layerMode(1)
test.layerEffect(2)
test.layer(1)
time.sleep(0.05)
test.drawMainScreen()
test.layer(0)
test.snapPic(35,0)
test.write_PODNEAR(110,110)
test.clearWindow(1)
#testing redraw
test.draw2X(130, 100, 15, 0x0000)
test.drawTrianglePoint(100,100,15,0x0000)
test.drawPoint(100,130,100,0x0000)
| Pynq_Neil_Copy_05.23/jupyter_notebooks/base/arduino/Testing/.ipynb_checkpoints/Display+Cam_cleaned-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 00: Preliminaries
#
# <img src="../meta/images/python-logo.png" style='float:right;'/>
#
# Week 00 will focus on introducing the class and setting up the tools we'll use.
#
# * [Course Overview](00-Overview.ipynb)
# * [Installation](00-Installation.ipynb)
# * [Exercises](00-Exercises.ipynb)
#
# ## Bonus Material
#
# * [Why Python?](00-Why-Python.ipynb)
# * [Why Sean?](00-Why-Sean.ipynb)
| 00-Preliminaries/00-Index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright 2021 The TensorFlow Similarity Authors.
# +
# @title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# # Tensorflow Similarity Sampler I/O Cookbook
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/similarity/blob/master/examples/sampler_io_cookbook.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/similarity/blob/master/examples/sampler_io_cookbook.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# Tensorflow Similarity's Samplers ensure that each batch contains a target number of examples per class per batch. This ensures that the loss functions are able to construct tuples of anchor, positive, and negatives within each batch of examples.
#
#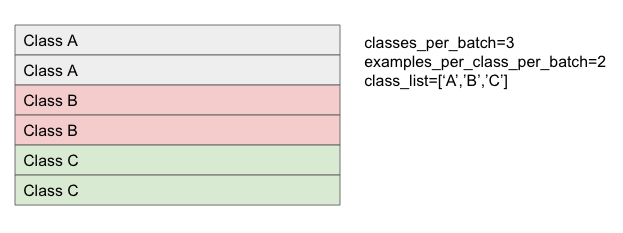 
#
# In this notebook you will learn how to use the:
#
# * `MultiShotMemorySampler()` for fitting to a sequence of data, such as a dataset.
# * `SingleShotMemorySampler()` to treat each example as a seperate class and generate augmented versions within each batch.
# * `TFDatasetMultiShotMemorySampler()` to directly integrate with the Tensorflow dataset catalog.
# ## Imports
# +
import os
import random
from typing import Tuple
import numpy as np
from matplotlib import pyplot as plt
# INFO messages are not printed.
# This must be run before loading other modules.
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "1"
# -
import tensorflow as tf
# install TF similarity if needed
try:
import tensorflow_similarity as tfsim # main package
except ModuleNotFoundError:
# !pip install tensorflow_similarity
import tensorflow_similarity as tfsim
tfsim.utils.tf_cap_memory() # Avoid GPU memory blow up
# <hr>
# # MultiShotMemorySampler: Load Random Numpy Arrays
# The following cell loads random numpy data using TensorFlow similarity`MultiShotMemorySampler()`.
#
# Using a sampler is required to ensure that each batch contains at least N samples for each class included in a batch.
#
# This batch strucutre is required for the contrastive loss to properly compute positive pairwise distances.
# +
num_ms_examples = 100000 # @param {type:"slider", min:1000, max:1000000}
num_ms_features = 784 # @param {type:"slider", min:10, max:1000}
num_ms_classes = 10 # @param {type:"slider", min:2, max:1000}
# We use random floats here to represent a dense feature vector
X_ms = np.random.rand(num_ms_examples, num_ms_features)
# We use random ints to represent N different classes
y_ms = np.random.randint(low=0, high=10, size=num_ms_examples)
# +
num_known_ms_classes = 5 # @param {type:"slider", min:2, max:1000}
ms_classes_per_batch = num_known_ms_classes
ms_examples_per_class_per_batch = 2 # @param {type:"integer"}
ms_class_list = random.sample(range(num_ms_classes), k=num_known_ms_classes)
ms_sampler = tfsim.samplers.MultiShotMemorySampler(
X_ms,
y_ms,
classes_per_batch=ms_classes_per_batch,
examples_per_class_per_batch=ms_examples_per_class_per_batch,
class_list=ms_class_list,
)
# -
# ## Generating Batches
# The Tensorflow Similarity memory samplers are a subclass of [tf.keras.utils.Sequence](https://www.tensorflow.org/api_docs/python/tf/keras/utils/Sequence), overriding the `__getitem__` and `__len__` methods.
#
# Additionally, Tensorflow Similarity provides a `generate_batch()` method that takes a batch ID and yields a single batch.
#
# We verify that the batch batch only conatins the classes defined in `CLASS_LIST` and that each class has `ms_classes_per_batch` * `ms_examples_per_class_per_batch` examples.
# +
X_ms_batch, y_ms_batch = ms_sampler.generate_batch(100)
print("#" * 10 + " X " + "#" * 10)
print(X_ms_batch)
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_ms_batch)
# Check that the batch size is equal to the target number of classes * target number of examples per class.
assert tf.shape(X_ms_batch)[0] == (ms_classes_per_batch * ms_examples_per_class_per_batch)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_ms_batch)[1] == (num_ms_features)
# Check that classes in the batch are from the allowed set in CLASS_LIST
assert set(tf.unique(y_ms_batch)[0].numpy()) - set(ms_class_list) == set()
# Check that we only have NUM_CLASSES_PER_BATCH
assert len(tf.unique(y_ms_batch)[0]) == ms_classes_per_batch
# -
# ## Sampler Sizes
#
# `MultiShotMemorySampler()` provides various attributes for accessing info about the data:
# * `__len__` provides the number of steps per epoch.
# * `num_examples` provides the total number of examples within the sampler.
# * `example_shape` provides the shape of the examples.
#
# The `num_examples` attribute represents the subset of X and y where y is in the `class_list` with each class limited to `num_examples_per_class`.
print(f"The sampler contains {len(ms_sampler)} steps per epoch.")
print(f"The sampler is using {ms_sampler.num_examples} examples out of the original {num_ms_examples}.")
print(f"Each examples has the following shape: {ms_sampler.example_shape}.")
# ## Accessing the Examples
#
# Additionaly, the `MultiShotMemorySampler()` provides `get_slice()` for manually accessing examples within the Sampler.
#
# **NOTE**: the examples are shuffled when creating the Sampler but will yield the same examples for each call to get_slice(begin, size).
# +
# Get 10 examples starting at example 200.
X_ms_slice, y_ms_slice = ms_sampler.get_slice(begin=200, size=10)
print("#" * 10 + " X " + "#" * 10)
print(X_ms_slice)
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_ms_slice)
# Check that the batch size is equal to our get_slice size.
assert tf.shape(X_ms_slice)[0] == 10
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_ms_slice)[1] == (num_ms_features)
# Check that classes in the batch are from the allowed set in CLASS_LIST
assert set(tf.unique(y_ms_slice)[0].numpy()) - set(ms_class_list) == set()
# -
# <hr>
# # SingleShotMemorySampler: Augmented MNIST Examples
#
# The following cell loads and augments MNIST examples using the `SingleShotMemorySampler()`.
#
# The Sampler treats each example as it's own class and adds augmented versions of each image to the batch.
#
# This means the final batch size is the number of `examples per batch * (1 + the number of augmentations)`.
(aug_x, _), _ = tf.keras.datasets.mnist.load_data()
# Normalize the image data.
aug_x = tf.cast(aug_x / 255.0, dtype="float32")
# +
aug_num_examples_per_batch = 18 # @param {type:"slider", min:18, max:512}
aug_num_augmentations_per_example = 1 # @param {type:"slider", min:1, max:3}
data_augmentation = tf.keras.Sequential(
[
tf.keras.layers.experimental.preprocessing.RandomRotation(0.12),
tf.keras.layers.experimental.preprocessing.RandomZoom(0.25),
]
)
def augmenter(
x: tfsim.types.FloatTensor, y: tfsim.types.IntTensor, examples_per_class: int, is_warmup: bool, stddev=0.025
) -> Tuple[tfsim.types.FloatTensor, tfsim.types.IntTensor]:
"""Image augmentation function.
Args:
X: FloatTensor representing the example features.
y: IntTensor representing the class id. In this case
the example index will be used as the class id.
examples_per_class: The number of examples per class.
Not used here.
is_warmup: If True, the training is still in a warm
up state. Not used here.
stddev: Sets the amount of gaussian noise added to
the image.
"""
_ = examples_per_class
_ = is_warmup
aug = tf.squeeze(data_augmentation(tf.expand_dims(x, -1)))
aug = aug + tf.random.normal(tf.shape(aug), stddev=stddev)
x = tf.concat((x, aug), axis=0)
y = tf.concat((y, y), axis=0)
idxs = tf.range(start=0, limit=tf.shape(x)[0])
idxs = tf.random.shuffle(idxs)
x = tf.gather(x, idxs)
y = tf.gather(y, idxs)
return x, y
aug_sampler = tfsim.samplers.SingleShotMemorySampler(
aug_x,
augmenter=augmenter,
examples_per_batch=aug_num_examples_per_batch,
num_augmentations_per_example=aug_num_augmentations_per_example,
)
# +
# Plot the first 36 examples
num_imgs = 36
num_row = num_col = 6
aug_batch_x, aug_batch_y = aug_sampler[0]
# Sort the class ids so we can see the original
# and augmented versions as pairs.
sorted_idx = np.argsort(aug_batch_y)
plt.figure(figsize=(10, 10))
for i in range(num_imgs):
idx = sorted_idx[i]
ax = plt.subplot(num_row, num_col, i + 1)
plt.imshow(aug_batch_x[idx])
plt.title(int(aug_batch_y[idx]))
plt.axis("off")
plt.tight_layout()
# -
# ## Sampler Sizes
#
# `SingleShotMemorySampler()` provides various attributes for accessing info about the data:
# * `__len__` provides the number of steps per epoch.
# * `num_examples` provides the number of examples within the sampler.
# * `example_shape` provides the shape of the examples.
#
# The `num_examples` attribute represents the unaugmented examples within the sampler.
print(f"The sampler contains {len(aug_sampler)} steps per epoch.")
print(f"The sampler is using {aug_sampler.num_examples} examples out of the original {len(aug_x)}.")
print(f"Each examples has the following shape: {aug_sampler.example_shape}.")
# ## Accessing the Examples
#
# Additionaly, the `SingleShotMemorySampler()` provides `get_slice()` for manually accessing examples within the Sampler.
#
# The method returns slice size plus the augmented examples returned by the augmenter function.
# +
# Get 10 examples starting at example 200.
X_aug_slice, y_aug_slice = aug_sampler.get_slice(begin=200, size=10)
print("#" * 10 + " X " + "#" * 10)
print(tf.reshape(X_aug_slice, (10, -1)))
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_aug_slice)
# Check that the batch size is double our get_slice size (original examples + augmented examples).
assert tf.shape(X_aug_slice)[0] == 10 + 10
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_aug_slice)[1] == (28)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_aug_slice)[2] == (28)
# -
# <hr>
# # TFDatasetMultiShotMemorySampler: Load data from TF Dataset
# The following cell loads data directly from the TensorFlow catalog using TensorFlow similarity
# `TFDatasetMultiShotMemorySampler()`.
#
# Using a sampler is required to ensure that each batch contains at least N samples of each class incuded in a batch. Otherwise the contrastive loss does not work properly as it can't compute positive distances.
# +
IMG_SIZE = 300 # @param {type:"integer"}
# preprocessing function that resizes images to ensure all images are the same shape
def resize(img, label):
with tf.device("/cpu:0"):
img = tf.cast(img, dtype="int32")
img = tf.image.resize_with_pad(img, IMG_SIZE, IMG_SIZE)
return img, label
# +
training_classes = 16 # @param {type:"slider", min:1, max:37}
tfds_examples_per_class_per_batch = 4 # @param {type:"integer"}
tfds_class_list = random.sample(range(37), k=training_classes)
tfds_classes_per_batch = max(16, training_classes)
print(f"Class IDs seen during training {tfds_class_list}\n")
# use the train split for training
print("#" * 10 + " Train Sampler " + "#" * 10)
train_ds = tfsim.samplers.TFDatasetMultiShotMemorySampler(
"oxford_iiit_pet",
splits="train",
examples_per_class_per_batch=tfds_examples_per_class_per_batch,
classes_per_batch=tfds_classes_per_batch,
preprocess_fn=resize,
class_list=tfds_class_list,
) # We filter train data to only keep the train classes.
# use the test split for indexing and querying
print("\n" + "#" * 10 + " Test Sampler " + "#" * 10)
test_ds = tfsim.samplers.TFDatasetMultiShotMemorySampler(
"oxford_iiit_pet", splits="test", total_examples_per_class=20, classes_per_batch=tfds_classes_per_batch, preprocess_fn=resize
)
# -
# ## Generating Batches
# The Tensorflow Similarity memory samplers are a subclass of [tf.keras.utils.Sequence](https://www.tensorflow.org/api_docs/python/tf/keras/utils/Sequence), overriding the `__getitem__` and `__len__` methods.
#
# Additionally, Tensorflow Similarity provides a `generate_batch()` method that takes a batch ID and yields a single batch.
#
# We verify that the batch batch only conatins the classes defined in `CLASS_LIST` and that each class has `tfds_classes_per_batch` * `tfds_examples_per_class_per_batch` examples.
# +
X_tfds_batch, y_tfds_batch = train_ds.generate_batch(100)
print("#" * 10 + " X " + "#" * 10)
print(f"Actual Tensor Shape {X_tfds_batch.shape}")
print(tf.reshape(X_tfds_batch, (len(X_tfds_batch), -1)))
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_tfds_batch)
# Check that the batch size is equal to the target number of classes * target number of examples per class.
assert tf.shape(X_tfds_batch)[0] == (tfds_classes_per_batch * tfds_examples_per_class_per_batch)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_batch)[1] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_batch)[2] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_batch)[3] == (3)
# Check that classes in the batch are from the allowed set in CLASS_LIST
assert set(tf.unique(y_tfds_batch)[0].numpy()) - set(tfds_class_list) == set()
# Check that we only have NUM_CLASSES_PER_BATCH
assert len(tf.unique(y_tfds_batch)[0]) == tfds_classes_per_batch
# -
# ## Sampler Sizes
#
# `TFDatasetMultiShotMemorySampler()` provides various attributes for accessing info about the data:
# * `__len__` provides the number of steps per epoch.
# * `num_examples` provides the number of examples within the sampler.
# * `example_shape` provides the shape of the examples.
print(f"The Train sampler contains {len(train_ds)} steps per epoch.")
print(f"The Train sampler is using {train_ds.num_examples} examples.")
print(f"Each examples has the following shape: {train_ds.example_shape}.")
print(f"The Test sampler contains {len(test_ds)} steps per epoch.")
print(f"The Test sampler is using {test_ds.num_examples} examples.")
print(f"Each examples has the following shape: {test_ds.example_shape}.")
# ## Accessing the Examples
#
# Additionaly, the `SingleShotMemorySampler()` provides `get_slice()` for manually accessing examples within the Sampler.
#
# The method returns slice size plus the augmented examples returned by the augmenter function.
# +
# Get 10 examples starting at example 200.
X_tfds_slice, y_tfds_slice = train_ds.get_slice(begin=200, size=10)
print("#" * 10 + " X " + "#" * 10)
print(f"Actual Tensor Shape {X_tfds_slice.shape}")
print(tf.reshape(X_tfds_slice, (len(X_tfds_slice), -1)))
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_tfds_slice)
# Check that the batch size.
assert tf.shape(X_tfds_slice)[0] == 10
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[1] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[2] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[3] == (3)
# +
# Get 10 examples starting at example 200.
X_tfds_slice, y_tfds_slice = test_ds.get_slice(begin=200, size=10)
print("#" * 10 + " X " + "#" * 10)
print(f"Actual Tensor Shape {X_tfds_slice.shape}")
print(tf.reshape(X_tfds_slice, (len(X_tfds_slice), -1)))
print("\n" + "#" * 10 + " y " + "#" * 10)
print(y_tfds_slice)
# Check that the batch size.
assert tf.shape(X_tfds_slice)[0] == 10
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[1] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[2] == (300)
# Check that the number of columns matches the number of expected features.
assert tf.shape(X_tfds_slice)[3] == (3)
| examples/sampler_io_cookbook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Horn of Africa initial basic accessibility
# %load_ext autoreload
# %autoreload 2
# +
import sys, os, inspect, logging, importlib
import geopandas as gpd
import pandas as pd
import numpy as np
import osmnx as ox
import networkx as nx
from shapely.ops import split, unary_union
from shapely.geometry import box, Point
import matplotlib.pyplot as plt
# -
# Get reference to GOSTNets
sys.path.append(r'../../../../GOSTnets')
import GOSTnets as gn
from GOSTnets.load_osm import *
# ## Create road network
# Countries for this analysis include Ethiopia, Kenya, Somalia, Djibouti, South Sudan, and Sudan. Initially the OSM pbf files were downloaded from Geofabrik for these countries then merged together using the Osmium command line tool. Unfortunately this resulted in the all the roads being able to be imported. Instead the whole OSM Planet file was downloaded, then Osmium was used to extract only roads within the horn of africa spatial extent.
import time
start_time = time.time()
f = r'D:\data\planet-210614-clipped-highways2.osm.pbf'
# create OSM_to_network object from the load_osm GOSTnets sub-module
horn_of_africa = OSM_to_network(f)
print(f"sec elapsed: {time.time() - start_time}")
# show the different highway types and counts
horn_of_africa.roads_raw.infra_type.value_counts()
# ## Decided to filter the primary roads
# +
#accepted_road_types = ['tertiary','road','secondary','primary','trunk','primary_link','trunk_link','tertiary_link','secondary_link']
# +
#horn_of_africa.filterRoads(acceptedRoads = accepted_road_types)
# +
#horn_of_africa.roads_raw.infra_type.value_counts()
# -
# load_osm GOSTnets sub-module intermediate step
horn_of_africa.generateRoadsGDF(verbose = False)
print(f"sec elapsed: {time.time() - start_time}")
# load_osm GOSTnets sub-module final step, creates the graph
horn_of_africa.initialReadIn()
print(nx.info(horn_of_africa.network))
print(f"sec elapsed: {time.time() - start_time}")
# ### save the graph, this creates a pickle which can be imported layer, it also saves the nodes and the edges as CSVs, which can be opened in QGIS
gn.save(horn_of_africa.network,"horn_of_africa_unclean5_full",r"temp")
# open horn_of_africa_unclean
G = nx.read_gpickle(os.path.join(r'temp', 'horn_of_africa_unclean5_full.pickle'))
print(nx.info(G))
# +
horn_of_africa_UTMZ = {'init': 'epsg:32638'}
WGS = {'init': 'epsg:4326'} # do not adjust. OSM natively comes in ESPG 4326
# -
# ### clean network processes, including simplifying edges between intersections
# +
# print('start: %s\n' % time.ctime())
# G_clean = gn.clean_network(G, UTM = horn_of_africa_UTMZ, WGS = {'init': 'epsg:4326'}, junctdist = 10, verbose = False)
# print('\nend: %s' % time.ctime())
# print('\n--- processing complete')
# -
# let's print info on our clean version
print(nx.info(G_clean))
# save and inspect graph
gn.save(G_clean,"G_cleaned_horn_of_africa5",r"temp")
# ### Identify only the largest graph
# +
# compatible with NetworkX 2.4
list_of_subgraphs = list(G.subgraph(c).copy() for c in nx.weakly_connected_components(G))
max_graph = None
max_edges = 0
# To sort the list in place...
list_of_subgraphs.sort(key=lambda x: x.number_of_edges(), reverse=True)
# +
for subgraph in list_of_subgraphs:
print(len(subgraph))
# to inspect
#gn.save(subgraph,f"{len(subgraph)}_inspect",r"temp", pickle = False, edges = True, nodes = False)
# -
G_largest = list_of_subgraphs[0]
len(G_largest)
# +
# # compatible with NetworkX 2.4
# list_of_subgraphs = list(G.subgraph(c).copy() for c in nx.strongly_connected_components(G))
# max_graph = None
# max_edges = 0
# for i in list_of_subgraphs:
# if i.number_of_edges() > max_edges:
# max_edges = i.number_of_edges()
# max_graph = i
# # set your graph equal to the largest sub-graph
# G_largest = max_graph
# -
# print info about the largest sub-graph
print(nx.info(G_largest))
len(G_largest)
# save and inspect graph
gn.save(G_largest,"G_largest_cleaned_horn_of_africa5",r"temp")
# load graph
G = nx.read_gpickle(os.path.join(r'temp', 'G_largest_cleaned_horn_of_africa5.pickle'))
# ## insert origins
# Origins are from the GHS SMOD rural categories, and they were converted to vector points in QGIS.
origins = gpd.read_file(r'C:\Users\war-machine\Documents\world_bank_work\horn_of_africa_analysis\horn_of_africa_ghs_rural_pts_4326.shp')
origins['osmid'] = 1110000000 + origins.index
origins
# find graph utm zone
G_utm = gn.utm_of_graph(G)
G_utm
# ### snap origins to the graph (snaps to the nearest node on the graph)
# %%time
#no need to do advanced_snap at this extent
G2, pois_meter, new_footway_edges = gn.advanced_snap(G, origins, u_tag = 'stnode', v_tag = 'endnode', node_key_col='node_ID', poi_key_col='osmid', path=None, threshold=2000, measure_crs=G_utm, factor=1000)
# +
# #%%time
#no need to do advanced_snap at this extent
#G2, pois_meter, new_footway_edges = gn.advanced_snap(G, origins, u_tag = 'stnode', v_tag = 'endnode', node_key_col='node_ID', poi_key_col='osmid', path=None, threshold=2000, measure_crs=G_utm)
#snapped_origins = gn.pandana_snap(G, origins, source_crs = 'epsg:4326', target_crs = G_utm)
# -
new_footway_edges
new_footway_edges
# ## Snap destinations to the road graph
# Destinations are centroids created in QGIS of the GHS urban centers
# insert destinations
destinations = gpd.read_file(r'C:\Users\war-machine\Documents\world_bank_work\horn_of_africa_analysis\horn_of_africa_ghs_stat_centroids2.shp')
destinations
snapped_destinations = gn.pandana_snap(G, destinations, source_crs = 'epsg:4326', target_crs = G_utm)
snapped_destinations
destinationNodes = list(snapped_destinations['NN'].unique())
destinationNodes
import pickle
with open("destinationNodes_horn_of_africa_6.txt", "wb") as fp: #Pickling
pickle.dump(destinationNodes, fp)
gn.example_edge(G2)
# ### add time to the graph edges as an attribute
# +
# for u, v, data in G2.edges(data=True):
# orig_len = data[distance_tag] * factor
# # Note that this is a MultiDiGraph so there could
# # be multiple indices here, I naively assume this is not
# # the case
# data['length'] = orig_len
# # get appropriate speed limit
# if graph_type == 'walk':
# -
# %%time
# note that the length above is in km, therefore set factor to 1000
G2_time = gn.convert_network_to_time(G2, 'length', road_col = 'infra_type', factor = 1000)
# +
## Note to improve: Pandana advanced snap calculates the length for new lines, but it is in meters, and also Euclidean distance distance. Fix this?
# -
gn.example_edge(G2)
# ## calculate OD matrix
print(nx.info(G2))
print(nx.info(G))
# save and inspect graph
gn.save(G2,"G_largest_horn_of_africa6_adv_snap",r"temp")
pois_meter
pois_meter2 = pois_meter[['osmid','VALUE','geometry']]
pois_meter2
G2.nodes[1110000001]
G2.nodes[1110000004]
nx.shortest_path(G2,1110000001,1110000004)
nx.shortest_path_length(G2,1110000001,1110000004, weight='length')
destinationNodes
G2.nodes[2469733]
nx.shortest_path_length(G2,1110000001,2469733, weight='length')
# save new_nodes
pois_meter2.to_csv(r"temp/clipped_origin_nodes_horn_of_africa6.csv")
pois_meter2['osmid'] = pd.to_numeric(pois_meter2['osmid'])
pois_meter2['VALUE'] = pd.to_numeric(pois_meter2['VALUE'])
originNodes_list = list(pois_meter2['osmid'])
originNodes_list
# +
#print(f"sec elapsed: {time.time() - start_time}")
# -
# %%time
OD_matrix = gn.calculate_OD(G2, originNodes_list, destinationNodes, fail_value=-1, weight='length')
OD_matrix
# ## calculate accessibility
# ### For each row, the closest facility is the smallest value in the row
closest_facility_per_origin = OD_matrix.min(axis=1)
results = pd.DataFrame([originNodes_list, closest_facility_per_origin]).transpose()
colName = "travel_time_to_closest_facility"
results.columns = ['osmid', colName]
results[:5]
output2 = pd.merge(pois_meter2, results, on="osmid")
output2
output2[output2['travel_time_to_closest_facility']==-1]
output2[output2['travel_time_to_closest_facility']>0]
# +
#snapped_origins
# +
#output2 = pd.merge(snapped_origins, results, on="NN")
# +
#output2[:5]
# +
#output2['travel_time_to_closest_facility'] = output2['travel_time_to_closest_facility'].astype('int64')
# +
#output2.dtypes
# -
# ## save a shapefile..
# ### Then in QGIS it can be opened and symbolized based on choosing a colored ramp based on the travel_time_to_closest_facility attribute
destinations_gpd = gpd.GeoDataFrame(output2, crs = "epsg:4326", geometry = 'geometry')
destinations_gpd.to_file('horn_of_africa_rural_accessibility6.shp')
| Implementations/FY22/ACC_horn_of_africa_accessibility/horn_of_africa_accessibility_analysis2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Scrape crowsourced labels of Exchange Ethereum addresses
import requests
from bs4 import BeautifulSoup
import time
def find_tag(address):
url = 'https://etherscan.io/address/{}'.format(address)
html = requests.get(url).content
soup = BeautifulSoup(html, 'html.parser')
#find tag
tag = soup.find('font', title='Public Name Tag (viewable by anyone)')
if type(tag) == type(None):
t = 'unknown'
else:
t = tag.decode_contents()
#find category
tag = soup.find('a', href="/accounts?l=Exchange")
if type(tag) == type(None):
c = 'unknown'
else:
c = tag.decode_contents()
time.sleep(2)
return t,c
find_tag("0x05f51AAb068CAa6Ab7eeb672f88c180f67F17eC7")
def find_label(address):
url = 'https://etherscan.io/address/{}'.format(address)
html = requests.get(url).content
soup = BeautifulSoup(html, 'html.parser')
#find tag
if soup.find('a',{'class':'mb-1 mb-sm-0 u-label u-label--xs u-label--secondary'}):
print(ok)
label = soup.find('a',{'class':'mb-1 mb-sm-0 u-label u-label--xs u-label--secondary'}).attrs['href']
return label
else:
return "unknown"
find_label("0x05f51AAb068CAa6Ab7eeb672f88c180f67F17eC7")
find_label(0x05f51AAb068CAa6Ab7eeb672f88c180f67F17eC7)
<a class="mb-1 mb-sm-0 u-label u-label--xs u-label--secondary" href="/accounts/label/exchange">Exchange</a>
def grab_all():
#get top 10k addresses with labels
res = []
for i in range(1,10):
url = 'https://etherscan.io/accounts/{}?ps=100'.format(i)
html = requests.get(url).content
soup = BeautifulSoup(html, 'html.parser')
table = soup.find('tbody')
table_rows = table.find_all('tr')
for tr in table_rows:
td = tr.find_all('td')
row = [td[2].text.strip(),td[3].text.strip()]
if row:
res.append(row)
df = pd.DataFrame(res, columns=['address', 'label'])
return df
grab_all()
def grab():
#get top 10k addresses with labels
url = 'https://etherscan.io/accounts/label/exchange?subcatid=undefined&size=500&start=1&col=1&order=asc'
html = requests.get(url).content
soup = BeautifulSoup(html, 'html.parser')
print(soup.get_text())
table = soup.find('tbody')
table_rows = table.find_all('tr')
for tr in table_rows:
td = tr.find_all('td')
row = [td[2].text.strip(),td[3].text.strip()]
if row:
res.append(row)
df = pd.DataFrame(res, columns=['address', 'label'])
return df
grab()
https://etherscan.io/accounts?l=Exchange
| notebooks/Drafts/scrape_labels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow-GPU
# language: python
# name: tf-gpu
# ---
# ## Instagram API Access
# +
CLIENT_ID = 'e06ea50d76524d9aa66e758b2e1b1a10'
CLIENT_SECRET = 'f240ee2ed3d049058487f5eb83b69bea'
REDIRECT_URI = 'http://anagno.com/'
base_url = 'https://api.instagram.com/oauth/authorize/'
url = '{0}?client_id={1}&redirect_uri={2}&response_type=code&scope=public_content'.format(base_url, CLIENT_ID, REDIRECT_URI)
print('Click the following URL, which will taken you to the REDIRECT_URI you set in creating the APP.')
print('You may need to log into Instagram.')
print()
print(url)
# +
import requests # pip install requests
CODE = '78be864b693146ed89d35667128f9ae6'
payload = dict(client_id=CLIENT_ID,
client_secret=CLIENT_SECRET,
grant_type='authorization_code',
redirect_uri=REDIRECT_URI,
code=CODE)
response = requests.post(
'https://api.instagram.com/oauth/access_token',
data = payload)
# -
response.json()
ACCESS_TOKEN = response.json()['access_token']
ACCESS_TOKEN
# +
response = requests.get('https://api.instagram.com/v1/users/self/?access_token='+ACCESS_TOKEN)
print(response.text)
# -
# ## Retrieving your Feed
# +
from IPython.display import display, Image
response = requests.get('https://api.instagram.com/v1/users/self/media/recent/?access_token='+ACCESS_TOKEN)
recent_posts = response.json()
def display_image_feed(feed, include_captions=True):
for post in feed['data']:
display(Image(url=post['images']['standard_resolution']['url']))
print(post['images']['standard_resolution']['url'])
if include_captions: print(post['caption']['text'])
print()
# -
# ## Anatomy of an Instagram Post
# +
import json
response = requests.get('https://api.instagram.com/v1/users/self/media/recent/?access_token='+ACCESS_TOKEN)
recent_posts = response.json()
#print(json.dumps(recent_posts, indent=1))
# -
print(recent_posts.keys())
print(recent_posts['pagination'])
print(recent_posts['meta'])
# +
#print(json.dumps(recent_posts['data'], indent=1))
# -
print(json.dumps(recent_posts['data'][0], indent=1))
# ## Artificial Neural Networks
# +
from sklearn import datasets, metrics
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split
digits = datasets.load_digits()
# Rescale the data and split into training and test sets
X, y = digits.data / 255., digits.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
mlp = MLPClassifier(hidden_layer_sizes=(100,), max_iter=100, alpha=1e-4,
solver='adam', verbose=10, tol=1e-4, random_state=1,
learning_rate_init=.1)
mlp.fit(X_train, y_train)
print()
print("Training set score: {0}".format(mlp.score(X_train, y_train)))
print("Test set score: {0}".format(mlp.score(X_test, y_test)))
# +
import matplotlib.pyplot as plt
# %matplotlib inline
fig, axes = plt.subplots(10,10)
fig.set_figwidth(20)
fig.set_figheight(20)
for coef, ax in zip(mlp.coefs_[0].T, axes.ravel()):
ax.matshow(coef.reshape(8, 8), cmap=plt.cm.gray, interpolation='bicubic')
ax.set_xticks(())
ax.set_yticks(())
plt.show()
# +
import numpy as np # pip install numpy
predicted = mlp.predict(X_test)
for i in range(5):
image = np.reshape(X_test[i], (8,8))
plt.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
plt.axis('off')
plt.show()
print('Ground Truth: {0}'.format(y_test[i]))
print('Predicted: {0}'.format(predicted[i]))
# -
| Instagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import mayavi as maya
from mayavi import mlab
from functools import partial
mlab.init_notebook('x3d',800,800)
# +
MZ = 12
MT = 12
def read_w7x(filename):
R = np.zeros((25, 12))
Z = np.zeros((25, 12))
with open(filename) as f:
_ = f.readline()
_, Nfp, _ = f.readline().split()
Nfp = int(Nfp)
_ = f.readline()
_ = f.readline()
for i in range(288):
n, m, rc, _, _, zs = f.readline().split()
n = int(n)
m = int(m)
rc = float(rc)
zs = float(zs)
R[n + 12, m] = rc
Z[n + 12, m] = zs
return R, Z, Nfp
def get_xyz(R, Z, zeta, theta, Nfp):
r = np.zeros((zeta.shape[0], theta.shape[0]))
z = np.zeros((zeta.shape[0], theta.shape[0]))
for mz in range(-MZ, MZ + 1):
for mt in range(MT):
r += R[mz + MZ, mt] * np.cos( mt * theta[np.newaxis, :] - mz * Nfp * zeta[:, np.newaxis] )
z += Z[mz + MZ, mt] * np.sin( mt * theta[np.newaxis, :] - mz * Nfp * zeta[:, np.newaxis] )
x = r * np.cos(zeta)[:, np.newaxis]
y = r * np.sin(zeta)[:, np.newaxis]
return np.concatenate((x[:, :, np.newaxis], y[:, :, np.newaxis], z[:, :, np.newaxis]), axis = -1)
def plot_surface(R, Z, NZ, NT, Nfp):
zeta = np.linspace(0,2 * np.pi, NZ + 1)
theta = np.linspace(0, 2 * np.pi, NT + 1)
r = get_xyz(R, Z, zeta, theta, Nfp)
x = r[:,:,0]
y = r[:,:,1]
z = r[:,:,2]
p = mlab.mesh(x,y,z,color=(0.8,0.0,0.0))
return p
def compute_drdz(R, Z, zeta, theta, Nfp):
x = np.zeros((zeta.shape[0], theta.shape[0]))
y = np.zeros((zeta.shape[0], theta.shape[0]))
z = np.zeros((zeta.shape[0], theta.shape[0]))
for mz in range(-MZ, MZ + 1):
for mt in range(MT):
coeff = R[mz + MZ, mt]
z_coeff = Z[mz + MZ, mt]
arg = mt * theta[np.newaxis, :] - mz * Nfp * zeta[:, np.newaxis]
x += coeff * np.cos(arg) * (-np.sin(zeta[:, np.newaxis])) + coeff * (mz * Nfp) * np.cos(zeta[:, np.newaxis]) * np.sin(arg)
y += coeff * np.cos(arg) * np.cos(zeta[:, np.newaxis]) + coeff * (mz * Nfp) * np.sin(arg) * np.sin(zeta[:, np.newaxis])
z += z_coeff * np.cos(arg) * -(mz * Nfp)
return np.concatenate((x[:, :, np.newaxis], y[:, :, np.newaxis], z[:, :, np.newaxis]), axis = -1)
def compute_drdt(R, Z, zeta, theta, Nfp):
x = np.zeros((zeta.shape[0], theta.shape[0]))
y = np.zeros((zeta.shape[0], theta.shape[0]))
z = np.zeros((zeta.shape[0], theta.shape[0]))
for mz in range(-MZ, MZ + 1):
for mt in range(MT):
coeff = R[mz + MZ, mt]
z_coeff = Z[mz + MZ, mt]
arg = mt * theta[np.newaxis, :] - mz * Nfp * zeta[:, np.newaxis]
x += coeff * np.sin(arg) * -mt * np.cos(zeta[:, np.newaxis])
y += coeff * np.sin(arg) * -mt * np.sin(zeta[:, np.newaxis])
z += z_coeff * np.cos(arg) * mt
return np.concatenate((x[:, :, np.newaxis], y[:, :, np.newaxis], z[:, :, np.newaxis]), axis = -1)
def get_w7x_data(R, Z, NZ, NT, Nfp):
zeta = np.linspace(0,2 * np.pi, NZ + 1)
theta = np.linspace(0, 2 * np.pi, NT + 1)
r = get_xyz(R, Z, zeta, theta, Nfp)
drdz = compute_drdz(R, Z, zeta, theta, Nfp)
drdt = compute_drdt(R, Z, zeta, theta, Nfp)
N = np.cross(drdz, drdt)
sg = np.linalg.norm(N, axis=-1)
nn = N / sg[:,:,np.newaxis]
return r, nn, sg
def plot_coils(r_coils):
for ic in range(r_coils.shape[0]):
p = mlab.plot3d(r_coils[ic,:,0], r_coils[ic,:,1], r_coils[ic,:,2], tube_radius=0.04, color = (0.0, 0.0, 0.8))#, line_width = 0.01,)
return p
# -
R, Z, Nfp = read_w7x("../../focusadd/initFiles/w7x/plasma.boundary_std")
mlab.clf()
p = plot_surface(R, Z, 128, 32, Nfp)
p
r, nn, sg = get_w7x_data(R, Z, 5 * 30, 32, Nfp)
mlab.clf()
x = r[:,:,0]
y = r[:,:,1]
z = r[:,:,2]
p = mlab.mesh(x,y,z,color=(0.8,0.0,0.0))
nn_x = nn[:,:,0]
nn_y = nn[:,:,1]
nn_z = nn[:,:,2]
s = mlab.quiver3d(x[::,::],y[::,::],z[::,::],sg*nn_x[::,::],sg*nn_y[::,::],sg*nn_z[::,::])
s
def read_coils(filename):
r = np.zeros((50, 96, 3))
with open(filename) as f:
_ = f.readline()
_ = f.readline()
_ = f.readline()
for i in range(50):
for s in range(96):
x, y, z, _ = f.readline().split()
r[i, s, 0] = float(x)
r[i, s, 1] = float(y)
r[i, s, 2] = float(z)
_ = f.readline()
return r
r_c = read_coils("../../focusadd/initFiles/w7x/coils.std_mc")
p = plot_coils(r_c)
p
def compute_coil_fourierSeries(NF, r_centroid):
NC = r_centroid.shape[0]
NS = r_centroid.shape[1]
x = r_centroid[:, :, 0]
y = r_centroid[:, :, 1]
z = r_centroid[:, :, 2]
xc = np.zeros((NC, NF))
yc = np.zeros((NC, NF))
zc = np.zeros((NC, NF))
xs = np.zeros((NC, NF))
ys = np.zeros((NC, NF))
zs = np.zeros((NC, NF))
xc[:,0] += np.sum(x, axis=1) / NS
yc[:,0] += np.sum(y, axis=1) / NS
zc[:,0] += np.sum(z, axis=1) / NS
theta = np.linspace(0, 2 * np.pi, NS + 1)[0:NS]
for m in range(1, NF):
xc[:, m] += 2.0 * np.sum(x * np.cos(m * theta), axis=1) / NS
yc[:, m] += 2.0 * np.sum(y * np.cos(m * theta), axis=1) / NS
zc[:, m] += 2.0 * np.sum(z * np.cos(m * theta), axis=1) / NS
xs[:, m] += 2.0 * np.sum(x * np.sin(m * theta), axis=1) / NS
ys[:, m] += 2.0 * np.sum(y * np.sin(m * theta), axis=1) / NS
zs[:, m] += 2.0 * np.sum(z * np.sin(m * theta), axis=1) / NS
return np.asarray([xc, yc, zc, xs, ys, zs]) # 6 x NC x NF
fc = compute_coil_fourierSeries(10, r_c)
def compute_r_centroid(NS, fc):
theta = np.linspace(0, 2 * np.pi, NS + 1)
NC = fc.shape[1]
NF = fc.shape[2]
xc = fc[0]
yc = fc[1]
zc = fc[2]
xs = fc[3]
ys = fc[4]
zs = fc[5]
x = np.zeros((NC, NS + 1))
y = np.zeros((NC, NS + 1))
z = np.zeros((NC, NS + 1))
for m in range(NF):
arg = m * theta
carg = np.cos(arg)
sarg = np.sin(arg)
x += (
xc[:, np.newaxis, m] * carg[np.newaxis, :]
+ xs[:, np.newaxis, m] * sarg[np.newaxis, :]
)
y += (
yc[:, np.newaxis, m] * carg[np.newaxis, :]
+ ys[:, np.newaxis, m] * sarg[np.newaxis, :]
)
z += (
zc[:, np.newaxis, m] * carg[np.newaxis, :]
+ zs[:, np.newaxis, m] * sarg[np.newaxis, :]
)
return np.concatenate(
(x[:, :, np.newaxis], y[:, :, np.newaxis], z[:, :, np.newaxis]), axis=2
)
r_c_2 = compute_r_centroid(96, fc)
mlab.clf()
p = plot_surface(R, Z, 128, 32, Nfp)
p = plot_coils(r_c_2)
p
with open('w7x_r_surf.npy', 'wb') as f:
np.save(f, r)
with open('w7x_nn_surf.npy', 'wb') as f:
np.save(f, nn)
with open('w7x_sg_surf.npy', 'wb') as f:
np.save(f, sg)
with open('w7x_fc.npy', 'wb') as f:
np.save(f, fc)
| plotting/w7x/.ipynb_checkpoints/read_w7x-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="x61SCwI468MZ" outputId="8c928db4-22ce-4091-e788-ed9cb9650736"
import pandas as pd
d_path = "a.txt"
with open(d_path, 'r') as reader:
a = reader.readlines()
no_integers = [x for x in a[1:] if not isinstance(x, int)]
no_integers
# + [markdown] id="VkeSHFhm4xov"
# # New Section
# + colab={"base_uri": "https://localhost:8080/"} id="V8KkjqSQG240" outputId="4922447c-42e9-42f0-d816-7960492c5b82"
d_path = "a.txt"
d_path=open(d_path, 'r')
u = d_path.read().replace('\n','\n')
u=u.split()
u
# + colab={"base_uri": "https://localhost:8080/"} id="VoHXtpuYG243" outputId="f544d750-074a-4dd3-896f-888155064a0f"
ing=[]
for x in u:
if not x.isdigit():
ing.append(x)
ing
# + colab={"base_uri": "https://localhost:8080/"} id="vHaG-nLi7Kq_" outputId="4aafaff6-ce41-43c4-fc0a-653a30696d6d"
digits = [int(elem) for elem in u[4:] if elem.isdigit()]
print(digits)
new2=[]
y=0
for x in digits:
new=[]
for i in range(0,x):
new.append(ing[y])
y=y+1
new2.append(new)
new2
# + colab={"base_uri": "https://localhost:8080/"} id="FWiwFam07OO5" outputId="47071b35-b87d-4dd2-950e-098d38ab28bb"
pizzas_num = int(u[0])
print(pizzas_num)
# + colab={"base_uri": "https://localhost:8080/"} id="pqpLTAAv7OVG" outputId="2a7b1f1b-5521-4d86-e4fc-ad67223d7e21"
for x in new2:
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="R6JU0Qy-7OcQ" outputId="00140fe3-3a7e-4c03-92df-9d53645932b3"
dic={pizza:new2[pizza] for pizza in range(0,pizzas_num)}
dic
# + colab={"base_uri": "https://localhost:8080/"} id="1jUXqpA0JF_v" outputId="6a5e847b-3dbe-4d92-f882-776473e7b7c5"
dic[0]
# + colab={"base_uri": "https://localhost:8080/"} id="PyIN4WKCG244" outputId="37f9fe75-1d43-4c2b-c20e-50980987271a"
t2=int(u[1])
t3=int(u[2])
t4=int(u[3])
type(int(t4))
# + id="fUVVGqlEG245"
# Function to create combinations
# without itertools
def combinations_fnc(lst, n):
if n == 0:
return [[]]
l =[]
for i in range(0, len(lst)):
m = lst[i]
remLst = lst[i + 1:]
for p in combinations_fnc(remLst, n-1):
l.append([m]+p)
return l
# + id="Nfre-fnSZB6D"
import itertools
# + colab={"base_uri": "https://localhost:8080/"} id="t2obKrs9G246" outputId="19f8b3fb-8417-4def-efdd-4e6013b9d938"
pizzas=list(range(0,pizzas_num))
print(pizzas)
comb_t2=combinations_fnc(pizzas,2)
comb_t3=combinations_fnc(pizzas,3)
comb_t4=combinations_fnc(pizzas,4)
#comb_t2=[item for item in itertools.combinations(pizzas,2)]
#comb_t3=[item for item in itertools.combinations(pizzas,3)]
#comb_t4=[item for item in itertools.combinations(pizzas,4)]
# + id="dYA3BU0wbj1a"
import itertools
from itertools import combinations
test_list=range(0,500)
i,j=2,4
res=[]
for sub in range(j):
if sub >= (i-1):
res.extend(combinations(test_list,sub+1))
print(res)
# + colab={"base_uri": "https://localhost:8080/"} id="mmQGxqOoG248" outputId="42db1637-6af8-46d9-b876-5c4310fdbcf4"
total=comb_t2 + comb_t3 + comb_t4
total
# + id="sj57PkWZG249"
def allUnique(x):
seen = list()
return not any(i in seen or seen.append(i) for i in x)
# + id="ukS279XeQGeY"
from itertools import combinations
# + id="CAq9ANzdJSFO"
#len([item for item in itertools.combinations(total,2)])
# + id="DCIWVSHDG25A"
m=[]
for i,j in [item for item in itertools.combinations(total,2)]:
if allUnique(i+j)==True:
m.append([i,j])
# + colab={"base_uri": "https://localhost:8080/"} id="_RLwVLtoTG_u" outputId="3ce96080-cef3-4efc-d120-23963979d826"
m
# + id="eSy0Cb6GTJFB"
n=[]
for i,j in m:
if(len(i)+len(j)<=pizzas_num)==True:
n.append([i,j])
# + colab={"base_uri": "https://localhost:8080/"} id="uCBS_ZkLVFr2" outputId="e2d6de08-51e4-458b-9f67-c9af9295f7f2"
n
# + id="uMap04MvVJi1" colab={"base_uri": "https://localhost:8080/"} outputId="2ad3deb4-4fff-445e-f48e-9e3471d0c67c"
#all unique possible combinations taking
for i,j in n:
print(i,j)
# + colab={"base_uri": "https://localhost:8080/"} id="PN0QlOSPJkQ7" outputId="4e04b9fd-f566-4b72-f88c-7e3b8615a895"
temp2=[]
z=[]
for i,j in n:
temp=[]
z=[]
for x in i:
z=z+dic[x]
temp.append(z)
temp2.append(z)
temp2
# + colab={"base_uri": "https://localhost:8080/"} id="8pgJnd8tJm9o" outputId="92244abf-52b5-4598-f169-0e7e52d9afb5"
import numpy as np
words=['onion', 'pepper', 'olive', 'chicken', 'mushroom', 'pepper']
len(np.unique(words))
# + colab={"base_uri": "https://localhost:8080/"} id="O-bYofHGJqJp" outputId="d0ced942-a743-4d93-bc21-9ffeaa9569ec"
for x in temp2:
print(len(np.unique(x)))
# + colab={"base_uri": "https://localhost:8080/"} id="naLF4vk5J3m6" outputId="db67c877-22ea-4a7d-fc99-0aede7e34559"
col1=[]
for x in temp2:
col1.append(len(np.unique(x)))
len(col1)
# + colab={"base_uri": "https://localhost:8080/"} id="HV8J_eSI4jh2" outputId="8ded9602-0a05-4a1e-ac91-98c9efe491d0"
temp3=[]
z=[]
for i,j in n:
temp=[]
z=[]
for x in j:
z=z+dic[x]
temp.append(z)
temp3.append(z)
temp3
# + colab={"base_uri": "https://localhost:8080/"} id="r77Je6Q84jsM" outputId="56d63bb8-de90-498d-a7d5-4b4d515450b5"
col2=[]
for x in temp3:
col2.append(len(np.unique(x)))
(col2)
# + colab={"base_uri": "https://localhost:8080/"} id="WijNXrTY4jvY" outputId="2383e734-108a-4009-eee4-f9bd2708f3bd"
final_list = []
# Choose the smaller list to iterate
list_to_iterate = len(col1) < len(col2) and col1 or col2
for i in range(0, len(list_to_iterate)):
final_list.append(pow(col1[i],2) +pow(col2[i],2))
final_list
#list_to_iterate
# + colab={"base_uri": "https://localhost:8080/"} id="bXlyDDI_4jza" outputId="287c60d2-56fd-477d-affd-3d971a309f43"
#indices of the max value
m = max(final_list)
indices=[i for i, j in enumerate(final_list) if j == m]
indices
# + colab={"base_uri": "https://localhost:8080/"} id="KwSWxKl54j3G" outputId="e55e3565-6d56-461d-fea8-6b7d56bace45"
x=[]
for i in indices:
x.append(n[i])
print(x[0])
# + colab={"base_uri": "https://localhost:8080/"} id="3TWkft4dZbHH" outputId="35b69fc1-4a34-4b34-f87b-88efbc2cd979"
print(len(x[0][0]))
# + id="orhSYYx-JCZ6" colab={"base_uri": "https://localhost:8080/"} outputId="95b275f7-4044-4599-849f-ed88b8900db4"
print(len(x[0]))
for i in x[0]:
print(len(i),end=" ")
for j in i:
print(j, end=" ")
print()
# + id="3BuwhZIIXksR"
| Google Hashcode 2021/Google_Hash_Code_Solution_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.4 64-bit
# name: python3
# ---
# +
import sys
import os
from json import dump, load
sys.path.append(os.path.abspath(os.path.join('..')))
from scripts.logger_creator import CreateLogger
# -
# Initializing Logger
logger = CreateLogger('AlphabetsBuilder', handlers=1)
logger = logger.get_default_logger()
class AlphabetsBuilder():
def __init__(self,file_name: str, alphabets_type: int = 2, train_labels: str = '../data/train_labels.json', test_labels: str = '../data/test_labels.json') -> None:
try:
self.file_name = file_name
self.alphabets_type = alphabets_type
self.train_labels = train_labels
self.test_labels = test_labels
self.alphabets_data = {}
logger.info('Successfully Created Alphabets Builder Class Object')
except Exception as e:
logger.exception("Failed to create Alphabets Builder Class Object")
def get_supported_alphabets(self):
try:
# Method 1
# Conside the entire Amharic Alphabets
if(self.alphabets_type == 1):
# Defining Entire Amharic Alphabets
self.supported_alphabets = """
ሀ ሁ ሂ ሃ ሄ ህ ሆ ለ ሉ ሊ ላ ሌ ል ሎ ሏ ሐ ሑ ሒ ሓ ሔ ሕ ሖ ሗ መ ሙ ሚ ማ ሜ ም ሞ ሟ ሠ ሡ ሢ ሣ ሤ ሥ ሦ ሧ
ረ ሩ ሪ ራ ሬ ር ሮ ሯ ሰ ሱ ሲ ሳ ሴ ስ ሶ ሷ ሸ ሹ ሺ ሻ ሼ ሽ ሾ ሿ ቀ ቁ ቂ ቃ ቄ ቅ ቆ ቇ ቋ ቐ ቐ ቑ ቒ ቓ ቔ ቕ ቖ
በ ቡ ቢ ባ ቤ ብ ቦ ቧ ቨ ቩ ቪ ቫ ቬ ቭ ቮ ቯ ተ ቱ ቲ ታ ቴ ት ቶ ቷ ቸ ቹ ቺ ቻ ቼ ች ቾ ቿ ኀ ኁ ኂ ኃ ኄ ኅ ኆ ኇ ኋ
ነ ኑ ኒ ና ኔ ን ጓ ኖ ኗ ኘ ኙ ኚ ኛ ኜ ኝ ኞ ኟ አ ኡ ኢ ኣ ኤ እ ኦ ኧ ከ ኩ ኪ ካ ኬ ክ ኮ ኯ ኰ ኳ ኲ
ኸ ኹ ኺ ኻ ኼ ኽ ኾ ወ ዉ ዊ ዋ ዌ ው ዎ ዐ ዑ ዒ ዓ ዔ ዕ ዖ ዘ ዙ ዚ ዛ ዜ ዝ ዞ ዟ ዠ ዡ ዢ ዣ ዤ ዥ ዦ ዧ
የ ዩ ዪ ያ ዬ ይ ዮ ዯ ደ ዱ ዲ ዳ ዴ ድ ዶ ዷ ጀ ጁ ጂ ጃ ጄ ጅ ጆ ጇ ገ ጉ ጊ ጋ ጌ ግ ጐ ጎ ጏ ጔ ጠ ጡ ጢ ጣ ጤ ጥ ጦ ጧ ጨ ጩ ጪ ጫ ጬ ጭ ጮ ጯ
ጰ ጱ ጲ ጳ ጴ ጵ ጶ ጷ ጸ ጹ ጺ ጻ ጼ ጽ ጾ ጿ ፀ ፁ ፂ ፃ ፄ ፅ ፆ ፇ ፈ ፉ ፊ ፋ ፌ ፍ ፎ ፏ ፐ ፑ ፒ ፓ ፔ ፕ ፖ ፗ
""".split()
# Adding space
self.supported_alphabets.insert(0, '<space>')
logger.info('Successfully retrieved alphabets from the entire Amharic Language')
else:
# Method 2
# Conside Characters only from the train and test transcriptions
# Reading Train Labels
with open(self.train_labels, 'r', encoding='UTF-8') as label_file:
train_labels = load(label_file)
# Reading Test Labels
with open(self.test_labels, 'r', encoding='UTF-8') as label_file:
test_labels = load(label_file)
# Creating an Alphabet Character Set
char_set = set()
# Reading from each Labels to extract alphabets
# Extracting from Train Labels
for label in train_labels.values():
characters = [char for char in label]
char_set.update(characters)
# Extracting from Test Labels
for label in test_labels.values():
characters = [char for char in label]
char_set.update(characters)
# Creating Alphabets List
self.supported_alphabets = list(char_set)
# Removing Space and Inserting as <space>
self.supported_alphabets.remove(' ')
self.supported_alphabets.insert(0, '<space>')
logger.info('Successfully retrieved alphabets from train and test transcriptions')
except Exception as e:
logger.exception('Failed To retrieve supported alphabets')
def construct_conversion_dicts(self):
try:
# Constructing Alphabet to num conversion dict
alphabet_to_num = {}
index = 0
# Iterating through alphabets and appending to the conversion dictionary
for alphabet in self.supported_alphabets:
alphabet_to_num[alphabet] = index
index += 1
# Constructing Alphabet to num conversion dict
# Iterating through alphabets to num dictionary to create the reverse
num_to_alphabet = {v: k for k, v in alphabet_to_num.items()}
self.alphabets_data['char_to_num'] = alphabet_to_num
self.alphabets_data['num_to_char'] = num_to_alphabet
self.alphabets_data['alphabet_size'] = len(self.supported_alphabets)
logger.info('Successfully constructed conversion dictionaries')
except Exception as e:
logger.exception('Failed to construct conversion dictionaries')
def save_alphabets_dict(self):
try:
with open(self.file_name, "w", encoding='UTF-8') as export_file:
dump(self.alphabets_data, export_file, indent=4, sort_keys=True, ensure_ascii=False)
logger.info(f'Successfuly Saved Generated Alphabets Dictionary in: {self.file_name}')
except Exception as e:
logger.exception('Failed to Save Generated Alphabets Dictionary')
def generate_and_save_alphabets(self):
self.get_supported_alphabets()
self.construct_conversion_dicts()
self.save_alphabets_dict()
alphabet_builder = AlphabetsBuilder('../data/alphabets_data.json')
alphabet_builder.generate_and_save_alphabets()
| notebooks/AlphabetsGenerator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from drawui import setup
import ipywidgets as widgets
import pandas as pd
filename = "ranges.xlsx"
df = pd.read_excel(filename,index_col=0)
levers = {var[0] : widgets.FloatSlider(description = var[1]["name"],
value = int((var[1]["maximum"]-var[1]["minimum"])/(2*var[1]["step"]))*var[1]["step"] + var[1]["minimum"],
min = var[1]["minimum"], max = var[1]["maximum"],
step=var[1]["step"]) \
for var in df.iterrows()}
display(setup(levers))
| draw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime
import pandas as pd
import spacy
import re
import string
from spacy.tokens import Token
from tqdm import tqdm
from textblob import TextBlob
from textblob import Word
import html
# -
#### Importing the file ####
Path="src/"
Filename='projects_TranslateV2.0.csv'
Cat_File="category_hier.csv"
df=pd.read_csv(Path+Filename)
Cat_data=pd.read_csv(Path+Cat_File)
#Filtering the empty abstracts.
df=df[df["Description"].str.strip()!="No abstract available"]
df['Translates'].head()
df["Translates"]=df["Translates"].apply(str).apply(html.unescape)
# +
nlp = spacy.load("en_core_web_sm")
ManualStopWords=['e.g.','i.e.','etc.','et al.','ibid.']
def remove_spell_errors(doc):
bdoc = TextBlob(str(doc))
## Correcting the words
return str(bdoc.correct())
def stop(doc):
#return [token for token in doc if not token.text in ManualStopWords and len(token.text)>1 and not token.is_digit and not token.is_stop and ( token.text.isalpha() or not token.text.isalnum())]
return [token for token in doc if not token.text in ManualStopWords and len(token.text)>1 and not token.is_digit and not token.is_stop ]
def lemmatize(doc):
return [token.lemma_.lower() if token.lemma_ != "-PRON-" else token.text.lower() for token in doc]
def remove_line_breaks(doc):
return [token.replace("\n", " ").replace("\r", " ") for token in doc]
nlp.add_pipe(stop)
nlp.add_pipe(lemmatize)
nlp.add_pipe(remove_line_breaks)
# +
print(str(datetime.datetime.now())+" : Started preprocessing")
docs = df['Translates'].apply(str).to_list()
processed_docs = []
with tqdm(total=len(docs)) as bar:
for doc in nlp.pipe(docs):
line = " ".join(doc)
## Removing the punctuation
line=line.translate(str.maketrans('','',string.punctuation))
## Removing numbers
line=" ".join(list(filter(lambda w : not w.isdigit(), line.split())))
processed_docs.append(line)
bar.update(1)
df["PreProcessedDescription"] = processed_docs
print(str(datetime.datetime.now())+" : Preprocessing completed")
# -
df.to_csv(Path+'projects_Preprocessed.csv', index=False)
| 2.2.PreProcessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HWIxg0g3Rtrj"
# #Multi-objective Optimization Test Case - Kursawe Function
# In this is example we will be solving the [Kursawe function](https://drive.google.com/file/d/1yiCQysLgRxc-MdeCbOTuMGkl3hadgS4f/view?usp=sharing) using NSGA 3. Kursawe is a popular test function used to test different multi-objective optimizations
#
# This is what the solution should look like 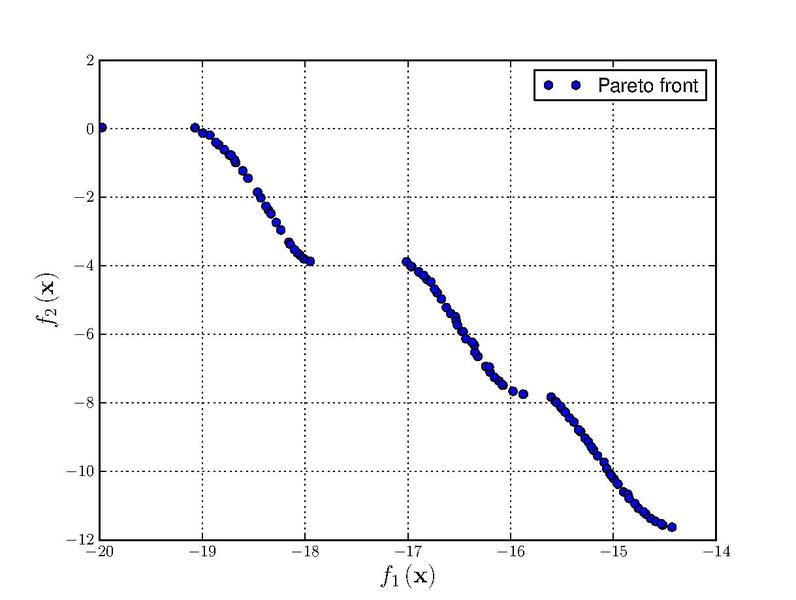
#
# + [markdown] id="kzZFUYXQTz62"
# ###Step 1: Cloning the Project (House keeping)
# Lets clone the test project in Glennopt. We will need the test folder located in `GlennOPT/test/kur/serial`
# + id="-gE-Ez1qtyIA" outputId="b48b7c42-4411-4bff-c5e2-489e8a41babb" colab={"base_uri": "https://localhost:8080/"}
# Clone the source code for GlennOPT
# !git clone https://github.com/nasa/GlennOPT.git
# Little Housekeeping
# !cp -r GlennOPT/test/KUR/serial/Evaluation . # Copy the folder we need
# !rm GlennOPT/ -r # Deletes GlennOPT source code. We don't need this anymore
# + id="G82fsy9HS71f" outputId="052a7bbd-ef64-40c7-fbe5-41ea252d800f" colab={"base_uri": "https://localhost:8080/"}
# Install GlennOPT
# !python --version
# !pip install glennopt
# Restart the runtime
import os
os.kill(os.getpid(), 9)
# + [markdown] id="v85eBtEUhf6m"
# #Folder Structure and Evaluation Script
# + [markdown] id="yiA7Q1xxhnyk"
# ###Initial folder structure
# ```
# Evaluation
# | - evaluate.py (Gets copied to each individual directory)
# multi-objective_example.ipynb
# machinefile.txt (Optional, add this if you want to breakdown hosts per evaluation)
# ```
#
# ###After optimization
# ```
# Calculation
# | - DOE
# | -- IND000
# | ----- input.txt (Generated by optimizer)
# | ----- evaluate.py (Executes the cfd and reads results)
# | ----- output.txt (Generated by evaluate.py)
# | -- ...
# | -- IND128
# | - POP000
# | -- IND000
# | -- ...
# | -- IND039
# Evaluation
# | - evaluate.py (Gets copied to each individual directory)
# multi-objective_example.ipynb
# machinefile.txt (Optional, add this if you want to break down hosts per evaluation)
# ```
# Note: Glennopt constructs the calculation folder automatically. Each population is saved as DOE or as POPXXX. This is done so that when you are running a Computational Fluid Simulation or any kind of optimization where crashes could occur, you can investigate why the simulation crashed.
#
# Also if there are any additional post processing that needs to be done, by saving the evaluations this way, it is also possible to re-process the files differently and restart the optimization. **Restarts** will be shown in a later section.
#
# + [markdown] id="kHenLEAjoIMT"
# ##Evaluation Script
# You may have noticed that when in earler codes the github folder is cloned and part of it was copied and the rest of it was deleted. This is done to import the evaluation script found in Evaluation/evaluate.py
#
# The purpose of this script is to call the Kursawe function (kur.py) to perform a single execution. The inputs will be read from an input.dat file and output to output.txt. See `read_input` and `print_output` functions
#
# ---
# ```
# # evaluation.py
# def read_input(input_filename):
# x = []
# with open(input_filename, "r") as f:
# for line in f:
# split_val = line.split('=')
# if len(split_val)==2: # x1 = 2 # Grab the 2
# x.append(float(split_val[1]))
# return x
#
# def print_output(y):
# with open("output.txt", "w") as f:
# f.write('objective1 = {0:.6f}\n'.format(y[0])) # Output should contain [Name of the Objective/Parameter] = [value] This is read by the optimizer
# f.write('objective2 = {0:.6f}\n'.format(y[1]))
# f.write('p1 = {0:.6f}\n'.format(y[2]))
# f.write('p2 = {0:.6f}\n'.format(y[3]))
# f.write('p3 = {0:.6f}\n'.format(y[4]))
# # f.write('Objective2 = {0:.6f}\n'.format(y))
#
# if __name__ == '__main__':
# x = read_input("input.dat")
# # Call Rosebrock test function
# import kur as kur
# y = kur.KUR(x[0],x[1],x[2])
# print_output(y)
# ```
# ## Kursawe Function
# This is the kursawe function copied from wikipedia
# ```
# # kur.py
# import math
#
# def KUR(x1,x2,x3):
# '''
# Kursawe Function
# mutiple output
# '''
# f1 = (-10*math.exp(-0.2*math.sqrt(x1*x1+x2*x2))) + (-10*math.exp(-0.2*math.sqrt(x2*x2+x3*x3)))
#
# f2 = (math.pow(abs(x1),0.8)+5*math.sin(x1*x1*x1))+(math.pow(abs(x2),0.8)+5*math.sin(x2*x2*x2))+(math.pow(abs(x3),0.8)+5*math.sin(x3*x3*x3))
# # Performance Parameter
# p1 = x1 + x2 + x3
# p2 = x1*x2*x3
# p3 = x1 - x2 - x3
# return f1,f2,p1,p2,p3
# ```
#
#
#
# + [markdown] id="gsvodYmMhZEB"
# #Optimization
# + [markdown] id="Mm7Az99Hm0ZA"
# Import relevant libraries
# + id="hvsEjIEQURPF"
import sys,os
from glennopt.base import Parameter
from glennopt.helpers import mutation_parameters, de_mutation_type
from glennopt.optimizers import NSGA3
from glennopt.DOE import Default,CCD,FullFactorial,LatinHyperCube
# + id="X5pwbjp6STEL" outputId="df4cc451-e6ad-4989-c35c-6820e324df94" colab={"base_uri": "https://localhost:8080/"}
# Clean up
# !rm -r Calculation # Remove the calculation folder, lets start from scratch
# !rm *.csv *.log # Remove the restart file and any log files generated
# + [markdown] id="X8CFj5xpg7j3"
# **Evaluation Parameters**: are required. This is the vector of variables that goes into your objective function.
#
# **Objectives**: number of objectives to solve for
#
# **Performance Parameters**: this is not required but I set this up for reference
# + id="arNMkiHynevz"
# Initialize the DOE
# doe = CCD()
doe = FullFactorial(levels=8)
# doe = LatinHyperCube(128)
doe.add_parameter(name="x1",min_value=-5,max_value=5)
doe.add_parameter(name="x2",min_value=-5,max_value=5)
doe.add_parameter(name="x3",min_value=-5,max_value=5)
doe.add_objectives(name='objective1')
doe.add_objectives(name='objective2')
# No performance Parameters
doe.add_perf_parameter(name='p1')
doe.add_perf_parameter(name='p2')
doe.add_perf_parameter(name='p2')
# + id="B6O8CIpZRBhT"
# Initialize the Optimizer
pop_size=32
current_dir = os.getcwd()
ns = NSGA3(eval_command = "python evaluation.py", eval_folder="Evaluation",pop_size=pop_size,optimization_folder=current_dir)
ns.add_objectives(objectives=doe.objectives)
ns.add_eval_parameters(eval_params=doe.eval_parameters)
ns.add_performance_parameters(performance_params= doe.perf_parameters)
# + [markdown] id="8VeKNnzFq2hZ"
# **If you want other mutaton strategies or crossover strategies just add an issue to the github and provide as many references and examples as possible. We will try to incorporate them. **
# + id="o8vGLFkHnmAz"
# Set the mutation parameters
ns.mutation_params.mutation_type = de_mutation_type.de_rand_1_bin # Choice of de_best_1_bin (single objective) or de_rand_1_bin (multi-objective)
ns.mutation_params.F = 0.8
ns.mutation_params.C = 0.7
# Parallel Settings (You don't need to run this block if you only want serial execution)
ns.parallel_settings.concurrent_executions = 8 # Change to 1 for serial
ns.parallel_settings.cores_per_execution: 1
ns.parallel_settings.execution_timeout = 0.2 # minutes
# + [markdown] id="rZTXHADuN8O6"
# Enable Parallel Execution (OPTIONAL)
# + id="VMoA-2bbpN8u"
# Parallel Settings (You don't need to run this block if you only want serial execution)
ns.parallel_settings.concurrent_executions = 8 # Change to 1 for serial
ns.parallel_settings.cores_per_execution: 1
ns.parallel_settings.execution_timeout = 0.2 # minutes
# + [markdown] id="qhFjdeCXRr2c"
# ##Run the Design of Experiments
# Design of experiments is used to sample the evaluation space. Say you have 5 variables and f(x[1-5]) = y[1,2] and each x1 through x5 have min and max bounds. The design of experiments is used to evaluate different combinations of x1 to x5 which are used as the starting population (pop_start=-1)
# + id="quXqm1SLqpD4" outputId="db622f13-9ee9-4bf2-ea74-9cd9df5ed11b" colab={"base_uri": "https://localhost:8080/"}
# Generate the DOE and start the optimization
ns.start_doe(doe.generate_doe())
# + id="CkmTt9tDV8rg"
# Execute the Optimization
ns.optimize_from_population(pop_start=-1,n_generations=25) # Start from the DOE and iterate from pop 0 to 24
# + [markdown] id="9yA36UO9rD_5"
# ###Restarting the Simulation from a previous restart file
#
# + id="TDK2O8fArmhc"
ns.optimize_from_population(pop_start=24,n_generations=25) # Start from the DOE and iterate from pop 24 to 49
# + [markdown] id="vlfUROt0rsre"
# ##Generating a restart file
# Just in case you accidentally deleted it
# + id="S2k2y2LLryXn"
# !rm restart_file.csv
ns.create_restart() # Appends/creates a new restart file
# + [markdown] id="RfaRcZ8vux67"
# ##Plotting the Pareto Front
#
#
#
#
# + [markdown] id="iu3PWtg9JOHk"
# You can make the plots using the glennopt or you can create animated plots of the pareto front
# + id="Xij92h7Cu5aR" outputId="1ba2c6e0-a961-44a9-9449-03e2de3c42f0" colab={"base_uri": "https://localhost:8080/", "height": 825}
ns.read_calculation_folder()
ns.plot_2D('objective1','objective2',[-20,0],[-15,20])
# + [markdown] id="fyfv9uvxJYpo"
# ###Animated Plot
# This is an example of an animated plot. It may take a while to run depending on the number of frames and the interval
# + id="bQ_ybbsTyfaA" outputId="3d23599e-6849-43ca-fd7e-3c7993191e85" colab={"base_uri": "https://localhost:8080/", "height": 914}
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from matplotlib import animation, rc
from IPython.display import HTML
import numpy as np
num_frames = 150
xlim = [-20,0]
ylim = [-15,20]
obj1_name = 'objective1'
obj2_name = 'objective2'
ns.read_calculation_folder()
fig,ax = plt.subplots(figsize=(10,6), dpi=80, facecolor='w', edgecolor='k')
keys = list(ns.pandas_cache.keys())
keys.sort()
nPop = len(list(ns.pandas_cache.keys()))
divisor = num_frames/nPop
def init():
data = get_data(0)
pop_scatter = ax.scatter(data[:,0],data[:,1],s=20,alpha=0.5)
ax.set_xlim(xlim[0],xlim[1])
ax.set_ylim(ylim[0],ylim[1])
ax.set_xlabel(obj1_name)
ax.set_ylabel(obj2_name)
#pop_scatter.set_array(np.array([]))
return pop_scatter,
def get_data(pop):
key = keys[pop]
obj_data = list()
for index, row in ns.pandas_cache[key].iterrows():
obj_data.append([row[obj1_name],row[obj2_name]])
return np.array(obj_data)
def animate(pop):
pop = int(pop/divisor)
data = get_data(pop)
pop_scatter = ax.scatter(data[:,0],data[:,1],s=20,alpha=0.5)
ax.legend([str(pop)])
return pop_scatter,
anim = animation.FuncAnimation(fig, animate, init_func=init,
frames=num_frames, interval=10, blit=True)
HTML(anim.to_html5_video())
# + [markdown] id="omYSOHD4trDs"
# #Visualization
#
# + [markdown] id="vc9sZaGPuFQe"
# Getting the best objective value vs population. This is useful to see if the objective is changing as population increases
# + id="mcCSLcK9tzz1" outputId="6e89a1cb-d230-4b5f-d179-33c5042e56f6" colab={"base_uri": "https://localhost:8080/"}
from glennopt.base import Individual
from glennopt.helpers import get_best,get_pop_best
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
individuals = ns.read_calculation_folder()
best_individuals, best_fronts = get_pop_best(individuals)
# pop size is the size of population to keep track of when applying non-dimensional sorting. Larger the size = greater the slow down
objectives, pop, best_fronts = get_best(individuals,pop_size=30)
print(objectives.shape)
# + [markdown] id="p8RONFaKWgYV"
# ## Plotting the best objective vs population
# + id="TxfyHYiqWjuF" outputId="b6865f5e-5440-4f93-892f-a835ea091a19" colab={"base_uri": "https://localhost:8080/", "height": 279}
objective_index = 1
_, ax = plt.subplots()
ax.scatter(pop, objectives[:,objective_index],color='blue',s=10)
ax.set_xlabel('Population')
ax.set_ylabel('Objective {0} Value'.format(objective_index))
plt.show()
# + [markdown] id="-4YtLpZGWyWR"
# #Plotting the best individual in each population
# + id="PDdAiojKW1Sr" outputId="5a5b3e04-4f05-4195-9a3b-2d48242950c1" colab={"base_uri": "https://localhost:8080/", "height": 295}
nobjectives = len(best_individuals[0][0].objectives)
objective_data = list()
for pop,best_individual in best_individuals.items():
objective_data.append(best_individual[objective_index].objectives[objective_index])
_,ax = plt.subplots()
colors = cm.rainbow(np.linspace(0, 1, len(best_individuals.keys())))
ax.scatter(list(best_individuals.keys()), objective_data, color='blue',s=10)
ax.set_xlabel('Population')
ax.set_ylabel('Objective {0} Value'.format(objective_index))
ax.set_title('Best individual at each population')
plt.show()
# + [markdown] id="aISd53HiW7S1"
# ##Plot the Pareto Front
# The pareto front shows a trade off between two objective values. As the optimization advances to the next population, the individuals get closer to the minimum of both objectives and yields designs that are a compromise between the two.
# + id="axlDva1iW-0n" outputId="f0de33e7-b842-43a4-87dc-4e3ece6ac04a" colab={"base_uri": "https://localhost:8080/", "height": 820}
best_fronts = [front for _,front in sorted(zip(pop,best_fronts))] # dictionary keys aren't sorted. this cleans up some stuff.
pop = sorted(pop)
fig,ax = plt.subplots(figsize=(10,8))
colors = cm.rainbow(np.linspace(0, 1, len(best_fronts)))
indx = 0
legend_labels = []
# Scan the pandas file, grab objectives for each population
for ind_list in best_fronts:
obj1_data = []
obj2_data = []
c=colors[indx]
for ind in ind_list[0]:
obj1_data.append(ind.objectives[0])
obj2_data.append(ind.objectives[1])
# Plot the gathered data
ax.scatter(obj1_data, obj2_data, color=c, s=20,alpha=0.5)
legend_labels.append(pop[indx])
indx+=1
ax.set_xlabel('Objective 1')
ax.set_ylabel('Objective 2')
ax.set_title('Non-dimensional sorting: Best Front for each population')
ax.legend(legend_labels)
fig.canvas.draw()
fig.canvas.flush_events()
plt.show()
| test/kur-nsopt/multi_objective_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Iris Training and Prediction with Sagemaker Scikit-learn
# This tutorial shows you how to use [Scikit-learn](https://scikit-learn.org/stable/) with Sagemaker by utilizing the pre-built container. Scikit-learn is a popular Python machine learning framework. It includes a number of different algorithms for classification, regression, clustering, dimensionality reduction, and data/feature pre-processing.
#
# The [sagemaker-python-sdk](https://github.com/aws/sagemaker-python-sdk) module makes it easy to take existing scikit-learn code, which we will show by training a model on the IRIS dataset and generating a set of predictions. For more information about the Scikit-learn container, see the [sagemaker-scikit-learn-containers](https://github.com/aws/sagemaker-scikit-learn-container) repository and the [sagemaker-python-sdk](https://github.com/aws/sagemaker-python-sdk) repository.
#
# For more on Scikit-learn, please visit the Scikit-learn website: <http://scikit-learn.org/stable/>.
#
# ### Table of contents
# * [Upload the data for training](#upload_data)
# * [Create a Scikit-learn script to train with](#create_sklearn_script)
# * [Create the SageMaker Scikit Estimator](#create_sklearn_estimator)
# * [Train the SKLearn Estimator on the Iris data](#train_sklearn)
# * [Using the trained model to make inference requests](#inferece)
# * [Deploy the model](#deploy)
# * [Choose some data and use it for a prediction](#prediction_request)
# * [Endpoint cleanup](#endpoint_cleanup)
# * [Batch Transform](#batch_transform)
# * [Prepare Input Data](#prepare_input_data)
# * [Run Transform Job](#run_transform_job)
# * [Check Output Data](#check_output_data)
# First, lets create our Sagemaker session and role, and create a S3 prefix to use for the notebook example.
# +
# S3 prefix
prefix = 'Scikit-iris'
import sagemaker
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
# Get a SageMaker-compatible role used by this Notebook Instance.
role = get_execution_role()
# -
# ## Upload the data for training <a class="anchor" id="upload_data"></a>
#
# When training large models with huge amounts of data, you'll typically use big data tools, like Amazon Athena, AWS Glue, or Amazon EMR, to create your data in S3. For the purposes of this example, we're using a sample of the classic [Iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set), which is included with Scikit-learn. We will load the dataset, write locally, then write the dataset to s3 to use.
# +
import numpy as np
import os
from sklearn import datasets
# Load Iris dataset, then join labels and features
iris = datasets.load_iris()
joined_iris = np.insert(iris.data, 0, iris.target, axis=1)
# Create directory and write csv
os.makedirs('./data', exist_ok=True)
np.savetxt('./data/iris.csv', joined_iris, delimiter=',', fmt='%1.1f, %1.3f, %1.3f, %1.3f, %1.3f')
# -
# Once we have the data locally, we can use use the tools provided by the SageMaker Python SDK to upload the data to a default bucket.
# +
WORK_DIRECTORY = 'data'
train_input = sagemaker_session.upload_data(WORK_DIRECTORY, key_prefix="{}/{}".format(prefix, WORK_DIRECTORY) )
# -
# ## Create a Scikit-learn script to train with <a class="anchor" id="create_sklearn_script"></a>
# SageMaker can now run a scikit-learn script using the `SKLearn` estimator. When executed on SageMaker a number of helpful environment variables are available to access properties of the training environment, such as:
#
# * `SM_MODEL_DIR`: A string representing the path to the directory to write model artifacts to. Any artifacts saved in this folder are uploaded to S3 for model hosting after the training job completes.
# * `SM_OUTPUT_DIR`: A string representing the filesystem path to write output artifacts to. Output artifacts may include checkpoints, graphs, and other files to save, not including model artifacts. These artifacts are compressed and uploaded to S3 to the same S3 prefix as the model artifacts.
#
# Supposing two input channels, 'train' and 'test', were used in the call to the `SKLearn` estimator's `fit()` method, the following environment variables will be set, following the format `SM_CHANNEL_[channel_name]`:
#
# * `SM_CHANNEL_TRAIN`: A string representing the path to the directory containing data in the 'train' channel
# * `SM_CHANNEL_TEST`: Same as above, but for the 'test' channel.
#
# A typical training script loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model to model_dir so that it can be hosted later. Hyperparameters are passed to your script as arguments and can be retrieved with an `argparse.ArgumentParser` instance. For example, the script that we will run in this notebook is the below:
#
# ```python
# import argparse
# import pandas as pd
# import os
#
# from sklearn import tree
# from sklearn.externals import joblib
#
#
# if __name__ == '__main__':
# parser = argparse.ArgumentParser()
#
# # Hyperparameters are described here. In this simple example we are just including one hyperparameter.
# parser.add_argument('--max_leaf_nodes', type=int, default=-1)
#
# # Sagemaker specific arguments. Defaults are set in the environment variables.
# parser.add_argument('--output-data-dir', type=str, default=os.environ['SM_OUTPUT_DATA_DIR'])
# parser.add_argument('--model-dir', type=str, default=os.environ['SM_MODEL_DIR'])
# parser.add_argument('--train', type=str, default=os.environ['SM_CHANNEL_TRAIN'])
#
# args = parser.parse_args()
#
# # Take the set of files and read them all into a single pandas dataframe
# input_files = [ os.path.join(args.train, file) for file in os.listdir(args.train) ]
# if len(input_files) == 0:
# raise ValueError(('There are no files in {}.\n' +
# 'This usually indicates that the channel ({}) was incorrectly specified,\n' +
# 'the data specification in S3 was incorrectly specified or the role specified\n' +
# 'does not have permission to access the data.').format(args.train, "train"))
# raw_data = [ pd.read_csv(file, header=None, engine="python") for file in input_files ]
# train_data = pd.concat(raw_data)
#
# # labels are in the first column
# train_y = train_data.ix[:,0]
# train_X = train_data.ix[:,1:]
#
# # Here we support a single hyperparameter, 'max_leaf_nodes'. Note that you can add as many
# # as your training my require in the ArgumentParser above.
# max_leaf_nodes = args.max_leaf_nodes
#
# # Now use scikit-learn's decision tree classifier to train the model.
# clf = tree.DecisionTreeClassifier(max_leaf_nodes=max_leaf_nodes)
# clf = clf.fit(train_X, train_y)
#
# # Print the coefficients of the trained classifier, and save the coefficients
# joblib.dump(clf, os.path.join(args.model_dir, "model.joblib"))
#
#
# def model_fn(model_dir):
# """Deserialized and return fitted model
#
# Note that this should have the same name as the serialized model in the main method
# """
# clf = joblib.load(os.path.join(model_dir, "model.joblib"))
# return clf
# ```
# Because the Scikit-learn container imports your training script, you should always put your training code in a main guard `(if __name__=='__main__':)` so that the container does not inadvertently run your training code at the wrong point in execution.
#
# For more information about training environment variables, please visit https://github.com/aws/sagemaker-containers.
# ## Create SageMaker Scikit Estimator <a class="anchor" id="create_sklearn_estimator"></a>
#
# To run our Scikit-learn training script on SageMaker, we construct a `sagemaker.sklearn.estimator.sklearn` estimator, which accepts several constructor arguments:
#
# * __entry_point__: The path to the Python script SageMaker runs for training and prediction.
# * __role__: Role ARN
# * __train_instance_type__ *(optional)*: The type of SageMaker instances for training. __Note__: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.
# * __sagemaker_session__ *(optional)*: The session used to train on Sagemaker.
# * __hyperparameters__ *(optional)*: A dictionary passed to the train function as hyperparameters.
#
# To see the code for the SKLearn Estimator, see here: https://github.com/aws/sagemaker-python-sdk/tree/master/src/sagemaker/sklearn
# +
from sagemaker.sklearn.estimator import SKLearn
script_path = 'scikit_learn_iris.py'
sklearn = SKLearn(
entry_point=script_path,
train_instance_type="ml.c4.xlarge",
role=role,
sagemaker_session=sagemaker_session,
hyperparameters={'max_leaf_nodes': 30})
# -
# ## Train SKLearn Estimator on Iris data <a class="anchor" id="train_sklearn"></a>
# Training is very simple, just call `fit` on the Estimator! This will start a SageMaker Training job that will download the data for us, invoke our scikit-learn code (in the provided script file), and save any model artifacts that the script creates.
sklearn.fit({'train': train_input})
# ## Using the trained model to make inference requests <a class="anchor" id="inference"></a>
#
# ### Deploy the model <a class="anchor" id="deploy"></a>
#
# Deploying the model to SageMaker hosting just requires a `deploy` call on the fitted model. This call takes an instance count and instance type.
predictor = sklearn.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
# ### Choose some data and use it for a prediction <a class="anchor" id="prediction_request"></a>
#
# In order to do some predictions, we'll extract some of the data we used for training and do predictions against it. This is, of course, bad statistical practice, but a good way to see how the mechanism works.
# +
import itertools
import pandas as pd
shape = pd.read_csv("data/iris.csv", header=None)
a = [50*i for i in range(3)]
b = [40+i for i in range(10)]
indices = [i+j for i,j in itertools.product(a,b)]
test_data = shape.iloc[indices[:-1]]
test_X = test_data.iloc[:,1:]
test_y = test_data.iloc[:,0]
# -
# Prediction is as easy as calling predict with the predictor we got back from deploy and the data we want to do predictions with. The output from the endpoint return an numerical representation of the classification prediction; in the original dataset, these are flower names, but in this example the labels are numerical. We can compare against the original label that we parsed.
print(predictor.predict(test_X.values))
print(test_y.values)
# ### Endpoint cleanup <a class="anchor" id="endpoint_cleanup"></a>
#
# When you're done with the endpoint, you'll want to clean it up.
sklearn.delete_endpoint()
# ## Batch Transform <a class="anchor" id="batch_transform"></a>
# We can also use the trained model for asynchronous batch inference on S3 data using SageMaker Batch Transform.
# Define a SKLearn Transformer from the trained SKLearn Estimator
transformer = sklearn.transformer(instance_count=1, instance_type='ml.m4.xlarge')
# ### Prepare Input Data <a class="anchor" id="prepare_input_data"></a>
# We will extract 10 random samples of 100 rows from the training data, then split the features (X) from the labels (Y). Then upload the input data to a given location in S3.
# + language="bash"
# # Randomly sample the iris dataset 10 times, then split X and Y
# mkdir -p batch_data/XY batch_data/X batch_data/Y
# for i in {0..9}; do
# cat data/iris.csv | shuf -n 100 > batch_data/XY/iris_sample_${i}.csv
# cat batch_data/XY/iris_sample_${i}.csv | cut -d',' -f2- > batch_data/X/iris_sample_X_${i}.csv
# cat batch_data/XY/iris_sample_${i}.csv | cut -d',' -f1 > batch_data/Y/iris_sample_Y_${i}.csv
# done
# -
# Upload input data from local filesystem to S3
batch_input_s3 = sagemaker_session.upload_data('batch_data/X', key_prefix=prefix + '/batch_input')
# ### Run Transform Job <a class="anchor" id="run_transform_job"></a>
# Using the Transformer, run a transform job on the S3 input data.
# Start a transform job and wait for it to finish
transformer.transform(batch_input_s3, content_type='text/csv')
print('Waiting for transform job: ' + transformer.latest_transform_job.job_name)
transformer.wait()
# ### Check Output Data <a class="anchor" id="check_output_data"></a>
# After the transform job has completed, download the output data from S3. For each file "f" in the input data, we have a corresponding file "f.out" containing the predicted labels from each input row. We can compare the predicted labels to the true labels saved earlier.
# Download the output data from S3 to local filesystem
batch_output = transformer.output_path
# !mkdir -p batch_data/output
# !aws s3 cp --recursive $batch_output/ batch_data/output/
# Head to see what the batch output looks like
# !head batch_data/output/*
# + language="bash"
# # For each sample file, compare the predicted labels from batch output to the true labels
# for i in {1..9}; do
# diff -s batch_data/Y/iris_sample_Y_${i}.csv \
# <(cat batch_data/output/iris_sample_X_${i}.csv.out | sed 's/[["]//g' | sed 's/, \|]/\n/g') \
# | sed "s/\/dev\/fd\/63/batch_data\/output\/iris_sample_X_${i}.csv.out/"
# done
# -
| sagemaker-python-sdk/scikit_learn_iris/Scikit-learn Estimator Example With Batch Transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HenriqueCCdA/bootCampAluraDataScience/blob/master/modulo4/extra/Explorando_ML.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Vnu2cuROHA_a"
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
import numpy as np
# + id="aTXuMIRx1zBs"
def colunas_selecionadas(colunas: str,
colunas_excluidas=[],
f_diff: bool=True)->str:
colunas_excluidas+=["ICU", "WINDOW"]
colunas_selecionadas = []
for name in colunas:
if ("DIFF" in name) and f_diff:
continue
elif name in colunas_excluidas:
continue
else:
colunas_selecionadas.append(name)
return colunas_selecionadas
def proporcao_y(y, name:str):
p = y.value_counts(normalize=True)
print(f"Proporcao do {name}")
for l , v in zip(p.index, p.values):
print(f"Campo {l} -> {v:.2f}")
def lendo_dados(path):
dados = pd.read_csv(path)
nl, nc = dados.shape
print(f"Número de linhas :{nl}")
print(f"Número de colunas :{nc}")
return dados
def score(y_pre, y_test, verbose: bool = True)-> (float, float):
tx = sum(y_pre == y_test)/len(y_test)
if verbose:
print("*************************")
print(f"Taxa de acerto {tx:.2f}")
print("*************************")
return tx
def run_LogisticRegression(dados,
seed,
colunas_excluidas= [],
f_diff: bool = True,
verbose: bool = True)-> float:
x = dados[colunas_selecionadas(dados.columns, colunas_excluidas, f_diff)]
y = dados.iloc[:,-1]
np.random.seed(seed)
x_train, x_test, y_train, y_test = train_test_split(x, y, stratify=y)
if verbose:
print(x.columns.tolist())
print("********************************************")
print(f"Numero de variaveis = {x.shape[1]}")
print(f"Numero de amostras de treino = {x.shape[0]}")
print("********************************************")
if verbose:
proporcao_y(y , "y")
proporcao_y(y_test , "y_test")
proporcao_y(y_train, "y_train")
modelo = LogisticRegression(max_iter=5000, verbose=0, tol=1e-8, C=0.1)
modelo.fit(x_train, y_train)
y_hat = modelo.predict(x_train)
y_pred = modelo.predict(x_test)
s_train = score(y_hat , y_train, verbose)
s_test = score(y_pred, y_test, verbose)
return s_test, s_train
# + id="5MPsMeja0-1r" outputId="ada29315-b824-4f1f-c0ec-408bb104aa28" colab={"base_uri": "https://localhost:8080/"}
dados = lendo_dados("dados_tratados_por_paciente.csv")
# + id="DKnpUoQe5FZh" outputId="f90cee25-c413-4b82-bcde-1b4bdebecb8e" colab={"base_uri": "https://localhost:8080/", "height": 289}
dados.head()
# + id="6acbSEwPN5F5" outputId="ae70f2df-a5f4-40a7-fa64-4a01cac08051" colab={"base_uri": "https://localhost:8080/"}
run_LogisticRegression(dados, seed=146731, f_diff=False, verbose = True)
# + id="EIvAD8RfM2wS"
lista_sementes = [ 61367, 78668, 76592, 49310, 2825,
34136, 14747, 65643, 94788, 31840,
54546, 71546, 95580, 32717, 1918,
20892, 18289, 6295, 8375, 1648,
65323, 58650, 70938, 41914, 7960,
73762, 76050, 22706, 39405, 70837]
# + id="lUTq0eWMLh5A"
res_treino = []
res_teste = []
for semente in lista_sementes:
test1, train1 = run_LogisticRegression(dados, seed=semente, f_diff=False, verbose = False)
test2, train2 = run_LogisticRegression(dados, seed=semente,
colunas_excluidas=["PATIENT_VISIT_IDENTIFIER"],
f_diff=True, verbose = False)
test3, train3 = run_LogisticRegression(dados, seed=semente,
colunas_excluidas=["PATIENT_VISIT_IDENTIFIER", "AGE_ABOVE65"],
f_diff=True, verbose = False)
test4, train4 = run_LogisticRegression(dados, seed=semente,
colunas_excluidas=["PATIENT_VISIT_IDENTIFIER", "AGE_PERCENTIL"],
f_diff=True, verbose = False)
res_teste.append((test1, test2, test3, test4))
res_treino.append((train1, train2, train3, train4))
res_treino = np.array(res_treino)
res_teste = np.array(res_teste)
# + id="pxp9IUQJDtsA" outputId="dec6128c-38b7-44fd-e5f7-7b3442c3980c" colab={"base_uri": "https://localhost:8080/", "height": 205}
res = pd.DataFrame(data={'sementes':lista_sementes,
'l1_test' :res_teste[:,0],
'l1_train' :res_treino[:,0],
'l2_test' :res_teste[:,1],
'l2_train' :res_treino[:,1],
'l3_test' :res_teste[:,2],
'l3_train' :res_treino[:,2],
'l4_test' :res_teste[:,3],
'l4_train' :res_treino[:,3]
})
res.head()
# + id="wu1mgLiIQAE7"
col = res.columns[1:]
medias = res[col].mean()
medianas = res[col].median()
desvio_padrao = res[col].std()
# + id="efTHliiCQB5w" outputId="5abcd647-147f-411d-eff0-b9eb31b2a9bb" colab={"base_uri": "https://localhost:8080/", "height": 143}
pd.DataFrame(data={'medias' :medias,
'mediana' :medianas,
'desvio_padrao':desvio_padrao
}).T
# + id="upYCo9hvSkKN"
| modulo4/extra/Explorando_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from astropy import table
from astropy.table import Table
from astropy.io import ascii
SAGA_DIR = os.environ['SAGA_DIR']
# -
from palettable.colorbrewer.qualitative import Dark2_8
if 'plt' in locals() and hasattr(plt, 'rcParams'):
plt.rcParams['lines.linewidth'] = 2.0
plt.rcParams['font.size'] = 15.0
plt.rcParams['font.family'] = 'serif'
plt.rcParams['axes.prop_cycle'] = plt.cycler(color=Dark2_8.mpl_colors)
plt.rcParams['legend.fontsize'] = 'medium'
plt.rcParams['legend.frameon'] = False
plt.rcParams['figure.dpi'] = 100
plt.rcParams['figure.figsize'] = 7, 6
plt.rcParams['xtick.major.size'] = 6
plt.rcParams['xtick.minor.size'] = 4
plt.rcParams['ytick.major.size'] = 6
plt.rcParams['ytick.minor.size'] = 4
SAGA_DIR = os.environ['SAGA_DIR']
# READ SPECTRA
file = SAGA_DIR +'/data/saga_spectra_clean.fits.gz'
allspec = Table.read(file)
# +
# FIND GRI GALAXIES BETWEEN 17 < r < 20.75
umag = allspec['u'] - allspec['EXTINCTION_U']
gmag = allspec['g'] - allspec['EXTINCTION_G']
rmag = allspec['r'] - allspec['EXTINCTION_R']
imag = allspec['i'] - allspec['EXTINCTION_I']
ug = umag - gmag
gr = gmag - rmag
ri = rmag - imag
grerr = np.sqrt(allspec['g_err']**2 + allspec['r_err']**2)
rierr = np.sqrt(allspec['r_err']**2 + allspec['i_err']**2)
ugerr = np.sqrt(allspec['u_err']**2 + allspec['g_err']**2)
cgmr = gr - 2.*grerr
crmi = ri - 2.*rierr
# GRI CUTS
gri1 = cgmr < 0.85
gri2 = crmi < 0.55
# UGRI CUT
#ug < 1.5*gr-0.25
#ugri = ug > 1.5*gr
ugri = (ug+2.*ugerr) > (1.5*(gr - 2.*grerr))
# +
# MAKE CUTS ON MAIN SAMPLE
m_rmv = allspec['REMOVE'] == -1
m_fib = allspec['FIBERMAG_R'] <= 23
m_sg = allspec['PHOT_SG'] == 'GALAXY'
m_boss = allspec['survey'] !='boss'
m_qual = m_rmv & m_sg & m_fib
m_ody = allspec['HOST_NSAID'] == 147100
m_mag = (17.7 < rmag) & (rmag < 20.75)
# -
# SAGA GRI SAMPLE
#print 1.*np.sum(gri1 & gri2 & m_qual & m_mag & ugri)/np.sum(gri1 & gri2 & m_qual & m_mag)
#print np.sum(gri1 & gri2 & m_qual & m_mag & ugri)
all = allspec[m_qual & m_mag & m_ody]
gri = allspec[gri1 & gri2 & m_qual & m_mag]
ugri = allspec[gri1 & gri2 & m_qual & m_mag & ugri]
## SET BIN SIZE
binwidth = 0.1
b=np.arange(0, 1.1 + binwidth, binwidth)
# +
# BOSS GALAXIES
file = SAGA_DIR + '/data/cmass_full.fits'
cmass = Table.read(file)
c_gr = cmass['gmag']-cmass['rmag']
c_ri = cmass['rmag']-cmass['imag']
m1 = cmass['Column1'] == 0
m2 = cmass['zerr'] !=0
m3 = (cmass['rmag'] > 17.7) & (cmass['rmag'] < 21)
m4 = cmass['z'] > 0.005 # no stars!
mlowz = cmass['z'] < 0.015
boss = cmass['z'][m1&m2&m3&m4]
c_gr = cmass['gmag']-cmass['rmag']
c_ri = cmass['rmag']-cmass['imag']
plt.plot(c_gr[m1&m2&m3&m4],c_ri[m1&m2&m3&m4],'ko',ms=1)
plt.plot(c_gr[m1&m2&m3&m4&mlowz],c_ri[m1&m2&m3&m4&mlowz],'ro',ms=5)
xl = [-0.3,1.7]
yl=[-0.55,1.1]
tgr = 0.85
tri = 0.55
mgri = (c_gr > tgr) | (c_ri > tri)
for obj in cmass[m1&m2&m3&m4&mlowz&mgri]:
print obj['rmag'],obj['ra'],obj['dec']
print 'All CMASS = ',np.sum(m1&m2&m3&m4)
print 'Non-gri in CMASS = ',np.sum(m1&m2&m3&m4&mgri)
print 'LOWz in CMASS = ',np.sum(m1&m2&m3&m4&mlowz)
print 'LOWz in CMASS, not passing gri = ',np.sum(m1&m2&m3&m4&mlowz&mgri)
plt.xlim(xl)
plt.ylim(yl)
plt.axvline(tgr, c='w')
plt.axvline(tgr, c='k', ls=':')
plt.axhline(tri, c='w')
plt.axhline(tri, c='k', ls=':')
# -
plt.hist(cmass['rmag'][m1&m2&m3&m4],bins=100)
def hist_norm_height(n,bins,const):
''' Function to normalise bin height by a constant.
Needs n and bins from np.histogram or ax.hist.'''
n = np.repeat(n,2)
n = n / const
new_bins = [bins[0]]
new_bins.extend(np.repeat(bins[1:],2))
return n,new_bins[:-1]
# +
# ALL GALAXIES
n, bins, patches = plt.hist(all['SPEC_Z'], bins = b)
c1 = np.max(n)
nall,new_all = hist_norm_height(n,bins,c1)
# GRI
n, bins, patches = plt.hist(gri['SPEC_Z'], bins = b)
c1 = np.max(n)
ngri,new_gri = hist_norm_height(n,bins,c1)
#UGRI
n, bins, patches = plt.hist(ugri['SPEC_Z'], bins = b)
c = np.max(n)
nugri,new_ugri = hist_norm_height(n,bins,c1)
#BOSS
n, bins, patches = plt.hist(boss, bins = b)
c = np.max(n)
nboss,new_boss = hist_norm_height(n,bins,c)
# +
plt.plot([-1],[-1],'.',ms=0,label='$17.7 < r_o < 20.75$')
plt.step(new_all,nall,color='k',label='All Galaxies')
plt.step(new_gri,ngri,color='r',label='SAGA gri')
plt.step(new_ugri,nugri,color='b',label='SAGA ugri')
plt.step(new_gri,ngri,color='r',label='_nolabel_',alpha=0.8)
#plt.step(new_boss,nboss,color='g',label='BOSS')
plt.xlabel('Spectroscopic Redshift')
plt.ylabel('Normalized Galaxy Counts')
plt.xlim(0,0.9)
plt.ylim(0,1.05)
plt.legend(fontsize=12)
plt.savefig('fig_redshift.pdf')
plt.show()
# +
plt.plot([-1],[-1],'.',ms=0,label='$17.7 < r_o < 20.75$')
#plt.step(new_all,nall,color='k',label='All Galaxies')
plt.step(new_gri,ngri,color='b',label='SAGA gri')
plt.step(new_ugri,nugri,color='g',label='SAGA ugri')
plt.step(new_gri,ngri,color='b',label='_nolabel_',alpha=0.8)
plt.step(new_boss,nboss,color='r',label='BOSS')
plt.xlabel('Spectroscopic Redshift')
plt.ylabel('Normalized Galaxy Counts')
plt.xlim(0,0.9)
plt.ylim(0,1.05)
plt.legend(fontsize=12)
#plt.show()
#plt.savefig('fig_redshift.pdf')
# -
| plot_Fig7_redshift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
from IPython.core.display import HTML
css = open('style-table.css').read() + open('style-notebook.css').read()
HTML('<style>{}</style>'.format(css))
titles = pd.read_csv('data/titles.csv')
titles.head()
cast = pd.read_csv('data/cast.csv')
cast.head()
# ### Using groupby(), plot the number of films that have been released each decade in the history of cinema.
# ### Use groupby() to plot the number of "Hamlet" films made each decade.
# ### How many leading (n=1) roles were available to actors, and how many to actresses, in each year of the 1950s?
# ### In the 1950s decade taken as a whole, how many total roles were available to actors, and how many to actresses, for each "n" number 1 through 5?
# ### Use groupby() to determine how many roles are listed for each of the Pink Panther movies.
# ### List, in order by year, each of the films in which <NAME> has played more than 1 role.
# ### List each of the characters that <NAME> has portrayed at least twice.
| pycon-pandas-tutorial/Exercises-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Carcione et al. (2007), Figures 6-9
#
# Reproduced by <NAME> ([@prisae](https://github.com/prisae)).
#
# > **<NAME>., <NAME>, and <NAME>, 2007**
# > Cross-property relations between electrical conductivity and the seismic velocity of rocks.
# > Geophysics, 72, E193-E204; DOI: [10.1190/1.2762224](https://doi.org/10.1190/1.2762224).
#
#
# ### Requirements
# - `NumPy`
# - `SciPy`
# - `IPython`
# - `Jupyter`
# - `matplotlib`
#
# **NOTE:** I created these scripts in the early stage of my PhD, somewhen in 2010/2011 (if you are interested in my thesis you can find it [here](https://werthmuller.org/research), it comes with all source code, unfortunately without the real data due to copyrights). It was my first go at Python, so don't be too harsh ;). Many things would probably be included in `bruges`, `welly`, or another package by now, I don't know. The only thing I did at this point was to extract the required functions and translate them from Python 2 to Python 3.
# +
import numpy as np
from copy import deepcopy as dc
import matplotlib.pyplot as plt
import vel2res
# -
# See the notes above: I quick and dirty translated the Python 2 code to Python 3. By doing this, there might have happened funny things (0- and NaN-checks I did etc, which were not properly translated). To not clutter this notebooks with warnings I ignore all warnings here. To work properly with all the functions one would have to be a bit more careful...
np.seterr(all='ignore')
# Plot-style adjustments
# %matplotlib inline
plt.rcParams['figure.dpi'] = 100
# ## Figure 6
#
# ### Calculation figures 6 and 8
# +
data = vel2res.carc_tab1('shale')
vel2res.carc_der(data, 500)
data['a_k'] = np.array(-3)
data['a_f'] = np.array(1.)
data['p_e'] = np.array(.15)
rho_b = data['rho_b']
rho_0 = data['rho_0']
vp_b = data['vp_b']
tdata = dc(data)
tdata['rho_b'] = rho_0
# Calculation
rho_gt = vel2res.in2por2out(data, vel2res.por_v_harm, vel2res.rho_glov)
rho_ht = vel2res.in2por2out(data, vel2res.por_v_harm, vel2res.rho_herm)
rho_st = vel2res.in2por2out(data, vel2res.por_v_harm, vel2res.rho_self)
vel_ar = vel2res.in2por2out(tdata, vel2res.por_r_arch, vel2res.vp_raym)
sig_ar = np.nan_to_num(1./vel2res.in2por2out(data, vel2res.por_v_raym,
vel2res.rho_arch))
vel_br = vel2res.in2por2out(tdata, vel2res.por_r_hsub2, vel2res.vp_raym)
sig_br = np.nan_to_num(1./vel2res.in2por2out(data, vel2res.por_v_raym,
vel2res.rho_hsub2))
sig_at = np.nan_to_num(1./vel2res.in2por2out(data, vel2res.por_v_harm,
vel2res.rho_arch))
vel_gar = vel2res.in2por2out(tdata, vel2res.por_r_arch, vel2res.vp_gass)
sig_gar = np.nan_to_num(1./vel2res.in2por2out(data, vel2res.por_v_gass,
vel2res.rho_arch))
vel_ghe = vel2res.in2por2out(data, vel2res.por_r_herm, vel2res.vp_gass)
rho_ghe = vel2res.in2por2out(data, vel2res.por_v_gass, vel2res.rho_herm)
vel_gcr = vel2res.in2por2out(data, vel2res.por_r_crim, vel2res.vp_gass)
rho_gcr = vel2res.in2por2out(data, vel2res.por_v_gass, vel2res.rho_crim)
vel_gss = vel2res.in2por2out(data, vel2res.por_r_self, vel2res.vp_gass)
rho_gss = vel2res.in2por2out(data, vel2res.por_v_gass, vel2res.rho_self)
vel_ghm = vel2res.in2por2out(data, vel2res.por_r_hslb, vel2res.vp_gass)
vel_ghp = vel2res.in2por2out(data, vel2res.por_r_hsub, vel2res.vp_gass)
rho_ghm = vel2res.in2por2out(data, vel2res.por_v_gass, vel2res.rho_hsub)
rho_ghp = vel2res.in2por2out(data, vel2res.por_v_gass, vel2res.rho_hslb)
# -
# ### Plot
# +
fig6 = plt.figure(6)
plt.axvline(data['vp_s'], linewidth=1, color='k')
plt.axvline(data['vp_f'], linewidth=1, color='k')
plt.plot(vp_b, sig_at, 'b-', label='Archie/time-average')
plt.plot(vel_ar, 1./rho_0, 'g--', linewidth=1)
plt.plot(vp_b, sig_ar, 'g-', label='Archie/Raymer')
plt.plot(vp_b[1:], 1./rho_gt[1:], 'r-', label='Glover/time-average')
plt.plot(vp_b, 1./rho_ht, 'c-', label='Hermance/time-average')
plt.plot(vp_b, 1./rho_st, 'm-', label='Self-similar/time-average')
plt.plot(vel_br, 1./rho_0, 'y--', linewidth=1)
plt.plot(vp_b, sig_br, 'y-', label='HS/Raymer')
plt.legend()
plt.title("Carcione et al., 2007, Figure 6")
plt.xlabel("Velocity (km/s)")
plt.ylabel("Conductivity (S/m)")
plt.axis([1.0, 4.2, 0.0, 0.45])
plt.show()
# -
# Figure 6. Cross-property relations for different models of the overburden (shale saturated with brine).
#
# ### Original Figure 6
#
# 
#
# ## Figure 7
#
# **Important**: Equation (49) in Carcione et al is wrong. It is given as
# $$
# v_P = 2.2888\left(Z\frac{\sigma}{\sigma_f}\right)^{1/6}\ ,
# $$
# where the P-wave velocity $v_P$ is in km/s, depth $Z$ in km, and the conductivities in S/m.
#
# However, the correct equation is
# $$
# v_P = 2.2888\left(Z\frac{\sigma_f}{\sigma}\right)^{1/6}\ ,
# $$
# as for instance given in *The Rock Physics Handbook* by Mavko et al., 2009.
#
#
# Looking at the figures you might think hey, the curve from the equation in Carcione et al (original figure) looks much better then the curve from the equation by Mavko et al (my figure). It is misleading. The Faust-equation is a function of depth. So the curve will change depending at which depth you are. The other curves are not. So the different curves can not be compared just like that without taking other aspects into the analysis too.
#
# ### Calculation
# +
data2 = vel2res.carc_tab1('sand')
vel2res.carc_der(data2, 500)
data2['a_k'] = np.array(-3)
data2['a_f'] = np.array(1.)
data2['m_e'] = np.array(2.)
data2['p_e'] = np.array(.15)
data2['depth'] = np.array(2.)
rho_b2 = data2['rho_b']
vp_b2 = data2['vp_b']
# Calculation
rho_gt2 = vel2res.in2por2out(data2, vel2res.por_v_harm, vel2res.rho_glov)
rho_ht2 = vel2res.in2por2out(data2, vel2res.por_v_harm, vel2res.rho_herm)
rho_st2 = vel2res.in2por2out(data2, vel2res.por_v_harm, vel2res.rho_self)
vel_ghe2 = vel2res.in2por2out(data2, vel2res.por_r_herm, vel2res.vp_gass)
rho_ghe2 = vel2res.in2por2out(data2, vel2res.por_v_gass, vel2res.rho_herm)
vel_gcr2 = vel2res.in2por2out(data2, vel2res.por_r_crim, vel2res.vp_gass)
rho_gcr2 = vel2res.in2por2out(data2, vel2res.por_v_gass, vel2res.rho_crim)
vel_gss2 = vel2res.in2por2out(data2, vel2res.por_r_self, vel2res.vp_gass)
rho_gss2 = vel2res.in2por2out(data2, vel2res.por_v_gass, vel2res.rho_self)
vel_ghm2 = vel2res.in2por2out(data2, vel2res.por_r_hslb, vel2res.vp_gass)
vel_ghp2 = vel2res.in2por2out(data2, vel2res.por_r_hsub, vel2res.vp_gass)
rho_ghm2 = vel2res.in2por2out(data2, vel2res.por_v_gass, vel2res.rho_hsub)
rho_ghp2 = vel2res.in2por2out(data2, vel2res.por_v_gass, vel2res.rho_hslb)
rho_ft2 = vel2res.rho_faus(data2)
# -
# ### Plot
# +
# PLOT NON-GASSMANN RELATIONS
fig7 = plt.figure(7)
plt.axvline(data2['vp_s'], linewidth=1, color='k')
plt.axvline(data2['vp_f'], linewidth=1, color='k')
plt.plot(vp_b2, 1000./rho_gt2, '-', label='Glover/time-average')
plt.plot(vp_b2, 1000./rho_ht2, '-', label='Hermance/time-average')
plt.plot(vp_b2, 1000./rho_st2, '-', label='Self-similar/time-average')
plt.plot(vp_b2, 1000./rho_ft2, '-', label='Faust')
plt.legend()
plt.title("Carcione et al., 2007, Figure 7")
plt.xlabel("Velocity (km/s)")
plt.ylabel("Conductivity (mS/m)")
plt.axis([0.0, 6.0, 0.0, 1])
plt.show()
# -
# Figure 7. Cross-property relations for different models of the reservoir (sandstone saturated with oil). Archie-based relations are not shown because the conductivity is negligible (it is assumed that $\sigma_s$ = 0). The Faust curve corresponds to 2-km depth.
#
#
# **Note:** Y-axis in Figures 7 and 8 by a factor 10 different from Carcione. -> This is a typo in the paper, either in the plot or in Table 1. And again, the Faust-curve by Carcione is wrong, see my comment above.
#
# ### Original Figure 7
# 
#
# ## Figure 8
#
# ### Calculation
# Was done above together with Figure 6.
#
# ### Plot
# +
# PLOT GASSMANN RELATIONS
fig8 = plt.figure(8)
plt.axvline(data['vp_s'], linewidth=1, color='k')
plt.axvline(data['vp_f'], linewidth=1, color='k')
plt.plot(vel_gar, 1./rho_0, 'b--', linewidth=1)
plt.plot(vp_b[1:], sig_gar[1:], 'b-', label='Archie')
plt.plot(vel_ghe, 1./rho_b, 'g--', linewidth=1)
plt.plot(vp_b[1:], 1./rho_ghe[1:], 'g-', label='Hermance')
plt.plot(vel_gcr, 1./rho_b, 'r--', linewidth=1)
plt.plot(vp_b[1:], 1./rho_gcr[1:], 'r-', label='CRIM')
plt.plot(vel_gss, 1./rho_b, 'c--', linewidth=1)
plt.plot(vp_b[1:], 1./rho_gss[1:], 'c-', label='Self-similar')
plt.plot(vel_ghm, 1./rho_b, 'm--', linewidth=1)
plt.plot(vp_b[1:], 1./rho_ghm[1:], 'm-', label='HS-')
plt.plot(vel_ghp, 1./rho_b, 'y--', linewidth=1)
plt.plot(vp_b[1:], 1./rho_ghp[1:], 'y-', label='HS+')
plt.legend(loc=1)
plt.text(1.7, .36, 'Gassmann relations')
plt.title("Carcione et al., 2007, Figure 8")
plt.xlabel("Velocity (km/s)")
plt.ylabel("Conductivity (S/m)")
plt.axis([1.0, 4.5, 0.0, 0.4])
plt.show()
# -
# Figure 8. Cross-property relations for different conductivity models of the overburden (shale saturated with brine), combined with the Gassmann equation. The dashed lines correspond to the HS bounds.
#
# ### Original Figure 8
# 
#
# ## Figure 9
#
#
# ### Calculation
# Was done above together with Figure 7.
# ### Plot
# +
# PLOT GASSMANN RELATIONS
fig9 = plt.figure(9)
plt.axvline(data2['vp_s'], linewidth=1, color='k')
plt.axvline(data2['vp_f'], linewidth=1, color='k')
plt.plot(vel_ghe2, 1000./rho_b2, 'b--', linewidth=1)
plt.plot(vp_b2[1:], 1000./rho_ghe2[1:], 'b-', label='Hermance')
plt.plot(vel_gcr2, 1000./rho_b2, 'g--', linewidth=1)
plt.plot(vp_b2[1:], 1000./rho_gcr2[1:], 'g-', label='CRIM')
plt.plot(vel_gss2, 1000./rho_b2, 'r--', linewidth=1)
plt.plot(vp_b2[1:], 1000./rho_gss2[1:], 'r-', label='Self-similar')
plt.plot(vel_ghm2, 1000./rho_b2, 'm--')
plt.plot(vp_b2[1:], 1000./rho_ghm2[1:], 'm-', label='HS-')
plt.plot(vel_ghp2, 1000./rho_b2, 'y--')
plt.plot(vp_b2[1:], 1000./rho_ghp2[1:], 'y-', label='HS+')
plt.legend(loc=5)
plt.text(.2, .93, 'Gassmann relations')
plt.title("Carcione et al., 2007, Figure 9")
plt.xlabel("Velocity (km/s)")
plt.ylabel("Conductivity (mS/m)")
plt.axis([0.0, 6.0, 0.0, 1])
plt.show()
# -
# Figure 9. Cross-property relations for different conductivity models of the reservoir (sandstone saturated with oil), combined with the Gassmann equation. The dashed lines correspond to the HS bounds.
#
# ### Original Figure 9
#
# 
| carcione-etal-2007/Carcione-etal-2007_Figures-6-9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1 – Fibonacci numbers
# The goal of this exercise is to cythonize a function.
#
# Use `%cython` and `%cython -a` to do type annotations. Measure speedup with `%timeit`.
# +
def fib(n):
"""Calculate Fibonacci number n.
https://en.wikipedia.org/wiki/Fibonacci_number
"""
a, b = 0, 1
for i in range(n):
a, b = b, a + b
return a
fib(1000)
# -
# %timeit fib(10_000)
# %load_ext cython
# +
# your code here
# -
# ## Is the "cythonized" function correct?
#
# Cythonized code uses numbers that have fixed width. For a sufficiently large number, the result will be wrong. Find out the smallest number for which an incorrect result is returned.
# ## Using floating point numbers
#
# "Cythonize" the function again, this time using floating point numbers (`double`) to hold the partial results. Does this work better? Is the result exact?
| cython-fibbo/exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import IPython
import os
import json
import random
import numpy as np
import requests
from io import BytesIO
import base64
from math import trunc
from PIL import Image as PILImage
from PIL import ImageDraw as PILImageDraw
# +
class CocoDataset():
def __init__(self, annotation_path, image_dir):
self.annotation_path = annotation_path
self.image_dir = image_dir
self.colors = colors = ['blue', 'purple', 'red', 'green', 'orange', 'salmon', 'pink', 'gold',
'orchid', 'slateblue', 'limegreen', 'seagreen', 'darkgreen', 'olive',
'teal', 'aquamarine', 'steelblue', 'powderblue', 'dodgerblue', 'navy',
'magenta', 'sienna', 'maroon']
json_file = open(self.annotation_path)
self.coco = json.load(json_file)
json_file.close()
# self.process_info()
# self.process_licenses()
self.process_categories()
self.process_images()
self.process_segmentations()
# print(self.segmentations)
def display_info(self):
print('Dataset Info:')
print('=============')
for key, item in self.info.items():
print(' {}: {}'.format(key, item))
requirements = [['description', str],
['url', str],
['version', str],
['year', int],
['contributor', str],
['date_created', str]]
for req, req_type in requirements:
if req not in self.info:
print('ERROR: {} is missing'.format(req))
elif type(self.info[req]) != req_type:
print('ERROR: {} should be type {}'.format(req, str(req_type)))
print('')
def display_licenses(self):
print('Licenses:')
print('=========')
requirements = [['id', int],
['url', str],
['name', str]]
for license in self.licenses:
for key, item in license.items():
print(' {}: {}'.format(key, item))
for req, req_type in requirements:
if req not in license:
print('ERROR: {} is missing'.format(req))
elif type(license[req]) != req_type:
print('ERROR: {} should be type {}'.format(req, str(req_type)))
print('')
print('')
def display_categories(self):
print('Categories:')
print('=========')
for sc_key, sc_val in self.super_categories.items():
print(' super_category: {}'.format(sc_key))
for cat_id in sc_val:
print(' id {}: {}'.format(cat_id, self.categories[cat_id]['name']))
print('')
def display_image(self, image_id, show_polys=True, show_bbox=True, show_crowds=True, use_url=False):
print('Image:')
print('======')
if image_id == 'random':
image_id = random.choice(list(self.images.keys()))
# Print the image info
image = self.images[image_id]
for key, val in image.items():
print(' {}: {}'.format(key, val))
# Open the image
if use_url:
image_path = image['coco_url']
response = requests.get(image_path)
image = PILImage.open(BytesIO(response.content))
else:
image_path = os.path.join(self.image_dir, image['file_name'])
image = PILImage.open(image_path)
buffer = BytesIO()
image.save(buffer, format='PNG')
buffer.seek(0)
data_uri = base64.b64encode(buffer.read()).decode('ascii')
image_path = "data:image/png;base64,{0}".format(data_uri)
# Calculate the size and adjusted display size
max_width = 600
image_width, image_height = image.size
adjusted_width = min(image_width, max_width)
adjusted_ratio = adjusted_width / image_width
adjusted_height = adjusted_ratio * image_height
# Create list of polygons to be drawn
polygons = {}
bbox_polygons = {}
rle_regions = {}
poly_colors = {}
print(' segmentations ({}):'.format(len(self.segmentations[image_id])))
for i, segm in enumerate(self.segmentations[image_id]):
polygons_list = []
if segm['iscrowd'] != 0:
# Gotta decode the RLE
px = 0
x, y = 0, 0
rle_list = []
for j, counts in enumerate(segm['segmentation']['counts']):
if j % 2 == 0:
# Empty pixels
px += counts
else:
# Need to draw on these pixels, since we are drawing in vector form,
# we need to draw horizontal lines on the image
x_start = trunc(trunc(px / image_height) * adjusted_ratio)
y_start = trunc(px % image_height * adjusted_ratio)
px += counts
x_end = trunc(trunc(px / image_height) * adjusted_ratio)
y_end = trunc(px % image_height * adjusted_ratio)
if x_end == x_start:
# This is only on one line
rle_list.append({'x': x_start, 'y': y_start, 'width': 1 , 'height': (y_end - y_start)})
if x_end > x_start:
# This spans more than one line
# Insert top line first
rle_list.append({'x': x_start, 'y': y_start, 'width': 1, 'height': (image_height - y_start)})
# Insert middle lines if needed
lines_spanned = x_end - x_start + 1 # total number of lines spanned
full_lines_to_insert = lines_spanned - 2
if full_lines_to_insert > 0:
full_lines_to_insert = trunc(full_lines_to_insert * adjusted_ratio)
rle_list.append({'x': (x_start + 1), 'y': 0, 'width': full_lines_to_insert, 'height': image_height})
# Insert bottom line
rle_list.append({'x': x_end, 'y': 0, 'width': 1, 'height': y_end})
if len(rle_list) > 0:
rle_regions[segm['id']] = rle_list
else:
# Add the polygon segmentation
for segmentation_points in segm['segmentation']:
segmentation_points = np.multiply(segmentation_points, adjusted_ratio).astype(int)
polygons_list.append(str(segmentation_points).lstrip('[').rstrip(']'))
polygons[segm['id']] = polygons_list
if i < len(self.colors):
poly_colors[segm['id']] = self.colors[i]
else:
poly_colors[segm['id']] = 'white'
bbox = segm['bbox']
bbox_points = [bbox[0], bbox[1], bbox[0] + bbox[2], bbox[1],
bbox[0] + bbox[2], bbox[1] + bbox[3], bbox[0], bbox[1] + bbox[3],
bbox[0], bbox[1]]
bbox_points = np.multiply(bbox_points, adjusted_ratio).astype(int)
bbox_polygons[segm['id']] = str(bbox_points).lstrip('[').rstrip(']')
# Print details
print(' {}:{}:{}'.format(segm['id'], poly_colors[segm['id']], self.categories[segm['category_id']]))
# Draw segmentation polygons on image
html = '<div class="container" style="position:relative;">'
html += '<img src="{}" style="position:relative;top:0px;left:0px;width:{}px;">'.format(image_path, adjusted_width)
html += '<div class="svgclass"><svg width="{}" height="{}">'.format(adjusted_width, adjusted_height)
if show_polys:
for seg_id, points_list in polygons.items():
fill_color = poly_colors[seg_id]
stroke_color = poly_colors[seg_id]
for points in points_list:
html += '<polygon points="{}" style="fill:{}; stroke:{}; stroke-width:1; fill-opacity:0.5" />'.format(points, fill_color, stroke_color)
if show_crowds:
for seg_id, rect_list in rle_regions.items():
fill_color = poly_colors[seg_id]
stroke_color = poly_colors[seg_id]
for rect_def in rect_list:
x, y = rect_def['x'], rect_def['y']
w, h = rect_def['width'], rect_def['height']
html += '<rect x="{}" y="{}" width="{}" height="{}" style="fill:{}; stroke:{}; stroke-width:1; fill-opacity:0.5; stroke-opacity:0.5" />'.format(x, y, w, h, fill_color, stroke_color)
if show_bbox:
for seg_id, points in bbox_polygons.items():
fill_color = poly_colors[seg_id]
stroke_color = poly_colors[seg_id]
html += '<polygon points="{}" style="fill:{}; stroke:{}; stroke-width:1; fill-opacity:0" />'.format(points, fill_color, stroke_color)
html += '</svg></div>'
html += '</div>'
html += '<style>'
html += '.svgclass { position:absolute; top:0px; left:0px;}'
html += '</style>'
return html
def process_info(self):
self.info = self.coco['info']
def process_licenses(self):
self.licenses = self.coco['licenses']
def process_categories(self):
self.categories = {}
self.super_categories = {}
for category in self.coco['categories']:
cat_id = category['id']
super_category = category['supercategory']
# Add category to the categories dict
if cat_id not in self.categories:
self.categories[cat_id] = category
else:
print("ERROR: Skipping duplicate category id: {}".format(category))
# Add category to super_categories dict
if super_category not in self.super_categories:
self.super_categories[super_category] = {cat_id} # Create a new set with the category id
else:
self.super_categories[super_category] |= {cat_id} # Add category id to the set
def process_images(self):
self.images = {}
for image in self.coco['images']:
image_id = image['id']
if image_id in self.images:
print("ERROR: Skipping duplicate image id: {}".format(image))
else:
self.images[image_id] = image
def process_segmentations(self):
self.segmentations = {}
for segmentation in self.coco['annotations']:
image_id = segmentation['image_id']
if image_id not in self.segmentations:
self.segmentations[image_id] = []
self.segmentations[image_id].append(segmentation)
# +
annotation_path = '/Users/mackim/datasets/garment_images/via_project_coco.json'
image_dir = '/Users/mackim/datasets/garment_images/images'
coco_dataset = CocoDataset(annotation_path, image_dir)
# coco_dataset.display_info()
# coco_dataset.display_licenses()
coco_dataset.display_categories()
# -
html = coco_dataset.display_image(400, use_url=False)
IPython.display.HTML(html)
with open(annotation_path) as json_file:
lines = json_file.readlines()
for line in lines:
print(line)
| coco_image_viewer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding Gamma and Beta
#
# This notebook is for calculating the gamma and beta correction factors from multiple loaded datasets. It is possible to find these parameters from a single data set so long as the distribution of apparent FRET efficiencies is dominated by a breadth of real underlying distances, however in this case we load multiple data sets each containing a single FRET state.
#
# Gamma describes the inequality between how well the donor and acceptor fluorophores can be detected by your system, whereas beta describes the inequality between how well they can be excited. They are both found by an equation which describes the slope and height (in S) of the doubly labelled population.
#
# You should find the alpha and delta factors first before this step.
# # Import packages
from fretbursts import *
sns = init_notebook()
import lmfit
import phconvert
import os
from fretbursts.burstlib_ext import burst_search_and_gate
# # Name and Load in data
#
# Name the data files, note that it will look for files starting from the folder this notebook is in.
files = ["definitiveset/1a1.hdf5", "definitiveset/1a2.hdf5",
"definitiveset/1b1.hdf5", "definitiveset/1b2.hdf5",
"definitiveset/1c1.hdf5", "definitiveset/1c2.hdf5"]
# Set the spectral cross talk parameters, load them, and sort them
# +
alpha = 0.081
delta = 0.076
datasets = []
for file in files:
datasets.append(loader.photon_hdf5(file))
for dataset in datasets:
dataset.leakage = alpha
dataset.dir_ex = delta
for i in range(0, len(dataset.ph_times_t)): #sorting code
indices = dataset.ph_times_t[i].argsort()
dataset.ph_times_t[i], dataset.det_t[i] = dataset.ph_times_t[i][indices], dataset.det_t[i][indices]
# -
datasets
# # Apply alternation cycle
for dataset in datasets:
#dataset.add(det_donor_accept = (0, 1),
#alex_period = 10000,
#offset = 0,
#D_ON = (0, 4500),
#A_ON = (5000, 9500))
loader.alex_apply_period(dataset)
# # Background Estimation
# We background correct all datasets as before
threshold = 1500
recalctime = 300
for dataset in datasets:
dataset.calc_bg(bg.exp_fit, time_s = recalctime, tail_min_us=(threshold))
dplot(dataset, hist_bg, show_fit=True)
# Now let's find bursts and plot them. We will use a dual channel burst search this time as we are looking for strictly doubly labelled bursts.
colourscheme= "viridis_r"
burstsets = []
for dataset in datasets:
dataset = burst_search_and_gate(dataset, F=45, m=10, mute=True)
dataset = dataset.select_bursts(select_bursts.size, th1=50)
burstsets.append(dataset)
for burstset in burstsets:
alex_jointplot(burstset, cmap=colourscheme, marginal_color=20, vmax_fret=False)
# Now let's extract the E and S positions from these plots with gaussian fits. We could plot the fitted curves, but here the graphs are not shown to save space.
def ESgauss(dataset):
dataset.E_fitter.fit_histogram(model=mfit.factory_gaussian(), verbose=False, pdf=False)
params = dataset.E_fitter.params
Efit = params.to_dict()
E = Efit['center'][0]
dataset.S_fitter.fit_histogram(model=mfit.factory_gaussian(), verbose=False, pdf=False)
params = dataset.S_fitter.params
Sfit = params.to_dict()
S = Sfit['center'][0]
return(E, S)
x = []
y = []
for dataset in burstsets:
E, S = ESgauss(dataset)
x.append(E)
y.append(S)
# Now we can fit to the equation:
#
# S = 1/(1+bg+(1-g)*b*E)
model = lmfit.models.ExpressionModel('1 / ( 1 + b * g + ( 1 - g ) * b * E )',
independent_vars=["E"], nan_policy = "omit")
params = model.make_params(b=0.5, g=1.0)
fit = model.fit(y, params, E=E)
fit.plot_fit(0)
print(fit.fit_report())
pars = fit.params
beta = pars['b']
beta=beta.value
gamma = pars['g']
gamma=gamma.value
print("gamma = ", gamma)
print("beta = ", beta)
| Jnotebooks/Correction Factor Finder Gamma-Beta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('../../src/')
import json
import os
from pathlib import Path
# ## HisDB
import experiment.data as exp
import datasets.divahisdb as hisdb
env = exp.Environment()
img_path = Path('../../doc/figures/')
dataset = hisdb.HisDBDataset(env.dataset(exp.Datasets.diva.value))
samples = [dataset[2], dataset[22], dataset[42]]
# !pwd
scale = 0.2
xpos, ypos, size = 600,1200, 600
box = (xpos,ypos,xpos+size,ypos+size)
for idx, data in enumerate(samples):
img, gt = data
fig = img.resize((int(img.width * scale), int(img.height * scale)))
name = img_path / 'datasets' / ('HisDBSample' + str(idx) + '.jpeg')
cropped = img_path / 'datasets' / ('HisDBSampleBox' + str(idx) + '.jpeg')
fig.save(name.open('wb'))
img.crop(box).save(cropped.open('wb'))
print(name, cropped)
# ## Ground truth
dataset = hisdb.HisDBDataset(env.dataset(exp.Datasets.diva.value), gt=True)
original, gt = dataset[5]
gt_color = Image.fromarray(hisdb.color_gt(gt))
gt_color = Image.fromarray(hisdb.color_gt(gt))
scale = 0.3
m, n, size = 0, 3000, 2000
for idx, gt_img in enumerate([original, gt_color]):
gt_img = gt_img.crop(box=[m,n,gt_img.width,n+size,])
gt_img = gt_img.resize((int(gt_img.width * scale), int(gt_img.height * scale)))
gt_img.save(str(img_path/ 'datasets' / 'gt_example{}.jpeg'.format(idx)))
# gt_img
# ## HisDB Processed
# %matplotlib inline
from PIL import Image
import matplotlib.pyplot as plt
import skimage.io
import skimage.segmentation as seg
from skimage.util import img_as_float, img_as_ubyte
import numpy as np
split = hisdb.Splits.training.value
dataset = hisdb.Processed(env.dataset(exp.Datasets.processd.value), split=split, load=['img', 'slic', 'tiles', 'y', 'meta'])
i = 0
img, slic, tiles, y, meta = dataset[i]
marked = Image.fromarray(img_as_ubyte(np.array(seg.mark_boundaries(img, slic,color=(1,0,0)))))
p = 300
box = (p,p,p+500,p+500)
marked.crop(box=box).save(img_path / 'mark_boundaries.jpg')
# # !pwd
marked.crop(box=box)
plt.imshow(slic,cmap='flag')
colored = np.zeros(slic.shape,dtype=np.int)
for idx in range(len(y)):
tile_id = slic[meta[idx][0],meta[idx][1]]
gt = y[idx]
colored[np.where(slic == tile_id)] = hisdb.LABEL_DICT[gt]
col_img = Image.fromarray(hisdb.color_gt(colored))
marked = Image.fromarray(img_as_ubyte(np.array(seg.mark_boundaries(col_img, slic,color=(1,0,0)))))
col_img = col_img.crop(box)
col_img
alpha = 0.2
Image.fromarray(((np.array(img.crop(box=box))*(1-alpha) + alpha*np.array(marked.crop(box))) ).astype(np.ubyte)).save(img_path/'colored_slic.jpg')
# marked.crop(box)
slic.shape
| notebooks/figures/HisDBExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import shutil
import zipfile
from os.path import join, getsize
def unzip_file(zip_src, dst_dir):
r = zipfile.is_zipfile(zip_src)
if r:
fz = zipfile.ZipFile(zip_src, 'r')
for file in fz.namelist():
fz.extract(file, dst_dir)
else:
print('This is not zip')
zip_src = '/mnt/马总/816nojiequ_zq.zip'
dst_dir = '/mnt/马总/'
unzip_file(zip_src, dst_dir)
# +
from nets import *
import torch
import numpy as np
import torchvision
import os
from torchvision import transforms,datasets,models
from torch.utils.data import DataLoader
import torch.nn as nn
from tools import save_model, show_accuracy, show_loss, show_img
from collections import Counter
import matplotlib.pyplot as plt
from datasets import LoadData
from torchsummary import summary
from tqdm import tqdm
import torch.nn.functional as F
import pretrainedmodels
if torch.cuda.is_available():
device = 'cuda'
else:
device = 'cpu'
LR = 0.0001
EPOCHS = 10
BATCH_SIZE = 32
traindataSize = 198776
testdataSize = 2008
# 训练和验证
criteration = nn.CrossEntropyLoss()
def train(model, device, dataset, optimizer, epoch):
model.train()
correct = 0
for i, (x, y) in tqdm(enumerate(dataset)):
x, y = x.to(device), y.to(device)
optimizer.zero_grad()
output = model(x)
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(y.view_as(pred)).sum().item()
loss = F.cross_entropy(output, y)
#loss = nn.CrossEntropyLoss(output,y)
loss.backward()
optimizer.step()
print("Epoch {} Loss {:.4f} Accuracy {}/{} ({:.0f}%)".format(epoch, loss, correct, traindataSize,
100 * correct / traindataSize))
def vaild(model, device, dataset):
model.eval()
correct = 0
with torch.no_grad():
for i, (x, y) in tqdm(enumerate(dataset)):
x, y = x.to(device), y.to(device)
output = model(x)
#loss = nn.CrossEntropyLoss(output, y)
loss = F.cross_entropy(output, y)
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(y.view_as(pred)).sum().item()
print(
"Test Loss {:.4f} Accuracy {}/{} ({:.0f}%)".format(loss, correct, testdataSize, 100. * correct / testdataSize))
if __name__ == '__main__':
print('------Starting------')
# 确定是否使用GPU
testloader, trainloader = LoadData(BATCH_SIZE)
# print(train_dataset.shape)
print('------Dataset initialized------')
#定义模型
#model = pretrainedmodels.senet154(num_classes=1000, pretrained=None)
model = pretrainedmodels.se_resnet50(num_classes=1000, pretrained=None)
#model = pretrainedmodels.pnasnet5large(num_classes=1001, pretrained=None)
#model = models.resnet34(pretrained=True)
#加载参数
#model.load_state_dict(torch.load('/mnt/马总/senet154-c7b49a05.pth'))
model.load_state_dict(torch.load('/mnt/马总/se_resnet50-ce0d4300.pth'))
#model.load_state_dict(torch.load('/mnt/马总/pnasnet5large-bf079911.pth'))
#增加全连接层
model.fc = nn.Sequential(
nn.Linear(512,2)
)
model.to(device)
summary(model,(3,224,224))
#optimizer = torch.optim.SGD(model.parameters(), lr = LR, momentum = 0.09)
optimizer = torch.optim.Adam(model.parameters(), lr = LR)
for epoch in range(1, EPOCHS + 1):
train(model, device, trainloader, optimizer, epoch)
vaild(model, device, testloader)
save_model(model.state_dict(),'MODELnojiequ_zq','modelsenet_99_1_10_32adam-0.00005.pt')
# print('------Network load successfully------')
# # trainset 第一个维度控制第几张图片,第二个维度控制是data还是label
#
#
# Loss_crossEntropy = nn.CrossEntropyLoss().to(device)
# print('------Loss created------')
#
# optimizer= torch.optim.RMSprop(newmodel.parameters(), lr=1e-3)
# train_loss_history = []
# train_acc_history = []
# test_loss_history = []
# test_acc_history = []
# MAX = 0.0
# tag = 0
# for epoch in range(EPOCHS):
# epoch_train_loss = 0.0
# epoch_train_acc = 0.0
# epoch_test_loss = 0.0
# epoch_test_acc = 0.0
# # train 阶段
# train_step = 0
# for i, data in enumerate(trainloader):
# correct = 0
# train_step += 1
# images, labels = data
# images = images.to(device)
# labels = labels.to(device)
# output = newmodel(images)
# # 梯度清零
# optimizer.zero_grad()
# # print(output)
# # 加上正则化的loss
# loss = Loss_crossEntropy(output, labels.to(device))
# loss.backward()
# optimizer.step()
# _, predicted = torch.max(output.data, 1)
# correct += (predicted == labels.to(device)).sum().item()
# acc = correct / BATCH_SIZE
#
# epoch_train_loss += loss.item()
# epoch_train_acc += acc
#
# print('Train-%d-%d, loss: %.3f, acc: %.3f' % (epoch, i, loss, acc))
# train_loss_history.append(epoch_train_loss / train_step)
# train_acc_history.append(epoch_train_acc / train_step)
# # test 阶段
# # 不跟踪梯度
# with torch.no_grad():
# test_step = 0
# for test_data in testloader:
# test_step += 1
# test_correct = 0
# test_images, test_labels = test_data
#
# test_output = newmodel(test_images.to(device))
# test_loss = Loss_crossEntropy(test_output, test_labels.to(device))
# _, test_predicted = torch.max(test_output.data, 1)
# test_correct += (test_predicted == test_labels.to(device)).sum().item()
# test_acc = test_correct / 64
#
# epoch_test_loss += test_loss.item()
# epoch_test_acc += test_acc
# print('Test epoch: %d, loss: %.3f, acc: %.3f' % (epoch, test_loss, test_acc))
# test_loss_history.append(epoch_test_loss / test_step)
# test_acc_history.append(epoch_test_acc / test_step)
# save_model(newmodel.state_dict(),'SE_MobileNet','model.pt')
# show_loss(train_loss_history, test_loss_history)
# show_accuracy(train_acc_history, test_acc_history)
# +
import numpy as np
import matplotlib as plt
import cv2
import tifffile as tiff
import torch
from torchvision import transforms,datasets,models
import os
from skimage import io, transform
import torch.nn as nn
from PIL import Image
from collections import OrderedDict
from torchsummary import summary
from nets import *
import torch
import numpy as np
import torchvision
import os
from torchvision import transforms,datasets,models
from torch.utils.data import DataLoader
import torch.nn as nn
from tools import save_model, show_accuracy, show_loss, show_img
from collections import Counter
import matplotlib.pyplot as plt
from datasets import LoadData
from torchsummary import summary
from tqdm import tqdm
import torch.nn.functional as F
from collections import OrderedDict
import pretrainedmodels
from skimage import transform,data
#model = pretrainedmodels.senet154(num_classes=1000, pretrained=None)
model = pretrainedmodels.se_resnet50(num_classes=1000, pretrained=None)
#model = models.resnet34(pretrained=True)
model.fc = nn.Sequential(
nn.Linear(512,2)
)
model.load_state_dict(torch.load('/mnt/马总/MODELnojiequenvi/modelsenet_9_1_10_32adam-0.00005.pt',map_location='cpu'))
model.eval()
# test
# k是19个地区/文件夹
for k in range(19):
# 预先产生16*16的数组,用于存放单个地区的预测结果
a = np.ones((16, 16))
for i in range(16):
for j in range(16):
# 读入小图
img_path = "816_L2A_Val_predict/" + str(k + 1) + "/" + str(i * 16 + j + 1) + ".tif"
# 用tiff读入数据,转为numpy数组
# img = np.array(tiff.imread(img_path))
# img = torch.tensor(img)
# 将数组转为224*224
# img = transform.resize(img, (224, 224))
tf = transforms.Compose([
#lambda x: Image.open(x).convert("RGB"), # string path => image data
lambda x: transform.resize(np.array(tiff.imread(x)), (224, 224)),
#transforms.Resize((int(224 * 1.25), int(224 * 1.25))),
#transforms.RandomRotation(15),
#transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
])
img = tf(img_path)
img = img.type(torch.FloatTensor)
img = np.expand_dims(img, axis=0) # 为batch添加第四维
# print(img.shape)
img = torch.tensor(img)
# print(model(img))
# 调用模型,输出预测结果
label = np.argmax(model(img).detach().numpy())
if label == 0:
# print('有人')
a[i, j] = 1
else:
# print('没人')
a[i, j] = 0
# 讲结果转为uint8
a = a.astype(np.uint8)
path = '预测结果senet'
if not os.path.exists(path):
os.mkdir(path)
# 保存结果
print(path+'/' + str(k + 1) + '.tif', a)
tiff.imsave(path+'/' + str(k + 1) + '.tif', a)
# plt.imshow(a, cmap='gray')
# +
import numpy as np
import matplotlib as plt
import cv2
import tifffile as tiff
import torch
from torchvision import transforms,datasets,models
import os
from skimage import io, transform
import torch.nn as nn
from PIL import Image
from collections import OrderedDict
from torchsummary import summary
from nets import *
import torch
import numpy as np
import torchvision
import os
from torchvision import transforms,datasets,models
from torch.utils.data import DataLoader
import torch.nn as nn
from tools import save_model, show_accuracy, show_loss, show_img
from collections import Counter
import matplotlib.pyplot as plt
from datasets import LoadData
from torchsummary import summary
from tqdm import tqdm
import torch.nn.functional as F
from collections import OrderedDict
#model.load_state_dict(torch.load('0.81.pth',map_location='cpu'))
#model.load_state_dict(torch.jit.load('/mnt/马总/0.81.pth',map_location='cpu'))
model_dir = '0.81-1.pth'
dict = torch.load(model_dir)
older_val = dict['fc.bias']
# 修改参数名
dict['fc.0.bias'] = dict.pop('fc.bias')
torch.save(dict, '0.81-2.pth')
# +
import torch
#from Resnet_remote.model_building import resnet34
from PIL import Image
from torchvision import transforms
import matplotlib.pyplot as plt
import json
import numpy as np
import os
from skimage import io
from torchvision import transforms,datasets,models
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print("using {} device.".format(device))
data_transform = transforms.Compose(
[
transforms.Resize(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])])
X = np.zeros((1, 800, 800, 3))
X_input = np.zeros((256, 50, 50, 3))
model = models.resnet34(pretrained=True)
model.fc = nn.Sequential(
nn.Linear(512,2)
)
#model = resnet34(num_classes=2)
model_weight_path = "/mnt/马总/0.81/0.81.pth"
model.load_state_dict(torch.load(model_weight_path, map_location=device))
model.eval()
path = '/mnt/马总/0.81/L2A_Val_predict'
land_types = os.listdir(path)
tif_out = np.zeros((256, 1)).astype(np.uint8)
for tidx, lt in enumerate(land_types):
for i in range(256):
img = Image.open(path + '/' + lt + '/' + str(i+1) + '.tif')
img = data_transform(img)
img = torch.unsqueeze(img, dim=0)
with torch.no_grad():
output = torch.squeeze(model(img))
predict = torch.softmax(output, dim=0)
predict_cla = torch.argmax(predict).numpy()
tif_out[i] = predict_cla
print(tif_out)
tif_result = tif_out.reshape(16, 16).astype(np.uint8)
print(tif_result)
io.imsave('/mnt/马总/0.81/' + lt + '.tif', tif_result)
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# pg 58 This will evaluate the prediction from section 2.1
from sklearn.datasets import load_boston
from sklearn.linear_model import LinearRegression
boston = load_boston()
lr = LinearRegression()
lr.fit(boston.data, boston.target)
predictions = lr.predict(boston.data)
import matplotlib.pyplot as plt
import numpy as np
f = plt.figure(figsize=(7,5))
ax = f.add_subplot(111)
ax.hist(boston.target - predictions, bins=50)
ax.set_title('Histogram of Residuals.')
# look at the mean of the residuals (closer to 0 is best)
np.mean(boston.target - predictions)
# +
# Look at the Q-Q plot.
# -
from scipy.stats import probplot
f = plt.figure(figsize=(7,5))
ax = f.add_subplot(111)
probplot(boston.target - predictions, plot=ax)
ax
# +
# Created Mean Squared Error (MSE) and Mean Absolute Deviation(MAD)
# in msemad.py for this next part and later in the book.
# -
from msemad import MSE, MAD
MSE(boston.target, predictions)
MAD(boston.target, predictions)
n_bootstraps = 100
len_boston = len(boston.target)
subsample_size = np.int(0.5*len_boston)
subsample = lambda: np.random.choice(np.arange(0, len_boston), size=subsample_size)
coefs = np.ones(n_bootstraps)
for i in range(n_bootstraps):
subsample_idx = subsample()
subsample_X = boston.data[subsample_idx]
subsample_y = boston.target[subsample_idx]
lr.fit(subsample_X, subsample_y)
coefs[i] = lr.coef_[0]
import matplotlib.pyplot as plt
f = plt.figure(figsize=(7,5))
ax = f.add_subplot(111)
ax.hist(coefs, bins=50)
ax.set_title("Histogram of the lr.coef_[0]")
np.percentile(coefs, [2.5, 97.5])
| Chapter 2/2.2 Evaluating the Linear Regression Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cnaseeb/AIHub/blob/master/Handwriting_Recognition.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5WMO5Zq73y9O" colab_type="text"
# Exercise 2
# In the course you learned how to do classificaiton using Fashion MNIST, a data set containing items of clothing. There's another, similar dataset called MNIST which has items of handwriting -- the digits 0 through 9.
# Write an MNIST classifier that trains to 99% accuracy or above, and does it without a fixed number of epochs -- i.e. you should stop training once you reach that level of accuracy.
# Some notes:
# It should succeed in less than 10 epochs, so it is okay to change epochs= to 10, but nothing larger
# When it reaches 99% or greater it should print out the string "Reached 99% accuracy so cancelling training!"
# If you add any additional variables, make sure you use the same names as the ones used in the class
# I've started the code for you below -- how would you finish it?
# + id="BRG5iYd331Jg" colab_type="code" colab={}
import tensorflow as tf
from os import path, getcwd, chdir
# DO NOT CHANGE THE LINE BELOW. If you are developing in a local
# environment, then grab mnist.npz from the Coursera Jupyter Notebook
# and place it inside a local folder and edit the path to that location
path = f"{getcwd()}/../Downloads/mnist.npz" #specify the correct path
# + id="70Nzs3Ta34Uq" colab_type="code" colab={}
ACCURACY_THRESHOLD = 0.99
# GRADED FUNCTION: train_mnist
def train_mnist():
# Please write your code only where you are indicated.
# please do not remove # model fitting inline comments.
#YOUR CODE SHOULD START HERE
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('acc')> ACCURACY_THRESHOLD):
print("\nReached 99% accuracy so cancelling training!")
self.model.stop_training = True
# YOUR CODE SHOULD END HERE
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data(path=path)
# YOUR CODE SHOULD START HERE
x_train, x_test = x_train / 255.0, x_test / 255.0
callbacks = myCallback()
# YOUR CODE SHOULD END HERE
model = tf.keras.models.Sequential([
# YOUR CODE SHOULD START HERE
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation = tf.nn.relu),
tf.keras.layers.Dense(10, activation = tf.nn.softmax)
# YOUR CODE SHOULD END HERE
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# model fitting
history = model.fit(# YOUR CODE SHOULD START HERE
x_train, y_train, epochs=10, callbacks=[callbacks]
# YOUR CODE SHOULD END HERE
)
# model fitting
return history.epoch, history.history['acc'][-1]
# + id="-ZdL31Y13zuj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 324} outputId="e392e191-9c12-42bc-ea80-db8934d3bd96"
train_mnist()
# + id="opKdb3N03_od" colab_type="code" colab={}
| Handwriting_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 1 Day 1 B7
#
# Landing palne safely
# +
# Landing plane safely
print("Please enter the landing altitude for plane")
speed=int(input())
if speed<=1000:
print("Plane is safe to land")
elif speed<=5000:
print("Bring down the plane to 1000 ft")
else:
print("Turn it over and attempt later")
# -
| Assignment 1 Day 1 B7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf22
# language: python
# name: tf22
# ---
# ### Inpainting with Posterior Analysis
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 411, "status": "ok", "timestamp": 1568330149594, "user": {"displayName": "", "photoUrl": "", "userId": "18062987068597777273"}, "user_tz": 420} id="D-Fe5G8m1FTC" outputId="481e97a2-895c-40e1-c3f8-aaac7fb27b5d"
import os
# %pylab inline
import pickle
# + colab={} colab_type="code" id="sZkaGpCR1kVS"
PARAMS_PATH = '../../params'
param_file = 'params_fmnist_-1_40_infoGAN_AE_best_params_noaugment_full_sigma'
params = pickle.load(open(os.path.join(PARAMS_PATH,param_file+'.pkl'),'rb'))
# + colab={} colab_type="code" id="-AEYmOsH1FTI"
import tensorflow.compat.v1 as tf
tf.disable_eager_execution()
import tensorflow_probability as tfp
import tensorflow_hub as hub
tfd = tfp.distributions
tfb = tfp.bijectors
# + colab={} colab_type="code" id="8puPFE90P0aD"
generator_path = os.path.join(params['module_dir'],'PAE_decoder')
encoder_path = os.path.join(params['module_dir'],'PAE_encoder')
nvp_path = os.path.join(params['module_dir'],'PAE_flow')
# + colab={} colab_type="code" id="nFzYYSxY1FTL"
from pae.load_data import load_fmnist
load_func = load_fmnist
x_train, y_train, x_valid, y_valid, x_test, y_test = load_func(params['data_dir'],flatten=False)
if np.all(x_test)==None:
x_test=x_valid
def add_noise(x,sigma=0.1):
nn = np.random.normal(size=np.shape(x))
x = x+nn*sigma
return x
x_train = add_noise(x_train)/256.-0.5
x_test = add_noise(x_test)/256.-0.5
x_valid = add_noise(x_valid)/256.-0.5
# -
for ii in [1470]:
plt.imshow(x_test[ii,:,:,0],'gray')
plt.axis('off')
plt.show()
# + colab={} colab_type="code" id="ft_jIh-W1FTN"
data_dim = 28*28
data_size = params['batch_size']
sigma_n = (params['full_sigma']+1e-4).reshape((1,28*28,1)).astype(np.float32)
noise_n = sigma_n
hidden_size = params['latent_size']
n_channels = 1
seed = 767
print(params['batch_size'])
# settings for reconstruction with uncorrupted data
# corr_type = 'none'
# num_mnist = 6
# label = 'uncorrupted'
# noise_level = 0.0
# num_comp = 2
# settings for reconstrcution with rectangular mask
corr_type = 'noise+mask'
num_mnist = 3055
label = 'PAE_solidmask_%d'%num_mnist
noise_level = 0.1
num_comp = 32
#settings for reconstruction with sparse mask
# corr_type = 'sparse mask'
# num_mnist = 1
# label = 'sparse95'
# noise_level = 0.
# num_comp = 2
# settings for reconstruction with noise
# corr_type = 'noise'
# num_mnist = 6
# label = 'noise05'
# noise_level = 0.5
# num_comp = 4
# settings for reconstruction with noise and mask
# corr_type = 'noise+mask'
# num_mnist = 6
# label = 'masknoise05'
# noise_level = 0.5
# num_comp = 2
# + colab={} colab_type="code" id="nxcZOE0MLGJ1"
def plot_image(image, save=True, directory='./plots/',filename='plotted_image', title='image',vmin=None,vmax=None, mask=None, cmap='gray', colorbar=False):
if np.any(mask==None):
mask=np.ones((28,28))
mask = np.reshape(mask,(28,28))
plt.figure()
plt.imshow((image).reshape((28,28))*mask,cmap=cmap,vmin=vmin, vmax=vmax)
plt.axis('off')
if colorbar:
plt.colorbar()
if save:
plt.savefig(directory+filename+'.pdf',bbox_inches='tight')
plt.show()
return True
def get_custom_noise(shape, signal_dependent=False, signal =None, sigma_low=0.07, sigma_high=0.22, threshold=0.02 ):
sigma = sigma_n
if signal_dependent:
for ii in range(data_size):
sigma[ii][np.where(signal[ii]<=threshold)]= sigma_low
sigma[ii][np.where(signal[ii]>threshold)]= sigma_high
data_noise = np.ones_like(sigma)*noise_level
sigma = np.sqrt(sigma**2+data_noise**2)
return sigma
def make_corrupted_data(x_true, corr_type='mask'):
mask = np.ones((28,28))
if corr_type=='mask':
minx = 2
maxx = 14
miny = 2
maxy = 12
mask[miny:maxy,minx:maxx]=0.
corr_data = x_true*mask[None,:,:,None]
elif corr_type=='sparse mask':
mask = np.ones(data_dim, dtype=int)
percent = 95
np.random.seed(seed+2)
indices = np.random.choice(np.arange(data_dim), replace=False,size=int(percent/100.*data_dim))
print('precentage masked:', len(indices)/data_dim)
mask[indices] =0
corr_data = x_true*mask[None,:,:]
elif corr_type=='noise':
np.random.seed(seed+2)
noise = np.random.randn(data_dim*data_size)*noise_level
corr_data = x_true+noise
elif corr_type=='noise+mask':
np.random.seed(seed+2)
noise = np.random.randn(data_dim)*noise_level
noise = np.tile(noise,[data_size,1])
print(noise.shape)
noise = noise.reshape(x_true.shape)
minx = 2
maxx = 14
miny = 2
maxy = 12
mask[miny:maxy,minx:maxx]=0.
mask[miny:maxy,minx:maxx]=0.
corr_data = x_true+noise
corr_data = corr_data*mask[None,:,:,None]
elif corr_type=='none':
corr_data = x_true
corr_data = np.expand_dims(corr_data,-1)
mask = mask.flatten()
return corr_data, mask
# -
data_size
# +
plot_path = params['plot_dir']+'/%d/'%num_mnist
if not os.path.isdir(plot_path):
os.makedirs(plot_path)
truth = x_test[num_mnist:num_mnist+1]
truth = np.tile(truth, [params['batch_size'],1,1,1])
plot_image(truth[0], directory=plot_path, filename='truth_%s'%label, title='truth')
print(corr_type, truth.shape)
data, custom_mask = make_corrupted_data(truth, corr_type=corr_type)
print(data.shape)
plot_image(data[3], directory=plot_path, filename='input_data_%s'%label, title='data')
plot_image(custom_mask, directory=plot_path, filename='mask_data_%s'%label, title='mask')
noise = get_custom_noise(data.shape, signal_dependent=False, signal=truth)
plot_image(noise[0], directory=plot_path, filename='noise_%s'%label, title='noise', cmap='summer', colorbar=True)
data = np.reshape(data,(-1,28*28,1))
custom_mask = np.reshape(custom_mask,(28*28))
#noise = noise_n
# + colab={} colab_type="code" id="9TIQArTJHE87"
def fwd_pass(generator,nvp,z,mask):
print(z)
fwd_z = nvp({'z_sample':np.zeros((1,hidden_size)),'sample_size':1, 'u_sample':z},as_dict=True)['fwd_pass']
fwd_z = generator({'z':z},as_dict=True)['x']
gen_z = tf.boolean_mask(tf.reshape(fwd_z,[data_size,data_dim,n_channels]),mask, axis=1)
return gen_z
def get_likelihood(generator,nvp,z,sigma,mask):
gen_z = fwd_pass(generator,nvp,z,mask)
sigma = tf.boolean_mask(sigma,mask, axis=1)
likelihood = tfd.Independent(tfd.MultivariateNormalDiag(loc=gen_z,scale_diag=sigma))
return likelihood
def get_prior():
return tfd.MultivariateNormalDiag(tf.zeros([data_size,hidden_size]), scale_identity_multiplier=1.0, name ='prior')
def get_log_posterior(z,x,generator,nvp,sigma,mask, beta):
likelihood = get_likelihood(generator,nvp,z,sigma,mask)
prior = get_prior()
masked_x = tf.boolean_mask(x,mask, axis=1)
log_posterior = prior.log_prob(z)+likelihood.log_prob(masked_x)
return log_posterior
def get_recon(generator,nvp, z,sigma,mask):
prob = get_likelihood(generator,nvp, z,sigma,mask)
recon= prob.mean()
return recon
def get_hessian(func, z):
hess = tf.hessians(func,z)
hess = tf.gather(hess, 0)
return(tf.reduce_sum(hess, axis = 2 ))
def get_GN_hessian(generator,nvp,z,mask,sigma):
gen_z = fwd_pass(generator,nvp,z,mask)
sigma = tf.boolean_mask(sigma,mask, axis=1)
grad_g = tf.gather(tf.gradients(gen_z/(sigma),z),0)
grad_g2 = tf.einsum('ij,ik->ijk',grad_g,grad_g)
one = tf.linalg.eye(hidden_size, batch_shape=[data_size],dtype=tf.float32)
hess_GN = one+grad_g2
return hess_GN
def compute_covariance(hessian):
hessian = transform_diagonal(hessian, None, 1e-4)
cov = tf.linalg.inv(hessian)
cov = (cov+tf.linalg.transpose(cov))*0.5
return cov
# + colab={} colab_type="code" id="QGyi6PVWx1qd"
def minimize_posterior(initial_value, x, custom_mask, noise, my_sess, annealing =True):
ini = np.reshape(initial_value,[data_size,hidden_size])
my_sess.run(MAP_reset,feed_dict={input_data: x, MAP_ini:ini, mask:custom_mask,sigma_corr:noise})
pos_def = False
posterior_loss = []
for lrate, numiter in zip([1e-1,1e-2,1e-3],[10000,12000,12000]):
print('lrate', lrate)
for jj in range(numiter):
if annealing and lrate==1e-1:
inv_T= np.round(0.5*np.exp(-(1.-jj/numiter)),decimals=1)
else:
inv_T= 1.
_, ll = my_sess.run([opt_op_MAP,loss_MAP],feed_dict={input_data: x, mask:custom_mask, sigma_corr:noise, lr: lrate, inverse_T:inv_T})
posterior_loss.append(ll)
if jj%1000==0:
print('iter', jj, 'loss', ll,r'inverse T', inv_T)
z_value = my_sess.run(MAP,feed_dict={input_data: x, mask:custom_mask, sigma_corr:noise})
eig = my_sess.run(tf.linalg.eigvalsh(hessian[0]),feed_dict={input_data: x, mask:custom_mask,sigma_corr:noise})
hess = my_sess.run(hessian[0],feed_dict={input_data: x, mask:custom_mask,sigma_corr:noise})
hessGN = my_sess.run(hessian_GN[0],feed_dict={input_data: x, mask:custom_mask,sigma_corr:noise})
print('eig', eig)
if np.all(eig>0.):
pos_def = True
loss = ll
plt.figure()
plt.plot(posterior_loss)
plt.ylabel('loss')
plt.xlabel('iteration')
plt.show()
return z_value, loss, pos_def, hess, hessGN
# + colab={} colab_type="code" id="oiAie-wjUcHN"
def get_laplace_sample(num,map_value,x,mymask,noise,my_sess):
my_sess.run(MAP_reset,feed_dict={MAP_ini:map_value})
my_sess.run(update_mu)
my_sess.run(update_TriL,feed_dict={input_data: x, mask: mymask, sigma_corr:noise})
samples=[]
for ii in range(num):
my_sess.run(posterior_sample,feed_dict={input_data: x, sigma_corr:noise})
samples.append(my_sess.run(recon,feed_dict={input_data: x, sigma_corr:noise}))
samples=np.asarray(samples)
return samples
def get_gmm_sample(num,x,mymask,noise,my_sess):
samples=[]
for ii in range(num):
samples.append(my_sess.run(gmm_recon,feed_dict={input_data: x, sigma_corr:noise}))
samples=np.asarray(samples)
return samples
# -
def get_encoded(data, nvp, encoder):
data = tf.reshape(data,(params['batch_size'],28,28,1))
encoded, _ = tf.split(encoder({'x':data},as_dict=True)['z'], 2, axis=-1)
encoded = nvp({'z_sample':encoded,'sample_size':1, 'u_sample':np.zeros((1,hidden_size))},as_dict=True)['bwd_pass']
return encoded
# + colab={} colab_type="code" id="SXhLJToHcp7b"
def plot_samples(samples, mask, title='samples', filename='samples'):
plt.figure()
plt.title(title)
for i in range(min(len(samples),16)):
subplot(4,4,i+1)
imshow(np.reshape(samples[i,:],(28,28)),vmin=-0.2,vmax=1.2, cmap='gray')
axis('off')
plt.savefig(plot_path+filename+'.pdf',bbox_inches='tight')
plt.show()
if corr_type in ['mask', 'sparse mask', 'noise+mask']:
plt.figure()
plt.title('masked'+title)
for i in range(min(len(samples),16)):
subplot(4,4,i+1)
imshow(np.reshape(samples[i,0,:,0]*mask,(28,28)),vmin=-0.2,vmax=1.2, cmap='gray')
axis('off')
plt.savefig(plot_path+filename+'masked.pdf',bbox_inches='tight')
plt.show()
# + colab={} colab_type="code" id="LYAt6f7MQSpa"
def get_random_start_values(num, my_sess):
result=[]
for ii in range(num):
result.append(my_sess.run(get_prior().sample()))
return result
# + colab={} colab_type="code" id="BaFIFXBnQ3o-"
def get_chi2(sigma,data,mean,masking=True, mask=None,threshold=0.02):
if masking:
mask = np.reshape(mask,data.shape)
data = data[np.where(mask==1)]
mean = mean[np.where(mask==1)]
sigma= sigma[np.where(mask==1)]
low = min(sigma.flatten())
high= max(sigma.flatten())
chi2_tot = np.sum((data-mean)**2/sigma**2)
dof_tot = len(np.squeeze(data))
if corr_type not in ['noise','noise+mask']:
chi2_low = np.sum((data[np.where(data<=threshold)]-mean[np.where(data<=threshold)])**2/sigma[np.where(data<=threshold)]**2)
dof_low = len(np.squeeze(data[np.where(data<=threshold)]))
chi2_high= np.sum((data[np.where(data>threshold)]-mean[np.where(data>threshold)])**2/sigma[np.where(data>threshold)]**2)
dof_high = len(np.squeeze(data[np.where(data>threshold)]))
else:
chi2_low = None
dof_low = None
chi2_high= None
dof_high = None
return chi2_tot, dof_tot, chi2_low, dof_low, chi2_high, dof_high, masking
# + colab={} colab_type="code" id="yGtEbpIZ2vhx"
def plot_minima(minima, losses, var):
plt.figure()
plt.title('Minimization result')
plt.plot(np.arange(len(losses)),losses,ls='',marker='o')
plt.xlabel('# iteration')
plt.ylabel('loss')
plt.savefig(plot_path+'minimzation_results_%s.pdf'%(label),bbox_inches='tight')
plt.show()
colors = matplotlib.colors.Normalize(vmin=min(losses), vmax=max(losses))
cmap = matplotlib.cm.get_cmap('Spectral')
var = np.squeeze(var)
plt.figure()
plt.title('value of hidden variables at minima')
for ii in range(len(minima)):
yerr_= np.sqrt(var[ii])
plt.errorbar(np.arange(hidden_size),np.squeeze(minima)[ii], marker='o',ls='', c=cmap(colors(losses[ii])), mew=0, yerr=yerr_, label ='%d'%losses[ii])
plt.legend(ncol=4, loc=(1.01,0))
plt.xlabel('# hidden variable')
plt.ylabel('value')
plt.savefig(plot_path+'hidden_values_at_minima_%s.pdf'%(label),bbox_inches='tight')
plt.show()
# + colab={} colab_type="code" id="qrZDSLzEIKrn"
def get_gmm_parameters(minima, x, noise, mymask, offset):
mu =[]
w =[]
sigma=[]
print(len(minima), num_comp)
for ii in range(num_comp):
# do Laplace approximation around this minimum
mu+=[minima[ii][0]]
sess.run(MAP_reset,feed_dict={MAP_ini:minima[ii]})
sigma+=[sess.run(update_TriL,feed_dict={input_data: x, sigma_corr:noise, mask: mymask})]
# correct weighting of different minima according to El20 procedure, with samples at the maxima and well seperated maxima
logdet = sess.run(tf.linalg.logdet(covariance[0]),feed_dict={input_data: x, sigma_corr:noise, mask: mymask})
logprob = sess.run(nlPost_MAP,feed_dict={input_data: x, sigma_corr:noise, mask: mymask})
w+=[np.exp(0.5*logdet+logprob+offset)]
print(np.asarray(w).shape)
print('weights of Gaussian mixtures:', np.asarray(w)[:,0]/np.sum(np.asarray(w)[:,0]))
mu = np.reshape(np.asarray(mu),[1,num_comp,hidden_size])
sigma = np.reshape(np.asarray(sigma)[:,0],[1,num_comp,hidden_size,hidden_size])
w = np.squeeze(np.asarray(w)[:,0])
return mu, sigma, w
# + colab={} colab_type="code" id="gQXNNSN7TecV"
def plot_prob_2D_GMM(samples, indices):
samples = samples[:,0,:]
samples = np.hstack((np.expand_dims(samples[:,indices[0]],-1),np.expand_dims(samples[:,indices[1]],-1)))
figure=corner.corner(samples)
axes = np.array(figure.axes).reshape((2, 2))
axes[1,0].set_xlabel('latent space variable %d'%indices[0])
axes[1,0].set_ylabel('latent space variable %d'%indices[1])
plt.savefig(plot_path+'posterior_contour_GMM_%s_latent_space_dir_%d_%d.pdf'%(label,indices[0],indices[1]),bbox_inches='tight')
plt.show()
# -
def transform_diagonal(matrix, transform=tf.nn.softplus, add=0):
diag = tf.linalg.diag_part(matrix)
if transform is not None:
diag = transform(diag)
transformed_diag = diag+add
new_matrix = tf.linalg.set_diag(matrix, transformed_diag)
return new_matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 353} colab_type="code" executionInfo={"elapsed": 14338, "status": "ok", "timestamp": 1558592190485, "user": {"displayName": "", "photoUrl": "", "userId": "18062987068597777273"}, "user_tz": 420} id="yvTEYw44O_5q" outputId="31ee8455-c44d-493e-e04f-e1d42abffdf7"
tf.reset_default_graph()
sigma_corr = tf.placeholder_with_default(noise_n,shape=[1,data_dim,n_channels])
mask = tf.placeholder_with_default(np.ones([data_dim], dtype='float32'),shape=[data_dim])
input_data = tf.placeholder(shape=[data_size,data_dim,n_channels], dtype=tf.float32)
inverse_T = tf.placeholder_with_default(1., shape=[])
lr = tf.placeholder_with_default(0.001,shape=[])
encoder = hub.Module(encoder_path, trainable=False)
generator = hub.Module(generator_path, trainable=False)
nvp_funcs = hub.Module(nvp_path, trainable=False)
MAP_ini = tf.placeholder_with_default(tf.zeros([data_size,hidden_size]),shape=[data_size,hidden_size])
MAP = tf.Variable(MAP_ini)
MAP_reset = tf.stop_gradient(MAP.assign(MAP_ini))
nlPost_MAP = get_log_posterior(MAP, input_data, generator,nvp_funcs, sigma_corr,mask, inverse_T)
loss_MAP = -nlPost_MAP[0]
optimizer = tf.train.AdamOptimizer(learning_rate=lr)
opt_op_MAP = optimizer.minimize(loss_MAP, var_list=[MAP])
recon_MAP = get_recon(generator,nvp_funcs, MAP,sigma_corr,mask)
hessian = get_hessian(-nlPost_MAP,MAP)
hessian_GN = get_GN_hessian(generator,nvp_funcs,MAP,mask,sigma_corr)
covariance = compute_covariance(hessian_GN)
variance = tf.linalg.diag_part(covariance)[0]
encoded = get_encoded(input_data, nvp_funcs,encoder)
# print(covariance)
approx_posterior_laplace = tfd.MultivariateNormalFullCovariance(loc=MAP[0],covariance_matrix=covariance[0])
#update_TriL = TriL_update.assign(tf.expand_dims(tf.linalg.cholesky(covariance[0]),0))
posterior_sample = approx_posterior_laplace.sample(params['batch_size'])
recon = get_recon(generator,nvp_funcs, posterior_sample ,sigma_corr,mask)
ini_val2 = np.ones((1,num_comp,(hidden_size *(hidden_size +1)) // 2),dtype=np.float32)
with tf.variable_scope("corrupted/gmm",reuse=tf.AUTO_REUSE):
mu_gmm = tf.Variable(np.ones((1,num_comp,hidden_size)), dtype=np.float32)
sigma_gmm = tf.Variable(tfp.math.fill_triangular(ini_val2))
w_gmm = tf.Variable(np.ones((num_comp))/num_comp, dtype=np.float32)
sigma_gmmt = transform_diagonal(sigma_gmm)
w_positive = tf.math.softplus(w_gmm)
w_rescaled = tf.squeeze(w_positive/tf.reduce_sum(w_positive))
gmm = tfd.MixtureSameFamily(mixture_distribution=tfd.Categorical(probs=w_rescaled),components_distribution=tfd.MultivariateNormalTriL(loc=mu_gmm,scale_tril=sigma_gmmt))
mu_ini = tf.placeholder_with_default(tf.zeros([1,num_comp,hidden_size]),shape=[1,num_comp,hidden_size])
sigma_ini = tf.placeholder_with_default(tf.ones([1,num_comp,hidden_size, hidden_size]),shape=[1,num_comp,hidden_size, hidden_size])
w_ini = tf.placeholder_with_default(tf.ones([num_comp])/num_comp,shape=[num_comp])
update_w = tf.stop_gradient(w_gmm.assign(tfp.math.softplus_inverse(w_ini)))
update_mugmm = tf.stop_gradient(mu_gmm.assign(mu_ini))
update_TriLgmm= tf.stop_gradient(sigma_gmm.assign(transform_diagonal(sigma_ini)))
gmm_sample = gmm.sample()
gmm_sample = tf.repeat(gmm_sample, params['batch_size'], axis=0)
# print(gmm_sample)
gmm_recon = get_recon(generator,nvp_funcs, gmm_sample ,sigma_corr,mask)
# +
minima_path = '../minimas/fmnist%d_PAE_2/'%num_mnist
if not os.path.isdir(minima_path):
os.makedirs(minima_path)
label_old = 'solidmask_2'
# -
# + colab={"base_uri": "https://localhost:8080/", "height": 4721} colab_type="code" executionInfo={"elapsed": 131558, "status": "ok", "timestamp": 1558592307817, "user": {"displayName": "", "photoUrl": "", "userId": "18062987068597777273"}, "user_tz": 420} id="Soh1tnGH1FTW" outputId="48802731-2a32-48fc-fd12-071300e5c583"
sess = tf.Session()
sess.run(tf.global_variables_initializer())
tf.random.set_random_seed(seed)
try:
minima, min_loss, min_var,recons, hesss, hesssGN = pickle.load(open(minima_path+'minima_%s.pkl'%label_old,'rb'))
except:
inits = get_random_start_values(20, sess)
minima =[]
min_loss=[]
min_var =[]
recons =[]
hesss =[]
hesssGN =[]
for jj,init in enumerate(inits):
print('progress in %', jj/len(inits)*100)
min_z, min_l, pos_def,hess,hessGN = minimize_posterior(init, data,custom_mask,noise,sess)
rec = sess.run(recon_MAP, feed_dict={sigma_corr:noise})
var = sess.run(variance, feed_dict={input_data: data,mask:custom_mask,sigma_corr:noise})
plot_image(rec[0], directory=plot_path, filename='recon_%s_minimum%d'%(label,jj), title='reconstruction with loss %.1f'%min_l)
if pos_def:
print('hessian postive definite')
minima.append(min_z)
min_loss.append(min_l)
min_var.append(var)
recons.append(rec)
hesss.append(hess)
hesssGN.append(hessGN)
order = np.argsort(min_loss)
min_loss = np.asarray(min_loss)[order]
minima = np.asarray(minima)[order]
min_var = np.asarray(min_var)[order]
hesss = np.asarray(hesss)[order]
hesssGN = np.asarray(hesssGN)[order]
pickle.dump([minima, min_loss, min_var,recons, hesss, hesssGN],open(minima_path+'minima_%s.pkl'%label,'wb'))
# + active=""
#
# +
try:
min_z_, min_z_, min_l_, pos_def_,hess_,hessGN_ = pickle.load(open(minima_path+'minima_from_true_%s.pkl'%label_old,'rb'))
except:
enc = sess.run(encoded, feed_dict={input_data: np.reshape(truth, (params['batch_size'],28*28,1))})
init= sess.run(MAP_reset, feed_dict={MAP_ini:enc})
min_z_, min_l_, pos_def_,hess_,hessGN_ = minimize_posterior(init,data,np.ones((28*28)),noise,sess,annealing =False)
pickle.dump([min_z_, min_z_, min_l_, pos_def_,hess_,hessGN_],open(minima_path+'minima_from_true_%s.pkl'%label,'wb'))
_ = sess.run(MAP_reset, feed_dict={MAP_ini:min_z_, sigma_corr:noise,input_data: data,sigma_corr:noise})
recs = sess.run(recon_MAP, feed_dict={MAP_ini:min_z_, sigma_corr:noise,input_data: data,sigma_corr:noise})
plot_image(recs[0], directory=plot_path, filename='start_from_truth_reconstruction', title='reconstruction')
# -
plt.scatter(np.arange(len(min_loss)),min_loss)
plt.scatter([0],min_l_)
print(min_loss)
# +
try:
min_z_, min_z_, min_l_, pos_def_,hess_,hessGN_ = pickle.load(open(minima_path+'minima_unamortized_%s.pkl'%label_old,'rb'))
except:
enc = sess.run(encoded, feed_dict={input_data: np.reshape(truth, (params['batch_size'],28*28,1))})
init= sess.run(MAP_reset, feed_dict={MAP_ini:enc})
min_z_, min_l_, pos_def_,hess_,hessGN_ = minimize_posterior(init,np.reshape(truth, (params['batch_size'],28*28,1)),np.ones((28*28)),noise,sess)
pickle.dump([min_z_, min_z_, min_l_, pos_def_,hess_,hessGN_],open(minima_path+'minima_unamortized_%s.pkl'%label,'wb'))
_ = sess.run(MAP_reset, feed_dict={MAP_ini:min_z_, sigma_corr:noise,input_data: data,sigma_corr:noise})
recs = sess.run(recon_MAP, feed_dict={MAP_ini:min_z_, sigma_corr:noise,input_data: data,sigma_corr:noise})
plot_image(recs[0], directory=plot_path, filename='true_reconstruction', title='reconstruction')
# -
print(plot_path)
w=[]
ind = np.arange(len(min_loss))
for ii in ind:
hess = hesss[ii]
cov = np.linalg.inv(hess)
_, logdet = np.linalg.slogdet(cov)
logprob = -min_loss[ii]
print(logprob,logdet)
w+=[np.exp(0.5*logdet+logprob-500)]
w=np.asarray(w)/np.sum(np.asarray(w))
#plt.plot(w)
index_gauss =np.sum(np.random.multinomial(1, w,500000),axis=0)
_=plt.scatter(np.arange(len(index_gauss)),index_gauss/500000)
print(index_gauss[0:10])
samples = []
for ii, n in enumerate(ind):
for jj in range(index_gauss[ii]):
cov = np.linalg.inv(hesss[n])
samples.append(np.dot(np.linalg.cholesky(cov),np.random.randn(params['latent_size']))+minima[n][0])
samples=np.asarray(samples)
np.random.shuffle(samples)
samples.shape
# +
ii = 0
_ = sess.run(MAP_reset, feed_dict={MAP_ini:samples[0:256], sigma_corr:noise,input_data: data,sigma_corr:noise})
recs = sess.run(recon_MAP, feed_dict={MAP_ini:samples[0:256], sigma_corr:noise,input_data: data,sigma_corr:noise})
recs = np.reshape(recs,(-1,28,28))+0.5
#plot_samples(recs+0.5, custom_mask, title='Samples from Laplace approximation', filename='samples_laplace_deepest_minimum_%s'%label)
plt.figure(figsize=(5,5))
for ii in range(25):
#plt.subplot(4,15,ii+1)
plt.subplot(5,5,ii+1)
plt.imshow((recs[ii]).reshape(28,28),cmap='gray')
plt.axis('off')
# for ii in range(30):
# plt.subplot(4,15,30+ii+1)
# masked = recs[ii]
# masked[np.where(custom_mask.reshape(28,28)==0)]=0
# plt.imshow(masked,cmap='gray')
# plt.axis('off')
# plt.tight_layout()
plt.savefig(os.path.join(plot_path+'posterior_samples.pdf'),bbox_inches='tight')
# plt.figure(figsize=(5,5))
# for ii in range(32):
# plt.subplot(8,8,ii+1)
# plt.axis('off')
# plt.tight_layout()
# plt.savefig(os.path.join(params['plot_dir']+'posterior_samples_maksed_example1.pdf'),bbox_inches='tight')
# -
recs = sess.run(MAP_reset, feed_dict={MAP_ini:minima[0], sigma_corr:noise,input_data: data,sigma_corr:noise})
recs = sess.run(recon_MAP, feed_dict={MAP_ini:minima[0], sigma_corr:noise,input_data: data,sigma_corr:noise})
plot_image(recs[0], directory=plot_path, filename='best_reconstruction', title='best_reconstruction')
from mcmcplot import mcmcplot as mcp
samples_cut=np.vstack((samples[:,7],samples[:,5])).T
samples_cut.shape
fjd, used_settings = mcp.plot_joint_distributions(
chains=samples_cut,names=['', ''],
settings=None,
return_settings=True)
a = fjd.ax_joint
tmp = a.yaxis.get_label()
tmp.set_fontsize(20)
tmp = a.xaxis.get_label()
tmp.set_fontsize(20)
plt.savefig(os.path.join(plot_path+'distribution.pdf'),bbox_inches='tight')
| notebooks/ImageRestoration/ImageCorruptionFMNIST-solidmask_PAE_3055.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# +
# Read CSV
unemployed_data_one = pd.read_csv("../Resources/unemployment_2010-2011.csv")
unemployed_data_two = pd.read_csv("../Resources/unemployment_2012-2014.csv")
# Merge our two data frames together
combined_unemployed_data = pd.merge(unemployed_data_one, unemployed_data_two, on="Country Name")
combined_unemployed_data.head()
# -
# Delete the duplicate 'Country Code' column and rename the first one back to 'Country Code'
del combined_unemployed_data['Country Code_y']
combined_unemployed_data = combined_unemployed_data.rename(columns={"Country Code_x":"Country Code"})
combined_unemployed_data.head()
# Set the 'Country Code' to be our index for easy referencing of rows
combined_unemployed_data = combined_unemployed_data.set_index("Country Code")
# +
# Collect the mean unemployment rates for the world
average_unemployment = combined_unemployed_data.mean()
# Collect the years where data was collected
years = average_unemployment.keys()
years
# +
# Plot the world average as a line chart
world_avg, = plt.plot(years, average_unemployment, color="blue", label="World Average" )
# Plot the unemployment values for a single country
country_one, = plt.plot(years, combined_unemployed_data.loc['USA',["2010","2011","2012","2013","2014"]],
color="green",label=combined_unemployed_data.loc['USA',"Country Name"])
# Create a legend for our chart
plt.legend(handles=[world_avg, country_one], loc="best")
# Show the chart
plt.show()
# -
average_unemployment.plot(label="World Average")
combined_unemployed_data.loc['USA', "2010":"2014"].plot(label="United States")
plt.legend()
plt.show()
| PandasMultiLine/unemploy_chart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
N = np.random.rand(3, 5)
N
Z = np.zeros((2,4)) # zeros는 괄호를 꼭 두번 입력해야한다.
Z
N[0][2] # 1행 3열
A = np.array([[0,1,2,-1,],
[1,2,1,2],
[-1,3,0,4]])
A.T
AA = np.array([[0,1,-1],
[1,2,3],
[2,1,0],
[-1,2,4]])
AA
B = np.array([[0,2],
[1,1],
[-1,2]])
B[0][1]
AA[1][1]
AA[3][0]
print(AA)
print(AA[1][1])
print(AA[3][0])
# ---
# ### 요소별 곱하기
# +
A = np.array([
[1, 2, 3],
[4, 5, 6],
[7, 8, 9]
])
B = np.array([
[0, 1, 2],
[2, 0, 1],
[1, 2, 0]
])
# -
A * B
# ### 내적곱
np.dot(A,B)
A @ B
# ---
# * 숫자/행렬 스칼라 곱 : *
# * 행렬 덧셈 : +
# * 행렬 곱셈 : @
A @ B + (A + 2 * B)
# A
A = np.array([
[1, -1, 2],
[3, 2, 2]
])
# B
B = np.array([
[0, 1],
[-1, 1],
[5, 2]
])
# C
C = np.array([
[2, -1],
[-3, 3]
])
# D
D = np.array([
[-5, 1],
[2, 0]
])
result = 2*A @ (-1*B) @ (3*C + D)
result
# ---
array_A = np.arange(10)
array_A
array_B = np.arange(10, 20)
array_B
array_A * 2
array_A / 2
# * array * 2
# * array + 2
# * array / 2
# * array ** 2
array_A + array_B
# ### 전치 행렬
# #### transpose
# * 행과 열을 바꿈
A = np.array([
[1, 2, 3],
[4, 5, 6]
])
A.transpose()
np.transpose(A)
A.T
# ### 단위 행렬
# #### identity matrix
# * 단위 행렬은 항상 정사각형의 모양
# * 어떤 행렬이든 간에 단위 행렬을 곱하면 기존 행렬이 그대로 유지
I = np.eye(3)
I
I = np.identity(3)
I
I = np.array([
[1, 0],
[0, 1]
])
D = np.array([
[-5, 1],
[2, 0]
])
D @ I
# ### 역행렬
# #### inverse matrix
# * A에 곱했을 때 단위 행렬 I가 나오도록 하는 행렬
# * 역행렬은 꼭 정사각형이어야 한다
# * 모든 행렬에 역행렬이 있는 건 아님
A = np.array([
[3, 4],
[1, 2]
])
# #### 선형대수(Linear Algebra) 함수
A_inverse = np.linalg.pinv(A)
A_inverse
np.linalg.inv(A)
A_inverse @ A
# ---
# +
A = np.array([
[1, -1, 2],
[3, 2, 2]
])
B = np.array([
[0, 1],
[-1, 1],
[5, 2]
])
C = np.array([
[2, -1],
[-3, 3]
])
D = np.array([
[-5, 1],
[2, 0]
])
# -
result = B.T @ (2 * A.T) @ (3 * np.linalg.pinv(C) + D.T)
result
| practice/numpy_nparray.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
# # BioInformatics Tools kit
#
# > Summarized some useful tools in it.
# BITK will help us manipulate data in different formats.
# ## Install
# `pip install bitk`
# ## How to use
# ### dedim.py
#
#
# ```bash
# usage: dedim.py [-h] [--sep SEP] [--dedimensions-method DEDIMENSIONS_METHOD]
# [--cluster-method CLUSTER_METHOD]
# [--assess-method ASSESS_METHOD] [--dimensions DIMENSIONS]
# [--cluster-number CLUSTER_NUMBER] [-r] [--no-row-feature]
# [--annotation ANNOTATION] [--size SIZE] [--style STYLE]
# [-t TITLE] [-f FIG] [--version]
# matrix prefix
#
# positional arguments:
# matrix matrix table, if row represents feature, please note to add '--row-feature' option
# prefix output prefix
#
# optional arguments:
# -h, --help show this help message and exit
# --sep SEP separation
# (default: )
# --dedimensions-method DEDIMENSIONS_METHOD
# de-dimensions method
# (default: PCA)
# --cluster-method CLUSTER_METHOD
# cluster method
# (default: MiniBatchKMeans)
# --assess-method ASSESS_METHOD
# assess methods for best cluster number
# (default: silhouette_score)
# --dimensions DIMENSIONS
# reduce to n dimensions
# (default: 3)
# --cluster-number CLUSTER_NUMBER
# cluster number, if not specific it, it will be the best cluster number infered
# (default: None)
# -r, --row-feature row in the matrix represents feature
# (default: True)
# --no-row-feature
# --annotation ANNOTATION
# annotation file, sep should be ','
# (default: None)
# --size SIZE size column in annotation file
# (default: None)
# --style STYLE style column in annotation file
# (default: None)
# -t TITLE, --title TITLE
# figure title
# (default: None)
# -f FIG, --fig FIG png/pdf
# (default: png)
# --version show program's version number and exit
# ```
#
#
# #### example
#
# ```bash
# dedim.py tests/test.csv tests/test_result \
# --sep , --dimensions 2 \
# --no-row-feature --annotation tests/test_anno.csv \
# --style targets --title 'test PCA' \
# --dedimensions-method TSNE --fig png
# ```
#
# 
# ### fc_rename.py
# ```
# usage: fc_rename.py [-h] [-s SAMPLE_TITLE] [-b BAM_TITLE] [-c COUNT_TITLE]
# [--version]
# featurecounts clinical prefix
#
# featurecounts will use the bamfile name as the sample name. This script rename the featurecounts by the given table.
#
# positional arguments:
# featurecounts featurecounts table file
# clinical clinical table file with header
# prefix output prefix, there will be two outputs, one is count output (prefix_count.txt), and the other one is the rename featurecounts (prefix_fc.txt)
#
# optional arguments:
# -h, --help show this help message and exit
# -s SAMPLE_TITLE, --sample-title SAMPLE_TITLE
# the column name of the sample name
# (default: sample)
# -b BAM_TITLE, --bam-title BAM_TITLE
# the column name of the bamfile name
# (default: bam)
# -c COUNT_TITLE, --count-title COUNT_TITLE
# the column name used as identity in count data
# (default: Geneid)
# --version show program's version number and exit
# ```
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Parse odometry and localization data
import json
import numpy as np
import matplotlib.pyplot as plt
# ### Read recorded log file
with open('../dataset/test/odometry.json') as json_data:
odom_raw = json.load(json_data)
odom_raw[:2]
# ### Parse position and velocity
odom = odom_raw
for i in range(len(odom_raw)):
pose = np.array( odom_raw[i]['pose_m'][1:-1].split(',') ).astype(np.float)
odom[i]['pose_m'] = pose
vel = np.array( odom_raw[i]['vel_m_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['vel_m_sec'] = vel
acc = np.array( odom_raw[i]['acc_m_sec_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['acc_m_sec_sec'] = acc
orient_quat = np.array( odom_raw[i]['orient_quat'][1:-1].split(',') ).astype(np.float)
odom[i]['orient_quat'] = orient_quat
ang_vel_deg_sec = np.array( odom_raw[i]['ang_vel_deg_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['ang_vel_deg_sec'] = ang_vel_deg_sec
ang_acc_deg_sec_sec = np.array( odom_raw[i]['ang_acc_deg_sec_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['ang_acc_deg_sec_sec'] = ang_acc_deg_sec_sec
odom[:2]
# ### Plot results
# Take into account that Unity has the following rigid body coordinate system,
# [reference](https://docs.nvidia.com/isaac/isaac/doc/simulation/unity3d.html#coordinates):
# - X - right
# - Y - up
# - Z - forward
# +
poses = []
for i in range(len(odom)):
poses.append( odom[i]['pose_m'] )
poses = np.array(poses)
plt.figure(figsize=(10,10))
plt.title('Player trajectory on XZ-plane')
plt.plot(poses[:,0], poses[:,2])
plt.grid()
plt.xlabel('X, [m]')
plt.ylabel('Z, [m]');
# +
vels = []
for i in range(len(odom)):
vels.append( odom[i]['vel_m_sec'] )
vels = np.array(vels)
plt.figure(figsize=(10,5))
plt.title('Horizontal velocities')
plt.plot(vels[:,0], label='V_x, [m/sec]')
plt.plot(vels[:,2], label='V_z, [m/sec]')
plt.legend()
plt.grid()
plt.ylim([-15, 15]);
# -
# ## Parse detections data
with open('../dataset/test/detections.json') as json_data:
dets_raw = json.load(json_data)
dets_raw[:1]
dets = dets_raw
for i in range(len(dets_raw)):
tags1 = dets_raw[i]['tags'].split(', ')
dets[i]['tags'] = tags1
poses1 = dets_raw[i]['poses_m'][1:-1].split('), (')
poses1 = [np.array(p.split(',')).astype(np.float) for p in poses1]
dets[i]['poses_m'] = poses1
bbox_sizes1 = dets_raw[i]['bbox_sizes_m'][1:-1].split('), (')
bbox_sizes1 = [np.array(p.split(',')).astype(np.float) for p in bbox_sizes1]
dets[i]['bbox_sizes_m'] = bbox_sizes1
orients_quat1 = dets_raw[i]['orients_quat'][1:-1].split('), (')
orients_quat1 = [np.array(p.split(',')).astype(np.float) for p in orients_quat1]
dets[i]['orients_quat'] = orients_quat1
# +
t = 17 #np.random.choice(len(dets))
poses1 = dets[t]['poses_m']
tags1 = dets[t]['tags']
plt.figure(figsize=(10,10))
plt.grid()
for pose, tag in zip(poses1, tags1):
if tag == 'TrafficCar':
color = 'b'
elif tag == 'Pedestrian':
color = 'r'
plt.plot(pose[0], pose[2], 'x', color=color)
# draw camera position
camera_pose = odom[t]['pose_m']
plt.plot(camera_pose[0], camera_pose[2], 'ro', label='camera_pose')
# draw circle of detection lookup radius
R_lookup = 50.0
lookup_area = plt.Circle((camera_pose[0], camera_pose[2]), R_lookup, color='g', label='detections area')
plt.gca().add_artist(lookup_area)
plt.gca().set_aspect('equal')
plt.xlim([camera_pose[0]-(R_lookup+5.), camera_pose[0]+(R_lookup+5.)])
plt.ylim([camera_pose[2]-(R_lookup+5.), camera_pose[2]+(R_lookup+5.)])
plt.legend();
# -
# ## Combine detections and odometry to form a dataset
import os
import cv2
import pyquaternion as pyq
import math
class Dataset:
def __init__(self, data_path, odom_filename='odometry.json', dets_filename='detections.json'):
files = os.listdir(data_path)
self.image_files = []
for file in files:
if file.endswith('.png'):
self.image_files.append(os.path.join(data_path, file))
self.odom_file = os.path.join(data_path, odom_filename)
self.dets_file = os.path.join(data_path, dets_filename)
self.odom = self.parse_odom()
self.dets = self.parse_dets()
# parameters
self.fov = {'horizontal': 70., 'vertical': 40.}
self.R_lookup = 50.0 # [m]
def parse_odom(self):
# check if odometry file exists
assert os.path.isfile(self.odom_file)
with open(self.odom_file) as json_data:
odom_raw = json.load(json_data)
odom = odom_raw
for i in range(len(odom_raw)):
pose = np.array( odom_raw[i]['pose_m'][1:-1].split(',') ).astype(np.float)
odom[i]['pose_m'] = pose
vel = np.array( odom_raw[i]['vel_m_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['vel_m_sec'] = vel
acc = np.array( odom_raw[i]['acc_m_sec_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['acc_m_sec_sec'] = acc
orient_quat = np.array( odom_raw[i]['orient_quat'][1:-1].split(',') ).astype(np.float)
odom[i]['orient_quat'] = orient_quat
ang_vel_deg_sec = np.array( odom_raw[i]['ang_vel_deg_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['ang_vel_deg_sec'] = ang_vel_deg_sec
ang_acc_deg_sec_sec = np.array( odom_raw[i]['ang_acc_deg_sec_sec'][1:-1].split(',') ).astype(np.float)
odom[i]['ang_acc_deg_sec_sec'] = ang_acc_deg_sec_sec
return odom
def parse_dets(self):
# check if detections file exists
assert os.path.isfile(self.dets_file)
with open(self.dets_file) as json_data:
dets_raw = json.load(json_data)
dets = dets_raw
for i in range(len(dets_raw)):
tags1 = dets_raw[i]['tags'].split(', ')
dets[i]['tags'] = tags1
poses1 = dets_raw[i]['poses_m'][1:-1].split('), (')
poses1 = [np.array(p.split(',')).astype(np.float) for p in poses1]
dets[i]['poses_m'] = poses1
bbox_sizes1 = dets_raw[i]['bbox_sizes_m'][1:-1].split('), (')
bbox_sizes1 = [np.array(p.split(',')).astype(np.float) for p in bbox_sizes1]
dets[i]['bbox_sizes_m'] = bbox_sizes1
orients_quat1 = dets_raw[i]['orients_quat'][1:-1].split('), (')
orients_quat1 = [np.array(p.split(',')).astype(np.float) for p in orients_quat1]
dets[i]['orients_quat'] = orients_quat1
return dets
@staticmethod
def quat_orient_diff(q1, q2):
"""
Outputs orientation difference of quaternion `q2` relative to `q1`.
Result is a quaternion `qd`.
Reference: https://stackoverflow.com/questions/57063595/how-to-obtain-the-angle-between-two-quaternions
"""
pyq1 = pyq.Quaternion(q1)
pyq2 = pyq.Quaternion(q2)
qd = pyq1 * pyq2.conjugate
return qd
def __getitem__(self, i):
assert len(self.image_files) > 0
# read image
image = cv2.imread(self.image_files[0][:-10]+str(i+1)+'_img.png')
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# get detections data relative to the camera
output = {}
output['tags'] = self.dets[i]['tags']
output['poses'] = self.dets[i]['poses_m'] - self.odom[i]['pose_m']
output['bbox_sizes_m'] = self.dets[i]['bbox_sizes_m']
output['orients_quat'] = []
for det_quat in self.dets[i]['orients_quat']:
# convert Unity quaternion notation: q_unity = [x,y,z,w]
# to pyquaternion notation: q_pyq = [w,x,y,z]
q1 = [det_quat[3], det_quat[0], det_quat[1], det_quat[2]]
camera_quat = self.odom[i]['orient_quat']
q2 = [camera_quat[3], camera_quat[0], camera_quat[1], camera_quat[2]]
quat_relative_to_cam = self.quat_orient_diff(q1, q2)
output['orients_quat'].append(quat_relative_to_cam)
# useful parameters
output['camera_fov'] = self.fov
output['lookup_radius'] = self.R_lookup
output['camera_quat'] = pyq.Quaternion(q2).normalised
return image, output
def __len__(self):
return len(self.image_files)
def quat_to_euler(q):
q = q.normalised
# Calculate Euler angles from pyquaternion
phi = math.atan2( 2 * (q.w * q.x + q.y * q.z), 1 - 2 * (q.x**2 + q.y**2) )
theta = math.asin ( np.clip(2 * (q.w * q.y - q.z * q.x), -1, 1) )
psi = math.atan2( 2 * (q.w * q.z + q.x * q.y), 1 - 2 * (q.y**2 + q.z**2) )
return phi, theta, psi
# +
dataset = Dataset('../dataset/test/')
ind = np.random.choice(len(dataset))
img, out = dataset[ind]
#print('Camera heading: {:.1f} deg'.format(math.degrees(quat_to_euler(out['camera_quat'])[1])))
plt.figure(figsize=(20, 5))
plt.subplot(1,2,1)
plt.imshow(img);
plt.subplot(1,2,2)
plt.grid()
for pose, tag in zip(out['poses'], out['tags']):
if tag == 'TrafficCar':
color = 'b'
elif tag == 'Pedestrian':
color = 'r'
plt.plot(pose[0], pose[2], 'x', color=color)
# draw camera position
camera_pose = [0, 0, 0]
plt.plot(camera_pose[0], camera_pose[2], 'ro', label='camera_pose')
# draw circle of detection lookup radius
R_lookup = out['lookup_radius']
lookup_area = plt.Circle((camera_pose[0], camera_pose[2]), R_lookup, color='g', label='detections area')
plt.gca().add_artist(lookup_area)
plt.gca().set_aspect('equal')
plt.xlim([camera_pose[0]-(R_lookup+5.), camera_pose[0]+(R_lookup+5.)])
plt.ylim([camera_pose[2]-(R_lookup+5.), camera_pose[2]+(R_lookup+5.)])
# TODO: draw camera FOV
psi = quat_to_euler(out['camera_quat'])[1]
psi_left = quat_to_euler(out['camera_quat'])[1] - math.radians(out['camera_fov']['horizontal']/2.)
psi_right = quat_to_euler(out['camera_quat'])[1] + math.radians(out['camera_fov']['horizontal']/2.)
plt.arrow(camera_pose[0],camera_pose[2], R_lookup*np.sin(psi), R_lookup*np.cos(psi))
plt.arrow(camera_pose[0],camera_pose[2], R_lookup*np.sin(psi_left), R_lookup*np.cos(psi_left))
plt.arrow(camera_pose[0],camera_pose[2], R_lookup*np.sin(psi_right), R_lookup*np.cos(psi_right))
plt.legend();
| tools/parse_odometry_and_detections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### ESERCIZIO 3 - Calcolo dell'area di un rettangolo
#
# Date le misure (intere o decimali) dei lati di un rettangolo, calcolarne l'area
| lezione1/testo-esercizi/ipynb/ES3-AreaRettangolo-testo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
train_df = pd.read_csv(Path('Resources/2019loans.csv'))
test_df = pd.read_csv(Path('Resources/2020Q1loans.csv'))
train_df.columns
test_df.columns
# +
# Convert categorical data to numeric and separate target feature for training data
X_train = train_df[['loan_amnt', 'int_rate', 'installment', 'home_ownership', 'annual_inc',
'verification_status', 'pymnt_plan', 'dti', 'delinq_2yrs',
'inq_last_6mths', 'open_acc', 'pub_rec', 'revol_bal', 'total_acc',
'initial_list_status', 'out_prncp', 'out_prncp_inv', 'total_pymnt',
'total_pymnt_inv', 'total_rec_prncp', 'total_rec_int',
'total_rec_late_fee', 'recoveries', 'collection_recovery_fee',
'last_pymnt_amnt', 'collections_12_mths_ex_med', 'policy_code',
'application_type', 'acc_now_delinq', 'tot_coll_amt', 'tot_cur_bal',
'open_acc_6m', 'open_act_il', 'open_il_12m', 'open_il_24m',
'mths_since_rcnt_il', 'total_bal_il', 'il_util', 'open_rv_12m',
'open_rv_24m', 'max_bal_bc', 'all_util', 'total_rev_hi_lim', 'inq_fi',
'total_cu_tl', 'inq_last_12m', 'acc_open_past_24mths', 'avg_cur_bal',
'bc_open_to_buy', 'bc_util', 'chargeoff_within_12_mths', 'delinq_amnt',
'mo_sin_old_il_acct', 'mo_sin_old_rev_tl_op', 'mo_sin_rcnt_rev_tl_op',
'mo_sin_rcnt_tl', 'mort_acc', 'mths_since_recent_bc',
'mths_since_recent_inq', 'num_accts_ever_120_pd', 'num_actv_bc_tl',
'num_actv_rev_tl', 'num_bc_sats', 'num_bc_tl', 'num_il_tl',
'num_op_rev_tl', 'num_rev_accts', 'num_rev_tl_bal_gt_0', 'num_sats',
'num_tl_120dpd_2m', 'num_tl_30dpd', 'num_tl_90g_dpd_24m',
'num_tl_op_past_12m', 'pct_tl_nvr_dlq', 'percent_bc_gt_75',
'pub_rec_bankruptcies', 'tax_liens', 'tot_hi_cred_lim',
'total_bal_ex_mort', 'total_bc_limit', 'total_il_high_credit_limit',
'hardship_flag', 'debt_settlement_flag']]
y_train = train_df[['target']]
X_train = pd.get_dummies(X_train)
#y_train = pd.get_dummies(y_train)
# -
# Convert categorical data to numeric and separate target feature for testing data
X_test = test_df[['loan_amnt', 'int_rate', 'installment', 'home_ownership', 'annual_inc',
'verification_status', 'pymnt_plan', 'dti', 'delinq_2yrs',
'inq_last_6mths', 'open_acc', 'pub_rec', 'revol_bal', 'total_acc',
'initial_list_status', 'out_prncp', 'out_prncp_inv', 'total_pymnt',
'total_pymnt_inv', 'total_rec_prncp', 'total_rec_int',
'total_rec_late_fee', 'recoveries', 'collection_recovery_fee',
'last_pymnt_amnt', 'collections_12_mths_ex_med', 'policy_code',
'application_type', 'acc_now_delinq', 'tot_coll_amt', 'tot_cur_bal',
'open_acc_6m', 'open_act_il', 'open_il_12m', 'open_il_24m',
'mths_since_rcnt_il', 'total_bal_il', 'il_util', 'open_rv_12m',
'open_rv_24m', 'max_bal_bc', 'all_util', 'total_rev_hi_lim', 'inq_fi',
'total_cu_tl', 'inq_last_12m', 'acc_open_past_24mths', 'avg_cur_bal',
'bc_open_to_buy', 'bc_util', 'chargeoff_within_12_mths', 'delinq_amnt',
'mo_sin_old_il_acct', 'mo_sin_old_rev_tl_op', 'mo_sin_rcnt_rev_tl_op',
'mo_sin_rcnt_tl', 'mort_acc', 'mths_since_recent_bc',
'mths_since_recent_inq', 'num_accts_ever_120_pd', 'num_actv_bc_tl',
'num_actv_rev_tl', 'num_bc_sats', 'num_bc_tl', 'num_il_tl',
'num_op_rev_tl', 'num_rev_accts', 'num_rev_tl_bal_gt_0', 'num_sats',
'num_tl_120dpd_2m', 'num_tl_30dpd', 'num_tl_90g_dpd_24m',
'num_tl_op_past_12m', 'pct_tl_nvr_dlq', 'percent_bc_gt_75',
'pub_rec_bankruptcies', 'tax_liens', 'tot_hi_cred_lim',
'total_bal_ex_mort', 'total_bc_limit', 'total_il_high_credit_limit',
'hardship_flag', 'debt_settlement_flag']]
y_test = test_df[['target']]
X_test = pd.get_dummies(X_test)
#y_test = pd.get_dummies(y_test)
# add missing dummy variables to testing set
missing = set(X_train)-set(X_test)
missing
X_test['debt_settlement_flag_Y'] = 0
X_test
y_test
y_train = LabelEncoder().fit_transform(y_train['target'])
y_train
y_test = LabelEncoder().fit_transform(y_test['target'])
y_test
# Train the Logistic Regression model on the unscaled data and print the model score
l = LogisticRegression(random_state=1234, max_iter=3000).fit(X_train, y_train)
print(f'Training Score: {l.score(X_train, y_train)}')
print(f'Testing Score: {l.score(X_test, y_test)}')
# +
# Train a Random Forest Classifier model and print the model score
r = RandomForestClassifier(random_state=1234).fit(X_train, y_train)
print(f'Training Score: {r.score(X_train, y_train)}')
print(f'Testing Score: {r.score(X_test, y_test)}')
# -
# Scale the data
scaler = StandardScaler().fit(X_train)
X_train_scaled = scaler.transform(X_train)
X_test_scaled = scaler.transform(X_test)
# Train the Logistic Regression model on the scaled data and print the model score
l = LogisticRegression(random_state=1234, max_iter=3000).fit(X_train, y_train)
print(f'Training Score: {l.score(X_train_scaled, y_train)}')
print(f'Testing Score: {l.score(X_test_scaled, y_test)}')
# Train a Random Forest Classifier model on the scaled data and print the model score
r = RandomForestClassifier(random_state=1234).fit(X_train, y_train)
print(f'Training Score: {r.score(X_train_scaled, y_train)}')
print(f'Testing Score: {r.score(X_test_scaled, y_test)}')
| Credit Risk Evaluator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interfacing PowerModels.jl with pandapower
# pandapower now has an interface to PowerModels.jl that can be used for efficient power system optimization.
#
# ### What is PowerModels.jl and why should I use it?
#
# - [PowerModels.jl](https://lanl-ansi.github.io/PowerModels.jl/stable/) is a package for steady-state power network optimization
# - It is based on the relatively new language [Julia](https://julialang.org/) which is gaining popularity in scientific applications
# - PowerModels uses Julia/JuMP for the optimization, which [clearly outperforms the Python alternative Pyomo](http://yetanothermathprogrammingconsultant.blogspot.com/2015/05/model-generation-in-julia.html)
# - PowerModels has a great modular design that allows you to define [different formulations for optimization problems](https://lanl-ansi.github.io/PowerModels.jl/stable/specifications/) based on different [network formulations](https://lanl-ansi.github.io/PowerModels.jl/stable/formulations/) as well as use several [relaxation schemes](https://lanl-ansi.github.io/PowerModels.jl/stable/relaxations/). You can then solve the problem using many open source as well as commercial solvers through [JuMP](http://www.juliaopt.org/JuMP.jl/0.18/installation.html#getting-solvers)
#
# ### Well then why do I still need pandapower?
#
# Because pandapower:
#
# - allows you to easily define power systems with nameplate parameters and standard types
# - comes with thouroughly validated element models of transformers with tap changers, three-winding transformers, switches/breakers, extended ward equivalents and many more
# - keeps all data in tables (pandas DataFrames), which makes data management and analysis very comfortable
# - provides different power system analysis functions, such as a (very fast) power flow, short-circuit calculation, state estimation, graph searches and a plotting library that can be used on the same grid models
# - allows you to do all pre- and postprocessing in Python, which still has a much richer environment of free libraries than Julia (currently 157,755 packages on PyPI vs. 1,906 libraries on Pkg)
#
# So using pandapower to define the grid models and then using PowerModels for the optimization really gives you the best of all worlds - you can use the rich environment of Python libraries, the sophisticated element models of pandapower, the modular optimization framework of PowerModels and the efficient mathematical modeling of JuMP.
# ### Let's get started
#
# So here is an example of how it works. First, we create a grid in pandapower. Here, we create a meshed 110kV grid with four buses that is fed from an 220kV network through a 3-Winding transformer.
# +
import pandapower as pp
import numpy as np
net = pp.create_empty_network()
min_vm_pu = 0.95
max_vm_pu = 1.05
#create buses
bus1 = pp.create_bus(net, vn_kv=220., geodata=(5,9), min_vm_pu=min_vm_pu, max_vm_pu=max_vm_pu)
bus2 = pp.create_bus(net, vn_kv=110., geodata=(6,10), min_vm_pu=min_vm_pu, max_vm_pu=max_vm_pu)
bus3 = pp.create_bus(net, vn_kv=110., geodata=(10,9), min_vm_pu=min_vm_pu, max_vm_pu=max_vm_pu)
bus4 = pp.create_bus(net, vn_kv=110., geodata=(8,8), min_vm_pu=min_vm_pu, max_vm_pu=max_vm_pu)
bus5 = pp.create_bus(net, vn_kv=110., geodata=(6,8), min_vm_pu=min_vm_pu, max_vm_pu=max_vm_pu)
#create 220/110/110 kV 3W-transformer
pp.create_transformer3w_from_parameters(net, bus1, bus2, bus5, vn_hv_kv=220, vn_mv_kv=110,
vn_lv_kv=110, vk_hv_percent=10., vk_mv_percent=10.,
vk_lv_percent=10., vkr_hv_percent=0.5,
vkr_mv_percent=0.5, vkr_lv_percent=0.5, pfe_kw=10,
i0_percent=0.1, shift_mv_degree=0, shift_lv_degree=0,
sn_hv_mva=100, sn_mv_mva=50, sn_lv_mva=50)
#create 110 kV lines
l1 = pp.create_line(net, bus2, bus3, length_km=70., std_type='149-AL1/24-ST1A 110.0')
l2 = pp.create_line(net, bus3, bus4, length_km=50., std_type='149-AL1/24-ST1A 110.0')
l3 = pp.create_line(net, bus4, bus2, length_km=40., std_type='149-AL1/24-ST1A 110.0')
l4 = pp.create_line(net, bus4, bus5, length_km=30., std_type='149-AL1/24-ST1A 110.0')
#create loads
pp.create_load(net, bus2, p_mw=60)
pp.create_load(net, bus3, p_mw=70)
pp.create_load(net, bus4, p_mw=10)
#create generators
g1 = pp.create_gen(net, bus1, p_mw=40, min_p_mw=0, max_p_mw=200, vm_pu=1.01, slack=True)
pp.create_poly_cost(net, g1, 'gen', cp1_eur_per_mw=1)
g2 = pp.create_gen(net, bus3, p_mw=40, min_p_mw=0, max_p_mw=200, vm_pu=1.01)
pp.create_poly_cost(net, g2, 'gen', cp1_eur_per_mw=3)
g3 = pp.create_gen(net, bus4, p_mw=50, min_p_mw=0, max_p_mw=200, vm_pu=1.01)
pp.create_poly_cost(net, g3, 'gen', cp1_eur_per_mw=3)
net
# -
# Note that PowerModels does not have a 3W-transformer model, but since pandapower includes the equations to calculates the equivalent branches for the 3W-transformers, it is possible to optimize grids with 3W-transformers in PowerModels through the pandapower interface. The same is true for other complex transformer models, switches/breaker, extended ward equivalents etc.
#
# Let's have a look at the grid we created with pandapowers plotting module:
import pandapower.plotting as plot
# %matplotlib inline
plot.simple_plot(net)
# Now lets run an OPF through PowerModels and look at the results (Note that the first time the runpm function is called, Julia is started in the background, which may take some time):
pp.runpm_ac_opf(net)
# Since Generator 1 has the lowest cost, all required power is supplied through this generator:
net.res_gen
# This however leeds to an overload in the three-winding transformer, through which g1 is connected:
net.res_trafo3w.loading_percent
# Let's set some constraints for the 3W-transformer and the lines and rerun the OPF:
net.trafo3w["max_loading_percent"] = 50
net.line["max_loading_percent"] = 20
pp.runpm_ac_opf(net)
# The constraints are complied with for all lines and the 3W transformer:
net.res_trafo3w.loading_percent
net.res_line.loading_percent
# The power is now generated by a mixture of the generators:
net.res_gen
# ## Accessing the full functionality of PowerModels.jl
# Apart from the AC OPF used in the example above, pandapower also has an interface to run the DC OPF:
pp.runpm_dc_opf(net)
net.res_bus
# The julia file that is used to do that can be found in pandapower/pandapower/opf/run_powermodels_dc.jl and looks like this:
"""
using PowerModels
using Ipopt
using PP2PM
function run_powermodels(json_path)
pm = PP2PM.load_pm_from_json(json_path)
result = PowerModels.run_dc_opf(pm, Ipopt.IpoptSolver(),
setting = Dict("output" => Dict("branch_flows" => true)))
return result
end
""";
# Of course PowerModels is a great modular tool that allows you to do much more than that. You might want to use a different OPF formulation, a relaxation method or a different solver. You might even want to use one of the variants of PowerModels that are being developed, such as [PowerModelsACDC.jl](https://github.com/hakanergun/PowerModelsACDC.jl) or [PowerModelsReliability.jl](https://github.com/frederikgeth/PowerModelsReliability.jl).
#
# To do that, you can switch out the standard file with your own custom .jl file. Lets say we want to run a power flow instead of an OPF. There is a custom julia file for that in pandapower/tutorials/run_powermodels_custom.jl that looks like this:
"""
using PowerModels
using Ipopt
import JSON
function run_powermodels(json_path)
pm = PP2PM.load_pm_from_json(json_path)
result = PowerModels.run_pf(pm, ACPPowerModel, Ipopt.IpoptSolver(),
setting = Dict("output" => Dict("branch_flows" => true)))
return result
end
""";
# We point the runpm function to this file, and as we can see by the flat voltage values, the OPF is now run with a DC network formulation:
pp.runpm(net, julia_file="run_powermodels_custom.jl")
net.res_bus
# The PowerModels data structure that was passed to Julia can be accessed like this:
net._pm["bus"]
# There is also a callback that allows you to add additional data to the PowerModels data structure in case it is not already added by the pandapower/PowerModels interface. In the callback you can add any data from the net, ppc or any source:
# +
def add_data(net, ppc, pm):
pm["gen"]["1"]["bar"] = "foo"
pm["f_hz"] = net.f_hz
pp.runpm(net, julia_file="run_powermodels_custom.jl", pp_to_pm_callback=add_data)
print(net._pm["gen"]["1"]["bar"])
print(net._pm["f_hz"])
# -
# These variables can now also be accessed on the Julia side, in case you need some more variables for custom optimizations.
#
# Keep in mind that indices in PowerModels are 1-based so that the indices are shifted by one between ppc and pm. Furthermore, the net might contain some elements that are not in the ppc, as they are out of service or disconnected, which is why the indices of all elements have to be identified through the lookup tables in `net._pd2ppc_lookups`.
#
# Some notes on the internal data structure can be found in [internal_datastructure.ipynb](internal_datastructure.ipynb)
# ## Timings
# Comparing the runopp function (that runs an OPF through PYPOWER) and the runpm function shows that PowerModels is much more performant:
# %timeit pp.runopp(net)
# %timeit pp.runpm_ac_opf(net)
| tutorials/opf_powermodels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import networkx as nx
import scipy.io
import anndata
import scanpy as sc
from networkx.algorithms.bipartite import biadjacency_matrix
import scglue
# -
# # scRNA-seq
# ## Read data
rna_counts = pd.read_table("../../data/download/Ma-2020/GSM4156608_skin.late.anagen.rna.counts.txt.gz", index_col=0)
rna_obs = pd.DataFrame(index=rna_counts.columns)
pd.DataFrame(index=rna_counts.index)
rna_obs.index = rna_obs.index.str.replace(",", ".")
rna_var = pd.DataFrame(index=rna_counts.index)
rna_obs.index.name, rna_var.index.name = "cells", "genes"
rna = anndata.AnnData(
X=scipy.sparse.csr_matrix(rna_counts.to_numpy().T),
obs=rna_obs,
var=rna_var
)
rna
# ## Process meta
rna.obs["domain"] = "scRNA-seq"
rna.obs["protocol"] = "SHARE-seq"
rna.obs["dataset"] = "Ma-2020-RNA"
scglue.data.get_gene_annotation(
rna, gtf="../../data/genome/gencode.vM25.chr_patch_hapl_scaff.annotation.gtf.gz", gtf_by="gene_name"
)
rna.var["genome"] = "mm10"
# # ATAC
# ## Read data
atac_counts = scipy.io.mmread("../../data/download/Ma-2020/GSM4156597_skin.late.anagen.counts.txt.gz")
atac_obs = pd.read_table(
"../../data/download/Ma-2020/GSM4156597_skin.late.anagen.barcodes.txt.gz",
header=None, names=["Cells"], index_col=0
)
atac_var = pd.read_table(
"../../data/download/Ma-2020/GSM4156597_skin.late.anagen.peaks.bed.gz",
header=None, names=["chrom", "chromStart", "chromEnd"]
)
atac_obs.index.name, atac_var.index.name = "cells", "peaks"
atac = anndata.AnnData(
X=atac_counts.T.tocsr(),
obs=atac_obs,
var=atac_var
)
atac
# ## Process meta
atac.obs["domain"] = "scATAC-seq"
atac.obs["protocol"] = "SHARE-seq"
atac.obs["dataset"] = "Ma-2020-ATAC"
atac.var.index = pd.Index(
atac.var["chrom"] + ":" +
atac.var["chromStart"].astype(str) + "-" +
atac.var["chromEnd"].astype(str),
name=atac.var.index.name
)
atac.var["genome"] = "mm10"
# # FRAGS2RNA
frags2rna = scglue.data.bedmap2anndata("../../data/download/Ma-2020/GSM4156597_skin.late.anagen.atac.fragments.bedmap.gz")
frags2rna.obs.index = frags2rna.obs.index.str.replace(",", ".")
frags2rna.obs.index.name, frags2rna.var.index.name = "cells", "genes"
frags2rna
# # Pair samples & add cell types
cell_type = pd.read_table("../../data/download/Ma-2020/celltype_v2.txt")
cell_type.shape
cell_type["celltype"] = cell_type["celltype"].replace({
"Dermal Fibrobalst": "Dermal Fibroblast",
"Hair Shaft-cuticle.cortex": "Hair Shaft-Cuticle/Cortex",
"K6+ Bulge Companion Layer": "K6+ Bulge/Companion Layer",
"ahighCD34+ bulge": "ahigh CD34+ bulge",
"alowCD34+ bulge": "alow CD34+ bulge"
})
cell_type = cell_type.query("celltype != 'Mix'")
cell_type.shape
# ATAC barcodes do not match, need some conversion...
# +
atac_bc_map = {
"04": "53",
"05": "53",
"06": "54",
"07": "55",
"08": "56"
}
@np.vectorize
def map_atac_bc(x):
xs = x.split(".")
xs[-1] = atac_bc_map[xs[-1]]
return ".".join(xs)
cell_type["atac.bc.mapped"] = map_atac_bc(cell_type["atac.bc"])
# -
rna = rna[cell_type["rna.bc"].to_numpy(), :]
rna.obs["cell_type"] = cell_type["celltype"].to_numpy()
atac = atac[cell_type["atac.bc.mapped"].to_numpy(), :]
atac.obs["cell_type"] = cell_type["celltype"].to_numpy()
frags2rna = frags2rna[cell_type["atac.bc"].to_numpy(), :]
frags2rna.obs["cell_type"] = cell_type["celltype"].to_numpy()
frags2rna.obs.index = atac.obs.index
# # Clean data
retained_genes = rna.var.dropna(subset=["chrom", "chromStart", "chromEnd"]).index
rna = rna[:, retained_genes]
rna.var = rna.var.astype({"chromStart": int, "chromEnd": int})
rna
sc.pp.filter_genes(rna, min_counts=1)
rna
blacklist_overlap = scglue.genomics.window_graph(
scglue.genomics.Bed(atac.var.assign(name=atac.var_names)),
"../genome/Blacklist/lists/mm10-blacklist.v2.bed.gz",
window_size=0
)
retained_peaks = np.asarray(biadjacency_matrix(
blacklist_overlap, atac.var_names
).sum(axis=1)).ravel() == 0
atac = atac[:, retained_peaks]
atac.var = atac.var.astype({"chromStart": int, "chromEnd": int})
atac
sc.pp.filter_genes(atac, min_counts=1)
atac
missing_vars = list(set(rna.var_names).difference(frags2rna.var_names))
frags2rna = anndata.concat([
frags2rna, anndata.AnnData(
X=scipy.sparse.csr_matrix((frags2rna.shape[0], len(missing_vars))),
obs=pd.DataFrame(index=frags2rna.obs_names), var=pd.DataFrame(index=missing_vars)
)
], axis=1, merge="first")
frags2rna = frags2rna[:, rna.var_names].copy() # Keep the same features as RNA
frags2rna
# # Process data
sc.pp.highly_variable_genes(rna, n_top_genes=2000, flavor="seurat_v3")
rna.var.highly_variable.sum()
# # Save data
rna.write("../../data/dataset/Ma-2020-RNA.h5ad", compression="gzip")
atac.write("../../data/dataset/Ma-2020-ATAC.h5ad", compression="gzip")
frags2rna.write("../../data/dataset/Ma-2020-FRAGS2RNA.h5ad", compression="gzip")
| data/collect/Ma-2020.ipynb |
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + [markdown] cell_id="dcf6b326-6955-49c9-9cc2-bedea44ab81d" deepnote_cell_type="markdown" tags=[]
/ # Procesamiento de datos - ETL
/ + [markdown] cell_id="53c33bbe-3018-4fc5-a698-c383541e69b9" deepnote_cell_type="markdown" tags=[]
/ # Extract
/ + cell_id="273f43e0-7ea3-4218-b9cf-458cb76017a7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=147 execution_start=1646000242506 source_hash="820cf9ee" tags=[]
import numpy as np
import pandas as pd
import requests
import os
from zipfile import ZipFile
from sqlalchemy import create_engine
/ + [markdown] cell_id="1dadc07e-4e34-4a45-b7d0-c6c533e8aa40" deepnote_cell_type="markdown" tags=[]
/ Se extrae la data de la encuesta de 2021 del sitio [oficial de Stack Overflow](https://insights.stackoverflow.com/survey) con web scraping.
/ + cell_id="7cffa243-8303-422e-b49c-7c886f7d325f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=9251 execution_start=1646000242696 source_hash="63800a23" tags=[]
# Get the 2021 survey (in a zip file) from Stack Overflow
path = 'https://info.stackoverflowsolutions.com/rs/719-EMH-566/images/stack-overflow-developer-survey-2021.zip'
response = requests.get(path)
print(response.status_code)
# Save the file locally
local_path = os.path.join(os.getcwd(), os.pardir, 'data', 'raw', 'survey_2021.zip')
with open(local_path, "wb") as f:
f.write(response.content)
/ + cell_id="117fc44d-86e3-48bf-b2bb-be32d7da93b7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=486 execution_start=1646000251777 source_hash="bca40b07" tags=[]
# Get the list of files
path_save = os.path.join(os.getcwd(), os.pardir, 'data', 'raw')
with ZipFile(local_path, "r") as f:
file_names = f.namelist()
print(file_names)
csv_file_path_1 = f.extract(file_names[2], path_save)
print(csv_file_path_1)
csv_file_path_2 = f.extract(file_names[3], path_save)
print(csv_file_path_2)
/ + [markdown] cell_id="a5c27b88-dc2c-4ced-bcfa-8a0a7da7f82a" deepnote_cell_type="markdown" tags=[]
/ # Transform
/ + cell_id="31a5dd80-5c4b-4aae-b4a6-94c2e8e2a7cd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1170 execution_start=1646000252265 source_hash="a1e324eb" tags=[]
# Read de data
path = os.path.join(os.getcwd(), os.pardir, 'data', 'raw')
df_schema = pd.read_csv(os.path.join(path, 'survey_results_schema.csv'))
df_survey = pd.read_csv(os.path.join(path, 'survey_results_public.csv'))
/ + [markdown] cell_id="2ae9495a-b87d-471b-8692-7d909e98250d" deepnote_cell_type="markdown" tags=[]
/ ### Filtro de datos
/
/ El `df_schema` contiene las preguntas usadas en la encuesta.
/ + cell_id="c751c12e-9b4a-400f-8d26-bc1101db8991" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1646000253450 source_hash="366e7cf0" tags=[]
# Explore what data to use
cols = ['qname', 'question']
df_schema[cols]
/ + cell_id="737489f0-3b4a-4848-b0c1-74ab74469ea8" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=96 execution_start=1646000253470 source_hash="77e67714" tags=[]
df_survey.sample(5)
/ + [markdown] cell_id="560413b1-9f1c-4445-bc54-dede3316235e" deepnote_cell_type="markdown" tags=[]
/ Seleccionar las columnas que pueden ayudar a contestar las preguntas (mostradas en el tercer notebook). Y también los países hispanohablantes para filtrar la data (también se incluyó a Brasil).
/ + cell_id="ad6bc086-52e3-45b5-9e56-1fef3e644e22" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=11 execution_start=1646000253574 source_hash="90ffcd9b" tags=[]
# Select columns
columns = ['ResponseId', 'Age', 'Gender', 'Sexuality', 'Country',
'EdLevel', 'LearnCode', 'YearsCode', 'YearsCodePro',
'Employment', 'ConvertedCompYearly']
# Filter registers by countries
latam = ['Peru', 'Colombia', 'Chile', 'Argentina', 'Costa Rica', 'Bolivia',
'Uruguay', 'Mexico', 'Venezuela, Bolivarian Republic of...'
'Dominican Republic', 'Ecuador', 'Guatemala', 'Paraguay', 'Panama',
'El Salvador', 'Nicaragua', 'Brazil', 'Spain']
# New dataset
in_latam = df_survey.Country.isin(latam)
df = df_survey[in_latam][columns].set_index('ResponseId')
/ + cell_id="49fdfe8a-344a-44bb-ab16-01603888694d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1646000253586 source_hash="cba0f1cb" tags=[]
s = df.shape
p = s[0] / df_survey.shape[0] * 100
print(f'Registros: {s[0]}')
print(f'Porcentaje total del dataset: {round(p,2)}%')
print(f'Columnas: {s[1]}')
/ + [markdown] cell_id="d576ea7e-f1ec-4ca4-8a50-993acbdbd1d2" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1645585403819 source_hash="1d89ab00" tags=[]
/ Veamos los tipos de datos y qué contienen las variables
/ + cell_id="16ee751e-5197-4c1e-9470-e73178fb5fe7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1646000253599 source_hash="de1e323c" tags=[]
df.info()
/ + cell_id="53827ed4-8dba-488e-a0c1-3fe976d55865" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=53 execution_start=1646000253608 source_hash="b1cecf2d" tags=[]
df.describe(include='all')
/ + [markdown] cell_id="d4c043e7-c1d3-434e-bb82-1afa071bd6de" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1645582744382 source_hash="98a4e821" tags=[]
/ Hay algunas columnas que podrían ser numéricas, pero están como categóricas, ¿por qué?
/ + cell_id="c585b081-f82d-4fe1-8c36-cbedfeb47f11" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1646000253665 source_hash="781be1ef" tags=[]
df.YearsCode.value_counts().sort_values().head()
/ + cell_id="94420460-adb6-4126-aeb3-bbbb5541e286" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1646000253697 source_hash="f55248f5" tags=[]
df.YearsCodePro.value_counts().sort_values().head()
/ + [markdown] cell_id="5e8229fa-bbe2-4c8d-8288-e84b87947c54" deepnote_cell_type="markdown" tags=[]
/ Como solo hay 1 valor de "más de 50 años" para cada columna, se lo reemplazará con `50`. Y los valores de "menos de 1 año" con `0.5`. De esa manera se podrá tener columnas numéricas.
/ + cell_id="c20c7104-ea5d-4508-af49-8cfecead88bb" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1646000253698 source_hash="a99a61c7" tags=[]
df['YearsCode'] = pd.to_numeric(df.YearsCode.replace(['More than 50 years', 'Less than 1 year'], [50, 0.5]))
df['YearsCodePro'] = pd.to_numeric(df.YearsCodePro.replace(['More than 50 years', 'Less than 1 year'], [50, 0.5]))
/ + [markdown] cell_id="90d67c94-bd47-4b31-9cd3-9a8947bd85fd" deepnote_cell_type="markdown" tags=[]
/ ### Reducir categorías
/
/ Hay varias preguntas que tienen múltiples opciones, pero son bajas porcentualmente. Se las reducirá a menos categorías para simplificar el análisis.
/ + [markdown] cell_id="27a2fd08-434a-4f26-825d-622da5cabb0c" deepnote_cell_type="markdown" tags=[]
/ #### Género
/ + cell_id="ed40c769-dae5-4d3b-9be6-2de75b91f695" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1646000253699 source_hash="7b71ac80" tags=[]
df.Gender.value_counts(normalize=True)
/ + cell_id="c19e60b4-2440-49a5-8e79-d0a0bd18b153" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1646000253701 source_hash="b840ec0e" tags=[]
df.Gender.where(df.Gender.isin(['Man', 'Woman']), 'Other', inplace=True)
df.Gender.value_counts(normalize=True)
/ + [markdown] cell_id="c6d5bf25-b727-4ce3-af60-72d0eb2b43f8" deepnote_cell_type="markdown" tags=[]
/ #### Edad
/ + cell_id="3761515e-ee8e-4d90-a0a4-eff97e962a3a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1646000253730 source_hash="98aff8ca" tags=[]
df.Age.value_counts(normalize=True)
/ + cell_id="90564676-8467-4804-9daf-8e463fb6c903" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1646000253731 source_hash="ba1ee9f9" tags=[]
df.Age.where(~(df.Age.isin(['45-54 years old', '55-64 years old', '65 years or older'])), '> 45 years old', inplace=True)
df.drop(df.Age[df.Age == 'Prefer not to say'].index, inplace=True)
df.Age.value_counts(normalize=True)
/ + [markdown] cell_id="e2eb8e87-9cf4-4d17-ba00-2c83473284f0" deepnote_cell_type="markdown" tags=[]
/ #### Sexuality
/ + cell_id="bd3f318d-292b-4d97-8fb9-4de911af2f2c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1646000253732 source_hash="c715ee4a" tags=[]
df.Sexuality.value_counts(normalize=True)
/ + cell_id="8fdc9d72-cb95-4645-8ea9-92277295a10e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1646000253737 source_hash="600337db" tags=[]
df.Sexuality.where(df.Sexuality.isin(['Straight / Heterosexual']), 'LGBT / Non-hetero', inplace=True)
df.Sexuality.value_counts(normalize=True)
/ + [markdown] cell_id="c6a4d989-f82b-464c-8909-8a20207de040" deepnote_cell_type="markdown" tags=[]
/ #### Empleo
/ + cell_id="c482204e-088b-40dc-9074-0277cc82b4e4" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1646000253743 source_hash="c1ebf45b" tags=[]
df.Employment.value_counts(normalize=True)
/ + cell_id="ac75eadf-a363-4c81-a129-19afeeb44b36" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=9 execution_start=1646000253753 source_hash="96df75a1" tags=[]
df.Employment.where(~(df.Employment.isin(['Employed full-time', 'Employed part-time'])), 'Employed', inplace=True)
df.Employment.where(~(df.Employment.isin(['Student, full-time', 'Student, part-time'])), 'Student', inplace=True)
df.Employment.where(~(df.Employment.isin(['Not employed, but looking for work', 'Not employed, and not looking for work', 'Retired'])), \
'Not employed', inplace=True)
df.drop(df.Employment[df.Employment == 'I prefer not to say'].index, inplace=True)
df.Employment.value_counts(normalize=True)
/ + [markdown] cell_id="55fefee8-688a-4fe0-9d21-81840b0e25e9" deepnote_cell_type="markdown" tags=[]
/ #### Dónde aprendió a programar
/
/ Se cambia las categorías por 3: `Tradicional`, `No tradicional`, `Ambos`
/ + cell_id="84f377f0-c8c0-4d96-a4dc-d9ce8382f846" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1646000253768 source_hash="7650561b" tags=[]
df.LearnCode.value_counts(normalize=True)
/ + cell_id="80c388e5-cd78-4a4f-b070-f20b95c005df" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1646000253775 source_hash="8afe11ce" tags=[]
typeEdu = []
for i in list(df.LearnCode.values):
if pd.isnull(i):
typeEdu.append(np.nan)
elif i == ('School'):
typeEdu.append('Traditional')
elif str(i).find('School') == -1:
typeEdu.append('Non-traditional')
else:
typeEdu.append('Both')
df['LearnCode'] = typeEdu
/ + [markdown] cell_id="ce8e0f15-eebb-44d3-8006-5a226cad2d2a" deepnote_cell_type="markdown" tags=[]
/ ### Agregar data
/
/ Columna para conocer cuántos años tomó desde que se aprendió a programar hasta que se lo hizo profesionalmente.
/ + cell_id="b2b748ef-0b57-4e5a-ac25-67bbd4424854" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1646000253796 source_hash="4b5c1427" tags=[]
df['YearsLearnPro'] = abs(df.YearsCode - df.YearsCodePro)
df.drop(columns=['YearsCode', 'YearsCodePro'], inplace=True)
/ + [markdown] cell_id="87ab5a36-c5a4-43f6-a8f0-ef2fa8ac34da" deepnote_cell_type="markdown" deepnote_to_be_reexecuted=false execution_millis=13 execution_start=1645721960540 source_hash="dd1b531b" tags=[]
/ Conocer si tiene o no un título universitario.
/ + cell_id="7104969c-38b3-4018-b764-19b7e029e4f9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1646000253797 source_hash="16e615e9" tags=[]
df.EdLevel.value_counts()
/ + cell_id="9afa0b7e-148f-4235-9d2b-225c7318cbb4" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1646000253798 source_hash="481457fa" tags=[]
not_degree = ['Some college/university study without earning a degree', 'Primary/elementary school',
'Secondary school (e.g. American high school, German Realschule or Gymnasium, etc.)', 'Something else']
df['Degree'] = df.EdLevel.where(df.EdLevel.isin(not_degree), 'Yes')
df['Degree'] = df.Degree.where(df.Degree == 'Yes', 'No')
/ + cell_id="fd940d8a-da3b-431c-a39f-393fc1817b2f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=3 execution_start=1646000253801 source_hash="ad988381" tags=[]
df.Degree.value_counts(normalize=True)
/ + [markdown] cell_id="1b8f430d-2e07-4091-95f7-dcc71e55f575" deepnote_cell_type="markdown" tags=[]
/ # Load
/ + [markdown] cell_id="6a1bcd29-5c40-4408-80a9-62da9f2bd069" deepnote_cell_type="markdown" tags=[]
/ El dataset está listo para poder trabajar. Así que se lo exportará a dos lugares distintos:
/ + [markdown] cell_id="31690acd-fe41-4f80-8634-39e4637bd2e2" deepnote_cell_type="markdown" tags=[]
/ #### 1. A una base de datos PostgreSQL en la nube:
/ + cell_id="c49760fe-42ad-4334-80f4-ca61b900a12c" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1646000253847 source_hash="f72ba7c6" tags=[]
# Environment variables
HOST = os.environ["HOST"]
DATABASE = os.environ["DATABASE"]
USER = os.environ["USER"]
PORT = int(os.environ["PORT"])
PASSWORD = os.<PASSWORD>["PASSWORD"]
/ + cell_id="74403bc4-01cf-443d-87c5-4c7171504a30" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=621 execution_start=1646000433197 source_hash="74130066" tags=[]
# Create the engine (created with environment variables)
engine = create_engine(f"postgresql://{USER}:{PASSWORD}@{HOST}:{PORT}/{DATABASE}")
df.to_sql('survey_s_2021', con=engine, index=True, index_label='ResponseId', if_exists='replace')
/ + [markdown] cell_id="465aeb66-25c5-4bde-b8f5-613bf1ce8a3f" deepnote_cell_type="markdown" tags=[]
/ #### 2. Como archivo csv al directorio `/data/processed/`:
/ + cell_id="b866c922-e4d7-4d8c-86e3-5b90f36c9fd2" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1646000312536 source_hash="29073edd" tags=[]
path_processed = os.path.join(os.getcwd(), os.pardir, 'data', 'processed')
df.to_csv(os.path.join(path_processed, 'survey.csv'))
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=09491c61-3767-4289-98fd-88aee19bb45d' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='<KEY> > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| notebooks/1-etl-process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Aujourd'hui on roule sur les mecs de l'ENS
#
#
# https://challengedata.ens.fr/en/challenge/39/prediction_of_transaction_claims_status.html
# # Imports des librairies de bases
#
# On ajoutera celles qui manquent au fur et à mesure de nos besoins
# +
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import pandas as pd
import os, gc
# -
# # Définition de la seed pour le random
#
# Très important pour qu'on voit les mêmes choses entre nos deux ordis
RANDOM_SEED = 42;
np.random.seed(RANDOM_SEED)
# # Définition des paramètres pour Matplot
#
# Rien de bien intéréssant
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# # Set des variables globales
#
# Attention, je n'utilise les variables globales pour la gestion des fichiers. Sinon, c'est mort
# Where to save the figures
PROJECT_ROOT_DIR = ".."
DATA_PROCESSED = os.path.join(PROJECT_ROOT_DIR, "data_processed")
# # Fonction pour load les libraires
#
# En vrai, on a juste besoin de pd.read_csv, mais c'était pour faire joli
def load_data(file,data_path=DATA_PROCESSED, sep=';'):
csv_path = os.path.join(data_path, file)
return pd.read_csv(csv_path, sep=';')
# # On load les jeux de données
TX_data = load_data(file = "train.csv");
TX_data.drop(['CARD_PAYMENT','COUPON_PAYMENT','RSP_PAYMENT','WALLET_PAYMENT'], axis = 1, inplace = True)
TX_data.info() # 42 colonnes, c'est un nombre qui fait plaisir
from sklearn.model_selection import train_test_split
train_set, test_set = train_test_split(TX_data,
test_size=0.3,
random_state=RANDOM_SEED,
stratify=TX_data["CLAIM_TYPE"]
)
del TX_data;
# # Jointure entre les X et Y
def datapreprocess(data):
data=data.apply(pd.to_numeric, errors='ignore')
# Y and X
try :
Y=data["CLAIM_TYPE"]
X=data.drop("CLAIM_TYPE", axis=1,inplace=False)
except:
Y=0
X=data
# Exclude Objets
X=X.select_dtypes(exclude=['object']) # j'exclude les variables catégorielles que j'ai oublié
# Work on fare
from sklearn.preprocessing import Imputer
imp = Imputer(missing_values='NaN',strategy='median', axis=1)
X=pd.DataFrame(imp.fit_transform(X),columns=X.columns.values)
return X, Y
# +
X_train, Y_train = datapreprocess(train_set)
X_test, Y_test = datapreprocess(test_set)
gc.collect()
# -
def multiclass_roc_auc_score(truth, pred):
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import LabelBinarizer
lb = LabelBinarizer()
lb.fit(truth)
return roc_auc_score(lb.transform(truth), lb.transform(pred), average="weighted")
from sklearn.metrics import confusion_matrix
def plot_confusion_matrix(matrix):
"""If you prefer color and a colorbar"""
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111)
cax = ax.matshow(matrix)
fig.colorbar(cax)
# # MODEL!
# ## LDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
model = LinearDiscriminantAnalysis()
kfold = KFold(n_splits=3, random_state=7)
result = cross_val_score(model, X, y, cv=kfold, scoring='accuracy')
print(result.mean())
from xgboost import XGBClassifier
# ### XGBoost solo
# +
params_XGB={
# General Parameters - the overall functioning
'booster':'gbtree',
'silent':0,
#'nthread':4, # Je le commente, puisque il détecte automatiquement le nombre de cores qu'il peut utiliser.
'n_estimators' : 1000,
# Booster Parameters - the individual booster (tree/regression) at each step
'learning_rate' : 0.1,
'min_child_weight' : 1, #A smaller value is chosen because it is a highly imbalanced class problem and leaf nodes can have smaller size groups.
'max_depth' : 3,
#'max_leaf_nodes':None, #If this is defined, GBM will ignore max_depth.
'gamma' : 0.3,
'max_delta_step':4, #it might help in logistic regression when class is extremely imbalanced/ 1-10 might help control the update
'subsample' : 0.55,
'colsample_bytree' : 0.85,
'colsample_bylevel':1, #default
'reg_lambda' : 1, #default
'reg_alpha':0,
'scale_pos_weight' : sample_weight_arr,
# Learning Task Parameters - the optimization performed
'objective' : 'multi:softmax', # you also need to set an additional num_class (number of classes)
'num_class' : len(Y_train.unique()),
'eval_metric':"auc",
'seed' : RANDOM_SEED,
}
# -
xgb_clf = XGBClassifier(**params_XGB)
xgb_clf.fit(
X=X_train,
y=Y_train,
sample_weight=sample_weight_arr,
eval_set=None,
eval_metric='auc',
early_stopping_rounds=None,
verbose=True,
xgb_model=None
)
y_pred_xgb_train = xgb_clf.predict(X_train)
y_pred_xgb = xgb_clf.predict(X_test)
train_mAUC = multiclass_roc_auc_score(Y_train, y_pred_xgb_train)
test_mAUC = multiclass_roc_auc_score(Y_test, y_pred_xgb)
print("Performance sur le train : {}".format(train_mAUC))
print("Performance sur le test : {}".format(test_mAUC))
# Performance sur le train : 0.6589482571844301
#
# Performance sur le test : 0.6180906043249655
#
conf_mx = confusion_matrix(Y_test, y_pred_xgb)
row_sums = conf_mx.sum(axis=1, keepdims=True)
norm_conf_mx = conf_mx / row_sums
plot_confusion_matrix(norm_conf_mx)
# #### C'est un beau score pour le XBoost
#
# Cependant, j'ai optimisé pour la mauvaise métrique, et j'ai toujours pas fait le Grid Search
pd.DataFrame(xgb_clf.feature_importances_, index=X_train.columns, columns=["Feature"]).sort_values(by="Feature", ascending=False)
# ### XGBoost Grid Search Iterate
#
# Comme mon PC est pourri, je vais chercher les paramètres de façon iterative
from sklearn.model_selection import GridSearchCV
# #### Paramètres pour le XGB qui ne changent pas
# +
params_XGB={
# General Parameters - the overall functioning
'booster':'gbtree',
'silent':0,
#'nthread':4, # Je vais le commenter, puisque il détecte automatiquement le nombre de cores qu'il peut utiliser.
'n_estimators' : 100,
# Booster Parameters - the individual booster (tree/regression) at each step
'learning_rate' : 0.1,
'colsample_bylevel':1, #default
'reg_lambda' : 1, #default
'scale_pos_weight' : sample_weight_arr,
# Learning Task Parameters - the optimization performed
'objective' : 'multi:softmax',
'num_class' : len(Y_train.unique()),
'eval_metric':"auc",
'seed' : RANDOM_SEED,
}
# -
# #### Paramètres pour la méthode `fit` de XGB qui ne changent pas
fit_params_xgb_cv={
'sample_weight': sample_weight_arr,
'eval_set' : None,
'eval_metric' : 'auc',
'early_stopping_rounds' : None,
'verbose':True,
'xgb_model':None
}
# ### Optim 1 : max_depth, min_child_weight et max_delta_step
# Paramètres pour le GridSearch
params_XGB_CV = {
'max_depth':range(2,5),
'min_child_weight':range(1,6),
'max_delta_step':list(range(1,5)),
}
xgb_gs_cv = GridSearchCV(XGBClassifier(**params_XGB),
params_XGB_CV,
n_jobs=-1,
verbose=1)
xgb_gs_cv.fit(
X = X_train,
y=Y_train,
groups=None,
**fit_params_xgb_cv
)
print(xgb_gs_cv.best_estimator_)
print("ROC score : {}".format(multiclass_roc_auc_score(Y_test, xgb_gs_cv.predict(X_test))))
# Donc les paramètres optimaux sont:
# 1. `max_depth` :
# 2. `min_child_weight` :
# 3. `max_delta_step` :
#
# ### Optim 2 : gamma, et subsample
# Paramètres pour le GridSearch
params_XGB_CV = {
'gamma':[i/10.0 for i in range(0,5)],
'subsample':[i/10.0 for i in range(6,10)],
}
xgb_gs_cv = GridSearchCV(XGBClassifier(**params_XGB),
params_XGB_CV,
n_jobs=-1,
verbose=1)
xgb_gs_cv.fit(
X = X_train,
y=Y_train,
groups=None,
**fit_params_xgb_cv
)
print(xgb_gs_cv.best_estimator_)
print("ROC score : {}".format(multiclass_roc_auc_score(Y_test, xgb_gs_cv.predict(X_test))))
# Donc les paramètres optimaux sont:
# 1. `gamma` :
# 2. `subsample` :
# ### Optim 3 : colsample_bytree, et reg_alpha
# Paramètres pour le GridSearch
params_XGB_CV = {
'colsample_bytree':[i/10.0 for i in range(6,10)],
'reg_alpha':[0, 0.001, 0.005, 0.01, 0.05]
}
xgb_gs_cv = GridSearchCV(XGBClassifier(**params_XGB),
params_XGB_CV,
n_jobs=-1,
verbose=1)
xgb_gs_cv.fit(
X = X_train,
y=Y_train,
groups=None,
**fit_params_xgb_cv
)
print(xgb_gs_cv.best_estimator_)
print("ROC score : {}".format(multiclass_roc_auc_score(Y_test, xgb_gs_cv.predict(X_test))))
# Donc les paramètres optimaux sont:
# 1. `colsample_bytree` :
# 2. `reg_alpha` :
conf_mx = confusion_matrix(Y_test, y_pred_xgb_cv)
row_sums = conf_mx.sum(axis=1, keepdims=True)
norm_conf_mx = conf_mx / row_sums
plot_confusion_matrix(norm_conf_mx)
| Models/.ipynb_checkpoints/LDA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# metadata:
# interpreter:
# hash: 31f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# name: python3
# ---
# https://adventofcode.com/2015/day/3
#
inp = open("../input/03.txt").read().strip()
# + tags=[]
# Answer to part 1
p = (0, 0)
seen = set()
mov = {'>': (1,0), '<': (-1,0), '^': (0,1), 'v': (0, -1)}
for c in inp:
seen.add(p)
m = mov[c]
p = (p[0] + m[0], p[1] + m[1])
print(len(seen))
# -
# ## Part 2
# + tags=[]
# inst = '^>v<'
inst = inp
pp = [(0, 0), (0,0)]
seen = set([(0,0)])
mov = {'>': (1,0), '<': (-1,0), '^': (0,1), 'v': (0, -1)}
turn = 0
for c in inst:
m = mov[c]
p = pp[turn]
pp[turn] = (p[0] + m[0], p[1] + m[1])
# print(turn, pp[turn])
seen.add(pp[turn])
turn = 1 - turn
print(len(seen))
# -
# 2342 is too high
| 2015/py/03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + slideshow={"slide_type": "skip"}
import numpy, scipy, matplotlib.pyplot as plt, pandas, librosa
# + [markdown] slideshow={"slide_type": "skip"}
# [← Back to Index](index.html)
# + [markdown] slideshow={"slide_type": "slide"}
# # Onset Detection
# + [markdown] slideshow={"slide_type": "notes"}
# Automatic detection of musical events in an audio signal is one of the most fundamental tasks in music information retrieval. Here, we will show how to detect an *onset*, the start of a musical event.
#
# For more reading, see [this tutorial on onset detection by <NAME>](https://files.nyu.edu/jb2843/public/Publications_files/2005_BelloEtAl_IEEE_TSALP.pdf).
# + [markdown] slideshow={"slide_type": "notes"}
# Load the audio file `simpleLoop.wav` into the NumPy array `x` and sampling rate `fs`.
# + slideshow={"slide_type": "slide"}
x, fs = librosa.load('simpleLoop.wav', sr=44100)
print x.shape
# + [markdown] slideshow={"slide_type": "skip"}
# Plot the signal:
# + slideshow={"slide_type": "fragment"}
librosa.display.waveplot(x, fs)
# + [markdown] slideshow={"slide_type": "skip"}
# Listen:
# + slideshow={"slide_type": "subslide"}
from IPython.display import Audio
Audio(x, rate=fs)
# + [markdown] slideshow={"slide_type": "slide"}
# ## `librosa.onset.onset_detect`
# + [markdown] slideshow={"slide_type": "notes"}
# [`librosa.onset.onset_detect`](http://bmcfee.github.io/librosa/generated/librosa.onset.onset_detect.html) returns the frame indices for estimated onsets in a signal:
# + slideshow={"slide_type": "subslide"}
onsets = librosa.onset.onset_detect(x, fs)
print onsets # frame numbers of estimated onsets
# + [markdown] slideshow={"slide_type": "notes"}
# Plot the onsets on top of a spectrogram of the audio:
# + slideshow={"slide_type": "subslide"}
S = librosa.stft(x)
logS = librosa.logamplitude(S)
librosa.display.specshow(logS, fs, alpha=0.75, x_axis='time')
plt.vlines(onsets, 0, logS.shape[0], color='r')
# + [markdown] slideshow={"slide_type": "slide"}
# ## `essentia.standard.OnsetRate`
# + [markdown] slideshow={"slide_type": "notes"}
# The easiest way in Essentia to detect onsets given a time-domain signal is using [`OnsetRate`](http://essentia.upf.edu/documentation/reference/std_OnsetRate.html). It returns a list of onset times and the onset rate, i.e. number of onsets per second.
# + slideshow={"slide_type": "subslide"}
from essentia.standard import OnsetRate
find_onsets = OnsetRate()
onset_times, onset_rate = find_onsets(x)
print onset_times
print onset_rate
# + [markdown] slideshow={"slide_type": "slide"}
# ## `essentia.standard.AudioOnsetsMarker`
# + [markdown] slideshow={"slide_type": "notes"}
# To verify our results, we can use [`AudioOnsetsMarker`](http://essentia.upf.edu/documentation/reference/std_AudioOnsetsMarker.html) to add a sound at the moment of each onset.
# + slideshow={"slide_type": "subslide"}
from essentia.standard import AudioOnsetsMarker
onsets_marker = AudioOnsetsMarker(onsets=onset_times, type='beep')
x_beeps = onsets_marker(x)
Audio(x_beeps, rate=fs)
# + [markdown] slideshow={"slide_type": "notes"}
# Sounds good!
#
# For more control over the onset detection algorithm, see [`OnsetDetection`](http://essentia.upf.edu/documentation/reference/std_OnsetDetection.html), [`OnsetDetectionGlobal`](http://essentia.upf.edu/documentation/reference/std_OnsetDetectionGlobal.html), and [`Onsets`](http://essentia.upf.edu/documentation/reference/std_Onsets.html).
# + [markdown] slideshow={"slide_type": "skip"}
# [← Back to Index](index.html)
| onset_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# ## Differentiation and Derivatives
# So far in this course, you've learned how to evaluate limits for points on a line. Now you're going to build on that knowledge and look at a calculus technique called *differentiation*. In differentiation, we use our knowledge of limits to calculate the *derivative* of a function in order to determine the rate of change at an individual point on a line.
#
# Let's remind ourselves of the problem we're trying to solve, here's a function:
#
# \begin{equation}f(x) = x^{2} + x\end{equation}
#
# We can visualize part of the line that this function defines using the folllowing Python code:
# +
# %matplotlib inline
# Here's the function
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
plt.show()
# -
# Now, we know that we can calculate the average rate of change for a given interval on the line by calculating the slope for a secant line that connects two points on the line. For example, we can calculate the average change for the interval between x=4 and x=6 by dividing the change (or *delta*, indicated as Δ) in the value of *f(x)* by the change in the value of *x*:
#
# \begin{equation}m = \frac{\Delta{f(x)}}{\Delta{x}} \end{equation}
#
# The delta for *f(x)* is calculated by subtracting the *f(x)* values of our points, and the delta for *x* is calculated by subtracting the *x* values of our points; like this:
#
# \begin{equation}m = \frac{f(x_{2}) - f(x_{1})}{x_{2} - x_{1}} \end{equation}
#
# So for the interval between x=4 and x=6, that's:
#
# \begin{equation}m = \frac{f(6) - f(4)}{6 - 4} \end{equation}
#
# We can calculate and plot this using the following Python:
# +
# %matplotlib inline
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the a values
x1 = 4
x2 = 6
# Get the corresponding f(x) values
y1 = f(x1)
y2 = f(x2)
# Calculate the slope by dividing the deltas
a = (y2 - y1)/(x2 - x1)
# Create an array of x values for the secant line
sx = [x1,x2]
# Use the function to get the y values
sy = [f(i) for i in sx]
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the interval points
plt.scatter([x1,x2],[y1,y2], c='red')
# Plot the secant line
plt.plot(sx,sy, color='magenta')
# Display the calculated average rate of change
plt.annotate('Average change =' + str(a),(x2, (y2+y1)/2))
plt.show()
# -
# The average rate of change for the interval between x=4 and x=6 is <sup>11</sup>/<sub>1</sub> (or simply 11), meaning that for every **1** added to *x*, *f(x)* increases by **11**. Put another way, if x represents time in seconds and f(x) represents distance in meters, the average rate of change for distance over time (in other words, *velocity*) for the 4 to 6 second interval is 11 meters-per-second.
#
# So far, this is just basic algebra; but what if instead of the average rate of change over an interval, we want to calculate the rate of change at a single point, say, where x = 4.5?
#
# One approach we could take is to create a secant line between the point at which we want the slope and another point on the function line that is infintesimally close to it. So close in fact that the secant line is actually a tangent that goes through both points. We can then calculate the slope for the secant line as before. This would look something like the graph produced by the following code:
# +
# %matplotlib inline
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11, 1))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the x1 point, arbitrarily 5
x1 = 4.5
y1 = f(x1)
# Set the x2 point, very close to x1
x2 = 5.000000001
y2 = f(x2)
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the point
plt.scatter(x1,y1, c='red')
plt.annotate('x' + str(x1),(x1,y1), xytext=(x1-0.5, y1+3))
# Approximate the tangent slope and plot it
m = (y2-y1)/(x2-x1)
xMin = x1 - 3
yMin = y1 - (3*m)
xMax = x1 + 3
yMax = y1 + (3*m)
plt.plot([xMin,xMax],[yMin,yMax], color='magenta')
plt.show()
# -
# ## Calculating a Derivative
# In the Python code above, we created the (almost) tangential secant line by specifying a point that is very close to the point at which we want to calculate the rate of change. This is adequate to show the line conceptually in the graph, but it's not a particularly generalizable (or accurate) way to actually calculate the line so that we can get the rate of change at any given point.
#
# If only we knew of a way to calculate a point on the line that is as close as possible to point with a given *x* value.
#
# Oh wait, we do! It's a *limit*.
#
# So how do we apply a limit in this scenario? Well, let's start by examining our general approach to calculating slope in a little more detail.Our tried and tested approach is to plot a secant line between two points at different values of x, so let's plot an arbitrary (*x,y*) point, and then add an arbitrary amount to *x*, which we'll call *h*. Then we know that we can plot a secant line between (*x,f(x)*) and (*x+h,f(x+h)*) and find its slope.
#
# Run the cell below to see these points:
# +
# %matplotlib inline
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the x point
x1 = 3
y1 = f(x1)
# set the increment
h = 3
# set the x+h point
x2 = x1+h
y2 = f(x2)
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the x point
plt.scatter(x1,y1, c='red')
plt.annotate('(x,f(x))',(x1,y1), xytext=(x1-0.5, y1+3))
# Plot the x+h point
plt.scatter(x2,y2, c='red')
plt.annotate('(x+h, f(x+h))',(x2,y2), xytext=(x2+0.5, y2))
plt.show()
# -
# As we saw previously, our formula to calculate slope is:
#
# \begin{equation}m = \frac{\Delta{f(x)}}{\Delta{x}} \end{equation}
#
# The delta for *f(x)* is calculated by subtracting the *f(x + h)* and *f(x)* values of our points, and the delta for *x* is just the difference between *x* and *x + h*; in other words, *h*:
#
# \begin{equation}m = \frac{f(x + h) - f(x)}{h} \end{equation}
#
# What we actually need is the slope at the shortest possible distance between x and x+h, so we're looking for the smallest possible value of *h*. In other words, we need the limit as *h* approaches 0.
#
# \begin{equation}\lim_{h \to 0} \frac{f(x + h) - f(x)}{h} \end{equation}
#
# This equation is generalizable, and we can use it as the definition of a function to help us find the slope at any given value of *x* on the line, and it's what we call the *derivative* of our original function (which in this case is called *f*). This is generally indicated in *Lagrange* notation like this:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{f(x + h) - f(x)}{h} \end{equation}
#
# You'll also sometimes see derivatives written in *Leibniz's* notation like this:
#
# \begin{equation}\frac{d}{dx}f(x) = \lim_{h \to 0} \frac{f(x + h) - f(x)}{h} \end{equation}
#
# ***Note:*** *Some textbooks use **h** to symbolize the difference between **x<sub>0</sub>** and **x<sub>1</sub>**, while others use **Δx**. It makes no diffrerence which symbolic value you use.*
# #### Alternate Form for a Derivative
# The formula above shows the generalized form for a derivative. You can use the derivative function to get the slope at any given point, for example to get the slope at point *a* you could just plug the value for *a* into the generalized derivative function:
#
# \begin{equation}f'(\textbf{a}) = \lim_{h \to 0} \frac{f(\textbf{a} + h) - f(\textbf{a})}{h} \end{equation}
#
# Or you could use the alternate form, which is specific to point *a*:
#
# \begin{equation}f'(a) = \lim_{x \to a} \frac{f(x) - f(a)}{x - a} \end{equation}
#
# These are mathematically equivalent.
# ### Finding the Derivative for a Specific Point
# It's easier to understand differentiation by seeing it in action, so let's use it to find the derivitive for a specific point in the function ***f***.
#
# Here's the definition of function ***f***:
#
# \begin{equation}f(x) = x^{2} + x\end{equation}
#
# Let's say we want to find ***f'(2)*** (the derivative for ***f*** when ***x*** is 2); so we're trying to find the slope at the point shown by the following code:
# +
# %matplotlib inline
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the point
x1 = 2
y1 = f(x1)
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the point
plt.scatter(x1,y1, c='red')
plt.annotate('(x,f(x))',(x1,y1), xytext=(x1-0.5, y1+3))
plt.show()
# -
# Here's our generalized formula for finding a derivative at a specific point (*a*):
#
# \begin{equation}f'(a) = \lim_{h \to 0} \frac{f(a + h) - f(a)}{h} \end{equation}
#
# So let's just start by plugging our *a* value in:
#
# \begin{equation}f'(\textbf{2}) = \lim_{h \to 0} \frac{f(\textbf{2} + h) - f(\textbf{2})}{h} \end{equation}
#
# We know that ***f(x)*** encapsulates the equation ***x<sup>2</sup> + x***, so we can rewrite our derivative equation as:
#
# \begin{equation}f'(2) = \lim_{h \to 0} \frac{((2+h)^{2} + 2 + h) - (2^{2} + 2)}{h} \end{equation}
#
# We can apply the distribution property to ***(2 + h)<sup>2</sup>*** using the rule that *(a + b)<sup>2</sup> = a<sup>2</sup> + b<sup>2</sup> + 2ab*:
#
# \begin{equation}f'(2) = \lim_{h \to 0} \frac{(4 + h^{2} + 4h + 2 + h) - (2^{2} + 2)}{h} \end{equation}
#
# Then we can simplify 2<sup>2</sup> + 2 (2<sup>2</sup> is 4, plus 2 gives is 6):
#
# \begin{equation}f'(2) = \lim_{h \to 0} \frac{(4 + h^{2} + 4h + 2 + h) - 6}{h} \end{equation}
#
# We can combine like terms on the left side of the numerator to make things a little clearer:
#
# \begin{equation}f'(2) = \lim_{h \to 0} \frac{(h^{2} + 5h + 6) - 6}{h} \end{equation}
#
# Which combines even further to get rid of the *6*:
#
# \begin{equation}f'(2) = \lim_{h \to 0} \frac{h^{2} + 5h}{h} \end{equation}
#
# And finally, we can simplify the fraction:
#
# \begin{equation}f'(2) = \lim_{h \to 0} h + 5 \end{equation}
#
# To get the limit when *h* is approaching 0, we can use direct substitution for h:
#
# \begin{equation}f'(2) = 0 + 5 \end{equation}
#
# so:
#
# \begin{equation}f'(2) = 5 \end{equation}
#
# Let's draw a tangent line with that slope on our graph to see if it looks right:
# +
# %matplotlib inline
def f(x):
return x**2 + x
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the point
x1 = 2
y1 = f(x1)
# Specify the derivative we calculated above
m = 5
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the point
plt.scatter(x1,y1, c='red')
plt.annotate('(x,f(x))',(x1,y1), xytext=(x1-0.5, y1+3))
# Plot the tangent line using the derivative we calculated
xMin = x1 - 3
yMin = y1 - (3*m)
xMax = x1 + 3
yMax = y1 + (3*m)
plt.plot([xMin,xMax],[yMin,yMax], color='magenta')
plt.show()
# -
# ### Finding a Derivative for Any Point
# Now let's put it all together and define a function that we can use to find the derivative for any point in the ***f*** function:
#
# Here's our general derivative function again:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{f(x + h) - f(x)}{h} \end{equation}
#
# We know that ***f(x)*** encapsulates the equation ***x<sup>2</sup> + x***, so we can rewrite our derivative equation as:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{((x+h)^{2} + x + h) - (x^{2} + x)}{h} \end{equation}
#
# We can apply the distribution property to ***(x + h)<sup>2</sup>*** using the rule that *(a + b)<sup>2</sup> = a<sup>2</sup> + b<sup>2</sup> + 2ab*:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{(x^{2} + h^{2} + 2xh + x + h) - (x^{2} + x)}{h} \end{equation}
#
# Then we can use the distributive property to expand ***- (x<sup>2</sup> + x)***, which is the same thing as *-1(x<sup>2</sup> + x)*, to ***- x<sup>2</sup> - x***:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{x^{2} + h^{2} + 2xh + x + h - x^{2} - x}{h} \end{equation}
#
# We can combine like terms on the numerator to make things a little clearer:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{h^{2} + 2xh + h}{h} \end{equation}
#
# And finally, we can simplify the fraction:
#
# \begin{equation}f'(x) = \lim_{h \to 0} 2x + h + 1 \end{equation}
#
# To get the limit when *h* is approaching 0, we can use direct substitution for h:
#
# \begin{equation}f'(x) = 2x + 0 + 1 \end{equation}
#
# so:
#
# \begin{equation}f'(x) = 2x + 1 \end{equation}
#
# Now we have a function for the derivative of ***f***, which we can apply to any *x* value to find the slope of the function at ***f(x***).
#
# For example, let's find the derivative of ***f*** with an *x* value of 5:
#
# \begin{equation}f'(5) = 2\cdot5 + 1 = 10 + 1 = 11\end{equation}
#
# Let's use Python to define the ***f(x)*** and ***f'(x)*** functions, plot ***f(5)*** and show the tangent line for ***f'(5)***:
# +
# %matplotlib inline
# Create function f
def f(x):
return x**2 + x
# Create derivative function for f
def fd(x):
return (2 * x) + 1
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(0, 11))
# Use the function to get the y values
y = [f(i) for i in x]
# Set the point
x1 = 5
y1 = f(x1)
# Calculate the derivative using the derivative function
m = fd(x1)
# Set up the graph
plt.xlabel('x')
plt.ylabel('f(x)')
plt.grid()
# Plot the function
plt.plot(x,y, color='green')
# Plot the point
plt.scatter(x1,y1, c='red')
plt.annotate('(x,f(x))',(x1,y1), xytext=(x1-0.5, y1+3))
# Plot the tangent line using the derivative we calculated
xMin = x1 - 3
yMin = y1 - (3*m)
xMax = x1 + 3
yMax = y1 + (3*m)
plt.plot([xMin,xMax],[yMin,yMax], color='magenta')
plt.show()
# -
# ## Differentiability
# It's important to realize that a function may not be *differentiable* at every point; that is, you might not be able to calculate the derivative for every point on the function line.
#
# To be differentiable at a given point:
# - The function must be *continuous* at that point.
# - The tangent line at that point cannot be vertical
# - The line must be *smooth* at that point (that is, it cannot take on a sudden change of direction at the point)
#
# For example, consider the following (somewhat bizarre) function:
#
# \begin{equation}
# q(x) = \begin{cases}
# \frac{40,000}{x^{2}}, & \text{if } x < -4, \\
# (x^{2} -2) \cdot (x - 1), & \text{if } x \ne 0 \text{ and } x \ge -4 \text{ and } x < 8, \\
# (x^{2} -2), & \text{if } x \ne 0 \text{ and } x \ge 8
# \end{cases}
# \end{equation}
# +
# %matplotlib inline
# Define function q
def q(x):
if x != 0:
if x < -4:
return 40000 / (x**2)
elif x < 8:
return (x**2 - 2) * x - 1
else:
return (x**2 - 2)
# Plot output from function g
from matplotlib import pyplot as plt
# Create an array of x values
x = list(range(-10, -5))
x.append(-4.01)
x2 = list(range(-4,8))
x2.append(7.9999)
x2 = x2 + list(range(8,11))
# Get the corresponding y values from the function
y = [q(i) for i in x]
y2 = [q(i) for i in x2]
# Set up the graph
plt.xlabel('x')
plt.ylabel('q(x)')
plt.grid()
# Plot x against q(x)
plt.plot(x,y, color='purple')
plt.plot(x2,y2, color='purple')
plt.scatter(-4,q(-4), c='red')
plt.annotate('A (x= -4)',(-5,q(-3.9)), xytext=(-7, q(-3.9)))
plt.scatter(0,0, c='red')
plt.annotate('B (x= 0)',(0,0), xytext=(-1, 40))
plt.scatter(8,q(8), c='red')
plt.annotate('C (x= 8)',(8,q(8)), xytext=(8, 100))
plt.show()
# -
# The points marked on this graph are non-differentiable:
# - Point **A** is non-continuous - the limit from the negative side is infinity, but the limit from the positive side ≈ -57
# - Point **B** is also non-continuous - the function is not defined at x = 0.
# - Point **C** is defined and continuous, but the sharp change in direction makes it non-differentiable.
# ## Derivatives of Equations
#
# We've been talking about derivatves of *functions*, but it's important to remember that functions are just named operations that return a value. We can apply what we know about calculating derivatives to any equation, for example:
#
# \begin{equation}\frac{d}{dx}(2x + 6)\end{equation}
#
# Note that we generally switch to *Leibniz's* notation when finding derivatives of equations that are not encapsulated as functions; but the approach for solving this example is exactly the same as if we had a hypothetical function with the definition *2x + 6*:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = \lim_{h \to 0} \frac{(2(x+h) + 6) - (2x + 6)}{h} \end{equation}
#
# After factoring out the* 2(x+h)* on the left and the *-(2x - 6)* on the right, this is:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = \lim_{h \to 0} \frac{2x + 2h + 6 - 2x - 6}{h} \end{equation}
#
# We can simplify this to:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = \lim_{h \to 0} \frac{2h}{h} \end{equation}
#
# Now we can factor *h* out entirely, so at any point:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = 2 \end{equation}
#
# If you run the Python code below to plot the line created by the equation, you'll see that it does indeed have a constant slope of 2:
# +
# %matplotlib inline
from matplotlib import pyplot as plt
# Create an array of x values from 0 to 10 to plot
x = list(range(1, 11))
# Use the function to get the y values
y = [(2*i) + 6 for i in x]
# Set up the graph
plt.xlabel('x')
plt.xticks(range(1,11, 1))
plt.ylabel('y')
plt.yticks(range(8,27, 1))
plt.grid()
# Plot the function
plt.plot(x,y, color='purple')
plt.show()
# -
# ## Derivative Rules and Operations
# When working with derivatives, there are some rules, or shortcuts, that you can apply to make your life easier.
#
# ### Basic Derivative Rules
# Let's start with some basic rules that it's useful to know.
#
# - If *f(x)* = *C* (where *C* is a constant), then *f'(x)* = 0
#
# This makes sense if you think about it for a second. No matter what value you use for *x*, the function returns the same constant value; so the graph of the function will be a horizontal line. There's no rate of change in a horiziontal line, so its slope is 0 at all points. This is true of any constant, including symbolic constants like *π* (pi).
#
# So, for example:
#
# \begin{equation}f(x) = 6 \;\; \therefore \;\; f'(x) = 0 \end{equation}
#
# Or:
#
# \begin{equation}f(x) = \pi \;\; \therefore \;\; f'(x) = 0 \end{equation}
#
# - If *f(x)* = *Cg(x)*, then *f'(x)* = *Cg'(x)*
#
# This rule tells us that if a function is equal to a second function multiplied by a constant, then the derivative of the first function will be equal to the derivative of the second function multiplied by the same constant. For example:
#
# \begin{equation}f(x) = 2g(x) \;\; \therefore \;\; f'(x) = 2g'(x) \end{equation}
#
# - If *f(x)* = *g(x)* + *h(x)*, then *f'(x)* = *g'(x)* + *h'(x)*
#
# In other words, if a function is the sum of two other functions, then the derivative of the first function is the sum of the derivatives of the other two functions. For example:
#
# \begin{equation}f(x) = g(x) + h(x) \;\; \therefore \;\; f'(x) = g'(x) + h'(x) \end{equation}
#
# Of course, this also applies to subtraction:
#
# \begin{equation}f(x) = k(x) - l(x) \;\; \therefore \;\; f'(x) = k'(x) - l'(x) \end{equation}
#
# As discussed previously, functions are just equations encapsulated as a named entity that return a value; and the rules can be applied to any equation. For example:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = \frac{d}{dx} 2x + \frac{d}{dx} 6\end{equation}
#
# So we can take advantage of the rules to make the calculation a little easier:
#
# \begin{equation}\frac{d}{dx}(2x) = \lim_{h \to 0} \frac{2(x+h) - 2x}{h} \end{equation}
#
# After factoring out the* 2(x+h)* on the left, this is:
#
# \begin{equation}\frac{d}{dx}(2x) = \lim_{h \to 0} \frac{2x + 2h - 2x}{h} \end{equation}
#
# We can simplify this to:
#
# \begin{equation}\frac{d}{dx}(2x) = \lim_{h \to 0} \frac{2h}{h} \end{equation}
#
# Which gives us:
#
# \begin{equation}\frac{d}{dx}(2x) = 2 \end{equation}
#
# Now we can turn our attention to the derivative of the constant 6 with respect to *x*, and we know that the derivative of a constant is always 0, so:
#
# \begin{equation}\frac{d}{dx}(6) = 0\end{equation}
#
# We add the two derivatives we calculated:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = 2 + 0\end{equation}
#
# Which gives us our result:
#
# \begin{equation}\frac{d}{dx}(2x + 6) = 2\end{equation}
#
#
# ### The Power Rule
# The *power rule* is one of the most useful shortcuts in the world of differential calculus. It can be stated like this:
#
# \begin{equation}f(x) = x^{n} \;\; \therefore \;\; f'(x) = nx^{n-1}\end{equation}
#
# So if our function for *x* returns *x* to the power of some constant (which we'll call *n*), then the derivative of the function for *x* is *n* times *x* to the power of *n* - 1.
#
# It's probably helpful to look at a few examples to see how this works:
#
# \begin{equation}f(x) = x^{3} \;\; \therefore \;\; f'(x) = 3x^{2}\end{equation}
#
# \begin{equation}f(x) = x^{-2} \;\; \therefore \;\; f'(x) = -2x^{-3}\end{equation}
#
# \begin{equation}f(x) = x^{2} \;\; \therefore \;\; f'(x) = 2x\end{equation}
#
# In each of these examples, the exponential of *x* in the function definition becomes the coefficient for *x* in the derivative definition, with the exponential is decremented by 1.
#
# Here's a worked example to find the derivative of the following function:
#
# \begin{equation}f(x) = x^{2}\end{equation}
#
# So we start with the general derivative function:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{f(x + h) - f(x)}{h} \end{equation}
#
# We can plug in our definition for *f*:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{(x + h)^{2} - x^{2}}{h} \end{equation}
#
# Now we can factor out the perfect square binomial on the left:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{x^{2} + h^{2} + 2xh - x^{2}}{h} \end{equation}
#
# The x<sup>2</sup> terms cancel each other out so we get to:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{h^{2} + 2xh}{h} \end{equation}
#
# Which simplifies to:
#
# \begin{equation}f'(x) = \lim_{h \to 0} h + 2x \end{equation}
#
# With *h* approaching 0, this is:
#
# \begin{equation}f'(x) = 0 + 2x \end{equation}
#
# So our answer is:
#
# \begin{equation}f'(x) = 2x \end{equation}
#
# Note that we could have achieved the same result by simply applying the power rule and transforming x<sup>2</sup> to 2x<sup>1</sup> (which is the same as 2x).
#
# ### The Product Rule
# The product rule can be stated as:
#
# \begin{equation}\frac{d}{dx}[f(x)g(x)] = f'(x)g(x) + f(x)g'(x) \end{equation}
#
# OK, let's break that down. What it's saying is that the derivative of *f(x)* multiplied by *g(x)* is equal to the derivative of *f(x)* multiplied by the value of *g(x)* added to the value of *f(x)* multiplied by the derivative of *g(x*).
#
# Let's see an example based on the following two functions:
#
# \begin{equation}f(x) = 2x^{2} \end{equation}
#
# \begin{equation}g(x) = x + 1 \end{equation}
#
# Let's start by calculating the derivative of *f(x)*:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{2(x + h)^{2} - 2x^{2}}{h} \end{equation}
#
# This factors out to:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{2x^{2} + 2h^{2} + 4xh - 2x^{2}}{h} \end{equation}
#
# Which when we cancel out the 2x<sup>2</sup> and -2x<sup>2</sup> is:
#
# \begin{equation}f'(x) = \lim_{h \to 0} \frac{2h^{2} + 4xh}{h} \end{equation}
#
# Which simplifies to:
#
# \begin{equation}f'(x) = \lim_{h \to 0} 2h + 4x \end{equation}
#
# With *h* approaching 0, this is:
#
# \begin{equation}f'(x) = 4x \end{equation}
#
# Now let's look at *g'(x)*:
#
# \begin{equation}g'(x) = \lim_{h \to 0} \frac{(x + h) + 1 - (x + 1)}{h} \end{equation}
#
# We can just remove the brackets on the left and factor out the *-(x + 1)* on the right:
#
# \begin{equation}g'(x) = \lim_{h \to 0} \frac{x + h + 1 - x - 1}{h} \end{equation}
#
# Which can be cleaned up to:
#
# \begin{equation}g'(x) = \lim_{h \to 0} \frac{h}{h} \end{equation}
#
# Enabling us to factor *h* out completely to give a constant derivative of *1*:
#
# \begin{equation}g'(x) = 1 \end{equation}
#
# So now we can calculate the derivative for the product of these functions by plugging the functions and the derivatives we've calculated for them into the product rule equation:
#
# \begin{equation}\frac{d}{dx}[f(x)g(x)] = f'(x)g(x) + f(x)g'(x) \end{equation}
#
# So:
#
# \begin{equation}\frac{d}{dx}[f(x)g(x)] = (4x \cdot (x + 1)) + (2x^{2} \cdot 1) \end{equation}
#
# Which can be simplified to:
#
# \begin{equation}\frac{d}{dx}[f(x)g(x)] = (4x^{2} + 4x) + 2x^{2} \end{equation}
#
# Which can be further simplified to:
#
# \begin{equation}\frac{d}{dx}[f(x)g(x)] = 6x^{2} + 4x \end{equation}
# ### The Quotient Rule
# The *quotient rule* applies to functions that are defined as a quotient of one expression divided by another; for example:
#
# \begin{equation}r(x) = \frac{s(x)}{t(x)} \end{equation}
#
# In this situation, you can apply the following quotient rule to find the derivative of *r(x)*:
#
# \begin{equation}r'(x) = \frac{s'(x)t(x) - s(x)t'(x)}{[t(x)]^{2}} \end{equation}
#
# Here are our definitions for *s(x)* and *t(x)*:
#
# \begin{equation}s(x) = 3x^{2} \end{equation}
#
# \begin{equation}t(x) = 2x\end{equation}
#
# Let's start with *s'(x)*:
#
# \begin{equation}s'(x) = \lim_{h \to 0} \frac{3(x + h)^{2} - 3x^{2}}{h} \end{equation}
#
# This factors out to:
#
# \begin{equation}s'(x) = \lim_{h \to 0} \frac{3x^{2} + 3h^{2} + 6xh - 3x^{2}}{h} \end{equation}
#
# Which when we cancel out the 3x<sup>2</sup> and -3x<sup>2</sup> is:
#
# \begin{equation}s'(x) = \lim_{h \to 0} \frac{3h^{2} + 6xh}{h} \end{equation}
#
# Which simplifies to:
#
# \begin{equation}s'(x) = \lim_{h \to 0} 3h + 6x \end{equation}
#
# With *h* approaching 0, this is:
#
# \begin{equation}s'(x) = 6x \end{equation}
#
# Now let's look at *t'(x)*:
#
# \begin{equation}t'(x) = \lim_{h \to 0} \frac{2(x + h) - 2x}{h} \end{equation}
#
# We can just factor out the *2(x + h)* on the left:
#
# \begin{equation}t'(x) = \lim_{h \to 0} \frac{2x + 2h - 2x}{h} \end{equation}
#
# Which can be cleaned up to:
#
# \begin{equation}t'(x) = \lim_{h \to 0} \frac{2h}{h} \end{equation}
#
# Enabling us to factor *h* out completely to give a constant derivative of *2*:
#
# \begin{equation}t'(x) = 2 \end{equation}
#
# So now we can calculate the derivative for the quotient of these functions by plugging the function definitions and the derivatives we've calculated for them into the quotient rule equation:
#
# \begin{equation}r'(x) = \frac{(6x \cdot 2x) - (3x^{2} \cdot 2)}{[2x]^{2}} \end{equation}
#
# We can factor out the numerator terms like this:
#
# \begin{equation}r'(x) = \frac{12x^{2} - 6x^{2}}{[2x]^{2}} \end{equation}
#
# Which can then be combined:
#
# \begin{equation}r'(x) = \frac{6x^{2}}{[2x]^{2}} \end{equation}
#
# The denominator is [2x]<sup>2</sup> (note that this is different from 2x<sup>2</sup>. [2x]<sup>2</sup> is 2x • 2x, whereas 2x<sup>2</sup> is 2 • x<sup>2</sup>):
#
# \begin{equation}r'(x) = \frac{6x^{2}}{4x^{2}} \end{equation}
#
# Which simplifies to:
#
# \begin{equation}r'(x) = 1\frac{1}{2} \end{equation}
#
# So the derivative of *r(x)* is 1.5.
# ### The Chain Rule
#
# The *chain rule* takes advantage of the fact that equations can be encapsulated as functions, and since functions contain equations, it's possible to nest one function within another.
#
# For example, consider the following function:
#
# \begin{equation}u(x) = 2x^{2} \end{equation}
#
# We could view the definition of *u(x)* as a composite of two functions,; an *inner* function that calculates x<sup>2</sup>, and an *outer* function that multiplies the result of the inner function by 2.
#
# \begin{equation}u(x) = \widehat{\color{blue}2\color{blue}(\underline{\color{red}x^{\color{red}2}}\color{blue})} \end{equation}
#
# To make things simpler, we can name these inner and outer functions:
#
# \begin{equation}i(x) = x^{2} \end{equation}
#
# \begin{equation}o(x) = 2x \end{equation}
#
# Note that *x* indicates the input for each function. Function *i* takes its input and squares it, and function *o* takes its input and multiplies it by 2. When we use these as a composite function, the *x* value input into the outer function will be the output from the inner function.
#
# Let's take a look at how we can apply these functions to get back to our original *u* function:
#
# \begin{equation}u(x) = o(i(x)) \end{equation}
#
# So first we need to find the output of the inner *i* function so we can use at as the input value for the outer *o* function. Well, that's easy, we know the definition of *i* (square the input), so we can just plug it in:
#
# \begin{equation}u(x) = o(x^{2}) \end{equation}
#
# We also know the definition for the outer *o* function (multiply the input by 2), so we can just apply that to the input:
#
# \begin{equation}u(x) = 2x^{2} \end{equation}
#
# OK, so now we know how to form a composite function. The *chain rule* can be stated like this:
#
# \begin{equation}\frac{d}{dx}[o(i(x))] = o'(i(x)) \cdot i'(x)\end{equation}
#
# Alright, let's start by plugging the output of the inner *i(x)* function in:
#
# \begin{equation}\frac{d}{dx}[o(i(x))] = o'(x^{2}) \cdot i'(x)\end{equation}
#
# Now let's use that to calculate the derivative of *o*, replacing each *x* in the equation with the output from the *i* function (*x<sup>2</sup>*):
#
# \begin{equation}o'(x) = \lim_{h \to 0} \frac{2(x^{2} + h) - 2x^{2}}{h} \end{equation}
#
# This factors out to:
#
# \begin{equation}o'(x) = \lim_{h \to 0} \frac{2x^{2} + 2h - 2x^{2}}{h} \end{equation}
#
# Which when we cancel out the 2x<sup>2</sup> and -2x<sup>2</sup> is:
#
# \begin{equation}o'(x) = \lim_{h \to 0} \frac{2h}{h} \end{equation}
#
# Which simplifies to:
#
# \begin{equation}o'(x) = 2 \end{equation}
#
# Now we can calculate *i'(x)*. We know that the definition of *i(x)* is x<sup>2</sup>, and we can use the power rule to determine that *i'(x)* is therefore 2x.
#
# So our equation at this point is:
#
# \begin{equation}\frac{d}{dx}[o(i(x))] = 2 \cdot 2x\end{equation}
#
# Which is:
#
# \begin{equation}\frac{d}{dx}[o(i(x))] = 4x\end{equation}
#
# Commonly, the chain rule is stated using a slighly different notation that you may find easier to work with. In this case, we can take our equation:
#
# \begin{equation}\frac{d}{dx}[o(i(x))] = o'(i(x)) \cdot i'(x)\end{equation}
#
# and rewrite it as
#
# \begin{equation}\frac{du}{dx} = \frac{do}{di}\frac{di}{dx}\end{equation}
#
# We can then complete the calculations like this:
#
# \begin{equation}\frac{du}{dx} = 2 \cdot 2x = 4x\end{equation}
| MathsToML/Module02-Derivatives and Optimization/02-03-Differentiation and Derivatives.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import time
import pyperclip
import urllib.request
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import os
from pprint import pprint
from tqdm import tqdm, notebook
# +
path = r'C:\Users\재욱\Downloads\chromedriver.exe'
driver = webdriver.Chrome(path)
driver.implicitly_wait(3)
country_url = "https://www.brandsoftheworld.com/logos/countries"
driver.get(country_url)
xpath = '//*[@id="primaryInner"]/div/ul'
division = driver.find_element_by_xpath(xpath)
country_list = division.find_elements_by_tag_name('a') # List형
country_url_list = []
for country_url in country_list:
country_url_list.append(country_url.get_attribute('href'))
# -
# ## Old Code
XPATH_COUNTRY_TABLE = '//*[@id="primaryInner"]/div/div[1]'
XPATH_COUNTRY_NAME = '//*[@id="primary"]/h1/span'
for country_path in country_url_list:
page_num = 0
while True:
driver.get(country_path + '?page=' + str(page_num))
division = driver.find_element_by_xpath(XPATH_COUNTRY_TABLE) # 빼면 안됨
name_list = division.find_elements_by_tag_name('li') # List형
image_url_list = division.find_elements_by_tag_name('img') # List형
img_folder_path = './brands_of_the_world/' + division.text + '_company_logo/'
company_name_list = []
for name in name_list:
company_name_list.append(name.text)
# if company_name_list == ['« first', '‹ previous', '1', '2']:
if len(image_url_list) == 0:
break
page_num +=1
i = 0
for image_url in image_url_list:
try:
with urllib.request.urlopen(image_url.get_attribute('src')) as f:
with open(img_folder_path + company_name_list[i] +'.jpg','wb') as h:
image = f.read()
h.write(image)
except Exception as e:
print(e)
i+=1
# ## New Code (joowon)
# +
try:
XPATH_COUNTRY_TABLE = '//*[@id="primaryInner"]/div/div[1]'
XPATH_COUNTRY_NAME = '//*[@id="primary"]/h1/span'
COUNTRY_NUMBER = 0
for country_path in notebook.tqdm(country_url_list, desc='iterate range 100'):
driver.get(f'{country_path}?page=0')
driver.implicitly_wait(2)
country_name = driver.find_element_by_xpath(XPATH_COUNTRY_NAME)
img_folder_path = './brands_of_the_world/' + country_name.text + '_company_logo/'
COUNTRY_NUMBER +=1
print('%d 번째 나라 : [%s]' %(COUNTRY_NUMBER,country_name.text))
if os.path.isdir(img_folder_path) is True:
continue
os.makedirs(img_folder_path, exist_ok=True)
page_num = 0
while True:
# driver.get(country_path + '?page=' + str(page_num))
driver.get(f'{country_path}?page={page_num}') #f string
division = driver.find_element_by_xpath(XPATH_COUNTRY_TABLE) # 빼면 안됨
name_list = division.find_elements_by_tag_name('li') # List형
image_url_list = division.find_elements_by_tag_name('img') # List형
if len(image_url_list) == 0: # ['« first', '‹ previous', '1', '2']
break
for i, image_url in enumerate(image_url_list):
try:
with urllib.request.urlopen(image_url.get_attribute('src')) as f:
with open(img_folder_path + name_list[i].text +'.jpg','wb') as h:
image = f.read()
h.write(image)
except Exception as e:
print(e)
print('Images of page number[%d] had saved completely!' %(page_num))
page_num +=1
except Exception as e:
print(e)
except KeyboardInterrupt:
print("KeyboardInterrupt")
except:
print('Other error')
# -
| status_not_saved.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import minimal requirements
import requests
import json
import re
# +
# Set the base URL for the ARAX reasoner and its endpoint
endpoint_url = 'https://arax.rtx.ai/api/rtx/v1/query'
# Create a dict of the request, specifying a start previous Message and the list of DSL commands
query = {"previous_message_processing_plan": {"processing_actions": [
"add_qnode(name=acetaminophen, id=n0)",
"add_qnode(type=protein, id=n1)",
"add_qedge(source_id=n0, target_id=n1, id=e0)",
"expand(edge_id=e0)",
"resultify(ignore_edge_direction=true)",
"filter_results(action=limit_number_of_results, max_results=10)",
"return(message=true, store=true)",
]}}
# -
# Send the request to RTX and check the status
print(f"Executing query at {endpoint_url}\nPlease wait...")
response_content = requests.post(endpoint_url, json=query, headers={'accept': 'application/json'})
status_code = response_content.status_code
if status_code != 200:
print("ERROR returned with status "+str(status_code))
print(response_content.json())
else:
print(f"Response returned with status {status_code}")
# Unpack respsonse from JSON and display the information log
response_dict = response_content.json()
for message in response_dict['log']:
if message['level'] >= 20:
print(message['prefix']+message['message'])
# These URLs provide direct access to resulting data and GUI
if 'id' in response_dict and response_dict['id'] is not None:
print(f"Data: {response_dict['id']}")
match = re.search(r'(\d+)$', response_dict['id'])
if match:
print(f"GUI: https://arax.rtx.ai/?m={match.group(1)}")
else:
print("No id was returned in response")
# Or you can view the entire Translator API response Message
print(json.dumps(response_dict, indent=2, sort_keys=True))
| code/ARAX/Examples/ARAX_Example1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np
import pandas as pd
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import random
sns.set()
# -
train = pd.read_csv('fashionmnist/fashion-mnist_train.csv')
test = pd.read_csv('fashionmnist/fashion-mnist_test.csv')
train.head()
test.head()
train.shape
test.shape
training_array = np.array(train,dtype='float32')
testing_array = np.array(test,dtype = 'float32')
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
i = random.randint(1,60000)
plt.figure()
plt.imshow(training_array[i,1:].reshape(28,28))
plt.grid(False)
plt.show()
label = int(training_array[i,0])
print(f'The image is for : {class_names[label]}')
# +
W_grid = 15
L_grid = 15
# subplot return the figure and axes object
# And by using axes object we can plot specific figure at various location
fig , axes = plt.subplots(L_grid,W_grid,figsize=(17,17))
axes = axes.ravel() #Flaten the 15 * 15 matrix into 255 array
n_training = len(training_array) #get the length of training dataset
for i in np.arange(0,L_grid*W_grid):
index = np.random.randint(0,n_training)
axes[i].imshow(training_array[index,1:].reshape(28,28))
axes[i].set_title(class_names[int(training_array[index,0])],fontsize=8)
axes[i].axis('off')
plt.subplots_adjust(hspace=0.4)
# -
# * We scale these values to a range of 0 to 1 before feeding to the neural network model. For this, we divide the values by 255. It's important that the *training set* and the *testing set* are preprocessed in the same way:
X_train = training_array[:,1:]/255
y_train = training_array[:,0]
X_test = testing_array[:,1:]/255
y_test = testing_array[:,0]
from sklearn.model_selection import train_test_split
X_train ,X_validate , y_train,y_validate = train_test_split(X_train,y_train,test_size = 0.2,random_state = 12345)
X_train = X_train.reshape(X_train.shape[0],28,28,1)
X_test = X_test.reshape(X_test.shape[0],28,28,1)
X_validate = X_validate.reshape(X_validate.shape[0],28,28,1)
print(f'shape of X train : {X_train.shape}')
print(f'shape of X test : {X_test.shape}')
print(f'shape of X validate : {X_validate.shape}')
print(f'shape of y train : {y_train.shape}')
print(f'shape of y test : {y_test.shape}')
print(f'shape of y validate : {y_validate.shape}')
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import MaxPool2D,Flatten,Dense
from tensorflow.keras.callbacks import EarlyStopping,TensorBoard
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.utils import to_categorical
y_cat_train = to_categorical(y_train,num_classes=10)
y_cat_test = to_categorical(y_test,num_classes=10)
y_cat_validate = to_categorical(y_validate,num_classes=10)
print(f'shape of y train : {y_cat_train.shape}')
print(f'shape of y test : {y_cat_test.shape}')
print(f'shape of y validate : {y_cat_validate.shape}')
# # **Start Building CNN Models**
# +
model = Sequential()
# CONVOLUTIONAL LAYER
model.add(Conv2D(filters=32, kernel_size=(4,4),input_shape=(28, 28, 1), activation='relu',))
# POOLING LAYER
model.add(MaxPool2D(pool_size=(2, 2)))
# FLATTEN IMAGES FROM 28 by 28 to 764 BEFORE FINAL LAYER
model.add(Flatten())
# 128 NEURONS IN DENSE HIDDEN LAYER (YOU CAN CHANGE THIS NUMBER OF NEURONS)
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
# LAST LAYER IS THE CLASSIFIER, THUS 10 POSSIBLE CLASSES
model.add(Dense(10, activation='softmax'))
# https://keras.io/metrics/
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']) # we can add in additional metrics https://keras.io/metrics/
# -
model.summary()
early_stopping = EarlyStopping(monitor='val_loss',patience=1)
model.fit(X_train,
y_cat_train,
epochs=50,
verbose=1,
validation_data=(X_validate,y_cat_validate),
callbacks=[early_stopping])
model_history = pd.DataFrame(model.history.history)
model_history
model_history[['accuracy','val_accuracy']].plot()
model_history[['loss','val_loss']].plot()
evalution = model.evaluate(X_test,y_cat_test)
print(f'Test Accuracy : {evalution[1]}')
predict_class = model.predict_classes(X_test)
predict_class.shape
i = random.randint(0,predict_class.shape[0])
print(class_names[predict_class[i]])
print(class_names[int(y_test[i])])
# +
w_gird = 5
l_gird = 5
fig,axes = plt.subplots(l_gird,w_gird,figsize=(12,12))
axes = axes.ravel()
for i in np.arange(0,l_gird*w_gird):
axes[i].imshow(X_test[i].reshape(28,28))
axes[i].set_title(f'{i}.Predict Class : {class_names[predict_class[i]]} \n True Class : {class_names[int(y_test[i])]}')
axes[i].axis('off')
plt.subplots_adjust(wspace=0.9,hspace=0.7)
# -
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test,predict_class)
fig = plt.figure(figsize=(12,12))
sns.heatmap(cm,annot=True,cmap='viridis',fmt='d')
# +
from sklearn.metrics import classification_report
num_classes = 10
target_names = ['Class {}'.format(i) for i in range(num_classes)]
print(classification_report(y_test,predict_class,target_names=target_names))
# -
| FASHION_MNIST/Fashion MNIST Classification with keras2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # load data
# +
# import load_iris function from datasets module
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
import random
import numpy as np
import pandas as pd
from sklearn.ensemble import ExtraTreesClassifier
# +
data = pd.read_csv("../../../accuracy_drop_proj/datasets/EEG_data_Epileptic_Seizure_Recognition.csv")
data = data.drop(['Unnamed: 0'], axis=1)
data.shape
# +
y = data.y.values
del data["y"] # remove rings from data, so we can convert all the dataframe to a numpy 2D array.
X = data.values.astype(np.float)
# -
y.shape
# +
#y = data['y']
# data = data.drop(['y'],axis=1)
# data.head()
# -
x_train, x_test, y_train, y_test = train_test_split(data, y, test_size=0.30,random_state=4)
print(x_test.shape)
print(x_train.shape)
# +
#find the order of the feature according to information gain
model = ExtraTreesClassifier()
model.fit(data, y)
information_gain = {}
for i in range(len(model.feature_importances_)):
information_gain.update({i: model.feature_importances_[i]})
# -
information_gain
col_sorted=sorted(information_gain.items(), key=lambda x:x[1],reverse=True)
select_col=[]
for i in col_sorted:
select_col.append(i[0])
print(select_col)
# +
from mlxtend.feature_selection import SequentialFeatureSelector as SFS
sfs1 = SFS(model,
k_features=4,
forward=True,
floating=False,
verbose=2,
scoring='accuracy',
cv=0)
sfs1 = sfs1.fit(data, y)
# -
sfs1.k_feature_idx_
temp = data.iloc[:,[9, 52, 168, 33, 157, 107, 159, 1, 54, 166,178]]
temp.head()
# +
y = temp.y.values
del temp["y"] # remove rings from data, so we can convert all the dataframe to a numpy 2D array.
X = temp.values.astype(np.float)
# -
y = temp['y']
temp = temp.drop(['y'],axis=1)
temp.shape
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.25,random_state=4)
# +
from sklearn import svm
model_svm= svm.SVC(probability=True)
model_svm.fit(x_train, y_train)
y_predsvm = model_svm.predict(x_test)
print('Accuracy of svm classifier on train set: {:.2f}'.format(accuracy_score(y_test,y_predsvm)))
# +
from sklearn import tree
model1=tree.DecisionTreeClassifier()
print("train tree")
model1.fit(x_train,y_train)
y_pred1 = model1.predict(x_test)
print('Accuracy of tree classifier on train set: {:.2f}'.format(accuracy_score(y_test,y_pred1)))
# -
model_svm.predict([[-14,-61,-29,-1,44,18,-45,-14,1,16]])
# +
import heapq
import numpy as np
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.feature_selection import chi2
from sklearn.ensemble import ExtraTreesClassifier
from accuracy_drop_proj.strategies.change_combination.change_combination import Change_Combination
"""
this method according the user request find the difference of the probability for each class and then sort the rows according to them.
for example if we have [0.1 0.6 0.3] and request of user be [0,1] we compute 0.5 for this row and sort the row Ascending
"""
class Change_ProbabilityDistance_RankFeature(object):
def __init__(self):
pass
def change(self,x_train, y_train, percetage, mnb, change_plan):
number_change_requested = int(percetage / 100 * x_train.shape[0])
print("{} percentage error is equal to {} change \n".format(percetage, number_change_requested))
used_row ={}
occurred_change = 0
all_changed = 1
change_done = False
x_train_changed = np.copy(x_train)
#---------------------find the order of the feature according to information gain-----------------------
model = ExtraTreesClassifier()
model.fit(x_train, y_train)
print("combination of feature")
information_gain = {}
for i in range(len(model.feature_importances_)):
information_gain.update({i: model.feature_importances_[i]})
ranked_information_dic = {}
sum_gain = 0
for L in range(0,x_train.shape[1] + 1):
for subset in Change_Combination.combinations_index(self,information_gain.keys(), L):
if not subset:
pass
else:
for key in subset:
sum_gain = sum_gain + information_gain.get(key)
ranked_information_dic.update({tuple(subset): sum_gain})
sum_gain = 0
print("create all subset")
all_subset = sorted(ranked_information_dic.items(), key=lambda item: len(item[0]) * 1000 - item[1], reverse=False)
probability = mnb.predict_proba(x_train)
probability_distance={}
#----------------------------------------------changing--------------------------------------------------
for i in range(len(change_plan["key"])):
occurred_change = 0
indices = [t for t, x in enumerate(y_train) if x == change_plan["key"][i][0]]
# print(indices)
print("{} rows have target {} \n".format(len(indices), change_plan["key"][i][0]))
probability_distance.clear()
probability_distance_sorted=[]
# find the distance probability between the class that user need to change
for elements in indices:
probability_distance.update({elements:np.abs(probability[elements][change_plan["key"][i][0]]- probability[elements][change_plan["key"][i][1]])})
# ---------------------------finding the order of the row according to probability distance-------------------------
# sort the row according the distance probability
probability_distance_sorted = sorted(probability_distance.items(), key=lambda x: x[1], reverse=False)
indices=[]
for j in probability_distance_sorted:
indices.append(j[0])
print(indices)
print("try in indices")
for p in range(len(indices)):
if (all_changed == number_change_requested + 1):
print("your requests have been done :)")
break
# print(mnb.predict([x_train[indices[p]]]))
if y_train[indices[p]] == mnb.predict([x_train[indices[p]]]) and indices[p] not in used_row:
change_done = False
for subset in all_subset:
if change_done:
break
else:
if (occurred_change == change_plan["number"][i]):
# print("part of your request has been done :))))")
break
print("try to change, with changing index {} on row {}".format(list(subset[0]),indices[p]))
#######################################################
# impose Outlier insted of 0
########################################################
x_train_changed[indices[p]][list(subset[0])] = 0
if (change_plan["key"][i][1] == mnb.predict([x_train_changed[indices[p]]])[0]):
print(x_train[indices[p]], mnb.predict([x_train[indices[p]]])[0])
print(x_train_changed[indices[p]],
mnb.predict([x_train_changed[indices[p]]])[0])
print(" \n change number {} \n".format(all_changed))
used_row.update({indices[p]: indices[p]})
occurred_change = occurred_change + 1
change_done = True
all_changed = all_changed + 1
break
else:
x_train_changed[indices[p]] = np.copy(x_train[indices[p]])
if (all_changed <= number_change_requested):
print("your request doesn't complete! please change your plan")
return np.copy(x_train_changed)
# -
change_plan={"key":[[4,2]],"number":[1]}
confus
classmodel = Change_ProbabilityDistance_RankFeature()
classmodel.change(x_test, y_test,10,model_svm, change_plan)
# +
import heapq
import numpy as np
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.feature_selection import chi2
from sklearn.ensemble import ExtraTreesClassifier
from accuracy_drop_proj.strategies.change_combination.change_combination import Change_Combination
"""
this method first sort the rows according to uncetainity and features according to information gain and then pick
first rows that classifer is not sure about them and feature are more likely to change
"""
class Change_Uncertainty_Rankfeatures(object):
def __init__(self):
pass
def change(self,x_train, y_train, percetage, mnb, change_plan):
number_change_requested = int(percetage / 100 * x_train.shape[0])
print("{} percentage error is equal to {} change \n".format(percetage, number_change_requested))
used_row ={}
occurred_change = 0
all_changed = 1
change_done = False
x_train_changed = np.copy(x_train)
#---------------------find the order of the feature according to information gain-----------------------
model = ExtraTreesClassifier()
model.fit(x_train, y_train)
print("combinatio of feature")
information_gain = {}
for i in range(len(model.feature_importances_)):
information_gain.update({i: model.feature_importances_[i]})
ranked_information_dic = {}
sum_gain = 0
for L in range(0,x_train.shape[1] + 1):
for subset in Change_Combination.combinations_index(self,information_gain.keys(), L):
if not subset:
pass
else:
for key in subset:
sum_gain = sum_gain + information_gain.get(key)
ranked_information_dic.update({tuple(subset): sum_gain})
sum_gain = 0
print("create all subset")
all_subset = sorted(ranked_information_dic.items(), key=lambda item: len(item[0]) * 1000 - item[1], reverse=False)
#---------------------------finding the order of the row according to uncertainity-------------------------
probability = mnb.predict_proba(x_train)
print("finding uncertainity")
uncertainty={}
for index,roww in enumerate(probability):
largest_val =heapq.nlargest(2, roww)
uncertainty.update({index:1-(np.abs(np.subtract(largest_val[0],largest_val[1])))})
largest_val=[]
# print(index,row,np.subtract(largest_val[0],largest_val[1]))
print(len(probability))
print(len(uncertainty))
#sort the uncertainty
uncertainty_sorted=sorted(uncertainty.items(), key=lambda x:x[1],reverse=True)
print("changing")
#---------------------------------------------changing--------------------------------------------
for i in range(len(change_plan["key"])):
occurred_change = 0
#sort the row according to uncertainty
indices=[]
for key_dic in uncertainty_sorted:
if y_train[key_dic[0]] == change_plan["key"][i][0]:
indices.append(key_dic[0])
print(indices)
#this is normal indices
# indices_2 = [t for t, x in enumerate(y_train) if x == change_plan["key"][i][0]]
print("{} rows have target {} \n".format(len(indices), change_plan["key"][i][0]))
for p in range(len(indices)):
print("try in indices")
if (all_changed == number_change_requested + 1):
print("your requests have been done :)")
break
if y_train[indices[p]] == mnb.predict([x_train[indices[p]]]) and indices[p] not in used_row:
change_done = False
for subset in all_subset:
if change_done:
break
else:
if (occurred_change == change_plan["number"][i]):
# print("part of your request has been done :))))")
break
print("try to change, with change index {} on row {}".format(list(subset[0]),indices[p]))
x_train_changed[indices[p]][list(subset[0])] = 0
if (change_plan["key"][i][1] == mnb.predict([x_train_changed[indices[p]]])[0]):
print(x_train[indices[p]], mnb.predict([x_train[indices[p]]])[0])
print(x_train_changed[indices[p]],
mnb.predict([x_train_changed[indices[p]]])[0])
print(" \n change number {} \n".format(all_changed))
used_row.update({indices[p]: indices[p]})
occurred_change = occurred_change + 1
change_done = True
all_changed = all_changed + 1
break
else:
x_train_changed[indices[p]] = np.copy(x_train[indices[p]])
if (all_changed <= number_change_requested):
print("your request doesn't complete! please change your plan")
return np.copy(x_train_changed)
# -
model2= Change_Uncertainty_Rankfeatures()
model2.change(x_test, y_test,10,model1, change_plan)
# # plotting
# - increase number of the rows
alg1=[0.356,0.424,1.069,1.121,2.764,2.861,4.963,3.568,4.540,4.029,3.993]
alg2=[0.115,0.242,0.848,0.763,2.166,2.387,2.986,2.668,3.887,2.758,2.970]
alg3=[0.224,0.211,1.099,0.637,1.747,3.280,2.866,3.429,3.541,2.662,2.983]
alg4=[0.208,0.202,0.980,0.701,2.444,2.343,3.070,2.864,3.794,2.773,4.555]
# +
from matplotlib.pyplot import figure
figure(num=None, figsize=(10, 8), dpi=80, facecolor='w', edgecolor='k')
plt.grid(True)
x=[500,1000,2000,3000,4000,5000,6000,7000,8000,9000,10000]
plt.yticks(np.arange(0, 5, 0.1))
plt.xticks(x)
plt.plot(x,alg1,c='red',linewidth=3,label= 'Algorithm 1')
plt.plot(x,alg2,c= 'blue',linewidth=3,label= 'Algorithm 2')
plt.plot(x,alg3,c='green',linewidth=3,label= 'Algorithm 3')
plt.plot(x,alg4,c= 'black',linewidth=3,label= 'Algorithm 4')
plt.xlabel('Number of the Rows')
plt.ylabel('Time (sec)')
plt.legend()
# -
# # plotting
# - increase number of column(Features)
alg1_col =[0.356,17.164,7200,7200,7200,7200,7200,7200,7200,7200]
alg2_col =[0.115,25.774,7200,7200,7200,7200,7200,7200,7200,7200]
alg3_col =[0.224,6.793,7200,7200,7200,7200,7200,7200,7200,7200]
alg4_col =[0.208,7.418,7200,7200,7200,7200,7200,7200,7200,7200]
# +
from matplotlib.pyplot import figure
figure(num=None, figsize=(10, 8), dpi=80, facecolor='w', edgecolor='k')
plt.grid(True)
x=[10,20,30,40,50,60,70,80,90,100]
#plt.yticks(np.arange(0, 5, 0.1))
plt.xticks(x)
plt.plot(x,alg1_col,c='red',linewidth=6,label= 'Algorithm 1')
plt.plot(x,alg2_col,c= 'blue',linewidth=4,label= 'Algorithm 2')
plt.plot(x,alg3_col,c='green',linewidth=2,label= 'Algorithm 3')
plt.plot(x,alg4_col,c= 'black',linewidth=1,label= 'Algorithm 4')
plt.xlabel('Number of the columns(Features)')
plt.ylabel('Time (sec)')
plt.legend()
# -
# # plotting
# - prunning columns
alg1_col_prunning =[0.373,2.437,143.573,269.429,3028.437,3028.437]
alg2_col_prunning =[0.217,0.742,2.029,3.315,5.455,4.870]
alg3_col_prunning =[0.189,3.051,30.391,62.347,254.160,609.44]
alg4_col_prunning =[0.192,1.910,10.427,29.151,261.219,281.683]
# +
from matplotlib.pyplot import figure
figure(num=None, figsize=(10, 8), dpi=80, facecolor='w', edgecolor='k')
plt.grid(True)
x=[10,20,30,40,50,60]
#plt.yticks(np.arange(0, 5, 0.1))
plt.xticks(x)
plt.plot(x,alg1_col_prunning,c='red',linewidth=3,label= 'Algorithm 1')
plt.plot(x,alg2_col_prunning,c= 'blue',linewidth=3,label= 'Algorithm 2')
plt.plot(x,alg3_col_prunning,c='green',linewidth=3,label= 'Algorithm 3')
plt.plot(x,alg4_col_prunning,c= 'black',linewidth=3,label= 'Algorithm 4')
plt.xlabel('Number of the columns(Features)')
plt.ylabel('Time (sec)')
plt.legend()
# -
# # after fix bugs
alg1_col_increase =[0.192,0.235,]
alg2_col_increase =[0.070,0.083,]
alg3_col_increase =[0.043,4.350,]
alg4_col_increase =[0.042,4.367]
| error_generator/strategies/adversarial_machine_learning/compare 4 Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Using Convolutional Neural Networks
# ## Use a pretrained VGG model with our **Vgg16** class
# %matplotlib inline
from __future__ import division, print_function
import os, json
from glob import glob
import numpy as np
np.set_printoptions(precision=4, linewidth=1)
from matplotlib import pyplot as plt
import utils; reload(utils)
from utils import plots
import vgg16; reload(vgg16)
from vgg16 import Vgg16
vgg = Vgg16()
batch_size = 64
path = "dogs_cats/data/sample/"
batches = vgg.get_batches(path + 'train', batch_size = batch_size)
val_batches = vgg.get_batches(path + 'valid', batch_size = (batch_size * 2))
vgg.finetune(batches)
vgg.fit(batches, val_batches, nb_epoch=1)
# ## Use Vgg16 for basic image recognition
vgg = Vgg16()
batches = vgg.get_batches(path+'train', batch_size = 2)
imgs,labels = next(batches)
vgg.predict(imgs, True)
# ## Use our Vgg16 class to finetune a Dogs vs Cats model
batches = vgg.get_batches(path + 'train', batch_size = batch_size)
val_batches = vgg.get_batches(path + 'valid', batch_size = batch_size)
vgg.finetune(batches)
vgg.fit(batches, val_batches, nb_epoch=4)
imgs,labels = next(batches)
vgg.predict(imgs, True)
# # Questions
#
# *finetune* - Modifies data such that it will be trained based on data in batches provided (dog, cat)
#
# * So instead of categorizing based on specific category, it groups it by a larger category "dogs" and "cats". Does this mean the original model already has a concept of dogs and cats? It would seem like it has to, otherwise it would be hard to map
#
# `German Shepherd -> Dog`
#
# Otherwise the original training data would be useless.
#
# * so finetune really __adjusts the specificity of a given category__?
# * How does it get more accurate after running *fit* multiple times over the *same* data?? What is new?
# * What is the difference between *finetune* and *fit*
# # Create a VGG model from scratch in Keras
# +
from numpy.random import random, permutation
from scipy import misc, ndimage
from scipy.ndimage.interpolation import zoom
import keras
from keras import backend as K
from keras.utils.data_utils import get_file
from keras.models import Sequential, Model
from keras.layers.core import Flatten, Dense, Dropout, Lambda
from keras.layers import Input
from keras.layers.convolutional import Convolution2D, MaxPooling2D, ZeroPadding2D
from keras.optimizers import SGD, RMSprop
from keras.preprocessing import image
# +
FILES_PATH = 'http://www.platform.ai/models/'; CLASS_FILE='imagenet_class_index.json'
fpath = get_file(CLASS_FILE, FILES_PATH+CLASS_FILE, cache_subdir='models')
with open(fpath) as f: class_dict = json.load(f)
classes = [class_dict[str(i)][1] for i in range(len(class_dict))]
# +
vgg_mean = np.array([123.68, 116.779, 103.939]).reshape((3,1,1))
def vgg_preprocess(x):
x = x - vgg_mean
return x[:, ::-1]
def FCBlock(model):
model.add(Dense(4096, activation='relu'))
model.add(Dropout(0.5))
def ConvBlock(layers, model, filters):
for i in range(layers):
model.add(ZeroPadding2D((1,1)))
model.add(Convolution2D(filters, 3, 3, activation='relu'))
model.add(MaxPooling2D((2,2), strides=(2,2)))
#Adding different layers?
#What this stuff doing?
def VGG_16():
model = Sequential()
model.add(Lambda(vgg_preprocess, input_shape=(3,224,224)))
ConvBlock(2, model, 64)
ConvBlock(2, model, 128)
ConvBlock(3, model, 256)
ConvBlock(3, model, 512)
ConvBlock(3, model, 512)
model.add(Flatten())
FCBlock(model)
FCBlock(model)
#Calling this twice?
model.add(Dense(1000, activation='softmax'))
return model
model = VGG_16()
fpath = get_file('vgg16.h5', FILES_PATH+'vgg16.h5', cache_subdir='models')
model.load_weights(fpath)
# +
def get_batches(dirname, gen=image.ImageDataGenerator(), shuffle=True, batch_size=4, class_mode='categorical'):
return gen.flow_from_directory(path+dirname,target_size=(224, 224), class_mode=class_mode, shuffle=shuffle, batch_size=batch_size)
batches = get_batches('train', batch_size = 4)
imgs,labels = next(batches)
plots(imgs, titles=labels)
# +
def pred_batch(imgs):
preds = model.predict(imgs)
idxs = np.argmax(preds, axis = 1)
print('Shape: {}'.format(preds.shape))
print('First 5 classes: {}'.format(classes[:5]))
print('First 5 probabilities: {}\n'.format(preds[0, :5]))
print('Predictions prob/class: ')
for i in range(len(idxs)):
idx = idxs[i]
print(' {:.4f}/{}'.format(preds[i, idx], classes[idx]))
pred_batch(imgs)
# -
# # Comments
#
# * The model architecture is created by a series of "blocks" to a "Sequential" model
# * The model is then trained on a set of data, resulting in a calculation of a set of __weights__
# * By making the weights available, we can just download the weights file and use the trained model without retraining it
# * __trained model__ = weights + architecture ??
| deeplearning1/nbs/lesson1_jvans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # <NAME> WINE QUALITY PREDICTION
# ### Loading Packages
# +
# # %%writefile train.py
#####################################################################################
#####################################################################################
# Loading the iconic trio 🔥
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('fivethirtyeight')
# Importing model_selection to get access to some dope functions like GridSearchCV()
from sklearn import model_selection
from sklearn.preprocessing import StandardScaler
from imblearn.over_sampling import SVMSMOTE
# Loading models
from sklearn import tree
from sklearn import ensemble
import xgboost
from sklearn import linear_model
# custom
from custom import helper
# Loading evaluation metrics
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import r2_score
import pickle
#####################################################################################
#####################################################################################
# +
####################
### Loading Data ###
####################
preprocessed_data = pd.read_csv("data/preprocessed_data.csv")
# preprocessed_data.head()
# +
##########################################
### Creating a Copy of the Loaded Data ###
##########################################
data_with_targets = preprocessed_data.copy()
# data_with_targets.head()
# +
#################################
### Dropping `quality` Column ###
#################################
data_with_targets = data_with_targets.drop(['quality'], axis=1)
# data_with_targets.head()
# +
###############################################################
### Splitting the Data into Feature Matrix and Target Label ###
###############################################################
target_variable = 'quality_rate'
# Unscaled Features
X = data_with_targets.drop([target_variable], axis=1)
# Target Variable
y = data_with_targets[target_variable]
# +
#####################################################
### SMOTE Sampling to deal with imbalance classes ###
#####################################################
# Setting Seed Value
seed = 81
smote = SVMSMOTE(random_state=seed)
resampled_X, resampled_y = smote.fit_sample(X, y)
# +
##########################################
### Splitting into Train and Test sets ###
##########################################
X_train, X_test, y_train, y_test = model_selection.train_test_split(resampled_X, resampled_y, test_size=0.3, stratify=resampled_y, random_state=seed)
# +
####################################
### Scalling Train and Test sets ###
####################################
column_names = list(X.columns.values)
scaler = StandardScaler()
normalized_X_train = pd.DataFrame(
scaler.fit_transform(X_train),
columns=column_names,
index=X_train.index
)
normalized_X_test = pd.DataFrame(
scaler.transform(X_test),
columns=column_names,
index=X_test.index
)
# +
#################
### Modelling ###
#################
# Instantiating baseline models
models = [
("Decision Tree", tree.DecisionTreeClassifier(random_state=seed)),
("Random Forest", ensemble.RandomForestClassifier(random_state=seed)),
("AdaBoost", ensemble.AdaBoostClassifier(random_state=seed)),
("ExtraTree", ensemble.ExtraTreesClassifier(random_state=seed)),
("GradientBoosting", ensemble.GradientBoostingClassifier(random_state=seed)),
("XGBOOST", xgboost.XGBClassifier(random_state=seed)),
]
feature_importance_of_models, df_model_features_with_importance, model_summary = helper.baseline_performance(
models=models,
X_train=normalized_X_train,
y_train=y_train,
X_test=normalized_X_test,
y_test=y_test,
column_names=list(X.columns.values),
csv_path='csv_tables',
save_model_summary=True,
save_feature_importance=True,
save_feature_imp_of_each_model=True,
)
# +
##########################################
### Plotting Train and Test Accuracies ###
##########################################
helper.plot_model_summary(
model_summary=model_summary,
figsize=(20, 14),
dpi=600,
transparent=True,
save_visualization=True,
figure_name='Train and Test Accuracies',
figure_path='figures',
)
# +
###################################
### Plotting Feature Importance ###
###################################
helper.plot_feature_importance(
df_model_features_with_importance,
figsize=(20, 15),
dpi=600,
transparent=True,
annotate_fontsize='xx-large',
save_plot=True,
path='figures',
)
# +
##############################
### Getting Top 6 Features ###
##############################
top_6_features = list(feature_importance_of_models['GradientBoosting'].head(6))
# top_6_features
normalized_X_train_new = normalized_X_train[top_6_features]
# normalized_X_train_new.head()
normalized_X_test_new = normalized_X_test[top_6_features]
# normalized_X_test_new.head()
# X_train_new, X_test_new, y_train_new, y_test_new = model_selection.train_test_split(X_new, y, test_size=0.2, stratify=y, random_state=80)
# +
############################
### Creating New Folders ###
############################
helper.create_folder('./new_csv_tables/')
helper.create_folder('./new_figures/')
# +
###################################
### Modelling on Top 6 Features ###
###################################
feature_importance_of_models_new, model_features_with_importance_new, model_summary_new = helper.baseline_performance(
models=models,
X_train=normalized_X_train_new,
y_train=y_train,
X_test=normalized_X_test_new,
y_test=y_test,
column_names=list(top_6_features),
csv_path='new_csv_tables',
save_model_summary=True,
save_feature_importance=True,
save_feature_imp_of_each_model=True
)
# +
##########################################
### Plotting Train and Test Accuracies ###
##########################################
helper.plot_model_summary(
model_summary=model_summary_new,
figsize=(20, 14),
dpi=300,
transparent=True,
save_visualization=True,
figure_name='Train and Test Accuracies_new',
figure_path='new_figures',
)
# +
###################################
### Plotting Feature Importance ###
###################################
helper.plot_feature_importance(
feature_importance=model_features_with_importance_new,
figsize=(20, 14),
dpi=600,
transparent=True,
annotate_fontsize='xx-large',
save_plot=True,
path='new_figures',
)
# +
###############################
### Cross Validating Models ###
###############################
# Splitting data into 10 folds
cv_kfold = model_selection.KFold(n_splits=10, shuffle=True, random_state=150)
scorer = "f1"
# Instantiating model_names as an empty list to keep the names of the models
model_names = []
# Instantiating cv_mean_scores as an empty list to keep the cross validation mean score of each model
cv_mean_scores = []
# Instantiating cv_std_scores as an empty list to keep the cross validation standard deviation score of each model
cv_std_scores = []
# Looping through the baseline models and cross validating each model
for model_name, model in models:
model_scores = model_selection.cross_val_score(
model, X, y, cv=cv_kfold, scoring=scorer, n_jobs=-1, verbose=1,
)
print(
f"{model_name} Score: %0.2f (+/- %0.2f)"
% (model_scores.mean(), model_scores.std() * 2)
)
# Appending model names to model_name
model_names.append(model_name)
# Appending cross validation mean score of each model to cv_mean_score
cv_mean_scores.append(model_scores.mean())
# Appending cross validation standard deviation score of each model to cv_std_score
cv_std_scores.append(model_scores.std())
# Parsing model_names, cv_mean_scores and cv_std_scores and a pandas DataFrame object
cv_results = pd.DataFrame({"model_name": model_names, "mean_score": cv_mean_scores, "std_score": cv_std_scores})
# Sorting the Dataframe in descending order
cv_results.sort_values("mean_score", ascending=False, inplace=True,)
# Saving the DataFrame as a csv file
cv_results.to_csv("csv_tables/cross_validation_results.csv", index=True)
# Showing the final results
# cv_results
# +
#######################################
### Choosing Classifier to Evaluate ###
#######################################
# classifier = models[2][1]
classifier = models[4][1]
classifier.fit(normalized_X_train_new, y_train)
# +
############################################################
### Getting Train and Test Accuracy of the Choosen Model ###
############################################################
train_accuracy = classifier.score(normalized_X_train_new, y_train)
test_accuracy = classifier.score(normalized_X_test_new, y_test)
# +
#############################
### Evaluating Classifier ###
#############################
helper.evaluate_classifier(
estimator=classifier,
X_test=normalized_X_test_new,
y_test=y_test,
save_figure=True,
figure_path='figures',
transparent=True,
dpi=600,
cmap="Purples",
)
| White Wine Quality Prediction/White Wine Quality Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Import Library
# Import General Packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import confusion_matrix, classification_report
import pickle
from pathlib import Path
import warnings
#warnings.filterwarnings('ignore')
# import dataset
df_load = pd.read_csv('https://dqlab-dataset.s3-ap-southeast-1.amazonaws.com/dqlab_telco_final.csv')
# Show the shape of the dataset
df_load.shape
# Show top 5 records
df_load.head()
# Show number of unique IDs
df_load.customerID.nunique()
# # Exploratory Data Analysis (EDA)
#
# Dalam kasus ini, Saya diminta untuk melihat persebaran dari:
# - Prosentase persebaran data Churn dan tidaknya dari seluruh data
# - Persebarang data dari variable predictor terhadap label (Churn)
#
# see univariate data visualization related to the percentage of churn data from customers
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.axis('equal')
labels = ['Yes','No']
churn = df_load.Churn.value_counts()
ax.pie(churn, labels=labels, autopct='%.0f%%')
plt.show()
# +
# choose a numeric variable predictor and make a bivariate plot, then interpret it
# creating bin in chart
numerical_features = ['MonthlyCharges','TotalCharges','tenure']
fig, ax = plt.subplots(1, 3, figsize=(15, 6))
# use the following code to plot two overlays of histogram per each numerical_features,
# use a color of blue and orange, respectively
df_load[df_load.Churn == 'No'][numerical_features].hist(bins=20, color='blue', alpha=0.5, ax=ax)
df_load[df_load.Churn == 'Yes'][numerical_features].hist(bins=20, color='orange', alpha=0.5, ax=ax)
plt.show()
# +
# choose a categorical predictor variable and make a bivariate plot, then interpret it
fig, ax = plt.subplots(3, 3, figsize=(14, 12))
sns.set(style='darkgrid')
sns.countplot(data=df_load, x='gender', hue='Churn', ax=ax[0][0])
sns.countplot(data=df_load, x='Partner', hue='Churn', ax=ax[0][1])
sns.countplot(data=df_load, x='SeniorCitizen', hue='Churn', ax=ax[0][2])
sns.countplot(data=df_load, x='PhoneService', hue='Churn', ax=ax[1][0])
sns.countplot(data=df_load, x='StreamingTV', hue='Churn', ax=ax[1][1])
sns.countplot(data=df_load, x='InternetService', hue='Churn', ax=ax[1][2])
sns.countplot(data=df_load, x='PaperlessBilling', hue='Churn', ax=ax[2][1])
plt.tight_layout()
plt.show()
# -
# **Conclusion**
#
# Based on the results and analysis above, it can be concluded:
#
# - At the first step, we know that the data distribution as a whole, the customer does not churn, with details on Churn as much as 26% and No Churn as much as 74%.
# - At the second step, we can see that for MonthlyCharges there is a tendency that the smaller the value of the monthly fees charged, the smaller the tendency to do Churn. For TotalCharges there doesn't seem to be any inclination towards Churn customers. For tenure, there is a tendency that the longer the customer subscribes, the less likely it is to churn.
# - At the third step, we know that there is no significant difference for people doing churn in terms of gender and telephone service (Phone Service). However, there is a tendency that people who churn are people who do not have a partner (partner: No), people whose status is a senior citizen (Senior Citizen: Yes), people who have streaming TV services (StreamingTV: Yes) , people who have Internet service (internetService: Yes) and people who have paperless bills (PaperlessBilling: Yes).
# # Pre-Processing Data
df_load.head()
#Remove the unnecessary columns customerID & UpdatedAt
cleaned_df = df_load.drop(['customerID','UpdatedAt'], axis=1)
cleaned_df.head()
cleaned_df.describe()
# +
# Encoding Data
#Convert all the non-numeric columns to numerical data types
for column in cleaned_df.columns:
if cleaned_df[column].dtype == np.number: continue
# Perform encoding for each non-numeric column
cleaned_df[column] = LabelEncoder().fit_transform(cleaned_df[column])
# -
cleaned_df.describe()
# +
# Splitting Dataset
# Predictor and Target
X = cleaned_df.drop('Churn', axis = 1)
y = cleaned_df['Churn']
# Splitting train and test
# Splitting train and test
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# -
# Print according to the expected result
print('The number of rows and columns of x_train is: ', x_train.shape, ', while the number of rows and columns of y_train is:', y_train.shape)
print('\nChurn percentage in training data is:')
print(y_train.value_counts(normalize=True))
print('\nThe number of rows and columns of x_test is:', x_test.shape,', while the number of rows and columns of y_test is:', y_test.shape)
print('\nChurn percentage in Testing data is:')
print(y_test.value_counts(normalize=True))
# **Conclusion**
#
# After we analyzed it further, it turned out that there were columns that were not needed in the model, namely the customer ID number (customerID) & the data collection period (UpdatedAt), so this needs to be deleted. Then we continue to change the value of the data which is still in the form of a string into numeric through encoding, after this is done, it can be seen in the data distribution, especially the min and max columns of each variable have changed to 0 & 1. The last step is to divide the data into 2 parts for modeling purposes After it is done, it can be seen that the number of rows and columns of each data is appropriate & the percentage of the churn column is also the same as the data at the beginning, this indicates that the data is separated properly and correctly.
# # LogisticRegression
# +
# Create a model using the LogisticRegression Algorithm
warnings.filterwarnings('ignore')
log_model = LogisticRegression().fit(x_train, y_train)
# -
# LogisticRegression Model
log_model
# Predict
y_train_pred = log_model.predict(x_train)
# Print classification report
print(classification_report(y_train, y_train_pred))
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_train, y_train_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# +
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Training Model\n(Logistic Regression)', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# +
# Performance Data Testing - Displays Metrics
# Predict
y_test_pred = log_model.predict(x_test)
# Print classification report
print(classification_report(y_test, y_test_pred))
# +
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_test, y_test_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Testing Model\n(Logistic Regression)\n', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# -
# **Conclusion**
#
# From the results and analysis above, then:
#
# * From the training data, it can be seen that the model is able to predict data with an accuracy of 79%, with details of the churn guess which is actually correct, the churn guess is 636, the churn guess which is actually not churn is 3227, the churn guess which is actually correct is 654 and the churn guess that is actually correct the actual churn is 348.
# * From the testing data, it can be seen that the model is able to predict the data with an accuracy of 79%, with details of the churn guess which is actually true churn is 263, the non-churn guess that actually doesn't churn is 1390, the churn guess which is actually true churn is 283 and the churn guess which is actually correct actually no churn is 149.
# # Random Forest Classifier
# Create a model using RandomForestClassifier
rdf_model = RandomForestClassifier().fit(x_train, y_train)
rdf_model
# +
# Predict
y_train_pred = rdf_model.predict(x_train)
# Print classification report
print(classification_report(y_train, y_train_pred))
# +
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_train, y_train_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Training Model\n(Random Forest)', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# +
# Performance Data Testing - Displays Metrics
# Predict
y_test_pred = rdf_model.predict(x_test)
# Print classification report
print(classification_report(y_test, y_test_pred))
# +
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_test, y_test_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Testing Model\n(Random Forest)\n', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# -
# **Kesimpulan**
#
# Dari hasil dan analisa di atas, maka:
#
# - Jika kita menggunakan menggunakan algoritma Random Forest dengan memanggil RandomForestClassifier() dari sklearn tanpa menambahi parameter apapun, maka yang dihasilkan adalah model dengan seting default dari sklearn, untuk detilnya bisa dilihat di dokumentasinya.
# - Dari data training terlihat bahwasannya model mampu memprediksi data dengan menghasilkan akurasi sebesar 100%, dengan detil tebakan churn yang sebenernya benar churn adalah 1278, tebakan tidak churn yang sebenernya tidak churn adalah 3566, tebakan tidak churn yang sebenernya benar churn adalah 12 dan tebakan churn yang sebenernya tidak churn adalah 9.
# - Dari data testing terlihat bahwasannya model mampu memprediksi data dengan menghasilkan akurasi sebesar 78%, dengan detil tebakan churn yang sebenernya benar churn adalah 262, tebakan tidak churn yang sebenernya tidak churn adalah 1360, tebakan tidak churn yang sebenernya benar churn adalah 284 dan tebakan churn yang sebenernya tidak churn adalah 179.
#
# # Gradient Boosting Classifier
# +
#Train the model
gbt_model = GradientBoostingClassifier().fit(x_train, y_train)
gbt_model
# +
# Predict
y_train_pred = gbt_model.predict(x_train)
# Print classification report
print(classification_report(y_train, y_train_pred))
# +
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_train, y_train_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Training Model\n(Gradient Boosting)', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# +
# Predict
y_test_pred = gbt_model.predict(x_test)
# Print classification report
print(classification_report(y_test, y_test_pred))
# +
# Form confusion matrix as a DataFrame
confusion_matrix_df = pd.DataFrame((confusion_matrix(y_test, y_test_pred)), ('No churn', 'Churn'), ('No churn', 'Churn'))
# Plot confusion matrix
plt.figure()
heatmap = sns.heatmap(confusion_matrix_df, annot=True, annot_kws={'size': 14}, fmt='d', cmap='YlGnBu')
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=0, ha='right', fontsize=14)
plt.title('Confusion Matrix for Testing Model\n(Gradient Boosting)', fontsize=18, color='darkblue')
plt.ylabel('True label', fontsize=14)
plt.xlabel('Predicted label', fontsize=14)
plt.show()
# -
# **Kesimpulan**
#
# Dari hasil dan analisa di atas, maka:
#
# - Jika kita menggunakan menggunakan algoritma Gradient Boosting dengan memanggil GradientBoostingClassifier() dari package sklearn tanpa menambahi parameter apapun, maka yang dihasilkan adalah model dengan seting default dari sklearn, untuk detilnya bisa dilihat di dokumentasinya.
# - Dari data training terlihat bahwasannya model mampu memprediksi data dengan menghasilkan akurasi sebesar 82%, dengan detil tebakan churn yang sebenernya benar churn adalah 684, tebakan tidak churn yang sebenernya tidak churn adalah 3286, tebakan tidak churn yang sebenernya benar churn adalah 606 dan tebakan churn yang sebenernya tidak churn adalah 289.
# - Dari data testing terlihat bahwasannya model mampu memprediksi data dengan menghasilkan akurasi sebesar 79%, dengan detil tebakan churn yang sebenernya benar churn adalah 261, tebakan tidak churn yang sebenernya tidak churn adalah 1394, tebakan tidak churn yang sebenernya benar churn adalah 285 dan tebakan churn yang sebenernya tidak churn adalah 145.
# Save model
pickle.dump(log_model, open('best_model_churn.pkl', 'wb'))
# Kesimpulan
#
# Berdasarkan pemodelan yang telah dilakukan dengan menggunakan Logistic Regression, Random Forest dan Extreme Gradiant Boost, maka dapat disimpulkan untuk memprediksi churn dari pelanggan telco dengan menggunakan dataset ini model terbaiknya adalah menggunakan algortima Logistic Regression. Hal ini dikarenakan performa dari model Logistic Regression cenderung mampu memprediksi sama baiknya di fase training maupun testing (akurasi training 80%, akurasi testing 79%), dilain sisi algoritma lainnya cenderung Over-Fitting performanya. Akan tetapi hal ini tidak menjadikan kita untuk menarik kesimpulan bahwsannya jika untuk melakukan pemodelan apapun maka digunakan Logistic Regression, kita tetap harus melakukan banyak percobaan model untuk menentukan mana yang terbaik.
| modelClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
import numpy as np
from keras.models import Model
from keras.layers import Input
from keras.layers.pooling import MaxPooling2D
from keras import backend as K
def format_decimal(arr, places=6):
return [round(x * 10**places) / 10**places for x in arr]
# ### MaxPooling2D
# **[pooling.MaxPooling2D.0] input 6x6x3, pool_size=(2, 2), strides=None, border_mode='valid', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='valid', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(270)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.1] input 6x6x3, pool_size=(2, 2), strides=(1, 1), border_mode='valid', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=(1, 1), border_mode='valid', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(271)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.2] input 6x7x3, pool_size=(2, 2), strides=(2, 1), border_mode='valid', dim_ordering='tf'**
# +
data_in_shape = (6, 7, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=(2, 1), border_mode='valid', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(272)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.3] input 6x6x3, pool_size=(3, 3), strides=None, border_mode='valid', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=None, border_mode='valid', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(273)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.4] input 6x6x3, pool_size=(3, 3), strides=(3, 3), border_mode='valid', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=(3, 3), border_mode='valid', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(274)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.5] input 6x6x3, pool_size=(2, 2), strides=None, border_mode='same', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='same', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(275)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.6] input 6x6x3, pool_size=(2, 2), strides=(1, 1), border_mode='same', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=(1, 1), border_mode='same', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(276)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.7] input 6x7x3, pool_size=(2, 2), strides=(2, 1), border_mode='same', dim_ordering='tf'**
# +
data_in_shape = (6, 7, 3)
L = MaxPooling2D(pool_size=(2, 2), strides=(2, 1), border_mode='same', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(277)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.8] input 6x6x3, pool_size=(3, 3), strides=None, border_mode='same', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=None, border_mode='same', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(278)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.9] input 6x6x3, pool_size=(3, 3), strides=(3, 3), border_mode='same', dim_ordering='tf'**
# +
data_in_shape = (6, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=(3, 3), border_mode='same', dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(279)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.10] input 5x6x3, pool_size=(3, 3), strides=(2, 2), border_mode='valid', dim_ordering='th'**
# +
data_in_shape = (5, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=(2, 2), border_mode='valid', dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(280)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.11] input 5x6x3, pool_size=(3, 3), strides=(1, 1), border_mode='same', dim_ordering='th'**
# +
data_in_shape = (5, 6, 3)
L = MaxPooling2D(pool_size=(3, 3), strides=(1, 1), border_mode='same', dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(281)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[pooling.MaxPooling2D.12] input 4x6x4, pool_size=(2, 2), strides=None, border_mode='valid', dim_ordering='th'**
# +
data_in_shape = (4, 6, 4)
L = MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='valid', dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(282)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
| notebooks/layers/pooling/MaxPooling2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing Personalization: LOSO vs Within-Subjects
# ## Set up the Notebook
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import importlib, sys, os
sys.path.insert(0, os.path.abspath('..'))
if(importlib.util.find_spec("mFlow") is None):
# !git clone https://github.com/mlds-lab/mFlow.git
# !pip install ./mFlow
else:
print("mFlow module found")
# ## Import modules
# +
from mFlow.Blocks.data_loader_extrasensory import extrasensory_data_loader
from mFlow.Blocks.filter import MisingLabelFilter, MisingDataColumnFilter, Take
from mFlow.Blocks.imputer import Imputer
from mFlow.Blocks.normalizer import Normalizer
from mFlow.Blocks.experimental_protocol import ExpTrainTest, ExpCV, ExpWithin
from mFlow.Blocks.results_analysis import ResultsConcat, ResultsCVSummarize, DataYieldReport
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score
from sklearn.linear_model import LogisticRegression
import matplotlib.pyplot as plt
from mFlow.Workflow.workflow import workflow
import mFlow.Workflow.compute_graph
# -
# ## Define the workflow
#
# This workflow compares a within-subject experimental design to the leave-one-subject-out (LOSO) design. In the first case, one model is trained per subject using only that subject's data. In the second case, for each subject, a model is trained using data from all other subjects and then applied to the target subject. The within-subject design thus corresponds to a personalized model fit to an individual subject, while the LOSO design corresponds to using a model fit to other subjects. We use the ExtraSensory data set sleeping prediction task using a subset of the first 50,000 instances. The model used is logistic regression with a default regularization hyper-parameters.
#
# The workflow includes a column filter that screens out feature dimensions that are less than 20% observed, and a missing label filter that removes instances without labels. Next, the workflow performs mean imputation followed by feature normalization. Finally, each of the two experimental protocols are applied. Results are displayed per-subject.
#
# In this particular case, we see that the accuracy measured by the within-subject protocol is slightly better than that of the LOSO design, but the difference does not look significant. Further, it is important to remember that the LOSO and within-subjects test sets are not the same due to the fact that some data for each subject are used for training in the within-subject design, so more work is needed to conclusively state that personalization provides an advantage for this task.
#
# +
metrics = [accuracy_score, f1_score, precision_score, recall_score]
df_raw = extrasensory_data_loader(label="SLEEPING");
df_sub = Take(df_raw, 50000)
df_cf = MisingDataColumnFilter(df_sub);
df_lf = MisingLabelFilter(df_cf);
df_imp = Imputer(df_lf)
df_norm = Normalizer(df_imp);
report = DataYieldReport(df_norm, names=["Norm"])
flow = workflow({"yield":report});
flow.draw(); plt.show();
output = flow.run();
num = output['yield']['report']["#Individuals with Data"]["Norm"]
display(output['yield']['report'])
estimators1 = {"Within-LR": LogisticRegression(solver="lbfgs",max_iter=100)}
res_within = ExpWithin(df_norm, estimators1, metrics=metrics, n_folds=num);
estimators2 = {"LOSO-LR": LogisticRegression(solver="lbfgs",max_iter=100)}
res_loso = ExpCV(df_norm, estimators2, metrics=metrics, n_folds=num);
res_cat = ResultsConcat(res_loso, res_within)
summary = ResultsCVSummarize(res_cat)
flow=workflow({"results":summary})
output=flow.run(backend="sequential", monitor=True,from_scratch=True);
# -
output["results"]["report"]
| Examples/ExtraSensory-ComparingPersonalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Bernstein-Vazirani
# %matplotlib inline
from random import randint
from qiskit import *
# Step 0: Generate a secret string
secret_string = bin(randint(0, 2**7 - 1))[2:]
N = len(secret_string)
# Step 1: Prepare the state with superpositions
prep_circuit = QuantumCircuit(N+1, N)
prep_circuit.x(N) # working qubit starts with |1>
prep_circuit.h(range(N+1))
prep_circuit.barrier()
prep_circuit.draw(output="mpl")
# +
# Step 2: send the query to the quantum oracle. Here, we prepare the quantum oracle
oracle_circuit = QuantumCircuit(N+1, N, name="Blackbox")
for index, value in enumerate(secret_string):
if value == "1":
oracle_circuit.cx(index, N) # XOR with working qubit
oracle_circuit.barrier()
oracle_circuit.draw(output="mpl")
# -
# Step 3: Apply Hadamard gates and measure
measure_circuit = QuantumCircuit(N+1, N)
measure_circuit.h(range(N))
measure_circuit.measure(
list(range(N)),
list(reversed(range(N)))
)
measure_circuit.draw(output="mpl")
# We now merge the circuit together
circuit = QuantumCircuit(N+1, N)
circuit += prep_circuit
circuit.append(oracle_circuit.to_instruction(), range(N+1)) # We do this to "hide" the oracle from the viewer
circuit.barrier()
circuit += measure_circuit
circuit.draw(output="mpl")
# We simulate to discover the hidden string
simulator = Aer.get_backend("qasm_simulator")
job = execute(circuit, simulator, shots=512)
results = job.result()
visualization.plot_histogram(results.get_counts())
# Was the guess right? Let's discover
print("Hidden string {}\nGuess {}".format(
secret_string, list(results.get_counts().keys())[0])
)
if list(results.get_counts().keys())[0] == secret_string:
print("The guess was correct!")
else:
print("The guess was not correct")
# For purposes of reproducibility, the Qiskit version is
qiskit.__qiskit_version__
| Algorithms/Bernstein-Vazirani.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://landlab.github.io"><img style="float: left" src="../../../landlab_header.png"></a>
# # Components for modeling overland flow erosion
#
# *(<NAME>, July 2021)*
#
# There are two related components that calculate erosion resulting from surface-water flow, a.k.a. overland flow: `DepthSlopeProductErosion` and `DetachmentLtdErosion`. They were originally created by <NAME> to work with the `OverlandFlow` component, which solves for water depth across the terrain. They are similar to the `StreamPowerEroder` and `FastscapeEroder` components in that they calculate erosion resulting from water flow across a topographic surface, but whereas these components require a flow-routing algorithm to create a list of node "receivers", the `DepthSlopeProductErosion` and `DetachmentLtdErosion` components only require a user-identified slope field together with an at-node depth or discharge field (respectively).
# ## `DepthSlopeProductErosion`
#
# This component represents the rate of erosion, $E$, by surface water flow as:
#
# $$E = k_e (\tau^a - \tau_c^a)$$
#
# where $k_e$ is an erodibility coefficient (with dimensions of velocity per stress$^a$), $\tau$ is bed shear stress, $\tau_c$ is a minimum bed shear stress for any erosion to occur, and $a$ is a parameter that is commonly treated as unity.
#
# For steady, uniform flow,
#
# $$\tau = \rho g H S$$,
#
# with $\rho$ being fluid density, $g$ gravitational acceleration, $H$ local water depth, and $S$ the (postive-downhill) slope gradient (an approximation of the sine of the slope angle).
#
# The component uses a user-supplied slope field (at nodes) together with the water-depth field `surface_water__depth` to calculate $\tau$, and then the above equation to calculate $E$. The component will then modify the `topographic__elevation` field accordingly. If the user wishes to apply material uplift relative to baselevel, an `uplift_rate` parameter can be passed on initialization.
#
# We can learn more about this component by examining its internal documentation. To get an overview of the component, we can examine its *header docstring*: internal documentation provided in the form of a Python docstring that sits just below the class declaration in the source code. This text can be displayed as shown here:
# +
from landlab.components import DepthSlopeProductErosion
print(DepthSlopeProductErosion.__doc__)
# -
# A second useful source of internal documentation for this component is its *init docstring*: a Python docstring that describes the component's class `__init__` method. In Landlab, the init docstrings for components normally provide a list of that component's parameters. Here's how to display the init docstring:
print(DepthSlopeProductErosion.__init__.__doc__)
# ### Example
#
# In this example, we load the topography of a small drainage basin, calculate a water-depth field by running overland flow over the topography using the `KinwaveImplicitOverlandFlow` component, and then calculating the resulting erosion.
#
# Note that in order to accomplish this, we need to identify which variable we wish to use for slope gradient. This is not quite as simple as it may sound. An easy way to define slope is as the slope between two adjacent grid nodes. But using this approach means that slope is defined on the grid *links* rathter than *nodes*. To calculate slope magnitude at *nodes*, we'll define a little function below that uses Landlab's `calc_grad_at_link` method to calculate gradients at grid links, then use the `map_link_vector_components_to_node` method to calculate the $x$ and $y$ vector components at each node. With that in hand, we just use the Pythagorean theorem to find the slope magnitude from its vector components.
#
# First, though, some imports we'll need:
import copy
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from landlab import imshow_grid
from landlab.components import KinwaveImplicitOverlandFlow
from landlab.grid.mappers import map_link_vector_components_to_node
from landlab.io import read_esri_ascii
def slope_magnitude_at_node(grid, elev):
# calculate gradient in elevation at each link
grad_at_link = grid.calc_grad_at_link(elev)
# set the gradient to zero for any inactive links
# (those attached to a closed-boundaries node at either end,
# or connecting two boundary nodes of any type)
grad_at_link[grid.status_at_link != grid.BC_LINK_IS_ACTIVE] = 0.0
# map slope vector components from links to their adjacent nodes
slp_x, slp_y = map_link_vector_components_to_node(grid, grad_at_link)
# use the Pythagorean theorem to calculate the slope magnitude
# from the x and y components
slp_mag = (slp_x * slp_x + slp_y * slp_y) ** 0.5
return slp_mag, slp_x, slp_y
# (See [here](https://landlab.readthedocs.io/en/latest/reference/grid/gradients.html#landlab.grid.gradients.calc_grad_at_link) to learn how `calc_grad_at_link` works, and [here](https://landlab.readthedocs.io/en/latest/reference/grid/raster_mappers.html#landlab.grid.raster_mappers.map_link_vector_components_to_node_raster) to learn how
# `map_link_vector_components_to_node` works.)
#
# Next, define some parameters we'll need.
#
# To estimate the erodibility coefficient $k_e$, one source is:
#
# [http://milford.nserl.purdue.edu/weppdocs/comperod/](http://milford.nserl.purdue.edu/weppdocs/comperod/)
#
# which reports experiments in rill erosion on agricultural soils. Converting their data into $k_e$, its values are on the order of 1 to 10 $\times 10^{-6}$ (m / s Pa), with threshold ($\tau_c$) values on the order of a few Pa.
# +
# Process parameters
n = 0.1 # roughness coefficient, (s/m^(1/3))
dep_exp = 5.0 / 3.0 # depth exponent
R = 72.0 # runoff rate, mm/hr
k_e = 4.0e-6 # erosion coefficient (m/s)/(kg/ms^2)
tau_c = 3.0 # erosion threshold shear stress, Pa
# Run-control parameters
rain_duration = 240.0 # duration of rainfall, s
run_time = 480.0 # duration of run, s
dt = 10.0 # time-step size, s
dem_filename = "../hugo_site_filled.asc"
# Derived parameters
num_steps = int(run_time / dt)
# set up arrays to hold discharge and time
time_since_storm_start = np.arange(0.0, dt * (2 * num_steps + 1), dt)
discharge = np.zeros(2 * num_steps + 1)
# -
# Read an example digital elevation model (DEM) into a Landlab grid and set up the boundaries so that water can only exit out the right edge, representing the watershed outlet.
# +
# Read the DEM file as a grid with a 'topographic__elevation' field
(grid, elev) = read_esri_ascii(dem_filename, name="topographic__elevation")
# Configure the boundaries: valid right-edge nodes will be open;
# all NODATA (= -9999) nodes will be closed.
grid.status_at_node[grid.nodes_at_right_edge] = grid.BC_NODE_IS_FIXED_VALUE
grid.status_at_node[np.isclose(elev, -9999.0)] = grid.BC_NODE_IS_CLOSED
# -
# Now we'll calculate the slope vector components and magnitude, and plot the vectors as quivers on top of a shaded image of the topography:
slp_mag, slp_x, slp_y = slope_magnitude_at_node(grid, elev)
imshow_grid(grid, elev)
plt.quiver(grid.x_of_node, grid.y_of_node, slp_x, slp_y)
# Let's take a look at the slope magnitudes:
imshow_grid(grid, slp_mag, colorbar_label="Slope gradient (m/m)")
# Now we're ready to instantiate a `KinwaveImplicitOverlandFlow` component, with a specified runoff rate and roughness:
# Instantiate the component
olflow = KinwaveImplicitOverlandFlow(
grid, runoff_rate=R, roughness=n, depth_exp=dep_exp
)
# The `DepthSlopeProductErosion` component requires there to be a field called `slope_magnitude` that contains our slope-gradient values, so we will we will create this field and assign `slp_mag` to it (the `clobber` keyword says it's ok to overwrite this field if it already exists, which prevents generating an error message if you run this cell more than once):
grid.add_field("slope_magnitude", slp_mag, at="node", clobber=True)
# Now we're ready to instantiate a `DepthSlopeProductErosion` component:
dspe = DepthSlopeProductErosion(grid, k_e=k_e, tau_crit=tau_c, slope="slope_magnitude")
# Next, we'll make a copy of the starting terrain for later comparison, then run overland flow and erosion:
# +
starting_elev = elev.copy()
for i in range(num_steps):
olflow.run_one_step(dt)
dspe.run_one_step(dt)
slp_mag[:], slp_x, slp_y = slope_magnitude_at_node(grid, elev)
# -
# We can visualize the instantaneous erosion rate at the end of the run, in m/s:
imshow_grid(grid, dspe._E, colorbar_label="erosion rate (m/s)")
# We can also inspect the cumulative erosion during the event by differencing the before and after terrain:
imshow_grid(grid, starting_elev - elev, colorbar_label="cumulative erosion (m)")
# Note that because this is a bumpy DEM, much of the erosion has occurred on (probably digital) steps in the channels. But we can see some erosion across the slopes as well.
# ## `DetachmentLtdErosion`
#
# This component is similar to `DepthSlopeProductErosion` except that it calculates erosion rate from discharge and slope rather than depth and slope. The vertical incision rate, $I$ (equivalent to $E$ in the above; here we are following the notation in the component's documentation) is:
#
# $$I = K Q^m S^n - I_c$$
#
# where $K$ is an erodibility coefficient (with dimensions of velocity per discharge$^m$; specified by parameter `K_sp`), $Q$ is volumetric discharge, $I_c$ is a threshold with dimensions of velocity, and $m$ and $n$ are exponents. (In the erosion literature, the exponents are sometimes treated as empirical parameters, and sometimes set to particular values on theoretical grounds; here we'll just set them to unity.)
#
# The component uses the fields `surface_water__discharge` and `topographic__slope` for $Q$ and $S$, respectively. The component will modify the `topographic__elevation` field accordingly. If the user wishes to apply material uplift relative to baselevel, an `uplift_rate` parameter can be passed on initialization.
#
# Here are the header and constructor docstrings:
# +
from landlab.components import DetachmentLtdErosion
print(DetachmentLtdErosion.__doc__)
# -
print(DetachmentLtdErosion.__init__.__doc__)
# The example below uses the same approach as the previous example, but now using `DetachmentLtdErosion`. Note that the value for parameter $K$ (`K_sp`) is just a guess. Use of exponents $m=n=1$ implies the use of total stream power.
# +
# Process parameters
n = 0.1 # roughness coefficient, (s/m^(1/3))
dep_exp = 5.0 / 3.0 # depth exponent
R = 72.0 # runoff rate, mm/hr
K_sp = 1.0e-7 # erosion coefficient (m/s)/(m3/s)
m_sp = 1.0 # discharge exponent
n_sp = 1.0 # slope exponent
I_c = 0.0001 # erosion threshold, m/s
# Run-control parameters
rain_duration = 240.0 # duration of rainfall, s
run_time = 480.0 # duration of run, s
dt = 10.0 # time-step size, s
dem_filename = "../hugo_site_filled.asc"
# Derived parameters
num_steps = int(run_time / dt)
# set up arrays to hold discharge and time
time_since_storm_start = np.arange(0.0, dt * (2 * num_steps + 1), dt)
discharge = np.zeros(2 * num_steps + 1)
# +
# Read the DEM file as a grid with a 'topographic__elevation' field
(grid, elev) = read_esri_ascii(dem_filename, name="topographic__elevation")
# Configure the boundaries: valid right-edge nodes will be open;
# all NODATA (= -9999) nodes will be closed.
grid.status_at_node[grid.nodes_at_right_edge] = grid.BC_NODE_IS_FIXED_VALUE
grid.status_at_node[np.isclose(elev, -9999.0)] = grid.BC_NODE_IS_CLOSED
# -
slp_mag, slp_x, slp_y = slope_magnitude_at_node(grid, elev)
grid.add_field("topographic__slope", slp_mag, at="node", clobber=True)
# Instantiate the component
olflow = KinwaveImplicitOverlandFlow(
grid, runoff_rate=R, roughness=n, depth_exp=dep_exp
)
dle = DetachmentLtdErosion(
grid, K_sp=K_sp, m_sp=m_sp, n_sp=n_sp, entrainment_threshold=I_c
)
# +
starting_elev = elev.copy()
for i in range(num_steps):
olflow.run_one_step(dt)
dle.run_one_step(dt)
slp_mag[:], slp_x, slp_y = slope_magnitude_at_node(grid, elev)
# -
imshow_grid(grid, starting_elev - elev, colorbar_label="cumulative erosion (m)")
# <hr>
# <small>For more Landlab tutorials, click here: <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html</a></small>
# <hr>
| notebooks/tutorials/overland_flow/overland_flow_erosion/ol_flow_erosion_components.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
# -
from get_aq_data import get_flo_data_new, ID_to_name, TICKS_TWO_HOURLY, annotate_boxplot
data, hourly_mean, daily_mean = get_flo_data_new()
lockdown = daily_mean['2020-03-23':'2020-06-01'].mean(axis=1)
nonlockdown = daily_mean['2019-03-23':'2019-06-01'].mean(axis=1)
lockdown.index = ((lockdown.index-lockdown.index[0]) / pd.Timedelta(1.0,unit='D')) + 1
nonlockdown.index = ((nonlockdown.index-nonlockdown.index[0]) / pd.Timedelta(1.0,unit='D')) + 1
ax = lockdown.plot(figsize=(10, 6), label="2020 (Lockdown)")
nonlockdown.plot(ax=ax, label="2019 (No Lockdown)")
plt.legend()
plt.xlabel("Days since 23rd March")
plt.ylabel(r'$\mathrm{PM}_{2.5}$ ($\mu g / m^3$)')
plt.grid()
plt.tight_layout()
plt.title("Lockdown comparison of daily mean $\mathrm{PM}_{2.5}$")
plt.savefig("graphs2020/Lockdown_Comparison_Timeseries.png", dpi=300)
lockdown = data['2020-03-23':'2020-06-01'].mean(axis=1)
nonlockdown = data['2019-03-23':'2019-06-01'].mean(axis=1)
plt.figure(figsize=(10, 6))
bpdict = plt.boxplot([lockdown, nonlockdown], labels=["2020 (Lockdown)", "2019 (No Lockdown)"], showfliers=False)
annotate_boxplot(bpdict, x_offset=0.07)
figsize=(10, 6)
plt.grid()
plt.tight_layout()
plt.savefig("graphs2020/Lockdown_Comparison_Boxplot.png", dpi=300)
stats = pd.DataFrame({'lockdown': lockdown.describe(percentiles=[0.05, 0.25, 0.5, 0.75, 0.95]), 'non-lockdown': nonlockdown.describe(percentiles=[0.05, 0.25, 0.5, 0.75, 0.95])})
stats.drop(["std", "count"], axis=0, inplace=True)
stats.to_csv('graphs2020/Lockdown_Comparison_Stats.csv')
stats['lockdown'] - stats['non-lockdown']
| New2020 - Plot Lockdown.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assessment - Object-oriented programming
# In this exercise, we'll create a few classes to simulate a server that's taking connections from the outside and then a load balancer that ensures that there are enough servers to serve those connections.
# <br><br>
# To represent the servers that are taking care of the connections, we'll use a Server class. Each connection is represented by an id, that could, for example, be the IP address of the computer connecting to the server. For our simulation, each connection creates a random amount of load in the server, between 1 and 10.
# <br><br>
# Run the following code that defines this Server class.
# +
#Begin Portion 1#
import random
class Server:
def __init__(self):
"""Creates a new server instance, with no active connections."""
self.connections = {}
def add_connection(self, connection_id):
"""Adds a new connection to this server."""
connection_load = random.random()*10+1
# Add the connection to the dictionary with the calculated load
self.connections[connection_id] = connection_load
def close_connection(self, connection_id):
"""Closes a connection on this server."""
# Remove the connection from the dictionary
if connection_id in self.connections:
del self.connections[connection_id]
def load(self):
"""Calculates the current load for all connections."""
total = 0
for load in self.connections.values():
total += load
# Add up the load for each of the connections
return total
def __str__(self):
"""Returns a string with the current load of the server"""
return "{:.2f}%".format(self.load())
#End Portion 1#
# -
# Now run the following cell to create a Server instance and add a connection to it, then check the load:
# +
server = Server()
server.add_connection("192.168.1.1")
print(server.load())
# -
# After running the above code cell, if you get a **<font color =red>NameError</font>** message, be sure to run the Server class definition code block first.
#
# The output should be 0. This is because some things are missing from the Server class. So, you'll need to go back and fill in the blanks to make it behave properly.
# <br><br>
# Go back to the Server class definition and fill in the missing parts for the `add_connection` and `load` methods to make the cell above print a number different than zero. As the load is calculated randomly, this number should be different each time the code is executed.
# <br><br>
# **Hint:** Recall that you can iterate through the values of your connections dictionary just as you would any sequence.
# Great! If your output is a random number between 1 and 10, you have successfully coded the `add_connection` and `load` methods of the Server class. Well done!
# <br><br>
# What about closing a connection? Right now the `close_connection` method doesn't do anything. Go back to the Server class definition and fill in the missing code for the `close_connection` method to make the following code work correctly:
server.close_connection("192.168.1.1")
print(server.load())
# You have successfully coded the `close_connection` method if the cell above prints 0.
# <br><br>
# **Hint:** Remember that `del` dictionary[key] removes the item with key *key* from the dictionary.
# Alright, we now have a basic implementation of the server class. Let's look at the basic LoadBalancing class. This class will start with only one server available. When a connection gets added, it will randomly select a server to serve that connection, and then pass on the connection to the server. The LoadBalancing class also needs to keep track of the ongoing connections to be able to close them. This is the basic structure:
#Begin Portion 2#
class LoadBalancing:
def __init__(self):
"""Initialize the load balancing system with one server"""
self.connections = {}
self.servers = [Server()]
def add_connection(self, connection_id):
"""Randomly selects a server and adds a connection to it."""
server = random.choice(self.servers)
# Add the connection to the dictionary with the selected server
# Add the connection to the server
server.add_connection(connection_id)
self.ensure_availability()
def close_connection(self, connection_id):
"""Closes the connection on the the server corresponding to connection_id."""
# Find out the right server
# Close the connection on the server
# Remove the connection from the load balancer
for server in self.servers:
if connection_id in server.connections:
server.close_connection(connection_id)
break
def avg_load(self):
"""Calculates the average load of all servers"""
# Sum the load of each server and divide by the amount of servers
total_load = 0
total_server = 0
for server in self.servers:
total_load += server.load()
total_server += 1
return total_load/total_server
def ensure_availability(self):
"""If the average load is higher than 50, spin up a new server"""
if self.avg_load() > 50:
self.servers.append(Server())
def __str__(self):
"""Returns a string with the load for each server."""
loads = [str(server) for server in self.servers]
return "[{}]".format(",".join(loads))
#End Portion 2#
# As with the Server class, this class is currently incomplete. You need to fill in the gaps to make it work correctly. For example, this snippet should create a connection in the load balancer, assign it to a running server and then the load should be more than zero:
l = LoadBalancing()
l.add_connection("fdca:fd00:c2b6:b24b:be67:2827:688d:e6a1:6a3b")
print(l.avg_load())
# After running the above code, the output is 0. Fill in the missing parts for the `add_connection` and `avg_load` methods of the LoadBalancing class to make this print the right load. Be sure that the load balancer now has an average load more than 0 before proceeding.
# What if we add a new server?
l.servers.append(Server())
print(l.avg_load())
# The average load should now be half of what it was before. If it's not, make sure you correctly fill in the missing gaps for the `add_connection` and `avg_load` methods so that this code works correctly.
# <br><br>
# **Hint:** You can iterate through the all servers in the *self.servers* list to get the total server load amount and then divide by the length of the *self.servers* list to compute the average load amount.
# Fantastic! Now what about closing the connection?
l.close_connection("fdca:fd00:c2b6:b24b:be67:2827:688d:e6a1:6a3b")
print(l.avg_load())
# Fill in the code of the LoadBalancing class to make the load go back to zero once the connection is closed.
# <br><br>
# Great job! Before, we added a server manually. But we want this to happen automatically when the average load is more than 50%. To make this possible, fill in the missing code for the `ensure_availability` method and call it from the `add_connection` method after a connection has been added. You can test it with the following code:
for connection in range(20):
l.add_connection(connection)
print(l)
# The code above adds 20 new connections and then prints the loads for each server in the load balancer. If you coded correctly, new servers should have been added automatically to ensure that the average load of all servers is not more than 50%.
# <br><br>
# Run the following code to verify that the average load of the load balancer is not more than 50%.
print(l.avg_load())
# Awesome! If the average load is indeed less than 50%, you are all done with this assessment.
| Week5_C1M5_Object_Oriented_Programming_V7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pathlib
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
from tensorflow.keras.layers.experimental import preprocessing
from tensorflow.keras import layers
from tensorflow.keras import models
from IPython import display
import pyaudio
# -
model = tf.keras.models.load_model('simple_audio_model.sav')
print('loaded saved model.')
print(model.summary())
# ## Get Audio Input
# get pyaudio input device
def getInputDevice(p):
index = None
nDevices = p.get_device_count()
print('Found %d devices:' % nDevices)
for i in range(nDevices):
deviceInfo = p.get_device_info_by_index(i)
#print(deviceInfo)
devName = deviceInfo['name']
print(devName)
# look for the "input" keyword
# choose the first such device as input
# change this loop to modify this behavior
# maybe you want "mic"?
if not index:
if 'input' in devName.lower():
index = i
# print out chosen device
if index is not None:
devName = p.get_device_info_by_index(index)["name"]
#print("Input device chosen: %s" % devName)
return index
# initialize pyaudio
p = pyaudio.PyAudio()
getInputDevice(p)
# Now let's try plotting 1 second of Mic Input
# +
def get_spectrogram(waveform):
# Padding for files with less than 16000 samples
zero_padding = tf.zeros([16000] - tf.shape(waveform), dtype=tf.float32)
# Concatenate audio with padding so that all audio clips will be of the
# same length
waveform = tf.cast(waveform, tf.float32)
equal_length = tf.concat([waveform, zero_padding], 0)
spectrogram = tf.signal.stft(
equal_length, frame_length=255, frame_step=128)
spectrogram = tf.abs(spectrogram)
return spectrogram
def plot_spectrogram(spectrogram, ax):
# Convert to frequencies to log scale and transpose so that the time is
# represented in the x-axis (columns).
log_spec = np.log(spectrogram.T)
height = log_spec.shape[0]
X = np.arange(16000, step=height + 1)
Y = range(height)
ax.pcolormesh(X, Y, log_spec)
# +
# set sample rate
NSEC = 1
sampleRate = 16000 # #48000
sampleLen = NSEC*sampleRate
print('opening stream...')
stream = p.open(format = pyaudio.paInt16,
channels = 1,
rate = sampleRate,
input = True,
frames_per_buffer = 4096,
input_device_index = -1)
# read a chunk of data - discard first
data = stream.read(sampleLen)
print(type(data))
p.close(stream)
waveform = tf.cast(tf.io.decode_raw(data, "int16"), "float32")/32768.0
print(waveform)
spectrogram = get_spectrogram(waveform)
#spectrogram = tf.reshape(spectrogram, (spectrogram.shape[0], spectrogram.shape[1], 1))
print(spectrogram.shape)
fig, axes = plt.subplots(2, figsize=(12, 8))
timescale = np.arange(waveform.shape[0])
axes[0].plot(timescale, waveform.numpy())
axes[0].set_title('Waveform')
axes[0].set_xlim([0, 16000])
axes[0].set_ylim([-1, 1])
plot_spectrogram(spectrogram.numpy(), axes[1])
axes[1].set_title('Spectrogram')
plt.show()
# +
commands = ['go', 'down', 'up', 'stop', 'yes', 'left', 'right', 'no']
print(spectrogram.shape)
spectrogram1= tf.reshape(spectrogram, (-1, spectrogram.shape[0], spectrogram.shape[1], 1))
print(spectrogram1.shape)
prediction = model(spectrogram1)
print(prediction)
sm = tf.nn.softmax(prediction[0])
am = tf.math.argmax(sm)
print(sm)
print(commands[am])
#plt.bar(commands, tf.nn.softmax(prediction[0]))
#plt.title(f'Predictions for "{commands[label[0]]}"')
#plt.show()
# -
| cooker_whistle/simple_audio_mic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"source_hidden": true}
import json
import svgling
from svgling.figure import Caption, SideBySide, RowByRow
from IRVVisualisationUtils import treeListToTuple, parseAssertions, printAssertions, buildRemainingTreeAsLists, buildPrintedResults, printTrees
a_file = open("../AssertionJSON/SF2019Nov8Assertions.json")
auditfile = json.load(a_file)
(apparentWinner, apparentNonWinners, WOLosers,IRVElims) = parseAssertions(auditfile)
elimTrees = buildPrintedResults(apparentWinner, apparentNonWinners, WOLosers,IRVElims)
print("Built "+str(len(elimTrees))+" trees to visualise excluded alternate winners.")
# -
# # RAIRE example assertion parser and visualizer
#
# This notebook parses and visualizes RAIRE assertions.
# Right now it's hardcoded to read RAIRE_sample_audit1.json, but you can change that.
# Start by executing the rectangle above to understand the election and the apparent winner.
# The audit needs to exclude all the other possible winners, though we don't care about other elimination orders in which the apparent winner still wins.
# Execute the next code snippet to see the trees of possible alternative elimination orders.
# Each tree will be pruned according to RAIRE's assertions, with each pruned branch tagged with the assertion that allowed us to exclude it.
# You (the auditor) need to check that all the leaves end in an assertion, which shows that they have been excluded.
#
# + jupyter={"source_hidden": true}
Caption(printTrees(elimTrees),"Trees showing how other winners are excluded.")
# -
# Now print all the assertions. This gives you an explanation of the meaning of each one.
#
# + jupyter={"source_hidden": true}
printAssertions(WOLosers,IRVElims)
# -
# Now the audit begins! We now apply a Risk Limiting Audit to test each of the assertions above.
# For each assertion, we consider the opposite hypothesis, that candidate C *can* be eliminated at that point. We then try to audit until that hypothesis can be rejected. If all the hypotheses are rejected, the election result is declared correct. At any time, if the audit has failed to reject all the hypotheses, a full manual recount can be conducted.
#
| Code/.ipynb_checkpoints/RAIREExampleDataParsing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reverse a String
#
# This interview question requires you to reverse a string using recursion. Make sure to think of the base case here.
#
# Again, make sure you use *recursion* to accomplish this. **Do not slice (e.g. string[::-1]) or use iteration, there must be a recursive call for the function.**
#
# ____
#
# ### Fill out your solution below
def reverse(s):
if(len(s)<=1):
return s
else:
m = int(len(s)/2)
return reverse(s[m:]) + (reverse((s[:m])))
pass
reverse('hello world')
# # Test Your Solution
#
# Run the cell below to test your solution against the following cases:
#
# string = 'hello'
# string = 'hello world'
# string = '123456789'
# +
'''
RUN THIS CELL TO TEST YOUR FUNCTION AGAINST SOME TEST CASES
'''
from nose.tools import assert_equal
class TestReverse(object):
def test_rev(self,solution):
assert_equal(solution('hello'),'olleh')
assert_equal(solution('hello world'),'dlrow olleh')
assert_equal(solution('123456789'),'987654321')
print ('PASSED ALL TEST CASES!')
# Run Tests
test = TestReverse()
test.test_rev(reverse)
# -
# **Good Luck!**
| code/algorithms/course_udemy_1/Recursion/Recursion Interview Problems/Recursion Problems - PRACTICE/Recursion Problem 1 - Reverse String .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# Ricordati di eseguire questa cella con Control+Invio
import sys;
sys.path.append('../');
import jupman;
# + [markdown] colab_type="text" id="dW_7xwCZAFP5"
# # Introduzione rapida a Python
#
# ## [Scarica zip esercizi](../_static/generated/quick-intro.zip)
#
# [Naviga file online](https://github.com/DavidLeoni/softpython-it/tree/master/quick-intro)
#
# <div class="alert alert-warning">
#
# **REQUISITI:**
#
# * **QUESTO FOGLIO E' PER CHI HA GIA' COMPETENZE DI PROGRAMMAZIONE**, e in 3-4h vuole farsi rapidamente un'idea di Python
# * **Aver installati Python 3 e Jupyter:** se non hai già provveduto, guarda [Installazione](https://it.softpython.org/installation.html)
# </div>
#
# <div class="alert alert-warning">
#
# **SE SEI UN PRINCIPIANTE**: Salta questo foglio e fai invece i tutorial che trovi nella sezione [Fondamenti](https://it.softpython.org/index.html#A---Fondamenti), a partire da [Strumenti e script](https://it.softpython.org/tools/tools-sol.html)
# </div>
#
#
# -
#
# ### Che fare
#
# - scompatta lo zip in una cartella, dovresti ottenere qualcosa del genere:
#
# ```
#
# quick-intro
# quick-intro.ipynb
# quick-intro-sol.ipynb
# jupman.py
# ```
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Per essere visualizzato correttamente, il file del notebook DEVE essere nella cartella szippata.
# </div>
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: In questo libro usiamo **SOLO PYTHON 3** <br/>
#
# Se per caso ottieni comportamenti inattesi, controlla di usare Python 3 e non il 2. Se per caso il tuo sistema operativo scrivendo `python` fa partire il 2, prova ad eseguire il tre scrivendo il comando: `python3`
# </div>
#
#
# - apri il Jupyter Notebook da quella cartella. Due cose dovrebbero aprirsi, prima una console e poi un browser. Il browser dovrebbe mostrare una lista di file: naviga la lista e apri il notebook `quick-intro.ipynb`
# - Prosegui leggendo il file degli esercizi, ogni tanto al suo interno troverai delle scritte **ESERCIZIO**, che ti chiederanno di scrivere dei comandi Python nelle celle successive. Gli esercizi sono graduati per difficoltà, da una stellina ✪ a quattro ✪✪✪✪
#
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Ricordati di eseguire sempre la prima cella dentro il notebook. Contiene delle istruzioni come `import jupman` che dicono a Python quali moduli servono e dove trovarli. Per eseguirla, vedi le seguenti scorciatoie
# </div>
#
#
#
# Scorciatoie da tastiera:
#
# * Per eseguire il codice Python dentro una cella di Jupyter, premi `Control+Invio`
# * Per eseguire il codice Python dentro una cella di Jupyter E selezionare la cella seguente, premi `Shift+Invio`
# * Per eseguire il codice Python dentro una cella di Jupyter E creare una nuova cella subito dopo, premi `Alt+Invio`
# * Se per caso il Notebook sembra inchiodato, prova a selezionare `Kernel -> Restart`
#
#
#
#
# ## Proviamo Jupyter
#
# Vediamo brevemente come funzionano i fogli Jupyter.
# **ESERCIZIO**: Proviamo a inserire un comando Python: scrivi nella cella qua sotto `3 + 5`, e poi mentre sei in quella cella premi i tasti speciali `Control+Invio`. Come risultato, dovresti vedere apparire il numero 8
# **ESERCIZIO**: in Python possiamo scrivere commenti iniziando una riga con un cancelletto `#`. Come prima, scrivi nella cella sotto `3 + 5` ma questa volta scrivilo nella riga sotto la scritta `# scrivi qui`:
# scrivi qui
# **ESERCIZIO**: Jupyter per ogni cella mostra il risultato solo dell'ultima riga eseguita in quella cella. Prova a inserire questo codice nella cella sotto ed esegui premendo `Control+Invio`. Che risultato appare?
#
# ```python
# 3 + 5
# 1 + 1
# ```
# +
# scrivi qui
# -
# **ESERCIZIO**: Proviamo adesso a creare noi una nuova cella.
#
# * Mentre sei con il cursore in questa cella, premi `Alt+Invio`. Si dovrebbe creare una nuova cella dopo la presente.
#
# * Nella cella appena creata, inserisci `2 + 3` e poi premi `Shift+Invio`. Cosa succede al cursore? prova la differenze con `Control+Invio`. Se non capisci la differenza, prova a premere ripetutamente `Shif+Invio` e vedi che succede.
# + [markdown] colab_type="text" id="vEzZXT8MAFP9"
#
# + [markdown] colab_type="text" id="uZyXs9KLAFQB"
# ## Principali tipi di dati Python
#
# Dato che il tema del corso è il trattamento dei dati, per cominciare ci focalizzeremo sui tipi di dati in Python.
#
# **Riferimenti**:
#
# - [Pensare in Python, Capitolo 1, Lo scopo del programma](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2002.html)
# - [Pensare in Python, Capitolo 2, Variabili, istruzioni ed espressioni](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2003.html)
#
#
# Quando leggiamo delle informazioni da una fonte esterna come un file, poi dovremo inevitabilmente incastonare i dati letti in qualche combinazione di questi tipi:
#
# + [markdown] colab_type="text" id="1ecI-hRCAFQG"
#
# | Tipo | Esempio 1 | Esempio 2 | Esempio 3|
# |--------|-----|---|----|
# |int| `0`|`3`| `-5`|
# |numero in virgola mobile float| `0.0`| `3.7` | `-2.3`|
# |bool|False|True||
# |string |`""`| `"Buon giorno"`|`'come stai?'`|
# |list | `[]`|`[5, 8, 10]`|[`"qualcosa", 5, "altro ancora"]`|
# | dict |`{}`|`{'chiave 1':'valore 1', 'chiave 2':'valore 2'}`|`{5:'un valore stringa', 'chiave stringa':7}`|
#
#
# + [markdown] colab_type="text" id="GBvgbl0xAFQL"
# A volte, useremo tipi più complessi, per esempio i valori temporali si potrebbero mettere in oggetti di tipo `datetime` che oltre alla data vera e propria possono contenere anche il fuso orario.
#
# In quel che segue, forniremo alcuni esempi rapidi su quello che si può fare sui vari tipi di dato, mettendo i riferimenti alle spiegazioni più dettagliate dal libro Pensare in Python.
#
# + [markdown] colab_type="text" id="p_V71t9BAFQQ"
# ## Numeri interi e con virgola
#
# Nel libro 'Pensare in Python' non c'è un vero e proprio capitolo dedicato ai calcoli numerici, che sono sparse nei primi capitoli segnalati prima. Metto qua due note velocissime:
#
#
# In Python abbiamo numeri interi:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="--cJxUntAFQT" outputId="6a261e4f-6f7f-481d-9f4e-02f951c18010"
3 + 5
# + [markdown] colab_type="text" id="rEV6BEORAFQo"
# La somma tra interi ovviamente ci da un intero:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="Vk7aiqO5AFQr" outputId="6408cc81-e7db-4778-c929-c42fdf178b0f"
type(8)
# + [markdown] colab_type="text" id="Z49LC8g3AFRQ"
# E se dividiamo interi? Ci troveremo con il tipo in virgola mobile _float_:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="fGqrVdg6AFRT" outputId="ead8a066-1d37-4ccb-8cf5-d7bf6bb7b5ad"
3 / 4
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="AuREnwBmAFRn" outputId="96b737a9-d30f-4696-9805-8974a1b731d3"
type(0.75)
# + [markdown] colab_type="text" id="DIcwRoa7AFR4"
# <div class="alert alert-warning">
#
# **ATTENZIONE al punto !**
#
# In Python e in molti formati dati, al posto della nostra virgola si usa il formato inglese con il punto '.' <br/>
# </div>
# -
# **✪ ESERCIZIO**: Prova a scrivere qua sotto `3.14` con il punto, e poi `3,14` con la virgola ed esegui con Ctrl+Invio. Cosa appare nei due casi?
# +
# scrivi qui con il punto
# +
# scrivi qui con la virgola
# + [markdown] colab_type="text" id="ivjG8uzWAFR7"
# **✪ ESERCIZIO**: Prova a scrivere qua sotto `3 + 1.0` ed esegui con Ctrl+Invio. Di quale tipo sarà il risultato? Controlla anche usando il comando `type`.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="IsFJ8O-IAFR8"
# scrivi qui i comandi
# + [markdown] colab_type="text" id="22kAoaiYAFSD"
# **✪ ESERCIZIO**: Qualche professore di matematica ti avrà sicuramente intimato di non dividere mai per zero. Neanche a Python piace molto, prova a scrivere nella cella qua sotto `1 / 0` e poi premi Ctrl+Invio per eseguire la cella. Nota come Python riporterà la riga dove è accaduto l'errore:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="UfI3UixcAFSE"
# scrivi qui sotto il codice
# + [markdown] colab_type="text" id="WMaObpo6AFSP"
#
# ## Booleani - bool
#
# I booleani rappresentano valori veri e falsi, e si possono usare per verificare quando certe condizioni avvengono.
#
# **Riferimenti**
#
# * Softpython: [Basi - booleani](https://it.softpython.org/basics/basics-sol.html#Booleani)
# - Softpython: [controllo di flusso - if](https://it.softpython.org/control-flow/flow1-if-sol.html)
# -
# Per indicare i booleani Python ci fornisce due costanti `True` e `False`. Cosa ci possiamo fare?
#
#
# ### and
#
# Potremmo usarle per segnare in variabili se un certo fatto è avvenuto, per esempio possiamo fare un programma che al mattino ci dice che possiamo uscire di casa solo dopo aver fatto entrambe (`and`) colazione e e lavato i denti:
# +
fatto_colazione = True
lavato_denti = True
if fatto_colazione and lavato_denti:
print("fatto tutto !")
print("posso uscire di casa")
else:
print("NON posso uscire di casa")
# -
# **✪ ESERCIZIO**: prova a scrivere qui sotto a mano il programma riportato nella cella precedente, ed eseguilo con Control+Invio. Prova a cambiare i valori da `True` a `False` e guarda che succede.
#
# Assicurati di provare tutti i casi:
#
# - True True
# - True False
# - False True
# - False False
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Ricordati i `:` alla fine della riga con `if` !!!!
#
# </div>
# +
# scrivi qui
# -
# Si può anche mettere un `if` dentro l'altro (_nested if_). Per esempio, questo programma qui funziona esattamente come il precedente:
# +
fatto_colazione = True
lavato_denti = True
if fatto_colazione:
if lavato_denti: # NOTA: Questo blocco if è indentato rispetto
print("fatto tutto !") # all' if fatto_colazione
print("posso uscire di casa!") #
else:
print("NON posso uscire di casa")
else:
print("NON posso uscire di casa")
# -
# **✪ ESERCIZIO**: Prova a modificare il programma riportato nella cella precedente per fargli riportare lo stato delle varie azioni compiute. Elenchiamo qui i possibili casi e i risultati attesi:
#
# - True False
#
# ```
# ho fatto colazione
# non ho lavato i denti
# NON posso uscire di casa
# ```
#
# - False True
# - False False
#
# ```
# non ho fatto colazione
# NON posso uscire di casa
# ```
#
# - True True
# ```
# ho fatto colazione
# ho lavato i denti
# fatto tutto !
# posso uscire di casa!
# ```
#
#
#
#
#
# +
# scrivi qui
fatto_colazione = True
lavato_denti = True
if fatto_colazione:
print("ho fatto colazione")
if lavato_denti: # NOTA: Questo blocco if è indentato rispetto
print("ho lavato i denti") #
print("fatto tutto !") # all' if fatto_colazione
print("posso uscire di casa!") #
else:
print("non ho lavato i denti")
print("NON posso uscire di casa")
else:
print("non ho fatto colazione")
print("NON posso uscire di casa")
# -
# ### or
#
# Per verificare se almeno di due condizioni si è verificata, si usa `or`. Per esempio, possiamo stabilire che per fare colazione ci serve avere del latte intero o scremato (nota: se li abbiamo tutte e due riusciamo a fare colazione lo stesso !)
#
# +
ho_latte_intero = True
ho_latte_scremato = False
if ho_latte_intero or ho_latte_scremato:
print("posso fare colazione !")
else:
print("NON posso fare colazione :-(")
# -
# **✪ ESERCIZIO**: prova a scriverecqui sotto a mano il programma riportato nella cella presedente, ed eseguilo con Control+Invio. Prova a cambiare i valori da `True` a `False` e guarda che succede:
#
# Assicurati di provare tutti i casi:
#
# - True True
# - True False
# - False True
# - False False
# +
# scrivi qui
# -
# **✪✪ ESERCIZIO**: prova a fare un programma che ti dice se puoi uscire di casa solo se hai fatto colazione (per cui devi avere hai almeno un tipo di latte) e lavato i denti
# +
ho_latte_intero = False
ho_latte_scremato = True
lavato_denti = False
# scrivi qui
if ho_latte_intero or ho_latte_scremato:
print("posso fare colazione !")
fatto_colazione = True
else:
print("NON posso fare colazione :-(")
fatto_colazione = False
if fatto_colazione and lavato_denti:
print("posso uscire di casa")
else:
print("NON posso uscire di casa")
# -
# ### not
#
# Per le negazioni, puoi usare il `not`:
#
not True
not False
# +
fatto_colazione = False
if not fatto_colazione:
print("Ho fame !")
else:
print("che buoni che erano i cereali")
# +
fatto_colazione = True
if not fatto_colazione:
print("Ho fame !")
else:
print("che buoni che erano i cereali")
# -
# ✪✪ ESERCIZIO: prova a fare un programma che ti dice se puoi nuotare se NON hai fatto colazione E hai il salvagente
#
# Assicurati di provare tutti i casi:
#
# - True True
# - True False
# - False True
# - False False
# +
hai_salvagente = True
fatto_colazione = True
# scrivi qui
if hai_salvagente and not fatto_colazione:
print("puoi nuotare")
else:
print("NON puoi nuotare")
# -
# ### Non solo True e False
# + [markdown] colab_type="text" id="Wtyg2lXuAFSU"
# <div class="alert alert-warning">
#
# **ATTENZIONE ai booleani diversi da True e False !**
#
# In Python, il numero `0` e altri oggetti 'nulli' (come l'oggetto `None`, la stringa vuota `""` e la lista vuota `[]`) sono considerati `False`, e tutto ciò che non è 'nullo' è considerato `True`!
# </div>
# + [markdown] colab_type="text" id="BT44SZeQAFSX"
# Facciamo degli esempi per mostrare quanto sopra riportato:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="VVd4Ikp3AFSY"
if True:
print("questo")
print("sarà")
print("stampato")
else:
print("quest'altro")
print("non sarà stampato")
# + [markdown] colab_type="text" id="AlntXiK3AFSc"
# Tutto ciò che non è 'nullo' è considerato `True`, verifichiamolo per esempio con la stringa `"ciao"`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="6GWu1jIUAFSd"
if "ciao":
print("anche questo")
print("sarà stampato!!")
else:
print("e questo no")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="HsEUjY8sAFSs"
if False:
print("io non sarò stampato")
else:
print("io sì")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="DG7nyxf5AFS8"
if 0:
print("anche questo non sarà stampato")
else:
print("io sì")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="bRhHcpEjAFTE"
if None:
print("neppure questo sarà stampato")
else:
print("io sì")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="5mo5mXdbAFTN" outputId="1991e1dd-4cdc-41f3-cbfc-53cafff54fe0"
if "": # stringa vuota
print("Neanche questo sarà stampato !!!")
else:
print("io sì")
# + [markdown] colab_type="text" id="FgE9xpe6AFTZ"
# **✪ ESERCIZIO**: Copia qua sotto l'`if` una stringa con uno spazio dentro `" "`nella condizione dell'`if`. Cosa succederà?
#
# - prova anche a mettere una lista vuota `[]`, che succede?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="81YonZhpAFTa"
# scrivi qui l'if
if " ": # spazio
print("Niente stampa!")
else:
print("io sì")
if []: # lista vuota
print("Niente stampa!")
else:
print("io sì")
# + [markdown] colab_type="text" id="AUCCneEoAFTw"
# ## Stringhe - string
#
# Le stringhe sono sequenze _immutabili_ di caratteri.
#
# **Riferimenti**:
#
# SoftPython:
#
# - [stringhe 1 - introduzione](https://it.softpython.org/strings/strings1-sol.html)
# - [stringhe 2 - operatori](https://it.softpython.org/strings/strings2-sol.html)
# - [stringhe 3 - metodi](https://it.softpython.org/strings/strings3-sol.html)
# - [stringhe 4 - altri esercizi](https://it.softpython.org/strings/strings4-sol.html)
#
# **Concatenare stringhe**
#
# Una delle cose che si fanno più frequentemente è concatenre delle stringhe:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="HUNg73x-AFTx"
"ciao " + "mondo"
# + [markdown] colab_type="text" id="5Dm0z8SOAFT0"
# Ma nota che quando concateniamo una stringa e un numero, Python si arrabbia:
# + [markdown] colab_type="text" id="vEDchuM9AFT2"
# ```python
# "ciao " + 5
# ```
# + [markdown] colab_type="text" id="jKoofPiyAFT3"
# ```bash
# ---------------------------------------------------------------------------
# TypeError Traceback (most recent call last)
# <ipython-input-38-e219e8205f7d> in <module>()
# ----> 1 "ciao " + 5
#
# TypeError: Can't convert 'int' object to str implicitly
# ```
# + [markdown] colab_type="text" id="_dPea14WAFT6"
# Questo succede perchè Python vuole che convertiamo esplicitamente il numero '5' in una stringa. Porrà simili lamentela anche con altri tipi di oggetti. Quindi, quando concatenate oggetti che non sono stringhe, per evitare problemi racchiudete l'oggetto da convertire nella funzione `str` come qui:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="-xndn9-FAFT7"
"ciao " + str(7)
# + [markdown] colab_type="text" id="SdgLz37QAFUC"
# Un modo alternativo e più veloce è usare l'operatore di formattazione percentuale `%`, che sostituisce alle occorrenze di `%s` quello che mettete dopo un `%` dopo la stringa :
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="tkupKFSeAFUD"
"ciao %s" % 7
# + [markdown] colab_type="text" id="ZMIOdHJbAFUK"
# Meglio ancora, il `%s` può stare all'interno della stringa e venire ripetuto. Per ogni occorrenza si può passare un sostituto diverso, come per esempio nella tupla `("bello", "Python")` (una tupla è semplicemente una sequenza immutabile di elementi racchiusi tra parentesi tonde e separati da virgole)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="01RTe3XzAFUM" outputId="c0c70ece-bcd8-482d-8478-1f77d0c056b3"
"Che %s finalmente imparo %s" % ("bello", "Python")
# + [markdown] colab_type="text" id="3ah1CeDPAFUV"
# **✪ ESERCIZIO**: il `%s` funziona con le stringhe ma anche con quasiasi altro tipo di dato, per esempio un intero. Scrivi qua sotto il comando sopra, aggiunendo un `%s` alla fine della stringa, e aggiungendo alla fine della tupla il numero `3` (separandolo dagli altri con una virgola).
#
# Domanda: i `%s` possono stare uno dopo l'altro senza spazi tra di loro? Prova.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="K1pTbCATAFUW"
# scrivi qui
# + [markdown] colab_type="text" id="F5GKXLjqAFYH"
# ### Usare metodi degli oggetti
#
# Quasi tutto in Python è un oggetto, faremo qui una velocissima introduzione tanto per dare l'idea.
#
# **Riferimenti**
#
# - [Pensare in Python, Capitolo 15, Classi e oggetti](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2016.html)
# - [Pensare in Python, Capitolo 16, Classi e funzioni](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2017.html)
# - [Pensare in Python, Capitolo 17, Classi e metodi](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2017.html)
#
# Quasi tutto in Python è un oggetto. Per esempio, le stringhe sono oggetti. Ogni tipo di oggetto ha delle azioni chiamati _metodi_ che si possono eseguire su quello oggetto. Per esempio, per le stringhe che rappresentano nomi potremmo voler rendere in maiuscolo la prima lettera: a tal fine possiamo cercare se per le stringhe ci sono giò metodi esistenti che fanno questo. Proviamo il metodo esistente `capitalize()` sulla _stringa_ `"trento"` (notare che la stringa è tutta in minuscolo e _capital_ in inglese vuol anche dire 'maiuscolo' ):
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="L59SZPKMAFYI" outputId="21f7d2b9-43d6-41a0-c02c-5c1a961f778f"
"trento".capitalize()
# + [markdown] colab_type="text" id="1eyxqR-PAFYU"
# Python ci ha appena fatto la cortesia di rendere la prima lettera della parola in maiuscolo `'Trento'`
#
# + [markdown] colab_type="text" id="YSb82zlxAFYW"
# **✪ ESERCIZIO**: Scrivi nella cella qua sotto `"trento".` e premi TAB: Jupyter dovrebbe suggerirti dei metodi disponibili per la stringa. Prova il metodo `upper()` e `count("t")`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="r5_aFtbdAFYY"
# scrivi qui
# + [markdown] colab_type="text" id="-YkY_K8AAFUc"
# ## Liste - list
#
# Una lista in python è una sequenza di elementi eterogenei, in cui possiamo mettere gli oggetti che vogliamo.
#
# **Riferimenti - SoftPython:**
#
# - [Liste 1 - introduzione](https://it.softpython.org/lists/lists1-sol.html)
# - [Liste 2 - operatori](https://it.softpython.org/lists/lists2-sol.html)
# - [Liste 3 - metodi](https://it.softpython.org/lists/lists3-sol.html)
# - [Liste 4 - iterazione e funzioni](https://it.softpython.org/lists/lists4-sol.html)
#
# Creiamo una lista di stringhe:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Ozn-Qt0yAFUc"
x = ["ciao", "soft", "python"]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="04EWaDNyAFUf" outputId="f478db2d-544a-47bc-e578-b0d5f1b9a3f1"
x
# + [markdown] colab_type="text" id="U6OlmaHaAFUn"
# Le liste sono sequenze di oggetti possibilmente eterogenei, quindi dentro ci potete buttare di tutto, interi, stringhe, dizionari ...:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="YRnNkTnbAFUn"
x = ["ciao", 123, {"a":"b"}]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="YBHGVy51AFUq" outputId="9ba3a0b6-36ca-4642-ac87-41d9f97c066f"
x
# + [markdown] colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="perY24wzAFUx"
# Per accedere ad un elemento in particolare dentro una lista, si può usare un indice tra parentesi quadre che indica un elemento:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="vWOiXQlNAFU0" outputId="ca5fe8cc-0788-4591-d13e-febc5fee6f7c"
# primo elemento
x[0]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="09ciP4W0AFU8" outputId="d7d2eacb-7afd-4e6f-da32-cfd5a12f65b4"
# secondo elemento
x[1]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="pHFc8586AFVD" outputId="400f7957-c1ca-4c5d-c762-809daf0fbe82"
# terzo elemento
x[2]
# + [markdown] colab_type="text" id="A6rRmAJ-AFVL"
# In una lista possiamo cambiare gli elementi con l'assegnazione:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="V2LYVi8CAFVL"
# Cambiamo il _secondo_ elemento:
x[1] = "soft"
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="8vf5xnm8AFVT" outputId="958c853d-3288-4d74-df26-e0998d98c79f"
x
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="QZ0Rg6QdAFVZ"
x[2] = "python"
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="0fh1vyFmAFVc" outputId="b754b8e7-9521-4e34-bf8d-82c58d44209c"
x
# + [markdown] colab_type="text" id="qghMxSoDAFVf"
# Per ottenere la lunghezza di una lista, possiamo usare `len`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="B5qka4xoAFVh" outputId="4b5f0e3c-2749-48ab-ad29-38ad7f8ad638"
x = ["ciao", "soft", "python"]
len(x)
# + [markdown] colab_type="text" id="1oK8VQhqAFVl"
# **✪ ESERCIZIO**: prova ad accedere ad un elemento fuori dalla lista, e vedi che succede.
#
# - `x[3]` è dentro o fuori dalla lista?
# - C'è una qualche lista `x` per cui possiamo scrivere `x[len(x)]` senza problemi ?
# - se usi indici negativi, che succede? Prova -1, -2, -3, -4 ...
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="SP2tU81tAFVm"
# scrivi qui
# + [markdown] colab_type="text" id="ZSY_jqK_AFVv"
# Possiamo aggiungere elementi alla fine di una lista usando il comando `append`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="QxpNlcbHAFVv"
x = []
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="dXKCs033AFVz" outputId="0c1d2151-929e-47ff-efe9-30fa1d805cff"
x
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="w6B-rfZpAFV2"
x.append("ciao")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="M8roNVZPAFV5" outputId="b61ddfa5-115b-43cc-df02-450435f69d10"
x
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="4zXn5AnYAFV9"
x.append("soft")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="1YuQeTO8AFWF" outputId="6e819883-ced0-4382-a24e-c6984a0fe3e7"
x
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="efAjHzyfAFWK"
x.append("python")
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="vSSZvL52AFWO" outputId="54946ba0-ef21-4f9b-ac8f-afddc65bb84f"
x
# + [markdown] colab_type="text" id="TrimbgQ8i6zE"
# ### Ordinamento liste
#
# Le liste possono ordinare comodamente con il metodo `.sort`, che funziona su tutti gli oggetti ordinabili. Per esempio, possiamo ordinare i numeri:
#
# **IMPORTANTE**: `.sort()` modifica la lista su cui è chiamato, _non_ ne genera una nuova !
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="xNFr7Up3jt9l"
x = [ 8, 2, 4]
x.sort()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="bxLaBfwaj3bM"
x
# + [markdown] colab_type="text" id="IQWomkTPj5P4"
# Come altro esempio, possiamo ordinare le stringhe:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 484, "status": "ok", "timestamp": 1519035909727, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="Z2kdDpxNjcMg" outputId="fbf06717-b6d4-4e00-a440-36628440c398"
x = [ 'mondo', 'python', 'ciao',]
x.sort()
x
# + [markdown] colab_type="text" id="tyK4Pgo7lcay"
# Se non volessimo modificare la lista originale e invece volessimo generarne una nuova, useremmo la funzione `sorted()`. **NOTA**: `sorted` è una _funzione_, non un _metodo_:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 826, "status": "ok", "timestamp": 1519035925999, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="7x9keLzIlbeY" outputId="975525d4-ba5f-4075-96bc-9de778fac761"
x = [ 'mondo', 'python', 'ciao',]
sorted(x)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 477, "status": "ok", "timestamp": 1519036050423, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="L87O-fjMlwoz" outputId="98291939-58ee-494c-a7b6-5caf071741a3"
# l'x originale non è cambiato:
x
# + [markdown] colab_type="text" id="nwsN6H3bkGrN"
# **✪ ESERCIZIO**: Che succede se ordini stringhe contenenti gli stessi caratteri ma maiscoli invece di minuscoli? Come vengono ordinati? Fai delle prove.
#
# +
# scrivi qui
# -
# **✪ ESERCIZIO**: Che succede se nella stessa lista metti sia stringhe che numeri e provi ad ordinarla? Fai delle prove.
#
# scrivi qui
# + [markdown] colab_type="text" id="OPTT3kHSpp3m"
# **Ordine rovesciato**
#
# Supponiamo di voler ordinare la lista alla rovescia usando `sorted`. Per fare cioò possiamo indicare a Python il parametro booleano `reverse` e il suo valore, che in questo caso sarà `True`. Questo ci permette di notare come Python consenta di usare parametri opzionali specificandoli _per nome_ :
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 662, "status": "ok", "timestamp": 1519036904044, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="1uL7iYVxp0Jc" outputId="0c2ff50a-f5a7-4a9b-9615-47a4bfd61df9"
sorted(['mondo', 'python', 'ciao'], reverse=True)
# + [markdown] colab_type="text" id="J4NE2J80qNj9"
# **✪ ESERCIZIO**: Per cercare informazioni su `sorted`, avremmo potuto chiedere a Python dell'aiuto. Per fare ciò Python mette a disposizione una comoda funzione chiamata `help`, che potresti usare così `help(sorted)`. Prova ad eseguirla nella cella qua sotto. A volte l'help è piuttosto complesso, e sta a noi sforzarci un po' per individuare i parametri di interesse
#
# +
# scrivi qui
# + [markdown] colab_type="text" id="BYKOKlw3rTXo"
# **Rovesciare liste non ordinate**
#
# E se volessimo rovesciare una lista così com'è, senza ordinarla in senso decrescente, per esempio per passare da`[6,2,4]` a `[2,4,6]` Cercando un po' nella libreria di Python, vediamo che c'è una comoda funzione `reversed()` che prende come paramatro la lista che vogliamo rovesciare e ne genera una nuova rovesciata.
#
# **✪ ESERCIZIO**: Prova ad eseguire `reversed([6,2,4])` nella cella qua sotto, e guarda che output ottieni. E' quello che ti aspetti ? In genere, e specialmente in Python 3, quando ci aspettiamo una lista e per caso vediamo invece un oggetto col nome `iterator`, possiamo risolvere passando il risultato come parametro della funzione `list()`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ajY8faRvsBIu"
# scrivi qui il codice
# + [markdown] colab_type="text" id="poPVz2M0AFWT"
#
# ## Dizionari - dict
#
# I dizionari sono dei contenitori che ci consentono di abbinare dei _valori_ a delle voci dette _chiavi_. Qua faremo un esempio rapidissimo per dare un'idea.
#
#
# **Riferimenti**:
#
# - [Pensare in Python, Capitolo 11, Dizionari](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2012.html)
#
# - SoftPython - dizionari:
# 1. [introduzione](https://it.softpython.org/dictionaries/dictionaries1-sol.html)
# 2. [operatori](https://it.softpython.org/dictionaries/dictionaries2-sol.html)
# 3. [metodi](https://it.softpython.org/dictionaries/dictionaries3-sol.html)
# 4. [iterazione e funzioni](https://it.softpython.org/dictionaries/dictionaries4-sol.html)
# 5. [strutture composte](https://it.softpython.org/dictionaries/dictionaries5-sol.html)
#
# Possiamo creare un dizionario con le graffe `{` `}`, separando le chiavi dai valori con i due punti `:`, e separando le coppie chiave/valore con la virgola `,`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="TBTib8uDAFWT"
d = { 'chiave 1':'valore 1', 'chiave 2':'valore 2'}
# + [markdown] colab_type="text" id="-KbnLHG4AFWY"
# Per accedere ai valori, possiamo usare le chiavi tra parentesi quadre:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="uOtygGF-AFWY" outputId="424158a1-1e06-4101-dfb8-3c5ea0439140"
d['chiave 1']
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="USOz4GrKAFWh" outputId="c19e78b1-5506-4ae2-9e52-123c6a36c24c"
d['chiave 2']
# + [markdown] colab_type="text" id="Y6g0wQOoAFWn"
# **Valori**: come valori nei dizionari possiamo mettere quello che ci pare, numeri, stringhe, tuple, liste, altri dizionari ..
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="n_jOaUL7AFWn"
d['chiave 3'] = 123
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="4rW2qESsAFWr" outputId="a4d78101-ffa3-4061-f359-c9c9e50c6841"
d
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="5xIZB-EnAFWw"
d['chiave 4'] = ('io','sono', 'una', 'tupla')
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="npAcGHM8AFW7" outputId="0ea2e4b3-8693-4d97-ee75-019c901b3b99"
d
# + [markdown] colab_type="text" id="ULq3n4TiAFXB"
# **✪ ESERCIZIO**: prova ad inserire nel dizionario delle coppie chiave valore con chiavi stringa e come valori delle liste e altri dizionari,
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="aqL7HMKjAFXC"
# scrivi qui:
# + [markdown] colab_type="text" id="9IpoJd3WAFXP"
# **Chiavi**: Per le chiavi abbiamo qualche restrizione in più. Dentro i dizionari possiamo anche inserire dei numeri interi come chiavi:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="2GGqOz7XAFXS"
d[123] = 'valore 3'
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="xT8hKlKWAFXW" outputId="8f04e1e5-9e98-4765-b639-4720aa4cd863"
d
# + [markdown] colab_type="text" id="HwOmHE59AFXo"
# o persino delle sequenze, purchè siano _immutabili_ come le tuple:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="yjUVpZA9AFXp"
d[('io','sono','una','tupla')] = 'valore 4'
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="cfFQsiTAAFXs" outputId="3d298239-01e2-417c-b5a6-84f684fc6bd6"
d
# + [markdown] colab_type="text" id="oDjgDSrdAFX7"
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Non tutti i tipi vanno bene a Python come chiavi. Senza andare nei dettagli, in genere non puoi inserire nei dizionari dei tipi che possono essere modificati dopo che sono stati creati.
#
# </div>
#
# **✪ ESERCIZIO**:
#
# - Prova ad inserire in un dizionario una lista tipo `['a','b']`come chiave, e come valore metti quello che vuoi. Python non dovrebbe permetterti di farlo, e ti dovrebbe mostrare la scritta `TypeError: unhashable type: 'list'`
# - prova anche ad inserire un dizionario come chiave (per es. anche il dizionario vuoto `{}`). Che risultato ottieni?
#
# -
# ## Visualizzare l'esecuzione con Python Tutor
#
#
# Abbiamo visto i principali tipi di dati. Prima di procedere oltre, è bene vedere gli strumenti giusti per comprendere al meglio cosa succede quando si esegue il codice.
# [Python tutor](http://pythontutor.com/) è un ottimo sito online per visualizzare online l’esecuzione di codice Python, permettendo di andare avanti e _indietro_ nell'esecuzione del codice. Sfruttatelo più che potete, dovrebbe funzionare con parecchi degli esempi che tratteremo a lezione. Vediamo un esempio.
#
# **Python tutor 1/4**
#
# Vai sul sito [pythontutor.com](http://pythontutor.com/) e seleziona _Python 3_
# 
# **Python tutor 2/4**
#
# 
# **Python tutor 3/4**
#
# 
# **Python tutor 4/4**
#
# 
# ### Debuggare codice in Jupyter
#
# Python tutor è fantastico, ma quando esegui del codice in Jupyter e non funziona, come si può fare? Per ispezionare l'esecuzione, gli editor di solito mettono a disposizione uno strumento chiamato _debugger_, che permette di eseguire le istruzioni una per una. Al momento (Agosto 2018), il debugger di Jupyter che si chiama [pdb](https://davidhamann.de/2017/04/22/debugging-jupyter-notebooks/) è estramamente limitato . Per superarne le limitazioni, in questo corso ci siamo inventati una soluzione di ripiego, che sfrutta Python Tutor.
#
# Se inserisci del codice Python in una cella, e poi **alla fine della cella** scrivete l'istruzione `jupman.pytut()`, come per magia il codice precedente verrà visualizzato all'interno del foglio Jupyter con il debugger di Python Tutor.
#
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: `jupman` è una collezione di funzioni di supporto che ci siamo inventati apposta per questo corso.
#
# Quando vedi comandi che iniziano con `jupman`, affinchè funzionino devi prima eseguire la cella in cima al documento. Riportiamo tale cella qua per comodità. Se non lo hai già fatto, eseguila adesso.
#
# </div>
#
# Ricordati di eseguire questa cella con Control+Invio
# Questi comandi dicono a Python dove trovare il file jupman.py
import sys;
sys.path.append('../');
import jupman;
# Adesso siamo pronti a provare Python tutor con la funzione magica `jupman.pytut()`:
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Per usare Python tutor dentro Jupyter dovete essere online. Una volta eseguita la cella successivo, dopo qualche secondo dovrebbe apparire il debugger di Python tutor:
#
# </div>
#
# +
x = 5
y = 7
z = x + y
jupman.pytut()
# -
# #### Python Tutor : Limitazione 1
#
# Python tutor è comodo, ma ci sono importanti limitazioni:
#
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Python Tutor guarda dentro una cella sola!
#
# Quando usi Python tutor dentro Jupyter, l'unico codice che viene considerato da Python tutor è quello dentro la cella dove sta il comando jupman.pytut().
# </div>
#
# Quindi per esempio in queste due celle che seguono, solo `print(w)` apparirà dentro Python tutor senza il `w = 3`. Se proverai a cliccare _Forward_ in Python tutor, ti verrà segnalato che non è stata definita `w`
w = 3
# +
print(w)
jupman.pytut()
# -
# Per avere tutto in Python tutor devi mettere tutto il codice nella stessa cella:
# +
w = 3
print(w)
jupman.pytut()
# -
# #### Python Tutor : Limitazione 2
#
# Un'altra limitazione è la seguente:
#
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Python tutor usa solo funzioni dalla distribuzione standard di Python**
#
# Python Tutor va bene per ispezionare semplici algoritmi che usano funzioni di base di Python.
# </div>
#
# Se usate qualche libreria tipo `numpy`, potete provare **solo online** a selezionare `Python 3.6 with anaconda`
#
# 
#
# + [markdown] colab_type="text" id="y1_ipl04AFYh"
# ## Iterazione
#
# Spesso è utile compiere azioni su ogni elemento di una sequenza.
#
# **Riferimenti**
#
# - SoftPython:
# - [controllo di flusso - cicli for](https://it.softpython.org/control-flow/flow2-for-sol.html)
# - [controllo di flusso - cicli while](https://it.softpython.org/control-flow/flow3-for-sol.html)
# - [Pensare in Python, Capitolo 7, Iterazione](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2008.html)
# - [<NAME>, Lezione 8, L'istruzione while](http://ncassetta.altervista.org/Tutorial_Python/Lezione_08.html)
#
#
# **Cicli for**
#
# Tra i vari modi per farlo, uno è usare i cicli `for`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="dvT2rnbBAFYi" outputId="e5deee14-f728-4236-9a21-0cb13531897e"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
for animale in animali:
print("Nella lista ci sono:")
print(animale)
# + [markdown] colab_type="text" id="aYoeEiXoAFYn"
# Qua abbiamo definito la variabile `animale` (avremmo potuto chiamarla con qualunque nome, anche `pippo`). Per ogni elemento nella lista `animali`, vengono eseguite le istruzioni dentro il blocco. Ogni volta che le istruzioni vengono eseguite, la variabile `animale` assume uno dei valori della lista `animali`
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: RICORDATI I DUE PUNTI `:` ALLA FINE DELLA LINEA DEL FOR !!!
# </div>
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: Per indentare il codice, usa SEMPRE sequenze di 4 spazi bianchi. Sequenze di 2 soli spazi per quanto consentite non sono raccomandate.
# </div>
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: A seconda dell'editor che usi, premendo TAB potresti ottenere una sequenza di spazi bianchi come accade in Jupyter (4 spazi che sono raccomandati), oppure un carattere speciale di tabulazione (da evitare)! Per quanto noiosa questa distinzione ti possa apparire, ricordatela perchè potrebbe generare errori molto difficili da scoprire.
# </div>
#
# +
# Guardiamo cosa succede con Python tutor:
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
for animale in animali:
print("Nella lista ci sono:")
print(animale)
jupman.pytut()
# + [markdown] colab_type="text" id="wnbHtwz7AFYo"
# **✪ ESERCIZIO**: Proviamo a capire meglio tutti gli _Attenzione_ qua sopra. Scrivi qui sotto il `for` con gli animali di prima (niente copia e incolla!), vedi se funziona. Ricordati di usare 4 spazi per le indentazioni.
#
# - Poi prova a togliere i due punti alla fine e vedi che errore ti da Python
# - Riaggiungi i due punti, e adesso prova a variare l'indentazione. Prova a mettere due spazi all'inizio di entrambi i print, vedi se esegue
# - Adesso prova a mettere due spazi prima del primo print e 4 spazi prima del secondo, e vedi se esegue
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="STgILuUEAFYr"
# scrivi qui - copia il for di sopra
# + [markdown] colab_type="text" id="4JM8nJYtAFYx"
# ### for in range
#
# Un'altra iterazione molto comune è incrementare un contatore ad ogni ciclo. Python rispetto ad altri linguaggi offre un sistema piuttosto particolare che usa la funzione `range(n)`, che ritorna una sequenza con i primi numeri da 0 incluso a `n` _escluso_. La possiamo usare così:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="AKaswfGQAFYy" outputId="7110d199-e29a-4c79-9ff6-c04a1b64bb63"
for indice in range(3):
print(indice)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} colab_type="code" id="kpkqkSuIAFY7" outputId="49723053-062b-4dca-b64c-b5e2ae2c8ae4"
for indice in range(6):
print(indice)
# -
# Guardiamo meglio con Python tutor:
# +
for indice in range(6):
print(indice)
jupman.pytut()
# + [markdown] colab_type="text" id="Z5jrBwStAFZQ"
# Quindi possiamo usare questo stile come un'alternativa per listare i nostri animali:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 121, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 591, "status": "ok", "timestamp": 1519033207873, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="u5UoFcHtAFZV" outputId="695b71fa-dff0-4787-c950-43d4872a55ae"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
for indice in range(3):
print("Nella lista ci sono:")
print(animali[indice])
# -
# Guardiamo meglio con Python tutor:
# +
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
for indice in range(3):
print("Nella lista ci sono:")
print(animali[indice])
jupman.pytut()
# + [markdown] colab_type="text" id="BeFkQO8VH16f"
# ## Funzioni
#
# Una funzione prende dei parametri e li usa per produrre o riportare qualche risultato.
#
# **Riferimenti**
#
# - [Pensare in Python, Capitolo 3, Funzioni](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2004.html)
# - [Pensare in Python, Capitolo 6, Funzioni produttive](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2007.html) puoi fare tutto saltando la parte 6.5 sulla ricorsione.
# **NOTA**: nel libro viene usato il termine strano 'funzioni produttive' per quelle funzioni che ritornano un valore, ed il termine ancora più strano 'funzioni vuote' per funzioni che non ritornano nulla ma fanno qualche effetto tipo stampa a video: ignora e dimentica questi termini !
#
# - [Nicola Cassetta, Lezione 4, Funzioni](http://ncassetta.altervista.org/Tutorial_Python/Lezione_04.html)
# - [SoftPython, Esercizi sulle funzioni](https://it.softpython.org/functions/functions-sol.html)
#
#
# Per definire una funzione, possiamo usare la parola chiave `def`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="JHBncVarIG0a"
def mia_stampa(x,y): # RICORDATI I DUE PUNTI ':' ALLA FINE DELLA RIGA !!!!!
print('Ora stamperemo la somma di due numeri')
print('La somma è %s' % (x + y))
# + [markdown] colab_type="text" id="4qODhc9qIQVZ"
# Possiamo chiamare la funzione così:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 52, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 477, "status": "ok", "timestamp": 1519045245334, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="I0xzUz41IIpG" outputId="91e139d0-6249-41a4-d6d8-de59ab9c00ba"
mia_stampa(3,5)
# -
# Vediamo meglio che succede con Python Tutor:
# +
def mia_stampa(x,y): # RICORDATI I DUE PUNTI ':' ALLA FINE DELLA RIGA !!!!!
print('Ora stamperemo la somma di due numeri')
print('La somma è %s' % (x + y))
mia_stampa(3,5)
jupman.pytut()
# + [markdown] colab_type="text" id="TNftj3qzJWgv"
# La funzione appena dichiarata stampa dei valori, ma non ritorna nulla. Per fare una funzione che ritorni un valore, dobbiamo usare la parola chiava `return`
#
# Trovi questo tipo di funzioni anche sul libro [Pensare in Python, Capitolo 6, Funzioni produttive](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2007.html), di cui puoi fare tutto saltando la parte 6.5 sulla ricorsione. NOTA: nel libro viene usato il termine strano “funzioni produttive” per quelle funzioni che ritornano un valore, ed il termine ancora più strano “funzioni vuote” per funzioni che non ritornano nulla ma fanno qualche effetto tipo stampa a video: ignora questi strani termini !
#
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="FN9Ek84QIZdK"
def mia_somma(x,y):
s = x + y
return s
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 488, "status": "ok", "timestamp": 1519045262248, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="vGaE6W6bIOn3" outputId="6bac1cdf-285a-41ea-d335-6703e1ee9cff"
mia_somma(3,5)
# +
# Vediamo meglio che succede con Python Tutor:
def mia_somma(x,y):
s = x + y
return s
print(mia_somma(3,5))
jupman.pytut()
# + [markdown] colab_type="text" id="ctEd5oqiJ5Pn"
# **✪ ESERCIZIO**: Se proviamo ad assegnare ad una variabile `x` il valore di ritorno delle funzione `mia_stampa` che apparantemente non ritorna nulla, che valore ci sarà in `x`? Prova a capirlo qua sotto:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Wdy1FRAcKH0i"
# scrivi qui
# + [markdown] colab_type="text" id="zP0UHumoKreN"
# **✪ ESERCIZIO**: Scrivi qua sotto una funzione `media` che calcola e ritorna la media tra due numeri x e y in input
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Wl2MS0-1K0ug"
# scrivi qui
def media(x, y):
return (x + y) / 2
# + [markdown] colab_type="text" id="2nMcWnwHLQte"
# **✪✪ ESERCIZIO**: Scrivi qua sotto una funzione che chiameremo `iniziab` che prende una stringa `x` in ingresso. Se la stringa inizia con la lettera 'b', per esempio `'bianco'` la funzione stampa la scritta `bianco inizia con b`, altrimenti stampa `bianco non inizia con la b`.
#
# * Per controllare se il primo carattere è uguale alla `'b'`, usa l'operatore `==` (ATTENZIONE: è un DOPPIO uguale !)
# * Se la stringa è vuota la tua funzione può avere problemi? Come potresti risolverli? (Per separare più condizioni nell'`if`, usa l'operatore `and` oppure `or` a seconda di come hai costruito l'if).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="2-rCBLAYMA3Y"
# scrivi qui
def iniziab(x):
if len(x) != 0 and x[0] == 'b':
print(x + ' inizia con b')
else:
print(x + ' non inizia con b')
iniziab('bianco')
iniziab('verde')
iniziab('')
# + [markdown] colab_type="text" id="4X1oP2z2Nz4z"
# ### Funzioni lambda
#
# In Python una variabile può contenere una funzione. Per esempio, sappiamo che `len("ciao")` ci dà la lunghezza della stringa `"ciao"`
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 500, "status": "ok", "timestamp": 1519046536040, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="El31mlUNOoRX" outputId="24ca6750-254e-4062-f24a-6850eb7af742"
len("ciao")
# + [markdown] colab_type="text" id="ENEMhfSiOrUM"
# Proviamo a creare una variabile `mia_variabile` che punta alla funzione `len`.
#
# **NOTA**: _non_ abbiamo aggiunto parametri a `len`!
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="6S-wDPTYOJUO"
mia_variabile = len
# + [markdown] colab_type="text" id="PX1hxxoOOUJg"
# Adesso possiamo usare `mia_variabile` esattamente come usiamo la funzione `len`, che da la lunghezza di sequenze come le stringhe
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 649, "status": "ok", "timestamp": 1519046587953, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="Lh6V_SAeOS3O" outputId="efd81d20-9aa9-40b4-8540-ac5d6f45ea6d"
mia_variabile("ciao")
# + [markdown] colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="DN4BJnksO3He"
# Possiamo anche riassegnare `mia_variabile` ad altre funzioni, per esempio `sorted`. Vediamo che succede:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="39ad-1FIOKPB"
mia_variabile = sorted
# + [markdown] colab_type="text" id="fIb3WcLyPCx_"
# chiamando `mia_variabile`, ci aspettiamo di vedere i caratteri di `"ciao"` in ordine alfabetico:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 494, "status": "ok", "timestamp": 1519047027841, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="PmR_4VeMPK9L" outputId="125162e5-a0a0-4c52-9728-4c04fe539fd1"
mia_variabile("ciao")
# + [markdown] colab_type="text" id="Q3CoQlvZPWTG"
# In Python possiamo definire funzioni in una sola riga, con le cosiddette _funzioni lambda_:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="uXGgflBePBlW"
mia_f = lambda x: x + 1
# + [markdown] colab_type="text" id="tdpFbGTsPcZb"
# Cosa fa `mia_f` ? Prende un parametro `x` e ritorna il risultato di calcolare l'espressione `x + 1`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 496, "status": "ok", "timestamp": 1519047256022, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="5wJ9R1oJPj8L" outputId="91aeee26-af36-4dfe-b34e-f149a91dbce2"
mia_f(5)
# + [markdown] colab_type="text" id="sWYLGHWJP0MY"
# Possiamo anche passare due parametri:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="mazLpnywPaDz"
mia_somma = lambda x,y: x + y
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 506, "status": "ok", "timestamp": 1519046858574, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="Fv3LhBAOPrqs" outputId="0efb7db1-538e-42cd-e9c9-3eef80d50ce5"
mia_somma(3,5)
# + [markdown] colab_type="text" id="B5ZCVVazQE8Q"
# **✪ ESERCIZIO**: Prova a definire qua sotto una funzione lambda per calcolare la media tra due numeri `x` e `y`, e assegnala alla variabile `media`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="FAhMilOBQJGK"
# scrivi qui
media = lambda x,y: (x + y)/2
media(2,7)
# + [markdown] colab_type="text" id="wqYdlbArcIo9"
# ## Trasformazioni sulle liste
#
# Supponiamo di voler prendere la lista di animali e generarne una nuova in cui tutti i nomi iniziano con la lettera maiuscola. Questa che vogliamo fare di fatto è la creazione di una nuova lista operando una trasformazione sulla precedente. Per fare ciò esistono diversi modi, quello più semplice è usare un ciclo `for` così
#
# ### Trasfomazioni con il for
#
#
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="SnnvFBIlcNWo"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
nuova_lista = []
for animale in animali: # ad ogni ciclo la variabile 'animale' contiene un nome preso dalla lista 'animali'
nuova_lista.append(animale.capitalize()) # aggiungiamo alla nuova lista il nome dell'animale corrente, con la prima lettera maiuscola
nuova_lista
#vediamo che succede con Python tutor
jupman.pytut()
# + [markdown] colab_type="text" id="6vRDoWxidjgX"
# Nota importante: i metodi sulle stringhe non modificano mai la stringa originale, ma ne generano sempre una nuova. Quindi la lista originale `animali` conterrà ancora le stringhe originale senza modifiche:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 560, "status": "ok", "timestamp": 1519034975589, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="vC58tUlqdxgf" outputId="6d32c5bc-782d-4d95-9755-2eab9ee93c05"
animali
# + [markdown] colab_type="text" id="45lFek_khbEX"
# **✪ ESERCIZIO**: Prova a scrivere qua sotto un ciclo `for` (senza usare il copia e incolla!) che scorre la lista dei nomi degli animali e crea un'altra lista che chiameremo `m` in cui tutti i caratteri dei nomi degli animali sono in maiuscolo (usa il metodo `.upper()`)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="3W5QjOZ6G_D-"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
# scrivi qui
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
# Scrivi qua
m = []
for animale in animali: # ad ogni ciclo la variabile 'animale' contiene un nome preso dalla lista 'animali'
m.append(animale.upper()) # aggiungiamo alla nuova lista il nome dell'animale corrente, con la prima lettera maiuscola
m
# + [markdown] colab_type="text" id="BbNGWT5PeX_x"
# ### Trasformazioni con le _list comprehension_
#
# **Riferimenti**: [SoftPython - Sequenze](https://it.softpython.org/sequences/sequences-sol.html#List-comprehensions)
#
# La stessa identica trasformazione di sopra si potrebbe attuare con una cosiddetta _list comprehension_, che servono per generare nuove liste eseguendo la stessa operazione su tutti gli elementi di una lista esistente di partenza. Come sintassi imitano le liste, infatti iniziano e finiscono con le parentesi quadre, ma dentro contengono un for speciale per ciclare dentro un sequenza :
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="I7fl60rudyTU"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
nuova_lista = [animale.capitalize() for animale in animali]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 483, "status": "ok", "timestamp": 1519033974548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="fSywtAYFdgyY" outputId="f9131bc7-11a0-4cb8-f708-5d101e51d6b1"
nuova_lista
# -
# Vediamo che succede con Python tutor:
# +
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
nuova_lista = [animale.capitalize() for animale in animali]
jupman.pytut()
# + [markdown] colab_type="text" id="G1isjIcitppx"
# **✪ ESERCIZIO**: Prova qua sotto ad usare una list comprehension per mettere tutti i caratteri in maiuscolo
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="eoNE416HdIEf"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
# scrivi qui
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
nuova_lista = [animale.upper() for animale in animali]
# + [markdown] colab_type="text" id="wJkXW0NulL8I"
# **Filtrare con le comprehension**: Volendo, possiamo anche filtrare i dati usando un `if` speciale da mettere alla fine della comprehension. Per esempio potremmo selezionare solo gli animali la cui lunghezza del nome è di 4 caratteri:
#
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 562, "status": "ok", "timestamp": 1519052514136, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="3j3N8Z16lKDW" outputId="ee544608-9e3a-4918-f6a3-716081ad8b8a"
[animale.upper() for animale in animali if len(animale) == 4]
# + [markdown] colab_type="text" id="VnneWysUHt02"
# ### Trasformazioni con le map
#
# Un'altro modo ancora per trasformare una lista in una nuova è usare l'operazione `map`, che a partire da una lista, ne genera un'altra applicando ad ogni elemento della lista di partenza una funzione `f` che passiamo come parametro. Per risolvere lo stesso esercizio precedente, si potrebbe per esempio creare al volo con `lambda` una funzione `f` che mette la prima lettera di una stringa in maiuscolo, e poi si potrebbe chiamare la `map` passando la `f` appena creata:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 478, "status": "ok", "timestamp": 1519047521588, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="b_0fe_rQHct3" outputId="e9797827-6f8b-4d18-a067-571aa70a265b"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
f = lambda animale: animale.capitalize()
map(f, animali)
# + [markdown] colab_type="text" id="N0EHix0jSjRL"
# Purtroppo il risultato non è esattamente la lista che volevamo. Il problema è che Python 3 aspetta a ritornarci una lista vera e propria, e invece ci ritorna un _iteratore_. Come mai? Python 3 per ragioni di efficienza spera che non non andremo mai ad usare nessun elemento della nuova lista, evitandogli di fatto la fatica di applicare la funzione a tutti gli elementi della lista originale. Ma noi possiamo forzarlo a darci la lista che vogliamo usando la funzione `list`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 484, "status": "ok", "timestamp": 1519047866872, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="BDT49G82TJMH" outputId="a095d6f0-9a99-4765-9bc1-f5e74515e4cb"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
f = lambda animale: animale.capitalize()
list(map(f, animali))
# + [markdown] colab_type="text" id="dh3LFNNqTyuK"
# Per avere un esempio totalmente equivalente ai precedenti, possimao assegnare il risultato a `nuova_lista`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="JM96quneT9lV"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
f = lambda animale: animale.capitalize()
nuova_lista = list(map(f, animali))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 601, "status": "ok", "timestamp": 1519048055236, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="xI7TtsFmSbFg" outputId="eb2dda9f-0855-4347-a83d-5afce1a1390d"
nuova_lista
# + [markdown] colab_type="text" id="DkdrP-hVUsNf"
# Un vero hacker Python probabilmente preferirà scrivere tutto in una sola linea, così:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="u-HHQz8BUjE6"
animali = ['cani', 'gatti', 'scoiattoli', 'alci']
nuova_lista = list(map(lambda animale: animale.capitalize(), animali))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 409, "status": "ok", "timestamp": 1519048159745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="fnNMSHV2U2QC" outputId="045c8d0b-ff7e-4a39-fde7-f8202b17e6ef"
nuova_lista
# + [markdown] colab_type="text" id="t6vIHoNwUaC5"
# **✪ ESERCIZIO**: la lista originale `animali` è cambiata? Controlla.
# + [markdown] colab_type="text" id="7lmjhzyxU_-p"
# **✪ ESERCIZIO**: Data una lista di numeri `numeri = [3, 5, 2, 7]` prova a scrivere una `map` che genera una nuova lista con i numeri raddoppiati, come `[6, 10, 4, 14]`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="4tf9IOLKVhUQ"
numeri = [3, 5, 2, 7]
# scrivi qui
numeri = [3,5,2,7]
list(map(lambda x: x * 2, numeri))
# + [markdown] colab_type="text" id="BCmNz8C6AFZY"
# ## Matrici
#
# Finita la presentazione, è ora di sforzarsi un po' di più. Vediamo brevemente matrici come liste di liste. Per approfondire, guardare i riferimenti.
#
# **Riferimenti**:
#
# - [SoftPython - matrici come liste di liste](https://it.softpython.org/matrices-lists/matrices-lists-sol.html)
# - [SoftPython - matrici numpy](https://it.softpython.org/matrices-numpy/matrices-numpy-sol.html)
#
#
# **✪✪ ESERCIZIO**: Date le due liste con nomi di animali e corrispondente aspettativa di vita in anni:
#
# ```python
# animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
# anni = [12,14,30,6,25]
# ```
#
# Scrivere nella cella sotto del codice che generi una lista di liste da due elementi, così:
#
# `[
# ['cane', 12],
# ['gatto', 14],
# ['pellicano', 30],
# ['scoiattolo', 6],
# ['aquila', 25]
# ]`
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="WSlJUPE5AFZY"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui
coppie = []
for i in range(len(animali)):
coppie.append([animali[i], anni[i]])
coppie
# + [markdown] colab_type="text" id="bd2Ap-uOAFZc"
# **✪✪ ESERCIZIO** modificare scrivendolo qua sotto il codice dell'esercizio precedente nella versione con il normale ciclo `for` per filtrare solo le specie con aspettativa di vita superiore ai 13 anni, così da ottenere questo risultato:
#
# ```python
# [['gatto', 14], ['pellicano', 30], ['aquila', 25]]
# ```
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="M4d3V2JjnocX"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
coppie = []
for i in range(len(animali)):
if anni[i] > 13:
coppie.append([animali[i], anni[i]])
coppie
# + [markdown] colab_type="text" id="LvVoUJ1mAFZg"
# **ESERCIZIO** Scrivi qua sotto del codice con un normale ciclo `for` filtri solo le specie con aspettativa di vita superiore ai 10 anni e inferiore ai 27, così da ottenere questo risultato:
#
# ```python
# [['cane', 12], ['gatto', 14], ['aquila', 25]]
# ```
#
# **SUGGERIMENTO**: in Pyhton per imporre due condizioni in un if si usa la parola chiave **and**
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ikN8wQ6oAFZg"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
coppie = []
for i in range(len(animali)):
if anni[i] > 10 and anni[i] < 27 :
coppie.append([animali[i], anni[i]])
coppie
# + [markdown] colab_type="text" id="itNZWFEYh4Oe"
# ### Funzione zip
#
# La funzione `zip` prende due liste e ne restituisce una terza nuova, in cui mette coppie di elementi come tuple (che ricordiamo sono come le liste ma immutabilii), abbinando il primo elemento della prima lista a primo elemento della seconda lista, il secondo elemento della prima lista al secondo elemento della secondo e così via :
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 505, "status": "ok", "timestamp": 1519053345513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="FDHB64Q5olmu" outputId="50b8e861-9f06-4215-a7fb-d3ba21d8dfa4"
list(zip(['a','b','c'], [5,2,7]))
# + [markdown] colab_type="text" id="eWfvYP15oqwu"
# Perchè abbiamo messo anche `list` nell'esempio? Perchè `zip` ha lo stesso problema della map, cioè non ritorna subito una lista come vorremmo noi:
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 538, "status": "ok", "timestamp": 1519053414986, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="jBWLb2vZjPKv" outputId="ebd86242-51e3-46f9-af11-85eb92aa54d1"
zip(['a','b','c'], [5,2,7])
# + [markdown] colab_type="text" id="kcbDFCyOjl7E"
# **✪✪✪ ESERCIZIO**: Come vedi con la `zip` abbiamo ottenuto un risultato simile a quello del precedente esercizio, ma abbiamo tuple con parentesi tonde invece di liste con parentesi quadre. Riusciresti aggiungendo una `list comprehension` o una `map` ad ottenere lo stesso identico risultato?
#
# - per convertire una tupla in una lista, usa la funzione `list`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 573, "status": "ok", "timestamp": 1519054370413, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="XNP1gY9roEbM" outputId="dfb74688-0995-47cc-870b-a5934fa0a23c"
list( ('ciao', 'soft', 'python') ) # all'interno abbiamo messo una tupla, delimitata da parentesi tonde
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="m_rDAV-Qnd86"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui - soluzione con list comprehension
[ list(c) for c in zip(animali, anni) ]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 104, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 416, "status": "ok", "timestamp": 1519054478279, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="z0PWbUtnnGAK" outputId="32110d5c-9e25-419d-9a66-88d6d1b789c1"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui - soluzione con map
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
list(map(list, zip(animali, anni)))
# + [markdown] colab_type="text" id="JS4ZOkHjqRE8"
# **✪✪✪ ESERCIZIO**: svolgi l'esercizio precedente filtrando gli animali con aspettativa di vita superiore ai 13 anni, usando la `zip` e una list comprehension
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="lmNN4HYfnZXi"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
[ list(c) for c in zip(animali, anni) if c[1] > 13 ]
# + [markdown] colab_type="text" id="gmluxI1sAFZn"
# **✪✪ ESERCIZIO**: Date le due liste con nomi di animali e corrispondente aspettativa di vita in anni come sopra, scrivere nella cella sotto del codice che con un normale ciclo `for` generi un dizionario che associa alla specie l'aspettativa di vita, così:
#
# ```python
# {
# 'aquila': 25,
# 'cane': 12,
# 'gatto': 14,
# 'pellicano': 30,
# 'scoiattolo': 6
# }
# ```
#
# <div class="alert alert-warning">
#
# **ATTENZIONE**: A seconda della versione esatta di Python che avete e di come il dizionario viene creato, l'ordine dei campi quando viene stampato potrebbe differire dall'esempio. Questo è perfettamente normale, perchè le chiavi di un dizionario sono da intendersi come un insieme senza ordine particolare. Se volete essere sicuri di trovare le chiavi stampate nell'ordine in cui sono state inserite, dovete usare un OrderedDict
#
# </div>
#
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="8aomrV6KAFZn"
animali = ['cane', 'gatto', 'pellicano', 'scoiattolo', 'aquila']
anni = [12,14,30,6,25]
# scrivi qui
d = {}
for i in range(len(animali)):
d[animali[i]] = anni[i]
d
# + [markdown] colab_type="text" id="y2Lk387pvSGo"
# Per ottenere lo stesso risultato in una linea, è possibile usare la funzione `zip` come fatto negli esercizi precedenti, e poi la funzione `dict` che crea un dizionario a partire dalla lista di coppie di elementi generata dallo `zip`:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 524, "status": "ok", "timestamp": 1519055238687, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102920557909426557439"}, "user_tz": -60} id="9DB78dCBcHPt" outputId="158d49f3-b0ff-4939-d42c-b8e4510cdf3b"
dict(zip(animali, anni))
# + [markdown] colab_type="text" id="jbX7XEmRAFZ8"
# **✪✪ ESERCIZIO**: Data una lista di prodotti contenente a sua volta liste ciascuna con tipologia, marca e quantità di confezioni vendute:
#
#
# ```python
# vendite = [
# ['pomodori', 'Santini', 5],
# ['pomodori', 'Cirio', 1],
# ['pomodori', 'Mutti', 2],
# ['cereali', 'Kelloggs', 3],
# ['cereali', 'Choco Pops', 8],
# ['cioccolata','Novi', 9],
# ['cioccolata','Milka', 4],
# ]
#
# ```
#
# Usando un normale ciclo for, scrivere del codice Python nella cella sotto per creare un dizionario in cui le chiavi sono le tipologie e i valori sono la somma delle confezioni vendute per quella categoria:
#
# ```python
# {
# 'cereali': 11,
# 'cioccolata': 13,
# 'pomodori': 8
# }
# ```
#
#
# **SUGGERIMENTO**: fare attenzione ai due casi, quando il dizionario da ritornare ancora non contiene la tipologia estratta dalla lista correntemente in esame e quando invece già la contiene:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Si8Tt6WHAFZ8"
vendite = [
['pomodori', 'Santini', 5],
['pomodori', 'Cirio', 1],
['pomodori', 'Mutti', 2],
['cereali', 'Kelloggs', 3],
['cereali', '<NAME>', 8],
['cioccolata','Novi', 9],
['cioccolata','Milka', 4],
]
# scrivi qui
vendite = [
['pomodori', 'Santini', 5],
['pomodori', 'Cirio', 1],
['pomodori', 'Mutti', 2],
['cereali', 'Kelloggs', 3],
['cereali', '<NAME>', 8],
['cioccolata','Novi', 9],
['cioccolata','Milka', 4],
]
d = {}
for vendita in vendite:
if vendita[0] in d:
d[vendita[0]] += vendita[2]
else:
d[vendita[0]] = vendita[2]
d
# -
# ## Approfondimenti
#
# **Strumenti e script**: Se vuoi approfondire come eseguire il codice in editor diversi da Jupyter e avere un'idea più precisa dell'architettura di Python, ti invitiamo a leggere [Strumenti e script](https://it.softpython.org/tools/tools-sol.html)
#
# **Gestione errori e testing**: Per capire come si gestiscono le situazioni di errore in genere guarda il foglio separato [Gestione errori e testing](https://it.softpython.org/exercises/errors-and-testing/errors-and-testing-sol.html), serve anche per capire come eseguire alcuni esercizi della Parte A - Fondamenti.
| quick-intro/quick-intro-sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Barnase-Barstar
import molsysmt as msm
molsys = msm.convert('pdb_id:1BRS')
msm.info(molsys)
msm.view(molsys)
| docs/contents/Barnase-Barstar/Barnase_Barstar_vacuum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
@author: mdigi14
"""
import pandas as pd
import matplotlib.pyplot as plt
import datetime
import time
import requests
"""
Update Parameters Here
"""
COLLECTION = "MekaVerse"
CONTRACT = "0x9A534628B4062E123cE7Ee2222ec20B86e16Ca8F"
YEAR = "2021"
MONTH = "10"
DAY = "13"
HOUR = "15"
MINUTE = "00"
REVEAL_TIME = "{}-{}-{}T{}:{}:00".format(YEAR, MONTH, DAY, HOUR, MINUTE) # have to double check if OS uses UTC time
DATETIME_REVEAL_TIME = datetime.datetime.strptime(REVEAL_TIME,"%Y-%m-%dT%H:%M:%S")
ETHER_UNITS = 1e18
LIMIT = 200
SLEEP = 5
LIMIT = 200
MAX_OFFSET = 10000
METHOD = "raritytools"
"""
Helper Functions
"""
def getOpenseaEvents(contract,offset=0,occurred_before=REVEAL_TIME):
print(offset)
url = "https://api.opensea.io/api/v1/events"
querystring = {
"asset_contract_address" : contract,
"only_opensea" : "false",
"offset" : str(offset),
"limit" : LIMIT,
"event_type" : 'bid_entered',
"occurred_before" : REVEAL_TIME
}
headers = {"Accept" : "application/json"}
response = requests.request("GET", url, headers=headers, params=querystring)
return response.json()
# +
RARITY_CSV = "../metadata/rarity_data/{}_{}.csv".format(COLLECTION, METHOD)
RARITY_DB = pd.read_csv(RARITY_CSV)
bids = []
events = []
offset = 0
while offset < MAX_OFFSET:
new_events = getOpenseaEvents(CONTRACT, offset = offset)
for event in new_events['asset_events']:
events.append(event)
offset += LIMIT
if len(new_events['asset_events']) < LIMIT:
break
print("total bids ", len(events))
for event in events:
bid_time = datetime.datetime.strptime(event['created_date'],"%Y-%m-%dT%H:%M:%S.%f")
if bid_time < DATETIME_REVEAL_TIME:
try:
tokenId = int(event['asset']['token_id'])
bid = dict()
bid['TOKEN_ID'] = tokenId
bid['USER'] = event['from_account']['address']
bid['OFFER'] = float(event['bid_amount']) / ETHER_UNITS
bid['DATE'] = event['created_date']
bid['RANK'] = int(RARITY_DB[RARITY_DB['TOKEN_ID'] == tokenId]['Rank'])
bids.append(bid)
except:
continue
bidding_df = pd.DataFrame(bids)
bidding_df.to_csv("pre-reveal_bids/{}_pre-reveal_bids.csv".format(COLLECTION))
ax = bidding_df.plot.scatter(x='TOKEN_ID', y='RANK', alpha=.25, title= "{} - {}".format(COLLECTION, "Pre-reveal Bids"), figsize=(14, 7))
ax.set_xlabel("Token ID")
ax.set_ylabel("Rarity Rank")
| fair_drop/prereveal_bids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing
# ## Introduction
# ### A few reasons not to do testing
# Sensibility | Sense
# ------------------------------------ | -------------------------------------
# **It's boring** | *Maybe*
# **Code is just a one off throwaway** | *As with most research codes*
# **No time for it** | *A bit more code, a lot less debugging*
# **Tests can be buggy too** | *See above*
# **Not a professional programmer** | *See above*
# **Will do it later** | *See above*
# ### A few reasons to do testing
#
# * **lazyness** *testing saves time*
# * **peace of mind** *tests (should) ensure code is correct*
# * **runnable specification** *best way to let others know what a function should do and
# not do*
# * **reproducible debugging** *debugging that happened and is saved for later reuse*
# * code structure / **modularity** *since the code is designed for at least two situations*
# * easier to modify *since results can be tested*
# ### Not a panacea
#
# > Trying to improve the quality of software by doing more testing is like trying to lose weight by
# > weighting yourself more often.
# - <NAME>
# * Testing won't corrrect a buggy code
# * Testing will tell you were the bugs are...
# * ... if the test cases *cover* the bugs
# ### Tests at different scales
#
# Level of test |Area covered by test
# -------------------------- |----------------------
# **Unit testing** |smallest logical block of work (often < 10 lines of code)
# **Component testing** |several logical blocks of work together
# **Integration testing** |all components together / whole program
#
#
# <br>
# <div class="fragment fade-in">
# Always start at the smallest scale!
#
# <div class="fragment grow">
# If a unit test is too complicated, go smaller.
# </div>
# </div>
# ### Legacy code hardening
#
# * Very difficult to create unit-tests for existing code
# * Instead we make a **regression test**
# * Run program as a black box:
#
# ```
# setup input
# run program
# read output
# check output against expected result
# ```
#
# * Does not test correctness of code
# * Checks code is a similarly wrong on day N as day 0
# ### Testing vocabulary
#
# * **fixture**: input data
# * **action**: function that is being tested
# * **expected result**: the output that should be obtained
# * **actual result**: the output that is obtained
# * **coverage**: proportion of all possible paths in the code that the tests take
# ### Branch coverage:
# + [markdown] attributes={"classes": [" python"], "id": ""}
# ```python
# if energy > 0:
# # ! Do this
# else:
# # ! Do that
# ```
# -
# Is there a test for both `energy > 0` and `energy <= 0`?
| ch03tests/01testingbasics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
from scipy import sparse
from scipy import linalg
import scipy.sparse.linalg
from sklearn.cluster import KMeans
routes = pd.read_csv('data/routes.dat', sep=',', header = None, encoding='utf-8')
routes.columns = ['Airline','AirlineID','SourceAirport','SourceAirportID','DestinationAirport','DestinationAirportID','Codeshare','Stops','Equipment']
routes = routes.drop(columns=['AirlineID','SourceAirportID','DestinationAirportID','Stops','Equipment','Codeshare'])
print(routes.head())
print(routes.duplicated().any())
alliances = pd.read_csv('data/alliances.dat', sep=',', header = None, encoding='utf-8')
alliances.columns = ['Alliance','IATA','Region']
print(alliances.head())
print(alliances.duplicated().any())
# +
airlines = pd.read_csv('data/airlines.dat', sep=',', header = None, encoding='utf-8')
airlines.columns = ['Airline ID', 'Name', 'Alias', 'IATA', 'ICAO','Callsign','Country','Active']
airlines = airlines.drop(columns=['Airline ID','Alias','ICAO','Callsign','Active','Country'])
airlines = airlines[~airlines.IATA.isnull()]
airlines = airlines[airlines.IATA != '-']
airlines = airlines[~airlines.Name.isnull()]
airlines = airlines.drop_duplicates()
airlines = airlines.drop_duplicates('IATA')
print(airlines.head())
print(airlines.duplicated(['IATA']).any())
airlineID = routes[['Airline']].rename(columns={'Airline':'IATA'})
airlineID = airlineID.drop_duplicates().reset_index().drop(columns=['index'])
print(airlineID.head())
print(airlineID.duplicated().any())
airlineID = pd.merge(airlineID,alliances,left_on='IATA',right_on='IATA',how='right')
airlineID = pd.merge(airlineID,airlines,left_on='IATA',right_on='IATA',how='left')
airlineID = airlineID.reset_index().rename(columns={'index':'airlineID'})
print(airlineID.head())
print(airlineID.duplicated().any())
# -
routesID = pd.merge(routes,airlineID,left_on='Airline',right_on='IATA',how='right')
# +
source_airports = routesID[['SourceAirport']]
source_airports = source_airports.rename(columns={'SourceAirport':'Airport'})
dest_airports = routesID[['DestinationAirport']]
dest_airports = dest_airports.rename(columns={'DestinationAirport':'Airport'})
airports = pd.concat([source_airports,dest_airports]).drop_duplicates().reset_index().drop(columns=['index']).reset_index()
airports = airports.set_index('Airport').rename(columns={'index':'airportsID'})
print(airports.head())
print(airports.duplicated().any())
# -
routesID = pd.merge(routesID,airports,left_on='SourceAirport',right_on='Airport',how='left')
routesID = routesID.rename(columns={'airportsID':'SourceAirportID'})
routesID = pd.merge(routesID,airports,left_on='DestinationAirport',right_on='Airport',how='left')
routesID = routesID.rename(columns={'airportsID':'DestinationAirportID'})
print(routesID.head())
connections = routesID
connections = connections.drop(columns=['Airline','SourceAirport','DestinationAirport'])
connections = pd.merge(connections,connections,left_on='DestinationAirportID',right_on='SourceAirportID',how='inner')
connections = connections[connections.airlineID_x != connections.airlineID_y]
print(connections.head())
# +
grouped = connections[['airlineID_x','airlineID_y']].groupby(['airlineID_x','airlineID_y'])
group_sizes = grouped.size()
n_airlines = len(airlineID)
adjacency_airlines = np.zeros((n_airlines,n_airlines))
for name,group in grouped:
adjacency_airlines[name[0],name[1]] += group_sizes.loc[name[0],name[1]]
adjacency_airlines[name[1],name[0]] += group_sizes.loc[name[0],name[1]]
for i in range(n_airlines):
for j in range(n_airlines):
if airlineID.loc[i].Region == airlineID.loc[j].Region:
adjacency_airlines[i,j] = 0
# -
adjacency = np.copy(adjacency_airlines)
for i in range(n_airlines):
adjacency[i] = adjacency[i]/np.sum(adjacency[i])
for i in range(n_airlines):
for j in range(n_airlines):
adjacency[i,j] = max(adjacency[i,j],adjacency[j,i])
adjacency[j,i] = adjacency[i,j]
sqrt_inv_degree_matrix
degrees = np.sum(adjacency, axis = 0)
degree_matrix = np.diag(degrees)
laplacian_combinatorial = degree_matrix - adjacency;
sqrt_inv_degree_matrix = np.diag(np.sqrt(1/degrees))
laplacian_normalized = np.dot(np.dot(sqrt_inv_degree_matrix,laplacian_combinatorial),sqrt_inv_degree_matrix)
# +
[eigenvalues, eigenvectors] = np.linalg.eig(laplacian_normalized)
sortID = np.argsort(eigenvalues)
eigenvalues = eigenvalues[sortID]
eigenvectors = eigenvectors[:,sortID]
print(eigenvalues)
# +
k = 3; d = 3
H = eigenvectors[:,:d];
clusters3 = KMeans(n_clusters=k, random_state=0).fit_predict(H)
print("----- For k=",k," and d=",d," -----")
print("Number of elements in clusters :")
for i in range(k):
cnt = 0
for j in clusters3:
if j == i:
cnt +=1
print("Cluster ",i+1,":",cnt)
# -
print(airlineID[clusters3 == 0][['IATA','Alliance','Name']])
print(airlineID[clusters3 == 1][['IATA','Alliance','Name']])
print(airlineID[clusters3 == 2][['IATA','Alliance','Name']])
| .ipynb_checkpoints/main_gabor_plot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importando as bibliotecas
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import metrics
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import KFold
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.ensemble import AdaBoostRegressor
from xgboost import XGBRegressor
from sklearn.model_selection import cross_val_score
from sklearn import model_selection
from sklearn.model_selection import RandomizedSearchCV
from math import sqrt
from warnings import simplefilter
simplefilter(action='ignore', category=FutureWarning)
# Carregando os dados
train = pd.read_csv('Database/train.csv')
test = pd.read_csv('Database/test.csv')
# Verificando os tipos dos dados.
train.info()
# Variáveis numéricas.
train.describe()
# Variáveis categóricas.
train.describe(include=['O'])
# # Exploratory Data Analysis (EDA)
# Começaremos a nossa análise verificando a distribuição da nossa variável alvo (Purchase)
plt.style.use('fivethirtyeight')
plt.figure(figsize=(8,6))
sns.distplot(train.Purchase, bins = 25)
plt.xlabel('Valor gasto na compra')
plt.ylabel('Número de compradores')
plt.title('Distribuição dos valores')
# Até o momento a única informação que temos é que existe uma maior concentração entre os valores de 5000 a 10000 dólares. Seria mais interessante plotarmos um gráfico de boxplot para uma melhor avaliação da distribuição desses dados, além disso podemos facilmente descobrir se temos outliers.
train['Purchase'].plot.box()
# E olha só esse gráfico acaba de nos dar uma imformação muito importante, temos outliers na nossa variável alvo. Podemos facilmente afirmar que os valores de compra acima de 20000 dólares são outliers.
# # Análise por Sexo
train['Gender'].value_counts()
sns.countplot(train.Gender)
# valor médio da compra para cada sexo
train[['Gender','Purchase']].groupby(['Gender'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('Gender', 'Purchase', data=train)
# Bom, o que podemos obsevar é que temos um número bem maior de usuários do sexo masculino e que o valor da compra também tende a ser um pouco maior que a do sexo feminino.
# # Análise por Idade
train['Age'].value_counts()
sns.countplot(train.Age)
# Observe que a faixa etária entre os usuários se concentra entre 26 a 30 anos, mais ainda não conseguimos saber se nessa faixa etária temos mais homens ou mulheres. O código a seguir agrupa os dados por sexo e faz a contagem dos registros por faixa etária.
train.groupby('Gender')['Age'].value_counts()
# Agora sim podemos dizer que temos mais homens que mulheres na faixa etária de 26 a 30 anos.
# valor médio de compra para cada faixa etária
train[['Age','Purchase']].groupby(['Age'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('Age', 'Purchase', data=train)
# O valor médio da compra está bem distribuido entre as idades, com o valor um pouco menor na faixa etária de 0 a 17 anos.
# # Análise por Ocupação
train['Occupation'].value_counts()
sns.countplot(train.Occupation)
train[['Occupation','Purchase']].groupby(['Occupation'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('Occupation', 'Purchase', data=train)
# Temos algumas ocupações com maior concentração de valores, só não sabemos que ocupações são essas, já que a empresa preferiu mascarar essa informação.
# # Análise por Cidade
train['City_Category'].value_counts()
sns.countplot(train.City_Category)
# Média de compra por cidade
train[['City_Category','Purchase']].groupby(['City_Category'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('City_Category', 'Purchase', data=train)
# Chegamos a seguinte conclusão: o maior número de usuários pertence a cidade de categoria B, porém a cidade de categoria C tem o maior valor médio de compra.
# # Análise por tempo de moradia na cidade
train['Stay_In_Current_City_Years'].value_counts()
sns.countplot(train.Stay_In_Current_City_Years)
# média do valor da compra de acordo com o número de anos que o usuário mora na cidade atual
train[['Stay_In_Current_City_Years','Purchase']].groupby(['Stay_In_Current_City_Years'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('Stay_In_Current_City_Years', 'Purchase', data=train)
# De acordo com os nossos gráficos os usuários na sua grande maioria moram há 1 ano na cidade atual.
# # Análise por estado civil
train['Marital_Status'].value_counts()
sns.countplot(train.Marital_Status)
# valor médio de compra para cada estado civil
train[['Marital_Status','Purchase']].groupby(['Marital_Status'], as_index = True).mean().sort_values(by='Purchase',ascending=False)
sns.boxplot('Marital_Status', 'Purchase', data=train)
# 0 não é casado e 1 é casado. Dito isso fica claro que os usuários em sua maioria não são casados.
# # Análise por categoria de produto
train['Product_Category_1'].value_counts()
train[['Product_Category_1','Purchase']].groupby(['Product_Category_1'], as_index = True).mean().sort_values(by='Purchase',ascending=True)
train['Product_Category_1'].plot.box()
# Os produtos da categoria 1 que pertencem ao grupo 19 e 20 são outliers.
train['Product_Category_2'].value_counts()
train[['Product_Category_2','Purchase']].groupby(['Product_Category_2'], as_index = True).mean().sort_values(by='Purchase',ascending=True)
train['Product_Category_2'].plot.box()
# Já na categoria 2, temos uma maior quantidade de produtos do grupo 8. Podemos observar também que o produto do grupo 10 teve o maior valor médio de compra.
train['Product_Category_3'].value_counts()
train[['Product_Category_3','Purchase']].groupby(['Product_Category_3'], as_index = True).mean().sort_values(by='Purchase',ascending=True)
train['Product_Category_3'].plot.box()
# Na categoria 3 temos o produto do grupo 3 com a menor quantidade. Dando uma verificada mais detalhada, chegamos a uma informação importante, na categoria 2 e 3 temos um valor médio de compra bem próximo para os produtos do grupo 10.
# +
# Distribuição dos valores para cada categoria.
plt.figure(figsize=(14,4))
plt.subplot(131)
sns.countplot(train.Product_Category_1)
plt.xticks(rotation=90)
plt.subplot(132)
sns.countplot(train.Product_Category_2)
plt.xticks(rotation=90)
plt.subplot(133)
sns.countplot(train.Product_Category_3)
plt.xticks(rotation=90)
plt.show()
# -
# matriz de correlação
matrix = train.corr()
sns.set(rc={'axes.facecolor':'white', 'figure.facecolor':'white'})
f, ax = plt.subplots(figsize = (8,6))
sns.heatmap(matrix, vmax=.8,annot_kws={'size': 10}, annot=True, fmt='.2f')
plt.show()
# Um ponto importante a se observar é a alta correlação entre a categoria dos produtos. Isso já era esperado uma vez que foi dito que um produto pode pertencer a mais de uma categoria.
# Meu objetivo principal foi mostrar como podemos extrair importantes informações através de técnicas estatísticas e visualização de dados.
# Abaixo vou resumir o que foi descoberto.
# - A maioria dos usuários é do sexo masculino
# - Com idade entre 26 a 30 anos
# - Não são casados
# - Residem na cidade de categoria B
# - Moram 1 ano na cidade atual
# - E compraram mais produtos da categoria 1.
# + [markdown] colab={} colab_type="code" id="8bR9vcWXo7e-"
# # Criando o Modelo
# + colab={} colab_type="code" id="UB6L_8u2zcOV"
# Carregando os dados de treino e teste
df_train = pd.read_csv('Database/train.csv')
df_test = pd.read_csv('Database/test.csv')
# + colab={} colab_type="code" id="7oezcDKy0IDA"
# Criando o DataFrame de submissão
submission = pd.DataFrame()
submission[['User_ID','Product_ID']] = df_test[['User_ID','Product_ID']]
# + [markdown] colab_type="text" id="trm1PLkBpsNW"
# Durante a análise exploratória de dados (EDA), ficou claro que na coluna 'Product_Category_1' temos dois outliers pontuais, vamos removê-los.
# + colab={"base_uri": "https://localhost:8080/", "height": 374} colab_type="code" id="8Vkc1ehMF1Sg" outputId="c76e4f64-c265-45f8-b029-95e68c18b697"
df_train['Product_Category_1'].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 340} colab_type="code" id="ewzKE6DrF4_N" outputId="f9f0dad5-e1ca-489c-8d7e-fe5e68e84fda"
df_test['Product_Category_1'].value_counts()
# + [markdown] colab_type="text" id="Cxv5lLYlF8Gd"
# Como podemos observar acima só temos esses indíces no dataset de treino, então teremos que removê-los antes de concatenar esses datasets.
# + colab={} colab_type="code" id="-3mlkf3G898H"
# Removendo os outliers do dataset de treino
df_train.drop(df_train[(df_train.Product_Category_1 == 19) | (df_train.Product_Category_1 == 20)].index,inplace=True)
# + [markdown] colab_type="text" id="pZJ7tAH1vMUB"
# Todas as transformações devem ser feitas em ambos datasets e afim de facilitar esse processo vamos concatena-los sem a variável target(Purchase), assim não precisamos fazer essa tarefa 2 vezes.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="E2mZ3kLV3NFR" outputId="9c3011e9-30b3-4521-f771-0cad8c0bc5ab"
train = df_train
test = df_test
df = pd.concat([train.drop('Purchase', axis=1),test],ignore_index = True, sort = False)
target = train['Purchase']
# Verificando a dimensão dos datasets.
train.shape, test.shape, df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 292} colab_type="code" id="xzvjuhtf3NFb" outputId="f75861e9-1e37-4e74-85d4-b5dda5e07794"
df.head()
# + [markdown] colab_type="text" id="R6AT_-lbpXpM"
# Informações sobre as colunas
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="ymQqN9GX3NFe" outputId="c8d062ab-64e2-406d-c7da-59c130164bfb"
df.info()
# + [markdown] colab_type="text" id="_dKYVNDfaepl"
# Verificando a quantidade de valores nulos no dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} colab_type="code" id="lUKxzVRJ3NFy" outputId="be8b958a-e37e-446b-d4e4-910dadeb85b1"
total = df.isnull().sum().sort_values(ascending=False)
percent = df.isnull().sum()/df.isnull().count().sort_values(ascending=False)
missing_data = pd.concat([total, percent], axis=1, sort=False, keys=['total', 'percent'])
missing_data[missing_data['percent']!=0]*100
# + [markdown] colab_type="text" id="26kQHT4PVIzb"
# Preeenchendo os valores nulos da coluna 'Product_Category_2' e transformando seus valores em inteiro.
# + colab={} colab_type="code" id="s-NhTtZRkf8p"
df['Product_Category_2'] = df['Product_Category_2'].fillna(-999).astype(int)
# + [markdown] colab_type="text" id="QUj6Eg1PDFmI"
# Opa! Mais temos duas colunas com valores nulos e só estamos preenchendo uma, isso parece meio estranho. Vamos dar uma olhada nos valores únicos de cada coluna.
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="UuHM2LnaF8HK" outputId="6ba9308d-71b7-4763-df6d-0f8b8ec3b4d9"
df.apply(lambda x: len(x.unique()))
# + [markdown] colab_type="text" id="Mnn6tPtCGOgf"
# **Podemos observar que a coluna 'Product_Category_3' tem um número menor de valores únicos comparado com as outras duas colunas referentes a categoria do produto. Conforme foi informado na descrição do problema os produtos da categoria 2 e 3 podem pertencer a outra categoria também, mais isso não se aplica aos produtos da categoria 1 que é a categoria principal. Podemos observar que 2 produtos da categoria principal não pertence a categoria 3, então isso pode nos indicar que essa categoria de produtos não Influenciará no valor final da compra.**
# + [markdown] colab_type="text" id="_oZmQSgXIM-X"
# Temos vários IDs(1000001,1000002,1000004 - que inclusive se repetem) e cada um
# desses IDs está relacionado com vários produtos.
# Vamos verificar a quantidade de usuários por ID e a quantidade de produtos. Para efetuar essa contagem vamos utilizar o **transform** que faz a contagem mais retorna a mesma quantidade de dados diferentemente do aggregate que retorna uma versão reduzida dos dados. Por último vamos ver qual o valor médio de compra de cada produto.
# + colab={} colab_type="code" id="yY6psXI9pnhO"
# Número de usuários por ID.
df['User_count'] = df.User_ID.groupby(df.User_ID).transform('count')
# Quantidade de produtos.
df['Product_count'] = df.Product_ID.groupby(df.Product_ID).transform('count')
# Valor médio de compra de cada produto.
df['Product_mean'] = df['Product_ID'].map(target.groupby(train['Product_ID']).mean())
df['Product_mean'] = df['Product_mean'].replace(np.nan, 0)
# + [markdown] colab_type="text" id="pzoRIq1oVAmi"
# Transformando os dados da coluna 'Age' em numéricos com a média aritmética entre os valores.
# + colab={} colab_type="code" id="T5f5Z7Zu3NGA"
age_dict = {'0-17':17, '18-25':21, '26-35':30, '36-45':40, '46-50':48, '51-55':53, '55+':60}
df["Age"] = df["Age"].apply(lambda x: age_dict[x])
# + [markdown] colab_type="text" id="HCq0e0JyU1IL"
# Na coluna 'Stay_In_Current_City_Years' temos uma string acompanhada de um caracter especial, vamos trocar o valor dessa string por 5 já que ela está como '4+' ao meu ver isso significa que é mais de 4 por isso vou preencher com o valor 5 e transformar seus dados para inteiros.
# + colab={} colab_type="code" id="V_OqGM5za5qs"
df['Stay_In_Current_City_Years'] = df['Stay_In_Current_City_Years'].replace('4+', 5).astype(int)
# + [markdown] colab_type="text" id="vGu2GxEvB0D3"
# Transformando as variáveis categóricas nominais em numéricas. Estou usando o **drop_first** para evitar o problema de Dummy Variable trap.
# + colab={} colab_type="code" id="ClRYEbS1eEfz"
df = pd.get_dummies(df, columns=['Gender','City_Category'], drop_first=True)
# + [markdown] colab_type="text" id="ZCZxCM-eaqgZ"
# Visualizando se os dados foram preenchidos corretamente.
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="jynNtAvD3NF5" outputId="920dbdde-2aee-4d5a-c930-9ce292f44ac3"
df.head()
# + [markdown] colab_type="text" id="lqxfJ-M0j9Ve"
# Agora vamos iremos remover as colunas 'User_ID' e 'Product_ID', pois elas só contém a identificação dos usuários e dos produtos, e a coluna 'Product_Category_3' que como vimos acima não tem uma relação total com a categoria principal e não serão necessárias no nosso modelo.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 689} colab_type="code" id="j3aucnI-3NGD" outputId="18f50667-cf7c-437d-dab6-cb214670a314"
def remove_features(lista_features):
for i in lista_features:
df.drop(i, axis=1, inplace=True)
remove_features(['User_ID','Product_ID','Product_Category_3'])
df.head(n=20)
# + [markdown] colab_type="text" id="AmOsua7pQJTa"
# Verificando se todas as variáveis foram transformadas em numéricas.
# + colab={"base_uri": "https://localhost:8080/", "height": 238} colab_type="code" id="8FzbKil_3NGN" outputId="5277deec-a308-459b-fe31-4ed19f92f2da"
df.dtypes
# + [markdown] colab_type="text" id="L7lZa7GTRc4I"
# Depois de fazer todas as transformações necessárias nos dados
# vamos dividir o dataset em dados de treino e teste novamente.
# + colab={} colab_type="code" id="KJ_kHPob3NGg"
X_train = df[:545915]
X_test = df[545915:]
# + [markdown] colab_type="text" id="9H-V2V0UWR9b"
# Dataset de treino
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="VfR9iin83NGk" outputId="e9bcd2fd-b71e-46a2-efbd-9ed8abc0287a"
X_train.head()
# + [markdown] colab_type="text" id="m8NTvKMKWUe6"
# Dataset de teste
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="L74YgMLm3NGn" outputId="0b5adc7b-cfa4-4c26-a428-185f2ff4e01e"
X_test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="eA1Q89VA3NGq" outputId="d1e40072-681d-4dc6-dfde-da7de8a259e7"
X_train.shape, X_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="sJchOKZQ3NG0" outputId="424506cf-ba53-4d5c-a725-c1f48cfef7cd"
target.values
# + [markdown] colab_type="text" id="VUMLvznbIP-9"
# #Avaliação dos Algoritmos
# + [markdown] colab_type="text" id="C53j9ajWL1bc"
# Vamos criar pipelines afim de automatizar o processo de treinamento dos nossos modelos e aplicar a **Standartization** (padronização) ao conjunto de dados (colocando-os na mesma escala)
# Nesta técnica, os dados serão transformados de modo que estejam com uma distrbuição normal, com média igual a zero e
# desvio padrão igual a 1.
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="Ko9fElTo3NHM" outputId="631a356a-2fc1-48d7-84c9-52cf741df5fe"
pipelines = []
pipelines.append(('Scaled-LR', Pipeline([('Scaler', StandardScaler()),('LR', LinearRegression())])))
pipelines.append(('Scaled-LASSO', Pipeline([('Scaler', StandardScaler()),('LASSO', Lasso())])))
pipelines.append(('Scaled-EN', Pipeline([('Scaler', StandardScaler()),('EN', ElasticNet())])))
pipelines.append(('Scaled-CART', Pipeline([('Scaler', StandardScaler()),('CART', DecisionTreeRegressor())])))
pipelines.append(('Scaled-XGBoost', Pipeline([('Scaler', StandardScaler()),('XG', XGBRegressor(objective ='reg:squarederror'))])))
pipelines.append(('Scaled-AB', Pipeline([('Scaler', StandardScaler()),('AB', AdaBoostRegressor())])))
pipelines.append(('Scaled-GBM', Pipeline([('Scaler', StandardScaler()),('GBM', GradientBoostingRegressor())])))
pipelines.append(('Scaled-RF', Pipeline([('Scaler', StandardScaler()),('RF', RandomForestRegressor())])))
pipelines.append(('Scaled-ET', Pipeline([('Scaler', StandardScaler()),('ET', ExtraTreesRegressor())])))
resultados = []
nomes = []
# Percorrendo cada um dos modelos
for nome, modelo in pipelines:
kfold = model_selection.KFold(n_splits=5, shuffle=True, random_state = 7)
cross_val_result = model_selection.cross_val_score(modelo, X_train, target, cv = kfold, scoring = 'neg_mean_squared_error')
resultados.append(cross_val_result)
nomes.append(nome)
texto = "%s: %f (%f)" % (nome, np.sqrt(np.abs(cross_val_result)).mean(), np.sqrt(-cross_val_result).std())
print(texto)
# + [markdown] colab_type="text" id="xBZqGDLSB_m0"
# O GradientBoostingRegressor e o XGBRegressor apresentaram a menor taxa de erro entre os modelos
# + [markdown] colab_type="text" id="z_2W4pBeJFb4"
# Vamos agora fazer o tuning do modelo utilizando o RandomizedSearchCV, que de acordo com a documentação do scikit-learn tem o tempo de execução
# drasticamente menor que o GridSearchCV. Foi testado vários valores para os parâmetros e deixei aqui só os valores que apareceram com mais frequência (pra ajudar a diminuir o tempo de processamento).
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="Bv8-7lsJjL2C" outputId="37d66960-9eec-4ea7-e1fc-788b7c747865"
# Definindo a escala
X_train = StandardScaler().fit_transform(X_train)
# Testando valores para o estimador.
valores_grid = {'n_estimators' : [100,200,300,400], 'max_depth' : [7,9],'min_child_weight': [5,7], 'gamma': [0.1,0.3]}
# Criando o modelo
modelo = XGBRegressor(objective ='reg:squarederror', n_jobs=4)
# Definindo k
kfold = model_selection.KFold(5, True, random_state = 7)
# Testando a combinação de parâmetros
grid = RandomizedSearchCV(modelo, valores_grid, cv = kfold, scoring = 'neg_mean_squared_error')
grid_result = grid.fit(X_train, target)
# Print do resultado
print("Grid scores on development set:")
means = grid.cv_results_['mean_test_score'].round(5)
stds = grid.cv_results_['std_test_score'].round(5)
for mean, std, params in zip((means), stds, grid.cv_results_['params']):
print(f'mean:{mean},std:{std},params:{params}')
print()
print(f'Melhor parâmetro:{grid.best_params_}, Score:{grid.best_score_}')
# + [markdown] colab_type="text" id="7UGXrbfZFbAQ"
# **É claro que não me deixei levar pelas vantagens do XGBRegressor e também fiz o tuning do GradientBoostingRegressor, e de fato obtive uma melhor pontuação com o XGBRegressor.
# Como o processo é um pouco demorado optei por mostrar somente um dos processos.**
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="5eD3ZzpKKDdS" outputId="90bd376e-ceee-4d68-b7b4-81d5e38fac6a"
# Preparando a versão final do modelo
X_train = StandardScaler().fit_transform(X_train)
modelo_xbg = XGBRegressor(objective ='reg:squarederror', n_estimators = 300,
min_child_weight=5,
max_depth=7,
gamma=0.3,
random_state=7,
n_jobs=4)
modelo_xbg.fit(X_train, target)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="yw5GRiOCRTDu" outputId="d4d961e6-1ea6-4ce9-f8b3-417328f37a6b"
# Aplicando o modelo aos dados de teste
X_test = StandardScaler().fit_transform(X_test)
previsoes = modelo_xbg.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="OlS5PYhF0Vmo" outputId="76e24e24-3137-422b-f6c2-44267639a312"
# Visualizando os resultados
submission['User_ID','Product_ID'] = np.around(previsoes,2)
submission = submission.rename(columns={('User_ID','Product_ID'):'Purchase'})
submission.head()
# + colab={} colab_type="code" id="UlxmlO9X7chA"
# Submissão
submission.to_csv("sample_submission.csv", index=False)
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="FpHPaEEoVSUn" colab_type="text"
# # DCGAN sample using tensorflow
#
# tensorflow を利用して MNIST で DCGAN を実行するサンプルです。
#
# - [Deep Convolutional Generative Adversarial Network][tutorial]
#
# [tutorial]: https://www.tensorflow.org/tutorials/generative/dcgan
# + [markdown] id="lcmZkCYOLQvk" colab_type="text"
# ## 環境の確認
# + id="LnMGmEyyLT2f" colab_type="code" outputId="45144fc0-8267-4774-f710-2089ad258944" colab={"base_uri": "https://localhost:8080/", "height": 53}
# !cat /etc/issue
# + id="UDMUXlaTLpG2" colab_type="code" outputId="913796cf-f9a4-4bf2-8ae6-a0529466b97e" colab={"base_uri": "https://localhost:8080/", "height": 71}
# !free -h
# + id="WHn3f12zLsTw" colab_type="code" outputId="b16b4deb-3d0f-4fdf-a0df-7280ffdc9911" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !cat /proc/cpuinfo
# + id="3FUZPKAcL4jO" colab_type="code" outputId="fe8f507b-fad4-455e-87e0-de49a1de987f" colab={"base_uri": "https://localhost:8080/", "height": 323}
# !nvidia-smi
# + id="w2V3sVtNPYdT" colab_type="code" outputId="a6fbfacf-2f49-420a-a82b-7a0c9125b066" colab={"base_uri": "https://localhost:8080/", "height": 35}
# !python --version
# + id="Zyf3UfsENhOf" colab_type="code" colab={}
from logging import Logger
def get_logger() -> Logger:
import logging
logger = logging.getLogger(__name__)
fmt = "%(asctime)s %(levelname)s %(name)s :%(message)s"
logging.basicConfig(level=logging.INFO, format=fmt)
return logger
logger = get_logger()
# + id="EppKsLuHMYjR" colab_type="code" outputId="82552424-df85-48da-f25a-6b66654880aa" colab={"base_uri": "https://localhost:8080/", "height": 35}
def check_tf_version() -> None:
import tensorflow as tf
logger.info(tf.__version__)
check_tf_version()
# + [markdown] id="SKyYk7oMY21S" colab_type="text"
# ## ソースコードの取得
# + id="PjgDKO_RV_U5" colab_type="code" outputId="a80b4b94-64d3-450f-84f6-c65ba3889e30" colab={"base_uri": "https://localhost:8080/", "height": 395}
# 対象のコードを取得
# !git clone -n https://github.com/iimuz/til.git
# %cd til
# !git checkout fdfa134
# %cd python/dcgan_tensorflow
# + [markdown] id="_vUlewGLYtT1" colab_type="text"
# ## 実行
# + [markdown] id="J4llcBuDKMus" colab_type="text"
# ### 事前準備
# + id="H8Ng5oAwIevW" colab_type="code" colab={}
import tensorflow.compat.v1 as tfv1
tfv1.enable_eager_execution()
# + [markdown] id="NDZFpP_IKKCr" colab_type="text"
# ### データセットの確認
# + id="8AwAXG4UHZ7V" colab_type="code" outputId="ce17000a-df4a-48f1-dd97-d4895f0cf8e3" colab={"base_uri": "https://localhost:8080/", "height": 599}
# %run -i dataset.py
# + id="nv1OyOv3I9f4" colab_type="code" outputId="4c17d097-54b0-45e8-9592-50f57c0e91e2" colab={"base_uri": "https://localhost:8080/", "height": 91}
import dataset
raw_train, raw_test = dataset.get_batch_dataset()
# + [markdown] id="leX-NiQxKRHh" colab_type="text"
# ### ネットワークの確認
# + id="sLKS5JeWHdNl" colab_type="code" outputId="5d339bc2-6492-4a75-daa5-59bfdd2dd6a4" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %run -i network.py
# + [markdown] id="lpsueOcOJRrK" colab_type="text"
# ### 学習の実行
# + id="H60exsmWJWPo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="0ada8b18-fcb6-4632-fe63-5ccc76fee25d"
import train
train.train(raw_train, batch_size=256, epochs=100, gen_input_dim=100, disc_input_shape=(28, 28, 1))
# + [markdown] id="0QiTCqIlJtMG" colab_type="text"
# ### 結果
# + id="J83wMaOaJvSA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="5bbc75d7-a678-47fd-80a1-249a955ff5bc"
import utils
import IPython
from IPython import display
def show_generated_images():
filepath = "_data/dcgan.gif"
utils.save_gif("_data/", "image_at_epoch_*", filepath)
try:
from google.colab import files
except ImportError:
pass
else:
files.download(filepath)
show_generated_images()
| machine_learning/tf_dcgan/dcgan_tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/proyectosRVyderivados/cristina/blob/main/VAN_TIR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="lTY98AgnoQgV" outputId="ba7c1458-fd75-4dd5-994f-37e83eab7d15"
pip install numpy-financial #instalo libreria financiera.
# + [markdown] id="xCYZ0ULHoni9"
# #VAN y TIR:
# * VAN = NPV
# * TIR = IRR
#
# Calcular el VAN y la TIR partiendo de los flujos de caja. (El guión bajo lo puedo utilizar como separador de miles).
#
# Flujos = [-desemboldo inicial, flujos que recibo] = [-600_000, 100_000, 150_000, 200_000, 250_000, 300_000]
#
# Rentabilidad = tasa de descuento = r = 0.1 (hay que ponerlo con punto que es el separador anglosajón)
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="a8nVVml-pT5i" outputId="68a62969-8d9a-4167-a19e-65e8c86de5c7"
import numpy_financial as npf #importo libreria y pongo abreviatura (para esta librería será npf)
cash_flows = [-600_000, 100_000, 150_000, 200_000, 250_000, 300_000]
tasa_descuento = 0.1
#VAN con libreria:
van = npf.npv(tasa_descuento, cash_flows)
print(f"Valor actual neto ({tasa_descuento:.2%}) = {van:,.2f} €")
#TIR con libreria:
tir = npf.irr(cash_flows)
print(f"TIR: {tir:.2%}")
# + colab={"base_uri": "https://localhost:8080/"} id="rHah9_cwtSdi" outputId="fde24df7-f971-468c-f3b9-fce1e013b176"
Desembolso_inicial = cash_flows[0] # guardamos en una variable el desembolso inicial, que esta en la posición 0 de la lista.
cash_flows[0] = 0 # en la lista de cash flows hacemos cero el desembolso inicial
van = Desembolso_inicial + npf.npv(tasa_descuento, cash_flows)
print(f"VAN({tasa_descuento:.2%}) = {van:,.2f} €") # obtenemos el van esperado, ahora sin formato
# + [markdown] id="prDbtLG1rzZr"
# #Programar el VAN:
# En una lista: l=[10,20,30]
#
# posición: 0 1 2
# Vamor a hacer la función que se hará primero, luego pondremos los parámetros y finalmente nos devolverá la solución.
#
# + colab={"base_uri": "https://localhost:8080/"} id="ts3yn1T5sIFQ" outputId="d0074ac5-2c53-4c7d-fbd6-0394fa8e7e1d"
#función:
def van(tasa, flujos):
total = 0
for i, flujo in enumerate(flujos):
total += flujo / (1 + tasa)**(i)
return total
#Pongo los parámetros:
tasa = 0.1
flujos = [-600_000, 100_000, 150_000, 200_000, 250_000, 300_000]
#Imprimo el resultado:
print(f"VAN = {van(tasa, flujos):,.2f} €")
# + [markdown] id="PIg38PZav-WR"
# #Función Pago (pmt)
# Préstamo de tipo francés.
# npf.pmt(rate, nper, pv, fv, when = ‘end’)
#
# rate: tipo de interes
#
# nper: número periodos
#
# pv: desembolso inicial
#
# fv: el valor residual o final que tienes que pagar para liquidar el prestamo
#
# when =: (pospagable) 'end'
#
# * Calcular el pago mensual periódico necesario para amortizar un préstamo de 350.000 €, a 20 años, al 6% TIN (Tipo de Interés Nominal) con Excel: =PAGO(6%/12;20*12;-350000)
# + colab={"base_uri": "https://localhost:8080/"} id="Oe3sFUns4I36" outputId="82b854d3-1586-42a0-bcd4-d54fd5bb8a8d"
#importo librería
import numpy_financial as npf
pago = npf.pmt(0.06/12, 20 * 12, 350_000)
print(f"Pago mensual: {-pago:,.2f} €" )
| VAN_TIR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Synapse PySpark
# name: synapse_pyspark
# ---
# # Using Azure Open Datasets in Synapse - Enrich NYC Green Taxi Data with Holiday and Weather
#
# Synapse has [Azure Open Datasets](https://azure.microsoft.com/en-us/services/open-datasets/) package pre-installed. This notebook provides examples of how to enrich NYC Green Taxi Data with Holiday and Weather with focusing on :
# - read Azure Open Dataset
# - manipulate the data to prepare for further analysis, including column projection, filtering, grouping and joins etc.
# - create a Spark table to be used in other notebooks for modeling training
# ## Data loading
# Let's first load the [NYC green taxi trip records](https://azure.microsoft.com/en-us/services/open-datasets/catalog/nyc-taxi-limousine-commission-green-taxi-trip-records/). The Open Datasets package contains a class representing each data source (NycTlcGreen for example) to easily filter date parameters before downloading.
# +
from azureml.opendatasets import NycTlcGreen
from datetime import datetime
from dateutil import parser
end_date = parser.parse('2018-06-06')
start_date = parser.parse('2018-05-01')
nyc_tlc = NycTlcGreen(start_date=start_date, end_date=end_date)
nyc_tlc_df = nyc_tlc.to_spark_dataframe()
# +
# Display 5 rows
nyc_tlc_df.show(5, truncate = False)
# -
# Now that the initial data is loaded. Let's do some projection on the data to
# - create new columns for the month number, day of month, day of week, and hour of day. These info is going to be used in the training model to factor in time-based seasonality.
# - add a static feature for the country code to join holiday data.
# +
# Extract month, day of month, and day of week from pickup datetime and add a static column for the country code to join holiday data.
import pyspark.sql.functions as f
nyc_tlc_df_expand = nyc_tlc_df.withColumn('datetime',f.to_date('lpepPickupDatetime'))\
.withColumn('month_num',f.month(nyc_tlc_df.lpepPickupDatetime))\
.withColumn('day_of_month',f.dayofmonth(nyc_tlc_df.lpepPickupDatetime))\
.withColumn('day_of_week',f.dayofweek(nyc_tlc_df.lpepPickupDatetime))\
.withColumn('hour_of_day',f.hour(nyc_tlc_df.lpepPickupDatetime))\
.withColumn('country_code',f.lit('US'))
# -
# Remove some of the columns that won't need for modeling or additional feature building.
#
#
#
# +
# Remove unused columns from nyc green taxi data
columns_to_remove = ["lpepDropoffDatetime", "puLocationId", "doLocationId", "pickupLongitude",
"pickupLatitude", "dropoffLongitude","dropoffLatitude" ,"rateCodeID",
"storeAndFwdFlag","paymentType", "fareAmount", "extra", "mtaTax",
"improvementSurcharge", "tollsAmount", "ehailFee", "tripType "
]
nyc_tlc_df_clean = nyc_tlc_df_expand.select([column for column in nyc_tlc_df_expand.columns if column not in columns_to_remove])
# -
# Display 5 rows
nyc_tlc_df_clean.show(5)
# ## Enrich with holiday data
# Now that we have taxi data downloaded and roughly prepared, add in holiday data as additional features. Holiday-specific features will assist model accuracy, as major holidays are times where taxi demand increases dramatically and supply becomes limited.
#
# Let's load the [public holidays](https://azure.microsoft.com/en-us/services/open-datasets/catalog/public-holidays/) from Azure Open datasets.
#
# +
from azureml.opendatasets import PublicHolidays
hol = PublicHolidays(start_date=start_date, end_date=end_date)
hol_df = hol.to_spark_dataframe()
# Display data
hol_df.show(5, truncate = False)
# -
# Rename the countryRegionCode and date columns to match the respective field names from the taxi data, and also normalize the time so it can be used as a key.
# +
hol_df_clean = hol_df.withColumnRenamed('countryRegionCode','country_code')\
.withColumn('datetime',f.to_date('date'))
hol_df_clean.show(5)
# -
# Next, join the holiday data with the taxi data by performing a left-join. This will preserve all records from taxi data, but add in holiday data where it exists for the corresponding datetime and country_code, which in this case is always "US". Preview the data to verify that they were merged correctly.
# +
# enrich taxi data with holiday data
nyc_taxi_holiday_df = nyc_tlc_df_clean.join(hol_df_clean, on = ['datetime', 'country_code'] , how = 'left')
nyc_taxi_holiday_df.show(5)
# +
# Create a temp table and filter out non empty holiday rows
nyc_taxi_holiday_df.createOrReplaceTempView("nyc_taxi_holiday_df")
spark.sql("SELECT * from nyc_taxi_holiday_df WHERE holidayName is NOT NULL ").show(5, truncate = False)
# -
# ## Enrich with weather data¶
#
# Now we append NOAA surface weather data to the taxi and holiday data. Use a similar approach to fetch the [NOAA weather history data](https://azure.microsoft.com/en-us/services/open-datasets/catalog/noaa-integrated-surface-data/) from Azure Open Datasets.
# +
from azureml.opendatasets import NoaaIsdWeather
isd = NoaaIsdWeather(start_date, end_date)
isd_df = isd.to_spark_dataframe()
# -
isd_df.show(5, truncate = False)
# +
# Filter out weather info for new york city, remove the recording with null temperature
weather_df = isd_df.filter(isd_df.latitude >= '40.53')\
.filter(isd_df.latitude <= '40.88')\
.filter(isd_df.longitude >= '-74.09')\
.filter(isd_df.longitude <= '-73.72')\
.filter(isd_df.temperature.isNotNull())\
.withColumnRenamed('datetime','datetime_full')
# +
# Remove unused columns
columns_to_remove_weather = ["usaf", "wban", "longitude", "latitude"]
weather_df_clean = weather_df.select([column for column in weather_df.columns if column not in columns_to_remove_weather])\
.withColumn('datetime',f.to_date('datetime_full'))
weather_df_clean.show(5, truncate = False)
# -
# Next group the weather data so that you have daily aggregated weather values.
#
# +
# Enrich weather data with aggregation statistics
aggregations = {"snowDepth": "mean", "precipTime": "max", "temperature": "mean", "precipDepth": "max"}
weather_df_grouped = weather_df_clean.groupby("datetime").agg(aggregations)
# -
weather_df_grouped.show(5)
# +
# Rename columns
weather_df_grouped = weather_df_grouped.withColumnRenamed('avg(snowDepth)','avg_snowDepth')\
.withColumnRenamed('avg(temperature)','avg_temperature')\
.withColumnRenamed('max(precipTime)','max_precipTime')\
.withColumnRenamed('max(precipDepth)','max_precipDepth')
# -
# Merge the taxi and holiday data you prepared with the new weather data. This time you only need the datetime key, and again perform a left-join of the data. Run the describe() function on the new dataframe to see summary statistics for each field.
# enrich taxi data with weather
nyc_taxi_holiday_weather_df = nyc_taxi_holiday_df.join(weather_df_grouped, on = 'datetime' , how = 'left')
nyc_taxi_holiday_weather_df.cache()
nyc_taxi_holiday_weather_df.show(5)
# +
# Run the describe() function on the new dataframe to see summary statistics for each field.
display(nyc_taxi_holiday_weather_df.describe())
# -
# The summary statistics shows that the totalAmount field has negative values, which don't make sense in the context.
#
#
# Remove invalid rows with less than 0 taxi fare or tip
final_df = nyc_taxi_holiday_weather_df.filter(nyc_taxi_holiday_weather_df.tipAmount > 0)\
.filter(nyc_taxi_holiday_weather_df.totalAmount > 0)
# ## Cleaning up the existing Database
#
# First we need to drop the tables since Spark requires that a database is empty before we can drop the Database.
#
# Then we recreate the database and set the default database context to it.
spark.sql("DROP TABLE IF EXISTS NYCTaxi.nyc_taxi_holiday_weather");
spark.sql("DROP DATABASE IF EXISTS NYCTaxi");
spark.sql("CREATE DATABASE NYCTaxi");
spark.sql("USE NYCTaxi");
# ## Creating a new table
# We create a nyc_taxi_holiday_weather table from the nyc_taxi_holiday_weather dataframe.
#
# +
from pyspark.sql import SparkSession
from pyspark.sql.types import *
final_df.write.saveAsTable("nyc_taxi_holiday_weather");
spark.sql("SELECT COUNT(*) FROM nyc_taxi_holiday_weather").show();
| Sample/OpenDatasets/Notebooks/01-UsingOpenDatasetsSynapse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:basepair]
# language: python
# name: conda-env-basepair-py
# ---
from __future__ import division, print_function
from importlib import reload
import abstention
reload(abstention)
reload(abstention.calibration)
reload(abstention.label_shift)
from abstention.calibration import TempScaling, ConfusionMatrix, softmax
from abstention.label_shift import EMImbalanceAdapter, BBSEImbalanceAdapter, ShiftWeightFromImbalanceAdapter
import glob
import gzip
import json
import numpy as np
from scipy.spatial import distance
from collections import defaultdict
import matplotlib.pyplot as plt
# %matplotlib inline
loaded_dicts = json.loads(gzip.open("label_shift_adaptation_results.json.gz").read())
metric_to_samplesize_to_calibname_to_unshiftedvals =\
loaded_dicts['metric_to_samplesize_to_calibname_to_unshiftedvals']
# +
x = np.arange(4)
methods = ['TS', 'NBVS', 'BCTS', 'VS']
font = {'family' : 'normal',
'weight' : 'bold',
'size' : 22}
plt.rc('font', **font)
for metric in metric_to_samplesize_to_calibname_to_unshiftedvals:
for size in metric_to_samplesize_to_calibname_to_unshiftedvals[metric]:
print(metric)
print(size)
y = [np.mean(np.array(metric_to_samplesize_to_calibname_to_unshiftedvals[metric][size][method])) for method in methods]
error = [np.std(np.array(metric_to_samplesize_to_calibname_to_unshiftedvals[metric][size][method])) for method in methods]
fig, ax = plt.subplots()
ax.bar(x, y, yerr=error, align='center', alpha=0.5, ecolor='black', capsize=10)
ax.set_ylabel('JS Divergence')
ax.set_xticks(x)
ax.set_xticklabels(methods)
ax.set_title('CIFAR10')
ax.yaxis.grid(True)
plt.tight_layout()
plt.show()
| cifar10/intro_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import gensim
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import pickle
from tensorflow.keras.models import Sequential, load_model, Model
from tensorflow.keras.layers import Dense
from tensorflow.keras.callbacks import ModelCheckpoint, EarlyStopping
from tensorflow.keras.layers import Dropout, Input
from tensorflow.keras.regularizers import l1
ROOT = os.path.dirname(os.getcwd())
#path_data = os.path.join(ROOT, 'data')
path_data = 'C:\\Users\\joris\\Documents\\eScience_data\\data'
sys.path.insert(0, ROOT)
sys.path.insert(0, "C:\\Users\\joris\\Documents\\eScience_data\\spec2vec_gnps_data_analysis\\custom_functions")
# -
# ## Loading data ready for training
# -Creation of this data can be seen in 3b
# training set
outfile = os.path.join(path_data, 'nn_prep_training_found_matches_s2v_2dec.pickle')
print(outfile)
if os.path.exists(outfile):
with open(outfile, 'rb') as inf:
nn_prep_training_found_matches_s2v_2dec = pickle.load(inf)
else:
nn_prep_training_found_matches_s2v_2dec = find_info_matches(old_and_unique_found_matches_s2v_2dec_top20,
old_and_unique_documents_library_s2v_2dec,
old_and_unique_documents_query_s2v_2dec,
max_parent_mass=max_parent_mass)
with open(outfile, 'wb') as outf:
pickle.dump(nn_prep_training_found_matches_s2v_2dec, outf)
outfile = os.path.join(path_data, 'nn_prep_testing_found_matches_s2v_2dec.pickle')
print(outfile)
if os.path.exists(outfile):
with open(outfile, 'rb') as inf:
nn_prep_testing_found_matches_s2v_2dec = pickle.load(inf)
else:
nn_prep_testing_found_matches_s2v_2dec = find_info_matches(new_and_unique2_found_matches_s2v_2dec_top20,
new_and_unique2_documents_library_s2v_2dec,
new_and_unique2_documents_query_s2v_2dec,
max_parent_mass=max_parent_mass)
with open(outfile, 'wb') as outf:
pickle.dump(nn_prep_testing_found_matches_s2v_2dec, outf)
nn_prep_testing_found_matches_s2v_2dec[2].iloc[:5]
# ## Add matches together for each query
# +
# add all the found matches together in one big df
nn_training_found_matches_s2v_2dec = nn_prep_training_found_matches_s2v_2dec[0].append(
nn_prep_training_found_matches_s2v_2dec[1:])
nn_training_found_matches_s2v_2dec = nn_training_found_matches_s2v_2dec.sample(frac=1)
nn_testing_full_found_matches_s2v_2dec = nn_prep_testing_found_matches_s2v_2dec[0].append(
nn_prep_testing_found_matches_s2v_2dec[1:])
nn_testing_full_found_matches_s2v_2dec = nn_testing_full_found_matches_s2v_2dec.sample(frac=1)
# take 300 (randomly chosen) query matches for validation set
np.random.seed(42)
first_half = list(np.random.choice(range(0,1000), 250, replace=False))
np.random.seed(42)
second_half = list(np.random.choice(range(1000,2000), 250, replace=False))
val_set = first_half + second_half
nn_val_found_matches_s2v_2dec = pd.DataFrame()
nn_testing_found_matches_s2v_2dec = pd.DataFrame()
for i in range(len(nn_prep_testing_found_matches_s2v_2dec)):
if i in val_set:
nn_val_found_matches_s2v_2dec = nn_val_found_matches_s2v_2dec.append(nn_prep_testing_found_matches_s2v_2dec[i])
else:
nn_testing_found_matches_s2v_2dec = nn_testing_found_matches_s2v_2dec.append(nn_prep_testing_found_matches_s2v_2dec[i])
nn_val_found_matches_s2v_2dec = nn_val_found_matches_s2v_2dec.sample(frac=1)
nn_testing_found_matches_s2v_2dec = nn_testing_found_matches_s2v_2dec.sample(frac=1)
# -
second_half
plt.hist(nn_training_found_matches_s2v_2dec['similarity'],
alpha=0.5, bins = np.arange(0,1,0.05), label = 'training set')
plt.hist(nn_testing_found_matches_s2v_2dec['similarity'],
alpha=0.5, bins = np.arange(0,1,0.05), label = 'test subset')
plt.hist(nn_val_found_matches_s2v_2dec['similarity'],
alpha=0.5, bins = np.arange(0,1,0.05), label = 'validation set')
plt.legend()
plt.show()
X_train = nn_training_found_matches_s2v_2dec.drop(['similarity', 'label'], axis = 1)
y_train = nn_training_found_matches_s2v_2dec['similarity']
X_test = nn_testing_found_matches_s2v_2dec.drop(['similarity', 'label'], axis = 1)
y_test = nn_testing_found_matches_s2v_2dec['similarity']
X_val = nn_val_found_matches_s2v_2dec.drop(['similarity', 'label'], axis = 1)
y_val = nn_val_found_matches_s2v_2dec['similarity']
# ## Load model from notebook 3
# +
#nn function
def train_nn(X_train, y_train, X_test, y_test, layers = [12, 12, 12, 12, 12, 1],
model_loss = 'binary_crossentropy', activations = 'relu',
last_activation = 'sigmoid', model_epochs = 20, model_batch_size = 16,
save_name = False):
'''Train a keras deep NN and test on test data, returns (model, history, accuracy, loss)
X_train: matrix like object like pd.DataFrame, training set
y_train: list like object like np.array, training labels
X_test: matrix like object like pd.DataFrame, test set
y_test: list like object like np.array, test labels
layers: list of ints, the number of layers is the len of this list while the elements
are the amount of neurons per layer, default: [12, 12, 12, 12, 12, 1]
model_loss: str, loss function, default: binary_crossentropy
activations: str, the activation of the layers except the last one, default: relu
last_activation: str, activation of last layer, default: sigmoid
model_epochs: int, number of epochs, default: 20
model_batch_size: int, batch size for updating the model, default: 16
save_name: str, location for saving model, optional, default: False
Returns:
model: keras sequential
history: dict, training statistics
accuracy: float, accuracy on test set
loss, float, loss on test set
If save_name is not False and save_name exists this function will load existing model
'''
if os.path.exists(save_name) and save_name:
print('\nLoading existing model')
nn_model = load_model(save_name)
with open(save_name + '_train_hist.pickle', 'rb') as hist_inf:
history = pickle.load(hist_inf)
else:
# define the keras model
nn_model = Sequential()
#add first layer
nn_model.add(Dense(layers[0], input_dim = X_train.shape[1], activation = activations))
#add other layers
for i in range(1,len(layers)-1): #skip first and last one
nn_model.add(Dense(layers[i], activation = activations))
#add last layer
nn_model.add(Dense(layers[-1], activation = last_activation))
# compile the keras model
nn_model.compile(loss = model_loss, optimizer='adam', metrics=['accuracy'])
# fit the keras model on the dataset
hist = nn_model.fit(X_train, y_train, epochs = model_epochs, batch_size = model_batch_size)
history = hist.history
#training set
print('Training loss: {:.4f}\n'.format(history['loss'][-1]))
#test set
loss, accuracy = nn_model.evaluate(X_test, y_test)
print('Test accuracy: {:.2f}'.format(accuracy*100))
print('Test loss: {:.4f}'.format(loss))
if save_name and not os.path.exists(save_name):
print('Saving model at:', save_name)
nn_model.save(save_name)
with open(save_name + '_train_hist.pickle', 'wb') as hist_outf:
pickle.dump(history, hist_outf)
return nn_model, history, accuracy, loss
# +
test_layers = [10,24,1]
#model_name = os.path.join(path_data, 'nn_2000_queries_top20_1')
model_name = os.path.join(path_data, 'nn_2000_queries_top20_layers_opt_1') #this one tested best in notebook 3
nn_2000_queries_top20_1 = train_nn(X_train, y_train, X_test, y_test, layers = test_layers,
model_loss = 'mean_squared_error', activations = 'relu', last_activation = None,
model_epochs = 50, model_batch_size = 16, save_name = model_name)
# -
# ## Model trimming
# - Simple model
# - L1 regularisation
# - Dropout regularistion
# +
layers = [48,48,1]
model_name = os.path.join(path_data, "nn_2000_queries_trimming")
# define the keras model
nn_model = Sequential()
#add first layer
nn_model.add(Dense(layers[0], input_dim = X_train.shape[1], activation = 'relu'))
#add other layers
for i in range(1,len(layers)-1): #skip first and last one
nn_model.add(Dense(layers[i], activation = 'relu'))
#add last layer
nn_model.add(Dense(layers[-1], activation = None))
# compile the keras model
nn_model.compile(loss = 'mean_squared_error', optimizer='adam', metrics=['mae'])
# save best model
checkpointer = ModelCheckpoint(filepath= model_name + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=10, verbose=1) # patience - try x more epochs to improve val_loss
# fit the keras model on the dataset
hist = nn_model.fit(X_train, y_train, epochs = 100, batch_size = 24, validation_data=(X_val, y_val),
callbacks = [checkpointer, earlystopper])
history = hist.history
# +
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True, figsize=(12,8), dpi=100)
ax1.plot(hist.history['mae'], "o--", label='Acuracy (training data)')
ax1.plot(hist.history['val_mae'], "o--", label='Acuracy (validation data)')
ax1.set_title('MAE loss')
ax1.set_ylabel("MAE")
ax1.legend()
ax2.plot(hist.history['loss'], "o--", label='training data')
ax2.plot(hist.history['val_loss'], "o--", label='validation data')
ax2.set_title('MSE loss')
ax2.set_ylabel("MSE loss")
ax2.set_xlabel("epochs")
ax2.legend()
model_name_1 = model_name + ".hdf5"
model_1 = load_model(model_name_1)
model_1.evaluate(X_test, y_test)
# +
#Some layer optimisation again
testing_layers = [[10,1],
[16,1],
[24,1],
[48,1],
[10,10,1],
[16,16,1],
[24,24,1],
[10,24,1],
[10,48,1],
[24,48,1],
[48,48,1],
[10,96,1],
[48,96,1],
[10,10,10,1],
[24,48,8,1],
[24,24,24,1],
[10,48,10,1],
[10,24,48,1],
[24,24,48,1],
[24,48,48,1],
[48,48,48,1],
[16,16,16,16,1],
[10,24,24,10,1],
[10,24,24,10,1]]
base_model_name = os.path.join(path_data, "nn_2000_queries_trimming_simple")
simple_models = []
for i, layers in enumerate(testing_layers):
model_name_loop = f"{base_model_name}_{i}"
# define the keras model
nn_model_simple = Sequential()
#add first layer
nn_model_simple.add(Dense(layers[0], input_dim = X_train.shape[1], activation = 'relu'))
#add other layers
for i in range(1,len(layers)-1): #skip first and last one
nn_model_simple.add(Dense(layers[i], activation = 'relu'))
#add last layer
nn_model_simple.add(Dense(layers[-1], activation = None))
# compile the keras model
nn_model_simple.compile(loss = 'mean_squared_error', optimizer='adam', metrics=['mae'])
# save best model
checkpointer = ModelCheckpoint(filepath= model_name_loop + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=10, verbose=1) # patience - try x more epochs to improve val_loss
# fit the keras model on the dataset
hist_simple = nn_model_simple.fit(X_train, y_train, epochs = 100, batch_size = 24, validation_data=(X_val, y_val),
callbacks = [checkpointer, earlystopper])
simple_models.append((nn_model_simple, hist_simple, hist_simple.history['loss'][-1], hist_simple.history['val_loss'][-10]))
# +
min_val_loss = (0, 1)
for i, layers in enumerate(testing_layers):
simp_model, hist_simp, train_loss, val_loss = simple_models[i]
if val_loss < min_val_loss[1]:
min_val_loss = (i, val_loss)
print(f"Model {i}, Train loss: {hist_simp.history['loss'][-10]:.4f}, Val loss: {val_loss:.4f}")
print(f"Best model: {min_val_loss[0]}, it has {testing_layers[min_val_loss[0]]} layers")
# -
# ### Best simple model
# load the best model: #10
base_model_name = os.path.join(path_data, "nn_2000_queries_trimming_simple")
best_model_name = f"{base_model_name}_10.hdf5"
print(best_model_name)
print("Evaluation on test set")
best_model = load_model(best_model_name)
best_model.evaluate(X_test, y_test)
# +
# save best model hist
base_model_name = os.path.join(path_data, "nn_2000_queries_trimming_simple")
hist_save = f"{base_model_name}_10_hist.pickle"
print(hist_save)
if not os.path.isfile(hist_save):
with open(hist_save, 'wb') as outf:
pickle.dump(simple_models[10][1].history, outf)
else:
with open(hist_save, 'rb') as inf:
best_model_hist = pickle.load(inf)
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True, figsize=(12,8), dpi=100)
ax1.plot(best_model_hist['mae'], "o--", label='Acuracy (training data)')
ax1.plot(best_model_hist['val_mae'], "o--", label='Acuracy (validation data)')
ax1.set_title('MAE loss')
ax1.set_ylabel("MAE")
ax1.legend()
ax2.plot(best_model_hist['loss'], "o--", label='training data')
ax2.plot(best_model_hist['val_loss'], "o--", label='validation data')
ax2.set_title('MSE loss')
ax2.set_ylabel("MSE loss")
ax2.set_xlabel("epochs")
ax2.legend()
# -
# ### L1 regularisation
# +
layersl1 = [48,48,48,48,48,1]
model_namel1 = os.path.join(path_data, "nn_2000_queries_trimming_l1")
l1_val = 0.001
# define the keras model
nn_modell1 = Sequential()
#add first layer
nn_modell1.add(Dense(layersl1[0], input_dim = X_train.shape[1], activation = 'relu', activity_regularizer=l1(l1_val)))
#add other layers
for i in range(1,len(layersl1)-1): #skip first and last one
nn_modell1.add(Dense(layersl1[i], activation = 'relu', activity_regularizer=l1(l1_val)))
#add last layer
nn_modell1.add(Dense(layersl1[-1], activation = None,activity_regularizer=l1(l1_val)))
# compile the keras model
nn_modell1.compile(loss = 'mean_squared_error', optimizer='adam', metrics=['mae'])
# save best model
checkpointer = ModelCheckpoint(filepath= model_namel1 + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=10, verbose=1) # patience - try x more epochs to improve val_loss
# fit the keras model on the dataset
histl1 = nn_modell1.fit(X_train, y_train, epochs = 100, batch_size = 30, validation_data=(X_val, y_val),
callbacks = [checkpointer, earlystopper])
historyl1 = histl1.history
# +
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True, figsize=(12,8), dpi=100)
ax1.plot(histl1.history['mae'], "o--", label='MAE (training data)')
ax1.plot(histl1.history['val_mae'], "o--", label='MAE (validation data)')
ax1.set_title('MAE loss')
ax1.set_ylabel("MAE")
ax1.legend()
ax2.plot(histl1.history['loss'], "o--", label='training data')
ax2.plot(histl1.history['val_loss'], "o--", label='validation data')
ax2.set_title('MSE loss')
ax2.set_ylabel("MSE loss")
ax2.set_xlabel("epochs")
ax2.legend()
# -
# ### Dropout regularisation
# +
layers2 = [500,500,1]
model_name2 = os.path.join(path_data, "nn_2000_queries_trimming_big_dropout")
drop_rate = 0.25
# define the keras model
#nn_model2 = Sequential()
nn_input = Input(shape=X_train.shape[1])
#add first layer
nn_layers = Dense(layers2[0], activation = 'relu')(nn_input)
nn_layers = Dropout(drop_rate)(nn_layers, training = True) # 'hack': let dropout layer believe its always training mode
#add other layers
for i in range(1,len(layers2)-1): #skip first and last one
nn_layers = Dense(layers2[i], activation = 'relu')(nn_layers)
nn_layers = Dropout(drop_rate)(nn_layers, training = True)
#add last layer
nn_layers = Dense(layers2[-1], activation = None)(nn_layers)
# compile the keras model
nn_model2 = Model(inputs=[nn_input], outputs=[nn_layers])
nn_model2.compile(loss = 'mean_squared_error', optimizer='adam', metrics=['mae'])
# save best model
checkpointer = ModelCheckpoint(filepath= model_name2 + ".hdf5", monitor='val_loss', verbose=1, save_best_only=True)
earlystopper = EarlyStopping(monitor='val_loss', patience=10, verbose=1) # patience - try x more epochs to improve val_loss
# fit the keras model on the dataset
hist2 = nn_model2.fit(X_train, y_train, epochs = 100, batch_size = 24, validation_data=(X_val, y_val),
callbacks = [checkpointer, earlystopper])
history2 = hist2.history
# +
fig, (ax1, ax2) = plt.subplots(2, 1, sharex=True, figsize=(12,8), dpi=100)
ax1.plot(hist2.history['mae'], "o--", label='MAE (training data)')
ax1.plot(hist2.history['val_mae'], "o--", label='MAE (validation data)')
ax1.set_title('MAE loss')
ax1.set_ylabel("MAE")
ax1.legend()
ax2.plot(hist2.history['loss'], "o--", label='training data')
ax2.plot(hist2.history['val_loss'], "o--", label='validation data')
ax2.set_title('MSE loss')
ax2.set_ylabel("MSE loss")
ax2.set_xlabel("epochs")
ax2.legend()
| .ipynb_checkpoints/3c-model-trimming-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Simple Recursive Feature Elimination Notebook
#
# This code uses LGBMClassifier, since RFECV works with that (and not with vanilla LightGBM)
#
# References:
# - https://www.kaggle.com/tilii7/recursive-feature-elimination/code
# - https://www.kaggle.com/tilii7/features-we-don-t-need-no-stinking-features
# - https://www.kaggle.com/nroman/recursive-feature-elimination
# +
import os
import sys
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import lightgbm as lgb
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import RFECV
from sklearn.metrics import make_scorer
from sklearn.metrics import roc_auc_score
pd.options.display.max_rows = 1000
# -
sys.path.insert(0, "/opt/vssexclude/personal/kaggle/kaggle_tab_mar/src")
# %load_ext autoreload
# %autoreload 2
import munging.process_data_util as process_data
import common.com_util as common
import config.constants as constants
import modeling.train_util as model
# +
SEED = 42
TARGET = 'target'
LOGGER_NAME = 'main'
logger = common.get_logger(LOGGER_NAME)
common.set_seed(SEED)
# -
train_df, test_df, sample_submission_df = process_data.read_processed_data(
logger, constants.PROCESSED_DATA_DIR, train=True, test=True, sample_submission=True, frac=0.1)
# +
combined_df = pd.concat([train_df.drop('target', axis=1), test_df])
target = train_df[TARGET]
cat_fetaures = [name for name in train_df.columns if "cat" in name]
logger.info("Label Encoding the categorcal features")
for name in cat_fetaures:
lb = LabelEncoder()
combined_df[name] = lb.fit_transform(combined_df[name])
train_df = combined_df.loc[train_df.index]
train_df[TARGET] = target
test_df = combined_df.loc[test_df.index]
train_X = train_df.drop([TARGET], axis=1)
train_Y = train_df[TARGET]
test_X = test_df
# -
features = train_X.columns
# +
MODEL_TYPE = "lgb"
OBJECTIVE = "binary"
BOOSTING_TYPE = "gbdt"
METRIC = "auc"
VERBOSE = 100
N_THREADS = -1
NUM_LEAVES = 31
MAX_DEPTH = -1
N_ESTIMATORS = 10000
LEARNING_RATE = 0.1
EARLY_STOPPING_ROUNDS = 100
lgb_params = {
'objective': OBJECTIVE,
'boosting_type': BOOSTING_TYPE,
'learning_rate': LEARNING_RATE,
'num_leaves': NUM_LEAVES,
'tree_learner': 'serial',
'n_jobs': N_THREADS,
'seed': SEED,
'max_depth': MAX_DEPTH,
'max_bin': 255,
'metric': METRIC,
'verbose': -1,
'n_estimators': N_ESTIMATORS
}
model = lgb.LGBMClassifier(**lgb_params)
# -
rfecv = RFECV(estimator=model,
step=2,
cv=StratifiedKFold(
n_splits=2,
shuffle=False),
scoring=make_scorer(score_func=roc_auc_score),
verbose=10)
rfecv.fit(train_X, train_Y)
# How many features were selected?
print('Optimal number of features:', rfecv.n_features_)
# What are the selected features
selected_features = list(train_X.loc[:, rfecv.get_support()].columns)
selected_features
# What are the dropped features
dropped_features = set(train_X.columns) - set(selected_features)
dropped_features
# ### Plot number of features vs CV scores
# I am not sure what does it show. Number of optimal features is 28, but this plot shows only 16 features.
#
# As per the documentation:
#
# > The size of grid_scores_ is equal to ceil((n_features - min_features_to_select) / step) + 1, where step is the number of features removed at each iteration.
#
plt.figure(figsize=(14, 8))
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
# ### All features with Rank 1 should be considered
# There were two features which were dropped. Below DF shows that those two features are of rank 2.
# Save sorted feature rankings
ranking = pd.DataFrame({'feature': features})
ranking['rank'] = rfecv.ranking_
ranking = ranking.sort_values(['rank'], )
ranking
# ### Make a prediction
# Highest Score
score = round(np.max(rfecv.grid_scores_), 5)
score
# +
predictions = rfecv.predict_proba(test_X)[:, -1]
sample_submission_df.target = predictions
sample_submission_df.head()
# +
# score = round(-np.max(rfecv.grid_scores_), 3)
# test['loss'] = rfecv.predict(Xt)
# test = test[['id', 'loss']]
# now = datetime.now()
# sub_file = 'submission_5xRFECV-RandomForest_' + str(score) + '_' + str(
# now.strftime("%Y-%m-%d-%H-%M")) + '.csv'
# print("\n Writing submission file: %s" % sub_file)
# test.to_csv(sub_file, index=False)
| notebooks/feature_selection/recursive_feature_elimination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Running roberta-movie-sentiment model
#
# This tutorial shows how to run the roberta-movie-sentiment model on Onnxruntime.
#
# To see how the roberta-movie-sentiment model was converted from tensorflow to onnx look at [roBERTatutorial.ipynb](https://github.com/SeldonIO/seldon-models/blob/master/pytorch/moviesentiment_roberta/pytorch-roberta-onnx.ipynb)
# # Step 1 - Preprocess
#
# Extract parameters from the given input and convert it into features.
# +
import torch
import numpy as np
from simpletransformers.model import TransformerModel
from transformers import RobertaForSequenceClassification, RobertaTokenizer
text = "This film is so good"
tokenizer = RobertaTokenizer.from_pretrained('roberta-base')
input_ids = torch.tensor(tokenizer.encode(text, add_special_tokens=True)).unsqueeze(0) # Batch size 1
# -
# # Step 2 - Run the ONNX model under onnxruntime
#
# Create an onnx inference session and run the model
# +
import onnxruntime
# Start from ORT 1.10, ORT requires explicitly setting the providers parameter if you want to use execution providers
# other than the default CPU provider (as opposed to the previous behavior of providers getting set/registered by default
# based on the build flags) when instantiating InferenceSession.
# For example, if NVIDIA GPU is available and ORT Python package is built with CUDA, then call API as following:
# onnxruntime.InferenceSession(path/to/model, providers=['CUDAExecutionProvider'])
ort_session = onnxruntime.InferenceSession("roberta-sequence-classification-9.onnx")
def to_numpy(tensor):
return tensor.detach().cpu().numpy() if tensor.requires_grad else tensor.cpu().numpy()
ort_inputs = {ort_session.get_inputs()[0].name: to_numpy(input_ids)}
ort_out = ort_session.run(None, ort_inputs)
# -
# # Step 3 - Postprocessing
#
# Print the results
pred = np.argmax(ort_out)
if(pred == 0):
print("Prediction: negative")
elif(pred == 1):
print("Prediction: positive")
| text/machine_comprehension/roberta/dependencies/roberta-sequence-classification-inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Shahid1993/pytorch-notebooks/blob/master/01_ImageClassifier_using_CNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vM_yO0e87OTv" colab_type="text"
# # Link : [Build an Image Classification Model using Convolutional Neural Networks in PyTorch](https://www.analyticsvidhya.com/blog/2019/10/building-image-classification-models-cnn-pytorch/?utm_source=blog&utm_medium=introduction-to-pytorch-from-scratch)
# + id="oCnmdxRwo383" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a199ad78-f057-4873-8580-2c4f2cf479cb"
from google.colab import drive
drive.mount("/content/drive")
# + [markdown] id="v05oghOz7I5d" colab_type="text"
# # Importing Libraries
# + id="97wgXD7btGCZ" colab_type="code" colab={}
import pandas as pd
import numpy as np
# for reading and displaying images
from skimage.io import imread
import matplotlib.pyplot as plt
# %matplotlib inline
# for creating validation set
from sklearn.model_selection import train_test_split
# for evaluating the model
from sklearn.metrics import accuracy_score
from tqdm import tqdm
# PyTorch Libraries and modules
import torch
from torch.autograd import Variable
from torch.nn import Linear, ReLU, CrossEntropyLoss, Sequential, Conv2d, MaxPool2d, Module, Softmax, BatchNorm2d, Dropout
from torch.optim import Adam, SGD
# + [markdown] id="ARPDP-oQ8xIZ" colab_type="text"
# # Loading the dataset
# + id="VaKC8VP4E4nN" colab_type="code" colab={}
# import zipfile
# with zipfile.ZipFile("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/test_ScVgIM0.zip","r") as zip_ref:
# zip_ref.extractall("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/test_ScVgIM0")
# import zipfile
# with zipfile.ZipFile("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/train_LbELtWX.zip","r") as zip_ref:
# zip_ref.extractall("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/train_LbELtWX")
# + id="GtEGnc3j8teq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="477764e2-3648-4e3c-f758-ba57598d611d"
train = pd.read_csv("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/train_LbELtWX/train.csv")
test = pd.read_csv("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/test_ScVgIM0/test.csv")
sample_submission = pd.read_csv("/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/sample_submission_I5njJSF.csv")
train.head()
# + [markdown] id="TtZZJJp_-MJH" colab_type="text"
# # Loading Images
# + id="2QF9LftC9c4I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="344083db-be44-4921-f934-f0b076e040a7"
# loading training images
train_img = []
for img_name in tqdm(train['id']):
img_path = '/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/train_LbELtWX/train/' + str(img_name) + '.png'
# reading image
img = imread(img_path, as_gray=True)
# normalizing the pixel values
img /= 255.0
#converting the type of pixel to float32
img = img.astype('float32')
train_img.append(img)
# converting the list to numpy array
train_x = np.array(train_img)
# defining the target
train_y = train['label'].values
train_x.shape
# + [markdown] id="XOwBeQ-hPP0y" colab_type="text"
# # Visualizing a few images
# + id="s4_10AGx_nBz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 627} outputId="3acdc734-cbe1-4dc6-efe1-8d9700f86f23"
# visualizing images
i = 0
plt.figure(figsize=(10,10))
plt.subplot(221), plt.imshow(train_x[i], cmap='gray')
plt.subplot(222), plt.imshow(train_x[i+25], cmap='gray')
plt.subplot(223), plt.imshow(train_x[i+50], cmap='gray')
plt.subplot(224), plt.imshow(train_x[i+75], cmap='gray')
# + [markdown] id="xgUQaYewVjXt" colab_type="text"
# # Creating a validation set and preprocessing the images
# + id="GN4UvVArVkO4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6b6a4669-fedd-4b1c-fef8-1ef7d3531cde"
# create validation set
train_x, val_x, train_y, val_y = train_test_split(train_x, train_y, test_size = 0.1)
(train_x.shape, train_y.shape), (val_x.shape, val_y.shape)
# + id="YdxNKRqQWN7I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a63874f9-fca1-4348-ab49-9476e3adc094"
# converting training images into torch format
train_x = train_x.reshape(54000, 1, 28, 28)
train_x = torch.from_numpy(train_x)
# converting the target into torch format
train_y = train_y.astype(int);
train_y = torch.from_numpy(train_y)
# shape of training data
train_x.shape, train_y.shape
# + id="8JI_yVpYYFrt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9b2999fd-5df9-41c1-f2ce-7e477859454a"
# converting validation images into torch format
val_x = val_x.reshape(6000, 1, 28, 28)
val_x = torch.from_numpy(val_x)
# converting the target into torch format
val_y = val_y.astype(int);
val_y = torch.from_numpy(val_y)
# shape of validation data
val_x.shape, val_y.shape
# + [markdown] id="o8BZCy8xbZ0H" colab_type="text"
# # Implementing CNNs using PyTorch
#
# We will use a very simple CNN architecture with just 2 convolutional layers to extract features from the images. We’ll then use a fully connected dense layer to classify those features into their respective categories.
# + [markdown] id="DoggGLXsbc0S" colab_type="text"
# ## Define the architecture
# + id="aJZCTvfQbNYs" colab_type="code" colab={}
class Net(Module):
def __init__(self):
super(Net, self).__init__()
self.cnn_layers = Sequential(
# Defining a 2D convolutional layer
Conv2d(1, 4, kernel_size=3, stride=1, padding=1),
BatchNorm2d(4),
ReLU(inplace=True),
MaxPool2d(kernel_size=2, stride=2),
# Defining another 2D convolutional layer
Conv2d(4, 4, kernel_size=3, stride=1, padding=1),
BatchNorm2d(4),
ReLU(inplace=True),
MaxPool2d(kernel_size=2, stride=2)
)
self.linear_layers = Sequential(
Linear(4 * 7 * 7, 10)
)
# Defining the forward pass
def forward(self, x):
x = self.cnn_layers(x)
x = x.view(x.size(0), -1)
x = self.linear_layers(x)
return x
# + id="RNRuzdQuescU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 280} outputId="7b14a271-1398-4960-b51d-128c5c1865da"
# defining the model
model = Net()
# defining the optimizer
optimizer = Adam(model.parameters(), lr=0.07)
# defining the loss function
criterion = CrossEntropyLoss()
# checking if GPU is available
if torch.cuda.is_available():
model = model.cuda()
criterion = criterion.cuda()
print(model)
# + [markdown] id="tT0vDJR-f79G" colab_type="text"
# ## Defining a function to train the model
# + id="bj3FyUjPfYep" colab_type="code" colab={}
def train(epoch):
model.train()
tr_loss = 0
# getting the training set
x_train, y_train = Variable(train_x), Variable(train_y)
# getting the validation set
x_val, y_val = Variable(val_x), Variable(val_y)
# converting the data into GPU format
if torch.cuda.is_available():
x_train = x_train.cuda()
y_train = y_train.cuda()
x_val = x_val.cuda()
y_val = y_val.cuda()
# clearing the gradients of the model parameters
optimizer.zero_grad()
# prediction for training and validation set
output_train = model(x_train)
output_val = model(x_val)
# computing the training and validatoin loss
loss_train = criterion(output_train, y_train)
loss_val = criterion(output_val, y_val)
train_losses.append(loss_train)
val_losses.append(loss_val)
# computed the updated weights of all the model parameters
loss_train.backward()
optimizer.step()
tr_loss = loss_train.item()
if epoch%2 == 0:
# printing the validation loss
print('Epoch : ', epoch+1, '\t', 'loss : ', loss_val)
# + [markdown] id="0LYOzKtNiBrx" colab_type="text"
# ### Training the model
# + id="chgXXfzXh5My" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 245} outputId="5963ef51-20f6-4bc0-8576-133e608ca355"
# defining the number of epochs
n_epochs = 25
# empty list to store training losses
train_losses = []
# empty list to store validation losses
val_losses = []
# training the model
for epoch in range(n_epochs):
train(epoch)
# + [markdown] id="-VWeRHrniP42" colab_type="text"
# ## Visualizing the training and validation losses
# + id="_ZPvCbOLiTXn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="1a789de4-beab-4e73-bdf4-80fbe12c843d"
# plotting the training and validation loss
plt.plot(train_losses, label='Training loss')
plt.plot(val_losses, label='Validation loss')
plt.legend()
plt.show()
# + [markdown] id="lSCEuO6riobO" colab_type="text"
# ## Checking model accuracy on training and validation sets
# + id="ivNf20OsiuQg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ceef8a2a-baa5-4e18-cf48-72f2eac0ac9b"
# prediction for training set
with torch.no_grad():
output = model(train_x.cuda())
softmax = torch.exp(output).cpu()
prob = list(softmax.numpy())
predictions = np.argmax(prob, axis=1)
# accuracy on training set
accuracy_score(train_y, predictions)
# + id="dtLeKM9xi9rQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="88a4197f-f0ea-4c8b-cd30-94b0a4d81ca4"
# prediction for validation set
with torch.no_grad():
output = model(val_x.cuda())
softmax = torch.exp(output).cpu()
prob = list(softmax.numpy())
predictions = np.argmax(prob, axis=1)
# accuracy on validation set
accuracy_score(val_y, predictions)
# + [markdown] id="wiPEktssjCvo" colab_type="text"
# # Generating predictions for the test set
# + id="sfac8AwTi-w2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="4ce285b4-d546-4593-d760-f28582c8c864"
# loading test images
test_img = []
for img_name in tqdm(test['id']):
# defining the image path
image_path = '/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/test_ScVgIM0/test/' + str(img_name) + '.png'
# reading the image
img = imread(image_path, as_gray=True)
# normalizing the pixel values
img /= 255.0
# converting the type of pixel to float 32
img = img.astype('float32')
# appending the image into the list
test_img.append(img)
# converting the list to numpy array
test_x = np.array(test_img)
test_x.shape
# + id="iXWwE1i5jQPg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="19699a11-48b9-4e42-e5ff-1cbcd48589d1"
# converting test images into torch format
test_x = test_x.reshape(10000, 1, 28, 28)
test_x = torch.from_numpy(test_x)
test_x.shape
# + id="Wul4eFS5ja27" colab_type="code" colab={}
# generating predictions for test set
with torch.no_grad():
output = model(test_x.cuda())
softmax = torch.exp(output).cpu()
prob = list(softmax.numpy())
predictions = np.argmax(prob, axis=1)
# + [markdown] id="8yTeYuA6jkLh" colab_type="text"
# Replace the labels in the sample submission file with the predictions and finally save the file and submit it on the leaderboard:
# + id="ZG0AQ6vojiOP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="f444477c-55fb-4078-e366-a1c415825b1f"
# replacing the label with prediction
sample_submission['label'] = predictions
sample_submission.head()
# + id="o3OtRD1KkS_5" colab_type="code" colab={}
# saving the file
sample_submission.to_csv('/content/drive/My Drive/ML/data/Identify the Apparels - AnalyticsVidhya/submission_cnn.csv', index=False)
# + id="2AejfG8Qkped" colab_type="code" colab={}
| 01_ImageClassifier_using_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import collections
import os
import json
import logging
import string
import re
from scipy.stats import entropy
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sqlalchemy import create_engine
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
import networkx as nx
if os.getcwd().endswith('notebook'):
os.chdir('..')
from rna_learn.codon_bias.process_domains import compute_protein_domain_scores
# -
sns.set(palette='colorblind', font_scale=1.3)
palette = sns.color_palette()
logging.basicConfig(level=logging.INFO, format="%(asctime)s (%(levelname)s) %(message)s")
logger = logging.getLogger(__name__)
db_path = os.path.join(os.getcwd(), 'data/db/seq.db')
engine = create_engine(f'sqlite+pysqlite:///{db_path}')
# ## Match
def compute_match_score(engine):
q = """
select assembly_accession from assembly_source
"""
assembly_accessions = pd.read_sql(q, engine)['assembly_accession'].values
count = 0
for assembly in assembly_accessions:
protein_domains_path = os.path.join(
os.getcwd(),
f'data/domains/codon_bias/pfam/{assembly}_protein_domains.csv',
)
if os.path.isfile(protein_domains_path):
count += 1
return count, 100 * count / len(assembly_accessions)
compute_match_score(engine)
# ## Thermotoga maritima
species_taxid = 2336
q = """
select assembly_accession, species_taxid from assembly_source
where species_taxid = ?
"""
assembly_accession = pd.read_sql(q, engine, params=(species_taxid,))['assembly_accession'].iloc[0]
assembly_accession
protein_domains_path = os.path.join(
os.getcwd(),
f'data/domains/codon_bias/pfam/{assembly_accession}_protein_domains.csv',
)
thermotoga_domains = pd.read_csv(protein_domains_path)
thermotoga_domains.head(20)
v = thermotoga_domains[thermotoga_domains['below_threshold']]
100 * len(v) / len(thermotoga_domains)
all_counts = thermotoga_domains[['pfam_query', 'pfam_accession']].groupby('pfam_query').count()
all_counts.columns = ['count_all']
all_counts = all_counts.sort_values('count_all', ascending=False)
all_counts.head()
below_threshold_counts = thermotoga_domains[
thermotoga_domains['below_threshold']
][['pfam_query', 'pfam_accession']].groupby('pfam_query').count()
below_threshold_counts.columns = ['count_below']
below_threshold_counts = below_threshold_counts.sort_values('count_below', ascending=False)
below_threshold_counts.head()
counts = pd.merge(
all_counts,
below_threshold_counts,
how='left',
on='pfam_query',
)
counts['count_below'] = counts['count_below'].fillna(0).astype(int)
counts.head()
counts['frequency_weight'] = counts['count_below'] / counts['count_all']
counts['species_score'] = np.sqrt(counts['count_below']) * counts['frequency_weight']
counts.sort_values('species_score', ascending=False).head(20)
# ## All species
def compute_protein_domain_score(engine, query_type):
if query_type not in ('pfam', 'tigr'):
raise ValueError('Query type must be one of (pfam, tigr)')
q = """
select a.assembly_accession, s.phylum from assembly_source as a
left join species_traits as s on s.species_taxid = a.species_taxid
"""
df = pd.read_sql(q, engine)
assembly_accessions = df['assembly_accession'].values
phyla = df['phylum'].values
logger.info(f'Counting protein domains for {len(assembly_accessions):,} assemblies')
n_assemblies = len(assembly_accessions)
n_assemblies_present = 0
domain_to_phyla = collections.defaultdict(set)
domain_to_score = collections.defaultdict(int)
domain_count = collections.defaultdict(int)
domain_count_top = collections.defaultdict(int)
for i, assembly in enumerate(assembly_accessions):
if (i+1) % 200 == 0:
logger.info(f'{i+1:,} / {n_assemblies:,}')
protein_domains_path = os.path.join(
os.getcwd(),
f'data/domains/codon_bias/{query_type}/{assembly}_protein_domains.csv',
)
if not os.path.isfile(protein_domains_path):
continue
else:
n_assemblies_present += 1
protein_domains = pd.read_csv(protein_domains_path)
phylum = phyla[i]
all_counts = protein_domains[['pfam_query', 'pfam_accession']].groupby('pfam_query').count()
all_counts.columns = ['count_all']
below_threshold_counts = protein_domains[
protein_domains['below_threshold']
][['pfam_query', 'pfam_accession']].groupby('pfam_query').count()
below_threshold_counts.columns = ['count_below']
counts = pd.merge(
all_counts,
below_threshold_counts,
how='left',
on='pfam_query',
)
counts['count_below'] = counts['count_below'].fillna(0).astype(int)
counts['frequency_weight'] = counts['count_below'] / counts['count_all']
counts['assembly_score'] = np.sqrt(counts['count_below']) * counts['frequency_weight']
for pfam_query in counts.index:
domain_to_phyla[pfam_query].add(phylum)
domain_to_score[pfam_query] += counts.loc[pfam_query, 'assembly_score']
domain_count[pfam_query] += 1
if counts.loc[pfam_query, 'count_below'] > 0:
domain_count_top[pfam_query] += 1
query_key = f'{query_type}_query'
sorted_queries = sorted(domain_to_score.keys())
data = {
query_key: sorted_queries,
'n_phylum': [len(domain_to_phyla[k]) for k in sorted_queries],
'assembly_score_sum': [domain_to_score[k] for k in sorted_queries],
'assembly_count': [domain_count[k] for k in sorted_queries],
'assembly_count_top': [domain_count_top[k] for k in sorted_queries],
'score': [domain_to_score[k] / n_assemblies_present for k in sorted_queries],
}
output_df = pd.DataFrame.from_dict(data).set_index(query_key)
return output_df.sort_values(['score', 'assembly_count'], ascending=False)
def compute_query_to_most_common_label(engine, query_type):
if query_type not in ('pfam', 'tigr'):
raise ValueError('Query type must be one of (pfam, tigr)')
q = """
select assembly_accession from assembly_source
"""
assembly_accessions = pd.read_sql(q, engine)['assembly_accession'].values
logger.info(f'Finding most common protein labels per query for {len(assembly_accessions):,} assemblies')
query_to_protein_labels = {}
for i, assembly in enumerate(assembly_accessions):
if (i+1) % 200 == 0:
logger.info(f'{i+1:,} / {len(assembly_accessions):,}')
protein_domains_path = os.path.join(
os.getcwd(),
f'data/domains/codon_bias/{query_type}/{assembly}_protein_domains.csv',
)
if not os.path.isfile(protein_domains_path):
continue
protein_domains = pd.read_csv(protein_domains_path)
for tpl in protein_domains.itertuples():
query, label = tpl.pfam_query, tpl.protein_label
if pd.isnull(label):
label = 'Unknown'
label = label.strip()
if query not in query_to_protein_labels:
query_to_protein_labels[query] = {
label: 1,
}
elif label not in query_to_protein_labels[query]:
query_to_protein_labels[query][label] = 1
else:
query_to_protein_labels[query][label] += 1
query_to_most_common_label = {}
for query in sorted(query_to_protein_labels.keys()):
label_counts = [(k, v) for k, v in query_to_protein_labels[query].items()]
sorted_labels = sorted(label_counts, key=lambda t: t[1], reverse=True)
query_to_most_common_label[query] = sorted_labels[0][0]
return query_to_most_common_label
# %%time
pfam_counts = compute_protein_domain_score(engine, query_type='pfam')
pfam_query_to_most_common_label = compute_query_to_most_common_label(engine, query_type='pfam')
pfam_labels = [pfam_query_to_most_common_label[k] for k in pfam_counts.index]
pfam_counts['label'] = pfam_labels
pfam_counts.head(50)
100 * len(pfam_counts[pfam_counts['score'] >= 0.1]) / len(pfam_counts)
_, ax = plt.subplots(1, 1, figsize=(12, 6))
ax.hist(pfam_counts['score'].values, bins=50, log=True);
pfam_threshold = np.percentile(pfam_counts['score'].values, 95)
ax.axvline(pfam_threshold, color='red')
ax.set_xlabel('Score (%)')
ax.set_ylabel('Pfam entry count')
ax.set_title('Distribution of scores for all Pfam entries');
# %%time
tigr_counts = compute_protein_domain_score(engine, query_type='tigr')
100 * len(tigr_counts[tigr_counts['score'] > 0.1]) / len(tigr_counts)
tigr_query_to_most_common_label = compute_query_to_most_common_label(engine, query_type='tigr')
tigr_labels = [tigr_query_to_most_common_label[k] for k in tigr_counts.index]
tigr_counts['label'] = tigr_labels
tigr_counts.head(20)
_, ax = plt.subplots(1, 1, figsize=(12, 6))
ax.hist(tigr_counts['score'].values, bins=20, log=True);
tigr_thresold = np.percentile(tigr_counts['score'].values, 95)
ax.axvline(tigr_thresold, color='red')
ax.set_xlabel('Score (%)')
ax.set_ylabel('TIGR entry count')
ax.set_title('Distribution of scores for all TIGR entries');
score_threshold = 0.05
base_path = os.path.join(os.getcwd(), 'data/domains/codon_bias/')
pfam_counts[pfam_counts['score'] >= pfam_threshold].to_excel(os.path.join(base_path, 'pfam_top.xlsx'))
tigr_counts[tigr_counts['score'] >= tigr_thresold].to_excel(os.path.join(base_path, 'tigr_top.xlsx'))
# ## Validation: Protein id match
#
# Let's make sure IDs properly match and we are not simply seeing an artefact of proteins that joined
def check_protein_matching(engine, query_type):
if query_type not in ('pfam', 'tigr'):
raise ValueError('Query type must be one of (pfam, tigr)')
q = """
select assembly_accession from assembly_source
"""
assembly_accessions = pd.read_sql(q, engine)['assembly_accession'].values
logger.info(f'Checking {query_type} protein ID match for {len(assembly_accessions):,} assemblies')
matching_scores = {}
for i, assembly in enumerate(assembly_accessions):
if (i+1) % 200 == 0:
logger.info(f'{i+1:,} / {len(assembly_accessions):,}')
protein_domains_path = os.path.join(
os.getcwd(),
f'data/domains/codon_bias/{query_type}/{assembly}_protein_domains.csv',
)
if not os.path.isfile(protein_domains_path):
continue
protein_domains = pd.read_csv(protein_domains_path)
protein_query = """
select metadata_json from sequences where sequence_type = 'CDS' and assembly_accession = ?
"""
cds_metadata_df = pd.read_sql(protein_query, engine, params=(assembly,))
metadata = [json.loads(v) for v in cds_metadata_df['metadata_json'].values if not pd.isnull(v)]
cds_protein_ids = {
m['protein_id'].strip() for m in metadata
if m.get('protein_id') is not None
}
query_protein_ids = set([p.strip() for p in protein_domains['protein_id'].values if not pd.isnull(p)])
matching_score = 100 * len(cds_protein_ids & query_protein_ids) / len(query_protein_ids)
matching_scores[assembly] = matching_score
return matching_scores
pfam_matching_scores = check_protein_matching(engine, query_type='pfam')
tigr_matching_scores = check_protein_matching(engine, query_type='tigr')
outlier_threshold = 90
outlier_assemblies = {a for a in pfam_matching_scores.keys() if pfam_matching_scores[a] < outlier_threshold}
outlier_assemblies |= {a for a in tigr_matching_scores.keys() if tigr_matching_scores[a] < outlier_threshold}
len(outlier_assemblies)
sorted(outlier_assemblies)
q = """
select a.assembly_accession, s.species_taxid, s.species, s.phylum from assembly_source as a
left join species_traits as s on s.species_taxid = a.species_taxid
"""
df = pd.read_sql(q, engine, index_col='assembly_accession')
ix = set(df.index.tolist()) - set(outlier_assemblies)
phyla = df.loc[ix]['phylum'].unique()
len(phyla)
df[df['phylum'].isnull()]
# ## Top Gene Ontology (GO) categories
pfam2go_path = os.path.join(os.getcwd(), 'data/domains/Pfam2go.txt')
pfam_results = pd.read_excel(
os.path.join(os.getcwd(), f'data/domains/codon_bias/pfam_top.xlsx'),
index_col='pfam_query',
)
def parse_pfam_to_go_file(path):
line_re = r'^Pfam:([^\s]+) ([^>]+) > GO:([^;]+) ; GO:([0-9]+)$'
domain_to_go = collections.defaultdict(list)
with open(path, 'r') as f:
for line in f:
if not line.strip() or line.startswith('!'):
continue
m = re.match(line_re, line)
if m:
pfam_id = m[1].strip()
query = m[2].strip()
go_label = m[3].strip()
go_id = m[4].strip()
domain_to_go[query].append((go_id, go_label))
return dict(domain_to_go)
domain_to_go = parse_pfam_to_go_file(pfam2go_path)
domain_to_go['Helicase_C_2']
def compute_top_go_categories(pfam_results, domain_to_go):
data = {
'go_id': [],
'go_label': [],
'count': [],
}
matching = 0
go_id_count = collections.defaultdict(int)
go_id_to_label = {}
for domain in pfam_results.index:
if domain not in domain_to_go:
continue
else:
matching += 1
go_data = domain_to_go[domain]
for go_id, go_label in go_data:
go_id_count[go_id] += 1
go_id_to_label[go_id] = go_label
for go_id in sorted(go_id_count.keys()):
data['go_id'].append(go_id)
data['go_label'].append(go_id_to_label[go_id])
data['count'].append(go_id_count[go_id])
print(f'{matching} / {len(pfam_results)} ({100 * matching / len(pfam_results):.0f}%) matching domains with go')
return pd.DataFrame.from_dict(data).set_index('go_id').sort_values('count', ascending=False)
go_df = compute_top_go_categories(pfam_results, domain_to_go)
go_df.head(20)
go_df.to_excel(os.path.join(os.getcwd(), 'data/domains/codon_bias/go_labels.xlsx'))
# ## tRNA adaptation index
trnai = pd.read_csv(os.path.join(os.getcwd(), 'data/trn_adaptation_index/GCA_000005825.2_tai.csv'))
trnai.head()
def score_fn(trnai):
mean = trnai['adaptation_index'].mean()
std = trnai['adaptation_index'].std()
def fn(adaptation_index):
if adaptation_index > mean + std:
return 'over expressed'
elif adaptation_index < mean - std:
return 'under expressed'
else:
return 'normally expressed'
return fn
trnai['expression'] = trnai['adaptation_index'].apply(score_fn(trnai))
trnai.head()
trnai['expression'].hist();
# ## AAA domains scale in numbers with genome size
#
# How does it affect our scoring?
species_traits = pd.read_sql(
'select species_taxid, species, genome_size from species_traits',
engine,
index_col='species_taxid',
)
species_traits.head()
species_traits.loc[[2336]]
thermotoga_maritima_domains = compute_protein_domain_scores(engine, ['GCA_000008545.1'], 'pfam')
aaa_domains = sorted([d for d in thermotoga_maritima_domains.index if 'AAA' in d])
thermotoga_maritima_domains.loc[aaa_domains].sort_values('score', ascending=False)
# ## Count unique Pfam
# +
# %%time
import pathlib
pfam_folder = '/Users/srom/workspace/rna_learn/data/domains/tri_nucleotide_bias/pfam'
protein_domains = set()
paths = pathlib.Path(pfam_folder).glob('*.csv')
for p in paths:
with p.open() as f:
df = pd.read_csv(f)
protein_domains |= set(df['pfam_query'].unique())
print(len(protein_domains))
# -
n_pfam_domains = len(protein_domains)
100 * 240 / n_pfam_domains
| notebook/Codon Bias vs Pfam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### <NAME>, MBA+MS (Analytics), Boston University (May, 2019)
# ## Recommendation system for e-commerce businesses
# A well developed recommendation system will help businesses improve their shopper's experience on website and result in better customer acquisition and retention.
#
# The recommendation system, I have designed below is based on the journey of a new customer from the time he/she lands on the business’s website for the first time to when he/she makes repeat purchases.
#
# The recommendation system is designed in 3 parts based on the business context:
#
# Recommendation system part I: Product pupularity based system targetted at new customers
#
# Recommendation system part II: Model-based collaborative filtering system based on customer's purchase history and ratings provided by other users who bought items similar items
#
# Recommendation system part III: When a business is setting up its e-commerce website for the first time withou any product rating
#
# When a new customer without any previous purchase history visits the e-commerce website for the first time, he/she is recommended the most popular products sold on the company's website. Once, he/she makes a purchase, the recommendation system updates and recommends other products based on the purchase history and ratings provided by other users on the website. The latter part is done using collaborative filtering techniques.
# ## Recommendation System - Part I
# ### Product popularity based recommendation system targeted at new customers
# Popularity based are a great strategy to target the new customers with the most popular products sold on a business's website and is very useful to cold start a recommendation engine.
# ## Amazon product review dataset
#
# Source: https://www.kaggle.com/skillsmuggler/amazon-ratings
# #### Importing libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# # %matplotlib inline
plt.style.use("ggplot")
import sklearn
from sklearn.decomposition import TruncatedSVD
# -
# #### Loading the dataset
amazon_ratings = pd.read_csv('ratings_Beauty.csv')
amazon_ratings = amazon_ratings.dropna()
amazon_ratings.head()
amazon_ratings.shape
popular_products = pd.DataFrame(amazon_ratings.groupby('ProductId')['Rating'].count())
most_popular = popular_products.sort_values('Rating', ascending=False)
most_popular.head(10)
most_popular.head(30).plot(kind = "bar")
# ##### Analysis:
#
# The above graph gives us the most popular products (arranged in descending order) sold by the business.
#
# For eaxmple, product, ID # B001MA0QY2 has sales of over 7000, the next most popular product, ID # B0009V1YR8 has sales of 3000, etc.
# ## Recommendation System - Part II
# ### Model-based collaborative filtering system
#
# Recommend items to users based on purchase history and similarity of ratings provided by other users who bought items to that of a particular customer.
# A model based collaborative filtering technique is closen here as it helps in making predictinfg products for a particular user by identifying patterns based on preferences from multiple user data.
# #### Utility Matrix based on products sold and user reviews
# Utility Matrix
# An utlity matrix is consists of all possible user-item preferences (ratings) details represented as a matrix. The utility matrix is sparce as none of the users would buy all teh items in the list, hence, most of the values are unknown.
# +
# Subset of Amazon Ratings
amazon_ratings1 = amazon_ratings.head(10000)
# -
ratings_utility_matrix = amazon_ratings1.pivot_table(values='Rating', index='UserId', columns='ProductId', fill_value=0)
ratings_utility_matrix.head()
# As expected, the utility matrix obtaned above is sparce, I have filled up the unknown values wth 0.
ratings_utility_matrix.shape
# Transposing the matrix
X = ratings_utility_matrix.T
X.head()
X.shape
# Unique products in subset of data
X1 = X
# ### Decomposing the Matrix
SVD = TruncatedSVD(n_components=10)
decomposed_matrix = SVD.fit_transform(X)
decomposed_matrix.shape
# ### Correlation Matrix
correlation_matrix = np.corrcoef(decomposed_matrix)
correlation_matrix.shape
# correlation_matrix
# ### Isolating Product ID # 6117036094 from the Correlation Matrix
#
# Assuming the customer buys Product ID # 6117036094 (randomly chosen)
X.index[99]
# Index # of product ID purchased by customer
# +
i = "6117036094"
product_names = list(X.index)
product_ID = product_names.index(i)
product_ID
# -
# Correlation for all items with the item purchased by this customer based on items rated by other customers people who bought the same product
correlation_product_ID = correlation_matrix[product_ID]
correlation_product_ID.shape
# ### Recommending top 10 highly correlated products in sequence
# +
Recommend = list(X.index[correlation_product_ID > 0.90])
# Removes the item already bought by the customer
Recommend.remove(i)
Recommend[0:9]
# -
# Product Id #
# Here are the top 10 products to be displayed by the recommendation system to the above customer based on the purchase history of other customers in the website.
# ## Recommendation System - Part III
# For a business without any user-item purchase history, a search engine based recommendation system can be designed for users. The product recommendations can be based on textual clustering analysis given in product description.
# +
# Importing libraries
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn.neighbors import NearestNeighbors
from sklearn.cluster import KMeans
from sklearn.metrics import adjusted_rand_score
# -
# ## Home Depot's dataset with product dataset:
#
# https://www.kaggle.com/c/home-depot-product-search-relevance/data
# ### Item to item based recommendation system based on product description
#
# Applicable when business is setting up its E-commerce website for the first time
product_descriptions = pd.read_csv('product_descriptions.csv')
product_descriptions.shape
# #### Checking for missing values
# +
# Missing values
product_descriptions = product_descriptions.dropna()
product_descriptions.shape
product_descriptions.head()
# +
product_descriptions1 = product_descriptions.head(500)
# product_descriptions1.iloc[:,1]
product_descriptions1["product_description"].head(10)
# -
# #### Feature extraction from product descriptions
#
# Converting the text in product description into numerical data for analysis
vectorizer = TfidfVectorizer(stop_words='english')
X1 = vectorizer.fit_transform(product_descriptions1["product_description"])
X1
# #### Visualizing product clusters in subset of data
# +
# Fitting K-Means to the dataset
X=X1
kmeans = KMeans(n_clusters = 10, init = 'k-means++')
y_kmeans = kmeans.fit_predict(X)
plt.plot(y_kmeans, ".")
plt.show()
# -
# #### Top words in each cluster based on product description
# +
# # Optimal clusters is
true_k = 10
model = KMeans(n_clusters=true_k, init='k-means++', max_iter=100, n_init=1)
model.fit(X1)
print("Top terms per cluster:")
order_centroids = model.cluster_centers_.argsort()[:, ::-1]
terms = vectorizer.get_feature_names()
for i in range(true_k):
print("Cluster %d:" % i),
for ind in order_centroids[i, :10]:
print(' %s' % terms[ind]),
print
# -
# #### Predicting clusters based on key search words
# cutting tool
print("Cluster ID:")
Y = vectorizer.transform(["cutting tool"])
prediction = model.predict(Y)
print(prediction)
# spray paint
print("Cluster ID:")
Y = vectorizer.transform(["spray paint"])
prediction = model.predict(Y)
print(prediction)
# steel drill
print("Cluster ID:")
Y = vectorizer.transform(["steel drill"])
prediction = model.predict(Y)
print(prediction)
# In case a word appears in multiple clusters, the algorithm chooses the cluster with the highest frequency of occurance of the word.
# water
print("Cluster ID:")
Y = vectorizer.transform(["water"])
prediction = model.predict(Y)
print(prediction)
# Once a cluster is identified based on the user's search words, the recommendation system can display items from the corresponding product clusters based on the product descriptions.
# #### Summary:
#
# This works best if a business is setting up its e-commerce website for the first time and does not have user-item purchase/rating history to start with initally. This recommendation system will help the users get a good recommendation to start with and once the buyers have a purchased history, the recommendation engine can use the model based collaborative filtering technique.
| _source/raw/amazon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Saving and loading models for inference
# In the end, we train models because we want to use them for inference, that is, using them to generate predictions on new targets. The general formula for doing this in FastAI.jl is to first train a `model` for a `method`, for example using [`fitonecycle!`](#) or [`finetune!`](#) and then save the model and the learning method configuration to a file using [`savemethodmodel`](#). In another session you can then use [`loadmethodmodel`](#) to load both. Since the learning method contains all preprocessing logic we can then use [`predict`](#) and [`predictbatch`](#) to generate predictions for new inputs.
#
# Let's fine-tune an image classification model (see [here](./fitonecycle.ipynb) for more info) and go through that process.
using FastAI
using Metalhead
# +
dir = joinpath(datasetpath("dogscats"), "train")
data = loadfolderdata(dir, filterfn=isimagefile, loadfn=(loadfile, parentname))
classes = unique(eachobs(data[2]))
method = BlockMethod(
(Image{2}(), Label(classes)),
(
ProjectiveTransforms((196, 196)),
ImagePreprocessing(),
OneHot()
)
)
backbone = Metalhead.ResNet50(pretrain=true).layers[1][1:end-1]
learner = methodlearner(method, data; backbone=backbone, callbacks=[ToGPU(), Metrics(accuracy)])
finetune!(learner, 3)
# -
# Now we can save the model using [`savemethodmodel`](#).
savemethodmodel("catsdogs.jld2", method, learner.model, force = true)
# In another session we can now use [`loadmethodmodel`](#) to load both model and learning method from the file. Since the model weights are transferred to the CPU before being saved, we need to move them to the GPU manually if we want to use that for inference.
method, model = loadmethodmodel("catsdogs.jld2")
model = gpu(model);
# Finally, let's select 9 random images from the dataset and see if the model classifies them correctly:
# use it for inference
x, y =
samples = [getobs(data, i) for i in rand(1:nobs(data), 9)]
images = [sample[1] for sample in samples]
labels = [sample[2] for sample in samples]
preds = predictbatch(method, model, images; device = gpu, context = Validation())
acc = sum(labels .== preds) / length(preds)
using CairoMakie
plotsamples(method, collect(zip(images, preds)))
| notebooks/serialization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-datascience
# language: python
# name: venv-datascience
# ---
# # Pandas Practice
#
# This notebook is dedicated to practicing different tasks with pandas. The solutions are available in a solutions notebook, however, you should always try to figure them out yourself first.
#
# It should be noted there may be more than one different way to answer a question or complete an exercise.
#
# Exercises are based off (and directly taken from) the quick introduction to pandas notebook.
#
# Different tasks will be detailed by comments or text.
#
# For further reference and resources, it's advised to check out the [pandas documnetation](https://pandas.pydata.org/pandas-docs/stable/).
# Import pandas
import pandas as pd
# Create a series of three different colours
colours_series = pd.Series(['Blue', 'Green', 'Red'])
# View the series of different colours
colours_series
# Create a series of three different car types and view it
car_series = pd.Series(['BMW', 'Mercedes-Benz', 'Maserati'])
car_series
# Combine the Series of cars and colours into a DataFrame
cars = pd.DataFrame({
'Brand': car_series,
'Colour': colours_series,
})
cars
# Import "../data/car-sales.csv" and turn it into a DataFrame
car_sales = pd.read_csv('car-sales.csv')
car_sales
# **Note:** Since you've imported `../data/car-sales.csv` as a DataFrame, we'll now refer to this DataFrame as 'the car sales DataFrame'.
# Export the DataFrame you created to a .csv file
car_sales.to_csv('exported-car-sales.csv')
# Find the different datatypes of the car data DataFrame
car_sales.dtypes
# Describe your current car sales DataFrame using describe()
car_sales.describe()
# Get information about your DataFrame using info()
car_sales.info()
# What does it show you?
# Create a Series of different numbers and find the mean of them
my_series = pd.Series([23, 3455435,12,24543])
my_series.mean()
# Create a Series of different numbers and find the sum of them
my_series.sum()
# List out all the column names of the car sales DataFrame
car_sales.columns
# Find the length of the car sales DataFrame
len(car_sales)
# Show the first 5 rows of the car sales DataFrame
car_sales.head()
# Show the first 7 rows of the car sales DataFrame
car_sales.head(7)
# Show the bottom 5 rows of the car sales DataFrame
car_sales.tail(5)
# Use .loc to select the row at index 3 of the car sales DataFrame
car_sales.loc[3]
# Use .iloc to select the row at position 3 of the car sales DataFrame
car_sales.iloc[3]
# Notice how they're the same? Why do you think this is?
#
# Check the pandas documentation for [.loc](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.loc.html) and [.iloc](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.iloc.html). Think about a different situation each could be used for and try them out.
# Select the "Odometer (KM)" column from the car sales DataFrame
car_sales['Odometer (KM)']
# Find the mean of the "Odometer (KM)" column in the car sales DataFrame
car_sales['Odometer (KM)'].mean()
# Select the rows with over 100,000 kilometers on the Odometer
car_sales[car_sales['Odometer (KM)'] > 100000]
# Create a crosstab of the Make and Doors columns
pd.crosstab(car_sales.Make, car_sales.Doors)
# Group columns of the car sales DataFrame by the Make column and find the average
car_sales.groupby('Make').mean()
# Import Matplotlib and create a plot of the Odometer column
# Don't forget to use %matplotlib inline
# %matplotlib inline
import matplotlib.pyplot as plt
car_sales['Odometer (KM)'].plot()
# Create a histogram of the Odometer column using hist()
car_sales['Odometer (KM)'].hist()
# Try to plot the Price column using plot()
car_sales.Price.plot()
# Why didn't it work? Can you think of a solution?
#
# You might want to search for "how to convert a pandas string columb to numbers".
#
# And if you're still stuck, check out this [Stack Overflow question and answer on turning a price column into integers](https://stackoverflow.com/questions/44469313/price-column-object-to-int-in-pandas).
#
# See how you can provide the example code there to the problem here.
# Remove the punctuation from price column
car_sales['Price'] = car_sales['Price'].str.replace('[\$\,\.]', '')
# Check the changes to the price column
car_sales['Price']
# Remove the two extra zeros at the end of the price column
car_sales['Price'] = car_sales['Price'].str[:-2]
# Check the changes to the Price column
car_sales['Price']
# Change the datatype of the Price column to integers
car_sales['Price'] = car_sales['Price'].astype(int)
car_sales.dtypes
# Lower the strings of the Make column
car_sales['Make'].str.lower()
# If you check the car sales DataFrame, you'll notice the Make column hasn't been lowered.
#
# How could you make these changes permanent?
#
# Try it out.
# Make lowering the case of the Make column permanent
car_sales['Make'] = car_sales['Make'].str.lower()
# Check the car sales DataFrame
car_sales
# Notice how the Make column stays lowered after reassigning.
#
# Now let's deal with missing data.
# +
# Import the car sales DataFrame with missing data ("../data/car-sales-missing-data.csv")
car_sales_missing_data = pd.read_csv('car-sales-missing-data.csv')
# Check out the new DataFrame
car_sales_missing_data
# -
# Notice the missing values are represented as `NaN` in pandas DataFrames.
#
# Let's try fill them.
# Fill the Odometer (KM) column missing values with the mean of the column inplace
car_sales_missing_data['Odometer'].fillna(car_sales_missing_data['Odometer'].mean(), inplace = True)
# View the car sales missing DataFrame and verify the changes
car_sales_missing_data
# Remove the rest of the missing data inplace
car_sales_missing_data.dropna(inplace=True)
# Verify the missing values are removed by viewing the DataFrame
car_sales_missing_data
# We'll now start to add columns to our DataFrame.
# Create a "Seats" column where every row has a value of 5
car_sales['Seats'] = 5
car_sales
# Create a column called "Engine Size" with random values between 1.3 and 4.5
# Remember: If you're doing it from a Python list, the list has to be the same length
# as the DataFrame
car_sales['Engine Size'] = [1.3, 2.5, 3.6, 4.5, 3.3, 2.6, 3.8, 4.3, 1.2, 2.9]
# Create a column which represents the price of a car per kilometer
# Then view the DataFrame
car_sales['Price per KM'] = car_sales['Price'] / car_sales['Odometer (KM)']
car_sales
# Remove the last column you added using .drop()
car_sales.drop('Price per KM', axis = 1, inplace = True)
car_sales
# Shuffle the DataFrame using sample() with the frac parameter set to 1
# Save the the shuffled DataFrame to a new variable
shuffed_car_sales = car_sales.sample(frac=1)
shuffed_car_sales
# Notice how the index numbers get moved around. The [`sample()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.sample.html) function is a great way to get random samples from your DataFrame. It's also another great way to shuffle the rows by setting `frac=1`.
# Reset the indexes of the shuffled DataFrame
shuffed_car_sales.reset_index()
# Notice the index numbers have been changed to have order (start from 0).
# Change the Odometer values from kilometers to miles using a Lambda function
# Then view the DataFrame
car_sales['Odometer (KM)'] = car_sales['Odometer (KM)'].apply(lambda x : x / 1.6)
# Change the title of the Odometer (KM) to represent miles instead of kilometers
car_sales.rename(columns={'Odometer (KM)' : 'Odometer (Miles)'}, inplace = True)
car_sales
# ## Extensions
#
# For more exercises, check out the pandas documentation, particularly the [10-minutes to pandas section](https://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html).
#
# One great exercise would be to retype out the entire section into a Jupyter Notebook of your own.
#
# Get hands-on with the code and see what it does.
#
# The next place you should check out are the [top questions and answers on Stack Overflow for pandas](https://stackoverflow.com/questions/tagged/pandas?sort=MostVotes&edited=true). Often, these contain some of the most useful and common pandas functions. Be sure to play around with the different filters!
#
# Finally, always remember, the best way to learn something new to is try it. Make mistakes. Ask questions, get things wrong, take note of the things you do most often. And don't worry if you keep making the same mistake, pandas has many ways to do the same thing and is a big library. So it'll likely take a while before you get the hang of it.
| Complete Machine Learning and Data Science - Zero to Mastery - AN/06.Pandas Data Analysis/pandas-exercises-MySolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [各期獎號下載 (Historical Winning Number Download)](https://www.taiwanlottery.com.tw/info/download_history.asp): 2004-2020 各種樂透的獎號, in zip
# +
import requests
for year in range(103, 111):
url = f'https://www.taiwanlottery.com.tw/info/download/{year}.zip'
print(f"Downloading year {year}...")
r = requests.get(url, allow_redirects=True)
if r.status_code == 200:
with open(f'../output/win_nums_{year}.zip', 'wb') as f:
f.write(r.content)
else:
print("ERROR!! status_code != 200")
else:
print("All downloads complete")
# -
| notebook/get_lotto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wilselby/diy_driverless_car_ROS/blob/ml-model/rover_ml/utils/Image_Regression_with_CNNs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EPxyR5SargY9" colab_type="text"
# #Environment Setup
#
# ##Set Flag for Local or Hosted Runtime
# + id="LoJxV8hsrnhh" colab_type="code" colab={}
local = 0
# + [markdown] id="7vy5iiwR19nJ" colab_type="text"
# ## Confirm TensorFlow can see the GPU
#
# Simply select "GPU" in the Accelerator drop-down in Notebook Settings (either through the Edit menu or the command palette at cmd/ctrl-shift-P).
# + id="_0h6Afcy2E9l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f52ea28e-9fbd-4a20-caa5-2963f4a6e008"
if not local:
import tensorflow as tf
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + [markdown] id="NI-UBX3-2WCU" colab_type="text"
# ## Import Google Drive
# + id="LnbgHIgn2XsB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="a3d17a40-7c03-4896-83ad-a1bcd0f8639f"
if not local:
# Load the Drive helper and mount
from google.colab import drive
# This will prompt for authorization.
drive.mount('/content/drive')
# + [markdown] id="lZbZCuOD2JLH" colab_type="text"
# ## Import Dependencies
# + id="AqUpKnu52L5S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="d27ffdd3-afc8-42e1-ca67-9624e8e6d3be"
if not local:
import os
import csv
import cv2
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
from keras import backend as K
from keras.models import Model, Sequential
from keras.layers import Dense, GlobalAveragePooling2D, MaxPooling2D, Lambda, Cropping2D
from keras.layers.convolutional import Convolution2D
from keras.layers.core import Flatten, Dense, Dropout, SpatialDropout2D
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, TensorBoard
from keras.preprocessing.image import ImageDataGenerator
import sklearn
from sklearn.model_selection import train_test_split
import pandas as pd
# Improve progress bar display
import tqdm
import tqdm.auto
tqdm.tqdm = tqdm.auto.tqdm
print(tf.__version__)
else:
import os
import csv
import cv2
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from keras import backend as K
from keras.models import Model, Sequential
from keras.layers import Dense, GlobalAveragePooling2D, MaxPooling2D, Lambda, Cropping2D
from keras.layers.convolutional import Convolution2D
from keras.layers.core import Flatten, Dense, Dropout, SpatialDropout2D
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint
import sklearn
from sklearn.model_selection import train_test_split
import pandas as pd
# + [markdown] id="-rEBDAIHHWTD" colab_type="text"
# ## Setup Tensorboard
# + id="iKdshk3PRvMe" colab_type="code" outputId="8afc86fe-b27e-4b47-f4e7-7e1478046ebc" colab={"base_uri": "https://localhost:8080/", "height": 68}
if not local:
# Launch Tensorboard
# TODO https://www.tensorflow.org/tensorboard/r2/get_started
from tensorboardcolab import *
tbc = TensorBoardColab()
# + [markdown] id="H2sPUxOR3FlN" colab_type="text"
# # Load Dataset
# + [markdown] id="8YsJN12w3aug" colab_type="text"
# ## Extract Dataset
# + id="ndnCqXG-3I7R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d50b1940-66f9-4e75-9ef3-46c20abe901d"
if not local:
path = '/content/drive/My Drive/research/diy_driverless_car_ROS/rover_ml/output'
data_set = 'office_delay_offset'
tar_file = data_set + ".tar.gz"
os.chdir(path)
# Unzip the .tgz file
# x for extract
# -v for verbose
# -z for gnuzip
# -f for file (should come at last just before file name)
# -C to extract the zipped contents to a different directory
# !if [ -d $data_set ]; then echo 'Directory Exists'; else tar -xvzf $tar_file ; fi
else:
path = '/content/drive/My Drive/research/diy_driverless_car_ROS/rover_ml/output'
data_set = 'office_2'
tar_file = data_set + ".tar.gz"
# + [markdown] id="9FogCRzr3sUF" colab_type="text"
# ## Parse CSV File
# + id="KeQ8c-9s3v8y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="fd91d75f-2505-4f16-be4d-5961437bb2df"
# Define path to csv file
if not local:
csv_path = data_set + '/interpolated.csv'
else:
csv_path = path + '/' + data_set + '/interpolated.csv'
# Load the CSV file into a pandas dataframe
df = pd.read_csv(csv_path, sep=",")
# Print the dimensions
print("Dataset Dimensions:")
print(df.shape)
# Print the first 5 lines of the dataframe for review
print("\nDataset Summary:")
df.head(5)
# + [markdown] id="3BnBpo6-F2oR" colab_type="text"
# # Clean and Pre-process the Dataset
# + [markdown] id="ub8CkShSJEkV" colab_type="text"
# ## Remove Unneccessary Columns
# + id="PDth_K-3JINP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="2141db42-e217-41f9-c4d9-78f0e0db9576"
# Remove 'index' and 'frame_id' columns
df.drop(['index','frame_id'],axis=1,inplace=True)
# Verify new dataframe dimensions
print("Dataset Dimensions:")
print(df.shape)
# Print the first 5 lines of the new dataframe for review
print("\nDataset Summary:")
print(df.head(5))
# + [markdown] id="o-zof8fUDz2C" colab_type="text"
# ## Detect Missing Data
# + id="ffOXGmmQD2om" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="6be4a7d7-3420-4901-c4a6-a1b96f4928d2"
# Detect Missing Values
print("Any Missing Values?: {}".format(df.isnull().values.any()))
# Total Sum
print("\nTotal Number of Missing Values: {}".format(df.isnull().sum().sum()))
# Sum Per Column
print("\nTotal Number of Missing Values per Column:")
print(df.isnull().sum())
# + [markdown] id="cKBJ4sODIFOC" colab_type="text"
# ## Remove Zero Throttle Values
# + id="QAk-fsbkIJrh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="d48098ac-2567-4e11-d1c8-d179de6637a9"
# Determine if any throttle values are zeroes
print("Any 0 throttle values?: {}".format(df['speed'].eq(0).any()))
# Determine number of 0 throttle values:
print("\nNumber of 0 throttle values: {}".format(df['speed'].eq(0).sum()))
# Remove rows with 0 throttle values
if df['speed'].eq(0).any():
df = df.query('speed != 0')
# Reset the index
df.reset_index(inplace=True)
# Verify new dataframe dimensions
print("\nNew Dataset Dimensions:")
print(df.shape)
# + [markdown] id="6XZJLwqKCbE7" colab_type="text"
# ## View Label Statistics
# + id="AgOG94fnCeDB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="b7317ec7-d78f-49cb-8e7d-7b692652ce3c"
# Steering Command Statistics
print("\nSteering Command Statistics:")
print(df['angle'].describe())
print("\nThrottle Command Statistics:")
# Throttle Command Statistics
print(df['speed'].describe())
# + [markdown] id="nae5TmmFFJ5T" colab_type="text"
# ## View Histogram of Steering Commands
# + id="JvPMKVGfFNdy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="c03eca9c-f7d5-46e4-88b7-a1922d039b28"
num_bins = 25
hist, bins = np.histogram(df['angle'], num_bins)
print(bins)
# + id="Bh3DSasZQKCi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="fb4e2915-ddac-4619-e15c-6f9b537e2700"
num_bins = 25
samples_per_bin = 50
hist, bins = np.histogram(df['angle'], num_bins)
center = (bins[:-1]+ bins[1:]) * 0.5
plt.bar(center, hist, width=0.05)
plt.plot((np.min(df['angle']), np.max(df['angle'])), (samples_per_bin, samples_per_bin))
# + id="cwunrGrDQweC" colab_type="code" colab={}
# Normalize the histogram
#print('total data:', len(df))
#remove_list = []
#for j in range(num_bins):
# list_ = []
# for i in range(len(df['angle'])):
# if df.loc[i,'angle'] >= bins[j] and df.loc[i,'angle'] <= bins[j+1]:
# list_.append(i)
# list_ = shuffle(list_)
# list_ = list_[samples_per_bin:]
# remove_list.extend(list_)
#print('removed:', len(remove_list))
#data.drop(data.index[remove_list], inplace=True)
#print('remaining:', len(data))
#hist, _ = np.histogram(data['steering'], (num_bins))
#plt.bar(center, hist, width=0.05)
#plt.plot((np.min(data['steering']), np.max(data['steering'])), (samples_per_bin, samples_per_bin))
# + [markdown] id="1uqfxZ4uGoNX" colab_type="text"
# ## View a Sample Image
# + id="L2nwHnC4Gq1m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="65f9acad-2488-417d-9f85-4ae625ea70ee"
# Crop values
y = 100
h = 100
x = 0
w = 320
image[y:y+h, x:x+w]
# View a Single Image
num = 105
img_name = path + '/' + data_set + '/' + df.loc[num,'filename']
angle = df.loc[num,'angle']
print(img_name)
center_image = cv2.imread(img_name)
crop_img = center_image[y:y+h, x:x+w]
center_image_mod = cv2.resize(center_image, (320,180)) #resize from 720x1280 to 180x320
plt.subplot(2,1,1)
plt.imshow(center_image_mod)
plt.grid(False)
plt.xlabel('angle: {:.4}'.format(angle))
plt.show()
plt.subplot(2,1,2)
plt.imshow(crop_img)
plt.grid(False)
plt.xlabel('angle: {:.4}'.format(angle))
plt.show()
# + [markdown] id="gTQywtOyGvLv" colab_type="text"
# ## View Multiple Images
# + id="ODmdWWpsGxK2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 253} outputId="75cb8cab-e9d6-4971-94f3-a342f2086c19"
# Number of Images to Display
num_images = 4
# Display the images
i = 0
for i in range (0,num_images):
image_path = df.loc[i,'filename']
angle = df.loc[i,'angle']
img_name = path + '/' + data_set + '/' + image_path
image = cv2.imread(img_name)
image = cv2.resize(image, (320,180))
plt.subplot(num_images/2,num_images/2,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(image, cmap=plt.cm.binary)
plt.xlabel('angle: {:.3}'.format(angle))
i += 1
# + [markdown] id="jg1aMw8mGQ8o" colab_type="text"
# ## Load the Images
# + id="xkmkVIiKGStZ" colab_type="code" colab={}
# Define image loading function
def load_images(dataframe):
# initialize images array
images = []
for i in df.index.values:
name = path + '/' + data_set + '/' + dataframe.loc[i,'filename']
center_image = cv2.imread(name)
crop_img = center_image[100:200, 0:320]
center_image = cv2.resize(crop_img, (320,180))
images.append(center_image)
return np.array(images)
# Load images
images = load_images(df)
# Normalize image values
images = images / 255.0
# + [markdown] id="BgFvPAZl9vfP" colab_type="text"
# # Split the Dataset
# + [markdown] id="t2ibXOio_saZ" colab_type="text"
# ## Create the feature set
# + id="Up2wjjIt-B1a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="97271a9f-1d37-4183-cac6-4540321093a6"
# partition the data into training and testing splits using 75% of
# the data for training and the remaining 25% for testing
split = train_test_split(df, images, test_size=0.25, random_state=42)
(trainAttrX, testAttrX, trainImagesX, testImagesX) = split
print("Number of training samples: {}".format(trainAttrX.shape[0]))
print("Number of validation samples: {}".format(testAttrX.shape[0]))
# + [markdown] id="dSHgIhOy_Cxd" colab_type="text"
# ## Create the label set
# + id="OMbWR4YW_Oe2" colab_type="code" colab={}
trainY = trainAttrX["angle"]
testY = testAttrX["angle"]
# + [markdown] id="m3t9Inb5G7lp" colab_type="text"
# ## Define a Batch Generator
# + id="qh3kRBYuG9ed" colab_type="code" colab={}
def csv_image_generator(dataframe, batch_size, mode="train", aug=None):
num_samples = dataframe.shape[0]
# loop indefinitely
while True:
for offset in range(0, num_samples, batch_size):
batch_samples = dataframe[offset:offset+batch_size]
# initialize our batches of images and labels
#print("\nLoaded batch {0}\n".format(1+(offset/batch_size)))
images = []
print("Init {}".format(type(images)))
labels = []
index = 0
for index in range(0,batch_samples.shape[0]):
if batch_samples.loc[offset,'filename'] != "filename":
name = path + '/' + data_set + '/' + dataframe.loc[offset,'filename']
center_image = cv2.imread(name)
center_image = cv2.resize(center_image, (320,180))
label = dataframe.loc[offset,'angle']
print("Loop {}".format(type(images)))
images.append(center_image)
labels.append(label)
index += 1
# if the data augmentation object is not None, apply it
if aug is not None:
(images, labels) = next(aug.flow(np.array(images), labels, batch_size=batch_size))
print("Aug {}".format(type(images)))
# yield the batch to the calling function
yield (np.array(images), labels)
# + [markdown] id="PAVmOpT8HEg0" colab_type="text"
# ## Define an Image Augmentation Data Generator
# + id="Gu67P9z2HIZV" colab_type="code" colab={}
# Construct the training image generator for data augmentation
aug = ImageDataGenerator(rotation_range=20, zoom_range=0.15,
width_shift_range=0.2, height_shift_range=0.2, shear_range=0.15,
vertical_flip=True, fill_mode="nearest")
# + [markdown] id="yA2di_mIdhvW" colab_type="text"
# ## Initialize Data Generators
# + id="dvhng7hsdiJQ" colab_type="code" colab={}
# Define a batch size
batch_size = 32
# initialize both the training and testing image generators
trainGen = csv_image_generator(df, batch_size, mode="train", aug=aug)
testGen = csv_image_generator(df, batch_size, mode="train", aug=None)
# + [markdown] id="-_WW4C27_4HO" colab_type="text"
# # Train the Model
# + [markdown] id="Xq73r642_6iI" colab_type="text"
# ## Preprocess the Input Image
# + id="95obLCcw_8lT" colab_type="code" colab={}
# Initialize the model
model = Sequential()
# trim image to only see section with road
# (top_crop, bottom_crop), (left_crop, right_crop)
model.add(Cropping2D(cropping=((50,20), (0,0)), input_shape=(180,320,3)))
# + [markdown] id="Z62Wkaj4ADbB" colab_type="text"
# ## Build the Model
# + id="AK578kaYAE1_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 785} outputId="5dd9b1cd-0d2e-4905-f697-21d20a44557b"
# Nvidia model
model.add(Convolution2D(24, (5, 5), activation="relu", name="conv_1", strides=(2, 2)))
model.add(Convolution2D(36, (5, 5), activation="relu", name="conv_2", strides=(2, 2)))
model.add(Convolution2D(48, (5, 5), activation="relu", name="conv_3", strides=(2, 2)))
model.add(SpatialDropout2D(.5, dim_ordering='default'))
model.add(Convolution2D(64, (3, 3), activation="relu", name="conv_4", strides=(1, 1)))
model.add(Convolution2D(64, (3, 3), activation="relu", name="conv_5", strides=(1, 1)))
model.add(Flatten())
model.add(Dense(1164))
model.add(Dropout(.5))
model.add(Dense(100, activation='relu'))
model.add(Dropout(.5))
model.add(Dense(50, activation='relu'))
model.add(Dropout(.5))
model.add(Dense(10, activation='relu'))
model.add(Dropout(.5))
model.add(Dense(1))
model.compile(loss='mse', optimizer='adam', metrics=['mse','mape'])
# Print model sumamry
model.summary()
# + [markdown] id="USKQYVmhAMaJ" colab_type="text"
# ## Setup Checkpoints
# + id="BS6R3FVoAOPc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c49e31c3-7ab8-4711-d896-c1de968e9ba5"
# checkpoint
model_path = data_set + '/model'
# !if [ -d $model_path ]; then echo 'Directory Exists'; else mkdir $model_path; fi
filepath = path + '/' + model_path + "/weights-improvement-{epoch:02d}-{val_loss:.2f}.hdf5"
checkpoint = ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True, mode='auto', period=1)
#model.load_weights(model_path = '/')
# + [markdown] id="BVIGONgjSy7V" colab_type="text"
# ## Setup Tensorboard
# + id="-YWTAEeyS0tw" colab_type="code" colab={}
tbCallBack = TensorBoard(log_dir='./log', histogram_freq=1,
write_graph=True,
write_grads=True,
batch_size=batch_size,
write_images=True)
# + [markdown] id="71O-pWk9AQy3" colab_type="text"
# ## Training
# + id="3nkossmrAUo2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0598b3db-75fd-4e99-e3f4-2395c06ea9f2"
# Define number of epochs
n_epoch = 35
# Define callbacks
callbacks_list = [TensorBoardColabCallback(tbc)]
# Fit the model
#history_object = model.fit_generator(train_generator, steps_per_epoch=(len(train_samples) / batch_size_value), validation_data=validation_generator, validation_steps=(len(validation_samples)/batch_size_value), callbacks=callbacks_list, epochs=n_epoch)
#history_object = model.fit_generator(
# trainGen,
# steps_per_epoch=trainAttrX.shape[0] // batch_size,
# validation_data=testGen,
# validation_steps=testAttrX.shape[0] // batch_size,
# callbacks=callbacks_list,
# epochs=n_epoch)
# Fit the model
history_object = model.fit(trainImagesX, trainY, validation_data=(testImagesX, testY), callbacks=callbacks_list, epochs=n_epoch, batch_size=batch_size)
# + [markdown] id="OdaGfWNxBT4T" colab_type="text"
# ## Save the Model
# + id="jWE--9r4BZEk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bcd16fa0-b50f-4521-ac8d-8bd3dae74c8d"
# Save model
model_path_full = path + '/' + model_path + '/'
model.save(model_path_full + 'model.h5')
with open(model_path_full + 'model.json', 'w') as output_json:
output_json.write(model.to_json())
# Save TensorFlow model
tf.train.write_graph(K.get_session().graph.as_graph_def(), logdir=model_path_full, name='model.pb', as_text=False)
# + [markdown] id="Y-pCmYO1A89_" colab_type="text"
# # Evaluate the Model
# + id="podNBS9cBRNW" colab_type="code" colab={}
#scores = model.evaluate(testImagesX, testY, verbose=0, batch_size=32)
#print("Loss: {}".format(scores*100))
#score = model.evaluate_generator(validation_generator, len(validation_samples)/batch_size_value,use_multiprocessing=True)
#print("Loss: {:.2}".format(score))
# + [markdown] id="caA5cztpBPTM" colab_type="text"
# ## Compute the loss
# + id="9FpUcNFu_5z6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="96450b57-af5c-4dac-8095-6568199b5d49"
#TODO: Remove 0s in dataset
preds = model.predict(testImagesX)
diff = preds.flatten() - testY
percentDiff = (diff / testY) * 100
absPercentDiff = np.abs(percentDiff)
#print(diff/testY)
# compute the mean and standard deviation of the absolute percentage
# difference
mean = np.mean(absPercentDiff)
std = np.std(absPercentDiff)
print("[INFO] mean: {:.2f}%, std: {:.2f}%".format(mean, std))
# + [markdown] id="5jVE6WfNGDEM" colab_type="text"
# ## Plot a Prediction
# + [markdown] id="O7o-6SbBD9zx" colab_type="text"
# ## Plot the Results
# + id="t-M9fEvWD_ZJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="0e1c52af-3ffd-4811-8e3c-0bb4d4b80766"
model_path_full = path + '/' + model_path + '/'
# Plot the training and validation loss for each epoch
print('Generating loss chart...')
plt.plot(history_object.history['loss'])
plt.plot(history_object.history['val_loss'])
plt.title('model mean squared error loss')
plt.ylabel('mean squared error loss')
plt.xlabel('epoch')
plt.legend(['training set', 'validation set'], loc='upper right')
plt.savefig(model_path_full + 'model.png')
# Done
print('Done.')
# + [markdown] id="QMGEoYbVHnaJ" colab_type="text"
# # References:
# https://www.pyimagesearch.com/2019/01/28/keras-regression-and-cnns/
# https://www.pyimagesearch.com/2019/01/21/regression-with-keras/
# https://www.pyimagesearch.com/2018/12/24/how-to-use-keras-fit-and-fit_generator-a-hands-on-tutorial/
# https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l04c01_image_classification_with_cnns.ipynb#scrollTo=7MqDQO0KCaWS
| rover_ml/utils/Image_Regression_with_CNNs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ty3117/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/Tyler_Sheppatd_LS_DS_123_Make_Explanatory_Visualizations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="EW8Asex3ApCE" colab_type="code" outputId="724d8136-ad74-4110-e6e6-23e79e605587" colab={"base_uri": "https://localhost:8080/", "height": 277}
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.style.use('fivethirtyeight')
fake = pd.Series([38, 3, 2, 1, 2, 4, 6, 5, 5, 33])
fake.plot.bar(color='C1', width=0.9);
# + id="Sig0blGxBoJ3" colab_type="code" outputId="3c6f46cd-ad5f-4cc5-8135-2ad45109a5a4" colab={"base_uri": "https://localhost:8080/", "height": 289}
fake2=pd.Series(
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
2, 2, 2,
3, 3, 3,
4, 4,
5, 5, 5,
6, 6, 6, 6,
7, 7, 7, 7, 7,
8, 8, 8, 8,
9, 9, 9, 9,
10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10])
fake2.value_counts().sort_index().plot.bar(color='C1', width=0.9);
# + id="j4gG1uGgB_0z" colab_type="code" outputId="6150c600-0a01-4564-be5b-ef2753dac318" colab={"base_uri": "https://localhost:8080/", "height": 438}
from IPython.display import display, Image
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=500)
display(example)
# + id="Q2D9PsKODGV_" colab_type="code" outputId="f43c49c0-dfd8-42d2-bd0b-f2d47c8723a6" colab={"base_uri": "https://localhost:8080/", "height": 374}
counts = [38, 3, 2, 1, 2, 4, 6, 5, 5, 33]
data_list = []
for i, c in enumerate(counts, 1):
data_list = data_list + [i]*c
fake2 = pd.Series(data_list)
plt.style.use('fivethirtyeight')
fake2.value_counts().sort_index().plot.bar(color='C1', width=0.9, rot=0);
plt.text(x=-1,
y=50,
fontsize=16,
fontweight='bold',
s="An Inconvenient Sequel: Truth To Power' is divisive")
plt.text(x=-1,
y=45,
fontsize=16,
s="IMDb ratings for the film as of Aug. 29")
plt.xlabel('Rating')
plt.ylabel('Percent of Votes')
plt.yticks(range(0, 50, 10));
# + id="8DGecv9DGMPm" colab_type="code" outputId="914b6f55-2bdc-40be-9ecf-4fafd7c9fbd7" colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/inconvenient-sequel/ratings.csv')
df.shape
# + id="ujlG70UGIbZJ" colab_type="code" outputId="70a27329-ea15-4fe9-abd8-3757f6ed2094" colab={"base_uri": "https://localhost:8080/", "height": 299}
df.head()
# + id="s-pp1W18IfSD" colab_type="code" outputId="846b5764-3aee-4db7-e3c0-91cc61795236" colab={"base_uri": "https://localhost:8080/", "height": 846}
df.sample(1).T
# + id="g4O21qheIloH" colab_type="code" outputId="030c6b56-b257-46c6-eb06-9eb6d4fd41a9" colab={"base_uri": "https://localhost:8080/", "height": 299}
df['timestamp'] = pd.to_datetime(df['timestamp'])
df.head()
# + id="M-WOSwstJMW2" colab_type="code" outputId="5a91132f-a797-45f5-c883-2e6da97a9362" colab={"base_uri": "https://localhost:8080/", "height": 329}
df = df.set_index('timestamp')
df.head()
# + id="840YPJ3JJ0Zd" colab_type="code" outputId="3b710adb-a9ce-410c-c2af-bf613c0feb35" colab={"base_uri": "https://localhost:8080/", "height": 353}
df['category'].value_counts()
# + id="ApaFuEXoJ9EW" colab_type="code" outputId="eeac66ff-d0d8-40c4-c31a-e8bc3044d4f3" colab={"base_uri": "https://localhost:8080/", "height": 34}
df_imdb = df[df['category'] == 'IMDb users']
df_imdb.shape
# + id="4Lf1tCNjKlii" colab_type="code" outputId="f2335704-bcf6-4d8e-8d07-3a0a4977e964" colab={"base_uri": "https://localhost:8080/", "height": 313}
lastday = df['2017-08-29']
lastday[lastday['category'] == 'IMDb users']['respondents'].plot()
# + id="vYfksIfYLro3" colab_type="code" outputId="4f9227a2-010e-4577-ccae-bf3246e25d93" colab={"base_uri": "https://localhost:8080/", "height": 143}
df = df.sort_index()
df_imdb = df[df['category'] == 'IMDb users']
final = df_imdb.tail(1)
final
# + id="Pzow1AJVMNLx" colab_type="code" outputId="d5a9dd24-4b02-4367-fe67-7bde26869579" colab={"base_uri": "https://localhost:8080/", "height": 106}
columns = ['%s_pct' % i for i in range(1,11)]
final[columns]
# + id="RIgRL1o4MrqS" colab_type="code" outputId="8ee5d0c1-15b5-47ae-d4f3-fa4c7405ca2e" colab={"base_uri": "https://localhost:8080/", "height": 375}
data = final[columns].T
data.index = range(1,11)
plt.style.use('fivethirtyeight')
data.plot.bar(color='#ed713a', width=0.9, legend=False)
plt.text(x=-2,
y=50,
fontsize=16,
fontweight='bold',
s="An Inconvenient Sequel: Truth To Power' is divisive")
plt.text(x=-2,
y=45,
fontsize=16,
s="IMDb ratings for the film as of Aug. 29")
plt.xlabel('Rating')
plt.ylabel('Percent of Votes')
plt.yticks(range(0, 50, 10));
# + id="fpB_W8v7O6uw" colab_type="code" outputId="799020ab-d0d3-44f3-94d9-afb36467a820" colab={"base_uri": "https://localhost:8080/", "height": 438}
display(example)
# + [markdown] id="7RFm4q9iRIUu" colab_type="text"
# **Stretch Goals**
# + id="gc9o7ol7PThH" colab_type="code" outputId="7f2a4055-f3a9-4b9d-d357-fdab1892ecb4" colab={"base_uri": "https://localhost:8080/", "height": 639}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/checking-our-work-data/master/nba_games.csv')
df.head(20)
# + id="0fkM_Zt9RSMO" colab_type="code" colab={}
# + [markdown] id="p5hS1IGHasr6" colab_type="text"
# I couldn't find a dataset to make a good visualization with :(
# + id="1o7IW7JQRTsx" colab_type="code" colab={}
| Tyler_Sheppatd_LS_DS_123_Make_Explanatory_Visualizations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf-lattice
# language: python
# name: tf-lattice
# ---
# # Basics of lattice models
# In this notebook, we'll explain a lattice model, an interpolated lookup table.
# In addition, we'll show how monotonicity and smooth regularizers can change the model.
# First we need to import libraries we're going to use.
# !pip install tensorflow_lattice
import tensorflow as tf
import tensorflow_lattice as tfl
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
# # Lattice model visualization
# Now, let us define helper functions for visualizing the surface of 2d lattice.
# +
# Hypercube (multilinear) interpolation in a 2 x 2 lattice.
# params[0] == lookup value at (0, 0)
# params[1] == lookup value at (0, 1)
# params[2] == lookup value at (1, 0)
# params[3] == lookup value at (1, 1)
def twod(x1, x2, params):
y = ((1 - x1) * (1 - x2) * params[0]
+ (1 - x1) * x2 * params[1]
+ x1 * (1 - x2) * params[2]
+ x1 * x2 * params[3])
return y
# This function will generate 3d plot for lattice function values.
# params uniquely characterizes the lattice lookup values.
def lattice_surface(params):
print('Lattice params:')
print(params)
# %matplotlib inline
fig = plt.figure()
ax = fig.gca(projection='3d')
# Make data.
n = 50
xv, yv = np.meshgrid(np.linspace(0.0, 1.0, num=n),
np.linspace(0.0, 1.0, num=n))
zv = np.zeros([n, n])
for k1 in range(n):
for k2 in range(n):
zv[k1, k2] = twod(xv[k1, k2], yv[k1, k2], params)
# Plot the surface.
surf = ax.plot_surface(xv, yv, zv, cmap=cm.coolwarm)
# Customize the z axis.
ax.set_zlim(0.0, 1.0)
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5,)
# -
# Let's draw a surface of 2d lattice model.
# This model represents an "XOR" function.
# This will plot the surface plot.
lattice_surface([0.0, 1.0, 1.0, 0.0])
# # Train XOR function
# We'll provide a synthetic data that represents the "XOR" function, that is
#
# f(0, 0) = 0
# f(0, 1) = 1
# f(1, 0) = 1
# f(1, 1) = 0
#
# and check whether a lattice can __learn__ this function.
# +
# Reset the graph.
tf.reset_default_graph()
# Prepare the dataset.
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [0.0]]
# Define placeholders.
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, _, _) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1)
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# Iterate 100 times
for _ in range(100):
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw the surface!
lattice_surface(lattice_param_val[0])
# -
# # Train with monotonicity
# Now we'll set monotonicity in a lattice model. We'll use the same synthetic data generated by "XOR" function, but now we'll require full monotonicity in both directions, x1 and x2. Note that the data does not contain monotonicity, since "XOR" function value decreases, i.e., f(1, 0) > f(1, 1) and f(0, 1) > f(1, 1).
# So the trained model will do its best to fit the data while satisfying the monotonicity.
# +
tf.reset_default_graph()
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [0.0]]
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, projection_op, _) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1, is_monotone=True)
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# Iterate 100 times
for _ in range(100):
# Apply gradient.
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Then projection.
sess.run(projection_op)
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw it!
# You can see that the prediction does not decrease.
lattice_surface(lattice_param_val[0])
# -
# # Train with partial monotonicity
# Now we'll set partial monotonicity. Here only one input is constrained to be monotonic.
# +
tf.reset_default_graph()
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [0.0]]
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, projection_op, _) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1, is_monotone=[True, False])
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# Iterate 100 times
for _ in range(100):
# Apply gradient.
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Then projection.
sess.run(projection_op)
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw it!
# You can see that the prediction does not decrease in one direction.
lattice_surface(lattice_param_val[0])
# -
# # Training OR function
# Now we switch to a synthetic dataset generated by "OR" function to illustrate other regularizers.
# f(0, 0) = 0
# f(0, 1) = 1
# f(1, 0) = 1
# f(1, 1) = 1
# +
tf.reset_default_graph()
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [1.0]]
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, _, _) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1)
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# Iterate 100 times
for _ in range(100):
# Apply gradient.
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw it!
lattice_surface(lattice_param_val[0])
# -
# # Laplacian regularizer
# Laplacian regularizer puts the penalty on lookup value changes. In other words, it tries to make the slope of each face as small as possible.
# +
tf.reset_default_graph()
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [1.0]]
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, _, regularization) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1, l2_laplacian_reg=[0.0, 1.0])
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
loss += regularization
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# Iterate 100 times
for _ in range(1000):
# Apply gradient.
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw it!
# With heavy Laplacian regularization along the second axis, the second axis's slope becomes zero.
lattice_surface(lattice_param_val[0])
# -
# # Torsion regularizer
# Torsion regularizer penalizes nonlinear interactions in the feature.
# +
tf.reset_default_graph()
x_data = [[0.0, 0.0], [0.0, 1.0], [1.0, 0.0], [1.0, 1.0]]
y_data = [[0.0], [1.0], [1.0], [1.0]]
x = tf.placeholder(dtype=tf.float32, shape=(None, 2))
y_ = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# 2 x 2 lattice with 1 output.
# lattice_param is [output_dim, 4] tensor.
lattice_sizes = [2, 2]
(y, lattice_param, _, regularization) = tfl.lattice_layer(
x, lattice_sizes=[2, 2], output_dim=1, l2_torsion_reg=1.0)
# Sqaured loss
loss = tf.reduce_mean(tf.square(y - y_))
loss += regularization
# Minimize!
train_op = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(loss)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# Iterate 1000 times
for _ in range(1000):
# Apply gradient.
sess.run(train_op, feed_dict={x: x_data, y_: y_data})
# Fetching trained lattice parameter.
lattice_param_val = sess.run(lattice_param)
# Draw it!
# With heavy Torsion regularization, the model becomes a linear model.
lattice_surface(lattice_param_val[0])
| extras/tensorflow_lattice/04_lattice_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="WcarqQUXNrGB"
# # <font color='red'>Table of Contents</font>
#
# + [markdown] colab_type="text" id="xK3qVL4pNt19"
# [15. Multi Task Learning](#section15)<br>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="s0-OQReUNxm9" outputId="4c3d5b56-ff6e-4519-d172-c382e6a5edce"
# Research Kernel Link - https://arxiv.org/pdf/2003.02261.pdf
import pandas as pd
import numpy as np
import itertools
import os
import sys
from prettytable import PrettyTable
import pickle
import multiprocessing
from multiprocessing.pool import ThreadPool
from tqdm import tqdm_notebook
print(multiprocessing.cpu_count()," CPU cores")
import seaborn as sns
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams["axes.grid"] = False
from sklearn.metrics import confusion_matrix, cohen_kappa_score,accuracy_score
from PIL import Image
import cv2
import keras
from keras import applications
from keras.preprocessing.image import ImageDataGenerator
from keras import optimizers,Model,Sequential
from keras.layers import Input,GlobalAveragePooling2D,Dropout,Dense,Activation,BatchNormalization,GlobalMaxPooling2D,concatenate,Flatten
from keras.callbacks.callbacks import EarlyStopping,ReduceLROnPlateau,Callback
from keras.initializers import random_normal
from keras.models import load_model
from keras.losses import binary_crossentropy,categorical_crossentropy,mean_squared_error
from keras import backend as K
import tensorflow as tf
#import shap
# Colab Libs...
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
# + [markdown] colab_type="text" id="gsWsEmxVNzmI"
# # <a id = 'section15'> <font color='red'> 15. Multi task learning </font> </a>
# + [markdown] colab_type="text" id="3ghH0prCN6_F"
# ### <font color='red'> 15.1 Setup Colab Environment </font>
#
# + colab={} colab_type="code" id="YSpwm4uBCwe3"
'''
The codes below uses Authentication URL to allow loading of Files and Images from Google Drive to Colab Memory
'''
# + colab={} colab_type="code" id="DaZG-E-IOE7F"
# Importing Libraries
#ref - https://buomsoo-kim.github.io/colab/2018/04/16/Importing-files-from-Google-Drive-in-Google-Colab.md/
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="EbafLLivOG06" outputId="8dfee3c5-b3cb-4cbf-c86c-ba12c6d25e08"
from google.colab import drive
drive.mount('/content/gdrive',force_remount = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="hmdV3sZDOOsH" outputId="c12a203d-60de-4702-b612-9bfa5a3c2703"
import os
os.chdir('/content/gdrive/My Drive/aptos2019')
print("We are currently in the folder of ",os.getcwd())
# + [markdown] colab_type="text" id="iqbNUH2r6dTa"
# ### <font color='red'> 15.2 Load data and preprocess</font>
#
# + colab={} colab_type="code" id="kG55l44tCzV1"
'''
Loading Train and Validation Data from Previously split Train/Test data
'''
# + colab={} colab_type="code" id="a1y2H02GOQXu"
def load_data():
file = open('df_train_train', 'rb')
df_train_train = pickle.load(file)
file.close()
file = open('df_train_test', 'rb')
df_train_test = pickle.load(file)
file.close()
return df_train_train,df_train_test
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" id="AsdpEXif7Cto" outputId="69a0a7fe-a4e9-4a8a-efca-8379bb79acc3"
df_train_train,df_train_test = load_data()
print(df_train_train.shape,df_train_test.shape,'\n')
df_train_train.head(6)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="cwTSL-XLIYPv" outputId="2bbf64d0-8382-4ddd-9b8d-8c729e2cc2b0"
print(len(os.listdir("./train_images_resized_preprocessed/")),len(os.listdir("./test_images_resized_preprocessed/")))
# + colab={} colab_type="code" id="L_-O--AhOXXD"
def crop_image_from_gray(img,tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
# print(img1.shape,img2.shape,img3.shape)
img = np.stack([img1,img2,img3],axis=-1)
# print(img.shape)
return img
def circle_crop(img, sigmaX = 30):
"""
Create circular crop around image centre
"""
img = crop_image_from_gray(img)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
height, width, depth = img.shape
x = int(width/2)
y = int(height/2)
r = np.amin((x,y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x,y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image_from_gray(img)
img=cv2.addWeighted(img,4, cv2.GaussianBlur( img , (0,0) , sigmaX) ,-4 ,128)
return img
def preprocess_image(file):
input_filepath = os.path.join('./','train_images_resized','{}.png'.format(file))
output_filepath = os.path.join('./','train_images_resized_preprocessed','{}.png'.format(file))
img = cv2.imread(input_filepath)
img = circle_crop(img)
cv2.imwrite(output_filepath, cv2.resize(img, (IMG_SIZE,IMG_SIZE)))
# + colab={} colab_type="code" id="xviDidlMLQtM"
'''
Defining Global Variables to be used in this notebook
'''
FACTOR = 4
BATCH_SIZE = 8 * FACTOR
IMG_SIZE = 512
EPOCHS = 20
WARMUP_EPOCHS = 5
LEARNING_RATE = 1e-4 * FACTOR
WARMUP_LEARNING_RATE = 1e-3 * FACTOR
HEIGHT = 320
WIDTH = 320
CANAL = 3
N_CLASSES = df_train_train['diagnosis'].nunique()
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
LR_WARMUP_EPOCHS_1st = 2
LR_WARMUP_EPOCHS_2nd = 5
STEP_SIZE = len(df_train_train) // BATCH_SIZE
TOTAL_STEPS_1st = WARMUP_EPOCHS * STEP_SIZE
TOTAL_STEPS_2nd = EPOCHS * STEP_SIZE
WARMUP_STEPS_1st = LR_WARMUP_EPOCHS_1st * STEP_SIZE
WARMUP_STEPS_2nd = LR_WARMUP_EPOCHS_2nd * STEP_SIZE
# + [markdown] colab_type="text" id="YQQrFrG8CrSS"
# ### <font color='red'> 15.3 Custom Image Data generator</font>
#
# + colab={} colab_type="code" id="tnKuoUQUC5Pc"
'''
This Function creates a custom Image Data generator,
Since this is a Multi Output Model, a custom Image data generator is used (which yeild's outputs to next iterator)
'''
# + colab={} colab_type="code" id="-tlnoyOdJ0MM"
# custom generator ref - https://classifai.net/blog/multiple-outputs-keras/
# ref - https://stackoverflow.com/questions/54143458/convert-categorical-data-back-to-numbers-using-keras-utils-to-categorical
def multiple_outputs(generator,dataframe, image_dir, batch_size, height,width, subset):
gen = generator.flow_from_dataframe(
dataframe = dataframe,
x_col = "file_name",
y_col = "diagnosis",
directory = image_dir,
target_size=(height, width),
batch_size=batch_size,
class_mode='categorical',
subset=subset)
mlb = MultiLabelBinarizer(classes = range(N_CLASSES))
while True:
gnext = gen.next()
yield gnext[0], [np.argmax(gnext[1],axis = -1),gnext[1],mlb.fit_transform([list(range(x+1)) for x in np.argmax(gnext[1],axis = -1)])]
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="n8YzFFqHK15Q" outputId="3093a6cb-2614-48c7-8830-07bde108edb3"
train_datagen=ImageDataGenerator(rescale=1./255, rotation_range=360,brightness_range=[0.5, 1.5],
zoom_range=[1, 1.2],zca_whitening=True,horizontal_flip=True,
vertical_flip=True,fill_mode='constant',cval=0.,validation_split = 0.0)
train_generator = multiple_outputs(generator = train_datagen,dataframe = df_train_train,
image_dir="./train_images_resized_preprocessed/",
batch_size=BATCH_SIZE,height = HEIGHT,width = WIDTH,
subset='training')
valid_generator = multiple_outputs(generator = train_datagen,dataframe = df_train_test,
image_dir="./test_images_resized_preprocessed/",
batch_size=BATCH_SIZE,height = HEIGHT,width = WIDTH,
subset='validation')
# + [markdown] colab_type="text" id="VCFNVYNoOdhB"
# ### <font color='red'> 15.4 Stage 1 (Pre Training) using ResNet50</font>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="Mwx2DeXP8MCO" outputId="d3bc4b98-0bad-440b-e75f-e0e265ae08c8"
'''Implementing this Stage 1 as mentioned in the Research paper'''
input_tensor = Input(shape=(HEIGHT, WIDTH, CANAL))
base_model = applications.ResNet50(weights=None, include_top=False,input_tensor=input_tensor)
base_model.load_weights('resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5')
x1 = GlobalAveragePooling2D()(base_model.output)
x1 = BatchNormalization()(x1)
x2 = GlobalMaxPooling2D()(base_model.output)
x2 = BatchNormalization()(x2)
x = concatenate([x1,x2])
# Regression Head
xr = Dense(2048, activation='relu')(x)
xr = Dropout(0.5)(xr)
xr = Dense(1,activation = 'linear',name = 'regression_output')(xr)
# Classification Head
xc = Dense(2048, activation='relu')(x)
xc = Dropout(0.5)(xc)
xc = Dense(N_CLASSES,activation = 'softmax',name = 'classification_output')(xc)
# Ordinal Regression Head
xo = Dense(2048, activation='relu')(x)
xo = Dropout(0.5)(xo)
xo = Dense(N_CLASSES,activation = 'softmax',name = 'ordinal_regression_output')(xo)
model = Model(inputs = [input_tensor], outputs = [xr,xc,xo])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="DehalZUM9Sf1" outputId="57eefeda-a4ef-415b-8bbb-ea14027c9f67"
# Train all Layers
for layer in model.layers:
layer.trainable = True
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="i4NV7v51KIeR" outputId="77becb7a-618f-49e2-ad15-4689fb485a4c"
STEP_SIZE_TRAIN = len(df_train_train)//BATCH_SIZE
STEP_SIZE_VALID = len(df_train_test)//BATCH_SIZE
print(STEP_SIZE_TRAIN,STEP_SIZE_VALID)
# + colab={} colab_type="code" id="AGk1j-cmEkCo"
''' This Code part includes Class & Function Implementatin of the Cosine Learning Rate Scheduler'''
# ref - https://github.com/dimitreOliveira/APTOS2019BlindnessDetection/blob/master/Best%20solution%20(Bronze%20medal%20-%20163rd%20place)/233%20-%20EfficientNetB5-Reg-Img224%200%2C5data%20Fold1.ipynb
def cosine_decay_with_warmup(global_step,
learning_rate_base,
total_steps,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0):
"""
Cosine decay schedule with warm up period.
In this schedule, the learning rate grows linearly from warmup_learning_rate
to learning_rate_base for warmup_steps, then transitions to a cosine decay
schedule.
:param global_step {int}: global step.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param global_step {int}: global step.
:Returns : a float representing learning rate.
:Raises ValueError: if warmup_learning_rate is larger than learning_rate_base, or if warmup_steps is larger than total_steps.
"""
if total_steps < warmup_steps:
raise ValueError('total_steps must be larger or equal to warmup_steps.')
learning_rate = 0.5 * learning_rate_base * (1 + np.cos(
np.pi *
(global_step - warmup_steps - hold_base_rate_steps
) / float(total_steps - warmup_steps - hold_base_rate_steps)))
if hold_base_rate_steps > 0:
learning_rate = np.where(global_step > warmup_steps + hold_base_rate_steps,
learning_rate, learning_rate_base)
if warmup_steps > 0:
if learning_rate_base < warmup_learning_rate:
raise ValueError('learning_rate_base must be larger or equal to warmup_learning_rate.')
slope = (learning_rate_base - warmup_learning_rate) / warmup_steps
warmup_rate = slope * global_step + warmup_learning_rate
learning_rate = np.where(global_step < warmup_steps, warmup_rate,
learning_rate)
return np.where(global_step > total_steps, 0.0, learning_rate)
class WarmUpCosineDecayScheduler(Callback):
"""Cosine decay with warmup learning rate scheduler"""
def __init__(self,
learning_rate_base,
total_steps,
global_step_init=0,
warmup_learning_rate=0.0,
warmup_steps=0,
hold_base_rate_steps=0,
verbose=0):
"""
Constructor for cosine decay with warmup learning rate scheduler.
:param learning_rate_base {float}: base learning rate.
:param total_steps {int}: total number of training steps.
:param global_step_init {int}: initial global step, e.g. from previous checkpoint.
:param warmup_learning_rate {float}: initial learning rate for warm up. (default: {0.0}).
:param warmup_steps {int}: number of warmup steps. (default: {0}).
:param hold_base_rate_steps {int}: Optional number of steps to hold base learning rate before decaying. (default: {0}).
:param verbose {int}: quiet, 1: update messages. (default: {0}).
"""
super(WarmUpCosineDecayScheduler, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.global_step = global_step_init
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.hold_base_rate_steps = hold_base_rate_steps
self.verbose = verbose
self.learning_rates = []
def on_batch_end(self, batch, logs=None):
self.global_step = self.global_step + 1
lr = K.get_value(self.model.optimizer.lr)
self.learning_rates.append(lr)
def on_batch_begin(self, batch, logs=None):
lr = cosine_decay_with_warmup(global_step=self.global_step,
learning_rate_base=self.learning_rate_base,
total_steps=self.total_steps,
warmup_learning_rate=self.warmup_learning_rate,
warmup_steps=self.warmup_steps,
hold_base_rate_steps=self.hold_base_rate_steps)
K.set_value(self.model.optimizer.lr, lr)
if self.verbose > 0:
print('\nBatch %02d: setting learning rate to %s.' % (self.global_step + 1, lr))
# + colab={} colab_type="code" id="yzSnpqUmFYtD"
# Use Cosine LR Scheduler as callback
cosine_lr = WarmUpCosineDecayScheduler(learning_rate_base = LEARNING_RATE,
total_steps=TOTAL_STEPS_1st,
warmup_learning_rate=0.0,
warmup_steps=TOTAL_STEPS_1st,
hold_base_rate_steps=(2 * STEP_SIZE))
callback_list = [cosine_lr_1st]
# + colab={"base_uri": "https://localhost:8080/", "height": 776} colab_type="code" id="Aki2wj75JNuA" outputId="e61b66a0-59c9-4bd7-c0eb-edfc0035dfab"
# ref - https://keras.io/getting-started/functional-api-guide/
model.compile(optimizer = optimizers.SGD(lr=LEARNING_RATE),
loss={'regression_output': 'mean_absolute_error',
'classification_output': 'categorical_crossentropy',
'ordinal_regression_output' : 'binary_crossentropy'
},
metrics = ['accuracy'])
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=20,
callbacks = callback_list,
verbose=1).history
model.save("model_pre_training.h5")
f = open("history_pre_training","wb")
pickle.dump(history,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="j2rduAwJeWFF" outputId="045fe918-d3a0-4ae6-ace3-5053424d3811"
plt.figure(figsize=(8,5))
plt.plot(history['regression_output_loss'])
plt.plot(history['val_regression_output_loss'])
plt.title('Regression Model Loss - Pre Training')
plt.ylabel('Loss (MAE)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,21))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="-MhYaj8VfbGo" outputId="b27560ce-7a28-40cb-a0bb-1065d6aa4b46"
plt.figure(figsize=(8,5))
plt.plot(history['classification_output_loss'])
plt.plot(history['val_classification_output_loss'])
plt.title('Classification Model Loss - Pre Training')
plt.ylabel('Loss (Categorical Cross Entropy)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,21))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="Cr75LfqLfbs1" outputId="79d1272d-310b-4bb8-9440-16d0f15464ea"
plt.figure(figsize=(8,5))
plt.plot(history['ordinal_regression_output_loss'])
plt.plot(history['val_ordinal_regression_output_loss'])
plt.title('Ordinal Regression Model Loss - Pre Training')
plt.ylabel('Loss (Binary Cross Entropy)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,21))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + [markdown] colab_type="text" id="WnYIX-05RkIb"
# ### <font color='red'> 15.5 Stage 2 (Main Training) </font>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="CF2lgRdKQPbn" outputId="318d7da7-3581-49cc-a2a6-026150bdbdbe"
# Freezing the Encoder Layers Training..(Only Last 14 layers are trainable) for 5 Epochs (Warming up the Weights for Main Training)
for layer in model.layers:
layer.trainable = False
for i in range(-14,0):
model.layers[i].trainable = True
model.summary()
# + colab={} colab_type="code" id="5dJgeOMnRl3G"
# ref - https://github.com/umbertogriffo/focal-loss-keras/blob/master/losses.py
'''Below Functions create custom loss functions - Categorical Focal Loss, Binary Focal Loss'''
def binary_focal_loss(gamma=2., alpha=.25):
"""
Binary form of focal loss.
FL(p_t) = -alpha * (1 - p_t)**gamma * log(p_t)
where p = sigmoid(x), p_t = p or 1 - p depending on if the label is 1 or 0, respectively.
References:
https://arxiv.org/pdf/1708.02002.pdf
Usage:
model.compile(loss=[binary_focal_loss(alpha=.25, gamma=2)], metrics=["accuracy"], optimizer=adam)
"""
def binary_focal_loss_fixed(y_true, y_pred):
"""
:param y_true: A tensor of the same shape as `y_pred`
:param y_pred: A tensor resulting from a sigmoid
:return: Output tensor.
"""
pt_1 = tf.where(tf.equal(y_true, 1), y_pred, tf.ones_like(y_pred))
pt_0 = tf.where(tf.equal(y_true, 0), y_pred, tf.zeros_like(y_pred))
epsilon = K.epsilon()
# clip to prevent NaN's and Inf's
pt_1 = K.clip(pt_1, epsilon, 1. - epsilon)
pt_0 = K.clip(pt_0, epsilon, 1. - epsilon)
return -K.mean(alpha * K.pow(1. - pt_1, gamma) * K.log(pt_1)) \
-K.mean((1 - alpha) * K.pow(pt_0, gamma) * K.log(1. - pt_0))
return binary_focal_loss_fixed
# + colab={} colab_type="code" id="JDC7DXOGtaa-"
def categorical_focal_loss(gamma=2., alpha=.25):
"""
Softmax version of focal loss.
m
FL = ∑ -alpha * (1 - p_o,c)^gamma * y_o,c * log(p_o,c)
c=1
where m = number of classes, c = class and o = observation
Parameters:
alpha -- the same as weighing factor in balanced cross entropy
gamma -- focusing parameter for modulating factor (1-p)
Default value:
gamma -- 2.0 as mentioned in the paper
alpha -- 0.25 as mentioned in the paper
References:
Official paper: https://arxiv.org/pdf/1708.02002.pdf
https://www.tensorflow.org/api_docs/python/tf/keras/backend/categorical_crossentropy
Usage:
model.compile(loss=[categorical_focal_loss(alpha=.25, gamma=2)], metrics=["accuracy"], optimizer=adam)
"""
def categorical_focal_loss_fixed(y_true, y_pred):
"""
:param y_true: A tensor of the same shape as `y_pred`
:param y_pred: A tensor resulting from a softmax
:return: Output tensor.
"""
# Scale predictions so that the class probas of each sample sum to 1
y_pred /= K.sum(y_pred, axis=-1, keepdims=True)
# Clip the prediction value to prevent NaN's and Inf's
epsilon = K.epsilon()
y_pred = K.clip(y_pred, epsilon, 1. - epsilon)
# Calculate Cross Entropy
cross_entropy = -y_true * K.log(y_pred)
# Calculate Focal Loss
loss = alpha * K.pow(1 - y_pred, gamma) * cross_entropy
# Compute mean loss in mini_batch
return K.mean(loss, axis=1)
return categorical_focal_loss_fixed
# + colab={"base_uri": "https://localhost:8080/", "height": 272} colab_type="code" id="tyP6URxCtk_G" outputId="30cc874e-4bea-4ab2-8c4f-dd541d062c23"
# ref - https://keras.io/getting-started/functional-api-guide/
model.compile(optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE),
loss={'regression_output': mean_squared_error,
'classification_output': categorical_focal_loss(alpha=.25, gamma=2) ,
'ordinal_regression_output' : binary_focal_loss(alpha=.25, gamma=2)
},
metrics = ['accuracy'])
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=5,
callbacks = callback_list,
verbose=1).history
model.save("model_main_training.h5")
f = open("history_main_training","wb")
pickle.dump(history,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="HGzThk-2QeP7" outputId="a1e74209-0ab0-468a-f3fb-03cde94e8311"
# Now Unfreeze all Layers and make all layers as trainable = True (Train for 45 Epochs)
for layer in model.layers:
layer.trainable = True
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="qpmGwpjCQec9" outputId="0a3e573e-e8b1-4767-8af4-881c30a979b6"
# ref - https://keras.io/getting-started/functional-api-guide/
model.compile(optimizer = optimizers.Adam(lr=LEARNING_RATE),
loss={'regression_output': mean_squared_error,
'classification_output': categorical_focal_loss(alpha=.25, gamma=2) ,
'ordinal_regression_output' : binary_focal_loss(alpha=.25, gamma=2)
},
metrics = ['accuracy'])
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=45,
callbacks = callback_list,
verbose=1).history
model.save("model_main_training.h5")
f = open("history_main_training","wb")
pickle.dump(history,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="PBIobuYSNDbu" outputId="2558bd40-9a2f-4324-eff9-48acf4855698"
plt.figure(figsize=(15,5))
plt.plot(history['regression_output_loss'])
plt.plot(history['val_regression_output_loss'])
plt.title('Regression Model Loss - Main Training')
plt.ylabel('Loss (MSE)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,46))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="XQlUK5u8NDjt" outputId="3eec29b9-abf5-409a-9aac-c41fd31c9fb0"
plt.figure(figsize=(15,5))
plt.plot(history['classification_output_loss'])
plt.plot(history['val_classification_output_loss'])
plt.title('Classification Model Loss - Main Training')
plt.ylabel('Loss (Categorical Focal Loss)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,46))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="tgYxfJGxNDqi" outputId="c7aba244-c445-4034-9615-4a9f537c6bfb"
plt.figure(figsize=(15,5))
plt.plot(history['ordinal_regression_output_loss'])
plt.plot(history['val_ordinal_regression_output_loss'])
plt.title('Ordinal Regression Model Loss - Main Training')
plt.ylabel('Loss (Binary Focal Loss)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,46))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + [markdown] colab_type="text" id="pOZFyljJRnMS"
# ### <font color='red'> 15.6 Stage 3 (Post Training) </font>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 104} colab_type="code" id="TPItySzSb_de" outputId="ffbae475-c00f-4c3c-ffdb-a68ad41aad5d"
'''
Get the Outputs from the 3 Heads - (Classification,Regression,Ordinal Regression) to pass the outputs to a new Model
'''
complete_datagen = ImageDataGenerator(rescale=1./255, rotation_range=360,brightness_range=[0.5, 1.5],
zoom_range=[1, 1.2],zca_whitening=True,horizontal_flip=True,
vertical_flip=True,fill_mode='constant')
complete_generator = complete_datagen.flow_from_dataframe(dataframe=df_train_train,
directory = "./train_images_resized_preprocessed/",
x_col="file_name",
target_size=(HEIGHT, WIDTH),
batch_size=1,
shuffle=False,
class_mode=None)
STEP_SIZE_COMPLETE = complete_generator.n//complete_generator.batch_size
print(complete_generator.n)
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="5S3C2x2Hwo8-" outputId="6179394a-08aa-4b4f-fcfa-ac1e2b3eb77e"
test_generator = complete_datagen.flow_from_dataframe(dataframe=df_train_test,
directory = "./test_images_resized_preprocessed/",
x_col="file_name",
target_size=(HEIGHT, WIDTH),
batch_size=1,
shuffle=False,
class_mode=None)
STEP_SIZE_TEST = test_generator.n//test_generator.batch_size
print(test_generator.n)
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="jPydELR1b_gF" outputId="7addea96-6e1c-46a3-c9ec-c1acd3e890fe"
model = load_model("model_main_training.h5",custom_objects={'categorical_focal_loss_fixed':categorical_focal_loss(alpha=.25, gamma=2),
'binary_focal_loss_fixed' : binary_focal_loss(alpha=.25, gamma=2)})
train_preds = model.predict_generator(complete_generator, steps=STEP_SIZE_COMPLETE,verbose = 1)
f = open("train_preds","wb")
pickle.dump(train_preds,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 121} colab_type="code" id="Snsubop3wzXg" outputId="f30b7332-888c-4f10-808a-4419f32f56ff"
test_preds = model.predict_generator(test_generator, steps=STEP_SIZE_TEST,verbose = 1)
f = open("test_preds","wb")
pickle.dump(test_preds,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 67} colab_type="code" id="mCPQGqWJqrIX" outputId="66cc6979-049d-4f49-d614-9897096570bb"
print(train_preds[0].shape,train_preds[1].shape,train_preds[2].shape)
train_output_regression = np.array(train_preds[0]).reshape(-1,1)
train_output_classification = np.array(np.argmax(train_preds[1],axis = -1)).reshape(-1,1)
train_output_ordinal_regression = np.array(np.sum(train_preds[2],axis = -1)).reshape(-1,1)
print(train_output_regression.shape,train_output_classification.shape,train_output_ordinal_regression.shape)
X_train = np.hstack((train_output_regression,train_output_classification,train_output_ordinal_regression))
print(X_train.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 67} colab_type="code" id="MDQmh3iaw68C" outputId="9c963b7d-26f7-43f1-f5fb-9d311325997e"
print(test_preds[0].shape,test_preds[1].shape,test_preds[2].shape)
test_output_regression = np.array(test_preds[0]).reshape(-1,1)
test_output_classification = np.array(np.argmax(test_preds[1],axis = -1)).reshape(-1,1)
test_output_ordinal_regression = np.array(np.sum(test_preds[2],axis = -1)).reshape(-1,1)
print(test_output_regression.shape,test_output_classification.shape,test_output_ordinal_regression.shape)
X_test = np.hstack((test_output_regression,test_output_classification,test_output_ordinal_regression))
print(X_test.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 185} colab_type="code" id="woVkwW_2Oicp" outputId="c2271671-2e11-4f2b-e5c4-55ee47957af0"
model_post = Sequential()
model_post.add(Dense(1, activation='linear', input_shape=(3,)))
model_post.compile(optimizer=optimizers.SGD(lr=LEARNING_RATE), loss='mean_squared_error', metrics=['mean_squared_error'])
model_post.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="c8Tnmh0FVWKV" outputId="904b600f-e4a1-4565-da2e-50b0348d9d5b"
history = model_post.fit(X_train,np.array(df_train_train.diagnosis.values),
batch_size=BATCH_SIZE,
epochs=50,
verbose=1,
validation_data = (X_test,np.array(df_train_test.diagnosis.values)))
model_post.save("model_post_training.h5")
f = open("history_post_training","wb")
pickle.dump(history,f)
f.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 342} colab_type="code" id="FtrJDNeCVWMo" outputId="6dd9acb8-458d-44ea-b087-c48915338f21"
plt.figure(figsize=(25,5))
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Linear Regression Model Loss - Post Training (50 epochs)')
plt.ylabel('Loss (MSE)')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='best')
plt.xticks(range(1,51))
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
plt.show()
# + [markdown] colab_type="text" id="UnpWnUyW4nuA"
# ### <font color='red'> 15.7 Evaluate Model performance (Test data)</font>
# + colab={} colab_type="code" id="Tb0PBdUbVWPJ"
# ref- https://github.com/dimitreOliveira/APTOS2019BlindnessDetection/blob/master/Model%20backlog/ResNet50/63%20-%20ResNet50%20-%20Regression%20-%20RGB%20scale.ipynb
'''This Function does nearest integer rounding for regression output from the post training model'''
def classify(x):
if x < 0.5:
return 0
elif x < 1.5:
return 1
elif x < 2.5:
return 2
elif x < 3.5:
return 3
return 4
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qySwX-6dVWRT" outputId="901fa894-e1b5-4e37-fece-ae2414fdf9e9"
train_labels = model_post.predict(X_train,batch_size=BATCH_SIZE,verbose = 1)
train_labels = np.apply_along_axis(classify, 1, train_labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="JGVHhia6VWT2" outputId="41d1be1b-33b0-4346-a520-442ec3653e83"
test_labels = model_post.predict(X_test,batch_size=BATCH_SIZE,verbose = 1)
test_labels = np.apply_along_axis(classify, 1, test_labels)
# + colab={} colab_type="code" id="iiQE5CcB4LSF"
# Plot Confusion Matrix..
def plot_conf_matrix(true,pred,classes):
cf = confusion_matrix(true, pred)
df_cm = pd.DataFrame(cf, range(len(classes)), range(len(classes)))
plt.figure(figsize=(8,5.5))
sns.set(font_scale=1.4)
sns.heatmap(df_cm, annot=True, annot_kws={"size": 16},xticklabels = classes ,yticklabels = classes,fmt='g')
#sns.heatmap(df_cm, annot=True, annot_kws={"size": 16})
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 485} colab_type="code" id="G-BTE9Cn4LVy" outputId="503e6077-479a-4840-fb31-ff4fa55f8da4"
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
plot_conf_matrix(list(df_train_test['diagnosis'].astype(int)),test_labels,labels)
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="d70ssg2u4sZ2" outputId="636ea908-970c-43d8-e4fd-51bc0ff1c934"
cnf_matrix = confusion_matrix(df_train_test['diagnosis'].astype('int'), test_labels)
cnf_matrix_norm = cnf_matrix.astype('float') / cnf_matrix.sum(axis=1)[:, np.newaxis]
df_cm = pd.DataFrame(cnf_matrix_norm, index=labels, columns=labels)
plt.figure(figsize=(16, 7))
sns.heatmap(df_cm, annot=True, fmt='.2f', cmap="Blues")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="l7c3uhJK432J" outputId="8919bd78-879f-41e5-db7c-5e292fcf503d"
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_labels, df_train_train['diagnosis'].astype('int'), weights='quadratic'))
print("Train Accuracy score : %.3f" % accuracy_score(df_train_train['diagnosis'].astype('int'),train_labels))
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="KjckvWfv4sc-" outputId="bbab7e29-0a52-4938-c2d1-d65ec1fd915a"
print("Test Cohen Kappa score: %.3f" % cohen_kappa_score(test_labels, df_train_test['diagnosis'].astype('int'), weights='quadratic'))
print("Test Accuracy score : %.3f" % accuracy_score(df_train_test['diagnosis'].astype('int'),test_labels))
| Extra/research_paper_implementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datascience import *
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# -
# ## Comparison ##
3 > 1
type(3 > 1)
True
true
3 = 3
3 == 3.0
10 != 2
x = 14
y = 3
x > 15
12 < x
x < 20
12 < x < 20
10 < x-y < 13
x > 13 and y < 3.14159
# ## Comparisons with arrays
pets = make_array('cat', 'cat', 'dog', 'cat', 'dog', 'rabbit')
pets == 'cat'
1 + 1 + 0 + 1 + 0 + 0
sum(make_array(True, True, False, True, False, False))
sum(pets == 'dog')
np.count_nonzero(pets == 'dog')
x = np.arange(20, 31)
x > 28
# ## Simulation
# Let's play a game. We roll a die once.
#
# If it comes up 1 or 2: you pay me a dollar.
#
# If it comes up 3 or 4: we do nothing.
#
# If it comes up 5 or 6: I pay you a dollar.
# ### Conditional Statements
# Work in progress
def one_round(roll):
if roll <= 2:
return 1
one_round(1)
one_round(4)
# Final correct version
def one_round(roll):
""" Meant for values from 1-6 (inclusive) """
if roll <= 2:
return 1
elif roll <= 4:
return 0
elif roll <= 6:
return -1
one_round(4)
one_round(6)
one_round(15)
# ### Random Selection
experiment_groups = make_array('treatment', 'control')
np.random.choice(experiment_groups)
np.random.choice(experiment_groups)
np.random.choice(experiment_groups)
np.random.choice(experiment_groups, 10)
sum(np.random.choice(experiment_groups, 10) == 'control')
sum(np.random.choice(experiment_groups, 10) == 'treatment')
individuals = np.random.choice(experiment_groups, 10)
individuals
sum(individuals == 'treatment')
sum(individuals == 'control')
die_faces = np.arange(1, 7)
np.random.choice(die_faces)
# Simulate one die roll and the subsequent payment in our game
def simulate_one_roll():
return one_round(np.random.choice(die_faces))
simulate_one_roll()
# ### Appending Arrays
first = np.arange(4)
second = np.arange(10, 17)
np.append(first, 6)
first
np.append(first, second)
first
second
# ### Repeated Betting ###
results = make_array()
results = np.append(results, simulate_one_roll())
results
# ## `For` Statements
for pet in make_array('cat', 'dog', 'rabbit'):
print('I love my ' + pet)
# +
# Unroll the above for loop to understand what happened:
pet = make_array('cat', 'dog', 'rabbit').item(0)
print('I love my ' + pet)
pet = make_array('cat', 'dog', 'rabbit').item(1)
print('I love my ' + pet)
pet = make_array('cat', 'dog', 'rabbit').item(2)
print('I love my ' + pet)
# +
game_outcomes = make_array()
for i in np.arange(5):
game_outcomes = np.append(game_outcomes, simulate_one_roll())
game_outcomes
# +
game_outcomes = make_array()
for i in np.arange(5000):
game_outcomes = np.append(game_outcomes, simulate_one_roll())
game_outcomes
# -
len(game_outcomes)
results = Table().with_column('Net Gain', game_outcomes)
results
results.group('Net Gain').barh(0)
# ### Simulate the Number of Heads in 100 Tosses ###
coin = make_array('heads', 'tails')
# Write a line (or two) of code to toss a coin 100 times, and count the number of heads.
np.count_nonzero(np.random.choice(coin, 100) == 'heads')
sum(np.random.choice(coin, 100) == 'heads')
# +
# Simulate one outcome
def num_heads():
return sum(np.random.choice(coin, 100) == 'heads')
# +
# Decide how many times you want to repeat the experiment
repetitions = 10000
# +
# Simulate that many outcomes
outcomes = make_array()
for i in np.arange(repetitions):
outcomes = np.append(outcomes, num_heads())
# -
heads = Table().with_column('Heads', outcomes)
heads
heads.hist(bins = np.arange(29.5, 70.6), normed = False)
# ## Optional: Advanced `where` ##
ages = make_array(16, 22, 18, 15, 19, 15, 16, 21)
age = Table().with_column('Age', ages)
age
age.where('Age', are.above_or_equal_to(18))
voter = ages >= 18
voter
age.where(voter)
is_voter = are.above_or_equal_to(18)
type(is_voter)
is_voter(22)
is_voter(3)
age.apply(is_voter, 'Age')
ages >= 18
voter
def my_voter_function(x):
return x >= 18
age.where('Age', are.above_or_equal_to(18))
age.where(voter)
age.where('Age', my_voter_function)
| lec/lec13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="5h66_taj8gkC" executionInfo={"status": "ok", "timestamp": 1638843014098, "user_tz": -330, "elapsed": 23649, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="5dbc9281-18c0-4396-bd32-bbc88076ebe0"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="XnL10uZ9_elu" executionInfo={"status": "ok", "timestamp": 1638843125689, "user_tz": -330, "elapsed": 111601, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="9e81358e-b18e-4893-a185-61cb6ee71b50"
''' install the desired pytorch version '''
# !pip3 install torch==1.2.0+cu92 torchvision==0.4.0+cu92 -f https://download.pytorch.org/whl/torch_stable.html
# + id="2CfVIE_6_hjj" executionInfo={"status": "ok", "timestamp": 1638843125692, "user_tz": -330, "elapsed": 30, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
''' importing the neccessary libraries '''
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
import numpy as np
import cv2
import matplotlib.pyplot as plt
from collections import OrderedDict
import pdb
from math import sqrt, ceil
from torch.autograd import Variable
from PIL import Image
from functools import partial
import sys
# + colab={"base_uri": "https://localhost:8080/"} id="UVOcVg-I_uWV" executionInfo={"status": "ok", "timestamp": 1638843659315, "user_tz": -330, "elapsed": 2205, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="6ab31034-c277-4836-d3a0-11dac4223c55"
''' load the CIFAR - 10 '''
''' taken from Pytorch Docs '''
## transforms are used to convert the data to tensors for easy use with CNN models
transform = transforms.Compose([
transforms.Resize(size=(32, 32)),
transforms.ToTensor(),
transforms.Normalize(
(0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010)
)
])
batch_size = 32
# downloading the training data through Pytorch API
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
#note num_workers attribute tells the number of workers(threads) that performs the task
#other attributes are pretty self-explanatory
#loading the training data
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size,
shuffle=True, num_workers=2)
# downloading the test data through Pytorch API
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
#loading the test data
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# + id="AZxYXoxjALw_" executionInfo={"status": "ok", "timestamp": 1638843538473, "user_tz": -330, "elapsed": 1007, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
#now the main part ,the code for the actual model
# We use the VGG16 Architecture (Zisserman et al)
class SMAI(nn.Module):
def __init__(self):
#not sure exactly how this works.Something to do with inheritance,
super(SMAI , self).__init__()
#we use default values of stride = 1
#and padding = 0
#our input is (3,32,32) C,H,W
#set out_channel as 64,so we get 32 kernels in the layer
# therefore output dimension are (64,32,32) C,H,W
#self.conv1 = nn.Conv2d(in_channels = 1,out_channels = 32 , kernel_size = 3)
#we want two linear layers
#first hidden layer
#self.d1 = nn.Linear(26*26*32,128)
#ouput layer
# 10 is the output class
#self.d2 = nn.Linear(128,10)
#the sequential is a container
#takes an input
#chains the input - output to the next layer
#returns the final output
#input BATCH_SIZE * 1 * 28 * 28 for MNIST
#commenting out some of the maxpooling Blocks to make sure kernel spatial dimension reduces to 7*7 and not below
self.features = nn.Sequential(
nn.Conv2d(3,16,3,padding = 1),
nn.ReLU(),
nn.MaxPool2d(2, stride = 2, return_indices = True) ,
#output 32,16,16,16
nn.Conv2d(16,32,3,padding = 1),
nn.ReLU(),
nn.MaxPool2d(2, stride = 2, return_indices = True) ,
#output 32,32,8,8
nn.Conv2d(32,32,3,padding = 1),
nn.ReLU()
#output 32,32,8,8
)
self.classifier = nn.Sequential(
nn.Linear(2048,128),
nn.Dropout(p=0.2),
# Linear layer with 10 output features
nn.Linear(128,10)
)
self.feature_maps = OrderedDict() #used to store all feature maps
self.max_pooling_locations = OrderedDict() #used to store the location of the values from max pools
self.Convolution_layers_indices = [0,3,6]
self.Relu_layers_indices = [1,4,7]
self.pooling_layers_indices = [2,5]
#a function to initialise with a pretrained model
#has to have the same architecture as the one define above
#must be compatible with above defined structure of features and classifier
def initialise_weights(self,model):
#copy the parameter values for features
for index,layer in enumerate(model.features):
#only relevant for learnable parameters,not for Pooling,Relu etc
if isinstance(layer,nn.conv2d):
self.features[index].weight.data = layer.weight.data
self.features[index].bias.data = layer.bias.data
#copy the parameter values for classifier
for index,layer in enumerate(model.classifier):
#only relevant for FC Linear parameters
if isinstance(layer,nn.Linear):
self.classifier[index].weight.data = layer.weight.data
self.classifier[index].bias.data = layer.bias.data
#function for forward propagation
def forward(self,x):
#neccessary for storing the max pool location for inverse pooling
for index,layer in enumerate(self.features):
if isinstance(layer,nn.MaxPool2d):
x, locs = layer(x)
else:
x = layer(x)
#converts a (N,C,H,W) input to (N,C*H*W)
x = x.view(x.size()[0],-1)
output = self.classifier(x)
output = torch.nn.functional.softmax(output,dim = 1)
return output
#this is the twin Deconvnet of the above VGG16 called Roy Net
class SMAI_Deconv(nn.Module) :
def __init__(self):
super(SMAI_Deconv, self).__init__()
self.features = nn.Sequential(
nn.ReLU(),
nn.ConvTranspose2d(32, 32, 3, padding = 1),
nn.MaxUnpool2d(2, stride = 2),
nn.ReLU(),
nn.ConvTranspose2d(32, 16, 3, padding = 1),
nn.MaxUnpool2d(2, stride = 2),
nn.ReLU(),
nn.ConvTranspose2d(16, 3, 3, padding = 1),
)
#this maps the indices of the layer of in conv net architecture with the deconvnet architecture
#{forward_ind : backward_ind}
self.conv_deconv_layer_index_mapping = {0:7 , 3:4 , 6:1}
#this maps the bias of the conv layer with the deconv layer.For some reason this does not
#perfectly allign with above transformation because the first backward layer is randomly set
#{forward_ind : backward_ind}
self.conv_deconv_layer_bias_index_mapping = {0:4,3:1}
#this maps the forward and backward relu layer
#{forward_ind : backward_ind}
self.relu_relu_layer_index_mapping = {1:6 , 4:3 , 7:0}
#this maps the forward maxpool layer index to the backward maxunpool netword
#{backward_ind : forward}
self.unmaxpool_maxpool_layer_index_mapping = {5:2, 2:5}
#this updates the parameters of the deconv model from the passed parameter which is the forward model
def update_weights(self, model):
#update the paramters
for index,layer in enumerate(model.features):
if isinstance(layer,nn.Conv2d):
self.features[self.conv_deconv_layer_index_mapping[index]].weight.data = layer.weight.data
if index in self.conv_deconv_layer_bias_index_mapping:
self.features[self.conv_deconv_layer_bias_index_mapping[index]].bias.data = layer.bias.data
#print("do")
#forward pass to obtain the inverse image
#pool locs is useful for maxunpooling
#layer is the index of the layer from which x is passed(I think)
#I am not too sure
#The input layer should be a RELU or CONV
#Not sure why Maxpooling layer not allowed
def forward(self, x , layer , pool_locs):
#print("Forward")
if layer in self.conv_deconv_layer_index_mapping:
start_index = self.conv_deconv_layer_index_mapping[layer]
elif layer in self.relu_relu_layer_index_mapping:
start_index = self.relu_relu_layer_index_mapping[layer]
else:
print("The given layer is not RELU or Conv,exiting")
#sys.exit(0)
for index in range(start_index,len(self.features)):
if isinstance(self.features[index] , nn.MaxUnpool2d):
print("Unpool")
x = self.features[index](x,pool_locs[self.unmaxpool_maxpool_layer_index_mapping[index]])
else:
if isinstance(self.features[index],nn.ConvTranspose2d):
print("Transpose")
if isinstance(self.features[index],nn.ReLU):
print("RElu")
x = self.features[index](x)
return x
# + id="kCYAsGBPBQCY" executionInfo={"status": "ok", "timestamp": 1638843150933, "user_tz": -330, "elapsed": 507, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
#Function to visualize a feature map in a grid
#feature map is the shape (C,W,H,N)
#this part is I dont understand that well
#utility function for visualization
def visualize_feature_map(feature_map):
(C,W,H,N) = feature_map.shape
cnt = int(ceil(sqrt(C)))
G = np.ones((cnt*H + cnt,cnt*W + cnt,N),feature_map.dtype)#the extra cnt is for black colored spacing
G *= np.min(feature_map)
n = 0
for row in range(cnt):
for col in range(cnt):
if n < C:
#additional cnt for spacing
G[row*H + row : (row + 1)*H + row, col*W + col : (col + 1)*W + col,:] = feature_map[n,:,:,:]
n += 1
#Normalize G
G = (G - G.min())/(G.max() - G.min())
return G
#function to visualize a layer represented by a grid
#again do not understand this part much
def visualize_layer(feature_layer_grid):
plt.clf() #clears figure
plt.subplot(121)
plt.imshow(feature_layer_grid[:,:,0],cmap = 'gray')
#transform a deconvolutional image to a normal one(Also normalize)
#input is the output of the deconvolated image
def transform_deconvolved_image(output) :
d_img = output.data.numpy()[0].transpose(1,2,0) #gets it to (H,W,C) format
d_img = (d_img - d_img.min())/(d_img.max() - d_img.min()) #normalize
return d_img.astype(np.uint8)
#function to store all the feature maps and max_pooling locations
def store_all_feature_maps(model):
#this is a function that is to be passed to another function
#It will be called everytime forward is called on model
def hook(module,input,output,key):
if isinstance(module,nn.MaxPool2d):
#remember the maxpool layer returns two values x,loc because return_indices is set to true
model.feature_maps[key] = output[0] #stores the feature map to the ordereddict
model.max_pooling_locations[key] = output[1] #stores the location
else:
model.feature_maps[key] = output
for index,layer in enumerate(model._modules.get('features')):
#register forward hook makes the hook to be called whenever forward is called
#partial allows hook to be called hook(module,input,output,key = index)
layer.register_forward_hook(partial(hook,key = index))
# + id="8uSSC8EeA-os" executionInfo={"status": "ok", "timestamp": 1638843164397, "user_tz": -330, "elapsed": 497, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
Layers = [0,1,2,3,5]
Activation_Index = [3,10]
def unormalize(im):
r = im[:,:,0] * 0.2023 + 0.4914
g = im[:,:,1] * 0.1994 + 0.4822
b = im[:,:,2] * 0.2010 + 0.4465
new_im = np.zeros(im.shape)
new_im[:,:,0] = r
new_im[:,:,1] = g
new_im[:,:,2] = b
return new_im
# + [markdown] id="xb5bS74TA7Sy"
# EPOCH 1 VISUALIZATION
#
# + colab={"base_uri": "https://localhost:8080/"} id="1ePc25aGAtJH" executionInfo={"status": "ok", "timestamp": 1638843502912, "user_tz": -330, "elapsed": 513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="fe0bc601-ae66-463f-c1de-9034c5e25a6a"
''' load the model'''
model = SMAI()
PATH = "./drive/MyDrive/Models/SMAI_MODEL_EPOCH_1.pth"
#print(checkpoint)
# original saved file with DataParallel
#state_dict = torch.load('myfile.pth.tar')
# create new OrderedDict that does not contain `module.`
# load params
model.load_state_dict(torch.load(PATH))
# + id="_pEmQK0kEknz"
# + colab={"base_uri": "https://localhost:8080/"} id="rytbrywlEgDe" executionInfo={"status": "ok", "timestamp": 1638843507329, "user_tz": -330, "elapsed": 615, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="672d8cae-f639-40cb-897e-7778e70dbdf2"
next(model.parameters()).is_cuda # returns a boolean
# + colab={"base_uri": "https://localhost:8080/"} id="mV_mPKPWDgrf" executionInfo={"status": "ok", "timestamp": 1638843507907, "user_tz": -330, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="b0e8a178-be12-4ac6-f4fd-fd3e1470fb25"
model.eval()
device = 'cuda' if torch.cuda.is_available() else 'cpu'
model.to(device)
next(model.parameters()).is_cuda # returns a boolean
# + colab={"base_uri": "https://localhost:8080/"} id="a0Rk0bmKFNnk" executionInfo={"status": "ok", "timestamp": 1638843544912, "user_tz": -330, "elapsed": 511, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="86d6e7af-8cca-4082-8f7d-f3a7b6217dfb"
deconv = SMAI_Deconv()
deconv.update_weights(model)
deconv = deconv.to(device)
deconv.eval()
# + colab={"base_uri": "https://localhost:8080/"} id="3q3fTM3aFrs8" executionInfo={"status": "ok", "timestamp": 1638843567754, "user_tz": -330, "elapsed": 633, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="00f56f77-c52e-481a-e754-c6b6f73aed3b"
next(deconv.parameters()).is_cuda # returns a boolean
# + id="A31NFMyuFzUl" executionInfo={"status": "ok", "timestamp": 1638843569263, "user_tz": -330, "elapsed": 5, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
store_all_feature_maps(model)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="dtGl-UJ1F8Fs" executionInfo={"status": "ok", "timestamp": 1638843733507, "user_tz": -330, "elapsed": 26262, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="7abe9ffc-f5af-442d-cbf8-8fcfe352ef51"
layer = 0
epoch = 1
activation_index = 3
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
# + [markdown] id="7Xj77cnlIJwL"
# EPOCH 1 LAYER 0 ACTIVATION INDEX = 3
# + colab={"base_uri": "https://localhost:8080/", "height": 484} id="v5-D-ahdG7WJ" executionInfo={"status": "ok", "timestamp": 1638843741231, "user_tz": -330, "elapsed": 528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="6e8456e9-abb1-4efd-f340-2f940fefa4d8"
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
# + id="to93gQBrIgnJ" executionInfo={"status": "ok", "timestamp": 1638843746981, "user_tz": -330, "elapsed": 721, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}}
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_0_index_3.jpg")
# + [markdown] id="0Oihs3UYJCUV"
# EPOCH : 1 LAYER : 0 ACTIVATION INDEX : 10
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="PJP6jEJTA5HE" executionInfo={"status": "ok", "timestamp": 1638843977229, "user_tz": -330, "elapsed": 27417, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="45005e0a-509a-44e1-ec15-c2ad2ffa7cf1"
layer = 0
epoch = 1
activation_index = 10
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_0_index_10.jpg")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="zqGeogm4O8Bo" executionInfo={"status": "ok", "timestamp": 1638844043237, "user_tz": -330, "elapsed": 26645, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="c81b84f0-49d0-4636-9a25-123b61bca4f1"
layer = 1
epoch = 1
activation_index = 3
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_1_index_3.jpg")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="rTk80i1fPNt9" executionInfo={"status": "ok", "timestamp": 1638844071518, "user_tz": -330, "elapsed": 26459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="c59ac77d-b571-457e-f0a8-ef77e0f18bb0"
layer = 1
epoch = 1
activation_index = 10
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_1_index_10.jpg")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="BqEuSdHoOZvB" executionInfo={"status": "ok", "timestamp": 1638844141495, "user_tz": -330, "elapsed": 23884, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="f1e50f38-a772-4f8b-f774-69e2b5ea4df5"
layer = 3
epoch = 1
activation_index = 3
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_3_index_3.jpg")
# + [markdown] id="BGV1x8hZOqFZ"
# EPOCH 1
# LAYER 2
# ACTIVATION INDEX 10
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="0ZQ4gohfOxQ0" executionInfo={"status": "ok", "timestamp": 1638844225130, "user_tz": -330, "elapsed": 23385, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="74be7e08-28c8-416a-e7f9-dd47eb5e39eb"
layer = 3
epoch = 1
activation_index = 10
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_3_index_10.jpg")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="PNiqeLSaJ3lx" executionInfo={"status": "ok", "timestamp": 1638844259168, "user_tz": -330, "elapsed": 24143, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="2adb9b2a-06c1-4712-d855-f3aeed84e19a"
layer = 4
epoch = 1
activation_index = 3
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_4_index_3.jpg")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="xDGdNY4NKxXz" executionInfo={"status": "ok", "timestamp": 1638844297541, "user_tz": -330, "elapsed": 22835, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06999756730187012596"}} outputId="0348e103-4d1d-4970-a937-2b2e64a3e0ae"
layer = 4
epoch = 1
activation_index = 10
print("Calling Feature Visualization part for layer : {0} , epoch : {1} and activation_index : {2}".format(layer,epoch,activation_index))
train_size = 45000
total_samples_to_be_used = train_size
samples_till_now = 0
batch_size = 32
highest_activate_image = np.zeros((32,32,3))
highest_activation = -np.inf
final_inp = None
deconv_image = np.zeros((32,32,3))
for i,(images,labels) in enumerate(trainloader):
if samples_till_now >= total_samples_to_be_used :
break
else:
samples_till_now += batch_size
im = images.to(device)
output = model(im)
feature_map = model.feature_maps[layer].cpu().data.numpy().transpose(1, 2, 3, 0) # (N, C, H, W) -> (C, H, W, N)
fmap = np.copy(feature_map)
for j in range(batch_size):
ft_mp = fmap[activation_index,:,:,j]
activation = np.linalg.norm(ft_mp,2)
if(activation >= highest_activation):
highest_activation = activation
src_image = np.transpose(images.cpu().numpy()[j],(1,2 ,0))
src_image = unormalize(src_image)
final_inp = im[j]
final_inp = final_inp[None] # (N,C,H,W)
#print("progress of computing the highest activation")
print("progress : {0} / {1} = {2}".format(samples_till_now,total_samples_to_be_used,samples_till_now/total_samples_to_be_used))
final_inp = final_inp.to(device)
out = model(final_inp)
new_map = model.feature_maps[layer].clone() # (1,C,H,W)
if activation_index == 0:
new_map[:,1:,:,:] = 0
else:
new_map[:,:activation_index,:,:] = 0
#print(new_map.shape)
if activation_index < new_map.shape[1] - 1:
new_map[:,activation_index + 1 :,:,:] = 0
#print("Deconvolution")
deconv_output = deconv(new_map , layer , model.max_pooling_locations)
r = deconv_output.cpu()[0][0][:][:]
decov_final = r.detach().numpy()
'''
file_path = "./drive/MyDrive/outputs/"
act_name1 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Deconv.npy"
act_name2 = file_path + "Activation_" + str(activation_index) + "_Layer_" + str(layer) + "_Epoch_" + str(epoch) + "_Src.npy"
print("Saving Image")
np.save(act_name1,decov_final)
np.save(act_name2,src_image)
'''
fig = plt.figure(figsize = (25,25))
fig.add_subplot(1,2,1)
plt.imshow(src_image)
plt.axis("off")
plt.title("The Original Image")
fig.add_subplot(1,2,2)
plt.imshow(decov_final)
plt.axis("off")
plt.title("Deconvoluted Image")
fig.savefig("./drive/MyDrive/outputs/SMAI_Epoch_1_Layer_4_index_10.jpg")
| experiments/Feature Evolution/src/small_epoch_1_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # AdaGrad算法
#
# 在之前介绍过的优化算法中,目标函数自变量的每一个元素在相同时间步都使用同一个学习率来自我迭代。举个例子,假设目标函数为$f$,自变量为一个二维向量$[x_1, x_2]^\top$,该向量中每一个元素在迭代时都使用相同的学习率。例如,在学习率为$\eta$的梯度下降中,元素$x_1$和$x_2$都使用相同的学习率$\eta$来自我迭代:
#
# $$
# x_1 \leftarrow x_1 - \eta \frac{\partial{f}}{\partial{x_1}}, \quad
# x_2 \leftarrow x_2 - \eta \frac{\partial{f}}{\partial{x_2}}.
# $$
#
# 在[“动量法”](./momentum.ipynb)一节里我们看到,当$x_1$和$x_2$的梯度值有较大差别时,需要选择足够小的学习率使得自变量在梯度值较大的维度上不发散。但这样会导致自变量在梯度值较小的维度上迭代过慢。动量法依赖指数加权移动平均使得自变量的更新方向更加一致,从而降低发散的可能。本节我们介绍AdaGrad算法,它根据自变量在每个维度的梯度值的大小来调整各个维度上的学习率,从而避免统一的学习率难以适应所有维度的问题 [1]。
#
#
# ## 算法
#
# AdaGrad算法会使用一个小批量随机梯度$\boldsymbol{g}_t$按元素平方的累加变量$\boldsymbol{s}_t$。在时间步0,AdaGrad将$\boldsymbol{s}_0$中每个元素初始化为0。在时间步$t$,首先将小批量随机梯度$\boldsymbol{g}_t$按元素平方后累加到变量$\boldsymbol{s}_t$:
#
# $$\boldsymbol{s}_t \leftarrow \boldsymbol{s}_{t-1} + \boldsymbol{g}_t \odot \boldsymbol{g}_t,$$
#
# 其中$\odot$是按元素相乘。接着,我们将目标函数自变量中每个元素的学习率通过按元素运算重新调整一下:
#
# $$\boldsymbol{x}_t \leftarrow \boldsymbol{x}_{t-1} - \frac{\eta}{\sqrt{\boldsymbol{s}_t + \epsilon}} \odot \boldsymbol{g}_t,$$
#
# 其中$\eta$是学习率,$\epsilon$是为了维持数值稳定性而添加的常数,如$10^{-6}$。这里开方、除法和乘法的运算都是按元素运算的。这些按元素运算使得目标函数自变量中每个元素都分别拥有自己的学习率。
#
# ## 特点
#
# 需要强调的是,小批量随机梯度按元素平方的累加变量$\boldsymbol{s}_t$出现在学习率的分母项中。因此,如果目标函数有关自变量中某个元素的偏导数一直都较大,那么该元素的学习率将下降较快;反之,如果目标函数有关自变量中某个元素的偏导数一直都较小,那么该元素的学习率将下降较慢。然而,由于$\boldsymbol{s}_t$一直在累加按元素平方的梯度,自变量中每个元素的学习率在迭代过程中一直在降低(或不变)。所以,当学习率在迭代早期降得较快且当前解依然不佳时,AdaGrad算法在迭代后期由于学习率过小,可能较难找到一个有用的解。
#
# 下面我们仍然以目标函数$f(\boldsymbol{x})=0.1x_1^2+2x_2^2$为例观察AdaGrad算法对自变量的迭代轨迹。我们实现AdaGrad算法并使用和上一节实验中相同的学习率0.4。可以看到,自变量的迭代轨迹较平滑。但由于$\boldsymbol{s}_t$的累加效果使学习率不断衰减,自变量在迭代后期的移动幅度较小。
# + attributes={"classes": [], "id": "", "n": "2"}
# %matplotlib inline
import d2lzh as d2l
import math
from mxnet import nd
def adagrad_2d(x1, x2, s1, s2):
g1, g2, eps = 0.2 * x1, 4 * x2, 1e-6 # 前两项为自变量梯度
s1 += g1 ** 2
s2 += g2 ** 2
x1 -= eta / math.sqrt(s1 + eps) * g1
x2 -= eta / math.sqrt(s2 + eps) * g2
return x1, x2, s1, s2
def f_2d(x1, x2):
return 0.1 * x1 ** 2 + 2 * x2 ** 2
eta = 0.4
d2l.show_trace_2d(f_2d, d2l.train_2d(adagrad_2d))
# -
# 下面将学习率增大到2。可以看到自变量更为迅速地逼近了最优解。
# + attributes={"classes": [], "id": "", "n": "3"}
eta = 2
d2l.show_trace_2d(f_2d, d2l.train_2d(adagrad_2d))
# -
# ## 从零开始实现
#
# 同动量法一样,AdaGrad算法需要对每个自变量维护同它一样形状的状态变量。我们根据AdaGrad算法中的公式实现该算法。
# + attributes={"classes": [], "id": "", "n": "4"}
features, labels = d2l.get_data_ch7()
def init_adagrad_states():
s_w = nd.zeros((features.shape[1], 1))
s_b = nd.zeros(1)
return (s_w, s_b)
def adagrad(params, states, hyperparams):
eps = 1e-6
for p, s in zip(params, states):
s[:] += p.grad.square()
p[:] -= hyperparams['lr'] * p.grad / (s + eps).sqrt()
# -
# 与[“小批量随机梯度下降”](minibatch-sgd.ipynb)一节中的实验相比,这里使用更大的学习率来训练模型。
# + attributes={"classes": [], "id": "", "n": "5"}
d2l.train_ch7(adagrad, init_adagrad_states(), {'lr': 0.1}, features, labels)
# -
# ## 简洁实现
#
# 通过名称为“adagrad”的`Trainer`实例,我们便可使用Gluon提供的AdaGrad算法来训练模型。
# + attributes={"classes": [], "id": "", "n": "6"}
d2l.train_gluon_ch7('adagrad', {'learning_rate': 0.1}, features, labels)
# -
# ## 小结
#
# * AdaGrad算法在迭代过程中不断调整学习率,并让目标函数自变量中每个元素都分别拥有自己的学习率。
# * 使用AdaGrad算法时,自变量中每个元素的学习率在迭代过程中一直在降低(或不变)。
#
# ## 练习
#
# * 在介绍AdaGrad算法的特点时,我们提到了它可能存在的问题。你能想到什么办法来解决这个问题?
# * 在实验中尝试使用其他的初始学习率,结果有什么变化?
#
#
#
#
#
# ## 参考文献
#
# [1] <NAME>., <NAME>., & <NAME>. (2011). Adaptive subgradient methods for online learning and stochastic optimization. Journal of Machine Learning Research, 12(Jul), 2121-2159.
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/2273)
#
# 
| 深度学习/d2l-zh-1.1/chapter_optimization/adagrad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# <NAME> - Odisha ROI Transformation from VoTT Raw format
# Takes filename of VoTT raw format file and generates ROI configuration
# v1.5 saral ocr version
import uuid
import json
def get_annotation(filename):
with open(filename) as f:
data = json.load(f)
f.close()
return data['regions']
def get_rois(regions,tagGroup):
rois = []
index = 0
roiIndex = 1
for region in regions:
if region['tags'][0].startswith(tagGroup):
rois.append({
# "annotationId": region['id'],
"annotationTag": region['tags'][0],
"extractionMethod": "NUMERIC_CLASSIFICATION",
"roiId": roiIndex,
"index": index,
"rect": {
"top": int(region['boundingBox']['top']),
"left": int(region['boundingBox']['left']),
"bottom": int(region['boundingBox']['top']) + int(region['boundingBox']['height']),
"right": int(region['boundingBox']['left']) + int(region['boundingBox']['width'])
}
})
index = index + 1
roiIndex = roiIndex + 1
return rois
def get_cells(regions,tagGroups,validationInfo):
cells_data = []
renderIndex = 1
cellIndex = 1
for tagGroup in tagGroups:
try:
validRegExp = validationInfo[str(tagGroup.rstrip('_'))]['regExp']
validName = validationInfo[str(tagGroup.rstrip('_'))]['name']
validErrorMsg = validationInfo[str(tagGroup.rstrip('_'))]['errorMessage']
validSource = validationInfo[str(tagGroup.rstrip('_'))]['source']
except KeyError as ke:
validRegExp = ""
validName = ""
validErrorMsg = ""
validSource = ""
cells_data.append({
"cellId": cellIndex,
"rois": get_rois(regions,tagGroup),
"render": {
"index": renderIndex
},
"format": {
"name": tagGroup.rstrip('_'),
"value": tagGroup.replace("_", "")
},
"validate": {
"name": validName,
"regExp": validRegExp,
"errorMessage": validErrorMsg,
"source": validSource
}
})
renderIndex = renderIndex +1
cellIndex = cellIndex + 1
return cells_data
def get_layout(cells,responseExcludeFields):
layout_data = []
layout_data.append({
"layout": {
"version": "1.0",
"name": "Odisha SAT 20 Questions Exam Sheet Form",
"type": "SAT_20_MARKSHEET",
"tolerance": {
"predictionMin": 0.95,
"roiMinWidth": 15,
"roiMinHeight": 15
},
"excludeFieldsInResponse": responseExcludeFields,
#"identifiers": [{"name":"teacherId","value":"2321121"}],
"cells": cells
}
})
return layout_data[0]
def pp_json(json_thing, sort=True, indents=4):
if type(json_thing) is str:
print(json.dumps(json.loads(json_thing), sort_keys=sort, indent=indents))
else:
print(json.dumps(json_thing, sort_keys=sort, indent=indents))
return None
regions=get_annotation("sat_odisha_vottraw.json")
#regions
# +
# Validation info may not be pre configured. These validations can be specific to school , exam.
# So these can be injected from backend during scanning time. format.name can be used as key ingest these validations.
validationInfo = {
'STUDENTID': { 'name': 'Between 10 to 15 Digits' , 'regExp': '^[1-9][0-9]{9,14}$' , 'errorMessage': 'Should be 10 to 15 Digits', 'source': 'BACKEND_SCHOOL' },
'QUESTION1': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION1 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION2': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION2 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION3': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION3 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION4': { 'name': 'Between 0 to 10 Marks' , 'regExp': '[0-10]' , 'errorMessage': 'QUESTION4 should range from 0 to 10 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION5': { 'name': 'Between 0 to 10 Marks' , 'regExp': '[0-10]' , 'errorMessage': 'QUESTION5 should range from 0 to 10 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION6': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION6 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION7': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION7 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION8': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION8 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION9': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION9 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION10': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION10 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION11': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION11 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION12': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION12 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION13': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION13 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION14': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION14 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION15': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION15 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION16': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION16 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION17': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION17 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION18': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION18 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION19': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION19 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'QUESTION20': { 'name': 'Between 0 to 5 Marks' , 'regExp': '[0-5]' , 'errorMessage': 'QUESTION20 should range from 0 to 5 Marks', 'source': 'BACKEND_EXAM' },
'MAX_MARKS': { 'name': 'Should be 110 Marks' , 'regExp': '^110$' , 'errorMessage': 'should be 110 Marks', 'source': 'BACKEND_EXAM' },
'MARKS_OBTAINED': { 'name': 'Between 0 to 110 Marks' , 'regExp': '\b([0-9]|[1-9][0-9]|110)\b' , 'errorMessage': 'Should be MAX of 110 Marks', 'source': 'BACKEND_EXAM' },
}
#validationInfo['STUDENTID']
#validationInfo['STUDENTID']['regExp']
# -
# Not all fields needed in response sent to backend for analytics/insights.
# So list of fields to be excluded in response can be defined for each layout.
responseExcludeFields= ['rois','validate','render']
tagGroups = ["STUDENTID", "QUESTION1", "QUESTION2","QUESTION3","QUESTION4","QUESTION5","QUESTION6","QUESTION7","QUESTION8","QUESTION9","QUESTION10","QUESTION11","QUESTION12","QUESTION13","QUESTION14","QUESTION15","QUESTION16","QUESTION17","QUESTION18","QUESTION19","QUESTION20","MAX_MARKS","MARKS_OBTAINED"]
#rois=get_rois(regions,tagGroups[0])
cells=get_cells(regions,tagGroups,validationInfo)
pp_json(get_layout(cells,responseExcludeFields))
| specs/v2/jupyter-notebook/transform_sat_odisha_vott_to_roi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Category Plots, aka Bar Charts
from beakerx.plots import *
# +
bars = CategoryBars(value= [[1, 2, 3], [1, 3, 5]])
plot = CategoryPlot()
plot.add(bars)
plot
# -
cplot = CategoryPlot(initWidth= 400, initHeight= 200)
bars = CategoryBars(value=[[1, 2, 3], [1, 3, 5]])
cplot.add(bars)
cplot = CategoryPlot(title= "Hello CategoryPlot!",
xLabel= "Categories",
yLabel= "Values")
cplot.add(CategoryBars(value=[[1, 2, 3], [1, 3, 5]]))
cplot = CategoryPlot(categoryNames= ["Helium", "Neon", "Argon"])
cplot.add(CategoryBars(value= [[1, 2, 3], [1, 3, 5]]))
CategoryPlot().add(CategoryBars(value= [[1, 2, 3], [1, 3, 5]],
seriesNames= ["Gas", "Liquid"]))
bars = CategoryBars(value= [[1, 2], [3, 4], [5, 6], [7, 8]],
seriesNames= ["Gas", None, "", "Liquid"])
CategoryPlot().add(bars)
plot = CategoryPlot(showLegend= True) # force legend display
bars = CategoryBars(value= [[1, 2, 3], [1, 3, 5]])
# since no display names were provided, default names "series0" etc will be used.
plot.add(bars)
plot = CategoryPlot(orientation= PlotOrientationType.HORIZONTAL)
plot.add(CategoryBars(value=[[1, 2, 3], [1, 3, 5]]))
import math
plot = CategoryPlot(categoryNames= ["Acid", "Neutral", "Base"],
categoryNamesLabelAngle= -1/4 * math.pi)
plot.add(CategoryBars(value= [[1, 2, 3], [4, 5, 6]]))
CategoryPlot(categoryMargin= 2).add(CategoryBars(value= [[1, 2, 3], [4, 5, 6]]))
# +
#bars = CategoryBars(value= (1..4) + 2)
#CategoryPlot().add(bars)
# -
bars = CategoryBars(value= [[1, 2], [3, 4], [5, 6]], color= Color.pink)
CategoryPlot().add(bars)
colors = [Color.red, Color.gray, Color.blue]
bars = CategoryBars(value= [[1, 2], [3, 4], [5, 6]], color= colors)
CategoryPlot().add(bars)
colors = [[Color.red, Color.gray],
[Color.gray, Color.gray],
[Color.blue, Color.pink]]
bars = CategoryBars(value= [[1, 2], [3, 4], [5, 6]], color= colors)
CategoryPlot().add(bars)
colors = [Color.pink, [Color.red, Color.gray, Color.blue]]
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 6]], color= colors)
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 6]], base= -2)
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 4]], base= [-1, -3])
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 6]],
base= [[0, 1, 1], [-4, -5, -6]])
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 6]],
width= [[0.3, 0.6, 1.7], 1.0])
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 6]],
fill= [[True, True, False], [True, False, True]],
drawOutline= [[True, False, True], [True, True, False]],
outlineColor= [Color.black, Color.red])
CategoryPlot().add(bars)
bars = CategoryBars(value= [[1, 2, 3], [4, 5, 8], [10, 9, 10]],
base= [0, [1, 2, 3], [4, 5, 8]],
centerSeries= True,
itemLabel= [[1, 2, 3], [4, 5, 8], [10, 9, 10]]
)
CategoryPlot().add(bars)
ss = [StrokeType.DASH, StrokeType.LONGDASH]
cs = [Color.black, Color.red]
CategoryPlot().add(CategoryStems(value= [[1, 2, 4], [4, 5, 8]], color= cs, style= ss))
CategoryPlot().add(CategoryPoints(value= [[1, 2, 4], [4, 5, 8]]))
ss = [StrokeType.DASH, StrokeType.DOT]
CategoryPlot().add(CategoryLines(value= [[1, 2, 4], [4, 5, 8]], style= ss,
seriesNames=["Lanthanide", "Actinide"]))
s1 = [1, 2, 4]
s2 = [4, 5, 8]
lines = CategoryLines(value= [s1, s2], centerSeries= True)
points = CategoryPoints(value= [s1, s2], centerSeries= True)
stems = CategoryStems(value=[s1], base= [s2], style= StrokeType.DOT, color= Color.gray)
#plot = CategoryPlot().add(lines).add(points).add(stems)
plot = CategoryPlot()
plot.add(lines)
plot.add(points)
plot.add(stems)
plot = CategoryPlot(initWidth= 500, initHeight= 400,
title= "Bar Chart Demo",
xLabel= "Alkali", yLabel= "Temperature ° Celcius",
categoryNames=["Lithium", "Sodium", "Potassium", "Rubidium"])
s1 = [[10, 15, 13, 7], [22, 18, 28, 17]]
high = [[12.4, 19.5, 15.1, 8.2], [24.3, 23.3, 30.1, 18.2]]
low = [[7.6, 10.5, 10.9, 5.8], [19.7, 12.7, 25.9, 15.8]]
color = [Color(247, 150, 70), Color.orange, Color(155, 187, 89)]
plot.add(CategoryBars(value= s1, color= color, seriesNames= ["Solid", "Liquid"]))
plot.add(CategoryStems(value= high, base= low, color= color[2]))
plot.add(CategoryPoints(value= high, outlineColor= color[2], size=12))
plot.add(CategoryPoints(value= low, outlineColor= color[2], size=12))
p = CategoryPlot(title= "Multiple Y Axes Demo",
yLabel= "Price",
categoryNames= ["Q1", "Q2", "Q3", "Q4"])
p.add(YAxis(label= "Volume", upperMargin= 1))
p.add(CategoryBars(value= [[1500, 2200, 2500, 4000]], width= 0.6,
color= Color.PINK, yAxis= "Volume", showItemLabel= True,
labelPosition= LabelPositionType.VALUE_INSIDE))
p.add(CategoryLines(value= [[5, 2, 3.5, 4]], color= Color.GRAY,
showItemLabel=True))
p.add(CategoryPoints(value=[[5, 2, 3.5, 4]], color=Color.GRAY))
CategoryPlot().add(CategoryStems(value=[[-3, 2, 4], [4, 5, 8]],
width= 10, showItemLabel= True))
bars = CategoryBars(value= [[-5, 2, 3], [1, 3, 5]],
showItemLabel= True)
CategoryPlot().add(bars)
bars = CategoryBars(value= [[-5, 2, 3], [1, 3, 5]],
labelPosition= LabelPositionType.BASE_OUTSIDE,
showItemLabel= True)
CategoryPlot().add(bars)
plot = CategoryPlot(title= "Move mouse cursor over bars")
bars = CategoryBars(value= [[-5, 2, 3], [1, 3, 5]], useToolTip= False)
plot.add(bars)
# +
#table = [[close=11.59, high=13.15, low=11.92, open=11.92],
# [close=12.76, high=15.44, low=11.88, open=12.42],
# [close=18.19, high=20.96, low=17.93, open=18.56]]
#v = table.collect { it.values().toList() }
#p = CategoryPlot(categoryNames= table[0].keySet().toList())
#p.add(new CategoryBars(value= v, seriesNames= ["A", "B", "C"]))
# +
cs = [Color.orange]
cp = CategoryPlot()
cp.add(CategoryArea(value= [[1, 3, 2]], base= [[0.5, 1, 0]]))
cp.add(CategoryArea(value= [[2, 1, 0.5]], color= cs))
| doc/CategoryPlot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('nbdev')
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/gtbook/gtsam-examples/blob/main/Pose2SLAMExample.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Pose2 SLAM
#
# A simple way to do Simultaneous Localization and Mapping is to just fuse **relative pose measurements** between successive robot poses. This landmark-less SLAM variant is often called "Pose SLAM"
# %pip -q install gtbook # also installs latest gtsam pre-release
# +
import math
import gtsam
import gtsam.utils.plot as gtsam_plot
import matplotlib.pyplot as plt
# -
PRIOR_NOISE = gtsam.noiseModel.Diagonal.Sigmas(gtsam.Point3(0.3, 0.3, 0.1))
ODOMETRY_NOISE = gtsam.noiseModel.Diagonal.Sigmas(
gtsam.Point3(0.2, 0.2, 0.1))
#
# 1. Create a factor graph container and add factors to it
#
graph = gtsam.NonlinearFactorGraph()
# 2a. Add a prior on the first pose, setting it to the origin
#
# A prior factor consists of a mean and a noise ODOMETRY_NOISE (covariance matrix)
graph.add(gtsam.PriorFactorPose2(1, gtsam.Pose2(0, 0, 0), PRIOR_NOISE))
# 2b. Add odometry factors
# Create odometry (Between) factors between consecutive poses
#
graph.add(gtsam.BetweenFactorPose2(1, 2, gtsam.Pose2(2, 0, 0), ODOMETRY_NOISE))
graph.add(gtsam.BetweenFactorPose2(2, 3, gtsam.Pose2(2, 0, math.pi / 2), ODOMETRY_NOISE))
graph.add(gtsam.BetweenFactorPose2(3, 4, gtsam.Pose2(2, 0, math.pi / 2), ODOMETRY_NOISE))
graph.add(gtsam.BetweenFactorPose2(4, 5, gtsam.Pose2(2, 0, math.pi / 2),ODOMETRY_NOISE))
# 2c. Add the loop closure constraint
# This factor encodes the fact that we have returned to the same pose. In real
# systems, these constraints may be identified in many ways, such as appearance-based
# techniques with camera images. We will use another Between Factor to enforce this constraint:
#
graph.add( gtsam.BetweenFactorPose2(5, 2, gtsam.Pose2(2, 0, math.pi / 2), ODOMETRY_NOISE))
print("\nFactor Graph:\n{}".format(graph))
# 3. Create the data structure to hold the initial_estimate estimate to the
# solution. For illustrative purposes, these have been deliberately set to incorrect values
#
initial_estimate = gtsam.Values()
initial_estimate.insert(1, gtsam.Pose2(0.5, 0.0, 0.2))
initial_estimate.insert(2, gtsam.Pose2(2.3, 0.1, -0.2))
initial_estimate.insert(3, gtsam.Pose2(4.1, 0.1, math.pi / 2))
initial_estimate.insert(4, gtsam.Pose2(4.0, 2.0, math.pi))
initial_estimate.insert(5, gtsam.Pose2(2.1, 2.1, -math.pi / 2))
print("\nInitial Estimate:\n{}".format(initial_estimate))
# 4. Optimize the initial values using a Gauss-Newton nonlinear optimizer
# The optimizer accepts an optional set of configuration parameters,
# controlling things like convergence criteria, the type of linear
# system solver to use, and the amount of information displayed during
# optimization. We will set a few parameters as a demonstration.
#
parameters = gtsam.GaussNewtonParams()
# Stop iterating once the change in error between steps is less than this value
#
parameters.setRelativeErrorTol(1e-5)
# Do not perform more than N iteration steps
#
parameters.setMaxIterations(100)
# Create the optimizer ...
#
optimizer = gtsam.GaussNewtonOptimizer(graph, initial_estimate, parameters)
# ... and optimize
#
result = optimizer.optimize()
print("Final Result:\n{}".format(result))
# 5. Calculate and print marginal covariances for all variables
#
marginals = gtsam.Marginals(graph, result)
for i in range(1, 6):
print("X{} covariance:\n{}\n".format(i, marginals.marginalCovariance(i)))
# +
for i in range(1, 6):
gtsam_plot.plot_pose2(0, result.atPose2(i), 0.5,
marginals.marginalCovariance(i))
plt.axis('equal')
plt.show()
| Pose2SLAMExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img style="float: left; margin: 30px 15px 15px 15px;" src="https://pngimage.net/wp-content/uploads/2018/06/logo-iteso-png-5.png" width="300" height="500" />
#
#
# ### <font color='navy'> Simulación de procesos financieros.
#
# **Nombres:** <NAME> y <NAME>
#
# **Fecha:** 15 de Marzo del 2020.
#
# **Expediente** : if721470 if721215
# **Profesor:** <NAME>.
#
# **Link Github**:
#
# # Tarea 7: Clase 13.MetodosDeReduccionDeVarianza
# ## Enunciado de tarea
#
# > Aproxime el valor de la siguiente integral usando el método monte carlo crudo y método de reducción de varianza de muestreo estratíficado
#
# $$I=\int_{0}^{1}x^2\text{d}x=\left.\frac{x^3}{3}\right|_{x=0}^{x=1}=\frac{1}{3}\approx 0.33333$$
#
# Pasos
# 1. Cree una función que realice el método de muestreo estratíficado, recibiendo como único parámetro de entrada la cantidad de estratos y retornando las variables estratíficadas correspondientes.
# 2. Reporte los resultados de la aproximación de la integral usando montecarlo crudo y muestreo estratíficado, en un Dataframe con la información mostrada en la siguiente imagen:
# 
# ## SOLICION CRISTINA
# Ejercicio 1: Cree una función que realice el método de muestreo estratíficado, recibiendo como único parámetro de entrada la cantidad de estratos y retornando las variables estratíficadas correspondientes.
import numpy as np
from functools import reduce
import time
import matplotlib.pyplot as plt
import scipy.stats as st # Librería estadística
import pandas as pd
#Defino una función que guarde el método estratificado
def estratificado(N:'Cantidad de Estratos'):
B = N
U = np.random.rand(N) #Distribucion entre 0 y 1
i = np.arange(0, B)
m_estratificado = (U + i) / B
return m_estratificado
#Crear un array con los N
N= np.logspace(1,7,7,dtype=int)
#llamo la función estratificada para generar U
u = list(map(lambda y : estratificado(y),N.tolist()))
#Uso u para evaluarlo en la función a integrar
I_m = list(map(lambda x:x**2,u))
# Ejercicio 2: Reporte los resultados de la aproximación de la integral usando montecarlo crudo y muestreo estratíficado, en un Dataframe con la información mostrada en la imagen anterior
#Saco el promedio de cada array para conocer el valor aproximado de la integral
sol = list(map(lambda x: sum(x)/len(x),I_m))
sol
# Integración montecarlo
def int_montecarlo(f:'Función a integrar',
a:'Límite inferior de la integral',
b:'Límite superior de la integral',
U:'Muestra de números U~[a,b]'):
return (b-a)/len(U)*np.sum(f(U))
# +
I = 1/3
a = 0; b = 1
# Cantidad de términos, en escala logarítmica
N = np.logspace(1,7,7,dtype=int)
# Crear data frame
df = pd.DataFrame(index=N,columns=['Montecarlo Crudo', 'Error_relativo%','Muestreo Estratificado',
'Error_relativo2%'], dtype='float')
df.index.name = "Cantidad_terminos"
# Números aleatorios dependiente de la cantidad de términos N
ui = list(map(lambda N:np.random.uniform(a,b,N),N))
# Calculamos la aproximación por montecarlo dependiendo de la cantidad de
# términos que hayamos creado con ui
I_m2 = list(map(lambda Y:int_montecarlo(lambda x:x**2,a,b,Y),ui))
# Mostramos los resultados en la tabla previamente creada
df.loc[N,"Montecarlo Crudo"] = I_m2
df.loc[N,"Error_relativo%"] = np.abs(df.loc[N,"Montecarlo Crudo"]-I)*100/I
df.loc[N,"Muestreo Estratificado"] = sol
df.loc[N,"Error_relativo2%"] = np.abs(df.loc[N,"Muestreo Estratificado"]-I)*100/I
df
# -
# ## SOLUCION DAYANA
# Ejercicio 1: Cree una función que realice el método de muestreo estratíficado, recibiendo como único parámetro de entrada la cantidad de estratos y retornando las variables estratíficadas correspondientes.
# Código de solución estudiante 2
.
.
.
.
# Ejercicio 2: Reporte los resultados de la aproximación de la integral usando montecarlo crudo y muestreo estratíficado, en un Dataframe con la información mostrada en la imagen anterior
| TAREA7_NavarroValenciaDayana_VazquezVargasCristina.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
# # Analysing Audioscrobbler Data
#
# In this notebook we want to create two very simple statistics on artists from data provided by Audioscrobbler. We are working with three related data sets:
#
# 1. A list of users containing their number of plays per artist
# 2. A list which maps a generic artist-id to its real (band) name
# 3. A list which maps bad artist ids to good ones (fixing typing errors)
#
# Then we will ask two simple questions:
#
# 1. Which artists have the most listeners (in terms of number of unique users)
# 2. Which artists are played most often (in terms of total play counts)
# # 1 Load Data
#
# First of all we have to load the data from S3.
# ## 1.1 Read User-Artist Data
#
# First we read in the most important data set, containing the information how often a user played songs of a specific artist. This information is stored in the file at `s3://dimajix-training/data/audioscrobbler/user_artist_data/`. The file has the following characteristics:
#
# * Format: CSV (kind of)
# * Separator: Space (” “)
# * Header: No
# * Fields: user_id (INT), artist_id(INT), play_count(INT)
#
# So we need to read in this file and store it in a local variable user_artist_data. Since we do not have any header contained in the file itself, we need to specify the schema explicitly.
# +
from pyspark.sql.types import *
schema = StructType([
StructField("user_id", IntegerType()),
StructField("artist_id", IntegerType()),
StructField("play_count", IntegerType())
])
user_artist_data = spark.read \
.option("sep", " ") \
.option("header", False) \
.schema(schema) \
.csv("s3://dimajix-training/data/audioscrobbler/user_artist_data/")
# -
# ### Peek inside
# Let us have a look at the first 5 records
user_artist_data.limit(5).toPandas()
# ## 1.2 Read in Artist aliases
#
# Now we read in a file containing mapping of bad artist IDs to good IDs. This fixes typos in the artists names and thereby enables us to merge information with different artist IDs belonging to the same band. This information is stored in the file at `s3://dimajix-training/data/audioscrobbler/artist_alias/`. The file has the following characteristics:
#
# * Format: CSV (kind of)
# * Separator: Tab (“\t”)
# * Header: No
# * Fields: bad_id (INT), good_id(INT)
#
# So we need to read in this file and store it in a local variable `artist_alias`. Since we do not have any header contained in the file itself, we need to specify the schema explicitly.
# +
schema = # YOUR CODR HERE
artist_alias = # YOUR CODR HERE
# -
# ### Peek inside
# +
# YOUR CODR HERE
# -
# ## 1.3 Read in Artist names
#
# The third file contains the artists name for his/her id. We also use this file in order to be able to display results with the artists names instead of their IDs. This information is stored in the file at `s3://dimajix-training/data/audioscrobbler/artist_data/`. The file has the following characteristics:
#
# * Format: CSV (kind of)
# * Separator: Tab (“\t”)
# * Header: No
# * Fields: artist_id (INT), artist_name(STRING)
#
# So we need to read in this file and store it in a local variable `artist_data`. Since we do not have any header contained in the file itself, we need to specify the schema explicitly.
# +
schema = # YOUR CODR HERE
artist_data = # YOUR CODR HERE
# -
# ### Peek inside
# +
# YOUR CODR HERE
# -
# # 2 Clean Data
#
# Before continuing with the analysis, we first create a cleaned version of the `user_artist_data` DataFrame with the `artist_alias` mapping applied. This means that we need to lookup each artist-id in the original data set in the alias data set and see if we find have a matching `bad_id` entry with a replacement `good_id`. The result should be stored in a variable `cleaned_user_artist_data`. This DataFrame should contain the columns Finally select only the columns `user_id`, `artist_id` (the corrected one) and `play_count`.
#
# Hints:
#
# 1. Join the user artist data DataFrame with the artist alias DataFrame containing fixes for some artists. Which join type is appropriate?
# 2. Replace the artist-id in the user artists data with the `good_id` from the artist alias DataFrame - if a match is found on `bad_id`
# 3. Finally select only the columns `user_id`, `artist_id` (the corrected one) and `play_count`
# +
from pyspark.sql.functions import *
cleaned_user_artist_data = # YOUR CODR HERE
cleaned_user_artist_data.limit(5).toPandas()
# -
# # 3 Question 1: Artist with most unique listeners
#
# Who are the artist with the most unique listeners? For this question, it is irrelevant how often an individul user has played songs of a specific artist. It is only important how many different users have played each artists work. Of course we do not want to see the artist-id, but their real names as provided in the DataFrame `artist_data`!
#
# Hints:
# 1. Group cleaned data by `artist_id`
# 2. Perform aggregation by counting unique user ids
# 3. Join `artist_data`
# 4. Lookup the artists name
# 5. Sort result by descending user counts
# +
result = # YOUR CODR HERE
result.limit(10).toPandas()
# -
# # 4 Question 2: Artist with most Plays
#
# Now we also take into account how often each user played the work of individual artists. That is, we also include the `play_coun` column into our calculations. So which artists have the most plays in total? Of course we do not want to see the artist-id, but their real names as provided in the DataFrame `artist_data`!
#
# Hints:
# 1. Group cleaned data by `artist_id`
# 2. Perform aggregation by summing up play count
# 3. Join `artist_data`
# 4. Lookup the artists name
# 5. Sort result by descending play counts
#
# +
result = # YOUR CODR HERE
result.limit(5).toPandas()
| spark-training/spark-python/jupyter-audioscrobbler/Audioscrobbler - Skeleton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
### IMPORTANT
### The most part of this code was created by Coursera on Applied Machine Learning with Python course from University of Michigan
### This repository is only to make safe some code implemented by me
### be happy :)
# %matplotlib notebook
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
path = "dataset/fruits/"
fruits = pd.read_table(path+"fruit_dataset.txt")
fruits.head()
#Using zip with .unique() to agregate de label value with the name of fruit
lookup_fruit_name = dict(zip(fruits.fruit_label.unique(), fruits.fruit_name.unique()))
lookup_fruit_name
#Another way to see missing data or something like that is using pd.info()
fruits.info()
# +
# plotting a scatter matrix
from matplotlib import cm
from sklearn.model_selection import train_test_split
from pandas.plotting import scatter_matrix
X = fruits[['height', 'width', 'mass', 'color_score']]
y = fruits['fruit_label']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
cmap = cm.get_cmap('gnuplot')
scatter = scatter_matrix(X_train, c= y_train, marker = 'o', s=40, hist_kwds={'bins':15}, figsize=(8,8), cmap=cmap)
# +
# plotting a 3D scatter plot
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection = '3d')
ax.scatter(X_train['width'], X_train['height'], X_train['color_score'], c = y_train, marker = 'o', s=100)
ax.set_xlabel('width')
ax.set_ylabel('height')
ax.set_zlabel('color_score')
plt.show()
# +
# For this example, we use the mass, width, and height features of each fruit instance
X = fruits[['mass', 'width', 'height']]
y = fruits['fruit_label']
# default is 75% / 25% train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors = 5)
# -
knn.fit(X_train, y_train)
knn.score(X_test, y_test)
# +
k_range = range(1,20)
scores = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors = k)
knn.fit(X_train, y_train)
scores.append(knn.score(X_test, y_test))
plt.figure()
plt.xlabel('k')
plt.ylabel('accuracy')
plt.scatter(k_range, scores)
plt.xticks([0,5,10,15,20]);
# +
t = [0.8, 0.7, 0.6, 0.5, 0.4, 0.3, 0.2]
knn = KNeighborsClassifier(n_neighbors = 5)
plt.figure()
for s in t:
scores = []
for i in range(1,1000):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 1-s)
knn.fit(X_train, y_train)
scores.append(knn.score(X_test, y_test))
plt.plot(s, np.mean(scores), 'bo')
plt.xlabel('Training set proportion (%)')
plt.ylabel('accuracy');
# -
| fruits/fruits_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import csv
from settings import *
from clustering import *
from objects import *
from sklearn.cluster import DBSCAN
hr = HR(data)
# -
def draw(df_prime, var_x, var_y, eps, min_samples, metric, title, colors):
pp.cla()
pp.clf()
figure, axes = pp.subplots()
for i, color in zip(set(df_prime["cluster"]),colors):
points = df_prime[df_prime["cluster"] == i][[var_x, var_y]]
#if points.shape[0] <= 50:
# pass
axes.scatter(points[var_x], points[var_y], label=str(i), color=color)
axes.set_xlabel(labels_pretty_print[var_x])
axes.set_ylabel(labels_pretty_print[var_y])
axes.legend()
pp.title(labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] +
"\neps := " + str(eps) + " k := " + str(min_samples))
pp.savefig(title + ".svg")
pp.show()
# +
with open("pickles/dbscan/average_montly_hours,number_project-euclidean-200dbscan.eps.graphs", "rb") as log:
dbscan_graph = pickle.load(log)
var_x = "average_montly_hours"
var_y = "number_project"
i = 10000
j = 12000
distances = dbscan_graph[i:j]
sorted_distances = sorted(distances)
sorted_set_distances = sorted(set(distances))
# -
filtered_sorted_set_distances = list(filter(lambda x:x < .2, sorted_set_distances))
filtered_sorted_set_distances
# +
eps = 0.10012
min_points = 200
distances = dbscan_graph
#distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
# Preprocess
df_prime = hr.normal[[var_x, var_y]]
#df_prime = df_prime.drop_duplicates()
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df_prime)
df_prime = df_prime.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
title = labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] + "\neps := " + str(eps) + " k := " + str(min_points)
colors_keys = ["red", "yellow", "black", "orange", "blue", "silver", "green"]
colors = [large_palette_full[key] for key in colors_keys]
#draw(df_prime, var_x, var_y, eps, min_points, "euclidean", title, colors)
# Cross variables
colors_keys = ["yellow", "red", "orange", "green", "blue"]
colors = [large_palette_full[key] for key in colors_keys]
pp.cla()
pp.clf()
df_prime = df_prime.assign(left = hr.normal["left"])
df_prime = df_prime.assign(Years = hr.data["time_spend_company"])
df_prime = df_prime.assign(promoted = hr.data["promotion_last_5years"])
phi = pd.crosstab(df_prime["Years"], df_prime["cluster"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True, color=colors)
pp.title("Cluster heterogeneity per years spent in company")
#pp.show()
pp.savefig("average_montly_hours-number_project-time.svg")
phi
phi_ptg
# +
clusters_prime = [df_prime[df_prime["cluster"] == i] for i in range(-1,4)]
means_prime = [df["time"].mean() for df in clusters_prime]
std_prime = [df["time"].std() for df in clusters_prime]
means = {i: mean for mean,i in zip(means_prime, range(-1,4))}
stds = {i: std for std,i in zip(std_prime, range(-1,4))}
print(str(means))
print(str(stds))
# +
# Cross variables
df_prime = df_prime.assign(time = hr.discrete["time_spend_company"])
df_prime = df_prime.assign(left = hr.discrete["left"])
df_prime = df_prime.assign(wage = hr.discrete["salary"])
df_prime = df_prime.assign(promoted = hr.discrete["promotion_last_5years"])
colors_keys = ["yellow", "red", "orange", "green", "blue"]
colors = [large_palette_full[key] for key in colors_keys]
phi = pd.crosstab(df_prime["time"], df_prime["cluster"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True, color=colors)
#pp.show()
pp.savefig(var_x + "-" + var_y + "-time.svg")
phi
phi_ptg
# +
with open("pickles/dbscan/average_montly_hours,number_project-cosine-200dbscan.eps.graphs", "rb") as log:
dbscan_graph = pickle.load(log)
var_x = "average_montly_hours"
var_y = "number_project"
eps = 0.1
min_points = 4
distances = dbscan_graph
distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
# Preprocess
df_prime = hr.normal[[var_x, var_y]]
df_prime = df_prime.drop_duplicates()
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df_prime)
df_prime = df_prime.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
title = labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] + "\neps := " + str(eps) + " k := " + str(min_points)
draw(df_prime, "satisfaction_level", "average_montly_hours", eps, min_points, "euclidean", title)
# Cross variables
df_prime = df_prime.assign(left = hr.normal["left"])
phi = pd.crosstab(df_prime["cluster"], df_prime["left"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True)
pp.savefig(title + "-left" + ".svg")
phi
phi_ptg
# +
with open("pickles/dbscan/satisfaction_level,average_montly_hours-chebyshev-4dbscan.eps.graphs", "rb") as log:
dbscan_graph = pickle.load(log)
var_x = "satisfaction_level"
var_y = "average_montly_hours"
eps = 0.07
min_points = 4
distances = dbscan_graph[:]
distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
# Preprocess
df_prime = hr.normal[[var_x, var_y]]
df_prime = df_prime.drop_duplicates()
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df_prime)
df_prime = df_prime.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
title = labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] + "\neps := " + str(eps) + " k := " + str(min_points)
color_keys = ["orange", "silver", "blue", "green"]
colors = [large_palette_full[color] for color in color_keys]
draw(df_prime, "satisfaction_level", "average_montly_hours", eps, min_points, "euclidean", title, colors=colors)
# Cross variables
df_prime = df_prime.assign(left = hr.normal["left"])
phi = pd.crosstab(df_prime["cluster"], df_prime["left"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True)
pp.savefig(title + "-left" + ".svg")
phi
phi_ptg
#pp.show()
# +
p = pd.crosstab(df_prime["cluster"], df_prime["left"])
p = p.div(p.sum(1).astype(float), axis=0)
sizes = [df_prime[df_prime["cluster"] == i].shape[0] for i in set(df_prime["cluster"])]
df_prime_ = df_prime.assign(s=pd.Series(sizes))
#df_prime_[df_prime_["s"] >= 50][["cluster", "s"]]
df_prime_ = df_prime_[df_prime_["cluster"].isin([0, 1, 2, 70])]
p = pd.crosstab(df_prime_["cluster"], df_prime_["left"])
p = p.div(p.sum(1).astype(float), axis=0)
p
# +
eps = 0.05
distances = dbscan_graph[:]
distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
df_prime = hr.normal[[var_x, var_y]]
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df_prime)
df_prime = df_prime.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
title = labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] + "\neps := " + str(eps)
+ " k := " + str(min_points)
draw(df_prime, "satisfaction_level", "average_montly_hours", eps, min_points, "euclidean", title)
df_prime = df_prime.assign(left = hr.normal["left"])
phi = pd.crosstab(df_prime["cluster"], df_prime["left"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True)
pp.savefig(title + "left" + ".svg")
phi
phi_ptg
pp.show()
# post processing
sizes = list(map(lambda x: x.shape[0], [df_prime[df_prime["cluster"] == i] for i in set(clusters)]))
sizes = list(zip(sizes, set(clusters)))
sorted_sizes = sorted(set(sizes), reverse=True)
sorted_filtered_sizes = list(filter(lambda x: x[0] >= 50, sorted_sizes))
filtered_df_prime = df_prime[df_prime["cluster"].isin(list(map(lambda x: x[1], sorted_filtered_sizes)))]
draw(filtered_df_prime, "satisfaction_level", "average_montly_hours", eps, min_points,
"euclidean", "filtered." + title)
filtered_df_prime = filtered_df_prime.assign(left = hr.normal["left"])
# +
eps = 0.05
distances = dbscan_graph[:]
distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
df_prime = hr.normal[[var_x, var_y]]
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df_prime)
df_prime = df_prime.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
title = labels_pretty_print[var_x] + " " + labels_pretty_print[var_y] + "\neps := " + str(eps)
+ " k := " + str(min_points)
draw(df_prime, "satisfaction_level", "average_montly_hours", eps, min_points, "euclidean", title)
df_prime = df_prime.assign(left = hr.normal["left"])
phi = pd.crosstab(df_prime["cluster"], df_prime["left"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True)
pp.savefig(title + "left" + ".svg")
phi
phi_ptg
pp.show()
# post processing
sizes = list(map(lambda x: x.shape[0], [df_prime[df_prime["cluster"] == i] for i in set(clusters)]))
sizes = list(zip(sizes, set(clusters)))
sorted_sizes = sorted(set(sizes), reverse=True)
sorted_filtered_sizes = list(filter(lambda x: x[0] >= 50, sorted_sizes))
filtered_df_prime = df_prime[df_prime["cluster"].isin(list(map(lambda x: x[1], sorted_filtered_sizes)))]
draw(filtered_df_prime, "satisfaction_level", "average_montly_hours", eps, min_points,
"euclidean", "filtered." + title)
filtered_df_prime = filtered_df_prime.assign(left = hr.normal["left"])
phi = pd.crosstab(filtered_df_prime["cluster"], filtered_df_prime["left"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="barh", stacked=True)
pp.savefig("filtered_" + title + "left" + ".svg")
phi
phi_ptg
pp.show()
phi = pd.crosstab(filtered_df_prime["left"], filtered_df_prime["cluster"])
phi_ptg = phi.div(phi.sum(1).astype(float), axis=0)
phi_ptg.plot(kind="bar", stacked=True)
pp.savefig("filtered_" + title + "left" + ".svg")
phi
phi_ptg
pp.show()
list(map(lambda x: (x[0]/13958, x[1]), sorted_filtered_sizes))
phi_ptg
# +
with open("satisfaction_level,average_montly_hours-euclidean-dbscan.eps.graphs", "rb") as log:
dbscan_graph = pickle.load(log)
var_x = "satisfaction_level"
var_y = "average_montly_hours"
metric = "euclidean"
k = 4
eps = 0.05
# Dataset without obvious clusters
df_prime = df[~df["cluster"].isin([0,1,2,3,23,-1])]
entry = var_x + "," + var_y
df_prime = df_prime[[var_x, var_y]]
m = df_prime.shape[0]
entries = [("satisfaction_level", "average_montly_hours")]
for (var_x, var_y) in entries:
entry = var_x + "," + var_y
for metric in ["euclidean"]:
distance_matrix = cdist(df_prime, df_prime, metric=metric)
# sort rows, then sort columns
distance_matrix.sort(axis=1)
distance_matrix.sort(axis=0)
for k in [4]:
figure, axes = pp.subplots()
distances = distance_matrix[:, k]
axes.plot(range(df_prime.shape[0]), distances)
axes.set_title("Distance for " + str(k) + "th neighbor [" + metric + "]" + " " + entry)
axes.set_xlabel("Points at distance eps")
axes.set_ylabel("Distance")
axes.grid()
pp.savefig(entry + ":" + str(k) + " [" + metric + "]" + " " + entry + ".svg")
pp.clf()
pp.cla()
pp.close(figure)
distances = dbscan_graph[:]
distances = np.clip(distances,0, 1)
sorted_distances = sorted(distances)
set_sorted_distances = sorted(set(distances))
df = hr.normal[[var_x, var_y]]
dbscan = DBSCAN(eps=eps, min_samples=min_points, metric="euclidean")
clusters = dbscan.fit_predict(df)
df = df.assign(cluster = clusters)
hist, bins = np.histogram(dbscan.labels_, bins=range(-1, len(set(dbscan.labels_)) + 1))
# + active=""
#
# -
| notebooks/DBscan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PDF Extraction
#
# 1. First run the rename function below
# 2. Then extract the yml files.
# +
# %load_ext autoreload
# %autoreload 2
import sys
import os
import re
import pandas as pd
import shutil
from pathlib import Path
#from dateutil.parser import parse
#from fuzzyparsers import parse_date
#import timelib
sys.path.append('../..')
import data.dataframe_preparation as preparation
purge_existing_folder = False
extract_text = True
reports_input_dir = "/Volumes/Mac_Backup/Data/fin-disclosures-nlp/stoxx600/unprocessed"
master_output_file = os.path.join(reports_input_dir, "Firm_AnnualReport_TrainingV4.csv")
random_seed = 1
# -
# ### Script for renaming files to AR_YYYY.pdf format
# Handles cases such as 2018-2019 (will take the higher year), and 123232019 (will ignore).
company_paths = preparation.get_company_paths(reports_input_dir)
for company_dir in company_paths:
company_files = preparation.get_reports_paths(company_dir.path)
for p in company_files:
filename = str(p.stem)
if purge_existing_folder:
try:
shutil.rmtree(os.path.join(company_dir.path, filename))
except:
print("No file found for: ", filename)
potential_years = re.findall(r"(?<!\d)((?:199|200|201|202)[0-9]{1})(?!\d)", filename)
if len(potential_years) != 1:
if len(potential_years) and abs(int(potential_years[0]) - int(potential_years[1])) == 1:
year = max(potential_years)
print(filename, ": found a range of years, taking the higher one...")
else:
print("Ambigious years found for ", filename)
print(potential_years)
print("=========")
break
else:
year = potential_years[0]
if year:
os.rename(p, Path(p.parent, f"AR_{year}.pdf"))
cleaned_company_name = re.sub(r'\W+', '', str(company_dir.name))
if cleaned_company_name != company_dir.name:
new_folder = os.path.join(os.path.dirname(company_dir), cleaned_company_name)
os.rename(company_dir, new_folder)
# ### Extract text from PDF
if extract_text:
# ! python ../../data/pipeline.py "$reports_input_dir"
master_df = preparation.get_df(input_path=reports_input_dir, include_text=False, include_page_no=False, include_toc=False)
master_df['should_label'] = True
master_df['is_labelled'] = False
master_df = master_df.sample(frac=1, random_state=1)
# +
master_df.to_csv(master_output_file)
# -
| v2/notebooks/PDF Extraction and Additional Labelling Preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="-gx4xAqY2ZK9" colab_type="code" colab={}
import pandas as pd
from sklearn.cluster import KMeans
# + id="2DzuW-5N2ey6" colab_type="code" colab={}
file_url = 'https://raw.githubusercontent.com/TrainingByPackt/The-Data-Science-Workshop/master/Chapter05/DataSet/taxstats2015.csv'
# + id="PWCdoz-p2m__" colab_type="code" colab={}
df = pd.read_csv(file_url, usecols=['Postcode', 'Average net tax', 'Average total deductions'])
# + id="vbPPpVxX2qcE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="d3c857f7-4a0e-4548-eb50-647af010e146" executionInfo={"status": "ok", "timestamp": 1568616929586, "user_tz": -330, "elapsed": 745, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13925024237489964117"}}
df.head()
# + id="0C4qfNoI2tWT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="9224b564-fde2-4375-df5f-b0f2107bb5b9" executionInfo={"status": "ok", "timestamp": 1568616954508, "user_tz": -330, "elapsed": 738, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13925024237489964117"}}
df.tail()
# + id="yj2pf5ah2zbn" colab_type="code" colab={}
kmeans = KMeans(random_state=42)
# + id="rgg3mGRP249m" colab_type="code" colab={}
X = df[['Average net tax', 'Average total deductions']]
# + id="TAMAoRem26KF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 73} outputId="b4585f92-866f-4e8a-c173-b0886be20859" executionInfo={"status": "ok", "timestamp": 1568616987998, "user_tz": -330, "elapsed": 957, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13925024237489964117"}}
kmeans.fit(X)
# + id="r_okd9dU27jv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="dba9a263-ca31-491e-9aaa-f266a07c5301" executionInfo={"status": "ok", "timestamp": 1568617009513, "user_tz": -330, "elapsed": 762, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13925024237489964117"}}
y_preds = kmeans.predict(X)
y_preds
# + id="4ZAl3a4Z3A3L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="f06bdafb-8c38-4eba-8d1c-08960dc37d84" executionInfo={"status": "ok", "timestamp": 1568617030824, "user_tz": -330, "elapsed": 750, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13925024237489964117"}}
df['cluster'] = y_preds
df.head()
# + id="FR-Bqa3x3GEM" colab_type="code" colab={}
| Chapter05/Exercise5.1/Exercise5.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import Files
import pandas as pd
vanorder = pd.read_csv('vanorder.csv')
vaninterest = pd.read_csv('vaninterest.csv')
# Exploratory Data Analysis
vanorder.head()
vaninterest.head()
vanorder.describe()
# Convert date into Pandas datetime object
vanorder.txCreate = pd.to_datetime(vanorder.txCreate)
vaninterest.txCreate = pd.to_datetime(vaninterest.txCreate)
vanorder.head()
# Q) 5 : What is the order fulfillment rate, i.e. percentage of orders that was completed ?
len(vanorder[vanorder.order_status == 2])/len(vanorder)
# Order Fulfillment rate = 94%
# Subset Order type- A
vanorderA = vanorder[vanorder.order_subset == 'A']
vaninterestA = vaninterest[vaninterest.order_subset_assigned == 'A']
# Create a new column matchtime i.e difference of time accepted and time created
vanorderA['txAccept'] = vaninterestA.txCreate
vanorderA['matchtime'] = vanorderA.txAccept - vanorderA.txCreate
vanorderA.head()
# Subset Advanced/Immediate orders
van_advanced_orders = vanorderA[(vanorderA.matchtime.between('00:00:00', '01:00:00') )]
van_advanced_orders = van_advanced_orders[van_advanced_orders.matchtime < '01:00:00']
van_advanced_orders.head()
# Q)6 (a) What is the average match time, by immediate/advanced orders?
van_advanced_orders.matchtime.mean()
# Average matchtime is 8 Minutes 59 sec
# Q)6 (b) What is the median match time, by immediate/advanced orders?
van_advanced_orders.matchtime.median()
# Median matchtime is 5 Minutes 06 sec
# (c) Which of the above one do you think provides a better representation the data, i.e. a better metric for tracking our performance in matching?
# Median gives a better metric because it doesn't get affected by outliers.(In this case midnight orders).
#
# However mean of binned hours would provide better insights.(Avg matchtime of morning hours,afternnon,evening and night)
# Export the file as a csv to prepeare dashnoard(Tableau)
van_advanced_orders.to_csv('1.csv')
# Dashboard for Management
# <img src="dashboard_for_management.jpg">
# Dashboard for Operations :
# <img src="dashboard_for_operations.jpg">
| Data Analysis Using Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualização de dados com Python
# ## 2 - Visualizações com mais de 2 dimensões
# *<NAME>*, [**DataLearningHub**](http://datalearninghub.com)
# Nesta lição veremos como fornecer visualizações com mais de duas dimensões de dados.
# [](https://www.lcm.com.br/site/#livros/busca?term=cleuton)
# ## Dispersão tridimensional
# Em casos que temos três características mensuráveis e, principalmente, plotáveis (dentro da mesma escala - ou podemos ajustar a escala), é interessante ver um gráfico de dispersão para podermos avaliar visualmente a distribuição das amostras. É o que veremos com a bilbioteca Matplotlib Toolkits, em especial a MPlot3D, que tem o objeto Axes3D para geração de gráficos tridimensionais.
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d, Axes3D # Objetos que usaremos em nosso gráfico
# %matplotlib inline
df = pd.read_csv('../datasets/evasao.csv') # Dados de evasão escolar que coletei
df.head()
# Algumas explicações. Para começar, vejamos as colunas deste dataset:
# - "periodo": Período em que o aluno está;
# - "bolsa": Percentual de bolsa que o aluno recebe;
# - "repetiu": Quantidade de disciplinas nas quais o aluno foi reprovado;
# - "ematraso": Se o aluno está com mensalidades em atraso;
# - "disciplinas": Disciplinas que o aluno está cursando atualmente;
# - "desempenho": Média acadêmica até agora;
# - "abandonou": Se o aluno abandonou o curso depois da medição ou não.
#
# Para podermos plotar um gráfico, precisamos reduzir a quantidade de dimensões, ou seja, as características. Farei isso da maneira mais "naive" possível, selecionando três características que mais influenciaram no resultado final, ou seja o abandono do aluno (Churn).
df2 = df[['periodo','repetiu','desempenho']][df.abandonou == 1]
df2.head()
fig = plt.figure()
#ax = fig.add_subplot(111, projection='3d')
ax = Axes3D(fig) # Para Matplotlib 0.99
ax.scatter(xs=df2['periodo'],ys=df2['repetiu'],zs=df2['desempenho'], c='r',s=8)
ax.set_xlabel('periodo')
ax.set_ylabel('repetiu')
ax.set_zlabel('desempenho')
plt.show()
# Simplesmente usei o Axes3D para obter um objeto gráfico tridimensional. O método "scatter" recebe três dimensões (xs, ys e zs), cada uma atribuída a uma das colunas do novo dataframe. O parâmetro "c" é a cor e o "s" é o tamanho de cada ponto. Informei os rótulos de cada eixo e pronto! Temos um gráfico 3D mostrando a distribuição espacial dos abandonos de curso, com relação às três variáveis.
#
# Podemos avaliar muito melhor a tendência de dados, se olharmos em visualizações 3D. Vejamos um exemplo sintético. Vamos gerar alguns valores 3D:
import numpy as np
np.random.seed(42)
X = np.linspace(1.5,3.0,num=100)
Y = np.array([x**4 + (np.random.rand()*6.5) for x in X])
Z = np.array([(X[i]*Y[i]) + (np.random.rand()*3.2) for i in range(0,100)])
# Primeiramente veremos como ficaria isso em visualização 2D:
fig = plt.figure()
ax = fig.add_subplot(111)
ax.scatter(X, Y, c='b', s=20)
ax.set_xlabel('X')
ax.set_ylabel('Y')
plt.show()
# Ok... Nada demais... Uma correlação não linear positiva, certo? Mas agora, vejamos isso com a matriz Z incluída:
fig = plt.figure()
ax = Axes3D(fig)
ax.scatter(X, Y, Z, c='r',s=8)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
plt.show()
# E isso fica mais interessante quando sobrepomos uma predição sobre os dados reais. Vamos usar um Decision Tree Regressor para criar um modelo preditivo para estes dados:
# +
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import train_test_split
features = pd.DataFrame({'X':X, 'Z':Z})
labels = pd.DataFrame({'Y':Y})
X_train, X_test, y_train, y_test = train_test_split(features, labels, test_size=0.33, random_state=42)
dtr3d = DecisionTreeRegressor(max_depth=4, random_state=42)
dtr3d.fit(X_train,y_train)
print('R2',dtr3d.score(X_train,y_train))
# +
yhat3d = dtr3d.predict(X_test)
fig = plt.figure()
ax = ax = fig.add_subplot(111, projection='3d')
ax.scatter(X, Y, Z, c='r',s=8)
ax.scatter(X_test['X'], yhat3d, X_test['Z'], c='k', marker='*',s=100)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
plt.show()
# -
# Plotamos as predições usando marker do tipo estrela. Ficou bem interessante, não?
# ## Mais de 3 dimensões
#
# As vezes queremos demonstrar informações com mais de 3 dimensões, mas como fazer isso? Vamos supor que queiramos também incluir o percentual de bolsa como uma variável em nosso exemplo de evasão escolar. Como faríamos?
# Uma abordagem possível seria manipular os markers para que representem a bolsa. Podemos usar cores, por exemplo. Vejamos, primeiramente, precisamos saber quais faixas de bolsa existem no dataset:
#
print(df.groupby("bolsa").count())
# Podemos criar uma tabela de cores, indexada pelo percentual de bolsa:
from decimal import Decimal
bolsas = {0.00: 'b',0.05: 'r', 0.10: 'g', 0.15: 'm', 0.20: 'y', 0.25: 'k'}
df['cor'] = [bolsas[float(round(Decimal(codigo),2))] for codigo in df['bolsa']]
df.head()
# Essa "maracutaia" merece uma explicação. Criei um dicionário indexado pelo valor da bolsa. Assim, pegamos o código da cor correspondente. Só que preciso incluir uma coluna no dataframe com esse valor, de modo a usar no gráfico. Só tem um problema: O dataset original está "sujo" (algo que acontece frequentemente) e o percentual 0.15 está como 0.1500000002. Posso retirar isso convertendo o falor de "float" para "Decimal", arredondanto e convertendo novamente em float.
#
# Quando plotarmos, vamos procurar a cor no dicionário:
fig = plt.figure()
#ax = fig.add_subplot(111, projection='3d')
ax = Axes3D(fig) # Para Matplotlib 0.99
ax.scatter(xs=df['periodo'],ys=df['repetiu'],zs=df['desempenho'], c=df['cor'],s=50)
ax.set_xlabel('periodo')
ax.set_ylabel('repetiu')
ax.set_zlabel('desempenho')
plt.show()
# Pronto! Temos ai a cor da bola dando a quarta dimensão: O percentual de bolsa
# Vemos que já uma concentração de alunos com bolsa de 25% (cor preta) com poucas repetições, mas baixo desempenho, em todos os períodos.
# Assim como mexemos com a cor, podemos mexer com o tamanho, criando algo como um "mapa de calor". Vamos transformar essa visão em 2D, colocando o "desempenho" com tamanho diferenciado.
fig, ax = plt.subplots()
ax.scatter(df['periodo'],df['repetiu'], c='r',s=df['desempenho']*30)
ax.set_xlabel('periodo')
ax.set_ylabel('repetiu')
plt.show()
# Isso nos mostra um fato curioso. Temos alunos com bom desempenho (bolas grandes) em todos os períodos, sem repetir nenhuma disciplina, que abandonaram. O que os teria feito fazer isto? Talvez sejam condições financeiras, ou insatisfação com o curso. Um fato a ser investigado, que só foi revelado graças a esta visualização.
# ## Georreferenciamento
# Muitas vezes temos datasets com informações geográficas e precisamos plotar os dados sobre um mapa. Vou mostrar aqui como fazer isso com um exemplo do dataset dos casos de Dengue de 2018 no Rio de Janeiro. Fonte: Data Rio: http://www.data.rio/datasets/fb9ede8d588f45b48b985e62c817f062_0
#
# Eu criei um dataset georreferenciado, que está na pasta desta demonstração. Ele está em formato CSV, separado por ponto e vírgula, com separador decimal em português (vírgula):
df_dengue = pd.read_csv('./dengue2018.csv',decimal=',', sep=';')
df_dengue.head()
# Um simples gráfico de dispersão já dá uma boa noção do problema:
fig, ax = plt.subplots()
ax.scatter(df_dengue['longitude'],df_dengue['latitude'], c='r',s=15)
plt.show()
# Podemos colocar o tamanho do ponto proporcional à quantidade de casos, aumentando a dimensão das informações:
fig, ax = plt.subplots()
ax.scatter(df_dengue['longitude'],df_dengue['latitude'], c='r',s=5+df_dengue['quantidade'])
plt.show()
# Podemos manipular a cor e intensidade para criar um "mapa de calor" da Dengue:
# +
def calcular_cor(valor):
cor = 'r'
if valor <= 10:
cor = '#ffff00'
elif valor <= 30:
cor = '#ffbf00'
elif valor <= 50:
cor = '#ff8000'
return cor
df_dengue['cor'] = [calcular_cor(codigo) for codigo in df_dengue['quantidade']]
# -
df_dengue.head()
# E vamos ordenar para que as maiores quantidades fiquem por último:
dfs = df_dengue.sort_values(['quantidade'])
dfs.head()
fig, ax = plt.subplots()
ax.scatter(dfs['longitude'],dfs['latitude'], c=dfs['cor'],s=10+dfs['quantidade'])
plt.show()
# Pronto! Um mapa de calor da Dengue em 2018. Mas está faltando algo certo? Cadê o mapa do Rio de Janeiro?
# Muita gente usa o **geopandas** e baixa arquivos de mapas. Eu prefiro usar o Google Maps. Ele tem uma API chamada Static Maps que permite baixar mapas. Primeiramente, vou instalar o **requests**:
# !pip install requests
# Agora, vem uma parte um pouco mais "esperta". Eu tenho as coordenadas do centro do Rio de Janeiro (centro geográfico, não o centro da cidade). Vou montar um request à API Static Map para baixar um mapa. Veja bem, você tem que cadastrar uma API Key para usar esta API. Eu omiti a minha propositalmente. Aqui você tem as instruções para isto: https://developers.google.com/maps/documentation/maps-static/get-api-key
import requests
latitude = -22.9137528
longitude = -43.526409
zoom = 10
size = 800
scale = 1
apikey = "**INFORME SUA API KEY**"
gmapas = "https://maps.googleapis.com/maps/api/staticmap?center=" + str(latitude) + "," + str(longitude) + \
"&zoom=" + str(zoom) + \
"&scale=" + str(scale) + \
"&size=" + str(size) + "x" + str(size) + "&key=" + apikey
with open('mapa.jpg', 'wb') as handle:
response = requests.get(gmapas, stream=True)
if not response.ok:
print(response)
for block in response.iter_content(1024):
if not block:
break
handle.write(block)
# 
# Bom, o mapa foi salvo, agora eu preciso saber as coordenadas dos limites. A API do Google só permite que você informe o centro (latitude e longitude) e as dimensões da imagem em pixels. Mas, para ajustar o mapa às coordenadas em latitudes e longitudes, é preciso saber as coordenadas do retângulo da imagem.
# Há vários exemplos de como calcular isso e eu uso um exemplo Javascript que converti para Python há algum tempo.
# Este cálculo é baseado no script de: https://jsfiddle.net/1wy1mm7L/6/
# +
import math
_C = { 'x': 128, 'y': 128 };
_J = 256 / 360;
_L = 256 / (2 * math.pi);
def tb(a):
return 180 * a / math.pi
def sb(a):
return a * math.pi / 180
def bounds(a, b, c):
if b != None:
a = max(a,b)
if c != None:
a = min(a,c)
return a
def latlonToPt(ll):
a = bounds(math.sin(sb(ll[0])), -(1 - 1E-15), 1 - 1E-15);
return {'x': _C['x'] + ll[1] * _J,'y': _C['y'] + 0.5 * math.log((1 + a) / (1 - a)) * - _L}
def ptToLatlon(pt):
return [tb(2 * math.atan(math.exp((pt['y'] - _C['y']) / -_L)) - math.pi / 2),(pt['x'] - _C['x']) / _J]
def calculateBbox(ll, zoom, sizeX, sizeY, scale):
cp = latlonToPt(ll)
pixelSize = math.pow(2, -(zoom + 1));
pwX = sizeX*pixelSize;
pwY = sizeY*pixelSize;
return {'ne': ptToLatlon({'x': cp['x'] + pwX, 'y': cp['y'] - pwY}),'sw': ptToLatlon({'x': cp['x'] - pwX, 'y': cp['y'] + pwY})}
limites = calculateBbox([latitude,longitude],zoom, size, size, scale)
print(limites)
# -
# A função "calculateBbox" retorna um dicionário contendo os pontos Nordeste e Sudoeste, com a latitude e longitude de cada um.
# Para usar isso no matplotlib, eu preciso usar o método **imshow**, só que eu preciso informar a escala, ou seja, qual é o intervalo de latitudes (vertical) e longitudes (horizontal) que o mapa representa. Assim, a plotagem de pontos ficará correta.
# Eu vou usar a biblioteca **mpimg** para ler o arquivo de imagem que acabei de baixar.
# Só que a função **imshow** usa as coordenadas no atributo **extent** na ordem: ESQUERDA, DIREITA, BAIXO, TOPO. Temos que organizar a passagem dos parâmetros para ela.
import matplotlib.image as mpimg
fig, ax = plt.subplots(figsize=(10, 10))
rio_mapa=mpimg.imread('./mapa.jpg')
plt.imshow(rio_mapa, extent=[limites['sw'][1],limites['ne'][1],limites['sw'][0],limites['ne'][0]], alpha=1.0)
ax.scatter(dfs['longitude'],dfs['latitude'], c=dfs['cor'],s=10+dfs['quantidade'])
plt.ylabel("Latitude", fontsize=14)
plt.xlabel("Longitude", fontsize=14)
plt.show()
# Pronto! Ai está! Um mapa de calor georreferenciado da Dengue em 2018 no Rio de Janeiro
| datavisualization/data_visualization_python_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Y2fkP_np3Zpl"
# <!--BOOK_INFORMATION-->
# <img align="left" style="width:80px;height:98px;padding-right:20px;" src="https://raw.githubusercontent.com/joe-papa/pytorch-book/main/files/pytorch-book-cover.jpg">
#
# This notebook contains an excerpt from the [PyTorch Pocket Reference](http://pytorchbook.com) book by [<NAME>](http://joepapa.ai); content is available [on GitHub](https://github.com/joe-papa/pytorch-book).
# + [markdown] id="CPNoDUafIa9c"
# [](https://colab.research.google.com/github/joe-papa/pytorch-book/blob/main/08_02_PyTorch_Ecosystem_TorchText.ipynb)
# + [markdown] id="yw9xjxnv7KUZ"
# # Chapter 8 - PyTorch Ecosystem
# + [markdown] id="ENv8CU1R5ya5"
# ## TorchText for NLP
# + colab={"base_uri": "https://localhost:8080/"} id="ngd00fwqH4PB" executionInfo={"status": "ok", "timestamp": 1615499990985, "user_tz": 300, "elapsed": 14636, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00487850786587503652"}} outputId="8d1526fc-198e-4e27-85f0-1f44992e14d9"
from torchtext.datasets import IMDB
train_iter, test_iter = \
IMDB(split=('train', 'test'))
next(train_iter)
# out:
# ('neg',
# 'I rented I AM CURIOUS-YELLOW ...)
# + id="cz1y5XkCIEb6"
from torchtext.data.utils \
import get_tokenizer
tokenizer = get_tokenizer('basic_english')
# + id="I-wx1JvOIcWT"
from collections import Counter
from torchtext.vocab import Vocab
train_iter = IMDB(split='train')
counter = Counter()
for (label, line) in train_iter:
counter.update(tokenizer(line))
vocab = Vocab(counter,
min_freq=10,
specials=('<unk>',
'<BOS>',
'<EOS>',
'<PAD>'))
# + colab={"base_uri": "https://localhost:8080/"} id="ubBrI5CoIpN9" executionInfo={"status": "ok", "timestamp": 1615500002137, "user_tz": 300, "elapsed": 25770, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00487850786587503652"}} outputId="053d578c-c0ef-4974-81b3-58f518da1c20"
text_transform = lambda x: [vocab['<BOS>']] \
+ [vocab[token] \
for token in tokenizer(x)] + [vocab['<EOS>']]
label_transform = lambda x: 1 \
if x == 'pos' else 0
print(text_transform("programming is awesome"))
# out: [1, 8320, 12, 1156, 2]
# + id="2D3NNfAKIzI0"
from torch.utils.data import DataLoader
train_iter = IMDB(split='train')
train_dataloader = DataLoader(
list(train_iter),
batch_size=8,
shuffle=True)
# for (text, label) in train_dataloader:
# 'Train your NN'
| 08_02_PyTorch_Ecosystem_TorchText.ipynb |